code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 32-bit
# name: python3
# ---
# + [markdown] id="ohoT6RjYmeHM"
# #**Visualizations**
# + id="fCVt5xlQmVF_"
import matplotlib.pyplot as plt
import numpy as np
# + id="HMT3CRtEmVGA" outputId="9349699a-79f5-4ff4-febb-556bc5649a37" colab={"base_uri": "https://localhost:8080/", "height": 297}
labels = ['G1', 'G2', 'G3', 'G4', 'G5']
men_means = [20, 34, 30, 35, 27]
women_means = [25, 32, 34, 20, 25]
x = np.arange(len(labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(x - width/2, men_means, width, label='Men')
rects2 = ax.bar(x + width/2, women_means, width, label='Women')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Scores')
ax.set_title('Scores by group and gender')
ax.set_xticks(x)
ax.set_xticklabels(labels)
ax.legend()
# ax.bar_label(rects1, padding=3)
# ax.bar_label(rects2, padding=3)
fig.tight_layout()
plt.show()
# + id="g8zvc2eomnTw"
|
Visualizations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DU-ds/pyspark_udemy/blob/main/Section4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Jply4-bFg8Hb" outputId="b85f47b1-6896-4ba9-d06e-e851fe6a52fb"
# ! apt update
# ! apt install openjdk-8-jdk-headless -qq > /dev/null
# ! wget -q http://archive.apache.org/dist/spark/spark-2.3.1/spark-2.3.1-bin-hadoop2.7.tgz
# ! tar xf spark-2.3.1-bin-hadoop2.7.tgz
# ! pip install -q findspark
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-2.3.1-bin-hadoop2.7"
# ! ls
import findspark
findspark.init()
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.getOrCreate()
spark
from pyspark.sql import types
# + colab={"base_uri": "https://localhost:8080/"} id="XJ5WFq5ghBoR" outputId="acf899f1-e0c3-4eb0-a052-f6aa0e1e6ee9"
df = spark.read.csv("challenge.csv", header=True)
df.show()
# + colab={"base_uri": "https://localhost:8080/"} id="EpMam8Eph55g" outputId="3b27acff-c0da-4f66-fe75-6437a7fb1c33"
from pyspark.sql import functions as sqlf
df_mex = df.withColumn("mexico", sqlf.when(df.Country == "Mexico", "YES").otherwise("NO"))
df_mex.show()
# + colab={"base_uri": "https://localhost:8080/"} id="8aZl-U_Fh5yU" outputId="2b8ef131-3dc9-4f8d-8586-5f21743b7fb6"
df_mex.groupBy("mexico").agg(sqlf.sum(df.Bytes_used)).alias("Bytes_Mexico").show()
# + colab={"base_uri": "https://localhost:8080/"} id="PLn2I6YAh5vR" outputId="fefcac29-fe93-4dfd-d895-fd427d9cd3bc"
df_ip_country = df.groupBy("Country").agg(sqlf.countDistinct("ip_address").alias("ips_per_country"))
df_ip_country.sort(col("ips_per_country").desc()).show()
# + id="QezAqCK8h5gn"
|
Section4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# #!/usr/bin/env python2
#Inspire du fichier train_fcn8.py
import os
import argparse
import time
from getpass import getuser
from distutils.dir_util import copy_tree
import pickle
import numpy as np
import random
import theano
import theano.tensor as T
from theano import config
import lasagne
from lasagne.regularization import regularize_network_params
from lasagne.objectives import categorical_crossentropy
import PIL.Image as Image
from matplotlib import pyplot as plt
from matplotlib import colors
from matplotlib import gridspec
from fcn_1D_general import buildFCN_1D
from metrics import jaccard, accuracy, crossentropy
from data_loader.cortical_layers import CorticalLayersDataset
from simple_model import build_simple_model
from profile_functions import profile2indices
#Stuff for profiling functions
#os.environ['CUDA_LAUNCH_BLOCKING']='1'
#theano.config.profile= True
##theano.config.profile_memory=True
#theano.config.profile_optimizer = True
# %matplotlib inline
# + deletable=true editable=true
_FLOATX = config.floatX
SAVEPATH = '/Tmp/larocste/cortical_layers'
LOADPATH = '/data/lisatmp4/larocste/cortical_layers'
WEIGHTS_PATH = LOADPATH
# + [markdown] deletable=true editable=true
# ## Hyperparameters
#
# ### Model hyperparameters
# - n_filters : int, nb of filters for each convLayer
# - filter_size : list of odd int (to fit with the pad='same'), len(filter_size) = nb of convLayer in simple_model, each of these layer with the corresponding filter_size
# - depth : int, depth of the network (how many stacked convolution)
#
# ### Training loop parameters
# - weight_decay : not implemented yet
# - num_epochs : int, max number of epochs
# - max_patience : int, max nb of epochs without improvement in the jaccard accuracy (jacc_valid) on the validation set
# - learning rate : defined later as a theano shared variable
#
# ### Hyperparameters for the dataset loader
# - batch_size=[training_batch_size, valid_batch_size, test_batch_size]
# - smooth_or_raw : 'smooth' or 'raw', whether to use smooth OR raw data
# - shuffle_at_each_epoch : boolean (keep it to True)
# - minibatches_subset : int, if>0 : get only that number of minibatch instead of all training dataset.
#
# + deletable=true editable=true
#Model hyperparameters
n_filters = 64
filter_size = [7,15,25,49]
depth = 4
data_augmentation={} #{'horizontal_flip': True, 'fill_mode':'constant'}
#Training loop hyperparameters
weight_decay=0.001
num_epochs=500
max_patience=25
resume=False
learning_rate_value = 0.0005 #learning rate is defined below as a theano variable.
#Hyperparameters for the dataset loader
batch_size=[500,500,1]
smooth_or_raw = 'both'
shuffle_at_each_epoch = True
minibatches_subset = 0
# + deletable=true editable=true
#
# Prepare load/save directories
#
savepath=SAVEPATH
loadpath=LOADPATH
exp_name = 'simple_model'
exp_name += '_lrate=' + str(learning_rate_value)
exp_name += '_fil=' + str(n_filters)
exp_name += '_fsizes=' + str(filter_size)
exp_name += '_depth=' + str(depth)
exp_name += '_data=' + smooth_or_raw
exp_name += '_decay=' + str(weight_decay)
exp_name += '_pat=' + str(max_patience)
exp_name += ('_noshuffle'+str(minibatches_subset)+'batch') if not shuffle_at_each_epoch else ''
#exp_name += 'test'
dataset = 'cortical_layers'
savepath = os.path.join(savepath, dataset, exp_name)
loadpath = os.path.join(loadpath, dataset, exp_name)
print 'Savepath : '
print savepath
print 'Loadpath : '
print loadpath
if not os.path.exists(savepath):
os.makedirs(savepath)
else:
print('\033[93m The following folder already exists {}. '
'It will be overwritten in a few seconds...\033[0m'.format(
savepath))
print('Saving directory : ' + savepath)
with open(os.path.join(savepath, "config.txt"), "w") as f:
for key, value in locals().items():
f.write('{} = {}\n'.format(key, value))
# + deletable=true editable=true
# + deletable=true editable=true
#
# Define symbolic variables
#
input_var = T.tensor3('input_var') #n_example*nb_in_channels*ray_size
target_var = T.ivector('target_var') #n_example*ray_size
learn_step= theano.shared(np.array(learning_rate_value, dtype=theano.config.floatX))
# + deletable=true editable=true
#
# Build dataset iterator
#
if smooth_or_raw =='both':
nb_in_channels = 2
use_threads = False
else:
nb_in_channels = 1
use_threads = True
train_iter = CorticalLayersDataset(
which_set='train',
smooth_or_raw = smooth_or_raw,
batch_size=batch_size[0],
data_augm_kwargs=data_augmentation,
shuffle_at_each_epoch = shuffle_at_each_epoch,
return_one_hot=False,
return_01c=False,
return_list=False,
use_threads=use_threads)
val_iter = CorticalLayersDataset(
which_set='valid',
smooth_or_raw = smooth_or_raw,
batch_size=batch_size[1],
shuffle_at_each_epoch = shuffle_at_each_epoch,
return_one_hot=False,
return_01c=False,
return_list=False,
use_threads=use_threads)
test_iter = None
n_batches_train = train_iter.nbatches
n_batches_val = val_iter.nbatches
n_batches_test = test_iter.nbatches if test_iter is not None else 0
n_classes = train_iter.non_void_nclasses
void_labels = train_iter.void_labels
#nb_in_channels = train_iter.data_shape[0]
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
#
# Build network
#
simple_net_output, net = build_simple_model(input_var,
filter_size = filter_size,
n_filters = n_filters,
depth = depth,
nb_in_channels = nb_in_channels,
n_classes = n_classes)
#simple_net_output = last layer of the simple_model net
#net = dictionary containing the names of every layer (used to visualize data)
# To print each layer, uncomment this:
# lays = lasagne.layers.get_all_layers(simple_net_output)
# for l in lays:
# print l, l.output_shape
# #print simple_net_output[l], simple_net_output[l].output_shape, l
# print '---------------------------'
# print 'simple_net_output :', simple_net_output
# print '---------------------------'
# #print 'net :', net
# + deletable=true editable=true
# + deletable=true editable=true
#
# Define and compile theano functions
#
print "Defining and compiling training functions"
prediction = lasagne.layers.get_output(simple_net_output[0])
loss = categorical_crossentropy(prediction, target_var)
loss = loss.mean()
if weight_decay > 0:
weightsl2 = regularize_network_params(
simple_net_output, lasagne.regularization.l2)
loss += weight_decay * weightsl2
train_acc, train_sample_acc = accuracy(prediction, target_var, void_labels)
params = lasagne.layers.get_all_params(simple_net_output, trainable=True)
updates = lasagne.updates.adam(loss, params, learning_rate=learn_step)
train_fn = theano.function([input_var, target_var], [loss, train_acc, train_sample_acc],
updates=updates)#, profile=True)
print "Done"
# + deletable=true editable=true
print "Defining and compiling valid functions"
valid_prediction = lasagne.layers.get_output(simple_net_output[0],
deterministic=True)
valid_loss = categorical_crossentropy(valid_prediction, target_var)
valid_loss = valid_loss.mean()
#valid_loss = crossentropy(valid_prediction, target_var, void_labels)
valid_acc, valid_sample_acc = accuracy(valid_prediction, target_var, void_labels)
valid_jacc = jaccard(valid_prediction, target_var, n_classes)
valid_fn = theano.function([input_var, target_var],
[valid_loss, valid_acc, valid_sample_acc, valid_jacc])#,profile=True)
print "Done"
# + deletable=true editable=true
#Function computing the prediction with current parameters (for visualization)
pred = theano.function([input_var], lasagne.layers.get_output(net['probs_reshape'],
deterministic=True))
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
#To visualize the ray
def make_2Darray(arr, height = 25):
arr = np.reshape(arr, (1,arr.shape[0]))
x = np.repeat(arr, height, 0)
return x
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
def plot_true_predicted_labels(X_batch, L_batch, idx_batch, smooth_or_raw='raw', index = -1, which_set=''):
#Get random sample
if index != -1:
idx = index
else:
idx = random.randint(0,X_batch.shape[0]-1)
real_idx = idx_batch[idx]
ray = X_batch[idx][0]
true_labels = L_batch[idx*200:(idx+1)*200]
predicted_labels = np.argmax(pred(X_batch), axis=2)[idx]
true_vs_predicted = np.concatenate((make_2Darray(true_labels),
make_2Darray(predicted_labels)), axis = 0)
if smooth_or_raw=='both':
plt.title('Smooth vs raw data ' + which_set + ' idx = '+ str(real_idx))
data = np.concatenate((make_2Darray(X_batch[idx][0]),
make_2Darray(X_batch[idx][1])), axis=0)
plt.imshow(data, cmap='gray', interpolation='none')
plt.figure()
else :
plt.title(smooth_or_raw + ' data ' + which_set + ' idx = '+ str(real_idx))
plt.imshow(make_2Darray(X_batch[idx][0]), cmap='gray', interpolation='none')
plt.figure()
plt.title('Ground truth vs predicted labels '+ which_set + ' idx = '+ str(real_idx))
plt.imshow(true_vs_predicted, interpolation='none', vmin=0, vmax=n_classes)
plt.figure()
plt.show()
# + deletable=true editable=true
def index_worse_than(sample_acc_batch, idx_batch, treshold=0.7):
worse_batch_idx = np.array([i for i in range(len(sample_acc_batch)) if sample_acc_batch[i]<treshold], dtype=int)
if worse_batch_idx==[]:
return []
else :
return idx_batch[worse_batch_idx]
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# Uncomment this if only 1 minibatch (smaller dataset)
# and comment 2 lines in training loop to avoid getting new minibatches
# X_train_batch, L_train_batch = train_iter.next()
# X_val_batch, L_val_batch = val_iter.next()
# minibatches_subset = 1
# print X_train_batch.shape
# print L_train_batch.shapedata
# + deletable=true editable=true
# + deletable=true editable=true
#whether to plot labels prediction or not during training
#(1 random example of the last minibatch for each epoch)
plot_results_train = True #from the training set
plot_results_valid = True #from the validation set
treshold = 0.7 # for extracting the very incorrect labelled samples
ratios=[0.80,0.85, 0.90] #ratios for the per sample accuracy
# + deletable=true editable=true
#
# Train loop
#
err_train = []
acc_train = []
sample_acc_train_tot = []
worse_indices_train = []
already_seen_idx = []
err_valid = []
acc_valid = []
jacc_valid = []
sample_acc_valid_tot = []
patience = 0
worse_indices_valid =[]
# Training main loop
print "Start training"
for epoch in range(num_epochs):
#learn_step.set_value((learn_step.get_value()*0.99).astype(theano.config.floatX))
# Single epoch training and validation
start_time = time.time()
#Cost train and acc train for this epoch
cost_train_epoch = 0
acc_train_epoch = 0
sample_acc_train_epoch = np.array([0.0 for i in range(len(ratios))])
worse_indices_train_epoch = []
# Train
if minibatches_subset > 0:
n_batches_val = minibatches_subset
n_batches_train = minibatches_subset
for i in range(n_batches_train):
# Get minibatch (comment the next line if only 1 minibatch in training)
train_batch = train_iter.next()
X_train_batch, L_train_batch, idx_train_batch = train_batch['data'], train_batch['labels'], train_batch['indices'][0]
L_train_batch = np.reshape(L_train_batch, np.prod(L_train_batch.shape))
# Training step
cost_train_batch, acc_train_batch, sample_acc_train_batch = train_fn(
X_train_batch, L_train_batch)
sample_acc_train_batch_mean = [np.mean([(i>=ratio)
for i in sample_acc_train_batch]) for ratio in ratios]
worse_indices_train_batch = index_worse_than(sample_acc_train_batch,
idx_train_batch, treshold=treshold)
#print i, 'training batch cost : ', cost_train_batch, ' batch accuracy : ', acc_train_batch
#Update epoch results
cost_train_epoch += cost_train_batch
acc_train_epoch += acc_train_batch
sample_acc_train_epoch += sample_acc_train_batch_mean
worse_indices_train_epoch = np.hstack((worse_indices_train_epoch,worse_indices_train_batch))
#Add epoch results
err_train += [cost_train_epoch/n_batches_train]
acc_train += [acc_train_epoch/n_batches_train]
sample_acc_train_tot += [sample_acc_train_epoch/n_batches_train]
worse_indices_train += [worse_indices_train_epoch]
if plot_results_train: #select random example from the last minibatch and plot it
plot_true_predicted_labels(X_train_batch, L_train_batch,idx_train_batch, smooth_or_raw,which_set='TRAINING')
# Validation
cost_val_epoch = 0
acc_val_epoch = 0
sample_acc_valid_epoch = np.array([0.0 for i in range(len(ratios))])
jacc_val_epoch = np.zeros((2, n_classes))
worse_indices_val_epoch = []
for i in range(n_batches_val):
# Get minibatch (comment the next line if only 1 minibatch in training)
val_batch = val_iter.next()
X_val_batch, L_val_batch, idx_val_batch = val_batch['data'], val_batch['labels'], val_batch['indices'][0]
L_val_batch = np.reshape(L_val_batch, np.prod(L_val_batch.shape))
# Validation step
cost_val_batch, acc_val_batch, sample_acc_valid_batch, jacc_val_batch = valid_fn(X_val_batch, L_val_batch)
#print i, 'validation batch cost : ', cost_val_batch, ' batch accuracy : ', acc_val_batch
worst_index = np.argmin(sample_acc_valid_batch)
sample_acc_valid_batch_mean = [np.mean([(i>=ratio)
for i in sample_acc_valid_batch]) for ratio in ratios]
worse_indices_val_batch = index_worse_than(sample_acc_valid_batch,
idx_val_batch, treshold=treshold)
#Update epoch results
cost_val_epoch += cost_val_batch
acc_val_epoch += acc_val_batch
sample_acc_valid_epoch += sample_acc_valid_batch_mean
jacc_val_epoch += jacc_val_batch
worse_indices_val_epoch = np.hstack((worse_indices_val_epoch, worse_indices_val_batch))
if plot_results_valid: #select random example from the last minibatch and plot it
plot_true_predicted_labels(X_val_batch, L_val_batch, idx_val_batch, smooth_or_raw, index=worst_index, which_set='VALIDATION WORST')
plot_true_predicted_labels(X_val_batch, L_val_batch, idx_val_batch, smooth_or_raw, which_set='VALIDATION')
#Add epoch results
err_valid += [cost_val_epoch/n_batches_val]
acc_valid += [acc_val_epoch/n_batches_val]
sample_acc_valid_tot += [sample_acc_valid_epoch/n_batches_val]
jacc_perclass_valid = jacc_val_epoch[0, :] / jacc_val_epoch[1, :]
jacc_valid += [np.mean(jacc_perclass_valid)]
worse_indices_valid += [worse_indices_val_epoch]
#Print results (once per epoch)
out_str = "EPOCH %i: Avg cost train %f, acc train %f"+\
", cost val %f, acc val %f, jacc val %f took %f s"
out_str = out_str % (epoch, err_train[epoch],
acc_train[epoch],
err_valid[epoch],
acc_valid[epoch],
jacc_valid[epoch],
time.time()-start_time)
out_str2 = 'Per sample accuracy (ratios ' + str(ratios) + ') '
out_str2 += ' train ' +str(sample_acc_train_tot[epoch])
out_str2 += ' valid ' + str(sample_acc_valid_tot[epoch])
print out_str
print out_str2
# Early stopping and saving stuff
with open(os.path.join(savepath, "fcn1D_output.log"), "a") as f:
f.write(out_str + "\n")
if epoch == 0:
best_jacc_val = jacc_valid[epoch]
elif epoch > 1 and jacc_valid[epoch] > best_jacc_val:
print('saving best (and last) model')
best_jacc_val = jacc_valid[epoch]
patience = 0
np.savez(os.path.join(savepath, 'new_fcn1D_model_best.npz'),
*lasagne.layers.get_all_param_values(simple_net_output))
np.savez(os.path.join(savepath , "fcn1D_errors_best.npz"),
err_train=err_train, acc_train=acc_train,
err_valid=err_valid, acc_valid=acc_valid, jacc_valid=jacc_valid)
np.savez(os.path.join(savepath, 'new_fcn1D_model_last.npz'),
*lasagne.layers.get_all_param_values(simple_net_output))
np.savez(os.path.join(savepath , "fcn1D_errors_last.npz"),
err_train=err_train, acc_train=acc_train,
err_valid=err_valid, acc_valid=acc_valid, jacc_valid=jacc_valid)
else:
patience += 1
print('saving last model')
np.savez(os.path.join(savepath, 'new_fcn1D_model_last.npz'),
*lasagne.layers.get_all_param_values(simple_net_output))
np.savez(os.path.join(savepath , "fcn1D_errors_last.npz"),
err_train=err_train, acc_train=acc_train,
err_valid=err_valid, acc_valid=acc_valid, jacc_valid=jacc_valid)
# Finish training if patience has expired or max nber of epochs reached
if patience == max_patience or epoch == num_epochs-1:
if savepath != loadpath:
print('Copying model and other training files to {}'.format(loadpath))
copy_tree(savepath, loadpath)
break
# + deletable=true editable=true
# + deletable=true editable=true
worse_train = np.sort(worse_indices_train[4])
for i in worse_train:
print int(i)
# + deletable=true editable=true
worse_valid = np.sort(worse_indices_valid[4])
for i in worse_valid:
print int(i)
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
#####################################################################################
# Stuff for computational graphs and profiling functions (to analyse theano behavior)
#####################################################################################
# with open('/u/larocste/cortical_layers/train_fct_debugprint_stephanie.txt', 'w') as save_file:
# theano.printing.debugprint(train_fn, file=save_file)
# train_fn.profile.summary()
# with open('/u/larocste/cortical_layers/test_fct_debugprint_stephanie.txt', 'w') as save_file:
# theano.printing.debugprint(valid_fn, file=save_file)
# valid_fn.profile.summary()
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
def show_feature_maps(net, layers_name, subset=25, type = 'grayscale'):
#Show feature maps for the net's layers_name in grayscale or regular plot
#(example below)
p = lasagne.layers.get_all_param_values(net[layers_name])
nb = min(subset, p[0].shape[0])
if type =='grayscale':
for i in range(subset):
plt.imshow(make_2Darray(p[0][i, 0, :], height = 5), cmap='gray')
plt.figure()
elif type =='regular':
for i in range(subset):
plt.plot(p[0][i, 0, :])
plt.show()
# + deletable=true editable=true
show_feature_maps(net, layers_name = 'relu3_3',subset=10, type='grayscale')
# + deletable=true editable=true
show_feature_maps(net, 'relu3_3',subset =10, type='regular')
# + deletable=true editable=true
# + deletable=true editable=true
# + deletable=true editable=true
def plot_error_acc_curves(err_train = err_train, err_valid = err_valid,
acc_train = acc_train, acc_valid=acc_valid, jacc_valid =jacc_valid):
plt.title('Cost error')
plt.plot(err_train, 'blue', label='cost train')
plt.plot(err_valid, 'orange', label='cost valid')
#plt.ylim((0,2))
plt.legend()
plt.figure()
plt.title('Accuracy')
plt.plot(acc_train, 'blue', label='acc train')
plt.plot(acc_valid, 'orange', label='acc valid')
plt.plot(jacc_valid, 'green', label='jaccard valid')
plt.legend()
# + deletable=true editable=true
plot_error_acc_curves()
# + [markdown] deletable=true editable=true
# ## Ground truth labels colors :
#
# - Purple : label 0 (padding)
# - Purple/blue : label 1 (layers 1)
# - blue : label 2 (layer 2-3)
# - turkoise : label 3 (layers 4 ... almost never present in the predicted labels)
# - green : label 4 (layer 5-6)
# - yellow : label 5 (nonsense)
# + [markdown] deletable=true editable=true
# ### Thoughts after version 2
#
# - Batch normalization (BN) helped a lot with the accuracy. 25% to 80% accuracy by adding BN to each convlayer
# - Label 3 seems really hard to predict. Often predicted as label 4
# - true labels distribution (per class) [4607, 1539, 5637, 976, 6041, 1200]
# - pred labels distribution (per class) [4980, 1470, 6702, 0, 6188, 660]
# - Can't learn (yet) the nonsense label
# - Learning rate 0.005 seems too big --lot of fluctuations in the cost error (0.001 seems fine, maybe lower could be good too)
# + [markdown] deletable=true editable=true
# ### Thoughts after version 3
#
# - Can now achieve 93-95% training accuracy, 90-91% validation accuracy
# - Can now predict class 3 and nonsense labels (better than in version 2)
#
# - Per sample accuracy as early stopping criterion?
# - Use raw+smooth? (instead of just raw)
# - Try to simplify the model (simpler the better)
# + [markdown] deletable=true editable=true
# ### Thoughts after version 4
#
# - Using raw+smooth (i.e. 2 in_channels) requires use_threads=False. If set to True, each epoch takes ~500sec instead of ~300sec. Back to ~300sec when set to False. When only 1 in_channel, what is faster? (Does it change something).
# + [markdown] deletable=true editable=true
# ## Best hyper-parameters so far
#
# - n_filters = 64
# - filter_size = [7, 15, 25, 49]
# - depth = 4
# - weight_decay = 0.001
# - num epochs max = 500
# - max patience = 25
# - learning rate initial value = 0.0005
# - smooth or raw ? raw
# - batch size = [500, 500, 1]
# - shuffle at each epoch ? True n_subset = 0
#
#
# ### Achieves
#
# - Training cost : 0.22
# - Validation cost : 0.23
#
# - Training accuracy (per pixel) : 94.1%
# - Validation accuracy (per pixel) : 91.4%
# - Validation jaccard accuracy : 80.6%
#
# ## Better hyperparameters have been found (in terms of % of the sample that achives >90%)
# + deletable=true editable=true
|
cortical_layers/train_simple_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy
import os
# +
# MNIST 데이터 불러오기
(X_train, Y_train), (X_test, Y_test) = tf.keras.datasets.mnist.load_data()
# +
X_train = X_train.reshape(X_train.shape[0], 784).astype('float32')
X_test = X_test.reshape(X_test.shape[0], 784).astype('float32')
Y_train = tf.keras.utils.to_categorical(Y_train, 10)
Y_test = tf.keras.utils.to_categorical(Y_test, 10)
# +
#모델 설계
# method 1
input_Layer = tf.keras.layers.Input(shape=(784,))
x = tf.keras.layers.Dense(512, activation='sigmoid')(input_Layer)
Out_Layer= tf.keras.layers.Dense(10, activation='softmax')(x)
model = tf.keras.Model(inputs=[input_Layer], outputs=[Out_Layer])
model.summary()
# +
# 모델 컴파일
loss=tf.keras.losses.categorical_crossentropy
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
metric=tf.keras.metrics.categorical_accuracy
model.compile(loss=loss,
optimizer=optimizer,
metrics=[metric])
# 베스트 모델 저장을 위한 디렉토리 선언
MODEL_DIR = './MNIST_model/'
if not os.path.exists(MODEL_DIR):
os.mkdir(MODEL_DIR)
## 저장한 모델의 주소와 이름.
modelpath="./MNIST_model/{epoch:02d}-{val_loss:.4f}.hdf5"
## 사용할 callback 함수 선언.
callback_list=[tf.keras.callbacks.ModelCheckpoint(filepath=modelpath, monitor='val_loss', verbose=1, save_best_only=True),
tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)]
# 모델의 실행
# validation_data 옵션으로 테스트 데이터만 넣어주어서 검증 데이터 분류가 가능
history = model.fit(X_train, Y_train, validation_split=0.1 , epochs=30, batch_size=200, verbose=1, callbacks=callback_list)
# 테스트 정확도 출력
print("\n Test Accuracy: %.4f" % (model.evaluate(X_test, Y_test)[1]))
# 테스트 셋의 오차
y_vloss = history.history['val_loss']
# 학습셋의 오차
y_loss = history.history['loss']
# 그래프로 표현
x_len = numpy.arange(len(y_loss))
plt.plot(x_len, y_vloss, marker='.', c="red", label='Testset_loss')
plt.plot(x_len, y_loss, marker='.', c="blue", label='Trainset_loss')
# 그래프에 그리드를 주고 레이블을 표시
plt.legend(loc='upper right')
# plt.axis([0, 20, 0, 0.35])
plt.grid()
plt.xlabel('epoch')
plt.ylabel('loss')
plt.show()
|
tensorflow/day4/answer/.ipynb_checkpoints/A_04_01_NN_MNIST-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import math
import os
from itertools import count, groupby
from operator import itemgetter
import matplotlib.pyplot as plt
import numpy as np
import torch
from torch import nn, optim
from torch.distributions import Bernoulli, Normal
from torch.nn import functional as F
from torch.utils.data import DataLoader
from torchvision.datasets import MNIST
from torchvision.transforms import ToTensor
from pytorch_lightning import LightningModule, Trainer
# +
@torch.no_grad()
def get_digit_samples():
by_digit = itemgetter(1)
mnist = MNIST(os.getcwd(), transform=ToTensor())
mnist = sorted(mnist, key=by_digit)
mnist = groupby(mnist, key=by_digit)
samples = []
for digit, grp in mnist:
x, y = next(grp)
samples.append(x.view(-1))
return torch.stack(samples)
@torch.no_grad()
def sweep_variable_across_samples(vae, samples, i, sweep):
"""Sweeps a single latent variable
Arguments
---------
vae : torch.Module
A VAE module; must have a decode method
samples : n-by-z array-like
Contains n samples of z latent variables
i : int < z
The latent variable to sweep
sweep : array
The values to use in sweeping z
"""
# XXX dumb, unvectorized version
recons = []
for sample in samples:
recons.append([])
for val in sweep:
sample[i] = val
# Use just means as image
img, _ = vae.decode(sample)
recons[-1].append(img.detach().numpy())
return np.array(recons)
@torch.no_grad()
def plot_sweep_grid(origs, recons, sweepvals):
idx = count(1)
fig = plt.figure(figsize=(15, 13))
fig.subplots_adjust(hspace=0, wspace=0)
for i in range(10):
plt.subplot(10, 11, next(idx))
plt.imshow(origs[i].reshape(28, 28))
plt.xticks([])
plt.yticks([])
if i == 0:
plt.title('Orig')
for j in range(10):
plt.subplot(10, 11, next(idx))
plt.imshow(recons[i][j].reshape(28, 28))
plt.xticks([])
plt.yticks([])
if i == 0:
plt.title(f'{sweepvals[j]:.2f}')
plt.show()
@torch.no_grad()
def plot_all_sweeps(model):
digits = get_digit_samples()
digit_encodings, *_ = model(digits)
sweep_range = torch.linspace(-4, 4, steps=10)
return digit_encodings, sweep_range
# for i in range(20):
for i in range(1):
print(f'Sweeping reconstructions over latent variable no. {i}')
recons_by_var = sweep_variable_across_samples(model,
digit_encodings.clone(),
i,
sweep_range)
plot_sweep_grid(digits.detach().numpy(), recons_by_var, sweep_range)
return digit_encodings, sweep_range
@torch.no_grad()
def zeroth_mu_sigma(enc, model):
m, s = model.decode(enc)
s = F.softplus(s)
m0, s0 = m[0], s[0]
plt.subplot(221)
plt.imshow(m0.reshape(28, 28), norm=None, cmap='gray', vmin=0.0, vmax=1.0)
plt.xticks([])
plt.yticks([])
plt.subplot(222)
plt.imshow(s0.reshape(28, 28), norm=None, cmap='gray', vmin=0.0, vmax=1.0)
plt.xticks([])
plt.yticks([])
plt.subplot(223)
plt.imshow(m0.reshape(28, 28))
plt.xticks([])
plt.yticks([])
plt.subplot(224)
plt.imshow(s0.reshape(28, 28))
plt.xticks([])
plt.yticks([])
plt.show()
return m, s
# -
# !ls -la betavae_tests
from betavae_tests.train import MnistBetaVAE
# # Recons
with torch.no_grad():
digits = get_digit_samples()
swp = torch.linspace(-4, 4, steps=10)
# +
model_b1 = model = MnistBetaVAE.load_from_checkpoint('betavae_tests/checkpoints/model-beta-1.0.ckpt')
with torch.no_grad():
enc, *_ = model(digits)
recons = sweep_variable_across_samples(model, enc.clone(), 0, swp)
plot_sweep_grid(digits.detach().numpy(), recons, swp)
# +
model_b0 = model = MnistBetaVAE.load_from_checkpoint('betavae_tests/checkpoints/model-beta-0.0.ckpt')
with torch.no_grad():
enc, *_ = model(digits)
recons = sweep_variable_across_samples(model, enc.clone(), 0, swp)
plot_sweep_grid(digits.detach().numpy(), recons, swp)
# +
model_b_small = model = MnistBetaVAE.load_from_checkpoint('betavae_tests/checkpoints/model-beta-0.0001.ckpt')
with torch.no_grad():
enc, *_ = model(digits)
recons = sweep_variable_across_samples(model, enc.clone(), 0, swp)
plot_sweep_grid(digits.detach().numpy(), recons, swp)
# -
with torch.no_grad():
mnist = MNIST(os.getcwd(), transform=ToTensor())
mnist = torch.stack([x.view(-1) for x, _ in mnist])
z_b1, z_b1_mu, z_b1_scale, x, x_b1_mu, x_b1_scale = model_b1(mnist)
z_b1_std = 1e-6 + F.softplus(z_b1_scale)
x_b1_std = 1e-6 + F.softplus(x_b1_scale)
z_b0, z_b0_mu, z_b0_scale, x, x_b0_mu, x_b0_scale = model_b0(mnist)
z_b0_std = 1e-6 + F.softplus(z_b0_scale)
x_b0_std = 1e-6 + F.softplus(x_b0_scale)
z_bsmall, z_bsmall_mu, z_bsmall_scale, x, x_bsmall_mu, x_bsmall_scale = model_b_small(mnist)
z_bsmall_std = 1e-6 + F.softplus(z_bsmall_scale)
x_bsmall_std = 1e-6 + F.softplus(x_bsmall_scale)
plt.figure(figsize=(16, 8))
plt.title('Z vector values flattened')
plt.hist([z_b1.view(-1).numpy(),
z_b0.view(-1).numpy(),
z_bsmall.view(-1).numpy()],
bins=100,
histtype='step',
label=['beta=1', 'beta=0', 'beta=0.0001'])
plt.legend()
plt.show()
plt.figure(figsize=(16, 8))
plt.title('Z means')
plt.hist([z_b1_mu.view(-1).numpy(),
z_b0_mu.view(-1).numpy(),
z_bsmall_mu.view(-1).numpy()],
bins=100,
histtype='step',
label=['beta=1', 'beta=0', 'beta=0.0001'])
plt.legend()
plt.show()
plt.figure(figsize=(16, 8))
plt.title('Z stddevs')
plt.hist([z_b1_std.view(-1).numpy(),
z_b0_std.view(-1).numpy(),
z_bsmall_std.view(-1).numpy()],
bins=100,
histtype='step',
label=['beta=1', 'beta=0', 'beta=0.0001'])
plt.legend()
plt.show()
plt.figure(figsize=(16, 8))
plt.title('X means')
plt.hist([x_b1_mu.view(-1).numpy(),
x_b0_mu.view(-1).numpy(),
x_bsmall_mu.view(-1).numpy()],
bins=20,
label=['beta=1', 'beta=0', 'beta=0.0001'])
plt.legend()
plt.show()
plt.figure(figsize=(16, 8))
plt.title('X stddevs')
plt.hist([x_b1_std.view(-1).numpy(),
x_b0_std.view(-1).numpy(),
x_bsmall_std.view(-1).numpy()],
bins=20,
label=['beta=1', 'beta=0', 'beta=0.0001'])
plt.legend()
plt.show()
|
8_Beta_Tests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
### uncomment to display figures
# %matplotlib inline
# # Demonstrates benchmarking functionality - SASMAS Soil Moisture
# #### Benchmarking dataset information
# The soil moisture content (in mm) dataset consists of time-series of soil water content reflectometer measurements at various depths within the profile (top:0-5cm, shallow:0-30cm, middle:30-60cm, deep:60-90cm) within the Upper Hunter River
# <br>
# ##### Source:
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. and <NAME>., 2007. Goulburn River experimental catchment data set. Water Resources Research, 43(10): W10403.
# This notebook goes through the following steps:
#
# 1. Import required libraries
# 2. Set up benchmarking configuration <br>
# 2.1 Catchments to be benchmarked<br>
# 2.2 Define observation inputs<br>
#
# 3. Create benchmark object<br>
# 4. Add models to be benchmarked <br>
# 4.1 Select or unselect models<br>
# 5. View benchmarking statistics<br>
# 6. View benchmarking plots<br>
# 7. Statistics plotting<br>
# ### 1. Import required libraries
# +
from awrams.benchmarking.benchmark import BenchmarkSoilMoisture
from awrams.utils import datetools as dt
import awrams.benchmarking.meta.sasmas as sasmas
from awrams.utils import config_manager
sys_profile = config_manager.get_system_profile().get_settings()
TRAINING_DATA_PATH = sys_profile['DATA_PATHS']['TRAINING_DATA']
# -
# ### 2. Set up benchmarking configuration
# Comparison against observed streamflow <br>
# You can use your own data in csv form similar to the example provided. <br>
# It just needs to have column names matching the names used in extracting AWRA data
# #### 2.1 Sites to be benchmarked
# Soil moisture comparisons at SASMAS sites
# +
sasmas_data_path = TRAINING_DATA_PATH + '/benchmarking/sasmas/' # # the sasmas data has been pre-processed into 5 files [top, shallow, middle, deep, profile]
site_list = ['G1', 'G2', 'G3', 'G4', 'G5', 'G6', 'K1', 'K2', 'K3', 'K4', 'K5', 'K6', 'M1', 'M2', 'M3',
'M4', 'M5', 'M6', 'M7', 'S1', 'S2', 'S3', 'S4', 'S5', 'S6', 'S7']
mod_site_list = ['SASMAS Soil moisture_' + site for site in site_list]
# -
# ### 3. Create the benchmark object:<br>
#
# An object of "Benchmark" class is created by defining what variable is to be benchmarked.
# Everything else gets progressively added, and statistics are calculated when the observation and model outputs are added.
# +
sas = BenchmarkSoilMoisture("SASMAS", "soil moisture", sasmas.meta)
# Specify benchmarking period
sas.period = dt.dates('2003-2011')
# Add observations and catchment subset [the id list needs to be present in the column names of the observation file]
sas.load(sasmas_data_path,mod_site_list,convert_units=100.)
# -
sorted(sas.sites)
sys_profile['BENCHMARKING']['MONTHLY_REJECTION_THRESHOLD'] = 15 # Minimum number of available obs days before monthly stats are calculated
sys_profile['BENCHMARKING']['SM_MODEL_VARNAMES']
sys_profile['BENCHMARKING']['SM_MODEL_LAYERS']
# ### 4. Add models to be benchmarked
# Any number of models can be simulataneously compared
# This step processes the data and calculates all the statistics [can take a while]
path = sasmas_data_path+"/awral_${v}.csv"
sas.add_model("AWRAMSI.v4_0.AWRAL", csv_path=path)
path = sasmas_data_path+"/AWRAMSI_v5QES_AWRAL_SASMAS_${v}.csv"
sas.add_model("AWRAMSI.v5_0.AWRAL", csv_path=path)
# #### 4.1 Show list of loaded or selected models
# list of loaded models is available with <tab> activated dropdown by typing "et.models."<br>
# can "select" or "unselect" models for displaying
sas.benchmark.top.selection
# ### 5. View benchmarking statistics
# Summary percentiles can be printed out by specifying a statistic from: <br>
# "grand_f", "nse", "bias_relative", "pearsons_r" (default), "mean" <br>
# to the 'stat_percentiles' function<br>
# The timeframe defaults to monthly, but can be specified
#
# These tables are pandas dataframes, so they can be exported to csv
sas.benchmark.deep.stat_percentiles('fobj',freq='m')
sas.benchmark.deep.stat_percentiles('grand_f',freq='m')
sas.benchmark.top.stat_percentiles('bias')
sas.benchmark.shallow.stat_percentiles('nse','daily')
sas.benchmark.shallow.data_percentiles()
# ### 6. View benchmarking plots [time series, Regression]
#
# specify frequency by "freq=d" for daily, "freq=m" for monthly, "freq=y" for yearly<br>
# can customise titles, labels, scaling etc using standard matplotlib keyword arguments
sas.benchmark.top.selection
p = sas.benchmark.shallow.plot_timeseries('G1','raw')
p = sas.benchmark.top.plot_timeseries('G1')
p = sas.benchmark.shallow.plot_timeseries('G1')
p = sas.benchmark.middle.plot_timeseries('G1')
p = sas.benchmark.deep.plot_timeseries('G1')
p = sas.benchmark.profile.plot_timeseries('G1')
p = sas.benchmark.shallow.plot_regression('M1',xlim=[-1,60],ylim=[-1,60])
# ### 7. Statistics plotting
# specify statistic type from "fobj", "nse", "rmse", "bias_relative", "pearsons_r" (default), "mean" and <br> frequency from 'd', 'm', 'y'
p = sas.benchmark.shallow.plot_box('pearsons_r','daily')
p = sas.benchmark.shallow.plot_cdf('pearsons_r')
p = sas.benchmark.shallow.plot_box('nse','daily',ylim=[-1,1])
|
Training/Benchmarking/BenchmarkingDemo_SM_SASMAS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python385jvsc74a57bd027c864c23e117f40a7a4f2f1a3201837d7329c7e3e4f87e64f2d40e2cb9d4f12
# ---
#1: Find the location of 'B' in my_string.
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.index('B'))
# ----------------------
#2: Find the number of occourances of the letter 'o' in the string
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.count('o'))
# ---
#3: Convert all the letters of the string to uppercase.
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.upper())
# ---
#4: Find the index at which the string 'Bitcoin' starts
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.find('Bitcoin'))
# ---
#5: Check if the string starts with letter 'X'
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.startswith('X'))
# ---
#6: Convert all the uppercase letters to lowercase and lowercase to uppercase.
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.swapcase())
# ---
#7: Remove all the spaces from the string
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.replace(' ', ''))
# ---
#8: Replace all the characters 'i' with 'btc'
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.replace('i', 'btc'))
# ---
#9: Split the strings into two parts using ',' as the delimiter
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.split(','))
# ---
#10: Concatenate the two strings
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
my_other_string = "Poor guy!"
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
my_other_string = "Poor guy!"
print(my_string+my_other_string)
# ---
#11: Convert first letter of each word in the string to uppercase
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string.title())
# ---
#12: Return first 12 characters of the string
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string)
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string[:12])
# ---
#13: Return every 7th character of the string starting with the first one.
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print()
# Solution:
my_string = "In 2010, someone paid 10k Bitcoin for two pizzas."
print(my_string[::7])
# ---
|
interviewer/250_python_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import json
from qiskit.finance.applications.ising import portfolio_diversification
from qiskit.providers.ibmq.runtime import UserMessenger, ProgramBackend
def get_portfoliodiversification_solution(n, result):
v = result.eigenstate
if isinstance(v, StateFn):
v = v.to_matrix()
N = n ** 2 + n
index_value = [x for x in range(len(v)) if v[x] == max(v)][0]
string_value = "{0:b}".format(index_value)
while len(string_value) < N:
string_value = '0' + string_value
x_state = list()
for elements in string_value:
if elements == '0':
x_state.append(0)
else:
x_state.append(1)
x_state = np.flip(x_state, axis=0)
return x_state
def diversify_portfolio(backend: ProgramBackend, user_messenger: UserMessenger, **kwargs):
"""Function that does classical-quantum calculation."""
rho = kwargs.pop('rho')
n = kwargs.pop('num_assets')
q = kwargs.pop('num_clusters')
qubitOp = portfolio_diversification.get_operator(rho, n, q)
optimizer = kwargs.pop('optimizer')
initial_point = kwargs.pop('initial_point')
ansatz = kwargs.pop('ansatz')
vqe_inputs = {
'ansatz': ansatz,
'operator': qubitOp,
'optimizer': optimizer,
'initial_point': initial_point,
'measurement_error_mitigation': True,
'shots': 1024'
}
backend_options = {
'backend_name': backend.name()
}
job = provider.runtime.run(
program_id='vqe',
inputs=vqe_inputs,
options=backend_options,
callback=raw_callback
)
raw_result = job.result()
quantum_solution = get_portfoliodiversification_solution(n,raw_result)
ground_level = portfolio_diversification.get_portfoliodiversification_value(rho, n, q, quantum_solution)
return quantum_solution, ground_level
def raw_callback(*args):
intermediate_info = {
'nfev': [],
'parameters': [],
'energy': [],
'stddev': []
}
job_id, (nfev, parameters, energy, stddev) = args
intermediate_info['nfev'].append(nfev)
intermediate_info['parameters'].append(parameters)
intermediate_info['energy'].append(energy)
intermediate_info['stddev'].append(stddev)
def main(backend: ProgramBackend, user_messenger: UserMessenger, **kwargs):
"""This is the main entry point of a runtime program.
The name of this method must not change. It also must have ``backend``
and ``user_messenger`` as the first two positional arguments.
Args:
backend: Backend for the circuits to run on.
user_messenger: Used to communicate with the program user.
kwargs: User inputs.
"""
result = diversify_portfolio(backend, user_messenger, **kwargs)
user_messenger.publish(result, final=True)
# +
import numpy as np
from qiskit.finance.data_providers import *
from qiskit.circuit.library import TwoLocal
stocks = ["TICKER1", "TICKER2"]
n = len(stocks)
rho = np.ones((n,n))
rho[0,1] = 0.8
rho[1,0] = 0.8
data = RandomDataProvider(tickers = stocks,
start = datetime.datetime(2016,1,1),
end = datetime.datetime(2016,1,30))
data.run()
rho = data.get_similarity_matrix()
rho = -1 * rho
q = 1
ansatz = TwoLocal(qubitOp.num_qubits, 'ry', 'cz', reps=5, entanglement='full')
np.random.seed(10) # seed for reproducibility
initial_point = np.random.random(ansatz.num_parameters)
# +
import sys
sys.path.insert(0, '..') # Add qiskit_runtime directory to the path
from qiskit.providers.ibmq.runtime.utils import RuntimeEncoder, RuntimeDecoder
from qiskit.providers.ibmq.runtime import UserMessenger
inputs = {'rho':rho,
'num_assets':n,
'num_clusters':q,
'optimizer':{'name':'COBYLA','max_iter':50},
'ansatz':ansatz,
'initial_point':initial_point
}
backend = Aer.get_backend('qasm_simulator')
user_messenger = UserMessenger()
serialized_inputs = json.dumps(inputs, cls=RuntimeEncoder)
unserialized_inputs = json.loads(serialized_inputs, cls=RuntimeDecoder)
portfolio_diversification.main(backend,user_messenger,**unserialized_inputs)
|
qiskit-runtime/tutorials/Portfolio_Diversification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quantile Regression
#
# [scikit-learn](http://scikit-learn.org/stable/) does not have a quantile regression. [mlinsights](http://www.xavierdupre.fr/app/mlinsights/helpsphinx/index.html) implements a version of it.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# %matplotlib inline
import warnings
warnings.simplefilter("ignore")
# ## Simple example
# We generate some dummy data.
import numpy
X = numpy.random.random(1000)
eps1 = (numpy.random.random(900) - 0.5) * 0.1
eps2 = (numpy.random.random(100)) * 10
eps = numpy.hstack([eps1, eps2])
X = X.reshape((1000, 1))
Y = X.ravel() * 3.4 + 5.6 + eps
from sklearn.linear_model import LinearRegression
clr = LinearRegression()
clr.fit(X, Y)
from mlinsights.mlmodel import QuantileLinearRegression
clq = QuantileLinearRegression()
clq.fit(X, Y)
from pandas import DataFrame
data= dict(X=X.ravel(), Y=Y, clr=clr.predict(X), clq=clq.predict(X))
df = DataFrame(data)
df.head()
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1, figsize=(10, 4))
choice = numpy.random.choice(X.shape[0]-1, size=100)
xx = X.ravel()[choice]
yy = Y[choice]
ax.plot(xx, yy, '.', label="data")
xx = numpy.array([[0], [1]])
y1 = clr.predict(xx)
y2 = clq.predict(xx)
ax.plot(xx, y1, "--", label="L2")
ax.plot(xx, y2, "--", label="L1")
ax.set_title("Quantile (L1) vs Square (L2)");
ax.legend();
# The L1 is clearly less sensible to extremas. The optimization algorithm is based on [Iteratively reweighted least squares](https://en.wikipedia.org/wiki/Iteratively_reweighted_least_squares). It estimates a linear regression with error L2 then reweights each oberservation with the inverse of the error L1.
clq = QuantileLinearRegression(verbose=True, max_iter=20)
clq.fit(X, Y)
clq.score(X,Y)
# ## Regression with various quantiles
import numpy
X = numpy.random.random(1200)
eps1 = (numpy.random.random(900) - 0.5) * 0.5
eps2 = (numpy.random.random(300)) * 2
eps = numpy.hstack([eps1, eps2])
X = X.reshape((1200, 1))
Y = X.ravel() * 3.4 + 5.6 + eps + X.ravel() * X.ravel() * 8
fig, ax = plt.subplots(1, 1, figsize=(10, 4))
choice = numpy.random.choice(X.shape[0]-1, size=100)
xx = X.ravel()[choice]
yy = Y[choice]
ax.plot(xx, yy, '.', label="data")
ax.set_title("Almost linear dataset");
clqs = {}
for qu in [0.1, 0.25, 0.5, 0.75, 0.9]:
clq = QuantileLinearRegression(quantile=qu)
clq.fit(X, Y)
clqs['q=%1.2f' % qu] = clq
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1, figsize=(10, 4))
choice = numpy.random.choice(X.shape[0]-1, size=100)
xx = X.ravel()[choice]
yy = Y[choice]
ax.plot(xx, yy, '.', label="data")
xx = numpy.array([[0], [1]])
for qu in sorted(clqs):
y = clqs[qu].predict(xx)
ax.plot(xx, y, "--", label=qu)
ax.set_title("Various quantiles");
ax.legend();
|
_doc/notebooks/sklearn/quantile_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Libraries and Download the Dataset
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
from tensorflow.keras import Sequential
from tensorflow.keras import layers
import tensorflow_datasets as tfds
from tensorflow import keras
import tensorflow as tf
import matplotlib.pyplot as plt
import numpy as np
ds, info = tfds.load('tf_flowers',
# take 80% for training, 10% for validation, and 10% for testing
split=["train[:80%]", "train[80%:90%]", "train[90%:100%]"],
as_supervised=True,
with_info=True)
train_set, valid_set, test_set = ds
# -
info.splits['train'].num_examples
class_names = info.features['label'].names
class_names
# # Visualize the images
plt.figure(figsize=(10, 10))
i = 0
for image, label in train_set.take(9):
plt.subplot(3, 3, i + 1)
image = tf.image.resize(image, (224, 224))
plt.imshow(image.numpy().astype("uint8"))
plt.title(class_names[label])
plt.axis("off")
i += 1
# # Prepare the Dataset
# +
# declare some variables
batch_size = 32
img_height = 224
img_width = 224
AUTOTUNE = tf.data.AUTOTUNE
# We will use this layer to standardize the pixel values
rescaling_layer = layers.Rescaling(1./255)
def create_dataset(ds, shuffle=False):
if shuffle:
ds = ds.shuffle(1000)
# resize the images
ds = ds.map(
lambda x, y: (tf.image.resize(x, (img_height, img_width)), y)
)
# standardize the pixel values to the [0 ,1] range
ds = ds.map(lambda x, y: (rescaling_layer(x), y))
ds = ds.batch(batch_size)
ds = ds.prefetch(AUTOTUNE)
return ds
train_set = create_dataset(train_set, shuffle=True)
valid_set = create_dataset(valid_set)
test_set = create_dataset(test_set)
# -
# # Extract the Features
# +
from keras.applications.vgg16 import VGG16
img_size = (img_height, img_width, 3)
conv_base = VGG16(input_shape=(img_size),
include_top=False,
weights='imagenet')
# -
conv_base.summary()
# +
def extract_features(dataset):
# here we will store the extracted features and their labels
features = []
labels = []
total_batchs = tf.data.experimental.cardinality(dataset)
current_batch = 1
# loop over the dataset to get batches of images and their labels
for images_batch, labels_batch in dataset:
print("[INFO] processing batch {}/{}".format(current_batch, total_batchs))
# extract the features using the predict method
# the shape will be (32, 7, 7, 512)
features_batch = conv_base.predict(images_batch)
# store the current batch of features and labels in a list
features.append(features_batch)
labels.append(labels_batch)
current_batch += 1
features = np.vstack(features) # shape: (2936, 7, 7, 512)
labels = np.hstack(labels) # shape: (2936,)
# flatten the features
features = features.reshape(features.shape[0], 7 * 7 * 512)
return features, labels
print('[INFO] extracting features from training dataset ...')
train_features, train_labels = extract_features(train_set)
print('[INFO] extracting features from validation dataset ...')
validation_features, validation_labels = extract_features(valid_set)
# -
# # Train a New FC Classifier on the Extracted Features
# +
model = Sequential([
layers.Dense(128, activation='relu', input_dim=7 * 7 * 512),
layers.Dense(5, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(train_features, train_labels,
epochs=10,
validation_data=(validation_features, validation_labels))
# -
# # Visualize the Results
# +
def plot_learning_curves():
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.figure(figsize=(10, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label="Training Accuracy")
plt.plot(val_acc, label="Validation Accuracy")
plt.legend()
plt.grid(True)
plt.subplot(2, 1, 2)
plt.plot(loss, label="Training Loss")
plt.plot(val_loss, label="Validation Loss")
plt.legend()
plt.grid(True)
plt.show()
plot_learning_curves()
# -
# # Transfer Learning with Data Augmentation
# +
data_augmentation = Sequential([
layers.RandomFlip("horizontal", input_shape=(img_size)),
layers.RandomRotation(0.2),
layers.RandomZoom(0.2),
])
base_model = tf.keras.applications.VGG16(
input_shape=img_size,
include_top=False,
weights="imagenet",
)
# freeze all layers of the base model
base_model.trainable = False
model = Sequential([
data_augmentation,
base_model,
layers.Dropout(0.5),
layers.Flatten(),
layers.Dense(128, activation='relu'),
layers.Dense(5)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
epochs=10
history = model.fit(
train_set,
validation_data=valid_set,
epochs=epochs
)
# -
plot_learning_curves()
|
transfer-learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jmuddappa/DeepClassificationBot/blob/master/roberta_cdqa.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="hmGbRIVP6C4l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e7bf5ed5-3b16-4c2f-edbe-5436f05b485c"
# !git clone https://github.com/pytorch/fairseq
# + id="sORWamKjD1Gs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="eafd9510-6260-4002-c37c-447e47892c30"
# %cd fairseq
# !pip install --editable .
# + id="wj2LqLGB6P8m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="084eae04-5798-47ca-ef37-52a792915158"
# !wget https://dl.fbaipublicfiles.com/fairseq/models/roberta.large.tar.gz
# !tar -xvf /content/fairseq/roberta.large.tar.gz
# !bash examples/roberta/commonsense_qa/download_cqa_data.sh
# + id="PqVPGBQf6WoM" colab_type="code" colab={}
# %env MAX_UPDATES=3000
# %env WARMUP_UPDATES=150
# %env LR=1e-05
# %env MAX_SENTENCES=16
# %env SEED=1
# %env ROBERTA_PATH=/roberta.large/model.pt
# %env DATA_DIR=/content/fairseq/data/CommonsenseQA
# %env FAIRSEQ_PATH=/content/fairseq
# %env FAIRSEQ_USER_DIR=/content/fairseq/examples/roberta/commonsense_qa
# !CUDA_VISIBLE_DEVICES=0 fairseq-train --fp16 --ddp-backend=no_c10d $DATA_DIR --user-dir $FAIRSEQ_USER_DIR \
# --restore-file $ROBERTA_PATH \
# --reset-optimizer --reset-dataloader --reset-meters \
# --no-epoch-checkpoints --no-last-checkpoints --no-save-optimizer-state \
# --best-checkpoint-metric accuracy --maximize-best-checkpoint-metric \
# --task commonsense_qa --init-token 0 --bpe gpt2 \
# --arch roberta_large --max-positions 512 \
# --dropout 0.1 --attention-dropout 0.1 --weight-decay 0.01 \
# --criterion sentence_ranking --num-classes 5 \
# --optimizer adam --adam-betas '(0.9, 0.98)' --adam-eps 1e-06 --clip-norm 0.0 \
# --lr-scheduler polynomial_decay --lr $LR \
# --warmup-updates $WARMUP_UPDATES --total-num-update $MAX_UPDATES \
# --max-sentences $MAX_SENTENCES \
# --max-update $MAX_UPDATES \
# --log-format simple --log-interval 25 \
# --seed $SEED
|
roberta_cdqa.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="aBtvWJNQhBM5" colab_type="code" colab={}
from sklearn import model_selection, datasets, neighbors
# load the data
cancer = datasets.load_breast_cancer()
# target
y = cancer.target
# features
X = cancer.data
# + id="p6fQASEChP03" colab_type="code" colab={}
# initalize the estimator
knn = neighbors.KNeighborsClassifier()
# + id="cADtTvf6hb3O" colab_type="code" colab={}
# grid contains k and the weight function
grid = {
'n_neighbors': [1, 3, 5, 7],
'weights': ['uniform', 'distance']
}
# + id="ZZ_uK5A0ho67" colab_type="code" colab={}
# set up the grid search with scoring on precsions and number of folds = 10
gscv = model_selection.GridSearchCV(estimator=knn, param_grid=grid, scoring='precision', cv=10)
# + id="tLqJ_3fBh2Bh" colab_type="code" outputId="bf075dfd-0a91-4171-951e-6735b6885c98" executionInfo={"status": "ok", "timestamp": 1571314594262, "user_tz": -660, "elapsed": 1591, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAp-Td-yKvu76Tg0Swzal8U17btuwNIXFmWVwZo=s64", "userId": "11337101975325054847"}} colab={"base_uri": "https://localhost:8080/", "height": 263}
# start the search
gscv.fit(X, y)
# + id="yPZkBBZGiWuC" colab_type="code" outputId="ae372db9-8dee-4a68-a564-48dc61d693b8" executionInfo={"status": "ok", "timestamp": 1571314594263, "user_tz": -660, "elapsed": 1583, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAp-Td-yKvu76Tg0Swzal8U17btuwNIXFmWVwZo=s64", "userId": "11337101975325054847"}} colab={"base_uri": "https://localhost:8080/", "height": 454}
# view the results
print(gscv.cv_results_)
# + id="H6n1RzMiifkZ" colab_type="code" outputId="cb139684-1c11-4e63-827a-0d8b2dd496e7" executionInfo={"status": "ok", "timestamp": 1571314594265, "user_tz": -660, "elapsed": 1576, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAp-Td-yKvu76Tg0Swzal8U17btuwNIXFmWVwZo=s64", "userId": "11337101975325054847"}} colab={"base_uri": "https://localhost:8080/", "height": 121}
import pandas as pd
# convert the results dictionary to a dataframe
results = pd.DataFrame(gscv.cv_results_)
# select just the hyperparameterizations tried, the mean test scores, order by score and show the top 5 models
print(
results.loc[:,['params','mean_test_score']].sort_values('mean_test_score', ascending=False).head(5)
)
# + id="I92n0QuvpM03" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="28021da8-996b-4b0b-c821-8ac571492b1b" executionInfo={"status": "ok", "timestamp": 1571314633729, "user_tz": -660, "elapsed": 1323, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAp-Td-yKvu76Tg0Swzal8U17btuwNIXFmWVwZo=s64", "userId": "11337101975325054847"}}
# visualize the result
results.loc[:,['params','mean_test_score']].plot.barh(x = 'params')
|
Chapter08/tuning_using_gridsearchcv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What we'll see:
# - Introduction to pandas
# - Preprocessing
# - Regression
# - Time series
# - Machine learning
# - Feature engineering
# - Hyper parameter optimization, model validation
# - Clustering
# - Anomaly detection
# - Neural networks
# Acknowledgement:
# - [Data science complete tutorial](https://github.com/zekelabs/data-science-complete-tutorial)
# - [10 steps to become a data scientist](https://github.com/mjbahmani/10-steps-to-become-a-data-scientist.git)
# - [Python data science handbook](https://github.com/jakevdp/PythonDataScienceHandbook.git)
|
data_mining/0-what_we_will_see.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZRODhCkhxORJ"
# ## **Building a 5 Layered deep neural network for credit card fraud detection using a imbalanced data**
# + id="3CPFf8ZeTkbJ"
import pandas as pd
import numpy as np
import seaborn as sn
# + id="eo4w52Y5UpCl"
data = pd.read_csv('creditcard.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 223} id="a1UljQVaU_Nu" outputId="4fbbef97-0d04-4f44-9673-a24ccf1799e2"
data.head()
# + id="mZy_UhfbXtE_"
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score, precision_score, f1_score, recall_score
# + [markdown] id="53h12hD0w_9A"
# ## Standardization
# + id="YVzMX1EuXVWf"
scaler = StandardScaler()
data['NormalizedAmount'] = scaler.fit_transform(data['Amount'].values.reshape(-1, 1))
# + id="DoJ_QSRdYFOC"
data = data.drop(['Amount', 'Time'], axis = 1)
y = data['Class']
X = data.drop(['Class'], axis = 1)
# + colab={"base_uri": "https://localhost:8080/"} id="RXnbPd96YUAG" outputId="40689dc1-b4dd-4fef-89da-63192a7100c9"
y.head()
# + [markdown] id="t2gsT30fvmEG"
# ## Split Data
# + id="1nzGm5IEYYNG"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 0)
# + id="f6chJeQiYjaT"
train_identity = X_train.index
test_identity = X_test.index
# + id="gkSwlLuoYzgY"
X_train = np.array(X_train)
X_test = np.array(X_test)
y_train = np.array(y_train)
y_test = np.array(y_test)
# + id="zDAvY9TTZCl9"
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
# + [markdown] id="DmkSv4gxv_JF"
# ## Build Deep Neural Network
# + id="oUVWiJm8Y2za"
model = Sequential()
#add input layer
model.add(Dense(input_dim = 29, units = 16, activation = 'relu'))
#add 2nd hidden layer
model.add(Dense(units = 24, activation = 'relu'))
#add dropout layer
model.add(Dropout(0.5))
#add 3rd hidden layer
model.add(Dense(units = 20, activation = 'relu'))
#add 4th hidden layer
model.add(Dense(units = 24, activation = 'relu'))
#add ouptut layer
model.add(Dense(units = 1, activation = 'sigmoid'))
# + colab={"base_uri": "https://localhost:8080/"} id="KKKjxysuY9n5" outputId="c1418253-8951-4a7f-daf6-fbe2704b2618"
model.summary()
# + [markdown] id="Fwl3SzLUwJW9"
# ## Fit Model
# + colab={"base_uri": "https://localhost:8080/"} id="6sW3ql0jZIrA" outputId="4f28d16c-c44a-4f55-c415-aa0a50b04d66"
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
model.fit(X_train, y_train, batch_size = 15, epochs = 5)
# + colab={"base_uri": "https://localhost:8080/"} id="4xgnu9kwZM5i" outputId="bdc32f91-26e6-44f5-cfb5-59d38fdafad0"
model.evaluate(X_test, y_test)
# + id="GzFcpXlKa5vb"
y_pred = model.predict(X_test)
# + id="lTWt-z7ceeji"
import matplotlib.pyplot as plt
from mlxtend.plotting import plot_confusion_matrix
from sklearn.metrics import confusion_matrix
# + [markdown] id="3EwNz3XwzzbX"
# ## Plotting Confusion Matrix
# + colab={"base_uri": "https://localhost:8080/"} id="b_tFNcfNevpG" outputId="8a8ed061-96f1-4d98-aeac-109d7f725a87"
conf_matrix = confusion_matrix(y_test, y_pred.round())
print(conf_matrix)
# + colab={"base_uri": "https://localhost:8080/", "height": 501} id="dIVwWUXsgz9Z" outputId="2d511e66-eb2b-4910-8f3b-2cce94754d8f"
fig, ax = plt.subplots(figsize=(7.5, 7.5))
ax.matshow(conf_matrix, cmap=plt.cm.Blues, alpha=0.3)
for i in range(conf_matrix.shape[0]):
for j in range(conf_matrix.shape[1]):
ax.text(x=j, y=i,s=conf_matrix[i, j], va='center', ha='center', size='xx-large')
plt.xlabel('Predictions', fontsize=18)
plt.ylabel('Actuals', fontsize=18)
plt.title('Confusion Matrix', fontsize=18)
plt.show()
# + [markdown] id="1IfRlR4SwoW8"
# ## Calculating Precision, Recall and F1 Score
# + colab={"base_uri": "https://localhost:8080/"} id="u4quzOdElR5E" outputId="e11c30d3-663a-4f80-9b25-52b1b9aeafd8"
print(precision_score(y_test, y_pred.round()))
print(recall_score(y_test, y_pred.round()))
print(f1_score(y_test, y_pred.round()))
# + [markdown] id="iBx2SSUAxhs0"
# Since the data was imbalanced so accuracy is not a correct parameter to measure the performance of a model so, we used precision and recall to see how well this model performs.
# Still the precision is around 85% that means 15% of non frauds are classified as frauds and recall is around 79% which means 21% of frauds were misclassified as non fruads.
# So, there is still enough space for improvement.
#
|
Credit_Card_Fraud_using_DNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: 181f68e503c14848ca7b19efffda8e928d91ce23811e7019b779e3a955f1e780
# name: python3
# ---
# # Exercise 1
# ### Step 1. Go to https://www.kaggle.com/openfoodfacts/world-food-facts/data
# ### Step 2. Download the dataset to your computer and unzip it.
# ### Step 3. Use the tsv file and assign it to a dataframe called food
# +
import pandas as pd
food = pd.read_csv('~/Desktop/projects/pandas_data/en.openfoodfacts.org.products.tsv', sep='\t')
# -
# ### Step 4. See the first 5 entries
food.head()
# ### Step 5. What is the number of observations in the dataset?
food.shape
# There are 356,027 observations in this dataset.
# ### Step 6. What is the number of columns in the dataset?
print(food.shape[1])
# There are 163 columns in this dataste.
# ### Step 7. Print the name of all the columns.
food.columns
# ### Step 8. What is the name of 105th column?
food.columns[104]
# ### Step 9. What is the type of the observations of the 105th column?
food.dtypes['-glucose_100g']
# ### Step 10. How is the dataset indexed?
food.index
# ### Step 11. What is the product name of the 19th observation?
food.values[18][7]
|
01_Getting_&_Knowing_Your_Data/1. World Food Facts/Exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''.venv'': venv)'
# language: python
# name: python3
# ---
# # 2021, Day 10: Syntax Scoring
from advent import get
from advent.y2021 import d10
# ## Tests
d10.solve(get.sample(2021, 10))
# ## Solution
d10.solve(get.input(2021, 10))
# ## Benchmarking
# %timeit d10.solve(get.input(2021, 10))
|
main/2021/d10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Generating C code for the right-hand-side of the scalar wave equation, in ***curvilinear*** coordinates, using a reference metric formalism
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). In addition, all expressions have been validated against a trusted code (the [original SENR/NRPy+ code](https://bitbucket.org/zach_etienne/nrpy)).
#
# ### NRPy+ Source Code for this module: [ScalarWaveCurvilinear/ScalarWaveCurvilinear_RHSs.py](../edit/ScalarWaveCurvilinear/ScalarWaveCurvilinear_RHSs.py)
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 0. [Preliminaries](#prelim): Reference Metrics and Picking Best Coordinate System to Solve the PDE
# 1. [Example](#example): The scalar wave equation in spherical coordinates
# 1. [Step 1](#contracted_christoffel): Contracted Christoffel symbols $\hat{\Gamma}^i = \hat{g}^{ij}\hat{\Gamma}^k_{ij}$ in spherical coordinates, using NRPy+
# 1. [Step 2](#rhs_scalarwave_spherical): The right-hand side of the scalar wave equation in spherical coordinates, using NRPy+
# 1. [Step 3](#code_validation): Code Validation against `ScalarWave.ScalarWaveCurvilinear_RHSs` NRPy+ Module
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='prelim'></a>
#
# # Preliminaries: Reference Metrics and Picking Best Coordinate System to Solve the PDE \[Back to [top](#toc)\]
# $$\label{prelim}$$
#
# Recall from [NRPy+ tutorial notebook on the Cartesian scalar wave equation](Tutorial-ScalarWave.ipynb), the scalar wave equation in 3D Cartesian coordinates is given by
#
# $$\partial_t^2 u = c^2 \nabla^2 u \text{,}$$
# where $u$ (the amplitude of the wave) is a function of time and Cartesian coordinates in space: $u = u(t,x,y,z)$ (spatial dimension as-yet unspecified), and subject to some initial condition
# $$u(0,x,y,z) = f(x,y,z),$$
#
# with suitable (sometimes approximate) spatial boundary conditions.
#
# To simplify this equation, let's first choose units such that $c=1$. Alternative wave speeds can be constructed
# by simply rescaling the time coordinate, with the net effect being that the time $t$ is replaced with time in dimensions of space; i.e., $t\to c t$:
#
# $$\partial_t^2 u = \nabla^2 u.$$
#
# As we learned in the [NRPy+ tutorial notebook on reference metrics](Tutorial-Reference_Metric.ipynb), reference metrics are a means to pick the best coordinate system for the PDE we wish to solve. However, to take advantage of reference metrics requires first that we generalize the PDE. In the case of the scalar wave equation, this involves first rewriting in [Einstein notation](https://en.wikipedia.org/wiki/Einstein_notation) (with implied summation over repeated indices) via
#
# $$(-\partial_t^2 + \nabla^2) u = \eta^{\mu\nu} u_{,\ \mu\nu} = 0,$$
#
# where $u_{,\mu\nu} = \partial_\mu \partial_\nu u$, and $\eta^{\mu\nu}$ is the contravariant flat-space metric tensor with components $\text{diag}(-1,1,1,1)$.
#
# Next we apply the "comma-goes-to-semicolon rule" and replace $\eta^{\mu\nu}$ with $\hat{g}^{\mu\nu}$ to generalize the scalar wave equation to an arbitrary reference metric $\hat{g}^{\mu\nu}$:
#
# $$\hat{g}^{\mu\nu} u_{;\ \mu\nu} = \hat{g}^{\mu\nu} \hat{\nabla}_{\mu} \hat{\nabla}_{\nu} u = 0,$$
#
# where $\hat{\nabla}_{\mu}$ denotes the [covariant derivative](https://en.wikipedia.org/wiki/Covariant_derivative) with respect to the reference metric basis vectors $\hat{x}^{\mu}$, and $\hat{g}^{\mu \nu} \hat{\nabla}_{\mu} \hat{\nabla}_{\nu} u$ is the covariant
# [D'Alembertian](https://en.wikipedia.org/wiki/D%27Alembert_operator) of $u$.
#
# For example, suppose we wish to model a short-wavelength wave that is nearly spherical. In this case, if we were to solve the wave equation PDE in Cartesian coordinates, we would in principle need high resolution in all three cardinal directions. If instead we chose spherical coordinates centered at the center of the wave, we might need high resolution only in the radial direction, with only a few points required in the angular directions. Thus choosing spherical coordinates would be far more computationally efficient than modeling the wave in Cartesian coordinates.
#
# Let's now expand the covariant scalar wave equation in arbitrary coordinates. Since the covariant derivative of a scalar is equivalent to its partial derivative, we have
# \begin{align}
# 0 &= \hat{g}^{\mu \nu} \hat{\nabla}_{\mu} \hat{\nabla}_{\nu} u \\
# &= \hat{g}^{\mu \nu} \hat{\nabla}_{\mu} \partial_{\nu} u.
# \end{align}
#
# $\partial_{\nu} u$ transforms as a one-form under covariant differentiation, so we have
# $$\hat{\nabla}_{\mu} \partial_{\nu} u = \partial_{\mu} \partial_{\nu} u - \hat{\Gamma}^\tau_{\mu\nu} \partial_\tau u,$$
# where
#
# $$\hat{\Gamma}^\tau_{\mu\nu} = \frac{1}{2} \hat{g}^{\tau\alpha} \left(\partial_\nu \hat{g}_{\alpha\mu} + \partial_\mu \hat{g}_{\alpha\nu} - \partial_\alpha \hat{g}_{\mu\nu} \right)$$
# are the [Christoffel symbols](https://en.wikipedia.org/wiki/Christoffel_symbols) associated with the reference metric $\hat{g}_{\mu\nu}$.
#
# Then the scalar wave equation is written:
# $$0 = \hat{g}^{\mu \nu} \left( \partial_{\mu} \partial_{\nu} u - \hat{\Gamma}^\tau_{\mu\nu} \partial_\tau u\right).$$
#
# Define the contracted Christoffel symbols:
# $$\hat{\Gamma}^\tau = \hat{g}^{\mu\nu} \hat{\Gamma}^\tau_{\mu\nu}.$$
#
# Then the scalar wave equation is given by
# $$0 = \hat{g}^{\mu \nu} \partial_{\mu} \partial_{\nu} u - \hat{\Gamma}^\tau \partial_\tau u.$$
#
# The reference metrics we adopt satisfy
# $$\hat{g}^{t \nu} = -\delta^{t \nu},$$
# where $\delta^{t \nu}$ is the [Kronecker delta](https://en.wikipedia.org/wiki/Kronecker_delta). Therefore the scalar wave equation in curvilinear coordinates can be written
# \begin{align}
# 0 &= \hat{g}^{\mu \nu} \partial_{\mu} \partial_{\nu} u - \hat{\Gamma}^\tau \partial_\tau u \\
# &= -\partial_t^2 u + \hat{g}^{i j} \partial_{i} \partial_{j} u - \hat{\Gamma}^i \partial_i u \\
# \implies \partial_t^2 u &= \hat{g}^{i j} \partial_{i} \partial_{j} u - \hat{\Gamma}^i \partial_i u,
# \end{align}
# where repeated Latin indices denote implied summation over *spatial* components only. This module implements the bottom equation for arbitrary reference metrics satisfying $\hat{g}^{t \nu} = -\delta^{t \nu}$. To gain an appreciation for what NRPy+ accomplishes automatically, let's first work out the scalar wave equation in spherical coordinates by hand:
# <a id='example'></a>
#
# # Example: The scalar wave equation in spherical coordinates \[Back to [top](#toc)\]
# $$\label{example}$$
#
# For example, the spherical reference metric is written
#
# $$\hat{g}_{\mu\nu} = \begin{pmatrix}
# -1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & r^2 & 0 \\
# 0 & 0 & 0 & r^2 \sin^2 \theta \\
# \end{pmatrix}.
# $$
#
# Since the inverse of a diagonal matrix is simply the inverse of the diagonal elements, we can write
# $$\hat{g}^{\mu\nu} = \begin{pmatrix}
# -1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & \frac{1}{r^2} & 0 \\
# 0 & 0 & 0 & \frac{1}{r^2 \sin^2 \theta} \\
# \end{pmatrix}.$$
#
# The scalar wave equation in these coordinates can thus be written
# \begin{align}
# 0 &= \hat{g}^{\mu \nu} \partial_{\mu} \partial_{\nu} u - \hat{\Gamma}^\tau \partial_\tau u \\
# &= \hat{g}^{tt} \partial_t^2 u + \hat{g}^{rr} \partial_r^2 u + \hat{g}^{\theta\theta} \partial_\theta^2 u + \hat{g}^{\phi\phi} \partial_\phi^2 u - \hat{\Gamma}^\tau \partial_\tau u \\
# &= -\partial_t^2 u + \partial_r^2 u + \frac{1}{r^2} \partial_\theta^2
# u + \frac{1}{r^2 \sin^2 \theta} \partial_\phi^2 u - \hat{\Gamma}^\tau \partial_\tau u\\
# \implies \partial_t^2 u &= \partial_r^2 u + \frac{1}{r^2} \partial_\theta^2
# u + \frac{1}{r^2 \sin^2 \theta} \partial_\phi^2 u - \hat{\Gamma}^\tau \partial_\tau u
# \end{align}
#
# The contracted Christoffel symbols
# $\hat{\Gamma}^\tau$ can then be computed directly from the metric $\hat{g}_{\mu\nu}$.
#
# It can be shown (exercise to the reader) that the only nonzero
# components of $\hat{\Gamma}^\tau$ in static spherical polar coordinates are
# given by
# \begin{align}
# \hat{\Gamma}^r &= -\frac{2}{r} \\
# \hat{\Gamma}^\theta &= -\frac{\cos\theta}{r^2 \sin\theta}.
# \end{align}
#
# Thus we have found the Laplacian in spherical coordinates is simply:
#
# \begin{align}
# \nabla^2 u &=
# \partial_r^2 u + \frac{1}{r^2} \partial_\theta^2 u + \frac{1}{r^2 \sin^2 \theta} \partial_\phi^2 u - \hat{\Gamma}^\tau \partial_\tau u\\
# &= \partial_r^2 u + \frac{1}{r^2} \partial_\theta^2 u + \frac{1}{r^2 \sin^2 \theta} \partial_\phi^2 u + \frac{2}{r} \partial_r u + \frac{\cos\theta}{r^2 \sin\theta} \partial_\theta u
# \end{align}
# (cf. http://mathworld.wolfram.com/SphericalCoordinates.html; though note that they defined the angle $\phi$ as $\theta$ and $\theta$ as $\phi$.)
# <a id='contracted_christoffel'></a>
#
# # Step 1: Contracted Christoffel symbols $\hat{\Gamma}^i = \hat{g}^{ij}\hat{\Gamma}^k_{ij}$ in spherical coordinates, using NRPy+ \[Back to [top](#toc)\]
# $$\label{contracted_christoffel}$$
#
# Let's next use NRPy+ to derive the contracted Christoffel symbols
# $$\hat{g}^{ij} \hat{\Gamma}^k_{ij}$$
# in spherical coordinates, where $i\in\{1,2,3\}$ and $j\in\{1,2,3\}$ are spatial indices.
#
# As discussed in the [NRPy+ tutorial notebook on reference metrics](Tutorial-Reference_Metric.ipynb), several reference-metric-related quantities in spherical coordinates are computed in NRPy+ (provided the parameter **`reference_metric::CoordSystem`** is set to **`"Spherical"`**), including the inverse spatial spherical reference metric $\hat{g}^{ij}$ and the Christoffel symbols from this reference metric $\hat{\Gamma}^{i}_{jk}$.
# +
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functionality for handling numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
# reference_metric::CoordSystem can be set to Spherical, SinhSpherical, SinhSphericalv2,
# Cylindrical, SinhCylindrical, SinhCylindricalv2, etc.
# See reference_metric.py and NRPy+ tutorial notebook on
# reference metrics for full list and description of how
# to extend.
par.set_parval_from_str("reference_metric::CoordSystem","Spherical")
par.set_parval_from_str("grid::DIM",3)
rfm.reference_metric()
contractedGammahatU = ixp.zerorank1()
for k in range(3):
for i in range(3):
for j in range(3):
contractedGammahatU[k] += rfm.ghatUU[i][j] * rfm.GammahatUDD[k][i][j]
for k in range(3):
print("contracted GammahatU["+str(k)+"]:")
print(sp.simplify(contractedGammahatU[k]))
# Sadly pretty_print results in garbage output in the generated PDF at the bottom of this notebook.
# sp.pretty_print(sp.simplify(contractedGammahatU[k]))
if k<2:
print("\n\n")
# -
# <a id='rhs_scalarwave_spherical'></a>
#
# # Step 2: The right-hand side of the scalar wave equation in spherical coordinates, using NRPy+ \[Back to [top](#toc)\]
# $$\label{rhs_scalarwave_spherical}$$
#
# Following our [implementation of the scalar wave equation in Cartesian coordinates](Tutorial-ScalarWave.ipynb), we will introduce a new variable $v=\partial_t u$ that will enable us to split the second time derivative into two first-order time derivatives:
#
# \begin{align}
# \partial_t u &= v \\
# \partial_t v &= \hat{g}^{ij} \partial_{i} \partial_{j} u - \hat{\Gamma}^i \partial_i u.
# \end{align}
#
# Adding back the sound speed $c$, we have a choice of a single factor of $c$ multiplying both right-hand sides, or a factor of $c^2$ multiplying the second equation only. We'll choose the latter:
#
# \begin{align}
# \partial_t u &= v \\
# \partial_t v &= c^2 \left(\hat{g}^{ij} \partial_{i} \partial_{j} u - \hat{\Gamma}^i \partial_i u\right).
# \end{align}
#
# Now let's generate the C code for the finite-difference representations of the right-hand sides of the above "time evolution" equations for $u$ and $v$. Since the right-hand side of $\partial_t v$ contains implied sums over $i$ and $j$ in the first term, and an implied sum over $k$ in the second term, we'll find it useful to split the right-hand side into two parts
#
# \begin{equation}
# \partial_t v = c^2 \left(
# {\underbrace {\textstyle \hat{g}^{ij} \partial_{i} \partial_{j} u}_{\text{Part 1}}}
# {\underbrace {\textstyle -\hat{\Gamma}^i \partial_i u}_{\text{Part 2}}}\right),
# \end{equation}
#
# and perform the implied sums in two pieces:
import NRPy_param_funcs as par # NRPy+: Parameter interface
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import grid as gri # NRPy+: Functions having to do with numerical grids
import finite_difference as fin # NRPy+: Finite difference C code generation module
import reference_metric as rfm # NRPy+: Reference metric support
from outputC import lhrh # NRPy+: Core C code output module
# +
# The name of this module ("scalarwave") is given by __name__:
thismodule = __name__
# Step 0: Read the spatial dimension parameter as DIM.
DIM = par.parval_from_str("grid::DIM")
# Step 1: Set the finite differencing order to 4.
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",4)
# Step 2a: Reset the gridfunctions list; below we define the
# full complement of gridfunctions needed by this
# tutorial. This line of code enables us to re-run this
# tutorial without resetting the running Python kernel.
gri.glb_gridfcs_list = []
# Step 2b: Register gridfunctions that are needed as input
# to the scalar wave RHS expressions.
uu, vv = gri.register_gridfunctions("EVOL",["uu","vv"])
# Step 3a: Declare the rank-1 indexed expression \partial_{i} u,
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
uu_dD = ixp.declarerank1("uu_dD")
# Step 3b: Declare the rank-2 indexed expression \partial_{ij} u,
# which is symmetric about interchange of indices i and j
# Derivative variables like these must have an underscore
# in them, so the finite difference module can parse the
# variable name properly.
uu_dDD = ixp.declarerank2("uu_dDD","sym01")
# Step 4: Define the C parameter wavespeed. The `wavespeed`
# variable is a proper SymPy variable, so it can be
# used in below expressions. In the C code, it acts
# just like a usual parameter, whose value is
# specified in the parameter file.
wavespeed = par.Cparameters("REAL",thismodule,"wavespeed", 1.0)
# Step 5: Define right-hand sides for the evolution.
uu_rhs = vv
# Step 5b: The right-hand side of the \partial_t v equation
# is given by:
# \hat{g}^{ij} \partial_i \partial_j u - \hat{\Gamma}^i \partial_i u.
# ^^^^^^^^^^^^ PART 1 ^^^^^^^^^^^^^^^^ ^^^^^^^^^^ PART 2 ^^^^^^^^^^^
vv_rhs = 0
for i in range(DIM):
# PART 2:
vv_rhs -= contractedGammahatU[i]*uu_dD[i]
for j in range(DIM):
# PART 1:
vv_rhs += rfm.ghatUU[i][j]*uu_dDD[i][j]
vv_rhs *= wavespeed*wavespeed
# Step 6: Generate C code for scalarwave evolution equations,
# print output to the screen (standard out, or stdout).
fin.FD_outputC("stdout",
[lhrh(lhs=gri.gfaccess("rhs_gfs","uu"),rhs=uu_rhs),
lhrh(lhs=gri.gfaccess("rhs_gfs","vv"),rhs=vv_rhs)])
# -
# <a id='code_validation'></a>
#
# # Step 3: Code Validation against `ScalarWave.ScalarWaveCurvilinear_RHSs` NRPy+ Module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for the RHSs of the Curvilinear Scalar Wave equation (i.e., uu_rhs and vv_rhs) between
#
# 1. this tutorial and
# 2. the NRPy+ [ScalarWave.ScalarWaveCurvilinear_RHSs](../edit/ScalarWaveCurvilinear/ScalarWaveCurvilinear_RHSs.py) module.
#
# By default, we analyze the RHSs in Spherical coordinates, though other coordinate systems may be chosen.
# +
# Step 7: We already have SymPy expressions for uu_rhs and vv_rhs in
# terms of other SymPy variables. Even if we reset the list
# of NRPy+ gridfunctions, these *SymPy* expressions for
# uu_rhs and vv_rhs *will remain unaffected*.
#
# Here, we will use the above-defined uu_rhs and vv_rhs to
# validate against the same expressions in the
# ScalarWaveCurvilinear/ScalarWaveCurvilinear module,
# to ensure consistency between the tutorial and the
# module itself.
#
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# Step 8: Call the ScalarWaveCurvilinear_RHSs() function from within the
# ScalarWaveCurvilinear/ScalarWaveCurvilinear_RHSs.py module,
# which should do exactly the same as in Steps 1-6 above.
import ScalarWave.ScalarWaveCurvilinear_RHSs as swcrhs
swcrhs.ScalarWaveCurvilinear_RHSs()
# Step 9: Consistency check between the tutorial notebook above
# and the ScalarWaveCurvilinear_RHSs() function from within the
# ScalarWaveCurvilinear/ScalarWaveCurvilinear_RHSs.py module.
print("Consistency check between ScalarWaveCurvilinear tutorial and NRPy+ module:")
if sp.simplify(uu_rhs - swcrhs.uu_rhs) != 0:
print("TEST FAILED: uu_ID_SphericalGaussian - swid.uu_ID = "+str(sp.simplify(uu_rhs - swcrhs.uu_rhs))+"\t\t (should be zero)")
sys.exit(1)
if sp.simplify(vv_rhs - swcrhs.vv_rhs) != 0:
print("TEST FAILED: vv_ID_SphericalGaussian - swid.vv_ID = "+str(sp.simplify(vv_rhs - swcrhs.vv_rhs))+"\t\t (should be zero)")
sys.exit(1)
print("TESTS PASSED!")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ScalarWaveCurvilinear.pdf](Tutorial-ScalarWaveCurvilinear.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-ScalarWaveCurvilinear")
|
Tutorial-ScalarWaveCurvilinear.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
boston = datasets.load_boston()
# +
x = boston.data
y = boston.target
x=x[y<50]
y=y[y<50]
# -
x.shape
# +
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state=666)
# -
# ### sklearn 中的线性回归
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(x_train, y_train)
lin_reg.coef_
lin_reg.intercept_
lin_reg.score(x_test, y_test)
# ### KNN regressor
from sklearn.neighbors import KNeighborsRegressor
knn_reg = KNeighborsRegressor()
knn_reg.fit(x_train, y_train)
knn_reg.score(x_test, y_test)
from sklearn.model_selection import GridSearchCV
param_grid = [
{
"weights": ["uniform"],
"n_neighbors": [i for i in range(1, 11)]
},
{
"weights": ["distance"],
"n_neighbors": [i for i in range(1, 11)],
"p": [i for i in range(1, 6)]
}
]
knn_reg = KNeighborsRegressor()
grid_search = GridSearchCV(knn_reg, param_grid, n_jobs=4, verbose=1)
grid_search.fit(x_train, y_train)
grid_search.best_params_
grid_search.best_score_
grid_search.best_estimator_.score(x_test, y_test)
|
data/mulitple_linear_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iued-uni-heidelberg/DAAD-Training-2021/blob/main/cwb2021experimentsV05.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3KsREtgj4KGB"
# # Working with CWB on Colab
# Author: <NAME>, IÜD, Heidelberg University
#
# Modifying CWB installations and packages to work with colab environment
#
# ### Downloading packages and data
# + id="zAn_DUV0xv3Y"
# !wget https://heibox.uni-heidelberg.de/f/7f1e8929352b4cf4b13a/?dl=1
# + id="RISPCuS9ywar"
# !mv index.html?dl=1 cwb-3.4.22-source.tar.gz
# + id="-9q2g4tHy526"
# !tar xvfz cwb-3.4.22-source.tar.gz
# + [markdown] id="LTjz5aJb_Kxs"
# ### Installing the parser generator 'bison'
# + id="xMCPjvHC2e0_" colab={"base_uri": "https://localhost:8080/"} outputId="06c5efc1-b51b-4b50-ee5c-079df4cc999d"
# !apt-get install flex bison
# + [markdown] id="JtRYAaTI_QVJ"
# ### Replacing the configuration file
# Using correct environment and 'standard' location for installation (otherwise python bindings do not work)
# + id="YF7LROot_cOr"
# !wget https://heibox.uni-heidelberg.de/f/67bb38a210064bc5961e/?dl=1
# !mv /content/cwb-3.4.22/config.mk /content/cwb-3.4.22/config.mk.old.01
# !mv index.html?dl=1 /content/cwb-3.4.22/config.mk
# + id="1b-ylUzM_gTc"
# alternative: editing the file at the line numbers
# !awk '{ if (NR == 42) print "PLATFORM=linux-64"; else print $0}' /content/cwb-3.4.22/config.mk > /content/cwb-3.4.22/config.mk.TMP
# !awk '{ if (NR == 63) print "SITE=standard"; else print $0}' /content/cwb-3.4.22/config.mk.TMP > /content/cwb-3.4.22/config.mk
# + [markdown] id="6iM4NBfQBdyw"
# ### Changing into installation directory and running installation scripts
# + id="ppn0btNhzB1O" outputId="1f1380e9-ff2a-497a-e552-30973da17c0a" colab={"base_uri": "https://localhost:8080/"}
# %cd /content/cwb-3.4.22/
# + id="7zyvBPLDzFT8" outputId="053d9355-279f-46e3-e599-6b40a6fafa6d" colab={"base_uri": "https://localhost:8080/"}
# !pwd
# + id="voZ90vnn7WTi"
# !mkdir -p /usr/local/share/cwb/registry/
# + id="8kt7QWlDzK7M"
# !sudo ./install-scripts/config-basic
# !sudo ./install-scripts/install-linux
# + colab={"base_uri": "https://localhost:8080/"} id="CVuSe3hQ_BDj" outputId="9b1258bf-8f25-47b0-b95c-1ee78717df70"
# %cd /content/
# + [markdown] id="rNjdhnI0By8g"
# ### Downloading and relocating the register of a sample corpus
# The register is placed into the standard cwb location
# + id="oGgO8XnS8mp1"
# !wget https://heibox.uni-heidelberg.de/f/dd3538603aa84dd09a76/?dl=1
# !mv index.html?dl=1 Dickens-1.0.tar.gz
# !tar xvzf Dickens-1.0.tar.gz
# + id="0UkrOMJw8ubE"
# !cp /content/Dickens-1.0/registry/dickens /content/Dickens-1.0/registry/dickens.old.01
# !awk '{ if (NR == 10) print "HOME /content/Dickens-1.0/data"; else print $0}' /content/Dickens-1.0/registry/dickens > /content/Dickens-1.0/registry/dickens.TMP
# !awk '{ if (NR == 12) print "INFO /content/Dickens-1.0/data/.info"; else print $0}' /content/Dickens-1.0/registry/dickens.TMP > /content/Dickens-1.0/registry/dickens
# + id="1zXK1Z9v8zob"
# !mv /content/Dickens-1.0/registry/dickens /usr/local/share/cwb/registry
# + [markdown] id="6RlHVMGcCJFk"
# ### Updating path (only needed if installing into a non-standard location
#
# + id="Zlu6uAee9eJ1"
# # !echo $PATH
# + id="nsUAyqA79-PO"
# # %env PATH=/usr/local/cwb-3.4.22/bin:/usr/local/nvidia/bin:/usr/local/cuda/bin:/usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin:/tools/node/bin:/tools/google-cloud-sdk/bin:/opt/bin
# + id="XwVe-fIX_BCO"
# # !echo $PATH
# + id="6TBYYIgvkcVc"
# # %env PATH=/usr/local/nvidia/bin:/usr/local/cuda/bin:/usr/local/sbin:/usr/local/bin:/usr/sbin:/usr/bin:/sbin:/bin:/tools/node/bin:/tools/google-cloud-sdk/bin:/opt/bin
# + id="-SegqQOf_a4H"
# !pwd
# + [markdown] id="E4GIzdN_Di-v"
# ### Testing interactive Corpus Query Processor (CQP)
#
# + colab={"base_uri": "https://localhost:8080/"} id="OmUjHSSfz41T" outputId="c5af8084-14f2-4012-8e89-4c3710476d4d"
# try these commands in the interactive prompt (just copy and paste them):
# DICKENS;
# "question";
# q
# exit;
# !cqp -e
# + colab={"base_uri": "https://localhost:8080/"} id="Wbbdaxst_020" outputId="1c2aa293-b068-44bd-a417-8e768977c29a"
# !cwb-describe-corpus -h
# + id="iCmNuT3A0XPn"
# !cwb-describe-corpus -s dickens
# + id="L27_yMizBI_k"
# # !cwb-describe-corpus -s -r registry dickens
# + id="HGZpuvwZFLVf"
# %cd /content/
# + [markdown] id="f6nQadtzEgp8"
# ## Installing python interface to CWB
# cwb-ccc
# + id="56JFOM66E1Kt"
# # !python -m pip install cwb-ccc
# !python -m pip install cwb-ccc
# + [markdown] id="cUBCq5DtEtu_"
# ## pandas versions are incompatible
# ERROR: pip's dependency resolver does not currently take into account all the packages that are installed. This behaviour is the source of the following dependency conflicts.
#
# google-colab 1.0.0 requires pandas~=1.1.0; python_version >= "3.0", but you have pandas 1.3.5 which is incompatible.
#
# Successfully installed association-measures-0.2.0 cwb-ccc-0.10.1 pandas-1.3.5 pyyaml-6.0 unidecode-1.3.2
#
# - we go for a compromise, which works for both so far...
# + id="_GKMon8G9xSo"
# !pip show pandas
# + id="UI2vEM5OCI4r"
# !pip install pandas==1.1.5
# click [restart runtime] button!
# + [markdown] id="6lulNEGl9zlR"
# ### Experiments with cwb-ccc
# From the webpage
#
# + id="_ls1FlVLl-U9"
from ccc import Corpora
corpora = Corpora(registry_path="/usr/local/share/cwb/registry/")
# corpora = Corpora("/content/Dickens-1.0/registry")
print(corpora)
corpora.show() # returns a DataFrame
# + id="CE4ETsZY05pa"
corpus = corpora.activate(corpus_name="DICKENS")
# + id="5SvuTHWO2biQ"
from ccc import Corpus
corpus = Corpus(
corpus_name="DICKENS",
registry_path="/usr/local/share/cwb/registry/"
)
# + id="bG0YV4eJ2nxd"
query = r'[word="[A-Z0-9][A-Z0-9][A-Z0-9]+"]'
dump = corpus.query(query)
# + id="SEduaqw7HJop"
dump.df
# + id="Huvnn6YNCnJK"
corpus.attributes_available
# + id="aXdYciaqDRZU"
query = r'"question"'
dump = corpus.query(query)
# + id="TdAvtzWAEBJG"
dump.df
# + id="eEP_kTV5Elik"
dump = corpus.query(
cqp_query=query,
context=20,
context_break='s'
)
# + id="aXiyEK_wFrIe"
dump.df
# + id="vnRPzZXwGtxb"
dump.set_context(
context_left=5,
context_right=10,
context_break='s'
)
# + id="wav8P_RPGwZ2"
dump.df
# + id="9y-q9SdRGz05"
dump.breakdown()
# + id="Ds3ZK761L3iY"
dump.concordance()
# + id="ikj5RzFeMqWt"
dump.concordance(p_show=["word", "lemma"], s_show=["text_id"])
# + id="fESAxu9uM792"
dump.concordance(form="kwic")
# + id="BG0UltJXNFLD"
lines = dump.concordance(
p_show=['word', 'pos', 'lemma'],
form='dataframe'
)
# + id="sCI08tY8T94n"
lines.iloc[0]['dataframe']
# + id="kShVkg2CfgiB"
type(lines.iloc[2]['dataframe'])
# + id="Btv0RCr-iMq-"
lines = dump.concordance(
p_show=['word', 'pos', 'lemma'],
form='dict'
)
# + id="mqjx_VY1qsms"
lines.iloc[0]['dict']
# + id="A0CF_QFyrCoh"
lines = dump.concordance(
p_show=['word', 'pos', 'lemma'],
form='slots'
)
# + id="h9RjA-pGHuZF"
lines.iloc[0]
# + id="pWWpjMrnrNdj"
lines.iloc[0]['lemma']
# + id="aRnleZo6rvgN"
dump = corpus.query(
cqp_query=r'@1[pos="D.*"] @2[pos="NN"] @3[word="question"]',
context=None,
context_break='s',
match_strategy='longest'
)
lines = dump.concordance(form='dataframe')
# + id="MnqLwcSHtIvK"
lines.iloc[1]['dataframe']
# + id="7zLLZRAGts3t"
lines = dump.concordance(form='dict')
# + id="J--lQCDktzDR"
lines.iloc[1]['dict']
# + id="TEqoBL7KuA0Q"
lines = dump.concordance(
form='slots',
p_show=['word', 'lemma'],
slots={"article": [1], "np": [2, 3]}
)
# + id="K6agGHvkupFa"
lines
# + id="GxMthnEXu4RS"
dump.correct_anchors({2: -2, 3: +1})
lines = dump.concordance(
form='slots',
slots={"art": [1],
"np": [2, 3]}
)
# + id="n5Hky4vSvZ6E"
lines
# + id="L_cz5_mny4XZ"
dump = corpus.query(
'[lemma="question"]',
context=10,
context_break='s'
)
# + id="RLk9NlIrzS1m"
dump.collocates(order='log_likelihood')
# + id="Mx5JnkG40YfS"
dump = corpus.query(
'[lemma="answer"]',
context=10,
context_break='s'
)
# + id="ZvbF3NMgz-ma"
dump.collocates(p_query=['lemma'], order='conservative_log_ratio')
# ['lemma', 'pos']
# + id="DSj0WaM6zhmp"
corpus.query('[lemma="question" & pos="N.*"]').breakdown()
# + id="d3vWtAL-v0Y4"
# https://pypi.org/project/cwb-ccc/#anchored-queries
# + colab={"base_uri": "https://localhost:8080/"} id="u9c1bxAHseaX" outputId="665c3484-e243-4f4a-a2e8-a325ec894dfd"
# %tb
# + id="KqxsSReWsTnd"
# # !export CWB_DIR=/usr/local/cwb-3.4.10
# /usr/local/cwb-3.4.22/bin
# + id="GdqTG_2BvkTq"
# # !python --version
# + [markdown] id="Ou8wY0JrIvmK"
# ### todo:
# 1. to add corpus lemmatization & encoding parts
# 2. to add generation of interesting collocations, exporting them as lists
# 3. to add parallel corpus functionality
#
|
cwb2021experimentsV05.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1.乖离率
# G60 (close-60均线)/60均线
# G40 (close-40均线)/40均线
# G90 (close-90均线)/90均线
# G105 (close-105均线)/105均线
2.macd
|
机器学习/算法/框架准备--找特征.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualizing LUAD Gene Expression
# ### importing libraries
import pandas as pd
import numpy as np
# ### reading data
df = pd.read_csv('LUAD_rsem_normal_tumor.sample.txt', sep='\t')
# ### exploring the dataframe
df.head()
display(df.head())
display(df.shape)
# + [markdown] heading_collapsed=true
# ### problems w the data
# 1. index is random
# 2. normal and tumor separated by trial readings
# 3. gene expression is not tidy
# -
# ### we want something like || gene || normal or tumor || expression level ||
# ### get columns
# +
df.columns.tolist()
# -
# ### reshape or "melt" data to make it "stacked" (long format instead of wide format)
# ?pd.DataFrame.melt
df_stack = pd.melt(df, id_vars=['Gene'], value_vars = ['Normal',
'Normal.1',
'Normal.2',
'Normal.3',
'Normal.4',
'Normal.5',
'Normal.6',
'Normal.7',
'Normal.8',
'Normal.9',
'Normal.10',
'Normal.11',
'Normal.12',
'Normal.13',
'Normal.14',
'Tumor',
'Tumor.1',
'Tumor.2',
'Tumor.3',
'Tumor.4',
'Tumor.5',
'Tumor.6',
'Tumor.7',
'Tumor.8',
'Tumor.9',
'Tumor.10',
'Tumor.11',
'Tumor.12',
'Tumor.13',
'Tumor.14'], var_name='Cancerous', value_name='Expression')
df_stack.shape
df_stack.tail()
# ### remove the numbers in 'Cancerous'
for index, data in df_stack.iterrows():
if (data.Cancerous.startswith('Normal')):
df_stack.at[index, 'Cancerous'] = 'Normal'
else:
df_stack.at[index, 'Cancerous'] = 'Tumor'
df_stack
# ### using groupby to calculate mean for normal and tumor
df_stack.groupby(['Cancerous'])['Expression'].mean()
# ### log transform gene expression to normalize, ie. put normal and tumor values on the same scale
expr = df_stack['Expression']
df_stack['Expression_Norm'] = np.log2(expr+1)
df_stack.head()
# ### query
df_stack.query('Expression_Norm > 1.47 & Expression_Norm <1.5').get('Gene').unique()
# ### select a few genes
# say: ARID1A,ARID2,ARNT,ASPSCR1,GNA11,GNAQ,PML,PMX1,PNUTL1,STAG2,STAT5B,ZNF384,ZNF521
# +
selected = ["ARID1A", "ARID2", "ARNT", "ASPSCR1", "GNA11", "GNAQ", "PML", "PMX1", "PNUTL1", "STAG2", "STAT5B", "ZNF384", "ZNF521"]
df_selected = df_stack[df_stack.get('Gene').isin(selected)]
# -
# ### making a boxplot
# ### with matplotlib
# +
import matplotlib.pyplot as plt
df_selected.boxplot(column='Expression_Norm', by='Cancerous')
plt.suptitle("")
plt.title("Lung Adenocarcinoma Comparison For Selected Genes")
plt.xlabel("Cancerous")
plt.ylabel("Log Normalized Gene Expression")
plt.show()
# -
# ### with seaborn
# +
import seaborn as sns
axes = sns.boxplot(x = 'Cancerous', y='Expression_Norm', data=df_selected, )
#axes = sns.swarmplot(x = 'Cancerous', y='Expression_Norm', data=df_selected,color="grey")
plt.title("Lung Adenocarcinoma Comparison For Selected Genes")
plt.xlabel("Cancerous")
plt.ylabel("Log Normalized Gene Expression")
plt.show()
# -
# ### with plotly
# +
import plotly.offline as py
from plotly import graph_objs as go
py.init_notebook_mode(connected=True)
py.iplot({
"data": [go.graph_objs.Box(x=df_selected['Cancerous'],y=df_selected['Expression_Norm'])],
"layout" : go.Layout(dict(title="Lung Adenocarcinoma Comparison For Selected Genes",
xaxis=dict(title="Cancerous"),
yaxis=dict(title="Log Normalized Gene Expression")))
})
# -
|
Pandas & Data Visualization/Visualizing LUAD Gene Expression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
import os
import zipfile
# !bash setup.sh
import sagemaker
from sagemaker_predictive_maintenance import config, preprocess, utils
role = config.role
# -
# ## Data Preparation
# Data gathering, storing, denoising, alignment, curating and querying is THE most complicated aspect of predictive maintenance. Fault reports can for example be buried in pdf scans, fault types being misdiagnosed, fault only discovered weeks after they actually occured, or reported in a different timezone without the timezone information. These are a few of a thousands of pitfalls that may
# await you when trying to collate such real-world dataset at scale. In this scenario we are using a hypothetical dataset where the data is cleaned and aligned.
#
#
# ### Background
# NASA’s Prognostic Center of Excellence established a repository with datasets to be used for benchmarking prognostics and predictive maintenance related algorithms. Among these datasets involves data from a turbofan engine simulation model C-MAPPS (or Commercial Modular Aero Propulsion System Simulation). The references section contains details about the over 100 publications using this dataset. C-MAPPS is a tool used to generate health, control and engine parameters from a simulated turbofan engine. A custom code wrapper was used to inject synthetic faults and continuous degradation trends into a time series of sensor data. Some high level characteristics of this dataset are as follows:
# The data obtained is from a high fidelity simulation of a turbofan engine, but closely models the sensor values of an actual engine. Synthetic noise was added to the dataset to replicate real-world scenarios. The effects of faults are masked due to operational conditions, which is a common trait of most real world systems.
# +
source_data_location = 's3://sagemaker-solutions-us-west-2/Predictive-maintenance-using-machine-learning/data'
#local data folder
data_folder = 'data'
# !aws s3 cp --recursive $source_data_location $data_folder
with zipfile.ZipFile(os.path.join(data_folder, 'CMAPSSData.zip'), "r") as zip_ref:
zip_ref.extractall(data_folder)
# -
# ### Process Data
#
# There are 4 different training dataset, corresponding to 4 different engines, and we will train our models on each individually. But first we must preprocess the dataset. Since the example dataset we're using is small we can run the preprocessing code located in `preprocess.py` on this notebook instance. The preprocessing script computes the RUL (Remaining Useful Life) column for each engine data in the training dataset. We also apply min-max scaling to normalize our sensor readings to between 0 and 1 for each column. We read and preprocess the test data as well. It consists of sensor readings and the actual RUL. See the code in `preprocess.py` for more. To adapt this to your own dataset, you would need to implement your custom preprocessing for your dataset. We plot a few columns from the training data for visualization purposes.
# +
train_df, test_df, columns = preprocess.preprocess_data(data_folder)
o = train_df[0][columns[2:10]][train_df[0]['id'] == 3].plot(subplots=True, sharex=True, figsize=(20,10), title="Train: 8 sensors of Engine 1 before failure")
# -
# ## SageMaker MXNet Estimator
#
# Now we will go over the steps needed to define the MXNet model and train with SageMaker.
# ### Upload processed data to S3 for training
#
# We have to upload the processed data to a location in S3 so that the SageMaker training instance can access the data from that location. We will also, at the same time, upload the test data to the S3 bucket so that we can use that as an input to the trained model for scheduled inference.
# +
from sagemaker.s3 import S3Uploader
bucket = config.solution_bucket
prefix = config.s3_prefix
s3_data_prefix = "s3://{}/{}/{}".format(bucket, prefix, 'data')
# Make local folder for processed data
# ! mkdir -p processed-data
# Upload processed test data for inference
for i in range(len(test_df)):
local_test_file = 'processed-data/test-{}.csv'.format(i)
test_df[i].to_csv(local_test_file)
S3Uploader.upload(local_test_file, s3_data_prefix)
# Upload processed data for training
for i in range(len(train_df)):
local_train_file = 'processed-data/train-{}.csv'.format(i)
train_df[i].to_csv(local_train_file)
S3Uploader.upload(local_train_file, s3_data_prefix)
print('uploaded training data location: {}'.format(s3_data_prefix))
# -
# ### Set model output location
output_location = 's3://{}/{}/output'.format(bucket, prefix)
print('training artifacts will be uploaded to: {}'.format(output_location))
# ### MXNet Model Training script
#
# Training MXNet models using MXNet Estimators is a two-step process. First, you prepare your training script, then second, you run this on SageMaker via an MXNet Estimator. The training script we have prepared for the model is located in the `sagemaker_predictive_maintenance_entry_point` folder.
#
# The training script contains functions to create the model for training and for inference. We also have functions to convert our dataframes into a Gluon Dataset so that it can be efficiently prefetched, transformed into numerical features used by the network and padded so that we can learn from multiple samples in batches.
#
# For more information on how to setup a training script for SageMaker using the MXNet estimator see: https://sagemaker.readthedocs.io/en/stable/using_mxnet.html#preparing-the-mxnet-training-script
# !pygmentize sagemaker_predictive_maintenance/sagemaker_predictive_maintenance_entry_point/sagemaker_predictive_maintenance_entry_point.py
# ### Train MXNet Estimator
#
# Now, we can start the SageMaker training job by creating an MXNet estimator. We pass in the required arguments such as the `entry_point`, `role`, `train_instance_type`, and `train_instance_count` into the MXNet Estimator constructor.
#
# Then we start the training script by calling `fit` on the MXNet Estimator. `fit` takes both required and optional arguments. The required argument here is the S3 location of the training data passed in as a dictionary. We are also adding an optional argument for the job name. This is important because when the training job is complete and SageMaker needs to create a SageMaker Model for real-time inference or batch transformation.
# +
from sagemaker.mxnet import MXNet
training_job_name = config.training_job_name
train_instance_type = 'ml.p3.2xlarge'
m = MXNet(entry_point='sagemaker_predictive_maintenance_entry_point.py',
source_dir='sagemaker_predictive_maintenance/sagemaker_predictive_maintenance_entry_point',
py_version='py3',
role=role,
train_instance_count=1,
train_instance_type=train_instance_type,
output_path=output_location,
hyperparameters={'num-datasets' : len(train_df),
'num-gpus': 1,
'epochs': 200,
'optimizer': 'adam',
'batch-size':1,
'log-interval': 100},
input_mode='File',
train_max_run=7200,
framework_version='1.6.0')
m.fit({'train': s3_data_prefix}, job_name=training_job_name)
# -
# ### Create Transformer Model
#
# We can now call the `transformer` function to create a SageMaker Model with the trained model. The SageMaker Model will have the same name as the training job that just completed. This will ensure that SageMaker stores a reference to the trained model which can be used for predictions later on.
batch_output = 's3://{}/{}/{}'.format(bucket, prefix, 'batch-inference')
transformer = m.transformer(instance_count=1, instance_type='ml.m4.xlarge', output_path=batch_output)
# ### Transform test data using the transformer model
#
# Using the `transformer` SageMaker Model, we can run a [SageMaker Batch Transform](https://docs.aws.amazon.com/sagemaker/latest/dg/batch-transform.html) job to get some predictions on test dataset for the model. Here we have a function that takes some test data in S3 and copies it to a new location where it's used as the input to the `transform` function of the Batch Transformer.
# +
s3_test_key = "{}/data/test-0.csv".format(prefix)
s3_transform_input = "{}/batch-transform-input".format(prefix)
job_name, input_key = utils.get_transform_input(bucket, config.solution_prefix, s3_test_key, s3_transform_input)
transformer.transform(input_key, wait=True)
# -
# ### View model prediction results
#
# Once the transform job terminates, we can see the models predictions for the fractional remaining useful life left for the sensor readings in the `data/test-0.csv`. The predictions are a fraction of `MAX_RUL` which is `130.0`, therefore the Remaining Useful Life predictions can be obtained by multiplying the output with 130
utils.get_transform_output(bucket, prefix, job_name)
|
source/notebooks/sagemaker_predictive_maintenance.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="rwxGnsA92emp"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="CPII1rGR2rF9"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="JtEZ1pCPn--z"
# # Custom training: walkthrough
# + [markdown] colab_type="text" id="GV1F7tVTN3Dn"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/beta/tutorials/eager/custom_training_walkthrough"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/custom_training_walkthrough.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/custom_training_walkthrough.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/eager/custom_training_walkthrough.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="LDrzLFXE8T1l"
# This guide uses machine learning to *categorize* Iris flowers by species. It uses TensorFlow to:
# 1. Build a model,
# 2. Train this model on example data, and
# 3. Use the model to make predictions about unknown data.
#
# ## TensorFlow programming
#
# This guide uses these high-level TensorFlow concepts:
#
# * Use TensorFlow's default [eager execution](https://www.tensorflow.org/guide/eager) development environment,
# * Import data with the [Datasets API](https://www.tensorflow.org/guide/datasets),
# * Build models and layers with TensorFlow's [Keras API](https://keras.io/getting-started/sequential-model-guide/).
#
# This tutorial is structured like many TensorFlow programs:
#
# 1. Import and parse the data sets.
# 2. Select the type of model.
# 3. Train the model.
# 4. Evaluate the model's effectiveness.
# 5. Use the trained model to make predictions.
# + [markdown] colab_type="text" id="yNr7H-AIoLOR"
# ## Setup program
# + [markdown] colab_type="text" id="1J3AuPBT9gyR"
# ### Configure imports
#
# Import TensorFlow and the other required Python modules. By default, TensorFlow uses [eager execution](https://www.tensorflow.org/guide/eager) to evaluate operations immediately, returning concrete values instead of creating a [computational graph](https://www.tensorflow.org/guide/graphs) that is executed later. If you are used to a REPL or the `python` interactive console, this feels familiar.
# + colab={} colab_type="code" id="jElLULrDhQZR"
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="bfV2Dai0Ow2o"
# !pip install tensorflow==2.0.0-beta1
import tensorflow as tf
# + colab={} colab_type="code" id="g4Wzg69bnwK2"
print("TensorFlow version: {}".format(tf.__version__))
print("Eager execution: {}".format(tf.executing_eagerly()))
# + [markdown] colab_type="text" id="Zx7wc0LuuxaJ"
# ## The Iris classification problem
#
# Imagine you are a botanist seeking an automated way to categorize each Iris flower you find. Machine learning provides many algorithms to classify flowers statistically. For instance, a sophisticated machine learning program could classify flowers based on photographs. Our ambitions are more modest—we're going to classify Iris flowers based on the length and width measurements of their [sepals](https://en.wikipedia.org/wiki/Sepal) and [petals](https://en.wikipedia.org/wiki/Petal).
#
# The Iris genus entails about 300 species, but our program will only classify the following three:
#
# * Iris setosa
# * Iris virginica
# * Iris versicolor
#
# <table>
# <tr><td>
# <img src="https://www.tensorflow.org/images/iris_three_species.jpg"
# alt="Petal geometry compared for three iris species: Iris setosa, Iris virginica, and Iris versicolor">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://commons.wikimedia.org/w/index.php?curid=170298">Iris setosa</a> (by <a href="https://commons.wikimedia.org/wiki/User:Radomil">Radomil</a>, CC BY-SA 3.0), <a href="https://commons.wikimedia.org/w/index.php?curid=248095">Iris versicolor</a>, (by <a href="https://commons.wikimedia.org/wiki/User:Dlanglois">Dlanglois</a>, CC BY-SA 3.0), and <a href="https://www.flickr.com/photos/33397993@N05/3352169862">Iris virginica</a> (by <a href="https://www.flickr.com/photos/33397993@N05"><NAME></a>, CC BY-SA 2.0).<br/>
# </td></tr>
# </table>
#
# Fortunately, someone has already created a [data set of 120 Iris flowers](https://en.wikipedia.org/wiki/Iris_flower_data_set) with the sepal and petal measurements. This is a classic dataset that is popular for beginner machine learning classification problems.
# + [markdown] colab_type="text" id="3Px6KAg0Jowz"
# ## Import and parse the training dataset
#
# Download the dataset file and convert it into a structure that can be used by this Python program.
#
# ### Download the dataset
#
# Download the training dataset file using the [tf.keras.utils.get_file](https://www.tensorflow.org/api_docs/python/tf/keras/utils/get_file) function. This returns the file path of the downloaded file.
# + colab={} colab_type="code" id="J6c7uEU9rjRM"
train_dataset_url = "https://storage.googleapis.com/download.tensorflow.org/data/iris_training.csv"
train_dataset_fp = tf.keras.utils.get_file(fname=os.path.basename(train_dataset_url),
origin=train_dataset_url)
print("Local copy of the dataset file: {}".format(train_dataset_fp))
# + [markdown] colab_type="text" id="qnX1-aLors4S"
# ### Inspect the data
#
# This dataset, `iris_training.csv`, is a plain text file that stores tabular data formatted as comma-separated values (CSV). Use the `head -n5` command to take a peak at the first five entries:
# + colab={} colab_type="code" id="FQvb_JYdrpPm"
# !head -n5 {train_dataset_fp}
# + [markdown] colab_type="text" id="kQhzD6P-uBoq"
# From this view of the dataset, notice the following:
#
# 1. The first line is a header containing information about the dataset:
# * There are 120 total examples. Each example has four features and one of three possible label names.
# 2. Subsequent rows are data records, one *[example](https://developers.google.com/machine-learning/glossary/#example)* per line, where:
# * The first four fields are *[features](https://developers.google.com/machine-learning/glossary/#feature)*: these are characteristics of an example. Here, the fields hold float numbers representing flower measurements.
# * The last column is the *[label](https://developers.google.com/machine-learning/glossary/#label)*: this is the value we want to predict. For this dataset, it's an integer value of 0, 1, or 2 that corresponds to a flower name.
#
# Let's write that out in code:
# + colab={} colab_type="code" id="9Edhevw7exl6"
# column order in CSV file
column_names = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
feature_names = column_names[:-1]
label_name = column_names[-1]
print("Features: {}".format(feature_names))
print("Label: {}".format(label_name))
# + [markdown] colab_type="text" id="CCtwLoJhhDNc"
# Each label is associated with string name (for example, "setosa"), but machine learning typically relies on numeric values. The label numbers are mapped to a named representation, such as:
#
# * `0`: Iris setosa
# * `1`: Iris versicolor
# * `2`: Iris virginica
#
# For more information about features and labels, see the [ML Terminology section of the Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/framing/ml-terminology).
# + colab={} colab_type="code" id="sVNlJlUOhkoX"
class_names = ['Iris setosa', 'Iris versicolor', 'Iris virginica']
# + [markdown] colab_type="text" id="dqPkQExM2Pwt"
# ### Create a `tf.data.Dataset`
#
# TensorFlow's [Dataset API](https://www.tensorflow.org/guide/datasets) handles many common cases for loading data into a model. This is a high-level API for reading data and transforming it into a form used for training. See the [Datasets Quick Start guide](https://www.tensorflow.org/get_started/datasets_quickstart) for more information.
#
#
# Since the dataset is a CSV-formatted text file, use the [make_csv_dataset](https://www.tensorflow.org/api_docs/python/tf/data/experimental/make_csv_dataset) function to parse the data into a suitable format. Since this function generates data for training models, the default behavior is to shuffle the data (`shuffle=True, shuffle_buffer_size=10000`), and repeat the dataset forever (`num_epochs=None`). We also set the [batch_size](https://developers.google.com/machine-learning/glossary/#batch_size) parameter.
# + colab={} colab_type="code" id="WsxHnz1ebJ2S"
batch_size = 32
train_dataset = tf.data.experimental.make_csv_dataset(
train_dataset_fp,
batch_size,
column_names=column_names,
label_name=label_name,
num_epochs=1)
# + [markdown] colab_type="text" id="gB_RSn62c-3G"
# The `make_csv_dataset` function returns a `tf.data.Dataset` of `(features, label)` pairs, where `features` is a dictionary: `{'feature_name': value}`
#
# These `Dataset` objects are iterable. Let's look at a batch of features:
# + colab={} colab_type="code" id="iDuG94H-C122"
features, labels = next(iter(train_dataset))
print(features)
# + [markdown] colab_type="text" id="E63mArnQaAGz"
# Notice that like-features are grouped together, or *batched*. Each example row's fields are appended to the corresponding feature array. Change the `batch_size` to set the number of examples stored in these feature arrays.
#
# You can start to see some clusters by plotting a few features from the batch:
# + colab={} colab_type="code" id="me5Wn-9FcyyO"
plt.scatter(features['petal_length'],
features['sepal_length'],
c=labels,
cmap='viridis')
plt.xlabel("Petal length")
plt.ylabel("Sepal length")
plt.show()
# + [markdown] colab_type="text" id="YlxpSyHlhT6M"
# To simplify the model building step, create a function to repackage the features dictionary into a single array with shape: `(batch_size, num_features)`.
#
# This function uses the [tf.stack](https://www.tensorflow.org/api_docs/python/tf/stack) method which takes values from a list of tensors and creates a combined tensor at the specified dimension.
# + colab={} colab_type="code" id="jm932WINcaGU"
def pack_features_vector(features, labels):
"""Pack the features into a single array."""
features = tf.stack(list(features.values()), axis=1)
return features, labels
# + [markdown] colab_type="text" id="V1Vuph_eDl8x"
# Then use the [tf.data.Dataset.map](https://www.tensorflow.org/api_docs/python/tf/data/dataset/map) method to pack the `features` of each `(features,label)` pair into the training dataset:
# + colab={} colab_type="code" id="ZbDkzGZIkpXf"
train_dataset = train_dataset.map(pack_features_vector)
# + [markdown] colab_type="text" id="NLy0Q1xCldVO"
# The features element of the `Dataset` are now arrays with shape `(batch_size, num_features)`. Let's look at the first few examples:
# + colab={} colab_type="code" id="kex9ibEek6Tr"
features, labels = next(iter(train_dataset))
print(features[:5])
# + [markdown] colab_type="text" id="LsaVrtNM3Tx5"
# ## Select the type of model
#
# ### Why model?
#
# A *[model](https://developers.google.com/machine-learning/crash-course/glossary#model)* is a relationship between features and the label. For the Iris classification problem, the model defines the relationship between the sepal and petal measurements and the predicted Iris species. Some simple models can be described with a few lines of algebra, but complex machine learning models have a large number of parameters that are difficult to summarize.
#
# Could you determine the relationship between the four features and the Iris species *without* using machine learning? That is, could you use traditional programming techniques (for example, a lot of conditional statements) to create a model? Perhaps—if you analyzed the dataset long enough to determine the relationships between petal and sepal measurements to a particular species. And this becomes difficult—maybe impossible—on more complicated datasets. A good machine learning approach *determines the model for you*. If you feed enough representative examples into the right machine learning model type, the program will figure out the relationships for you.
#
# ### Select the model
#
# We need to select the kind of model to train. There are many types of models and picking a good one takes experience. This tutorial uses a neural network to solve the Iris classification problem. *[Neural networks](https://developers.google.com/machine-learning/glossary/#neural_network)* can find complex relationships between features and the label. It is a highly-structured graph, organized into one or more *[hidden layers](https://developers.google.com/machine-learning/glossary/#hidden_layer)*. Each hidden layer consists of one or more *[neurons](https://developers.google.com/machine-learning/glossary/#neuron)*. There are several categories of neural networks and this program uses a dense, or *[fully-connected neural network](https://developers.google.com/machine-learning/glossary/#fully_connected_layer)*: the neurons in one layer receive input connections from *every* neuron in the previous layer. For example, Figure 2 illustrates a dense neural network consisting of an input layer, two hidden layers, and an output layer:
#
# <table>
# <tr><td>
# <img src="https://www.tensorflow.org/images/custom_estimators/full_network.png"
# alt="A diagram of the network architecture: Inputs, 2 hidden layers, and outputs">
# </td></tr>
# <tr><td align="center">
# <b>Figure 2.</b> A neural network with features, hidden layers, and predictions.<br/>
# </td></tr>
# </table>
#
# When the model from Figure 2 is trained and fed an unlabeled example, it yields three predictions: the likelihood that this flower is the given Iris species. This prediction is called *[inference](https://developers.google.com/machine-learning/crash-course/glossary#inference)*. For this example, the sum of the output predictions is 1.0. In Figure 2, this prediction breaks down as: `0.02` for *Iris setosa*, `0.95` for *Iris versicolor*, and `0.03` for *Iris virginica*. This means that the model predicts—with 95% probability—that an unlabeled example flower is an *Iris versicolor*.
# + [markdown] colab_type="text" id="W23DIMVPQEBt"
# ### Create a model using Keras
#
# The TensorFlow [tf.keras](https://www.tensorflow.org/api_docs/python/tf/keras) API is the preferred way to create models and layers. This makes it easy to build models and experiment while Keras handles the complexity of connecting everything together.
#
# The [tf.keras.Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) model is a linear stack of layers. Its constructor takes a list of layer instances, in this case, two [Dense](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense) layers with 10 nodes each, and an output layer with 3 nodes representing our label predictions. The first layer's `input_shape` parameter corresponds to the number of features from the dataset, and is required.
# + colab={} colab_type="code" id="2fZ6oL2ig3ZK"
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation=tf.nn.relu, input_shape=(4,)), # input shape required
tf.keras.layers.Dense(10, activation=tf.nn.relu),
tf.keras.layers.Dense(3)
])
# + [markdown] colab_type="text" id="FHcbEzMpxbHL"
# The *[activation function](https://developers.google.com/machine-learning/crash-course/glossary#activation_function)* determines the output shape of each node in the layer. These non-linearities are important—without them the model would be equivalent to a single layer. There are many [available activations](https://www.tensorflow.org/api_docs/python/tf/keras/activations), but [ReLU](https://developers.google.com/machine-learning/crash-course/glossary#ReLU) is common for hidden layers.
#
# The ideal number of hidden layers and neurons depends on the problem and the dataset. Like many aspects of machine learning, picking the best shape of the neural network requires a mixture of knowledge and experimentation. As a rule of thumb, increasing the number of hidden layers and neurons typically creates a more powerful model, which requires more data to train effectively.
# + [markdown] colab_type="text" id="2wFKnhWCpDSS"
# ### Using the model
#
# Let's have a quick look at what this model does to a batch of features:
# + colab={} colab_type="code" id="xe6SQ5NrpB-I"
predictions = model(features)
predictions[:5]
# + [markdown] colab_type="text" id="wxyXOhwVr5S3"
# Here, each example returns a [logit](https://developers.google.com/machine-learning/crash-course/glossary#logits) for each class.
#
# To convert these logits to a probability for each class, use the [softmax](https://developers.google.com/machine-learning/crash-course/glossary#softmax) function:
# + colab={} colab_type="code" id="_tRwHZmTNTX2"
tf.nn.softmax(predictions[:5])
# + [markdown] colab_type="text" id="uRZmchElo481"
# Taking the `tf.argmax` across classes gives us the predicted class index. But, the model hasn't been trained yet, so these aren't good predictions.
# + colab={} colab_type="code" id="-Jzm_GoErz8B"
print("Prediction: {}".format(tf.argmax(predictions, axis=1)))
print(" Labels: {}".format(labels))
# + [markdown] colab_type="text" id="Vzq2E5J2QMtw"
# ## Train the model
#
# *[Training](https://developers.google.com/machine-learning/crash-course/glossary#training)* is the stage of machine learning when the model is gradually optimized, or the model *learns* the dataset. The goal is to learn enough about the structure of the training dataset to make predictions about unseen data. If you learn *too much* about the training dataset, then the predictions only work for the data it has seen and will not be generalizable. This problem is called *[overfitting](https://developers.google.com/machine-learning/crash-course/glossary#overfitting)*—it's like memorizing the answers instead of understanding how to solve a problem.
#
# The Iris classification problem is an example of *[supervised machine learning](https://developers.google.com/machine-learning/glossary/#supervised_machine_learning)*: the model is trained from examples that contain labels. In *[unsupervised machine learning](https://developers.google.com/machine-learning/glossary/#unsupervised_machine_learning)*, the examples don't contain labels. Instead, the model typically finds patterns among the features.
# + [markdown] colab_type="text" id="RaKp8aEjKX6B"
# ### Define the loss and gradient function
#
# Both training and evaluation stages need to calculate the model's *[loss](https://developers.google.com/machine-learning/crash-course/glossary#loss)*. This measures how off a model's predictions are from the desired label, in other words, how bad the model is performing. We want to minimize, or optimize, this value.
#
# Our model will calculate its loss using the `tf.keras.losses.SparseCategoricalCrossentropy` function which takes the model's class probability predictions and the desired label, and returns the average loss across the examples.
# + colab={} colab_type="code" id="QOsi6b-1CXIn"
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# + colab={} colab_type="code" id="tMAT4DcMPwI-"
def loss(model, x, y):
y_ = model(x)
return loss_object(y_true=y, y_pred=y_)
l = loss(model, features, labels)
print("Loss test: {}".format(l))
# + [markdown] colab_type="text" id="3IcPqA24QM6B"
# Use the [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) context to calculate the *[gradients](https://developers.google.com/machine-learning/crash-course/glossary#gradient)* used to optimize our model.
# + colab={} colab_type="code" id="x57HcKWhKkei"
def grad(model, inputs, targets):
with tf.GradientTape() as tape:
loss_value = loss(model, inputs, targets)
return loss_value, tape.gradient(loss_value, model.trainable_variables)
# + [markdown] colab_type="text" id="lOxFimtlKruu"
# ### Create an optimizer
#
# An *[optimizer](https://developers.google.com/machine-learning/crash-course/glossary#optimizer)* applies the computed gradients to the model's variables to minimize the `loss` function. You can think of the loss function as a curved surface (see Figure 3) and we want to find its lowest point by walking around. The gradients point in the direction of steepest ascent—so we'll travel the opposite way and move down the hill. By iteratively calculating the loss and gradient for each batch, we'll adjust the model during training. Gradually, the model will find the best combination of weights and bias to minimize loss. And the lower the loss, the better the model's predictions.
#
# <table>
# <tr><td>
# <img src="https://cs231n.github.io/assets/nn3/opt1.gif" width="70%"
# alt="Optimization algorithms visualized over time in 3D space.">
# </td></tr>
# <tr><td align="center">
# <b>Figure 3.</b> Optimization algorithms visualized over time in 3D space.<br/>(Source: <a href="http://cs231n.github.io/neural-networks-3/">Stanford class CS231n</a>, MIT License, Image credit: <a href="https://twitter.com/alecrad"><NAME></a>)
# </td></tr>
# </table>
#
# TensorFlow has many [optimization algorithms](https://www.tensorflow.org/api_guides/python/train) available for training. This model uses the [tf.train.GradientDescentOptimizer](https://www.tensorflow.org/api_docs/python/tf/train/GradientDescentOptimizer) that implements the *[stochastic gradient descent](https://developers.google.com/machine-learning/crash-course/glossary#gradient_descent)* (SGD) algorithm. The `learning_rate` sets the step size to take for each iteration down the hill. This is a *hyperparameter* that you'll commonly adjust to achieve better results.
# + [markdown] colab_type="text" id="XkUd6UiZa_dF"
# Let's setup the optimizer:
# + colab={} colab_type="code" id="8xxi2NNGKwG_"
optimizer = tf.keras.optimizers.Adam(learning_rate=0.01)
# + [markdown] colab_type="text" id="pJVRZ0hP52ZB"
# We'll use this to calculate a single optimization step:
# + colab={} colab_type="code" id="rxRNTFVe56RG"
loss_value, grads = grad(model, features, labels)
print("Step: {}, Initial Loss: {}".format(optimizer.iterations.numpy(),
loss_value.numpy()))
optimizer.apply_gradients(zip(grads, model.trainable_variables))
print("Step: {}, Loss: {}".format(optimizer.iterations.numpy(),
loss(model, features, labels).numpy()))
# + [markdown] colab_type="text" id="7Y2VSELvwAvW"
# ### Training loop
#
# With all the pieces in place, the model is ready for training! A training loop feeds the dataset examples into the model to help it make better predictions. The following code block sets up these training steps:
#
# 1. Iterate each *epoch*. An epoch is one pass through the dataset.
# 2. Within an epoch, iterate over each example in the training `Dataset` grabbing its *features* (`x`) and *label* (`y`).
# 3. Using the example's features, make a prediction and compare it with the label. Measure the inaccuracy of the prediction and use that to calculate the model's loss and gradients.
# 4. Use an `optimizer` to update the model's variables.
# 5. Keep track of some stats for visualization.
# 6. Repeat for each epoch.
#
# The `num_epochs` variable is the number of times to loop over the dataset collection. Counter-intuitively, training a model longer does not guarantee a better model. `num_epochs` is a *[hyperparameter](https://developers.google.com/machine-learning/glossary/#hyperparameter)* that you can tune. Choosing the right number usually requires both experience and experimentation.
# + colab={} colab_type="code" id="AIgulGRUhpto"
## Note: Rerunning this cell uses the same model variables
# keep results for plotting
train_loss_results = []
train_accuracy_results = []
num_epochs = 201
for epoch in range(num_epochs):
epoch_loss_avg = tf.keras.metrics.Mean()
epoch_accuracy = tf.keras.metrics.SparseCategoricalAccuracy()
# Training loop - using batches of 32
for x, y in train_dataset:
# Optimize the model
loss_value, grads = grad(model, x, y)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# Track progress
epoch_loss_avg(loss_value) # add current batch loss
# compare predicted label to actual label
epoch_accuracy(y, model(x))
# end epoch
train_loss_results.append(epoch_loss_avg.result())
train_accuracy_results.append(epoch_accuracy.result())
if epoch % 50 == 0:
print("Epoch {:03d}: Loss: {:.3f}, Accuracy: {:.3%}".format(epoch,
epoch_loss_avg.result(),
epoch_accuracy.result()))
# + [markdown] colab_type="text" id="2FQHVUnm_rjw"
# ### Visualize the loss function over time
# + [markdown] colab_type="text" id="j3wdbmtLVTyr"
# While it's helpful to print out the model's training progress, it's often *more* helpful to see this progress. [TensorBoard](https://www.tensorflow.org/guide/summaries_and_tensorboard) is a nice visualization tool that is packaged with TensorFlow, but we can create basic charts using the `matplotlib` module.
#
# Interpreting these charts takes some experience, but you really want to see the *loss* go down and the *accuracy* go up.
# + colab={} colab_type="code" id="agjvNd2iUGFn"
fig, axes = plt.subplots(2, sharex=True, figsize=(12, 8))
fig.suptitle('Training Metrics')
axes[0].set_ylabel("Loss", fontsize=14)
axes[0].plot(train_loss_results)
axes[1].set_ylabel("Accuracy", fontsize=14)
axes[1].set_xlabel("Epoch", fontsize=14)
axes[1].plot(train_accuracy_results)
plt.show()
# + [markdown] colab_type="text" id="Zg8GoMZhLpGH"
# ## Evaluate the model's effectiveness
#
# Now that the model is trained, we can get some statistics on its performance.
#
# *Evaluating* means determining how effectively the model makes predictions. To determine the model's effectiveness at Iris classification, pass some sepal and petal measurements to the model and ask the model to predict what Iris species they represent. Then compare the model's prediction against the actual label. For example, a model that picked the correct species on half the input examples has an *[accuracy](https://developers.google.com/machine-learning/glossary/#accuracy)* of `0.5`. Figure 4 shows a slightly more effective model, getting 4 out of 5 predictions correct at 80% accuracy:
#
# <table cellpadding="8" border="0">
# <colgroup>
# <col span="4" >
# <col span="1" bgcolor="lightblue">
# <col span="1" bgcolor="lightgreen">
# </colgroup>
# <tr bgcolor="lightgray">
# <th colspan="4">Example features</th>
# <th colspan="1">Label</th>
# <th colspan="1" >Model prediction</th>
# </tr>
# <tr>
# <td>5.9</td><td>3.0</td><td>4.3</td><td>1.5</td><td align="center">1</td><td align="center">1</td>
# </tr>
# <tr>
# <td>6.9</td><td>3.1</td><td>5.4</td><td>2.1</td><td align="center">2</td><td align="center">2</td>
# </tr>
# <tr>
# <td>5.1</td><td>3.3</td><td>1.7</td><td>0.5</td><td align="center">0</td><td align="center">0</td>
# </tr>
# <tr>
# <td>6.0</td> <td>3.4</td> <td>4.5</td> <td>1.6</td> <td align="center">1</td><td align="center" bgcolor="red">2</td>
# </tr>
# <tr>
# <td>5.5</td><td>2.5</td><td>4.0</td><td>1.3</td><td align="center">1</td><td align="center">1</td>
# </tr>
# <tr><td align="center" colspan="6">
# <b>Figure 4.</b> An Iris classifier that is 80% accurate.<br/>
# </td></tr>
# </table>
# + [markdown] colab_type="text" id="z-EvK7hGL0d8"
# ### Setup the test dataset
#
# Evaluating the model is similar to training the model. The biggest difference is the examples come from a separate *[test set](https://developers.google.com/machine-learning/crash-course/glossary#test_set)* rather than the training set. To fairly assess a model's effectiveness, the examples used to evaluate a model must be different from the examples used to train the model.
#
# The setup for the test `Dataset` is similar to the setup for training `Dataset`. Download the CSV text file and parse that values, then give it a little shuffle:
# + colab={} colab_type="code" id="Ps3_9dJ3Lodk"
test_url = "https://storage.googleapis.com/download.tensorflow.org/data/iris_test.csv"
test_fp = tf.keras.utils.get_file(fname=os.path.basename(test_url),
origin=test_url)
# + colab={} colab_type="code" id="SRMWCu30bnxH"
test_dataset = tf.data.experimental.make_csv_dataset(
test_fp,
batch_size,
column_names=column_names,
label_name='species',
num_epochs=1,
shuffle=False)
test_dataset = test_dataset.map(pack_features_vector)
# + [markdown] colab_type="text" id="HFuOKXJdMAdm"
# ### Evaluate the model on the test dataset
#
# Unlike the training stage, the model only evaluates a single [epoch](https://developers.google.com/machine-learning/glossary/#epoch) of the test data. In the following code cell, we iterate over each example in the test set and compare the model's prediction against the actual label. This is used to measure the model's accuracy across the entire test set.
# + colab={} colab_type="code" id="Tw03-MK1cYId"
test_accuracy = tf.keras.metrics.Accuracy()
for (x, y) in test_dataset:
logits = model(x)
prediction = tf.argmax(logits, axis=1, output_type=tf.int32)
test_accuracy(prediction, y)
print("Test set accuracy: {:.3%}".format(test_accuracy.result()))
# + [markdown] colab_type="text" id="HcKEZMtCOeK-"
# We can see on the last batch, for example, the model is usually correct:
# + colab={} colab_type="code" id="uNwt2eMeOane"
tf.stack([y,prediction],axis=1)
# + [markdown] colab_type="text" id="7Li2r1tYvW7S"
# ## Use the trained model to make predictions
#
# We've trained a model and "proven" that it's good—but not perfect—at classifying Iris species. Now let's use the trained model to make some predictions on [unlabeled examples](https://developers.google.com/machine-learning/glossary/#unlabeled_example); that is, on examples that contain features but not a label.
#
# In real-life, the unlabeled examples could come from lots of different sources including apps, CSV files, and data feeds. For now, we're going to manually provide three unlabeled examples to predict their labels. Recall, the label numbers are mapped to a named representation as:
#
# * `0`: Iris setosa
# * `1`: <NAME>icolor
# * `2`: Iris virginica
# + colab={} colab_type="code" id="kesTS5Lzv-M2"
predict_dataset = tf.convert_to_tensor([
[5.1, 3.3, 1.7, 0.5,],
[5.9, 3.0, 4.2, 1.5,],
[6.9, 3.1, 5.4, 2.1]
])
predictions = model(predict_dataset)
for i, logits in enumerate(predictions):
class_idx = tf.argmax(logits).numpy()
p = tf.nn.softmax(logits)[class_idx]
name = class_names[class_idx]
print("Example {} prediction: {} ({:4.1f}%)".format(i, name, 100*p))
# + colab={} colab_type="code" id="JliO3dfQRcbg"
|
site/en/r2/tutorials/eager/custom_training_walkthrough.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/salmanhiro/MALT90-Clumps-Classification-Deep-Learning/blob/main/Albumentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="NnHW3BrMU-PP" outputId="622f06be-587d-40be-c8f6-ff3786d7ba17"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="s3G_N9cXU_H6" outputId="c4d5bd8a-c0b5-47d9-999d-2826f8b4a00b"
# %cd /content/drive/MyDrive/malt90
# + colab={"base_uri": "https://localhost:8080/"} id="JqJY_N_BVPNH" outputId="e47d2193-64c5-4662-ec6a-e31674571c33"
# !pip install -U albumentations
# !pip uninstall opencv-python-headless
# !pip install opencv-python-headless==4.1.2.30
# + id="qSuzggq5VYOb"
import albumentations as A
import cv2
transform = A.Compose([
A.RandomCrop(width=600, height=600),
A.HorizontalFlip(p=0.5),
A.VerticalFlip(p=0.5),
A.SafeRotate(limit=45),
A.RGBShift(r_shift_limit=10),
A.RandomBrightnessContrast(p=0.2),
])
# + id="pKWCL85KV5m3"
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="Wc-UuTuMWC-Q" outputId="8c2346b8-618e-4cb5-9fce-8db658ebe824"
import matplotlib.pyplot as plt
image = cv2.imread("Gim_ml_split_Gim/train/A/AG0031_Gim.png")
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = image[15:700,110:-10]
plt.figure()
plt.imshow(image)
plt.axis("off")
# + id="3Y0r_iN1WOMP"
img_list = []
for i in range(5):
transformed = transform(image=image)
transformed_image = transformed["image"]
img_list.append(transformed_image)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="AHKGZRxaWRX8" outputId="72ed33b2-ccab-46ff-e506-2d14646d5989"
import matplotlib.pyplot as plt
plt.imshow(img_list[4])
# + id="heetnHtQWZ5l"
# !mkdir Gim_ml_split_Gim/train_augmented
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="l0aNbNpdXzUK" outputId="f4c7c16e-15f4-48d0-dce5-2a68bd84d206"
import os
subset = "test"
dest = "test_crop"
#classes = os.listdir(f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{subset}")
classes = ['A', 'P', 'Q', 'H', 'C']
for class_img in classes:
os.makedirs(f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{dest}/{class_img}", exist_ok=True)
imgs = os.listdir(f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{subset}/{class_img}")
count = 0
print(len(imgs))
for img in imgs:
try:
imagename = f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{subset}/{class_img}/{img}"
image = cv2.imread(imagename)
image = image[15:700,110:-10]
cv2.imwrite(f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{dest}/{class_img}/{img}", image)
#if 700//len(imgs) > 1:
# for i in range(700//len(imgs)):
#
# #image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# transformed = transform(image=image)
# transformed_image = transformed["image"]
# #transformed_image = cv2.cvtColor(transformed_image, cv2.COLOR_BGR2RGB)
# cv2.imwrite(f"/content/drive/MyDrive/malt90/Gim_ml_split_Gim/{dest}/{class_img}/{i}_{img}", transformed_image)
# print(f"{imagename} finished")
# count += 1
# print(f"{count}/{len(imgs)}")
except ValueError:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="fZSfzr4mX61y" outputId="ecef936f-a6a7-4ef2-eb28-491051c39152"
classes
# + id="QgnXnbFXX76y"
|
Albumentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cap_env
# language: python
# name: cap_env
# ---
# # In this notebook the datsets for the predictor will be generated.
# +
# Basic imports
import os
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import datetime as dt
import scipy.optimize as spo
import sys
from time import time
from sklearn.metrics import r2_score, median_absolute_error
# %matplotlib inline
# %pylab inline
pylab.rcParams['figure.figsize'] = (20.0, 10.0)
# %load_ext autoreload
# %autoreload 2
sys.path.append('../../')
import predictor.feature_extraction as fe
import utils.preprocessing as pp
# -
# ## Let's first define the list of parameters to use in each dataset.
# +
# Input values
GOOD_DATA_RATIO = 0.99 # The ratio of non-missing values for a symbol to be considered good
SAMPLES_GOOD_DATA_RATIO = 0.9 # The ratio of non-missing values for an interval to be considered good
train_val_time = -1 # In real time days (-1 is for the full interval)
''' Step days will be fixed. That means that the datasets with longer base periods will have samples
that are more correlated. '''
step_days = 7 # market days
base_days = [7, 14, 28, 56, 112] # In market days
ahead_days = [7, 14, 28, 56] # market days
# -
datasets_params_list_df = pd.DataFrame([(x,y) for x in base_days for y in ahead_days],
columns=['base_days', 'ahead_days'])
datasets_params_list_df
# ## Now, let's define the function to generate each dataset.
def generate_one_set(params):
# print(('-'*70 + '\n {}, {} \n' + '-'*70).format(params['base_days'].values, params['ahead_days'].values))
return params
# ## Finally, let's parallellize the generation of all the datasets, and generate them. (took some code and suggestions from here: http://www.racketracer.com/2016/07/06/pandas-in-parallel/#comments)
# +
from multiprocessing import Pool
num_partitions = datasets_params_list_df.shape[0] #number of partitions to split dataframe
num_cores = 4 #number of cores on your machine
def parallelize_dataframe(df, func):
df_split = np.array_split(df, num_partitions)
pool = Pool(num_cores)
df = pd.concat(pool.map(func, df_split))
pool.close()
pool.join()
return df
# -
parallelize_dataframe(datasets_params_list_df, generate_one_set)
|
notebooks/prod/.ipynb_checkpoints/n00_datasets_generation-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 10.1 - Solution
# ## Signal Classification using Dynamic Graph Convolutional Neural Networks
# After a long journey through the universe before reaching the earth, the cosmic particles interact with the galactic magnetic field $B$.
# As these particles carry a charge $q$ they are deflected in the field by the Lorentz force $F = q \cdot v × B$.
# Sources of cosmic particles are located all over the sky, thus arrival distributions of the cosmic particles are isotropic in general. However, particles originating from the same source generate on top of the isotropic
# arrival directions, street-like patterns from galactic magnetic field deflections.
#
# In this tasks we want to classify whether a simulated set of $500$ arriving cosmic particles contains street-like patterns (signal), or originates from an isotropic background.
#
# Training graph networks can be computationally demanding, thus, we recommend to use a GPU for this task.
# +
from tensorflow import keras
import numpy as np
from matplotlib import pyplot as plt
layers = keras.layers
print("keras", keras.__version__)
# -
# #### Download EdgeConv Layer
# +
import gdown
import os
url = "https://github.com/DeepLearningForPhysicsResearchBook/deep-learning-physics/blob/main/edgeconv.py"
output = 'edgeconv.py'
if os.path.exists(output) == False:
gdown.download(url, output, quiet=False)
from edgeconv import EdgeConv
# -
# ### Download Data
# +
url = "https://drive.google.com/u/0/uc?export=download&confirm=HgGH&id=1XKN-Ik7BDyMWdQ230zWS2bNxXL3_9jZq"
output = 'cr_sphere.npz'
if os.path.exists(output) == False:
gdown.download(url, output, quiet=True)
# -
f = np.load(output)
n_train = 10000
x_train, x_test = f['data'][:-n_train], f['data'][-n_train:]
labels = keras.utils.to_categorical(f['label'], num_classes=2)
y_train, y_test = labels[:-n_train], labels[-n_train:]
print("x_train.shape", x_train.shape)
print("y_train.shape", y_train.shape)
# +
# define coordinates for very first EdgeConv
train_points, test_points = x_train[..., :3], x_test[..., :3]
# Use normalized Energy as features for convolutional layers
train_features, test_features = x_train[..., -1, np.newaxis], x_test[..., -1, np.newaxis]
train_features = np.concatenate([train_features, train_points], axis=-1)
train_input_data = [train_points, train_features]
test_input_data = [test_points, test_features]
# -
# ### Plot an example sky map
# +
def scatter(v, c=None, zlabel="", title="", **kwargs):
def vec2ang(v):
x, y, z = np.asarray(v)
phi = np.arctan2(y, x)
theta = np.arctan2(z, (x * x + y * y) ** .5)
return phi, theta
lons, lats = vec2ang(v)
lons = -lons
fig = plt.figure(figsize=kwargs.pop('figsize', [12, 6]))
ax = fig.add_axes([0.1, 0.1, 0.85, 0.9], projection="hammer")
events = ax.scatter(lons, lats, c=c, s=12, lw=2)
cbar = plt.colorbar(events, orientation='horizontal', shrink=0.85, pad=0.05, aspect=30, label=zlabel)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
return fig
test_id = 0
example_map = x_test[test_id]
fig = scatter(example_map[:, 0:3].T, c=example_map[:, 3], zlabel="Energy (normed)", title = "Event %i" % test_id)
# -
# ### Design DGCNN
# #### Start with defining a kernel network
# Design a kernel network. The input to the kernel network is the central pixel coordinate and the neighborhood pixel coordinates.
# Hint: using `layers.BatchNormalization` can help to stabilize the training process of a DGCNN.
#
# You can make use of the code snippet below.
#
# Note that the output of the DNN should be `(None, nodes)`, where `None` is a placeholder for the batch size.
#
# <em> In this case, we perform subtraction and concatenate the result with the central pixel value to combine translational invariance with local information. </em>
def kernel_nn(data, nodes=16):
d1, d2 = data # get xi ("central" pixel) and xj ("neighborhood" pixels)
dif = layers.Subtract()([d1, d2]) # perform substraction for translational invariance
x = layers.Concatenate(axis=-1)([d1, dif]) # add information on the absolute pixel value
x = layers.Dense(nodes, use_bias=False, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(nodes, use_bias=False, activation="relu")(x)
x = layers.BatchNormalization()(x)
x = layers.Dense(nodes, use_bias=False, activation="relu")(x)
x = layers.BatchNormalization()(x)
return x
# #### Build complete graph network model
# In the first layer, it might be advantageous to choose the next neighbors using the coordinates of the cosmic ray but perform the convolution using their energies also.
# Thus, we input `y = EdgeConv(...)[points_input, feats_input]` into the first EdgeConv layer.
# If we later want to perform a dynamic EdgeConv (we want to update the graph), we simply input `z = EdgeConv(...)(y)`.
#
# To specify the size of the "convolutional filter", make use of the `next_neighbors` argument (searches for $k$ next neighbors for each cosmic ray).
# +
points_input = layers.Input((500, 3))
feats_input = layers.Input((500, 4))
x = EdgeConv(lambda a: kernel_nn(a, nodes=8), next_neighbors=8)([points_input, feats_input]) # conv with fixed graph
x = layers.Activation("relu")(x)
x = EdgeConv(lambda a: kernel_nn(a, nodes=16), next_neighbors=8)([points_input, x]) # conv with fixed graph
x = layers.Activation("relu")(x)
x = EdgeConv(lambda a: kernel_nn(a, nodes=32), next_neighbors=8)([x, x]) # conv with dynamic graph
x = layers.Activation("relu")(x)
x = layers.GlobalAveragePooling1D(name="embedding")(x)
out = layers.Dense(2, name="classification", activation="softmax")(x)
model = keras.models.Model([points_input, feats_input], out)
print(model.summary())
# -
# You can inspect the kernel network using:
model.layers[2].kernel_func.summary()
# <em> The kernel network maps the energies an positions of 2 cosmic rays (the central and the neighbor comsic ray) to 8 features. </em>
# The kernel network in the third layer maps from 16 extracted features (of 2 cosmic rays) to 32 new features and looks like this:
model.layers[6].kernel_func.summary()
# ### Train the model
model.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.Adam(3E-3, decay=1E-4),
metrics=['acc'])
# If you don't have `networkx` or `sklearn` install it by executing:
history = model.fit(train_input_data, y_train, batch_size=64, epochs=4)
# ## Visualization of the underlying graph
# To inspect the changing neighborhood relation (we used a dynamic layer) of the nodes, we visualize the underlying graph structure.
#
# Note that plotting may take some time, so be a bit patient.
# To perform the relative complex plotting, we make use of networkx and sklearn.
# If you don't have installed the packages yet, run the cell below.
import sys
# !{sys.executable} -m pip install sklearn
# !{sys.executable} -m pip install networkx
# +
import tensorflow.keras.backend as K
from sklearn.neighbors import kneighbors_graph
import networkx as nx
edge_layers = [l for l in model.layers if "edge_conv" in l.name]
coord_mask = [np.sum(np.linalg.norm(inp_d[test_id], axis=-1)) == 500 for inp_d in train_input_data]
assert True in coord_mask, "For plotting the spherical graph at least one input has to have 3 dimensions XYZ"
fig, axes = plt.subplots(ncols=len(edge_layers), figsize=(5 * len(edge_layers), 5))
for i, e_layer in enumerate(edge_layers):
points_in, feats_in = model.inputs
coordinates = e_layer.get_input_at(0)
functor = K.function(model.inputs, coordinates)
sample_input = [inp[np.newaxis, test_id] for inp in train_input_data]
if type(e_layer.input) == list:
layer_points, layer_features = functor(sample_input)
else:
layer_points = functor(sample_input)
layer_points = np.squeeze(layer_points)
adj = kneighbors_graph(layer_points, e_layer.next_neighbors)
g = nx.DiGraph(adj)
for c, s in zip(coord_mask, sample_input):
if c == True:
pos = s
break
axes[i].set_title("Graph in %s" % e_layer.name)
nx.draw(g, cmap=plt.get_cmap('viridis'), pos=pos.squeeze()[:, :-1],
node_size=10, width=0.5, arrowsize=5, ax=axes[i])
axes[i].axis('equal')
|
Exercise_10_1_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# ## Importing required packages
# + deletable=true editable=true
import os
import numpy as np
from collections import Counter
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import LinearSVC
from sklearn.metrics import confusion_matrix
# + [markdown] deletable=true editable=true
# ## Function for creating a word dictionary
# + deletable=true editable=true
def make_Dictionary(train_dir):
email_files = [os.path.join(train_dir,f) for f in sorted(os.listdir(train_dir))]
stop_words = set(stopwords.words('english'))
all_words = []
for mail in email_files:
with open(mail) as m:
for i,line in enumerate(m):
if i == 2: #Body of email is only 3rd line of text file
list_words = word_tokenize(line)
all_words += list_words
dictionary = Counter(all_words)
list_to_remove = dictionary.keys()
for item in list_to_remove:
if item.isalpha() == False:
del dictionary[item]
elif len(item) == 1:
del dictionary[item]
elif item in stop_words:
del dictionary[item]
dictionary = dictionary.most_common(5000)
return dictionary
# + [markdown] deletable=true editable=true
# ## Function for the process of Feature Extraction
# + deletable=true editable=true
def extract_features(mail_dir):
email_files = [os.path.join(mail_dir,f) for f in sorted(os.listdir(mail_dir))]
features_matrix = np.zeros((len(email_files),5000))
docID = 0
for mail in email_files:
with open(mail) as m:
for i,line in enumerate(m):
if i == 2:
words = word_tokenize(line)
for word in words:
wordID = 0
for i,d in enumerate(dictionary):
if d[0] == word:
wordID = i
features_matrix[docID,wordID] = words.count(word)
docID = docID + 1
return features_matrix
# + [markdown] deletable=true editable=true
# ## Create a dictionary of words with its frequency
# + deletable=true editable=true
train_dir = 'train-mails'
dictionary = make_Dictionary(train_dir)
# + [markdown] deletable=true editable=true
# ## Prepare feature vectors of each training mail and its labels
# + deletable=true editable=true
train_labels = np.zeros(702)
train_labels[351:701] = 1
train_matrix = extract_features(train_dir)
# + [markdown] deletable=true editable=true
# ## Training Naive Bayes and SVM classifier
# + deletable=true editable=true
NBmodel = MultinomialNB()
NBmodel.fit(train_matrix,train_labels)
SVMmodel = LinearSVC()
SVMmodel.fit(train_matrix,train_labels)
# + [markdown] deletable=true editable=true
# ## Predicting the type of mail of test dataset using our models
# + deletable=true editable=true
test_dir = 'test-mails'
test_labels = np.zeros(260)
test_labels[130:260] = 1
test_matrix = extract_features(test_dir)
NBmodelresult = NBmodel.predict(test_matrix)
SVMmodelresult = SVMmodel.predict(test_matrix)
# + [markdown] deletable=true editable=true
# ## Finding out accuracy of our models
# + deletable=true editable=true
print 'Confusion Matrix of our Naive Bayes Classifier is:'
print confusion_matrix(test_labels,NBmodelresult)
print '\nConfusion Matrix of our SVM Classifier is:'
print confusion_matrix(test_labels,SVMmodelresult)
|
SpamDetection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Random forest fraction correct analysis
# ### Table of contents
# 1. [Data preprocessing](#Data-preprocessing)
# 2. [Fitting random forest](#Fit-random-forest-and-run-10-fold-CV-validation)
# 3. [Feature importance](#Feature-importance)
import sys
sys.path.append('/home/jbourbeau/cr-composition')
print('Added to PYTHONPATH')
# +
import argparse
from collections import defaultdict
import itertools
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import seaborn.apionly as sns
from sklearn.metrics import accuracy_score
from sklearn.model_selection import cross_val_score, StratifiedShuffleSplit, KFold
import composition as comp
import composition.analysis.plotting as plotting
# Plotting-related
sns.set_palette('muted')
sns.set_color_codes()
color_dict = defaultdict()
for i, composition in enumerate(['light', 'heavy', 'total']):
color_dict[composition] = sns.color_palette('muted').as_hex()[i]
# %matplotlib inline
# -
# ## Data preprocessing
# 1. Load simulation dataframe and apply specified quality cuts
# 2. Extract desired features from dataframe
# 3. Get separate testing and training datasets
# +
df, cut_dict = comp.load_sim(return_cut_dict=True)
selection_mask = np.array([True] * len(df))
standard_cut_keys = ['lap_reco_success', 'lap_zenith', 'num_hits_1_30', 'IT_signal',
'max_qfrac_1_30', 'lap_containment', 'energy_range_lap']
for key in standard_cut_keys:
selection_mask *= cut_dict[key]
df = df[selection_mask]
feature_list, feature_labels = comp.get_training_features()
print('training features = {}'.format(feature_list))
X_train, X_test, y_train, y_test, le = comp.get_train_test_sets(df, feature_list, comp_class=True)
print('number training events = ' + str(y_train.shape[0]))
print('number testing events = ' + str(y_test.shape[0]))
# -
# ## Fit random forest and run 10-fold CV validation
pipeline = comp.get_pipeline('RF')
clf_name = pipeline.named_steps['classifier'].__class__.__name__
print('=' * 30)
print(clf_name)
scores = cross_val_score(estimator=pipeline, X=X_train, y=y_train, cv=10, n_jobs=20)
print('CV score: {:.2%} (+/- {:.2%})'.format(scores.mean(), scores.std()))
print('=' * 30)
def get_frac_correct(X_train, X_test, y_train, y_test, comp_list):
pipeline = comp.get_pipeline('RF')
pipeline.fit(X_train, y_train)
test_predictions = pipeline.predict(X_test)
correctly_identified_mask = (test_predictions == y_test)
# Energy-related variables
energy_bin_width = 0.1
energy_bins = np.arange(6.2, 8.1, energy_bin_width)
energy_midpoints = (energy_bins[1:] + energy_bins[:-1]) / 2
log_energy = X_test[:, 0]
# Construct MC composition masks
MC_comp_mask = {}
for composition in comp_list:
MC_comp_mask[composition] = (le.inverse_transform(y_test) == composition)
# Get number of MC comp in each reco energy bin
num_MC_energy, num_MC_energy_err = {}, {}
for composition in comp_list:
num_MC_energy[composition] = np.histogram(log_energy[MC_comp_mask[composition]],
bins=energy_bins)[0]
num_MC_energy_err[composition] = np.sqrt(num_MC_energy[composition])
num_MC_energy['total'] = np.histogram(log_energy, bins=energy_bins)[0]
num_MC_energy_err['total'] = np.sqrt(num_MC_energy['total'])
# Get number of correctly identified comp in each reco energy bin
num_reco_energy, num_reco_energy_err = {}, {}
for composition in comp_list:
num_reco_energy[composition] = np.histogram(
log_energy[MC_comp_mask[composition] & correctly_identified_mask],
bins=energy_bins)[0]
num_reco_energy_err[composition] = np.sqrt(num_reco_energy[composition])
num_reco_energy['total'] = np.histogram(log_energy[correctly_identified_mask], bins=energy_bins)[0]
num_reco_energy_err['total'] = np.sqrt(num_reco_energy['total'])
# Calculate correctly identified fractions as a function of MC energy
reco_frac, reco_frac_err = {}, {}
for composition in comp_list:
# print(composition)
reco_frac[composition], reco_frac_err[composition] = comp.ratio_error(
num_reco_energy[composition], num_reco_energy_err[composition],
num_MC_energy[composition], num_MC_energy_err[composition])
frac_correct_folds[composition].append(reco_frac[composition])
reco_frac['total'], reco_frac_err['total'] = comp.ratio_error(
num_reco_energy['total'], num_reco_energy_err['total'],
num_MC_energy['total'], num_MC_energy_err['total'])
return reco_frac, reco_frac_err
# ## Compute systematic in fraction correct via CV
comp_list = ['light', 'heavy']
# Split data into training and test samples
kf = KFold(n_splits=10)
frac_correct_folds = defaultdict(list)
fold_num = 0
for train_index, test_index in kf.split(X_train):
fold_num += 1
print('Fold number {}...'.format(fold_num))
X_train_fold, X_test_fold = X_train[train_index], X_train[test_index]
y_train_fold, y_test_fold = y_train[train_index], y_train[test_index]
reco_frac, reco_frac_err = get_frac_correct(X_train_fold, X_test_fold,
y_train_fold, y_test_fold,
comp_list)
for composition in comp_list:
frac_correct_folds[composition].append(reco_frac[composition])
frac_correct_folds['total'].append(reco_frac['total'])
frac_correct_sys_err = {key: np.std(frac_correct_folds[key], axis=0) for key in frac_correct_folds}
# +
reco_frac, reco_frac_sterr = get_frac_correct(X_train, X_test,
y_train, y_test,
comp_list)
# Energy-related variables
energy_bin_width = 0.1
energy_bins = np.arange(6.2, 8.1, energy_bin_width)
energy_midpoints = (energy_bins[1:] + energy_bins[:-1]) / 2
step_x = energy_midpoints
step_x = np.append(step_x[0]-energy_bin_width/2, step_x)
step_x = np.append(step_x, step_x[-1]+energy_bin_width/2)
# Plot fraction of events vs energy
def plot_steps(x, y, y_err, ax, color, label):
step_x = x
x_widths = x[1:]-x[:-1]
if len(np.unique(x_widths)) != 1:
raise('Unequal bins...')
x_width = np.unique(x_widths)[0]
step_x = np.append(step_x[0]-x_width/2, step_x)
step_x = np.append(step_x, step_x[-1]+x_width/2)
step_y = y
step_y = np.append(step_y[0], step_y)
step_y = np.append(step_y, step_y[-1])
err_upper = y + y_err
err_upper = np.append(err_upper[0], err_upper)
err_upper = np.append(err_upper, err_upper[-1])
err_lower = y - y_err
err_lower = np.append(err_lower[0], err_lower)
err_lower = np.append(err_lower, err_lower[-1])
ax.step(step_x, step_y, where='mid',
marker=None, color=color, linewidth=1,
linestyle='-', label=label, alpha=0.8)
ax.fill_between(step_x, err_upper, err_lower,
alpha=0.15, color=color,
step='mid', linewidth=1)
return step_x, step_y
fig, ax = plt.subplots()
for composition in comp_list + ['total']:
err = np.sqrt(frac_correct_sys_err[composition]**2+reco_frac_sterr[composition]**2)
plot_steps(energy_midpoints, reco_frac[composition], err, ax, color_dict[composition], composition)
plt.xlabel('$\log_{10}(E_{\mathrm{reco}}/\mathrm{GeV})$')
ax.set_ylabel('Fraction correctly identified')
ax.set_ylim([0.0, 1.0])
ax.set_xlim([6.2, 8.0])
ax.grid()
leg = plt.legend(loc='upper center',
bbox_to_anchor=(0.5, # horizontal
1.1),# vertical
ncol=len(comp_list)+1, fancybox=False)
# set the linewidth of each legend object
for legobj in leg.legendHandles:
legobj.set_linewidth(3.0)
# place a text box in upper left in axes coords
textstr = '$\mathrm{\underline{Training \ features}}$: \n'
for i, label in enumerate(feature_labels):
if (i == len(feature_labels)-1):
textstr += '{}) '.format(i+1) + label
else:
textstr += '{}) '.format(i+1) + label + '\n'
props = dict(facecolor='white', linewidth=0)
ax.text(1.025, 0.855, textstr, transform=ax.transAxes, fontsize=8,
verticalalignment='top', bbox=props)
cvstr = '$\mathrm{\underline{CV \ score}}$:\n' + '{:0.2f}\% (+/- {:.2}\%)'.format(scores.mean()*100, scores.std()*100)
print(cvstr)
props = dict(facecolor='white', linewidth=0)
ax.text(1.025, 0.9825, cvstr, transform=ax.transAxes, fontsize=8,
verticalalignment='top', bbox=props)
plt.show()
# +
df, cut_dict = comp.load_sim(return_cut_dict=True)
selection_mask = np.array([True] * len(df))
standard_cut_keys = ['lap_reco_success', 'lap_zenith', 'num_hits_1_30', 'IT_signal',
'max_qfrac_1_30', 'lap_containment', 'energy_range_lap']
for key in standard_cut_keys:
selection_mask *= cut_dict[key]
df = df[selection_mask]
# feature_list, feature_labels = comp.get_training_features()
feature_list = np.array(['lap_log_energy', 'InIce_log_charge_1_30', 'lap_cos_zenith',
'log_NChannels_1_30', 'log_s125', 'StationDensity', 'charge_nchannels_ratio',
'stationdensity_charge_ratio', 'lap_likelihood'])
label_dict = {'reco_log_energy': '$\log_{10}(E_{\mathrm{reco}}/\mathrm{GeV})$',
'lap_log_energy': '$\log_{10}(E_{\mathrm{Lap}}/\mathrm{GeV})$',
'log_s125': '$\log_{10}(S_{\mathrm{125}})$',
'lap_likelihood': '$r\log_{10}(l)$',
'InIce_charge_1_30': 'InIce charge (top 50\%)',
'InIce_log_charge_1_30': '$\log_{10}$(InIce charge (top 50\%))',
'lap_cos_zenith': '$\cos(\\theta_{\mathrm{Lap}})$',
'LLHlap_cos_zenith': '$\cos(\\theta_{\mathrm{Lap}})$',
'LLHLF_cos_zenith': '$\cos(\\theta_{\mathrm{LLH+COG}})$',
'lap_chi2': '$\chi^2_{\mathrm{Lap}}/\mathrm{n.d.f}$',
'NChannels_1_30': 'NChannels (top 50\%)',
'log_NChannels_1_30' : '$\log_{10}$(NChannels (top 50\%))',
'StationDensity': 'StationDensity',
'charge_nchannels_ratio': 'Charge/NChannels',
'stationdensity_charge_ratio': 'StationDensity/Charge',
}
feature_labels = np.array([label_dict[feature] for feature in feature_list])
print('training features = {}'.format(feature_list))
X_train, X_test, y_train, y_test, le = comp.get_train_test_sets(
df, feature_list, train_he=True, test_he=True)
print('number training events = ' + str(y_train.shape[0]))
print('number testing events = ' + str(y_test.shape[0]))
fig, axarr = plt.subplots(2, 2, sharex=True, sharey=True)
max_depth_list = [3, 5, 6, 7]
for max_depth, ax in zip(max_depth_list, axarr.flatten()):
pipeline = comp.get_pipeline('RF')
params = {'classifier__max_depth': max_depth}
pipeline.set_params(**params)
sbs = comp.analysis.SBS(pipeline, k_features=2)
sbs.fit(X_train, y_train)
# plotting performance of feature subsets
k_feat = [len(k) for k in sbs.subsets_]
ax.plot(k_feat, sbs.scores_, marker='.', linestyle=':')
# plt.ylim([0.5, 1.1])
ax.set_xlim([sorted(k_feat)[0]-1, sorted(k_feat)[-1]+1])
ax.set_ylabel('Accuracy')
ax.set_xlabel('Number of features')
ax.set_title('max depth = {}'.format(max_depth))
ax.grid()
plt.tight_layout()
plt.show()
# -
print(sbs.subsets_)
print(len(feature_list))
gen = (feature_list[i] for i in sbs.subsets_)
for i in sbs.subsets_:
print(', '.join(feature_list[np.array(i)]))
print('\n')
feature_list[ np.array((0, 3, 4))]
# ## Feature importance
# +
num_features = len(feature_list)
pipeline = comp.get_pipeline('RF')
pipeline.fit(X_train, y_train)
importances = pipeline.named_steps['classifier'].feature_importances_
indices = np.argsort(importances)[::-1]
fig, ax = plt.subplots()
for f in range(num_features):
print('{}) {}'.format(f + 1, importances[indices[f]]))
plt.ylabel('Feature Importances')
plt.bar(range(num_features),
importances[indices],
align='center')
plt.xticks(range(num_features),
feature_labels[indices], rotation=90)
plt.xlim([-1, len(feature_list)])
# plt.ylim([0, .40])
plt.show()
# -
probs = pipeline.named_steps['classifier'].predict_proba(X_test)
prob_1 = probs[:, 0][MC_iron_mask]
prob_2 = probs[:, 1][MC_iron_mask]
# print(min(prob_1-prob_2))
# print(max(prob_1-prob_2))
# plt.hist(prob_1-prob_2, bins=30, log=True)
plt.hist(prob_1, bins=np.linspace(0, 1, 50), log=True)
plt.hist(prob_2, bins=np.linspace(0, 1, 50), log=True)
probs = pipeline.named_steps['classifier'].predict_proba(X_test)
dp1 = (probs[:, 0]-probs[:, 1])[MC_proton_mask]
print(min(dp1))
print(max(dp1))
dp2 = (probs[:, 0]-probs[:, 1])[MC_iron_mask]
print(min(dp2))
print(max(dp2))
fig, ax = plt.subplots()
# plt.hist(prob_1-prob_2, bins=30, log=True)
counts, edges, pathes = plt.hist(dp1, bins=np.linspace(-1, 1, 100), log=True, label='Proton', alpha=0.75)
counts, edges, pathes = plt.hist(dp2, bins=np.linspace(-1, 1, 100), log=True, label='Iron', alpha=0.75)
plt.legend(loc=2)
plt.show()
pipeline.named_steps['classifier'].classes_
print(pipeline.named_steps['classifier'].classes_)
le.inverse_transform(pipeline.named_steps['classifier'].classes_)
pipeline.named_steps['classifier'].decision_path(X_test)
comp_list = ['P', 'He', 'O', 'Fe']
# test_probs = defaultdict(list)
fig, ax = plt.subplots()
test_probs = pipeline.predict_proba(X_test)
for class_ in pipeline.classes_:
composition = le.inverse_transform(class_)
plt.hist(test_probs[:, class_], bins=np.linspace(0, 1, 50),
histtype='step', label=composition,
color=color_dict[composition], alpha=0.8, log=True)
plt.ylabel('Counts')
plt.xlabel('Testing set class probabilities')
plt.legend()
plt.grid()
plt.show()
# +
pipeline = comp.get_pipeline('RF')
pipeline.fit(X_train, y_train)
test_predictions = pipeline.predict(X_test)
comp_list = ['P', 'He', 'O', 'Fe']
fig, ax = plt.subplots()
test_probs = pipeline.predict_proba(X_test)
fig, axarr = plt.subplots(2, 2, sharex=True, sharey=True)
for composition, ax in zip(comp_list, axarr.flatten()):
comp_mask = (le.inverse_transform(y_test) == composition)
probs = np.copy(test_probs[comp_mask])
print('probs = {}'.format(probs.shape))
weighted_mass = np.zeros(len(probs))
for class_ in pipeline.classes_:
c = le.inverse_transform(class_)
weighted_mass += comp.simfunctions.comp2mass(c) * probs[:, class_]
print('min = {}'.format(min(weighted_mass)))
print('max = {}'.format(max(weighted_mass)))
ax.hist(weighted_mass, bins=np.linspace(0, 5, 100),
histtype='step', label=None, color='darkgray',
alpha=1.0, log=False)
for c in comp_list:
ax.axvline(comp.simfunctions.comp2mass(c), color=color_dict[c],
marker='None', linestyle='-')
ax.set_ylabel('Counts')
ax.set_xlabel('Weighted atomic number')
ax.set_title('MC {}'.format(composition))
ax.grid()
plt.tight_layout()
plt.show()
# +
pipeline = comp.get_pipeline('RF')
pipeline.fit(X_train, y_train)
test_predictions = pipeline.predict(X_test)
comp_list = ['P', 'He', 'O', 'Fe']
fig, ax = plt.subplots()
test_probs = pipeline.predict_proba(X_test)
fig, axarr = plt.subplots(2, 2, sharex=True, sharey=True)
for composition, ax in zip(comp_list, axarr.flatten()):
comp_mask = (le.inverse_transform(y_test) == composition)
probs = np.copy(test_probs[comp_mask])
weighted_mass = np.zeros(len(probs))
for class_ in pipeline.classes_:
c = le.inverse_transform(class_)
ax.hist(probs[:, class_], bins=np.linspace(0, 1, 50),
histtype='step', label=c, color=color_dict[c],
alpha=1.0, log=True)
ax.legend(title='Reco comp', framealpha=0.5)
ax.set_ylabel('Counts')
ax.set_xlabel('Testing set class probabilities')
ax.set_title('MC {}'.format(composition))
ax.grid()
plt.tight_layout()
plt.show()
# -
comp_list = np.unique(df['MC_comp'])
test_probs = defaultdict(list)
fig, ax = plt.subplots()
# test_probs = pipeline.predict_proba(X_test)
for event in pipeline.predict_proba(X_test):
composition = le.inverse_transform(np.argmax(event))
test_probs[composition].append(np.amax(event))
for composition in comp_list:
plt.hist(test_probs[composition], bins=np.linspace(0, 1, 100),
histtype='step', label=composition,
color=color_dict[composition], alpha=0.8, log=False)
plt.ylabel('Counts')
plt.xlabel('Testing set class probabilities')
plt.legend(title='Reco comp')
plt.grid()
plt.show()
|
notebooks/legacy/lightheavy/fraction-correct-RF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 3</font>
#
#
# ## Download: http://github.com/dsacademybr
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# ### Loop For
# Criando uma tupla e imprimindo cada um dos valores
tp = (2,3,4)
for i in tp:
print(i)
# Criando uma lista e imprimindo cada um dos valores
ListaDoMercado = ["Leite", "Frutas", "Carne"]
for i in ListaDoMercado:
print(i)
# Imprimindo os valores no intervalo entre 0 e 5 (exclusive)
for contador in range(0,5):
print(contador)
# Imprimindo na tela os números pares da lista de números
lista = [1,2,3,4,5,6,7,8,9,10]
for num in lista:
if num % 2 == 0:
print(num)
# Listando os números no intervalo entre 0 e 101, com incremento em 2
for i in range(0,101,2):
print(i)
# Listando os números no intervalo entre 0 e 101, com incremento em 2
lista = []
for i in range(0,101,2):
lista.append(i)
print(lista)
# Strings também são sequências
for caracter in 'Python é uma linguagem de programação divertida!':
print (caracter)
# Strings também são sequências
lista = []
for caracter in 'Python é uma linguagem de programação divertida!':
lista.append(caracter)
print (lista)
# ### Loops Aninhados
# Loops aninhados
for i in range(0,5):
for a in range(0,5):
print(a)
# +
# Operando os valores de uma lista com loop for
listaB = [32,53,85,10,15,17,19]
soma = 0
for i in listaB:
double_i = 2 * i
soma += double_i
print(soma)
# -
# Loops em lista de listas
listas = [[1,2,3], [10,15,14], [10.1,8.7,2.3]]
for valor in listas:
print(valor)
# +
# Contando os itens de uma lista
lista = [5,6,10,13,17]
count = 0
for item in lista:
count += 1
print(count)
# +
# Contando o número de colunas
lst = [[1,2,3],[3,4,5],[5,6,7]]
primeira_linha = lst[0]
count = 0
for column in primeira_linha:
count = count + 1
print(count)
# +
# Pesquisando em listas
listaC = [5, 6, 7, 10, 50]
# Loop através da lista
for item in listaC:
if item == 10:
print(f"Número {item} encontrado na lista!")
else:
pass
print(item)
# -
# Listando as chaves de um dicionário
dict = {'k1':'Python','k2':'R','k3':'Scala'}
for item in dict:
print(item)
# Imprimindo chave e valor do dicionário. Usando o método items() para retornar os itens de um dicionário
for k,v in dict.items():
print (k,v)
# Imprimindo valor do dicionário. Usando o método values() para retornar os itens de um dicionário
for v in dict.values():
print (v)
# # Fim
# ### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
|
Cap03/Notebooks/DSA-Python-Cap03-02-For.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lustraka/Data_Analysis_Workouts/blob/main/Wrangle_Data/Try_custom_search.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6352eeb2-104a-41b6-bae5-80b36cbec408"
# - [Programmatically searching google in Python using custom search](https://stackoverflow.com/questions/37083058/programmatically-searching-google-in-python-using-custom-search)
# - [Creating a Programmable Search Engine](https://developers.google.com/custom-search/docs/tutorial/creatingcse)
# - [Programmable Search Engines](https://programmablesearchengine.google.com/controlpanel/all) + [Public Address](https://cse.google.com/cse?cx=bf1b8ef497991822f)
# + id="96a91f5a-8074-4481-b7ab-8d41b1429ba1"
from googleapiclient.discovery import build
import pprint
my_api_key = '' # creds.google_api_key
my_cse_id = "bf1b8ef497991822f"
def google_search(search_term, api_key, cse_id, **kwargs):
service = build("customsearch", "v1", developerKey=api_key) # type: googleapiclient.discovery.Resource
res = service.cse().list(q=search_term, cx=cse_id, **kwargs).execute()
return res
# + id="MJ9c5s9_g6eZ"
results = google_search('data analytics portfolio projects', my_api_key, my_cse_id, num=10) # num has to be in <1, 10>
# for result in results:
# pprint.pprint(result)
# + [markdown] id="7UepivmQ8RSJ"
# ```python
# [In] results.keys()
# [Out] dict_keys(['kind', 'url', 'queries', 'context', 'searchInformation', 'items'])
# ```
#
# ```python
# [In] results.items()
# [Out] dict_items([('kind', 'customsearch#search'), ('url', {'type': 'application/json', 'template': ...
# ```
#
# ```python
# [In] results.values()
# [Out] dict_values(['customsearch#search', {'type': 'application/json', 'template':
# ```
# Key | Content
# -|-
# kind | customsearch#search
# url | dict {'template': 'https://www.googleapis.com/customsearch/v1?q={searchTerms}&num={count?}&'... , 'type': 'application/json'} for a search
# queries | dict
# searchInformation | dict
# items | list of dicts
#
# **Dict 'queries'**
# ```python
# {'nextPage': [{ ### same as request except:
# 'startIndex': 11,
# }],
# 'request': [{'count': 10,
# 'cx': 'bf1b8ef497991822f',
# 'inputEncoding': 'utf8',
# 'outputEncoding': 'utf8',
# 'safe': 'off',
# 'searchTerms': 'data analytics portfolio projects',
# 'startIndex': 1,
# 'title': 'Google Custom Search - data analytics portfolio projects',
# 'totalResults': '196000000'}]}
# ```
#
# **Dict 'searchInformation'**
# ```python
# {'formattedSearchTime': '0.46',
# 'formattedTotalResults': '196,000,000',
# 'searchTime': 0.464201,
# 'totalResults': '196000000'}
# ```
#
# **Keys of item dict**
# ```
# dict_keys(['kind', 'title', 'htmlTitle', 'link', 'displayLink', 'snippet',
# 'htmlSnippet', 'cacheId', 'formattedUrl', 'htmlFormattedUrl', 'pagemap'])
# ```
# + colab={"base_uri": "https://localhost:8080/"} id="FjAsoP1E753R" outputId="366a3ecb-d660-4769-9d3b-2c1acba27208"
# Sample item
results['items'][0]
# + colab={"base_uri": "https://localhost:8080/"} id="JRAbFFl38Qug" outputId="ab5e9dd6-9753-4e43-85d6-556663820b2b"
for item in results['items']:
print(item['title'])
print('\t\t', item['link'])
# + id="XhRnDAcJ75wp"
|
Wrangle_Data/Try_custom_search.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
import torch.nn as nn
import torch
import pickle
import pandas as pd
from pandas import Series, DataFrame
from pandarallel import pandarallel
pandarallel.initialize(progress_bar=False)
from sklearn.metrics import roc_auc_score, roc_curve, accuracy_score, matthews_corrcoef, f1_score, precision_score, recall_score
import numpy as np
import torch.optim as optim
folder = "/data/AIpep-clean/"
import matplotlib.pyplot as plt
from vocabulary import Vocabulary
from datasethem import Dataset, collate_fn
from models import Classifier
import os
random.seed(0)
# # Load data
# +
df = pd.read_pickle(folder + "pickles/DAASP_RNN_dataset_with_hemolysis.plk")
df = df.query("isNotHemolytic==1 or isNotHemolytic==0").copy()
df_training = df[df["Set"]=="training"]
df_test = df[df["Set"]=="test"]
df_training_fool = df_training.copy()
isNotHemolytic = df_training_fool.isNotHemolytic.tolist()
random.shuffle(isNotHemolytic)
df_training_fool["isNotHemolytic"] = isNotHemolytic
df_test_fool = df_test.copy()
isNotHemolytic = df_test_fool.isNotHemolytic.tolist()
random.shuffle(isNotHemolytic)
df_test_fool["isNotHemolytic"] = isNotHemolytic
df_training = df_training_fool
df_test = df_test_fool
vocabulary = Vocabulary.get_vocabulary_from_sequences(df_training.Sequence.values)
if torch.cuda.is_available():
device = "cuda"
else:
device = "cpu"
# -
# # Define helper functions
# +
def randomChoice(l):
return l[random.randint(0, len(l) - 1)]
def categoryFromOutput(output):
top_n, top_i = output.topk(1)
category_i = top_i[0].item()
return category_i
def nan_equal(a,b):
try:
np.testing.assert_equal(a,b)
except AssertionError:
return False
return True
def models_are_equal(model1, model2):
model1.vocabulary == model2.vocabulary
model1.hidden_size == model2.hidden_size
for a,b in zip(model1.model.parameters(), model2.model.parameters()):
if nan_equal(a.detach().numpy(), b.detach().numpy()) == True:
print("true")
# -
# # Define model
def training(model, test_dataloader, training_dataloader, n_epoch, optimizer, filename):
roc_training = []
roc_test = []
for e in range(1, n_epoch + 1):
for i_batch, sample_batched in enumerate(training_dataloader):
seq_batched = sample_batched[0][0].to(model.device, non_blocking=True)
seq_lengths = sample_batched[0][1].to(model.device, non_blocking=True)
cat_batched = sample_batched[1].to(model.device, non_blocking=True)
output = model.evaluate(seq_batched, seq_lengths)
loss = criterion(output, cat_batched)
optimizer.zero_grad()
loss.backward()
torch.nn.utils.clip_grad_value_(model.model.parameters(), 2)
optimizer.step()
model.save(filename.format(e))
def _evaluate_ROC(data_loader):
cat_list = []
out_list = []
with torch.no_grad():
for i_batch, sample_batched in enumerate(data_loader):
seq_batched = sample_batched[0][0].to(model.device, non_blocking=True)
seq_lengths = sample_batched[0][1].to(model.device, non_blocking=True)
cat_list += sample_batched[1].to("cpu", non_blocking=True)
out_list += torch.exp(model.evaluate(seq_batched, seq_lengths))[: ,1].to("cpu", non_blocking=True)
cat_list = torch.stack(cat_list)
out_list = torch.stack(out_list)
roc = roc_auc_score(cat_list.cpu().numpy().astype(int), out_list.cpu().numpy())
return roc
roc_tr = _evaluate_ROC(training_dataloader)
roc_te = _evaluate_ROC(test_dataloader)
roc_training.append(roc_tr)
roc_test.append(roc_te)
print("epoch: " + str(e))
print("roc auc training: " + str(roc_tr))
print("roc auc test: " + str(roc_te))
if roc_training == 1.0:
break
return model, optimizer, roc_training, roc_test
learning_rate = 0.01
momentum = 0.9
batch_size = 20
n_epoch = 150
criterion = nn.NLLLoss()
# # Hyper parameters optimization
# + jupyter={"outputs_hidden": true}
n_embeddings = [2, 21, 42, 100]
n_hiddens = [50, 100, 200, 300, 400]
n_layerss = [1, 2,3]
if not os.path.exists(folder+"pickles/fool_classifier_hyperparameter_optimization_results_hem_.pkl"):
df_opt = df_training.copy()
# create an evaluation/training set only from the training set
# assign to training or evaluation set
df_opt["Set2"] = "test"
training_ = df_opt.sample(frac=0.75, random_state=0)
df_opt.loc[training_.index, "Set2"] = "training"
df_training = df_opt[df_opt["Set2"]=="training"]
df_test = df_opt[df_opt["Set2"]=="test"]
training_dataset = Dataset(df_training, vocabulary)
test_dataset = Dataset(df_test, vocabulary)
training_dict = {}
for n_embedding in n_embeddings:
for n_hidden in n_hiddens:
for n_layers in n_layerss:
if "em{}_hi{}_la{}".format(n_embedding, n_hidden, n_layers) in training_dict:
continue
print(f"dimensions of embedding {n_embedding}, dimensions of hidden {n_hidden}, number of layers {n_layers}")
model = Classifier(n_embedding, n_hidden, n_layers, vocabulary)
model.to(device)
optimizer = optim.SGD(model.model.parameters(), lr = learning_rate, momentum=momentum)
training_dataloader = torch.utils.data.DataLoader(training_dataset, batch_size=batch_size, shuffle=True, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
test_dataloader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
filename = folder+"models/RNN-classifier-fool-hem/em{}_hi{}_la{}_ep{{}}".format(n_embedding, n_hidden, n_layers)
model, optimizer, roc_training, roc_test = training(model, test_dataloader, training_dataloader, n_epoch, optimizer, filename)
training_dict["em{}_hi{}_la{}".format(n_embedding, n_hidden, n_layers)] = [roc_training, roc_test]
print(f"maximum roc auc for test set {max(roc_test)}")
with open(folder+"pickles/fool_classifier_hyperparameter_optimization_results_hem.pkl","bw") as fd:
pickle.dump(training_dict, fd)
else:
with open(folder+"pickles/fool_classifier_hyperparameter_optimization_results_hem.pkl",'rb') as fd:
training_dict = pickle.load(fd)
# -
# # Optimized hyper parameters
max_test = 0
for k,v in training_dict.items():
if max(v[1]) > max_test:
max_test = max(v[1])
best = k
best = best.split("_")
n_embedding = int(best[0].replace("em", ""))
n_hidden = int(best[1].replace("hi", ""))
n_layers = int(best[2].replace("la", ""))
print(n_embedding, n_hidden, n_layers)
# # Training
# + jupyter={"outputs_hidden": true}
training_dataset = Dataset(df_training, vocabulary)
test_dataset = Dataset(df_test, vocabulary)
print(f"dimensions of embedding {n_embedding}, dimensions of hidden {n_hidden}, number of layers {n_layers}")
model = Classifier(n_embedding, n_hidden, n_layers, vocabulary)
model.to(device)
optimizer = optim.SGD(model.model.parameters(), lr = learning_rate, momentum=momentum)
training_dataloader = torch.utils.data.DataLoader(training_dataset, batch_size=batch_size, shuffle=True, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
test_dataloader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=True, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
filename = folder + "/models/RNN-classifier-fool/hem_em{}_hi{}_la{}_ep{{}}".format(n_embedding, n_hidden, n_layers)
model, optimizer, roc_training, roc_test = training(model, test_dataloader, training_dataloader, n_epoch, optimizer, filename)
print(f"maximum roc auc for test set {max(roc_test)}")
# +
roc_test = np.array(roc_test)
epoch = np.argmax(roc_test) + 1
print(epoch)
training_dataset = Dataset(df_training, vocabulary)
test_dataset = Dataset(df_test, vocabulary)
filename = folder + "models/RNN-classifier-fool/hem_em{}_hi{}_la{}_ep{}".format(n_embedding, n_hidden, n_layers, epoch)
model = Classifier.load_from_file(filename)
model.to(device)
training_dataloader_eval = torch.utils.data.DataLoader(training_dataset, batch_size=1, shuffle=False, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
test_dataloader_eval = torch.utils.data.DataLoader(test_dataset, batch_size=1, shuffle=False, collate_fn = collate_fn, drop_last=True, pin_memory=True, num_workers=4)
# -
# # Evaluation
# +
def predict(data_loader):
cat_list = []
out_list = []
with torch.no_grad():
for i_batch, sample_batched in enumerate(data_loader):
seq_batched = sample_batched[0][0].to(model.device, non_blocking=True)
seq_lengths = sample_batched[0][1].to(model.device, non_blocking=True)
cat_list += sample_batched[1].to("cpu", non_blocking=True)
out_list += torch.exp(model.evaluate(seq_batched, seq_lengths))[: ,1].to("cpu", non_blocking=True)
cat_list = torch.stack(cat_list)
out_list = torch.stack(out_list)
return cat_list.cpu().numpy().astype(int), out_list.cpu().numpy()
def roc(y_true, y_score):
fpr, tpr, thresh = roc_curve(y_true, y_score)
roc = roc_auc_score(y_true, y_score)
return roc, fpr, tpr
def find_threshold(y_true, y_score, alpha = 0.049):
fpr, tpr, thresh = roc_curve(y_true, y_score)
for i, fp in enumerate(fpr):
if fp > alpha:
return thresh[i-1]
def calc_metrics(y_true, y_score, threshold = 0.5):
y_score = y_score > threshold
accuracy = accuracy_score(y_true, y_score)
f1 = f1_score(y_true, y_score)
mcc = matthews_corrcoef(y_true, y_score)
precision = precision_score(y_true, y_score)
recall = recall_score(y_true, y_score)
return accuracy, f1, mcc, precision, recall
# -
y_true, y_score = predict(test_dataloader_eval)
threshold = find_threshold(y_true, y_score)
threshold
accuracy, f1, mcc, precision, recall = calc_metrics(y_true, y_score, threshold)
print(f"accuracy: {accuracy}\nf1 score: {f1}\nmcc: {mcc}\nprecision: {precision}\nrecall: {recall}" )
# accuracy: 0.6078886310904872
# f1 score: 0.09625668449197859
# mcc: 0.012529783364402296
# precision: 0.4090909090909091
# recall: 0.05454545454545454
accuracy, f1, mcc, precision, recall = calc_metrics(y_true, y_score, 0.5)
print(f"accuracy: {accuracy}\nf1 score: {f1}\nmcc: {mcc}\nprecision: {precision}\nrecall: {recall}" )
# accuracy: 0.6078886310904872
# f1 score: 0.09625668449197859
# mcc: 0.012529783364402296
# precision: 0.4090909090909091
# recall: 0.05454545454545454
# +
Y_true, Y_score = predict(training_dataloader_eval)
roc_training, fpr_training, tpr_training = roc(Y_true, Y_score)
y_true, y_score = predict(test_dataloader_eval)
roc_test, fpr_test, tpr_test = roc(y_true, y_score)
with open("/data/AIpep/auc_files/fool_classifier_hem.pkl","bw") as fd:
pickle.dump((fpr_test, tpr_test), fd)
plt.figure()
name = "RNN fool-classifier"
plt.rcParams.update({'font.size': 15})
lw = 2
plt.plot(fpr_training, tpr_training, color='blue',
lw=lw, label='Training Set ROC AUC = %0.2f' % roc_training)
plt.plot(fpr_test, tpr_test, color='darkorange',
lw=lw, label='Test Set ROC AUC = %0.2f' % roc_test)
plt.plot([0, 1], [0, 1], color='navy', lw=lw, linestyle='--')
plt.xlim([-0.005, 1.0])
plt.ylim([-0.005, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title(name)
plt.legend(loc="lower right")
plt.tight_layout()
plt.savefig(folder+"plots/RNN-fool-classifier.svg")
plt.savefig("plots/RNN-fool-classifier.svg")
plt.show()
# -
|
04b-RNN-GRU-hemolysis-fool-classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''venv37'': virtualenv)'
# language: python
# name: python37564bitvenv37virtualenv5772f1f1f81449bd9fbfbe2ddfacfeee
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="D_PfLJqAgMIq" outputId="1fd2de6f-fbc5-40c9-ac1f-363ddcde0007"
import tsflex
print(tsflex.__version__)
# + [markdown] id="KXqEDPikmHc6"
# ## Get the data
# + id="bB1tBMOLggLv"
import pandas as pd
url = "https://github.com/predict-idlab/tsflex/raw/main/examples/data/empatica/"
df_tmp = pd.read_parquet(url+"tmp.parquet").set_index("timestamp")
df_acc = pd.read_parquet(url+"acc.parquet").set_index("timestamp")
df_gsr = pd.read_parquet(url+"gsr.parquet").set_index("timestamp")
df_ibi = pd.read_parquet(url+"ibi.parquet").set_index("timestamp")
# + colab={"base_uri": "https://localhost:8080/"} id="Qo6btlF8kn8v" outputId="2d572d8a-b2cb-4e24-ff22-679d85a25a90"
from pandas.tseries.frequencies import to_offset
data = [df_tmp, df_acc, df_gsr, df_ibi]
for df in data:
print("Time-series:", df.columns.values)
print(df.shape)
try:
print("Sampling rate:", 1 / pd.to_timedelta(to_offset(pd.infer_freq(df.index))).total_seconds(), "Hz")
except:
print("Irregular sampling rate")
print()
# + [markdown] id="G3JN03iomGui"
# ## Look at the data
# + colab={"base_uri": "https://localhost:8080/", "height": 817} id="HYLMtx7tjTtR" outputId="5619b18f-5ef9-49df-9f4b-53a0de934931"
import plotly.graph_objects as go
from plotly.subplots import make_subplots
fig = make_subplots(
rows=len(data), cols=1, shared_xaxes=True,
subplot_titles=[df.columns.values[0].split('_')[0] for df in data]
)
for plot_idx, df in enumerate(data, 1):
# Select first minute of data
sub_df = df.first('1min')
for col in df.columns:
fig.add_trace(
go.Scattergl(x=sub_df.index, y=sub_df[col].values, name=col, mode='markers'),
row=plot_idx, col=1
)
fig.update_layout(height=len(data)*200)
fig.show(renderer='iframe')
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="aJeTIxeupRu5" outputId="9c0b7085-623a-4d00-e73d-7f78006d753c"
import matplotlib.pyplot as plt
fig, axes = plt.subplots(nrows=1, ncols=4, figsize=(16,4))
for plot_idx, df in enumerate(data):
df.plot(kind='box', ax=axes[plot_idx])
plt.tight_layout()
# + [markdown] id="AKLw2bJxpCiE"
# These visualizations indicate that some preprocessing might be necessary for the signals (some sort of clipping)
# + [markdown] id="qlswfOOyr4zT"
# # tsflex processing
# -
# This is roughly identical to the processing of notebook containing the example code of the paper.
# + colab={"base_uri": "https://localhost:8080/"} id="XsAKFB3bkjQ8" outputId="c87cfc5b-fabe-47bf-d9d2-7bed9e731f8b"
from tsflex.processing import SeriesProcessor, SeriesPipeline
# Import / create the processing functions
import numpy as np
from scipy.signal import savgol_filter
def clip_quantiles(sig: pd.Series, lower_q=0.01, upper_q=0.99) -> np.ndarray:
# Note that this function induces a data leakage
quantile_vals = np.quantile(sig, q=[lower_q, upper_q])
return np.clip(sig, *quantile_vals)
def smv(*sigs) -> pd.Series:
sig_prefixes = set(sig.name.split('_')[0] for sig in sigs)
result = np.sqrt(np.sum([np.square(sig) for sig in sigs], axis=0))
return pd.Series(result, index=sigs[0].index, name='|'.join(sig_prefixes)+'_'+'SMV')
# Create the series processors (with their keyword arguments)
clipper_tmp = SeriesProcessor(clip_quantiles, series_names="TMP", lower_q=0, upper_q=0.999)
savgol_eda = SeriesProcessor(savgol_filter, "EDA", window_length=5, polyorder=2)
savgol_acc = SeriesProcessor(savgol_filter, ["ACC_x", "ACC_y", "ACC_z"], window_length=33, polyorder=2)
smv_processor = SeriesProcessor(smv, ("ACC_x", "ACC_y", "ACC_z"))
# Create the series pipeline
series_pipe = SeriesPipeline(
processors=[clipper_tmp, savgol_eda, savgol_acc, smv_processor]
)
series_pipe
# + id="TK64KF0h0HuT"
out_data = series_pipe.process(data, drop_keys=["ACC_x", "ACC_y", "ACC_z"])
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="jXUDDMbWxqkv" outputId="94c79711-9202-4296-c248-1b515cff1e4f"
import matplotlib.pyplot as plt
fig, axes = plt.subplots(nrows=1, ncols=4, figsize=(16,4))
for plot_idx, df in enumerate(out_data):
df.plot(kind='box', ax=axes[plot_idx])
plt.tight_layout()
# + [markdown] id="Fy0gYc961AAz"
# # tsflex feature extraction with [tsfresh](https://github.com/blue-yonder/tsfresh) integration
# + tags=[]
# !pip install tsfresh
# -
# > Useful links;
# > [List of all tsfresh features](https://tsfresh.readthedocs.io/en/latest/text/list_of_features.html)
# > [More detailed documentation of the tsfresh features](https://tsfresh.readthedocs.io/en/latest/api/tsfresh.feature_extraction.html#module-tsfresh.feature_extraction.feature_calculators)
# > [More detailed documentation of the tsfresh feature extraction settings](https://tsfresh.readthedocs.io/en/latest/text/feature_extraction_settings.html)
#
# [tsfresh feature extraction settings](https://github.com/blue-yonder/tsfresh/blob/main/tsfresh/feature_extraction/settings.py) is how tsfresh represents a collection of features (with their parameters).
# **=> requires wrapping this settings object in a `tsfresh_settings_wrapper` for interoperability with tsflex**.
#
# [tsfresh feature-funtions](https://github.com/blue-yonder/tsfresh/blob/main/tsfresh/feature_extraction/feature_calculators.py) are either of type `simple` or `combiner`.
# * `simple`: feature calculators which calculate a single number
# **=> integrates natively with tsflex**
# * `combiner`: feature calculates which calculate a bunch of features for a list of parameters. These features are returned as a list of (key, value) pairs for each input parameter.
# **=> requires wrapping the function to only extract the values of the returned tuples**
#
# Of course, feature functions that require other keyword arguments, should be wrapped in a `FuncWrapper`
# This wrapper handles tsfresh its feature extraction settings
from tsflex.features.integrations import tsfresh_settings_wrapper
# This wrappers handles tsfresh its combiner functions
from tsflex.features.integrations import tsfresh_combiner_wrapper
# + tags=[]
from tsflex.features import FeatureCollection, MultipleFeatureDescriptors
# -
# ## Using tsfresh feature extraction settings
# + tags=[]
# Import some preset feature extraction setting from tsfresh
from tsfresh.feature_extraction import MinimalFCParameters, EfficientFCParameters
# -
# Calculate the features for a tsfresh feature extraction setting.
# Note that;
# * `tsfresh_settings_wrapper` transforms this feature extraction settings object to a list of features that you can directly pass as the `function` argument of tsflex `MultipleFeatureDescriptors`.
# + tags=[]
simple_feats = MultipleFeatureDescriptors(
functions=tsfresh_settings_wrapper(MinimalFCParameters()),
series_names=["ACC_SMV", "EDA", "TMP"],
windows=["5min", "2.5min"],
strides=["2.5min"],
)
feature_collection = FeatureCollection(simple_feats)
feature_collection
# + tags=[]
features_df = feature_collection.calculate(out_data, return_df=True, show_progress=True)
features_df
# -
# Extract a lot more tsfresh features (& customize the settings, i.e., remove the slower functions)
# +
slow_funcs = [
"matrix_profile",
"number_cwt_peaks",
"augmented_dickey_fuller",
"partial_autocorrelation",
"agg_linear_trend",
"lempel_ziv_complexity",
"benford_correlation",
"ar_coefficient",
"permutation_entropy",
"friedrich_coefficients",
]
settings = EfficientFCParameters()
for f in slow_funcs:
del settings[f]
# + tags=[]
efficient_feats = MultipleFeatureDescriptors(
functions=tsfresh_settings_wrapper(settings),
series_names=["ACC_SMV", "EDA", "TMP"],
windows=["5min", "2.5min"],
strides=["2.5min"],
)
feature_collection = FeatureCollection(efficient_feats)
feature_collection
# + tags=[]
features_df = feature_collection.calculate(out_data, return_df=True, show_progress=True)
features_df
# + [markdown] id="c36Hw96oDPkV"
# ### Plot the EDA features
# + colab={"base_uri": "https://localhost:8080/", "height": 717} id="NxJKV1u0DVvg" outputId="853b0f4c-62b6-4978-e826-693cd411b9b8"
import plotly.graph_objects as go
from plotly.subplots import make_subplots
fig = make_subplots(
rows=2, cols=1, shared_xaxes=True,
subplot_titles=['Raw EDA data', 'EDA features']
)
fig.add_trace(
go.Scattergl(x=df_gsr.index[::4*5], y=df_gsr['EDA'].values[::4*5], name='EDA', mode='markers'),
row=1, col=1
)
ibi_feats = [c for c in features_df.columns if 'EDA_' in c and 'w=2m30s_' in c]
for col in ibi_feats:
sub_df = features_df[[col]].dropna()
if not np.issubdtype(sub_df.values.dtype, np.number):
continue
fig.add_trace(
go.Scattergl(x=sub_df.index, y=sub_df[col].values, name=col, mode='markers'),
row=2, col=1
)
fig.update_layout(height=2*350)
fig.show(renderer='iframe')
# -
# ## Using simple tsfresh features
# Integrates natively :)
# + colab={"base_uri": "https://localhost:8080/"} id="zwnMitvayEhd" outputId="c6ea7f18-e007-4bb9-bdc4-386475a7c3d0"
# Import some simple funtions
from tsfresh.feature_extraction.feature_calculators import (
abs_energy,
absolute_sum_of_changes,
cid_ce,
variance_larger_than_standard_deviation,
)
from tsflex.features import FeatureCollection, FuncWrapper, MultipleFeatureDescriptors
simple_feats = MultipleFeatureDescriptors(
functions=[
abs_energy,
absolute_sum_of_changes,
variance_larger_than_standard_deviation,
FuncWrapper(cid_ce, normalize=True),
],
series_names=["ACC_SMV", "EDA", "TMP"],
windows=["5min", "2.5min"],
strides="2min",
)
feature_collection = FeatureCollection(simple_feats)
feature_collection
# + colab={"base_uri": "https://localhost:8080/", "height": 640} id="ahXC5VxR2w0W" outputId="f7f8b9e0-937f-4986-80f4-eb6eb36c3093" tags=[]
features_df = feature_collection.calculate(out_data, return_df=True)
features_df
# + [markdown] id="c36Hw96oDPkV"
# ### Plot the EDA features
# + colab={"base_uri": "https://localhost:8080/", "height": 717} id="NxJKV1u0DVvg" outputId="853b0f4c-62b6-4978-e826-693cd411b9b8"
import plotly.graph_objects as go
from plotly.subplots import make_subplots
fig = make_subplots(
rows=2, cols=1, shared_xaxes=True,
subplot_titles=['Raw EDA data', 'EDA features']
)
fig.add_trace(
go.Scattergl(x=df_gsr.index[::4*5], y=df_gsr['EDA'].values[::4*5], name='EDA', mode='markers'),
row=1, col=1
)
ibi_feats = [c for c in features_df.columns if 'EDA_' in c and 'w=2m30s_' in c]
for col in ibi_feats:
sub_df = features_df[[col]].dropna()
fig.add_trace(
go.Scattergl(x=sub_df.index, y=sub_df[col].values, name=col, mode='markers'),
row=2, col=1
)
fig.update_layout(height=2*350)
fig.show(renderer='iframe')
# -
# ## Using combiner tsfresh features
# + tags=[]
# Import all combiner funcs
from tsfresh.feature_extraction.feature_calculators import (
agg_autocorrelation,
augmented_dickey_fuller,
cwt_coefficients,
fft_aggregated,
fft_coefficient,
index_mass_quantile,
linear_trend,
partial_autocorrelation,
spkt_welch_density,
symmetry_looking,
ar_coefficient,
friedrich_coefficients,
agg_linear_trend,
energy_ratio_by_chunks,
linear_trend_timewise,
matrix_profile,
query_similarity_count,
)
# -
# Calculate the features for some of tsfresh its combiner functions.
# Note that;
# * `param` is now passed to `tsfresh_combiner_wrapper` instead of the combiner function itself
# * combiner functions that require a `pd.Series` (with a `pd.DatetimeIndex`) are also handled by this wrapper
# + colab={"base_uri": "https://localhost:8080/"} id="zwnMitvayEhd" outputId="c6ea7f18-e007-4bb9-bdc4-386475a7c3d0"
from tsflex.features import FeatureCollection, MultipleFeatureDescriptors
combiner_feats = MultipleFeatureDescriptors(
functions=[
tsfresh_combiner_wrapper(index_mass_quantile, param=[{"q": v} for v in [0.15, 0.5, 0.75]]),
tsfresh_combiner_wrapper(linear_trend, param=[{"attr": v} for v in ["intercept", "slope", "stderr"]]),
tsfresh_combiner_wrapper(spkt_welch_density, param=[{"coeff": v} for v in range(5)]),
# This function requires a pd.Series with a pd.DatetimeIndex
tsfresh_combiner_wrapper(linear_trend_timewise, param=[{"attr": v} for v in ["intercept", "slope"]]),
],
series_names=["ACC_SMV", "EDA", "TMP"],
windows=["5min", "2.5min"],
strides=["2.5min"],
)
feature_collection = FeatureCollection(combiner_feats)
feature_collection
# + colab={"base_uri": "https://localhost:8080/", "height": 640} id="ahXC5VxR2w0W" outputId="f7f8b9e0-937f-4986-80f4-eb6eb36c3093"
features_df = feature_collection.calculate(out_data, return_df=True)
features_df
# + [markdown] id="c36Hw96oDPkV"
# ### Plot the EDA features
# + colab={"base_uri": "https://localhost:8080/", "height": 717} id="NxJKV1u0DVvg" outputId="853b0f4c-62b6-4978-e826-693cd411b9b8"
import plotly.graph_objects as go
from plotly.subplots import make_subplots
fig = make_subplots(
rows=2, cols=1, shared_xaxes=True,
subplot_titles=['Raw EDA data', 'EDA features']
)
fig.add_trace(
go.Scattergl(x=df_gsr.index[::4*5], y=df_gsr['EDA'].values[::4*5], name='EDA', mode='markers'),
row=1, col=1
)
ibi_feats = [c for c in features_df.columns if 'EDA_' in c and 'w=2m30s_' in c]
for col in ibi_feats:
sub_df = features_df[[col]].dropna()
fig.add_trace(
go.Scattergl(x=sub_df.index, y=sub_df[col].values, name=col, mode='markers'),
row=2, col=1
)
fig.update_layout(height=2*350)
fig.show(renderer='iframe')
|
examples/tsfresh_integration.ipynb
|
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # chemical_balance_lp
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/examples/chemical_balance_lp.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/python/chemical_balance_lp.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010-2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# We are trying to group items in equal sized groups.
# Each item has a color and a value. We want the sum of values of each group to
# be as close to the average as possible.
# Furthermore, if one color is an a group, at least k items with this color must
# be in that group.
from ortools.linear_solver import pywraplp
import math
# Data
max_quantities = [["N_Total", 1944], ["P2O5", 1166.4], ["K2O", 1822.5],
["CaO", 1458], ["MgO", 486], ["Fe", 9.7], ["B", 2.4]]
chemical_set = [["A", 0, 0, 510, 540, 0, 0, 0], ["B", 110, 0, 0, 0, 160, 0, 0],
["C", 61, 149, 384, 0, 30, 1,
0.2], ["D", 148, 70, 245, 0, 15, 1,
0.2], ["E", 160, 158, 161, 0, 10, 1, 0.2]]
num_products = len(max_quantities)
all_products = range(num_products)
num_sets = len(chemical_set)
all_sets = range(num_sets)
# Model
max_set = [
min(max_quantities[q][1] / chemical_set[s][q + 1] for q in all_products
if chemical_set[s][q + 1] != 0.0) for s in all_sets
]
solver = pywraplp.Solver("chemical_set_lp",
pywraplp.Solver.GLOP_LINEAR_PROGRAMMING)
set_vars = [solver.NumVar(0, max_set[s], "set_%i" % s) for s in all_sets]
epsilon = solver.NumVar(0, 1000, "epsilon")
for p in all_products:
solver.Add(
sum(chemical_set[s][p + 1] * set_vars[s]
for s in all_sets) <= max_quantities[p][1])
solver.Add(
sum(chemical_set[s][p + 1] * set_vars[s]
for s in all_sets) >= max_quantities[p][1] - epsilon)
solver.Minimize(epsilon)
print(("Number of variables = %d" % solver.NumVariables()))
print(("Number of constraints = %d" % solver.NumConstraints()))
result_status = solver.Solve()
# The problem has an optimal solution.
assert result_status == pywraplp.Solver.OPTIMAL
assert solver.VerifySolution(1e-7, True)
print(("Problem solved in %f milliseconds" % solver.wall_time()))
# The objective value of the solution.
print(("Optimal objective value = %f" % solver.Objective().Value()))
for s in all_sets:
print(
" %s = %f" % (chemical_set[s][0], set_vars[s].solution_value()),
end=" ")
print()
for p in all_products:
name = max_quantities[p][0]
max_quantity = max_quantities[p][1]
quantity = sum(
set_vars[s].solution_value() * chemical_set[s][p + 1] for s in all_sets)
print("%s: %f out of %f" % (name, quantity, max_quantity))
|
examples/notebook/examples/chemical_balance_lp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W1D4_GeneralizedLinearModels/student/W1D4_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a> <a href="https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D4_GeneralizedLinearModels/student/W1D4_Tutorial1.ipynb" target="_parent"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" alt="Open in Kaggle"/></a>
# -
# # Tutorial 1: GLMs for Encoding
# **Week 1, Day 4: Generalized Linear Models**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>
#
#
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# # Tutorial Objectives
#
# *Estimated timing of tutorial: 1 hour, 15 minutes*
#
# This is part 1 of a 2-part series about Generalized Linear Models (GLMs), which are a fundamental framework for supervised learning.
#
# In this tutorial, the objective is to model a retinal ganglion cell spike train by fitting a temporal receptive field. First with a Linear-Gaussian GLM (also known as ordinary least-squares regression model) and then with a Poisson GLM (aka "Linear-Nonlinear-Poisson" model). In the next tutorial, we’ll extend to a special case of GLMs, logistic regression, and learn how to ensure good model performance.
#
# This tutorial is designed to run with retinal ganglion cell spike train data from [Uzzell & Chichilnisky 2004](https://journals.physiology.org/doi/full/10.1152/jn.01171.2003?url_ver=Z39.88-2003&rfr_id=ori:rid:crossref.org&rfr_dat=cr_pub%20%200pubmed).
#
# *Acknowledgements:*
#
# - We thank <NAME> for providing the dataset. Please note that it is provided for tutorial purposes only, and should not be distributed or used for publication without express permission from the author (<EMAIL>).
# - We thank <NAME>, much of this tutorial is inspired by exercises asigned in his 'Statistical Modeling and Analysis of Neural Data' class.
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in all tutorials today
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/upyjz/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# # Setup
#
# + cellView="both"
# Imports
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import minimize
from scipy.io import loadmat
# + cellView="form"
#@title Figure settings
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
# + cellView="form"
# @title Plotting Functions
def plot_stim_and_spikes(stim, spikes, dt, nt=120):
"""Show time series of stim intensity and spike counts.
Args:
stim (1D array): vector of stimulus intensities
spikes (1D array): vector of spike counts
dt (number): duration of each time step
nt (number): number of time steps to plot
"""
timepoints = np.arange(nt)
time = timepoints * dt
f, (ax_stim, ax_spikes) = plt.subplots(
nrows=2, sharex=True, figsize=(8, 5),
)
ax_stim.plot(time, stim[timepoints])
ax_stim.set_ylabel('Stimulus intensity')
ax_spikes.plot(time, spikes[timepoints])
ax_spikes.set_xlabel('Time (s)')
ax_spikes.set_ylabel('Number of spikes')
f.tight_layout()
def plot_glm_matrices(X, y, nt=50):
"""Show X and Y as heatmaps.
Args:
X (2D array): Design matrix.
y (1D or 2D array): Target vector.
"""
from matplotlib.colors import BoundaryNorm
from mpl_toolkits.axes_grid1 import make_axes_locatable
Y = np.c_[y] # Ensure Y is 2D and skinny
f, (ax_x, ax_y) = plt.subplots(
ncols=2,
figsize=(6, 8),
sharey=True,
gridspec_kw=dict(width_ratios=(5, 1)),
)
norm = BoundaryNorm([-1, -.2, .2, 1], 256)
imx = ax_x.pcolormesh(X[:nt], cmap="coolwarm", norm=norm)
ax_x.set(
title="X\n(lagged stimulus)",
xlabel="Time lag (time bins)",
xticks=[4, 14, 24],
xticklabels=['-20', '-10', '0'],
ylabel="Time point (time bins)",
)
plt.setp(ax_x.spines.values(), visible=True)
divx = make_axes_locatable(ax_x)
caxx = divx.append_axes("right", size="5%", pad=0.1)
cbarx = f.colorbar(imx, cax=caxx)
cbarx.set_ticks([-.6, 0, .6])
cbarx.set_ticklabels(np.sort(np.unique(X)))
norm = BoundaryNorm(np.arange(y.max() + 1), 256)
imy = ax_y.pcolormesh(Y[:nt], cmap="magma", norm=norm)
ax_y.set(
title="Y\n(spike count)",
xticks=[]
)
ax_y.invert_yaxis()
plt.setp(ax_y.spines.values(), visible=True)
divy = make_axes_locatable(ax_y)
caxy = divy.append_axes("right", size="30%", pad=0.1)
cbary = f.colorbar(imy, cax=caxy)
cbary.set_ticks(np.arange(y.max()) + .5)
cbary.set_ticklabels(np.arange(y.max()))
def plot_spike_filter(theta, dt, **kws):
"""Plot estimated weights based on time lag model.
Args:
theta (1D array): Filter weights, not including DC term.
dt (number): Duration of each time bin.
kws: Pass additional keyword arguments to plot()
"""
d = len(theta)
t = np.arange(-d + 1, 1) * dt
ax = plt.gca()
ax.plot(t, theta, marker="o", **kws)
ax.axhline(0, color=".2", linestyle="--", zorder=1)
ax.set(
xlabel="Time before spike (s)",
ylabel="Filter weight",
)
def plot_spikes_with_prediction(
spikes, predicted_spikes, dt, nt=50, t0=120, **kws):
"""Plot actual and predicted spike counts.
Args:
spikes (1D array): Vector of actual spike counts
predicted_spikes (1D array): Vector of predicted spike counts
dt (number): Duration of each time bin.
nt (number): Number of time bins to plot
t0 (number): Index of first time bin to plot.
kws: Pass additional keyword arguments to plot()
"""
t = np.arange(t0, t0 + nt) * dt
f, ax = plt.subplots()
lines = ax.stem(t, spikes[:nt], use_line_collection=True)
plt.setp(lines, color=".5")
lines[-1].set_zorder(1)
kws.setdefault("linewidth", 3)
yhat, = ax.plot(t, predicted_spikes[:nt], **kws)
ax.set(
xlabel="Time (s)",
ylabel="Spikes",
)
ax.yaxis.set_major_locator(plt.MaxNLocator(integer=True))
ax.legend([lines[0], yhat], ["Spikes", "Predicted"])
plt.show()
# + cellView="form"
#@title Data retrieval and loading
import os
import hashlib
import requests
fname = "RGCdata.mat"
url = "https://osf.io/mzujs/download"
expected_md5 = "1b2977453020bce5319f2608c94d38d0"
if not os.path.isfile(fname):
try:
r = requests.get(url)
except requests.ConnectionError:
print("!!! Failed to download data !!!")
else:
if r.status_code != requests.codes.ok:
print("!!! Failed to download data !!!")
elif hashlib.md5(r.content).hexdigest() != expected_md5:
print("!!! Data download appears corrupted !!!")
else:
with open(fname, "wb") as fid:
fid.write(r.content)
# -
# ---
# # Section 1: Linear-Gaussian GLM
# + cellView="form"
# @title Video 1: Linear Gaussian model
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV17T4y1E75x", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="Yv89UHeSa9I", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Section 1.1: Load retinal ganglion cell activity data
#
# *Estimated timing to here from start of tutorial: 10 min*
#
# In this exercise we use data from an experiment that presented a screen which randomly alternated between two luminance values and recorded responses from retinal ganglion cell (RGC), a type of neuron in the retina in the back of the eye. This kind of visual stimulus is called a "full-field flicker", and it was presented at ~120Hz (ie. the stimulus presented on the screen was refreshed about every 8ms). These same time bins were used to count the number of spikes emitted by each neuron.
#
# The file `RGCdata.mat` contains three variablies:
#
# - `Stim`, the stimulus intensity at each time point. It is an array with shape $T \times 1$, where $T=144051$.
#
# - `SpCounts`, the binned spike counts for 2 ON cells, and 2 OFF cells. It is a $144051 \times 4$ array, and each column has counts for a different cell.
#
# - `dtStim`, the size of a single time bin (in seconds), which is needed for computing model output in units of spikes / s. The stimulus frame rate is given by `1 / dtStim`.
#
# Because these data were saved in MATLAB, where everything is a matrix, we will also process the variables to more Pythonic representations (1D arrays or scalars, where appropriate) as we load the data.
# +
data = loadmat('RGCdata.mat') # loadmat is a function in scipy.io
dt_stim = data['dtStim'].item() # .item extracts a scalar value
# Extract the stimulus intensity
stim = data['Stim'].squeeze() # .squeeze removes dimensions with 1 element
# Extract the spike counts for one cell
cellnum = 2
spikes = data['SpCounts'][:, cellnum]
# Don't use all of the timepoints in the dataset, for speed
keep_timepoints = 20000
stim = stim[:keep_timepoints]
spikes = spikes[:keep_timepoints]
# -
# Use the `plot_stim_and_spikes` helper function to visualize the changes in stimulus intensities and spike counts over time.
plot_stim_and_spikes(stim, spikes, dt_stim)
# ### Coding Exercise 1.1: Create design matrix
#
# Our goal is to predict the cell's activity from the stimulus intensities preceding it. That will help us understand how RGCs process information over time. To do so, we first need to create the *design matrix* for this model, which organizes the stimulus intensities in matrix form such that the $i$th row has the stimulus frames preceding timepoint $i$.
#
# In this exercise, we will create the design matrix $\mathbf{X}$ using $d=25$ time lags. That is, $\mathbf{X}$ should be a $T \times d$ matrix. $d = 25$ (about 200 ms) is a choice we're making based on our prior knowledge of the temporal window that influences RGC responses. In practice, you might not know the right duration to use.
#
# The last entry in row `t` should correspond to the stimulus that was shown at time `t`, the entry to the left of it should contain the value that was show one time bin earlier, etc. Specifically, $X_{ij}$ will be the stimulus intensity at time $i + d - 1 - j$.
#
# Note that for the first few time bins, we have access to the recorded spike counts but not to the stimulus shown in the recent past. For simplicity we are going to assume that values of `stim` are 0 for the time lags prior to the first timepoint in the dataset. This is known as "zero-padding", so that the design matrix has the same number of rows as the response vectors in `spikes`.
#
# Your task is is to complete the function below to:
#
# - make a zero-padded version of the stimulus
# - initialize an empty design matrix with the correct shape
# - **fill in each row of the design matrix, using the zero-padded version of the stimulus**
#
# To visualize your design matrix (and the corresponding vector of spike counts), we will plot a "heatmap", which encodes the numerical value in each position of the matrix as a color. The helper functions include some code to do this.
# +
def make_design_matrix(stim, d=25):
"""Create time-lag design matrix from stimulus intensity vector.
Args:
stim (1D array): Stimulus intensity at each time point.
d (number): Number of time lags to use.
Returns
X (2D array): GLM design matrix with shape T, d
"""
# Create version of stimulus vector with zeros before onset
padded_stim = np.concatenate([np.zeros(d - 1), stim])
#####################################################################
# Fill in missing code (...),
# then remove or comment the line below to test your function
raise NotImplementedError("Complete the make_design_matrix function")
#####################################################################
# Construct a matrix where each row has the d frames of
# the stimulus preceding and including timepoint t
T = len(...) # Total number of timepoints (hint: number of stimulus frames)
X = np.zeros((T, d))
for t in range(T):
X[t] = ...
return X
# Make design matrix
X = make_design_matrix(stim)
# Visualize
plot_glm_matrices(X, spikes, nt=50)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D4_GeneralizedLinearModels/solutions/W1D4_Tutorial1_Solution_03ed3adf.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=830.0 height=1115.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D4_GeneralizedLinearModels/static/W1D4_Tutorial1_Solution_03ed3adf_0.png>
#
#
# -
# ## Section 1.2: Fit Linear-Gaussian regression model
#
# *Estimated timing to here from start of tutorial: 25 min*
#
#
# First, we will use the design matrix to compute the maximum likelihood estimate for a linear-Gaussian GLM (aka "general linear model"). The maximum likelihood estimate of $\theta$ in this model can be solved analytically using the equation you learned about on Day 3:
#
# \begin{align}
# \boldsymbol{\hat \theta} = (\mathbf{X}^{\top}\mathbf{X})^{-1}\mathbf{X}^{\top}\mathbf{y}
# \end{align}
#
# Before we can apply this equation, we need to augment the design matrix to account for the mean of $y$, because the spike counts are all $\geq 0$. We do this by adding a constant column of 1's to the design matrix, which will allow the model to learn an additive offset weight. We will refer to this additional weight as $b$ (for bias), although it is alternatively known as a "DC term" or "intercept".
# +
# Build the full design matrix
y = spikes
constant = np.ones_like(y)
X = np.column_stack([constant, make_design_matrix(stim)])
# Get the MLE weights for the LG model
theta = np.linalg.inv(X.T @ X) @ X.T @ y
theta_lg = theta[1:]
# -
# Plot the resulting maximum likelihood filter estimate (just the 25-element weight vector $\theta$ on the stimulus elements, not the DC term $b$).
plot_spike_filter(theta_lg, dt_stim)
# ### Coding Exercise 1.2: Predict spike counts with Linear-Gaussian model
#
# Now we are going to put these pieces together and write a function that outputs a predicted spike count for each timepoint using the stimulus information.
#
# Your steps should be:
#
# - Create the complete design matrix
# - Obtain the MLE weights ($\boldsymbol{\hat \theta}$)
# - Compute $\mathbf{\hat y} = \mathbf{X}\boldsymbol{\hat \theta}$
# +
def predict_spike_counts_lg(stim, spikes, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
##########################################################################
# Fill in missing code (...) and then comment or remove the error to test
raise NotImplementedError("Complete the predict_spike_counts_lg function")
##########################################################################
# Create the design matrix
y = spikes
constant = ...
X = ...
# Get the MLE weights for the LG model
theta = ...
# Compute predicted spike counts
yhat = X @ theta
return yhat
# Predict spike counts
predicted_counts = predict_spike_counts_lg(stim, spikes)
# Visualize
plot_spikes_with_prediction(spikes, predicted_counts, dt_stim)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D4_GeneralizedLinearModels/solutions/W1D4_Tutorial1_Solution_823fa455.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1120.0 height=832.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D4_GeneralizedLinearModels/static/W1D4_Tutorial1_Solution_823fa455_0.png>
#
#
# -
# Is this a good model? The prediction line more-or-less follows the bumps in the spikes, but it never predicts as many spikes as are actually observed. And, more troublingly, it's predicting *negative* spikes for some time points.
#
# The Poisson GLM will help to address these failures.
#
#
# ### Bonus challenge
#
# The "spike-triggered average" falls out as a subcase of the linear Gaussian GLM: $\mathrm{STA} = \mathbf{X}^{\top} \mathbf{y} \,/\, \textrm{sum}(\mathbf{y})$, where $\mathbf{y}$ is the vector of spike counts of the neuron. In the LG GLM, the term $(\mathbf{X}^{\top}\mathbf{X})^{-1}$ corrects for potential correlation between the regressors. Because the experiment that produced these data used a white noise stimulus, there are no such correlations. Therefore the two methods are equivalent. (How would you check the statement about no correlations?)
# ---
# # Section 2: Linear-Nonlinear-Poisson GLM
#
# *Estimated timing to here from start of tutorial: 36 min*
#
# + cellView="form"
# @title Video 2: Generalized linear model
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="BV1mz4y1X7JZ", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="wRbvwdze4uE", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Section 2.1: Nonlinear optimization with `scipy.optimize`
#
# *Estimated timing to here from start of tutorial: 45 min*
#
# Before diving into the Poisson GLM case, let us review the use and importance of convexity in optimization:
# - We have seen previously that in the Linear-Gaussian case, maximum likelihood parameter estimate can be computed analytically. That is great because it only takes us a single line of code!
# - Unfortunately in general there is no analytical solution to our statistical estimation problems of interest. Instead, we need to apply a nonlinear optimization algorithm to find the parameter values that minimize some *objective function*. This can be extremely tedious because there is no general way to check whether we have found *the optimal solution* or if we are just stuck in some local minimum.
# - Somewhere in between these two extremes, the special case of convex objective function is of great practical importance. Indeed, such optimization problems can be solved very reliably (and usually quite rapidly too!) using some standard software.
#
# Notes:
# - a function is convex if and only if its curve lies below any chord joining two of its points
# - to learn more about optimization, you can consult the book of <NAME> and <NAME> [Convex Optimization](https://web.stanford.edu/~boyd/cvxbook/).
# Here we will use the `scipy.optimize` module, it contains a function called [`minimize`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html) that provides a generic interface to a large number of optimization algorithms. This function expects as argument an objective function and an "initial guess" for the parameter values. It then returns a dictionary that includes the minimum function value, the parameters that give this minimum, and other information.
#
# Let's see how this works with a simple example. We want to minimize the function $f(x) = x^2$:
# +
f = np.square
res = minimize(f, x0=2)
print(
f"Minimum value: {res['fun']:.4g}",
f"at x = {res['x']}",
)
# -
# When minimizing a $f(x) = x^2$, we get a minimum value of $f(x) \approx 0$ when $x \approx 0$. The algorithm doesn't return exactly $0$, because it stops when it gets "close enough" to a minimum. You can change the `tol` parameter to control how it defines "close enough".
#
# A point about the code bears emphasis. The first argument to `minimize` is not a number or a string but a *function*. Here, we used `np.square`. Take a moment to make sure you understand what's going on, because it's a bit unusual, and it will be important for the exercise you're going to do in a moment.
#
# In this example, we started at $x_0 = 2$. Let's try different values for the starting point:
# +
start_points = -1, 1.5
xx = np.linspace(-2, 2, 100)
plt.plot(xx, f(xx), color=".2")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
for i, x0 in enumerate(start_points):
res = minimize(f, x0)
plt.plot(x0, f(x0), "o", color=f"C{i}", ms=10, label=f"Start {i}")
plt.plot(res["x"].item(), res["fun"], "x", c=f"C{i}", ms=10, mew=2, label=f"End {i}")
plt.legend()
# -
# The runs started at different points (the dots), but they each ended up at roughly the same place (the cross): $f(x_\textrm{final}) \approx 0$. Let's see what happens if we use a different function:
# +
g = lambda x: x / 5 + np.cos(x)
start_points = -.5, 1.5
xx = np.linspace(-4, 4, 100)
plt.plot(xx, g(xx), color=".2")
plt.xlabel("$x$")
plt.ylabel("$f(x)$")
for i, x0 in enumerate(start_points):
res = minimize(g, x0)
plt.plot(x0, g(x0), "o", color=f"C{i}", ms=10, label=f"Start {i}")
plt.plot(res["x"].item(), res["fun"], "x", color=f"C{i}", ms=10, mew=2, label=f"End {i}")
plt.legend()
# -
# Unlike $f(x) = x^2$, $g(x) = \frac{x}{5} + \cos(x)$ is not *convex*. We see that the final position of the minimization algorithm depends on the starting point, which adds a layer of comlpexity to such problems.
# ### Coding Exercise 2.1: Fitting the Poisson GLM and prediction spikes
#
# In this exercise, we will use [`scipy.optimize.minimize`](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html) to compute maximum likelihood estimates for the filter weights in the Poissson GLM model with an exponential nonlinearity (LNP: Linear-Nonlinear-Poisson).
#
# In practice, this will involve filling out two functions.
#
# - The first should be an *objective function* that takes a design matrix, a spike count vector, and a vector of parameters. It should return a negative log likelihood.
# - The second function should take `stim` and `spikes`, build the design matrix and then use `minimize` internally, and return the MLE parameters.
#
# What should the objective function look like? We want it to return the negative log likelihood: $-\log P(y \mid \mathbf{X}, \theta).$
#
# In the Poisson GLM,
#
# \begin{align}
# \log P(\mathbf{y} \mid \mathbf{X}, \theta) = \sum_t \log P(y_t \mid \mathbf{x_t},\theta),
# \end{align}
#
# where
#
# \begin{align}
# P(y_t \mid \mathbf{x_t}, \theta) = \frac{\lambda_t^{y_t}\exp(-\lambda_t)}{y_t!} \text{, with rate } \lambda_t = \exp(\mathbf{x_t}^{\top} \theta).
# \end{align}
#
# Now, taking the log likelihood for all the data we obtain:
# $\log P(\mathbf{y} \mid X, \theta) = \sum_t( y_t \log\left(\lambda_t) - \lambda_t - \log(y_t !)\right).$
#
# Because we are going to minimize the negative log likelihood with respct to the parameters $\theta$, we can ignore the last term that does not depend on $\theta$. For faster implementation, let us rewrite this in matrix notation:
#
# \begin{align}
# \mathbf{y}^{\top} \log(\mathbf{\lambda}) - \mathbf{1}^{\top} \mathbf{\lambda} \text{, with rate } \mathbf{\lambda} = \exp(\mathbf{X} \theta)
# \end{align}
#
# Finally, don't forget to add the minus sign for your function to return the negative log likelihood.
# +
def neg_log_lik_lnp(theta, X, y):
"""Return -loglike for the Poisson GLM model.
Args:
theta (1D array): Parameter vector.
X (2D array): Full design matrix.
y (1D array): Data values.
Returns:
number: Negative log likelihood.
"""
#####################################################################
# Fill in missing code (...), then remove the error
raise NotImplementedError("Complete the neg_log_lik_lnp function")
#####################################################################
# Compute the Poisson log likeliood
rate = np.exp(X @ theta)
log_lik = y @ ... - ...
return ...
def fit_lnp(stim, spikes, d=25):
"""Obtain MLE parameters for the Poisson GLM.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
d (number): Number of time lags to use.
Returns:
1D array: MLE parameters
"""
#####################################################################
# Fill in missing code (...), then remove the error
raise NotImplementedError("Complete the fit_lnp function")
#####################################################################
# Build the design matrix
y = spikes
constant = np.ones_like(y)
X = np.column_stack([constant, make_design_matrix(stim)])
# Use a random vector of weights to start (mean 0, sd .2)
x0 = np.random.normal(0, .2, d + 1)
# Find parameters that minmize the negative log likelihood function
res = minimize(..., args=(X, y))
return ...
# Fit LNP model
theta_lnp = fit_lnp(stim, spikes)
# Visualize
plot_spike_filter(theta_lg[1:], dt_stim, color=".5", label="LG")
plot_spike_filter(theta_lnp[1:], dt_stim, label="LNP")
plt.legend(loc="upper left");
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D4_GeneralizedLinearModels/solutions/W1D4_Tutorial1_Solution_f09f1cc5.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1116.0 height=828.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D4_GeneralizedLinearModels/static/W1D4_Tutorial1_Solution_f09f1cc5_0.png>
#
#
# -
# Plotting the LG and LNP weights together, we see that they are broadly similar, but the LNP weights are generally larger. What does that mean for the model's ability to *predict* spikes? To see that, let's finish the exercise by filling out the `predict_spike_counts_lnp` function:
# +
def predict_spike_counts_lnp(stim, spikes, theta=None, d=25):
"""Compute a vector of predicted spike counts given the stimulus.
Args:
stim (1D array): Stimulus values at each timepoint
spikes (1D array): Spike counts measured at each timepoint
theta (1D array): Filter weights; estimated if not provided.
d (number): Number of time lags to use.
Returns:
yhat (1D array): Predicted spikes at each timepoint.
"""
###########################################################################
# Fill in missing code (...) and then remove the error to test
raise NotImplementedError("Complete the predict_spike_counts_lnp function")
###########################################################################
y = spikes
constant = np.ones_like(spikes)
X = np.column_stack([constant, make_design_matrix(stim)])
if theta is None: # Allow pre-cached weights, as fitting is slow
theta = fit_lnp(X, y, d)
yhat = ...
return yhat
# Predict spike counts
yhat = predict_spike_counts_lnp(stim, spikes, theta_lnp)
# Visualize
plot_spikes_with_prediction(spikes, yhat, dt_stim)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W1D4_GeneralizedLinearModels/solutions/W1D4_Tutorial1_Solution_ae48f475.py)
#
# *Example output:*
#
# <img alt='Solution hint' align='left' width=1120.0 height=832.0 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W1D4_GeneralizedLinearModels/static/W1D4_Tutorial1_Solution_ae48f475_0.png>
#
#
# -
# We see that the LNP model does a better job of fitting the actual spiking data. Importantly, it never predicts negative spikes!
#
# *Bonus:* Our statement that the LNP model "does a better job" is qualitative and based mostly on the visual appearance of the plot. But how would you make this a quantitative statement?
# ---
# # Summary
#
# *Estimated timing of tutorial: 1 hour, 15 minutes*
#
# In this first tutorial, we used two different models to learn something about how retinal ganglion cells respond to a flickering white noise stimulus. We learned how to construct a design matrix that we could pass to different GLMs, and we found that the Linear-Nonlinear-Poisson (LNP) model allowed us to predict spike rates better than a simple Linear-Gaussian (LG) model.
#
# In the next tutorial, we'll extend these ideas further. We'll meet yet another GLM — logistic regression — and we'll learn how to ensure good model performance even when the number of parameters `d` is large compared to the number of data points `N`.
# ---
# # Notation
#
# \begin{align}
# y &\quad \text{measurement or response, here: spike count}\\
# T &\quad \text{number of time points}\\
# d &\quad \text{input dimensionality}\\
# \mathbf{X} &\quad \text{design matrix, dimensions: } T \times d\\
# \theta &\quad \text{parameter}\\
# \hat \theta &\quad \text{estimated parameter}\\
# \hat y &\quad \text{estimated response}\\
# P(\mathbf{y} \mid \mathbf{X}, \theta) &\quad \text{probability of observing response } y \text{ given design matrix } \mathbf{X} \text{ and parameters } \theta \\
# \mathrm{STA} &\quad \text{spike-triggered average}\\
# b &\quad \text{bias weight, intercept}\\
# \end{align}
|
tutorials/W1D4_GeneralizedLinearModels/student/W1D4_Tutorial1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UJ7tbIrsI7Ij"
# K-means clustering
# + id="8FBA8626JDVi"
import numpy as np, pandas as pd, matplotlib.pyplot as plt
# %matplotlib inline
# + id="BZhLgw1sCd1v"
# K-means clustering Algorithm
import random
class K_means():
def __init__(self,X,n_clusters):
self.X = X
self.feature_cols = X.columns
self.cluster_count = n_clusters
self.random_init_points = pd.DataFrame([])
self.get_random_inits()
b=self.random_init_points[self.random_init_points.duplicated()]
if b.shape[0]>0:
self.get_random_inits()
self.cluster_tags = self.get_tags()
rows = self.X.shape[0]
self.X['assigned_cluster'] = np.zeros(rows)
for tag in self.cluster_tags:
s = "distance_to_" + tag
self.X[s] = np.zeros(rows)
self.random_init_points.reset_index(inplace=True)
self.random_init_points.drop("index",axis =1,inplace=True)
self.distance_tags =[]
for tag in self.cluster_tags:
s = "distance_to_" + tag
self.distance_tags.append(s)
def get_random_inits(self):
l = list(self.X.index.to_series().values)
temp =random.sample(l,self.cluster_count )
l=[]
for i in temp:
l.append(self.X.iloc[i])
self.random_init_points = pd.DataFrame(l)
def get_tags(self):
tag_dict= {0:"zero ", 1:"one ",2:"two ",3:"three ",4:"four ", 5:"five ", 6:"six ",7:"seven ",8:"eight ",9:"nine "}
l = []
if self.cluster_count < 2:
print ("[i] Number of clusters less than 2, this model is broken! reinitialise it with 2 or more clusters")
return
mod = self.cluster_count
for cluster_number in range (mod):
Temp = cluster_number
s=[]
w = ""
for digits in range(len(str(cluster_number))):
mod = Temp % 10
Temp = Temp / 10
s.append(tag_dict[mod])
s.reverse()
for i in s:
w = w + i
w = w.rstrip()
l.append(w)
return l
def get_euclidean(self):
for dist_to_point in self.distance_tags:
self.X[dist_to_point] = np.sqrt((np.square(self.X - self.random_init_points.iloc[self.distance_tags.index(dist_to_point)])).sum(axis=1))
return
def assign_cluster(self):
for m in self.X.index.to_series().values:
min_point=self.distance_tags.index(self.X[self.distance_tags].iloc[m][self.X[self.distance_tags].iloc[m] == self.X[self.distance_tags].iloc[m].min()].index[0])
self.X.iloc[m]['assigned_cluster'] = min_point
return
def shift_centroid(self):
for cluster in self.X.assigned_cluster.values:
self.random_init_points.iloc[int(cluster)]=self.X[self.feature_cols][self.X['assigned_cluster']== cluster].mean()
return
def cost(self):
c = 0
for cluster in range(len(self.distance_tags)):
c = c + self.X[self.distance_tags][self.X["assigned_cluster"]== cluster].values.sum()
return c
def create_clusters(self):
prev_cost =0
iter = 0
cost = 1
while prev_cost != cost:
# step 1 get euclidean distance
self.get_euclidean()
# step 2 centriod closest among all to this point becomes the centroid for this point
self.assign_cluster()
# calculate mean of all points in a cluster and shift centroid to mean
self.shift_centroid()
prev_cost = cost
cost = self.cost()
iter = iter + 1
print("[i] Iterations: ",iter)
print("[i] Cost: ",cost)
return
# + id="2t4S2F2wN1N2" outputId="9cc5ac08-b68f-4bee-f787-d197641afdcc" colab={"base_uri": "https://localhost:8080/", "height": 475}
X =pd.DataFrame([[0.5,8],[1,5.5],[1.7,8],[2,7],[3.3,7],[3.9,6],[4,7],[1,5.5],[2,2],[2.1,3.2],[1,5.5],[3,0.5],[1,5.5],[3,5.2],[1,5.5],[4.5,1.5],[5.1,5.5],[5.5,7.1],[5.9,7.5],[6,8.5],[6.1,7.9],[7,7],[6.9,5.1],[7.2,8],[7.5,6],[5.1,2],[5.1,3],[6.1,1.5],[6,2],[6,4.1],[6.1,1],[6.9,3],[6.9,4],[7,1.5],[7.2,3]])
plt.scatter(X[0],X[1])
plt.title("Scatter Plot")
plt.xlabel("X1")
plt.ylabel("X2")
plt.show()
df = pd.DataFrame(X.iloc[:,:].values)
K = K_means(df,5)
# every time the object is invoked random points are chosen as init
K.random_init_points
# + id="bABmqNaNpQNm" outputId="650a5c1e-1913-47de-9b78-7281f785c3ef" colab={"base_uri": "https://localhost:8080/", "height": 54}
K.create_clusters()
# + id="bZXpSxuLpxlK" outputId="32805613-9932-4378-d4fe-7bd1205e506b" colab={"base_uri": "https://localhost:8080/", "height": 1000}
K.X
# + id="6YUmh4pD3TAM" outputId="7886ea82-cd7d-4973-a23e-d6f96864b67e" colab={"base_uri": "https://localhost:8080/", "height": 295}
#Plot Visualization
plt.scatter(K.X[0][K.X['assigned_cluster']==0],K.X[1][K.X['assigned_cluster']==0])
plt.scatter(K.random_init_points.iloc[0][0],K.random_init_points.iloc[0][1],marker="X")
plt.scatter(K.X[0][K.X['assigned_cluster']==1],K.X[1][K.X['assigned_cluster']==1])
plt.scatter(K.random_init_points.iloc[1][0],K.random_init_points.iloc[1][1],marker="X")
plt.scatter(K.X[0][K.X['assigned_cluster']==2],K.X[1][K.X['assigned_cluster']==2])
plt.scatter(K.random_init_points.iloc[2][0],K.random_init_points.iloc[2][1],marker="X")
plt.scatter(K.X[0][K.X['assigned_cluster']==3],K.X[1][K.X['assigned_cluster']==3])
plt.scatter(K.random_init_points.iloc[3][0],K.random_init_points.iloc[3][1],marker="X")
plt.scatter(K.X[0][K.X['assigned_cluster']==4],K.X[1][K.X['assigned_cluster']==4])
plt.scatter(K.random_init_points.iloc[4][0],K.random_init_points.iloc[4][1],marker="X")
plt.title("K-means clustering")
plt.xlabel("X1")
plt.ylabel("X2")
plt.show()
|
Kmeans.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Open the session of Tensorflow
# bash-3.2$ source activate tensorflow
#
#
# (tensorflow) bash-3.2$ jupyter notebook
import tensorflow as tf
print(tf.__version__)
import numpy as np
# to make this notebook's output stable across runs
np.random.seed(42)
# # Regression
x = tf.placeholder(dtype=tf.float64, name='x')
y = tf.placeholder(dtype=tf.float64, name='y')
w = tf.Variable(0., dtype=tf.float64, name='w')
b = tf.Variable(0., dtype=tf.float64, name='b')
tf.summary.scalar('w', w);
tf.summary.scalar('b', b);
with tf.name_scope('y1'):
y1 = w*x + b
with tf.name_scope('loss'):
loss = tf.reduce_mean(tf.square(y - y1))
tf.summary.scalar('loss',loss)
optimizer = tf.train.GradientDescentOptimizer(0.5)
step = tf.Variable(0, name='step', trainable=False)
train = optimizer.minimize(loss, global_step=step)
initialize = tf.global_variables_initializer()
# +
from datetime import datetime
now = datetime.utcnow().strftime("%Y%m%d")
root_logdir = "tf_logs"
logdir = "{}/run-{}/".format(root_logdir, now)
summary = [tf.summary.merge_all(), step]
writer = tf.summary.FileWriter(logdir)
# -
logdir
# # tensorboard
# $ source activate tensorflow
#
#
# $ tensorboard --logdir tf_logs/run-20170719/
x_dat = np.random.rand(100)
y_dat = 3.*x_dat + 2. + 0.1*np.random.rand(100)
feed_dict = {x:x_dat, y:y_dat}
sess = tf.Session()
writer.add_graph(sess.graph)
sess.run(initialize)
for step in range(201):
sess.run(train, feed_dict)
if step%20 == 0:
print(step, sess.run([w,b,loss], feed_dict))
writer.add_summary(*sess.run(summary, feed_dict))
|
code/tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alirezash97/Cardio/blob/master/HeartBeat.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-aUFDobFVvj1" colab_type="code" colab={}
# !pip install kaggle
# !mkdir .kaggle
import json
token = {"username":"alirezashafaei97","key":"<KEY>"}
with open('/content/.kaggle/kaggle.json', 'w') as file:
json.dump(token, file)
# !mkdir ~/.kaggle
# !cp /content/.kaggle/kaggle.json ~/.kaggle/kaggle.json
# !kaggle config set -n path -v{/content}
# !chmod 600 /root/.kaggle/kaggle.json
# !kaggle datasets download -d shayanfazeli/heartbeat -p /content
# !unzip /content/heartbeat.zip -d /content/heartbeat
# + id="Sio7SWAwWBLj" colab_type="code" outputId="5e70d484-5ea4-484f-8cab-11d7fa9be8ea" colab={"base_uri": "https://localhost:8080/", "height": 34}
import pandas as pd
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.models import load_model, Sequential, Model
from tensorflow.keras.layers import (Input, Dense, LeakyReLU, Softmax, InputLayer, concatenate, Conv1D, MaxPool1D, Add, MaxPooling1D
, Flatten, Dropout, ReLU, BatchNormalization, GlobalAveragePooling1D)
from keras.utils.np_utils import to_categorical
from tensorflow.keras import backend as K
from tensorflow.keras import regularizers
from random import uniform
import random
from sklearn.preprocessing import OneHotEncoder
from scipy.sparse import csr_matrix
# + id="DTudiQO8WEXT" colab_type="code" colab={}
train_df=pd.read_csv('/content/heartbeat/mitbih_train.csv',header=None)
test_df=pd.read_csv('/content/heartbeat/mitbih_test.csv',header=None)
# + id="N2SBJpgbi9WG" colab_type="code" colab={}
# train_df = train_df[:186].astype('float16')
# test_df = test_df[:186].astype('float16')
# + id="i58wFx5vXQS7" colab_type="code" outputId="142bbe10-a1bf-4fde-96a2-85fa8d58209c" colab={"base_uri": "https://localhost:8080/", "height": 118}
train_df[187]=train_df[187].astype(int)
counter=train_df[187].value_counts()
print(counter)
# + id="asQqNZG2q59y" colab_type="code" colab={}
from sklearn.utils import resample
df_1=train_df[train_df[187]==1]
df_2=train_df[train_df[187]==2]
df_3=train_df[train_df[187]==3]
df_4=train_df[train_df[187]==4]
df_0=(train_df[train_df[187]==0]).sample(n=10000,random_state=42)
df_1_upsample=resample(df_1,replace=True,n_samples=10000,random_state=123)
df_2_upsample=resample(df_2,replace=True,n_samples=10000,random_state=124)
df_3_upsample=resample(df_3,replace=True,n_samples=10000,random_state=125)
df_4_upsample=resample(df_4,replace=True,n_samples=10000,random_state=126)
train_df=pd.concat([df_0,df_1_upsample,df_2_upsample,df_3_upsample,df_4_upsample])
# + id="bomIXpkCrozb" colab_type="code" colab={}
target_train=train_df[187]
target_test=test_df[187]
y_train=to_categorical(target_train)
y_test=to_categorical(target_test)
# + id="yYBctv4Tr58D" colab_type="code" outputId="31e5a711-6003-4415-bec1-d2ddf10c0969" colab={"base_uri": "https://localhost:8080/", "height": 118}
equilibre=train_df[187].value_counts()
print(equilibre)
# + id="vX3HJfHYr605" colab_type="code" outputId="9a34ad27-aafb-47d6-ff53-f8ccf06d5806" colab={"base_uri": "https://localhost:8080/", "height": 244}
c=train_df.groupby(187,group_keys=False).apply(lambda train_df : train_df.sample(1))
c
# + id="ie12gfa0cRQE" colab_type="code" colab={}
# del c
# + id="VwB80IzMo1iK" colab_type="code" colab={}
def add_gaussian_noise(signal):
noise=np.random.normal(0,0.05,186)
return (signal+noise)
# + id="PJB5fWu5sWNc" colab_type="code" outputId="0ce95d0e-46c2-4840-f4f8-2b6dd9b82332" colab={"base_uri": "https://localhost:8080/", "height": 265}
tempo=c.iloc[0,:186]
bruiter=add_gaussian_noise(tempo)
plt.subplot(2,1,1)
plt.plot(c.iloc[0,:186])
plt.subplot(2,1,2)
plt.plot(bruiter)
plt.show()
# + id="VG4fXuypr6vv" colab_type="code" colab={}
target_train=train_df[187]
target_test=test_df[187]
y_train=to_categorical(target_train)
y_test=to_categorical(target_test)
# + id="PUVlLpKPsQUz" colab_type="code" colab={}
train_df = train_df.astype('float16')
test_df = test_df.astype('float16')
X_train=train_df.iloc[:,:186].values
X_test=test_df.iloc[:,:186].values
# X_train = X_train.reshape(len(X_train), X_train.shape[1],1)
# X_test = X_test.reshape(len(X_test), X_test.shape[1],1)
# + id="TexzfSPiD34D" colab_type="code" outputId="389ab1a9-6086-4fa4-b6cb-9f630d800043" colab={"base_uri": "https://localhost:8080/", "height": 50}
print(X_train.shape)
print(X_test.shape)
# + id="GPfQkUVfnTx9" colab_type="code" colab={}
# data augmentation
def augmetation(X_train, y_train, chance):
augment_number = 0
XF_train = np.zeros((X_train.shape[0]*2, X_train.shape[1]))
yf_train = np.zeros((y_train.shape[0]*2, y_train.shape[1]))
pointer = 0
for index, row in enumerate(X_train):
XF_train[pointer, :] = row
yf_train[pointer, :] = y_train[index, :]
pointer += 1
rand_num = random.uniform(0, 1)
if chance > rand_num :
augment_number += 1
noise = np.random.normal(0,0.05,186)
new_signal = row + noise
XF_train[pointer, :] = new_signal
yf_train[pointer, :] = y_train[index, :]
pointer += 1
filled = X_train.shape[0] + augment_number
XFF_train = XF_train[:filled, :]
yff_train = yf_train[:filled, :]
return XFF_train, yff_train
# + id="4dXhm8thnj1x" colab_type="code" colab={}
X_train, y_train = augmetation(X_train, y_train, 0.1)
# + id="dzROJp1Fr6q6" colab_type="code" colab={}
# periodic signal extend
import pywt
XF_train = np.zeros((X_train.shape[0], 9000))
XF_test = np.zeros((X_test.shape[0], 9000))
for index, row in enumerate(X_train):
XF_train[index, :] = pywt.pad(row, 4407, 'periodic')
for index, row in enumerate(X_test):
XF_test[index, :] = pywt.pad(row, 4407, 'periodic')
# + id="cmANeCYtjmgE" colab_type="code" colab={}
XF_train = XF_train.reshape((XF_train.shape[0], 9000, 1))
XF_test = XF_test.reshape((XF_test.shape[0], 9000, 1))
# + id="vZNvkTDCjyoA" colab_type="code" outputId="eeec7109-224c-4337-ae8e-c119d3340e84" colab={"base_uri": "https://localhost:8080/", "height": 84}
print("X_train : ", XF_train.shape)
print("Y_train : ", y_train.shape)
print("X_test : ", XF_test.shape)
print("Y_test : ", y_test.shape)
# + id="zKvIW2sWkaoP" colab_type="code" outputId="c0250120-8f5f-4084-d120-a446e8ef0b21" colab={"base_uri": "https://localhost:8080/", "height": 1000}
X_input = Input(shape=(9000, 1))
Conv = Conv1D(filters=64, kernel_size=5, strides=3)(X_input)
### step 1
Conv1_1 = Conv1D(filters=64, kernel_size=9, strides=1, padding='same')(Conv)
Bn1_1 = BatchNormalization()(Conv1_1)
Act1_1 = LeakyReLU()(Bn1_1)
Conv1_2 = Conv1D(filters=64, kernel_size=7, strides=1, padding='same')(Act1_1)
Bn1_2 = BatchNormalization()(Conv1_2)
Act1_2 = LeakyReLU()(Bn1_2)
DO1_1 = Dropout(0.2)(Act1_2)
Conv1_3 = Conv1D(filters=64, kernel_size=9, strides=1, padding='same')(DO1_1)
Bn1_3 = BatchNormalization()(Conv1_3)
shortcut1_1 = Add()([Bn1_3, Conv])
Bn1_4 = BatchNormalization()(shortcut1_1)
Act1_3 = LeakyReLU()(Bn1_4)
##### auxiliary
Conv1_4 = Conv1D(filters=128, kernel_size=7, strides=3, padding='same')(Act1_3)
Bn1_5 = BatchNormalization()(Conv1_4)
Act1_4 = LeakyReLU()(Bn1_5)
###############
Max1_1 = MaxPooling1D(pool_size=5, strides=2)(Act1_4)
## step 2
Conv2_1 = Conv1D(filters=256, kernel_size=3, strides=1, padding='same')(Max1_1)
Bn2_1 = BatchNormalization()(Conv2_1)
Act2_1 = LeakyReLU()(Bn2_1)
Conv2_2 = Conv1D(filters=256, kernel_size=5, strides=1, padding='same')(Act2_1)
Bn2_2 = BatchNormalization()(Conv2_2)
Act2_2 = LeakyReLU()(Bn2_2)
DO2_1 = Dropout(0.2)(Act2_2)
Conv2_3 = Conv1D(filters=128, kernel_size=3, strides=1, padding='same')(DO2_1)
Bn2_3 = BatchNormalization()(Conv2_3)
shortcut2_1 = Add()([Bn2_3, Max1_1])
Bn2_4 = BatchNormalization()(shortcut2_1)
Act2_3 = LeakyReLU()(Bn2_4)
##### auxiliary
Conv2_4 = Conv1D(filters=512, kernel_size=7, strides=2, padding='same')(Act2_3)
Bn2_5 = BatchNormalization()(Conv2_4)
Act2_4 = LeakyReLU()(Bn2_5)
###############
Max2_1 = MaxPooling1D(pool_size=5, strides=3)(Act2_4)
Flat1 = Flatten()(Max2_1)
D1 = Dense(256)(Flat1)
A6 = LeakyReLU()(D1)
D_O = Dropout(0.15)(A6)
D2 = Dense(128)(D_O)
D3 = Dense(5)(D2)
A7 = Softmax()(D3)
model = Model(inputs=X_input, outputs=A7)
model.summary()
# + id="MSfZxMLK7xI6" colab_type="code" colab={}
lr_schedule = tf.keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=1e-3,
decay_steps=1719,
decay_rate=0.7)
opt = tf.keras.optimizers.Adam(learning_rate=lr_schedule)
# + id="J51SxpfQofzM" colab_type="code" outputId="c68dde1e-58cc-46a4-bbfb-079b76c62712" colab={"base_uri": "https://localhost:8080/", "height": 218}
# overfitting so augment more data and decrease initial learning rate to 1e-3
# compile model
model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['accuracy'])
# Early Stopping
es_callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=3)
# Fit the model
history = model.fit(XF_train, y_train, epochs=6, batch_size=64, validation_data=(XF_test, y_test), callbacks=[es_callback])
# + id="KrUPC94zqTiE" colab_type="code" outputId="15e76721-375a-4993-e207-30055522d4b8" colab={"base_uri": "https://localhost:8080/", "height": 229}
# evaluate the model
scores = model.evaluate(XF_test, y_test, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# save model and architecture to single file
model.save("/content/drive/My Drive/Cardio/HeartBeat.h5")
# + id="VCecYVrhtbNa" colab_type="code" colab={}
/*
# + id="ht3ltnGvtjlU" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
|
HeartBeat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cdei
# language: python
# name: cdei
# ---
# # Data reweighting by <NAME>
#
# This notebook contains an implementation of the pre-processing fairness intervention introduced in [Data preprocessing techniques for classification without discrimination](https://link.springer.com/article/10.1007/s10115-011-0463-8) by <NAME> (2012) as part of the IBM AIF360 fairness tool box github.com/IBM/AIF360.
#
# The intervention achieves demographic parity by attaching weights to the data so that certain types of observations are more influential during training, thereby balancing out the label distributions across different protected groups. The resulting weights can also be used to resample the data set with replacement to create a fair transformed data set.
# +
from pathlib import Path
import joblib
import numpy as np
import pandas as pd
from aif360.algorithms.preprocessing.reweighing import Reweighing
from aif360.datasets import StandardDataset
from fairlearn.metrics import demographic_parity_difference
from helpers.metrics import accuracy
from helpers.plot import group_box_plots
from sklearn.linear_model import LogisticRegression
# + tags=["export"]
from helpers import export_plot
# -
# ## Load data
#
# We have committed preprocessed data to the repository for reproducibility and we load it here. Check out hte preprocessing notebook for details on how this data was obtained.
artifacts_dir = Path("../../../artifacts")
# + tags=["export"]
# override data_dir in source notebook
# this is stripped out for the hosted notebooks
artifacts_dir = Path("../../../../artifacts")
# +
data_dir = artifacts_dir / "data" / "adult"
train = pd.read_csv(data_dir / "processed" / "train-one-hot.csv")
val = pd.read_csv(data_dir / "processed" / "val-one-hot.csv")
test = pd.read_csv(data_dir / "processed" / "test-one-hot.csv")
# -
# In order to process data for our fairness intervention we need to define special dataset objects which are part of every intervention pipeline within the IBM AIF360 toolbox. These objects contain the original data as well as some useful further information, e.g., which feature is the protected attribute as well as which column corresponds to the label.
train_sds = StandardDataset(
train,
label_name="salary",
favorable_classes=[1],
protected_attribute_names=["sex"],
privileged_classes=[[1]],
)
test_sds = StandardDataset(
test,
label_name="salary",
favorable_classes=[1],
protected_attribute_names=["sex"],
privileged_classes=[[1]],
)
val_sds = StandardDataset(
val,
label_name="salary",
favorable_classes=[1],
protected_attribute_names=["sex"],
privileged_classes=[[1]],
)
# Define which binary value goes with the (un-)privileged group
privileged_groups = [{"sex": 1.0}]
unprivileged_groups = [{"sex": 0.0}]
# ## Load original model
# For maximum reproducibility we can also load the baseline model from disk, but the code used to train can be found in the baseline model notebook.
# +
bl_model = joblib.load(artifacts_dir / "models" / "finance" / "baseline.pkl")
bl_test_probs = bl_model.predict_proba(test.drop("salary", axis=1))[:, 1]
bl_test_pred = bl_test_probs > 0.5
# -
# ## Demographic parity
#
# We learn the data transformation due to Kamiran and Claders on the training data. The transformation attaches fair weights to data it is applied to. A fair data set can then be generated via weighted sampling. We apply the transformation to the validation set, but instead of resampling according to the resulting weights, we train a logisitc regression model using the underlying weights in the validation set. Finally, we generate predictions for the test data based on the leanrnt fair logisitic regression and analyse the outcomes for fairness and accuracy.
#
# The intervention does not require any parameter tuning.
RW = Reweighing(
unprivileged_groups=unprivileged_groups,
privileged_groups=privileged_groups,
)
RW.fit(train_sds)
# Apply intervention on validation data.
val_sds_transf = RW.transform(val_sds)
# ## Train fair model
#
# We learn a logistic regression model on the validation set incorporating the learnt fair weights.
model_fair = LogisticRegression(max_iter=10000)
X_val = val_sds_transf.features
y_val = val_sds_transf.labels.flatten()
model_fair.fit(X_val, y_val, sample_weight=val_sds_transf.instance_weights)
# Apply fair model on test set.
#
# Note that the pre-processing intervention of the validation data happens in the model prediction since the model has been based on the weighting which was determined by the reweight transformed validation data.
test_sds_pred = test_sds.copy(deepcopy=True)
X_test = test_sds_pred.features
y_test = test_sds.labels
test_probs = model_fair.predict_proba(X_test)[:, 1]
test_pred = test_probs > 0.5
# Analyse fairness and accuracy
# +
mask = test.race_white == 1
bl_acc = accuracy(test.salary, bl_test_probs)
bl_dpd = demographic_parity_difference(
test.salary, bl_test_pred, sensitive_features=test.sex,
)
acc = accuracy(test.salary, test_probs)
dpd = demographic_parity_difference(
test.salary, test_pred, sensitive_features=test.sex,
)
print(f"Baseline model accuracy: {bl_acc:.3f}")
print(f"Model accuracy: {acc:.3f}")
print(f"Baseline demographic parity difference: {bl_dpd:.3f}")
print(f"Model demographic parity difference: {dpd:.3f}")
# -
dp_box = group_box_plots(
np.concatenate([bl_test_probs, test_probs]),
np.tile(test.sex.map({0: "Female", 1: "Male"}), 2),
groups=np.concatenate(
[
np.zeros_like(bl_test_probs),
np.ones_like(test_sds_pred.scores.flatten()),
]
),
group_names=["Baseline", "Kamiran-Calders"],
title="Score by sex for model and baseline",
xlabel="Score",
ylabel="Method",
)
dp_box
# + tags=["export"]
export_plot(dp_box, "kamiran-calders-dp.json")
|
src/notebooks/finance/interventions/kamiran_calders.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning Project - Traffic Signs Image Classification
import os, shutil
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
def save_hist_to_excel(filename):
df = pd.DataFrame(history.history)
df.to_excel(filename + '.xlsx')
# ## Overview of the training data
# +
#setting the training directory (the training images can be found there, already sorted by class into folders)
base_training_dir = "/Users/henriquevaz/NOVA IMS/YEAR 1/SPRING SEMESTER/DL/Project/my_notebooks/Data/Training/Final_Training/Images/"
#base_training_dir = "/Users/philippmetzger/Documents/GitHub/Deep_Learning_Project_Group_10/Data/GTSRB_Final_Training_Images/GTSRB/Final_Training/Images/"
#setting the directory where the selected training and validation images will be stored in
created_dir = "/Users/henriquevaz/NOVA IMS/YEAR 1/SPRING SEMESTER/DL/Project/my_notebooks/Data/Selected"
#created_dir = "/Users/philippmetzger/Documents/GitHub/Deep_Learning_Project_Group_10/Data/Selected"
#storing all the folder names that belong to the respective classes
all_classes = sorted(i for i in os.listdir(base_training_dir) if i.startswith("0"))
print("There are", len(all_classes), "different classes within the training data!")
# +
#getting the number of images within each class of the training data
amount_per_class = {}
for i in range(len(all_classes)):
directory = base_training_dir + "/" + all_classes[i]
amount_per_class[i] = len(sorted(i for i in os.listdir(directory) if i.startswith("0")))
amount_per_class_df = pd.DataFrame.from_dict(amount_per_class, orient='index').rename(columns={0:"amount"})
amount_per_class_df.index.name = 'class'
#remove the "#" of the following line to display the number of images within each class
#amount_per_class_df
# +
#displaying the number of images per class visually
plt.rcParams["figure.figsize"] = (20,8)
fig, ax = plt.subplots()
plt.bar(amount_per_class_df.index, amount_per_class_df.amount)
plt.title("Number of images per class", fontsize=20)
plt.xlabel('class', fontsize=18)
plt.ylabel('number of images', fontsize=18)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.xticks(amount_per_class_df.index)
plt.grid(axis="y")
plt.show()
# -
# As it can be seen in the bar chart, the dataset is highly unbalanced. Some classes have over 2000 instances, while others only have 210.
#displaying the 20 classes with the fewest images
amount_per_class_df.sort_values("amount").head(20).T
# Within each class, there are several groups of images, that belong together (these are basically all images of the very same traffic sign, that just differ in that they were made as a series while approaching the actual sign). These series of images shouldn't be splitted later when the data is split into training and validation sets, so some precautions might be useful:
# +
#checking the size(=amount of images) of these series and whether they are all having the same size
#running a for loop over all classes
for i in range(len(all_classes)):
directory = base_training_dir + "/" + all_classes[i]
#get the names of all images within a class
list_of_images = sorted(i for i in os.listdir(directory) if i.startswith("0"))
image_series = []
#store all the prefixes of the images (which correspond to the series they belong to)
for element in list_of_images:
image_series.append(element.split("_")[0])
#count the frequency of each prefix, which equals the size of each respective series
image_counts = pd.Series(image_series).value_counts().sort_index()
for element in image_counts.values.tolist():
if element != image_counts.values.tolist()[0]:
#this line will show if there is a series that contains not the same number of images than the others
print("There is a series whose number doesn't match the others in class", all_classes[i], ", with the prefix", image_counts.sort_values().index[0], "!\nIt only contains", element, "images.")
#apart from only one series, all others consist of exactly 30 images
print("All the other series of images contain exactly", image_counts.values.tolist()[0], "images!")
# -
# ## Split into training and validation datasets
# As already mentioned, it is important for the split into training and validation sets that the individual image series stay together. As a first approach, there will be 210 images used per class, as this number corresponds to the amount of images in the "smallest" class. This will fix the problem of imbalance in the dataset. Subsequently, the data will be splitted by a 5:2 (150:60) ratio into training and validation set, in order to guarantee that the series will stay together.
new_train_dir = "/Users/henriquevaz/NOVA IMS/YEAR 1/SPRING SEMESTER/DL/Project/my_notebooks/Data/Selected/train_all"
new_val_dir = "/Users/henriquevaz/NOVA IMS/YEAR 1/SPRING SEMESTER/DL/Project/my_notebooks/Data/Selected/val_all"
for i in range(len(all_classes)):
os.mkdir(new_train_dir + "/" + all_classes[i])
os.mkdir(new_val_dir + "/" + all_classes[i])
# +
import random
for i in range(len(all_classes)):
list_ = os.listdir(base_training_dir + "/" + all_classes[i])
random.shuffle(list_)
new_list = []
for element in list_:
if element.startswith("000"):
new_list.append(element)
new_list = sorted(new_list)
for image in range(len(new_list)):
#setting the cut off according to the next closest number in steps of 30 according to a 70:30 ratio
if len(new_list) == 210 or len(new_list) == 240:
cut = 60
elif len(new_list) == 270 or len(new_list) == 300 or len(new_list) == 330:
cut = 90
elif len(new_list) == 360 or len(new_list) == 390 or len(new_list) == 420:
cut = 120
elif len(new_list) == 450 or len(new_list) == 510 or len(new_list) == 540:
cut = 150
elif len(new_list) == 600 or len(new_list) == 630:
cut = 180
elif len(new_list) == 689:
cut = 210 - 1
elif len(new_list) == 780:
cut = 240
elif len(new_list) == 1110:
cut = 330
elif len(new_list) == 1200:
cut = 360
elif len(new_list) == 1320:
cut = 390
elif len(new_list) == 1410 or len(new_list) == 1440:
cut = 420
elif len(new_list) == 1470 or len(new_list) == 1500:
cut = 450
elif len(new_list) == 1860:
cut = 570
elif len(new_list) == 1980 or len(new_list) == 2010:
cut = 600
elif len(new_list) == 2070 or len(new_list) == 2100:
cut = 630
elif len(new_list) == 2160 or len(new_list) == 2220:
cut = 660
elif len(new_list) == 2250:
cut = 690
if image < len(new_list) - cut:
src = base_training_dir + "/" + all_classes[i] + "/" + new_list[image]
dst = new_train_dir + "/" + all_classes[i] + "/" + new_list[image]
shutil.copyfile(src, dst)
else:
src = base_training_dir + "/" + all_classes[i] + "/" + new_list[image]
dst = new_val_dir + "/" + all_classes[i] + "/" + new_list[image]
shutil.copyfile(src, dst)
# +
size_per_class_list = []
for i in range(len(all_classes)):
size_per_class_list.append([len(os.listdir(new_train_dir + "/" + all_classes[i])), len(os.listdir(new_val_dir + "/" + all_classes[i]))])
print("Class", i, ": training set size =", len(os.listdir(new_train_dir + "/" + all_classes[i])), "; validation set size:", len(os.listdir(new_val_dir + "/" + all_classes[i])))
# -
# Now, there should be exactly 150 images within the train folder of each class and 60 images within the validation folder. Having 43 different classes, this means that there are a total of 6,450 (150x43) training images and another 2,580 (60x43) validation images. A sanity check will show if this worked correctly:
# As it can be seen, it worked correctly!
# ## Data Preprocessing
# After running the upper part once, the notebook can be started from here from now on:
# +
#this cell is optional and the notebook should be runned from here once the upper part has been executed once
import os, shutil
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import tensorflow as tf
# -
train_dir = new_train_dir
validation_dir = new_val_dir
# +
from tensorflow.keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
train_dir,
target_size=(100,100),
batch_size=20,
class_mode="categorical",
color_mode="grayscale"
)
validation_generator = test_datagen.flow_from_directory(
validation_dir,
target_size=(100,100),
batch_size=20,
class_mode="categorical",
color_mode="grayscale"
)
# +
import sys
from PIL import Image
sys.modules['Image'] = Image
for data_batch, labels_batch in train_generator:
print('data batch shape:', data_batch.shape)
print('labels batch shape:', labels_batch.shape)
break
# -
# ## Draft of the model
# ### Base model
# +
from tensorflow.keras import layers
from tensorflow.keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (2, 2), activation='relu', input_shape=(100, 100, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.25))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.25))
model.add(layers.Conv2D(128, (4, 4), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.25))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense (43, activation='softmax'))
# -
model.summary()
# +
from tensorflow.keras import optimizers
#compilation
model.compile(loss="categorical_crossentropy", optimizer=optimizers.RMSprop(lr=1e-4), metrics=["acc"])
# -
history = model.fit(
train_generator,
epochs=50,
validation_data=validation_generator,
)
model.save('model_all_data_dropout_0_5_new_convlayer_grayscale_more_ropout_layers_up')
save_hist_to_excel('history_model_all_data_dropout_0_5_new_convlayer_grayscale_mored_ropout_layers_up')
# +
#Displaying curves of loss and accuracy during training
import matplotlib.pyplot as plt
acc = history.history["acc"]
val_acc = history.history["val_acc"]
loss = history.history["loss"]
val_loss = history.history["val_loss"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.legend()
plt.show()
plt.plot(epochs, loss, "bo", label="Training loss")
plt.plot(epochs, val_loss, "b", label="Validation loss")
plt.title("Training and validation loss")
plt.legend()
plt.show()
# -
# ### Reduce dropout rates
# +
from tensorflow.keras import layers
from tensorflow.keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (2, 2), activation='relu', input_shape=(100, 100, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.15))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.1))
model.add(layers.Conv2D(128, (4, 4), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.1))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dropout(0.3))
model.add(layers.Dense (43, activation='softmax'))
# -
model.summary()
# +
from tensorflow.keras import optimizers
#compilation
model.compile(loss="categorical_crossentropy", optimizer=optimizers.RMSprop(lr=1e-4), metrics=["acc"])
# -
history = model.fit(
train_generator,
epochs=50,
validation_data=validation_generator,
)
model.save('model_all_data_dropout_0_5_new_convlayer_grayscale_more_ropout_layers_down')
save_hist_to_excel('history_model_all_data_dropout_0_5_new_convlayer_grayscale_mored_ropout_layers_down')
# +
#Displaying curves of loss and accuracy during training
import matplotlib.pyplot as plt
acc = history.history["acc"]
val_acc = history.history["val_acc"]
loss = history.history["loss"]
val_loss = history.history["val_loss"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.legend()
plt.show()
plt.plot(epochs, loss, "bo", label="Training loss")
plt.plot(epochs, val_loss, "b", label="Validation loss")
plt.title("Training and validation loss")
plt.legend()
plt.show()
# -
# ### Remove one dropout layer
# +
from tensorflow.keras import layers
from tensorflow.keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (2, 2), activation='relu', input_shape=(100, 100, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.15))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.1))
model.add(layers.Conv2D(128, (4, 4), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dropout(0.5))
model.add(layers.Dense (43, activation='softmax'))
# -
model.summary()
# +
from tensorflow.keras import optimizers
#compilation
model.compile(loss="categorical_crossentropy", optimizer=optimizers.RMSprop(lr=1e-4), metrics=["acc"])
# -
history = model.fit(
train_generator,
epochs=70,
validation_data=validation_generator,
)
model.save('model_all_data_dropout_0_5_new_convlayer_grayscale_more_ropout_layers_down_removed1')
save_hist_to_excel('history_model_all_data_dropout_0_5_new_convlayer_grayscale_mored_ropout_layers_down_removed1')
# ### Reduce dropout rates again
# +
from tensorflow.keras import layers
from tensorflow.keras import models
model = models.Sequential()
model.add(layers.Conv2D(32, (2, 2), activation='relu', input_shape=(100, 100, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Dropout(0.1))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.05))
model.add(layers.Conv2D(128, (4, 4), activation='relu'))
model.add(layers.MaxPooling2D((2, 2) ))
model.add(layers.Dropout(0.05))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dropout(0.2))
model.add(layers.Dense (43, activation='softmax'))
model.summary()
# +
from tensorflow.keras import optimizers
#compilation
model.compile(loss="categorical_crossentropy", optimizer=optimizers.RMSprop(lr=1e-4), metrics=["acc"])
# -
history = model.fit(
train_generator,
epochs=150,
validation_data=validation_generator,
)
save_hist_to_excel('history_model_all_data_dropout_0_5_new_convlayer_grayscale_mored_ropout_layers_down_down')
# +
#Displaying curves of loss and accuracy during training
import matplotlib.pyplot as plt
acc = history.history["acc"]
val_acc = history.history["val_acc"]
loss = history.history["loss"]
val_loss = history.history["val_loss"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, "bo", label="Training acc")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.legend()
plt.show()
plt.plot(epochs, loss, "bo", label="Training loss")
plt.plot(epochs, val_loss, "b", label="Validation loss")
plt.title("Training and validation loss")
plt.legend()
plt.show()
# -
|
Additional (older) notebooks/2021-03-22_DL_Project_Group_10_HV_NewConvLayer_Grayscaling_AddingDropoutLayers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting up salinity results from SalishSeaCast using xarray
# This is testing code following a tutorial by Doug that is saved here:
# http://nbviewer.jupyter.org/urls/bitbucket.org/salishsea/analysis-doug/raw/tip/notebooks/SalishSeaCastVizDemo.ipynb
# ## First start with imports
import xarray as xr
import matplotlib.pyplot as plt
import numpy as np
import arrow
import cmocean
# %matplotlib inline
np.__version__
xr.__version__
# ## Load Data
ds = xr.open_dataset('https://salishsea.eos.ubc.ca/erddap/griddap/ubcSSg3DTracerFields1hV17-02')
ds.salinity
# ## Plot surface salinity
# Use "isel" to select location based on index [0 39]
SSS_t0_z0 = ds.salinity.isel(time=1, depth=1)
SSS_t0_z39= ds.salinity.isel(time=1, depth=39)
SSS_t0_z0
SSS_t0_z0.plot()
SSS_t0_z39.plot()
# Salinity is zero at the bottom because this isn't a sigma-coordinate model
SSS_t0_z30= ds.salinity.isel(time=1, depth=30)
SSS_t0_z30.plot()
# The nice thing about z-coordinate models is that they help you visualize topographic bariers more easily!
#
# ## Now let's try to make a pretty plot
# +
# This section includes code from Elise that makes use of a netcdf version
# of bathymetry. I haven't yet setup sshfs to allow for skookum access on
# local computer
# from salishsea_tools import evaltools as et, viz_tools
# import netCDF4 as nc
# grid = nc.Dataset('/data/vdo/MEOPAR/NEMO-forcing/grid/bathymetry_201702.nc')
# viz_tools.plot_coastline(ax, grid, coords = 'map')
# ax.set_ylim(48, 50.5)
# ax.legend()
# ax.set_xlim(-125.7, -122.5);
# +
# I have modified the above to work with bathy on ERDDAP
### need to install GSW #####
grid = xr.open_dataset('https://salishsea.eos.ubc.ca/erddap/griddap/ubcSSnBathymetryV17-02')
# This next bit is taken from Elise's code that she shared with us during group meeting
import salishsea_tools.evaltools as viz_tools
# import evaltools as viz_tools
fig, ax = plt.subplots(figsize = (6,6))
viz_tools.set_aspect(ax, coords = 'map')
viz_tools.plot_coastline(ax, grid, coords = 'map')
ax.set_ylim(48, 50.5)
ax.set_xlim(-125.7, -122.5);
ax.set_xlabel('Lon.')
ax.set_ylabel('Lat.')
# -
|
notebooks/learning/Plotting_Salinity.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Support Vector Classifier
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
style.use("ggplot")
from sklearn import svm
X = [1,5,1.5,8,1,9]
Y = [2,8,1.8,8,0.6,11]
plt.scatter(X,Y)
plt.show()
data = np.array(list(zip(X,Y)))
data
target = [0, 1, 0, 1, 0, 1]
classifier = svm.SVC(kernel="linear", C = 1.0)
classifier.fit(data, target)
p = np.array([10.32, 12.67]).reshape(1,2)
print(p)
classifier.predict(p)
# * Modelo: w0 . x + w1 . y + e = 0
# * Ecuación del hiperplano en 2D: y = a . x + b
w = classifier.coef_[0]
w
a = -w[0]/w[1]
a
b = - classifier.intercept_[0]/w[1]
b
xx = np.linspace(0,10)
yy = a * xx + b
plt.plot(xx, yy, 'k-', label = "Hiperplano de separación")
plt.scatter(X, Y, c = target)
plt.legend()
plt.plot()
|
notebooks/T8 - 1 - SVM - Linear SVC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Examples of the BioSCRAPE package
# ## Biocircuit Stochastic Simulation of Single Cell Reactions and Parameter Estimation
#
# The purpose of this Python notebook is twofold.
#
# 1. The first is to serve as a quick start guide where you should be able to get started with the package by simply looking at the examples here and copying them to your liking.
#
# 2. The second is as a unit testing replacement. It is hard to unit test stochastic algorithms as the output may not (and should not) be the same thing every time. Therefore, instead, if all the examples included below work well, then you can assume that the package installed correctly and is working fine.
#
# Before, getting started, we start by doing some basic plotting configuration and importing the numpy library. Advanced users can modify this to their liking.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
# #%config InlineBackend.figure_f.ormats=['svg']
mpl.rc('axes', prop_cycle=(mpl.cycler('color', ['r', 'k', 'b','g','y','m','c']) ))
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
import numpy as np
# -
# # 1. A Simple Model of Gene Expression
#
# We start with a simple model of gene expression that only contains 4 reactions: transcription, translation, mRNA degradation, and protein degradation. The rate of transcription is constant, and the rates of translation, mRNA degradation, and protein degradation are linear in mRNA, protein, and protein respectively. This model is included in models/gene_expression_with_delay.xml, where you can look at it. The model text is included below as well.
#
# <model>
# <reaction text="--" after="--mRNA">
# <propensity type="massaction" k="beta" species="" />
# <delay type="fixed" delay="tx_delay" />
# </reaction>
#
# <reaction text="mRNA--" after="--">
# <propensity type="massaction" k="delta_m" species="mRNA" />
# <delay type="none" />
# </reaction>
#
# <reaction text="--" after="--protein">
# <propensity type="massaction" k="k_tl" species="mRNA" />
# <delay type="gamma" k="tx_k" theta="tx_theta" />
# </reaction>
#
# <reaction text="protein--">
# <propensity type="massaction" k="delta_p" species="protein" />
# <delay type="none" />
# </reaction>
#
#
# <parameter name="beta" value="2.0" />
# <parameter name="delta_m" value="0.2" />
# <parameter name="k_tl" value="5.0" />
# <parameter name="delta_p" value="0.05" />
# <parameter name="tx_delay" value="10" />
# <parameter name="tl_delay" value="10" />
# <parameter name="tx_k" value="2" />
# <parameter name="tx_theta" value="5" />
#
# <species name="mRNA" value="0" />
# <species name="protein" value="0" />
# </model>
#
# From this code, you can see that there are 4 reactions. Each reaction specifies a text field which says what reactants go to what products. The after field specifies the delayed part of the reaction. Each reaction has a propensity and a delay. At the bottom, all the parameter values and initial species values are specified.
#
# Simulating this model with a regular DeterministicSimulator or a regular SSASimulator will result in the specified delays being ignored and assumed to be zero. In order for the delay to matter, you must do a simulation with a stochastic delay simulator.
#
# Let's start by trying out the model without delay in both the deterministic and stochastic case.
# ## 1a. Simple Model of Gene Expression without Delay
#
# Below, we load in the simple model of gene expression. We then simulate it determinstically and stochastically. The parameters have been chosen so that the mean mRNA level should be 10 and the mean protein level should be 1000.
#
# Thus, the deterministic simulation should quickly go to a steady state of mRNA = 10 and protein = 1000 and the stochastic simulation should bounce around that number.
# +
# Code for simple gene expression without delay
# Import relevant types
from bioscrape.types import Model
from bioscrape.simulator import DeterministicSimulator, SSASimulator
from bioscrape.simulator import ModelCSimInterface, SafeModelCSimInterface
import numpy as np
import pylab as plt
import time as pytime
# Load the model by creating a model with the file name containing the model
m = Model('models/gene_expression_with_delay.xml', input_printout = False)
# Expose the model's core characteristics for simulation. (i.e. stoichiometry,
# delays, and propensities)
s = ModelCSimInterface(m)
# This function uses sparsity to further optimize the speed of deterministic
# simulations. You must call it before doing deterministic simulations.
s.py_prep_deterministic_simulation()
# Set up our desired timepoints for which to simulate.
# Must match with initial time.
timepoints = np.linspace(0,10000,10000)
# Create a DeterministicSimulator as well as an SSASimulator
ssa_simulator = SSASimulator()
det_simulator = DeterministicSimulator()
# Set the initial simulation time
s.py_set_initial_time(0)
# This function uses sparsity to further optimize the speed of deterministic
# simulations. You must call it before doing deterministic simulations.
s.py_prep_deterministic_simulation()
# Simulate the model with both simulators for the desired timepoints
stoch_result = ssa_simulator.py_simulate(s,timepoints)
det_result = det_simulator.py_simulate(s,timepoints)
# Process the simulation output.
# py_get_result() returns a numpy 2d array of timepoints x species.
# Each row is one time point and each column is a species.
stoch_sim_output = stoch_result.py_get_result()
# Get the indices for each species of interest
# From the model, we can recover which column corresponds to which species, so
# we then know which column of the result array is which species.
X_ind = m.get_species_index('X')
# Plot the mRNA levels over time for both deterministic and stochastic simulation
plt.plot(timepoints,stoch_sim_output[:,X_ind])
plt.xlabel('Time')
plt.ylabel('X')
plt.show()
# -
# ### Results
#
# In the above plots, ideally you see for mRNA a smooth line going to a steady state of 10 with another line from the stochastic simulation bouncing around.
#
# For the protein, you should see something similar but with the value being 1000 at steady state.
# ## 1b. Simulate Gene Expression Model with Delay
#
# We can now try and simulate the model from before that we've already loaded while accounting for delays. This time, we will use a delay SSA simulator. From the model, there are only delays for transcription and translation. The transcription delay was specified as
#
# <delay type="fixed" delay="tx_delay" />
#
# In this term, the delay is of a fixed type meaning it's always a constant, and the length of the delay is given in the parameter tx_delay.
#
# The translation delay is specified as
#
# <delay type="gamma" k="tx_k" theta="tx_theta" />
#
# In this term, the delay is specified by a gamma distribution, which means the delay time for this reaction to happen is a gamma random variable with parameters $k$ and $\theta$, where those parameters are specified by tx_k and tx_theta.
#
# In addition to specifying the delays, we also need a DelayQueue in order to do delay simulations. The delay queue is a data structure that keeps track of what future reactions have been queued up to occur. Different implementations of the DelayQueue used can have severe tradeoffs in speed and accuracy.
#
#
# +
# Simulate the simple gene expression model WITH delay.
# import a couple of additional types that we need
from bioscrape.simulator import ArrayDelayQueue
from bioscrape.simulator import DelaySSASimulator
# Create a delay queue with setup_queue(num_reactions, num_timepoints, dt)
# so this delay queue will go up to 1500 * 0.01 = 15 time units in the future.
# You want to pick the delay queue resolution to be small for accuracy, and then
# have enough time points to capture the maximum length delay that could possibly
# occur.
q = ArrayDelayQueue.setup_queue(4,1500,0.01)
# Like before when we created an SSA simulator, now we create a DelaySSASimulator
delay_simulator = DelaySSASimulator()
# Simulate just like before, but now we need to pass the DelayQueue q as an
# extra argument. The delayqueue is part of the initial state as well, as any
# reactions already on the queue will occur.
# In this case, however, the queue is empty to begin with.
answer = delay_simulator.py_delay_simulate(s,q,timepoints)
# Recover the state trajctory from the simulation.
state = answer.py_get_result()
mrna_ind = m.get_species_index('mRNA')
prot_ind = m.get_species_index('protein')
# Plot the mRNA
plt.plot(timepoints,state[:,mrna_ind])
plt.xlim((0,120))
plt.xlabel('Time')
plt.ylabel('mRNA')
# Plot the Protein
plt.figure()
plt.plot(timepoints,state[:,prot_ind])
plt.xlim((0,120))
plt.xlabel('Time')
plt.ylabel('Protein')
# -
# ### Results
#
# In this case, the results should be a delay of 10 before transcription begins and then a final mRNA level of around 10, as well as a delay of about 20 before protein appears with a final protein level of around 1000. The translation delay is gamma distributed, so the protein turns on somewhat smoothly, while the mRNA transcription delay is fixed at 10 minutes, so the mRNA turns on sharply.
#
# We can then move on to the next example, which is testing cell lineages and doing simulations with volume.
#
# ## 1c. Simulating Lineages of Cells with Growth and Division
#
# Now we can add cell growth and division to the picture. When we simulate cell growth and division, we need to tell the simulator how the cells actually grow and divide. We do this by providing a volume model as well as a partitioning model for cell division. The volume model tells the cell how to grow over time and when to divide. The growth and moment of division can be deterministic or stochastic.
#
# The partitioning model tells the simulator how to split up the contents of the cells when the cells divide. For example, molecules with high counts should probably be split up binomially, while the genome should probably be divided equally.
# +
from bioscrape.types import StochasticTimeThresholdVolume
from bioscrape.simulator import VolumeSSASimulator, PerfectBinomialVolumeSplitter
from bioscrape.simulator import py_simulate_cell_lineage
# Reset the state and time to 0 for the model.
s.py_set_dt(0.01) # this is the time resolution at which the volume changes.
s.py_set_initial_time(0)
s.py_set_initial_state(np.array([0.0,0.0])) # the state must be of type double
# Specify the volume model to use. In this case, grow without noise and divide
# at a mean 33 minute division time, with a division volume of 2.
# The 0.05 is noise parameter that says how much noise to put into the division
# time.
v = StochasticTimeThresholdVolume(33,2,0.05)
# We need to initialize the volume with the initial time and volume, which we do
# here. The first two arguments are the state and parameter vectors, which don't
# matter here, so we pass in a junk argument for those.
junk = np.empty(1,)
v.py_initialize(junk, junk,0.0,1.0)
# Set up timepoints to simulate for.
timepoints = np.linspace(0,300,1000)
# Create a volume SSA simulator now.
vsim = VolumeSSASimulator()
# Need to create a splitter that will partition divided cells. This one splits
# the volume exactly 50/50 and partitions all species binomially with p = 0.5
vsplit = PerfectBinomialVolumeSplitter()
# Use the simulate cell lineage function to simulate the lineage.
# The arguments are the model s, volume model v, timepoints, volume simulator,
# and volume splitter
# l is a lineage object
l = py_simulate_cell_lineage(s,v,timepoints,vsim,vsplit)
# Go through the lineage object l which has l.py_size() entries
# Each entry is called a schnitz and each schnitz has a set of timepoints
# as well as a simulation trajectory and a volume trajectory. You can plot
# the protein concentration over time for each schnitz.
plt.figure()
for i in range(l.py_size()):
sch = l.py_get_schnitz(i)
plt.plot(sch.py_get_time(),
sch.py_get_data()[:,prot_ind] / sch.py_get_volume())
plt.title('Protein Concentration Over Time (nM)')
plt.xlabel('Time (min)')
plt.ylabel('Concentration (nm)')
# Can plot mRNA as well
plt.figure()
for i in range(l.py_size()):
sch = l.py_get_schnitz(i)
plt.plot(sch.py_get_time(),
sch.py_get_data()[:,mrna_ind] / sch.py_get_volume())
plt.title('mRNA Concentration Over Time (nM)')
plt.xlabel('Time (min)')
plt.ylabel('Concentration (nm)')
# -
# ### Results
#
# In this case, you should see the expression go up from zero and then as cells grow and divide, there will be many protein trajectories all hanging out between around 600 to 800 nm. For mRNA, the average value should be around 10 nM.
#
# Next, we will do some simulations with cell division and delay at the same time.
# ## 1d. Gene Expression with Delay and Cell Division
#
# Here, we simulate the same model again but incorporate both cell division and delay. This example is similar to 1c.
# +
from bioscrape.simulator import PerfectBinomialDelayVolumeSplitter
from bioscrape.simulator import DelayVolumeSSASimulator
from bioscrape.simulator import py_simulate_delay_cell_lineage
m = Model('models/gene_expression_with_delay.xml')
# Set the delay to be bigger than 1 cell cycle though.
m.set_params({"tx_delay": 33.0})
final_time = 180.0
s = ModelCSimInterface(m)
s.py_set_dt(0.01)
s.py_set_initial_time(0.0)
# Specify a delay queue.
q = ArrayDelayQueue.setup_queue(4,10000,0.01)
# Specify the volume model to use. In this case, grow without noise and divide
# at approximately 33 minutes
v = StochasticTimeThresholdVolume(33,2,0.05)
junk = np.zeros(10,)
v.py_initialize(junk, junk, 0.0, 1.0)
dvsplit = PerfectBinomialDelayVolumeSplitter()
dvsim = DelayVolumeSSASimulator()
timepoints = np.linspace(0,final_time,final_time)
l = py_simulate_delay_cell_lineage(s,q,v,timepoints,dvsim,dvsplit)
fig = plt.figure(0)
ax1 = fig.add_subplot(211)
ax2 = fig.add_subplot(212)
for i in range(l.py_size()):
sch = l.py_get_schnitz(i)
ax1.plot(sch.py_get_time(), sch.py_get_data()[:,mrna_ind] / sch.py_get_volume())
ax2.plot(sch.py_get_time(), sch.py_get_data()[:,prot_ind] / sch.py_get_volume())
ax1.set_xlabel('Time')
ax1.set_ylabel('mRNA Concentration')
ax2.set_xlabel('Time')
ax2.set_ylabel('Protein Concentration')
# -
# ### Results
#
# For this example, the mRNA average should be around 5 after coming on around 30 minutes, and the protein level should come on around 40 minutes and then end up around 300.
#
# The reason the steady state mRNA and protein levels are lower than in the lineage simulation with no delay is that when there is delay in production, the amount of mRNA/protein appearing at a given time is actually proportional to the number of cells that existed a long time ago, which will be only a fraction of the number of cells there are currently. This makes the effective instantaneous arrival rate of new mRNA's and proteins smaller per cell, which leads to a lower steady state.
# # SBML Compatibility
#
# The next cell imports a model from an SBML file and then simulates it using a deterministic simulation. There are limitations to SBML compatibility.
#
# 1. Cannot support delays or events when reading in SBML files. Events will be ignored and a warning will be printed out.
# 2. SBML reaction rates must be in a format such that when the reaction rates are converted to a string formula, sympy must be able to parse the formula. This will work fine for usual PEMDAS rates. This will fail for complex function definitions and things like that.
# 3. Species will be initialized to their initialAmount field when it is nonzero. If the initialAmount is zero, then the initialConcentration will be used instead.
# 4. Multiple compartments or anything related to having compartments will not be supported. No warnings will be provided for this.
# 5. Assignment rules are supported, but any other type of rule will be ignored and an associated warning will be printed out.
# 6. Parameter names must start with a letter and be alphanumeric, same for species names. Furthermore, log, exp, abs, heaviside, and other associated keywords for functions are not allowed to be variable names. When in doubt, just pick something else :)
#
# Below, we first plot out the simulation results for an SBML model where a species X0 goes to a final species X1 through an enymatic process.
# +
import bioscrape
m = bioscrape.types.read_model_from_sbml('models/sbml_test.xml')
s = bioscrape.simulator.ModelCSimInterface(m)
s.py_prep_deterministic_simulation()
s.py_set_initial_time(0)
sim = bioscrape.simulator.DeterministicSimulator()
timepoints = np.linspace(0,100,1000)
result = sim.py_simulate(s,timepoints)
plt.plot(timepoints,result.py_get_result())
plt.legend(m.get_species_list())
# -
# ## Deterministic and Stochastic Simulation of the Repressilator
#
# We plot out the repressilator model found <a href="http://www.ebi.ac.uk/biomodels-main/BIOMD0000000012">here</a>. This model generates oscillations as expected. Highlighting the utility of this package, we then with a single line of code switch to a stochastic simulation and note that the amplitudes of each burst become noisy.
# +
# Repressilator deterministic example
import bioscrape
plt.figure()
m = bioscrape.types.read_model_from_sbml('models/repressilator_sbml.xml')
s = bioscrape.simulator.ModelCSimInterface(m)
s.py_prep_deterministic_simulation()
s.py_set_initial_time(0)
sim = bioscrape.simulator.DeterministicSimulator()
timepoints = np.linspace(0,1000,10000)
result = sim.py_simulate(s,timepoints)
plt.plot(timepoints,result.py_get_result())
plt.legend(m.get_species_list())
plt.title('Repressilator Model')
plt.xlabel('Time')
plt.ylabel('Amount')
# -
# The plot above should show deterministic oscillations. If we want to switch to stochastic simulation, all we need to do is switch the type of simulator we are using to the stochastic simulator. This can be done with the single following line.
sim = bioscrape.simulator.SSASimulator()
# Now, we can run exactly the same code as above to generate a stochastic simulation trace.
# +
s.py_set_initial_time(0)
timepoints = np.linspace(0,1000,10000)
result = sim.py_simulate(s,timepoints)
plt.plot(timepoints,result.py_get_result())
plt.legend(m.get_species_list())
plt.title('Repressilator Model')
plt.xlabel('Time')
plt.ylabel('Amount')
# -
# Note that now the oscillations are non regular and stochastic.
# +
from bioscrape.simulator import PerfectBinomialDelayVolumeSplitter
from bioscrape.simulator import DelayVolumeSSASimulator
from bioscrape.simulator import ArrayDelayQueue
from bioscrape.simulator import DelaySSASimulator
from bioscrape.types import StochasticTimeThresholdVolume
from bioscrape.simulator import VolumeSSASimulator, PerfectBinomialVolumeSplitter
from bioscrape.simulator import py_simulate_cell_lineage
from bioscrape.simulator import ModelCSimInterface
from bioscrape.types import Model
import numpy as np
import pylab as plt
m = bioscrape.types.read_model_from_sbml('models/repressilator_sbml.xml')
# Set the delay to be bigger than 1 cell cycle though.
m.set_params({"tx_delay": 33.0})
final_time = 180.0
s = ModelCSimInterface(m)
s.py_set_dt(0.01)
s.py_set_initial_time(0.0)
# Specify a delay queue.
q = ArrayDelayQueue.setup_queue(4,10000,0.01)
# Specify the volume model to use. In this case, grow without noise and divide
# at approximately 33 minutes
v = StochasticTimeThresholdVolume(33,2,0.05)
junk = np.zeros(10,)
v.py_initialize(junk, junk, 0.0, 1.0)
dvsplit = PerfectBinomialDelayVolumeSplitter()
dvsim = DelayVolumeSSASimulator()
timepoints = np.linspace(0,final_time,final_time)
l = py_simulate_delay_cell_lineage(s,q,v,timepoints,dvsim,dvsplit)
fig = plt.figure(0)
ax1 = fig.add_subplot(211)
ax2 = fig.add_subplot(212)
mrna_ind = m.get_species_index('mRNA')
prot_ind = m.get_species_index('protein')
for i in range(l.py_size()):
sch = l.py_get_schnitz(i)
ax1.plot(sch.py_get_time(), sch.py_get_data()[:,mrna_ind] / sch.py_get_volume())
ax2.plot(sch.py_get_time(), sch.py_get_data()[:,prot_ind] / sch.py_get_volume())
ax1.set_xlabel('Time')
ax1.set_ylabel('mRNA Concentration')
ax2.set_xlabel('Time')
ax2.set_ylabel('Protein Concentration')
# -
|
examples/Advanced Examples - Direct Simulator Instantiation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Throughout this document the following imports and styles are used.
# %pylab inline
# # Goal
# In this document we will be testing a multiple exponential moving average crossover strategy. We will also create an entry position calculated with a Fibonacci between the point of the crossover and the low/high of the candle.
# ## Issues with our model
# This model still has a few unsolved issues:
#
# 1. We have created a target price for which the older should be opened. It is however opened on the close price for the candle where the target price is hit with the high or low. The assumption is that it gives minimal effect.
#
# 2. Short position P/L is perhaps not calculated correctly.
# # Imports
# ## General libraries
# These are the general Python libraries that are used.
from datetime import datetime, timedelta
import numpy as np
import pandas as pd
# ## Custom libraries
# These are the custom libraries that hold the framework to perform and analyze a backtest.
import poloniex as plnx
import ta_lib as ta
import signals as sg
import backtest as bt
# # Technical Analysis for Trading Signals
# To run a backtest trading simulation we will need to set it up with parameters. The parameters will be explained per section.
# ## Chart data
# We are using data from http://www.poloniex.com API. Our test is done on the `USDT_BTC` pair..
pair = 'USDT_BTC'
timeframe = 120 * 60
end = datetime.utcnow()
start = end - timedelta(days=365)
chart = plnx.get_chart(pair, timeframe, start, end)
# ## Technical Indicators
# We will be using the following technical indicators to generate signals:
ta.ema(chart, 8)
ta.ema(chart, 13)
ta.ema(chart, 21)
ta.ema(chart, 55)
chart[50:60]
# ## Signals
# We will be using multiple EMA's to generate crossover signals.
chart['open_long'] = chart['open_short'] = chart['close_long'] = chart['close_short'] = 0
def getFib(crossover_price, candle_high_low, lvl): return crossover_price-(crossover_price-candle_high_low)*lvl
# +
head = 56
ema1 = 'ema8'
ema2 = 'ema13'
ema3 = 'ema21'
ema4 = 'ema55'
chart['crossover'] = chart['target_price'] = chart['target_low'] = chart['dbg_long'] = chart['dbg_short'] = 0
state = 'none'
order_filled = False
for i in range(head, len(chart.index)):
p1 = chart.loc[i, ema1] - chart.loc[i, ema4]
p2 = chart.loc[i, ema2] - chart.loc[i, ema4]
p3 = chart.loc[i, ema3] - chart.loc[i, ema4]
if(p1 > 0 and p2 > 0 and p3 > 0): chart.loc[i, 'crossover'] = 1
if(p2 < 0 and p2 < 0 and p3 < 0): chart.loc[i, 'crossover'] = -1
prev = chart.loc[i-1, 'crossover']
current = chart.loc[i, 'crossover']
chart.loc[i, 'target_price'] = chart.loc[i-1, 'target_price']
if (prev == -1 or prev == 0) and current == 1:
if state == 'short' and order_filled:
chart.loc[i, 'close_short'] = 1
state = 'long'
chart.loc[i, 'target_price'] = getFib(chart.loc[i, ema4], chart.loc[i, 'low'],.702)
chart.loc[i, 'dbg_long'] = 1
order_filled = False
if (prev == 1 or prev == 0) and current == -1:
if state == 'long' and order_filled:
chart.loc[i, 'close_long'] = 1
state = 'short'
chart.loc[i, 'target_price'] = getFib(chart.loc[i, ema4], chart.loc[i, 'high'],.702)
chart.loc[i, 'dbg_short'] = 1
order_filled = False
if state == 'long' and not order_filled:
if chart.loc[i, 'target_price'] > chart.loc[i, 'low']:
chart.loc[i, 'open_long'] = 1
order_filled = True
if state == 'short' and not order_filled:
if chart.loc[i, 'target_price'] < chart.loc[i, 'high']:
chart.loc[i, 'open_short'] = 1
order_filled = True
# -
# ## Plotting strategy indicators
# If we plot all our technical indicators in a chart.
plt.figure(figsize(120,12))
plot(chart['high'], c='lime', lw=1, alpha=1)
plot(chart['low'], c='red', lw=1, alpha=1)
plot(chart['ema55'], c='magenta')
plot(chart.loc[chart['target_price'] > 0]['target_price'], c='b', ls='dashed')
dbg_long = chart.loc[chart['dbg_long'] > 0]
dbg_short = chart.loc[chart['dbg_short'] > 0]
scatter(dbg_long.index, dbg_long['close'], c='lime', lw=20, marker='x', alpha=0.5)
scatter(dbg_short.index, dbg_short['close'], c='red', lw=20, marker='x', alpha=0.5)
legend(['High', 'Low', 'EMA55', 'Target price', 'Long', 'Short'])
grid()
# ### Close and EMA55 distance indicator
plt.figure(figsize(20,6))
chart['dist'] = chart['close'] - chart['ema55']
plot(chart['dist'])
fill_between(range(len(chart.index)), chart['dist'], alpha=0.25)
axhline(0,c='black')
grid()
# ### EMA crossover indicator
plot(chart['crossover'])
fill_between(range(len(chart.index)), chart['crossover'], alpha=0.25)
axhline(0, ls='dashed', c='black')
grid();
# ### Signal indicator
plt.step(range(len(chart.index)), chart['open_long'],c='lime')
plt.step(range(len(chart.index)), chart['open_short'],c='r')
plt.step(range(len(chart.index)), chart['close_long'], c='lime', linestyle='dashed')
plt.step(range(len(chart.index)), chart['close_short'],c='r', linestyle='dashed')
plt.step(range(len(chart.index)), np.zeros(len(chart.index)))
plt.xlabel('t')
plt.ylabel('Signal On/Off')
plt.title('Signals')
plt.grid();
# # Simulation
# We will start with $\$1000$ and use the following parameters:
sim = bt.TradingSim(pd.DataFrame(chart))
sim.balance = 1000
sim.lot_size = 0.1
sim.fee_percentage = 0.00050
sim.leverage = 10
sim.stop_loss_type = 'limit'
sim.stop_loss_percentage = 0.1
sim.run()
print('Final balance: {}'.format(sim.balance))
# ## Optimal stop loss percentage
# +
results = {}
for slp in range(21):
slp /= 100
print('Simulating for {}'.format(slp))
sim = bt.TradingSim(pd.DataFrame(chart))
sim.balance = 1000
sim.lot_size = 0.1
sim.fee_percentage = 0.00050
sim.leverage = 10
sim.stop_loss_type = 'limit'
sim.stop_loss_percentage = slp
sim.run()
results[slp] = sim.balance
x,y = list(results.keys()), list(results.values())
plt.figure(figsize=(5,5))
plot(x,y)
plt.show()
print('Optimal solution at: {}'.format(x[y.index(max(y))]))
# -
# # Performance analysis
# +
analysis = bt.AnalyzeSim(sim)
analysis.sim.df['max_balance'] = 0
analysis.sim.df.loc[0, 'max_balance'] = analysis.sim.df.loc[0, 'balance']
for i in range(1, len(analysis.sim.df.index)):
analysis.sim.df.loc[i, 'max_balance'] = max(analysis.sim.df.loc[i-1, 'max_balance'], analysis.sim.df.loc[i, 'balance'])
p = analysis.equity_curve()
p.plot(analysis.sim.df['max_balance'], c='black', ls='dashed');
# -
plt.figure(figsize=(20,5))
drawdown = analysis.sim.df['max_balance'] - analysis.sim.df['balance']
plot(drawdown, lw=2)
fill_between(range(len(analysis.sim.df.index)), drawdown, alpha=0.25)
plt.show()
max_drawdown = max(drawdown)
print('Max drawdown: {:.2f}'.format(1-max_drawdown / analysis.sim.df.loc[drawdown.loc[drawdown == max_drawdown].index[0], 'balance']))
print('There are {} trades.'.format(len(analysis.trades)))
good_trades = analysis.trades.loc[analysis.trades['pl'] > 0]
bad_trades = analysis.trades.loc[analysis.trades['pl'] < 0]
print('Good trades: {}'.format(len(good_trades)))
print('Bad trades: {}'.format(len(bad_trades)))
print('Win ratio is {:.2f}%.'.format(len(good_trades)/(len(good_trades)+len(bad_trades))*100))
analysis.roe()
analysis.paid_fees()
analysis.trades
print('There are {} stopped trades.'.format(len(analysis.trades.loc[analysis.trades['state'] == 'stopped'])))
analysis.trades.loc[analysis.trades['state'] == 'stopped']
# # Fibonacci levels
# We can use this to calculate the Fibonacci level between two points.
a = 9174 # crossover price EMA55
b = 8135 # high of crossover candle if long, low of crossover if short
r = a-b
f5 = a-r*0.5
f618 = a-r*0.618
f702 = a-r*0.702
f702
# Actually it can be done for any level.
def getFib(crossover_price, candle_high_low, lvl): return crossover_price-(crossover_price-candle_high_low)*lvl
# We are using `0.702` which is a mid-level.
getFib(9174, 8135, .702) # Long
getFib(8135, 9174, .702) # Short
# Generating the Fibonacci sequence.
def fibo(n):
a = 1
b = 1
print(1)
print(1)
for _ in range(n):
c=a+b
a=b
b=c
yield c
for n in fibo(25): print(n)
|
Crypto/Strategy Backtesting/.ipynb_checkpoints/Multi Exponential Moving Average Strategy with Fibonacci Retrace Order Filling-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Wine Quality."
# ### _"Quality ratings of Portuguese white wines" (Classification task)._
# ## Table of Contents
#
#
# ## Part 0: Introduction
#
# ### Overview
# The dataset that's we see here contains 12 columns and 4898 entries of data about Portuguese white wines.
#
# **Метаданные:**
#
# * **fixed acidity**
#
# * **volatile acidity**
#
# * **citric acid**
#
# * **residual sugar**
#
# * **chlorides**
#
# * **free sulfur dioxide**
#
# * **total sulfur dioxide**
#
# * **density**
#
# * **pH**
#
# * **sulphates**
#
# * **alcohol**
#
# * **quality** - score between 3 and 9
#
#
# ### Questions:
#
# Predict which wines are 'Good/1' and 'Not Good/0' (use binary classification; check balance of classes; calculate perdictions; choose the best model)
#
#
# ## [Part 1: Import, Load Data](#Part-1:-Import,-Load-Data.)
# * ### Import libraries, Read data from ‘.csv’ file
#
# ## [Part 2: Exploratory Data Analysis](#Part-2:-Exploratory-Data-Analysis.)
# * ### Info, Head, Describe
# * ### Encoding 'quality' attribute
# * ### 'quality' attribute value counts and visualisation
# * ### Resampling of an imbalanced dataset
# * ### Random under-sampling of an imbalanced dataset
# * ### Random over-sampling of an imbalanced dataset
# * ### Initialisation of target
# * ### Drop column 'quality'
#
# ## [Part 3: Data Wrangling and Transformation](#Part-3:-Data-Wrangling-and-Transformation.)
# * ### StandardScaler
# * ### Creating datasets for ML part
# * ### 'Train\Test' splitting method
#
# ## [Part 4: Machine Learning](#Part-4:-Machine-Learning.)
# * ### Build, train and evaluate models without hyperparameters
# * #### Logistic Regression, K-Nearest Neighbors, Decision Trees
# * #### Classification report
# * #### Confusion Matrix
# * #### ROC-AUC score
# * ### Build, train and evaluate models with hyperparameters
# * #### Logistic Regression, K-Nearest Neighbors, Decision Trees
# * #### Classification report
# * #### Confusion Matrix
# * #### ROC-AUC score
#
# ## [Conclusion](#Conclusion.)
#
#
# ## Part 1: Import, Load Data.
# * ### Import libraries
# +
# import standard libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
from scipy.stats import norm
# %matplotlib inline
sns.set()
import sklearn.metrics as metrics
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, roc_auc_score
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
import warnings
warnings.filterwarnings('ignore')
# -
# * ### Read data from ‘.csv’ file
# read data from '.csv' file
dataset = pd.read_csv('winequality.csv')
# ## Part 2: Exploratory Data Analysis.
# * ### Info
# print the full summary of the dataset
dataset.info()
# Dataset consist of 4898 rows and 12 columns.
#
# has 2 datatypes: float64(11), integer64(1)
#
# has no missing values
# * ### Head
# preview of the first 5 lines of the loaded data
dataset.head()
# * ### Describe
dataset.describe()
# Задача классифицировать вина на плохие и хорошие. Атрибута "Y" подобного нет, четкого отета нет. Но есть атрибут "quality", который отечает за оценку качеста вина и из, которого мы можем создать атрибут "Y" с ответом для обучения модели. Атрибут "quality" содержит значения от 3 до 9 (посмотрела в CSV файле), где 3 "Not Good" и 9 - "Good", соответстенно чем выше число тем качество вина выше.
# * ### Encoding 'quality' attribute
# lambda function; wine quality from 3-6 == 0, from 7-9 == 1.
dataset['quality'] = dataset.quality.apply(lambda q: 0 if q <=6 else 1)
# preview of the first 5 lines of the loaded data
dataset.head()
# * ### 'quality' attribute value counts and visualisation
print ('Not good wine', round(dataset['quality'].value_counts()[0]/len(dataset) * 100,2), '% of the dataset')
print ('Good wine', round(dataset['quality'].value_counts()[1]/len(dataset) * 100,2), '% of the dataset')
dataset['quality'].value_counts()
# visualisation plotby bar
dataset['quality'].value_counts().plot(x = dataset['quality'], kind='bar')
# visualisation plot by pie
dataset['quality'].value_counts().plot(x = dataset['quality'], kind = 'pie')
# There are 78.36 % of "Not good" quality wines an only 21.64 % of "Good" quality wines in our dataset. So we can see that the dataset is imbalanced.
# * ### Resampling of an imbalanced dataset
# +
# class count
# divide by class
# -
# * ### Random under-sampling of an imbalanced dataset
# * ### Random over-sampling of an imbalanced dataset
# * ### Initialisation of target
# * ### Drop column 'quality'
# ## Part 3: Data Wrangling and Transformation.
# * ### StandardScaler
# StandardScaler
# * ### Creating datasets for ML part
# +
# set 'X' for features' and y' for the target ('quality').
# for under-sampling dataset
# for over-sampling dataset
# -
# preview of the first 5 lines of the loaded data
# * ### 'Train\Test' split
# apply 'Train\Test' splitting method
# print shape of X_train and y_train
# print shape of X_test and y_test
# ## Part 4: Machine Learning.
# * ### Build, train and evaluate models without hyperparameters
# * Logistic Regression
# * K-Nearest Neighbors
# * Decision Trees
#
# +
# Logistic Regression
# K-Nearest Neighbors
# Decision Tree
# -
# * ### Classification report
# * ### Confusion matrix
# * ### ROC-AUC score
# * ### Build, train and evaluate models with hyperparameters
# +
# Logistic Regression
# K-Nearest Neighbors
# Decision Tree
# -
# print the best hyper parameters set
# * ### Classification report
# * ### Confusion matrix
# +
# confusion matrix of DT model
# visualisation
# -
# * ### ROC-AUC score
# ## Conclusion.
# submission of .csv file with predictions
|
ML-101 Modules/Module 03/Lesson 02/Practice 2/.ipynb_checkpoints/Winequality - Homework-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Digit Recognition
# # Introduction
# In this notebook I will explain the how my Python script "digitrec.py" works and its preformance
# ## Imports
# +
import gzip
import keras as kr
import numpy as np
import sklearn.preprocessing as pre
import matplotlib.pyplot as plt
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense
from keras.models import load_model
# -
# These are the imports I used for my script
# ## Building the network
# The first thing to do is initialise the network. I used the sequential model, this means we can add layers to the initialised network.
# Start neural network
model = kr.models.Sequential()
#Build neural network
model = Sequential()
# The next step is to add the layers, I added three layers. 1000, 750 and 512. I set the activation function as ReLu. To determine which class to output, I used the SoftMax function
# +
# Neural Network with 3 layers (1000, 750, 512)
model.add(kr.layers.Dense(units=1000, activation='relu', input_dim=784))
model.add(kr.layers.Dense(units=750, activation='relu'))
model.add(kr.layers.Dense(units=512, activation='relu'))
# Compile model - Adam optimizer for our model
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# Add 10 output neurons, one for each
model.add(kr.layers.Dense(units=10, activation='softmax'))
# -
# input, output and hidden layers
model.summary()
# ## Unzip the files
# I used gzip to unzip all the training, testing images and labels
# +
# Read in the files
with gzip.open('data/t10k-images-idx3-ubyte.gz', 'rb') as f:
test_images = f.read()
with gzip.open('data/t10k-labels-idx1-ubyte.gz', 'rb') as f:
test_labels = f.read()
with gzip.open('data/train-images-idx3-ubyte.gz', 'rb') as f:
training_images = f.read()
with gzip.open('data/train-labels-idx1-ubyte.gz', 'rb') as f:
training_labels = f.read()
# -
# ## Save files to memory
# +
# Read all files and save to memory
training_images = ~np.array(list(training_images[16:])).reshape(60000, 28, 28).astype(np.uint8) / 255.0
training_labels = np.array(list(training_labels[8:])).astype(np.uint8)
test_images = ~np.array(list(test_images[16:])).reshape(10000, 784).astype(np.uint8) / 255.0
test_labels = np.array(list(test_labels[8:])).astype(np.uint8)
# -
# ## Images and Lables
# We must put each pixel into a corrisponding neuron in the inputs of the network, we have to flatten the datasets into single arrays
# Flatten the array , 784 neurons
inputs = training_images.reshape(60000, 784)
# We then set up the lables, we must encode the data and then turn all the label dataset into binary values in a 10x10 matrix
encoder = pre.LabelBinarizer()
encoder.fit(training_labels)
outputs = encoder.transform(training_labels)
# ## Program running
# Now that every thing is set up we can run the program
# The user will firs be greeted with a message asking what they would like to do.
print("-------------------Welcome---------------------------------")
print("Would you like to train a dataset? Or load the data you have?")
print("Enter Y to train data set")
print("Enter N to load your own data")
# When the user presses y they will proceed to train the model
model.fit(inputs, outputs, epochs=10, batch_size=100)
# As we ca see neural network is working, We will now test it manually to see if it works
# +
from random import randint
for i in range(10): #Run 10 tests
print("----------------------------------")
randIndex = randint(0, 9999) #Get a random index to pull an image from
test = model.predict(test_images[randIndex:randIndex+1]) #Pull the image from the dataset
result = test.argmax(axis=1) #Set result to the highest array value
print("The actual number: ", test_labels[randIndex:randIndex+1])
print("The network reads: ", result)
print("----------------------------------")
# -
# As we can see it is working very accurately. We will now test the metrics accuracy
#print out accuracy
metrics = model.evaluate(inputs, outputs, verbose=0)
print("Metrics(Test loss & Test Accuracy): ")
print(metrics)
# Check Error Rate %
# Evaluates and then prints error rate accuracy
scores = model.evaluate(inputs, outputs, verbose=2)
print("Error Rate: %.2f%%" % (100-scores[1]*100))
I think this script is fairly accurate and quite well put together
# # References
# - https://machinelearningmastery.com/handwritten-digit-recognition-using-convolutional-neural-networks-python-keras/
# - https://medium.com/coinmonks/handwritten-digit-prediction-using-convolutional-neural-networks-in-tensorflow-with-keras-and-live-5ebddf46dc8
# - https://www.tensorflow.org/tutorials/
|
digit-recognition-notebook/digit-recognition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wtsyang/dl-reproducibility-project/blob/master/Report.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zJKQLTZVTrE_" colab_type="text"
# # Striving for Simplicity: The All Convolutional Net
# ###### *Group 16: <NAME>, <NAME>, and <NAME>*
# ###### 20/4/2020
# In this notebook, we try to reproduce the TABLE 3 in the [original paper](https://arxiv.org/abs/1412.6806). The source code of one of the models on ```Pytorch``` and the training procedure on Google Colab can be found in [Github](https://github.com/StefOe/all-conv-pytorch). **We adopt the original training procedure and change it to the Python class. Also, we build the models from scratch on Pytorch.**
#
# ---
#
# # Brief Introduction
# The paper shows that replacing the max-pooling with the convolutional with increased strides can improve the performance. The authors prove it by training models with max-pooling, models removing max-pooling, and models replacing max-pooling with convolutional layers. The results show the models replacing max-pooling with convolutional layers with strides generally have better performance. We provide a detaied explanation as follows. The authors tested 12 networks by designing 3 model bases and 3 branches.
#
# ## *Base: Model A, Model B, and Model C*
# Since the design of convolutional layers would influence the performance, the authors test three model bases. **Model A** uses the 5x5 strides. **Model B** uses the 5x5 strides but also adds one convolutional layer with 1x1 strides after that. **Model C** uses two convolutional layers with 3x3 strides.
#
# <img src='https://drive.google.com/uc?id=1HKDGWePX-PkBqRbeb8J_mwO-DDULhU2q' width="600px"/>
#
#
# ## *Branch: Model, Strided-CNN, ConvPool-CNN, and ALL-CNN*
# Each model base has one original model and thee branches. **“Model”** is the model with max-pooling. **“Strided-CNN”** is the model removing max-pooling. **“All-CNN”** is the model replacing max-pooling with convolutional strides. The better performance of “All-CNN” might result from more parameters than “Model” and “Strided-CNN”. To solve it, “ConvPool-CNN” is proposed. **“ConvPool-CNN”** is the model with max-pooling and one more convolutional layer before the pooling. “ConvPool-CNN” should have the same number of parameters as “All-CNN”. Therefore, if “All-CNN” has a better performance than “ConvPool-CNN”, we can prove the better performance on “All-CNN” does not result from more parameters.
# We show architecture with the base of model C in the following image.
#
# <img src='https://drive.google.com/uc?id=1gzpTwoW_Xx8YrHZdvU0ZmFT3n-1ktx1v' width="600px"/>
#
# ---
# # Experiment Setup
# All 12 networks are trained on the CIFAR-10 with the stochastic gradient descent with a fixed momentum of 0.9 and 350 epochs. The learning rate γ is chosen from the set ∈ [0.25, 0.1, 0.05, 0.01]. It is also scheduled by multiplying with a fixed factor of 0.1 in the epoch S= [200, 250, 300]. The paper only presents the best performance among all learning rates. Based on the source code, the performance is directly evaluated on the CIFAR-10 test set. In other words, **the source code did not use the validation for hyper-parameter tuning!**
#
# ---
#
# # Reproduction Results
# Our reproduction results are different from the paper as shown in the following table. First, we perceive an error rate gap between the paper and our reproduction, which is around 5~7%. Secondly, we obtain the same ranking in **Model A**, but we fail to reproduce the same order in **Model B** and **Model C**. This is because Model B and Model C are difficult to converge than Model A. Also, **ALL-CNN-B** and **ALL-CNN-C** fail to converge to the right place. During the training, the first three learning rates seem too large for the models. Therefore, we also try another learning rate=0.001 and a longer epoch=400 in **ALL-CNN-B**, where we add the * mark. Unfortunately, the model still fails to converge to the right place.
#
#
# | Model | Error Rate of Paper | Error Rate of Ours|
# |-|-|-|
# | Model A | 12.47%|19.27%|
# | Strided-CNN-A |13.46% |20.27%|
# | **ConvPool-CNN-A** |**10.21%**|**15.46%**|
# | ALL-CNN-A |10.30% |15.60%|
# | | ||
# | **Model B** | 10.20%| **17.01%** |
# | Strided-CNN-B | 10.98%|23.20%|
# | ConvPool-CNN-B | 9.33%|18.22%|
# | **ALL-CNN-B** | **9.10%** | *29.48%|
# | | ||
# | **Model C** |9.74%| **13.07%** |
# | Strided-CNN-C | 10.19%|15.49%|
# | ConvPool-CNN-C | 9.31%|14.39%|
# | **ALL-CNN-C** | **9.08%** |17.89%|
#
#
# + id="hsojPAB1q75j" colab_type="code" colab={}
# The example of training procedure
# Choose the model A
training=Training(baseModel=[True,False,False])
# Create the dataset
training.createDataset()
# Choose the branch: All-CNN
training.modifiedModel=[False,False,False,True]
# Start Training
training.Procedure()
# + [markdown] id="eVksKoMlaxi6" colab_type="text"
# # Validation
# Since the source code did not use the validation set for hyper-parameters tuning, we conduct the validation in this section. We split 5% of the training data to create the validation set. We show the results in the following table. The performance on the test set does not drop too much from the counterpart on the validation set. However, if we compare the test error with the original test error, the test error with the validation is generally higher. The result reveals that the models might be overestimated.
#
# | Model | Validation Set Error | Test Error| Original Test Error|
# |-|-|-|-|
# | Model A | 21.20% |20.45%|19.27%|
# | Strided-CNN-A |21.72% |21.38%|20.27%|
# | **ConvPool-CNN-A** |**15.93%**|**17.04%**|**15.46%**|
# | ALL-CNN-A |17.57%|18.57%|15.60%|
# | | ||
# | **Model B** | **16.65%**|17.81%|**17.01%**|
# | Strided-CNN-B | 17.53%|18.68%|23.20%|
# | ConvPool-CNN-B | 17.53%|**17.51%**|18.22%|
# | ALL-CNN-B | *24.53% | *25.78%|*29.48%|
# | | ||
# | **Model C** | **14.13%**|**14.87%**|**13.07%**|
# | Strided-CNN-C | 20.89%|21.67%| 15.49%|
# | ConvPool-CNN-C | 17.81%|17.60%| 14.39%|
# | ALL-CNN-C | 20.41%|19.13%| 17.89%|
# + id="7zAdzu2mq8l9" colab_type="code" colab={}
# Validation can be conducted by settting validation equal to True
training=Training(validation=True,bestModel_allLR=True,baseModel=[True,False,False])
# + [markdown] id="hztDzk-5qvm7" colab_type="text"
# # DropOut and Batch Normalization
# Dropout is a simple method to prevent neural networks from overfitting. In our all convolutional net paper, the author stated that dropout was used to regularize all networks.
# Dropout was almost essential in all the state-of-the-art networks before the introduction of batch normalization(BN). With the introduction of BN, it has shown its effectiveness and practicability in the recent neural networks. However, there is evidence that when these two techniques are used combinedly in a network, the performance of the network actually becomes worse. (Ioffe & Szegedy, 2015). In our study, we will investigate the results using BN and Dropout independently and also the effect of equipping both BN and Dropout simultaneously.
#
# + [markdown] id="3ZXK-gvLJ5np" colab_type="text"
#
# ### BatchNorm *only*
# + id="Qh2N6fpLq9Er" colab_type="code" colab={}
model = Model(dropOut=False, BN=True)
# + [markdown] id="z695JJpVnmkh" colab_type="text"
# <img src='https://drive.google.com/uc?id=17zb3ZUMTgRVLyRa1HYKm1SXqd-M_b6ai' width="500px"/>
# + [markdown] id="yHxeNuDgKOkp" colab_type="text"
# ### BatchNorm with Dropout
# + id="CCcecSDaKXT7" colab_type="code" colab={}
model = Model(dropOut=True, BN=True)
# + [markdown] id="FP4P4SoCLLwD" colab_type="text"
# <img src='https://drive.google.com/uc?id=11rOzypoJjhbfdQsD6KND2SyqtLfOuh2n' width="500px"/>
# + [markdown] id="Nb10QUPVfAI-" colab_type="text"
# ### Dropout *only*
# + id="3GjPD1UqfKnj" colab_type="code" colab={}
model = Model(dropOut=True, BN=False)
# + [markdown] id="g-41_3aFfK5Z" colab_type="text"
# <img src='https://drive.google.com/uc?id=1eVqvPxWPCAovkP3lNT1G_odgyisbAlLr' width="500px"/>
# + [markdown] id="b5uNJZc_LtTc" colab_type="text"
# ### Without using BatchNorm or Dropout
# + id="4H7CEUUbL914" colab_type="code" colab={}
model = Model(dropOut=False, BN=False)
# + [markdown] id="vAqeF0EIMD7E" colab_type="text"
# <img src='https://drive.google.com/uc?id=1uzW63lONO39_Sy9JGhzLc6Qqc976qe00' width="500px"/>
# + [markdown] id="lVvoCDCwgBrq" colab_type="text"
# We compare the results of different combination of these two techniques and generated the table below:
# + [markdown] id="kegTz4wwVQm8" colab_type="text"
# | Model | BN only | BN + Dropout |Dropout only| No BN no Dropout |
# |-|-|-|-|-|
# | Model A | no converge |no converge|19.27%| 13.82%|
#
#
# + [markdown] id="dr4OCR_QZf5z" colab_type="text"
# As shown in the table above, we used general Model A to study the two techniques Batch Normalization (BN) and Dropout, and whether it increases or decreases our model performance in this case. We implemented BN layer between two convolution layers, right before feeding into ReLu activations. Dropout is also applied in between convolution layers, and it is used after the pooling layer. The original paper stated that they used Dropout only, and we found out that using BN without Dropout or combining both BN with Dropout will not let our model converge. Li, Xiang, et al. (2019) stated in their papers that the worse performance may be the result of variance shifts. However, using Dropout only does lead the model to converge, giving the result of 19.27%. But the performance is still not as good as 13.82% without using either BN or dropout. It might be due to that we did not have the time to tune hyperparameter, dropout rate, we only used the parameter from the original paper: 20% for dropping out inputs and 50% otherwise.
# + [markdown] id="RmXwCYC2qhCF" colab_type="text"
# # Optimizer
# The default optimizer used in the paper and in our reproduction is Stochastic Gradient Descent (SGD) with momentum. We experimented with different optimizers since Adaptive Moment Estimation (Adam) is one of the most popular optimization algorithms, we decided to run Adam instead of SGD optimizer in our model. However, the model did not converge with Adam under the specific setting that we tried to reproduce. So even though in theory Adam combines the advantage of two SGD variants, RMSProp and AdaGrad, it is not very consistent in our specific setting to converge to an optimal solution.
# + [markdown] id="-Pvxevz9n1DI" colab_type="text"
#
# SGD with momentum optimizer:
# + id="L24H7bgEq9kD" colab_type="code" colab={}
self.optimizer = optim.SGD(model.parameters(), lr=lr, momentum=0.9)
# + [markdown] id="2ro4Fnkxlfsi" colab_type="text"
# Adam optimizer:
# + id="mT3REOuhlbJH" colab_type="code" colab={}
self.optimizer = optim.Adam(self.model.parameters(), lr=self.lr)
# + [markdown] id="LqmHp_bNmE3z" colab_type="text"
# The optimizer is the main approach nowadays for training neural networks with minimizing its error rate. Although Adam is a newer and more popular optimizer in a lot of the project, the original paper still chose to use SGD with momentum. During our model investigation, we found out that Adam fails to converge to an optimal solution in our specific setting, SGD with momentum does in this case perform better. This might be due reasons that the original paper has already tuned the hyperparameters extensively for SGD optimizer, so they chose Learning rate γ from the set ∈ [0.25, 0.1, 0.05, 0.01], and used the momentum 0.9. We, on the other hand, used the same learning rates for the Adam optimizer without extensively tuning them.
# + [markdown] id="zG6UzCsfoEsu" colab_type="text"
# # Summary
# This project is a reproduction work of one table in the paper "Striving for Simplicity: The All Convolutional Net." Three models are recreated with four different kinds of branches. Results show that model A has a good consistency compared with data in the paper while model B and model C differ. The influence of dropout and batch normalization to the results are also analyzed with several experiments. It is found that only using batch normalization or combining batch normalization with dropout will not let the model converge in this specific setting.
#
# # Discussion
#
# From our results we obtained, All-CNN models performed much worse than the ConvPool and base models except for variant model A. And when we tried to use the validation, All-CNN models cannot converge to an optimal solution for model A and model C. We also realized that the 0.25 learning rate in the original paper will not let most models converge, so we decided to drop it and later even tried learning rates such as small as 0.001, we found out that smaller learning rates might not guarantee the fast converge, but it will generally ensure the converge of the models. In general, we couldn't reproduce the results that the AllConv Net paper claimed, and the hypermeters such as the learning rates it suggested even lead to some models fail to converge. In conclusion, for the reproducibility of the paper, it is a good idea to always publish the source code as well, otherwise, the authors should pay more attention to the hyperparameters they suggest in the papers.
# + [markdown] id="M_GV7IN7U_Yf" colab_type="text"
# # Further Reading
# If you want to see more reproduction projects and even another researches on the same paper, please go to the [website](https://reproducedpapers.org? ).
# + [markdown] id="pOx0yCttcaFv" colab_type="text"
# # Reference
#
# Springenberg, <NAME>, et al. "Striving for simplicity: The all convolutional net." arXiv preprint arXiv:1412.6806 (2014).
#
#
# Li, Xiang, et al. "Understanding the disharmony between dropout and batch normalization by variance shift." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. (2019).
#
# Ioffe, Sergey, and <NAME>. "Batch normalization: Accelerating deep network training by reducing internal covariate shift." arXiv preprint arXiv:1502.03167 (2015).
#
# + [markdown] id="982dZGSNat4a" colab_type="text"
# # Appendix
#
#
# * [Github Repository](https://github.com/wtsyang/dl-reproducibility-project)
# * [A Python Class to build all Models](https://github.com/wtsyang/dl-reproducibility-project/blob/master/Model.py)
# * [A Python Class for Training Procedure](https://github.com/wtsyang/dl-reproducibility-project/blob/master/Training.py)
# * [The Notebooks for Model A](https://github.com/wtsyang/dl-reproducibility-project/tree/master/modelA)
# * [The Notebooks for Model B](https://github.com/wtsyang/dl-reproducibility-project/tree/master/modelB)
# * [The Notebooks for Model C](https://github.com/wtsyang/dl-reproducibility-project/tree/master/modelC)
# * [The Notebooks for Validation](https://github.com/wtsyang/dl-reproducibility-project/tree/master/Validation)
# * [The Notebooks for DropOut and Batch Normalization](https://github.com/wtsyang/dl-reproducibility-project/tree/master/BN_Dropout)
# * [The Notebooks for Optimizers](https://github.com/wtsyang/dl-reproducibility-project/tree/master/optimizer)
#
#
#
#
#
#
#
#
#
|
Report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing re-orientation in shearflow + elongation.
#
# The result should match Figure 5 in
# <NAME>, <NAME>., Tucker, <NAME>: "An objective model for slow
# orientation kinetics in concentrated fiber suspensions: Theory and rheological
# evidence", Journal of Rheology, 52, 1179, 2008: DOI: 10.1122/1.2946437.
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
from fiberoripy.orientation import rsc_ode
# +
# geometric factor
xi = 1.0
# time steps
t = np.linspace(0, 400, 500)
# initial fiber orientation state
A0 = 1.0 / 3.0 * np.eye(3)
# +
fig, axes = plt.subplots(3, 1, figsize=(4, 6))
for ax, ratio in zip(axes, [0.1, 0.12, 0.2]):
def L(t):
"""Velocity gradient."""
return np.array(
[[-ratio, 0.0, 1.0], [0.0, ratio, 0.0], [0.0, 0.0, 0.0]]
)
# computed solution
A = odeint(rsc_ode, A0.ravel(), t, args=(xi, L, 0.01, 0.1))
ax.plot(t, A[:, 0], label="A11")
ax.plot(t, A[:, 4], label="A22")
ax.plot(t, A[:, 2], label="A13")
ax.set_xlabel("Time $t$ in s")
ax.set_ylim([0, 1])
ax.grid()
ax.legend()
plt.tight_layout()
plt.show()
# -
|
examples/orientation/rsc_combiload.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DS Automation Assignment - <NAME>
# +
import pandas as pd
df = pd.read_csv('prepped_churn_data.csv', index_col='Customer')
df
# -
df.drop('Tenure_Charges_ratio', inplace=True, axis=1)
conda install -c conda-forge pycaret -y
from pycaret.classification import setup, compare_models, predict_model, save_model, load_model
automl = setup(df, target='Churn')
automl[6]
best_model = compare_models()
best_model
df.iloc[-2:-1].shape
predict_model(best_model, df.iloc[-2:-1])
save_model(best_model, 'CBC')
new_data = df.iloc[-2:-1].copy()
loaded_CBC = load_model('CBC')
predict_model(loaded_CBC, new_data)
# +
from IPython.display import Code
Code('predict_churn.py')
# -
# %run predict_churn.py
# # Summary
# This assignment presented a number of challenges. While working through the autoML packages was straight forward, saving the model for future use and crafting a viable Python code required a fair amount of trial and error. One challenge was generated by the randomness of the best_model function. Every time the code was run, it selected a new model which meant changing all the labels and .py files. In the end, I left the model names as written ('CBC') even thought the last run selected logistic regression as the best. I also cut out the pickle read and write seciton because I could not get the code to run without an error. This was redundant anyway and I was able to complete the assignment requirements without this code. In the end the model seemed to do better at predicting no churn instead of churn. The scores were higher. This is useful in itself, given that the company could reduce its marketing efforts for this with a high score for no churn.
|
MSDS600_CGREEN_WEEK5_ASSIGNMENT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CoNLL_View_Doc.ipynb
#
# Notebook for viewing individual documents from the CoNLL corpus, for use
# alongside the other CoNLL related notebooks in this directory.
# +
# The Jupyter kernel for this notebook usually starts up inside the notebooks
# directory, but the text_extensions_for_pandas package code is in the parent
# directory. Add that parent directory to the front of the Python include path.
import sys
if ".." not in sys.path:
sys.path.insert(0, "..")
# Libraries
import numpy as np
import pandas as pd
# And of course we need the text_extensions_for_pandas library itself.
import text_extensions_for_pandas as tp
# Common code shared across notebooks comes from util.py
import util
# -
# Download and cache the data set.
# NOTE: This data set is licensed for research use only. Be sure to adhere
# to the terms of the license when using this data set!
data_set_info = util.get_conll_data()
data_set_info
# The raw dataset in its original tokenization
corpus_raw = {}
for fold_name, file_name in data_set_info.items():
df_list = tp.conll_2003_to_dataframes(file_name,
["pos", "phrase", "ent"],
[False, True, True])
corpus_raw[fold_name] = [
df.drop(columns=["pos", "phrase_iob", "phrase_type"])
for df in df_list
]
# Convert IOB2-tagged tokens to spans
all_spans = {
k: [tp.iob_to_spans(df) for df in v] for k, v in corpus_raw.items()
}
# Turn off the 60-row limit for displaying dataframes
pd.options.display.max_rows = None
fold = "dev"
doc_offset = 28
doc_df = all_spans[fold][doc_offset]
doc_df
doc_df["token_span"].values.repr_html_show_offsets = False
doc_df["token_span"].values
# + jupyter={"outputs_hidden": true}
# Dataframe of tokens for finding offsets
toks_df = corpus_raw[fold][doc_offset]
toks_df
# -
# ######
|
tutorials/corpus/CoNLL_View_Doc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solution Graded Exercise 8: Hopfield Network model of associative memory
# first name: ...
#
# last name: ...
#
# sciper: ...
#
# date: ...
#
# *Your teammate*
#
# first name of your teammate: ...
#
# last name of your teammate: ...
#
# sciper of your teammate: ...
#
#
# Note: You are allowed to discuss the concepts with your class mates. You are not allowed to share code. You have to understand every line of code you write in this notebook. We will ask you questions about your submission during a fraud detection session during the last week of the semester.
#
# ** Remember **
#
# If you are asked for plots: The appearance of the plots (labelled axes, ** useful scaling **, etc.) is important!
#
# If you are asked for discussions: Answer in a precise way and try to be concise.
#
#
# ** Submission **
#
# Rename this notebook to Ex8_FirstName_LastName_Sciper.ipynb and upload that single file on moodle before the deadline.
#
# ** Link to the exercise **
#
# http://neuronaldynamics-exercises.readthedocs.io/en/latest/exercises/hopfield-network.html
# ## \* \* \* \* Programming Notes : Copying and slicing lists and numpy arrays \* \* \* \*
# We would like to take the opportunity to bring to your attention certain features of Python, that might lead to unwanted behaviour and serious mistakes, if one is not aware of them.
# Please check the Python Cheat Sheet file on the moodle (https://moodle.epfl.ch/mod/page/view.php?id=981134 Part 4 of the ipynb file) for some examples of the following notes:
#
# * Assigning a list to a new variable does not create a copy of the list, but creates a variable that points to the list. This means that modifying the second variable, also modifies the original list.
# * Assigning a slice of a list to a new variable, creates a copy of the list. Any modification to the sliced list does not modify the original.
#
# Now when it comes to numpy arrays:
# * Assigning a numpy array to a new variable does not create a copy of the array, but creates a variable that points to the array. This means that modifying the second variable, also modifies the original array. (same as above)
# * Assigning a slice of a numpy array to a new variable creates a variable that points to the corresponding elements of the original array as well! (contrary to what we saw above!) This means that modifying the second variable, also modifies the original array!
# * To copy the original array and ensure that it is not modified by any modification of its copied version, the method copy() should be used.
# # Exercise 8.1. Getting started
# +
# %matplotlib inline
from neurodynex.hopfield_network import network, pattern_tools, plot_tools
pattern_size = 5
# create an instance of the class HopfieldNetwork
hopfield_net = network.HopfieldNetwork(nr_neurons= pattern_size**2)
# instantiate a pattern factory
factory = pattern_tools.PatternFactory(pattern_size, pattern_size)
# create a checkerboard pattern and add it to the pattern list
checkerboard = factory.create_checkerboard()
pattern_list = [checkerboard]
# add random patterns to the list
pattern_list.extend(factory.create_random_pattern_list(nr_patterns=3, on_probability=0.5))
plot_tools.plot_pattern_list(pattern_list)
# how similar are the random patterns and the checkerboard? Check the overlaps
overlap_matrix = pattern_tools.compute_overlap_matrix(pattern_list)
plot_tools.plot_overlap_matrix(overlap_matrix)
# let the hopfield network "learn" the patterns. Note: they are not stored
# explicitly but only network weights are updated !
hopfield_net.store_patterns(pattern_list)
# create a noisy version of a pattern and use that to initialize the network
noisy_init_state = pattern_tools.flip_n(checkerboard, nr_of_flips=4)
hopfield_net.set_state_from_pattern(noisy_init_state)
# from this initial state, let the network dynamics evolve.
states = hopfield_net.run_with_monitoring(nr_steps=4)
# each network state is a vector. reshape it to the same shape used to create the patterns.
states_as_patterns = factory.reshape_patterns(states)
# plot the states of the network
plot_tools.plot_state_sequence_and_overlap(states_as_patterns, pattern_list, reference_idx=0, suptitle="Network dynamics")
# -
# # 8.3. Exercise: N=4x4 Hopfield-network
# ## 8.3.1. Question: Storing a single pattern
# #### [2 + 2 + 3 points]
# +
# write your code here
# +
# Plot the sequence of network states along with the overlap of network state with the checkerboard
# -
# Now test whether the network can still retrieve the pattern if we increase the number of flipped pixels. What happens at nr_flipped_pixels = 8, what if nr_flipped_pixels > 8 ?
# ## 8.3.2. Question: the weights matrix
# #### [1 + 3 + 2 points]
# +
# write your code here: Bullet points 1-5
# +
# write your code here: Bullet points 6-11
# -
# How does this matrix compare to the two previous matrices?
#
# ## 8.3.3. Question (optional): Weights Distribution
# You can easily plot a histogram by adding the following two lines to your script.
#It assumes you have stored your network in the variable ‘hopfield_net’.
"""
plt.figure()
plt.hist(hopfield_net.weights.flatten())
"""
# # 8.4. Exercise: Capacity of an N=100 Hopfield-network
# ## 8.4.1. Associative memory.
# #### [2 points]
# A Hopfield network implements so called associative or content-adressable memory. Explain what this means. (max 4 lines)
#
# ## 8.4.2. Capacity of the network.
# #### [1 points]
# Using the value $C_{store}$
# given in the book, how many patterns can you store in a N=10x10 network? Use this number K in the next question:
#
# ## 8.4.3. Checkerboard and random patterns.
# #### [3 points]
# +
# write your code here
# -
# Rerun your script a few times. What do you observe?
# #### [2 points]
# # 8.5. Exercise: Non-random patterns
# ## 8.5.1. Alphabet.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from neurodynex.hopfield_network import network, pattern_tools, plot_tools
import numpy
# the letters we want to store in the hopfield network
letter_list = ['A', 'B', 'C', 'S', 'X', 'Y', 'Z']
# set a seed to reproduce the same noise in the next run
# numpy.random.seed(123)
abc_dictionary =pattern_tools.load_alphabet()
print("the alphabet is stored in an object of type: {}".format(type(abc_dictionary)))
# access the first element and get it's size (they are all of same size)
pattern_shape = abc_dictionary['A'].shape
print("letters are patterns of size: {}. Create a network of corresponding size".format(pattern_shape))
# create an instance of the class HopfieldNetwork
hopfield_net = network.HopfieldNetwork(nr_neurons= pattern_shape[0]*pattern_shape[1])
# create a list using Pythons List Comprehension syntax:
pattern_list = [abc_dictionary[key] for key in letter_list ]
plot_tools.plot_pattern_list(pattern_list)
# store the patterns
hopfield_net.store_patterns(pattern_list)
# # create a noisy version of a pattern and use that to initialize the network
noisy_init_state = pattern_tools.get_noisy_copy(abc_dictionary['A'], noise_level=0.2)
hopfield_net.set_state_from_pattern(noisy_init_state)
# from this initial state, let the network dynamics evolve.
states = hopfield_net.run_with_monitoring(nr_steps=4)
# each network state is a vector. reshape it to the same shape used to create the patterns.
states_as_patterns = pattern_tools.reshape_patterns(states, pattern_list[0].shape)
# plot the states of the network
plot_tools.plot_state_sequence_and_overlap(
states_as_patterns, pattern_list, reference_idx=0, suptitle="Network dynamics")
# -
# ## 8.5.2. Add a letter.
# #### [2 points]
# +
# write your code here
# -
# Is the pattern ‘A’ still a fixed point?
# Does the overlap between the network state and the reference pattern ‘A’ always decrease? (max 3 lines)
# #### [3 points]
# ## 8.5.3. Capacity.
# #### [3 points]
# +
# write your code here
# -
# Explain the discrepancy between the network capacity C (computed above) and your observation. (max 4 lines)
# #### [3 points]
# # 8.6. Implementing different types of dynamics
# ### *** Note:*** this exercise is NOT a bonus and it is graded
# ## 8.6.1. Exercise:
# Implement a Hopfield network with asynchronous dynamics (use the function set_dynamics_sign_async()). Store 5 random patterns (take inspiration from exercise 8.3). Fix a noisy initial state, run the network a few times and observe the performance over the different trials.
#
# Parameters: network size=5x5, number of rnd patterns=5, on probability=0.5, number of flips=8.
# #### [5 points]
# +
# write your code here
# -
# Comment on your results (max 3 lines).
# #### [3 points]
# ## 8.6.2. Stochastic neuron
# Implement a synchronous and stochastic neuron. Use the function HopfieldNetwork.set_dynamics_to_user_function() in order to pass _get_sigmoid_update_function(), which you should implement.
#
# In the stochastic network the dynamics depend on the so-called inverse temperature $\beta$. The temperature in this system can be seen as noise. The activity $S_i(t+1)$ in the next time-step follows the probability distribution:
# \begin{equation}
# P\left(S_i(t+1)=1\right) = \frac1{1+\exp\left[-h_i(t)\beta\right]} \qquad \textrm{ where } \qquad h_i(t) = \sum_j w_{ij} S_j(t)
# \end{equation}
# That is, it is more likely for a neuron $i$ to be set to +1 if its input $h_i$ is higher.
#
# Parameters: network size = 5x5, number of rnd patterns = 5, on probability = 0.5, number of flips = 8, $\beta$=10.
#
# *** Hint 1: *** For inspiration check out the source code of the function _get_sign_update_function()
#
# *** Hint 2: *** In order to implement stochasticity, we suggest to use the python-library numpy.random.
#
# #### [7 points]
# +
# Synchronous stochastic update
def _get_sigmoid_update_function(beta):
"""
for internal use
Returns:
A function implementing a synchronous state update using the sigmoid function (h)
"""
def upd(state_s0, weights):
# ???
return s1
return upd
beta = ???
hopfield_net_stoch = network.HopfieldNetwork(nr_neurons= pattern_size**2)
hopfield_net_stoch.set_dynamics_to_user_function(_get_sigmoid_update_function(beta=beta))
# -
# ## 8.6.3 Finite temperature
# Choose one of the stored patterns (let's say number 1) and flip 10 bits. Then plot the overlap between the network state and this pattern, $m^1$, over time for different values of $\beta$, $\beta\in \left[0.1,2,5,100\right]$.
#
# Parameters: network size = 10x10, number of rnd patterns = 5, on probability = 0.5, number of flips = 10, number of trials = 100, number of steps = 9.
#
# *** Hint 1:*** In order to plot $m^1$ over time, run the script several times and compute the mean and the standard deviation $\sigma$ of the overlap $m^1$ over trials. You can use the function plt.fill and its parameter $\alpha$ (the transparency), in order to fill with color the area between $+\sigma$ and $-\sigma$.
#
# *** Hint 2: *** Use the function pattern_tools.compute_overlap()
#
# *** Hint 3:*** We suggest to implement and test your code with a smaller network size before getting to the 10x10 units network.
# #### [7 points]
# +
def compute_overlap_over_time(hopfield_net, pattern_list, idx, init_state, nTrials, nr_steps):
""" For a given hopfield_net and a pattern_list, initialize it with the init_state and run it
Compute its overlap with the pattern pattern_list[idx] in time (for nr_steps)
Repeat the above over many trial (nTrials)
"""
overlap = np.zeros((nTrials,nr_steps+1))
for i in range(0,nTrials):
# learn patterns
# ???
# run a synch and stoch Hopfield model with the same initial state
# ???
# let the netwrok dynamics evolve
# ???
states_as_patterns = ???
for i_step in range(len(states_as_patterns)):
overlap[i,i_step] = ???
return overlap
# Create the patterns and store them
nTrials = ???
nr_steps= ???
betas = ???
overlap_mean = np.zeros((len(betas), nr_steps+1))
overlap_std = np.zeros((len(betas), nr_steps+1))
for j in range(0, len(betas)):
# ???
overlap_mean[j,:] = ???
overlap_std[j,:] = ???
# Plot
# ???
# -
# Comment on your plot (max 5 lines).
# #### [5 points]
# write your answer here
# ## 8.6.4 Asynchronous stochastic neuron
# Implement the stochastic neuron of exercise 8.6.2, but with asynchronous update.
#
# *** Hint: *** For help check out the source code of the function _get_async_sign_update_function()
# #### [4 points]
# +
def _get_asynch_sigmoid_update_function(beta):
"""
for internal use
Returns:
A function implementing a asynchronous state update using the sigmoid function
"""
def upd(state_s0, weights):
random_neuron_idx_list = np.random.permutation(len(state_s0))
state_s1 = state_s0.copy()
for i in range(len(random_neuron_idx_list)):
# ???
return state_s1
return upd
beta = ???
hopfield_net_asynch_stoch = network.HopfieldNetwork(nr_neurons= pattern_size**2)
hopfield_net_asynch_stoch.set_dynamics_to_user_function(_get_asynch_sigmoid_update_function(beta=beta))
# -
# # 8.7 Energy
#
# ## 8.7.1 Exercise
# Set up a deterministic and asynchronous Hopfield network and store 3 random patterns.
#
# The energy can be defined as $E=-\sum_i^N \sum_j^N w_{ij} S_i S_j$.
#
# Implement a function that calculates the energy according to the above definition.
#
# Parameters: network size = 10x10, number of rnd patterns = 3, on probability = 0.5.
#
# #### [4 points]
# +
# Set up your network
# ???
def energy(weight, state):
return ???
# -
# ## 8.7.2 Exercise
# Compute the energy of one of the stored patterns: $E (\xi^{\mu})$.
#
# Choose one pattern and gradually flip more and more bits. For that you need to implement a function flip_idx, after checking out the function pattern_tools.flip_n(). How does the energy change and why? (max 3 lines)
#
# *** Hint: *** You can get an intuition by plotting the energy of the state as a function of the number of flipped bits.
# #### [1 + 6 + 4 points]
# +
nr_flips = np.arange(0,10,1)
energy_flips = np.zeros(len(nr_flips))
for i in range(len(nr_flips)):
# ???
e_flips[i] = ???
# Plot
# ???
# -
# your answer
# ## 8.7.3 Question.
# Compute the energy of the reverse of one of the stored patterns $E(-\xi^1)$. What do you observe, what are the consequences on the dynamics of the system and why? (max 3 lines)
# #### [1 + 3 points]
# ## 8.7.4 Exercise.
# 1 - Initialize the network with the state $S_0 = sgn(\sum_{\mu}^3 \xi^{\mu})$ and run the dynamics. What is the energy of this state?
#
# Parameters: number steps = 4.
#
# #### [2 points]
# +
# write your code here
# -
# 2 - Consider the ensemble of all states that differ from $S_0$ in exactly one flip. Initialize the network in each of those states and check what is the final state of the dynamical evolution.
#
# *** Hint: *** Compare the final states with $S_0$.
# #### [8 points]
# +
# Function to flip one neuron
def flip_idx(template, idx):
"""
makes a copy of the template pattern and flips the idx-th state.
Args:
template:
nr_of_flips:
Returns:
a new pattern
"""
# ???
return ???
# Flip all neurons one after the other
for i in range(0, pattern_size**2):
s_0_flipped = flip_idx(s_0, i)
# ???
# -
# 3 - What can you tell about the stability of $S_0$? Briefly comment (max 5 lines).
#
# ***Hint:*** For help, refer to the pdf on associative memory on the MOODLE (https://moodle.epfl.ch/pluginfile.php/1091071/mod_resource/content/0/Lecture5/Hertz_Hopfield.pdf).
#
# #### [6 points]
# your answer
# ## 8.7.5 Question.
# Does your argumentation above hold in the case of $S_0 = sgn(\sum_{\mu}^2 \xi^{\mu})$? Why? (max 3 lines)
# #### [3 points]
# ## 8.7.6 Exercise.
# Use now the stochastic neuron with asynchronous update you implemented in 8.6.4. Initialize the network with the state $S_0 = sgn(\sum_{\mu}^3 \xi^{\mu})$. For $\beta$ varying in $\beta\in \left[1,2,8,10,100\right]$, plot the overlap of the network initialized with $S_0$ with each of the 3 stored patterns as a function of time.
#
# What do you observe? Comment on the stability of the spurious state $S_0$ in each case. (max 4 lines)
#
# Parameters: network size = 10x10, number of rnd patterns = 3, on probability = 0.5, number of steps = 9.
#
# *** Hint 1: *** Make 4 plots (or 1 plot with 4 subplots), one for each value of $\beta$.
#
# *** Hint 2: *** You can use the function pattern_tools.compute_overlap_list (or pattern_tools.compute_overlap).
# #### [5 + 4 points]
# +
# ???
for j in range(0, len(betas)):
# ???
# Compute
# Plot
# -
# your answer
|
project2/.ipynb_checkpoints/Ex8_firstName_lastName_000-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from thinkbayes2 import Pmf, Cdf, Suite, MakeJoint
import thinkplot
import pymc3 as pm
import numpy as np
import pandas as pd
from pymc3.math import exp, log
# -
def plotCdf(values, *args, **kwargs):
thinkplot.Cdf(Cdf(values), *args, **kwargs)
# # Problem Description
# In the 2018 season of Major League Baseball...
#
# We want to estimate which team is the best team, and ...
# To accomplish this, we build an object-oriented PyMC model that predicts the expected number of runs scored by a given team in a game against another team. The model also includes mechanisms for representing properties of Baseball as a game, allowing it to build a Bayesian estimate of the mean score in a baseball game and the additional runs that a team is expected to get from having home field advantage.
class Baseball(object):
"""
A Baseball object contains properties about fundamental properties of
the sport of Baseball itself, represented with PyMC3 distributions.
"""
def __init__(self, name='baseball'):
self.name = name
self.mean_score = pm.Normal(f'{name}.mean_score', mu=4.45, sd=1) # Based on 2018 MLB season
self.home_field_advantage = pm.Exponential(f'{name}.home_field_advantage', lam=5)
self.offense_weight = pm.Normal(f'{name}.offense_weight', mu=1, sd=0.5)
self.defense_weight = pm.Normal(f'{name}.defense_weight', mu=1, sd=0.5)
def get_runs(self, team, opponent):
return self.mean_score * exp(team.off_quality * self.offense_weight - opponent.def_quality * self.defense_weight)
# The model we use for runs is a Poisson process with parameter dependent on the skills of both teams, as well as which team has home field advantage. In particular, we model the expected number of runs a given team will score in a game as
#
# $$k_{baseball} * \frac{e^{a(offense_{you})}}{e^{b(defense_{opponent})}} + home\_field\_boost$$
#
# where
#
# \begin{cases}
# home\_field\_boost > 0 & \text{Team is home} \\
# home\_field\_boost = 0 & \text{Team is away} \\
# \end{cases}
#
# A team's expected runs scored is dependent on both the team's offensive (run creation) skill and the opponent's defensive (run prevention) skill, while `a` and `b` in the above equation represent the relative weights of those two parameters.
class Team(object):
def __init__(self, name, baseball):
self.name = name
self.baseball = baseball
self.off_quality = pm.Normal(f'{name}.off_quality', mu=0, sd=1)
self.def_quality = pm.Normal(f'{name}.def_quality', mu=0, sd=1)
def model_game(self, opponent, homefield : bool, date, observed_runs_scored=None, observed_runs_allowed=None):
expected_runs_scored = self.baseball.get_runs(self, opponent)
expected_runs_allowed = self.baseball.get_runs(opponent, self)
if homefield:
expected_runs_scored += self.baseball.home_field_advantage
else:
expected_runs_allowed += self.baseball.home_field_advantage
pm.Poisson(f'{date}_{self.name}', mu=expected_runs_scored, observed=observed_runs_scored)
pm.Poisson(f'{date}_{opponent.name}', mu=expected_runs_allowed, observed=observed_runs_allowed)
team_names = ['BOS', 'NYY', 'TBR', 'TOR', 'BAL']
team_games = {}
for team in team_names:
data = pd.read_csv(f'../data/{team.lower()}.csv')
df = data[['Date', 'Opp', 'Unnamed: 4', 'R', 'RA']]
df = df.loc[df['Opp'].isin(team_names)]
df = df.rename(index=str, columns={'Unnamed: 4': 'Home'})
df['Home'] = df['Home'].apply(lambda x: False if x == '@' else True)
team_games[team] = df
# +
modeled_games = {}
num_games = 0
runs = {}
allowed = {}
with pm.Model() as model:
bb = Baseball()
teams = {name: Team(name, bb) for name in team_names}
for name, games in team_games.items():
for index, game in games.head(20).iterrows():
date = game['Date']
opp = game['Opp']
if not (opp, date) in modeled_games:
modeled_games[(name, date)] = True
num_games += 1
teams[name].model_game(teams[opp], game['Home'], date, game['R'], game['RA'])
# +
# Inspect the prior distribution
with model:
prior_trace = pm.sample_prior_predictive()
# +
# Infer the posterior distributions
with model:
trace = pm.sample(2000, tune=1000, cores=4, nuts_kwargs=dict(target_accept=.95))
pm.traceplot(trace);
# -
for t in team_names:
plotCdf(prior_trace[f'{t}.off_quality'], label=f'{t} Offensive Quality Prior')
thinkplot.Show()
for t in team_names:
plotCdf(prior_trace[f'{t}.def_quality'], label=f'{t} Defensive Quality Prior')
thinkplot.Show()
# The respective posterior distributions align with what we would expect from these teams qualitatively (and from looking at their first 10 inter-division games, quantitatively).
for t in team_names:
plotCdf(trace[f'{t}.off_quality'], label=f'{t} Offensive Quality Posterior')
thinkplot.Show()
for t in team_names:
plotCdf(trace[f'{t}.def_quality'], label=f'{t} Defensive Quality Posterior')
thinkplot.Show()
plotCdf(prior_trace['baseball.mean_score'], label='prior')
plotCdf(trace['baseball.mean_score'], label='posterior')
thinkplot.Config(title='Mean runs per game')
plotCdf(prior_trace['baseball.home_field_advantage'], label='prior')
plotCdf(trace['baseball.home_field_advantage'], label='posterior')
thinkplot.Config(title='Home field advantage')
# +
def plot_better(better, worse, attr, attr_name):
plotCdf(prior_trace[f'{better}.{attr}'] - prior_trace[f'{worse}.{attr}'], label='prior', color='grey')
plotCdf(trace[f'{better}.{attr}'] - trace[f'{worse}.{attr}'], label='posterior')
thinkplot.Config(title='Quality differential')
win_chance=np.mean((trace[f'{better}.{attr}'] - trace[f'{worse}.{attr}'])>0)
print(f'{win_chance} chance that {better} {attr_name} is better than {worse} {attr_name}')
plot_better('BOS', 'NYY', 'off_quality', 'offense') # What really matters
# -
plot_better('BOS', 'NYY', 'def_quality', 'defense')
# +
# Logic borrowed from Eric's ranking project
best_off_teams = []
best_def_teams = []
for i in range(len(trace)):
best_off_teams.append(max(team_names, key=lambda name: trace[f'{name}.off_quality'][i]))
best_def_teams.append(max(team_names, key=lambda name: trace[f'{name}.def_quality'][i]))
from collections import Counter
for team, count in Counter(best_off_teams).most_common():
print(f'{team} offense is best with probability {count/len(trace)*100:.1f}%')
print()
for team, count in Counter(best_def_teams).most_common():
print(f'{team} defense is best with probability {count/len(trace)*100:.1f}%')
# -
|
code/bayesball_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Embedding Multiple Graphs
#
# This demo shows you how to simultaneously embed two graphs using omnibus embedding from two graphs sampled from different stochastic block models (SBM). We will also compare the results to that of adjacency spectral embedding, and show why it is useful to embed the graphs simultaneously.
# +
import graspologic
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
# ## Simulate two different graphs using stochastic block models (SBM)
#
# We sample 2-block SBMs (undirected, no self-loops) with 50 vertices, each block containing 25 vertices (n = [25, 25]), and the following block probabilities:
#
# \begin{align*}
# P_1 =
# \begin{bmatrix}0.3 & 0.1\\
# 0.1 & 0.7
# \end{bmatrix},~
# P_2 = \begin{bmatrix}0.3 & 0.1\\
# 0.1 & 0.3
# \end{bmatrix}
# \end{align*}
#
# The only difference between the two are the block probability for the second block. We sample $G_1 \sim \text{SBM}(n, P_1)$ and $G_2 \sim \text{SBM}(n, P_2)$.
# +
from graspologic.simulations import sbm
n = [25, 25]
P1 = [[.3, .1],
[.1, .7]]
P2 = [[.3, .1],
[.1, .3]]
np.random.seed(8)
G1 = sbm(n, P1)
G2 = sbm(n, P2)
# -
# ## Visualize the graphs using heatmap
#
# We visualize the sampled graphs using heatmap function. Heatmap will plot the adjacency matrix, where the colors represent the weight of the edge. In this case, we have binary graphs so the values will be either 0 or 1.
#
# There is clear block structure to the graphs, and we see that the second, lower right, block of $G_1$ has more edges than that of $G_2$.
# +
from graspologic.plot import heatmap
heatmap(G1, figsize=(7, 7), title='Visualization of Graph 1')
_ = heatmap(G2, figsize=(7, 7), title='Visualization of Graph 2')
# -
# ## Embed the two graphs using omnibus embedding
#
# The purpose of embedding graphs is to obtain a Euclidean representation, sometimes called latent positions, of the adjacency matrices. Again, we assume that the probability matrix of a graph is given by $P = XX^T$ and we are trying to estimate $X$. The benefit of omnibus embedding is that the latent positions of all embedded graphs live in the same canonical space, thus eliminating the need to align the results.
#
# We use all of the default parameters. Underneath, the select_dimension algorithm will automatically find the optimal embedding dimension for us. In this example, we get the following estimate,
#
# \begin{align*}
# \hat{Z} =
# \begin{bmatrix}
# \hat{X_1}\\
# \hat{X_2}
# \end{bmatrix}
# \end{align*}
#
# where the first block, $\hat{X_1}$, are the latent positions of the first graph, and the second block, $\hat{X_2}$, are the latent positions of the second graph.
# + tags=[]
from graspologic.embed import OmnibusEmbed
embedder = OmnibusEmbed()
Zhat = embedder.fit_transform([G1, G2])
print(Zhat.shape)
# -
# ## Visualize the latent positions
#
# Since the two graphs have clear block structures, we should see two "clusters" when we visualize the latent positions. The vertices that form the first block should be close together since they have the same block probabilities, while those that form the second block should be further apart since they have different block probabilities.
# +
Xhat1 = Zhat[0]
Xhat2 = Zhat[1]
# Plot the points
fig, ax = plt.subplots(figsize=(10, 10))
ax.scatter(Xhat1[:25, 0], Xhat1[:25, 1], marker='s', c='blue', label = 'Graph 1, Block 1')
ax.scatter(Xhat1[25:, 0], Xhat1[25:, 1], marker='o', c='blue', label = 'Graph 1, Block 2')
ax.scatter(Xhat2[:25, 0], Xhat2[:25, 1], marker='s', c='red', label = 'Graph 2, Block 1')
ax.scatter(Xhat2[25:, 0], Xhat2[25:, 1], marker='o', c='red', label= 'Graph 2, Block 2')
ax.legend()
# Plot lines between matched pairs of points
for i in range(50):
ax.plot([Xhat1[i, 0], Xhat2[i, 0]], [Xhat1[i, 1], Xhat2[i, 1]], 'black', alpha = 0.15)
_ = ax.set_title('Latent Positions from Omnibus Embedding', fontsize=20)
|
docs/tutorials/embedding/Omnibus.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Python 3
// language: python
// name: python3
// ---
// + [markdown] colab_type="text" id="view-in-github"
// <a href="https://colab.research.google.com/github/tgalkovskyi/deep-text-recognition-benchmark/blob/master/demo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
// + [markdown] colab_type="text" id="dHbHK60Tqn2c"
// This can be executed in https://colab.research.google.com "Python 3 / GPU" runtime.
// + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="Pi-PA14AhdK-" outputId="5d620547-6ce9-42b9-8206-20f88f23e567" pycharm={"name": "#%% bash\n"}
# !git clone https://github.com/clovaai/deep-text-recognition-benchmark
# %cd deep-text-recognition-benchmark
// + [markdown] colab_type="text" id="jX-ucvimlaFZ"
// Next, download large model files from Google Drive, using hack: https://stackoverflow.com/questions/20665881/direct-download-from-google-drive-using-google-drive-api/32742700#32742700
// + pycharm={"name": "#%% bash\n"}
# !pwd
# !ls
#// change according to your path
# %cd /home/selcuk/PycharmProjects/ocr_toolkit/recognition/deep_text_recognition
# %pwd
# !ls
// + colab={"base_uri": "https://localhost:8080/", "height": 833} colab_type="code" id="9eEhhPBshkjr" outputId="205d0bd8-340f-4e9b-ebfd-e9c8f7d327a0" pycharm={"name": "#%% bash\n"}
# Original
# models = {
# 'None-ResNet-None-CTC.pth': 'https://drive.google.com/open?id=1FocnxQzFBIjDT2F9BkNUiLdo1cC3eaO0',
# 'None-VGG-BiLSTM-CTC.pth': 'https://drive.google.com/open?id=1GGC2IRYEMQviZhqQpbtpeTgHO_IXWetG',
# 'None-VGG-None-CTC.pth': 'https://drive.google.com/open?id=1FS3aZevvLiGF1PFBm5SkwvVcgI6hJWL9',
# 'TPS-ResNet-BiLSTM-Attn-case-sensitive.pth': 'https://drive.google.com/open?id=1ajONZOgiG9pEYsQ-eBmgkVbMDuHgPCaY',
# 'TPS-ResNet-BiLSTM-Attn.pth': 'https://drive.google.com/open?id=1b59rXuGGmKne1AuHnkgDzoYgKeETNMv9',
# 'TPS-ResNet-BiLSTM-CTC.pth': 'https://drive.google.com/open?id=1FocnxQzFBIjDT2F9BkNUiLdo1cC3eaO0',
# }
# my google drive
models = {
'None-ResNet-None-CTC.pth': 'https://drive.google.com/open?id=1WF5XJvReLQ4DyYTbrvFzZ7zc-nFc7VGI',
'None-VGG-BiLSTM-CTC.pth': 'https://drive.google.com/open?id=1UcnA5eqGTj4Wq2lFp-qKOrjIbcJ0tVuP',
'None-VGG-None-CTC.pth': 'https://drive.google.com/open?id=1bbom7pjB37X-TqparKO4U-4cTfq01kC0',
'TPS-ResNet-BiLSTM-Attn-case-sensitive.pth': 'https://drive.google.com/open?id=10XlPutQuvhGR1tPgYwNAxiHNd1AvpbA9',
'TPS-ResNet-BiLSTM-Attn.pth': 'https://drive.google.com/open?id=1m4jUiTLFDkOhYA3ErPiPK9TGgUG_xQxz',
'TPS-ResNet-BiLSTM-CTC.pth': 'https://drive.google.com/open?id=1kZubKJij7hN4rERNnSd2R5e8Yfo9pinq',
}
for k, v in models.items():
doc_id = v[v.find('=')+1:]
# take basic html response not file itself. html response is about security warning.
# !curl -c /tmp/cookies "https://drive.google.com/uc?export=download&id=$doc_id" > /tmp/intermezzo.html
# download actual file using html security response
# !curl -L -b /tmp/cookies "https://drive.google.com$(cat /tmp/intermezzo.html | grep -Po 'uc-download-link" [^>]* href="\K[^"]*' | sed 's/\&/\&/g')" > $k
# !ls -al *.pth
# # !bash download_models.sh
// + pycharm={"name": "#%% bash\n"}
# !MKL_SERVICE_FORCE_INTEL=1
# !MKLTHREADING_LAYER=INTEL
# !export MKL_SERVICE_FORCE_INTEL=1
# !export MKLTHREADING_LAYER=INTEL
# output = !CUDA_VISIBLE_DEVICES=0 python3 demo.py \
# --Transformation TPS --FeatureExtraction ResNet --SequenceModeling BiLSTM --Prediction Attn \
# --image_folder demo_image/ \
# --saved_model TPS-ResNet-BiLSTM-Attn.pth
# # output = !CUDA_VISIBLE_DEVICES=0 python3 demo.py \
# # --Transformation TPS --FeatureExtraction ResNet --SequenceModeling BiLSTM --Prediction Attn \
# # --image_folder /home/selcuk/Desktop/idcard/idcard_crops/ \
# # --saved_model TPS-ResNet-BiLSTM-Attn.pth
print(output)
// + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="bJRPln2QlxlJ" outputId="a81e0b6b-8171-49a6-c59f-15649d7c6bb8" pycharm={"name": "#%% python\n"}
from IPython.core.display import display, HTML
from PIL import Image
import base64
import io
import pandas as pd
data = pd.DataFrame()
for ind, row in enumerate(output[output.index(
'image_path \tpredicted_labels \tconfidence score')+2:]):
row = row.split('\t')
filename = row[0].strip()
label = row[1].strip()
conf = row[2].strip()
img = Image.open(filename)
img_buffer = io.BytesIO()
img.save(img_buffer, format="PNG")
imgStr = base64.b64encode(img_buffer.getvalue()).decode("utf-8")
data.loc[ind, 'img'] = '<img src="data:image/png;base64,{0:s}">'.format(imgStr)
data.loc[ind, 'id'] = filename
data.loc[ind, 'label'] = label
data.loc[ind, 'conf'] = conf
html_all = data.to_html(escape=False)
display(HTML(html_all))
|
demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="pn6Z-EQkmVos"
# # Flight Delay Predictions
# + [markdown] id="OG7QXsh5xQ0M"
# <img src="https://www.webintravel.com/wp-content/uploads/2020/05/den-belitsky-GettyImages-854673918-scaled.jpg" width="700">
# + [markdown] id="nSxBf4--gC0n"
# ### Link to Dataset: [flights.csv](https://www.kaggle.com/usdot/flight-delays?select=flights.csv)
# + [markdown] id="5ffM0ge7FcKp"
# ### Importing Libraries
# + id="B8VRPIqIp8_V"
import pandas as pd
import seaborn as sb
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import roc_auc_score
# + [markdown] id="ZckrxAZjFiC0"
# ### Getting Data
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="bX5VWtiDnlR8" outputId="4c728ed8-89ea-4357-b75e-fdf172ee0579"
flights = pd.read_csv('flights.csv')
flights
# + [markdown] id="5djkFMqsGh8x"
# ### Insight into the Data
# + id="R7O_ogZ8p7F7"
# selecting a section from the dataframe
flights_needed_data = flights[0:100000]
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="cikjVPHFvEqT" outputId="328386c4-d46c-481b-e160-a8550bf0f5b4"
flights_needed_data
# + colab={"base_uri": "https://localhost:8080/"} id="wlm3Y2ePqnU6" outputId="5074fb81-9db9-47d2-ee36-18a8bbf96828"
flights_needed_data.info()
# + colab={"base_uri": "https://localhost:8080/"} id="0gyL1I0vq2y2" outputId="7b1bed43-e267-4248-c21b-a0ac21e1eaba"
# no. of flights which were diverted
flights_needed_data.value_counts('DIVERTED')
# + [markdown] id="H3LsxmwUG46-"
# ### Data Visualisation
# + colab={"base_uri": "https://localhost:8080/", "height": 458} id="I8tU0O_ArHtb" outputId="5768bb85-23cb-474d-ea16-669d2805cc6d"
sb.jointplot(data=flights_needed_data, x="SCHEDULED_ARRIVAL", y="ARRIVAL_TIME")
# + id="aSRS4HrFwxZF"
# using Pearson's correlation method
corr = flights_needed_data.corr(method='pearson')
# + colab={"base_uri": "https://localhost:8080/", "height": 390} id="J9_L4K8Zwu5B" outputId="1bcf2276-9af6-4094-b706-29e4bd68d0cf"
sb.heatmap(corr)
# + colab={"base_uri": "https://localhost:8080/", "height": 837} id="YhgZYhwKxHJJ" outputId="e438d733-a2ad-4b26-e4fe-a6b49301f47a"
corr
# + [markdown] id="pnx2TjlzHMEc"
# ### Data Preprocessing
# + id="pftA6NKayM4h"
flights_needed_data=flights_needed_data.drop(['YEAR','FLIGHT_NUMBER','AIRLINE','DISTANCE','TAIL_NUMBER','TAXI_OUT',
'SCHEDULED_TIME','DEPARTURE_TIME','WHEELS_OFF','ELAPSED_TIME',
'AIR_TIME','WHEELS_ON','DAY_OF_WEEK','TAXI_IN','CANCELLATION_REASON'],
axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="oSQQ10_P1gqY" outputId="ed6916f7-3a1a-4464-c850-f221ef48a76f"
flights_needed_data
# + id="I5IBw7ad0uxw"
# replacing NaN values with the mean of the attribute
flights_needed_data=flights_needed_data.fillna(flights_needed_data.mean())
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="IwDGKuZ04ImK" outputId="7681ba8f-fb64-45bc-bf95-10aec3b82919"
flights_needed_data
# + id="BjQ1rhrb1YFd"
result=[]
# + id="C2AFDcSz2bjO"
# if the delay in flight's arrival is more than 15 mins, then it's definitely delayed
for row in flights_needed_data['ARRIVAL_DELAY']:
if row > 15:
result.append(1)
else:
result.append(0)
# + id="Kh0J66aj3EzK"
flights_needed_data['result'] = result
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="uJ4QW_k33ZNu" outputId="7a0e9576-26a4-4217-81b4-8792712bfe55"
flights_needed_data
# + colab={"base_uri": "https://localhost:8080/"} id="krPajIiZ3_G5" outputId="91845772-d06b-404c-8bc8-fe64871403a7"
flights_needed_data.value_counts('result')
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="Ij79v7Y3_RgW" outputId="87325518-f689-42cc-b978-633e9169dda6"
flights_needed_data=flights_needed_data.drop(['ORIGIN_AIRPORT', 'DESTINATION_AIRPORT', 'ARRIVAL_TIME', 'ARRIVAL_DELAY'],axis=1)
flights_needed_data
# + [markdown] id="K3FRiemNH1ml"
# ### Splitting Data for Training and Testing
# + id="GwAUjhg-4Zn6"
data = flights_needed_data.values
X, y = data[:,:-1], data[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42) # splitting in the ratio 70:30
# + [markdown] id="mQgSjXYFudyY"
# ### Standardizing
# + id="6XsHnDLP7CFs"
scaled_features = StandardScaler().fit_transform(X_train, X_test)
# + [markdown] id="FOOgER-OujcI"
# ### Applying Decision Tree Classifier on Training Data
# + id="0AIkZKNdBEWS"
clf = DecisionTreeClassifier()
clf = clf.fit(X_train,y_train)
# + [markdown] id="mwXLSatCusMb"
# ### Making Predictions and Checking Accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="E3CWbnZOB2M0" outputId="3ec45653-a3ec-4ff0-d32f-343d8845d3cd"
pred_prob = clf.predict_proba(X_test)
auc_score = roc_auc_score(y_test, pred_prob[:,1])
auc_score
# + [markdown] id="8-zrkG_Hwsa7"
# # Predictions are 99.77% accurate.
|
Python/ml/Flight_Delay_Predictions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wiener Filter Implementation in TensorFlow
#
# To implement Wiener Filter functions, we are going to mimic its implementation in Scikit Image (see skimage.restoration.wiener) and in Tikhonet (see https://github.com/CosmoStat/ShapeDeconv/blob/master/python/DeepDeconv/utils/deconv_utils_FCS.py).
#
# ## Implementation in skimage:
# +
from skimage.restoration import uft
import numpy as np
def wiener(image, psf, balance, reg=None, is_real=True, clip=True):
if reg is None:
reg, _ = uft.laplacian(image.ndim, image.shape, is_real=is_real)
if not np.iscomplexobj(reg):
reg = uft.ir2tf(reg, image.shape, is_real=is_real)
if psf.shape != reg.shape:
trans_func = uft.ir2tf(psf, image.shape, is_real=is_real)
else:
trans_func = psf
wiener_filter = np.conj(trans_func) / (np.abs(trans_func) ** 2 +
balance * np.abs(reg) ** 2)
if is_real:
deconv = uft.uirfft2(wiener_filter * uft.urfft2(image),
shape=image.shape)
else:
deconv = uft.uifft2(wiener_filter * uft.ufft2(image))
if clip:
deconv[deconv > 1] = 1
deconv[deconv < -1] = -1
return deconv
#USAGE CASE OF tikhonov:
#deconv_img = tikhonov(noisy_img, psfs, 1./SNR_list)
def tikhonov(X, psf, tau):
if len(X.shape) == 2:
return(wiener(X, psf, tau))
tikho_list = []
for i in range(len(X)):
deconvolved = wiener(X[i], psf[i],tau[i])
tikho_list.append(deconvolved)
return np.asarray(tikho_list)
# -
# ## Minimal Required Parts of the `wiener` function:
def wiener(image, psf, balance):
trans_func = uft.ir2tf(psf, image.shape)
wiener_filter = np.conj(trans_func) / (np.abs(trans_func) ** 2 + balance)
deconv = uft.uirfft2(wiener_filter * uft.urfft2(image),
shape=image.shape)
deconv[deconv > 1] = 1
deconv[deconv < -1] = -1
return deconv
# ## TensorFlow Implementation
# +
import tensorflow as tf
def wiener_tf(image, psf, balance):
trans_func = tf.signal.rfft2d(tf.signal.ifftshift(psf))
wiener_filter = tf.math.conj(trans_func) / (tf.dtypes.cast(tf.math.abs(trans_func),'complex64') ** 2 + balance)
deconv = tf.signal.irfft2d(wiener_filter * tf.signal.rfft2d(image),fft_length=image.shape)
deconv = tf.keras.backend.clip(deconv, -1, 1)
return deconv
def tikhonov_tf(img, psf, tau):
if len(img.shape) == 2:
return(wiener_tf(img, psf, tau))
tikho_list = []
for i in range(img.shape[0]):
deconvolved = wiener_tf(img[i], psf[i], tau[i])
tikho_list += [deconvolved]
return tikho_list
# -
# ## Basic Test
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# create mock data
images = np.zeros((2,128,128))
psf = np.zeros((128,128))
# create square and rectangle images
size = 10
# square image
images[0,62-size:62+size,62-size:62+size] = 1
# rectangle image
images[1,62-5:62+5,62-3*size:62+3*size] = 1
# psf image
psf[43:48,65:70] = 1
psf[48:55,55:72] = 1
# show images
plt.figure(1)
plt.subplot(131)
plt.imshow(images[0])
plt.axis('off')
plt.title('square')
plt.subplot(132)
plt.imshow(images[1])
plt.axis('off')
plt.title('rectangle')
plt.subplot(133)
plt.imshow(psf)
plt.axis('off')
plt.title('psf')
# +
# deconvolved numpy images
tau = [1,1]
deconv_images_np = tikhonov(images, [psf,psf], tau)
# plot deconvolved images
plt.figure(2)
plt.subplot(121)
plt.imshow(deconv_images_np[0])
plt.colorbar()
plt.axis('off')
plt.title('deconvolved square')
plt.subplot(122)
plt.imshow(deconv_images_np[1])
plt.colorbar()
plt.axis('off')
plt.title('deconvolved rectangle')
# +
# convert arrays to tensors
images_tf = tf.convert_to_tensor(images,dtype='float32')
psf_tf = tf.convert_to_tensor(psf,dtype='float32')
# deconvolved tf images
deconv_images = tikhonov_tf(images_tf, [psf_tf,psf_tf], tau)
dec_1 = tf.keras.backend.eval(deconv_images[0])
dec_2 = tf.keras.backend.eval(deconv_images[1])
# plot deconvolved tensorflow images
plt.figure(3)
plt.subplot(121)
plt.imshow(dec_1)
plt.colorbar()
plt.axis('off')
plt.title('deconvolved TF square')
plt.subplot(122)
plt.imshow(dec_2)
plt.colorbar()
plt.axis('off')
plt.title('deconvolved TF rectangle')
# -
# ## Compare Test Results
# plot the difference of the deconvolved images
plt.figure(4)
plt.subplot(121)
plt.imshow(dec_1-deconv_images_np[0])
plt.colorbar()
plt.axis('off')
plt.title('differences 1')
plt.subplot(122)
plt.imshow(dec_2-deconv_images_np[1])
plt.colorbar()
plt.axis('off')
plt.title('differences 2')
# ## Fourier Transform Tensorflow Test
# +
square_fft = tf.signal.rfft2d(images_tf[0])
square_ifft = tf.signal.irfft2d(square_fft)
fft_array = tf.keras.backend.eval(square_fft)
ifft_array = tf.keras.backend.eval(tf.dtypes.cast(square_ifft,'float32'))
square_fft_np = uft.ir2tf(images[0], psf.shape)#np.fft.fft2(images[0])#
square_ifft_np = uft.uirfft2(square_fft_np, psf.shape)
# plot the real fft and ifft of the square
plt.figure(5, figsize=[10,10])
plt.subplot(231)
plt.imshow(np.abs(fft_array))
plt.colorbar()
plt.axis('off')
plt.title('square tf fft amp')
plt.subplot(232)
plt.imshow(np.angle(fft_array))
plt.colorbar()
plt.axis('off')
plt.title('square tf fft ang')
plt.subplot(233)
plt.imshow(ifft_array)
plt.colorbar()
plt.axis('off')
plt.title('square tf reconstruction')
plt.subplot(234)
plt.imshow(np.abs(square_fft_np))
plt.colorbar()
plt.axis('off')
plt.title('square uft fft amp')
plt.subplot(235)
plt.imshow(np.angle(square_fft_np))
plt.colorbar()
plt.axis('off')
plt.title('square uft fft ang')
plt.subplot(236)
plt.imshow(np.real(square_ifft_np))
plt.colorbar()
plt.axis('off')
plt.title('square uft reconstruction')
# -
# ## Wiener Filter Test
trans_func = uft.ir2tf(psf, psf.shape)
wiener_filter = np.conj(trans_func) / (np.abs(trans_func) ** 2 + 1)
wiener_recons = uft.uirfft2(wiener_filter,shape=psf.shape)
trans_func = tf.signal.rfft2d(psf_tf)
wiener_filter_tf = tf.math.conj(trans_func) / (tf.dtypes.cast(tf.square(tf.math.abs(trans_func)),'complex64') + 1)
wiener_filter_tf_np = tf.keras.backend.eval(wiener_filter_tf)
wiener_recons_tf = tf.signal.irfft2d(wiener_filter_tf,fft_length=psf_tf.shape)
wiener_recons_tf_np = tf.keras.backend.eval(wiener_recons_tf)
# plot the difference of the deconvolved images
plt.figure(6,figsize=[10,10])
plt.subplot(231)
plt.imshow(np.abs(wiener_filter))
plt.colorbar()
plt.axis('off')
plt.title('numpy filter amp')
plt.subplot(232)
plt.imshow(np.abs(wiener_filter_tf_np))
plt.colorbar()
plt.axis('off')
plt.title('tensorflow filter amp')
plt.subplot(233)
plt.imshow(np.log(np.abs(wiener_recons)))
plt.colorbar()
plt.axis('off')
plt.title('numpy filter recons')
plt.subplot(234)
plt.imshow(np.angle(wiener_filter))
plt.colorbar()
plt.axis('off')
plt.title('numpy filter ang')
plt.subplot(235)
plt.imshow(np.angle(wiener_filter_tf_np))
plt.colorbar()
plt.axis('off')
plt.title('tensorflow filter ang')
plt.subplot(236)
plt.imshow(np.log(np.abs(wiener_recons_tf_np)))
plt.colorbar()
plt.axis('off')
plt.title('tensorflow filter recons')
|
notebooks/Tikhonov_filter/WienerTensorFlow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Goal of this is to look at the sentences and correlations between correct predictions and other data fields.
# +
from keras.models import load_model
# Put any of the models here
blstm_model = load_model('./models/atta_blstm_dropout6.hdf5')
# +
from utils import *
from models import blstm_dropout
from keras.preprocessing.text import Tokenizer
from keras.utils import to_categorical
import pandas as pd
import numpy as np
atta_df = pd.read_csv('/scratch/gussteen/final_project/attasidor.csv')
atta_df['word'] = atta_df['word'].astype(str)
atta_all_sents = atta_df.groupby('sent_id')['word'].apply(lambda x: ' '.join(x))
print("Total sentences:", len(atta_all_sents))
max_words = 10000
tokenizer = Tokenizer(num_words=max_words, oov_token='UNK', filters='–—!"#$%&()*+,-./:;<=>?@[\]^_`{|}~')
tokenizer.fit_on_texts(atta_all_sents)
word_lookup = {v: k for k, v in tokenizer.word_index.items()}
sentence_list = atta_df.groupby('sent_id')['word'].apply(list)
pos_list = atta_df.groupby('sent_id')['pos'].apply(list)
X_ids = []
X_before = []
X_after = []
y = []
for words, w_pos, s_id in zip(list(sentence_list), list(pos_list), list(sentence_list.index)):
for ex_id, before, after, w in create_training_example(words, w_pos, s_id, tokenizer):
X_ids.append(ex_id)
X_before.append(before)
X_after.append(after)
y.append(w)
X_ids = np.array(X_ids)
X_before = np.array(X_before)
X_after = np.array(X_after)
y_cat = to_categorical(y, num_classes = max_words + 1)
# -
# Get the predictions for the training data
y_preds = blstm_model.predict([X_before, X_after])
cross_ent = - np.sum(y_cat * np.log(y_preds), axis = 1)
cross_ent.shape
pred_word = np.argmax(y_preds, axis=1)
pred_word
results_df = pd.DataFrame()
results_df['pred_word'] = pred_word
results_df['actual_word'] = y
results_df['cross_entropy'] = cross_ent
results_df['sent_id'] = X_ids
#results_df.index = X_ids
results_df['actual_word'] = results_df['actual_word'].apply(lambda i: word_lookup[i[0]])
results_df['pred_word'] = results_df['pred_word'].apply(lambda i: word_lookup[i])
sent_df = pd.DataFrame()
sent_df['sentence'] = atta_all_sents
sent_df['word_count'] = atta_df.groupby('sent_id')['word'].apply(len)
# Compute average cross_entropy for each sentence
sent_df['avg_cross_entropy'] = results_df.groupby('sent_id')['cross_entropy'].mean()
# +
import matplotlib.pyplot as plt
# Correlation and plot for entropy and sentence length
res = sent_df[sent_df['avg_cross_entropy'].notna()][['word_count', 'avg_cross_entropy']]
by_wordcount = res.groupby('word_count')['avg_cross_entropy'].agg(['mean','std'])
# -
# by_wordcount_small = by_wordcount[by_wordcount.index < 25]
plt.scatter(by_wordcount.index, by_wordcount['mean'])
plt.errorbar(by_wordcount.index, by_wordcount['mean'], by_wordcount['std'], ecolor='#d3d3d3')
plt.xlabel('Word Count')
plt.ylabel('Mean Cross Entropy')
plt.title('Cross Entropy by Word Count Group')
plt.savefig('./results/cross_entropy_word_count.png')
#plt.show()
# Gets the minimum entropy for each actual word
min_entropy_word = results_df.loc[results_df.groupby('actual_word')['cross_entropy'].idxmin()]
# Join the sentence
# Get the top 20 predictions as latex table
pd.set_option('display.max_colwidth', -1)
print(min_entropy_word.join(sent_df, on='sent_id').sort_values(by='cross_entropy')[['actual_word', 'sentence']].head(20).to_latex(index=False))
|
Explore Predictions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
random_seed = 42
import csv
import random
random.seed(random_seed)
import numpy as np
np.random.seed(random_seed)
import pandas as pd
pd.set_option('max_colwidth', 256)
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
def read_data(filename):
df = pd.read_json(filename, lines=True)
df.dropna()
####
df = df[['gold_label', 'sentence1', 'sentence2']]
df.columns = ['label', 'sentence1', 'sentence2']
df['label'] = df['label'].map({'contradiction': 0, 'neutral': 0, 'entailment': 1, '-': 3})
df = df[df['label']!=3]
####
df_entail = df[df['label']==1]
df_non_entail = df[df['label']==0].sample(len(df_entail), random_state=42) # 变成二分类
df = df_entail.append(df_non_entail)
####
return df
data_path = "data/SNLI/snli_1.0/"
train = read_data(data_path+"snli_1.0_{}.jsonl".format('train'))
train.info()
train['label'].value_counts()
pool = train.copy()
pool, train = train_test_split(pool, test_size=10000, shuffle=True, random_state=random_seed, stratify=pool['label'])
train.info()
train[train['sentence1'].str.contains("guy") & train['sentence2'].str.contains("male")]
train[train['sentence2'].str.contains("guy") & train['sentence1'].str.contains("male")]
train[train['sentence1'].str.contains("iPod")]
train[train['sentence1'].str.contains("iPod") & train['sentence2'].str.contains("mp3")]
train[train['sentence2'].str.contains("iPod") & train['sentence1'].str.contains("mp3")]
dev = read_data(data_path+"snli_1.0_{}.jsonl".format('dev'))
dev.info()
test = read_data(data_path+"snli_1.0_{}.jsonl".format('test'))
test.info()
train['sample_index'] = list(range(len(train)))
train.head()
train.tail()
train['label'].value_counts()
dev['sample_index'] = list(range(len(dev)))
dev.head()
dev['label'].value_counts()
test['sample_index'] = list(range(len(test)))
test.head()
test['label'].value_counts()
train.to_csv('data/train.csv', index=False)
dev.to_csv('data/dev.csv', index=False)
test.to_csv('data/test.csv', index=False)
total = len(train)
for percentage in range(0, 100, 10):
k = int(total*(percentage/100))
print(percentage, k)
tmp = train.sample(k,
random_state=0
)
tmp = train.drop(tmp.index)
print(tmp['label'].value_counts())
filename = "data/random_0/{}.csv".format(percentage)
os.makedirs(os.path.dirname(filename), exist_ok=True)
tmp[['label', 'sentence1', 'sentence2', 'sample_index']].to_csv(filename, index=False)
total = len(train)
for percentage in range(0, 100, 10):
k = int(total*(percentage/100))
print(percentage, k)
tmp = train.sample(k,
random_state=2
)
tmp = train.drop(tmp.index)
print(tmp['label'].value_counts())
filename = "data/random_2/{}.csv".format(percentage)
os.makedirs(os.path.dirname(filename), exist_ok=True)
tmp[['label', 'sentence1', 'sentence2', 'sample_index']].to_csv(filename, index=False)
total = len(train)
for percentage in range(0, 100, 10):
k = int(total*(percentage/100))
print(percentage, k)
tmp = train.sample(k,
random_state=42
)
tmp = train.drop(tmp.index)
print(tmp['label'].value_counts())
filename = "data/random/{}.csv".format(percentage)
os.makedirs(os.path.dirname(filename), exist_ok=True)
tmp[['label', 'sentence1', 'sentence2', 'sample_index']].to_csv(filename, index=False)
|
snli/.ipynb_checkpoints/00_EDA-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pnlp37
# language: python
# name: pnlp37
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/c-w-m/pnlp/blob/master/Ch02/05_Data_Augmentation_Using_NLPaug.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="yavI9mt4gayF"
# # Chapter 2.5: Data Augmentation Using __NLPaug__
# This notebook demostrate the usage of a character augmenter, word augmenter. There are other types such as augmentation for sentences, audio, spectrogram inputs etc. All of the types many before mentioned types and many more can be found at the [github repo](https://github.com/makcedward/nlpaug) and [docs](https://nlpaug.readthedocs.io/en/latest/) of nlpaug.
# + colab={"base_uri": "https://localhost:8080/"} id="cF5zJdr-kAPY" outputId="b60433c1-18fc-4eef-9ec4-180567a8c426"
#Installing the nlpaug package
# !pip install nlpaug==0.0.14
# + id="8yhkOl3cgZ28"
#this will be the base text which we will be using throughout this notebook
text="The quick brown fox jumps over the lazy dog ."
# + colab={"base_uri": "https://localhost:8080/"} id="ekFhzIWHUmoj" outputId="0259c3f9-5757-4f15-d90f-49e54f55daaf"
import nlpaug.augmenter.char as nac
import nlpaug.augmenter.word as naw
import nlpaug.augmenter.sentence as nas
import nlpaug.flow as nafc
from nlpaug.util import Action
import os
# !git clone https://github.com/makcedward/nlpaug.git
os.environ["MODEL_DIR"] = 'nlpaug/model/'
# + [markdown] id="-Xo3CzNhh-zU"
# ### Augmentation at the Character Level
#
#
# 1. OCR Augmenter: To read textual data from on image, we need an OCR(optical character recognition) model. Once the text is extracted from the image, there may be errors like; '0' instead of an 'o', '2' instead of 'z' and other such similar errors.
# 2. Keyboard Augmenter: While typing/texting typos are fairly common this augmenter simulates the errors by substituting characters in words with ones at a similar distance on a keyboard.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="lfAaokTmjzak" outputId="d7fb5bf6-f695-47bc-f401-cdfa62fd9fbf"
#OCR augmenter
#import nlpaug.augmenter.char as nac
aug = nac.OcrAug()
augmented_texts = aug.augment(text, n=3) #specifying n=3 gives us only 3 augmented versions of the sentence.
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
# + colab={"base_uri": "https://localhost:8080/"} id="fKQCpS35j9Ie" outputId="8b41774f-cc04-4e0a-e337-a7ec28440b55"
#Keyboard Augmenter
#import nlpaug.augmenter.word as naw
aug = nac.KeyboardAug()
augmented_text = aug.augment(text, n=3) #specifying n=3 gives us only 3 augmented versions of the sentence.
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="XbfPMwZWmper"
# There are other types of character augmenters too. Their details are avaiable in the links mentioned at the beginning of this notebook.
# + [markdown] id="MufLJXsQm4i1"
# ### Augmentation at the Word Level
#
# Augmentation is important at the word level as well , here we use word2vec to insert or substitute a similar word.
# + [markdown] id="Tc_K1-niTGFP"
# **Spelling** **augmentor**
#
# + colab={"base_uri": "https://localhost:8080/"} id="2Qzmv4QCYrJe" outputId="565df27c-9972-4c9e-a1da-7940b52af3b2"
#Downloading the required txt file
# !pip install wget
import wget
if not os.path.exists("spelling_en.txt"):
wget.download("https://raw.githubusercontent.com/makcedward/nlpaug/5238e0be734841b69651d2043df535d78a8cc594/nlpaug/res/word/spelling/spelling_en.txt")
else:
print("File already exists")
# + colab={"base_uri": "https://localhost:8080/"} id="gOHrgDIill2F" outputId="f12d1b2a-4afe-49c0-ff03-cfce4ce7dbf0"
#Substitute word by spelling mistake words dictionary
aug = naw.SpellingAug('spelling_en.txt')
augmented_texts = aug.augment(text)
print("Original:")
print(text)
print("Augmented Texts:")
print(augmented_texts)
# + [markdown] id="eaeQOtVqTQKG"
# **Word embeddings augmentor**
# + colab={"base_uri": "https://localhost:8080/"} id="wDrq-v-B5mAX" outputId="e9381e1e-33fb-4322-8e08-ed1d4f106a6d"
import gzip
import shutil
gn_vec_path = "GoogleNews-vectors-negative300.bin"
if not os.path.exists("GoogleNews-vectors-negative300.bin"):
if not os.path.exists("../Ch3/GoogleNews-vectors-negative300.bin"):
#Downloading the reqired model
if not os.path.exists("../Ch3/GoogleNews-vectors-negative300.bin.gz"):
if not os.path.exists("GoogleNews-vectors-negative300.bin.gz"):
wget.download("https://s3.amazonaws.com/dl4j-distribution/GoogleNews-vectors-negative300.bin.gz")
gn_vec_zip_path = "GoogleNews-vectors-negative300.bin.gz"
else:
gn_vec_zip_path = "../Ch3/GoogleNews-vectors-negative300.bin.gz"
#Extracting the required model
with gzip.open(gn_vec_zip_path, 'rb') as f_in:
with open(gn_vec_path, 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
else:
gn_vec_path = "../Ch3/" + gn_vec_path
print(f"Model: {gn_vec_path}")
# + [markdown] id="Jf_QHk-SgegN"
# Insert word randomly by word embeddings similarity
# + colab={"base_uri": "https://localhost:8080/"} id="ffUb6s-XTOsQ" outputId="9a0259fc-da4f-4de4-c008-17b4de20095b"
# model_type: word2vec, glove or fasttext
aug = naw.WordEmbsAug(
model_type='word2vec', model_path=gn_vec_path,
action="insert")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="kUB3Nd4Wghd0"
# Substitute word by word2vec similarity
#
# + colab={"base_uri": "https://localhost:8080/"} id="pSeZNfQRfy2l" outputId="fafd19d5-db59-4da8-f242-5d67ae0f3278"
aug = naw.WordEmbsAug(
model_type='word2vec', model_path=gn_vec_path,
action="substitute")
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
# + [markdown] id="reALNlOuDI9u"
# There are many more features which nlpaug offers you can visit the github repo and documentation for further details
|
Ch02/05_Data_Augmentation_Using_NLPaug.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Exercise 3
# Add the specified code for each code cell, running the cells _in order_.
# Define a function `add_three` that takes a single argument and returns a value 3 greater than the input.
def add_three(number):
return number+3
# Create and output a variable `ten` that is the result of passing `7` to your `add_three()` function.
ten=add_three(7)
print(ten)
# Create a variable `ten_str` that is the result of passing `"7"` to your `add_three()` function. What does this tell you about how the function should be described (e.g., in a doc string)?
ten_str=add_three("7")
argument should be integer
# Define a function `imperial_to_metric` that takes in two arguments: a number of feet and a number of inches. The function should return the total length in meters. _Include an appropriate doc string_.
def imperial_to_metric(num_feet,num_inches):
"""
summary
parameters:
argument1 : number of feet
argument2 : number of inches
returns:
total length in meters
"""
num_feet=num_feet*.3048
num_inches=num_inches*.0254
return num_feet+num_inches
# Create and output variable `height_in_meters` by passing your height in imperial to the `imperial_to_metric()` function.
height_in_meters=imperial_to_metric(5,4)
print(height_in_meters)
# Define a function `compare_str_length` that takes in 2 strings, and returns a sentence of the form
# ```
# "The difference in string lengths is N"
# ```
# Include an appropriate doc string.
#some changes
def compare_str_length(str1,str2):
"""
parameters:str1,str2 two strings
returns: difference in lengths
"""
N=abs(len(str1)-len(str2))
print("The difference in string lengths is ",N)
# Pass two strings of different lengths to your `compare_str_length()` function.
compare_str_length("two","three")
# Define a function `fraction_str()` that takes two parameters, a numerator and a denominator, and outputs a string version of that that fraction (e.g., `"3/4"`). Make the parameters be **keyword arguments** with default values of 1.
def fraction_str(num=1,denom=1):
"""
parameters:two numbers
returns:string version of the fraction
"""
return (str(num)+"/"+str(denom))
# Call the `fraction_str()` function with named arguments to produce the string `"5/11"`. Print the result.
# - For fun: try listing the denominator argument before the numerator argument! What happens?
fraction_str(denom=5,num=11)
# Call the `fraction_str()` function only specifying a denominator of `3`. Print the result
fraction_str(denom=3)
# Call the `fraction_str()` function using **positional arguments** (unnamed) to produce the string `"11/5"`. Print the result.
fraction_str(11,5)
|
exercise-3/exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CENG796 - Project Submission
# This notebook is about our implementation of ["A U-Net Based Discriminator for Generative Adversarial Networks"](https://arxiv.org/abs/2002.12655). (CVPR2020)
#
# All figures and tables in this notebook are from the original paper unless stated otherwise.
# ### Project Members
# <NAME>, 2171593, <EMAIL>
#
# <NAME>, 2310647, <EMAIL>
# ## A U-Net Based Discriminator for Generative Adversarial Networks
# ### Paper Abstract
#
# Among the major remaining challenges for generativeadversarial networks (GANs) is the capacity to synthesize globally and locally coherent images with object shapes and textures indistinguishable from real images. To target this issue we propose an alternative U-Net based discriminator architecture, borrowing the insights from the segmentation literature. The proposed U-Net based architecture allows to provide detailed per-pixel feedback to the generator while maintaining the global coherence of synthesized images, by providing the global image feedback as well. Empowered by the per-pixel response of the discriminator, we further propose a per-pixel consistency regularization technique based on the CutMix data augmentation, encouraging the U-Net discriminator to focus more on semantic and structural changes between real and fake images. This improves the U-Net discriminator training, further enhancing the quality of generated samples. The novel discriminator improves over the state of the art in terms of the standard distribution and image quality metrics, enabling the generator to synthesize images with varying structure, appearance and levels of detail, maintaining global and local realism. Compared to the BigGAN baseline, we achieve an average improvement of 2.7 FID points across FFHQ, CelebA, and the newly introduced COCO-Animals dataset.
# ### Standard GAN Model
#
# A ”vanilla” GAN consists of two networks: a generator $G$ and a discriminator $D$, trained by minimizing the following competing objectives in an alternating manner:
#
# $$ L_{D} = -E_{x}[logD(x)] - E_{z}[log(1 - D(G(z)))] $$
# $$ L_{G} = -E_{z}[logD(G(z))] $$
#
# Note that the $L_{G}$ loss is proposed as non-saturating GAN.
# Ordinarily, $G$ and $D$ are modeled as a decoder and an encoder convolutional network, respectively.
# ### U-Net GAN Model
#
# The authors propose to alter the $D$ architecture from a standard classification network to an encoder-decoder U-Net network. This new $D^{U}$ discriminator is able to make normal image classification at its encoder network $D^{U}_{enc}$, and it also outputs a per-pixel classification label through its decoder network $D^{U}_{dec}$. The generator network is not modified in any major contribution of the paper (BigGAN's generator for 128x128 is taken with minimal modifications). Overall, the U-Net GAN has the following design (Figure 2 in the original paper):
# 
# ### U-Net GAN Losses
#
# The U-Net GAN discriminator is designed as a U-Net $D^{U}$, where an encoder network $D^{U}_{enc}$ and decoder network $D^{U}_{dec}$ are connected via a bottleneck, as well as skip connections that copy and concatenate feature maps from the encoder to the decoder network.
#
# The encoder network classifies the input image as real or fake, and the decoder network performs this classification on a per-pixel basis. The overall decoder loss in this then equal to
# $$ L_{D^{U}} = L_{D^{U}_{enc}} + L_{D^{U}_{dec}} $$
#
# where we have the encoder loss as
# $$ L_{D^{U}_{enc}} = -E_{x}[logD^{U}_{enc}(x)] - E_{z}[log(1 - D^{U}_{enc}(G(z)))] $$
# and the decoder loss as the mean classification over all pixels:
# $$ D^{U}_{dec} = -E_{x}[\sum_{i, j}log[D^{U}_{dec}(x)]_{i, j}] - E_{z}[\sum_{i, j}log(1 - [D^{U}_{dec}(G(z))]_{i,j})] $$
#
# Correspondingly, the generator loss becomes:
# $$ L_{G} = -E_{z}[logD^{U}_{enc}(G(z)) + \sum_{i, j}log[D^{U}_{dec}(G(z))]_{i, j}] $$
# ### Consistency Regularization
#
# The authors propose consistancy regularization of the $D^{U}$ discriminator by encouraging the decoder module $D^{U}_{dec}$ to output preditions under CutMix transformations of real and fake samples.
#
# CutMix augmentation creates synthetic images via cutting and pasting patches from real images and generated synthetic images. In short, a mix ratio $r$ is sampled from $U[0, 1]$, and a mask is obtained by cutting out a patch of a real image and replacing it with another image, thereby maintaining the area ratio $r$. These samples are visualized below (Figure 3 in the original paper):
#
# 
# New training samples $\hat{x}$ are synthesized by mixing real sample $x$ and generated $G(z)$ with a sampled CutMix mask $M$:
#
# $$ \hat{x} = mix(x, G(z), M) $$
# $$ mix(x, G(z), M) = M \odot x + (1 - M) \odot G(z) $$
#
# The mask $M$ is a binary mask indicating if a pixel is from the original image $M_{i, j} = 1$ or from the fake iamge $M_{i, j} = 0$.
#
# When computing loss, CutMix images are treated differently:
# For $D^{U}_{enc}$, CutMix images are labeled as fake ($c = 0$).
# For $D^{U}_{dec}$, CutMix image classification ground truth labels are the mask $M$ itself.
# Following these, the authors introduce the consistency regularization loss term for the discriminator:
# $$ L^{cons}_{D^{U}_{dec}} = || D^{U}_{dec}(mix(x, G(z), M)) - mix(D^{U}_{dec}(x), D^{U}_{dec}(G(z)), M)||^{2}$$
#
# This loss is then added to the overall discriminator loss:
#
# $$ L_{D^{U}} = L_{D^{U}_{enc}} + L_{D^{U}_{dec}} + \lambda L^{cons}_{D^{U}_{dec}} $$
# where the authors find $\lambda = 1$ to be a good hyperparameter. The generator objective does not change under consistency regularization loss.
# ### Relevant experiments
#
# We focus our attention on the FID results (Section 4.2 - Table 3) reported on the CelebA dataset with cropped 128x128 image resolution.
# | Method | FID | IS |
# |-----------|------|------|
# | U-Net GAN | 2.95 | 3.43 |
#
# Alternatively, the authors also report FID calculated via the PyTorch implementation (on Appendix A - Table S1)
#
# | Method | FID | IS |
# |-----------|------|------|
# | U-Net GAN | 2.03 | 3.33 |
# ## Experiments
# The [experiments](./experiments) directory contains incremental implementations of each part of the paper, such as CutMix training, U-NetGAN discriminator loss, and scaling to 128x128 image sizes.
#
# See [experiments/README](./experiments/README.md) if interested in intermediate results.
#
# The cells below show the results obtained with the final trained network (assumed to be under `resources/trained.pth`, see README.md for a download link. Trained for ~32.5k iterations).
# Generator and Discriminator loss graphs, screenshot from tensorboard.
#
# <img src="./resources/gen.png" alt="generator" width="400"/>
#
# <img src="./resources/disc.png" alt="discriminator" width="400"/>
# +
import torch
import random
import numpy as np
import torchvision
import matplotlib.pyplot as plt
# %matplotlib inline
from unetgan import *
# +
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
def seed_generators(seed):
torch.manual_seed(seed)
random.seed(seed)
np.random.seed(seed)
seed_generators(796)
# +
trained_dis = BigGANDiscriminator().to(device)
trained_gen = BigGANGenerator(latent_dim=140, base_ch_width=64).to(device)
utils.load_pretrained_models(trained_dis, trained_gen, './resources/trained.pth')
# -
noise = torch.randn((4, 140), device=device)
sample = trained_gen(noise)
preview_samples(plt, sample, figure_size=(16,2))
# ## Quantitative evaluation
# Let's compute FID scores. To do this, we need to create a "fake" dataset and compare its statistics with an equivalent "real" dataset. So let's generate a "fake" dataset of 10k images using the trained generator (10k is the suggested minimum as a rule of thumb for FID)
# Create our "fake" dataset for FID stats
utils.create_sample_directory(trained_gen, sample_dir='./celeba_fake_samples',
num_samples=10000, device=device, overwrite=True)
# We should also create a subset of the real dataset which has the same characteristics (cropped size, centering, etc.):
# Create the target "real" dataset
utils.create_celeba_sample_directory(celeba_root_dir='.',
sample_dir='./celeba_real_samples',
num_samples=10000,
overwrite=True)
# We can now compute FID scores using these. Let's use the official PyTorch and TensorFlow implementations of FID (you will need to install `pytorch_fid` using `pip` or clone `TTUR` from GitHub to execute the cells below):
# !python -m pytorch_fid ./celeba_fake_samples ./celeba_real_samples
# !TTUR/fid.py ./celeba_fake_samples ./celeba_real_samples
# Our results are somewhat close to the officially reported FID values on CelebA, and maybe the difference in values can be attributed to the difference in training time (32k vs 800k).
# ### Challenges Encountered
#
# The main challenge that we've encountered is that for the reported FID scores and generated images, the authors train the proposed U-NetGAN (with BigGAN-like discriinator and generator) model for 800.000 iterations using a batch size of 50 given 128x128 resolution images. For us, it has been difficult to allocate computational resources to train and evaluate this model (We couldn't even fit the model + suggested batch size of images into a single GPU with 7GB of memory during training).
#
# For this reason, we had to start by using "lower-budget models" in our experiments to observe the contributions of the paper. In essence, the "U-Net GAN loss" does not require the original full-fledged GAN in the paper to be used, so we used a lesser model (relying on "good old DCGAN") to implement "U-Net GAN loss", per-pixel classification, consistancy loss and evaluation scores while preparing the main model.
#
# After we used these "training wheels" to verify the parts of the model, we started to train the full model. With our effort, we could only train the model for 100k iterations, which we used to report statistics in the Experiments section. The actual results in FID don't seem too bad (`2.03` vs `20.30`), and the qualitative results look "realistic enough" despite the difference in training.
# There was also some confusion on how to evaluate our models, as the original FID score reported in the paper is based on the TensorFlow Inceptionv3 model, and the FID scores are not the same when we use PyTorch. In the end, we decided that we can use both versions to output FID scores, since the authors also provide the PyTorch FID score as well.
|
UNetGAN/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook extracts syntatical features from the queries found in SWC and SQS, returning a data frame containing those features.
# # Import Libraries
# The following block of code loads all libraries needed for this notebook.
# +
import nltk
import os
import pickle
import re
import shlex
import stanza
import subprocess
import time
import pandas as pd
import numpy as np
from nltk import word_tokenize
from nltk.tokenize import SyllableTokenizer
from subprocess import Popen, PIPE
from tqdm import tqdm
# -
# # Declare Functions
#
# The following block of code declares functions used in this notebook.
# +
# This function generates n-grams generated from a string.
#
# param s: is the string passed into this function
# param n: is the n in n-grams
# returns: the n-grams
def generate_ngrams(s, n):
# Convert to lowercases
s = s.lower()
# Replace all none alphanumeric characters with spaces
s = re.sub(r'[^a-zA-Z0-9\s]', ' ', s)
# Break sentence in the token, remove empty tokens
tokens = [token for token in s.split(" ") if token != ""]
# Use the zip function to help us generate n-grams
# Concatentate the tokens into ngrams and return
ngrams = zip(*[tokens[i:] for i in range(n)])
return [" ".join(ngram) for ngram in ngrams]
# -
# # Load Data Sets
#
# This block of code loads the data sets and extracts all unique queries from both.
allSessions = pickle.load( open( "../Data/DataSets/SWC/SWC.p", "rb" ) )
allSessionsSQS = pickle.load( open( "../Data/DataSets/SQS/SQS.p", "rb" ) )
allQueries = allSessions['query'].tolist() + allSessionsSQS['query'].tolist()
setQueries = set(allQueries)
# # Extract D-Level Features
#
# The following block of code extracts D-Level features from each query. This code is extremely slow as it is making system calls which execute another block of code. I have encountered difficulties with getting this code to run before, as COLLINS-PARSER/code is compiled C code that may need to be recompiled to ensure compatibility with processor. The solution is to run the make clean, and then make again. Further information about this suite of code can be found at:
#
# http://www.personal.psu.edu/xxl13/downloads/d-level.html
# +
count = 0
input_file = 'DLA/data/lemmatize_pos_sentences.tagged'
loc_file = '../../data/lemmatize_pos_sentences.tagged'
processor_dict = {
'tokenize': 'gsd',
'pos': 'bnc',
'lemma': 'default'
}
nlp = stanza.Pipeline('en', processors=processor_dict)
from tqdm import tqdm
with tqdm(total = len(setQueries) ) as pbar:
for text in setQueries:
doc = nlp(text)
out = open(input_file, 'w')
for sentence in doc.sentences:
s = ''
l = 0
for word in sentence.words:
s+='{} {}'.format(word.lemma, word.xpos) + ' ' # needs to be xpos so it uses Penn Treebank
l+=1
out.write('{} {}\n'.format(l, s.strip()))
out.close()
cmd = 'cd DLA/d-level-analyzer/COLLINS-PARSER;'
cmd += ' code/parser {} models/model2/grammar 10000 1 1 1 1 > ../../data/parsed.m2;'.format(loc_file)
cmd += 'cd ..;'
cmd += 'python d-level.py ../data/parsed.m2 > ../data/dlevel.dla;'
proc = subprocess.Popen(cmd, stdout=PIPE, stderr=PIPE, shell=True).wait()
if count == 0:
dl = pd.read_csv('DLA/data/dlevel.dla')
dl['query'] = text
dLevel = dl
count += 1
else:
dl = pd.read_csv('DLA/data/dlevel.dla')
dl['query'] = text
dLevel = dLevel.append(dl)
pbar.update()
# -
# # Extract Part of Speech Features
#
# The following block of code first generates part of speech uni-gram, bi-gram, and tri-gram for each query, then takes the top 10 most common bi-grams and top 5 most common tri-grams (was determined be initial research); returning the ratio of all n-grams for each query.
# +
posData = []
for document in setQueries:
text = nltk.word_tokenize(document)
tags = np.array(nltk.pos_tag(text)).flatten()
posData.append(tags[1::2])
posMod = []
for pos in posData:
string = []
for entry in pos:
string += str(entry) + " "
posMod.append("".join(string))
posUni = []
posBi = []
posTri = []
for document in posMod:
doc = generate_ngrams(document,1)
posUni.append(doc)
for document in posMod:
doc = generate_ngrams(document,2)
posBi.append(doc)
for document in posMod:
doc = generate_ngrams(document,3)
posTri.append(doc)
posDF = pd.DataFrame(setQueries)
posDF['all'] = posMod
posDF['uniPos'] = posUni
posDF['biPos'] = posBi
posDF['triPos']= posTri
posDF = posDF.rename(columns={0: "query"})
allSessionsuni = pd.concat([posDF,pd.get_dummies(posDF['uniPos'].apply(pd.Series).stack()).sum(level=0)],axis=1).drop(['uniPos', 'all', 'biPos', 'triPos'],axis=1)
allSessionsbi = pd.concat([posDF,pd.get_dummies(posDF['biPos'].apply(pd.Series).stack()).sum(level=0)],axis=1).drop(['biPos', 'uniPos', 'all', 'triPos'],axis=1)
allSessionstri = pd.concat([posDF,pd.get_dummies(posDF['triPos'].apply(pd.Series).stack()).sum(level=0)],axis=1).drop(['uniPos', 'all', 'biPos', 'triPos'],axis=1)
# -
allSessionsbiLanding = allSessionsbi[[
'nn nn',
'jj nn',
'nn nns',
'to vb',
'jj nns',
'jj to',
'nn in',
'nns in',
'in nn',
'dt nn',
'query']]
allSessionstriLanding = allSessionstri[[
'jj nn nn',
'nn nn nn',
'jj to vb',
'nn nn nns',
'to vb nn',
'query']]
# +
synFeats = allSessionsuni.merge(allSessionsbiLanding)
synFeats = synFeats.merge(allSessionstriLanding)
synFeats = synFeats.merge(allSessionstriLanding)
synFeats = synFeats.fillna(0)
listCols = list(synFeats.columns)
listCols.pop(0) ##removes 'query' from the list of columns
synFeats['length'] = synFeats['query'].str.split().str.len()
for col in listCols:
synFeats[col] = synFeats[col]/synFeats['length']
# -
# # Return Feature Set
#
# Combines all data frames into one, preprocesses out extraneous information and returns the cleaned data frame.
synFeats = synFeats.merge(dLevel, on = 'query')
synFeats.drop(columns = [' Sentences', 'length', 'Filename'], inplace = True)
pickle.dump(synFeats, open( "Pickles/SynFeat.p", "wb" ) )
|
FeatureExtraction/ExtractSyntaticalFeatures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 1: Setting up the Model
# ### Import the required packages.
import numpy as np
import torch
import torch.nn.functional as F
from transformers import GPT2Tokenizer, GPT2LMHeadModel
from random import choice
# ### Load the OpenGPT2 Tokenizer and the Language Model
# download the pre-trained model
tok = GPT2Tokenizer.from_pretrained("gpt2")
model = GPT2LMHeadModel.from_pretrained("gpt2")
# ### Function which can input text to the model and generate a prediction.
#
# 1. tokenizing and encoding the input text from input_ids
# 2. ask our model to generate a logits vector for the next word/token
# 3. applying softmax and sorting these probabilities in descending order
# 4. vector idxs lists the indices of each token in our vocab in order by their respective probabilities
def get_pred(text, model, tok, p = 0.7):
input_ids = torch.tensor(tok.encode(text)).unsqueeze(0) # Natural Language Processing is => [35364, 15417, 28403, 318]
logits = model(input_ids)[0][:, -1] # [all columns, last line] vector of next toekn
probs = F.softmax(logits, dim = -1).squeeze() # [2.9178e-05, 1.6191e-05, ..., 1.0366e-08, 1.6604e-05]
idxs = torch.argsort(probs, descending = True) # [257, 262, 281, ..., 179, 22997, 36173]
res = []
cumsum = 0
for idx in idxs:
res.append(idx) # e.g : 257, ........, 1762
cumsum += probs[idx] # e.g : 0.0009
if cumsum > p:
pred_idx = idxs.new_tensor([choice(res)]) # e.g : 477
break
pred = tok.convert_ids_to_tokens(int(pred_idx)) # Ġeasy
return tok.convert_tokens_to_string(pred) # easy
# ### Let’s test out our prediction function
# Each time, there is a different result which is exactly what we expect. Our prediction function is now ready. Let’s build our web app!
get_pred("Natural Language Processing is", model, tok, p = 0.7)
# # Part 2: Building the Web Application
# ### Implementation
#
# Let’s first import panel and create the text input widget:
import panel as pn
pn.extension() # loading panel's extension for jupyter compatibility
pn.extension(sizing_mode='scale_width')
text_input = pn.widgets.TextInput()
# ### if we execute text_input in jupyter, we get the following:
text_input
generated_text = pn.pane.Markdown(object = text_input.value)
text_input.link(generated_text, value = 'object')
pn.Row(text_input, generated_text)
button = pn.widgets.Button(name="Generate", button_type = "success")
resetBtn = pn.widgets.Button(name="Reset", button_type = "danger")
def click_cb(event):
pred = get_pred(generated_text.object, model, tok)
generated_text.object += pred
def click_reset_btn(event):
text_input.value = ""
button.on_click(click_cb)
resetBtn.on_click(click_reset_btn)
app = pn.Column(text_input, button, resetBtn, generated_text); app
title = pn.pane.HTML('<h1 style="text-align: center;">Text generator</h1>')
desc = pn.pane.HTML("<marquee scrollamount='10'>Welcome to the text generator! enter some starting input text below, click generate</marquee>")
template = """
{% extends base %}
<!-- goes in body -->
{% block postamble %}
<link rel="stylesheet" href="https://stackpath.bootstrapcdn.com/bootstrap/4.3.1/css/bootstrap.min.css">
<style>
body { background-image: url(https://cdn.pixabay.com/photo/2020/01/02/10/15/background-image-4735444_1280.png) }
marquee, p { font-size: 24px; }
</style>
{% endblock %}
<!-- goes in body -->
{% block contents %}
<div class="container">
<div class="row mt-5">
<div class="col-12">
{{ embed(roots.title) }}
</div>
<div class="col-12">
{{ embed(roots.desc) }}
</div>
<div class="col-12">
{{ embed(roots.input_text) }}
</div>
<div class="col-12">
<div class="row">
<div class="col-6">
{{ embed(roots.button) }}
</div>
<div class="col-6">
{{ embed(roots.resetBtn) }}
</div>
</div>
</div>
<div class="col-12 pl-4">
{{ embed(roots.generated_text) }}
</div>
</div>
</div>
{% endblock %}
"""
# +
tmpl = pn.Template(template)
tmpl.add_panel('title', title)
tmpl.add_panel('desc', desc)
tmpl.add_panel('input_text', text_input)
tmpl.add_panel('button', button)
tmpl.add_panel('resetBtn', resetBtn)
tmpl.add_panel('generated_text', generated_text)
tmpl.show()
|
NLP-mini-project-text-generation/text-generation-openAI-GPT2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Merge metadata with ESCO Crosswalk
# <NAME> | 18.05.2021
#
# ## Core Analysis Goal(s)
# 1. merge onet greenness scores and ASHE wage & employment data to onet-esco
# crosswalk at isco 4-digit level
#
# ## Key Insight(s)
# 1. None
# +
import os
import sys
import logging
from pathlib import Path
import numpy as np
import scipy as sp
import statsmodels.api as sm
from statsmodels.formula.api import ols
# %load_ext autoreload
# %autoreload 2
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set_context("paper")
sns.set(rc={'figure.figsize': (16, 9.)})
sns.set_style("ticks")
import pandas as pd
pd.set_option("display.max_rows", 120)
pd.set_option("display.max_columns", 120)
logging.basicConfig(level=logging.INFO, stream=sys.stdout)
# + [markdown] pycharm={"name": "#%% md\n"}
# Define directory structure
# + pycharm={"name": "#%%\n"}
# project directory
abspath = os.path.abspath('')
project_dir = str(Path(abspath).parents[0])
# sub-directories
data_raw = os.path.join(project_dir, "data", "raw")
data_interim = os.path.join(project_dir, "data", "interim")
data_processed = os.path.join(project_dir, "data", "processed")
figure_dir = os.path.join(project_dir, "reports", "figures")
# + [markdown] pycharm={"name": "#%% md\n"}
# Read crosswalk
# + pycharm={"name": "#%%\n"}
crosswalk = pd.read_csv(
os.path.join(data_raw, "mcc_data", "processed", "ESCO_ONET_xwalk_full.csv")
)
# decompose isco 4-digit level
crosswalk["isco_level_1"] = crosswalk["isco_level_4"].astype(str).str[:1].astype(int)
crosswalk["isco_level_2"] = crosswalk["isco_level_4"].astype(str).str[:2].astype(int)
crosswalk["isco_level_3"] = crosswalk["isco_level_4"].astype(str).str[:3].astype(int)
# + pycharm={"name": "#%%\n"}
crosswalk.query("isco_level_4 == 8211")
# + [markdown] pycharm={"name": "#%% md\n"}
# Read greenness data
# + pycharm={"name": "#%%\n"}
greenness = pd.read_excel(
io=os.path.join(data_raw, "onet", "Onet_GreenTask_AppA.xlsx"),
sheet_name="Occupations"
)
greenness.columns
# + [markdown] pycharm={"name": "#%% md\n"}
# Read ASHE employment and earnings data (at 4-digit level)
# + pycharm={"name": "#%%\n"}
uk_employment = pd.read_csv(
os.path.join(data_raw, "mcc_data", "processed",
"linked_data", "ESCO_top_occupations_UK_employment.csv")
)
uk_employment # [["isco_level_4", "employment_share", "employment_count"]]
# + pycharm={"name": "#%%\n"}
uk_earnings = pd.read_csv(
os.path.join(data_raw, "mcc_data", "processed",
"linked_data", "ESCO_occupations_UK_earnings_and_hours_imputed.csv")
)
uk_earnings[["isco_level_4", "total_paid_hours", "annual_earnings"]]
# + pycharm={"name": "#%%\n"}
uk_earnings.query("isco_level_4 == 8211")
# + [markdown] pycharm={"name": "#%% md\n"}
# read onet job zone data
# + pycharm={"name": "#%%\n"}
job_zones = pd.read_csv(
os.path.join(data_raw, "mcc_data", "processed",
"linked_data", "ESCO_occupations_Job_Zones.csv")
)
job_zones
# + [markdown] pycharm={"name": "#%% md\n"}
# left-join greenness index
# + pycharm={"name": "#%%\n"}
df_merged_1 = pd.merge(
left=crosswalk,
right=greenness[[
'onet_code', 'occupation_type', 'n_new_green_tasks',
'n_existing_green_tasks', 'n_non_green_tasks', 'greenness_vona_2018',
'greenness_vona_2018_v2'
]],
on="onet_code",
how="left",
validate="many_to_one"
)
df_merged_1
# -
# left-join earnings data
# + pycharm={"name": "#%%\n"}
df_merged_2 = pd.merge(
left=df_merged_1,
right=uk_earnings[["concept_uri", "total_paid_hours", "annual_earnings"]],
on="concept_uri",
how="left"
)
fill_values = {
"occupation_type": "Non Green",
# "n_new_green_tasks": 0,
# "n_existing_green_tasks": 0,
# "n_non_green_tasks": 0,
# "greenness_vona_2018": 0
}
# fill empty values of green economy programme cols
df_merged_2.fillna(value=fill_values, inplace=True)
# + [markdown] pycharm={"name": "#%% md\n"}
# left-join employment data
# + pycharm={"name": "#%%\n"}
uk_employment[["concept_uri", "employment_share", "employment_count"]]
# + pycharm={"name": "#%%\n"}
df_merged_3 = pd.merge(
left=df_merged_2,
right=uk_employment[["concept_uri", "employment_share", "employment_count"]],
on="concept_uri",
how="left"
)
df_merged_3
# + [markdown] pycharm={"name": "#%% md\n"}
# left-join job zone data
# + pycharm={"name": "#%%\n"}
job_zones
# + pycharm={"name": "#%%\n"}
df_merged_4 = pd.merge(
left=df_merged_3,
right=job_zones[["concept_uri", "job_zone", "education_level", "related_work_experience", "on_the_job_training"]],
on="concept_uri",
how="left"
)
df_merged_4
# -
# Read ILO green transition scenario data
# + pycharm={"name": "#%%\n"}
fpath_ilo = os.path.join(data_raw, "ilo_scenarios", "ilo_empl_scenarios_2030.xlsx")
ilo_scenarios = ["energy_sustainability", "circular_economy"]
df_ilo = pd.read_excel(
io=fpath_ilo,
sheet_name=ilo_scenarios
)
# convert to millions and calculate net employment change
scale_factor = 1000
numeric_cols = ['new_jobs_absorbing', 'new_jobs_net', 'jobs_lost_net', 'jobs_lost_reallocated', 'net_change']
for ilo_scenario in ilo_scenarios:
df_ilo[ilo_scenario][numeric_cols] *= scale_factor
# df_ilo[ilo_scenario]["net_change"] = df_ilo[ilo_scenario].new_jobs_net + df_ilo[ilo_scenario].jobs_lost_net
# + pycharm={"name": "#%%\n"}
df_merged_5 = pd.merge(
left=df_merged_4,
right=df_ilo["energy_sustainability"].add_suffix('_ilo_2030_es'),
left_on="isco_level_2",
right_on="isco_code_ilo_2030_es",
how="left"
).drop(
columns=["isco_code_ilo_2030_es", "isco_occupation_ilo_2030_es"]
).rename(columns={"skill_level_ilo_2030_es": "skill_level_ilo"})
# + pycharm={"name": "#%%\n"}
df_merged_6 = pd.merge(
left=df_merged_5,
right=df_ilo["circular_economy"].add_suffix('_ilo_2030_ce'),
left_on="isco_level_2",
right_on="isco_code_ilo_2030_ce",
how="left"
).drop(
columns=["isco_code_ilo_2030_ce", "isco_occupation_ilo_2030_ce", "skill_level_ilo_2030_ce"]
)
# + pycharm={"name": "#%%\n"}
df_merged_6.to_csv(
os.path.join(data_interim, "ESCO_ONET_METADATA.csv")
)
# + pycharm={"name": "#%%\n"}
df_merged_6.columns
# + [markdown] pycharm={"name": "#%% md\n"}
# Explore relationships
# + pycharm={"name": "#%%\n"}
df = df_merged_6.copy()
df.greenness_vona_2018.isna().value_counts()
# + pycharm={"name": "#%%\n"}
# check if size of UK labour force meaningful: fine!
df[["employment_share", "employment_count"]].sum()
# + pycharm={"name": "#%%\n"}
# reduce to isco 2-digit means
cols = ['isco_level_2', 'greenness_vona_2018', 'greenness_vona_2018_v2', 'total_paid_hours',
'annual_earnings', 'employment_share', 'employment_count', 'job_zone',
'education_level', 'related_work_experience', 'on_the_job_training',
'net_change_ilo_2030_es', 'net_change_ilo_2030_ce']
metadata_isco_02 = df[cols].groupby("isco_level_2").mean() #.sort_values("greenness_vona_2018")
metadata_isco_02
# + pycharm={"name": "#%%\n"}
sns.regplot(
x="greenness_vona_2018_v2",
y="education_level",
robust=False,
data=metadata_isco_02
)
# + pycharm={"name": "#%%\n"}
from src.visualization.visualize import correlation_matrix_plot
correlation_matrix_plot(metadata_isco_02, figsize=(10,10), significance_level=0.01)
|
notebooks/04-fz-merge-metadata-to-esco-crosswalk.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # OT for image color adaptation with mapping estimation
#
# OT for domain adaptation with image color adaptation [6] with mapping
# estimation [8].
#
# [6] <NAME>., <NAME>., <NAME>., & <NAME>. (2014). Regularized
# discrete optimal transport. SIAM Journal on Imaging Sciences, 7(3), 1853-1882.
#
# [8] <NAME>, <NAME>, <NAME>, <NAME>, "Mapping estimation for
# discrete optimal transport", Neural Information Processing Systems (NIPS), 2016.
#
# +
# Authors: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: MIT License
# sphinx_gallery_thumbnail_number = 3
import os
from pathlib import Path
import numpy as np
from matplotlib import pyplot as plt
import ot
rng = np.random.RandomState(42)
def im2mat(img):
"""Converts and image to matrix (one pixel per line)"""
return img.reshape((img.shape[0] * img.shape[1], img.shape[2]))
def mat2im(X, shape):
"""Converts back a matrix to an image"""
return X.reshape(shape)
def minmax(img):
return np.clip(img, 0, 1)
# -
# ## Generate data
#
#
# +
# Loading images
this_file = os.path.realpath('__file__')
data_path = os.path.join(Path(this_file).parent.parent.parent, 'data')
I1 = plt.imread(os.path.join(data_path, 'ocean_day.jpg')).astype(np.float64) / 256
I2 = plt.imread(os.path.join(data_path, 'ocean_sunset.jpg')).astype(np.float64) / 256
X1 = im2mat(I1)
X2 = im2mat(I2)
# training samples
nb = 500
idx1 = rng.randint(X1.shape[0], size=(nb,))
idx2 = rng.randint(X2.shape[0], size=(nb,))
Xs = X1[idx1, :]
Xt = X2[idx2, :]
# -
# ## Domain adaptation for pixel distribution transfer
#
#
# +
# EMDTransport
ot_emd = ot.da.EMDTransport()
ot_emd.fit(Xs=Xs, Xt=Xt)
transp_Xs_emd = ot_emd.transform(Xs=X1)
Image_emd = minmax(mat2im(transp_Xs_emd, I1.shape))
# SinkhornTransport
ot_sinkhorn = ot.da.SinkhornTransport(reg_e=1e-1)
ot_sinkhorn.fit(Xs=Xs, Xt=Xt)
transp_Xs_sinkhorn = ot_sinkhorn.transform(Xs=X1)
Image_sinkhorn = minmax(mat2im(transp_Xs_sinkhorn, I1.shape))
ot_mapping_linear = ot.da.MappingTransport(
mu=1e0, eta=1e-8, bias=True, max_iter=20, verbose=True)
ot_mapping_linear.fit(Xs=Xs, Xt=Xt)
X1tl = ot_mapping_linear.transform(Xs=X1)
Image_mapping_linear = minmax(mat2im(X1tl, I1.shape))
ot_mapping_gaussian = ot.da.MappingTransport(
mu=1e0, eta=1e-2, sigma=1, bias=False, max_iter=10, verbose=True)
ot_mapping_gaussian.fit(Xs=Xs, Xt=Xt)
X1tn = ot_mapping_gaussian.transform(Xs=X1) # use the estimated mapping
Image_mapping_gaussian = minmax(mat2im(X1tn, I1.shape))
# -
# ## Plot original images
#
#
# +
plt.figure(1, figsize=(6.4, 3))
plt.subplot(1, 2, 1)
plt.imshow(I1)
plt.axis('off')
plt.title('Image 1')
plt.subplot(1, 2, 2)
plt.imshow(I2)
plt.axis('off')
plt.title('Image 2')
plt.tight_layout()
# -
# ## Plot pixel values distribution
#
#
# +
plt.figure(2, figsize=(6.4, 5))
plt.subplot(1, 2, 1)
plt.scatter(Xs[:, 0], Xs[:, 2], c=Xs)
plt.axis([0, 1, 0, 1])
plt.xlabel('Red')
plt.ylabel('Blue')
plt.title('Image 1')
plt.subplot(1, 2, 2)
plt.scatter(Xt[:, 0], Xt[:, 2], c=Xt)
plt.axis([0, 1, 0, 1])
plt.xlabel('Red')
plt.ylabel('Blue')
plt.title('Image 2')
plt.tight_layout()
# -
# ## Plot transformed images
#
#
# +
plt.figure(2, figsize=(10, 5))
plt.subplot(2, 3, 1)
plt.imshow(I1)
plt.axis('off')
plt.title('Im. 1')
plt.subplot(2, 3, 4)
plt.imshow(I2)
plt.axis('off')
plt.title('Im. 2')
plt.subplot(2, 3, 2)
plt.imshow(Image_emd)
plt.axis('off')
plt.title('EmdTransport')
plt.subplot(2, 3, 5)
plt.imshow(Image_sinkhorn)
plt.axis('off')
plt.title('SinkhornTransport')
plt.subplot(2, 3, 3)
plt.imshow(Image_mapping_linear)
plt.axis('off')
plt.title('MappingTransport (linear)')
plt.subplot(2, 3, 6)
plt.imshow(Image_mapping_gaussian)
plt.axis('off')
plt.title('MappingTransport (gaussian)')
plt.tight_layout()
plt.show()
|
master/_downloads/acdb5c8d9a410d04b44379453a1620f0/plot_otda_mapping_colors_images.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Connecting Networks
# **scikit-rf** supports the connection of arbitrary ports of N-port networks. It accomplishes this using an algorithm called sub-network growth[[1]](#References), available through the function `connect()`. Note that this function takes into account port impedances. If two connected ports have different port impedances, an appropriate impedance mismatch is inserted. This capability is illustrated here with situations often encountered.
import skrf as rf
# ## Cascading 2-port and 1-port Networks
# A common problem is to connect two Networks one to the other, also known as cascading Networks, which creates a new Network. The figure below illustrates sile simple situations, where the port numbers are identified in gray:
#
# <img src="figures/networks_connecting_2_2ports.svg" width="600">
#
# or,
#
# <img src="figures/networks_connecting_2port_1port.svg" width="600">
#
#
# Let's illustrate this by connecting a transmission line (2-port Network) to a short-circuit (1-port Network) to create a delay short (1-port Network):
#
# <img src="figures/networks_delay_short.svg" width="600">
#
# Cascading Networks being a frequent operation, it can done conveniently through the `**` operator or with the `cascade` function:
# +
line = rf.data.wr2p2_line # 2-port
short = rf.data.wr2p2_short # 1-port
delayshort = line ** short # --> 1-port Network
print(delayshort)
# -
# or, equivalently using the `cascade()` function:
delayshort2 = rf.cascade(line, short)
print(delayshort2 == delayshort) # the result is the same
# It is of course possible to connect two 2-port Networks together using the `connect()` function. The `connect()` function requires the Networks and the port numbers to connect together. In our example, the port 1 of the line is connected to the port 0 of the short:
delayshort3 = rf.connect(line, 1, short, 0)
print(delayshort3 == delayshort)
# One often needs to cascade a chain Networks together:
#
# <img src="figures/networks_connecting_N_2ports.svg" width="700">
# or,
# <img src="figures/networks_connecting_N_2ports_1port.svg" width="700">
#
#
# which can be realized using chained `**` or the convenient function `cascade_list`:
# +
line1 = rf.data.wr2p2_line # 2-port
line2 = rf.data.wr2p2_line # 2-port
line3 = rf.data.wr2p2_line # 2-port
line4 = rf.data.wr2p2_line # 2-port
short = rf.data.wr2p2_short # 1-port
chain1 = line1 ** line2 ** line3 ** line4 ** short
chain2 = rf.cascade_list([line1, line2, line3, line4, short])
print(chain1 == chain2)
# -
# ## Cascacing 2N-port Networks
# The cascading operator `**` also works for to 2N-port Networks, width the following port scheme:
#
# <img src="figures/networks_connecting_2_2Nports.svg" width="600">
#
# It also works for multiple 2N-port Network. For example, assuming you want to cascade three 4-port Network `ntw1`, `ntw2` and `ntw3`, you can use:
# ```
# resulting_ntw = ntw1 ** ntw2 ** ntw3
# ```
# This is illustrated in [this example on balanced Networks](../examples/networktheory/Balanced%20Network%20De-embedding.ipynb).
# ## Cascading Multi-port Networks
# To make specific connections between multi-port Networks, two solutions are available, which mostly depends of the complexity of the circuit one wants to build:
#
# * For reduced number of connection(s): the `connect()` function
#
# * For advanced connections between many arbitrary N-port Networks, the `Circuit` object is more relevant since it allows defining explicitly the connections between ports and Networks. For more information, please refer to the [Circuit documentation](Circuit.ipynb).
#
# As an example, terminating one of the port of an a 3-port Network, such as an ideal 3-way splitter:
#
# <img src="figures/networks_connecting_3port_1port.svg" width="600">
#
# can be done like:
tee = rf.data.tee
# To connect port `1` of the tee, to port `0` of the delay short,
#
terminated_tee = rf.connect(tee, 1, delayshort, 0)
terminated_tee
# In the previous example, the port #2 of the 3-port Network `tee` becomes the port #1 of the resulting 2-port Network.
# ## Multiple Connections of Multi-port Networks
# Keeping track of the port numbering when using multiple time the `connect` function can be tedious (this is the reason why the [Circuit object](Circuit.ipynb) can be simpler to use).
#
# Let's illustrate this with the following example: connecting the port #1 of a tee-junction (3-port) to the port #0 of a transmission line (2-port):
#
# <img src="figures/networks_connecting_3port_2port.svg" width="600">
#
# To keep track of the port scheme after the connection operation, let's change the port characteristic impedances (in red in the figure above):
tee.z0 = [1, 2, 3]
line.z0 = [10, 20]
# the resulting network is:
rf.connect(tee, 1, line, 0)
# ## References
#
#
# [1] <NAME>.; , "Perspectives in microwave circuit analysis," Circuits and Systems, 1989., Proceedings of the 32nd Midwest Symposium on , vol., no., pp.716-718 vol.2, 14-16 Aug 1989. URL: http://ieeexplore.ieee.org/stamp/stamp.jsp?tp=&arnumber=101955&isnumber=3167
|
doc/source/tutorials/Connecting_Networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pcsilcan/aed/blob/master/IdeasTF.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="MkJZa_k-fFzU" colab_type="code" colab={}
# %%writefile tableTest.cpp
#include <iostream>
#include "table.h"
void Table::indexar(string nombreColumna) {
if (columnas[nombreColumna].getTipo() == "int") { // yes, columnas es un map.
auto k = [=](Fila* fila) { return fila.getInt(nombreColumna); };
arboles[nombreColumna] = new ArbolInt(k);
} else if (columnas[nombreColumna].getTipo() == "string") {
auto k = [=](Fila* fila) { return fila.getString(nombreColumna); };
arboles[nombreColumna] = new ArbolString(k);
} // Así para todos los demás tipos
for (auto fila : filas) {
arboles[nombreColumna]->add(fila);
}
}
string Table::getString(int numFila, string nombreColumna) {
return columns[NombreColumna].getString(numFila);
}
class Column {
string nombre;
string tipo;
vector<int> intData;
vector<string> strData;
vector<float> floatData;
public:
void addInt(int dato) {
intData.push_back(dato);
}
};
string Column::getString(int 0) {
return strData[0];
}
void Fila::set(string nombreColumna, int dato) {
tabla->columnas[nombreColumna]->addInt(dato);
}
void Fila::set(string nombreColumna, string dato) {
tabla->columnas[nombreColumna]->addString(dato);
}
void Fila::set(string nombreColumna, float dato) {
tabla->columnas[nombreColumna]->addFloat(dato);
}
Fila* Table::createFila() {
return new Fila(this);
}
int main() {
Table* t1 = new Table();
Column* c1 = new Column("dni", "int");
t1->addColumn(c1);
Column* c2 = new Column("nombre", "string");
t1->addColumn(c2);
Column* c3 = new Column("apellido", "string");
t1->addColumn(c3);
Fila* f1 = t1->createFila();
f1.set("dni", 1);
f1.set("nombre", "Rosa");
f1.set("apellido", "Rosales");
Fila* f2 = t1->createFila();
f2.set("dni", 6);
f2.set("nombre", "Felipe");
f2.set("apellido", "Flores");
Table* t2 = t1->select({"dni", "apellido"})
Table* t3 = t1->filter("apellido", "empieza con", "F")
cout << t1->getString(0, "apellido") << endl; // Rosales
cout << t3->getString(0, "apellido") << endl; // Flores
t1->indexar("dni");
return 0;
}
|
IdeasTF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image Compression via K-means
# Source: CS229, PS3, Q5
import numpy as np
from matplotlib.image import imread
import matplotlib.pyplot as plt
# %matplotlib inline
A = imread('cs229_Data/mandrill-large.tiff')
plt.imshow(A)
# ## K-means clustering using a smaller image
B = imread('cs229_Data/mandrill-small.tiff')
# Vectorise the image X, there are 128x128 data in 3D color space
X = B.reshape((-1,3))/255
# See [k-Nearest neighbours](https://github.com/meichen91/MachineLearning-Snippets/blob/master/Week1/KNN_Vectorisation.ipynb) for the explanation of vectorised implementation of the following function
# Function to calculate the distance between X and K
def dist_cluster(X, C):
N = np.shape(X)[0]
K = np.shape(C)[0]
A = np.sum(np.square(X),axis = 1)
B = np.sum(np.square(C),axis = 1)
C = np.dot(X,C.T)
dists = np.sqrt(A[:,np.newaxis]+B[np.newaxis,:]-2*C)
return dists
# Function to calculate the mean for different clusters
def update_cluster_mean(X, A, K):
'''
X is the data
A is assigned_clusters
'''
K_list = np.unique(A)
C = np.zeros((K,np.shape(X)[1]))
for i in range(len(K_list)):
C[i,:] = np.mean(X[A==K_list[i]], axis = 0)
if len(K_list) < K:
print('the number of cluster {} is less than the assigned cluster number {}'.format(len(K_list),K))
# Add a new cluster center randomly
C[len(K_list):] = np.random.rand(K-len(K_list),3)
return C
# Initialise the cluster center
K = 16
C = np.random.rand(K,3)
diff = []
tol = 1
i = 0
# EM for Kmeans
while tol > 1e-8:
# E-step
C_old = C
dists = dist_cluster(X, C)
assigned_clusters = dists.argmin(axis = 1)
# M-step
C = update_cluster_mean(X, assigned_clusters, K)
# Track convergence
tol = np.linalg.norm(C_old - C)
diff.append(tol)
i = i + 1
print('finished at iter {0:d} final difference {1:.2e}'.format(i, tol))
# # Compression
# Compress the large picture using the clusters found from the small one
X2 = A.reshape((-1,3))/255
dists = dist_cluster(X2, C)
assigned_clusters = dists.argmin(axis = 1)
X2_compressed = np.zeros_like(X2)
for i in range(K):
X2_compressed[assigned_clusters==i] = C[i]
A_compressed = np.reshape(X2_compressed,(np.shape(A)))
plt.subplot(1,2,1)
plt.imshow(A_compressed)
plt.subplot(1,2,2)
plt.imshow(A)
# Great quality after compression! The most obvious defect is lack of details at the bottom left and the regions besides the noise.
#
# ## (d)
# If we represent the image with these reduced (16) colors, by (approximately) what factor have we compressed the image?
# The original image has $512\times 512 \times 3 \times 8$ bits in total
#
# The compressed image has $512\times 512 \times 4 + 16\times 3 \times 8$ bits in total (Instead of the 24 bits color, the information stored in each pixel is which of the 16 clusters it belongs to (i.e., 3 bits)
|
CS229_PS/PS3_Q5_KMeans_Compression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Behavioral Cloning**
#
# ## Writeup
#
# ### This writeup serves as the explanation of how I successfully run the car in autonomous mode within one loop without accident.
#
# ---
#
# **Behavioral Cloning Project**
#
# The goals / steps of this project are the following:
#
# * Use the simulator to collect data of good driving behavior
# * Build, a convolution neural network in Keras that predicts steering angles from images
# * Train and validate the model with a training and validation set
# * Test that the model successfully drives around track one without leaving the road
# * Summarize the results with a written report
#
#
# [//]: # (Image References)
#
# [image1]: ./output_images/histogram_origin.png
# [image2]: ./output_images/left.png
# [image3]: ./output_images/center.png
# [image4]: ./output_images/right.png
# [image5]: ./output_images/center_crop.png
# [image6]: ./output_images/crop.png
# [image7]: ./output_images/corner_resize.png
# [image8]: ./output_images/origin.png
# [image9]: ./output_images/corner_yuv.png
# [image10]: ./output_images/flip.png
# [image11]: ./output_images/his_blance.png
# [image12]: ./output_images/his.png
# [image13]: ./output_images/histogram0.15.png
# [image14]: ./output_images/HLSchannle.png
# [image15]: ./output_images/s_channle.png
# [image16]: ./output_images/s_channel.png
# [image17]: ./output_images/s_processed.png
#
#
#
#
#
# ## Rubric Points
# ### Here I will consider the [rubric points](https://review.udacity.com/#!/rubrics/432/view) individually and describe how I addressed each point in my implementation.
#
# ---
# ### Files Submitted & Code Quality
#
# #### Submission includes all required files and can be used to run the simulator in autonomous mode
#
# My project includes the following files:
# * model.py containing the script to create and train the model
# * drive.py for driving the car in autonomous mode
# * model.h5 containing a trained convolution neural network
# * writeup_report.md summarizing the results
# * video.mp4
#
#
#
#
# ### Data collection and balancing
#
# #### 1. I have tried various data set:
# 1. Self-collected data from simulator
# 2. Default data from Udacity
# 3. Recovery data from road edge to road center turn.
# 4. Zigzag running data
#
# However, among all these data, only Default data from Udacity worked well. I think it is the reason that the data was collected in joystick. By keyborads it is very diffculty to collect high quality data with continuous steering angle changing data. This experience has taught me that a good quality data makes difference in the deep learning model performance.
#
# #### 2. Label distribution
#
# The first thing to analyze is to check the label distribution. In the deep learning model the steering angles are the label that need to be learnt from the data. After drawing the histogram of steering angles distributions below, we can see that there zero angle consist most of the data.
#
#
# * The histogram of the original data
#
# ![alt text][image12]
# * The samples from the original data
#
# ![alt text][image8]
#
#
# #### 3. Data balancing
#
# We balance the data by randomly picking up 15% of the those data that the angles are zero. In the experiment, we see that with too many portion of zero angle steering data, the car are more likely to run stright. And the balanced data plays an important role in predicting the steering angle in the later on model training. The following function does the balancing:
#
# ```python
# def collect_data(path):
# lines=[]
# drive_log_path = path + "driving_log.csv"
# data_path = path + "IMG/"
# with open(drive_log_path) as csvfile:
# reader=csv.reader(csvfile)
# next(csvfile)
# for line in reader:
# lines.append(line)
# for line in lines:
# if float(line[3]) != 0:
# image_process(line,data_path)
# else:
# prob = np.random.uniform()
# if prob <= 0.15: #adjust the ratio to balance the data
# image_process(line,data_path)
# ```
#
#
# ### Image processing
#
# 1. In this project, I have done several image processing, such as
# 2. Image crop: to get rid of redundant information like trees, sky and front of car body
# 3. Image resize: to fit into the deep learning model designed by Nvidia
# 4. Image flip: To balance the turning right data
# 5. Image color channel change: to standout the lane in the image.
#
#
# * Image crop
# ![alt text][image6]
#
# * Image resize to 64x64x3
# ![alt text][image7]
#
# * Color channel change from RGB to HLS and show S channel (inspired by image segmentation)
# ![alt text][image16]
#
#
# * Process the S channel and thresh needs fine-tuning
# ```python
# thresh = (40, 255)
# hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
# S= hls[:,:,2]
# binary = np.zeros_like(S)
# binary[(S > thresh[0]) & (S <= thresh[1])] = 1
# plt.imshow(binary)
# ```
# ![alt text][image17]
#
# After fine-tuning the s channel to make the road lane standout in the image, we pad processed s channel with two other channels with zeros to make sure the image is 64x64x3, it looks like the following:
# ![alt text][image14]
#
# * Image flip, in the same time reverse the sign of associated angle.
# ![alt text][image10]
#
# After we the image process, we are able to balance the data with a balanced steering angles set
#
# * Blances data
# ![alt text][image13]
#
#
#
# ### Deep model construction
#
# I tested NVIDIA architecture but I did not do the normalization part for the data but it went well. I used keras API to build the model, compared to tensorflow we used in the previous section, keras API is much more easy to handle and easy to check the input-output parameters
#
#
# ```python
# model = Sequential()
# model.add(Lambda(lambda x: (x/127.5) -1.0, input_shape=(64, 64, 3)))
# model.add(Convolution2D(32, 3, 3, border_mode='same', subsample=(2, 2), activation='relu', name='Conv1'))
# model.add(MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='same'))
# model.add(Convolution2D(64, 3, 3, border_mode='same', subsample=(2, 2), activation='relu', name='Conv2'))
# model.add(MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='same'))
# model.add(Convolution2D(128, 3, 3, border_mode='same', subsample=(1, 1), activation='relu', name='Conv3'))
# model.add(MaxPooling2D(pool_size=(2, 2), strides=None, border_mode='same'))
# model.add(Convolution2D(128, 2, 2, border_mode='same', subsample=(1, 1), activation='relu', name='Conv4'))
#
# model.add(Flatten())
# model.add(Dropout(0.2))
# model.add(Dense(128, activation='relu', name='FC1'))
# model.add(Dropout(0.5))
# model.add(Dense(128, activation='relu', name='FC2'))
# model.add(Dropout(0.5))
# model.add(Dense(64, activation='relu', name='FC3'))
# model.add(Dense(1))
# ```
# In the above layers, dropout layer plays an important role in proventing the model to be overfitting.
# Besides the layers, I use Adam optimizer
# ```python
# adam = Adam(lr=1e-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
# model.compile(loss='mse',optimizer='adam')
# ```
# In order to save the memory and train the model faster, I use ``fit_generator`` API to iterate the data within a batch size. With its help my training time reduces a lot.
#
# #### Parameters that to be fine-tuned are shown in the following table
#
# After tried many times of fine-tuning the following two parameter, I found that with 9 Epochs and 64 batch size the model could be successful.
#
#
# | parameters | value |
# |:-------------:|:-------------:|
# | Epochs | 9 |
# | Batch_size | 64 |
#
# At the end I will share the project video that needs to be submitted here.
#
# [](https://youtu.be/bSUrWfuB280)
#
# ### Reflections
# As we can see from the video that the car was swinging in the road even though the car did not run out of the track. This is due to the right and left images data with correction on the steering angle that were used in the training. And I have tuned the correction value for many rounds and 0.15 was the value that makes the car run inside the lane but still now stable enough. I have also spent more than two weeks to train the model and process the image under various method, like change the brightness change the size, but those only helps were presented above in this note.
#
# Among all the 4 projects that I have done, this one is most challenging for me because I have failed many times in the big turn corner place where the car were always not able to turn big enough to stay inside the lane. But after many trials and errors, I am able to ensure the car runing insde the lane. Even though I am not very satisfied with the performance now but I have no other solutions for the moment now I will first submit first to wait for better suggested solutions.
#
# In the modified version, I processed the image in HLS channel, which I learnt during the project for advanced lane detection.
#
#
|
writeup_report.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import requests
from bs4 import BeautifulSoup
def get_content(url):
response = requests.get(url)
assert response.status_code == 200
return response.text
# +
## print out plot,
wiki = 'https://en.wikipedia.org/wiki/The_Gift_of_the_Magi'
soup = BeautifulSoup(get_content(wiki))
elements = soup.find_all(['p','h2'])
take_plot = False
plot = []
for element in elements:
if take_plot and element.name == 'p':
plot.append(element.text)
if element.find("span", {"id": 'Plot'}):
take_plot = True
print('\n'.join(plot))
# -
url = 'http://www.gutenberg.org/cache/epub/7256/pg7256.txt'
content = get_content(url)
# +
def clean_gift_of_the_magi(content):
start_of_ebook = '*** START OF THIS PROJECT GUTENBERG EBOOK THE GIFT OF THE MAGI ***'
end_of_ebook = 'End of the Project Gutenberg EBook of The Gift of the Magi, by <NAME>'
start_of_ebook_index = content.index(start_of_ebook) + len(start_of_ebook)
end_of_ebook_index = content.index(end_of_ebook)
text = content[start_of_ebook_index:end_of_ebook_index]
start_of_author = 'by <NAME>'
start_of_author_index = text.index(start_of_author) + len(start_of_author)
text = text[start_of_author_index:]
return re.sub(r'[\r\n]+', ' ', text, flags=re.MULTILINE).strip()
text = clean_gift_of_the_magi(content)
# -
import spacy
from collections import defaultdict
nlp = spacy.load('en_core_web_md')
doc = nlp(text)
# +
characters = defaultdict(int)
people = (ent for ent in doc.ents if ent.label_ == 'PERSON')
for ent in people:
person = ent.text.strip()
person_lower = person.lower()
if not 'mme' in person_lower:
characters[person] += 1
# -
characters
from afinn import Afinn
# +
interactions = []
afinn = Afinn('en')
tokens = doc
for index, token in enumerate(tokens):
if characters[token.text] > 0:
start = index - 15
end = index + 15
tokens_close_to = tokens[start:end]
for close in tokens_close_to:
if close.text == token.text:
continue
if characters[close.text] > 0:
sentence = ' '.join([
tk.text
for tk in tokens_close_to
])
interactions.append(
(token.text, close.text, afinn.score(sentence))
)
# -
interactions
import networkx as nx
import matplotlib.pyplot as plt
G = nx.Graph()
for interaction in interactions:
n1 = interaction[0]
n2 = interaction[1]
if G.has_edge(n1, n2):
G[n1][n2]['weight'] += .5
else:
G.add_edge(n1, n2, weight = .5)
positions = nx.spring_layout(G)
# +
nx.draw(G, pos = positions, node_color = 'lightblue', node_size = 3800)
nx.draw_networkx_labels(
G,
pos = positions,
font_size = 11,
font_color = 'red',
font_weight = 'bold'
)
for node1, node2, edge_attr in G.edges(data=True):
width = edge_attr['weight']
edgelist = [(node1, node2)]
nx.draw_networkx_edges(
G,
positions,
edgelist = edgelist,
width = width
)
# -
import pandas as pd
# +
data = {
'#': dict(G.degree),
'Degree': nx.degree_centrality(G),
'Closeness': nx.closeness_centrality(G),
'Betweenness': nx.betweenness_centrality(G),
'Pagerank': nx.pagerank(G)
}
pd.DataFrame(data)
# -
|
ner/Extracting Character Networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MNIST
from sklearn.datasets import fetch_openml
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
mnist = fetch_openml('mnist_784', version=1)
mnist.keys()
X, y = mnist["data"], mnist["target"]
X.shape
y.shape
# +
some_digit = X[0]
some_digit_image = some_digit.reshape(28, 28)
plt.imshow(some_digit_image, cmap="binary")
plt.axis("off")
# -
y = y.astype(np.uint8)
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# # Training a Binary Classifier
from sklearn.linear_model import SGDClassifier
y_train_5 = (y_train == 5) # True for all 5s, False for all other digits
y_test_5 = (y_test == 5)
sgd_clf = SGDClassifier(random_state=42)
sgd_clf.fit(X_train, y_train_5)
sgd_clf.predict([some_digit])
# # Measuring Accuracy Using Cross-Validation
from sklearn.model_selection import cross_val_score
y_scores_accuracy = cross_val_score(sgd_clf, X_train, y_train_5, cv=3, scoring="accuracy")
y_scores_accuracy
# # Confusiong Matrix
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
y_train_pred = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3)
confusion_matrix(y_train_5, y_train_pred)
y_train_perfect_predictions = y_train_5
confusion_matrix(y_train_5, y_train_perfect_predictions)
# # Precision and Recall
from sklearn.metrics import precision_score, recall_score, f1_score
precision_score(y_train_5, y_train_pred)
recall_score(y_train_5, y_train_pred)
f1_score(y_train_5, y_train_pred)
# # Precision/Recall Trade-off
from sklearn.metrics import precision_recall_curve
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision")
plt.plot(thresholds, recalls[:-1], "g-", label="Recall")
y_scores = sgd_clf.decision_function([some_digit])
y_scores
threshold = 0
y_some_digit_pred = (y_scores > threshold)
y_some_digit_pred
threshold = 8000
y_some_digit_pred = (y_scores > threshold)
y_some_digit_pred
y_scores = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3, method="decision_function")
y_scores
precisions, recalls, thresholds = precision_recall_curve(y_train_5, y_scores)
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.plot(recalls, precisions, "b--", label="Precisions vs. recalls")
index_90_precision = np.argmax(precisions >= 0.90)
threshold_90_precision = thresholds[index_90_precision]
y_scores
y_train_pred_90 = (y_scores >= threshold_90_precision)
precision_score(y_train_5, y_train_pred_90)
recall_score(y_train_5, y_train_pred_90)
# # The ROC Curve
from sklearn.metrics import roc_curve, roc_auc_score
from sklearn.ensemble import RandomForestClassifier
fpr, tpr, thresholds = roc_curve(y_train_5, y_scores)
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--') # Dashed diagonal
plot_roc_curve(fpr, tpr)
forest_clf = RandomForestClassifier(random_state=42)
y_probas_forest = cross_val_predict(forest_clf, X_train, y_train_5, cv=3, method="predict_proba")
y_scores_forest = y_probas_forest[:, 1]
fpr_forest, tpr_forest, thresholds_forest = roc_curve(y_train_5,y_scores_forest)
plt.plot(fpr, tpr, "b:", label="SGD")
plot_roc_curve(fpr_forest, tpr_forest, "Random Forest")
plt.legend(loc="lower right")
roc_auc_score(y_train_5, y_scores_forest)
# # Multiclass Classification
from sklearn.svm import SVC
from sklearn.multiclass import OneVsRestClassifier
from sklearn.preprocessing import StandardScaler
# + jupyter={"outputs_hidden": true}
svm_clf = SVC()
svm_clf.fit(X_train, y_train) # y_train, not y_train_5
svm_clf.predict([some_digit])
# -
svm_clf.predict(X_test)
y_pred = _
some_digit_scores = svm_clf.decision_function([some_digit])
some_digit_scores
np.argmax(some_digit_scores)
svm_clf.classes_
svm_clf.classes_[5]
ovr_clf = OneVsRestClassifier(SVC())
ovr_clf.fit(X_train, y_train)
ovr_clf.predict([some_digit])
len(ovr_clf.estimators_)
sgd_clf.fit(X_train, y_train)
sgd_clf.predict([some_digit])
sgd_clf.decision_function([some_digit])
cross_val_score(sgd_clf, X_train, y_train, cv=3, scoring="accuracy")
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train.astype(np.float64))
cross_val_score(sgd_clf, X_train_scaled, y_train, cv=3, scoring="accuracy")
# # Error Analysis
y_train_pred = cross_val_predict(sgd_clf, X_train_scaled, y_train, cv=3)
conf_mx = confusion_matrix(y_train, y_train_pred)
conf_mx
plt.matshow(conf_mx, cmap=plt.cm.gray)
row_sums = conf_mx.sum(axis=1, keepdims=True)
norm_conf_mx = conf_mx / row_sums
np.fill_diagonal(norm_conf_mx, 0)
plt.matshow(norm_conf_mx, cmap=plt.cm.gray)
# # Multilabel Classification
from sklearn.neighbors import KNeighborsClassifier
y_train_large = (y_train >= 7)
y_train_odd = (y_train % 2 == 1)
y_multilabel = np.c_[y_train_large, y_train_odd]
knn_clf = KNeighborsClassifier()
knn_clf.fit(X_train, y_multilabel)
knn_clf.predict([some_digit])
y_train_knn_pred = cross_val_predict(knn_clf, X_train, y_multilabel, cv=3)
f1_score(y_multilabel, y_train_knn_pred, average="macro")
# # Multioutput Classification
noise = np.random.randint(0, 100, (len(X_train), 784))
X_train_mod = X_train + noise
noise = np.random.randint(0, 100, (len(X_test), 784))
X_test_mod = X_test + noise
y_train_mod = X_train
y_test_mod = X_test
plt.imshow(X_train_mod[1].reshape(28, 28), cmap='gray')
plt.imshow(X_train[1].reshape(28, 28), cmap='gray')
plt.imshow(X_test_mod[1].reshape(28, 28), cmap='gray')
knn_clf.fit(X_train_mod, y_train_mod)
clean_digit = knn_clf.predict([X_test_mod[1]])
plt.imshow(clean_digit.reshape(28, 28), cmap='gray')
|
cd03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# language: python
# name: python36964bit8528f96367f74bd6857989b464e962ed
# ---
# # Temporal-Difference Methods
#
# In this notebook, you will write your own implementations of many Temporal-Difference (TD) methods.
#
# While we have provided some starter code, you are welcome to erase these hints and write your code from scratch.
#
# ---
#
# ### Part 0: Explore CliffWalkingEnv
#
# We begin by importing the necessary packages.
# +
import sys
import gym
import random
import numpy as np
from collections import defaultdict, deque
import matplotlib.pyplot as plt
# %matplotlib inline
import check_test
from plot_utils import plot_values
# -
# Use the code cell below to create an instance of the [CliffWalking](https://github.com/openai/gym/blob/master/gym/envs/toy_text/cliffwalking.py) environment.
env = gym.make('CliffWalking-v0')
# The agent moves through a $4\times 12$ gridworld, with states numbered as follows:
# ```
# [[ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11],
# [12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23],
# [24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35],
# [36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47]]
# ```
# At the start of any episode, state `36` is the initial state. State `47` is the only terminal state, and the cliff corresponds to states `37` through `46`.
#
# The agent has 4 potential actions:
# ```
# UP = 0
# RIGHT = 1
# DOWN = 2
# LEFT = 3
# ```
#
# Thus, $\mathcal{S}^+=\{0, 1, \ldots, 47\}$, and $\mathcal{A} =\{0, 1, 2, 3\}$. Verify this by running the code cell below.
print(env.action_space)
print(env.observation_space)
# In this mini-project, we will build towards finding the optimal policy for the CliffWalking environment. The optimal state-value function is visualized below. Please take the time now to make sure that you understand _why_ this is the optimal state-value function.
#
# _**Note**: You can safely ignore the values of the cliff "states" as these are not true states from which the agent can make decisions. For the cliff "states", the state-value function is not well-defined._
# +
# define the optimal state-value function
V_opt = np.zeros((4,12))
V_opt[0:13][0] = -np.arange(3, 15)[::-1]
V_opt[0:13][1] = -np.arange(3, 15)[::-1] + 1
V_opt[0:13][2] = -np.arange(3, 15)[::-1] + 2
V_opt[3][0] = -13
plot_values(V_opt)
# -
# ### Part 1: TD Control: Sarsa
#
# In this section, you will write your own implementation of the Sarsa control algorithm.
#
# Your algorithm has four arguments:
# - `env`: This is an instance of an OpenAI Gym environment.
# - `num_episodes`: This is the number of episodes that are generated through agent-environment interaction.
# - `alpha`: This is the step-size parameter for the update step.
# - `gamma`: This is the discount rate. It must be a value between 0 and 1, inclusive (default value: `1`).
#
# The algorithm returns as output:
# - `Q`: This is a dictionary (of one-dimensional arrays) where `Q[s][a]` is the estimated action value corresponding to state `s` and action `a`.
#
# Please complete the function in the code cell below.
#
# (_Feel free to define additional functions to help you to organize your code._)
# +
def get_epsilon_policy(bj_env, epsilon, Q, state, nA):
policy = np.ones(nA) * epsilon / nA
astar = np.argmax(Q[state])
policy[astar] = 1 - epsilon + (epsilon / nA)
action = np.random.choice(nA, p=policy) if state in Q else bj_env.action_space.sample()
return action
def get_greedy_policy(bj_env, Q, state):
return np.argmax(Q[state])
def get_expected_value(bj_env, epsilon, Q, state, nA):
policy = np.ones(nA) * epsilon / nA
astar = np.argmax(Q[state])
policy[astar] = 1 - epsilon + (epsilon / nA)
V = np.dot(Q[state],policy)
return V
# -
def sarsa(env, num_episodes, alpha, gamma=1.0):
# initialize action-value function (empty dictionary of arrays)
nA = env.action_space.n
Q = defaultdict(lambda: np.zeros(env.nA))
# initialize performance monitor
# loop over episodes
for i_episode in range(1, num_episodes+1):
# monitor progress
if i_episode % 100 == 0:
print("\rEpisode {}/{}".format(i_episode, num_episodes), end="")
sys.stdout.flush()
state = env.reset()
epsilon = 1 / i_episode
action = get_epsilon_policy(env, epsilon, Q, state, nA)
while True:
next_state, reward, done, info = env.step(action)
next_action = get_epsilon_policy(env, epsilon, Q, next_state, nA)
if done:
Q[state][action] += alpha * (reward - Q[state][action])
break
else:
Q[state][action] += alpha * (reward + gamma * Q[next_state][next_action] - Q[state][action])
state = next_state
action = next_action
return Q
# Use the next code cell to visualize the **_estimated_** optimal policy and the corresponding state-value function.
#
# If the code cell returns **PASSED**, then you have implemented the function correctly! Feel free to change the `num_episodes` and `alpha` parameters that are supplied to the function. However, if you'd like to ensure the accuracy of the unit test, please do not change the value of `gamma` from the default.
# +
# obtain the estimated optimal policy and corresponding action-value function
Q_sarsa = sarsa(env, 5000, .01)
# print the estimated optimal policy
policy_sarsa = np.array([np.argmax(Q_sarsa[key]) if key in Q_sarsa else -1 for key in np.arange(48)]).reshape(4,12)
check_test.run_check('td_control_check', policy_sarsa)
print("\nEstimated Optimal Policy (UP = 0, RIGHT = 1, DOWN = 2, LEFT = 3, N/A = -1):")
print(policy_sarsa)
# plot the estimated optimal state-value function
V_sarsa = ([np.max(Q_sarsa[key]) if key in Q_sarsa else 0 for key in np.arange(48)])
plot_values(V_sarsa)
# -
# ### Part 2: TD Control: Q-learning
#
# In this section, you will write your own implementation of the Q-learning control algorithm.
#
# Your algorithm has four arguments:
# - `env`: This is an instance of an OpenAI Gym environment.
# - `num_episodes`: This is the number of episodes that are generated through agent-environment interaction.
# - `alpha`: This is the step-size parameter for the update step.
# - `gamma`: This is the discount rate. It must be a value between 0 and 1, inclusive (default value: `1`).
#
# The algorithm returns as output:
# - `Q`: This is a dictionary (of one-dimensional arrays) where `Q[s][a]` is the estimated action value corresponding to state `s` and action `a`.
#
# Please complete the function in the code cell below.
#
# (_Feel free to define additional functions to help you to organize your code._)
def q_learning(env, num_episodes, alpha, gamma=1.0):
# initialize empty dictionary of arrays
Q = defaultdict(lambda: np.zeros(env.nA))
nA = env.nA
# loop over episodes
for i_episode in range(1, num_episodes+1):
# monitor progress
if i_episode % 100 == 0:
print("\rEpisode {}/{}".format(i_episode, num_episodes), end="")
sys.stdout.flush()
state = env.reset()
epsilon = 1 / i_episode
while True:
action = get_epsilon_policy(env, epsilon, Q, state, nA)
next_state, reward, done, info = env.step(action)
if done:
Q[state][action] += alpha * (reward - Q[state][action])
break
else:
greedy_action = get_greedy_policy(env, Q, next_state)
Q[state][action] += alpha * (reward + gamma * Q[next_state][greedy_action] - Q[state][action])
state = next_state
return Q
# Use the next code cell to visualize the **_estimated_** optimal policy and the corresponding state-value function.
#
# If the code cell returns **PASSED**, then you have implemented the function correctly! Feel free to change the `num_episodes` and `alpha` parameters that are supplied to the function. However, if you'd like to ensure the accuracy of the unit test, please do not change the value of `gamma` from the default.
# +
# obtain the estimated optimal policy and corresponding action-value function
Q_sarsamax = q_learning(env, 5000, .01)
# print the estimated optimal policy
policy_sarsamax = np.array([np.argmax(Q_sarsamax[key]) if key in Q_sarsamax else -1 for key in np.arange(48)]).reshape((4,12))
check_test.run_check('td_control_check', policy_sarsamax)
print("\nEstimated Optimal Policy (UP = 0, RIGHT = 1, DOWN = 2, LEFT = 3, N/A = -1):")
print(policy_sarsamax)
# plot the estimated optimal state-value function
plot_values([np.max(Q_sarsamax[key]) if key in Q_sarsamax else 0 for key in np.arange(48)])
# -
# ### Part 3: TD Control: Expected Sarsa
#
# In this section, you will write your own implementation of the Expected Sarsa control algorithm.
#
# Your algorithm has four arguments:
# - `env`: This is an instance of an OpenAI Gym environment.
# - `num_episodes`: This is the number of episodes that are generated through agent-environment interaction.
# - `alpha`: This is the step-size parameter for the update step.
# - `gamma`: This is the discount rate. It must be a value between 0 and 1, inclusive (default value: `1`).
#
# The algorithm returns as output:
# - `Q`: This is a dictionary (of one-dimensional arrays) where `Q[s][a]` is the estimated action value corresponding to state `s` and action `a`.
#
# Please complete the function in the code cell below.
#
# (_Feel free to define additional functions to help you to organize your code._)
def expected_sarsa(env, num_episodes, alpha, gamma=1.0):
# initialize empty dictionary of arrays
Q = defaultdict(lambda: np.zeros(env.nA))
# loop over episodes
for i_episode in range(1, num_episodes+1):
# monitor progress
if i_episode % 100 == 0:
print("\rEpisode {}/{}".format(i_episode, num_episodes), end="")
sys.stdout.flush()
state = env.reset()
epsilon = 0.005
while True:
action = get_epsilon_policy(env, epsilon, Q, state, env.nA)
next_state, reward, done, info = env.step(action)
if done:
Q[state][action] += alpha * (reward - Q[state][action])
break
else:
expected_value = get_expected_value(env, epsilon, Q, next_state, env.nA)
Q[state][action] += alpha * (reward + gamma * expected_value - Q[state][action])
state = next_state
return Q
# Use the next code cell to visualize the **_estimated_** optimal policy and the corresponding state-value function.
#
# If the code cell returns **PASSED**, then you have implemented the function correctly! Feel free to change the `num_episodes` and `alpha` parameters that are supplied to the function. However, if you'd like to ensure the accuracy of the unit test, please do not change the value of `gamma` from the default.
# +
# obtain the estimated optimal policy and corresponding action-value function
Q_expsarsa = expected_sarsa(env, 5000, 1)
# print the estimated optimal policy
policy_expsarsa = np.array([np.argmax(Q_expsarsa[key]) if key in Q_expsarsa else -1 for key in np.arange(48)]).reshape(4,12)
check_test.run_check('td_control_check', policy_expsarsa)
print("\nEstimated Optimal Policy (UP = 0, RIGHT = 1, DOWN = 2, LEFT = 3, N/A = -1):")
print(policy_expsarsa)
# plot the estimated optimal state-value function
plot_values([np.max(Q_expsarsa[key]) if key in Q_expsarsa else 0 for key in np.arange(48)])
# -
|
temporal-difference/Temporal_Difference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import eigensheep
# +
# %%eigensheep --layer arn:aws:lambda:us-east-1:972882471061:layer:z3:2
# IS SOCRATES MORTAL?
# TUNE IN NEXT EPISODE TO FIND OUT!
from z3 import *
Object = DeclareSort('Object')
Human = Function('Human', Object, BoolSort())
Mortal = Function('Mortal', Object, BoolSort())
# a well known philosopher
socrates = Const('socrates', Object)
# free variables used in forall must be declared Const in python
x = Const('x', Object)
axioms = [ForAll([x], Implies(Human(x), Mortal(x))),
Human(socrates)]
s = Solver()
s.add(axioms)
print("axioms are coherent", s.check()) # prints sat so axioms are coherents
# classical refutation
s.add(Not(Mortal(socrates)))
print("socrates is mortal", s.check()) # prints unsat so socrates is Mortal
# +
# %%eigensheep --layer arn:aws:lambda:us-east-1:972882471061:layer:z3:2
# Solve Sudoku
# Based on https://github.com/ppmx/sudoku-solver/blob/master/sudoku-z3.py
def cross(A, B):
return [(a + b) for a in A for b in B]
class Sudoku:
@staticmethod
def parse_grid(puzzle):
"""
A1 A2 A3 | A4 A5 A6 | A7 A8 A9
B1 B2 B3 | B4 B5 B6 | B7 B8 B9
C1 C2 C3 | C4 C5 C6 | C7 C8 C9
–––––––––+––––––––––+–––––––––
D1 D2 D3 | D4 D5 D6 | D7 D8 D9
E1 E2 E3 | E4 E5 E6 | E7 E8 E9
F1 F2 F3 | F4 F5 F6 | F7 F8 F9
–––––––––+––––––––––+–––––––––
G1 G2 G3 | G4 G5 G6 | G7 G8 G9
H1 H2 H3 | H4 H5 H6 | H7 H8 H9
I1 I2 I3 | I4 I5 I6 | I7 I8 I9
puzzle = 'A1A2A3A4...' and every element holds a value of '123456789.'
where the dot represents an empty cell.
"""
s = Sudoku()
if any(c not in "123456789." for c in puzzle) or len(puzzle) != 81:
raise Exception("got invalid puzzle format")
elements = cross("ABCDEFGHI", "123456789")
s.values = {e: v for e,v in zip(elements, puzzle)}
return s
def __init__(self, values=dict()):
# mapping cells -> "123456789." where the dot represents an empty cell
# cells = cross product of "ABCDEFGHI" and "123456789"
self.values = values
# we define some additional informations that may be used by a solving function:
rows, cols = "ABCDEFGHI", "123456789"
self.elements = cross(rows, cols)
self.unitlist = []
self.unitlist += [cross(rows, c) for c in cols]
self.unitlist += [cross(r, cols) for r in rows]
self.unitlist += [cross(rs, cs) for rs in ["ABC", "DEF", "GHI"] for cs in ["123", "456", "789"]]
self.units = {e: [u for u in self.unitlist if e in u] for e in self.elements}
def is_solved(self):
# assure that every cell holds a single value between 1 and 9:
if not all(k in "123456789" for k in self.values.values()):
return False
# assure that every cell of every unit is unique in the proper unit:
unitsolved = lambda u: set([self.values[e] for e in u]) == set("123456789")
return all(unitsolved(u) for u in self.unitlist)
def __str__(self):
lines, elements = [], cross("ABCDEFGHI", "123456789")
print("[+] Puzzle:", ''.join(self.values[e] for e in elements))
for index_row, row in enumerate("ABCDEFGHI"):
if index_row % 3 == 0:
lines.append("+–––––––––+–––––––––+–––––––––+")
line = ''
for index_col, col in enumerate("123456789"):
line += "{1} {0} ".format(self.values[row + col], '|' if index_col % 3 == 0 else '')
lines.append(line + '|')
lines.append("+–––––––––+–––––––––+–––––––––+")
return '\n'.join(lines) + '\n'
def Z3Solving(sudoku):
from z3 import Solver, Int, Or, Distinct, sat
elements = cross("ABCDEFGHI", "123456789")
symbols = {e: Int(e) for e in elements}
# first we build a solver with the general constraints for sudoku puzzles:
s = Solver()
# assure that every cell holds a value of [1,9]
for symbol in symbols.values():
s.add(Or([symbol == i for i in range(1, 10)]))
# assure that every row covers every value:
for row in "ABCDEFGHI":
s.add(Distinct([symbols[row + col] for col in "123456789"]))
# assure that every column covers every value:
for col in "123456789":
s.add(Distinct([symbols[row + col] for row in "ABCDEFGHI"]))
# assure that every block covers every value:
for i in range(3):
for j in range(3):
s.add(Distinct([symbols["ABCDEFGHI"[m + i * 3] + "123456789"[n + j * 3]] for m in range(3) for n in range(3)]))
# now we put the assumptions of the given puzzle into the solver:
for elem, value in sudoku.values.items():
if value in "123456789":
s.add(symbols[elem] == value)
if not s.check() == sat:
raise Exception("unsolvable")
model = s.model()
values = {e: model.evaluate(s).as_string() for e, s in symbols.items()}
return Sudoku(values)
def main(puzzle):
print("[+] processing puzzle:", puzzle)
s = Sudoku.parse_grid(puzzle)
print(s)
print("[+] trying to solve it with z3")
s_solved = Z3Solving(s)
print("[+] it is solved:", s_solved.is_solved())
main("4.....8.5.3..........7......2.....6.....8.4......1.......6.3.7.5..2.....1.4......")
# -
|
examples/Z3 SMT Solver.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Automated Machine Learning
# **BikeShare Demand Forecasting**
#
# ## Contents
# 1. [Introduction](#Introduction)
# 1. [Setup](#Setup)
# 1. [Compute](#Compute)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Featurization](#Featurization)
# 1. [Evaluate](#Evaluate)
# ## Introduction
# This notebook demonstrates demand forecasting for a bike-sharing service using AutoML.
#
# AutoML highlights here include built-in holiday featurization, accessing engineered feature names, and working with the `forecast` function. Please also look at the additional forecasting notebooks, which document lagging, rolling windows, forecast quantiles, other ways to use the forecast function, and forecaster deployment.
#
# Make sure you have executed the [configuration notebook](../../../configuration.ipynb) before running this notebook.
#
# Notebook synopsis:
# 1. Creating an Experiment in an existing Workspace
# 2. Configuration and local run of AutoML for a time-series model with lag and holiday features
# 3. Viewing the engineered names for featurized data and featurization summary for all raw features
# 4. Evaluating the fitted model using a rolling test
# ## Setup
#
# +
import azureml.core
import pandas as pd
import numpy as np
import logging
from azureml.core import Workspace, Experiment, Dataset
from azureml.train.automl import AutoMLConfig
from datetime import datetime
# -
# This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
print("This notebook was created using version 1.31.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
# As part of the setup you have already created a <b>Workspace</b>. To run AutoML, you also need to create an <b>Experiment</b>. An Experiment corresponds to a prediction problem you are trying to solve, while a Run corresponds to a specific approach to the problem.
# +
ws = Workspace.from_config()
# choose a name for the run history container in the workspace
experiment_name = 'automl-bikeshareforecasting'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['SKU'] = ws.sku
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Run History Name'] = experiment_name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ## Compute
# You will need to create a [compute target](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-set-up-training-targets#amlcompute) for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
#
# > Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
#
# #### Creation of AmlCompute takes approximately 5 minutes.
# If the AmlCompute with that name is already in your workspace this code will skip the creation process.
# As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your cluster.
amlcompute_cluster_name = "bike-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# ## Data
#
# The [Machine Learning service workspace](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-workspace) is paired with the storage account, which contains the default data store. We will use it to upload the bike share data and create [tabular dataset](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) for training. A tabular dataset defines a series of lazily-evaluated, immutable operations to load data from the data source into tabular representation.
datastore = ws.get_default_datastore()
datastore.upload_files(files = ['./bike-no.csv'], target_path = 'dataset/', overwrite = True,show_progress = True)
# Let's set up what we know about the dataset.
#
# **Target column** is what we want to forecast.
#
# **Time column** is the time axis along which to predict.
target_column_name = 'cnt'
time_column_name = 'date'
# +
dataset = Dataset.Tabular.from_delimited_files(path = [(datastore, 'dataset/bike-no.csv')]).with_timestamp_columns(fine_grain_timestamp=time_column_name)
# Drop the columns 'casual' and 'registered' as these columns are a breakdown of the total and therefore a leak.
dataset = dataset.drop_columns(columns=['casual', 'registered'])
dataset.take(5).to_pandas_dataframe().reset_index(drop=True)
# -
# ### Split the data
#
# The first split we make is into train and test sets. Note we are splitting on time. Data before 9/1 will be used for training, and data after and including 9/1 will be used for testing.
# select data that occurs before a specified date
train = dataset.time_before(datetime(2012, 8, 31), include_boundary=True)
train.to_pandas_dataframe().tail(5).reset_index(drop=True)
test = dataset.time_after(datetime(2012, 9, 1), include_boundary=True)
test.to_pandas_dataframe().head(5).reset_index(drop=True)
# ## Forecasting Parameters
# To define forecasting parameters for your experiment training, you can leverage the ForecastingParameters class. The table below details the forecasting parameter we will be passing into our experiment.
#
# |Property|Description|
# |-|-|
# |**time_column_name**|The name of your time column.|
# |**forecast_horizon**|The forecast horizon is how many periods forward you would like to forecast. This integer horizon is in units of the timeseries frequency (e.g. daily, weekly).|
# |**country_or_region_for_holidays**|The country/region used to generate holiday features. These should be ISO 3166 two-letter country/region codes (i.e. 'US', 'GB').|
# |**target_lags**|The target_lags specifies how far back we will construct the lags of the target variable.|
# |**freq**|Forecast frequency. This optional parameter represents the period with which the forecast is desired, for example, daily, weekly, yearly, etc. Use this parameter for the correction of time series containing irregular data points or for padding of short time series. The frequency needs to be a pandas offset alias. Please refer to [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#dateoffset-objects) for more information.
# ## Train
#
# Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
#
# |Property|Description|
# |-|-|
# |**task**|forecasting|
# |**primary_metric**|This is the metric that you want to optimize.<br> Forecasting supports the following primary metrics <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>
# |**blocked_models**|Models in blocked_models won't be used by AutoML. All supported models can be found at [here](https://docs.microsoft.com/en-us/python/api/azureml-train-automl-client/azureml.train.automl.constants.supportedmodels.forecasting?view=azure-ml-py).|
# |**experiment_timeout_hours**|Experimentation timeout in hours.|
# |**training_data**|Input dataset, containing both features and label column.|
# |**label_column_name**|The name of the label column.|
# |**compute_target**|The remote compute for training.|
# |**n_cross_validations**|Number of cross validation splits.|
# |**enable_early_stopping**|If early stopping is on, training will stop when the primary metric is no longer improving.|
# |**forecasting_parameters**|A class that holds all the forecasting related parameters.|
#
# This notebook uses the blocked_models parameter to exclude some models that take a longer time to train on this dataset. You can choose to remove models from the blocked_models list but you may need to increase the experiment_timeout_hours parameter value to get results.
# ### Setting forecaster maximum horizon
#
# The forecast horizon is the number of periods into the future that the model should predict. Here, we set the horizon to 14 periods (i.e. 14 days). Notice that this is much shorter than the number of days in the test set; we will need to use a rolling test to evaluate the performance on the whole test set. For more discussion of forecast horizons and guiding principles for setting them, please see the [energy demand notebook](https://github.com/Azure/MachineLearningNotebooks/tree/master/how-to-use-azureml/automated-machine-learning/forecasting-energy-demand).
forecast_horizon = 14
# ### Config AutoML
# +
from azureml.automl.core.forecasting_parameters import ForecastingParameters
forecasting_parameters = ForecastingParameters(
time_column_name=time_column_name,
forecast_horizon=forecast_horizon,
country_or_region_for_holidays='US', # set country_or_region will trigger holiday featurizer
target_lags='auto', # use heuristic based lag setting
freq='D' # Set the forecast frequency to be daily
)
automl_config = AutoMLConfig(task='forecasting',
primary_metric='normalized_root_mean_squared_error',
blocked_models = ['ExtremeRandomTrees'],
experiment_timeout_hours=0.3,
training_data=train,
label_column_name=target_column_name,
compute_target=compute_target,
enable_early_stopping=True,
n_cross_validations=3,
max_concurrent_iterations=4,
max_cores_per_iteration=-1,
verbosity=logging.INFO,
forecasting_parameters=forecasting_parameters)
# -
# We will now run the experiment, you can go to Azure ML portal to view the run details.
remote_run = experiment.submit(automl_config, show_output=False)
remote_run.wait_for_completion()
# ### Retrieve the Best Model
# Below we select the best model from all the training iterations using get_output method.
best_run, fitted_model = remote_run.get_output()
fitted_model.steps
# ## Featurization
#
# You can access the engineered feature names generated in time-series featurization. Note that a number of named holiday periods are represented. We recommend that you have at least one year of data when using this feature to ensure that all yearly holidays are captured in the training featurization.
fitted_model.named_steps['timeseriestransformer'].get_engineered_feature_names()
# ### View the featurization summary
#
# You can also see what featurization steps were performed on different raw features in the user data. For each raw feature in the user data, the following information is displayed:
#
# - Raw feature name
# - Number of engineered features formed out of this raw feature
# - Type detected
# - If feature was dropped
# - List of feature transformations for the raw feature
# Get the featurization summary as a list of JSON
featurization_summary = fitted_model.named_steps['timeseriestransformer'].get_featurization_summary()
# View the featurization summary as a pandas dataframe
pd.DataFrame.from_records(featurization_summary)
# ## Evaluate
# We now use the best fitted model from the AutoML Run to make forecasts for the test set. We will do batch scoring on the test dataset which should have the same schema as training dataset.
#
# The scoring will run on a remote compute. In this example, it will reuse the training compute.
test_experiment = Experiment(ws, experiment_name + "_test")
# ### Retrieving forecasts from the model
# To run the forecast on the remote compute we will use a helper script: forecasting_script. This script contains the utility methods which will be used by the remote estimator. We copy the script to the project folder to upload it to remote compute.
# +
import os
import shutil
script_folder = os.path.join(os.getcwd(), 'forecast')
os.makedirs(script_folder, exist_ok=True)
shutil.copy('forecasting_script.py', script_folder)
# -
# For brevity, we have created a function called run_forecast that submits the test data to the best model determined during the training run and retrieves forecasts. The test set is longer than the forecast horizon specified at train time, so the forecasting script uses a so-called rolling evaluation to generate predictions over the whole test set. A rolling evaluation iterates the forecaster over the test set, using the actuals in the test set to make lag features as needed.
# +
from run_forecast import run_rolling_forecast
remote_run = run_rolling_forecast(test_experiment, compute_target, best_run, test, target_column_name)
remote_run
# -
remote_run.wait_for_completion(show_output=False)
# ### Download the prediction result for metrics calcuation
# The test data with predictions are saved in artifact outputs/predictions.csv. You can download it and calculation some error metrics for the forecasts and vizualize the predictions vs. the actuals.
remote_run.download_file('outputs/predictions.csv', 'predictions.csv')
df_all = pd.read_csv('predictions.csv')
# +
from azureml.automl.core.shared import constants
from azureml.automl.runtime.shared.score import scoring
from sklearn.metrics import mean_absolute_error, mean_squared_error
from matplotlib import pyplot as plt
# use automl metrics module
scores = scoring.score_regression(
y_test=df_all[target_column_name],
y_pred=df_all['predicted'],
metrics=list(constants.Metric.SCALAR_REGRESSION_SET))
print("[Test data scores]\n")
for key, value in scores.items():
print('{}: {:.3f}'.format(key, value))
# Plot outputs
# %matplotlib inline
test_pred = plt.scatter(df_all[target_column_name], df_all['predicted'], color='b')
test_test = plt.scatter(df_all[target_column_name], df_all[target_column_name], color='g')
plt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)
plt.show()
# -
# For more details on what metrics are included and how they are calculated, please refer to [supported metrics](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-understand-automated-ml#regressionforecasting-metrics). You could also calculate residuals, like described [here](https://docs.microsoft.com/en-us/azure/machine-learning/how-to-understand-automated-ml#residuals).
#
#
# Since we did a rolling evaluation on the test set, we can analyze the predictions by their forecast horizon relative to the rolling origin. The model was initially trained at a forecast horizon of 14, so each prediction from the model is associated with a horizon value from 1 to 14. The horizon values are in a column named, "horizon_origin," in the prediction set. For example, we can calculate some of the error metrics grouped by the horizon:
from metrics_helper import MAPE, APE
df_all.groupby('horizon_origin').apply(
lambda df: pd.Series({'MAPE': MAPE(df[target_column_name], df['predicted']),
'RMSE': np.sqrt(mean_squared_error(df[target_column_name], df['predicted'])),
'MAE': mean_absolute_error(df[target_column_name], df['predicted'])}))
# To drill down more, we can look at the distributions of APE (absolute percentage error) by horizon. From the chart, it is clear that the overall MAPE is being skewed by one particular point where the actual value is of small absolute value.
# +
df_all_APE = df_all.assign(APE=APE(df_all[target_column_name], df_all['predicted']))
APEs = [df_all_APE[df_all['horizon_origin'] == h].APE.values for h in range(1, forecast_horizon + 1)]
# %matplotlib inline
plt.boxplot(APEs)
plt.yscale('log')
plt.xlabel('horizon')
plt.ylabel('APE (%)')
plt.title('Absolute Percentage Errors by Forecast Horizon')
plt.show()
|
how-to-use-azureml/automated-machine-learning/forecasting-bike-share/auto-ml-forecasting-bike-share.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="cIecysdeMIz2" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
# + id="FILI36XqYmLR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="62b5a38f-288c-4d77-cf05-d191d34ed9fb" executionInfo={"status": "ok", "timestamp": 1581628484171, "user_tz": -60, "elapsed": 1464, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
df = pd.read_csv('data/man_shoes.csv', low_memory=False)
df.shape
# + id="eQpNUXTCZZQH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="fcedef0a-8744-428e-8d13-53bab2431b79" executionInfo={"status": "ok", "timestamp": 1581628534899, "user_tz": -60, "elapsed": 578, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
df.columns
# + id="lQzmsPh3Z-rd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06bfbc4d-364a-4fb7-90bf-faf50426b1c4" executionInfo={"status": "ok", "timestamp": 1581628889951, "user_tz": -60, "elapsed": 651, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
mean_price = np.mean( df['prices_amountmin'] )
mean_price
# + id="6dSMtEwna8NX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d3bd2b81-d91b-4751-f5bc-6543ed1e132f" executionInfo={"status": "ok", "timestamp": 1581628973322, "user_tz": -60, "elapsed": 561, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
[1] * 5
# + id="yp6FjaR8bt54" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="70a107de-7c15-409a-da5d-b41ea6730dc0" executionInfo={"status": "ok", "timestamp": 1581629281253, "user_tz": -60, "elapsed": 700, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
y_true = df['prices_amountmin']
y_pred = [mean_price] * df.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="Q8Wj5wuEcH8Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="65080b67-0278-443a-b0ad-62a1c21e8c26" executionInfo={"status": "ok", "timestamp": 1581629544291, "user_tz": -60, "elapsed": 1063, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
df['prices_amountmin'].hist(bins=100)
# + id="_X5WZT8HdvTK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="5ac38002-d2c4-4316-e86f-582331a4b16f" executionInfo={"status": "ok", "timestamp": 1581629727184, "user_tz": -60, "elapsed": 746, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
np.log(df['prices_amountmin'] + 1).hist(bins=100)
# + id="Iv5kTN3Sel5Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ff31584-286f-4ea6-86a1-899f0978495d" executionInfo={"status": "ok", "timestamp": 1581629852383, "user_tz": -60, "elapsed": 956, "user": {"displayName": "<NAME>00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
y_true = df['prices_amountmin']
y_pred = [np.median(y_true)] * df.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="6gS66KV6fEZw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="175d7621-7016-4d19-e5b8-fd069f4e8ecb" executionInfo={"status": "ok", "timestamp": 1581630530279, "user_tz": -60, "elapsed": 611, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
y_true = df['prices_amountmin']
price_log_mean = np.expm1(np.mean( np.log1p(y_true) ))
y_pred = [price_log_mean] * y_true.shape[0]
mean_absolute_error(y_true, y_pred)
# + id="KAkVjYXagRpt" colab_type="code" colab={}
df['brand_cat'] = df['brand'].factorize()[0]
# + id="VOK-jFargsW1" colab_type="code" colab={}
def run_model(feats):
x = df[feats].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, x, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="fQWXIfadl0oj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2b3e25b-21c4-40a3-bcb4-5c353f4e824f" executionInfo={"status": "ok", "timestamp": 1581631672599, "user_tz": -60, "elapsed": 635, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
run_model(['brand_cat'])
# + id="tBZo2wnFl8fQ" colab_type="code" colab={}
df['manufacturer_cat'] = df['manufacturer'].factorize()[0]
# + id="mTrPVz9XmiEG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="224a0abc-bc1b-479f-af8b-482acb0e86a8" executionInfo={"status": "ok", "timestamp": 1581631868061, "user_tz": -60, "elapsed": 667, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
run_model(['manufacturer_cat'])
# + id="EycVhGXhmwmw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3a4d4f52-2ae5-4af2-b208-46c9b56cbdd3" executionInfo={"status": "ok", "timestamp": 1581631884412, "user_tz": -60, "elapsed": 542, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
run_model(['manufacturer_cat', 'brand_cat'])
# + id="manF26reorCa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3369fb4a-0364-4028-c2f0-48891f41e586" executionInfo={"status": "ok", "timestamp": 1581632465179, "user_tz": -60, "elapsed": 1837, "user": {"displayName": "<NAME>\u00f3rnisiewicz", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCHJCMEanP9_BT7pj2iBUxjszpDo4mppKcg8jBmKA=s64", "userId": "10504772661452646683"}}
# !git add
# + id="qmAuGtiEpCFS" colab_type="code" colab={}
|
matrix_one/Day_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="TBFXQGKYUc4X"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="1z4xy2gTUc4a" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="FE7KNzPPVrVV"
# # Image classification
# + [markdown] colab_type="text" id="KwQtSOz0VrVX"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/images/classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ja/tutorials/images/classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ja/tutorials/images/classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/ja/tutorials/images/classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="DjmULc9dAvL-" colab_type="text"
# Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [<EMAIL> メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。
# + [markdown] colab_type="text" id="gN7G9GFmVrVY"
# このチュートリアルでは、画像から猫または犬を分類する方法を示します。 `tf.keras.Sequential` モデルを使用して画像分類器を構築し、 `tf.keras.preprocessing.image.ImageDataGenerator` を使用してデータをロードします。このチュートリアルでは、以下のコンセプトにしたがって、実践的な経験と感覚を養います。
#
# * `tf.keras.preprocessing.image.ImageDataGenerator` クラスを使用して _データ入力パイプライン_ を構築し、モデルで使用するディスク上のデータを効率的に処理します。
# * _過学習(Overfitting)_ —過学習を識別および防止する方法。
# * _データ拡張(Data Augmentation)_ および _ドロップアウト(dropout)_ —データパイプラインおよび画像分類モデルに組み込むコンピュータービジョンタスクの過学習と戦うための重要なテクニック。
#
# このチュートリアルは、基本的な機械学習のワークフローに従います。
#
# 1. データの調査及び理解
# 2. 入力パイプラインの構築
# 3. モデルの構築
# 4. モデルの学習
# 5. モデルのテスト
# 6. モデルの改善とプロセスの繰り返し
# + [markdown] colab_type="text" id="zF9uvbXNVrVY"
# ## パッケージのインポート
# + [markdown] colab_type="text" id="VddxeYBEVrVZ"
# まずは必要なパッケージをインポートすることから始めましょう。 `os`パッケージはファイルとディレクトリ構造を読み込み、 NumPy は python リストの numpy 配列への変換と必要な行列演算の実行、 `matplotlib.pyplot` はグラフの描画や学習データおよび検証データに含まれる画像の表示、に利用します。
# + colab_type="code" id="rtPGh2MAVrVa" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# + [markdown] colab_type="text" id="Jlchl4x2VrVg"
# モデルの構築に必要な TensorFlow と Keras クラスをインポートします。
# + colab_type="code" id="E82grprdYPI0" colab={}
try:
# # %tensorflow_version は Colab にのみ存在します。
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + colab_type="code" id="L1WtoaOHVrVh" colab={}
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, Flatten, Dropout, MaxPooling2D
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import os
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] colab_type="text" id="UZZI6lNkVrVm"
# ## データの読み込み
# + [markdown] colab_type="text" id="DPHx8-t-VrVo"
# データセットのダウンロードから始めます。このチュートリアルでは、 Kaggle の <a href="https://www.kaggle.com/c/dogs-vs-cats/data" target="_blank">Dogs vs Cats</a> データセットをフィルタリングしたバージョンを使用します。データセットのアーカイブバージョンをダウンロードし、"/tmp/"ディレクトリに保存します。
# + colab_type="code" id="C1nqr-CYY6uw" colab={}
_URL = 'https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip'
path_to_zip = tf.keras.utils.get_file('cats_and_dogs.zip', origin=_URL, extract=True)
PATH = os.path.join(os.path.dirname(path_to_zip), 'cats_and_dogs_filtered')
# + [markdown] colab_type="text" id="Giv0wMQzVrVw"
# データセットのディレクトリ構造は次のとおりです:
#
# <pre>
# <b>cats_and_dogs_filtered</b>
# |__ <b>train</b>
# |______ <b>cats</b>: [cat.0.jpg, cat.1.jpg, cat.2.jpg ....]
# |______ <b>dogs</b>: [dog.0.jpg, dog.1.jpg, dog.2.jpg ...]
# |__ <b>validation</b>
# |______ <b>cats</b>: [cat.2000.jpg, cat.2001.jpg, cat.2002.jpg ....]
# |______ <b>dogs</b>: [dog.2000.jpg, dog.2001.jpg, dog.2002.jpg ...]
# </pre>
# + [markdown] colab_type="text" id="VpmywIlsVrVx"
# データの内容を抽出した後、学習および検証セットのための適切なファイルパスで変数を設定します。
# + colab_type="code" id="sRucI3QqVrVy" colab={}
train_dir = os.path.join(PATH, 'train')
validation_dir = os.path.join(PATH, 'validation')
# + colab_type="code" id="Utv3nryxVrV0" colab={}
train_cats_dir = os.path.join(train_dir, 'cats') # 学習用の猫画像のディレクトリ
train_dogs_dir = os.path.join(train_dir, 'dogs') # 学習用の犬画像のディレクトリ
validation_cats_dir = os.path.join(validation_dir, 'cats') # 検証用の猫画像のディレクトリ
validation_dogs_dir = os.path.join(validation_dir, 'dogs') # 検証用の犬画像のディレクトリ
# + [markdown] colab_type="text" id="ZdrHHTy2VrV3"
# ### データの理解
# + [markdown] colab_type="text" id="LblUYjl-VrV3"
# 学習および検証ディレクトリの中にある猫と犬の画像の数を見てみましょう:
# + colab_type="code" id="vc4u8e9hVrV4" colab={}
num_cats_tr = len(os.listdir(train_cats_dir))
num_dogs_tr = len(os.listdir(train_dogs_dir))
num_cats_val = len(os.listdir(validation_cats_dir))
num_dogs_val = len(os.listdir(validation_dogs_dir))
total_train = num_cats_tr + num_dogs_tr
total_val = num_cats_val + num_dogs_val
# + colab_type="code" id="g4GGzGt0VrV7" colab={}
print('total training cat images:', num_cats_tr)
print('total training dog images:', num_dogs_tr)
print('total validation cat images:', num_cats_val)
print('total validation dog images:', num_dogs_val)
print("--")
print("Total training images:", total_train)
print("Total validation images:", total_val)
# + [markdown] colab_type="text" id="8Lp-0ejxOtP1"
# 便宜上、データセットの前処理およびネットワークの学習中に使用する変数を設定します。
# + colab_type="code" id="3NqNselLVrWA" colab={}
batch_size = 128
epochs = 15
IMG_HEIGHT = 150
IMG_WIDTH = 150
# + [markdown] colab_type="text" id="INn-cOn1VrWC"
# ## データの準備
# + [markdown] colab_type="text" id="5Jfk6aSAVrWD"
# モデルにデータを送る前に、画像を適切に前処理された浮動小数点テンソルにフォーマットします。
#
# 1.ディスクから画像を読み取ります。
# 2.これらの画像のコンテンツをデコードし、RGB値にしたがって適切なグリッド形式に変換します。
# 3.それらを浮動小数点テンソルに変換します。
# 4.ニューラルネットワークは小さな入力値を扱う方が適しているため、テンソルを0〜255の値から0〜1の値にリスケーリングします。
#
# 幸い、これらすべてのタスクは、 `tf.keras` によって提供される `ImageDataGenerator` クラスで実行できます。この `ImageDataGenerator` はディスクから画像を読み取り、適切なテンソルに前処理を行います。さらに、これらの画像をテンソルのバッチに変換するジェネレータをセットアップします。これは、ネットワーク学習時に便利です。
# + colab_type="code" id="syDdF_LWVrWE" colab={}
train_image_generator = ImageDataGenerator(rescale=1./255) # 学習データのジェネレータ
validation_image_generator = ImageDataGenerator(rescale=1./255) # 検証データのジェネレータ
# + [markdown] colab_type="text" id="RLciCR_FVrWH"
# 学習および検証画像のジェネレータを定義したのち、 `flow_from_directory` メソッドはディスクから画像をロードし、リスケーリングを適用し、画像を必要な大きさにリサイズします。
# + colab_type="code" id="Pw94ajOOVrWI" colab={}
train_data_gen = train_image_generator.flow_from_directory(batch_size=batch_size,
directory=train_dir,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH),
class_mode='binary')
# + colab_type="code" id="2oUoKUzRVrWM" colab={}
val_data_gen = validation_image_generator.flow_from_directory(batch_size=batch_size,
directory=validation_dir,
target_size=(IMG_HEIGHT, IMG_WIDTH),
class_mode='binary')
# + [markdown] colab_type="text" id="hyexPJ8CVrWP"
# ### 学習用画像の可視化
# + [markdown] colab_type="text" id="60CnhEL4VrWQ"
# 学習用のジェネレータから画像バッチを抽出して可視化します。(この例では32個の画像を抽出し、そのうち5つを `matplotlib` で描画します。)
# + colab_type="code" id="3f0Z7NZgVrWQ" colab={}
sample_training_images, _ = next(train_data_gen)
# + [markdown] colab_type="text" id="49weMt5YVrWT"
# `next` 関数はデータセットからバッチを返します。 `next` 関数の返り値は `(x_train、y_train)` の形式で、 `x_train` は学習用の特徴量、 `y_train` はそのラベルです。ラベルを破棄して、学習用画像の可視化のみを行います。
# + colab_type="code" id="JMt2RES_VrWU" colab={}
# この関数は、1行5列のグリッド形式で画像をプロットし、画像は各列に配置されます。
def plotImages(images_arr):
fig, axes = plt.subplots(1, 5, figsize=(20,20))
axes = axes.flatten()
for img, ax in zip( images_arr, axes):
ax.imshow(img)
ax.axis('off')
plt.tight_layout()
plt.show()
# + colab_type="code" id="d_VVg_gEVrWW" colab={}
plotImages(sample_training_images[:5])
# + [markdown] colab_type="text" id="b5Ej-HLGVrWZ"
# ## モデルの構築
# + [markdown] colab_type="text" id="wEgW4i18VrWZ"
# モデルはmax pooling層を伴う3つの畳み込みブロックからなります。さらに `relu` 活性化関数によるアクティベーションを伴う512ユニットの全結合層があります。モデルは、シグモイド活性化関数による2値分類に基づいてクラスに属する確率を出力します。
# + colab_type="code" id="F15-uwLPVrWa" colab={}
model = Sequential([
Conv2D(16, 3, padding='same', activation='relu', input_shape=(IMG_HEIGHT, IMG_WIDTH ,3)),
MaxPooling2D(),
Conv2D(32, 3, padding='same', activation='relu'),
MaxPooling2D(),
Conv2D(64, 3, padding='same', activation='relu'),
MaxPooling2D(),
Flatten(),
Dense(512, activation='relu'),
Dense(1, activation='sigmoid')
])
# + [markdown] colab_type="text" id="PI5cdkMQVrWc"
# ### モデルのコンパイル
# このチュートリアルでは、 *ADAM* オプティマイザーと *binary cross entropy* 損失関数を選択します。各学習エポックの学習と検証の精度を表示するために、`metrics` 引数を渡します。
# + colab_type="code" id="6Mg7_TXOVrWd" colab={}
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="2YmQZ3TAVrWg"
# ### モデルの概要
#
# すべてのネットワークのレイヤーを見るには、モデルの `summary` メソッドを利用します:
# + colab_type="code" id="Vtny8hmBVrWh" colab={}
model.summary()
# + [markdown] colab_type="text" id="N06iqE8VVrWj"
# ### モデルの学習
# + [markdown] colab_type="text" id="oub9RtoFVrWk"
# `ImageDataGenerator` クラスの `fit_generator` メソッドを使用して、ネットワークを学習します。
# + colab_type="code" id="KSF2HqhDVrWk" colab={}
history = model.fit_generator(
train_data_gen,
steps_per_epoch=total_train // batch_size,
epochs=epochs,
validation_data=val_data_gen,
validation_steps=total_val // batch_size
)
# + [markdown] colab_type="text" id="ojJNteAGVrWo"
# ### 学習結果の可視化
# + [markdown] colab_type="text" id="LZPYT-EmVrWo"
# ネットワークを学習した後、結果を可視化します。
# + colab_type="code" id="K6oA77ADVrWp" colab={}
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(epochs)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
# + [markdown] colab_type="text" id="kDnr50l2VrWu"
# プロットからわかるように、学習セットの精度と検証セットの精度は大幅に外れており、モデルは検証セットで約70%の精度しか達成していません。
#
# 何がうまくいかなかったかを見て、モデル全体のパフォーマンスを向上してみましょう。
# + [markdown] colab_type="text" id="rLO7yhLlVrWu"
# ## 過学習
# + [markdown] colab_type="text" id="hNyx3Lp4VrWv"
# 上記のプロットでは、学習セットの精度は時間とともに直線的に向上していますが、検証セットの精度は学習プロセスの中で約70%あたりで頭打ちになっています。そして、学習と検証の精度の違いが顕著です。これは *過学習* のサインです。
#
# 学習サンプルが少ない場合、モデルは学習サンプルに含まれるノイズや不要な詳細から学習してしまい、これによって新しいサンプルに対するモデルの性能に悪影響を与えることがあります。この現象は、過学習として知られています。過学習とは、モデルが新しいデータセットに対して汎化するのが難しい状態をいいます。
#
# 学習プロセスにおいて過学習に対抗する手段はいくつかあります。このチュートリアルでは、*データ拡張(data Augmentation)* を使用し、さらにモデルに *ドロップアウト(dropout)* を追加します。
# + [markdown] colab_type="text" id="UOoVpxFwVrWy"
# ## データ拡張(Data augmentation)
# + [markdown] colab_type="text" id="Wn_QLciWVrWy"
# 過学習は一般に、学習サンプルが少ない場合に発生します。この問題を解決する方法の1つは、十分な数の学習サンプルが含まれるようにデータセットを拡張することです。データ拡張は、既存の学習サンプルに対してランダムな変換を行い、データセットとして利用できそうな画像を生成するアプローチをとります。このデータ拡張の目的は、学習中にモデルがまったくおなじ画像を2回利用しないようにすることです。これによってモデルをデータのより多くの特徴を利用し、より汎化することができます。
#
# `tf.keras` においては、このデータ拡張を `ImageDataGenerator` クラスを使用して実装します。データセットに対するさまざまな変換を指定することによって、学習プロセス中にそれが適用されます。
# + [markdown] colab_type="text" id="2uJ1G030VrWz"
# ### データの拡張と可視化
# + [markdown] colab_type="text" id="hvX7hHlgVrW0"
# 最初に、ランダムな水平反転による拡張をデータセットに適用し、それぞれの画像が変換後にどのように見えるかを確認します。
# + [markdown] colab_type="text" id="rlVj6VqaVrW0"
# ### 水平反転の適用
# + [markdown] colab_type="text" id="xcdvx4TVVrW1"
# このデータ拡張を適用するためには、 `ImageDataGenerator` クラスの引数として `horizontal_flip` を渡し、 `True`を設定します。
# + colab_type="code" id="Bi1_vHyBVrW2" colab={}
image_gen = ImageDataGenerator(rescale=1./255, horizontal_flip=True)
# + colab_type="code" id="zvwqmefgVrW3" colab={}
train_data_gen = image_gen.flow_from_directory(batch_size=batch_size,
directory=train_dir,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH))
# + [markdown] colab_type="text" id="zJpRSxJ-VrW7"
# 学習サンプルから1つのサンプル画像を取得する作業を5回繰り返して、おなじ画像に5回データ拡張が適用されるようにします。
# + colab_type="code" id="RrKGd_jjVrW7" colab={}
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
# + colab_type="code" id="EvBZoQ9xVrW9" colab={}
# 上で学習用画像の可視化のために定義、使用されたおなじカスタムプロット関数を再利用する
plotImages(augmented_images)
# + [markdown] colab_type="text" id="i7n9xcqCVrXB"
# ### 画像のランダムな回転
# + [markdown] colab_type="text" id="qXnwkzFuVrXB"
# 回転のデータ拡張を利用して学習用サンプルをランダムに左右45度の範囲で回転させてみましょう。
# + colab_type="code" id="1zip35pDVrXB" colab={}
image_gen = ImageDataGenerator(rescale=1./255, rotation_range=45)
# + colab_type="code" id="kVoWh4OIVrXD" colab={}
train_data_gen = image_gen.flow_from_directory(batch_size=batch_size,
directory=train_dir,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH))
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
# + colab_type="code" id="wmBx8NhrVrXK" colab={}
plotImages(augmented_images)
# + [markdown] colab_type="text" id="FOqGPL76VrXM"
# ### ズームによるデータ拡張の適用
# + [markdown] colab_type="text" id="NvqXaD8BVrXN"
# データセットにズームによるデータ拡張を適用して、画像をランダムに最大50%拡大します。
# + colab_type="code" id="tGNKLa_YVrXR" colab={}
image_gen = ImageDataGenerator(rescale=1./255, zoom_range=0.5)
# + colab_type="code" id="VOvTs32FVrXU" colab={}
train_data_gen = image_gen.flow_from_directory(batch_size=batch_size,
directory=train_dir,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH))
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
# + colab_type="code" id="-KQWw8IZVrXZ" colab={}
plotImages(augmented_images)
# + [markdown] colab_type="text" id="usS13KCNVrXd"
# ### すべてのデータ拡張を同時に利用する
# + [markdown] colab_type="text" id="OC8fIsalVrXd"
# ここまでで紹介したすべてのデータ拡張機能を適用します。ここでは、学習用画像に対して、リスケール、45度の回転、幅シフト、高さシフト、水平反転、ズームを適用しました。
# + colab_type="code" id="gnr2xujaVrXe" colab={}
image_gen_train = ImageDataGenerator(
rescale=1./255,
rotation_range=45,
width_shift_range=.15,
height_shift_range=.15,
horizontal_flip=True,
zoom_range=0.5
)
# + colab_type="code" id="K0Efxy7EVrXh" colab={}
train_data_gen = image_gen_train.flow_from_directory(batch_size=batch_size,
directory=train_dir,
shuffle=True,
target_size=(IMG_HEIGHT, IMG_WIDTH),
class_mode='binary')
# + [markdown] colab_type="text" id="AW-pV5awVrXl"
# これらのデータ拡張がデータセットにランダムに適用されたときに、一つの画像に対して5回の個別の適用を行った際にそれぞれどのように見えるかを可視化します。
# + colab_type="code" id="z2m68eMhVrXm" colab={}
augmented_images = [train_data_gen[0][0][0] for i in range(5)]
plotImages(augmented_images)
# + [markdown] colab_type="text" id="J8cUd7FXVrXq"
# ### 検証データジェネレータの構築
# + [markdown] colab_type="text" id="a99fDBt7VrXr"
# 一般に、データ拡張は学習サンプルのみに適用します。今回は、 `ImageDataGenerator` を使用して検証画像に対してリスケールのみを実施し、バッチに変換します。
# + colab_type="code" id="54x0aNbKVrXr" colab={}
image_gen_val = ImageDataGenerator(rescale=1./255)
# + colab_type="code" id="1PCHKzI8VrXv" colab={}
val_data_gen = image_gen_val.flow_from_directory(batch_size=batch_size,
directory=validation_dir,
target_size=(IMG_HEIGHT, IMG_WIDTH),
class_mode='binary')
# + [markdown] colab_type="text" id="yQGhdqHFVrXx"
# ## ドロップアウト(dropout)
# + [markdown] colab_type="text" id="2Iq5TAH_VrXx"
# 過学習を避けるもう一つの方法は、ネットワークに *ドロップアウト* を導入することです。これは、ネットワークにおいて重みを小さくする正則化の方式で、これによって重みの値の分布がより規則的になり、少ない学習データに対する過学習を減らすことができます。ドロップアウトはこのチュートリアルで利用される正則化手法の一つです。
#
# ドロップアウトをレイヤーに適用すると、学習プロセス中に適用されたレイヤーのうちランダムに出力ユニットをドロップアウト(ゼロに設定)します。ドロップアウトは、入力値として0.1、0.2、0.4といった形式の小数をとります。これは、適用されたレイヤーからランダムに出力単位の10%、20%、または40%をドロップアウトすることを意味します。
#
# 特定のレイヤーに0.1ドロップアウトを適用すると、各学習エポックにおいて出力ユニットの10%がランダムに0にされます。
#
# この新しいドロップアウト機能を使用したネットワークアーキテクチャを作成し、異なる畳み込みレイヤーや全接続レイヤーに適用してみましょう。
# + [markdown] colab_type="text" id="DyxxXRmVVrXy"
# ## ドロップアウトを追加した新しいネットワークの構築
# + [markdown] colab_type="text" id="1Ba2LjtkVrXy"
# ここでは、ドロップアウトを最初と最後の max pool 層に適用します。ドロップアウトを適用すると、各学習エポック中にニューロンの20%がランダムにゼロに設定されます。これにより、学習データセットに対する過学習を避けることができます。
# + colab_type="code" id="2fjio8EsVrXz" colab={}
model_new = Sequential([
Conv2D(16, 3, padding='same', activation='relu',
input_shape=(IMG_HEIGHT, IMG_WIDTH ,3)),
MaxPooling2D(),
Dropout(0.2),
Conv2D(32, 3, padding='same', activation='relu'),
MaxPooling2D(),
Conv2D(64, 3, padding='same', activation='relu'),
MaxPooling2D(),
Dropout(0.2),
Flatten(),
Dense(512, activation='relu'),
Dense(1, activation='sigmoid')
])
# + [markdown] colab_type="text" id="tpTgIxWAVrX0"
# ### モデルのコンパイル
# + [markdown] colab_type="text" id="1osvc_iTVrX1"
# ネットワークにドロップアウトを導入した後、モデルをコンパイルし、レイヤーの概要を表示します。
# + colab_type="code" id="OkIJhS-WVrX1" colab={}
model_new.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
model_new.summary()
# + [markdown] colab_type="text" id="7KiDshEUVrX6"
# ### モデルの学習
# + [markdown] colab_type="text" id="NFj0oVqVVrX6"
# 学習サンプルにデータ拡張を導入し、ネットワークにドロップアウトを追加した後、この新しいネットワークを学習します:
# + colab_type="code" id="GWxHs_luVrX7" colab={}
history = model_new.fit_generator(
train_data_gen,
steps_per_epoch=total_train // batch_size,
epochs=epochs,
validation_data=val_data_gen,
validation_steps=total_val // batch_size
)
# + [markdown] colab_type="text" id="bbdyqZdxVrYA"
# ### モデルの可視化
# + [markdown] colab_type="text" id="OgvF2nt7OtR7"
# 学習後に新しいモデルを可視化すると、過学習が前回よりも大幅に少ないことがわかります。より多くのエポックでモデルを学習すると、精度はさらに向上するはずです。
# + colab_type="code" id="7BTeMuNAVrYC" colab={}
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs_range = range(epochs)
plt.figure(figsize=(8, 8))
plt.subplot(1, 2, 1)
plt.plot(epochs_range, acc, label='Training Accuracy')
plt.plot(epochs_range, val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.title('Training and Validation Accuracy')
plt.subplot(1, 2, 2)
plt.plot(epochs_range, loss, label='Training Loss')
plt.plot(epochs_range, val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.title('Training and Validation Loss')
plt.show()
|
site/ja/tutorials/images/classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelo 2021-09-28_Lab_Madrid_CopaBor_FondoBlanco_Xception_100ep
#
# Problema de clasificación con 6 clases.
#
# Modelo: Xception con image_size = 528, batch_size=16, 100 épocas. Tarda unos 52 s por época.
#
# 528 imágenes en total: 369 train + 53 val + 106 test que se han distribuido de forma homogénea de manera que el dataset esté equilibrado (ya que había la mitad de imágenes de clase <= 50 mL con respecto al resto de clases).
#
# Mean RGB pixel: [145.7009020327461, 145.57541800835594, 143.40409009710942]
# Standard deviation of RGB pixel: [37.09825214810984, 38.16266727994038, 37.706027031863435]
# +
import os
import json
import numpy as np
import matplotlib.pylab as plt
from tensorflow.keras.models import load_model
from imgclas.data_utils import load_image, load_data_splits, load_class_names
from imgclas.test_utils import predict
from imgclas import paths, plot_utils, utils
# User parameters to set
TIMESTAMP = '2021-09-28_Lab_Madrid_CopaBor_FondoBlanco_Xception_100ep' # timestamp of the model
MODEL_NAME = 'final_model.h5' # model to use to make the prediction
TOP_K = 2 # number of top classes predictions to save
# Set the timestamp
paths.timestamp = TIMESTAMP
# Load the data
class_names = load_class_names(splits_dir=paths.get_ts_splits_dir())
# Load training configuration
conf_path = os.path.join(paths.get_conf_dir(), 'conf.json')
with open(conf_path) as f:
conf = json.load(f)
# Load the model
model = load_model(os.path.join(paths.get_checkpoints_dir(), MODEL_NAME), custom_objects=utils.get_custom_objects(), compile=False)
# -
# ## Predicciones
#
# ### Test Laboratorio/Madrid/copa Bor/Fondo blanco
# cd ..
# +
SPLIT_NAME = 'test' # data split to use (nombre del .txt que vas a leer, en este caso test)
# conf['general']['images_directory'] = '/media/ignacio/Datos/datasets/semillas/datasets/RJB' # custom the absolute path to the images directory, if needed
# Load the data
X, y = load_data_splits(splits_dir=paths.get_ts_splits_dir(),
im_dir=conf['general']['images_directory'],
split_name=SPLIT_NAME)
# Predict
pred_lab, pred_prob = predict(model, X, conf, top_K=TOP_K, filemode='local')
# Save the predictions
pred_dict = {'filenames': list(X),
'pred_lab': pred_lab.tolist(),
'pred_prob': pred_prob.tolist()}
if y is not None:
pred_dict['true_lab'] = y.tolist()
pred_path = os.path.join(paths.get_predictions_dir(), '{}+{}+top{}.json'.format(MODEL_NAME, SPLIT_NAME, TOP_K))
with open(pred_path, 'w') as outfile:
json.dump(pred_dict, outfile, sort_keys=True)
# +
from imgclas import test_utils
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
true_lab, pred_lab = np.array(pred_dict['true_lab']), np.array(pred_dict['pred_lab'])
top1 = test_utils.topK_accuracy(true_lab, pred_lab, K=1)
# top5 = test_utils.topK_accuracy(true_lab, pred_lab, K=5)
print('Top1 accuracy: {:.1f} %'.format(top1 * 100))
# print('Top5 accuracy: {:.1f} %'.format(top5 * 100))
labels = range(len(class_names))
print('\n')
print('Micro recall: {:.1f} %'.format(100 * recall_score(true_lab, pred_lab[:, 0], labels=labels, average='micro')))
print('Macro recall: {:.1f} %'.format(100 * recall_score(true_lab, pred_lab[:, 0], labels=labels, average='macro')))
print('Macro recall (no labels): {:.1f} %'.format(100 * recall_score(true_lab, pred_lab[:, 0], average='macro')))
print('Weighted recall: {:.1f} %'.format(100 * recall_score(true_lab, pred_lab[:, 0], labels=labels, average='weighted')))
print('\n')
print('Micro precision: {:.1f} %'.format(100 * precision_score(true_lab, pred_lab[:, 0], labels=labels, average='micro')))
print('Macro precision: {:.1f} %'.format(100 * precision_score(true_lab, pred_lab[:, 0], labels=labels, average='macro')))
print('Macro precision (no labels): {:.1f} %'.format(100 * precision_score(true_lab, pred_lab[:, 0], average='macro')))
print('Weighted precision: {:.1f} %'.format(100 * precision_score(true_lab, pred_lab[:, 0], labels=labels, average='weighted')))
print('\n')
print('Micro F1 score: {:.1f} %'.format(100 * f1_score(true_lab, pred_lab[:, 0], labels=labels, average='micro')))
print('Macro F1 score: {:.1f} %'.format(100 * f1_score(true_lab, pred_lab[:, 0], labels=labels, average='macro')))
print('Macro F1 score (no labels): {:.1f} %'.format(100 * f1_score(true_lab, pred_lab[:, 0], average='macro')))
print('Weighted F1 score: {:.1f} %'.format(100 * f1_score(true_lab, pred_lab[:, 0], labels=labels, average='weighted')))
# +
print("Confusion matrix")
import matplotlib.pylab as plt
import seaborn
from sklearn.metrics import confusion_matrix
def plt_conf_matrix(conf_mat, labels=False):
fig = plt.figure(figsize=(20, 20))
hm = seaborn.heatmap(conf_mat, annot=False, square=True, cbar_kws={'fraction':0.046, 'pad':0.04},
xticklabels=labels, yticklabels=labels)
fontsize = None
hm.yaxis.set_ticklabels(hm.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=fontsize)
hm.xaxis.set_ticklabels(hm.xaxis.get_ticklabels(), rotation=90, ha='right', fontsize=fontsize)
plt.ylabel('True label')
plt.xlabel('Predicted label')
y_true, y_pred = np.array(pred_dict['true_lab']), np.array(pred_dict['pred_lab'])[:, 0]
conf_mat = confusion_matrix(y_true, y_pred, labels=range(len(class_names)), sample_weight=None)
normed_conf = conf_mat / np.sum(conf_mat, axis=1)[:, np.newaxis]
# plt_conf_matrix(conf_mat)
plt_conf_matrix(normed_conf, labels=class_names)
|
notebooks/N_2021-09-28_Lab_Madrid_CopaBor_FondoBlanco_Xception_100ep.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# PREPARATION
import pandas as pd
import os
import re
# read file
data = pd.read_csv('stock_data.csv', encoding='utf-8')
countPos = 0
countNeg = 0
count = 0
def format_sentence(sentence) :
# split text into one sentence per line
sentenceSplit = sentence.split(". ")
newSentence = ''
# add white space around punctuation like periods, commas, and brackets.
for s in sentenceSplit:
s = re.sub('([.,!?()])', r' \1 ', s)
s = re.sub('\s{2,}', ' ', s)
newSentence += s + '.\n'
return newSentence
# all text convert to lowercase with lower()
for line in data.values:
if str(line[1]) == '1':
with open('stock twitter data - 2/all/pos/pos_{}.txt'.format(countPos),'a+', encoding='utf-8') as f:
f.write(format_sentence(str(line[0]).lower()))
countPos += 1
if str(line[1]) == '-1':
with open('stock twitter data - 2/all/neg/neg_{}.txt'.format(countNeg),'a+', encoding='utf-8') as f:
format_sentence(str(line[0]).lower())
f.write(format_sentence(str(line[0]).lower()))
countNeg += 1
count += 1
|
cleaning_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import random
import numpy as np
import pandas as pd
import time
import matplotlib as plt
def pick_weekly_topic():
topics = np.random.choice( 350, 6, replace=False ).tolist()
#generate 6 random topics
#first 5 are the user's topics, last one is the random one
if topics[ -1 ] * 2 <= ( 350 * 2 * .05 ):
return topics[ -1 ]
else:
return topics[ -2 ]
def iter_topics( topics ):
wt = pick_weekly_topic()
topics = topics[ 1: ] + [ wt ]
return topics
def instantiate_topics( topics = [] ):
for i in range( 3 ):
topics.append( pick_weekly_topic() )
return topics
topics = instantiate_topics( )
topics_permutations_counts = {}
topics_permutations = {}
topics_permutations[ tuple( topics ) ] = 1
#time stuff
start_time = time.time()
last_run = start_time
for j in range( 1000 ):
if j % 100 == 0:
print( j )
print( "--- %s seconds ---" % (time.time() - last_run ) )
last_run = time.time()
for i in range( 1000000 ):
topics = iter_topics( topics )
if tuple( topics ) not in topics_permutations:
topics_permutations[ tuple( topics ) ] = 1
else:
topics_permutations[ tuple( topics ) ] += 1
if len( topics_permutations ) not in topics_permutations_counts:
topics_permutations_counts[ len( topics_permutations ) ] = 1
else:
topics_permutations_counts[ len( topics_permutations ) ] += 1
# -
perms = [ perm for perm_sublist in [ [ key ] * value for key, value in topics_permutations_counts.items() ] for perm in perm_sublist ]
perms = pd.DataFrame( perms, columns=['permutations'] )
perms.hist(bins=50)
#the permutations
print( perms )
print( perms[ 'permutations'].mean() )
|
TAPI permutation count.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Assignment
#
# These are the top 10 most frequently ordered products. How many times was each ordered?
#
# 1. Banana
# 2. Bag of Organic Bananas
# 3. Organic Strawberries
# 4. Organic Baby Spinach
# 5. Organic Hass Avocado
# 6. Organic Avocado
# 7. Large Lemon
# 8. Strawberries
# 9. Limes
# 10. Organic Whole Milk
#
# First, write down which columns you need and which dataframes have them.
#
# Next, merge these into a single dataframe.
#
# Then, use pandas functions from the previous lesson to get the counts of the top 10 most frequently ordered products.
from numba import jit
from dask import compute, delayed
# !wget https://s3.amazonaws.com/instacart-datasets/instacart_online_grocery_shopping_2017_05_01.tar.gz
import dask.dataframe as ddf
from dask.distributed import Client
# !tar --gunzip --extract --verbose --file=instacart_online_grocery_shopping_2017_05_01.tar.gz
# %cd instacart_2017_05_01
# !ls -lh *.csv
import pandas as pd
# %time
order_products = ddf.read_csv('order_products*.csv')
products = pd.read_csv('products.csv')
# %time
order_products = ddf.merge(order_products, products[['product_id', 'product_name']])
top_10 = order_products['product_name'].value_counts()
print(top_10.head(10))
|
module1-aws-sagemaker/Dask Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# Based on pdf_1.ncl
a = np.random.normal(0, 50, (64, 128))
b = np.random.chisquare(2, 1000)
c = np.random.gamma(scale=1/75,shape=2, size=(50, 100))
# # CHI
#
# Note the NCL function is for a Chi distribution, but the numpy one is for Chi-Squared.
#
# # GAMMA
# The gamma distribution is slightly different between NCL and numpy. It looks like NCL's location parameter is the inverse of Numpy's scale parameter.
#
# NCL uses Fortran code that says the density is:
#
# (A**R)/Gamma(R) * X**(R-1) * Exp(-A*X)
#
# where A is the shape parameter and R is the location parameter. Whereas Numpy's documentation says the probability density for the Gamma distribution is
#
# \begin{equation}
# p(x) = x^{k-1}\frac{e^{-x/\theta}}{\theta^k\Gamma(k)}
# \end{equation}
#
# where k is the shape and \theta the scale, and \Gamma is the Gamma function.
#
#
# +
# construct the distribution....
# simplest way is to histogram:
# analogous to NCL:
# ap = pdfx(a, 0, False) ; default number of bins
# bp = pdfx(b, 0, False)
# # cp = pdfx(c, 0, False)
# pdfx returns values in percent. with default 25 bins
ah1, ah1_bin_edges = np.histogram(a, bins=25, density=True) # density=True means the INTEGRAL is 1 ... NOT the sum unless bin width=1
bh1, bh1_bin_edges = np.histogram(b, bins=25, density=True)
ch1, ch1_bin_edges = np.histogram(c, bins=25, density=True)
# can get the bin centers: [left_edge] + 0.5*bin_width
ah1_centers = ah1_bin_edges[0:-1] + 0.5*(ah1_bin_edges[1:] - ah1_bin_edges[0:-1])
bh1_centers = bh1_bin_edges[0:-1] + 0.5*(bh1_bin_edges[1:] - bh1_bin_edges[0:-1])
ch1_centers = ch1_bin_edges[0:-1] + 0.5*(ch1_bin_edges[1:] - ch1_bin_edges[0:-1])
# -
ah1_centers
# Visualize the distributions
fig, ax = plt.subplots(ncols=3, constrained_layout=True)
ax[0].plot(ah1_centers, ah1)
ax[1].plot(bh1_centers, bh1)
ax[2].plot(ch1_centers, ch1)
# +
# Show that the integrals are 1:
def dist_integral(dens, bin_edges):
dx = bin_edges[1:] - bin_edges[0:-1]
return np.sum(dens * dx)
a_int = dist_integral(ah1, ah1_bin_edges)
b_int = dist_integral(bh1, bh1_bin_edges)
c_int = dist_integral(ch1, ch1_bin_edges)
print(f"The integral of the Normal distribution: {a_int}\n The integral of the $\Chi^2$ distribution: {b_int}\n The integral of the gamma distribution: {c_int}")
# -
# To convert to just the percentage, you just plot the density times the bin width; which is what we summed to get 1.
fig, ax = plt.subplots(ncols=3, constrained_layout=True, sharey=True)
ax[0].plot(ah1_centers, ah1*(ah1_bin_edges[1:] - ah1_bin_edges[0:-1])*100)
ax[1].plot(bh1_centers, bh1*(bh1_bin_edges[1:] - bh1_bin_edges[0:-1])*100)
ax[2].plot(ch1_centers, ch1*(ch1_bin_edges[1:] - ch1_bin_edges[0:-1])*100)
ax[0].set_ylabel("PDF (%)")
# # Kernel Density Estimate
#
# Another way to estimate the density is to use a kernel density estimate. This usually results in a smoother estimate of the distribution than a simple histogram. Here we can use a Gaussian KDE from SciPy and compare with the density estimate from the histograms. You can see they are pretty similar, but the KDEs are smoother. I used the bin centers to evaluate the KDE, but you could use many more ponts to make the curves even more smooth.
# +
from scipy import stats
a_kernel = stats.gaussian_kde(a.ravel())
b_kernel = stats.gaussian_kde(b.ravel())
c_kernel = stats.gaussian_kde(c.ravel())
# +
# Visualize the distributions
fig, ax = plt.subplots(figsize=(12,4), ncols=3, constrained_layout=True)
ax[0].plot(ah1_centers, ah1)
ax[0].plot(ah1_centers, a_kernel.evaluate(ah1_centers))
ax[1].plot(bh1_centers, bh1)
ax[1].plot(bh1_centers, b_kernel.evaluate(bh1_centers))
ax[2].plot(ch1_centers, ch1)
ax[2].plot(ch1_centers, c_kernel.evaluate(ch1_centers))
# +
# pdf_2.ncl
# Not much interesting in this second example.
# Generate data of different sizes:
a2 = np.random.normal( 0, 75, 1000)
b2 = np.random.normal( 25, 20, (10, 40))
c2 = np.random.normal( 5, 50, 500)
# NCL example uses 40 bins to build the 3 pdfs.
# Spice things up. Let's use 40 bins on the first one.
# Second one, let's specify some non-uniform bins
# Third one, let's just make our own very simple histogram
ah2, ah2_bin_edges = np.histogram(a2, bins=40, density=True) # density=True means the INTEGRAL is 1 ... NOT the sum unless bin width=1
bbins = [-100, -50, -25, -15, -10, -5, 0, 5, 10, 15, 25, 50, 60, 70, 80, 200]
bh2, bh2_bin_edges = np.histogram(b2, bins=bbins, density=True) # If bins is a string, it defines the method used to calculate the optimal bin width, as defined by histogram_bin_edges.
# NOTE: The histogram is computed over the flattened array.
ch2_bin_edges = np.linspace(c2.min(), c2.max(), 40)
ch2 = np.zeros( len(ch2_bin_edges)-1)
for i in range(len(ch2_bin_edges)-1):
# print(f"i = {i}, checking for {ch2_bin_edges[i]} <= x < {ch2_bin_edges[i+1]}")
ch2[i] = np.sum(np.where((c2 >= ch2_bin_edges[i]) & (c2 < ch2_bin_edges[i+1]), 1, 0))
# normalize ch2 to convert from counts to fraction:
print(ch2)
print(f"The sum of the ch2 is : {ch2.sum()}, the length is {c2.shape}")
ch2 /= c2.shape[0]
print(ch2)
# +
ah2_ctr = ah2_bin_edges[0:-1] + 0.5*(ah2_bin_edges[1:] - ah2_bin_edges[0:-1])
bh2_ctr = bh2_bin_edges[0:-1] + 0.5*(bh2_bin_edges[1:] - bh2_bin_edges[0:-1])
ch2_ctr = ch2_bin_edges[0:-1] + 0.5*(ch2_bin_edges[1:] - ch2_bin_edges[0:-1])
fig2, ax2 = plt.subplots()
ax2.plot(ah2_ctr, ah2, label="A")
ax2.plot(bh2_ctr, bh2, label="B")
ax2.plot(ch2_ctr, ch2, label="C")
fig2.legend()
# -
# To be fair, let's make sure we do everything as percent
# To convert to just the percentage, you just plot the density times the bin width; which is what we summed to get 1.
fig2a, ax2a = plt.subplots()
ax2a.plot(ah2_ctr, ah2*(ah2_bin_edges[1:] - ah2_bin_edges[0:-1])*100)
ax2a.plot(bh2_ctr, bh2*(bh2_bin_edges[1:] - bh2_bin_edges[0:-1])*100)
ax2a.plot(ch2_ctr, ch2*100) # already in fraction, bin width didn't matter
ax2a.set_ylabel("PDF (%)")
# Since we have uneven bins, maybe better to do this as a bar chart
fig2b, ax2b = plt.subplots()
ax2b.bar(ah2_ctr, ah2*(ah2_bin_edges[1:] - ah2_bin_edges[0:-1])*100, width=(ah2_bin_edges[1:] - ah2_bin_edges[0:-1]), alpha=.5, edgecolor='C0')
ax2b.bar(bh2_ctr, bh2*(bh2_bin_edges[1:] - bh2_bin_edges[0:-1])*100, width=(bh2_bin_edges[1:] - bh2_bin_edges[0:-1]), alpha=.5, edgecolor='C1')
ax2b.bar(ch2_ctr, ch2*100, width=(ch2_bin_edges[1:] - ch2_bin_edges[0:-1]), alpha=.5, edgecolor='C2')
ax2b.set_ylabel("PDF (%)")
# +
# To mimic the "stepped" or "outlined" style of NCL, two options:
# Use Matplotlib's plt.hist() with kwarg histtype : {'bar', 'barstacked', 'step', 'stepfilled'} set to 'step'
# Or make the line step using drawstyle
fig2c, ax2c = plt.subplots()
ax2c.plot(ah2_ctr, ah2*(ah2_bin_edges[1:] - ah2_bin_edges[0:-1])*100, drawstyle='steps', label='A')
ax2c.plot(bh2_ctr, bh2*(bh2_bin_edges[1:] - bh2_bin_edges[0:-1])*100, drawstyle='steps', label='B')
ax2c.plot(ch2_ctr, ch2*100, drawstyle='steps', label='C')
ax2c.set_ylabel("PDF (%)")
fig2c.legend()
# NOTE: you can change where the step is with steps-pre, steps-mid, steps-post; I didn't confirm that we chose the right one here.
# +
# 2-dimensions
# pdf_3.ncl
xvals = np.random.normal(0, 5, 10000)
yvals = np.random.normal(40, 25, 10000)
# simplest, let matplotlib just do it:
fig3a, ax3a = plt.subplots()
ax3a.hist2d(xvals, yvals, bins=30, cmap='Blues');
# -
# More control using numpy
counts, xedges, yedges = np.histogram2d(xvals, yvals, bins=30, density=False)
xcenter = xedges[0:-1]+0.5*(xedges[1]-xedges[0])
ycenter = yedges[0:-1]+0.5*(yedges[1]-yedges[0])
xgrid, ygrid = np.meshgrid(xcenter, ycenter)
fig3b, ax3b = plt.subplots()
img = ax3b.contourf(xgrid, ygrid, counts/len(xvals.flatten()), cmap='Blues')
ctr = ax3b.contour(xgrid, ygrid, counts/len(xvals.flatten()), colors='gray')
clbs = ax3b.clabel(ctr, fontsize=10, colors='black', inline=True, fmt='%1.3f')
|
Examples/Notebooks/pdf_examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="TcYcW7FyCHL_"
# # 2.5 Common Runtimes
# + colab={"base_uri": "https://localhost:8080/", "height": 586} id="-bHQ_TETCJ2e" outputId="5f330d90-ce2c-4142-f5d1-b6aac1961cc8"
path = "/content/here/MyDrive/Data and Algorithms/ALGO02/big-O complexity chart.png"
show_img(path, source="Know Thy Complexities! from https://www.bigocheatsheet.com/", source_scale=0.375)
# + [markdown] id="jwkiwxxZ2GTG"
# - $O(1)$: constant.
# - first item in list;
# - is list length odd or even?
# - $O(n)$: linear.
# - $O(100)$ hundred times longer.
# - search linearly through list;
# - print every item in list
# - $O(n^2)$: polynomial.
# - the compute time goes up by the square.
# - loop nested within loop;
# - bubble sort
# - $O(\text{log }n)$: logarithmic.
# - very efficient time. While we increase data point exponentially, the runtime increases only linearly from 1 to 2 to 3.
# - One of the most efficient runtimes.
# - binary search
# - $O(2^n)$: exponential.
# - One of the worst runtimes.
# - Count combinations of list elements
# - $O(n\text{ log }n$
# - One of the most common
# - mergesort;
# - quick sort
# - $O(n!)$: factorial.
# - generate all permutations of a list;
# - simple Traveling Salesman solution
#
# + colab={"base_uri": "https://localhost:8080/", "height": 534} id="Lnnz9NIx8IWw" outputId="075e1309-e733-4040-c9f2-12b254c2750c"
path = "/content/here/MyDrive/Data and Algorithms/ALGO02/common data structur eoperations.png"
show_img(path, source="Know Thy Complexities! from https://www.bigocheatsheet.com/")
# + [markdown] id="PwihXntm2Id6"
# # 2.6 Best vs. Worst Case
# - Variance in Big O possible based on data
# - Example) for quick sort:
# - Best case is $O(n)$ if all items are equal
# - Worst case is $O(n^2)$ if "pivot" is always largest item.
# - Expected case (a.k.a. average case) is $O(n \text{ log }n)$
# - Somewhere in the middle
#
# <br/>
#
# Quick Sort
# - Best case: $O(n)$
# - Worst case: $O(n^2)$
# - Expected: $O(n \text{ log }n)$
#
# <br/>
#
# $O(2n) \rightarrow O(n)$
# - All that matters is the linearity level.
# - Constants are of little importance.
#
# $O(n^2 + n) \rightarrow O(n^2)$
# - Only the highest-degree term matters in polynomial time.
#
# Sometimes it is important to retain all the terms:
# - for example, in a situation where $m$ is laaaaarge
# - and $n$ is small; well, retain all of them.
# - $O(n^2 + m)$
# - $O(n \cdot m)$
# + [markdown] id="s_YUPloYF-SL"
# Why do we learn the Big O notation?
# - Because we can optimise the algorithm and runtime.
# + [markdown] id="elxDS14MZjIH"
# # Important Functions
#
# - `sns.lmplot(x='n', y='time', data=dataset, ci=None)`
# - lmplot = linear model (regression) plot
# - ci: confidence intreval
# - `pd.DataFrame(list(zip(list1, list2, ..., list_n))`
# - `list(zip(item1, item2 ... item_n))`
# - `time.process_time()`
# + [markdown] id="VG76405zM7fm"
# References
#
# My algorithm learning notebook following the live lesson series [**"Data Structures, Algorithms, and Machine Learning Optimization"**](https://learning.oreilly.com/videos/data-structures-algorithms/9780137644889/) by Dr. <NAME>. I adapted some and partially modified or added entirely new code. Notes largely based on and (some of them entirely) from Jon's notebooks and learning materials. The lesson and original notebook source code at:
#
# https://learning.oreilly.com/videos/data-structures-algorithms/9780137644889/
# https://github.com/jonkrohn/ML-foundations/blob/master/notebooks/7-algos-and-data-structures.ipynb
|
02 DSA and ML Optimisation/ALGO 02 Big O Notation 5 Common Runtime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Numpy Bicimlendirme
# * **ndim**: boyut sayisi
# * **shape**: boyut bilgisi
# * **size**: toplam eleman sayisi
# * **dtype**: array veri tipi
# +
import numpy as np
a = np.random.randint(10, size = 10)
#tek boyutlu bir array
a.ndim
a.shape
a.size
a.dtype
# -
b = np.random.randint(20, size = (10,5))
#iki boyutlu bir array(10 tane 5 uzunluğunda satır, 1den 10akadar)
b.ndim
b
b.size
b.dtype
a = np.random.randint(10, size = (3,5,2))
#3 boyutlu bir dizi
a.ndim
a
np.arange(1,10)
np.arange(1,10).reshape((3,3))
#yukarıdaki 1den 10a kadar olan arrayi
#reshape diyerek yeniden şekillendirip 3e3lük yaptık
#şeklini , dimensionunu (ndim)inini degistrdik
c = np.array([1,2,3])
c
c.ndim
d = c.reshape((1,3))
d
d.ndim
# +
#iki tane koseli parantezin icine yazılmıs olması sunu gosterir;
#goruldugu gibi reshape ile tek satırlı olan vektör olan
#arrayi de 2 boyutlu yaptık
#vektor formunda kaldı ama bilgisayar artık onu matris olarak dusunuyır
# -
c
c[np.newaxis, :]
# +
#bu seiklde de usttekinin aynisni yaptık
# -
c[:, np.newaxis]
# +
#aynisini tersten yazarak satır vektoru degil de
#sutun vektoru yaptik
# -
c.shape
|
numpy_array_bicimlendirme.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="jqVqT_Cxh4Ho"
# #Introduction to Neural Networks
# In this notebook you will learn how to create and use a neural network to classify articles of clothing. To achieve this, we will use a sub module of TensorFlow called *keras*.
#
# *This guide is based on the following TensorFlow documentation.*
#
# https://www.tensorflow.org/tutorials/keras/classification
#
#
#
# + [markdown] id="ZFQqW9r-ikJb"
# ##Keras
# Before we dive in and start discussing neural networks, I'd like to give a breif introduction to keras.
#
# From the keras official documentation (https://keras.io/) keras is described as follows.
#
# "Keras is a high-level neural networks API, written in Python and capable of running on top of TensorFlow, CNTK, or Theano. It was developed with a focus on enabling fast experimentation.
#
# Use Keras if you need a deep learning library that:
#
# - Allows for easy and fast prototyping (through user friendliness, modularity, and extensibility).
# - Supports both convolutional networks and recurrent networks, as well as combinations of the two.
# - Runs seamlessly on CPU and GPU."
#
# Keras is a very powerful module that allows us to avoid having to build neural networks from scratch. It also hides a lot of mathematical complexity (that otherwise we would have to implement) inside of helpful packages, modules and methods.
#
# In this guide we will use keras to quickly develop neural networks.
#
#
# + [markdown] id="Hivk879ZQhxU"
# ##What is a Neural Network
# So, what are these magical things that have been beating chess grandmasters, driving cars, detecting cancer cells and winning video games?
#
# A deep neural network is a layered representation of data. The term "deep" refers to the presence of multiple layers. Recall that in our core learning algorithms (like linear regression) data was not transformed or modified within the model, it simply existed in one layer. We passed some features to our model, some math was done, an answer was returned. The data was not changed or transformed throughout this process. A neural network processes our data differently. It attempts to represent our data in different ways and in different dimensions by applying specific operations to transform our data at each layer. Another way to express this is that at each layer our data is transformed in order to learn more about it. By performing these transformations, the model can better understand our data and therefore provide a better prediction.
#
#
# + [markdown] id="GOqUCZ2klTAq"
# ##How it Works
# Before going into too much detail I will provide a very surface level explination of how neural networks work on a mathematical level. All the terms and concepts I discuss will be defined and explained in more detail below.
#
# On a lower level neural networks are simply a combination of elementry math operations and some more advanced linear algebra. Each neural network consists of a sequence of layers in which data passes through. These layers are made up on neurons and the neurons of one layer are connected to the next (see below). These connections are defined by what we call a weight (some numeric value). Each layer also has something called a bias, this is simply an extra neuron that has no connections and holds a single numeric value. Data starts at the input layer and is trasnformed as it passes through subsequent layers. The data at each subsequent neuron is defined as the following.
#
# > $Y =(\sum_{i=0}^n w_i x_i) + b$
#
# > $w$ stands for the weight of each connection to the neuron
#
# > $x$ stands for the value of the connected neuron from the previous value
#
# > $b$ stands for the bias at each layer, this is a constant
#
# > $n$ is the number of connections
#
# > $Y$ is the output of the current neuron
#
# > $\sum$ stands for sum
#
# The equation you just read is called a weighed sum. We will take this weighted sum at each and every neuron as we pass information through the network. Then we will add what's called a bias to this sum. The bias allows us to shift the network up or down by a constant value. It is like the y-intercept of a line.
#
# But that equation is the not complete one! We forgot a crucial part, **the activation function**. This is a function that we apply to the equation seen above to add complexity and dimensionality to our network. Our new equation with the addition of an activation function $F(x)$ is seen below.
#
# > $Y =F((\sum_{i=0}^n w_i x_i) + b)$
#
# Our network will start with predefined activation functions (they may be different at each layer) but random weights and biases. As we train the network by feeding it data it will learn the correct weights and biases and adjust the network accordingly using a technqiue called **backpropagation** (explained below). Once the correct weights and biases have been learned our network will hopefully be able to give us meaningful predictions. We get these predictions by observing the values at our final layer, the output layer.
#
#
#
# + [markdown] id="o-oMh18_j5kl"
# ##Breaking Down The Neural Network!
#
# Before we dive into any code lets break down how a neural network works and what it does.
#
# 
# *Figure 1*
#
#
#
# + [markdown] id="-9hd-R1ulSdp"
# ###Data
# The type of data a neural network processes varies drastically based on the problem being solved. When we build a neural network, we define what shape and kind of data it can accept. It may sometimes be neccessary to modify our dataset so that it can be passed to our neural network.
#
# Some common types of data a neural network uses are listed below.
# - Vector Data (2D)
# - Timeseries or Sequence (3D)
# - Image Data (4D)
# - Video Data (5D)
#
# There are of course many different types or data, but these are the main categories.
#
#
# + [markdown] id="Xyxxs7oMlWtz"
# ###Layers
# As we mentioned earlier each neural network consists of multiple layers. At each layer a different transformation of data occurs. Our initial input data is fed through the layers and eventually arrives at the output layer where we will obtain the result.
# ####Input Layer
# The input layer is the layer that our initial data is passed to. It is the first layer in our neural network.
# ####Output Layer
# The output layer is the layer that we will retrive our results from. Once the data has passed through all other layers it will arrive here.
# ####Hidden Layer(s)
# All the other layers in our neural network are called "hidden layers". This is because they are hidden to us, we cannot observe them. Most neural networks consist of at least one hidden layer but can have an unlimited amount. Typically, the more complex the model the more hidden layers.
# ####Neurons
# Each layer is made up of what are called neurons. Neurons have a few different properties that we will discuss later. The important aspect to understand now is that each neuron is responsible for generating/holding/passing ONE numeric value.
#
# This means that in the case of our input layer it will have as many neurons as we have input information. For example, say we want to pass an image that is 28x28 pixels, thats 784 pixels. We would need 784 neurons in our input layer to capture each of these pixels.
#
# This also means that our output layer will have as many neurons as we have output information. The output is a little more complicated to understand so I'll refrain from an example right now but hopefully you're getting the idea.
#
# But what about our hidden layers? Well these have as many neurons as we decide. We'll discuss how we can pick these values later but understand a hidden layer can have any number of neurons.
# ####Connected Layers
# So how are all these layers connected? Well the neurons in one layer will be connected to neurons in the subsequent layer. However, the neurons can be connected in a variety of different ways.
#
# Take for example *Figure 1* (look above). Each neuron in one layer is connected to every neuron in the next layer. This is called a **dense** layer. There are many other ways of connecting layers but well discuss those as we see them.
#
#
# + [markdown] id="a_bM6nQ-PZBY"
# ###Weights
# Weights are associated with each connection in our neural network. Every pair of connected nodes will have one weight that denotes the strength of the connection between them. These are vital to the inner workings of a neural network and will be tweaked as the neural network is trained. The model will try to determine what these weights should be to achieve the best result. Weights start out at a constant or random value and will change as the network sees training data.
# + [markdown] id="XwYq9doXeIl-"
# ###Biases
# Biases are another important part of neural networks and will also be tweaked as the model is trained. A bias is simply a constant value associated with each layer. It can be thought of as an extra neuron that has no connections. The purpose of a bias is to shift an entire activation function by a constant value. This allows a lot more flexibllity when it comes to choosing an activation and training the network. There is one bias for each layer.
# + [markdown] id="F92rhvd6PcRI"
# ###Activation Function
# Activation functions are simply a function that is applied to the weighed sum of a neuron. They can be anything we want but are typically higher order/degree functions that aim to add a higher dimension to our data. We would want to do this to introduce more comolexity to our model. By transforming our data to a higher dimension, we can typically make better, more complex predictions.
#
# A list of some common activation functions and their graphs can be seen below.
#
# - Relu (Rectified Linear Unit)
#
# 
# - Tanh (Hyperbolic Tangent)
#
# 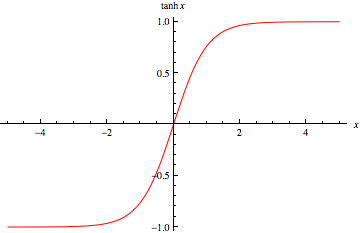
# - Sigmoid
#
# 
#
#
# + [markdown] id="Q2xNjpctlBUM"
# ###Backpropagation
# Backpropagation is the fundemental algorithm behind training neural networks. It is what changes the weights and biases of our network. To fully explain this process, we need to start by discussing something called a cost/loss function.
#
# ####Loss/Cost Function
# As we now know our neural network feeds information through the layers until it eventually reaches an output layer. This layer contains the results that we look at to determine the prediciton from our network. In the training phase it is likely that our network will make many mistakes and poor predicitions. In fact, at the start of training our network doesn't know anything (it has random weights and biases)!
#
# We need some way of evaluating if the network is doing well and how well it is doing. For our training data we have the features (input) and the labels (expected output), because of this we can compare the output from our network to the expected output. Based on the difference between these values we can determine if our network has done a good job or poor job. If the network has done a good job, we'll make minor changes to the weights and biases. If it has done a poor job our changes may be more drastic.
#
# So, this is where the cost/loss function comes in. This function is responsible for determining how well the network did. We pass it the output and the expected output, and it returns to us some value representing the cost/loss of the network. This effectively makes the networks job to optimize this cost function, trying to make it as low as possible.
#
# Some common loss/cost functions include.
# - Mean Squared Error
# - Mean Absolute Error
# - Hinge Loss
#
# ####Gradient Descent
# Gradient descent and backpropagation are closely related. Gradient descent is the algorithm used to find the optimal paramaters (weights and biases) for our network, while backpropagation is the process of calculating the gradient that is used in the gradient descent step.
#
# Gradient descent requires some pretty advanced calculus and linear algebra to understand so we'll stay away from that for now. Let's just read the formal definition for now.
#
# "Gradient descent is an optimization algorithm used to minimize some function by iteratively moving in the direction of steepest descent as defined by the negative of the gradient. In machine learning, we use gradient descent to update the parameters of our model." (https://ml-cheatsheet.readthedocs.io/en/latest/gradient_descent.html)
#
# And that's all we really need to know for now. I'll direct you to the video for a more in depth explination.
#
# 
#
#
# + [markdown] id="0KiTMDCKlBI7"
# ###Optimizer
# You may sometimes see the term optimizer or optimization function. This is simply the function that implements the backpropagation algorithm described above. Here's a list of a few common ones.
# - Gradient Descent
# - Stochastic Gradient Descent
# - Mini-Batch Gradient Descent
# - Momentum
# - Nesterov Accelerated Gradient
#
# *This article explains them quite well is where I've pulled this list from.*
#
# (https://medium.com/@sdoshi579/optimizers-for-training-neural-network-59450d71caf6)
# + [markdown] id="Kc5hFCLSiDNr"
# ##Creating a Neural Network
# Okay now you have reached the exciting part of this tutorial! No more math and complex explinations. Time to get hands on and train a very basic neural network.
#
# *As stated earlier this guide is based off of the following TensorFlow tutorial.*
# https://www.tensorflow.org/tutorials/keras/classification
#
# + [markdown] id="3io6gbUrjOQY"
# ###Imports
# + id="y8t_EdO8jEHz"
# %tensorflow_version 2.x # this line is not required unless you are in a notebook
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="p_iFN10li6V1"
# ###Dataset
# For this tutorial we will use the MNIST Fashion Dataset. This is a dataset that is included in keras.
#
# This dataset includes 60,000 images for training and 10,000 images for validation/testing.
# + id="eQmVmgOxjCOV"
fashion_mnist = keras.datasets.fashion_mnist # load dataset
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data() # split into tetsing and training
# + [markdown] id="AcIall2njfn1"
# Let's have a look at this data to see what we are working with.
# + id="WhLXRxOdjisI"
train_images.shape
# + [markdown] id="D2npdFHwjsLS"
# So we've got 60,000 images that are made up of 28x28 pixels (784 in total).
# + id="m280zyPqj3ws"
train_images[0,23,23] # let's have a look at one pixel
# + [markdown] id="GUciblEwkBe4"
# Our pixel values are between 0 and 255, 0 being black and 255 being white. This means we have a grayscale image as there are no color channels.
# + id="Rn78KO7fkQPJ"
train_labels[:10] # let's have a look at the first 10 training labels
# + [markdown] id="r90qZKsnkaW7"
# Our labels are integers ranging from 0 - 9. Each integer represents a specific article of clothing. We'll create an array of label names to indicate which is which.
# + id="pBiICD2tkne8"
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + [markdown] id="4rv06eD8krMR"
# Fianlly let's look at what some of these images look like!
# + id="Nfc8LV4Pkq0X"
plt.figure()
plt.imshow(train_images[1])
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] id="n_DC1b0grL1N"
# ##Data Preprocessing
# The last step before creating our model is to *preprocess* our data. This simply means applying some prior transformations to our data before feeding it the model. In this case we will simply scale all our greyscale pixel values (0-255) to be between 0 and 1. We can do this by dividing each value in the training and testing sets by 255.0. We do this because smaller values will make it easier for the model to process our values.
#
#
# + id="wHde8MYW0OQo"
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] id="dHOX6GqR0QuD"
# ##Building the Model
# Now it's time to build the model! We are going to use a keras *sequential* model with three different layers. This model represents a feed-forward neural network (one that passes values from left to right). We'll break down each layer and its architecture below.
# + id="XDxodHMv0xgG"
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)), # input layer (1)
keras.layers.Dense(128, activation='relu'), # hidden layer (2)
keras.layers.Dense(10, activation='softmax') # output layer (3)
])
# + [markdown] id="c-bL-I5w0414"
# **Layer 1:** This is our input layer and it will conist of 784 neurons. We use the flatten layer with an input shape of (28,28) to denote that our input should come in in that shape. The flatten means that our layer will reshape the shape (28,28) array into a vector of 784 neurons so that each pixel will be associated with one neuron.
#
# **Layer 2:** This is our first and only hidden layer. The *dense* denotes that this layer will be fully connected and each neuron from the previous layer connects to each neuron of this layer. It has 128 neurons and uses the rectify linear unit activation function.
#
# **Layer 3:** This is our output later and is also a dense layer. It has 10 neurons that we will look at to determine our models output. Each neuron represnts the probabillity of a given image being one of the 10 different classes. The activation function *softmax* is used on this layer to calculate a probabillity distribution for each class. This means the value of any neuron in this layer will be between 0 and 1, where 1 represents a high probabillity of the image being that class.
# + [markdown] id="-j1UF9QH21Ex"
# ###Compile the Model
# The last step in building the model is to define the loss function, optimizer and metrics we would like to track. I won't go into detail about why we chose each of these right now.
# + id="Msigq4Ja29QX"
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] id="7YYW5V_53OXV"
# ##Training the Model
# Now it's finally time to train the model. Since we've already done all the work on our data this step is as easy as calling a single method.
# + id="XmAtc4uI3_C7"
model.fit(train_images, train_labels, epochs=10) # we pass the data, labels and epochs and watch the magic!
# + [markdown] id="y6SRtNcF4K1O"
# ##Evaluating the Model
# Now it's time to test/evaluate the model. We can do this quite easily using another builtin method from keras.
#
# The *verbose* argument is defined from the keras documentation as:
# "verbose: 0 or 1. Verbosity mode. 0 = silent, 1 = progress bar."
# (https://keras.io/models/sequential/)
# + id="WqI0FEO54XN1"
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=1)
print('Test accuracy:', test_acc)
# + [markdown] id="nb4_EtfK5DuW"
# You'll likely notice that the accuracy here is lower than when training the model. This difference is reffered to as **overfitting**.
#
# And now we have a trained model that's ready to use to predict some values!
# + [markdown] id="Pv0XpgwJ7GlW"
# ##Making Predictions
# To make predictions we simply need to pass an array of data in the form we've specified in the input layer to ```.predict()``` method.
# + id="BMAkNWii7Ufj"
predictions = model.predict(test_images)
# + [markdown] id="LmRgxuEc7Xjc"
# This method returns to us an array of predictions for each image we passed it. Let's have a look at the predictions for image 1.
# + id="4y2eQtCr7fnd"
predictions[0]
# + [markdown] id="eiRNg9Yr7lCt"
# If we wan't to get the value with the highest score we can use a useful function from numpy called ```argmax()```. This simply returns the index of the maximium value from a numpy array.
# + id="NaagMfi671ci"
np.argmax(predictions[0])
# + [markdown] id="aWY4SKYm8h93"
# And we can check if this is correct by looking at the value of the cooresponding test label.
# + id="xVNepduo8nEy"
test_labels[0]
# + [markdown] id="Y8I1EqJu8qRl"
# ##Verifying Predictions
# I've written a small function here to help us verify predictions with some simple visuals.
# + id="-HJV4JF789aC"
COLOR = 'white'
plt.rcParams['text.color'] = COLOR
plt.rcParams['axes.labelcolor'] = COLOR
def predict(model, image, correct_label):
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
prediction = model.predict(np.array([image]))
predicted_class = class_names[np.argmax(prediction)]
show_image(image, class_names[correct_label], predicted_class)
def show_image(img, label, guess):
plt.figure()
plt.imshow(img, cmap=plt.cm.binary)
plt.title("Excpected: " + label)
plt.xlabel("Guess: " + guess)
plt.colorbar()
plt.grid(False)
plt.show()
def get_number():
while True:
num = input("Pick a number: ")
if num.isdigit():
num = int(num)
if 0 <= num <= 1000:
return int(num)
else:
print("Try again...")
num = get_number()
image = test_images[num]
label = test_labels[num]
predict(model, image, label)
# + [markdown] id="1HRzP5hCAijM"
# And that's pretty much it for an introduction to neural networks!
# + [markdown] id="PmbcLZZ0lo_2"
# ##Sources
#
# 1. <NAME>. “Various Optimization Algorithms For Training Neural Network.” Medium, Medium, 10 Mar. 2019, www.medium.com/@sdoshi579/optimizers-for-training-neural-network-59450d71caf6.
#
# 2. “Basic Classification: Classify Images of Clothing : TensorFlow Core.” TensorFlow, www.tensorflow.org/tutorials/keras/classification.
#
# 3. “Gradient Descent¶.” Gradient Descent - ML Glossary Documentation, www.ml-cheatsheet.readthedocs.io/en/latest/gradient_descent.html.
#
# 4. <NAME>. Deep Learning with Python. Manning Publications Co., 2018.
#
# 5. “Keras: The Python Deep Learning Library.” Home - Keras Documentation, www.keras.io/.
|
Neural_Networks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Introduction to Debugging
#
# In this book, we want to explore _debugging_ - the art and science of fixing bugs in computer software. In particular, we want to explore techniques that _automatically_ answer questions like: Where is the bug? When does it occur? And how can we repair it? But before we start automating the debugging process, we first need to understand what this process is.
#
# In this chapter, we introduce basic concepts of systematic software debugging and the debugging process, and at the same time get acquainted with Python and interactive notebooks.
# -
from bookutils import YouTubeVideo, quiz
YouTubeVideo("bCHRCehDOq0")
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Prerequisites**
#
# * The book is meant to be a standalone reference; however, a number of _great books on debugging_ are listed at the end,
# * Knowing a bit of _Python_ is helpful for understanding the code examples in the book.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from debuggingbook.Intro_Debugging import <identifier>
# ```
#
# and then make use of the following features.
#
#
# In this chapter, we introduce some basics of how failures come to be as well as a general process for debugging.
#
#
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## A Simple Function
# +
# ignore
# If this is the first time you are opening a notebook, please read this.
#
# Interaction basics
# ------------------
#
# * The notebook is composed of _code cells_ (like this one) and
# _text cells_ (like the one above).
#
# * To interact with the notebook, click into a cell (or type `Return`)
# and start editing. Type `Shift+Return` to execute the code, or
# `Esc` to leave without executing it.
#
# * Since code cells depend on earlier cells, you will have to execute
# those first - beginning from the top of the notebook.
#
# * You can change code (and text) at your leisure to try out alternatives.
# In Jupyter, use `b` to add a new cell below, and `Return` to enter it.
# Type `x` to delete (cut) a cell, and `z` to undo this.
#
# * There's a `Help` menu at the top of Jupyter Notebook. Enjoy!
#
#
# `# ignore` markers
# ------------------
#
# * In the notebook, there are some extra code cells starting with `# ignore`
# (like this one, actually). These are code blocks used to create diagrams,
# run tests, create or tear down special environments or more - code blocks
# that are not necessary for reading (but for creation).
#
#
# `# type: ignore` markers
# ------------------------
#
# * In the notebook, some lines come with a comment `# type: ignore`. This tells
# static code checkers to ignore any typing errors in that line. Most frequently,
# this is used to mark untyped Python code (as in our examples) such that the
# static code checker does not mark it as erroneous.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ### Your Task: Remove HTML Markup
#
# Let us start with a simple example. You may have heard of how documents on the Web are made out of text and HTML markup. HTML markup consists of _tags_ in angle brackets that surround the text, providing additional information on how the text should be interpreted. For instance, in the HTML text
#
# ```html
# This is <em>emphasized</em>.
# ```
#
# the word "emphasized" is enclosed in the HTML tags `<em>` (start) and `</em>` (end), meaning that it should be interpreted (and rendered) in an emphasized way – typically in italics. In your environment, the HTML text gets rendered as
#
# > This is <em>emphasized</em>.
#
# There's HTML tags for pretty much everything – text markup (<strong>bold</strong> text, <s>strikethrough</s> text), text structure (titles, lists), references (links) to other documents, and many more. These HTML tags shape the Web as we know it.
# -
# However, within all the HTML markup, it may become difficult to actually access the _text_ that lies within. We'd like to implement a simple function that removes _HTML markup_ and converts it into text. If our input is
#
# ```html
# Here's some <strong>strong argument</strong>.
# ```
# the output should be
#
# > Here's some strong argument.
# Here's a Python function which does exactly this. It takes a (HTML) string and returns the text without markup.
def remove_html_markup(s): # type: ignore
tag = False
out = ""
for c in s:
if c == '<': # start of markup
tag = True
elif c == '>': # end of markup
tag = False
elif not tag:
out = out + c
return out
# This function works, but not always. Before we start debugging things, let us first explore its code and how it works.
# ### Understanding Python Programs
#
# If you're new to Python, you might first have to understand what the above code does. We very much recommend the [Python tutorial](https://docs.python.org/3/tutorial/) to get an idea on how Python works. The most important things for you to understand the above code are these three:
#
# 1. Python structures programs through _indentation_, so the function and `for` bodies are defined by being indented;
# 2. Python is _dynamically typed_, meaning that the type of variables like `c`, `tag`, or `out` is determined at run-time.
# 3. Most of Python's syntactic features are inspired by other common languages, such as control structures (`while`, `if`, `for`), assignments (`=`), or comparisons (`==`, `!=`, `<`).
#
# With that, you can already understand what the above code does: `remove_html_markup()` takes a (HTML) string `s` and then iterates over the individual characters (`for c in s`). By default, these characters are added to the return string `out`. However, if `remove_html_markup()` finds a `<` character, it sets the `tag` flag, meaning that all further characters are ignored until a `>` character is found.
#
# In contrast to other languages, Python makes no difference between strings and characters – there's only strings. As in HTML, strings can be enclosed in single quotes (`'a'`) and in double quotes (`"a"`). This is useful if you want to specify a string that contains quotes, as in `'She said "hello", and then left'` or `"The first character is a 'c'"`
# ### Running a Function
#
# To find out whether `remove_html_markup()` works correctly, we can *test* it with a few values. For the string
#
# ```html
# Here's some <strong>strong argument</strong>.
# ```
#
# for instance, it produces the correct value:
remove_html_markup("Here's some <strong>strong argument</strong>.")
# ### Interacting with Notebooks
#
# If you are reading this in the interactive notebook, you can try out `remove_html_markup()` with other values as well. Click on the above cells with the invocation of `remove_html_markup()` and change the value – say, to `remove_html_markup("<em>foo</em>")`. Press <kbd>Shift</kbd>+<kbd>Enter</kbd> (or click on the play symbol) to execute it and see the result. If you get an error message, go to the above cell with the definition of `remove_html_markup()` and execute this first. You can also run _all_ cells at once; see the Notebook menu for details. (You can actually also change the text by clicking on it, and corect mistaks such as in this sentence.)
# Executing a single cell does not execute other cells, so if your cell builds on a definition in another cell that you have not executed yet, you will get an error. You can select `Run all cells above` from the menu to ensure all definitions are set.
# Also keep in mind that, unless overwritten, all definitions are kept across executions. Occasionally, it thus helps to _restart the kernel_ (i.e. start the Python interpreter from scratch) to get rid of older, superfluous definitions.
# ### Testing a Function
# Since one can change not only invocations, but also definitions, we want to ensure that our function works properly now and in the future. To this end, we introduce tests through _assertions_ – a statement that fails if the given _check_ is false. The following assertion, for instance, checks that the above call to `remove_html_markup()` returns the correct value:
assert remove_html_markup("Here's some <strong>strong argument</strong>.") == \
"Here's some strong argument."
# If you change the code of `remove_html_markup()` such that the above assertion fails, you will have introduced a bug.
# ## Oops! A Bug!
# As nice and simple as `remove_html_markup()` is, it is buggy. Some HTML markup is not properly stripped away. Consider this HTML tag, which would render as an input field in a form:
#
# ```html
# <input type="text" value="<your name>">
# ```
# If we feed this string into `remove_html_markup()`, we would expect an empty string as the result. Instead, this is what we get:
remove_html_markup('<input type="text" value="<your name>">')
# Every time we encounter a bug, this means that our earlier tests have failed. We thus need to introduce another test that documents not only how the bug came to be, but also the result we actually expected.
# The assertion we write now fails with an error message. (The `ExpectError` magic ensures we see the error message, but the rest of the notebook is still executed.)
from ExpectError import ExpectError
with ExpectError():
assert remove_html_markup('<input type="text" value="<your name>">') == ""
# With this, we now have our task: _Fix the failure as above._
# ## Visualizing Code
#
# To properly understand what is going on here, it helps drawing a diagram on how `remove_html_markup()` works. Technically, `remove_html_markup()` implements a _state machine_ with two states `tag` and `¬ tag`. We change between these states depending on the characters we process. This is visualized in the following diagram:
# + ipub={"ignore": true}
from graphviz import Digraph, nohtml
# + ipub={"ignore": true}
from IPython.display import display
# + ipub={"ignore": true}
# ignore
PASS = "✔"
FAIL = "✘"
PASS_COLOR = 'darkgreen' # '#006400' # darkgreen
FAIL_COLOR = 'red4' # '#8B0000' # darkred
STEP_COLOR = 'peachpuff'
FONT_NAME = 'Raleway'
# + ipub={"ignore": true}
# ignore
def graph(comment: str ="default") -> Digraph:
return Digraph(name='', comment=comment, graph_attr={'rankdir': 'LR'},
node_attr={'style': 'filled',
'fillcolor': STEP_COLOR,
'fontname': FONT_NAME},
edge_attr={'fontname': FONT_NAME})
# + ipub={"ignore": true}
# ignore
state_machine = graph()
state_machine.node('Start', )
state_machine.edge('Start', '¬ tag')
state_machine.edge('¬ tag', '¬ tag', label=" ¬ '<'\nadd character")
state_machine.edge('¬ tag', 'tag', label="'<'")
state_machine.edge('tag', '¬ tag', label="'>'")
state_machine.edge('tag', 'tag', label="¬ '>'")
# -
# ignore
display(state_machine)
# You see that we start in the non-tag state (`¬ tag`). Here, for every character that is not `'<'`, we add the character and stay in the same state. When we read a `'<'`, though, we end in the tag state (`tag`) and stay in that state (skipping characters) until we find a closing `'>'` character.
# ## A First Fix
#
# Let us now look at the above state machine, and process through our input:
#
# ```html
# <input type="text" value="<your name>">
# ```
# So what you can see is: We are interpreting the `'>'` of `"<your name>"` as the closing of the tag. However, this is a quoted string, so the `'>'` should be interpreted as a regular character, not as markup. This is an example of _missing functionality:_ We do not handle quoted characters correctly. We haven't claimed yet to take care of all functionality, so we still need to extend our code.
# So we extend the whole thing. We set up a special "quote" state which processes quoted inputs in tags until the end of the quoted string is reached. This is how the state machine looks like:
# + ipub={"ignore": true}
# ignore
state_machine = graph()
state_machine.node('Start')
state_machine.edge('Start', '¬ quote\n¬ tag')
state_machine.edge('¬ quote\n¬ tag', '¬ quote\n¬ tag',
label="¬ '<'\nadd character")
state_machine.edge('¬ quote\n¬ tag', '¬ quote\ntag', label="'<'")
state_machine.edge('¬ quote\ntag', 'quote\ntag', label="'\"'")
state_machine.edge('¬ quote\ntag', '¬ quote\ntag', label="¬ '\"' ∧ ¬ '>'")
state_machine.edge('quote\ntag', 'quote\ntag', label="¬ '\"'")
state_machine.edge('quote\ntag', '¬ quote\ntag', label="'\"'")
state_machine.edge('¬ quote\ntag', '¬ quote\n¬ tag', label="'>'")
# -
# ignore
display(state_machine)
# This is a bit more complex already. Proceeding from left to right, we first have the state `¬ quote ∧ ¬ tag`, which is our "standard" state for text. If we encounter a `'<'`, we again switch to the "tagged" state `¬ quote ∧ tag`. In this state, however (and only in this state), if we encounter a quotation mark, we switch to the "quotation" state `quote ∧ tag`, in which we remain until we see another quotation mark indicating the end of the string – and then continue in the "tagged" state `¬ quote ∧ tag` until we see the end of the string.
# Things get even more complicated as HTML allows both single and double quotation characters. Here's a revised implementation of `remove_html_markup()` that takes the above states into account:
def remove_html_markup(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
return out
# Now, our previous input works well:
remove_html_markup('<input type="text" value="<your name>">')
# and our earlier tests also pass:
assert remove_html_markup("Here's some <strong>strong argument</strong>.") == \
"Here's some strong argument."
assert remove_html_markup('<input type="text" value="<your name>">') == ""
# However, the above code still has a bug. In two of these inputs, HTML markup is still not properly stripped:
#
# ```html
# <b>foo</b>
# <b>"foo"</b>
# "<b>foo</b>"
# <"b">foo</"b">
# ```
#
# Can you guess which ones these are?
# Again, a simple assertion will reveal the culprits:
with ExpectError():
assert remove_html_markup('<b>foo</b>') == 'foo'
with ExpectError():
assert remove_html_markup('<b>"foo"</b>') == '"foo"'
with ExpectError():
assert remove_html_markup('"<b>foo</b>"') == '"foo"'
with ExpectError():
assert remove_html_markup('<"b">foo</"b">') == 'foo'
# So, unfortunately, we're not done yet – our function still has errors.
# ## The Devil's Guide to Debugging
#
# Let us now discuss a couple of methods that do _not_ work well for debugging. (These "devil's suggestions" are adapted from the 1993 book "Code Complete" from <NAME>Connell.)
# ### Printf Debugging
#
# When I was a student, never got any formal training in debugging, so I had to figure this out for myself. What I learned was how to use _debugging output_; in Python, this would be the `print()` function. For instance, I would go and scatter `print()` calls everywhere:
def remove_html_markup_with_print(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
print("c =", repr(c), "tag =", tag, "quote =", quote)
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
return out
# This way of inspecting executions is commonly called "Printf debugging", after the C `printf()` function. Then, running this would allow me to see what's going on in my code:
remove_html_markup_with_print('<b>"foo"</b>')
# Yes, one sees what is going on – but this is horribly inefficient! Think of a 1,000-character input – you'd have to go through 2,000 lines of logs. It may help you, but it's a total time waster. Plus, you have to enter these statements, remove them again... it's a maintenance nightmare.
# (You may even forget printf's in your code, creating a security problem: Mac OS X versions 10.7 to 10.7.3 would log the password in clear because someone had forgotten to turn off debugging output.)
# ### Debugging into Existence
# I would also try to _debug the program into existence._ Just change things until they work. Let me see: If I remove the conditions "and not quote" from the program, it would actually work again:
def remove_html_markup_without_quotes(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<': # and not quote:
tag = True
elif c == '>': # and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
return out
assert remove_html_markup_without_quotes('<"b">foo</"b">') == 'foo'
# Cool! Unfortunately, the function still fails on the other input:
with ExpectError():
assert remove_html_markup_without_quotes('<b>"foo"</b>') == '"foo"'
# So, maybe we can change things again, such that both work? And maybe the other tests we had earlier won't fail? Let's just continue to change things randomly again and again and again.
# Oh, and of course, I would never back up earlier versions such that I would be able to keep track of what has changed and when.
# ### Use the Most Obvious Fix
# My favorite: Use the most obvious fix. This means that you're fixing the symptom, not the problem. In our case, this would be something like:
def remove_html_markup_fixed(s): # type: ignore
if s == '<b>"foo"</b>':
return '"foo"'
...
# Miracle! Our earlier failing assertion now works! Now we can do the same for the other failing test, too, and we're done.
# (Rumor has it that some programmers use this technique to get their tests to pass...)
# ### Things to do Instead
#
# As with any devil's guide, you get an idea of how to do things by doing the _opposite._ What this means is:
#
# 1. Understand the code
# 2. Fix the problem, not the symptom
# 3. Proceed systematically
#
# which is what we will apply for the rest of this chapter.
# ## From Defect to Failure
#
# To understand how to systematically debug a program, we first have to understand how failures come to be. The typical debugging situation looks like this. We have a program (execution), taking some input and producing some output. The output is in *error* (✘), meaning an unwanted and unintended deviation from what is correct, right, or true.
# The input, in contrast, is assumed to be correct (✔). (Otherwise, we wouldn't search for the bug in our program, but in whatever produced its input.)
# ignore
from typing import List, Optional
# + ipub={"ignore": true}
# ignore
def execution_diagram(show_steps: bool = True, variables: List[str] = [],
steps: int = 3, error_step: int = 666,
until: int = 666, fault_path: List[str] = []) -> Digraph:
dot = graph()
dot.node('input', shape='none', fillcolor='white', label=f"Input {PASS}",
fontcolor=PASS_COLOR)
last_outgoing_states = ['input']
for step in range(1, min(steps + 1, until)):
step_color: Optional[str]
if step == error_step:
step_label = f'Step {step} {FAIL}'
step_color = FAIL_COLOR
else:
step_label = f'Step {step}'
step_color = None
if step >= error_step:
state_label = f'State {step} {FAIL}'
state_color = FAIL_COLOR
else:
state_label = f'State {step} {PASS}'
state_color = PASS_COLOR
state_name = f's{step}'
outgoing_states = []
incoming_states = []
if not variables:
dot.node(name=state_name, shape='box',
label=state_label, color=state_color,
fontcolor=state_color)
else:
var_labels = []
for v in variables:
vpath = f's{step}:{v}'
if vpath in fault_path:
var_label = f'<{v}>{v} ✘'
outgoing_states.append(vpath)
incoming_states.append(vpath)
else:
var_label = f'<{v}>{v}'
var_labels.append(var_label)
record_string = " | ".join(var_labels)
dot.node(name=state_name, shape='record',
label=nohtml(record_string), color=state_color,
fontcolor=state_color)
if not outgoing_states:
outgoing_states = [state_name]
if not incoming_states:
incoming_states = [state_name]
for outgoing_state in last_outgoing_states:
for incoming_state in incoming_states:
if show_steps:
dot.edge(outgoing_state, incoming_state,
label=step_label, fontcolor=step_color)
else:
dot.edge(outgoing_state, incoming_state)
last_outgoing_states = outgoing_states
if until > steps + 1:
# Show output
if error_step > steps:
dot.node('output', shape='none', fillcolor='white',
label=f"Output {PASS}", fontcolor=PASS_COLOR)
else:
dot.node('output', shape='none', fillcolor='white',
label=f"Output {FAIL}", fontcolor=FAIL_COLOR)
for outgoing_state in last_outgoing_states:
label = "Execution" if steps == 0 else None
dot.edge(outgoing_state, 'output', label=label)
display(dot)
# -
# ignore
execution_diagram(show_steps=False, steps=0, error_step=0)
# This situation we see above is what we call a *failure*: An externally visible _error_ in the program behavior, with the error again being an unwanted and unintended deviation from what is correct, right, or true.
# How does this failure come to be? The execution we see above breaks down into several program _states_, one after the other.
# ignore
for until in range(1, 6):
execution_diagram(show_steps=False, until=until, error_step=2)
# Initially, the program state is still correct (✔). However, at some point in the execution, the state gets an _error_, also known as a *fault*. This fault – again an unwanted and unintended deviation from what is correct, right, or true – then propagates along the execution, until it becomes externally visible as a _failure_.
# (In reality, there are many, many more states than just this, but these would not fit in a diagram.)
# How does a fault come to be? Each of these program states is produced by a _step_ in the program code. These steps take a state as input and produce another state as output. Technically speaking, the program inputs and outputs are also parts of the program state, so the input flows into the first step, and the output is the state produced by the last step.
# ignore
for until in range(1, 6):
execution_diagram(show_steps=True, until=until, error_step=2)
# Now, in the diagram above, Step 2 gets a _correct_ state as input and produces a _faulty_ state as output. The produced fault then propagates across more steps to finally become visible as a _failure_.
# The goal of debugging thus is to _search_ for the step in which the state first becomes faulty. The _code_ associated with this step is again an error – an unwanted and unintended deviation from what is correct, right, or true – and is called a _defect_. This is what we have to find – and to fix.
# Sounds easy, right? Unfortunately, things are not that easy, and that has something to do with the program state. Let us assume our state consists of three variables, `v1` to `v3`, and that Step 2 produces a fault in `v2`. This fault then propagates to the output:
# ignore
for until in range(1, 6):
execution_diagram(show_steps=True, variables=['v1', 'v2', 'v3'],
error_step=2,
until=until, fault_path=['s2:v2', 's3:v2'])
# The way these faults propagate is called a *cause-effect chain*:
#
# * The _defect_ in the code _causes_ a fault in the state when executed.
# * This _fault_ in the state then _propagates_ through further execution steps...
# * ... until it becomes visible as a _failure_.
# Since the code was originally written by a human, any defect can be related to some original _mistake_ the programmer made. This gives us a number of terms that all are more precise than the general "error" or the colloquial "bug":
#
# * A _mistake_ is a human act or decision resulting in an error.
# * A _defect_ is an error in the program code. Also called *bug*.
# * A _fault_ is an error in the program state. Also called *infection*.
# * A _failure_ is an externally visible error in the program behavior. Also called *malfunction*.
#
# The cause-effect chain of events is thus
#
# * Mistake → Defect → Fault → ... → Fault → Failure
#
# Note that not every defect also causes a failure, which is despite all testing, there can still be defects in the code looming around until the right conditions are met to trigger them. On the other hand, though, _every failure can be traced back to the defect that causes it_. Our job is to break the cause-effect chain.
# ## From Failure to Defect
#
# To find a defect from a failure, we _trace back_ the faults along their _propagation_ – that is, we find out which faults in the earlier state have caused the later faults. We start from the very end of the execution and then gradually progress backwards in time, examining fault after fault until we find a _transition_ from a correct state to a faulty state – that is, a
# step in which a correct state comes in and a faulty state comes out. At this point, we have found the origin of the failure – and the defect that causes it.
# What sounds like a straight-forward strategy, unfortunately, doesn't always work this way in practice. That is because of the following problems of debugging:
#
# * First, program states are actually _large_, encompassing dozens to thousands of variables, possibly even more. If you have to search all of these manually and check them for faults, you will spend a lot of time for a single state.
#
# * Second, you do not always know _whether a state is correct or not._ While most programs have some form of specification for their inputs and outputs, these do not necessarily exist for intermediate results. If one had a specification that could check each state for correctness (possibly even automatically), debugging would be trivial. Unfortunately, it is not, and that's partly due to the lack of specifications.
#
# * Third, executions typically do not come in a handful of steps, as in the diagrams above; instead, they can easily encompass _thousands to millions of steps._ This means that you will have to examine not just one state, but several, making the problem much worse.
#
# To make your search efficient, you thus have to _focus_ your search – starting with most likely causes and gradually progressing to the less probable causes. This is what we call a _debugging strategy_.
# ## The Scientific Method
#
# Now that we know how failures come to be, let's look into how to systematically find their causes. What we need is a _strategy_ that helps us search for how and when the failure comes to be. For this, we use a process called the *scientific method*.
# When we are debugging a program, we are trying to find the causes of a given effect – very much like natural scientists try to understand why things in nature are as they are and how they come to be. Over thousands of years, scientists have conducted _observations_ and _experiments_ to come to an understanding of how our world works. The process by which experimental scientists operate has been coined "The scientific method". This is how it works:
# 1. Formulate a _question_, as in "Why does this apple fall down?".
# 2. Invent a _hypothesis_ based on knowledge obtained while formulating the question, that may explain the observed behavior.
# 3. Determining the logical consequences of the hypothesis, formulate a _prediction_ that can _support_ or _refute_ the hypothesis. Ideally, the prediction would distinguish the hypothesis from likely alternatives.
# 4. _Test_ the prediction (and thus the hypothesis) in an _experiment_. If the prediction holds, confidence in the hypothesis increases; otherwise, it decreases.
# 5. Repeat Steps 2–4 until there are no discrepancies between hypothesis and predictions and/or observations.
# At this point, your hypothesis may be named a *theory* – that is, a predictive and comprehensive description of some aspect of the natural world. The gravitational theory, for instance, predicts very well how the moon revolves around the earth, and how the earth revolves around the sun. Our debugging problems are of a slightly lesser scale – we'd like a theory of how our failure came to be – but the process is pretty much the same.
# + ipub={"ignore": true}
# ignore
dot = graph()
dot.node('Hypothesis')
dot.node('Observation')
dot.node('Prediction')
dot.node('Experiment')
dot.edge('Hypothesis', 'Observation',
label="<Hypothesis<BR/>is <I>supported:</I><BR/>Refine it>",
dir='back')
dot.edge('Hypothesis', 'Prediction')
dot.node('Problem Report', shape='none', fillcolor='white')
dot.edge('Problem Report', 'Hypothesis')
dot.node('Code', shape='none', fillcolor='white')
dot.edge('Code', 'Hypothesis')
dot.node('Runs', shape='none', fillcolor='white')
dot.edge('Runs', 'Hypothesis')
dot.node('More Runs', shape='none', fillcolor='white')
dot.edge('More Runs', 'Hypothesis')
dot.edge('Prediction', 'Experiment')
dot.edge('Experiment', 'Observation')
dot.edge('Observation', 'Hypothesis',
label="<Hypothesis<BR/>is <I>rejected:</I><BR/>Seek alternative>")
# -
# ignore
display(dot)
# In debugging, we proceed the very same way – indeed, we are treating bugs as if they were natural phenomena. This analogy may sound far-fetched, as programs are anything but natural. Nature, by definition, is not under our control. But bugs are _out of our control just as well._ Hence, the analogy is not that far-fetched – and we can apply the same techniques for debugging.
# ### Finding a Hypothesis
# Let us apply the scientific method to our Python program which removes HTML tags. First of all, let us recall the problem – `remove_html_markup()` works for some inputs, but fails on others.
for i, html in enumerate(['<b>foo</b>',
'<b>"foo"</b>',
'"<b>foo</b>"',
'<"b">foo</"b">']):
result = remove_html_markup(html)
print("%-2d %-15s %s" % (i + 1, html, result))
# Input #1 and #4 work as expected, the others do not. We can write these down in a table, such that we can always look back at our previous results:
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |`<b>foo</b>`|`foo`|`foo`|✔|
# |`<b>"foo"</b>`|`"foo"`|`foo`|✘|
# |`"<b>foo</b>"`|`"foo"`|`<b>foo</b>`|✘|
# |`<"b">foo</"b">`|`foo`|`foo`|✔|
#
quiz("From the difference between success and failure,"
" we can already devise some observations about "
" what is wrong with the output."
" Which of these can we turn into general hypotheses?",
[
"Double quotes (`\"`) are stripped from the tagged input.",
"Tags in double quotes are not stripped.",
"The tag `<>` is always stripped from the input.",
"Four-letter words are stripped."
], '[298 % 33, 1234 % 616]')
# ### Testing a Hypothesis
#
# The hypotheses that remain are:
#
# 1. Double quotes are stripped from the tagged input.
# 2. Tags in double quotes are not stripped.
# These may be two separate issues, but chances are they are tied to each other. Let's focus on 1., because it is simpler. Does it hold for all inputs, even untagged ones? Our hypothesis becomes
#
# 1. Double quotes are stripped from the ~~tagged~~ input.
# Let's devise an experiment to validate this. If we feed the string
# ```html
# "foo"
# ```
# (including the double quotes) into `remove_html_markup()`, we should obtain
# ```html
# "foo"
# ```
# as result – that is, the output should be the unchanged input. However, if our hypothesis 1. is correct, we should obtain
# ```html
# foo
# ```
# as result – that is, "Double quotes are stripped from the input" as predicted by the hypothesis.
# We can very easily test this hypothesis:
remove_html_markup('"foo"')
# Our hypothesis is confirmed! We can add this to our list of observations.
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |`<b>foo</b>`|`foo`|`foo`|✔|
# |`<b>"foo"</b>`|`"foo"`|`foo`|✘|
# |`"<b>foo</b>"`|`"foo"`|`<b>foo</b>`|✘|
# |`<"b">foo</"b">`|`foo`|`foo`|✔|
# |`"foo"`|`"foo"`|`foo`|✘|
#
# You can try out the hypothesis with more inputs – and it remains valid. Any non-markup input that contains double quotes will have these stripped.
# Where does that quote-stripping come from? This is where we need to explore the cause-effect chain. The only place in `remove_html_markup()` where quotes are handled is this line:
#
# ```python
# elif c == '"' or c == "'" and tag:
# quote = not quote
# ```
#
# So, quotes should be removed only if `tag` is set. However, `tag` can be set only if the input contains a markup tag, which is not the case for a simple input like `"foo"`. Hence, what we observe is actually _impossible._ Yet, it happens.
# ### Refining a Hypothesis
#
# Debugging is a game of falsifying assumptions. You assume the code works – it doesn't. You assume the `tag` flag cannot be set – yet it may be. What do we do? Again, we create a hypothesis:
#
# 1. The error is due to `tag` being set.
# How do we know whether tag is being set? Let me introduce one of the most powerful debugging tools ever invented, the `assert` statement. The statement
# ```python
# assert cond
# ```
# evaluates the given condition `cond` and
#
# * if it holds: proceed as usual
# * if `cond` does not hold: throw an exception
#
# An `assert` statement _encodes our assumptions_ and as such, should never fail. If it does, well, then something is wrong.
# Using `assert`, we can check the value of `tag` all through the loop:
def remove_html_markup_with_tag_assert(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
assert not tag # <=== Just added
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
quote = not quote
elif not tag:
out = out + c
return out
# Our expectation is that this assertion would fail. So, do we actually get an exception? Try it out for yourself by uncommenting the following line:
# +
# remove_html_markup_with_tag_assert('"foo"')
# -
quiz("What happens after inserting the above assertion?",
[
"The program raises an exception. (i.e., `tag` is set)",
"The output is as before, i.e., `foo` without quotes."
" (which means that `tag` is not set)"
], 2)
# Here's the solution:
with ExpectError():
result = remove_html_markup_with_tag_assert('"foo"')
result
# ### Refuting a Hypothesis
#
# We did not get an exception, hence we reject our hypothesis:
#
# 1. ~~The error is due to `tag` being set.~~
#
# Again, let's go back to the only place in our code where quotes are handled:
#
# ```python
# elif c == '"' or c == "'" and tag:
# quote = not quote
# ```
#
# Because of the assertion, we already know that `tag` is always False. Hence, this condition should never hold either.
# But maybe there's something wrong with the condition such that it holds? Here's our hypothesis:
#
# 1. The error is due to the quote condition evaluating to true
#
# If the condition evaluates to true, then `quote` should be set. We could now go and assert that `quote` is false; but we only care about the condition. So we insert an assertion that assumes that setting the code setting the `quote` flag is never reached:
def remove_html_markup_with_quote_assert(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif c == '"' or c == "'" and tag:
assert False # <=== Just added
quote = not quote
elif not tag:
out = out + c
return out
# Our expectation this time again is that the assertion fails. So, do we get an exception this time? Try it out for yourself by uncommenting the following line:
# +
# remove_html_markup_with_quote_assert('"foo"')
# -
quiz("What happens after inserting the 'assert' tag?",
[
"The program raises an exception (i.e., the quote condition holds)",
"The output is still foo (i.e., the quote condition does not hold)"
], 29 % 7)
# Here's what happens now that we have the `assert` tag:
with ExpectError():
result = remove_html_markup_with_quote_assert('"foo"')
# From this observation, we can deduce that our hypothesis is _confirmed_:
#
# 1. The error is due to the quote condition evaluating to true (CONFIRMED)
#
# and the _condition is actually faulty._ It evaluates to True although `tag` is always False:
# ```python
# elif c == '"' or c == "'" and tag:
# quote = not quote
# ```
# But this condition holds for single and double quotes. Is there a difference?
# Let us see whether our observations generalize towards general quotes:
#
# 1. ~~Double~~ quotes are stripped from the input.
#
# We can verify these hypotheses with an additional experiment. We go back to our original implementation (without any asserts), and then check it:
remove_html_markup("'foo'")
# Surprise: Our hypothesis is rejected and we can add another observation to our table:
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |`'foo'`|`'foo'`|`'foo'`|✔|
#
# So, the condition
#
# * becomes True when a double quote is seen
# * becomes False (as it should) with single quotes
# At this point, you should have enough material to solve the problem. How do we have to fix the condition? Here are four alternatives:
#
# ```python
# c == "" or c == '' and tag # Choice 1
# c == '"' or c == "'" and not tag # Choice 2
# (c == '"' or c == "'") and tag # Choice 3
# ... # Something else
# ```
quiz("How should the condition read?",
[
'''`c == "" or c == '' and tag` (Choice 1)''',
'''`c == '"' or c == "'" and not tag` (Choice 2)''',
'''`(c == '"' or c == "'") and tag` (Choice 3)''',
"Something else"
],
'399 % 4')
# ## Fixing the Bug
# So, you have spotted the defect: In Python (and most other languages), `and` takes precedence over `or`, which is why the condition is wrong. It should read:
#
# ```python
# (c == '"' or c == "'") and tag
# ```
#
# (Actually, good programmers rarely depend on precedence; it is considered good style to use parentheses lavishly.)
# So, our hypothesis now has become
#
# 1. The error is due to the `quote` condition evaluating to True
#
# Is this our final hypothesis? We can check our earlier examples whether they should now work well:
#
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |`<b>foo</b>`|`foo`|`foo`|✔|
# |`<b>"foo"</b>`|`"foo"`|`foo`|✘|
# |`"<b>foo</b>"`|`"foo"`|`<b>foo</b>`|✘|
# |`<"b">foo</"b">`|`foo`|`foo`|✔|
# |`"foo"`|`'foo'`|`foo`|✘|
# |`'foo'`|`'foo'`|`'foo'`|✔|
#
# In all of these examples, the `quote` flag should now be set outside of tags; hence, everything should work as expected.
# In terms of the scientific process, we now have a *theory* – a hypothesis that
#
# * is consistent with all earlier observations
# * predicts future observations (in our case: correct behavior)
#
# For debugging, our problems are usually too small for a big word like theory, so we use the word *diagnosis* instead. So is our diagnosis sufficient to fix the bug? Let us check.
# ### Checking Diagnoses
#
# In debugging, you should start to fix your code if and only if you have a diagnosis that shows two things:
#
# 1. **Causality.** Your diagnosis should explain why and how the failure came to be. Hence, it induces a _fix_ that, when applied, should make the failure disappear.
# 2. **Incorrectness.** Your diagnosis should explain why and how the code is _incorrect_ (which in turn suggests how to _correct_ the code). Hence, the fix it induces not only applies to the given failure, but also to all related failures.
# Showing both these aspects requirements – _causality_ and _incorrectness_ – are crucial for a debugging diagnosis:
#
# * If you find that you can change some location to make the failure go away, but are not sure why this location is wrong, then your "fix" may apply only to the symptom rather than the source. Your diagnosis explains _causality_, but not _incorrectness_.
# * If you find that there is a defect in some code location, but do not verify whether this defect is related to the failure in question, then your "fix" may not address the failure. Your diagnosis addresses _incorrectness_, but not _causality_.
# When you do have a diagnosis that explains both causality (how the failure came to be), and incorrectness (how to correct the code accordingly), then (and only then!) is it time to actually _fix_ the code accordingly. After applying the fix, the failure should be gone, and no other failure should occur. If the failure persists, this should come as a surprise. Obviously, there is some other aspect that you haven't considered yet, so you have to go back to the drawing board and add another failing test case to the set of observations.
# ### Fixing the Code
# All these things considered, let us go and fix `remove_html_markup()`. We know how the defect _causes_ the failure (by erroneously setting `quote` outside of tags). We know that the line in question is _incorrect_ (as single and double of quotes should be treated similarly). So, our diagnosis shows both causality and incorrectness, and we can go and fix the code accordingly:
def remove_html_markup(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif (c == '"' or c == "'") and tag: # <-- FIX
quote = not quote
elif not tag:
out = out + c
return out
# We verify that the fix was successful by running our earlier tests. Not only should the previously failing tests now pass, the previously passing tests also should not be affected. Fortunately, all tests now pass:
assert remove_html_markup("Here's some <strong>strong argument</strong>.") == \
"Here's some strong argument."
assert remove_html_markup(
'<input type="text" value="<your name>">') == ""
assert remove_html_markup('<b>foo</b>') == 'foo'
assert remove_html_markup('<b>"foo"</b>') == '"foo"'
assert remove_html_markup('"<b>foo</b>"') == '"foo"'
assert remove_html_markup('<"b">foo</"b">') == 'foo'
# So, our hypothesis _was_ a theory, and our diagnosis was correct. Success!
# ### Alternate Paths
#
# A defect may have more than one hypothesis, and each diagnosis can be obtained by many ways. We could also have started with our other hypothesis
#
# 2. Tags in double quotes are not stripped
#
# and by reasoning and experiments, we would have reached the same conclusion that the condition is faulty:
#
# * To strip tags, the `tag` flag must be set (but it is not).
# * To set the `tag` flag, the `quote` variable must not be set (but it is).
# * The `quote` flag is set under the given condition (which thus must be faulty).
#
# This gets us to the same diagnosis as above – and, of course, the same fix.
# ## Homework after the Fix
# After having successfully validated the fix, we still have some homework to make.
# ### Check for further Defect Occurrences
# First, we may want to check that the underlying mistake was not made elsewhere, too.
#
# For an error as with `remove_html_markup()`, it may be wise to check other parts of the code (possibly written by the same programmer) whether Boolean formulas show proper precendence. Consider setting up a static program checker or style checker to catch similar mistakes.
# ### Check your Tests
#
# If the defect was not found through testing, now is a good time to make sure it will be found the next time. If you use automated tests, add a test that catches the bug (as well as similar ones), such that you can prevent regressions.
# ### Add Assertions
#
#
# To be 100% sure, we could add an assertion to `remove_html_markup()` that checks the final result for correctness. Unfortunately, writing such an assertion is just as complex as writing the function itself.
#
# There is one assertion, though, which could be placed in the loop body to catch this kind of errors, and which could remain in the code. Which is it?
quiz("Which assertion would have caught the problem?",
[
"`assert quote and not tag`",
"`assert quote or not tag`",
"`assert tag or not quote`",
"`assert tag and not quote`"
], '3270 - 3267')
# Indeed, the statement
#
# ```python
# assert tag or not quote
# ```
# is correct. This excludes the situation of ¬`tag` ∧ `quote` – that is, the `tag` flag is not set, but the `quote` flag is. If you remember our state machine from above, this is actually a state that should never exist:
# ignore
display(state_machine)
# Here's our function in its "final" state. As software goes, software is never final – and this may also hold for our function, as there is still room for improvement. For this chapter though, we leave it be.
def remove_html_markup(s): # type: ignore
tag = False
quote = False
out = ""
for c in s:
assert tag or not quote
if c == '<' and not quote:
tag = True
elif c == '>' and not quote:
tag = False
elif (c == '"' or c == "'") and tag:
quote = not quote
elif not tag:
out = out + c
return out
# ### Commit the Fix
# It may sound obvious, but your fix is worth nothing if it doesn't go into production. Be sure to commit your change to the code repository, together with your diagnosis. If your fix has to be approved by a third party, a good diagnosis on why and what happened is immensely helpful.
# ### Close the Bug Report
#
# If you [systematically track bugs](Tracking.ipynb), and your bug is properly tracked, now is the time to mark the issue as "resolved". Check for duplicates of the issue and check whether they are resolved, too. And now, you are finally done:
#
# 
#
# Time to relax – and look for the next bug!
# ## Become a Better Debugger
#
# We have now systematically fixed a bug. In this book, we will explore a number of techniques to make debugging easier – coming up with automated diagnoses, explanations, even automatic repairs, including for our example above. But there are also number of things _you_ can do to become a better debugger.
#
#
# ### Follow the Process
#
# If you're an experienced programmer, you may have spotted the problem in `remove_html_markup()` immediately, and start fixing the code right away. But this is dangerous and risky.
#
# Why is this so? Well, because you should first
#
# * try to understand the problem, and
# * have a full diagnosis before starting to fix away.
#
# You _can_ skip these steps, and jump right to your interactive debugger the very moment you see a failure, happily stepping through their program. This may even work well for simple problems, including this one. The risk, however, is that this narrows your view to just this one execution, which limits your ability to understand _all_ the circumstances of the problem. Even worse: If you start "fixing" the bug without exactly understanding the problem, you may end up with an incomplete solution – as illustrated in "The Devil's Guide to Debugging", above.
# ### Keep a Log
#
# A second risk of starting debugging too soon is that it lets you easily deviate from a systematic process. Remember how we wrote down every experiment in a table? How we numbered every hypothesis? This is not just for teaching. Writing these things down explicitly allow you to keep track of all your observations and hypotheses over time.
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |`<b>foo</b>`|`foo`|`foo`|✔|
#
# Every time you come up with a new hypothesis, you can immediately check it against your earlier observations, which will help you eliminating unlikely ones from the start. This is a bit like in the classic "Mastermind" board game, in which you have to guess some secret combination of pins, and in which you opponent gives you hints on whether and how your guesses are correct. At any time, you can see your previous guesses (experiments) and the results (observations) you got; any new guess (hypothesis) as to be consistent with the previous observations and experiments.
# 
# Keeping such a log also allows you to interrupt your debugging session at any time. You can be home in time, sleep over the problem, and resume the next morning with a refreshed mind. You can even hand over the log to someone else, stating your findings so far.
#
# The alternative to having a log is to _keep all in memory_. This only works for short amounts of time, as puts a higher and higher cognitive load on your memory as you debug along. After some time, you will forget earlier observations, which leads to mistakes. Worst of all, any interruption will break your concentration and make you forget things, so you can't stop debugging until you're done.
#
# Sure, if you are a real master, you can stay glued to the screen all night. But I'd rather be home in time, thank you.
# ### Rubberducking
#
# A great technique to revisit your observations and to come up with new hypotheses is to _explain the problem to someone else_. In this process, the "someone else" is important, but even more important is that _you are explaining the problem to yourself_! As Kernighan and Pike \cite{Kernighan1999} put it:
#
# > Sometimes it takes no more than a few sentences, followed by an embarrassed "Never mind. I see what's wrong. Sorry to bother you."
#
# The reason why this works is that teaching someone else forces you to take different perspectives, and these help you resolving the inconsistency between what you assume and what you actually observe.
#
# Since that "someone else" can be totally passive, you can even replace her with an inanimate object to talk to – even a rubber duck. This technique is called *rubber duck debugging* or *rubberducking* – the idea is that you explain your problem to a rubber duck first before interrupting one of your co-workers with the problem. Some programmers, when asked for advice, explicitly request that you "explain your problem to the duck first", knowing that this resolves a good fraction of problems.
# 
# ## The Cost of Debugging
#
# \todo{add recent stuff on how much time debugging takes}
#
# And it's not only that debugging takes time – the worst thing is that it is a search process, which can take anything between a few minutes and several hours, sometimes even days and weeks. But even if you never know how much time a bug will take, it's a bit of blessing to use a process which gradually gets you towards its cause.
# ## History of Debugging
#
# Engineers and programmers have long used the term "bug" for faults in their systems – as if it were something that crept into an otherwise flawless program to cause the effects that none could explain. And from a psychological standpoint, it is far easier to blame some "bug" rather than taking responsibility ourselves. In the end, though, we have to face the fact: We made the bugs, and they are ours to fix.
#
# Having said that, there has been one recorded instance where a real bug has crept into a system. That was on September 9, 1947, when a moth got stuck in the relay of a Harvard Mark II machine. This event was logged, and the log book is now on display at the Smithsonian Natural Museum of American History, as "First actual case of bug being found."
# 
# The actual term "bug", however, is much older. What do you think is its origin?
# + ipub={"ignore": true}
# ignore
import hashlib
# + ipub={"ignore": true}
# ignore
bughash = hashlib.md5(b"debug").hexdigest()
# -
quiz('Where has the name "bug" been used to denote disruptive events?',
[
'In the early days of Morse telegraphy, referring to a special key '
'that would send a string of dots',
'Among radio technicians to describe a device that '
'converts electromagnetic field variations into acoustic signals',
"In Shakespeare's " '"Henry VI", referring to a walking spectre',
'In Middle English, where the word "bugge" is the basis for terms '
'like "bugbear" and "bugaboo"'
], [bughash.index(i) for i in "d42f"])
# (Source: \cite{jargon}, \cite{wikipedia:debugging})
# ## Synopsis
# In this chapter, we introduce some basics of how failures come to be as well as a general process for debugging.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# 1. An _error_ is a deviation from what is correct, right, or true. Specifically,
# * A _mistake_ is a human act or decision resulting in an error.
# * A _defect_ is an error in the program code. Also called *bug*.
# * A _fault_ is an error in the program state. Also called *infection*.
# * A _failure_ is an externally visible error in the program behavior. Also called *malfunction*.
# 2. In a failing program execution, a mistake by the programmer results in a defect in the code, which creates a fault in the state, which propagates until it results in a failure. Tracing back fault propagation allows to identify the defect that causes the failure.
# 3. In debugging, the _scientific method_ allows to systematically identify failure causes by gradually refining and refuting hypotheses based on experiments and observations.
# 4. Before fixing the defect, have a complete _diagnosis_ that
# * shows _causality_ (how the defect causes the failure)
# * shows _incorrectness_ (how the defect is wrong)
# 5. You can become a better debugger by
# * Following a systematic process like the scientific method
# * Keeping a log of your observations and hypotheses
# * Making your observations and conclusions explicit by telling them somebody (or something).
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Next Steps
#
# In the next chapters, we will learn how to
#
# * [trace and observe executions](Tracer.ipynb)
# * [build your own interactive debugger](Debugger.ipynb)
# * [locate defects automatically by correlating failures and code coverage](StatisticalDebugger.ipynb)
# * [identify and simplify failure-inducing inputs](DeltaDebugger.ipynb)
#
# Enjoy!
# -
# ## Background
#
# There are several good books on debugging, but these three are especially recommended:
#
# * _Debugging_ by Agans \cite{agans2006-debugging} takes a pragmatic approach to debugging, highlighting systematic approaches that help for all kinds of application-specific problems;
# * _Why Programs Fail_ by Zeller \cite{zeller2009-why-programs-fail} takes a more academic approach, creating theories of how failures come to be and systematic debugging processes;
# * _Effective Debugging_ by Spinellis \cite{spinellis2016-effective-debugging} aims for a middle ground between the two, creating general recipes and recommendations that easily instantiate towards specific problems.
#
# All these books focus on _manual_ debugging and the debugging process, just like this chapter; for _automated_ debugging, simply read on :-)
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Exercises
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ### Exercise 1: Get Acquainted with Notebooks and Python
#
# Your first exercise in this book is to get acquainted with notebooks and Python, such that you can run the code examples in the book – and try out your own. Here are a few tasks to get you started.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# #### Beginner Level: Run Notebooks in Your Browser
#
# The easiest way to get access to the code is to run them in your browser.
#
# 1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `Edit as Notebook`.
# 2. After a short waiting time, this will open a Jupyter Notebook right within your browser, containing the current chapter as a notebook.
# 3. You can again scroll through the material, but you click on any code example to edit and run its code (by entering <kbd>Shift</kbd> + <kbd>Return</kbd>). You can edit the examples as you please.
# 4. Note that code examples typically depend on earlier code, so be sure to run the preceding code first.
# 5. Any changes you make will not be saved (unless you save your notebook to disk).
#
# For help on Jupyter Notebooks, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# #### Advanced Level: Run Python Code on Your Machine
#
# This is useful if you want to make greater changes, but do not want to work with Jupyter.
#
# 1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `Download Code`.
# 2. This will download the Python code of the chapter as a single Python .py file, which you can save to your computer.
# 3. You can then open the file, edit it, and run it in your favorite Python environment to re-run the examples.
# 4. Most importantly, you can [import it](Importing.ipynb) into your own code and reuse functions, classes, and other resources.
#
# For help on Python, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# #### Pro Level: Run Notebooks on Your Machine
#
# This is useful if you want to work with Jupyter on your machine. This will allow you to also run more complex examples, such as those with graphical output.
#
#
# 1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `All Notebooks`.
# 2. This will download all Jupyter Notebooks as a collection of .ipynb files, which you can save to your computer.
# 3. You can then open the notebooks in Jupyter Notebook or Jupyter Lab, edit them, and run them. To navigate across notebooks, open the notebook [`00_Table_of_Contents.ipynb`](00_Table_of_Contents.ipynb).
# 4. You can also download individual notebooks using Select `Resources` $\rightarrow$ `Download Notebook`. Running these, however, will require that you have the other notebooks downloaded already.
#
# For help on Jupyter Notebooks, from the [Web Page](__CHAPTER_HTML__), check out the `Help` menu.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# #### Boss Level: Contribute!
#
# This is useful if you want to contribute to the book with patches or other material. It also gives you access to the very latest version of the book.
#
# 1. From the [Web Page](__CHAPTER_HTML__), check out the menu at the top. Select `Resources` $\rightarrow$ `Project Page`.
# 2. This will get you to the GitHub repository which contains all sources of the book, including the latest notebooks.
# 3. You can then _clone_ this repository to your disk, such that you get the latest and greatest.
# 4. You can report issues and suggest pull requests on the GitHub page.
# 5. Updating the repository with `git pull` will get you updated.
#
# If you want to contribute code or text, check out the [Guide for Authors](Guide_for_Authors.ipynb).
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution2="hidden" solution2_first=true
# ### Exercise 2: More Bugs!
#
# You may have noticed that our `remove_html_markup()` function is still not working perfectly under all circumstances. The error has something to do with different quotes occurring in the input.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Part 1: Find the Problem
#
# What does the problem look like? Set up a test case that demonstrates the problem.
# + cell_style="center"
assert(...)
# + [markdown] solution2="hidden" solution2_first=true
# Set up additional test cases as useful.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** The remaining problem stems from the fact that in `remove_html_markup()`, we do not differentiate between single and double quotes. Hence, if we have a _quote within a quoted text_, the function may get confused. Notably, a string that begins with a double quote may be interpreted as ending when a single quote is seen, and vice versa. Here's an example of such a string:
#
# ```html
# <b title="<Shakespeare's play>">foo</b>
# ```
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# When we remove the HTML markup, the `>` in the string is interpreted as _unquoted_. Hence, it is interpreted as ending the tag, such that the rest of the tag is not removed.
# + slideshow={"slide_type": "skip"} solution2="hidden"
s = '<b title="<Shakespeare' + "'s play>" + '">foo</b>'
s
# + slideshow={"slide_type": "skip"} solution2="hidden"
remove_html_markup(s)
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
with ExpectError():
assert(remove_html_markup(s) == "foo")
# + [markdown] solution2="hidden" solution2_first=true
# #### Part 2: Identify Extent and Cause
#
# Using the scientific method, identify the extent and cause of the problem. Write down your hypotheses and log your observations, as in
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |(input)|(expectation)|(output)|(outcome)|
#
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** The first step is obviously
#
# |Input|Expectation|Output|Outcome|
# |-----|-----------|------|-------|
# |<b title="<Shakespeare's play>">foo</b>|foo|"foo|✘|
#
# + [markdown] solution2="hidden" solution2_first=true
# #### Part 3: Fix the Problem
#
# Design a fix for the problem. Show that it satisfies the earlier tests and does not violate any existing test.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution**. Here's an improved implementation that actually tracks the opening and closing quote by storing the quoting character in the `quote` variable. (If `quote` is `''`, we are not in a string.)
# + slideshow={"slide_type": "skip"} solution2="hidden"
def remove_html_markup_with_proper_quotes(s): # type: ignore
tag = False
quote = ''
out = ""
for c in s:
assert tag or quote == ''
if c == '<' and quote == '':
tag = True
elif c == '>' and quote == '':
tag = False
elif (c == '"' or c == "'") and tag and quote == '':
# beginning of string
quote = c
elif c == quote:
# end of string
quote = ''
elif not tag:
out = out + c
return out
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# Python enthusiasts may note that we could also write `not quote` instead of `quote == ''`, leaving most of the original code untouched. We stick to classic Boolean comparisons here.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# The function now satisfies the earlier failing test:
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
assert(remove_html_markup_with_proper_quotes(s) == "foo")
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# as well as all our earlier tests:
# + slideshow={"slide_type": "skip"} solution2="hidden"
assert remove_html_markup_with_proper_quotes(
"Here's some <strong>strong argument</strong>.") == \
"Here's some strong argument."
assert remove_html_markup_with_proper_quotes(
'<input type="text" value="<your name>">') == ""
assert remove_html_markup_with_proper_quotes('<b>foo</b>') == 'foo'
assert remove_html_markup_with_proper_quotes('<b>"foo"</b>') == '"foo"'
assert remove_html_markup_with_proper_quotes('"<b>foo</b>"') == '"foo"'
assert remove_html_markup_with_proper_quotes('<"b">foo</"b">') == 'foo'
|
notebooks/Intro_Debugging.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Files
#
# Python uses file objects to interact with external files on your computer. These file objects can be any sort of file you have on your computer, whether it be an audio file, a text file, emails, Excel documents, etc. Note: You will probably need to install certain libraries or modules to interact with those various file types, but they are easily available. (We will cover downloading modules later on in the course).
#
# Python has a built-in open function that allows us to open and play with basic file types. First we will need a file though. We're going to use some IPython magic to create a text file!
#
# ## IPython Writing a File
# #### This function is specific to jupyter notebooks!
# %%writefile test.txt
Hello, this is a quick test file.
# ## Python Opening a file
#
# We can open a file with the open() function. The open function also takes in arguments (also called parameters). Lets see how this is used:
# Open the text.txt we made earlier
my_file = open('test.txt')
# We can now read the file
my_file.read()
# But what happens if we try to read it again?
my_file.read()
# This happens because you can imagine the reading "cursor" is at the end of the file after having read it. So there is nothing left to read. We can reset the "cursor" like this:
# Seek to the start of file (index 0)
my_file.seek(0)
# Now read again
my_file.read()
# You can read a file line by line using the readlines method. Use caution with large files, since everything will be held in memory. We will learn how to iterate over large files later in the course.
# Readlines returns a list of the lines in the file
my_file.seek(0)
my_file.readlines()
# When you have finished using a file, it is always good practice to close it.
my_file.close()
# ## Writing to a File
#
# By default, the `open()` function will only allow us to read the file. We need to pass the argument `'w'` to write over the file. For example:
# +
# Add a second argument to the function, 'w' which stands for write.
# Passing 'w+' lets us read and write to the file
my_file = open('test.txt','w+')
# -
# ### <strong><font color='red'>Use caution!</font></strong>
# Opening a file with `'w'` or `'w+'` truncates the original, meaning that anything that was in the original file **is deleted**!
# Write to the file
my_file.write('This is a new line')
# Read the file
my_file.seek(0)
my_file.read()
my_file.close() # always do this when you're done with a file
# ## Appending to a File
# Passing the argument `'a'` opens the file and puts the pointer at the end, so anything written is appended. Like `'w+'`, `'a+'` lets us read and write to a file. If the file does not exist, one will be created.
my_file = open('test.txt','a+')
my_file.write('\nThis is text being appended to test.txt')
my_file.write('\nAnd another line here.')
my_file.seek(0)
print(my_file.read())
my_file.close()
# ### Appending with `%%writefile`
# We can do the same thing using IPython cell magic:
# +
# %%writefile -a test.txt
This is text being appended to test.txt
And another line here.
# -
# Add a blank space if you want the first line to begin on its own line, as Jupyter won't recognize escape sequences like `\n`
# ## Iterating through a File
#
# Lets get a quick preview of a for loop by iterating over a text file. First let's make a new text file with some IPython Magic:
# %%writefile test.txt
First Line
Second Line
# Now we can use a little bit of flow to tell the program to for through every line of the file and do something:
for line in open('test.txt'):
print(line)
# Don't worry about fully understanding this yet, for loops are coming up soon. But we'll break down what we did above. We said that for every line in this text file, go ahead and print that line. It's important to note a few things here:
#
# 1. We could have called the "line" object anything (see example below).
# 2. By not calling `.read()` on the file, the whole text file was not stored in memory.
# 3. Notice the indent on the second line for print. This whitespace is required in Python.
# Pertaining to the first point above
for asdf in open('test.txt'):
print(asdf)
# We'll learn a lot more about this later, but up next: Sets and Booleans!
|
Complete-Python-3-Bootcamp-master/00-Python Object and Data Structure Basics/07-Files.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center> <h1><font size=7> Case Study C</font> </h1> </center>
#
# # Predicting Student Performance
#
# In this case study you will aim to predict the *writing* test score of students taking an exam based on other factors.
#
# The features in the data set `/Data/student_performance.csv` are as follows:
#
# * gender
# * parent_education - the highest level of education achieved by the child's parent
# * lunch - whether or not the student receives free/reduced school lunches (True: receives free/reduced)
# * preparation_course - whether the student attended a test preparation course
# * math_score - the attained grade on their most recent math test
#
# Using a 80:20 train/test split your aim is to use a regression discussed in Chapter 3 in order to predict the *writing_score* target producing a `MSE < 32`.
#
# <div class="alert alert-block alert-info">
#
# <ul>
# <li>Attempt to complete the whole problem before looking at the model answer.
# </li>
# <li>Can you improve on the model answer's score?
# </li>
# <li>Is regularisation necessary for this problem?
# </li>
# </ul>
# </div>
#
#
# Write your code here
|
course_content/case_study/Case Study C.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9 (XPython)
# language: python
# name: xpython
# ---
# <center>
# <img src="xeus-python.png" width="50%">
# <h1>Python kernel based on xeus</h1>
# </center>
# # Simple code execution
a = 3
a
# +
b = 89
def sq(x):
return x * x
sq(b)
# -
print
# # Redirected streams
# +
import sys
print("Error !!", file=sys.stderr)
# -
# # Error handling
# +
"Hello"
def dummy_function():
import missing_module
# -
dummy_function()
# # Code completion
# ### press `tab` to see what is available in `sys` module
from sys import
# # Code inspection
# ### using the question mark
# ?print
# ### by pressing `shift+tab`
print(
# # Input support
name = input('Enter your name: ')
'Hello, ' + name
# # Rich representation
class Person:
def __init__(self, name="<NAME>", address="Paris", picture=""):
self.name = name
self.address = address
self.picture = picture
def _repr_mimebundle_(self, include=None, exclude=None):
return {
"text/html": """<img src="{}">
<div><i class='fa-user fa'></i>: {}</div>
<div><i class='fa-map fa'></i>: {}</div>""".format(self.picture, self.name, self.address)
}
james = Person("<NAME>", "Boston")
display(james)
marie = Person("<NAME>", "Poland", "./marie.png")
display(marie)
# # Matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure()
plt.plot(np.sin(np.linspace(0, 20, 100)));
# %matplotlib widget
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure()
plt.plot(np.sin(np.linspace(0, 20, 100)));
# # Widgets support
# ### Basic widgets
from ipywidgets import IntSlider
slider = IntSlider()
slider
slider.value
slider
slider.value = 36
# ### Widget interacts
from ipywidgets import interact
@interact
def foo(x = ['a', 'b'], n=(1, 10)):
print(x * n)
# ### Binary buffers support for widgets
from ipywidgets import Video
video = Video.from_file("Big.Buck.Bunny.mp4")
video
# ### Higher-level widgets libraries support
# +
import matplotlib
matplotlib.use("agg")
try:
from urllib.request import urlretrieve
except ImportError:
from urllib import urlretrieve
import os
import itk
from itkwidgets import view
# Download data
file_name = '005_32months_T2_RegT1_Reg2Atlas_ManualBrainMask_Stripped.nrrd'
if not os.path.exists(file_name):
url = 'https://data.kitware.com/api/v1/file/564a5b078d777f7522dbfaa6/download'
urlretrieve(url, file_name)
image = itk.imread(file_name)
view(image)
# -
# ## IPython.display module
from IPython.display import clear_output, display, update_display
from time import sleep
# ### Update display
# +
class Square:
color = 'PeachPuff'
def _repr_html_(self):
return '''
<div style="background: %s; width: 200px; height: 100px; border-radius: 10px;">
</div>''' % self.color
square = Square()
display(square, display_id='some-square')
# -
square.color = 'OliveDrab'
update_display(square, display_id='some-square')
# ### Clear output
print("hello")
sleep(3)
clear_output() # will flicker when replacing "hello" with "goodbye"
print("goodbye")
print("hello")
sleep(3)
clear_output(wait=True) # prevents flickering
print("goodbye")
# ### Display classes
from IPython.display import HTML
HTML('''
<div style="background: aliceblue; width: 200px; height: 100px; border-radius: 10px;">
</div>''')
from IPython.display import Math
Math(r'F(k) = \int_{-\infty}^{\infty} f(x) e^{2\pi i k} dx')
from IPython.display import Latex
Latex(r"""\begin{eqnarray}
\nabla \times \vec{\mathbf{B}} -\, \frac1c\, \frac{\partial\vec{\mathbf{E}}}{\partial t} & = \frac{4\pi}{c}\vec{\mathbf{j}} \\
\nabla \cdot \vec{\mathbf{E}} & = 4 \pi \rho \\
\nabla \times \vec{\mathbf{E}}\, +\, \frac1c\, \frac{\partial\vec{\mathbf{B}}}{\partial t} & = \vec{\mathbf{0}} \\
\nabla \cdot \vec{\mathbf{B}} & = 0
\end{eqnarray}""")
from IPython.display import SVG
SVG(url='https://jupyter.org/assets/main-logo.svg')
from IPython.display import SVG
SVG(filename='./logo.svg')
# +
from time import sleep
from IPython.display import ProgressBar
for i in ProgressBar(10):
sleep(0.1)
# -
from IPython.display import JSON
JSON(['foo', {'bar': ('baz', None, 1.0, 2)}], metadata={}, expanded=True, root='test')
from IPython.display import GeoJSON
GeoJSON(
data={
"type": "Feature",
"geometry": {
"type": "Point",
"coordinates": [11.8, -45.04]
}
}, url_template="http://s3-eu-west-1.amazonaws.com/whereonmars.cartodb.net/{basemap_id}/{z}/{x}/{y}.png",
layer_options={
"basemap_id": "celestia_mars-shaded-16k_global",
"attribution" : "Celestia/praesepe",
"tms": True,
"minZoom" : 0,
"maxZoom" : 5
}
)
|
notebooks/xeus-python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://drive.google.com/uc?id=1v7YY_rNBU2OMaPnbUGmzaBj3PUeddxrw" alt="ITI MCIT EPITA" style="width: 750px;"/>
#
# ---
# <img src="https://drive.google.com/uc?id=1R0-FYpJQW5YFy6Yv-RZ1rpyBslay0251" alt="Python Logo" style="width: 400px;"/>
#
# ___
#
# By: **<NAME>**, <EMAIL>
#
# # Session 02: Variables & Data Types
# ## Variables
#
# reserved location in the memory that stores a value. which can be easily retrieved later in the program.
#
# *python is not statically typed language* which means you don't need to declare the type or the variable before using them.
#
# every variable is an object (we will come to this point later)
#
# ### variable names
#
# the variable name must follow the following criteria:
# 1. must start with a letter or underscore
# 2. can't start with a number
# 3. only *alpha-numeric and underscore values* accepted (A-Z, a-z, 0-9, _ )
# 4. case sensitive (ITI is different from iti) (EPITA is a different variable than epita)
# 5. it can be one letter (x) or a name with a meaning (my_name)
#
# ### variable assignment
#
# Assignment is done with a single equals sign (=)
#
# ```x = 10``` means variable x is assigned the value of integer 10
#
#
year = 2021
print(year)
DOB = 1999
print(year-DOB)
x = True # valid
_y = True # valid
# 1u = False #not-valid starts with number
# u-i = 29 #not-valid starts with number
# z = X # X (capital letter) is not same as x (small letter)
# you can even change type after the variable has been set.
# or (re-declaring) the variable
year = "last year"
print(year)
# assign values to multiple variables in one line
ali, shaymaa, mohamed = "pass", "pass", "fail"
print(ali)
print(shaymaa)
print(mohamed)
# assign one value to multiple variables
mahmoud = zayed = saly = "pass"
print(mahmoud)
print(zayed)
print(saly)
x = 352
# Return the “identity” of an object.
# This is an integer which is guaranteed to be unique and constant for this object during its lifetime
print(id(x))
#Convert an integer number to a lowercase hexadecimal string prefixed with “0x”.
print(hex(id(x)))
z = 14
y = 14
print (id(x), id(z), id(y))
print (hex(id(x)), hex(id(z)), hex(id(y)))
# ### variable operators
#
# we can use variables in operators mentioned in session 01
x = 39
y = 74
print(x>y)
print(x/y)
print(x+y)
z = x-y
print(z)
print(x-y)
pi = 3.14
radius = 5
circumference = 2*pi*radius
print(circumference)
print(x)
s = 19
s += x
print(s)
# +
double_message = "welcome to the course" # double quotes
single_message = 'welcome to the course' # Single Quote
print(double_message)
print(single_message)
# -
print("It's AI-Pro 1st intake's 1st course !")
# String Concatenation with variables (needs to be of same type)
welcome = "Welcome, "
first_name = "student"
print(welcome+first_name)
print(welcome+"Mohamed.")
# ### Variable Types
#
# A variable is created when you first assign a value to it.
#
#
# variables declared before
print(type(x))
print(type(y))
print(type(pi))
print(type(welcome))
# #### Variable Type casting
#
# you can specify the data type of a variable
print(type(x))
x = str(x)
print(type(x))
x = 10
print(type(x))
print(x)
x = float(4)
print(x)
import sys
print("Float value information: ",sys.float_info)
print("\nInteger value information: ",sys.int_info)
print("\nMaximum size of an integer: ",sys.maxsize)
# +
# value error
#message = int("welcome") ## expected error
# -
# ## Strings
#
# variable assigned string of text (characters)
#
# string is quoted inside single quotation marks, or double quotation marks
#
# strings are arrays (we will discuss this in future session)
#
#
#
#
hello_world = "Hello, World!"
print(hello_world)
lorem_ipsum = """Lorem ipsum dolor sit amet,
consectetur adipiscing elit,
sed do eiusmod tempor incididunt
ut labore et dolore magna aliqua."""
print(lorem_ipsum)
print(len(lorem_ipsum)) #String Length
print("amet" in lorem_ipsum) #check if in string
print("github" not in lorem_ipsum) #check not in string
str02 = "Python is very flexible for your needs"
print(str02[:9])
print(str02[:9][:6])
# we can modify strings using built-in methods
#
# #### Modify strings
#
#
name = "introduction to python"
print(name.upper())
full_name = "<NAME> "
print(full_name.upper()) # returns uppercased string
print(full_name.capitalize()) # Converts first character to Capital Letter
print(full_name.strip()) # Removes Both Leading and Trailing Characters
print(full_name.title()) # Returns a Title Cased String
print(full_name.swapcase()) # Convert uppercase characters to lowercase and lowercase characters to uppercase.
print(full_name.casefold()) #Return a version of the string suitable for caseless comparisons.
full_name = full_name.strip()
print(full_name)
help('str.casefold')
help('str.swapcase')
## how we can make changes in the variable values
first_name = "moHAmed"
last_name = "AhMEd"
full_name = first_name.title() + " " + last_name.title()
print("Welcome, " + full_name + "!")
# #### Escape Character
#
# An escape character is a backslash \ followed by the character you want to insert. usually used when we want to use illegal character for example a single quote inside a string that's single-quoted.
text = "Hi, \"Student\" !"
print(text)
# some of commonly used escape character is
#
# \n for new line
# \t for horizontal tab
#
# ## Numbers in variables
#
# ### Integers
#
# a whole number
# positive or negative
# without decimals
#
#
x = 19
print(type(x))
x = 494937830948393784763
print(type(x))
x = -398
print(type(x))
x = 0b00011000 # binary
print(x)
y = 0o12 # octal
print(y)
z = 0x12 # hexadecimal
print(z)
print(type(z))
print(x+y+z)
# +
#leading zero's in non-zero int is not valid
# x = 07
# -
# no comma allowed
# Use underscore _ as a delimiter instead
x=1_234_567_890
print(x)
# ### floats
#
# positive or negative
# one or more decimals
y = 3.14
print(type(y))
y = 0.0000001
print(type(y))
y = -0.1
print(type(y))
print(3 * 100.e2)
# Floats has the maximum size depends on your system. The float beyond its maximum size referred as "inf", "Inf", "INFINITY", or "infinity". Float ```2e400``` will be considered as infinity for most systems.
f = 2e400
print(f)
# for scientific purposes we can use float with e to indicate the power of 10
sci_y = 1384e4
print(sci_y)
print(type(sci_y))
# number type conversions
x = 49
y = 7.74
x = float(x)
y = int(y)
print(type(x))
print(type(y))
print(x)
print(y)
print(round(87.93, 1))
print(max(74,87))
print(min(74,87,37,736))
#complex numbers
a=6+4j
b=3+2j
c=a*b
#c=(6+4j)*(3+2j)
#c=(18+12j+12j+8*-1)
#c=10+24j
print(c)
# ### Boolean
#
print(bool("text goes here"))
print(bool(1))
print(bool(0))
print(bool(""))
print(bool(True))
print(bool(False))
print(bool(None))
# +
x = True
y = False
z = True
print(x or y)
print(y or z)
print(x and y)
print(x and z)
print(not x)
print(not y)
print(x and not y)
# -
# #### isinstance()
#
# checks if the variable type is true or false
message = "welcome"
print(isinstance(message, int))
# #### concatenating 2 variables of different types
#
# ##### Using str() function
#
my_string = "my phone number is: "
my_decimal = 521452145
print("Welcome" + " " + "Guest")
#print(my_string + my_decimal)
print(my_string + str(my_decimal))
# ##### Using % Operator
my_string = "my phone number is: "
my_decimal = 521452145
print("%s%d" % (my_string, my_decimal))
print("%s%s" % (my_string, my_decimal))
# ##### Using format() function
print("{}{}".format(my_string, my_decimal))
# ##### Using f-strings
print(f'{my_string}{my_decimal}')
# +
name = 'Mohamed'
age = 38
f_example = f'My Name is {name} and my age is {age}'
print(f_example)
# -
# ### input
x = input()
print(int(x)+2)
str = input("Type Your Message Here")
print(str)
print('Welcome, ' + input('Enter Your Name: '))
a, b, c = input().split()
print(b)
a, b, c, d = map(float, input().split())
# +
first = input('Enter first number: ')
first = float(first)
# let's make it in a single line
second = float(input('Enter second number: '))
print(first, '+', second, '=', first + second)
print(first, '-', second, '=', first - second)
print(first, '*', second, '=', first * second)
print(first, '/', second, '=', first / second)
print("\nEnd of program")
# +
name1 = input("Enter the first stduent's name: ")
id1 = input("Enter the first stduent's ID: ")
grade1 = float(input("Enter the first stduent's grade: "))
name2 = input("\nEnter the second stduent's name: ")
id2 = input("Enter the second stduent's ID: ")
grade2 = float(input("Enter the second stduent's grade: "))
print('\n\nInformat for students and their "Math" grades')
msg = name1 + '(ID ' + id1 + ') got grade: ' + str(grade1)
print(msg)
msg = name2 + '(ID ' + id2 + ') got grade: ' + str(grade2)
print(msg)
average = (grade1 + grade2) / 2.0
print('Average math grade is', average)
# +
A = input()
B = input()
C = input()
combo = A + "'" + B + '"' + C
combo = combo * 10
print(combo)
# -
a, b, c, d = map(int, input().split())
print('a < b is = ', a<b)
|
Session 02 - Variables and Data Types.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # How to insert variables into markdown text
# + [markdown] kernel="SoS"
# * **Difficulty level**: easy
# * **Time need to lean**: 10 minutes or less
#
# + [markdown] kernel="SoS"
# ### Markdown cell and markdown kernel
# + [markdown] kernel="SoS"
# You can include headers, lists, figures, tables in your Jupyter notebook using markdown cells. These markdown cells are rendered by Jupyter itself and do not interact with the kernels. Consequently, it is not possible to pass information (e.g. results from analysis) to markdown cells to generate dynamic output. In contrast, RStudio/RMarkdown has long allowed the inclusion of expressions in markdown texts.
#
# To overcome this problem, you can install a markdown kernel with commands
#
# ```
# pip install markdown-kernel
# python -m markdown.kernel install
# ```
# and write markdown code in code cells with a markdown kernel.
# + kernel="markdown"
Hello, this is a **code cell in markdown kernel**, not a markdown cell.
# + [markdown] kernel="SoS"
# The significance of the markdown kernel is that you can pass information from SoS to it through the `%expand` magic. For example, suppose you have defined a function to calculate Fibonacci sequence,
# + kernel="SoS"
def fibo(n):
return n if n <= 1 else (fibo(n-1) + fibo(n-2))
# + [markdown] kernel="SoS"
# You can write use it in Python expressions as follows:
# + kernel="markdown"
# %expand
The Fibonacci sequence has value {fibo(1)} when `n=1` and {fibo(10)}
when `n=10`, which can be calculated recursively by
`fibo(10)=fibo(9) + fib(8)={fibo(9)}+{fibo(8)}`, and so on.
# + [markdown] kernel="SoS"
# ## Further reading
#
# *
|
src/user_guide/markdown_kernel.ipynb
|
# +
# Copyright 2010-2018 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Code sample to demonstrate how to build a NoOverlap constraint."""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from ortools.sat.python import cp_model
def NoOverlapSampleSat():
"""No overlap sample with fixed activities."""
model = cp_model.CpModel()
horizon = 21 # 3 weeks.
# Task 0, duration 2.
start_0 = model.NewIntVar(0, horizon, 'start_0')
duration_0 = 2 # Python cp/sat code accepts integer variables or constants.
end_0 = model.NewIntVar(0, horizon, 'end_0')
task_0 = model.NewIntervalVar(start_0, duration_0, end_0, 'task_0')
# Task 1, duration 4.
start_1 = model.NewIntVar(0, horizon, 'start_1')
duration_1 = 4 # Python cp/sat code accepts integer variables or constants.
end_1 = model.NewIntVar(0, horizon, 'end_1')
task_1 = model.NewIntervalVar(start_1, duration_1, end_1, 'task_1')
# Task 2, duration 3.
start_2 = model.NewIntVar(0, horizon, 'start_2')
duration_2 = 3 # Python cp/sat code accepts integer variables or constants.
end_2 = model.NewIntVar(0, horizon, 'end_2')
task_2 = model.NewIntervalVar(start_2, duration_2, end_2, 'task_2')
# Weekends.
weekend_0 = model.NewIntervalVar(5, 2, 7, 'weekend_0')
weekend_1 = model.NewIntervalVar(12, 2, 14, 'weekend_1')
weekend_2 = model.NewIntervalVar(19, 2, 21, 'weekend_2')
# No Overlap constraint.
model.AddNoOverlap(
[task_0, task_1, task_2, weekend_0, weekend_1, weekend_2])
# Makespan objective.
obj = model.NewIntVar(0, horizon, 'makespan')
model.AddMaxEquality(obj, [end_0, end_1, end_2])
model.Minimize(obj)
# Solve model.
solver = cp_model.CpSolver()
status = solver.Solve(model)
if status == cp_model.OPTIMAL:
# Print out makespan and the start times for all tasks.
print('Optimal Schedule Length: %i' % solver.ObjectiveValue())
print('Task 0 starts at %i' % solver.Value(start_0))
print('Task 1 starts at %i' % solver.Value(start_1))
print('Task 2 starts at %i' % solver.Value(start_2))
else:
print('Solver exited with nonoptimal status: %i' % status)
NoOverlapSampleSat()
|
examples/notebook/sat/no_overlap_sample_sat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import graphgallery
import matplotlib.pyplot as plt
# Set if memory growth should be enabled for ALL `PhysicalDevice`.
graphgallery.set_memory_growth()
# -
tf.__version__
graphgallery.__version__
# # Load the Datasets
# + cora
# + citeseer
# + pubmed
# +
from graphgallery.data import Planetoid
# set `verbose=False` to avoid these printed tables
data = Planetoid('cora', root="~/GraphData/datasets/", verbose=False)
graph = data.graph
idx_train, idx_val, idx_test = data.split()
# -
data.supported_datasets
from graphgallery.nn.models import GCN
model = GCN(graph, device='GPU', attr_transform="normalize_attr", seed=123)
model.build()
# train with validation
his = model.train(idx_train, idx_val, verbose=1, epochs=100)
# train without validation
# his = model.train(idx_train, verbose=1, epochs=100)
loss, accuracy = model.test(idx_test)
print(f'Test loss {loss:.5}, Test accuracy {accuracy:.2%}')
# ## Show model summary
model.summary()
# ## Visualization Training
import matplotlib.pyplot as plt
with plt.style.context(['science', 'no-latex']):
fig, axes = plt.subplots(1, 2, figsize=(15, 5))
axes[0].plot(his.history['acc'], label='Train accuracy')
axes[0].plot(his.history['val_acc'], label='Val accuracy')
axes[0].legend()
axes[0].set_title('Accuracy')
axes[0].set_xlabel('Epochs')
axes[0].set_ylabel('Accuracy')
axes[1].plot(his.history['loss'], label='Training loss')
axes[1].plot(his.history['val_loss'], label='Validation loss')
axes[1].legend()
axes[1].set_title('Loss')
axes[1].set_xlabel('Epochs')
axes[1].set_ylabel('Loss')
plt.autoscale(tight=True)
plt.show()
|
examples/TensorFlow/test_GCN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="WNUSBVDr0GEk" executionInfo={"status": "ok", "timestamp": 1610024717385, "user_tz": -420, "elapsed": 6399, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}}
# !pip install pennylane
from IPython.display import clear_output
clear_output()
# + id="ja8cpOuLz4Wv" executionInfo={"status": "ok", "timestamp": 1610024717388, "user_tz": -420, "elapsed": 6391, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}}
# This cell is added by sphinx-gallery
# It can be customized to whatever you like
# %matplotlib inline
# + [markdown] id="5n-x3KXMz4W2"
#
#
# Data-reuploading classifier
# ===========================
# *Author: <NAME> (<EMAIL>)*
#
# .. meta::
# :property="og:description": Implement a single-qubit universal quantum classifier using PennyLane.
# :property="og:image": https://pennylane.ai/qml/_images/universal_dnn1.png
#
# .. related::
#
# tutorial_variational_classifier Variational quantum classifier
# tutorial_multiclass_classification Multiclass margin classifier
# tutorial_expressivity_fourier_series Quantum models as Fourier series
#
# A single-qubit quantum circuit which can implement arbitrary unitary
# operations can be used as a universal classifier much like a single
# hidden-layered Neural Network. As surprising as it sounds,
# `<NAME>. (2019) <https://arxiv.org/abs/1907.02085>`_
# discuss this with their idea of 'data
# reuploading'. It is possible to load a single qubit with arbitrary
# dimensional data and then use it as a universal classifier.
#
# In this example, we will implement this idea with Pennylane - a
# python based tool for quantum machine learning, automatic
# differentiation, and optimization of hybrid quantum-classical
# computations.
#
# Background
# ----------
#
# We consider a simple classification problem and will train a
# single-qubit variational quantum circuit to achieve this goal. The data
# is generated as a set of random points in a plane $(x_1, x_2)$ and
# labeled as 1 (blue) or 0 (red) depending on whether they lie inside or
# outside a circle. The goal is to train a quantum circuit to predict the
# label (red or blue) given an input point’s coordinate.
#
# .. figure:: ../demonstrations/data_reuploading/universal_circles.png
# :scale: 65%
# :alt: circles
#
#
# Transforming quantum states using unitary operations
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# A single-qubit quantum state is characterized by a two-dimensional state
# vector and can be visualized as a point in the so-called Bloch sphere.
# Instead of just being a 0 (up) or 1 (down), it can exist in a
# superposition with say 30% chance of being in the $|0 \rangle$ and
# 70% chance of being in the $|1 \rangle$ state. This is represented
# by a state vector $|\psi \rangle = 0.3|0 \rangle + 0.7|1 \rangle$ -
# the probability "amplitude" of the quantum state. In general we can take
# a vector $(\alpha, \beta)$ to represent the probabilities of a qubit
# being in a particular state and visualize it on the Bloch sphere as an
# arrow.
#
# .. figure:: ../demonstrations/data_reuploading/universal_bloch.png
# :scale: 65%
# :alt: bloch
#
# Data loading using unitaries
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# In order to load data onto a single qubit, we use a unitary operation
# $U(x_1, x_2, x_3)$ which is just a parameterized
# matrix multiplication representing the rotation of the state vector in
# the Bloch sphere. E.g., to load $(x_1, x_2)$ into the qubit, we
# just start from some initial state vector, $|0 \rangle$,
# apply the unitary operation $U(x_1, x_2, 0)$ and end up at a new
# point on the Bloch sphere. Here we have padded 0 since our data is only
# 2D. Pérez-Salinas et al. (2019) discuss how to load a higher
# dimensional data point ($[x_1, x_2, x_3, x_4, x_5, x_6]$) by
# breaking it down in sets of three parameters
# ($U(x_1, x_2, x_3), U(x_4, x_5, x_6)$).
#
# Model parameters with data re-uploading
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# Once we load the data onto the quantum circuit, we want to have some
# trainable nonlinear model similar to a neural network as well as a way of
# learning the weights of the model from data. This is again done with
# unitaries, $U(\theta_1, \theta_2, \theta_3)$, such that we load the
# data first and then apply the weights to form a single layer
# $L(\vec \theta, \vec x) = U(\vec \theta)U(\vec x)$. In principle,
# this is just application of two matrix multiplications on an input
# vector initialized to some value. In order to increase the number of
# trainable parameters (similar to increasing neurons in a single layer of
# a neural network), we can reapply this layer again and again with new
# sets of weights,
# $L(\vec \theta_1, \vec x) L(\vec \theta_2, , \vec x) ... L(\vec \theta_L, \vec x)$
# for $L$ layers. The quantum circuit would look like the following:
#
# .. figure:: ../demonstrations/data_reuploading/universal_layers.png
# :scale: 75%
# :alt: Layers
#
#
# The cost function and "nonlinear collapse"
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# So far, we have only performed linear operations (matrix
# multiplications) and we know that we need to have some nonlinear
# squashing similar to activation functions in neural networks to really
# make a universal classifier (Cybenko 1989). Here is where things gets a
# bit quantum. After the application of the layers, we will end up at some
# point on the Bloch sphere due to the sequence of unitaries implementing
# rotations of the input. These are still just linear transformations of
# the input state. Now, the output of the model should be a class label
# which can be encoded as fixed vectors (Blue = $[1, 0]$, Red =
# $[0, 1]$) on the Bloch sphere. We want to end up at either of them
# after transforming our input state through alternate applications of
# data layer and weights.
#
# We can use the idea of the “collapse” of our quantum state into
# one or other class. This happens when we measure the quantum state which
# leads to its projection as either the state 0 or 1. We can compute the
# fidelity (or closeness) of the output state to the class label making
# the output state jump to either $| 0 \rangle$ or
# $|1\rangle$. By repeating this process several times, we can
# compute the probability or overlap of our output to both labels and
# assign a class based on the label our output has a higher overlap. This
# is much like having a set of output neurons and selecting the one which
# has the highest value as the label.
#
# We can encode the output label as a particular quantum state that we want
# to end up in and use Pennylane to find the probability of ending up in that
# state after running the circuit. We construct an observable corresponding to
# the output label using the `Hermitian <https://pennylane.readthedocs.io/en/latest/code/ops/qubit.html#pennylane.ops.qubit.Hermitian>`_
# operator. The expectation value of the observable gives the overlap or fidelity.
# We can then define the cost function as the sum of the fidelities for all
# the data points after passing through the circuit and optimize the parameters
# $(\vec \theta)$ to minimize the cost.
#
# \begin{align}\texttt{Cost} = \sum_{\texttt{data points}} (1 - \texttt{fidelity}(\psi_{\texttt{output}}(\vec x, \vec \theta), \psi_{\texttt{label}}))\end{align}
#
# Now, we can use our favorite optimizer to maximize the sum of the
# fidelities over all data points (or batches of datapoints) and find the
# optimal weights for classification. Gradient-based optimizers such as
# Adam (Kingma et. al., 2014) can be used if we have a good model of
# the circuit and how noise might affect it. Or, we can use some
# gradient-free method such as L-BFGS (<NAME>., and <NAME>., 1989)
# to evaluate the gradient and find the optimal weights where we can
# treat the quantum circuit as a black-box and the gradients are computed
# numerically using a fixed number of function evaluations and iterations.
# The L-BFGS method can be used with the PyTorch interface for Pennylane.
#
# Multiple qubits, entanglement and Deep Neural Networks
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# The Universal Approximation Theorem declares that a neural network with
# two or more hidden layers can serve as a universal function approximator.
# Recently, we have witnessed remarkable progress of learning algorithms using
# Deep Neural Networks.
#
# Pérez-Salinas et al. (2019) make a connection to Deep Neural Networks by
# describing that in their approach the
# “layers” $L_i(\vec \theta_i, \vec x )$ are analogous to the size
# of the intermediate hidden layer of a neural network. And the concept of
# deep (multiple layers of the neural network) relates to the number
# of qubits. So, multiple qubits with entanglement between them could
# provide some quantum advantage over classical neural networks. But here,
# we will only implement a single qubit classifier.
#
# .. figure:: ../demonstrations/data_reuploading/universal_dnn.png
# :scale: 35%
# :alt: DNN
#
# "Talk is cheap. Show me the code." - <NAME>
# ---------------------------------------------------
#
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="Pl7gFapEz4W4" executionInfo={"status": "ok", "timestamp": 1610024729216, "user_tz": -420, "elapsed": 8143, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}} outputId="27bccb5b-f877-4ab0-8917-592df0cf97bb"
import pennylane as qml
from pennylane import numpy as np
from pennylane.optimize import AdamOptimizer, GradientDescentOptimizer
qml.enable_tape()
import matplotlib.pyplot as plt
# Set a random seed
np.random.seed(42)
# Make a dataset of points inside and outside of a circle
def circle(samples, center=[0.0, 0.0], radius=np.sqrt(2 / np.pi)):
"""
Generates a dataset of points with 1/0 labels inside a given radius.
Args:
samples (int): number of samples to generate
center (tuple): center of the circle
radius (float: radius of the circle
Returns:
Xvals (array[tuple]): coordinates of points
yvals (array[int]): classification labels
"""
Xvals, yvals = [], []
for i in range(samples):
x = 2 * (np.random.rand(2)) - 1
y = 0
if np.linalg.norm(x - center) < radius:
y = 1
Xvals.append(x)
yvals.append(y)
return np.array(Xvals), np.array(yvals)
def plot_data(x, y, fig=None, ax=None):
"""
Plot data with red/blue values for a binary classification.
Args:
x (array[tuple]): array of data points as tuples
y (array[int]): array of data points as tuples
"""
if fig == None:
fig, ax = plt.subplots(1, 1, figsize=(5, 5))
reds = y == 0
blues = y == 1
ax.scatter(x[reds, 0], x[reds, 1], c="red", s=20, edgecolor="k")
ax.scatter(x[blues, 0], x[blues, 1], c="blue", s=20, edgecolor="k")
ax.set_xlabel("$x_1$")
ax.set_ylabel("$x_2$")
Xdata, ydata = circle(500)
fig, ax = plt.subplots(1, 1, figsize=(4, 4))
plot_data(Xdata, ydata, fig=fig, ax=ax)
plt.show()
# Define output labels as quantum state vectors
def density_matrix(state):
"""Calculates the density matrix representation of a state.
Args:
state (array[complex]): array representing a quantum state vector
Returns:
dm: (array[complex]): array representing the density matrix
"""
return state * np.conj(state).T
label_0 = [[1], [0]]
label_1 = [[0], [1]]
state_labels = [label_0, label_1]
# + colab={"base_uri": "https://localhost:8080/"} id="sNTrJHj80yvM" executionInfo={"status": "ok", "timestamp": 1610024731625, "user_tz": -420, "elapsed": 1966, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}} outputId="53f66bac-6023-41f0-87e7-e7e70a448368"
Xdata.shape, ydata.shape
# + [markdown] id="yfgAeUSgz4W5"
# Simple classifier with data reloading and fidelity loss
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
#
# + id="9YCQERdOz4W5" executionInfo={"status": "ok", "timestamp": 1610024755144, "user_tz": -420, "elapsed": 1551, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}}
dev = qml.device("default.qubit", wires=1)
# Install any pennylane-plugin to run on some particular backend
@qml.qnode(dev)
def qcircuit(params, x=None, y=None):
"""A variational quantum circuit representing the Universal classifier.
Args:
params (array[float]): array of parameters
x (array[float]): single input vector
y (array[float]): single output state density matrix
Returns:
float: fidelity between output state and input
"""
for i in range(len(params[0])):
qml.Rot(*(params[0][i]*x + params[1][i]), wires=0)
#qml.Rot(*params[1][i], wires=0)
return qml.expval(qml.Hermitian(y, wires=[0]))
def cost(params, x, y, state_labels=None):
"""Cost function to be minimized.
Args:
params (array[float]): array of parameters
x (array[float]): 2-d array of input vectors
y (array[float]): 1-d array of targets
state_labels (array[float]): array of state representations for labels
Returns:
float: loss value to be minimized
"""
# Compute prediction for each input in data batch
loss = 0.0
dm_labels = [density_matrix(s) for s in state_labels]
for i in range(len(x)):
f = qcircuit(params, x=x[i], y=dm_labels[y[i]])
loss = loss + (1 - f) ** 2
return loss / len(x)
# + [markdown] id="ZywIobJtz4W6"
# Utility functions for testing and creating batches
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
#
# + id="BES5HGAwz4W7" executionInfo={"status": "ok", "timestamp": 1610024763553, "user_tz": -420, "elapsed": 1408, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}}
def test(params, x, y, state_labels=None):
"""
Tests on a given set of data.
Args:
params (array[float]): array of parameters
x (array[float]): 2-d array of input vectors
y (array[float]): 1-d array of targets
state_labels (array[float]): 1-d array of state representations for labels
Returns:
predicted (array([int]): predicted labels for test data
output_states (array[float]): output quantum states from the circuit
"""
fidelity_values = []
dm_labels = [density_matrix(s) for s in state_labels]
predicted = []
for i in range(len(x)):
fidel_function = lambda y: qcircuit(params, x=x[i], y=y)
fidelities = [fidel_function(dm) for dm in dm_labels]
best_fidel = np.argmax(fidelities)
predicted.append(best_fidel)
fidelity_values.append(fidelities)
return np.array(predicted), np.array(fidelity_values)
def accuracy_score(y_true, y_pred):
"""Accuracy score.
Args:
y_true (array[float]): 1-d array of targets
y_predicted (array[float]): 1-d array of predictions
state_labels (array[float]): 1-d array of state representations for labels
Returns:
score (float): the fraction of correctly classified samples
"""
score = y_true == y_pred
return score.sum() / len(y_true)
def iterate_minibatches(inputs, targets, batch_size):
"""
A generator for batches of the input data
Args:
inputs (array[float]): input data
targets (array[float]): targets
Returns:
inputs (array[float]): one batch of input data of length `batch_size`
targets (array[float]): one batch of targets of length `batch_size`
"""
for start_idx in range(0, inputs.shape[0] - batch_size + 1, batch_size):
idxs = slice(start_idx, start_idx + batch_size)
yield inputs[idxs], targets[idxs]
# + [markdown] id="fPsEEwFEz4W7"
# Train a quantum classifier on the circle dataset
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="LZUxWb1Wz4W8" executionInfo={"status": "ok", "timestamp": 1610025589054, "user_tz": -420, "elapsed": 822294, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggpw7xw-lyk6u6l92QjpI7MlI7qjJuuciCpwrUd=s64", "userId": "03770692095188133952"}} outputId="380271d4-9a4f-4595-a510-45eab6f20beb"
# Generate training and test data
num_training = 200
num_test = 2000
Xdata, y_train = circle(num_training)
X_train = np.hstack((Xdata, np.zeros((Xdata.shape[0], 1))))
Xtest, y_test = circle(num_test)
X_test = np.hstack((Xtest, np.zeros((Xtest.shape[0], 1))))
# Train using Adam optimizer and evaluate the classifier
num_layers = 10
learning_rate = 0.6
epochs = 10
batch_size = 32
opt = AdamOptimizer(learning_rate, beta1=0.9, beta2=0.999)
# initialize random weights
theta = np.random.uniform(size=(num_layers, 3))
w = np.random.uniform(size=(num_layers, 3))
params = [w, theta]
predicted_train, fidel_train = test(params, X_train, y_train, state_labels)
accuracy_train = accuracy_score(y_train, predicted_train)
predicted_test, fidel_test = test(params, X_test, y_test, state_labels)
accuracy_test = accuracy_score(y_test, predicted_test)
# save predictions with random weights for comparison
initial_predictions = predicted_test
loss = cost(params, X_test, y_test, state_labels)
print(
"Epoch: {:2d} | Cost: {:3f} | Train accuracy: {:3f} | Test Accuracy: {:3f}".format(
0, loss, accuracy_train, accuracy_test
)
)
for it in range(epochs):
for Xbatch, ybatch in iterate_minibatches(X_train, y_train, batch_size=batch_size):
params = opt.step(lambda v: cost(v, Xbatch, ybatch, state_labels), params)
predicted_train, fidel_train = test(params, X_train, y_train, state_labels)
accuracy_train = accuracy_score(y_train, predicted_train)
loss = cost(params, X_train, y_train, state_labels)
predicted_test, fidel_test = test(params, X_test, y_test, state_labels)
accuracy_test = accuracy_score(y_test, predicted_test)
res = [it + 1, loss, accuracy_train, accuracy_test]
print(
"Epoch: {:2d} | Loss: {:3f} | Train accuracy: {:3f} | Test accuracy: {:3f}".format(
*res
)
)
# + [markdown] id="tslHNOLMz4W8"
# Results
# ~~~~~~~
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 471} id="IiQtaHyuz4W9" executionInfo={"status": "ok", "timestamp": 1609940233890, "user_tz": -420, "elapsed": 2042, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "03770692095188133952"}} outputId="ea2148fc-8390-40ef-b829-ccb7ec83ceae"
print(
"Cost: {:3f} | Train accuracy {:3f} | Test Accuracy : {:3f}".format(
loss, accuracy_train, accuracy_test
)
)
print("Learned weights")
for i in range(num_layers):
print("Layer {}: {}".format(i, params[i]))
fig, axes = plt.subplots(1, 3, figsize=(10, 3))
plot_data(X_test, initial_predictions, fig, axes[0])
plot_data(X_test, predicted_test, fig, axes[1])
plot_data(X_test, y_test, fig, axes[2])
axes[0].set_title("Predictions with random weights")
axes[1].set_title("Predictions after training")
axes[2].set_title("True test data")
plt.show()
# + [markdown] id="54KalcQKz4W-"
# References
# ----------
# [1] Pérez-Salinas, Adrián, et al. “Data re-uploading for a universal
# quantum classifier.” arXiv preprint arXiv:1907.02085 (2019).
#
# [2] Kingma, <NAME>., and <NAME>. "Adam: A method for stochastic
# optimization." arXiv preprint arXiv:1412.6980 (2014).
#
# [3] Liu, <NAME>., and <NAME>. "On the limited memory BFGS
# method for large scale optimization." Mathematical programming
# 45.1-3 (1989): 503-528.
#
#
|
PennyLane/Data Reuploading Classifier/tutorial_data_reuploading_classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3.6
# ---
# # Principal Components Analysis (PCA)
#
# PCA identifies the axes that correspond to the greatest variation in a dataset. Usually, most of the variation in a dataset can be summarized by a few principal components. Therefore, the structure of a dataset can be represented using only several principal components.
# ## 1. Import required Python libraries
# <div class="alert alert-info">
# - Click in the cell below
# - Execute the cell by doing **one** of the following:
# - Type `Shift-Enter`
# - Choose the Cell -> Run Cells menu option
# - <img align="left" src="https://github.com/genepattern/example-notebooks/blob/master/2017-11-07_CCMI_workshop/jupyter-run.png?raw=true"> <-- Click the Run icon on the navigation bar under the menu.
import numpy as np
import matplotlib.pyplot as plt
import re
import urllib.request
from matplotlib.ticker import FuncFormatter
# ## 2. Sign in to GenePattern
# <div class="alert alert-info">
# - If you haven't yet logged in, enter your credentials into the cell below and click Login:
# + genepattern={"server": "https://genepattern.broadinstitute.org/gp", "type": "auth"}
# Requires GenePattern Notebook: pip install genepattern-notebook
import gp
import genepattern
# Username and password removed for security reasons.
genepattern.GPAuthWidget(genepattern.register_session("https://genepattern.broadinstitute.org/gp", "", ""))
# -
# ## 3. Compute the principal components of the dataset
# <div class="alert alert-info">
# - Click and drag the following breast cancer dataset link to the **input filename** parameter below: [BRCA_HUGO_symbols.preprocessed.gct](https://datasets.genepattern.org/data/ccmi_tutorial/2017-12-15/BRCA_HUGO_symbols.preprocessed.gct)
# - Notice we are clustering by **columns**, which correspond to samples. This means we will be observing which samples cluster with one another.
# - Click **Run**
# + genepattern={"type": "task"}
pca_task = gp.GPTask(genepattern.get_session(0), 'urn:lsid:broad.mit.edu:cancer.software.genepattern.module.analysis:00017')
pca_job_spec = pca_task.make_job_spec()
pca_job_spec.set_parameter("input.filename", "")
pca_job_spec.set_parameter("cluster.by", "3")
pca_job_spec.set_parameter("output.file", "<input.filename_basename>")
genepattern.GPTaskWidget(pca_task)
# -
# When the job completes, you will see a new cell above with the title **#######.PCA**, where the ####### corresponds to the GenePattern job ID of your PCA analysis. You will also see 4 result files:
#
# Filename | Description
# :------------ | :-------------
# `<filename>_s.odf` | the **s matrix (eigenvectors)**
# `<filename>_t.odf` | the **t matrix (transformed original dataset)**
# `<filename>_u.odf` | the **u matrix (eigenvalues)**
# `gp_execution_log.txt` | the execution log - a record of the analysis run
# ## 4. Visualize the PCA results
# To visualize the results of the PCA analysis, we will read the **s matrix** and **u matrix files** into Python array structures, and create graphs based on the arrays. We do not need the **t matrix** for this analysis.
# ### a. Read file results into Python variables
# The GenePattern results are files on the GenePattern server. To read them into Python arrays, we will use the "`Send to code`" functionality in GenePattern Notebook.
# <div class="alert alert-info">
# 1. In the cell above titled **#######.PCA**, you will see a filename that ends in `_s.odf`. To the right of this file, you will see the following icon:  Click this icon.
# 2. You will see a menu of several choices. Select `Send to Code`.
# 3. You will see a new code cell appear below the **#######.PCA** job results cell.
# 4. In this cell, you will see a Python variable name such as `brca_hugo_symbols_preprocessed_s_odf_1597528`
# 5. Select and copy this variable name.
# 6. In the cell below, paste the variable name into the input field for **gp s matrix file**.
# 7. Repeat the above steps for the filename above that ends in `_u.odf`.
# 8. Execute the cell below by clicking **Run**.
# + genepattern={"output_variable": "result_matrices", "param_values": {"gp_file": "all_aml_train_preprocessed_u_odf_1569581", "gp_s_matrix_file": "", "gp_u_matrix_file": "", "matrix_array": "matrix_array"}, "show_code": false, "type": "uibuilder"}
def pca_results_to_arrays(gp_s_matrix_file, gp_u_matrix_file):
s_matrix_array = gp_matrix_odf_to_nparray(gp_s_matrix_file)
u_matrix_array = gp_matrix_odf_to_nparray(gp_u_matrix_file)
return s_matrix_array, u_matrix_array
def gp_matrix_odf_to_nparray(gp_file):
fh = gp_file.open()
# convert bytes->string->nparray
matrix_raw = fh.read()
matrix_bytes = matrix_raw.decode("utf-8")
# Remove header lines
matrix_string = re.sub(".*\n", '', matrix_bytes, count=5, flags=0)
matrix_string = re.sub("\t\n", '\n', matrix_string, count=0, flags=0)
# The final split leaves an extra line, which must be removed
matrix_list = matrix_string.split('\n')
matrix_list.pop(len(matrix_list)-1)
matrix_2dlist = [row.split('\t') for row in matrix_list]
# Populate the new array with contents of the list:
matrix_array = np.empty(shape=(len(matrix_2dlist),len(matrix_2dlist[0])))
for r in range(len(matrix_2dlist)):
for c in range(len(matrix_2dlist[0])):
matrix_array[r][c] = matrix_2dlist[r][c]
return(matrix_array)
genepattern.GPUIBuilder(pca_results_to_arrays,
name="Convert GenePattern ODF Matrix model result files to numpy arrays",
description="Take as input the S and U matrices resulting from a GenePattern PCA job " +
"and convert then to numpy arrays")
# -
# ### b. Read phenotype assignments to each sample
# We will next read the file that contains the phenotype assignments (e.g., tumor, normal, etc.) for the samples in our dataset. These are in the [CLS](http://software.broadinstitute.org/cancer/software/genepattern/file-formats-guide#CLS) file format.
# <div class="alert alert-info">
# - Click and drag the file containing the phenotype descriptions to the **cls file url** parameter below: [BRCA_HUGO_symbols.preprocessed.cls](https://datasets.genepattern.org/data/ccmi_tutorial/2017-12-15/BRCA_HUGO_symbols.preprocessed.cls)
# - Click **Run**
# + genepattern={"output_variable": "class_data", "param_values": {"cls_file_url": ""}, "show_code": false, "type": "uibuilder"}
def read_phenotype_assignments(cls_file_url):
cls_file = urllib.request.urlopen(cls_file_url)
l1 = cls_file.readline()
(num_samples, num_classes, one) = [int(i) for i in l1.split()]
l2 = cls_file.readline()
class_names = l2.split()
class_names.pop(0)
l3 = cls_file.readline()
class_assignments = [int(i) for i in l3.split()]
return (num_samples, num_classes, class_names, class_assignments)
genepattern.GPUIBuilder(read_phenotype_assignments,
name="Read a phenotype assignment file (cls format) from a url and return its data",
description="Take as input the url to a cls file and return the data it contains: " +
"number of samples, number of classes, class names, class assignments")
# -
# ### c. Set up Python variables for plotting
# <div class="alert alert-info">
# - Execute the cell below
# +
# Extract the s and u matrices from the results
(s_matrix, u_matrix) = result_matrices
# The principal components are the transpose of the u matrix:
pc = u_matrix.transpose()
# Convert eigenvectors from an array to a list
# The eigenvector matrix only has entries on the diagonal. Extract these into a list to facilitate processing:
evectors = [s_matrix[x][x] for x in range(len(s_matrix))]
# Compute percentage contribution of each eigenvector
ev_total = sum(evectors)
ev_percents = evectors/ev_total
# The `class_data` variable contains the class information - parse it out into variables:
(num_samples, num_classes, class_names, class_assignments) = class_data
# Create color map for up to 6 classes:
colormap = ["#ff0000","#0000ff", "#00ff00", "#00ffff", "#ff00ff", "#ffff00"]
colors = [colormap[class_assignments[i]] for i in range(len(class_assignments))]
# -
# ## Display scatter plot of first 2 principal components
# <div class="alert alert-info">
# - Execute the cell below
# +
plt.rcParams["figure.figsize"] = (10,10)
plt.clf()
plt.scatter(pc[0],pc[1],color=colors)
plt.xlabel("Principal Component 1")
plt.ylabel("Principal Component 2")
for i in range(num_classes):
plt.plot(pc[0][i],pc[1][i], marker='o', color=colormap[i], label=str(class_names[i].decode("utf-8")))
plt.legend(title="Classes", loc=0)
plt.show()
# -
# ## Display percentage of variance explained for each principal component
# <div class="alert alert-info">
# - Execute the cell below
# +
plt.clf()
def percents(x, pos):
'The two args are the value and tick position'
return '%1.1f%%' % (x * 100)
formatter = FuncFormatter(percents)
plt.title("Variance Explained Per Principal Component")
x_vals = [i for i in range(num_samples)]
bars = plt.bar(x_vals, ev_percents, 0.8)
plt.xlabel("Principal Component")
plt.ylabel("Variance Explained")
plt.show()
# -
# ## Extra credit
# - Perform the PCA analysis on the following files, which consist of 38 samples comprising two leukemia subtypes, ALL and AML. The rightmost column indicates where you should drag their urls.
#
# Filename | Description | Send to this notebook parameter
# :------------ | :------------- | :-------------
# [all_aml_preprocessed.gct](https://github.com/genepattern/example-notebooks/blob/master/2017-11-07_CCMI_workshop/all_aml_train.preprocessed.gct?raw=true) | Gene expression file | PCA analysis cell **input filename** parameter
# [all_aml_train.cls](https://raw.githubusercontent.com/genepattern/example-notebooks/master/2017-11-07_CCMI_workshop/all_aml_train.cls) | Phenotype assignments file | "Read a phenotype" analysis cell **cls file url** parameter
|
2017-12-15_CCMI_workshop/notebooks/2017-12-15_09_CCMI_Principal+Components+Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def removeDuplicates(self, s: str, k: int) -> str:
stack = []
for i, c in enumerate(s):
stack.append(c)
if len(stack) >= 3:
while len(stack) >= 3 and ''.join(stack[-k:]) == stack[-1] * k:
for _ in range(k):
stack.pop()
return ''.join(stack)
class Solution:
def removeDuplicates(self, s: str, k: int) -> str:
stack = []
n = len(s)
idx = 0
while idx < n:
c = s[idx]
if s[idx: idx+k] == c * k:
idx += k
continue
stack.append(c)
if len(stack) >= k and c == stack[-k]:
while len(stack) >= k and ''.join(stack[-k:]) == stack[-1] * k:
for _ in range(k):
stack.pop()
idx += 1
return ''.join(stack)
class Solution:
def removeDuplicates(self, s: str, k: int) -> str:
stack = []
for c in s:
if stack and stack[-1][0] == c:
if stack[-1][1] == k - 1:
stack.pop()
else:
stack[-1][1] += 1
else:
stack.append([c, 1])
print(stack)
result = ''
for c, n in stack:
result += c * n
return result
solution = Solution()
solution.removeDuplicates(s = "pbbcggttciiippooaais", k = 2)
a = [1, 2, 3, 4]
a[-2]
|
Stack/1223/1209. Remove All Adjacent Duplicates in String II.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automate Publishing of Jupyter Notebooks as Medium Blog Posts with jupyter_to_medium
#
# I am very excited to announce the official release of [jupyter_to_medium][0], a Python package that extends Jupyter Notebooks, allowing you to automate the process of publishing them as Medium blog posts.
#
# ## Motivation
#
# I've published dozens of blog posts on Medium myself with all of them beginning as Jupyter Notebooks. Manually converting them to Medium posts was a fairly lengthy, painstaking process. One particularly painful process was inserting tables, which Medium does not support, into my posts. Nearly all of my posts contain numerous pandas DataFrames ([such as this one][1], which has 40! DataFrames) which are represented as HTML tables within a notebook. I'd take screenshots of each one to insert them into my Medium posts.
#
# jupyter_to_medium automates the process of converting Jupyter Notebooks to Medium blog posts allowing you to save a substantial amount of time.
#
# ## Installation
#
# Install from PyPI with the following command:
#
# ```
# pip install jupyter_to_medium
# ```
#
# After installation, fire up a Jupyter Notebook that you'd like to publish on Medium. Make sure you start a brand new notebook session. Even if you don't have a notebook that is ready to publish, you can still test this extension and publish as a draft so that it remains private.
#
# ## Deploy as Medium Post
#
# Once you open your notebook, head into the **File** menu and notice that you'll have a new option **Deploy as -> Medium Post**.
#
# 
#
# [0]: https://www.dexplo.org/jupyter_to_medium
# [1]: https://medium.com/dunder-data/selecting-subsets-of-data-in-pandas-6fcd0170be9c
# ### If 'Deploy as' option is missing
#
# This new option should automatically show up in your notebook without doing anything. If it is missing, run the follow command to enable it.
#
# ```
# jupyter bundlerextension enable --py jupyter_to_medium._bundler --sys-prefix
# ```
# ### Fill out form
#
# Clicking on **Deploy as -> Medium Post** triggers a new browser tab with a short form that needs to be filled out before posting.
#
# 
# ### Medium Integration Token
#
# Before you can post to Medium, you'll need to request an integration token from them. Do this by emailing them at <a href="mailto:<EMAIL>"><EMAIL></a>. Please read the [entire instructions on how to get your integration token](https://github.com/Medium/medium-api-docs#21-self-issued-access-tokens).
#
# Once your request is granted, navigate to <a href="https://medium.com/me/settings">your Medium settings page.</a> Towards the bottom of the page exists the section on **Integration Tokens**. Enter a description for the token (`jupyter_to_medium` is a good choice) and then create the token.
#
# 
#
# ### Save your integration token
#
# Once you have your integration token, create the folder and file `.jupyter_to_medium/integration_token` in your home directory and save the token there. If you don't save it, you'll need to access it every time you wish to make a new post.
#
# ## Complete form
#
# Once you have your integration token, you can complete the rest of the form. Note that the 'Title' text box is not the actual title of the post, but a title that Medium uses for SEO purposes. Use an `H1` Markdown header in the first cell of your notebook to create the actual title of your post.
#
# ## Publish
#
# Once you've completed the form, click the **Publish** button. At this time, only publishing as a 'draft' is allowed as you probably want to review what your post looks like before publishing it publicly.
# ## Success
#
# If your post was successful, you'll see the following screen with the URL of your post.
#
# 
#
# ## Finalize on Medium
#
# As stated, `jupyter_to_medium` only allows for publishing as a draft. To make your post public, go to your post on Medium and publish there.
#
# 
# ## Features
#
# There are several features that jupyter_to_medium provides to ensure that your notebook appears in the
#
# ### All images in Markdown are found and uploaded to Medium
#
# All images in the Markdown cells, such as the five above are found and uploaded to Markdown. Medium limits the image types to png, jpeg, gif, and tiff.
#
#
# ### Pandas DataFrames are converted to images
#
# Medium does not support tables, such as those produced in output cells by pandas DataFrames. As a workaround, these tables are uploaded as images captured by the Chrome browser's screenshot ability. In the following code cell, the [bar_chart_race](https://www.dexplo.org/bar_chart_race/) package is used to import a pandas DataFrame containing deaths from COVID-19 in several countries. It is embedded in the Medium post as an image.
import bar_chart_race as bcr
df = bcr.load_dataset('covid19')
df = df.iloc[-15:-10, ::3]
df
# ### Styled pandas DataFrames
#
# Styled pandas DataFrames are also embedded in your post as images.
df.style.highlight_max()
# ### Matplotlib Plots
#
# All matplotlib plots, along with any other command that outputs an image will be safely embedded in your post.
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(4, 2.5), dpi=144)
df.loc['2020-04-12'].sort_values().plot(kind='barh', ax=ax, width=.8);
# ### Animated gifs
#
# Even animated gifs are supported by Medium. This one created by the `bar_chart_race` function.
from IPython.display import Image
bcr.bar_chart_race(df, 'docs/images/covid19.gif', figsize=(4, 2.5))
Image(filename='docs/images/covid19.gif')
# ## Citing Jupyter to Medium
#
# If you do use [jupyter_to_medium][0], please mention in the post that you've used it, so that others can find out about it.
#
# [0]: https://www.dexplo.org/jupyter_to_medium
# ## Master Python, Data Science and Machine Learning
#
# Immerse yourself in my comprehensive path for mastering data science and machine learning with Python. [Purchase the All Access Pass][0] to get lifetime access to all current and future courses. Some of the courses it contains:
#
# * [Exercise Python][1] A comprehensive introduction to Python (200+ pages, 100+ exercises)
# * [Master Data Analysis with Python][2] The most comprehensive course available to learn pandas. (800+ pages and 350+ exercises)
# * [Master Machine Learning with Python][3] A deep dive into doing machine learning with scikit-learn constantly updated to showcase the latest and greatest tools.
#
# [0]: https://www.dunderdata.com/all-access-pass
# [1]: https://www.dunderdata.com/exercise-python
# [2]: https://www.dunderdata.com/master-data-analysis-with-python
# [3]: https://www.dunderdata.com/master-machine-learning-with-python
|
Jupyter to Medium Initial Post.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#Importando librerias
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# +
#Configurando las visualizaciones que quiero aplicar
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns.set_style("whitegrid")
# +
#Leemos la base correspondiente
autos = pd.read_csv('dnrpa-robos-feb.csv')
# -
# ### Primeros conocimientos de la base
# +
#Formato de nuestra base
autos.shape
# -
# Nuestra base contiene 25 variables a analizar con 2814 casos
# #### Esto significa que tenemos 2814 robos en lo que va de febrero en Argentina
# +
#Columnas de nuestra base
autos.columns
# +
#Breve informacion de tipos de datos
autos.info()
# -
# Tenemos 7 variables numéricas y 15 variables categóricas
autos.isnull().sum().sum()
# Tenemos 349 valores nulos en total en toda la base de datos, ubicados en las siguientes columnas
autos.isnull().sum()
# +
#Primer vistazo a nuestra base
autos.head()
# -
# ### Limpieza de datos
#
# Se eliminan los datos correspondientes a columnas con valores nulos debido a la poca importancia que obtienen en el estudio general de robos de vehículos en Argentina.
# +
# Se realiza la limpieza total de la base
autos.dropna(inplace = True)
# +
#Chequeamos que no tengamos valores nulos en nuestra base
autos.isnull().sum()
# +
### Corregimos el valor de fecha para obtener resultados en un timeseries
autos['tramite_fecha'] = pd.to_datetime(autos['tramite_fecha'])
# -
# ### Primeros datos
# #### Cantidad de casos por día
robos_por_dia = autos.groupby('tramite_fecha')['titular_genero'].count().to_frame()
robos_por_dia.reset_index(inplace = True)
# +
#Graficando casos diarios
sns.lineplot(data = robos_por_dia, x = "tramite_fecha", y = "titular_genero")
plt.title('Cantidad de robos de autos por día en Argentina durante el mes de Febrero 2022', fontsize = 20)
plt.xlabel('Fecha')
plt.ylabel('Cantidad de robos')
plt.show()
# -
# Se observa un comportamiento extraño el dia 28 de febrero o la perdida en la carga de información por parte de la base de origen prevista por Datos.gob.ar
robos_por_dia.describe()
# . En Argentina se robaron en febrero unos **127** vehículos por día promedio </br>
# . El valor máximo de robos se registró el **día 02 de febrero** con un valor de **156** vehículos por día
# ### Cantidad de robos registrados por provincia
autos.registro_seccional_provincia.value_counts().plot.bar()
plt.title('Robos registrados por provincia - Febrero 2022 ', fontsize = 20)
plt.xlabel('Provincias')
plt.ylabel('Casos registrados')
plt.show()
# Buenos Aires registra la mayor cantidad de casos con 1695 casos. Lo siguen CABA, Córdoba y Mendoza.
autos.registro_seccional_provincia.value_counts(normalize = True)
# **Mendoza** ocupa el cuarto lugar con un 5.6% de los robos registrados en todo el país
# ### Modelos de autos robados en Argentina
autos['automotor_anio_modelo'].value_counts().head(5)
# Los modelos más elegidos por los delincuentes durante febrero 2022 fueron los autos modelo 2013, 2011, 2012, 2010 y 2008. Los modelos actuales parecen no estar en la mira de los delincuentes.
# ### Top 10 - Marcas de automóviles con mayor cantidad de robos
autos.automotor_marca_descripcion.value_counts().head(10).plot.bar()
plt.title('Marcas de automotores con mayor cantidad de robos Febrero 2022', fontsize= 20)
plt.xlabel("Marcas")
plt.ylabel('Cantidad de robos')
plt.show()
# Sin duda algunas , la marca alemana VW lidera el ranking acompañado por FIAT y Renault que cierran el top 3. Lo siguen Chevrolet, Ford y Peugeot.
# ### Los 10 autos más robados en Argentina durante febrero 2022
autos.automotor_modelo_descripcion.value_counts().head(10)
# 1. Gol 1.6 Classic
# 2. Gol Trend 1.6
# 3. Fox 1.6
# 4. Voyage 1.6
# 5. Corsa Classic 4p
# 6. Fiorino Fire
#
# Se observa la hegemonía de la marca alemana Volkswagen en el modelo de autos robados en Argentina durante febrero 2022
# ### Todo sobre las víctimas de robos en Argentina
autos.titular_genero.value_counts(normalize = True)
# El **52%** de los robos registrados en Argentina, las víctimas son de género masculino. El **27%** son mujeres y el **19%** no se identificaron.
sns.boxplot(autos.titular_anio_nacimiento)
plt.title("Distribución de robos por año de nacimiento de la víctima", fontsize = 20);
# Se observa que la concentración de robos se da en víctimas que tienen como año de nacimiento entre 1970 a 1990. Son casos ailsados los registrados a personas de tercera edad por debajo de 1940.
# ## Mendoza
# ### Autos más robados en Mendoza
mendoza = autos[autos['registro_seccional_provincia'] == "Mendoza"]
mendoza.automotor_modelo_descripcion.value_counts().head(5)
# A diferencia de la Nación, en **Mendoza**, el **FORD KA** es el auto más robado durante febrero 2022 seguidos por el Peugeot 206 y VW Gol en sus diferentes modelos.
mendoza.titular_genero.value_counts(normalize= True)
# El género femenino es víctima por encima de los 8 puntos porcentuales si lo comparamos con el porcentaje arrojado en Argentina por lo que este género es seriamente perjudicado en la provincia.
mendoza.titular_domicilio_localidad.value_counts(normalize= True).head(10)
# <NAME>, <NAME> (Guaymallén) y Las Heras ciudad son las localidades con mayor cantidad de robos de automoviles en Mendoza. Lo siguen de cerca otras dos localidades de Guaymallén como Dorrego y Rodeo de la Cruz.
|
robo-automoviles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Data Analysis
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# -
np.random.seed(1)
# load data
df = pd.read_csv('../input_data/heartdisease_data.csv',sep= ',')
df[0:10]
# The data contains 13 features:<br/>
# 0) age: Age (years) --> discrete <br/>
# 1) sex: Sex (1: male, 0: female) --> categorical <br/>
# 2) cp: Chest pain type (1: typical angina, 2: atypical angina, 3: non-anginal pain, 4: asymptomatic) --> categorical <br/>
# 3) trestbps: Resting blood pressure (mm Hg on admission to the hospital) --> continuous <br/>
# 4) chol: Cholesterol measurement (mg/dl) --> continuous <br/>
# 5) fbs: Fasting blood sugar (0: <120 mg/dl, 1: > 120 mg/dl) --> categorical <br/>
# 6) restecg: Resting electrocardiographic measurement (0: normal, 1: having ST-T wave abnormality, 2: showing probable or definite left ventricular hypertrophy by Estes' criteria) --> categorical <br/>
# 7) thalach: Maximum heart rate achieved --> continuous<br/>
# 8) exang: Exercise induced angina (1: yes; 0: no) --> categorical <br/>
# 9) oldpeak: ST depression induced by exercise relative to rest ('ST' relates to positions on the ECG plot) --> continuous<br/>
# 10) slope: The slope of the peak exercise ST segment (1: upsloping, 2: flat, 3: downsloping) --> categorical<br/>
# 11) ca: The number of major vessels (0-3) --> categorical <br/>
# 12) thal: Thalassemia (a type of blood disorder) (3: normal; 6: fixed defect; 7: reversable defect) --> categorical <br/>
#
# and 1 target: Heart disease (0: no, 1: yes) <br/>
# +
# select features and target:
df = np.array(df).astype(float)
# features:
X = df[:,:-1]
l,n = X.shape
print(l,n)
# target:
y = df[:,-1]
# -
# ### Features
"""
plt.figure(figsize=(11,6))
features = s[0,:8]
for j in range(2):
for i in range(4):
ii = j*4 + i
plt.subplot2grid((2,4),(j,i))
bins = np.linspace(min(X[:,ii]), max(X[:,ii]),10, endpoint=False)
plt.hist(X[:,ii],bins,histtype='bar',rwidth=0.8,normed=True)
plt.title('%s'%features[ii])
plt.tight_layout(h_pad=1, w_pad=1.5)
"""
# ### Target
plt.figure(figsize=(4,3))
plt.bar(0,sum(y==0)/float(l),width=0.8,color='blue',label='non disease')
plt.bar(1,sum(y==1)/float(l),width=0.8,color='red',label='disease')
plt.xlabel('0: non disease, 1: disease')
plt.title('target')
# ### 0) Age
ct = pd.crosstab(X[:,0], y)
ct.plot.bar(stacked=True,figsize=(12,3))
plt.xlabel('age')
# ### 1) Sex
ct = pd.crosstab(X[:,1], y)
ct.plot.bar(stacked=True,figsize=(4,3))
plt.xlabel('0: female, 1: male')
# ### 2) Chest pain type
ct = pd.crosstab(X[:,2], y)
ct.plot.bar(stacked=True,figsize=(8,3))
plt.xlabel('Chest pain type')
# ### 3) Resting blood pressure
# +
#ct = pd.crosstab(X[:,3], y)
#ct.plot.histo(stacked=True,figsize=(10,3))
#plt.xlabel('Resting blood pressure')
# -
# ### 5) Fasting blood sugar
pd.crosstab(X[:,5], y).plot.bar(stacked=True,figsize=(4,3))
plt.xlabel('0: <120 mg/dl, 1: > 120 mg/dl')
|
.ipynb_checkpoints/data_exploration_heart_disease-checkpoint.ipynb
|
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # earliness_tardiness_cost_sample_sat
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/sat/earliness_tardiness_cost_sample_sat.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/ortools/sat/samples/earliness_tardiness_cost_sample_sat.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010-2018 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Encodes an convex piecewise linear function."""
from ortools.sat.python import cp_model
class VarArraySolutionPrinter(cp_model.CpSolverSolutionCallback):
"""Print intermediate solutions."""
def __init__(self, variables):
cp_model.CpSolverSolutionCallback.__init__(self)
self.__variables = variables
self.__solution_count = 0
def on_solution_callback(self):
self.__solution_count += 1
for v in self.__variables:
print('%s=%i' % (v, self.Value(v)), end=' ')
print()
def solution_count(self):
return self.__solution_count
def earliness_tardiness_cost_sample_sat():
"""Encode the piecewise linear expression."""
earliness_date = 5 # ed.
earliness_cost = 8
lateness_date = 15 # ld.
lateness_cost = 12
# Model.
model = cp_model.CpModel()
# Declare our primary variable.
x = model.NewIntVar(0, 20, 'x')
# Create the expression variable and implement the piecewise linear function.
#
# \ /
# \______/
# ed ld
#
large_constant = 1000
expr = model.NewIntVar(0, large_constant, 'expr')
# First segment.
s1 = model.NewIntVar(-large_constant, large_constant, 's1')
model.Add(s1 == earliness_cost * (earliness_date - x))
# Second segment.
s2 = 0
# Third segment.
s3 = model.NewIntVar(-large_constant, large_constant, 's3')
model.Add(s3 == lateness_cost * (x - lateness_date))
# Link together expr and x through s1, s2, and s3.
model.AddMaxEquality(expr, [s1, s2, s3])
# Search for x values in increasing order.
model.AddDecisionStrategy([x], cp_model.CHOOSE_FIRST,
cp_model.SELECT_MIN_VALUE)
# Create a solver and solve with a fixed search.
solver = cp_model.CpSolver()
# Force the solver to follow the decision strategy exactly.
solver.parameters.search_branching = cp_model.FIXED_SEARCH
# Search and print out all solutions.
solution_printer = VarArraySolutionPrinter([x, expr])
solver.SearchForAllSolutions(model, solution_printer)
earliness_tardiness_cost_sample_sat()
|
examples/notebook/sat/earliness_tardiness_cost_sample_sat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python-public-policy] *
# language: python
# name: conda-env-python-public-policy-py
# ---
# + [markdown] id="br6IoqhOkfW8"
# # **HOMEWORK 4**
# + [markdown] colab_type="text" id="Rj4Wq1CrlSjj"
# # Coding
#
# Goal: Find complaint types that increased or decreased when COVID-19 hit New York Ciy: mid-March 2020.
# + [markdown] colab_type="text" id="n5pp9g_5v_8K"
# ## Step 0: Setup
#
# For this homework, instead of the data being provided, you will export it directly from the NYC Open Data Portal, as if you were working on your own project.
#
# 1. Download the data.
# 1. Visit the [311 data](https://data.cityofnewyork.us/Social-Services/311-Service-Requests-from-2010-to-Present/erm2-nwe9/data) page.
# 1. From that page, filter the data to `Created Date`s between `01/01/2020 12:00:00 AM` and `03/31/2020 11:59:59 PM`.
# 1. It should say "Showing 311 Service Requests 1-100 out of 475,943" (or close to that number) near the bottom of the screen.
# - It's ok if the total is slightly different.
# 1. Click `Export`.
# 1. Click `CSV`. It will start downloading a file.
# 1. Rename the file `311_covid.csv`.
# 1. Upload the CSV.
# 1. Read the data from `./<filename>.csv`.
#
# If the above is taking a long time due to have a slow network connection or whatever else, load the data from https://storage.googleapis.com/python-public-policy/data/311_covid.csv.zip.
# + [markdown] id="BtvfdWpGlLGP"
# ## Step 1: Load data
#
# Read the data into a DataFrame called `df_2020`.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} executionInfo={"elapsed": 10172, "status": "ok", "timestamp": 1605814693342, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="D7ESnwwTw0RW" outputId="2d2de5ad-4109-438e-cb14-c2aa111c1438"
# your code here
# + [markdown] id="JQo1Pa951V-P"
# ## Step 2: Convert dates
#
# Copy code from [Lecture 4](https://padmgp-4506001-fall.rcnyu.org/user-redirect/notebooks/class_materials/lecture_4.ipynb) to convert the `Created Date` to a `datetime`.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 13026, "status": "ok", "timestamp": 1605814696204, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="N4wk3UZV1gQT" outputId="556a137e-c38c-4f55-cd74-835dba9539e7"
# your code here
# + [markdown] id="_AISfaNylvfX"
# ## Step 3: Date counts
#
# Create a DataFrame called `date_counts` that has the count of complaints per Complaint Type per day, then display it.
# + colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"elapsed": 13232, "status": "ok", "timestamp": 1605814696426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="IICgp0fO4WSp" outputId="93482533-7f07-40ae-c145-06523be67067"
# your code here
# + [markdown] id="cnYXtBEGCxKp"
# ## Step 4: Plotting over time
#
# Create a line chart of the count of complaints over time, one line per `Complaint Type`.
# + colab={"base_uri": "https://localhost:8080/", "height": 542} executionInfo={"elapsed": 19575, "status": "ok", "timestamp": 1605814702785, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="zJ8C7CSQ4afz" outputId="7a888333-919b-4bba-a2e0-6608d2224743"
# your code here
# + [markdown] id="zJlm08A3CtJn"
# ---
#
# This has the information we need, but is a lot to look at. Let's only show complaint types that changed greatly (in March 2020) relative to the same period in the previous year (March 2019).
# + [markdown] id="blbSpASlq2BN"
# ## Step 5: March 2020 counts
#
# Create a DataFrame called `mar_counts` that has the count of each `Complaint Type` in March 2020 in a column called `2020`. Use [`.to_frame()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.to_frame.html) (instead of [`.reset_index()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.reset_index.html)) to use the `Complaint Type` as the index. It should end up looking something like this:
#
# Complaint Type | 2020
# --- | ---
# APPLIANCE | 824
# Abandoned Vehicle | 2500
# Air Quality | 657
# … | …
#
# _Note there is no numeric index._
# + colab={"base_uri": "https://localhost:8080/", "height": 455} executionInfo={"elapsed": 571, "status": "ok", "timestamp": 1605814871210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="_F-Ne4qy5hoE" outputId="490793ee-7523-4ec6-dbc5-186d502a9949"
# your code here
# + [markdown] id="Fr1wKrqVILOF"
# ## Step 6: Get March 2019 data
#
# Follow Steps 0-2 again, this time with 311 requests for all of March 2019. Name the DataFrame `mar_2019`.
#
# Similar to Step 0, if having trouble downloading, you can load from https://storage.googleapis.com/python-public-policy/data/311_mar_2019.csv.zip.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 4152, "status": "ok", "timestamp": 1605814882243, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="E-bLZa0EJi01" outputId="50ec800e-98d8-4e6d-cc80-f70a26574756"
# your code here
# + [markdown] id="g8lAFy31LfLy"
# ## Step 7: March 2019 counts
#
# 1. Get the `Complaint Type` counts for March 2019.
# 1. Add these to the `mar_counts` DataFrame as a column called `2019`.
# - Reminder that adding a Series as a new column to a DataFrame matches rows based on the index.
# + colab={"base_uri": "https://localhost:8080/", "height": 455} executionInfo={"elapsed": 292, "status": "ok", "timestamp": 1605814884908, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="YrS3QunUL7hI" outputId="e7f5f940-08f8-47cb-ed60-7f976ebac484"
# your code here
# + [markdown] id="NmfzxOwEuTWt"
# ## Step 8: Percent change
#
# Use `mar_counts` to calculate the percent change from March 2019 to March 2020 for each `Complaint Type`. Save as the `pct_change` column. Should result in something like this:
#
# Complaint Type | 2020 | 2019 | pct_change
# --- | --- | --- | ---
# APPLIANCE | 824 | 1042 | -0.20
# Abandoned Vehicle | 2500 | 1 | 2499.00
# Air Quality | 657 | 642 | 0.02
# … | … | … | …
# + colab={"base_uri": "https://localhost:8080/", "height": 455} executionInfo={"elapsed": 334, "status": "ok", "timestamp": 1605815115004, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="3KAd1tpI91ia" outputId="96cb58ee-a6d9-4c34-e4f9-b1164c0efdb7"
# your code here
# + [markdown] id="nu5-RoTCFRPr"
# ## Step 9: Filter
#
# Filter to `Complaint Type`s that both:
#
# - Occurred at least 50 times in March 2020
# - Changed (increased _or_ decreased) by more than 90%
#
# and save the DataFrame as `top_changed`. A couple of things that may be helpful:
#
# - [Selecting Subsets of Data in Pandas](https://medium.com/dunder-data/selecting-subsets-of-data-in-pandas-39e811c81a0c#0eb4), starting from "Multiple condition expression"
# - [Getting absolute values](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.abs.html)
# + colab={"base_uri": "https://localhost:8080/", "height": 332} executionInfo={"elapsed": 343, "status": "ok", "timestamp": 1605815607681, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="RdmzmAYGAfA4" outputId="e20be9fd-1929-406d-c2d8-f699aba982f7"
# your code here
# + [markdown] id="ENFrJrS3w7H_"
# ## Step 10: Top changed
#
# Filter the `date_counts` to only the `top_changed` `Complaint Type`s. Save as `top_changed_by_day`.
# + colab={"base_uri": "https://localhost:8080/", "height": 424} executionInfo={"elapsed": 348, "status": "ok", "timestamp": 1605815609438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="xnYwDc-v_kMA" outputId="df11239f-b79f-4328-cf5d-96c08edfd35c"
# your code here
# + [markdown] id="xJflRexSypwA"
# ## Step 11: Plotting changed complaints
#
# Make a similar plot to Step 4, but with only the top complaints (`top_changed_by_day`).
# + colab={"base_uri": "https://localhost:8080/", "height": 542} executionInfo={"elapsed": 1026, "status": "ok", "timestamp": 1605815612657, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj83fkoiPxKpQIphhpuOgId0Uq3vFHX-u5P9mGM=s64", "userId": "02302695983983360874"}, "user_tz": 300} id="8S2ztIF1yqQP" outputId="d1391e3a-69e8-4a55-b46d-058c8dd078a5"
# your code here
# + [markdown] colab_type="text" id="9X1Cjz1i0f_h"
# ## Question 0
#
# ***Did the change of any of the `Complaint Type`s in Step 10/11 surprise you? Why or why not? (Speak at least one specifically.)***
#
# YOUR RESPONSE HERE
# -
# Then, give these a read:
#
# - [NY Daily News article](https://www.nydailynews.com/coronavirus/ny-coronavirus-price-gouging-new-york-city-20200429-z5zs4ygfxbcmrpgzfrnlbxsnea-story.html)
# - [Press release from Department of Consumer and Worker Protection](https://www1.nyc.gov/site/dca/media/pr031720-DCWP-Emergency-Rule-Price-Gouging-Illegal.page)
#
# Overall caveat for this assignment: [**correlation does not imply causation**](https://www.khanacademy.org/math/probability/scatterplots-a1/creating-interpreting-scatterplots/v/correlation-and-causality).
# + [markdown] colab_type="text" id="-hYwWOH01LlC"
# ## Question 1
#
# ***Did you work with anyone else on this assignment?***
#
# YOUR RESPONSE HERE
# -
# ## Bonus: Charting against COVID-19 case counts
#
# Let's take a look at the `Consumer Complaint`s against the COVID-19 case numbers in NYC in the same graph. You'll need to:
#
# 1. Find data that provides the COVID-19 case counts for NYC by day.
# 1. Create a DataFrame with only the `Consumer Complaint` `Complaint Type` counts, by day.
# 1. Chart the two against each other for February through March.
#
# The result should look something like this (without the black box):
#
# 
#
# Some resources that may be helpful:
#
# - [Reading CSV data from GitHub](https://projectosyo.wixsite.com/datadoubleconfirm/single-post/2019/04/15/Reading-csv-data-from-Github---Python)
# - [Two Y Axes in plotly](https://plotly.com/python/multiple-axes/#two-y-axes)
# - Note that the `plotly.graph_objects` syntax is a bit different than the `plotly.express` syntax we've been using. With `go.Scatter()`, you don't provide the DataFrame and the names of the columns; you pass `x` and `y` as lists/Series of the values themselves.
# - [Setting the Range of Axes Manually in plotly](https://plotly.com/python/axes/#setting-the-range-of-axes-manually)
# +
# your code here
# -
# ### Bonus Question 1: What observations do you have?
#
# YOUR RESPONSE HERE
# + [markdown] colab_type="text" id="CbrIfq1J4oEU"
# # Tutorial
#
# In the videos below, don't get hung up on mentions of JavaScript, Node.js, or Twilio — those were technologies used for another course.
#
# 1. Watch:
# 1. [What are APIs?](https://www.youtube.com/watch?v=OVvTv9Hy91Q)
# 1. [APIs, Conceptually](https://drive.google.com/file/d/10VCtYI5Im9MnvDcn4vnUeWbqztF77tyL/view?usp=sharing)
# 1. Read [Understanding And Using REST APIs](https://www.smashingmagazine.com/2018/01/understanding-using-rest-api/)
# 1. Watch:
# 1. [Let's look at some data](https://drive.google.com/file/d/10_2UPxa0ThWus47jKKeefGji5ZZmnr-e/view?usp=sharing)
# 1. [Data formats](https://drive.google.com/file/d/10dR1oMt7V-Hk75mkIpnguq70mukSz6OA/view?usp=sharing)
# 1. [API documentation](https://drive.google.com/file/d/10fOxW42-ODIgHlLgLCP_wklrxB8G4K3A/view?usp=sharing)
# 1. Read [Python’s Requests Library (Guide)](https://realpython.com/python-requests/) through `The Message Body`
|
hw_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Covid19 Analysis
# Our goal is to built a feature that represent a hospital overload.
# +
# libraries
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from haversine import haversine, Unit
from haversine import haversine_vector
# +
# reading dataset
# https://opendatasus.saude.gov.br/dataset/bd-srag-2020
# reading dataset
df_2020 = pd.read_csv('/home/pedro/bkp/code/dataset/INFLUD-21-09-2020.csv',sep=';',encoding = "ISO-8859-1")
# Inputing constraint in the dataset
# Positive case:
df_2020 = df_2020[df_2020['PCR_SARS2']==1]
print(df_2020.shape)
# Hospitalized people:
df_2020 = df_2020[df_2020['PCR_SARS2']==1][df_2020['HOSPITAL']==1][df_2020['NU_IDADE_N']<=110]
print(df_2020.shape)
# Hospitalized people with age small than 110:
df_2020 = df_2020[df_2020['PCR_SARS2']==1][df_2020['HOSPITAL']==1][df_2020['NU_IDADE_N']<=110][df_2020['EVOLUCAO'] != 3][df_2020['EVOLUCAO'] != 9][df_2020['EVOLUCAO'].notnull()]
print(df_2020.shape)
# +
# Calculating the number of hospitalization for each hospital separated by epidemiological week
# CNES = hospital code
# +
# To build a dataframe
df_2020 = df_2020.groupby(['CO_UNI_NOT', 'SEM_NOT']).size().reset_index(name="Times")
# cheking
print(df_2020)
# size
print(df_2020.shape)
# Nan number
print(df_2020['CO_UNI_NOT'].isna().sum())
# +
# Times distribution
plt.hist(df_2020['Times'], 100, density=False, facecolor='g', alpha=0.75)
plt.show()
# +
# max value
print(df_2020['Times'].max())
# hospital code (hospital de campanha)
print(df_2020['CO_UNI_NOT'][df_2020['Times'] == 277])
# +
# reading dataset
# http://plataforma.saude.gov.br/coronavirus/dados-abertos/
# To analysing the hospitalization by SARS from 2019 to get a proxy of hospital absorption capacity.
df_2019 = pd.read_csv('/home/pedro/bkp/code/dataset/INFLUD19-16042020.csv',sep=';',encoding = "ISO-8859-1")
# +
# hospitalization for by hospital
# general data
print(df_2019.shape)
# Selecting hospitalized patients and code from hospital reporting unit.
df_2019 = df_2019['CO_UNI_NOT'][df_2019['HOSPITAL']==1]
print(df_2019.shape)
# +
# Number of hospitalization in 2019 for each hospital reporting unit.
df_2019 = df_2019.value_counts().rename_axis('CO_UNI_NOT').to_frame('counts')
print(df_2019.shape)
# +
# max value
print(df_2019['counts'].max())
# +
# Hospitalization by SARS distribution for 2019
plt.hist(df_2019['counts'], 100, density=False, facecolor='g', alpha=0.75) # .dropna(),
plt.show()
# +
# Merging both dataset
df = pd.merge(df_2020, df_2019, on='CO_UNI_NOT', how="left")
print(df.shape)
# +
# Nan number for each variable
print(df['Times'].isna().sum())
print(df['counts'].isna().sum())
# -
# To check
print(df['Times'].sum())
# +
# To defininf the Overcrowd feature
df['Overload'] = df['Times']/df['counts']
print(df['Overload'].isna().sum())
# +
# To building a catalog with reporting health unit, epidemiologic week, and Overcrowded features
df = df[['CO_UNI_NOT','SEM_NOT','Overload']]
print(df.shape)
# -
# the histogram of the data
plt.hist(df['Overload'], 100, density=False, facecolor='g', alpha=0.75) # .dropna(),
plt.xlabel('Overload')
plt.ylabel('Frequency')
plt.show()
# +
# To transform in csv dataset
df.to_csv('hospital_overcrowded.csv',index=False)
|
over_crowded.ipynb
|