code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # `satn_to_seq`
# Converts values of invasion saturation into sequence numbers
import numpy as np
import porespy as ps
import matplotlib.pyplot as plt
from edt import edt
# The arguments and default values for this function are:
import inspect
inspect.signature(ps.filters.satn_to_seq)
# Generate an image containing invasion sizes using the ``porosimetry`` function:
np.random.seed(0)
im = ps.generators.blobs([200, 200], porosity=0.5)
inv = ps.simulations.drainage(im=im, voxel_size=1, g=0)
# ## `satn`
seq = ps.filters.satn_to_seq(satn=inv.im_satn)
fig, ax = plt.subplots(1, 2, figsize=[12, 6])
ax[0].imshow(inv.im_satn/im, origin='lower', interpolation='none')
ax[0].set_title('Invasion map by saturation')
ax[0].axis(False)
ax[1].imshow(seq/im, origin='lower', interpolation='none')
ax[1].set_title('Invasion map by sequence')
ax[1].axis(False);
# ## `im`
# Passing the boolean image lets the function correctly determine voxels that are solid vs uninvaded, which are both labelled 0.
# +
fig, ax = plt.subplots(1, 2, figsize=[12, 6])
seq = ps.filters.satn_to_seq(satn=inv.im_satn)
ax[0].imshow(seq, origin='lower', interpolation='none')
ax[0].set_title('Invasion map by saturation')
ax[0].axis(False)
seq = ps.filters.satn_to_seq(satn=inv.im_satn, im=im)
ax[1].imshow(seq, origin='lower', interpolation='none')
ax[1].set_title('Invasion map by sequence')
ax[1].axis(False);
|
examples/filters/reference/satn_to_seq.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yohanesnuwara/reservoir-engineering/blob/master/docs/testing_matbal.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="2i6PesJ8vnoY" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="qSp2JQhqu7I3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="3d42bbfa-45dd-4ef5-e9a4-6f6f72bed675"
# !git clone https://github.com/yohanesnuwara/reservoir-engineering
# + id="h165vBcPwIZU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 118} outputId="d897c0f5-83a3-41ed-bed2-977328bccdd6"
# !git clone https://github.com/yohanesnuwara/pyreservoir
# + [markdown] id="Q2uflxWwvUHz" colab_type="text"
# ## Gas Condensate Material Balance
# + id="b5AwM0TZ-38Z" colab_type="code" colab={}
import sys
sys.path.append('/content/pyreservoir/pvt')
import gascorrelation
# + id="bFYi7m8FwQf8" colab_type="code" colab={}
import sys
sys.path.append('/content/pyreservoir/matbal')
from plot import *
# + [markdown] id="wH-_ax7-vwyf" colab_type="text"
# ### Data 1. Ideal data (Rs and Bo are known, measured)
# + id="GKSN4_tAvufu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 462} outputId="2af5e914-24bd-4979-c004-0e2734277e11"
columns = ['p', 'Np', 'Gp', 'Bg', 'Bo', 'Rs', 'Rv']
data = pd.read_csv('/content/reservoir-engineering/Unit 10 Gas-Condensate Reservoirs/data/Table 10.15-PVT and Production Data for Problem 10.5.csv', names=columns)
data
# + [markdown] id="qtuN_X60BfMG" colab_type="text"
# #### Plot 1
# + id="N-28O-O2AqaA" colab_type="code" colab={}
Pdp = 4400 # psia
p = data['p'].values # psia
Np = data['Np'].values # STB
Gp = data['Gp'].values # scf
Bg = data['Bg'].values # RB/scf
Bo = data['Bo'].values # RB/STB
Rs = data['Rs'].values # scf/STB
Rv = data['Rv'].values # STB/scf
# + id="O312aJF5wGx_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="311a65f5-5dda-493f-b690-f0b4abb830e5"
x = condensate()
F, Eg = condensate.plot1(x, Pdp, p, Rs, Rv, Rv[0], Bo, Bg, Bg[0], Np, Gp, Gp[0], Rs[0])
|
docs/testing_matbal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env37
# language: python
# name: env37
# ---
# +
# default_exp models.wavenet
# +
# hide
import sys
sys.path.append("..")
# -
# # Wavenet model
#
# > The WaveNet` architecture for time series forecasting. <https://arxiv.org/pdf/1609.03499.pdf>
#
# Mostly copied from <https://github.com/MSRDL/Deep4Cast>
# hide
from nbdev.showdoc import *
from fastcore.test import *
#export
from fastcore.utils import *
from fastcore.imports import *
from fastai.basics import *
# # Model
# +
# export
class ConcreteDropout(torch.nn.Module):
"""Applies Dropout to the input, even at prediction time and learns dropout probability
from the data.
In convolutional neural networks, we can use dropout to drop entire channels using
the 'channel_wise' argument.
Arguments:
* dropout_regularizer (float): Should be set to 2 / N, where N is the number of training examples.
* init_range (tuple): Initial range for dropout probabilities.
* channel_wise (boolean): apply dropout over all input or across convolutional channels.
"""
def __init__(self,
dropout_regularizer=1e-5,
init_range=(0.1, 0.3),
channel_wise=False):
super(ConcreteDropout, self).__init__()
self.dropout_regularizer = dropout_regularizer
self.init_range = init_range
self.channel_wise = channel_wise
# Initialize dropout probability
init_min = np.log(init_range[0]) - np.log(1. - init_range[0])
init_max = np.log(init_range[1]) - np.log(1. - init_range[1])
self.p_logit = torch.nn.Parameter(
torch.empty(1).uniform_(init_min, init_max))
def forward(self, x):
"""Returns input but with randomly dropped out values."""
# Get the dropout probability
p = torch.sigmoid(self.p_logit)
# Apply Concrete Dropout to input
out = self._concrete_dropout(x, p)
# Regularization term for dropout parameters
dropout_regularizer = p * torch.log(p)
dropout_regularizer += (1. - p) * torch.log(1. - p)
# The size of the dropout regularization depends on the kind of input
if self.channel_wise:
# Dropout only applied to channel dimension
input_dim = x.shape[1]
else:
# Dropout applied to all dimensions
input_dim = np.prod(x.shape[1:])
dropout_regularizer *= self.dropout_regularizer * input_dim
return out, dropout_regularizer.mean()
def _concrete_dropout(self, x, p):
# Empirical parameters for the concrete distribution
eps = 1e-7
temp = 0.1
# Apply Concrete dropout channel wise or across all input
if self.channel_wise:
unif_noise = torch.rand_like(x[:, :, [0]])
else:
unif_noise = torch.rand_like(x)
drop_prob = (torch.log(p + eps)
- torch.log(1 - p + eps)
+ torch.log(unif_noise + eps)
- torch.log(1 - unif_noise + eps))
drop_prob = torch.sigmoid(drop_prob / temp)
random_tensor = 1 - drop_prob
# Need to make sure we have the right shape for the Dropout mask
if self.channel_wise:
random_tensor = random_tensor.repeat([1, 1, x.shape[2]])
# Drop weights
retain_prob = 1 - p
x = torch.mul(x, random_tensor)
x /= retain_prob
return x
# -
# export
class WaveNet(torch.nn.Module):
"""Implements `WaveNet` architecture for time series forecasting. Inherits
from pytorch `Module <https://pytorch.org/docs/stable/nn.html#torch.nn.Module>`_.
Vector forecasts are made via a fully-connected linear layer.
References:
- `WaveNet: A Generative Model for Raw Audio <https://arxiv.org/pdf/1609.03499.pdf>`_
Arguments:
* input_channels (int): Number of covariates in input time series.
* output_channels (int): Number of target time series.
* horizon (int): Number of time steps to forecast.
* hidden_channels (int): Number of channels in convolutional hidden layers.
* skip_channels (int): Number of channels in convolutional layers for skip connections.
* n_layers (int): Number of layers per Wavenet block (determines receptive field size).
* n_blocks (int): Number of Wavenet blocks.
* dilation (int): Dilation factor for temporal convolution.
"""
def __init__(self,
input_channels,
output_channels,
horizon,
hidden_channels=64,
skip_channels=64,
n_layers=7,
n_blocks=1,
dilation=2):
"""Inititalize variables."""
super(WaveNet, self).__init__()
self.output_channels = output_channels
self.horizon = horizon
self.hidden_channels = hidden_channels
self.skip_channels = skip_channels
self.n_layers = n_layers
self.n_blocks = n_blocks
self.dilation = dilation
self.dilations = [dilation**i for i in range(n_layers)] * n_blocks
# Set up first layer for input
self.do_conv_input = ConcreteDropout(channel_wise=True)
self.conv_input = torch.nn.Conv1d(
in_channels=input_channels,
out_channels=hidden_channels,
kernel_size=1
)
# Set up main WaveNet layers
self.do, self.conv, self.skip, self.resi = [], [], [], []
for d in self.dilations:
self.do.append(ConcreteDropout(channel_wise=True))
self.conv.append(torch.nn.Conv1d(in_channels=hidden_channels,
out_channels=hidden_channels,
kernel_size=2,
dilation=d))
self.skip.append(torch.nn.Conv1d(in_channels=hidden_channels,
out_channels=skip_channels,
kernel_size=1))
self.resi.append(torch.nn.Conv1d(in_channels=hidden_channels,
out_channels=hidden_channels,
kernel_size=1))
self.do = torch.nn.ModuleList(self.do)
self.conv = torch.nn.ModuleList(self.conv)
self.skip = torch.nn.ModuleList(self.skip)
self.resi = torch.nn.ModuleList(self.resi)
# Set up nonlinear output layers
self.do_conv_post = ConcreteDropout(channel_wise=True)
self.conv_post = torch.nn.Conv1d(
in_channels=skip_channels,
out_channels=skip_channels,
kernel_size=1
)
self.do_linear_mean = ConcreteDropout()
self.do_linear_std = ConcreteDropout()
self.do_linear_df = ConcreteDropout()
self.linear_mean = torch.nn.Linear(
skip_channels, horizon*output_channels)
self.linear_std = torch.nn.Linear(
skip_channels, horizon*output_channels)
self.linear_df = torch.nn.Linear(
skip_channels, horizon*output_channels)
def forward(self, inputs):
"""Forward function."""
output, reg_e = self.encode(inputs)
output_df, reg_d = self.decode(output)
# Regularization
regularizer = reg_e + reg_d
return output_df # , 'loc': output_mean, 'scale': output_std, 'regularizer': regularizer}
def encode(self, inputs: torch.Tensor):
"""Returns embedding vectors.
Arguments:
* inputs: time series input to make forecasts for
"""
# Input layer
output, res_conv_input = self.do_conv_input(inputs)
output = self.conv_input(output)
# Loop over WaveNet layers and blocks
regs, skip_connections = [], []
for do, conv, skip, resi in zip(self.do, self.conv, self.skip, self.resi):
layer_in = output
output, reg = do(layer_in)
output = conv(output)
output = torch.nn.functional.relu(output)
skip = skip(output)
output = resi(output)
output = output + layer_in[:, :, -output.size(2):]
regs.append(reg)
skip_connections.append(skip)
# Sum up regularizer terms and skip connections
regs = sum(r for r in regs)
output = sum([s[:, :, -output.size(2):] for s in skip_connections])
# Nonlinear output layers
output, res_conv_post = self.do_conv_post(output)
output = torch.nn.functional.relu(output)
output = self.conv_post(output)
output = torch.nn.functional.relu(output)
output = output[:, :, [-1]]
output = output.transpose(1, 2)
# Regularization terms
regularizer = res_conv_input \
+ regs \
+ res_conv_post
return output, regularizer
def decode(self, inputs: torch.Tensor):
"""Returns forecasts based on embedding vectors.
Arguments:
* inputs: embedding vectors to generate forecasts for
"""
# Apply dense layer to match output length
output_mean, res_linear_mean = self.do_linear_mean(inputs)
output_std, res_linear_std = self.do_linear_std(inputs)
output_df, res_linear_df = self.do_linear_df(inputs)
output_mean = self.linear_mean(output_mean)
output_std = self.linear_std(output_std).exp()
output_df = self.linear_df(output_df).exp()
# Reshape the layer output to match targets
# Shape is (batch_size, output_channels, horizon)
batch_size = inputs.shape[0]
output_mean = output_mean.reshape(
(batch_size, self.output_channels, self.horizon)
)
output_std = output_std.reshape(
(batch_size, self.output_channels, self.horizon)
)
output_df = output_df.reshape(
(batch_size, self.output_channels, self.horizon)
)
# Regularization terms
regularizer = res_linear_mean + res_linear_std + res_linear_df
return output_df, regularizer
@property
def n_parameters(self):
"""Returns the number of model parameters."""
par = list(self.parameters())
s = sum([np.prod(list(d.size())) for d in par])
return s
@property
def receptive_field_size(self):
"""Returns the length of the receptive field."""
return self.dilation * max(self.dilations)
# +
class LogTransform(Transform):
r"""Natural logarithm of target covariate + `offset`.
.. math:: y_i = log_e ( x_i + \mbox{offset} )
Args:
* offset (float): amount to add before taking the natural logarithm
* targets (list): list of indices to transform.
Example:
>>> transforms.LogTransform(targets=[0], offset=1.0)
"""
def __init__(self, target_dim=None, offset=0.0):
self.offset = offset
self.target_dim = target_dim
def encodes(self, sample):
X = sample[0]
y = sample[1]
if self.target_dim:
X[:,self.target_dim, :] = torch.log(self.offset + X[:,self.target_dim, :])
y[:,self.target_dim, :] = torch.log(self.offset + y[:,self.target_dim, :])
else:
X = torch.log(self.offset + X)
y = torch.log(self.offset + y)
return X,y
def decodes(self, sample):
X, y = sample[0], sample[1]
if self.target_dim:
X[:, self.target_dim, :] = torch.exp(X[:, self.target_dim, :]) - self.offset
else:
X = torch.exp(X) - self.offset
y = torch.exp(y) - self.offset
return X,y
# -
tmf = LogTransform([0], offset=1.0)
x, y = tensor([[[0.0,1.0,2.1]]]), torch.randn(1,1,3)
_x,_y = tmf((x,y))
test_eq(_x, torch.log(1.0 + tensor([[[0.0,1.0,2.1]]])))
__a,_b = tmf.decode((_x,_y))
# # Learner
# +
#export
from fastai.callback.all import *
@delegates(WaveNet.__init__)
def wavelet_learner(dbunch, output_channels=None, metrics=None, hidden_channels=89, skip_channels =199, **kwargs):
"Build a dnn style learner"
output_channels = ifnone(output_channels,dbunch.train[0][0].shape[0])
model = WaveNet(input_channels=dbunch.train[0][0].shape[0],
output_channels=output_channels,
horizon = dbunch.train_dl.horizon,
hidden_channels=hidden_channels,
skip_channels=skip_channels,
**kwargs
)
dbunch.after_batch.add(LogTransform([0], offset=1.0))
learn = Learner(dbunch, model, loss_func=F.mse_loss, opt_func= Adam, metrics=L(metrics)+L(mae, smape),cbs=[ShowGraphCallback()])
return learn
# -
from fastseq.all import *
from fastseq.core import *
from fastai.basics import *
# +
# hide
path = untar_data(URLs.m4_daily)
data = TSDataLoaders.from_folder(path, horizon = 14, lookback = 128, bs=16, nrows=100, device='cpu')
test_eq(data.train_dl.one_batch()[0].is_cuda,False)
for o in data.valid_dl:
print(o[0].shape)
# -
path = untar_data(URLs.m4_daily)
data = TSDataLoaders.from_folder(path, horizon = 14, lookback = 128, bs=64, nrows=1000, device = 'cpu')
learn = wavelet_learner(data)
from fastai.callback.all import *
learn.lr_find()
learn.fit_one_cycle(1, 5e-4)
from fastai.callback.all import *
learn.lr_find()
learn.fit_one_cycle(3, 1e-3)
learn.validate()
# +
# hide
from nbdev.export import *
notebook2script()
# -
|
nbs/archive/_05_models.wavenet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_pirel_test
# language: python
# name: venv_pirel_test
# ---
# # ```pirel``` Tutorial : introduction and ```pcells```
#
# This is a tutorial for the [```pirel```](https://github.com/giumc/pirel) Python3 module.
#
# ```pirel``` stands for PIezoelectric REsonator Layout, and it is based on the module [```phidl```](https://github.com/amccaugh/phidl)
#
# There are ***four*** packages within ```pirel```: ```pcells``` , ```modifiers``` , ```sweeps``` and ```tools``` . Let's start with ```pcells```.
import pirel.pcells as pc
# ```pcells``` is a collection of classes that are commonly required in piezoelectric resonator design.
# all the cells defined here are derived from ```pirel.tools.LayoutPart``` and they share :
#
# * ```name``` attribute
# * ```set_params``` method
# * ```get_params``` method
# * ```view``` method
# * ```draw``` method
# * ```get_components``` method
#
# In general, these modules are growing pretty fast in size, so it wouldn't make sense to go through all the pcells/modifiers/sweeps/tools.
#
# Instead, try to ```help(pirel.pcells)```,```help(pirel.tools)``` when you are looking for information!
#
# An example of layout class is the InterDigiTated fingers ```pc.IDT```.
idt=pc.IDT(name='TutorialIDT')
# + [markdown] tags=[]
# You can get the parameters available to ```idt``` by just printing it!
# -
idt
# you can get these parameters in a ```dict``` by using ```get_params```.
#
# you can modify any of the parameters and then import them back in the object.
# + tags=[]
idt_params=idt.get_params()
idt_params["N"]=4
idt.set_params(idt_params)
idt
# -
# At any point, you can visualize the layout cell by calling ```view()```
# + tags=[]
import matplotlib.pyplot as plt
idt.view(blocking=True)
# -
# the output is showing a ```phidl.Device``` . These ```Device``` instances are powered up versions of ```gdspy.Cell``` instances.
#
# Refer to [```phidl```](https://github.com/amccaugh/phidl) if you want to learn how many cool things you can do with them.
#
# you can explicitly get this ```Device``` instance by calling the ```draw()``` method.
#
# At that point, you can play around with the cells by using the powerful tools in ```phidl```.
#
# In this example, we will align and distribute two ```idt``` Device using the ```phidl``` module.
# +
idt.coverage=0.3
cell1=idt.draw()
idt.coverage=0.8 ### yes you can set attributes like this, but you will have to find variable names from typing help(idt)
cell2=idt.draw()
import phidl
import phidl.device_layout as dl
from phidl import quickplot as qp
g=dl.Group([cell1,cell2])
g.distribute(direction='x',spacing=30)
g.align(alignment='y')
cell_tot=phidl.Device()
cell_tot<<cell1
cell_tot<<cell2
qp(cell_tot)
# -
# Feel free to look at ```help(pc)``` to figure out all the classes implemented in this module.
#
# Some classes in ``pc`` are created by subclassing , some other by composition of *unit* classes.
# For example, a Lateral Field Excitation RESonator (```pc.LFERes```) is built starting from some components:
#
# * ```pc.IDT```
# * ```pc.Bus```
# * ```pc.EtchPit```
# * ```pc.Anchor```
#
# For any class in ```pc```, you can find components by querying the ```get_components()``` method:
via=pc.Via(name='TutorialVia')
res=pc.LFERes(name='TutorialLFERes')
via.get_components()
res.get_components()
# Note that ```via``` has no components, ```res``` has four.
#
# All layout parameters of each component is also a layout parameter of the composed class.
#
# For example, this is the list of parameters that define ```LFERes```:
lfe_params=res.get_params()
res.view()
lfe_params
# Classes built from ```components``` can have also parameters of their own :
# the class ```FBERes``` (Floating Bottom Electrode Resonators) has a parameter that sets the margin of the floating bottom electrode:
# +
fbe=pc.FBERes(name="TutorialFBE")
params=fbe.get_params()
params["PlatePosition"]='in, long'
fbe.set_params(params)
fbe.view()
# -
# A useful feature of ```set_params``` is that functions can be passed.
#
# For example, when setting a resonator anchor, it might happen that some dimensions have to be derived from others.
#
# The resonator overall width can be found by querying the ```get_active_area``` method of the ```idt``` component of ```fbe```:
fbe.idt.active_area
# If you want to set a pitch, ```active area``` will be updated automatically.
# If you want to keep the anchor size a third of the active area, you can simply write
# +
params['AnchorSizeX']=lambda x : x.idt.active_area.x/3
params['AnchorMetalizedX'] = lambda x : x.anchor.size.x*0.8
fbe.set_params(params)
fbe.view()
# -
# Note that ```idt.active_area``` is a ```pt.Point``` instance . To checkout what you can do with ```pt.Point```, ```help (pt.Point)```!
#
# Now, scaling ```IDTPitch``` will scale ```AnchorSizeX``` and ```AnchorMetalizedX``` accordingly...
#
params['IDTPitch']=40
fbe.set_params(params)
fbe.view()
# If you are bothered by the ```phidl.Port``` labels, just pass the optional ```joined=True``` to the ```check``` method in ```pirel.tools```
# +
import pirel.tools as pt
fbe_cell=fbe.draw()
pt.check(fbe_cell,joined=True)
# -
# Check out the next tutorial on how to power up the ```LayoutPart``` classes using [```pirel.modifiers```](./Tutorial_modifiers.ipynb)
|
tutorials/Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Long-term Investment in SPY
# https://finance.yahoo.com/quote/SPY?p=SPY
# If you have time, is good to invest in SPY for long-term investment.
# ## SPY Market
# + outputHidden=false inputHidden=false
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
import seaborn as sns
from tabulate import tabulate
import math
from scipy.stats import norm
import warnings
warnings.filterwarnings("ignore")
# yfinance is used to fetch data
import yfinance as yf
yf.pdr_override()
# + outputHidden=false inputHidden=false
# input
symbol = 'SPY'
start = '2007-01-01'
end = '2019-01-01'
# Read data
df = yf.download(symbol,start,end)['Adj Close']
# View Columns
df.head()
# + outputHidden=false inputHidden=false
df.tail()
# + outputHidden=false inputHidden=false
df.min()
# + outputHidden=false inputHidden=false
df.max()
# + outputHidden=false inputHidden=false
from datetime import datetime
from dateutil import relativedelta
d1 = datetime.strptime(start, "%Y-%m-%d")
d2 = datetime.strptime(end, "%Y-%m-%d")
delta = relativedelta.relativedelta(d2,d1)
print('How many years of investing?')
print('%s years' % delta.years)
# -
# ### Starting Cash with 100k to invest in Bonds
# + outputHidden=false inputHidden=false
Cash = 100000
# + outputHidden=false inputHidden=false
print('Number of Shares:')
shares = int(Cash/df.iloc[0])
print('{}: {}'.format(symbol, shares))
# + outputHidden=false inputHidden=false
print('Beginning Value:')
shares = int(Cash/df.iloc[0])
Begin_Value = round(shares * df.iloc[0], 2)
print('{}: ${}'.format(symbol, Begin_Value))
# + outputHidden=false inputHidden=false
print('Current Value:')
shares = int(Cash/df.iloc[0])
Current_Value = round(shares * df.iloc[-1], 2)
print('{}: ${}'.format(symbol, Current_Value))
# + outputHidden=false inputHidden=false
returns = df.pct_change().dropna()
# + outputHidden=false inputHidden=false
returns.head()
# + outputHidden=false inputHidden=false
returns.tail()
# + outputHidden=false inputHidden=false
# Calculate cumulative returns
daily_cum_ret=(1+returns).cumprod()
print(daily_cum_ret.tail())
# + outputHidden=false inputHidden=false
# Print the mean
print("mean : ", returns.mean()*100)
# Print the standard deviation
print("Std. dev: ", returns.std()*100)
# Print the skewness
print("skew: ", returns.skew())
# Print the kurtosis
print("kurt: ", returns.kurtosis())
# + outputHidden=false inputHidden=false
# Calculate total return and annualized return from price data
total_return = (returns[-1] - returns[0]) / returns[0]
print(total_return)
# + outputHidden=false inputHidden=false
# Annualize the total return over 12 year
annualized_return = ((1+total_return)**(1/12))-1
# + outputHidden=false inputHidden=false
# Calculate annualized volatility from the standard deviation
vol_port = returns.std() * np.sqrt(250)
# + outputHidden=false inputHidden=false
# Calculate the Sharpe ratio
rf = 0.001
sharpe_ratio = (annualized_return - rf) / vol_port
print(sharpe_ratio)
# + outputHidden=false inputHidden=false
# Create a downside return column with the negative returns only
target = 0
downside_returns = returns.loc[returns < target]
# Calculate expected return and std dev of downside
expected_return = returns.mean()
down_stdev = downside_returns.std()
# Calculate the sortino ratio
rf = 0.01
sortino_ratio = (expected_return - rf)/down_stdev
# Print the results
print("Expected return: ", expected_return*100)
print('-' * 50)
print("Downside risk:")
print(down_stdev*100)
print('-' * 50)
print("Sortino ratio:")
print(sortino_ratio)
# + outputHidden=false inputHidden=false
# Calculate the max value
roll_max = returns.rolling(center=False,min_periods=1,window=252).max()
# Calculate the daily draw-down relative to the max
daily_draw_down = returns/roll_max - 1.0
# Calculate the minimum (negative) daily draw-down
max_daily_draw_down = daily_draw_down.rolling(center=False,min_periods=1,window=252).min()
# Plot the results
plt.figure(figsize=(15,15))
plt.plot(returns.index, daily_draw_down, label='Daily drawdown')
plt.plot(returns.index, max_daily_draw_down, label='Maximum daily drawdown in time-window')
plt.legend()
plt.show()
# + outputHidden=false inputHidden=false
# Box plot
returns.plot(kind='box')
# + outputHidden=false inputHidden=false
print("Stock returns: ")
print(returns.mean())
print('-' * 50)
print("Stock risk:")
print(returns.std())
# + outputHidden=false inputHidden=false
rf = 0.001
Sharpe_Ratio = ((returns.mean() - rf) / returns.std()) * np.sqrt(252)
print('Sharpe Ratio: ', Sharpe_Ratio)
# -
# ### Value-at-Risk 99% Confidence
# + outputHidden=false inputHidden=false
# 99% confidence interval
# 0.01 empirical quantile of daily returns
var99 = round((returns).quantile(0.01), 3)
# + outputHidden=false inputHidden=false
print('Value at Risk (99% confidence)')
print(var99)
# + outputHidden=false inputHidden=false
# the percent value of the 5th quantile
print('Percent Value-at-Risk of the 5th quantile')
var_1_perc = round(np.quantile(var99, 0.01), 3)
print("{:.1f}%".format(-var_1_perc*100))
# + outputHidden=false inputHidden=false
print('Value-at-Risk of 99% for 100,000 investment')
print("${}".format(int(-var99 * 100000)))
# -
# ### Value-at-Risk 95% Confidence
# + outputHidden=false inputHidden=false
# 95% confidence interval
# 0.05 empirical quantile of daily returns
var95 = round((returns).quantile(0.05), 3)
# + outputHidden=false inputHidden=false
print('Value at Risk (95% confidence)')
print(var95)
# + outputHidden=false inputHidden=false
print('Percent Value-at-Risk of the 5th quantile')
print("{:.1f}%".format(-var95*100))
# + outputHidden=false inputHidden=false
# VaR for 100,000 investment
print('Value-at-Risk of 99% for 100,000 investment')
var_100k = "${}".format(int(-var95 * 100000))
print("${}".format(int(-var95 * 100000)))
# + outputHidden=false inputHidden=false
mean = np.mean(returns)
std_dev = np.std(returns)
# + outputHidden=false inputHidden=false
returns.hist(bins=50, normed=True, histtype='stepfilled', alpha=0.5)
x = np.linspace(mean - 3*std_dev, mean + 3*std_dev, 100)
plt.plot(x, mlab.normpdf(x, mean, std_dev), "r")
plt.title('Histogram of Returns')
plt.show()
# + outputHidden=false inputHidden=false
VaR_90 = norm.ppf(1-0.9, mean, std_dev)
VaR_95 = norm.ppf(1-0.95, mean, std_dev)
VaR_99 = norm.ppf(1-0.99, mean, std_dev)
# + outputHidden=false inputHidden=false
print(tabulate([['90%', VaR_90], ['95%', VaR_95], ['99%', VaR_99]], headers=['Confidence Level', 'Value at Risk']))
|
Python_Stock/Portfolio_Strategies/Long_term_SPY.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The notebook is meant to help the user experiment with different models and features. This notebook assumes that there is a saved csv called 'filteredAggregateData.csv' somewhere on your local harddrive. The location must be specified below.
# The cell imports all of the relevant packages.
# +
############## imports
# general
import statistics
import datetime
from sklearn.externals import joblib # save and load models
import random
# data manipulation and exploration
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
## machine learning stuff
# preprocessing
from sklearn import preprocessing
# feature selection
from sklearn.feature_selection import SelectKBest, SelectPercentile
from sklearn.feature_selection import f_regression
# pipeline
from sklearn.pipeline import Pipeline
# train/testing
from sklearn.model_selection import train_test_split, KFold, GridSearchCV, cross_val_score
# error calculations
from sklearn.metrics import mean_squared_error, mean_absolute_error, r2_score
# models
from sklearn.linear_model import LinearRegression # linear regression
from sklearn.linear_model import BayesianRidge #bayesisan ridge regression
from sklearn.svm import SVR # support vector machines regression
from sklearn.gaussian_process import GaussianProcessRegressor # import GaussianProcessRegressor
from sklearn.neighbors import KNeighborsRegressor # k-nearest neightbors for regression
from sklearn.neural_network import MLPRegressor # neural network for regression
from sklearn.tree import DecisionTreeRegressor # decision tree regressor
from sklearn.ensemble import RandomForestRegressor # random forest regression
from sklearn.ensemble import AdaBoostRegressor # adaboost for regression
# saving models
# from sklearn.externals import joblib
import joblib
# -
# Imports the API. 'APILoc' is the location of 'API.py' on your local harddrive.
# +
# import the API
APILoc = r"C:\Users\thejo\Documents\school\AI in AG research\API"
import sys
sys.path.insert(0, APILoc)
from API import *
# -
# Load the dataset. Note that the location of the dataset must be specified.
# +
# get aggregate data
aggDataLoc = r'C:\Users\thejo\Documents\school\AI in AG research\experiment\aggregateData_RockSprings_PA.csv'
#aggDataLoc = r'C:\Users\thejo\Documents\school\AI in AG research\experiment\aggregateDataWithVariety.csv'
targetDataLoc = r'C:\Users\thejo\Documents\school\AI in AG research\experiment\aggregateData_GAonly_Annual_final.csv'
aggDf = pd.read_csv(aggDataLoc)
#aggDf = aggDf.drop("Unnamed: 0",axis=1)
targetDf = pd.read_csv(targetDataLoc)
#targetDf = targetDf.drop("Unnamed: 0",axis=1)
# -
# Test to see if the dataset was loaded properly. A table of the first 5 datapoints should appear.
aggDf.head()
#targetDf.head()
# Filter out features that will not be made available for feature selection. All of the features in the list 'XColumnsToKeep' will be made available for feature selection. The features to include are: <br>
# "Julian Day" <br>
# "Time Since Sown (Days)" <br>
# "Time Since Last Harvest (Days)" <br>
# "Total Radiation (MJ/m^2)" <br>
# "Total Rainfall (mm)" <br>
# "Avg Air Temp (C)" <br>
# "Avg Min Temp (C)" <br>
# "Avg Max Temp (C)"<br>
# "Avg Soil Moisture (%)"<br>
# "Day Length (hrs)"<br>
# "Percent Cover (%)"<br>
# +
# filter out the features that will not be used by the machine learning models
# the features to keep:
# xColumnsToKeep = ["Julian Day", "Time Since Sown (Days)", "Time Since Last Harvest (Days)", "Total Radiation (MJ/m^2)",
# "Total Rainfall (mm)", "Avg Air Temp (C)", "Avg Min Temp (C)", "Avg Max Temp (C)",
# "Avg Soil Moisture (%)", "Day Length (hrs)"], "Percent Cover (%)"]
xColumnsToKeep = ["Julian Day", "Time Since Sown (Days)", "Total Radiation (MJ/m^2)",
"Total Rainfall (mm)", "Avg Air Temp (C)", "Avg Min Temp (C)", "Avg Max Temp (C)",
"Avg Soil Moisture (%)"]
#xColumnsToKeep = ["Julian Day", "Time Since Sown (Days)", "Total Radiation (MJ/m^2)", "Total Rainfall (mm)"]
# the target to keep
yColumnsToKeep = ["Yield (tons/acre)"]
# get a dataframe containing the features and the targets
xDf = aggDf[xColumnsToKeep]
test_xDf = targetDf[xColumnsToKeep]
yDf = aggDf[yColumnsToKeep]
test_yDf = targetDf[yColumnsToKeep]
# reset the index
xDf = xDf.reset_index(drop=True)
yDf = yDf.reset_index(drop=True)
test_xDf = xDf.reset_index(drop=True)
test_yDf = yDf.reset_index(drop=True)
pd.set_option('display.max_rows', 2500)
pd.set_option('display.max_columns', 500)
xCols = list(xDf)
# -
# Test to see if the features dataframe and the target dataframe were successfully made.
xDf.head()
yDf.head()
# Lets now define the parameters that will be used to run the machine learning experiments. Note that parameter grids could be made that will allow sci-kit learn to use a 5-fold gridsearch to find the model's best hyperparameters. The parameter grids that are defined here will specify the possible values for the grid search. <br>
# <br>
# Once the parameter grids are defined, a list of tuples must also be defined. The tuples must take the form of: <br>
# (sci-kit learn model, appropriate parameter grid, name of the file to be saved). <br>
# <br>
# Then the number of iterations should be made. This is represented by the variable 'N'. Each model will be evaluated N times (via N-fold cross validation), and the average results of the models over those N iterations will be returned. <br>
# <br>
# 'workingDir' is the directory in which all of the results will be saved. <br>
# <br>
# 'numFeatures' is the number of features that will be selected (via feature selection).
# +
# hide the warnings because training the neural network caues lots of warnings.
import warnings
warnings.filterwarnings('ignore')
# make the parameter grids for sklearn's gridsearchcv
rfParamGrid = {
'model__n_estimators': [5, 10, 25, 50, 100], # Number of estimators
'model__max_depth': [5, 10, 15, 20], # Maximum depth of the tree
'model__criterion': ["mae"]
}
knnParamGrid ={
'model__n_neighbors':[2,5,10],
'model__weights': ['uniform', 'distance'],
'model__leaf_size': [5, 10, 30, 50]
}
svrParamGrid = {
'model__kernel': ['linear', 'poly', 'rbf', 'sigmoid'],
'model__C': [0.1, 1.0, 5.0, 10.0],
'model__gamma': ["scale", "auto"],
'model__degree': [2,3,4,5]
}
nnParamGrid = {
'model__hidden_layer_sizes':[(3), (5), (10), (3,3), (5,5), (7,7)],
'model__solver': ['sgd', 'adam'],
'model__learning_rate' : ['constant', 'invscaling', 'adaptive'],
'model__learning_rate_init': [0.1, 0.01, 0.001]
}
linRegParamGrid = {}
bayesParamGrid={
'model__n_iter':[100,300,500]
}
dtParamGrid = {
'model__criterion': ['mae'],
'model__max_depth': [5,10,25,50,100]
}
aModelList = [(MLPRegressor(), nnParamGrid, "nnTup.pkl")]
N = 10
workingDir = r"C:\Users\thejo\Documents\school\AI in AG research\experiment"
numFeatures = 8 # 11
# -
# This cell will run the tests and save the results.
saveMLResults(test_xDf, test_yDf, N, xDf, yDf, aModelList, workingDir, numFeatures, printResults=True)
|
notebooks/modelExperiments112320_PA_to_GA-NN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python37]
# language: python
# name: conda-env-python37-py
# ---
# +
# Millions to miss out on the net
# aust addresses un security council over iraq
# -
import requests
import os
from tqdm import tqdm
import spacy
nlp = spacy.load("en")
# +
host = "http://0.0.0.0:8890/correct"
def call_gec(data):
resp = requests.post(host, json=data)
res = resp.json()
return res
# -
data_path = '/home/citao/github/BBC-Dataset-News-Classification/dataset/data_files'
domains = ['business', 'entertainment', 'politics', 'sport', 'tech']
# +
corpus = {d: [] for d in domains}
for domain in domains:
domain_path = os.path.join(data_path, domain)
for f in os.listdir(domain_path):
full_path = os.path.join(domain_path, f)
try:
doc = nlp(open(full_path).read())
corpus[domain].extend([i.text for i in doc.sents])
except:
print(full_path)
print(domain, len(corpus[domain]))
# +
import json
json.dump(corpus, open('./bbc_news.json', 'w'))
# +
# text = "England coach <NAME> is already without centre <NAME> and flanker <NAME> while fly-half <NAME> is certain to miss at least the first two games."
# text = "Hi, Guibin! My namme is Citao. The marked was closed yestreday. (This email are sent from OnMail.)"
# text = "Millions to miss out on the net"
# text = "The marked is closed today."
text = """
Hi,
My namme is <NAME> (EID: cw39729) and I do admitted into the Computer Online Master prograssm 2021 spring. In my status panel, I saw a requirement about "Required immunizations".
My question is whether I still need to meets this requirement or not, even thoxugh my program is fully online.
If no, could you please help me to remove this requirements from my application?
Thanks,
"""
# text = "I want go school"
# text = "I I want go to school"
data = {
'text': text,
'iterations': 3,
'min_probability': 0.5,
'min_error_probability': 0.7,
'case_sensitive': True,
'languagetool_post_process': True,
'languagetool_call_thres': 0.7,
'whitelist': ['Citao', 'Guibin', 'Onmail'],
'with_debug_info': True,
}
result = call_gec(data)
print(result['debug_info'])
print(result['input'])
print(result['output'])
print(result['corrections'])
# -
for domain in domains:
print(domain)
right_data = record[domain]['pass']
wrong_data = record[domain]['wrong']
print(len(right_data), len(wrong_data))
for sent in record['tech']['wrong']:
print([sent])
break
# +
import json
total = json.load(open('./bbc_news_result_2_0.8_0.9_False.json'))
i = 0
for min_probability in [0.5, 0.6, 0.7, 0.8]:
for min_error_probability in [0.7, 0.8, 0.9]:
for add_spell_check in [True, False]:
fn = './result/bbc_news_result_2_{}_{}_{}.json'.format(min_probability, min_error_probability, str(add_spell_check))
tmp = {domain: total[domain][5000*i:5000*(i+1)] for domain in domains}
print(fn)
json.dump(tmp, open(fn, 'w'))
i+=1
# +
import pandas as pd
source_rows = []
for min_probability in [0.5, 0.6, 0.7, 0.8]:
for min_error_probability in [0.7, 0.8, 0.9]:
for add_spell_check in [True, False]:
fn = './result/bbc_news_result_2_{}_{}_{}.json'.format(min_probability, min_error_probability, str(add_spell_check))
tmp = json.load(open(fn))
result = {
'cased': {domain: {'right':[], 'wrong':[]} for domain in domains},
'uncased': {domain: {'right':[], 'wrong':[]} for domain in domains}
}
print(fn)
for domain in domains:
for ori_sent, cor_sent, correction in tmp[domain]:
if correction == []:
result['cased'][domain]['right'].append(ori_sent)
result['uncased'][domain]['right'].append(ori_sent)
else:
result['cased'][domain]['wrong'].append(ori_sent)
uncased_correction = [c for c in correction if c[0].lower()!=c[1].lower()]
if uncased_correction:
result['uncased'][domain]['wrong'].append(ori_sent)
else:
result['uncased'][domain]['right'].append(ori_sent)
for case_type in ['cased', 'uncased']:
row = {
'min_probability': min_probability,
'min_error_probability': min_error_probability,
'add_spell_check': add_spell_check,
'cased_type': case_type,
}
row.update({d: len(result[case_type][d]['right'])/5000.0 for d in domains})
source_rows.append(row)
# -
df = pd.DataFrame(source_rows)
df
df.to_csv('~/eval_pos.csv', index=False)
for domain in domains:
df[df['min_probability']==0.]
import numpy as np
1-np.average(df[df['cased_type']=='uncased'].iloc[:, 4:].to_numpy())
# +
path='/home/citao/github/gector/dataset/'
fn = 'wil.ABCN.dev.gold.bea19.0'
source_list = [i.strip() for i in open(path+fn+'.source').readlines()]
target_list = [i.strip() for i in open(path+fn+'.target').readlines()]
diff = 0
for s, t in zip(source_list, target_list):
if s!=t:
diff+=1
print(diff, len(source_list), len(target_list))
# -
4384-2819
for s, t in zip(source_list[:10], target_list):
print(s)
print(t)
print()
25000+1565
2819 / (2819+26565)
# +
import Levenshtein
from difflib import SequenceMatcher
from nltk.corpus import words
NLTK_COMMON_WORDS = {w:1 for w in words.words()}
# 1: spelling error
# 2: grammar error
def spell_or_grammer_error(correction):
ori, cor, _ = correction
if len(ori.split()) != len(cor.split()):
return 2
# dist = Levenshtein.distance(ori, cor)
# print(dist)
seq = SequenceMatcher(None, ori, cor)
ratio = seq.ratio()
print(ratio)
# if dist <= 3:
if ratio > 0.8 and ori not in NLTK_COMMON_WORDS:
return 1
else:
return 2
correction = ['are', 'is', [10, 15]]
spell_or_grammer_error(correction)
# -
|
eval_pos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
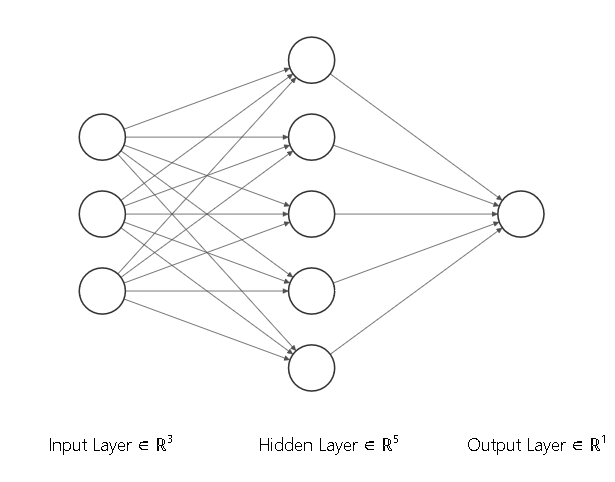
# # Buidling simple layers of a neural net
#
# ### Dr. <NAME><br><br>Fremont, CA 94536 <br><br>Nov 2019
import torch
import numpy as np
torch.manual_seed(7)
# ### Simple feature vector of dimension 3
features = torch.randn((1,3))
print(features)
# ### Define the size of each layer of the neural network
n_input = features.shape[1] # Must match the shape of the features
n_hidden = 5 # Number of hidden units
n_output = 1 # Number of output units (for example 1 for binary classification)
# ### Weights for the input layer to the hidden layer
W1 = torch.randn(n_input,n_hidden)
# ### Weights for the hidden layer to the output layer
W2 = torch.randn(n_hidden,n_output)
# ### Bias terms for the hidden and the output layer
B1 = torch.randn((1,n_hidden))
B2 = torch.randn((1,n_output))
# ### Define the activation function - sigmoid
# $$\sigma(x)=\frac{1}{1+exp(-x)}$$
def activation(x):
"""
Sigmoid activation function
"""
return 1/(1+torch.exp(-x))
# ### Check the shape of all the tensors
print("Shape of the input features: ",features.shape)
print("Shape of the first tensor of weights (between input and hidden layers): ",W1.shape)
print("Shape of the second tensor of weights (between hidden and output layers): ",W2.shape)
print("Shape of the bias tensor added to the hidden layer: ",B1.shape)
print("Shape of the bias tensor added to the output layer: ",B2.shape)
# 
# ### First layer output
# $$\mathbf{h_1} = sigmoid(\mathbf{W_1}\times\mathbf{feature}+\mathbf{B_1})$$
h1 = activation(torch.mm(features,W1)+B1)
print("Shape of the output of the hidden layer",h1.shape)
# ### Second layer output
# $$\mathbf{h_2} = sigmoid(\mathbf{W_2}\times\mathbf{h1}+\mathbf{B_1})$$
h2 = activation(torch.mm(h1,W2)+B2)
print("Shape of the output layer",h2.shape)
print(h2)
|
Building a simple NN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*Superposition*_
#
#
# The latest version of this notebook is available on https://github.com/qiskit/qiskit-tutorial.
#
# ***
# ### Contributors
# <NAME>, <NAME>, <NAME>, <NAME>
#
# ### Qiskit Package Versions
import qiskit
qiskit.__qiskit_version__
# ## Introduction
# Many people tend to think quantum physics is hard math, but this is not actually true. Quantum concepts are very similar to those seen in the linear algebra classes you may have taken as a freshman in college, or even in high school. The challenge of quantum physics is the necessity to accept counter-intuitive ideas, and its lack of a simple underlying theory. We believe that if you can grasp the following two Principles, you will have a good start:
# 1. A physical system in a definite state can still behave randomly.
# 2. Two systems that are too far apart to influence each other can nevertheless behave in ways that, though individually random, are somehow strongly correlated.
#
# In this tutorial, we will be discussing the first of these Principles, the second is discussed in [this other tutorial](entanglement_introduction.ipynb).
# +
# useful additional packages
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# importing Qiskit
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, execute
from qiskit import BasicAer, IBMQ
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# +
backend = BasicAer.get_backend('qasm_simulator') # run on local simulator by default
# Uncomment the following lines to run on a real device
#IBMQ.load_accounts()
#from qiskit.providers.ibmq import least_busy
#backend = least_busy(IBMQ.backends(operational=True, simulator=False))
#print("the best backend is " + backend.name())
# -
# ## Quantum States - Basis States and Superpositions<a id='section1'></a>
#
# The first Principle above tells us that the results of measuring a quantum state may be random or deterministic, depending on what basis is used. To demonstrate, we will first introduce the computational (or standard) basis for a qubit.
#
# The computational basis is the set containing the ground and excited state $\{|0\rangle,|1\rangle\}$, which also corresponds to the following vectors:
#
# $$|0\rangle =\begin{pmatrix} 1 \\ 0 \end{pmatrix}$$
# $$|1\rangle =\begin{pmatrix} 0 \\ 1 \end{pmatrix}$$
#
# In Python these are represented by
zero = np.array([[1],[0]])
one = np.array([[0],[1]])
# In our quantum processor system (and many other physical quantum processors) it is natural for all qubits to start in the $|0\rangle$ state, known as the ground state. To make the $|1\rangle$ (or excited) state, we use the operator
#
# $$ X =\begin{pmatrix} 0 & 1 \\ 1 & 0 \end{pmatrix}.$$
#
# This $X$ operator is often called a bit-flip because it exactly implements the following:
#
# $$X: |0\rangle \rightarrow |1\rangle$$
# $$X: |1\rangle \rightarrow |0\rangle.$$
#
# In Python this can be represented by the following:
X = np.array([[0,1],[1,0]])
print(np.dot(X,zero))
print(np.dot(X,one))
# Next, we give the two quantum circuits for preparing and measuring a single qubit in the ground and excited states using Qiskit.
# +
# Creating registers
qr = QuantumRegister(1)
cr = ClassicalRegister(1)
# Quantum circuit ground
qc_ground = QuantumCircuit(qr, cr)
qc_ground.measure(qr[0], cr[0])
# Quantum circuit excited
qc_excited = QuantumCircuit(qr, cr)
qc_excited.x(qr)
qc_excited.measure(qr[0], cr[0])
# -
qc_ground.draw(output='mpl')
qc_excited.draw(output='mpl')
# Here we have created two jobs with different quantum circuits; the first to prepare the ground state, and the second to prepare the excited state. Now we can run the prepared jobs.
circuits = [qc_ground, qc_excited]
job = execute(circuits, backend)
result = job.result()
# After the run has been completed, the data can be extracted from the API output and plotted.
plot_histogram(result.get_counts(qc_ground))
plot_histogram(result.get_counts(qc_excited))
# Here we see that the qubit is in the $|0\rangle$ state with 100% probability for the first circuit and in the $|1\rangle$ state with 100% probability for the second circuit. If we had run on a quantum processor rather than the simulator, there would be a difference from the ideal perfect answer due to a combination of measurement error, preparation error, and gate error (for the $|1\rangle$ state).
#
# Up to this point, nothing is different from a classical system of a bit. To go beyond, we must explore what it means to make a superposition. The operation in the quantum circuit language for generating a superposition is the Hadamard gate, $H$. Let's assume for now that this gate is like flipping a fair coin. The result of a flip has two possible outcomes, heads or tails, each occurring with equal probability. If we repeat this simple thought experiment many times, we would expect that on average we will measure as many heads as we do tails. Let heads be $|0\rangle$ and tails be $|1\rangle$.
#
# Let's run the quantum version of this experiment. First we prepare the qubit in the ground state $|0\rangle$. We then apply the Hadamard gate (coin flip). Finally, we measure the state of the qubit. Repeat the experiment 1024 times (shots). As you likely predicted, half the outcomes will be in the $|0\rangle$ state and half will be in the $|1\rangle$ state.
#
# Try the program below.
# +
# Quantum circuit superposition
qc_superposition = QuantumCircuit(qr, cr)
qc_superposition.h(qr)
qc_superposition.measure(qr[0], cr[0])
qc_superposition.draw()
# +
job = execute(qc_superposition, backend, shots = 1024)
result = job.result()
plot_histogram(result.get_counts(qc_superposition))
# -
# Indeed, much like a coin flip, the results are close to 50/50 with some non-ideality due to errors (again due to state preparation, measurement, and gate errors). So far, this is still not unexpected. Let's run the experiment again, but this time with two $H$ gates in succession. If we consider the $H$ gate to be analog to a coin flip, here we would be flipping it twice, and still expecting a 50/50 distribution.
# +
# Quantum circuit two Hadamards
qc_twohadamard = QuantumCircuit(qr, cr)
qc_twohadamard.h(qr)
qc_twohadamard.barrier()
qc_twohadamard.h(qr)
qc_twohadamard.measure(qr[0], cr[0])
qc_twohadamard.draw(output='mpl')
# +
job = execute(qc_twohadamard, backend)
result = job.result()
plot_histogram(result.get_counts(qc_twohadamard))
# -
# This time, the results are surprising. Unlike the classical case, with high probability the outcome is not random, but in the $|0\rangle$ state. *Quantum randomness* is not simply like a classical random coin flip. In both of the above experiments, the system (without noise) is in a definite state, but only in the first case does it behave randomly. This is because, in the first case, via the $H$ gate, we make a uniform superposition of the ground and excited state, $(|0\rangle+|1\rangle)/\sqrt{2}$, but then follow it with a measurement in the computational basis. The act of measurement in the computational basis forces the system to be in either the $|0\rangle$ state or the $|1\rangle$ state with an equal probability (due to the uniformity of the superposition). In the second case, we can think of the second $H$ gate as being a part of the final measurement operation; it changes the measurement basis from the computational basis to a *superposition* basis. The following equations illustrate the action of the $H$ gate on the computational basis states:
# $$H: |0\rangle \rightarrow |+\rangle=\frac{|0\rangle+|1\rangle}{\sqrt{2}}$$
# $$H: |1\rangle \rightarrow |-\rangle=\frac{|0\rangle-|1\rangle}{\sqrt{2}}.$$
# We can redefine this new transformed basis, the superposition basis, as the set {$|+\rangle$, $|-\rangle$}. We now have a different way of looking at the second experiment above. The first $H$ gate prepares the system into a superposition state, namely the $|+\rangle$ state. The second $H$ gate followed by the standard measurement changes it into a measurement in the superposition basis. If the measurement gives 0, we can conclude that the system was in the $|+\rangle$ state before the second $H$ gate, and if we obtain 1, it means the system was in the $|-\rangle$ state. In the above experiment we see that the outcome is mainly 0, suggesting that our system was in the $|+\rangle$ superposition state before the second $H$ gate.
#
#
# The math is best understood if we represent the quantum superposition state $|+\rangle$ and $|-\rangle$ by:
#
# $$|+\rangle =\frac{1}{\sqrt{2}}\begin{pmatrix} 1 \\ 1 \end{pmatrix}$$
# $$|-\rangle =\frac{1}{\sqrt{2}}\begin{pmatrix} 1 \\ -1 \end{pmatrix}$$
#
# A standard measurement, known in quantum mechanics as a projective or von Neumann measurement, takes any superposition state of the qubit and projects it to either the state $|0\rangle$ or the state $|1\rangle$ with a probability determined by:
#
# $$P(i|\psi) = |\langle i|\psi\rangle|^2$$
#
# where $P(i|\psi)$ is the probability of measuring the system in state $i$ given preparation $\psi$.
#
# We have written the Python function ```state_overlap``` to return this:
state_overlap = lambda state1, state2: np.absolute(np.dot(state1.conj().T,state2))**2
# Now that we have a simple way of going from a state to the probability distribution of a standard measurement, we can go back to the case of a superposition made from the Hadamard gate. The Hadamard gate is defined by the matrix:
#
# $$ H =\frac{1}{\sqrt{2}}\begin{pmatrix} 1 & 1 \\ 1 & -1 \end{pmatrix}$$
#
# The $H$ gate acting on the state $|0\rangle$ gives:
Hadamard = np.array([[1,1],[1,-1]],dtype=complex)/np.sqrt(2)
psi1 = np.dot(Hadamard,zero)
P0 = state_overlap(zero,psi1)
P1 = state_overlap(one,psi1)
plot_histogram({'0' : P0.item(0), '1' : P1.item(0)})
# which is the ideal version of the first superposition experiment.
#
# The second experiment involves applying the Hadamard gate twice. While matrix multiplication shows that the product of two Hadamards is the identity operator (meaning that the state $|0\rangle$ remains unchanged), here (as previously mentioned) we prefer to interpret this as doing a measurement in the superposition basis. Using the above definitions, you can show that $H$ transforms the computational basis to the superposition basis.
print(np.dot(Hadamard,zero))
print(np.dot(Hadamard,one))
# This is just the beginning of how a quantum state differs from a classical state. Please continue to [Amplitude and Phase](amplitude_and_phase.ipynb) to explore further!
|
terra/qis_intro/superposition.ipynb
|
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="a2782576-c8ad-483e-bd03-289dd656844c" extensions={"azuredatastudio": {"views": []}}
/ # Set up Azure SQL Database for catching the bus application
/
/ This is a SQL Notebook, which allows you to separate text and code blocks and save code results. Azure Data Studio supports several languages, referred to as kernels, including SQL, PowerShell, Python, and more.
/
/ In this activity, you'll learn how to import data into Azure SQL Database and create tables to store the route data, geofence data, and real-time bus information.
/
/ ## Connect to `bus-db`
/
/ At the top of the window, select **Select Connection** \> **Change Connection** next to "Attach to".
/
/ Under _Recent Connections_ select your `bus-db` connection.
/
/ You should now see it listed next to _Attach to_.
/ + [markdown] azdata_cell_guid="f348fdae-e69a-4271-907d-e6b4e0619151" extensions={"azuredatastudio": {"views": []}}
/ ## Part 1: Import the bus route data from Azure Blob Storage
/
/ The first step in configuring the database for the scenario is to import a CSV file that contains route information data. The following script will walk you through that process. Full documentation on "Accessing data in a CSV file referencing an Azure blob storage location" here: [https://docs.microsoft.com/sql/relational-databases/import-export/examples-of-bulk-access-to-data-in-azure-blob-storage](https://docs.microsoft.com/sql/relational-databases/import-export/examples-of-bulk-access-to-data-in-azure-blob-storage).
/
/ You need to first create a table and schema for data to be loaded into.
/ + azdata_cell_guid="c14329fd-fee4-4014-a77d-ed5b59785685" extensions={"azuredatastudio": {"views": []}}
CREATE TABLE [dbo].[Routes]
(
[Id] [int] NOT NULL,
[AgencyId] [varchar](100) NULL,
[ShortName] [varchar](100) NULL,
[Description] [varchar](1000) NULL,
[Type] [int] NULL
)
GO
ALTER TABLE [dbo].[Routes] ADD PRIMARY KEY CLUSTERED
(
[Id] ASC
)
GO
/ + [markdown] azdata_cell_guid="962dcc3f-be18-4cc3-bdfc-d670c962a0dc" extensions={"azuredatastudio": {"views": []}}
/ The next step is to create a master key.
/ + azdata_cell_guid="5a690353-b1f9-43b2-85e1-2a7467f7b3ad" extensions={"azuredatastudio": {"views": []}}
CREATE MASTER KEY ENCRYPTION BY PASSWORD = '<PASSWORD>!'
/ + [markdown] azdata_cell_guid="3a30a8e0-5054-471c-84f3-6e28dc47c694" extensions={"azuredatastudio": {"views": []}}
/ A master key is required to create a `DATABASE SCOPED CREDENTIAL` value because Blob storage is not configured to allow public (anonymous) access. The credential refers to the Blob storage account, and the data portion specifies the container for the store return data.
/
/ We use a shared access signature as the identity that Azure SQL knows how to interpret. The secret is the SAS token that you can generate from the Blob storage account. In this example, the SAS token for a storage account that you don't have access to is provided so you can access only the store return data.
/ + azdata_cell_guid="92c0d495-d239-4dd1-b095-05f8fa0a6cef" extensions={"azuredatastudio": {"views": []}}
CREATE DATABASE SCOPED CREDENTIAL AzureBlobCredentials
WITH IDENTITY = 'SHARED ACCESS SIGNATURE',
SECRET = 'sp=r&st=2021-03-12T00:47:24Z&se=2025-03-11T07:47:24Z&spr=https&sv=2020-02-10&sr=c&sig=BmuxFevKhWgbvo%2Bj8TlLYObjbB7gbvWzQaAgvGcg50c%3D' -- Omit any leading question mark
/ + [markdown] azdata_cell_guid="bca17886-bcfa-4eaa-87b1-94e94e801ce7" extensions={"azuredatastudio": {"views": []}}
/ Next, create an external data source to the container.
/ + azdata_cell_guid="aea81643-4d8a-44b2-a915-8d797f036be5" extensions={"azuredatastudio": {"views": []}}
CREATE EXTERNAL DATA SOURCE RouteData
WITH (
TYPE = blob_storage,
LOCATION = 'https://azuresqlworkshopsa.blob.core.windows.net/bus',
CREDENTIAL = AzureBlobCredentials
)
/ + [markdown] azdata_cell_guid="7658b21b-c9a0-4ffd-a2dc-7af3d0f87842" extensions={"azuredatastudio": {"views": []}}
/ Now you are ready to bring in the data.
/ + azdata_cell_guid="3f642673-43ec-4215-91da-a3a37776fb2f" extensions={"azuredatastudio": {"views": []}} tags=[]
DELETE FROM dbo.[Routes];
INSERT INTO dbo.[Routes]
([Id], [AgencyId], [ShortName], [Description], [Type])
SELECT
[Id], [AgencyId], [ShortName], [Description], [Type]
FROM
OPENROWSET
(
BULk 'routes.txt',
DATA_SOURCE = 'RouteData',
FORMATFILE = 'routes.fmt',
FORMATFILE_DATA_SOURCE = 'RouteData',
FIRSTROW=2,
FORMAT='csv'
) t;
/ + [markdown] azdata_cell_guid="622d6f84-6ece-4b52-a2a1-8666cec71128" extensions={"azuredatastudio": {"views": []}}
/ Finally, let's look at what's been inserted relative to the route we'll be tracking.
/ + azdata_cell_guid="48fa0ba2-64cb-4230-a9b2-e7264994d7d7" extensions={"azuredatastudio": {"views": []}}
SELECT * FROM dbo.[Routes] WHERE [Description] LIKE '%Education Hill%'
/ + [markdown] azdata_cell_guid="102d8657-f5ab-4921-809d-d77fc9b41ad2" extensions={"azuredatastudio": {"views": []}}
/ ## Part 2: Create necessary tables
/
/ ### Select a route to monitor
/
/ Now that you've added the route information, you can select the route to be a "Monitored Route". This will come in handy if you later choose to monitor multiple routes. For now, you will just add the "Education Hill - Crossroads - Eastgate" route.
/ + azdata_cell_guid="b8d596bf-0540-49fa-b206-ce52397c0459" extensions={"azuredatastudio": {"views": []}}
-- Create MonitoredRoutes table
CREATE TABLE [dbo].[MonitoredRoutes]
(
[RouteId] [int] NOT NULL
)
GO
ALTER TABLE [dbo].[MonitoredRoutes] ADD PRIMARY KEY CLUSTERED
(
[RouteId] ASC
)
GO
ALTER TABLE [dbo].[MonitoredRoutes]
ADD CONSTRAINT [FK__MonitoredRoutes__Router]
FOREIGN KEY ([RouteId]) REFERENCES [dbo].[Routes] ([Id])
GO
-- Monitor the "Education Hill - Crossroads - Eastgate" route
INSERT INTO dbo.[MonitoredRoutes] (RouteId) VALUES (100113);
/ + [markdown] azdata_cell_guid="a53ad031-a7c3-40a9-bf5b-695a148dbea9" extensions={"azuredatastudio": {"views": []}}
/ ### Select a GeoFence to monitor
/
/ In addition to monitoring specific bus routes, you will want to monitor certain GeoFences so you can ultimately get notified when your bus enters or exits where you are (i.e. the GeoFence). For now, you will add a small GeoFence that represents the area near the "Crossroads" bus stop.
/ + azdata_cell_guid="1cc9823a-4b83-4e91-a36f-8b503abf0347" extensions={"azuredatastudio": {"views": []}}
-- Create GeoFences table
CREATE SEQUENCE [dbo].[global]
AS INT
START WITH 1
INCREMENT BY 1
GO
CREATE TABLE [dbo].[GeoFences](
[Id] [int] NOT NULL,
[Name] [nvarchar](100) NOT NULL,
[GeoFence] [geography] NOT NULL
)
GO
ALTER TABLE [dbo].[GeoFences] ADD PRIMARY KEY CLUSTERED
(
[Id] ASC
)
GO
ALTER TABLE [dbo].[GeoFences] ADD DEFAULT (NEXT VALUE FOR [dbo].[global]) FOR [Id]
GO
CREATE SPATIAL INDEX [ixsp] ON [dbo].[GeoFences]
(
[GeoFence]
) USING GEOGRAPHY_AUTO_GRID
GO
-- Create a GeoFence
INSERT INTO dbo.[GeoFences]
([Name], [GeoFence])
VALUES
('Crossroads', 0xE6100000010407000000B4A78EA822CF4740E8D7539530895EC03837D51CEACE4740E80BFBE630895EC0ECD7DF53EACE4740E81B2C50F0885EC020389F0D03CF4740E99BD2A1F0885EC00CB8BEB203CF4740E9DB04FC23895EC068C132B920CF4740E9DB04FC23895EC0B4A78EA822CF4740E8D7539530895EC001000000020000000001000000FFFFFFFF0000000003);
GO
/ + [markdown] azdata_cell_guid="46ac87ea-ab53-4c87-9b41-57a3b96924a0" extensions={"azuredatastudio": {"views": []}}
/ ### Create table to track activity in the GeoFence
/
/ Next, create a system-versioned table to keep track of what activity is currently happening within the GeoFence. This means tracking buses entering, exiting, and staying within a given GeoFence. Another table within that table will serve as a histroical log for all activity.
/ + azdata_cell_guid="3e3da57b-2132-4788-b46c-20374d080b2f" extensions={"azuredatastudio": {"views": []}}
CREATE TABLE [dbo].[GeoFencesActive]
(
[Id] [int] IDENTITY(1,1) NOT NULL PRIMARY KEY CLUSTERED,
[VehicleId] [int] NOT NULL,
[DirectionId] [int] NOT NULL,
[GeoFenceId] [int] NOT NULL,
[SysStartTime] [datetime2](7) GENERATED ALWAYS AS ROW START NOT NULL,
[SysEndTime] [datetime2](7) GENERATED ALWAYS AS ROW END NOT NULL,
PERIOD FOR SYSTEM_TIME ([SysStartTime], [SysEndTime])
)
WITH ( SYSTEM_VERSIONING = ON ( HISTORY_TABLE = [dbo].[GeoFencesActiveHistory] ) )
GO
/ + [markdown] azdata_cell_guid="c8d2f9b4-af0e-48c2-a594-e21bb7e297a7" extensions={"azuredatastudio": {"views": []}}
/ ### Create a table to store real-time bus data
/
/ You'll need one last table to store the real-time bus data as it comes in.
/ + azdata_cell_guid="59797569-2711-4675-ab8a-532e0e2a7c22" extensions={"azuredatastudio": {"views": []}}
CREATE TABLE [dbo].[BusData](
[Id] [int] IDENTITY(1,1) NOT NULL,
[DirectionId] [int] NOT NULL,
[RouteId] [int] NOT NULL,
[VehicleId] [int] NOT NULL,
[Location] [geography] NOT NULL,
[TimestampUTC] [datetime2](7) NOT NULL,
[ReceivedAtUTC] [datetime2](7) NOT NULL
)
GO
ALTER TABLE [dbo].[BusData] ADD DEFAULT (SYSUTCDATETIME()) FOR [ReceivedAtUTC]
GO
ALTER TABLE [dbo].[BusData] ADD PRIMARY KEY CLUSTERED
(
[Id] ASC
)
GO
CREATE NONCLUSTERED INDEX [ix1] ON [dbo].[BusData]
(
[ReceivedAtUTC] DESC
)
GO
CREATE SPATIAL INDEX [ixsp] ON [dbo].[BusData]
(
[Location]
) USING GEOGRAPHY_AUTO_GRID
GO
/ + [markdown] azdata_cell_guid="48c43e3d-b0d0-462a-aaab-da6ca728b2de" extensions={"azuredatastudio": {"views": []}}
/ Confirm you've created the tables with the following.
/ + azdata_cell_guid="5619e4aa-306c-46e5-9779-699bb29e387a" extensions={"azuredatastudio": {"views": []}}
SELECT * FROM INFORMATION_SCHEMA.TABLES WHERE TABLE_TYPE = 'BASE TABLE' AND TABLE_SCHEMA = 'dbo'
|
database/notebooks/01-set-up-database.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''hinton'': conda)'
# name: python3
# ---
# # Breast cancer detection from thermal imaging
# The main purpose of this project is to develop a comprehensive decision support system for breast cancer screening.
# ## Library import
# In this section we will try to import the libraries that will be used throughout this model. Note that some of the libraries used in this program are declared in the files found in `src/scripts/*.py`.
# %reload_ext autoreload
# %autoreload 2
from scripts import *
computer.check_available_devices() # Check available devices
# ## Data selection
# To make this model work correctly it will be necessary to extract and save the images found in the `data` folder.
#
# In this folder there are two labeled folders that contain all the images to be used:
# ```
# data
# ├── healthy
# └── sick
# ```
# +
data = Data("./data/") # Data imported into a table
data.images.head(3) # Display first 3 rows
# -
# ## Transformation
# In the transformation stage, the data is adapted to find the solution to the problem to be solved.
# First of all, the data obtained previously will be divided to be able to use it for training and to check the results.
data.training, data.test = data.train_test_split(test_size=0.15, shuffle=True, stratify=True) # Split data into train and test
# The category distribution is shown for the original, training, and test data
data.count_labels(data.images, "Original")
data.count_labels(data.training, "Training")
data.count_labels(data.test, "Test")
# Once the data is divided, different transformation techniques are applied on it to expand the size of the dataset in real time while training the model.
train_generator, validation_generator, test_generator = data.image_generator(shuffle=False) # Image genearation
# +
filters = {
"original": lambda x: x,
"red": lambda x: data.getImageTensor(x, (330, 0, 0), (360, 255, 255)) + data.getImageTensor(x, (0, 0, 0), (50, 255, 255)),
"green": lambda x: data.getImageTensor(x, (60, 0, 0), (130, 255, 255)),
"blue": lambda x: data.getImageTensor(x, (180, 0, 0), (270, 255, 255)),
}
data.show_images(train_generator, filters, "Training") # Show some images from the training generator
# -
# ## Data Mining
# This section seeks to apply techniques that are capable of extracting useful patterns and then evaluate them.
# ### Model creation
# The model to be used for the next training is created.
red_model = Model("red", filter=filters["red"], new=True, summary=False, plot=False) # Red model creation
green_model = Model("green", filter=filters["green"], new=True, summary=False, plot=False) # Green model creation
blue_model = Model("blue", filter=filters["blue"], new=True, summary=False, plot=False) # Blue model creation
red_model.compile() # Compile the red model
green_model.compile() # Compile the green model
blue_model.compile() # Compile the blue model
# ### Model training
# The created model is trained indicating the times that are going to be used.
red_model.fit(train_generator, validation_generator, epochs=600, verbose=False, plot=False) # Train the red model
green_model.fit(train_generator, validation_generator, epochs=600, verbose=False, plot=False) # Train the green model
blue_model.fit(train_generator, validation_generator, epochs=600, verbose=False, plot=False) # Train the blue model
# ### Model evaluation
# The trained model is evaluated using the generators created before. In this case, the best weight matrix obtained in the training will be used.
red_model.evaluate(test_generator, path=None) # Evaluate the red model
green_model.evaluate(test_generator, path=None) # Evaluate the green model
blue_model.evaluate(test_generator, path=None) # Evaluate the blue model
# ### Grad-CAM
# An activation map of the predictions obtained by the convolutional network is displayed.
# +
join_models = Join(red_model, green_model, blue_model)
# The activation map is displayed
for index, image in data.test.iterrows():
join_models.visualize_heatmap(image)
|
src/model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp vision.core
#default_cls_lvl 3
# +
#export
from local.torch_basics import *
from local.test import *
from local.data.all import *
from local.notebook.showdoc import show_doc
from PIL import Image
# -
#export
_all_ = ['Image','ToTensor']
# +
#It didn't use to be necessary to add ToTensor in all but we don't have the encodes methods defined here otherwise.
#TODO: investigate
# -
# # Core vision
# > Basic image opening/processing functionality
# ## Helpers
im = Image.open(TEST_IMAGE).resize((30,20))
#export
@patch_property
def n_px(x: Image.Image): return x.size[0] * x.size[1]
# #### `Image.n_px`
#
# > `Image.n_px` (property)
#
# Number of pixels in image
test_eq(im.n_px, 30*20)
#export
@patch_property
def shape(x: Image.Image): return x.size[1],x.size[0]
# #### `Image.shape`
#
# > `Image.shape` (property)
#
# Image (height,width) tuple (NB: opposite order of `Image.size()`, same order as numpy array and pytorch tensor)
test_eq(im.shape, (20,30))
#export
@patch_property
def aspect(x: Image.Image): return x.size[0]/x.size[1]
# #### `Image.aspect`
#
# > `Image.aspect` (property)
#
# Aspect ratio of the image, i.e. `width/height`
test_eq(im.aspect, 30/20)
#export
@patch
def reshape(x: Image.Image, h, w, resample=0):
"`resize` `x` to `(w,h)`"
return x.resize((w,h), resample=resample)
show_doc(Image.Image.reshape)
test_eq(im.reshape(12,10).shape, (12,10))
#export
@patch
def resize_max(x: Image.Image, resample=0, max_px=None, max_h=None, max_w=None):
h,w = x.shape
if max_px and x.n_px>max_px: h,w = h*max_px/x.n_px,w*max_px/x.n_px
if max_h and h>max_h: h,w = h*max_h/h,w*max_h/h
if max_w and w>max_w: h,w = h*max_w/w,w*max_w/w
return x.reshape(round(h), round(w), resample=resample)
test_eq(im.resize_max(max_px=30*20).size, (30,20))
test_eq(im.resize_max(max_px=30*20/2).size, (15,10))
test_eq(im.resize_max(max_px=500, max_h=10, max_w=20).size, (15,10))
test_eq(im.resize_max(max_px=500, max_h=14, max_w=15).size, (15,10))
test_eq(im.resize_max(max_px=30*20/2, max_h=16, max_w=25).size, (15,10))
# +
#TODO function to resize_max all images in a path (optionally recursively) and save them somewhere (same relative dirs if recursive)
# -
# ## Basic types
# This section regroups the basic types used in vision with the transform that create objects of those types.
# +
# TODO: docs
# -
#export
def load_image(fn, mode=None, **kwargs):
"Open and load a `PIL.Image` and convert to `mode`"
im = Image.open(fn, **kwargs)
im.load()
im = im._new(im.im)
return im.convert(mode) if mode else im
#export
class PILBase(Image.Image, metaclass=BypassNewMeta):
default_dl_tfms = ByteToFloatTensor
_show_args = {'cmap':'viridis'}
_open_args = {'mode': 'RGB'}
@classmethod
def create(cls, fn, **kwargs)->None:
"Open an `Image` from path `fn`"
return cls(load_image(fn, **merge(cls._open_args, kwargs)))
def show(self, ctx=None, **kwargs):
"Show image using `merge(self._show_args, kwargs)`"
return show_image(self, ctx=ctx, **merge(self._show_args, kwargs))
#export
class PILImage(PILBase): pass
#export
class PILImageBW(PILImage): _show_args,_open_args = {'cmap':'Greys'},{'mode': 'L'}
im = PILImage.create(TEST_IMAGE)
test_eq(type(im), PILImage)
test_eq(im.mode, 'RGB')
im.resize((64,64))
ax = im.show(figsize=(1,1))
test_fig_exists(ax)
#export
class PILMask(PILBase): _open_args,_show_args = {'mode':'L'},{'alpha':0.5, 'cmap':'tab20'}
im = PILMask.create(TEST_IMAGE)
test_eq(type(im), PILMask)
test_eq(im.mode, 'L')
# ### Images
mnist = untar_data(URLs.MNIST_TINY)
fns = get_image_files(mnist)
mnist_fn = fns[0]; mnist_fn
timg = Transform(PILImageBW.create)
mnist_img = timg(mnist_fn)
test_eq(mnist_img.size, (28,28))
assert isinstance(mnist_img, PILImageBW)
mnist_img
# ### Segmentation masks
camvid = untar_data(URLs.CAMVID_TINY)
fns = get_image_files(camvid)
cam_fn = fns[0]
mask_fn = camvid/'labels'/f'{cam_fn.stem}_P{cam_fn.suffix}'
cam_img = timg(cam_fn)
test_eq(cam_img.size, (128,96))
tmask = Transform(PILMask.create)
mask = tmask(mask_fn)
test_eq(type(mask), PILMask)
test_eq(mask.size, (128,96))
_,axs = plt.subplots(1,3, figsize=(12,3))
cam_img.show(ctx=axs[0], title='image')
mask.show(alpha=1, ctx=axs[1], vmin=1, vmax=30, title='mask')
cam_img.show(ctx=axs[2], title='superimposed')
mask.show(ctx=axs[2], vmin=1, vmax=30);
# ### Points
# export
class TensorPoint(TensorBase):
"Basic type for points in an image"
_show_args = dict(s=10, marker='.', c='r')
@classmethod
def create(cls, t)->None:
"Convert an array or a list of points `t` to a `Tensor`"
return cls(tensor(t).view(-1, 2).float())
def show(self, ctx=None, **kwargs):
if 'figsize' in kwargs: del kwargs['figsize']
ctx.scatter(self[:, 0], self[:, 1], **{**self._show_args, **kwargs})
return ctx
# Points are expected to come as an array/tensor of shape `(n,2)` or as a list of lists with two elements. Unless you change the defaults in `PointScaler` (see later on), coordinates should go from 0 to width/height, with the first one being the column index (so from 0 to width) and the second one being the row index (so from 0 to height).
#
# > Note: This is differnt from the usual indeixing convention for arrays in numpy or in PyTorch, but it's the way points are expected by matplotlib or the internal functions in PyTorch like `F.grid_sample`.
pnt_img = TensorImage(mnist_img.resize((28,35)))
pnts = np.array([[0,0], [0,35], [28,0], [28,35], [9, 17]])
tfm = Transform(TensorPoint.create)
tpnts = tfm(pnts)
test_eq(tpnts.shape, [5,2])
test_eq(tpnts.dtype, torch.float32)
ctx = pnt_img.show(figsize=(1,1), cmap='Greys')
tpnts.show(ctx=ctx);
# ### Bounding boxes
# export
def get_annotations(fname, prefix=None):
"Open a COCO style json in `fname` and returns the lists of filenames (with maybe `prefix`) and labelled bboxes."
annot_dict = json.load(open(fname))
id2images, id2bboxes, id2cats = {}, collections.defaultdict(list), collections.defaultdict(list)
classes = {o['id']:o['name'] for o in annot_dict['categories']}
for o in annot_dict['annotations']:
bb = o['bbox']
id2bboxes[o['image_id']].append([bb[0],bb[1], bb[0]+bb[2], bb[1]+bb[3]])
id2cats[o['image_id']].append(classes[o['category_id']])
id2images = {o['id']:ifnone(prefix, '') + o['file_name'] for o in annot_dict['images'] if o['id'] in id2bboxes}
ids = list(id2images.keys())
return [id2images[k] for k in ids], [(id2bboxes[k], id2cats[k]) for k in ids]
#hide
#TODO explain and/or simplify this
coco = untar_data(URLs.COCO_TINY)
images, lbl_bbox = get_annotations(coco/'train.json')
annots = json.load(open(coco/'train.json'))
test_eq(images, [k['file_name'] for k in annots['images']])
for _ in range(5):
idx = random.randint(0, len(images)-1)
fn = images[idx]
i = 0
while annots['images'][i]['file_name'] != fn: i+=1
img_id = annots['images'][i]['id']
bbs = [ann for ann in annots['annotations'] if ann['image_id'] == img_id]
i2o = {k['id']:k['name'] for k in annots['categories']}
lbls = [i2o[bb['category_id']] for bb in bbs]
bboxes = [bb['bbox'] for bb in bbs]
bboxes = [[bb[0],bb[1], bb[0]+bb[2], bb[1]+bb[3]] for bb in bboxes]
test_eq(lbl_bbox[idx], [bboxes, lbls])
# +
# export
from matplotlib import patches, patheffects
def _draw_outline(o, lw):
o.set_path_effects([patheffects.Stroke(linewidth=lw, foreground='black'), patheffects.Normal()])
def _draw_rect(ax, b, color='white', text=None, text_size=14, hw=True, rev=False):
lx,ly,w,h = b
if rev: lx,ly,w,h = ly,lx,h,w
if not hw: w,h = w-lx,h-ly
patch = ax.add_patch(patches.Rectangle((lx,ly), w, h, fill=False, edgecolor=color, lw=2))
_draw_outline(patch, 4)
if text is not None:
patch = ax.text(lx,ly, text, verticalalignment='top', color=color, fontsize=text_size, weight='bold')
_draw_outline(patch,1)
# -
# export
class BBox(TupleBase):
"Basic type for a list of bounding boxes in an image"
def show(self, ctx=None, **kwargs):
for b,l in zip(self.bbox, self.lbl):
if l != '#bg': _draw_rect(ctx, b, hw=False, text=l)
return ctx
@classmethod
def create(cls, x): return cls(x)
bbox,lbl = add_props(lambda i,self: self[i])
# export
class TensorBBox(TupleBase):
"Basic type for a tensor of bounding boxes in an image"
@classmethod
def create(cls, x): return cls(tensor(x[0]).view(-1, 4).float(), x[1])
bbox,lbl = add_props(lambda i,self: self[i])
# Bounding boxes are expected to come as tuple with an array/tensor of shape `(n,4)` or as a list of lists with four elements adn a list of corresponding labels. Unless you change the defaults in `BBoxScaler` (see later on), coordinates for each bounding box should go from 0 to height/width, with the following convetion: top, left, bottom, right.
#
# > Note: We use the same convention as for points with y axis being before x.
coco = untar_data(URLs.COCO_TINY)
images, lbl_bbox = get_annotations(coco/'train.json')
idx=2
coco_fn,bbox = coco/'train'/images[idx],lbl_bbox[idx]
coco_img = timg(coco_fn)
tbbox = BBox(bbox)
ctx = coco_img.show(figsize=(3,3), cmap='Greys')
tbbox.show(ctx=ctx);
# ## Basic Transforms
# Unless specifically metioned, all the following transforms can be used as single-item transforms (in one of the list in the `tfms` you pass to a `TfmdDS` or a `Datasource`) or tuple transform (in the `tuple_tfms` you pass to a `TfmdDS` or a `Datasource`). The safest way that will work accross applications is to always use them as `tuple_tfms`. For instance, if you have points or bounding boxes as targets and use `ImageResizer` as a single-item transform, when you get to `PointScaler` or `BBoxScaler` (which are tuple transforms) you won't have the correct size of the image to properly scale your points.
class ImageResizer(Transform):
order=10
"Resize image to `size` using `resample"
def __init__(self, size, resample=Image.BILINEAR):
if not is_listy(size): size=(size,size)
self.size,self.resample = (size[1],size[0]),resample
def encodes(self, o:PILImage): return o.resize(size=self.size, resample=self.resample)
def encodes(self, o:PILMask): return o.resize(size=self.size, resample=Image.NEAREST)
# `size` can either be one integer (in which case images are resized to a square) or a tuple `height,width`.
#
# > Note: This is the usual convention for arrays or in PyTorch, but it's not the usual convention for PIL Image, which use the other way round.
# +
f = ImageResizer(14)
test_eq(f(mnist_img).size, (14,14))
test_eq(f(mask).size, (14,14))
f = ImageResizer((32,28))
test_eq(f(mnist_img).size, (28,32))#PIL has width first
test_eq(array(f(mnist_img)).shape, (32,28))#But numpy as height first and that is our convention
# -
# export
def image2byte(img):
"Transform image to byte tensor in `c*h*w` dim order."
res = torch.ByteTensor(torch.ByteStorage.from_buffer(img.tobytes()))
w,h = img.size
return res.view(h,w,-1).permute(2,0,1)
#export
@ToTensor
def encodes(self, o:PILImage): return TensorImage(image2byte(o))
@ToTensor
def encodes(self, o:PILImageBW): return TensorImageBW(image2byte(o))
@ToTensor
def encodes(self, o:PILMask): return TensorMask(image2byte(o)[0])
# Any data augmentation transform that runs on PIL Images must be run before this transform.
tfm = ToTensor()
test_eq(tfm(mnist_img).shape, (1,28,28))
test_eq(type(tfm(mnist_img)), TensorImageBW)
test_eq(tfm(mask).shape, (96,128))
test_eq(type(tfm(mask)), TensorMask)
# Let's confirm we can pipeline this with `PILImage.create`.
pipe_img = Pipeline([PILImageBW.create, ToTensor()])
img = pipe_img(mnist_fn)
pipe_img.show(img, figsize=(1,1));
def _cam_lbl(x): return mask_fn
cam_tds = TfmdDS([cam_fn], [[PILImage.create, ToTensor()], [_cam_lbl, PILMask.create, ToTensor()]])
cam_tds.show_at(0);
# +
#export
def _scale_pnts(x, y, do_scale=True,y_first=False):
if y_first: y = y.flip(1)
sz = [x.shape[-1], x.shape[-2]] if isinstance(x, Tensor) else x.size
return y * 2/tensor(sz).float() - 1 if do_scale else y
def _unscale_pnts(x, y):
sz = [x.shape[-1], x.shape[-2]] if isinstance(x, Tensor) else x.size
return (y+1) * tensor(sz).float()/2
# +
# export
#TODO: Transform on a whole tuple lose types, see if we can simplify that?
class PointScaler(ItemTransform):
"Scale a tensor representing points"
def __init__(self, do_scale=True, y_first=False): self.do_scale,self.y_first = do_scale,y_first
def encodes(self, o): return (o[0],TensorPoint(_scale_pnts(*o, self.do_scale, self.y_first)))
def decodes(self, o): return (o[0],TensorPoint(_unscale_pnts(*o)))
TensorPoint.default_ds_tfms = PointScaler
# -
# To work with data augmentation, and in particular the `grid_sample` method, points need to be represented with coordinates going from -1 to 1 (-1 being top or left, 1 bottom or right), which will be done unless you pass `do_scale=False`. We also need to make sure they are following our convention of points being x,y coordinates, so pass along `y_first=True` if you have your data in an y,x format to add a flip.
#
# > Warning: This transform needs to run on the tuple level, before any transform that changes the image size.
def _pnt_lbl(x): return TensorPoint.create(pnts)
def _pnt_open(fn): return PILImage(PILImage.create(fn).resize((28,35)))
pnt_tds = TfmdDS([mnist_fn], [_pnt_open, [_pnt_lbl]])
pnt_tdl = TfmdDL(pnt_tds, bs=1, after_item=[PointScaler(), ToTensor()])
x,y = pnt_tdl.one_batch()
#Scaling and flipping properly done
test_close(y[0], tensor([[-1., -1.], [-1., 1.], [1., -1.], [1., 1.], [9/14-1, 17/17.5-1]]))
a,b = pnt_tdl.decode_batch((x,y))[0]
test_eq(b, tensor(pnts).float())
#Check types
test_eq(type(x), TensorImage)
test_eq(type(y), TensorPoint)
test_eq(type(a), TensorImage)
test_eq(type(b), TensorPoint)
pnt_tdl.show_batch(figsize=(2,2), cmap='Greys');
# export
class BBoxScaler(PointScaler):
"Scale a tensor representing bounding boxes"
def encodes(self, o):
x,y = o
scaled_bb = _scale_pnts(x, y.bbox.view(-1,2), self.do_scale, self.y_first)
return (x,TensorBBox((scaled_bb.view(-1,4),y.lbl)))
def decodes(self, o):
x,y = o
scaled_bb = _unscale_pnts(x, y.bbox.view(-1,2))
return (x, TensorBBox((scaled_bb.view(-1,4), y.lbl)))
# +
# export
class BBoxCategorize(Transform):
"Reversible transform of category string to `vocab` id"
order,state_args=1,'vocab'
def __init__(self, vocab=None):
self.vocab = vocab
self.o2i = None if vocab is None else {v:k for k,v in enumerate(vocab)}
def setups(self, dsrc):
if not dsrc: return
dsrc = getattr(dsrc,'train',dsrc)
vals = set()
for bb in dsrc: vals = vals.union(set(bb.lbl))
self.vocab,self.otoi = uniqueify(list(vals), sort=True, bidir=True, start='#bg')
def encodes(self, o:BBox):
return TensorBBox.create((o.bbox,tensor([self.otoi[o_] for o_ in o.lbl if o_ in self.otoi])))
def decodes(self, o:TensorBBox):
return BBox((o.bbox,[self.vocab[i_] for i_ in o.lbl]))
BBox.default_type_tfms,BBox.default_ds_tfms = BBoxCategorize,BBoxScaler
# -
#export
#TODO tests
def bb_pad(samples, pad_idx=0):
"Function that collect `samples` of labelled bboxes and adds padding with `pad_idx`."
max_len = max([len(s[1][1]) for s in samples])
def _f(img,bbox,lbl):
bbox = torch.cat([bbox,bbox.new_zeros(max_len-bbox.shape[0], 4)])
lbl = torch.cat([lbl, lbl .new_zeros(max_len-lbl .shape[0])+pad_idx])
return img,TensorBBox((bbox,lbl))
return [_f(x,*y) for x,y in samples]
def _coco_lbl(x): return BBox(bbox)
tcat = BBoxCategorize()
coco_tds = TfmdDS([coco_fn], [PILImage.create, [_coco_lbl, tcat]])
coco_tdl = TfmdDL(coco_tds, bs=1, after_item=[BBoxScaler(), ToTensor()])
y
x,y = coco_tdl.one_batch()
y0 = y[0][0],y[1][0]
#Scaling and flipping properly done
test_close(y0[0], -1+tensor(bbox[0])/64)
test_eq(y0[1], tensor([1,1,1]))
a,b = coco_tdl.decode_batch((x,y))[0]
test_close(b[0], tensor(bbox[0]).float())
test_eq(b[1], bbox[1])
#Check types
test_eq(type(x), TensorImage)
test_eq(type(y), TensorBBox)
test_eq(type(a), TensorImage)
test_eq(type(b), BBox)
coco_tdl.show_batch();
# # Export -
#hide
from local.notebook.export import notebook2script
notebook2script(all_fs=True)
|
dev/07_vision_core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Submission Systems
# Submission system play an important role, if you want to develop your pygromos code. Many times, they are hidden in the Simulation_runner blocks. But maybe you want to develop something, where you need direct access on the submission system?
#
# This notebook will give you some examples, how you can use the submission systems.
# Note that all submission systems are write in the same ways, such you can exchange them quickly.
from pygromos.simulations.hpc_queuing.submission_systems import local # this executes your code in your local session.
from pygromos.simulations.hpc_queuing.submission_systems import lsf # this module can be used to submit to the lsf-queue (e.g. on euler)
from pygromos.simulations.hpc_queuing.submission_systems import dummy # this is a dummy system, that only prints the commands
# ## Local Submission
#
# This system executes the commands directly in your current session. This allows you to locally test or execute your code. Maybe if your process needs much more time, you want later to switch to a submission system for job-queueing.
sub_local = local.LOCAL()
sub_local.verbose = True
# +
bash_command = "sleep 2; echo \"WUHA\"; sleep 2"
job_id = sub_local.submit_to_queue(bash_command)
job_id
# -
#This is a dummy function, to not break the code!
sub_local.get_jobs_from_queue("FUN")
# ## LSF Submission
#
# The Lsf submission system allows to submit jobs to the IBM LSF-Queueing system.
#
# **Careful! This part requires a running LSF-Queueing System on your System**
#
# You can submit and kill jobs and arrays to the queue, as well as getting information from the queuing list.
# +
#Construct system:
sub_lsf = lsf.LSF(nmpi=1, job_duration = "24:00", max_storage=100)
sub_lsf.verbose = True
sub_lsf._refresh_job_queue_list_all_s = 0 #you must wait at least 1s to update job_queue list
# -
# ### Queue Checking:
sub_lsf.get_queued_jobs()
sub_lsf.job_queue_list
# ### Submission:
# here you can submit jobs to the queue as bash commands
# +
bash_command = "sleep 5; echo \"WUHA\"; sleep 2"
job_name = "Test1"
job_id = sub_lsf.submit_to_queue(command=bash_command, jobName=job_name)
# -
#search for the just submitted job in the queue
sub_lsf.search_queue_for_jobid(job_id)
sub_lsf.search_queue_for_jobname("Test1")
# ### Submitting multiple jobs
bash_command = "sleep 2; echo \"WUHA\"; sleep 2"
job_ids = []
for test in range(3):
job_name = "Test"+str(test)
job_id = sub_lsf.submit_to_queue(command=bash_command, jobName=job_name)
job_ids.append(job_id)
sub_lsf.search_queue_for_jobname("Te", regex=True)
# ### Killing a jobs
#
# Remove a job the job queue
sub_lsf.kill_jobs(job_ids=[job_id])
sub_lsf.search_queue_for_jobname("Te", regex=True)
|
examples/example_submission_systems.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mohammadmotiurrahman/mohammadmotiurrahman.github.io/blob/main/cse203/code/CSE203Lecture1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YGoUSIB4ESm_"
# ## Review of CSC 101 course taught at IUB .
#
# In order to run the code , click on the black and orange icon "Open in Colab" above. This will take you to Google Colaboratory. After that try the following key combination, Shift + Enter to run each cell.
#
#
# + [markdown] id="a8Ek3ghdYGxf"
# ## Basic Idea
# + [markdown] id="EJQl_gJNK6ig"
# So let us begin with the Hello World of CSC 101.
# + id="k8tTFQsXDkiC" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="89a6afe4-be74-4661-f1d6-7a61eb2c6aa8"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
int x = 6;
cout << "Hello World" << endl;
return 0;
}
# + id="6w2AoSydD_-e" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="20c3cf1b-6867-4360-8ab7-1bb708e6ba4a" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="3lesbP2vFnEQ"
# Let us move to something more interesting. Add two numbers.
# + id="UKYPa2dEFeJu" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2a7b7ba9-a9ec-4f91-e38f-c3d832d47bb6"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
int x = 6; int y = 5;
int z = x + y;
cout << "The summation is " << z << endl;
return 0;
}
# + id="yqhr6q7SHAiP" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="cace5228-bc75-465b-c3b6-1f361b83a732" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="ru736VuEHItD"
# Try to write a for-loop
# + id="sT3_IO6PHSA_" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="85faa027-042e-4238-c733-0a29a86b7403"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
for (int i = 0; i < 10; ++i) {
cout << 2 * i << " ";
}
return 0;
}
# + id="QPmWKpKQHiNw" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d9a7b7c4-6cd4-4bea-95c5-2a38dd195829" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="eBoUasp2INDJ"
# Write a while loop
# + id="0gjdCUfGIP5J" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="75e65585-f664-4d60-d823-77ebe4ab5fc1"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
int i = 0;
while (i < 10) {
cout << 2 * i << " ";
i = i + 1;
}
return 0;
}
# + id="B45fXIkMIdgu" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8d072157-6522-415b-d882-e4f555180106" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="vniJJkELI4Z8"
# Find whether a number is prime or not
# + id="JGiZr3b7I-YG" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4dc5a597-fc0a-4c32-f65e-cfc4f90190e4"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
bool isPrime = true;
int num = 7;
for (int i = 2; i < num ; ++i) {
if (num % i == 0) {
isPrime = false;
}
}
if (isPrime == false) {
cout << num << " is not Prime" << endl;
}
else {
cout << num << " is Prime" << endl;
}
return 0;
}
# + id="m8UuismrJclT" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f8e88797-b97f-4d09-fbc0-1e01e3e599cb" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="RC74eIxXYCGU"
# ## Functions
# + [markdown] id="SRI0Pd-hYg2w"
# Functions are components of a program that helps to modularize a program.
# Here is a function to print a list of odd numbers.
# + id="5OQigsLKZJHV" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="662f2acd-13f0-494b-cda8-81d89ceb77bb"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
void printOddNumbers(int x) {
for (int i = 0; i < x; ++i) {
if ( i % 2 ) {
cout << i << " ";
}
}
}
int main() {
printOddNumbers(10);
return 0;
}
# + id="tP1_RRLkaRIE" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="985177e8-6801-46cc-95a2-5721047e9135" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="2U5FHaBwatV3"
# Here is an function to print a list of prime numbers from 0 to 100.
# + id="5_Ry4zH6ay08" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="fb9e8ef3-fab4-4c30-f768-e362a89019a4"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
bool isPrimeNumber(int x) {
bool isPrime = true;
for (int i = 2; i < x ; ++i) {
if ( x % i == 0) {
isPrime = false;
}
}
return isPrime;
}
void printPrimeNumber(int x) {
for ( int i = 2 ; i < x ; ++i) {
// If a number i is a prime number
if (isPrimeNumber(i) == true) {
cout << i << " ";
}
}
}
int main() {
printPrimeNumber(100);
return 0;
}
# + id="5ytsO2oHaTn-" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="128de7b2-0b24-4b40-85a1-fa081138765a" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="umZs5HrBflur"
# Here is an another function to find LCM and HCF of given numbers
# + id="Rq1F1XJgf87Z" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="acb80c44-7499-4ea2-9f53-cea147ae326f"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int lcm(int a, int b) {
int x = 2;
while (!((x % a == 0) && (x % b == 0))) {
x = x + 1;
}
return x;
}
int hcf(int a, int b) {
return (a * b) / lcm(a, b);
}
int main() {
int a = 12; int b = 30;
cout << "LCM of " << a << " " << b << " is " << lcm(a, b) << endl;
cout << "HCF of " << a << " " << b << " is " << hcf(a, b) << endl;
return 0;
}
# + id="uETSzaZTg5cU" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="c5a7e4fb-c3fc-4a28-8a02-438577d867b4" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="AUF2bOKFor-a"
# ## Array
# + [markdown] id="T9PVHUAypaWu"
# Array is a collection of similar types of data
# + id="s9MDkrcxpesz" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="83e3d6aa-b410-438f-e28a-d8a30109acb0"
# %%writefile test.cpp
//This is where you will write your C++ code
#include <iostream>
using namespace std;
int main() {
int arr[10];
for (int i = 0; i < 10; ++i) {
arr[i] = i;
}
for (int i = 0; i < 10; ++i) {
cout << arr[i] << " ";
}
return 0;
}
# + id="LKm0lDRJq6Zd" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6e5d4a7e-93cf-4c82-d5b8-91a62217e739" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="oxIY_TCuvSKp"
# Something about ```const```
#
#
# + id="6kZSfpfRvcH8" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="221c23e6-fd15-4bff-c763-d452d34df8a5"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
//It is possible to change value of the variable a
int a = 30;
a = 10;
//It is not possible to change value of the variable a
//if the variable is change to const
const int a = 10;
a = 13;
return 0;
}
# + id="PsJjMV0ivk_N" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="71239502-e74b-4c8e-9c34-af1e72b26a36" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="-ukY5230xskY"
# While declaring an array in Visual Studio, the following norm is followed.
# + id="FavsoxACy2Fu" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a4f2ad0e-c784-4be5-9d0a-cf1e18f8f705"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
const int a = 5;
int arr[a];
for (int i = 0; i < a; ++i) {
arr[i] = i;
}
for (int i = 0; i < a; ++i ) {
cout << arr[i] << " ";
}
return 0;
}
# + id="HaZ32sMQzR5k" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6fe1eb63-6aef-4e8a-9220-a8dcb768a31e" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="QEUKFtl3ziXq"
# However, in compiler such as gcc, in which all of these codes are run such norms are not followed
# + id="DPOspxY8zyCN" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bb4d8d94-6652-446d-8512-590bfd0e7bf1"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
//Initially the size of the array has been set to 5
int a = 5;
int arr[a];
//However, it is possible to change value of the variable a
//in the middle of the code, which should lead to some
//unsavory problems. The first one is why should there be
//unnecessary spaces , if the value of a is less than the
//original. But more worrisome is when the value of a is
//more than the orignally allocated space. For example what
//if the value of a is 10 instead of 3.
a = 3;
for (int i = 0; i < a; ++i) {
arr[i] = i;
}
for (int i = 0; i < a; ++i ) {
cout << arr[i] << " ";
}
cout << endl;
a = 10;
for (int i = 0; i < a; ++i) {
arr[i] = i;
}
for (int i = 0; i < a; ++i ) {
cout << arr[i] << " ";
}
return 0;
}
# + id="KYZgBjdczzfd" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="af115079-8d40-4602-f820-9cfd499bbc1a" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="UJeOyM8o1nVB"
# Observe the 2nd line of the output, it should not happen. The initial array was designed to hold only 5 integer values. So when the value of the variable a was changed to 10, how can the array accomodate the extra 5 integer. At those moments the compiler steps into memory of other data structures which it shouldn't have the permission to access. Therefore it can easily be said that the compiler for Visual Studio is the best way to go. It ensures that the array data structure only access values that it was originally assigned to .
# + [markdown] id="UDT32EV__LG6"
# Somethings that want to use array for:
# 1. Average of numbers in an array
# + id="fMSmITql_3BD" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8fd93b30-b3f3-4f90-b13e-3eb15b2c29b7"
# %%writefile test.cpp
//Finding average of numbers in an array
#include <iostream>
using namespace std;
int main() {
//The size of the array has been set to 5
const int a = 5;
int arr[a] = {1, 2, 4, 5, 3};
float avg = 0;
for (int i = 0; i < a; ++i ) {
avg += arr[i];
}
cout << "The average of the values: " << avg / a << endl;
return 0;
}
# + id="Xi5aOGhyPs6E" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="423ac85a-d305-43fe-9d8f-68f151ed4d09" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="5uvZWmurQhPn"
# 2. Adding the value of two array's into a third array.
# + id="UWL9ir5DSTbS" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8c8748df-309e-4d19-f3f9-e4b133afd6b0"
# %%writefile test.cpp
//Adding the values of two array's and
//storing it in a third array
#include <iostream>
using namespace std;
int main() {
//The size of the array has been set to 5
const int a = 5;
int firstArray[a] = {1, 2, 4, 5, 3};
const int b = 5;
int secondArray[b] = {4, 5, 5, 6, 7};
const int c = 5;
int thirdArray[c] = {0, 0, 0, 0, 0};
for (int i = 0; i < a; ++i) {
thirdArray[i] = firstArray[i] + secondArray[i];
}
for (int i = 0; i < a; ++i) {
cout << thirdArray[i] << " ";
}
return 0;
}
# + id="yVhLK8exWrkn" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ba0043ec-2eec-4305-b43e-84b43baf9b88" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="_7h5MjpYW5_m"
# 3. Finding the freqeuncy of an element in an array
# + id="8fPA13yXZvWZ" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6a8835f6-4171-4373-cf1f-5144e75771e4"
# %%writefile test.cpp
//Finding the freqeuncy of a number
//in an array
#include <iostream>
using namespace std;
int main() {
//The size of the array has been set to 5
const int a = 5; int counter = 0;
int firstArray[a] = {1, 3, 3, 5, 3};
for (int i = 0; i < a; ++i) {
//Count the number of 3 in an array
if (firstArray[i] == 3) {
counter = counter + 1;
}
}
cout << "The number of 3 in the array is: " << counter << endl;
return 0;
}
# + id="8PaxlEgVba5m" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="269847e0-8cd9-452e-918f-70b39a63b619" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="Cpj8bRvOboMM"
# ## Two dimensional array
# So two dimensional array is another way to represent a lot of one dimensional arrays together. For example, if a student 4 subjects in the first semester, the final marks it can be represented in an array which contains 4 integers. If he takes another 4 subjects in the second semester his final marks can be represented in another array. What if we want to represent all the marks in the first semester and the second semester in an array. We should use two dimensional array.
# Two dimensional array is represented as : `int arr[row][col]`. The `row` is the number of semesters that the student has attended which is `2`. The `col` is the number of subjects that the student took in the semester which is `4`.
# While writing the code in Visual Studio, it is written like this:
# ```
# const int row = 2;
# const int col = 4;
# int arr[row][col];
# ```
# + id="hGQBfsN1buys" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3f21f6ec-dd6e-4cbb-be5e-c58f4968c167"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
int main() {
const int row = 2; const int col = 4;
int marks[row][col] = {
//Marks for 1st semester
{95, 80, 91, 100},
//Marks for 2nd semester
{100, 93, 90, 80}
};
for (int i = 0; i < row; ++i) {
cout << "Semester number: " << i + 1 << endl;
cout << "Final marks: ";
for (int j = 0; j < col; ++j) {
cout << marks[i][j] << " ";
}
cout << endl;
}
return 0;
}
# + id="AmJFGWbjxk_D" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="6c9b5094-0182-4020-b135-263f73e8ab60" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="jkKZoVSFhJsj"
# ### Matrix addition using two dimensional array
# So, a matrix can be a vector of dimension n x 1 or it can be a structure of dimension m x n. If it a vector of dimension n x 1 , it can be considered as an one dimensional array. Addition and subtraction of a vector will follow the same rules as addition and subtraction of a one dimensional array.
# Here I will show how addition and subtraction is done using two dimensional array.
# + id="GESwK5c9xopm" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c52a45cc-7541-44dd-e3c8-65cc27ac3c4a"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
int main() {
const int row = 2; const int col = 4;
int marks[row][col] = {
{95, 80, 91, 100},
{100, 93, 90, 80}
};
int extraAssignmentMarks[row][col] = {
{10, 5, 12, 8},
{0, 0, 0, 0}
};
int finalMarks[row][col];
//Add the marks
for (int i = 0; i < row; ++i) {
for (int j = 0; j < col; ++j) {
finalMarks[i][j] = marks[i][j] + extraAssignmentMarks[i][j];
}
}
//Display the marks
for (int i = 0; i < row; ++i) {
cout << "Semester number: " << i + 1 << endl;
cout << "Final grades: ";
for (int j = 0; j < col; ++j) {
cout << finalMarks[i][j] << " ";
} cout << endl;
}
return 0;
}
# + id="CVytnTqWt823" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="0b1fd6e6-f52c-427b-eaf9-86fa45c30036" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="AFnerxI50D9b"
# ### Matrix multiplication using two dimensional array
# 1. Matrix multiplication of matrices with the following dimensions m x m and m x m
# + id="U6vZxaFRlxYX" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="09f3332b-f2f9-4b5a-ddcd-e4afcd938f89"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
int main() {
const int row = 3; const int col = 3;
int arrA[row][col] = {
{10, 10, 10},
{20, 20, 20},
{30, 30, 30}
};
int arrB[row][col] = {
{10, 20, 30},
{10, 20, 30},
{10, 20, 30}
};
//This is where the multiplied value
//of the array will be kept
int arrC[row][col] = {
{0, 0, 0},
{0, 0, 0},
{0, 0, 0}
};
//Multiply the matrix
for (int i = 0; i < row; ++i) {
for (int j = 0; j < row; ++j) {
//Value of each row being multiplied
//by each column is being added
int val = 0;
for (int k = 0; k < row; ++k) {
val += arrA[i][k] * arrB[k][j];
}
//It is then stored in the proper
//index in arrC
arrC[i][j] = val;
}
}
//Display the marks
for (int i = 0; i < row; ++i) {
cout << "Row " << i << " : ";
for (int j = 0; j < col; ++j) {
cout << arrC[i][j] << " ";
} cout << endl;
}
return 0;
}
# + id="FUl26LWYl05K" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="88c8aa3c-f8f9-47f8-aa37-e24f83063b52" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="b5Fksilzl32X"
# 2. A more generic matrix multiplication is possible using when matrix has dimensions of the following size , m x a and a x n .
# + id="utkEjYtN2oI1" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b47cf536-784c-404f-d47f-f03a511a8894"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
int main() {
const int row1 = 2; const int col1 = 4;
int arrA[row1][col1] = {
{10, 10, 10, 10},
{20, 20, 20, 20}
};
const int row2 = 4; const int col2 = 2;
int arrB[row2][col2] = {
{10, 20},
{10, 20},
{10, 20},
{10, 20}
};
//This is where the multiplied value
//of the array will be kept
int arrC[row1][col2] = {
{0, 0},
{0, 0}
};
//Multiply the matrix
for (int i = 0; i < row1; ++i) {
for (int j = 0; j < col2; ++j) {
//Value of each row being multiplied
//by each column is being added
int val = 0;
for (int k = 0; k < col1; ++k) {
val += arrA[i][k] * arrB[k][j];
}
//It is then stored in the proper
//index in arrC
arrC[i][j] = val;
}
}
//Display the marks
for (int i = 0; i < row1; ++i) {
cout << "Row " << i << " : ";
for (int j = 0; j < col2; ++j) {
cout << arrC[i][j] << " ";
} cout << endl;
}
return 0;
}
# + id="z4UGCNPF2prA" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9cd8535f-52a0-456a-b4af-32da2c1cb381" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="jHvA5x_pqkW9"
# # Function with 1 dimensional array
# Functions as shown above is useful in the functions section were only used to modularize program with basic data types. Here functions will be used alongside with arrays to show data transfer with a program. Example is given below:
#
# * Printing an array
# + id="cQZBsZiWD4I6" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a30583f2-784a-44b2-88d9-13339b1f0952"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
void printArray(int arr[], int size) {
cout << "The array is: ";
for (int i = 0; i < size; ++i) {
cout << arr[i] << " ";
}
}
int main() {
const int N = 5;
int arr[N] = {1, 2, 3, 4, 5};
printArray(arr, N);
return 0;
}
# + id="Ckqk4PkMEGZS" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a25fd01d-896a-419d-86bf-315187121882" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="zNgPlju-J7MJ"
# * Print an array that contains odd numbers.
# + id="Zpl3d07-J0a3" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bbd5c4ae-b0ea-4d88-889b-0ef452c3a69b"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
void printOddNumArr(int arr[], int size) {
cout << "The odd numbers are: ";
for (int i = 0; i < size; ++i) {
if (arr[i] % 2 == 1) {
cout << arr[i] << " ";
}
}
}
int main() {
const int N = 5;
int arr[N] = {1, 2, 3, 4, 5};
printOddNumArr(arr, N);
return 0;
}
# + id="w44766cDKGbg" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f2dea6a1-cf1e-459d-cc95-ad828551217e" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="uGyJ7by-YKov"
# * Find the number of elements in an array that are less than zero.
# + id="cmLRqcErXJJ_" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6c56633c-31b8-4b42-8af2-3c2a37c19279"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
int numValLessThanZero(int arr[], int size) {
int count = 0;
for (int i = 0; i < size; ++i) {
if (arr[i] < 0) {
count = count + 1;
}
}
return count;
}
int main() {
const int N = 5;
int arr[N] = {2, 3, -12, 4, -1};
cout << "Number of value less than zeros is: " << numValLessThanZero(arr, N);
return 0;
}
# + id="b3tOSLaTZ_Ys" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="dbfc35aa-d0e2-442a-93b1-c889b3e52c49" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="BKkbLc7yLcfD"
# * Find the maximum number in an array.
# + id="VFhNRHf1KjYG" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="299ef971-40fe-4521-eb37-36d270668f90"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
int getMaxValue(int arr[], int size) {
for (int i = 0; i < size - 1; ++i) {
if (arr[i] > arr[i + 1]) {
int a = arr[i + 1];
arr[i + 1] = arr[i];
arr[i] = a;
}
}
return arr[size - 1];
}
int main() {
const int N = 5;
int arr[N] = {2, 3, 1, 4, -1};
cout << "Max value is: " << getMaxValue(arr, N);
return 0;
}
# + id="zZyly2q_Lntm" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8b1f0338-a7a8-409d-8101-1e0aeb772dc7" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="zhlVt6LCOOmO"
# So the code above needs a little bit of explanation.
#
# The array [2,3,1,4,-1] is passed from the main function to the function which has the name getMaxValue. The size of the array which is 5 is send along it.
# After that the following line of code runs on it.
# ```
# 1.for(int i = 0; i < size - 1; ++i){
# 2. if(arr[i] > arr[i+1]){
# 3. int a = arr[i+1];
# 4. arr[i+1] = arr[i];
# 5. arr[i] = a;
# 6. }
# 7. }
# ```
# The value of the variable `size` is `5`. However, it is not possible to access the value `arr[size]`, since indexing of an array starts from `0`. The maximum index that can be accessed is `4`. The `i < size` code snippet would ensure that effect. Howv, why is it `i < size - 1`. It is because the comparison of arr[i] and arr[i+1] would require us to access values which are not accessible. If value of `i` for example is `4` still less than `5`, will it be possible to access `i+1` which is `4+1 = 5` , no. So, `i` runs upto `size - 1`.
#
# In line 2 the comparison happens between `arr[i]` and `arr[i+1]` . If `arr[i]` is greater than `arr[i+1]` we need to do something. The thing we do is swapping. We swap value contained in `arr[i+1]` with the value of `arr[i]` , such that the value of `arr[i+1]` goes to `arr[i]` and the value of `arr[i]` goes to `arr[i+1]` . Line 3, Line 4 and Line 5 is responsible for the swapping. In Line 3 the value from `arr[i+1]` is stored into variable a. Remember value of `arr[i+1]` is less than `arr[i]`. In line 4 value of `arr[i]` which is greater than `arr[i+1]` is transferred to `arr[i+1]`. In Line 5 value of variable a which is smaller and was previously in `arr[i+1]` is transferred to `arr[i]`.
# This process continues till the 2nd last index of the array, swapping values of array, if one is smaller than the previous one. This allows for the maximum value to be transferred to the last index of the array.
#
# Lastly this piece of code return `arr[size-1]`; is necessary since we are only interested in returning the value of the last index of the array which is size-1( 5 - 1 = 4 in this case).
#
#
#
# + [markdown] id="vKtY_jY4iBUa"
# # Functions with 2 dimensional array
# 2 dimensional arrays can be utilized with functions in a similar way as 1 dimensional array with function. For example:
# * Printing the contents of the final marks for 2 semesters
# + id="oVzwQQnX3HTG" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d0e58898-a7df-458c-b6e3-044eb0f8d2f5"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
void printArray(int arr[][3], int row, int col) {
for (int i = 0; i < row; ++i) {
cout << "Semester: " << i + 1 << endl;
cout << "Final Marks: ";
for (int j = 0; j < col; ++j) {
cout << arr[i][j] << " ";
}
cout << endl;
}
}
int main() {
const int row = 2; const int col = 3;
int marks[row][col] = {
{90, 100, 90},
{100, 89, 90}
};
printArray(marks, row, col);
return 0;
}
# + id="cgCqCppq4wW7" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="5e15792f-aa00-4bbc-b651-68b93ab56320" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="bXDSkKun9BS2"
# Couple of things before jumping to the next example. Observe the parameters send to the function signature here: `void printArray(int arr[][3], int row, int col)`. There is a empty `[]` bracket after `arr`. If you any number out there it will not be a problem. Normally it is kept empty. Observe the second bracket of `arr`. It contains the value `3`. This signifies the number of columns that is present in the two dimensional array. That value should strictly be the value of the number of columns that is contained in the two dimensional array.
# + [markdown] id="tWrkmEHI52Iz"
# * Adding two matrices together using a function
# + id="3kAXpUfR6HX-" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d7394c38-f04d-48ea-c0d7-11f1e4c4c31e"
# %%writefile test.cpp
//Two dimensional array
#include <iostream>
using namespace std;
void printArray(int arrA[][3], int r1, int c1, int arrB[][3], int r2, int c2 ) {
for (int i = 0; i < r1; ++i) {
cout << "Semester: " << i + 1 << endl;
cout << "After adding bonus: ";
for (int j = 0; j < c1; ++j) {
cout << arrA[i][j] + arrB[i][j] << " ";
} cout << endl;
}
}
int main() {
const int row = 2; const int col = 3;
int marks[row][col] = {
{90, 100, 90},
{100, 89, 90}
};
int bonus[row][col] = {
{10, 4, 5},
{3, 5, 1}
};
printArray(marks, row, col, bonus, row, col);
return 0;
}
# + id="M3a4k7R38L2y" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="da11cf0f-accd-45e7-8858-4a22671ef7ab" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="D5os9A1X_KOy"
# # Return array from functions
# There are couple of ways in which array's can be returned from a function. Some of the ways that I have used in CSC 101 will be shown here.
# * There is no need to explicitly return arrays in C++. After calling the function with the array from the main function , if the array is printed again, the array changes its value. The value that the array takes, is due to the changes in the function which was called from the main function.
# + id="htgVI8W68MqI" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="17a2c35a-0139-4675-a019-7571b54a11bf"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
void addValueToArray(int arr[], int row) {
for (int i = 0; i < row; ++i) {
arr[i] = arr[i] + 10;
}
}
int main() {
const int row = 5 ;
int marks[row] = {89, 91, 91, 88, 90};
addValueToArray(marks, row);
cout << "Values in the array after calling the function: " << endl;
for (int i = 0 ; i < row; ++i) {
cout << marks[i] << " ";
}
return 0;
}
# + id="rXr3BBEWCtgr" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="16b6adbc-2c15-4b43-e7e9-b01b707f2889" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="toXC2jdQSdUE"
# One can modularize the code above further, but the idea will remain the same.
# + id="r1shrSNYSo1l" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0c1b458b-2180-4817-d85b-65225f0d3e3e"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
void addValueToArray(int arr[], int row) {
for (int i = 0; i < row; ++i) {
arr[i] = arr[i] + 10;
}
}
void printArray(int arr[], int row) {
for (int i = 0; i < row; ++i) {
cout << arr[i] << " ";
}
cout << endl;
}
void setValueToArray(int arr[], int row) {
for (int i = 0; i < row; ++i) {
arr[i] = 90 + i;
}
}
int main() {
const int row = 5 ;
int marks[row];
setValueToArray(marks, row);
cout << "Array content before: ";
printArray(marks, row);
//The value of the array changes
//due to the function below
addValueToArray(marks, row);
//The changes in the array is
//shown below
cout << "Array content after: ";
printArray(marks, row);
return 0;
}
# + id="o58gQCvfSrBm" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="90adae4d-834d-4e0c-df0e-94e3e3f8cbe8" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="1WF_E1dpVaQ3"
# Ideas presented in the above code block can be replicated for 2 dimensional arrays as well.
# + id="8Fnsbvg9TeNg" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="fb188a47-27bb-449c-f434-ad829ca01ca6"
# %%writefile test.cpp
//One dimensional array
#include <iostream>
using namespace std;
//Print the two dimensional array
void printArray(int arr[][3], int row, int col) {
for (int i = 0; i < row; ++i) {
for (int j = 0; j < col; ++j) {
cout << arr[i][j] << " ";
} cout << endl;
}
}
//Add bonus marks to the array
void addBonusToMarks(int marks[][3], int row1, int col1, int bonus[][3], int row2, int col2) {
for (int i = 0; i < row1; ++i) {
for (int j = 0 ; j < col1; ++j) {
marks[i][j] = marks[i][j] + bonus[i][j];
}
}
}
//Initialize the arrays
void initializeTheArrays(int marks[][3], int row1, int col1, int bonus[][3], int row2, int col2) {
for (int i = 0; i < row1; ++i) {
for (int j = 0; j < col1; ++j) {
marks[i][j] = 90 + i;
bonus[i][j] = 10 + i;
}
}
}
int main() {
const int row = 2 ; const int col = 3;
//Marks of the student in 2 semesters, each having 3 courses
int marks[row][col];
//Bonus marks of the students in each of the 2 semesters
//and each of the 3 courses
int bonus[row][col];
//Initialize the arrays with values
initializeTheArrays(marks, row, col, bonus, row, col);
cout << "Array content before adding bonus marks: " << endl;
printArray(marks, row, col);
//The value of the array changes due to the function below
addBonusToMarks(marks, row, col, bonus, row, col);
//The changes in the array is shown below
cout << "Array content after bonus marks are added: " << endl;
printArray(marks, row, col);
return 0;
}
# + id="2k9AHOia8C74" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="df52d964-b265-440d-8bed-91d63d373ac3" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="vkJnjfHyLj3a"
# # Memory
# So memory is an important topic in Computer Science. The examples that will be shown below hopefully illustrate that.
#
# So, let us begin with swapping numbers. For example in the code example below, after swapping values of variable `a` and `b` the values of the variable will change after swapping of the value.
# + id="bYVD2rAWN6JP" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="281627ba-1ce8-43a7-966c-14bfab677763"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Swapping of two numbers
int main() {
int a = 3; int b = 5;
cout << "Before swapping - a: " << a << " b: " << b << endl;
int temp = a;//Value of temp is 3 and value of a is 3
a = b; //Value of a is 5 and value of b is 5
b = temp; //Value of b is 3 because value of temp is 3
//So, printing the value of a and b after swapping is
cout << "After swapping - a: " << a << " b: " << b << endl;
return 0;
}
# + id="0a0oJNnGRoQi" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="7ac395c9-6bd8-4a12-9630-f75ee88fa2c8" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="zQQWaDPae_jo"
# Let us do swapping of the variable `a` and `b` using a function.
# + id="_GYii8WnclmG" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ff9c6f16-d785-4ead-8e0b-58b7c7439620"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Swapping of two numbers
void swapValues(int a , int b) {
int temp = a;//Value of temp is 3 and value of a is 3
a = b; //Value of a is 5 and value of b is 5
b = temp; //Value of b is 3 because value of temp is 3
}
int main() {
int a = 3; int b = 5;
cout << "Before swapping - a: " << a << " b: " << b << endl;
swapValues(a, b);
cout << "After swapping - a: " << a << " b: " << b << endl;
return 0;
}
# + id="12mWJYpjfPSE" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="212f4b4f-aea0-4f52-c7b1-ecf48255d647" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="0ipnUrn4fsKc"
# Look at the values of variables `a` and `b` after swapping using the function swapValues. The values of the variables `a` and `b` does not change. It is `3` and `5` even after the function is called. This is the result of scoping where the value of the function disappers after a function. If you rewrite the function like so ...
# ```
# void swapValues(int a , int b){
# cout<<"a: " << a << "b: " << b << endl;
# int temp = a;//Value of temp is 3 and value of a is 3
# a = b; //Value of a is 5 and value of b is 5
# b = temp; //Value of b is 3 because value of temp is 3
# cout<<"a: " << a << "b: " << b << endl;
# }
# ```
# ... you will see that the value actually changes inside the function.
#
# There are couple of ways to fix this issue with swapping.
# 1. Pass By Reference
# + id="WTazcPvpfqQ_" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8e4cb0ef-79e3-46f3-e96a-13b1b661fce3"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Swapping of two numbers
void swapValues(int &a , int &b) {
int temp = a;
a = b;
b = temp;
}
int main() {
int x = 3; int y = 5;
cout << "Before swapping - x: " << x << " y: " << y << endl;
swapValues(x, y);
cout << "After swapping - x: " << x << " y: " << y << endl;
return 0;
}
# + id="xZDsgTcz6t8p" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2874b83f-a47c-4458-d74f-aa6971921d50" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="kg6oopWI7GjP"
# 2. Pass by address
# + id="lHfgs3K17EAi" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3fea51cc-7a17-4968-ceea-c68a1b0be5e5"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Swapping of two numbers
void swapValues(int *a , int *b) {
int temp = *a;
*a = *b;
*b = temp;
}
int main() {
int x = 3; int y = 5;
cout << "Before swapping - x: " << x << " y: " << y << endl;
swapValues(&x, &y);
cout << "After swapping - x: " << x << " y: " << y << endl;
return 0;
}
# + id="OGKMtR0S8c00" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="5033cfd8-53e0-4054-baf3-1b96d70183d9" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="TcXLN0HDEgyh"
# In order to understand how pass by reference works one has to understand how memory works. So every variable in a program contains two things, an adress and a value. The address is where the variable lives in the RAM, and the value is what that variable contains in it.
#
# For example, in order to find the address of a variable, type something inside the main function like this:
#
#
# ```
# int a = 56;
# cout << a << endl; //Value of a
# cout << &a << endl; //Adress of a
# ```
# The number `56` which is transferred into `a` is the value of a amd `&a` is the address of `a`.
# The following program will illustrate it further.
#
# + id="eMd3q-AH8fEk" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="34bc861e-622f-44bb-84f8-478126d8d0a1"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
int a = 3;
cout << "The value of a is: " << a << endl;
cout << "The address of a is: " << &a << endl;
return 0;
}
# + id="6-TKXPDjId3K" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="ca04ba16-f4af-4ed4-d417-63df211bdaef" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="ASNBlwvlL8eo"
# Now the value of `&a` needs to be stored somewhere, something similar to how to values are stored in some variables. In the case of address of variables though, the addresses of variables are not stored little bit differently.
# The following way of storing is wrong:
# `int y = &a`. The correct way to store the address is `int *y = &a`. The `*` symbol denotes that `&a` is not a normal data type, instead it is of address data type, and it has to be stored in a different way.
# So, in order to find out the value of `a` and the address of `a` as it is stored in the variable `y`, one has to do the following:
# + id="0Nq4nxB1IekJ" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a2b52d40-59c1-4fd0-f7c3-2285c3488703"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
int a = 3;
int *y = &a;
cout << "y will give the address of the variable a" << endl;
cout << "y is : " << y << " and &a is : " << &a << endl;
cout << "*y will give the value of the variable a" << endl;
cout << "*y is : " << *y << " and a is : " << a << endl;
return 0;
}
# + id="B9tlk56ySZ-Y" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="175424f8-c3eb-4395-9e0f-cac0d25232cc" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="qtD7SVQZYwB_"
# One has to remember an important nomenclature while dealing with addresses of variables. In the following example:
# ```
# int x = 10;
# int *y = &x;
# cout << *y << endl; //Gets value of x
# cout << y << endl; //Gets address of x
#
# ```
# `*y` is known as dereferencing. There are couple of things to notice in the above code:
# ```
# int a = &x; //It is not allowed
# int y = 95 // It is also not allowed
# ```
#
# + [markdown] id="W4EC3SSBcJ5y"
# Pass by address as mentioned happens due to something known as aliasing.
# Here is an example how it works:
# + id="yKAii-laSbwY" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="644fdac7-2464-461e-cd12-349043f1957f"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
int a = 5;
/*
The line below replicates the passing of
the variable a to the parameter of the
function passByReference in the code block
for passByReference
*/
int &x = a;
/*
Here, when the above code is compiled, a very
important thing happens, the variable x and
a , shares the same value. So, if the following
is done to the value of x ...
*/
x = 56;
//The value of a also changes to 56
cout << "a is: " << a << endl;
cout << "x is: " << x << endl;
return 0;
}
# + id="HjPLiovcfhZl" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="a583a8f1-ea39-4338-e0c0-cda8bd9d8b46" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="tA-b4DgUk9dJ"
# Compare the above program with the program below:
# + id="eKSu14n-jhRq" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f7e60283-fc75-423d-9e31-b5feaa026a41"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
int a = 5;
int x = a;
x = 56;
cout << "a is: " << a << endl;
cout << "x is: " << x << endl;
return 0;
}
# + id="o_veu47BlJ5S" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="f05fb006-3a1a-4143-88b7-53eb4a263bad" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="QEZC8BGnlUZO"
# The value of the variable `a` does not change, only the value of the variable `x` change. Note this is a very important concept, we will revisit it in the later chapters.
# + [markdown] id="rO60wsjXmMe2"
# One of the important implications of pass by reference and pass by memory is that, it can used to return the value of more than one variable from a function.
# + id="Iz8BKSNNmkpI" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="935767aa-bfa3-4590-bc94-8b9803136b4f"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Using pass by reference
/*
Remember very carefully of "&" infront of "addition" and "multiplication"
This helps to relay the computation in the function addAndMult
to the main function.
*/
void addAndmult(int &addition, int &multiplication, int a , int b) {
addition = a + b; multiplication = a * b;
}
int main() {
int a = 5; int b = 6; int add = 0; int mult = 0;
addAndmult(add, mult, a, b);
cout << "After adding a and b: " << add << endl;
cout << "After multiplying a and b: " << mult << endl;
return 0;
}
# + id="fzFl5aokG-t_" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="fe1ec59e-7a1d-4c42-e4ca-6ec4e836dbae" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="4rdkgVZZHK1C"
# Another way of getting the same effect using pass by address concepts
# + id="9LNGr97wlKy7" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f154c78e-2b08-45b6-f4ae-158ed6e6070a"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Using pass by reference
/*
Remember very carefully of "&" infront of "addition" and "multiplication"
This helps to relay the computation in the function addAndMult
to the main function.
*/
void addAndmult(int *addition, int *multiplication, int a , int b) {
*addition = a + b; *multiplication = a * b;
}
int main() {
int a = 5; int b = 6; int add = 0; int mult = 0;
//Since this int *y = &x , therefore
addAndmult(&add, &mult, a, b);
cout << "After adding a and b: " << add << endl;
cout << "After multiplying a and b: " << mult << endl;
return 0;
}
# + id="5kQ4P53zHTf8" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="36b28d21-9e06-4ba2-c277-ba49e4e85163" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="5e2U7VL6KRam"
# These above two examples of "returning" more than one value from a C++ function.
# + [markdown] id="8thS0arXKqY2"
# # String
# In order to handle alphanumeric characters string is used in C++. Here only string class will be shown, array based implementation will be shown in more detail in the later chapters.
# + id="NAj-Yt_0Ieox" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="43f9f15f-8350-405f-bdeb-29f6724fe05f"
# %%writefile test.cpp
#include <iostream>
using namespace std;
int main() {
//So a string can be written something like this
string str = "hello today is 15th June";
//In order to find the length of a string, do something like
cout << "Length of the string is: " << str.length() << endl;
//You can iterate the string like so:
int len = str.length();
for (int i = 0; i < len; ++i) {
cout << str[i] << " ";
}
cout << endl;
//You can also find substring of a given string like this
int start = 5; int numOfChar = 6;
cout << "The substring is:" << str.substr(start, numOfChar);
return 0;
}
# + id="hKQh-zfIiCG9" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="c46fcaa7-624a-4de0-d7ef-4c8ec361f363" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="_HzLhfTAkNbQ"
# One of the application of pass by reference when it comes to string is given below:
# + id="-DgqJIrUiClv" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="719187cd-de41-47e3-a973-508a2ae44387"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Using pass by reference
/*
Count the number of vowels,consonants and space in a string
*/
void numOfVowConsSpace(string str, int &vowel, int &cons, int &space) {
int len = str.length();
for (int i = 0; i < len; ++i) {
if (str[i] == ' ') {
space = space + 1;
}
else if (str[i] == 'a' || str[i] == 'e' || str[i] == 'i' || str[i] == 'o' || str[i] == 'u') {
vowel = vowel + 1;
}
else {
cons = cons + 1;
}
}
}
int main() {
string str = "today is monday";
int vowel = 0; int cons = 0; int space = 0;
numOfVowConsSpace(str, vowel, cons, space);
cout << "Number of vowels : " << vowel << endl;
cout << "Number of consonant : " << cons << endl;
cout << "Number of space : " << space << endl;
return 0;
}
# + id="CAFnTrZJlzBi" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="84cd3568-e089-47a8-aef3-a9ef6e5153d2" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
#
# + [markdown] id="4lBwaqOJnMzj"
# One another interesting application of pass by reference is that the content of a string variable can change just like a variable of int, float or double while using aliasing.
#
# An example is shown below:
# + id="CXZGdRRsmb1x" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="05ad93ee-6dbc-4492-d393-235c00faf504"
# %%writefile test.cpp
#include <iostream>
using namespace std;
//Using pass by reference
/*
Count the number of vowels and consonants in a string
*/
void changeColor(string &color) {
color = "blue";
}
int main() {
string str = "red";
changeColor(str);
cout << "Color is now: " << str << endl;
return 0;
}
# + id="wBDmUcrMoPBa" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1b7f2efd-1e26-4169-e9ae-0f42efe80362" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
#
# + [markdown] id="5nJUjDbioSbw"
# There will be a comprehensive discussion about string and pass by reference in the coming lesson, for now appreciate that the strings can be changed using pass by reference, without any explicit return from a function.
# + [markdown] id="f7ii4wvPonYg"
# # File
# Writing to a file and reading to a file is important aspect of C++. It is important to read stored data from disk , as well write data to disk for later usage.
#
# **1. Reading a file from disk**
#
# Write something to a file named hello.txt
# + id="hfUMCkeU9YWU" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b17df61c-53a4-4f3c-e254-96f4815762da"
# %%writefile hello.txt
So, we meet again, after all these years.
It has been some time.
# + [markdown] id="oUrxE2_M-zcF"
# And then read from that text file
# + id="W4vWxdWWopgh" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9694c3ac-b002-40b9-c667-8960b5a5f410"
# %%writefile test.cpp
#include <iostream>
#include <fstream>
#include <string>
using namespace std;
int main() {
/*
Initialize the variable file with the filename
The file type is ifstream. ifstream is also
known as input file stream
*/
ifstream myfile("hello.txt");
//If the file is not open, do not do anything
if (!myfile.is_open()) {
cout << "Could not read" << endl;
return -1;
}
/*
Else read the file line by line from filestream
variable myfile to a string myStr
*/
string myStr;
while (getline(myfile, myStr)) {
cout << myStr << endl;
}
return 0;
}
# + id="Oz3FaVOI9u-B" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="16ac1cd9-6364-433e-ab80-ecc6951af1cb" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="AMkO13tvAEmU"
# **2. Writing a file into disk**
# + id="Or1IGKmH9xLJ" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b64c526d-1df7-43ec-ea00-fd0f0730acc1"
# %%writefile test.cpp
#include <iostream>
#include <fstream>
using namespace std;
int main() {
/*
Initialize the variable file with the filename
The file type is now ofstream. ofstream is also
known as output file stream
*/
ofstream myfile("hello.txt");
//If the file is not open, do not do anything
if (!myfile.is_open()) {
cout << "Could not read" << endl;
return -1;
}
/*
Else write something to the filestream
*/
myfile << "I see, it really has been sometime";
return 0;
}
# + id="3fmtRSwbBvTl" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="b5ezPXuuB4vi"
# **3. Appending text to a file**
#
#
# If you want to append to an existing file containing previous data.
# + id="eVjSMldgBxA-" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="de6b5105-bd8c-4228-d905-9560a10356a6"
# %%writefile test.cpp
#include <iostream>
#include <fstream>
using namespace std;
int main() {
/*
Initialize the variable file with the filename
The file type is now ofstream. ofstream is also
known as output file stream. If you want to append
to an exisiting file, you have to write like the
following...
*/
ofstream myfile("hello.txt", ios_base::out | ios_base::app);
//If the file is not open, do not do anything
if (!myfile.is_open()) {
cout << "Could not read" << endl;
return -1;
}
/*
Else write something to the filestream
*/
myfile << "I didn't realize that you will be waiting for me for so long";
return 0;
}
# + id="nl6NUyPjDNnA" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="s9LlhzokDRti"
# To test whether the file contains what you were expecting you can test the file reading program and see what is in the file `hello.txt`
# + [markdown] id="oYDibb9WGXBh"
# # Recursion
# This is one another way to write programs in C++. An example is given below:
# + id="e858kl-1JO9t" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="03db2924-7387-4785-bdae-a1654de19faa"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
for (int i = 0; i < num; ++i) {
cout << i << endl;
}
}
int main() {
int a = 5;
printNumber(a);
return 0;
}
# + id="FzaKfH2FLZ4m" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="740c1b70-04c1-4eb1-b75b-56a8003d468f" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="WdMo3vaiMCD1"
# It can be also be written like this:
# + id="85E2shMKL_S3" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2da2dd4a-f445-47b8-dd6d-ccafc42f2bd6"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
//What is the meaning of num-- , find it out
while (num--) {
cout << num << endl;
}
}
int main() {
int a = 5;
printNumber(a);
return 0;
}
# + id="j60El6kVMJUT" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="7c8d4995-5a2f-4f50-b34f-5d8d0de88042" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="hjyy6FzKNQGY"
# Something in the while loop can be done to make it more interesting.
# + id="AJISHIgeMUcw" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f98cd07f-ce2d-4f6e-bcde-18609c81ff61"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
//What is the meaning of num-- , find it out
while (num--) {
if (num == 5) return ;
else cout << num << endl;
}
}
int main() {
int a = 10;
printNumber(a);
return 0;
}
# + id="M6YD0kaVNsnC" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="bf402716-07c0-4970-cdf3-37511e64819f" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="-49vtgRhOSZ_"
# Finally it can be written be written like this:
# + id="0zs2wirSOGpA" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7e7eb72c-9e37-44ea-cdb9-753615235b84"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
if (num == 0) return ;
else cout << num << endl;
printNumber(--num);
}
int main() {
int a = 10;
printNumber(a);
return 0;
}
# + id="TOgnV60GO0WZ" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="124f335f-289f-4dff-cfba-56f81e32b318" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="05yqYzvuQMCI"
# Let us modify the code a little bit more.
# + id="n1hsB0RlP_uj" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f3e8dec4-01e0-445d-8fa6-eb16ade17c55"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
if (num == 0) return ;
else cout << num << endl;
num = num - 1;
printNumber(num);
}
int main() {
int a = 10;
printNumber(a);
return 0;
}
# + id="ySpp5v8uO2i3" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="51c7cbfc-18dc-4c68-b054-4d15f905982e" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="8XKuC6aNQ6LZ"
# What happens if the code is written like this?
# + id="ZxNud7-MQ1LR" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="14537977-4c56-4bf0-a46c-faede17baa33"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void printNumber(int num) {
if (num == 0) return ;
else cout << num << endl;
/*
Be careful before running this
Uncomment it before trying to run it
*/
//printNumber(num--);
}
int main() {
int a = 10;
printNumber(a);
return 0;
}
# + id="KPUfVQUcREAw" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6991b480-6320-4f77-8961-df800b7543a5" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="EYLbmbygRL-6"
# Anyhow the last code block that ran successfully is an example of recursive programming. Example of another program which can be written recursively is given below:
# + id="GylWcAIcVAn2" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9f0ab072-436a-4795-eb64-d17a054b43fb"
# %%writefile test.cpp
#include <iostream>
using namespace std;
void findEven(int i) {
if (i > 10)return ;
else if (i % 2 == 0) cout << i << endl;
findEven(++i);
}
int main() {
int a = 0;
findEven(a);
return 0;
}
# + id="V2WpygxDVWjM" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="4ad82dbf-29f9-4942-e31d-a390b4fe7c9d" magic_args="bash" language="script"
# g++ test.cpp -o test
# ./test
# + [markdown] id="r5YtqTTBVg00"
# More example of recursive programs will be shown in the upcoming chapters.
# + [markdown] id="4wxfP_aIVqOm"
# # Matrix transformation
# Topic such as linear transform, rotation, reflection of 2dimensional matrix will be written here.However , it is right now under construction.
#
# I will come back to it hopefully later.
# + id="EU9aYEqLVXSe"
|
cse203/code/CSE203Lecture1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="4JlLTP1Y-WHg"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="if-ujOZN-Par"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="Uq9kCbELjzgJ"
# # Efficient serving
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/recommenders/examples/efficient_serving"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/recommenders/blob/main/docs/examples/efficient_serving.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/recommenders/blob/main/docs/examples/efficient_serving.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/recommenders/docs/examples/efficient_serving.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="UlFcUNXT7hSF"
# [Retrieval models](https://www.tensorflow.org/recommenders/examples/basic_retrieval) are often built to surface a handful of top candidates out of millions or even hundreds of millions of candidates. To be able to react to the user's context and behaviour, they need to be able to do this on the fly, in a matter of milliseconds.
#
# Approximate nearest neighbour search (ANN) is the technology that makes this possible. In this tutorial, we'll show how to use ScaNN - a state of the art nearest neighbour retrieval package - to seamlessly scale TFRS retrieval to millions of items.
# + [markdown] id="Q_s_2UgUWA9u"
# ## What is ScaNN?
# + [markdown] id="GSvmiDQPsGmb"
# ScaNN is a library from Google Research that performs dense vector similarity search at large scale. Given a database of candidate embeddings, ScaNN indexes these embeddings in a manner that allows them to be rapidly searched at inference time. ScaNN uses state of the art vector compression techniques and carefully implemented algorithms to achieve the best speed-accuracy tradeoff. It can greatly outperform brute force search while sacrificing little in terms of accuracy.
# + [markdown] id="bTpnORU7WEPD"
# ## Building a ScaNN-powered model
# + [markdown] id="zXEZ3lZnWIVh"
# To try out ScaNN in TFRS, we'll build a simple MovieLens retrieval model, just as we did in the [basic retrieval](https://www.tensorflow.org/recommenders/examples/basic_retrieval) tutorial. If you have followed that tutorial, this section will be familiar and can safely be skipped.
#
# To start, install TFRS and TensorFlow Datasets:
# + id="mD2hiRviCxFE"
# #!pip install -q tensorflow-recommenders
# #!pip install -q --upgrade tensorflow-datasets
# + [markdown] id="oEbc-66nDJzc"
# We also need to install `scann`: it's an optional dependency of TFRS, and so needs to be installed separately.
# + id="daEivxsJDO0Y"
# #!pip install -q scann
# + [markdown] id="bDe054pgDQdp"
# Set up all the necessary imports.
# + id="6ekaJkcuHsiY"
from typing import Dict, Text
import os
import pprint
import tempfile
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
# + id="WdTPCz136mvc"
import tensorflow_recommenders as tfrs
# + [markdown] id="DfmRuUgJWlEQ"
# And load the data:
# + id="k-VF30hJn5-3"
# Load the MovieLens 100K data.
ratings = tfds.load(
"movielens/100k-ratings",
split="train"
)
# Get the ratings data.
ratings = (ratings
# Retain only the fields we need.
.map(lambda x: {"user_id": x["user_id"], "movie_title": x["movie_title"]})
# Cache for efficiency.
.cache(tempfile.NamedTemporaryFile().name)
)
# Get the movies data.
movies = tfds.load("movielens/100k-movies", split="train")
movies = (movies
# Retain only the fields we need.
.map(lambda x: x["movie_title"])
# Cache for efficiency.
.cache(tempfile.NamedTemporaryFile().name))
# + [markdown] id="SiVuNZ-lWv0R"
# Before we can build a model, we need to set up the user and movie vocabularies:
# + id="jw-iQKBBajnz"
user_ids = ratings.map(lambda x: x["user_id"])
unique_movie_titles = np.unique(np.concatenate(list(movies.batch(1000))))
unique_user_ids = np.unique(np.concatenate(list(user_ids.batch(1000))))
# + [markdown] id="yRbZCvWHWzPU"
# We'll also set up the training and test sets:
# + id="FqV8p7N8CrEg"
tf.random.set_seed(42)
shuffled = ratings.shuffle(100_000, seed=42, reshuffle_each_iteration=False)
train = shuffled.take(80_000)
test = shuffled.skip(80_000).take(20_000)
# + [markdown] id="Ok3-kzr1bI7U"
# ### Model definition
#
# Just as in the [basic retrieval](https://www.tensorflow.org/recommenders/examples/basic_retrieval) tutorial, we build a simple two-tower model.
# + id="yX_j4pEVbKIS"
class MovielensModel(tfrs.Model):
def __init__(self):
super().__init__()
embedding_dimension = 32
# Set up a model for representing movies.
self.movie_model = tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.StringLookup(
vocabulary=unique_movie_titles, mask_token=None),
# We add an additional embedding to account for unknown tokens.
tf.keras.layers.Embedding(len(unique_movie_titles) + 1, embedding_dimension)
])
# Set up a model for representing users.
self.user_model = tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.StringLookup(
vocabulary=unique_user_ids, mask_token=None),
# We add an additional embedding to account for unknown tokens.
tf.keras.layers.Embedding(len(unique_user_ids) + 1, embedding_dimension)
])
# Set up a task to optimize the model and compute metrics.
self.task = tfrs.tasks.Retrieval(
metrics=tfrs.metrics.FactorizedTopK(
candidates=movies.batch(128).cache().map(self.movie_model)
)
)
def compute_loss(self, features: Dict[Text, tf.Tensor], training=False) -> tf.Tensor:
# We pick out the user features and pass them into the user model.
user_embeddings = self.user_model(features["user_id"])
# And pick out the movie features and pass them into the movie model,
# getting embeddings back.
positive_movie_embeddings = self.movie_model(features["movie_title"])
# The task computes the loss and the metrics.
return self.task(user_embeddings, positive_movie_embeddings, compute_metrics=not training)
# + [markdown] id="JtO3lKR_XKkw"
# ### Fitting and evaluation
#
# A TFRS model is just a Keras model. We can compile it:
# + id="uOGTdwAAbuB6"
model = MovielensModel()
model.compile(optimizer=tf.keras.optimizers.Adagrad(learning_rate=0.1))
# + [markdown] id="4rGLyo-XXPmX"
# Estimate it:
# + id="uf_E4dIMcGnk"
model.fit(train.batch(8192), epochs=3)
# + [markdown] id="7xymbWgVXSrT"
# And evaluate it.
# + id="EMlIj741cIT8"
model.evaluate(test.batch(8192), return_dict=True)
# + [markdown] id="3RbHiBWqsFmf"
# ## Approximate prediction
#
# The most straightforward way of retrieving top candidates in response to a query is to do it via brute force: compute user-movie scores for all possible movies, sort them, and pick a couple of top recommendations.
#
# In TFRS, this is accomplished via the `BruteForce` layer:
# + id="x_L2yAPjpHsk"
brute_force = tfrs.layers.factorized_top_k.BruteForce(model.user_model)
brute_force.index(movies.batch(128).map(model.movie_model), movies)
# + [markdown] id="CzoNR28vXw7o"
# Once created and populated with candidates (via the `index` method), we can call it to get predictions out:
# + id="SBo1Nu0Grife"
# Get predictions for user 42.
_, titles = brute_force(np.array(["42"]), k=3)
print(f"Top recommendations: {titles[0]}")
# + [markdown] id="AzNECPifr6i6"
# On a small dataset of under 1000 movies, this is very fast:
# + id="w57iyu7Ir87Q"
# %timeit _, titles = brute_force(np.array(["42"]), k=3)
# + [markdown] id="u2AjJsdrsClR"
# But what happens if we have more candidates - millions instead of thousands?
#
# We can simulate this by indexing all of our movies multiple times:
# + id="AapJk84csTqV"
# Construct a dataset of movies that's 1,000 times larger. We
# do this by adding several million dummy movie titles to the dataset.
lots_of_movies = tf.data.Dataset.concatenate(
movies.batch(4096),
movies.batch(4096).repeat(1_000).map(lambda x: tf.zeros_like(x))
)
# We also add lots of dummy embeddings by randomly perturbing
# the estimated embeddings for real movies.
lots_of_movies_embeddings = tf.data.Dataset.concatenate(
movies.batch(4096).map(model.movie_model),
movies.batch(4096).repeat(1_000)
.map(lambda x: model.movie_model(x))
.map(lambda x: x * tf.random.uniform(tf.shape(x)))
)
# + [markdown] id="viCLP9qSYBQh"
# We can build a `BruteForce` index on this larger dataset:
# + id="mfY62oQbYA3Z"
brute_force_lots = tfrs.layers.factorized_top_k.BruteForce()
brute_force_lots.index(lots_of_movies_embeddings, lots_of_movies)
# + [markdown] id="OrkMt8O_xm-s"
# The recommendations are still the same
# + id="I9fIYUeYxjki"
_, titles = brute_force_lots(model.user_model(np.array(["42"])), k=3)
print(f"Top recommendations: {titles[0]}")
# + [markdown] id="wwF25ZzdseX8"
# But they take much longer. With a candidate set of 1 million movies, brute force prediction becomes quite slow:
# + id="oetK_wNxsdw0"
# %timeit _, titles = brute_force_lots(model.user_model(np.array(["42"])), k=3)
# + [markdown] id="mKF9yEeotbXQ"
# As the number of candidate grows, the amount of time needed grows linearly: with 10 million candidates, serving top candidates would take 250 milliseconds. This is clearly too slow for a live service.
#
# This is where approximate mechanisms come in.
#
# Using ScaNN in TFRS is accomplished via the `tfrs.layers.factorized_top_k.ScaNN` layer. It follow the same interface as the other top k layers:
# + id="SLgPmA90sbDL"
scann = tfrs.layers.factorized_top_k.ScaNN(num_reordering_candidates=1000)
scann.index(lots_of_movies_embeddings, lots_of_movies)
# + [markdown] id="qRI-qv7S2h97"
# The recommendations are (approximately!) the same
# + id="HCkRn1VnxuXn"
_, titles = scann(model.user_model(np.array(["42"])), k=3)
print(f"Top recommendations: {titles[0]}")
# + [markdown] id="iW1oBtcC2mb1"
# But they are much, much faster to compute:
# + id="ooJsLhpWstlf"
# %timeit _, titles = scann(model.user_model(np.array(["42"])), k=3)
# + [markdown] id="zOYk0zi12q-0"
# In this case, we can retrieve the top 3 movies out of a set of ~1 million in around 2 milliseconds: 15 times faster than by computing the best candidates via brute force. The advantage of approximate methods grows even larger for larger datasets.
# + [markdown] id="tE7eL7ZzDKtl"
# ## Evaluating the approximation
#
# When using approximate top K retrieval mechanisms (such as ScaNN), speed of retrieval often comes at the expense of accuracy. To understand this trade-off, it's important to measure the model's evaluation metrics when using ScaNN, and to compare them with the baseline.
#
# Fortunately, TFRS makes this easy. We simply override the metrics on the retrieval task with metrics using ScaNN, re-compile the model, and run evaluation.
#
# To make the comparison, let's first run baseline results. We still need to override our metrics to make sure they are using the enlarged candidate set rather than the original set of movies:
# + id="ZZtJRQqBep_5"
# Override the existing streaming candidate source.
model.task.factorized_metrics = tfrs.metrics.FactorizedTopK(
candidates=lots_of_movies_embeddings
)
# Need to recompile the model for the changes to take effect.
model.compile()
# %time baseline_result = model.evaluate(test.batch(8192), return_dict=True, verbose=False)
# + [markdown] id="HHFzcS5cQtB_"
# We can do the same using ScaNN:
# + id="5T-YxOqoKMje"
model.task.factorized_metrics = tfrs.metrics.FactorizedTopK(
candidates=scann
)
model.compile()
# We can use a much bigger batch size here because ScaNN evaluation
# is more memory efficient.
# %time scann_result = model.evaluate(test.batch(8192), return_dict=True, verbose=False)
# + [markdown] id="Y0gnjcUUZ6-v"
# ScaNN based evaluation is much, much quicker: it's over ten times faster! This advantage is going to grow even larger for bigger datasets, and so for large datasets it may be prudent to always run ScaNN-based evaluation to improve model development velocity.
#
# But how about the results? Fortunately, in this case the results are almost the same:
#
# + id="gcXUbZx3Fq4f"
print(f"Brute force top-100 accuracy: {baseline_result['factorized_top_k/top_100_categorical_accuracy']:.2f}")
print(f"ScaNN top-100 accuracy: {scann_result['factorized_top_k/top_100_categorical_accuracy']:.2f}")
# + [markdown] id="d2UJR-5nZ6YT"
# This suggests that on this artificial datase, there is little loss from the approximation. In general, all approximate methods exhibit speed-accuracy tradeoffs. To understand this in more depth you can check out <NAME>'s [ANN benchmarks](https://github.com/erikbern/ann-benchmarks).
# + [markdown] id="-jdPPOlV3JOr"
# ## Deploying the approximate model
#
# The `ScaNN`-based model is fully integrated into TensorFlow models, and serving it is as easy as serving any other TensorFlow model.
#
# We can save it as a `SavedModel` object
# + id="KnVI_6N53WU5"
# We re-index the ScaNN layer to include the user embeddings in the same model.
# This way we can give the saved model raw features and get valid predictions
# back.
scann = tfrs.layers.factorized_top_k.ScaNN(model.user_model, num_reordering_candidates=1000)
scann.index(lots_of_movies_embeddings, lots_of_movies)
# Need to call it to set the shapes.
_ = scann(np.array(["42"]))
with tempfile.TemporaryDirectory() as tmp:
path = os.path.join(tmp, "model")
scann.save(
path,
options=tf.saved_model.SaveOptions(namespace_whitelist=["Scann"])
)
loaded = tf.keras.models.load_model(path)
# + [markdown] id="O5vDZjro4lXG"
# and then load it and serve, getting exactly the same results back:
# + id="TXm8smCt3iFB"
_, titles = loaded(tf.constant(["42"]))
print(f"Top recommendations: {titles[0][:3]}")
# + [markdown] id="S0Doal2ETqU4"
# The resulting model can be served in any Python service that has TensorFlow and ScaNN installed.
#
# It can also be served using a customized version of TensorFlow Serving, available as a Docker container on [Docker Hub](https://hub.docker.com/r/google/tf-serving-scann). You can also build the image yourself from the [Dockerfile](https://github.com/google-research/google-research/tree/master/scann/tf_serving).
# + [markdown] id="0gQsvn5PYbR-"
# ## Tuning ScaNN
# + [markdown] id="918uqacB7sNH"
# Now let's look into tuning our ScaNN layer to get a better performance/accuracy tradeoff. In order to do this effectively, we first need to measure our baseline performance and accuracy.
#
# From above, we already have a measurement of our model's latency for processing a single (non-batched) query (although note that a fair amount of this latency is from non-ScaNN components of the model).
#
# Now we need to investigate ScaNN's accuracy, which we measure through recall. A recall@k of x% means that if we use brute force to retrieve the true top k neighbors, and compare those results to using ScaNN to also retrieve the top k neighbors, x% of ScaNN's results are in the true brute force results. Let's compute the recall for the current ScaNN searcher.
#
# First, we need to generate the brute force, ground truth top-k:
# + id="qgf_QuP-8EXb"
# Process queries in groups of 1000; processing them all at once with brute force
# may lead to out-of-memory errors, because processing a batch of q queries against
# a size-n dataset takes O(nq) space with brute force.
titles_ground_truth = tf.concat([
brute_force_lots(queries, k=10)[1] for queries in
test.batch(1000).map(lambda x: model.user_model(x["user_id"]))
], axis=0)
# + [markdown] id="LSZkWESc856P"
# Our variable `titles_ground_truth` now contains the top-10 movie recommendations returned by brute-force retrieval. Now we can compute the same recommendations when using ScaNN:
# + id="yUKtdf1X87mP"
# Get all user_id's as a 1d tensor of strings
test_flat = np.concatenate(list(test.map(lambda x: x["user_id"]).batch(1000).as_numpy_iterator()), axis=0)
# ScaNN is much more memory efficient and has no problem processing the whole
# batch of 20000 queries at once.
_, titles = scann(test_flat, k=10)
# + [markdown] id="JTsTDiAZ9F6h"
# Next, we define our function that computes recall. For each query, it counts how many results are in the intersection of the brute force and the ScaNN results and divides this by the number of brute force results. The average of this quantity over all queries is our recall.
# + id="PCtBew2C9Gv0"
def compute_recall(ground_truth, approx_results):
return np.mean([
len(np.intersect1d(truth, approx)) / len(truth)
for truth, approx in zip(ground_truth, approx_results)
])
# + [markdown] id="_tdxlKua9JR2"
# This gives us baseline recall@10 with the current ScaNN config:
# + id="nMi4VtJD9K9P"
print(f"Recall: {compute_recall(titles_ground_truth, titles):.3f}")
# + [markdown] id="gKpgkNseYWW8"
# We can also measure the baseline latency:
# + id="81mO-GS4VJLJ"
# %timeit -n 1000 scann(np.array(["42"]), k=10)
# + [markdown] id="UICnYQln9PAq"
# Let's see if we can do better!
#
# To do this, we need a model of how ScaNN's tuning knobs affect performance. Our current model uses ScaNN's tree-AH algorithm. This algorithm partitions the database of embeddings (the "tree") and then scores the most promising of these partitions using AH, which is a highly optimized approximate distance computation routine.
#
# The default parameters for TensorFlow Recommenders' ScaNN Keras layer sets `num_leaves=100` and `num_leaves_to_search=10`. This means our database is partitioned into 100 disjoint subsets, and the 10 most promising of these partitions is scored with AH. This means 10/100=10% of the dataset is being searched with AH.
#
# If we have, say, `num_leaves=1000` and `num_leaves_to_search=100`, we would also be searching 10% of the database with AH. However, in comparison to the previous setting, the 10% we would search will contain higher-quality candidates, because a higher `num_leaves` allows us to make finer-grained decisions about what parts of the dataset are worth searching.
#
# It's no surprise then that with `num_leaves=1000` and `num_leaves_to_search=100` we get significantly higher recall:
# + id="vq6L1Qtl9Qan"
scann2 = tfrs.layers.factorized_top_k.ScaNN(
model.user_model,
num_leaves=1000,
num_leaves_to_search=100,
num_reordering_candidates=1000)
scann2.index(lots_of_movies_embeddings, lots_of_movies)
_, titles2 = scann2(test_flat, k=10)
print(f"Recall: {compute_recall(titles_ground_truth, titles2):.3f}")
# + [markdown] id="G2WR8zPH9TtW"
# However, as a tradeoff, our latency has also increased. This is because the partitioning step has gotten more expensive; `scann` picks the top 10 of 100 partitions while `scann2` picks the top 100 of 1000 partitions. The latter can be more expensive because it involves looking at 10 times as many partitions.
# + id="Po0kb4Mf9VhX"
# %timeit -n 1000 scann2(np.array(["42"]), k=10)
# + [markdown] id="fCDzY0sc9Zgc"
# In general, tuning ScaNN search is about picking the right tradeoffs. Each individual parameter change generally won't make search both faster and more accurate; our goal is to tune the parameters to optimally trade off between these two conflicting goals.
#
# In our case, `scann2` significantly improved recall over `scann` at some cost in latency. Can we dial back some other knobs to cut down on latency, while preserving most of our recall advantage?
#
# Let's try searching 70/1000=7% of the dataset with AH, and only rescoring the final 400 candidates:
# + id="jBp8Yvdj9pMQ"
scann3 = tfrs.layers.factorized_top_k.ScaNN(
model.user_model,
num_leaves=1000,
num_leaves_to_search=70,
num_reordering_candidates=400)
scann3.index(lots_of_movies_embeddings, lots_of_movies)
_, titles3 = scann3(test_flat, k=10)
print(f"Recall: {compute_recall(titles_ground_truth, titles3):.3f}")
# + [markdown] id="3Isgpm7b9rgE"
# `scann3` delivers about a 3% absolute recall gain over `scann` while also delivering lower latency:
# + id="JiDEWwtr9sKG"
# %timeit -n 1000 scann3(np.array(["42"]), k=10)
# + [markdown] id="NwWKyQgt9uh1"
# These knobs can be further adjusted to optimize for different points along the accuracy-performance pareto frontier. ScaNN's algorithms can achieve state-of-the-art performance over a wide range of recall targets.
# + [markdown] id="UvlCsKyFU40k"
# ## Further reading
# + [markdown] id="0ikGqmNa9yRG"
# ScaNN uses advanced vector quantization techniques and highly optimized implementation to achieve its results. The field of vector quantization has a rich history with a variety of approaches. ScaNN's current quantization technique is detailed in [this paper](https://arxiv.org/abs/1908.10396), published at ICML 2020. The paper was also released along with [this blog article](https://ai.googleblog.com/2020/07/announcing-scann-efficient-vector.html) which gives a high level overview of our technique.
#
# Many related quantization techniques are mentioned in the references of our ICML 2020 paper, and other ScaNN-related research is listed at http://sanjivk.com/.
|
src/recommenders/efficient_serving.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pyspark
from operator import add
sc = pyspark.SparkContext()
# +
textFile = sc.textFile("data/shakespeare.txt").filter(lambda l: len(l) > 0)
words=textFile.flatMap(lambda l: l.split())
counts = words.map(lambda w: (w, 1)).reduceByKey(lambda x,y: x+y).map(lambda x: (x[1],x[0])).sortByKey(False) #(add)
# -
counts.collect()
sc
|
PySparkDemo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python Multi
# language: python
# name: multi
# ---
import feedparser
import MySQLdb
# 데이터베이스 연결하기
connection = MySQLdb.connect(
user="root",
passwd="<PASSWORD>",
host="localhost",
db="tip",
charset="utf8")
# 커서 생성하기
cursor = connection.cursor()
# +
# 실행할 때마다 같은 레코드가 중복되어 들어가지 않게 테이블을 제거해두기
#cursor.execute("DROP TABLE IF EXISTS books")
# 테이블 생성하기
cursor.execute("CREATE TABLE books (title text, url text)")
# -
# URL을 지정해서 FeedParserDict 객체 생성하기
rss = feedparser.parse("http://www.aladin.co.kr/rss/special_new/351")
# RSS 버전 확인하기
print(rss.version)
# 피드의 제목
print(rss["feed"]["title"])
# 반복 적용
for content in rss["entries"]:
# 데이터 저장하기
cursor.execute("INSERT INTO books VALUES(%s, %s)", (content["title"], content["link"]))
# +
# 커밋하기
connection.commit()
# 연결 종료하기
connection.close()
# -
|
Web_Crawling/python-crawler/chapter_4/mysql-rss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: ipykernel_py2
# ---
# ## Comparison Operators
# *Suggested Answers follow (usually there are multiple ways to solve a problem in Python).*
# Verify that 25 is smaller than 30.
25 < 30
# Verify that 5 multiplied by 3 is less than or equal to 5 to the power of 3.
5 * 3 <= 5 ** 3
# Verify that 100 is equal to 10 square.
100 == 10 ** 2
# Verify that 53 is not equal to 46.
53 != 46
|
11 - Introduction to Python/4_More on Operators/1_Comparison Operators (2:10)/Comparison Operators - Solution_Py2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# # Using Azure Machine Learning Pipelines for Batch Inference for CSV Files
#
# In this notebook, we will demonstrate how to make predictions on large quantities of data asynchronously using the ML pipelines with Azure Machine Learning. Batch inference (or batch scoring) provides cost-effective inference, with unparalleled throughput for asynchronous applications. Batch prediction pipelines can scale to perform inference on terabytes of production data. Batch prediction is optimized for high throughput, fire-and-forget predictions for a large collection of data.
#
# > **Tip**
# If your system requires low-latency processing (to process a single document or small set of documents quickly), use [real-time scoring](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-consume-web-service) instead of batch prediction.
#
# In this example we will take use a machine learning model already trained to predict different types of iris flowers and run that trained model on some of the data in a CSV file which has characteristics of different iris flowers. However, the same example can be extended to manipulating data to any embarrassingly-parallel processing through a python script.
#
# The outline of this notebook is as follows:
#
# - Create a DataStore referencing the CSV files stored in a blob container.
# - Register the pretrained model into the model registry.
# - Use the registered model to do batch inference on the CSV files in the data blob container.
#
# ## Prerequisites
# If you are using an Azure Machine Learning Notebook VM, you are all set. Otherwise, make sure you go through the configuration Notebook located at https://github.com/Azure/MachineLearningNotebooks first. This sets you up with a working config file that has information on your workspace, subscription id, etc.
#
# 
# ### Connect to workspace
# Create a workspace object from the existing workspace. Workspace.from_config() reads the file config.json and loads the details into an object named ws.
# +
from azureml.core import Workspace
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# -
# ### Create or Attach existing compute resource
# By using Azure Machine Learning Compute, a managed service, data scientists can train machine learning models on clusters of Azure virtual machines. Examples include VMs with GPU support. In this tutorial, you create Azure Machine Learning Compute as your training environment. The code below creates the compute clusters for you if they don't already exist in your workspace.
#
# **Creation of compute takes approximately 5 minutes. If the AmlCompute with that name is already in your workspace the code will skip the creation process.**
# +
import os
from azureml.core.compute import AmlCompute, ComputeTarget
from azureml.core.compute_target import ComputeTargetException
# choose a name for your cluster
compute_name = os.environ.get("AML_COMPUTE_CLUSTER_NAME", "cpu-cluster")
compute_min_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MIN_NODES", 0)
compute_max_nodes = os.environ.get("AML_COMPUTE_CLUSTER_MAX_NODES", 4)
# This example uses CPU VM. For using GPU VM, set SKU to STANDARD_NC6
vm_size = os.environ.get("AML_COMPUTE_CLUSTER_SKU", "STANDARD_D2_V2")
if compute_name in ws.compute_targets:
compute_target = ws.compute_targets[compute_name]
if compute_target and type(compute_target) is AmlCompute:
print('found compute target. just use it. ' + compute_name)
else:
print('creating a new compute target...')
provisioning_config = AmlCompute.provisioning_configuration(vm_size = vm_size,
min_nodes = compute_min_nodes,
max_nodes = compute_max_nodes)
# create the cluster
compute_target = ComputeTarget.create(ws, compute_name, provisioning_config)
# can poll for a minimum number of nodes and for a specific timeout.
# if no min node count is provided it will use the scale settings for the cluster
compute_target.wait_for_completion(show_output=True, min_node_count=None, timeout_in_minutes=20)
# For a more detailed view of current AmlCompute status, use get_status()
print(compute_target.get_status().serialize())
# -
# ### Create a datastore containing sample images
# The input dataset used for this notebook is CSV data which has attributes of different iris flowers. We have created a public blob container `sampledata` on an account named `pipelinedata`, containing iris data set. In the next step, we create a datastore with the name `iris_datastore`, which points to this container. In the call to `register_azure_blob_container` below, setting the `overwrite` flag to `True` overwrites any datastore that was created previously with that name.
#
# This step can be changed to point to your blob container by providing your own `datastore_name`, `container_name`, and `account_name`.
# +
from azureml.core.datastore import Datastore
account_name = "pipelinedata"
datastore_name="iris_datastore_data"
container_name="sampledata"
iris_data = Datastore.register_azure_blob_container(ws,
datastore_name=datastore_name,
container_name= container_name,
account_name=account_name,
overwrite=True)
# -
# ### Create a TabularDataset
# A [TabularDataSet](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.data.tabulardataset?view=azure-ml-py) references single or multiple files which contain data in a tabular structure (ie like CSV files) in your datastores or public urls. TabularDatasets provides you with the ability to download or mount the files to your compute. By creating a dataset, you create a reference to the data source location. If you applied any subsetting transformations to the dataset, they will be stored in the dataset as well. The data remains in its existing location, so no extra storage cost is incurred.
# +
from azureml.core.dataset import Dataset
iris_ds_name = 'iris_data'
path_on_datastore = iris_data.path('iris/')
input_iris_ds = Dataset.Tabular.from_delimited_files(path=path_on_datastore, validate=False)
registered_iris_ds = input_iris_ds.register(ws, iris_ds_name, create_new_version=True)
named_iris_ds = registered_iris_ds.as_named_input(iris_ds_name)
# -
# ### Intermediate/Output Data
# Intermediate data (or output of a Step) is represented by [PipelineData](https://docs.microsoft.com/en-us/python/api/azureml-pipeline-core/azureml.pipeline.core.pipelinedata?view=azure-ml-py) object. PipelineData can be produced by one step and consumed in another step by providing the PipelineData object as an output of one step and the input of one or more steps.
#
# **Constructing PipelineData**
# - name: [Required] Name of the data item within the pipeline graph
# - datastore_name: Name of the Datastore to write this output to
# - output_name: Name of the output
# - output_mode: Specifies "upload" or "mount" modes for producing output (default: mount)
# - output_path_on_compute: For "upload" mode, the path to which the module writes this output during execution
# - output_overwrite: Flag to overwrite pre-existing data
# +
from azureml.pipeline.core import PipelineData
datastore = ws.get_default_datastore()
output_folder = PipelineData(name='inferences', datastore=datastore)
# -
# ## Registering the Model with the Workspace
# Get the pretrained model from a publicly available Azure Blob container, then register it to use in your workspace
# +
model_container_name="iris-model"
model_datastore_name="iris_model_datastore"
model_datastore = Datastore.register_azure_blob_container(ws,
datastore_name=model_datastore_name,
container_name= model_container_name,
account_name=account_name,
overwrite=True)
# +
from azureml.core.model import Model
model_datastore.download('iris_model.pkl')
# register downloaded model
model = Model.register(model_path = "iris_model.pkl/iris_model.pkl",
model_name = "iris", # this is the name the model is registered as
tags = {'pretrained': "iris"},
workspace = ws)
# -
# ### Using your model to make batch predictions
# To use the model to make batch predictions, you need an **entry script** and a list of **dependencies**:
#
# #### An entry script
# This script accepts requests, scores the requests by using the model, and returns the results.
# - __init()__ - Typically this function loads the model into a global object. This function is run only once at the start of batch processing per worker node/process. init method can make use of following environment variables (ParallelRunStep input):
# 1. AZUREML_BI_OUTPUT_PATH – output folder path
# - __run(mini_batch)__ - The method to be parallelized. Each invocation will have one minibatch.<BR>
# __mini_batch__: Batch inference will invoke run method and pass either a list or Pandas DataFrame as an argument to the method. Each entry in min_batch will be - a filepath if input is a FileDataset, a Pandas DataFrame if input is a TabularDataset.<BR>
# __run__ method response: run() method should return a Pandas DataFrame or an array. For append_row output_action, these returned elements are appended into the common output file. For summary_only, the contents of the elements are ignored. For all output actions, each returned output element indicates one successful inference of input element in the input mini-batch.
# User should make sure that enough data is included in inference result to map input to inference. Inference output will be written in output file and not guaranteed to be in order, user should use some key in the output to map it to input.
#
#
# #### Dependencies
# Helper scripts or Python/Conda packages required to run the entry script or model.
#
# The deployment configuration for the compute target that hosts the deployed model. This configuration describes things like memory and CPU requirements needed to run the model.
#
# These items are encapsulated into an inference configuration and a deployment configuration. The inference configuration references the entry script and other dependencies. You define these configurations programmatically when you use the SDK to perform the deployment. You define them in JSON files when you use the CLI.
#
# ## Print inferencing script
# +
scripts_folder = "Code"
script_file = "iris_score.py"
# peek at contents
with open(os.path.join(scripts_folder, script_file)) as inference_file:
print(inference_file.read())
# -
# ## Build and run the batch inference pipeline
# The data, models, and compute resource are now available. Let's put all these together in a pipeline.
# ### Specify the environment to run the script
# Specify the conda dependencies for your script. This will allow us to install pip packages as well as configure the inference environment.
# +
from azureml.core import Environment
from azureml.core.runconfig import CondaDependencies
predict_conda_deps = CondaDependencies.create(pip_packages=[ "scikit-learn==0.20.3" ])
predict_env = Environment(name="predict_environment")
predict_env.python.conda_dependencies = predict_conda_deps
predict_env.docker.enabled = True
predict_env.spark.precache_packages = False
# -
# ### Create the configuration to wrap the inference script
# +
from azureml.contrib.pipeline.steps import ParallelRunStep, ParallelRunConfig
# In a real-world scenario, you'll want to shape your process per node and nodes to fit your problem domain.
parallel_run_config = ParallelRunConfig(
source_directory=scripts_folder,
entry_script=script_file, # the user script to run against each input
mini_batch_size='5MB',
error_threshold=5,
output_action='append_row',
environment=predict_env,
compute_target=compute_target,
node_count=3,
run_invocation_timeout=600)
# -
# ### Create the pipeline step
# Create the pipeline step using the script, environment configuration, and parameters. Specify the compute target you already attached to your workspace as the target of execution of the script. We will use ParallelRunStep to create the pipeline step.
distributed_csv_iris_step = ParallelRunStep(
name='example-iris',
inputs=[named_iris_ds],
output=output_folder,
parallel_run_config=parallel_run_config,
models=[model],
arguments=['--model_name', 'iris'],
allow_reuse=True
)
# ### Run the pipeline
# At this point you can run the pipeline and examine the output it produced. The Experiment object is used to track the run of the pipeline
# +
from azureml.core import Experiment
from azureml.pipeline.core import Pipeline
pipeline = Pipeline(workspace=ws, steps=[distributed_csv_iris_step])
pipeline_run = Experiment(ws, 'iris').submit(pipeline)
# -
# this will output a table with link to the run details in azure portal
pipeline_run
# ## View progress of Pipeline run
#
# The progress of the pipeline is able to be viewed either through azureml.widgets or a console feed from PipelineRun.wait_for_completion().
# GUI
from azureml.widgets import RunDetails
RunDetails(pipeline_run).show()
# Console logs
pipeline_run.wait_for_completion(show_output=True)
# ## View Results
# In the iris_score.py file above you can see that the Result with the prediction of the iris variety gets returned and then appended to the original input of the row from the csv file. These results are written to the DataStore specified in the PipelineData object as the output data, which in this case is called *inferences*. This contains the outputs from all of the worker nodes used in the compute cluster. You can download this data to view the results ... below just filters to a random 20 rows
# +
import pandas as pd
import shutil
shutil.rmtree("iris_results", ignore_errors=True)
prediction_run = next(pipeline_run.get_children())
prediction_output = prediction_run.get_output_data("inferences")
prediction_output.download(local_path="iris_results")
for root, dirs, files in os.walk("iris_results"):
for file in files:
if file.endswith('parallel_run_step.txt'):
result_file = os.path.join(root,file)
# cleanup output format
df = pd.read_csv(result_file, delimiter=" ", header=None)
df.columns = ["sepal.length", "sepal.width", "petal.length", "petal.width", "variety"]
print("Prediction has ", df.shape[0], " rows")
random_subset = df.sample(n=20)
random_subset.head(20)
# -
# ## Cleanup compute resources
# For re-occuring jobs, it may be wise to keep compute the compute resources and allow compute nodes to scale down to 0. However, since this is just a single run job, we are free to release the allocated compute resources.
# +
# uncomment below and run if compute resources are no longer needed
# compute_target.delete()
|
contrib/batch_inferencing/tabular-dataset-inference-iris.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] colab_type="text" id="7NB399aprajy"
# # Down the rabbit hole with Tensorflow
#
# 
#
# In this seminar, we're going to play with [Tensorflow](https://www.tensorflow.org/) and see how it helps you build deep learning models.
#
# If you're running this notebook outside the course environment, you'll need to install tensorflow:
# * `pip install tensorflow` should install cpu-only TF on Linux & Mac OS
# * If you want GPU support from offset, see [TF install page](https://www.tensorflow.org/install/)
# + colab={} colab_type="code" id="kdF7cq1_raj-"
import sys, os
if 'google.colab' in sys.modules and not os.path.exists('.setup_complete'):
# %tensorflow_version 1.x
if not os.path.exists('.setup_complete'):
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/setup_colab.sh -O- | bash
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/coursera/week1_intro/primer/mnist.py
# !touch .setup_complete
# This code creates a virtual display to draw game images on.
# It will have no effect if your machine has a monitor.
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# !bash ../xvfb start
os.environ['DISPLAY'] = ':1'
# + colab={} colab_type="code" id="HScNFE_ZrakL"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={} colab_type="code" id="Dvht4SuLrakX"
import tensorflow as tf
# session is main tensorflow object. You ask session to compute stuff for you.
sess = tf.InteractiveSession()
# + [markdown] colab_type="text" id="f0rOfFi6rakh"
# # Warming up
# For starters, let's implement a python function that computes the sum of squares of numbers from 0 to N-1.
# * Use numpy or python
# * An array of numbers 0 to N - numpy.arange(N)
# + colab={} colab_type="code" id="n5u1u_TErakh"
def sum_squares(N):
return <student.implement_me()>
# + colab={} colab_type="code" id="wncqxwvirakp"
# %%time
sum_squares(10**8)
# + [markdown] colab_type="text" id="FXJdviP7rak0"
# __Same with tensorflow__
# + colab={} colab_type="code" id="zQaqJy-Hrak3"
# "i will insert N here later"
N = tf.placeholder('int64', name="input_to_your_function")
# a recipe on how to produce {sum of squares of arange of N} given N
result = tf.reduce_sum((tf.range(N)**2))
# + colab={} colab_type="code" id="lWcRJ3Jirak8"
# %%time
# dear session, compute the result please. Here's your N.
print(sess.run(result, {N: 10**8}))
# hint: run it several times to let tensorflow "warm up"
# + [markdown] colab_type="text" id="iL_mEFjoralD"
# # How it works: computation graphs
#
#
# 1. create placeholders for future inputs;
# 2. define symbolic graph: a recipe for mathematical transformation of those placeholders;
# 3. compute outputs of your graph with particular values for each placeholder
# * ```sess.run(outputs, {placeholder1:value1, placeholder2:value2})```
# * OR output.eval({placeholder:value})
#
# Still confused? We gonna fix that.
# + [markdown] colab_type="text" id="Pg9ILbBzralE"
# __Placeholders and constants__
# + colab={} colab_type="code" id="bpFmPU20ralF"
# placeholder that can be arbitrary float32 scalar, vertor, matrix, etc.
arbitrary_input = tf.placeholder('float32')
# input vector of arbitrary length
input_vector = tf.placeholder('float32', shape=(None,))
# input vector that _must_ have 10 elements and integer type
fixed_vector = tf.placeholder('int32', shape=(10,))
# you can generally use None whenever you don't need a specific shape
input1 = tf.placeholder('float64', shape=(None, 100, None))
input2 = tf.placeholder('int32', shape=(None, None, 3, 224, 224))
# + [markdown] colab_type="text" id="LejYHRI4ralM"
# You can create new __tensors__ with arbitrary operations on placeholders, constants and other tensors.
#
# * tf.reduce_sum(tf.arange(N)\**2) are 3 sequential transformations of placeholder N
# * there's a tensorflow symbolic version for every numpy function
# * `a + b, a / b, a ** b, ...` behave just like in numpy
# * np.zeros -> tf.zeros
# * np.sin -> tf.sin
# * np.mean -> tf.reduce_mean
# * np.arange -> tf.range
#
# There are tons of other stuff in tensorflow, see the [docs](https://www.tensorflow.org/api_docs/python) or learn as you go with __shift+tab__.
# + colab={} colab_type="code" id="mQs5iLqnralN"
# elementwise multiplication
double_the_vector = input_vector * 2
# elementwise cosine
elementwise_cosine = tf.cos(input_vector)
# elementwise difference between squared vector and it's means - with some random salt
vector_squares = input_vector ** 2 - \
tf.reduce_mean(input_vector) + tf.random_normal(tf.shape(input_vector))
# + [markdown] colab_type="text" id="3EEidqElralQ"
# ## Practice 1: polar pretzels
# _inspired by [this post](https://www.quora.com/What-are-the-most-interesting-equation-plots)_
#
# There are some simple mathematical functions with cool plots. For one, consider this:
#
# $$ x(t) = t - 1.5 * cos( 15 t) $$
# $$ y(t) = t - 1.5 * sin( 16 t) $$
#
# + colab={} colab_type="code" id="Sg-9MuNJralR"
t = tf.placeholder('float32')
# compute x(t) and y(t) as defined above.
x = <YOUR CODE>
y = <YOUR CODE>
x_points, y_points = sess.run([x, y], {t: np.linspace(-10, 10, num=10000)})
plt.plot(x_points, y_points)
# + [markdown] colab_type="text" id="no2k2g-vralW"
# ### Visualizing graphs with Tensorboard
#
# It's often useful to visualize the computation graph when debugging or optimizing.
# Interactive visualization is where tensorflow really shines as compared to other frameworks.
#
# There's a special instrument for that, called Tensorboard. You can launch it from console:
#
# __```tensorboard --logdir=/tmp/tboard --port=7007```__
#
# If you're pathologically afraid of consoles, try this:
#
# __```import os; os.system("tensorboard --logdir=/tmp/tboard --port=7007 &")```__
#
# _(but don't tell anyone we taught you that)_
# + [markdown] colab_type="text" id="_MtVKFTBralX"
# One basic functionality of tensorboard is drawing graphs. One you've run the cell above, go to `localhost:7007` in your browser and switch to _graphs_ tab in the topbar.
#
# Here's what you should see:
#
# <img src="https://www.tensorflow.org/images/graph_vis_animation.gif" width=480>
#
# Tensorboard also allows you to draw graphs (e.g. learning curves), record images & audio ~~and play flash games~~. This is useful when monitoring learning progress and catching some training issues.
#
# One researcher said:
# ```
# If you spent last four hours of your worktime watching as your algorithm prints numbers and draws figures, you're probably doing deep learning wrong.
# ```
# + [markdown] colab_type="text" id="9nIV4rMzralX"
# You can read more on tensorboard usage [here](https://www.tensorflow.org/get_started/graph_viz)
# + [markdown] colab_type="text" id="uGEIkH4UralZ"
# # Practice 2: mean squared error
#
# + colab={} colab_type="code" id="yX1aRByoralZ"
# Quest #1 - implement a function that computes a mean squared error of two input vectors
# Your function has to take 2 vectors and return a single number
<student.define_inputs_and_transformations()>
mse = <student.define_transformation()>
compute_mse = lambda vector1, vector2: sess.run( < how to run you graph?> , {})
# + colab={} colab_type="code" id="mF0fQ8cxralc"
# Tests
from sklearn.metrics import mean_squared_error
for n in [1, 5, 10, 10 ** 3]:
elems = [np.arange(n), np.arange(n, 0, -1), np.zeros(n),
np.ones(n), np.random.random(n), np.random.randint(100, size=n)]
for el in elems:
for el_2 in elems:
true_mse = np.array(mean_squared_error(el, el_2))
my_mse = compute_mse(el, el_2)
if not np.allclose(true_mse, my_mse):
print('Wrong result:')
print('mse(%s,%s)' % (el, el_2))
print("should be: %f, but your function returned %f" %
(true_mse, my_mse))
raise ValueError, "Что-то не так"
print("All tests passed")
# + [markdown] colab_type="text" id="Z7-lFh25ralh"
# # Tensorflow variables
#
# The inputs and transformations have no value outside function call. That's a bit unnatural if you want your model to have parameters (e.g. network weights) that are always present, but can change their value over time.
#
# Tensorflow solves this with `tf.Variable` objects.
# * You can assign variable a value at any time in your graph
# * Unlike placeholders, there's no need to explicitly pass values to variables when `s.run(...)`-ing
# * You can use variables the same way you use transformations
#
# + colab={} colab_type="code" id="96R2_aCPrali"
# creating shared variable
shared_vector_1 = tf.Variable(initial_value=np.ones(5))
# initialize all variables with initial values
sess.run(tf.global_variables_initializer())
# + colab={} colab_type="code" id="0199csQVralm"
# evaluating shared variable (outside symbolicd graph)
print("initial value", sess.run(shared_vector_1))
# within symbolic graph you use them just as any other inout or transformation, not "get value" needed
# + colab={} colab_type="code" id="OjmEStuuralp"
# setting new value manually
sess.run(shared_vector_1.assign(np.arange(5)))
# getting that new value
print("new value", sess.run(shared_vector_1))
# + [markdown] colab_type="text" id="YJb_Rkhbralt"
# # tf.gradients - why graphs matter
# * Tensorflow can compute derivatives and gradients automatically using the computation graph
# * Gradients are computed as a product of elementary derivatives via chain rule:
#
# $$ {\partial f(g(x)) \over \partial x} = {\partial f(g(x)) \over \partial g(x)}\cdot {\partial g(x) \over \partial x} $$
#
# It can get you the derivative of any graph as long as it knows how to differentiate elementary operations
# + colab={} colab_type="code" id="rSXUsM9rralv"
my_scalar = tf.placeholder('float32')
scalar_squared = my_scalar ** 2
# a derivative of scalar_squared by my_scalar
derivative = tf.gradients(scalar_squared, [my_scalar])[0]
# + colab={} colab_type="code" id="NMTrmY2araly"
x = np.linspace(-3, 3)
x_squared, x_squared_der = sess.run(
[scalar_squared, derivative], {my_scalar: x})
plt.plot(x, x_squared, label="x^2")
plt.plot(x, x_squared_der, label="derivative")
plt.legend()
# + [markdown] colab_type="text" id="h25xVA_Qral1"
# ## Why autograd is cool
# + colab={} colab_type="code" id="08GB4KXFral9"
my_vector = tf.placeholder('float32', [None])
# Compute the gradient of the next weird function over my_scalar and my_vector
# warning! Trying to understand the meaning of that function may result in permanent brain damage
weird_psychotic_function = tf.reduce_mean((my_vector+my_scalar)**(1+tf.nn.moments(my_vector, [0])[1]) + 1. / tf.atan(my_scalar))/(my_scalar**2 + 1) + 0.01*tf.sin(
2*my_scalar**1.5)*(tf.reduce_sum(my_vector) * my_scalar**2)*tf.exp((my_scalar-4)**2)/(1+tf.exp((my_scalar-4)**2))*(1.-(tf.exp(-(my_scalar-4)**2))/(1+tf.exp(-(my_scalar-4)**2)))**2
der_by_scalar = <student.compute_grad_over_scalar()>
der_by_vector = <student.compute_grad_over_vector()>
# + colab={} colab_type="code" id="6IgTD8lCramA"
# Plotting your derivative
scalar_space = np.linspace(1, 7, 100)
y = [sess.run(weird_psychotic_function, {my_scalar: x, my_vector: [1, 2, 3]})
for x in scalar_space]
plt.plot(scalar_space, y, label='function')
y_der_by_scalar = [sess.run(der_by_scalar, {my_scalar: x, my_vector: [1, 2, 3]})
for x in scalar_space]
plt.plot(scalar_space, y_der_by_scalar, label='derivative')
plt.grid()
plt.legend()
# + [markdown] colab_type="text" id="qUv0CXvVramE"
# # Almost done - optimizers
#
# While you can perform gradient descent by hand with automatic grads from above, tensorflow also has some optimization methods implemented for you. Recall momentum & rmsprop?
# + colab={} colab_type="code" id="7VPIeAHTramF"
y_guess = tf.Variable(np.zeros(2, dtype='float32'))
y_true = tf.range(1, 3, dtype='float32')
loss = tf.reduce_mean((y_guess - y_true + tf.random_normal([2]))**2)
optimizer = tf.train.MomentumOptimizer(
0.01, 0.9).minimize(loss, var_list=y_guess)
# same, but more detailed:
# updates = [[tf.gradients(loss,y_guess)[0], y_guess]]
# optimizer = tf.train.MomentumOptimizer(0.01,0.9).apply_gradients(updates)
# + colab={} colab_type="code" id="u_KHCCVZramL"
from IPython.display import clear_output
sess.run(tf.global_variables_initializer())
guesses = [sess.run(y_guess)]
for _ in range(100):
sess.run(optimizer)
guesses.append(sess.run(y_guess))
clear_output(True)
plt.plot(*zip(*guesses), marker='.')
plt.scatter(*sess.run(y_true), c='red')
plt.show()
# + [markdown] colab_type="text" id="Epv8D8ZDramQ"
# # Logistic regression example
# Implement the regular logistic regression training algorithm
#
# We shall train on a two-class MNIST dataset.
#
# This is a binary classification problem, so we'll train a __Logistic Regression with sigmoid__.
# $$P(y_i | X_i) = \sigma(W \cdot X_i + b) ={ 1 \over {1+e^{- [W \cdot X_i + b]}} }$$
#
#
# The natural choice of loss function is to use binary crossentropy (aka logloss, negative llh):
# $$ L = {1 \over N} \underset{X_i,y_i} \sum - [ y_i \cdot log P(y_i | X_i) + (1-y_i) \cdot log (1-P(y_i | X_i)) ]$$
#
# Mind the minus :)
#
# + colab={} colab_type="code" id="Ao44ybVqramQ"
from sklearn.datasets import load_digits
X, y = load_digits(2, return_X_y=True)
print("y [shape - %s]:" % (str(y.shape)), y[:10])
print("X [shape - %s]:" % (str(X.shape)))
# + colab={} colab_type="code" id="PRzGx39NramV"
print('X:\n', X[:3, :10])
print('y:\n', y[:10])
plt.imshow(X[0].reshape([8, 8]))
# + colab={} colab_type="code" id="XJXHAfaUramZ"
# inputs and shareds
weights = <student.create_variable()>
input_X = <student.create_placeholder_matrix()>
input_y = <student.code_placeholder_vector()>
# + colab={} colab_type="code" id="mjX6I88Iramc"
predicted_y_proba = <predicted probabilities for input_X using weights>
loss = <logistic loss(scalar, mean over sample) between predicted_y_proba and input_y>
train_step = <operator that minimizes loss>
# + colab={} colab_type="code" id="hVsu0ldjrami"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# + colab={} colab_type="code" id="HLOgcfl_ramp"
from sklearn.metrics import roc_auc_score
for i in range(5):
loss_i, _ = sess.run([loss, train_step], # <YOUR CODE: feed values to placeholders>)
print("loss at iter %i: %.4f" % (i, loss_i))
print("train auc:", roc_auc_score(
y_train, sess.run(predicted_y_proba, {input_X: X_train})))
print("test auc:", roc_auc_score(
y_test, sess.run(predicted_y_proba, {input_X: X_test})))
print("resulting weights:")
plt.imshow(shared_weights.get_value().reshape(8, -1))
plt.colorbar();
# + [markdown] colab_type="text" id="gKbDbO66ramt"
# # Practice 3: my first tensorflow network
# Your ultimate task for this week is to build your first neural network [almost] from scratch and pure tensorflow.
#
# This time you will same digit recognition problem, but at a larger scale
# * images are now 28x28
# * 10 different digits
# * 50k samples
#
# Note that you are not required to build 152-layer monsters here. A 2-layer (one hidden, one output) NN should already have ive you an edge over logistic regression.
#
# __[bonus score]__
# If you've already beaten logistic regression with a two-layer net, but enthusiasm still ain't gone, you can try improving the test accuracy even further! The milestones would be 95%/97.5%/98.5% accuraсy on test set.
#
# __SPOILER!__
# At the end of the notebook you will find a few tips and frequently made mistakes. If you feel enough might to shoot yourself in the foot without external assistance, we encourage you to do so, but if you encounter any unsurpassable issues, please do look there before mailing us.
# + colab={} colab_type="code" id="-JDNsun_ramu"
from mnist import load_dataset
# [down]loading the original MNIST dataset.
# Please note that you should only train your NN on _train sample,
# _val can be used to evaluate out-of-sample error, compare models or perform early-stopping
# _test should be hidden under a rock untill final evaluation... But we both know it is near impossible to catch you evaluating on it.
X_train, y_train, X_val, y_val, X_test, y_test = load_dataset()
print(X_train.shape, y_train.shape)
# + colab={} colab_type="code" id="82BzUEV7ramy"
plt.imshow(X_train[0, 0])
# + colab={} colab_type="code" id="bet-BegTram1"
<this cell looks as if it wants you to create variables here>
# + colab={} colab_type="code" id="CT2fLkGtram5"
<you could just as well create a computation graph here - loss, optimizers, all that stuff>
# + colab={} colab_type="code" id="Sska5_7gram9"
<this may or may not be a good place to run optimizer in a loop>
# + colab={} colab_type="code" id="VqBtWQlXranD"
<this may be a perfect cell to write a training & evaluation loop in>
# + colab={} colab_type="code" id="RW4KuJZ2ranG"
<predict & evaluate on test here, right? No cheating pls.>
# + [markdown] colab_type="text" id="FnuB4X3granJ"
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
# ```
#
#
# # SPOILERS!
#
# Recommended pipeline
#
# * Adapt logistic regression from previous assignment to classify some number against others (e.g. zero vs nonzero)
# * Generalize it to multiclass logistic regression.
# - Either try to remember lecture 0 or google it.
# - Instead of weight vector you'll have to use matrix (feature_id x class_id)
# - softmax (exp over sum of exps) can implemented manually or as T.nnet.softmax (stable)
# - probably better to use STOCHASTIC gradient descent (minibatch)
# - in which case sample should probably be shuffled (or use random subsamples on each iteration)
# * Add a hidden layer. Now your logistic regression uses hidden neurons instead of inputs.
# - Hidden layer uses the same math as output layer (ex-logistic regression), but uses some nonlinearity (sigmoid) instead of softmax
# - You need to train both layers, not just output layer :)
# - Do not initialize layers with zeros (due to symmetry effects). A gaussian noize with small sigma will do.
# - 50 hidden neurons and a sigmoid nonlinearity will do for a start. Many ways to improve.
# - In ideal casae this totals to 2 .dot's, 1 softmax and 1 sigmoid
# - __make sure this neural network works better than logistic regression__
#
# * Now's the time to try improving the network. Consider layers (size, neuron count), nonlinearities, optimization methods, initialization - whatever you want, but please avoid convolutions for now.
|
Practical_Reinforcement_learning/week_1/primers/recap_tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AshuKV/DCGAN-tensorflow/blob/master/Simul1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="PO0WLW1e1NYb" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="5_JBOfVp1SUl" colab_type="code" colab={}
from keras.models import Sequential
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint
from keras.layers import Dense,Conv2D,MaxPooling2D,Dropout,Flatten,Lambda
# + [markdown] id="HnUk9UiL1Sti" colab_type="text"
#
# + id="42GDUZE66Chy" colab_type="code" outputId="d196de67-1ad1-4474-cdc3-fea56e23b192" colab={"base_uri": "https://localhost:8080/", "height": 54}
# !pip install --user --upgrade scikit-learn==0.18.0
# + id="_miH-gi94S8C" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# + id="zMnmTAOm4WkO" colab_type="code" colab={}
import argparse
import os
# + id="vGk5Q04Q7LML" colab_type="code" colab={}
np.random.seed(1)
# + [markdown] id="7Hsc8TZd8UeW" colab_type="text"
#
# + id="bLks-xSV7Twt" colab_type="code" colab={}
def load(arg):
df=pd.read_csv(os.path.join(os.getcwd(),args.data_dir,"Driving.csv"),names=['center','left','right','steer','throttle','reverse','speed'])
X=df[['cetre','left','right']].values
y=df['steering'].values
X_tr,X_va,Y_tr,Y_va=train_test_split(X,y,test_size=args.test_size,random_state=0)
return X_tarin,X_valid,y_train,y_valid
# + id="oh0LwLI-9ePx" colab_type="code" colab={}
def build(args):
model=Sequential()
model.add(ConvD(12,5,5,activation='elu',subsample=(2,2)))
model.add(ConvD(24,5,5,activation='elu',subsample=(2,2)))
model.add(ConvD(36,5,5,activation='elu',subsample=(2,2)))
model.add(ConvD(48,5,5,activation='elu'))
model.add(ConvD(64,5,5,activation='elu'))
model.add(Dropout(args.keep_prob))
model.add(Flatten())
model.add(Dense(100,activation='elu'))
model.add(Dense(50,activation='elu'))
model.add(Dense(10,activation='elu'))
moedl.add(DEnse(1))
model.summary()
return model
# + id="8ueA6zMTLJJq" colab_type="code" colab={}
def train(model,arg,X_train,X_valid,y_train,y_valid):
checkpoint=ModelCheckpoint('model-{epoch:03d}.h5',
monitor="val_loss",
verbose=0,
save_best_only=args.save_best_only,
mode='auto')
model.compile(loss="mean_squared_error",
optimizere=Adam(lr=args.learning_rate))
model.fit_generator(batch_generator(args.data_dir,X_tr,y_tr,args,batch_size,True),
args.samples_per_epoch,
args.nb_epoch,
max_q_size=1,
validation_data=batch_generator(args.data_dir,X_va,y_va,args,batch_size,False),
nb_val_samples=len(X_valid),
callbacks=[checkpoint],
verbose=1)
# + id="xHfV6wUmN3GF" colab_type="code" colab={}
|
Simul1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Object Detection
#
# *Object detection* is a form of computer vision in which a machine learning model is trained to classify individual instances of objects in an image, and indicate a *bounding box* that marks its location. You can think of this as a progression from *image classification* (in which the model answers the question "what is this an image of?") to building solutions where we can ask the model "what objects are in this image, and where are they?".
#
# 
#
# For example, a grocery store might use an object detection model to implement an automated checkout system that scans a conveyor belt using a camera, and can identify specific items without the need to place each item on the belt and scan them individually.
#
# The **Custom Vision** cognitive service in Microsoft Azure provides a cloud-based solution for creating and publishing custom object detection models.
#
# ## Create a Custom Vision resource
#
# To use the Custom Vision service, you need an Azure resource that you can use to train a model, and a resource with which you can publish it for applications to use. You can use the same resource for each of these tasks, or you can use different resources for each to allocate costs separately provided both resources are created in the same region. The resource for either (or both) tasks can be a general **Cognitive Services** resource, or a specific **Custom Vision** resource. Use the following instructions to create a new **Custom Vision** resource (or you can use an existing resource if you have one).
#
# 1. In a new browser tab, open the Azure portal at [https://portal.azure.com](https://portal.azure.com), and sign in using the Microsoft account associated with your Azure subscription.
# 2. Select the **+Create a resource** button, search for *custom vision*, and create a **Custom Vision** resource with the following settings:
# - **Create options**: Both
# - **Subscription**: *Your Azure subscription*
# - **Resource group**: *Select existing resource group with name AI900-deploymentID*
# - **Name**: *object-deploymentID*
# - **Training location**: *Choose any available region*
# - **Training pricing tier**: F0
# - **Prediction location**: *The same as the training location*
# - **Prediction pricing tier**: F0
#
# > **Note**: If you already have an F0 custom vision service in your subscription, select **S0** for this one.
#
# 3. Wait for the resource to be created.
#
# ## Create a Custom Vision project
#
# To train an object detection model, you need to create a Custom Vision project based on your training resource. To do this, you'll use the Custom Vision portal.
#
# 1. In a new browser tab, open the Custom Vision portal at [https://customvision.ai](https://customvision.ai), and sign in using the Microsoft account associated with your Azure subscription.
# 2. Create a new project with the following settings:
# - **Name**: Grocery Detection
# - **Description**: Object detection for groceries.
# - **Resource**: *The Custom Vision resource you created previously*
# - **Project Types**: Object Detection
# - **Domains**: General
# 3. Wait for the project to be created and opened in the browser.
#
# ## Add and tag images
#
# To train an object detection model, you need to upload images that contain the classes you want the model to identify, and tag them to indicate bounding boxes for each object instance.
#
# 1. Download and extract the training images from https://aka.ms/fruit-objects. The extracted folder contains a collection of images of fruit.
# 2. In the Custom Vision portal, in your object detection project, select **Add images** and upload all of the images in the extracted folder.
# 3. After the images have been uploaded, select the first one to open it.
# 4. Hold the mouse over any object in the image until an automatically detected region is displayed like the image below. Then select the object, and if necessary resize the region to surround it.
#
# 
#
# Alternatively, you can simply drag around the object to create a region.
#
# 5. When the region surrounds the object, add a new tag with the appropriate object type (*apple*, *banana*, or *orange*) as shown here:
#
# 
#
# 6. Select and tag each other object in the image, resizing the regions and adding new tags as required.
#
# 
#
# 7. Use the **>** link on the right to go to the next image, and tag its objects. Then just keep working through the entire image collection, tagging each apple, banana, and orange.
#
# 8. When you have finished tagging the last image, close the **Image Detail** editor and on the **Training Images** page, under **Tags**, select **Tagged** to see all of your tagged images:
#
# 
#
# ## Train and test a model
#
# Now that you've tagged the images in your project, you're ready to train a model.
#
# 1. In the Custom Vision project, click **Train** to train an object detection model using the tagged images. Select the **Quick Training** option.
# 2. Wait for training to complete (it might take ten minutes or so), and then review the *Precision*, *Recall*, and *mAP* performance metrics - these measure the prediction accuracy of the classification model, and should all be high.
# 3. At the top right of the page, click **Quick Test**, and then in the **Image URL** box, enter `https://aka.ms/apple-orange` and view the prediction that is generated. Then close the **Quick Test** window.
#
# ## Publish and consume the object detection model
#
# Now you're ready to publish your trained model and use it from a client application.
#
# 1. At the top left of the **Performance** page, click **🗸 Publish** to publish the trained model with the following settings:
# - **Model name**: detect-produce
# - **Prediction Resource**: *Your custom vision **prediction** resource*.
#
# ### (!) Check In
# Did you use the same model name: **detect-produce**?
#
# 2. After publishing, click the *settings* (⚙) icon at the top right of the **Performance** page to view the project settings. Then, under **General** (on the left), copy the **Project Id**. Scroll down and paste it into the code cell below step 5 replacing **YOUR_PROJECT_ID**.
#
# > (*if you used a **Cognitive Services** resource instead of creating a **Custom Vision** resource at the beginning of this exercise, you can copy its key and endpoint from the right side of the project settings, paste it into the code cell below, and run it to see the results. Otherwise, continue completing the steps below to get the key and endpoint for your Custom Vision prediction resource*).
#
# 3. At the top left of the **Project Settings** page, click the *Projects Gallery* (👁) icon to return to the Custom Vision portal home page, where your project is now listed.
#
# 4. On the Custom Vision portal home page, at the top right, click the *settings* (⚙) icon to view the settings for your Custom Vision service. Then, under **Resources**, expand your *prediction* resource (<u>not</u> the training resource) and copy its **Key** and **Endpoint** values to the code cell below step 5, replacing **YOUR_KEY** and **YOUR_ENDPOINT**.
#
# ### (!) Check In
# If you are using a **Custom Vision** resource, did you use the **prediction** resource (<u>not</u> the training resource)?
#
# 5. Run the code cell below by clicking the Run Cell <span>▷</span> button (at the top left of the cell) to set the variables to your project ID, key, and endpoint values.
# + gather={"logged": 1599692485387}
project_id = 'YOUR_PROJECT_ID' # Replace with your project ID
cv_key = 'YOUR_KEY' # Replace with your prediction resource primary key
cv_endpoint = 'YOUR_ENDPOINT' # Replace with your prediction resource endpoint
model_name = 'detect-produce' # this must match the model name you set when publishing your model iteration exactly (including case)!
print('Ready to predict using model {} in project {}'.format(model_name, project_id))
# -
# Now you can use your key and endpoint with a Custom Vision client to connect to your custom vision object detection model.
#
# Run the following code cell, which uses your model to detect individual produce items in an image.
#
# > **Note**: Don't worry too much about the details of the code. It uses the Python SDK for the Custom Vision service to submit an image to your model and retrieve predictions for detected objects. Each prediction consists of a class name (*apple*, *banana*, or *orange*) and *bounding box* coordinates that indicate where in the image the predicted object has been detected. The code then uses this information to draw a labelled box around each object on the image.
# + gather={"logged": 1599692585672}
from azure.cognitiveservices.vision.customvision.prediction import CustomVisionPredictionClient
from msrest.authentication import ApiKeyCredentials
from matplotlib import pyplot as plt
from PIL import Image, ImageDraw, ImageFont
import numpy as np
import os
# %matplotlib inline
# Load a test image and get its dimensions
test_img_file = os.path.join('data', 'object-detection', 'produce.jpg')
test_img = Image.open(test_img_file)
test_img_h, test_img_w, test_img_ch = np.array(test_img).shape
# Get a prediction client for the object detection model
credentials = ApiKeyCredentials(in_headers={"Prediction-key": cv_key})
predictor = CustomVisionPredictionClient(endpoint=cv_endpoint, credentials=credentials)
print('Detecting objects in {} using model {} in project {}...'.format(test_img_file, model_name, project_id))
# Detect objects in the test image
with open(test_img_file, mode="rb") as test_data:
results = predictor.detect_image(project_id, model_name, test_data)
# Create a figure to display the results
fig = plt.figure(figsize=(8, 8))
plt.axis('off')
# Display the image with boxes around each detected object
draw = ImageDraw.Draw(test_img)
lineWidth = int(np.array(test_img).shape[1]/100)
object_colors = {
"apple": "lightgreen",
"banana": "yellow",
"orange": "orange"
}
for prediction in results.predictions:
color = 'white' # default for 'other' object tags
if (prediction.probability*100) > 50:
if prediction.tag_name in object_colors:
color = object_colors[prediction.tag_name]
left = prediction.bounding_box.left * test_img_w
top = prediction.bounding_box.top * test_img_h
height = prediction.bounding_box.height * test_img_h
width = prediction.bounding_box.width * test_img_w
points = ((left,top), (left+width,top), (left+width,top+height), (left,top+height),(left,top))
draw.line(points, fill=color, width=lineWidth)
plt.annotate(prediction.tag_name + ": {0:.2f}%".format(prediction.probability * 100),(left,top), backgroundcolor=color)
plt.imshow(test_img)
# -
# View the resulting predictions, which show the objects detected and the probability for each prediction.
|
03 - Object Detection.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rose-n-dan/AlcoholPredictionRegression/blob/master/MOW.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9pzxS9119XFt" colab_type="text"
# # Wczytanie bibliotek
# + id="hDehd4kfxNEc" colab_type="code" outputId="ced47ff6-c0ca-4ae6-fc84-95d85b13547c" colab={"base_uri": "https://localhost:8080/", "height": 714}
library(ggplot2)
library(plyr)
library(dplyr)
install.packages("devtools")
library(devtools)
install.packages("gridExtra")
library(gridExtra)
install.packages("alluvial")
library(alluvial)
install.packages("extrafont")
library(extrafont)
install.packages("varhandle")
library(varhandle)
install.packages("caret")
library(caret)
devtools::install_github("mrdwab/splitstackshape", ref = "v2.0")
install.packages("splitstackshape")
### GLMNET LIBS ###
library(tidyverse)
library(broom)
install.packages("glmnet")
library(glmnet)
###################
### SVM LIBS ###
install.packages("e1071")
library(e1071)
################
# + [markdown] id="uTbU2ip33dPo" colab_type="text"
# # Wczytanie danych
# + [markdown] id="9qRLG-NbtTb4" colab_type="text"
# Analiza przeprowadzona zostanie na zbiorze danych udostępnionym na witrynie [kaggle](https://www.kaggle.com/uciml/student-alcohol-consumption). Dane
# ze zbioru zostały zebrane za pomocą ankiet wypełnianych przez uczniów kursów matematyki
# oraz języka portugalskiego w szkole średniej. Są podzielone według kursu, na którego uczniowie
# uczęszczali, i zawierają 30 kolumn, które reprezentują atrybuty informujące o płci, wynikach
# w nauce, statusie społecznym rodziców, aktywnościach czy relacjach z rówieśnikami każdego z
# ankietowanych.
# + id="oia_yxjsyKja" colab_type="code" colab={}
d_mat = read.table("student-mat.csv", sep=",", header=TRUE)
d_por = read.table("student-por.csv", sep=",", header=TRUE)
# + [markdown] id="DIZPOaJfdXzJ" colab_type="text"
# # Przygotowanie danych
# + [markdown] id="iSKtTmcStF0y" colab_type="text"
# Wstępne przetwarzanie atrybutów będzie odbywać się będzie przed dostarczeniem danych
# do modelu regresji wyznaczającej przewidywane spożycie alkoholu. Ponadto, niewykluczone, że
# niezbędnym będzie również ograniczenie liczby wykorzystywanych atrybutów poprzez scalenie.
# + [markdown] id="F6UfDysWj7FM" colab_type="text"
# ## Scalenie zbiorów danych
# + [markdown] id="R4iKHswOkg0-" colab_type="text"
# Scalono zbiory danych uczniów z języka portugalskiego oraz matematyki. Dodatkowo scalono kolumny `G1`, `G2` i `G3` (reprezentujące oceny z przedmiotu w kolejnych trymestrach) jedną kolumną `avggrades` reprezentującą średnią z trzech trymestrów.
# + id="qpDVkcP6B4g9" colab_type="code" outputId="8740b6bd-ff19-4025-e6e4-245b97a2128f" colab={"base_uri": "https://localhost:8080/", "height": 289}
data <- rbind(d_mat, d_por)
data$avggrades = rowMeans(cbind(data$G1, data$G2, data$G3))
data <- data[,-(31:33)]
print(head(data, n=3))
# + [markdown] id="1GNE3rzydf-B" colab_type="text"
# ## Przetworzenie typów danych
# + [markdown] id="MaQ0YrQOa-BT" colab_type="text"
# Zbiór danych zawiera zarówno atrybuty typu liczbowego, jak i słownikowego. Z uwagi na
# charakter parametrów przyjmowanych przez wybrane algorytmy regresji, badania przeprowadzone
# muszą zostać na zbiorach danych o numerycznych typach atrybutów, dlatego niezbędnym jest
# przetworzenie zbioru na wartości numeryczne.
#
# + id="sB2Hqg555_Pu" colab_type="code" colab={}
data <- unfactor(data)
schoolUp <- c("GP" = 0, "MS" = 1)
sexUp <- c("F" = 0, "M" = 1)
addressUp <- c("R" = 0, "U" = 1)
famsizeUp <- c("LE3" = 0, "GT3" = 1)
PstatusUp <- c("A" = 0, "T" = 1)
schoolsupUp <- c("no" = 0, "yes" = 1)
famsupUp <- c("no" = 0, "yes" = 1)
paidUp <- c("no" = 0, "yes" = 1)
activitiesUp <- c("no" = 0, "yes" = 1)
nurseryUp <- c("no" = 0, "yes" = 1)
higherUp <- c("no" = 0, "yes" = 1)
internetUp <- c("no" = 0, "yes" = 1)
romanticUp <- c("no" = 0, "yes" = 1)
data$school <- schoolUp[data$school]
data$sex <- sexUp[data$sex]
data$address <- addressUp[data$address]
data$famsize <- famsizeUp[data$famsize]
data$Pstatus <- PstatusUp[data$Pstatus]
data$schoolsup <- schoolsupUp[data$schoolsup]
data$famsup <- famsupUp[data$famsup]
data$paid <- paidUp[data$paid]
data$activities <- activitiesUp[data$activities]
data$nursery <- nurseryUp[data$nursery]
data$higher <- higherUp[data$higher]
data$internet <- internetUp[data$internet]
data$romantic <- romanticUp[data$romantic]
# + [markdown] id="zW-tsiUtdm83" colab_type="text"
# ## Rozbicie kolumn
# + [markdown] id="O88ZBiA6djXo" colab_type="text"
# W przypadku parametrów słownikowych o liczbie możliwych wartości `n > 2` wykonano zamianę ich na `n` parametrów typu binarnego.
# + id="KzVaa6gOSilx" colab_type="code" colab={}
## Split up the Fjob column
data = splitstackshape::concat.split.expanded(data, "Fjob", "-", type = "character",
fill = 0, drop = TRUE)
## Split up the Mjob column
data = splitstackshape::concat.split.expanded(data, "Mjob", "-", type = "character",
fill = 0, drop = TRUE)
## Split up the reason column
data = splitstackshape::concat.split.expanded(data, "reason", "-", type = "character",
fill = 0, drop = TRUE)
## Split up the guardian column
data = splitstackshape::concat.split.expanded(data, "guardian", "-", type = "character",
fill = 0, drop = TRUE)
# + [markdown] id="uBpvfOOP3jQN" colab_type="text"
# ## Sprawdzenie kompletności danych
# + [markdown] id="FyrnKKOhd4pi" colab_type="text"
# Wyświetlono rozmiar niekomplentych danych.
# + id="wXigEpBi2o7M" colab_type="code" outputId="7cefdc3b-3ed2-4367-b908-5291a640ee0d" colab={"base_uri": "https://localhost:8080/", "height": 34}
missing = data[!complete.cases(data),]
dim(missing)
# + [markdown] id="DY4vXn7_9QR8" colab_type="text"
# Wszystkie dane są kompletne.
# + [markdown] id="ww9KsfiWhfzf" colab_type="text"
# ## Podział danych
# + [markdown] id="H4Ay2LdWDNJv" colab_type="text"
# Dane podzielono na dwa zbiory reprezentujące dwa osobne zadania, czyli prognoza:
#
# * spożycia w dni powszednie
# * spożycia w dni weekendy
#
# Każdy z nich zawiera zestaw zmiennych objaśniających. Ostatnią kolumną zbioru danych jest zmienna prognozowana.
# Żadna ze zmiennych prognozowanych w którymkolwiek z zadań nie należy do zmiennych objaśniających w innym zadaniu.
#
#
# + id="2LGxjuYcGr9j" colab_type="code" outputId="83c43c6d-a31e-43bb-8420-21b7ea52039f" colab={"base_uri": "https://localhost:8080/", "height": 782}
str(data)
# + id="vDNUcMBFCxhP" colab_type="code" outputId="4fa103cb-e2d7-4455-8b43-341a1e2d3672" colab={"base_uri": "https://localhost:8080/", "height": 161}
# change columns order so input data is separated from output
data_wdays <- subset(data, select=c(1:22,25:44,23))
head(data_wdays, n=3)
# + id="s75u5gMVwDcg" colab_type="code" outputId="d513c97a-8905-4d22-fce3-ea44013f71bf" colab={"base_uri": "https://localhost:8080/", "height": 161}
# change columns order so input data is separated from output
data_wend <- subset(data, select=c(1:22,25:44,24))
head(data_wend, n=3)
# + [markdown] id="-31fZPS0_5bh" colab_type="text"
# # Modele
# + [markdown] id="qTmBVMPbtucI" colab_type="text"
# Celem projektu jest wnikliwa i szeroko zakrojona analiza działań zbudowanych modeli regresji w zadaniu przewidywania spożycia alkoholu przez studentów. Za pomocą dostępnych w R narzędzi stworzone zostaną odpowiednie modele pozwalające na przewidzenie ilości spożywanego alkoholu na podstawie wartości atrybutów opisujących każdego z uczniów. W wyniku przeprowadzonych eksperymentów stwierdzone zostanie, który ze zbudowanych modeli daje najlepsze wyniki w tak zdefiniowanym problemie.
#
# Predykcja przeprowadzona zostanie w dwóch częściach: predykcji spożycia weekendowego oraz tygodniowego (bez
# weekendu). W każdym z tych zadań dane o spożyciu alkoholu w jakimkolwiek z dwóch trybów nie będą uwzględniane jako dane wejściowe. Na przykład dane o spożyciu w weekend nie będą uwzględniane w budowaniu modelu predykcji spożycia tygodniowego.
# + [markdown] id="UlKD72vIIeai" colab_type="text"
# ### Definiowanie funkcji pomocniczych
#
# + [markdown] id="6YQNABV9i7lm" colab_type="text"
# Pierwszą z funkcji pomocniczych jest funkcja `rmse` obliczająca pierwiastek błędu średniokwadratowego dla danej predykcji. Średnia kwadratowa błędów ( RMSE ) lub piewiastkiem błędu średniokwadratowego ( MSE ) jest często stosowanym miernikiem różnic między wartościami (próbki lub wartości populacyjnych) przewidywanymi przez model lub estymatora a wartościami obserwowanymi. RMSE reprezentuje pierwiastek kwadratowy z drugiego próbnego momentu różnic między wartościami przewidywanymi a wartościami obserwowanymi lub średnią kwadratową tych różnic.
# + id="9nNZeY46Invl" colab_type="code" colab={}
rmse <- function(Y, predictedY) {
error <- predictedY - Y
rmse <- sqrt(mean(error^2))
return(rmse)
}
# + [markdown] id="dJNaw3dQur_E" colab_type="text"
# Kolejna z funkcji pomocniczych, 'confusion_matrix', używana jest do generowania macierzy pomyłek dla predykcji poszczególnych modeli. Każdy z wierszy macierzy reprezentuje obiekty z klasami predykowanej, natomiast każda z kolumn - obiekty rzeczywistej klasy. Macierz błędów wykorzystywana jest przede wszystkim w ocenie jakości klasyfikacji binarnej, więc aby zastosowanie macierzy pomyłek do oceny jakości modelu regresji było możliwe, należy najpierw zaokrąglić wynik dostarczany z modelu. Klasa rzeczywista ma pięć dyskretnych wartości, także będzie można określić, czy zaokrąglony wynik został sklasyfikowany poprawnie.
#
# Dodatkowo, funkcja, na podstawie wygenerowanej macierzy, wyznacza wskaźniki jakości rozwiązania, jak:
# * dokładność
# * czułość
# * precyzja
# * specyficzność
#
# + id="M87D_ElTRPOo" colab_type="code" colab={}
confusion_matrix <- function(Y, predictedY) {
predictedY <- round(predictedY, digits=0)
predictedY[predictedY < 1] <- 1
predictedY[predictedY > 5] <- 5
confusionMatrix(factor(Y, levels=1:5), factor(predictedY, levels=1:5))
}
# + [markdown] id="iuW1t28Lvd_v" colab_type="text"
# Kolejną funkcją pomocniczą jest funkcja `coefficient_of_determination`, wyznaczająca współczynik determinancji. Współczynnik ten informuje o tym, jaka część wariancji zmiennej objaśnianej w próbie pokrywa się z korelacjami
# ze zmiennymi zawartymi w modelu. Jest on więc miarą stopnia, w jakim model pasuje do próby. Współczynnik determinacji przyjmuje wartości z przedziału [0;1], a dopasowanie modelu jest tym lepsze, im wartość `R^2`
# jest bliższa jedności.
# + id="b1-YNodsbf9G" colab_type="code" colab={}
coefficient_of_determination <- function(Y, predictedY) {
mean <- mean(Y)
numerator <- sum((predictedY - Y) ^ 2)
denominator <- sum((Y - mean) ^ 2)
return(numerator / denominator)
}
# + [markdown] id="sc16o-jxvemH" colab_type="text"
# Ostatnią z funkcji pomocniczych jest funkcja `coefficient_of_variation`. Wyznacza ona współczynnik zmienności resztowej, który informuje o tym, jaką część wartości średniej zmiennej objaśnianej stanowi odchylenie standardowe składnika resztowego, czyli w jakim stopniu na zmienną objaśnianą mają wpływ czynniki losowe.
# + id="J_8rUsy0iG9t" colab_type="code" colab={}
coefficient_of_variation <- function(Y, predictedY, num_col) {
mean <- mean(Y)
error <- predictedY - Y
deviation <- sqrt(sum(error ^ 2) / (NROW(Y) - num_col - 1))
return(deviation / mean)
}
# + [markdown] id="QgUTni7X1Wr4" colab_type="text"
# Funkcja `evaluate_model` drukuje na ekran wyniki wszystkich opisanych funkcji pomocniczych.
# + id="hgmf0UAuy7KU" colab_type="code" colab={}
evaluate_model <- function(Y, predictedY, num_col) {
print(paste0("RMSE: ", rmse(Y, predictedY)))
print(confusion_matrix(Y, predictedY))
print(paste0("Coefficient of determination: ", coefficient_of_determination(Y, predictedY)))
print(paste0("Coefficient of variation: ", coefficient_of_variation(Y, predictedY, num_col)))
}
# + [markdown] id="nqZIhKEcBj7i" colab_type="text"
# ### Przewidywanie spożycia w dni powszednie
#
#
# + [markdown] id="ARWOuszEBdtH" colab_type="text"
# #### Regresja liniowa:
# + [markdown] id="G1Xr8zsmam7D" colab_type="text"
# Regresja liniowa (ang. Linear regression) to metoda oparta o liniowe kombinacje zmiennych i parametrów dopasowujących model do danych, jeden z najpopularniejszych typów regresji. Budowanie modelu regresji liniowej nie wykorzystuje żadnych parametrów.
#
# + id="VBcBqBHg4VE8" colab_type="code" outputId="49bcd0e6-2e80-4a79-e0b1-3dcbaba0ff95" colab={"base_uri": "https://localhost:8080/", "height": 1000}
linear_reg_model_wdays <- lm(Dalc ~ ., data=data_wdays[,1:43])
summary(linear_reg_model_wdays)
# + id="8Vl-2LWr7DdH" colab_type="code" outputId="5df4c702-b289-4575-dbd1-68e251fd7c9b" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(linear_reg_model_wdays)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="frRq1cOd0kHj" colab_type="text"
# #### Regresja logistyczna
# + [markdown] id="Svv9x9PMas58" colab_type="text"
# Regresja logistyczna (ang. Logistic Regression) jest uogólnionym modelem liniowym, w którym używa się funkcji logit jako funkcji wiążącej. Funkcja logit jest logarytmem szansy, jest to metoda bezparametrowa. Wynik predykcji należał będzie do przedziału [0, 1], zostanie więc przeskalowany na przedział [1, 5].
#
# + id="Qa7rgaOd0m9C" colab_type="code" outputId="a87dd1ad-dd80-4e26-be8f-f583019045a1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
logistic_reg_model_wdays <- glm(Dalc ~ ., data=data_wdays[,1:43])
summary(logistic_reg_model_wdays)
# + id="kO2TXFLeGAWE" colab_type="code" outputId="1cff7812-3111-4704-f9ee-e97154fb6765" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(logistic_reg_model_wdays)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="c0UgbEie4Ps7" colab_type="text"
# #### Regresja grzbietowa
# + [markdown] id="HgaSsMwJa5AR" colab_type="text"
# Regresja grzbietowa (ang. Ridge Regression) jest metodą regularyzacji szczególnie użyteczną w przypadku danych zawierających wiele liniowo zależnych zmiennych towarzyszących. Z tego powodu dobrze radzi sobie w przypadkach potencjalnie prowadzących do overfittingu z powodu zbyt dużej liczby parametrów w stosunku do liczby przykładów w zbiorze danych. Potencjalnym parametrem w tej metodzie jest mnożnik Lagrange’a λ. Jendakże pakiet R posiada zaimplementowaną metodę automatycznego dobierania parametru λ na podstawie metody Cule, która dobiera optymalną wartość parametru. Dla porównania przeprowadzony zostanie także eksperyment z dobraną arbitralnie wartością parametru λ.
# + [markdown] id="5Qo1dJ3c6njV" colab_type="text"
# Konwersja danych do formatu wymaganego przez funkcję `glmnet`
# + id="bFQtEY347CZ1" colab_type="code" colab={}
y_wdays <- data_wdays$Dalc
x_wdays <- data_wdays[,1:42] %>% data.matrix()
lambdas <- 10^seq(3, -2, by = -.1)
# + id="Laytf-qD4SRv" colab_type="code" outputId="7b19b2b9-c631-4908-a8a6-71ca2535d99d" colab={"base_uri": "https://localhost:8080/", "height": 238}
ridge_reg_model_wdays <- cv.glmnet(x_wdays, y_wdays, alpha=0, lambda=lambdas)
summary(ridge_reg_model_wdays)
opt_lambda <- ridge_reg_model_wdays$lambda.min
print(opt_lambda)
# + id="iiRKhJeD5wAQ" colab_type="code" outputId="fe7ab9cb-d3d3-470a-f04e-b658ce8bf44e" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(ridge_reg_model_wdays, newx=x_wdays, s=opt_lambda)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="neGU4_lkBUUe" colab_type="text"
# #### Regresja Lasso
# + [markdown] id="4EahjpGHbCW0" colab_type="text"
# Regresja Lasso (ang. Lasso Regression) jest metodą podobną do regresji grzbietowej oraz posiada takie same parametry (więc eksperymenty przeprowadzone z użyciem tej metody będą takie same). W przeciwieństwie do regresji grzbietowej, Lasso dokonuje selekcji zmiennych towarzyszących przez co zwiększa interpretowalność modelu.
# + id="rTyMH4kzBZmy" colab_type="code" outputId="9fcef5f6-80aa-4ae3-ba64-6d7c4e49181b" colab={"base_uri": "https://localhost:8080/", "height": 238}
lasso_reg_model_wdays <- cv.glmnet(x_wdays, y_wdays, alpha=1, lambda=lambdas)
summary(lasso_reg_model_wdays)
opt_lambda <- lasso_reg_model_wdays$lambda.min
print(opt_lambda)
# + id="2_mXesC4z3ks" colab_type="code" outputId="bef282d2-b8d0-432f-c65a-c8af710bf6e7" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(lasso_reg_model_wdays, newx=x_wdays, s=opt_lambda)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="kiWJQhr2Ce6H" colab_type="text"
# #### Regresja Elastic Net
# + [markdown] id="jKf4rGk6bF87" colab_type="text"
# Regresja Elastic Net (ang. Elastic Net Regression) jest kombinacją liniową dwóch poprzednich metod: regresji grzbietowej oraz Lasso. Parametrem, który będzie tu regulowany jest α - przyjmuje on wartości od 0 do 1 i mówi on o tym jak duży udział w modelu będzie miała każda ze składowych: grzbietowa oraz Lasso.
# + [markdown] id="WRhuTfo0ChwH" colab_type="text"
# Testy dla trzech wartości parametru alpha: 0.25, 0.5 i 0.75.
# + id="kQevLBwoC41P" colab_type="code" outputId="2eeb31b9-5808-462f-ace3-5f6accd419f9" colab={"base_uri": "https://localhost:8080/", "height": 238}
elastic_net_025_reg_model_wdays <- cv.glmnet(x_wdays, y_wdays, alpha=0.25, lambda=lambdas)
summary(elastic_net_025_reg_model_wdays)
opt_lambda <- elastic_net_025_reg_model_wdays$lambda.min
print(opt_lambda)
# + id="ZJ_met6ZzaY3" colab_type="code" outputId="22f34a99-05e8-499f-ef0f-2cee4580dd5f" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(elastic_net_025_reg_model_wdays, newx=x_wdays, s=opt_lambda)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + id="VzBu2jLIFLrt" colab_type="code" outputId="a5e9dad8-1bf2-41d2-bd06-e4a652da8995" colab={"base_uri": "https://localhost:8080/", "height": 238}
elastic_net_05_reg_model_wdays <- cv.glmnet(x_wdays, y_wdays, alpha=0.5, lamda=lambdas)
summary(elastic_net_05_reg_model_wdays)
opt_lambda <- elastic_net_05_reg_model_wdays$lambda.min
print(opt_lambda)
# + id="EHkuTslfzbHc" colab_type="code" outputId="86bf1ced-4fcf-4c86-8c9d-96ce40daaac5" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(ridge_reg_model_wdays, newx=x_wdays, s=opt_lambda)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + id="90GCVqHiFQey" colab_type="code" outputId="ce5379de-614c-46e8-a399-0389579d3e99" colab={"base_uri": "https://localhost:8080/", "height": 238}
elastic_net_075_reg_model_wdays <- cv.glmnet(x_wdays, y_wdays, alpha=0.75, lamda=lambdas)
summary(elastic_net_075_reg_model_wdays)
opt_lambda <- elastic_net_075_reg_model_wdays$lambda.min
print(opt_lambda)
# + id="W9nx6kmrzbxZ" colab_type="code" outputId="b9b9544c-51ba-4a84-bb64-e628924de2b2" colab={"base_uri": "https://localhost:8080/", "height": 612}
predictedY <- predict(ridge_reg_model_wdays, newx=x_wdays, s=opt_lambda)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="S147PLvYIYtL" colab_type="text"
# #### Regresja Wektorów Nośnych
# + [markdown] id="Y31R1VN7bMF-" colab_type="text"
# Regresja Wektorów Nośnych (ang. Support Vector Regression - SVR) jest to metoda regresji, która nie minimalizuje błędu średniokwadratowego każdego z przypadków, a bada czy leży w określonych przez parametr `epsilon` granicach. Każdy punkt nieleżący w tych granicach generuje dodatkową karę. W eksperymentach użyte zostaną dwa jądra: liniowe oraz radialnej funkcji bazowej.
# + [markdown] id="i79tYAPXqX2C" colab_type="text"
# Za pomocą funkcji `tune` dopasowano lokalnie optymalne wartości parametrów modelu regresji wektorów nośnych: `epsilon` i `cost`.
# + [markdown] id="RlW_8-coIlI2" colab_type="text"
# Dopasowano model korzystający z jądra liniowego.
# + id="gpLL-gHlIro1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="7a94b884-4a2e-4220-d270-ca3184dc121a"
tune_results <- tune(svm, x_wdays, y_wdays, svm = "regression", kernel="linear",
ranges = list(epsilon = seq(0, 1, 0.2), cost = 2^(1:5)))
plot(tune_results)
# + id="8itKre1LSp_L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="37bf920c-5cf3-40e3-e782-809d32a715e7"
svrl_model_wdays <- tune_results$best.model
summary(svrl_model_wdays)
# + id="unerD0c59e3X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="8bd8c91d-ce4e-4f74-8d1d-088cfda04d09"
predictedY <- predict(svrl_model_wdays, x=x_wdays)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="IJD_XycmvIZc" colab_type="text"
# Dopasowano model korzystający o radialnej funkcji bazowej.
#
# + id="6Tw9_DHavL1R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="6b07397f-ee62-44f8-9c90-7b11dfa6d3d0"
tune_results <- tune(svm, x_wdays, y_wdays, svm = "regression", kernel="radial",
ranges = list(epsilon = seq(0, 1.0, 0.1), cost = 2^(-1:5)))
plot(tune_results)
# + id="dfxGTX5tvc2f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="00596cf3-b2e2-4593-aa1c-faa5a6651dcb"
svrr_model_wdays <- tune_results$best.model
summary(svrr_model_wdays)
# + id="CepfuyLY9ra_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="62f74fd5-c6f8-42f0-e03b-52ee133ad936"
predictedY <- predict(svrr_model_wdays, x=x_wdays)
Y <- data_wdays$Dalc
evaluate_model(Y, predictedY, ncol(data_wdays))
# + [markdown] id="lnZ9CRjWCJon" colab_type="text"
# ### Przewidywanie spożycia weekendowego
#
# + [markdown] id="3F4PD7zebXrr" colab_type="text"
# Na tym etapie powtórzone zostaną operacje wykonane uprzednio w celu predykcji spożycia alkoholu w ciągu tygodnia roboczego, tym razem w celu predukcji spożycia w dni weekendowe.
# + [markdown] id="uwkMIrrUCZDF" colab_type="text"
# #### Regresja liniowa:
# + id="jw-lVMvL_8sH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="031f5c59-55c6-44df-839c-a66d8aef46c9"
linear_reg_model_wend <- lm(Walc ~ ., data=data_wend[,1:43])
summary(linear_reg_model_wend)
# + id="SzX52Mr7DGiX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="e25f68bd-728f-4a02-fe86-dcd06dbf5d9d"
predictedY <- predict(linear_reg_model_wend)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] colab_type="text" id="ROutf4uwoFbt"
# #### Regresja logistyczna
# + id="06U-MmaspFPo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3b4d6629-6699-40f4-94b9-28de68def53f"
logistic_reg_model_wend <- glm(Walc ~ ., data=data_wend[,1:43])
summary(logistic_reg_model_wend)
# + id="yosidGd5DO82" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="12b275ec-366d-48b0-ea97-c307911a957a"
predictedY <- predict(logistic_reg_model_wend)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] colab_type="text" id="cLilB_Ljn8QT"
# #### Regresja grzbietowa
# + id="71yxc8xTpQFu" colab_type="code" colab={}
y_wend <- data_wend$Walc
x_wend <- data_wend[,1:42] %>% data.matrix()
lambdas <- 10^seq(3, -2, by = -.1)
# + id="yWlyagvbpbwt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="4b62a6b2-e0cf-4e19-b3ce-31b777c8b9aa"
ridge_reg_model_wend <- cv.glmnet(x_wend, y_wend, alpha=0, lambda=lambdas)
summary(ridge_reg_model_wend)
opt_lambda <- ridge_reg_model_wend$lambda.min
print(opt_lambda)
# + id="vAcLBX5cDhax" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="7e32c8c0-2d37-4d3b-f537-54a0572b095e"
predictedY <- predict(ridge_reg_model_wend, newx=x_wdays, s=opt_lambda)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] colab_type="text" id="dxXZQKk-oOPk"
# #### Regresja Lasso
# + id="I5tJkTXIpjSA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="fdcc86d0-fc22-4fa3-9718-84b24713b1e8"
lasso_reg_model_wend <- cv.glmnet(x_wend, y_wend, alpha=1, lambda=lambdas)
summary(lasso_reg_model_wend)
opt_lambda <- lasso_reg_model_wend$lambda.min
print(opt_lambda)
# + id="tddGVQjpDxe0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="1de40a12-a9b4-40d5-8636-59b62277fafe"
predictedY <- predict(lasso_reg_model_wend, newx=x_wdays, s=opt_lambda)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] colab_type="text" id="yclpeljNoWpk"
# #### Regresja Elastic Net
# + id="wGa2yntnpqqQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="dd880535-c598-41ab-87a9-c0dd2691514a"
elastic_net_025_reg_model_wend <- cv.glmnet(x_wend, y_wend, alpha=0.25, lambda=lambdas)
summary(elastic_net_025_reg_model_wend)
opt_lambda <- elastic_net_025_reg_model_wend$lambda.min
print(opt_lambda)
# + id="PTLV71e4EIqz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="a8fb0b94-3b12-412d-86c9-4a5df9ab41e8"
predictedY <- predict(elastic_net_025_reg_model_wend, newx=x_wdays, s=opt_lambda)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + id="c20LrhZqpwnA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="14787c14-7904-4266-fe4d-582943deb4c7"
elastic_net_05_reg_model_wend <- cv.glmnet(x_wend, y_wend, alpha=0.5, lambda=lambdas)
summary(elastic_net_05_reg_model_wend)
opt_lambda <- elastic_net_05_reg_model_wend$lambda.min
print(opt_lambda)
# + id="Crc1pwzKEJnS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="3b06b2cf-e873-4baa-b296-439807310a32"
predictedY <- predict(elastic_net_05_reg_model_wend, newx=x_wdays, s=opt_lambda)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + id="arUULc6vp09O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="d947e5ff-40f8-4f7b-85d7-4415ee581289"
elastic_net_075_reg_model_wend <- cv.glmnet(x_wend, y_wend, alpha=0.75, lambda=lambdas)
summary(elastic_net_075_reg_model_wend)
opt_lambda <- elastic_net_075_reg_model_wend$lambda.min
print(opt_lambda)
# + id="t36a-IAcEKNW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="88f172c6-bdfe-449e-d072-7ee9a1269160"
predictedY <- predict(elastic_net_075_reg_model_wend, newx=x_wdays, s=opt_lambda)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] colab_type="text" id="jHnzv5P_oZr1"
# #### Regresja Wektorów Nośnych
# + id="EyQNpTq4qAPI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="0fd8e50b-f8b0-4e41-998f-83fca9e04647"
tune_results <- tune(svm, x_wend, y_wend, svm = "regression", kernel="linear",
ranges = list(epsilon = seq(0, 1, 0.2), cost = 2^(1:6)))
plot(tune_results)
# + id="G_ptkHIvqF0f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="72ba6617-f5ef-428a-d512-cc0270bcae2f"
svrl_model_wend <- tune_results$best.model
summary(svrl_model_wend)
# + id="ibfgdvWREXR_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="2ee56d07-de79-48c6-fc72-f73af71dae07"
predictedY <- predict(svrl_model_wend, x=x_wend)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + id="iNyA_8h4q1Pg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="c6d6155b-1612-4a30-9de8-e206a1f2686d"
tune_results <- tune(svm, x_wend, y_wend, svm = "regression", kernel="radial",
ranges = list(epsilon = seq(0, 1.0, 0.1), cost = 2^(-1:5)))
plot(tune_results)
# + id="4vzXbwS6q7yd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="fc6d3c9c-5080-4584-f8b0-155c78560b56"
svrr_model_wend <- tune_results$best.model
summary(svrr_model_wend)
# + id="iJtV4p4CEYLg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 612} outputId="327cb39c-57c9-4dea-cb56-693ab222fb72"
predictedY <- predict(svrr_model_wend, x=x_wend)
Y <- data_wend$Walc
evaluate_model(Y, predictedY, ncol(data_wend))
# + [markdown] id="pFU99u99Qf2t" colab_type="text"
# # Wnioski
# + [markdown] id="tKh_EHSFQo6g" colab_type="text"
#
# Podsumowując, należałoby ocenić, czy tak zbudowane modele posiadałyby jakąkolwiek użyteczną funkcjonalność w dzisiejszych czasach i wyciągnąć kilka wniosków:
# * Wartości atrybutów klasy `Dalc` w udostępnionym zbiorze danych były mocno niezbalansowane. Spośród 1043 rekordów, jedynie po 26 posiadało wartość 4 lub 5, co miało duży wpływ na predykcje modeli regresji.
# * Maszyny wektorów nośnych, choć realizujące w przeważającej liczbie przypadków zadania klasyfikacyjne, mogą z powodzeniem być wykorzystywane do budowy modelów regresorów.
# * Stopień skomplikowania i pojemność modelu jest niezależna od liczby wymiarów danych.
# * Model oparty na maszynie wektorów nośnych znajdowuje minimum globalne. Minimalizowana jest funkcja kwadratowa co gwarantuje znalezienie minimum. Algorytm jest bardzo wydajny i SVM generuje prawie optymalny regresor. Nie jest tez czuły na przetrenowanie.
# * Trening modelu przebiega bardzo powoli, w szczególności trening modelu, którego jądro oparte jest o funkcje wielomianową i przy dużej ilości danych użytych do treningu.
# * Dane zebrane w postaci anonimowych ankiet nie stanowią idealnej reprezentacji. Uczniowie mogli wypełniać je subiektywnie, przez co nie oddawały one rzeczywistości i nie pozwoliłyby na dobrej jakości predykcje.
# * Cechy, jakimi posłużono się do opisu poszczególnych uczniów były niewystarczająco dokładne. Wiele atrybutów mogło przyjąć ograniczoną liczbę wartości w wąskim przedziale, co prowadziło do utraty informacji.
# * Dodatkowo budowe modelu komplikował fakt, że niektóre z atrybutów, które wykraczały w rzeczywistości za minimalną lub maksymalną skale, w jakiej atrybuty były reprezentowane, zostały przycięte do tej skali.
# * Dane dotyczą jedynie uczniów dwóch róznych liceów. Model ten nie posiada zastosowania do oszacowania spożycia alkoholu przez uczniów z innych regionów czy krajów.
|
MOW.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sparkify Project Workspace
# This workspace contains a tiny subset (128MB) of the full dataset available (12GB). Feel free to use this workspace to build your project, or to explore a smaller subset with Spark before deploying your cluster on the cloud. Instructions for setting up your Spark cluster is included in the last lesson of the Extracurricular Spark Course content.
#
# You can follow the steps below to guide your data analysis and model building portion of this project.
# import libraries
from pyspark.sql import SparkSession
import pandas as pd
from pyspark.sql.functions import isnan, when, count, col, countDistinct, to_timestamp
from pyspark.sql import functions as F
import matplotlib.pyplot as plt
import seaborn as sns
from pyspark.ml.feature import MinMaxScaler, VectorAssembler
from pyspark.sql.types import IntegerType
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression, RandomForestClassifier, LinearSVC, GBTClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
# create a Spark session
spark = SparkSession \
.builder \
.appName("Python Spark SQL") \
.getOrCreate()
# # Load and Clean Dataset
# In this workspace, the mini-dataset file is `mini_sparkify_event_data.json`. Load and clean the dataset, checking for invalid or missing data - for example, records without userids or sessionids.
df = spark.read.json('mini_sparkify_event_data.json')
df.show(5)
print((df.count(), len(df.columns)))
df.select([count(when(isnan(c) | col(c).isNull(), c)).alias(c) for c in df.columns]).show()
df.select(col('location')).groupBy('location').count().count()
for column in df.columns:
if df.select(col(column)).groupBy(column).count().count()<30:
print('\033[1m' + column + '\033[0m') , print(df.select(col(column)).groupBy(column).count().show(30, False))
df.where(col("firstName").isNull()).select(col('auth')).groupBy('auth').count().show()
df.where(col("firstName").isNull()).select(col('level')).groupBy('level').count().show()
df.where(col("firstName").isNull()).select(col('page')).groupBy('page').count().show()
df.where(col("artist").isNotNull()).select(col('page')).groupBy('page').count().show()
df.where(col("artist").isNull()).select(col('page')).groupBy('page').count().show()
# We have 2 different types of missing values.
#
# 1. Missing user data for 8346 entries. From the analysis above it seems that the users that have null data are users that have not logged in the app yet. As these events cannot be correlated with the userId, we cannot use them, so we will drop them
# 2. Mssing song data for 58392 entries. From the analysis above it seems that the missing songs are reasonable. The song data are populated only in case the page is the NextSong page, so we will keep all these entries for now
df = df.na.drop(subset=["firstName"])
df.select([count(when(isnan(c) | col(c).isNull(), c)).alias(c) for c in df.columns]).show()
# # Exploratory Data Analysis
# When you're working with the full dataset, perform EDA by loading a small subset of the data and doing basic manipulations within Spark. In this workspace, you are already provided a small subset of data you can explore.
#
# ### Define Churn
#
# Once you've done some preliminary analysis, create a column `Churn` to use as the label for your model. I suggest using the `Cancellation Confirmation` events to define your churn, which happen for both paid and free users. As a bonus task, you can also look into the `Downgrade` events.
#
# ### Explore Data
# Once you've defined churn, perform some exploratory data analysis to observe the behavior for users who stayed vs users who churned. You can start by exploring aggregates on these two groups of users, observing how much of a specific action they experienced per a certain time unit or number of songs played.
df.createOrReplaceTempView("DATA")
spark.sql("""
SELECT count(distinct userId) FROM DATA
""").show(10, False)
spark.sql("""
SELECT distinct userId,page
FROM DATA
where page in ('Cancellation Confirmation','Downgrade')
order by userId,page
""").show(10, False)
spark.sql("""
SELECT page,to_timestamp(ts/1000) as ts,level
FROM DATA
where userId='100001'
order by ts
""").show(500, False)
spark.sql("""
SELECT page,to_timestamp(ts/1000) as ts,level
FROM DATA
where userId='100002'
order by ts
""").show(500, False)
# We can see that even that the user went to Downgrade page he remained paid. I assume that he should do a Submit Downgrade page to consider his downgrade valid
spark.sql("""
SELECT distinct userId,page
FROM DATA
where page in ('Cancellation Confirmation','Submit Downgrade')
order by userId,page
""").show(10, False)
spark.sql("""
SELECT page,to_timestamp(ts/1000) as ts,level
FROM DATA
where userId='100009'
order by ts
""").show(500, False)
# This user after the submit upgrade become paid and after the submit downgrade become free again
spark.sql("""
SELECT distinct userId,page
FROM DATA
where page in ('Cancellation Confirmation','Submit Downgrade')
order by userId,page
""").count()
df = df.withColumn("churn", when((col("page")=='Cancellation Confirmation') \
| (col("page")=='Submit Downgrade'),1).otherwise(0))
df.show(5)
df.createOrReplaceTempView("DATA")
spark.sql("""
SELECT distinct userId,page,churn
FROM DATA
where page in ('Cancellation Confirmation','Submit Downgrade')
order by userId,page
""").show(5, False)
# # Feature Engineering
# Once you've familiarized yourself with the data, build out the features you find promising to train your model on. To work with the full dataset, you can follow the following steps.
# - Write a script to extract the necessary features from the smaller subset of data
# - Ensure that your script is scalable, using the best practices discussed in Lesson 3
# - Try your script on the full data set, debugging your script if necessary
#
# If you are working in the classroom workspace, you can just extract features based on the small subset of data contained here. Be sure to transfer over this work to the larger dataset when you work on your Spark cluster.
spark.sql("""
SELECT max(to_timestamp(ts/1000)) as max_ts,min(to_timestamp(ts/1000)) as min_ts
FROM DATA
""").show(5, False)
df_dataset = spark.sql("""
SELECT DATA.userId,
case when gender='M' then 1 else 0 end as is_male_flag,
max(churn) as churn,
count(distinct ts_day) as days_in_app,
count(distinct song)/sum(case when song is not null then 1 else 0 end) as avg_songs,
count(distinct artist)/sum(case when song is not null then 1 else 0 end) as avg_artists,
round(sum(length/60)/sum(case when song is not null then 1 else 0 end),2) as avg_song_length,
count(1) as events_cnt,
count(1)/count(distinct ts_day) as avg_sessions_per_day,
sum(case when DATA.page='NextSong' then 1 else 0 end)/count(distinct ts_day) as avg_pg_song_cnt,
sum(case when DATA.page='Roll Advert' then 1 else 0 end)/count(distinct ts_day) as avg_pg_advert_cnt,
sum(case when DATA.page='Logout' then 1 else 0 end)/count(distinct ts_day) as avg_pg_logout_cnt,
sum(case when DATA.page='Thumbs Down' then 1 else 0 end)/count(distinct ts_day) as avg_pg_down_cnt,
sum(case when DATA.page='Thumbs Up' then 1 else 0 end)/count(distinct ts_day) as avg_pg_up_cnt,
sum(case when DATA.page='Add Friend' then 1 else 0 end)/count(distinct ts_day) as avg_pg_friend_cnt,
sum(case when DATA.page='Add to Playlist' then 1 else 0 end)/count(distinct ts_day) as avg_pg_playlist_cnt,
sum(case when DATA.page='Help' then 1 else 0 end)/count(distinct ts_day) as avg_pg_help_cnt,
sum(case when DATA.page='Home' then 1 else 0 end)/count(distinct ts_day) as avg_pg_home_cnt,
sum(case when DATA.page='Save Settings' then 1 else 0 end)/count(distinct ts_day) as avg_pg_save_settings_cnt,
sum(case when DATA.page='About' then 1 else 0 end)/count(distinct ts_day) as avg_pg_about_cnt,
sum(case when DATA.page='Settings' then 1 else 0 end)/count(distinct ts_day) as avg_pg_settings_cnt,
sum(case when DATA.page='Login' then 1 else 0 end)/count(distinct ts_day) as avg_pg_login_cnt,
sum(case when DATA.page='Submit Registration' then 1 else 0 end)/count(distinct ts_day) as avg_pg_sub_reg_cnt,
sum(case when DATA.page='Register' then 1 else 0 end)/count(distinct ts_day) as avg_pg_reg_cnt,
sum(case when DATA.page='Upgrade' then 1 else 0 end)/count(distinct ts_day) as avg_pg_upg_cnt,
sum(case when DATA.page='Submit Upgrade' then 1 else 0 end)/count(distinct ts_day) as avg_pg_sub_upg_cnt,
sum(case when DATA.page='Error' then 1 else 0 end)/count(distinct ts_day) as avg_pg_error_cnt
FROM DATA
LEFT JOIN
(
SELECT distinct DATE_TRUNC('day', to_timestamp(ts/1000)) as ts_day, userId FROM DATA
) day_ts
ON day_ts.userId=DATA.userId
GROUP BY DATA.userId,gender
""")
churn_cnt = df_dataset.select(col('churn'),col('userId')).groupby('churn').count().toPandas()
#churn_cnt.show()
sns.barplot('churn','count', data=churn_cnt)
plt.title('Churn Distribution')
plt.xticks(rotation = 90)
is_male_flag_dstr = df_dataset.select(col('is_male_flag'),col('churn')).groupby('is_male_flag','churn').agg(count("churn").alias("churn_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'churn_cnt', hue = 'is_male_flag', data=is_male_flag_dstr)
plt.title('Churn Distribution Per Gender')
plt.xticks(rotation = 90)
is_male_flag_dstr = df_dataset.select(col('is_male_flag'),col('churn')).groupby('is_male_flag').agg(F.mean("churn").alias("avg_churn_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('is_male_flag', 'avg_churn_cnt', data=is_male_flag_dstr)
plt.title('Average Churn Distribution Per Gender')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('days_in_app'),col('churn')).groupby('churn').agg(F.mean("days_in_app").alias("avg_days_in_app")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_days_in_app', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Days in App')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_sessions_per_day'),col('churn')).groupby('churn').agg(F.mean("avg_sessions_per_day").alias("avg_sessions_per_day")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_sessions_per_day', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Sessions Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_down_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_down_cnt").alias("avg_pg_thumbs_down")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_thumbs_down', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Thumbs Down Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_up_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_up_cnt").alias("avg_pg_thumps_up")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_thumps_up', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Thumbs Up Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_friend_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_friend_cnt").alias("avg_pg_friend_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_friend_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Add Friends Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_playlist_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_playlist_cnt").alias("avg_pg_playlist_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_playlist_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Add to Playlist Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_advert_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_advert_cnt").alias("avg_pg_advert_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_advert_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Average Advert Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_error_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_error_cnt").alias("avg_pg_error_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_error_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Error Per Day')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('events_cnt'),col('churn')).groupby('churn').agg(F.mean("events_cnt").alias("avg_events_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_events_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Events Average')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_song_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_song_cnt").alias("avg_pg_song_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_song_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Songs Average')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_logout_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_logout_cnt").alias("avg_pg_logout_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_logout_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per LogOut Average')
plt.xticks(rotation = 90)
days_in_app_dstr = df_dataset.select(col('avg_pg_sub_upg_cnt'),col('churn')).groupby('churn').agg(F.mean("avg_pg_sub_upg_cnt").alias("avg_pg_sub_upg_cnt")).toPandas()
#is_male_flag_dstr.show()
sns.barplot('churn', 'avg_pg_sub_upg_cnt', data=days_in_app_dstr)
plt.title('Churn Distribution Per Upgrade Average')
plt.xticks(rotation = 90)
# # Modeling
# Split the full dataset into train, test, and validation sets. Test out several of the machine learning methods you learned. Evaluate the accuracy of the various models, tuning parameters as necessary. Determine your winning model based on test accuracy and report results on the validation set. Since the churned users are a fairly small subset, I suggest using F1 score as the metric to optimize.
df_dataset = spark.sql("""
SELECT DATA.userId,
case when gender='M' then 1 else 0 end as is_male_flag,
max(churn) as churn,
count(distinct ts_day) as days_in_app,
count(distinct song)/sum(case when song is not null then 1 else 0 end) as avg_songs,
count(distinct artist)/sum(case when song is not null then 1 else 0 end) as avg_artists,
round(sum(length/60)/sum(case when song is not null then 1 else 0 end),2) as avg_song_length,
count(1) as events_cnt,
count(1)/count(distinct ts_day) as avg_sessions_per_day,
sum(case when DATA.page='NextSong' then 1 else 0 end)/count(distinct ts_day) as avg_pg_song_cnt,
sum(case when DATA.page='Roll Advert' then 1 else 0 end)/count(distinct ts_day) as avg_pg_advert_cnt,
sum(case when DATA.page='Logout' then 1 else 0 end)/count(distinct ts_day) as avg_pg_logout_cnt,
sum(case when DATA.page='Thumbs Down' then 1 else 0 end)/count(distinct ts_day) as avg_pg_down_cnt,
sum(case when DATA.page='Thumbs Up' then 1 else 0 end)/count(distinct ts_day) as avg_pg_up_cnt,
sum(case when DATA.page='Add Friend' then 1 else 0 end)/count(distinct ts_day) as avg_pg_friend_cnt,
sum(case when DATA.page='Add to Playlist' then 1 else 0 end)/count(distinct ts_day) as avg_pg_playlist_cnt,
sum(case when DATA.page='Help' then 1 else 0 end)/count(distinct ts_day) as avg_pg_help_cnt,
sum(case when DATA.page='Home' then 1 else 0 end)/count(distinct ts_day) as avg_pg_home_cnt,
sum(case when DATA.page='Save Settings' then 1 else 0 end)/count(distinct ts_day) as avg_pg_save_settings_cnt,
sum(case when DATA.page='About' then 1 else 0 end)/count(distinct ts_day) as avg_pg_about_cnt,
sum(case when DATA.page='Settings' then 1 else 0 end)/count(distinct ts_day) as avg_pg_settings_cnt,
sum(case when DATA.page='Login' then 1 else 0 end)/count(distinct ts_day) as avg_pg_login_cnt,
sum(case when DATA.page='Submit Registration' then 1 else 0 end)/count(distinct ts_day) as avg_pg_sub_reg_cnt,
sum(case when DATA.page='Register' then 1 else 0 end)/count(distinct ts_day) as avg_pg_reg_cnt,
sum(case when DATA.page='Upgrade' then 1 else 0 end)/count(distinct ts_day) as avg_pg_upg_cnt,
sum(case when DATA.page='Submit Upgrade' then 1 else 0 end)/count(distinct ts_day) as avg_pg_sub_upg_cnt,
sum(case when DATA.page='Error' then 1 else 0 end)/count(distinct ts_day) as avg_pg_error_cnt
FROM DATA
LEFT JOIN
(
SELECT distinct DATE_TRUNC('day', to_timestamp(ts/1000)) as ts_day, userId FROM DATA
) day_ts
ON day_ts.userId=DATA.userId
GROUP BY DATA.userId,gender
""")
#for column in ['days_in_app','events_cnt','avg_sessions_per_day','avg_pg_song_cnt','avg_pg_advert_cnt',
# 'avg_pg_friend_cnt','avg_pg_playlist_cnt','avg_songs','avg_artists','avg_song_length',
# 'avg_pg_logout_cnt','avg_pg_sub_upg_cnt','avg_pg_upg_cnt','avg_pg_down_cnt','avg_pg_up_cnt',
# 'avg_pg_error_cnt'
# ]:
for column in [ 'days_in_app',
'events_cnt',
'avg_sessions_per_day',
'avg_pg_song_cnt',
'avg_pg_advert_cnt',
'avg_pg_down_cnt',
'avg_pg_up_cnt',
'avg_pg_friend_cnt',
'avg_pg_playlist_cnt'
]:
# VectorAssembler Transformation - Converting column to vector type
vector_assempler = VectorAssembler(inputCols=[column],outputCol=column+"_vect")
# MinMaxScaler Transformation
scaler = MinMaxScaler(inputCol=column+"_vect", outputCol=column+"_scaled")
# Pipeline of VectorAssembler and MinMaxScaler
pipeline = Pipeline(stages=[vector_assempler, scaler])
# Fitting pipeline on dataframe
df_dataset = pipeline.fit(df_dataset).transform(df_dataset).drop(column+"_vect")
#features_vector_assempler = VectorAssembler(inputCols=['days_in_app_scaled','events_cnt_scaled',
# 'avg_sessions_per_day_scaled','avg_pg_song_cnt_scaled','avg_pg_advert_cnt_scaled',
# 'avg_pg_friend_cnt_scaled','avg_pg_playlist_cnt_scaled','avg_songs_scaled','avg_artists_scaled',
# 'avg_song_length_scaled','avg_pg_logout_cnt_scaled','avg_pg_sub_upg_cnt_scaled',
# 'avg_pg_upg_cnt_scaled','avg_pg_down_cnt_scaled','avg_pg_up_cnt_scaled',
# 'avg_pg_error_cnt_scaled'
# ],outputCol="features")
features_vector_assempler = VectorAssembler(inputCols=[
'days_in_app_scaled',
'events_cnt_scaled',
'avg_sessions_per_day_scaled',
'avg_pg_song_cnt_scaled',
'avg_pg_advert_cnt_scaled',
'avg_pg_down_cnt_scaled',
'avg_pg_up_cnt_scaled',
'avg_pg_friend_cnt_scaled',
'avg_pg_playlist_cnt_scaled'],outputCol="features")
df_dataset_model = features_vector_assempler.transform(df_dataset)
df_dataset_model = df_dataset_model.select(col("churn").alias("label"),col("features"))
#Test 1
train, test = df_dataset_model.randomSplit([0.8, 0.2], seed=7)
#sub_test, validation = test.randomSplit([0.5, 0.5], seed = 7)
print("Training Dataset Count: " + str(train.count()))
print("Test Dataset Count: " + str(test.count()))
gbt = GBTClassifier(featuresCol = 'features', labelCol = "label", maxIter = 10, maxDepth = 10, seed = 7)
gbt_fitted_model = gbt.fit(train)
predictions = gbt_fitted_model.transform(test)
f1 = MulticlassClassificationEvaluator(metricName = 'f1')
acc = MulticlassClassificationEvaluator(metricName = 'accuracy')
prec = MulticlassClassificationEvaluator(metricName = 'weightedPrecision')
rec = MulticlassClassificationEvaluator(metricName = 'weightedRecall')
gbt_f1_score = f1.evaluate(predictions)
gbt_acc_score = acc.evaluate(predictions)
gbt_prec_score = prec.evaluate(predictions)
gbt_rec_score = rec.evaluate(predictions)
print('GBT Accuracy: {}, GBT Precision: {}, GBT Recall: {}, GBT F1-Score: {}'.format(round(gbt_acc_score*100,2),round(gbt_prec_score*100,2),round(gbt_rec_score*100,2),round(gbt_f1_score*100,2)))
rf = RandomForestClassifier()
rf_fitted_model = rf.fit(train)
predictions = rf_fitted_model.transform(test)
f1 = MulticlassClassificationEvaluator(metricName = 'f1')
acc = MulticlassClassificationEvaluator(metricName = 'accuracy')
prec = MulticlassClassificationEvaluator(metricName = 'weightedPrecision')
rec = MulticlassClassificationEvaluator(metricName = 'weightedRecall')
rf_f1_score = f1.evaluate(predictions)
rf_acc_score = acc.evaluate(predictions)
rf_prec_score = prec.evaluate(predictions)
rf_rec_score = rec.evaluate(predictions)
print('Random Forest Accuracy: {}, Random Forest Precision: {}, Random Forest Recall: {}, Random Forest F1-Score: {}'.format(round(rf_acc_score*100,2),round(rf_prec_score*100,2),round(rf_rec_score*100,2),round(rf_f1_score*100,2)))
lr = LogisticRegression(featuresCol="features", labelCol="label", maxIter=10, regParam=0.01)
lr_fitted_model = lr.fit(train)
predictions = lr_fitted_model.transform(test)
f1 = MulticlassClassificationEvaluator(metricName = 'f1')
acc = MulticlassClassificationEvaluator(metricName = 'accuracy')
prec = MulticlassClassificationEvaluator(metricName = 'weightedPrecision')
rec = MulticlassClassificationEvaluator(metricName = 'weightedRecall')
lr_f1_score = f1.evaluate(predictions)
lr_acc_score = acc.evaluate(predictions)
lr_prec_score = prec.evaluate(predictions)
lr_rec_score = rec.evaluate(predictions)
print('Logistic Regression Accuracy: {}, Logistic Regression Precision: {}, Logistic Regression Recall: {}, Logistic Regression F1-Score: {}'.format(round(lr_acc_score*100,2),round(lr_prec_score*100,2),round(lr_rec_score*100,2),round(lr_f1_score*100,2)))
svm = LinearSVC(featuresCol="features", labelCol="label", maxIter=10, regParam=0.1)
svm_fitted_model = svm.fit(train)
predictions = svm_fitted_model.transform(test)
f1 = MulticlassClassificationEvaluator(metricName = 'f1')
acc = MulticlassClassificationEvaluator(metricName = 'accuracy')
prec = MulticlassClassificationEvaluator(metricName = 'weightedPrecision')
rec = MulticlassClassificationEvaluator(metricName = 'weightedRecall')
svm_f1_score = f1.evaluate(predictions)
svm_acc_score = acc.evaluate(predictions)
svm_prec_score = prec.evaluate(predictions)
svm_rec_score = rec.evaluate(predictions)
print('SVM Accuracy: {}, SVM Precision: {}, SVM Recall: {}, SVM F1-Score: {}'.format(round(svm_acc_score*100,2),round(svm_prec_score*100,2),round(svm_rec_score*100,2),round(svm_f1_score*100,2)))
# From the above executions and evaluations we will choosse as better performant algorythm the GBT one.
# This is the algorythm that we will use for the calculation of the churn score with these KPIs.
#
# Of course, the next step is to evaluate and validate the results running the code on the full dataset.
# If we are happy with the results we can deploy the churn calculation algorythm in production
# # Final Steps
# Clean up your code, adding comments and renaming variables to make the code easier to read and maintain. Refer to the Spark Project Overview page and Data Scientist Capstone Project Rubric to make sure you are including all components of the capstone project and meet all expectations. Remember, this includes thorough documentation in a README file in a Github repository, as well as a web app or blog post.
|
try_runs/Sparkify_6th_run.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: HSE-RL
# language: python
# name: hse-rl
# ---
# # Crossentropy method
#
# This notebook will teach you to solve reinforcement learning problems with crossentropy method. We'll follow-up by scaling everything up and using neural network policy.
# +
# In google collab, uncomment this:
# # !wget https://bit.ly/2FMJP5K -O setup.py && bash setup.py
# XVFB will be launched if you run on a server
# import os
# if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# # !bash ../xvfb start
# %env DISPLAY = : 1
# +
import gym
import numpy as np
import pandas as pd
env = gym.make("Taxi-v2")
env.reset()
env.render()
# +
n_states = env.observation_space.n
n_actions = env.action_space.n
print("n_states=%i, n_actions=%i" % (n_states, n_actions))
# -
# # Create stochastic policy
#
# This time our policy should be a probability distribution.
#
# ```policy[s,a] = P(take action a | in state s)```
#
# Since we still use integer state and action representations, you can use a 2-dimensional array to represent the policy.
#
# Please initialize policy __uniformly__, that is, probabililities of all actions should be equal.
#
policy = np.zeros([n_states, n_actions]) + 1.0 / n_actions
assert type(policy) in (np.ndarray, np.matrix)
assert np.allclose(policy, 1./n_actions)
assert np.allclose(np.sum(policy, axis=1), 1)
# # Play the game
#
# Just like before, but we also record all states and actions we took.
def generate_session(env, policy, t_max=10**4):
"""
Play game until end or for t_max ticks.
:param policy: an array of shape [n_states,n_actions] with action probabilities
:returns: list of states, list of actions and sum of rewards
"""
states, actions = [], []
total_reward = 0.
s = env.reset()
for t in range(t_max):
a = np.random.choice(np.arange(n_actions), p=policy[s])
new_s, r, done, info = env.step(a)
# Record state, action and add up reward to states,actions and total_reward accordingly.
states.append(s)
actions.append(a)
total_reward += r
s = new_s
if done:
break
return states, actions, total_reward
s, a, r = generate_session(env, policy)
assert type(s) == type(a) == list
assert len(s) == len(a)
assert type(r) in [float, np.float]
# +
# let's see the initial reward distribution
import matplotlib.pyplot as plt
# %matplotlib inline
sample_rewards = [generate_session(env, policy, t_max=1000)[-1] for _ in range(200)]
plt.hist(sample_rewards, bins=20)
plt.vlines([np.percentile(sample_rewards, 50)], [0], [
100], label="50'th percentile", color='green')
plt.vlines([np.percentile(sample_rewards, 90)], [0], [
100], label="90'th percentile", color='red')
plt.legend()
# -
# ### Crossentropy method steps (2pts)
def select_elites(states_batch, actions_batch, rewards_batch, percentile=50):
"""
Select states and actions from games that have rewards >= percentile
:param states_batch: list of lists of states, states_batch[session_i][t]
:param actions_batch: list of lists of actions, actions_batch[session_i][t]
:param rewards_batch: list of rewards, rewards_batch[session_i]
:returns: elite_states,elite_actions, both 1D lists of states and respective actions from elite sessions
Please return elite states and actions in their original order
[i.e. sorted by session number and timestep within session]
If you're confused, see examples below. Please don't assume that states are integers (they'll get different later).
"""
reward_threshold = np.percentile(rewards_batch, percentile)
sessions = rewards_batch > reward_threshold
elite_states = []
elite_actions = []
for it, reward in enumerate(rewards_batch):
if reward >= reward_threshold:
elite_states += states_batch[it]
elite_actions += actions_batch[it]
return elite_states, elite_actions
# +
states_batch = [
[1, 2, 3], # game1
[4, 2, 0, 2], # game2
[3, 1] # game3
]
actions_batch = [
[0, 2, 4], # game1
[3, 2, 0, 1], # game2
[3, 3] # game3
]
rewards_batch = [
3, # game1
4, # game2
5, # game3
]
test_result_0 = select_elites(
states_batch, actions_batch, rewards_batch, percentile=0)
test_result_40 = select_elites(
states_batch, actions_batch, rewards_batch, percentile=30)
test_result_90 = select_elites(
states_batch, actions_batch, rewards_batch, percentile=90)
test_result_100 = select_elites(
states_batch, actions_batch, rewards_batch, percentile=100)
assert np.all(test_result_0[0] == [1, 2, 3, 4, 2, 0, 2, 3, 1]) \
and np.all(test_result_0[1] == [0, 2, 4, 3, 2, 0, 1, 3, 3]),\
"For percentile 0 you should return all states and actions in chronological order"
assert np.all(test_result_40[0] == [4, 2, 0, 2, 3, 1]) and \
np.all(test_result_40[1] == [3, 2, 0, 1, 3, 3]),\
"For percentile 30 you should only select states/actions from two first"
assert np.all(test_result_90[0] == [3, 1]) and \
np.all(test_result_90[1] == [3, 3]),\
"For percentile 90 you should only select states/actions from one game"
assert np.all(test_result_100[0] == [3, 1]) and\
np.all(test_result_100[1] == [3, 3]),\
"Please make sure you use >=, not >. Also double-check how you compute percentile."
print("Ok!")
# -
def update_policy(elite_states, elite_actions):
"""
Given old policy and a list of elite states/actions from select_elites,
return new updated policy where each action probability is proportional to
policy[s_i,a_i] ~ #[occurences of si and ai in elite states/actions]
Don't forget to normalize policy to get valid probabilities and handle 0/0 case.
In case you never visited a state, set probabilities for all actions to 1./n_actions
:param elite_states: 1D list of states from elite sessions
:param elite_actions: 1D list of actions from elite sessions
"""
new_policy = np.zeros([n_states, n_actions])
for state, action in zip(elite_states, elite_actions):
new_policy[state][action] += 1
n_visits = np.sum(new_policy, axis=1)
new_policy[n_visits == 0, :] = 1
n_visits[n_visits == 0] = n_actions
new_policy /= n_visits[None].T
return new_policy
# +
elite_states, elite_actions = ([1, 2, 3, 4, 2, 0, 2, 3, 1],
[0, 2, 4, 3, 2, 0, 1, 3, 3])
new_policy = update_policy(elite_states, elite_actions)
assert np.isfinite(new_policy).all(
), "Your new policy contains NaNs or +-inf. Make sure you don't divide by zero."
assert np.all(
new_policy >= 0), "Your new policy can't have negative action probabilities"
assert np.allclose(new_policy.sum(
axis=-1), 1), "Your new policy should be a valid probability distribution over actions"
reference_answer = np.array([
[1., 0., 0., 0., 0.],
[0.5, 0., 0., 0.5, 0.],
[0., 0.33333333, 0.66666667, 0., 0.],
[0., 0., 0., 0.5, 0.5]])
assert np.allclose(new_policy[:4, :5], reference_answer)
print("Ok!")
# -
# # Training loop
# Generate sessions, select N best and fit to those.
# +
import itertools
from joblib import Parallel, delayed
# -
def generate_batch(env, n_sessions, *args, **kwargs):
return [generate_session(env, *args, **kwargs) for _ in range(n_sessions)]
# +
from IPython.display import clear_output
def show_progress(rewards_batch, log, percentile, reward_range=[-990, +10]):
"""
A convenience function that displays training progress.
No cool math here, just charts.
"""
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.plot(list(zip(*log))[0], label='Mean rewards')
plt.plot(list(zip(*log))[1], label='Reward thresholds')
plt.legend()
plt.grid()
plt.subplot(1, 2, 2)
plt.hist(rewards_batch, range=reward_range)
plt.vlines([np.percentile(rewards_batch, percentile)],
[0], [100], label="percentile", color='red')
plt.legend()
plt.grid()
plt.show()
# -
def solve_taxi(n_sessions, percentile, learning_rate, n_steps, t_max=1000, n_jobs=4):
assert n_sessions % n_jobs == 0, "Number of sessions should divide by number of jobs"
n_sessions_ = n_sessions // n_jobs
log = []
policy = np.ones([n_states, n_actions]) / n_actions
envs = [gym.make("Taxi-v2").env for _ in range(n_jobs)]
for i in range(n_steps):
with Parallel(n_jobs=n_jobs) as pool:
func = delayed(generate_batch)
sessions = pool(func(it, n_sessions_, policy, 1000) for it in envs)
sessions = list(itertools.chain(*sessions))
states_batch, actions_batch, rewards_batch = zip(*sessions)
elite_states, elite_actions = select_elites(states_batch, actions_batch, rewards_batch, percentile)
new_policy = update_policy(elite_states, elite_actions)
policy = learning_rate * new_policy + (1 - learning_rate) * policy
mean_reward = np.mean(rewards_batch)
threshold = np.percentile(rewards_batch, percentile)
log.append([mean_reward, threshold])
clear_output(True)
print("mean reward = %.3f, threshold=%.3f" % (mean_reward, threshold))
show_progress(rewards_batch, log, percentile)
solve_taxi(n_sessions=300, percentile=50, learning_rate=0.5, n_steps=100)
solve_taxi(n_sessions=300, percentile=70, learning_rate=0.5, n_steps=100)
solve_taxi(n_sessions=300, percentile=30, learning_rate=0.5, n_steps=100)
solve_taxi(n_sessions=300, percentile=25, learning_rate=0.3, n_steps=100)
solve_taxi(n_sessions=500, percentile=40, learning_rate=0.2, n_steps=100)
solve_taxi(n_sessions=500, percentile=25, learning_rate=0.2, n_steps=200)
solve_taxi(n_sessions=500, percentile=50, learning_rate=0.5, n_steps=100)
# # Digging deeper: approximate crossentropy with neural nets
#
# 
#
# In this section we will train a neural network policy for continuous state space game
# +
# if you see "<classname> has no attribute .env", remove .env or update gym
env = gym.make("CartPole-v0").env
env.reset()
n_actions = env.action_space.n
plt.imshow(env.render("rgb_array"))
# -
# create agent
from sklearn.neural_network import MLPClassifier
agent = MLPClassifier(hidden_layer_sizes=(20, 20),
activation='tanh',
warm_start=True, # keep progress between .fit(...) calls
max_iter=1 # make only 1 iteration on each .fit(...)
)
# initialize agent to the dimension of state an amount of actions
agent.fit([env.reset()]*n_actions, range(n_actions))
def generate_session(env, t_max=1000):
states, actions = [], []
total_reward = 0
s = env.reset()
for t in range(t_max):
# predict array of action probabilities
probs = agent.predict_proba([s])[0]
a = np.random.choice(np.arange(n_actions), p=probs)
new_s, r, done, info = env.step(a)
# record sessions like you did before
states.append(s)
actions.append(a)
total_reward += r
s = new_s
if done:
break
return states, actions, total_reward
# +
n_sessions = 100
percentile = 70
log = []
for i in range(100):
sessions = [generate_session(env) for _ in range(n_sessions)]
states_batch, actions_batch, rewards_batch = map(np.array, zip(*sessions))
elite_states, elite_actions = select_elites(states_batch, actions_batch, rewards_batch)
agent.fit(elite_states, elite_actions)
mean_reward = np.mean(rewards_batch)
threshold = np.percentile(rewards_batch, percentile)
log.append([mean_reward, threshold])
clear_output(True)
print("mean reward = %.3f, threshold=%.3f" % (mean_reward, threshold))
show_progress(rewards_batch, log, percentile, reward_range=[0, np.max(rewards_batch)])
if np.mean(rewards_batch) > 190:
print("You Win!")
break
# -
# # Results
# record sessions
import gym.wrappers
env = gym.wrappers.Monitor(gym.make("CartPole-v0"),
directory="videos", force=True)
sessions = [generate_session(env) for _ in range(100)]
env.close()
# +
# show video
from IPython.display import HTML
import os
video_names = list(filter(lambda s: s.endswith(".mp4"), os.listdir("./videos/")))
HTML("""
<video width="640" height="480" controls>
<source src="{}" type="video/mp4">
</video>
""".format("./videos/" + video_names[-1])) # this may or may not be _last_ video. Try other indices
# -
# # Homework part I
#
# ### Tabular crossentropy method
#
# You may have noticed that the taxi problem quickly converges from -100 to a near-optimal score and then descends back into -50/-100. This is in part because the environment has some innate randomness. Namely, the starting points of passenger/driver change from episode to episode.
#
# ### Tasks
# - __1.1__ (1 pts) Find out how the algorithm performance changes if you change different percentile and different n_samples.
# - __1.2__ (2 pts) Tune the algorithm to end up with positive average score.
#
# It's okay to modify the existing code.
#
# - More samples $\rightarrow$ more statistics $\rightarrow$ lower variance
# - Larger percentile $\rightarrow$ less statistics $\rightarrow$ higher variance
#
# As you can see from the experiments above, if the percentile is equal 30, the average score is higher than if it's equal 50 or 70. As I understand it happens because the learning rate and variances are too high.
# # Homework part II
#
# ### Deep crossentropy method
#
# By this moment you should have got enough score on [CartPole-v0](https://gym.openai.com/envs/CartPole-v0) to consider it solved (see the link). It's time to try something harder.
#
# * if you have any trouble with CartPole-v0 and feel stuck, feel free to ask us or your peers for help.
#
# ### Tasks
#
# * __2.1__ (3 pts) Pick one of environments: MountainCar-v0 or LunarLander-v2.
# * For MountainCar, get average reward of __at least -150__
# * For LunarLander, get average reward of __at least +50__
#
# See the tips section below, it's kinda important.
# __Note:__ If your agent is below the target score, you'll still get most of the points depending on the result, so don't be afraid to submit it.
#
#
# * __2.2__ (bonus: 4++ pt) Devise a way to speed up training at least 2x against the default version
# * Obvious improvement: use [joblib](https://www.google.com/search?client=ubuntu&channel=fs&q=joblib&ie=utf-8&oe=utf-8)
# * Try re-using samples from 3-5 last iterations when computing threshold and training
# * Experiment with amount of training iterations and learning rate of the neural network (see params)
# * __Please list what you did in anytask submission form__
#
#
# ### Tips
# * Gym page: [mountaincar](https://gym.openai.com/envs/MountainCar-v0), [lunarlander](https://gym.openai.com/envs/LunarLander-v2)
# * Sessions for MountainCar may last for 10k+ ticks. Make sure ```t_max``` param is at least 10k.
# * Also it may be a good idea to cut rewards via ">" and not ">=". If 90% of your sessions get reward of -10k and 20% are better, than if you use percentile 20% as threshold, R >= threshold __fails cut off bad sessions__ whule R > threshold works alright.
# * _issue with gym_: Some versions of gym limit game time by 200 ticks. This will prevent cem training in most cases. Make sure your agent is able to play for the specified __t_max__, and if it isn't, try `env = gym.make("MountainCar-v0").env` or otherwise get rid of TimeLimit wrapper.
# * If you use old _swig_ lib for LunarLander-v2, you may get an error. See this [issue](https://github.com/openai/gym/issues/100) for solution.
# * If it won't train it's a good idea to plot reward distribution and record sessions: they may give you some clue. If they don't, call course staff :)
# * 20-neuron network is probably not enough, feel free to experiment.
#
# ### Bonus tasks
#
# * __2.3 bonus__ Try to find a network architecture and training params that solve __both__ environments above (_Points depend on implementation. If you attempted this task, please mention it in anytask submission._)
#
# * __2.4 bonus__ Solve continuous action space task with `MLPRegressor` or similar.
# * Start with ["Pendulum-v0"](https://github.com/openai/gym/wiki/Pendulum-v0).
# * Since your agent only predicts the "expected" action, you will have to add noise to ensure exploration.
# * [MountainCarContinuous-v0](https://gym.openai.com/envs/MountainCarContinuous-v0), [LunarLanderContinuous-v2](https://gym.openai.com/envs/LunarLanderContinuous-v2)
# * 4 points for solving. Slightly less for getting some results below solution threshold. Note that discrete and continuous environments may have slightly different rules aside from action spaces.
#
#
# If you're still feeling unchallenged, consider the project (see other notebook in this folder).
# +
env = gym.make("MountainCar-v0").env
env.reset()
n_actions = env.action_space.n
plt.imshow(env.render("rgb_array"))
# +
agent = MLPClassifier(hidden_layer_sizes=(10, 10),
activation='tanh',
warm_start=True,
learning_rate_init=0.01,
max_iter=10)
agent.fit([env.reset()] * n_actions, range(n_actions))
# +
n_jobs = 4
n_sessions = 40
percentile = 70
n_batch_history = 5
assert n_sessions % n_jobs == 0, "Number of sessions should divide by number of jobs"
n_sessions_ = n_sessions // n_jobs
log = []
states = []
actions = []
rewards = []
envs = [gym.make("MountainCar-v0").env for _ in range(n_jobs)]
for i in range(300):
with Parallel(n_jobs=n_jobs) as pool:
sessions = pool(delayed(generate_batch)(it, n_sessions_, 10_000) for it in envs)
sessions = list(itertools.chain(*sessions))
states_batch, actions_batch, rewards_batch = map(np.array, zip(*sessions))
states_batch = [list(it) for it in states_batch]
actions_batch = [list(it) for it in actions_batch]
rewards_batch = list(rewards_batch)
states += states_batch
actions += actions_batch
rewards += rewards_batch
if i >= n_batch_history:
states = states[n_sessions:]
actions = actions[n_sessions:]
rewards = rewards[n_sessions:]
elite_states, elite_actions = select_elites(states, actions, rewards)
agent.fit(elite_states, elite_actions)
mean_reward = np.mean(rewards_batch)
threshold = np.percentile(rewards_batch, percentile)
log.append([mean_reward, threshold])
clear_output(True)
print("mean reward = %.3f, threshold=%.3f" % (mean_reward, threshold))
show_progress(rewards_batch, log, percentile, reward_range=[np.min(rewards_batch), np.max(rewards_batch)])
|
task01/crossentropy_method.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GENERATE TNC CURBSIDE DEMAND
#
# This module takes the arrival and departure demand along with the mode choice percentage and extracts the appropriate amount of demand from the people demand and converts it to cars for TNCs.
#
# ```python
# def create_sumo_demand_TNC_curbside(people,
# percentOfTNC=.255,
# stops=stop_dict = {
# "A":['A_top_1','A_top_2','A_top_3','A_bot_1'],
# "B":['B_top_1','B_top_2','B_top_3','B_bot_1'],
# "C":['C_top_1','C_top_2','C_top_3','C_bot_1'],
# "D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2'],
# "E":['E_top_1','E_top_2','E_top_3','E_bot_1'],
#
# },
# alt_stops = {
# "A":['A_top_1','A_top_2','A_top_3','A_bot_1','A_bot_2','A_bot_3'],
# "B":['B_top_1','B_top_2','B_top_3','B_bot_1','B_bot_2','B_bot_3'],
# "C":['C_top_1','C_top_2','C_top_3','C_bot_1','C_bot_2','C_bot_3'],
# "D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2','D_service'],
# "E":['E_top_1','E_top_2','E_top_3','E_bot_1','E_bot_2','E_bot_3'],
#
# },
# end_weight = [.2,.8],
# start_weights = [.225,.225,.275,.275],
# stop_duration = np.random.exponential(20,1000) + np.random.normal(60,5,1000),
# ):
# ```
#
# > - **people:** *DataFrame* - is the people demand for the
# > - **percentOFTNC** *Float* - perecent of TNCs going to curb
# > - **stops** *Dictionary* - informs the stops for the passenger vehicles
# > - **stops** *Dictionary* - informs the stops for the passenger vehicles
# > - **alt_stops** *Dictionary* - informs where the alterante stops for a policy implimentation
# > - **Policy** *Class/Function* - informs which policy function to impliment
# > - **end_weight** *List* - list of floats that add to 1. Informs the destination of each trip.
# > - **start_weights** *List or Array* - list of floats that add to 1. Informs the origin of each trip.
# > - **stop_duration** *List or Array* - distribution of dwell times on the curb
#
#
#
# +
from __future__ import print_function
import xml.etree.ElementTree as ET
from xml.etree.ElementTree import Element, SubElement, Comment
from lxml import etree
from copy import copy
import os
import inspect
from xml.dom import minidom
import pandas as pd
import numpy as np
import random
import sys
sys.path.insert(0, os.path.abspath('../Generate_SUMO_demand/'))
import DFW_gen_flow as gf
# -
Date = '2018-5-13'
# +
folder = "../Example_Files/Demand_CSV"
selected = pd.read_csv("selected_demand.csv")
level = selected.loc[selected['Date'] == Date]['Volume_Category'].item()
file_name = Date + '.' + level +".people.csv"
people = pd.read_csv(os.path.join(folder,file_name),index_col=[0])
people.head()
# -
def create_sumo_demand_TNC_curbside(people,
level,
percentOfTNC=.255,
peopleToCars=1.7,
stops= {
"A":['A_top_1','A_top_2','A_top_3','A_bot_1'],
"B":['B_top_1','B_top_2','B_top_3','B_bot_1'],
"C":['C_top_1','C_top_2','C_top_3','C_bot_1'],
"D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2'],
"E":['E_top_1','E_top_2','E_top_3','E_bot_1'],
},
alt_stops = {
"A":['A_top_1','A_top_2','A_top_3','A_bot_1','A_bot_2','A_bot_3'],
"B":['B_top_1','B_top_2','B_top_3','B_bot_1','B_bot_2','B_bot_3'],
"C":['C_top_1','C_top_2','C_top_3','C_bot_1','C_bot_2','C_bot_3'],
"D":['D_depart_1','D_depart_2','D_arrive_1','D_arrive_2','D_service'],
"E":['E_top_1','E_top_2','E_top_3','E_bot_1','E_bot_2','E_bot_3'],
},
ballpark = {
'Arrive':['TNC_1','TNC_2','TNC_3']
},
policy=None,
end_weight = [.2,.8],
start_weights = [.225,.225,.275,.275],
stop_duration = np.random.exponential(20,10000) + np.random.normal(60,5,10000),
):
end_weight_south = end_weight[::-1]
columns = ['Arrive_A_people','Arrive_B_people','Arrive_C_people','Arrive_D_people',
'Arrive_E_people','Depart_A_people','Depart_B_people',
'Depart_C_people','Depart_D_people','Depart_E_people']
starts = ['South_1', 'South_Plaza', 'North_Plaza', 'North_1']
ends = ['South_Exit', 'North_Exit']
routes = Element('routes')
routes.set('xmlns:xsi','http://www.w3.org/2001/XMLSchema-instance')
routes.set('xsi:noNamespaceSchemaLocation', 'http://sumo.dlr.de/xsd/routes_file.xsd')
people['seconds'] = np.array(people.index) * 30 * 60
count = 1
for column in columns:
column_string = column.split('_')
terminal = column_string[1]
for t,numberOfPeople in enumerate(people[column]):
numberOfVehicles = round((numberOfPeople/peopleToCars) * percentOfTNC)
for i in range(numberOfVehicles):
time = people['seconds'][t] + round(np.random.uniform(0,1800))
# make this basic from plaza to terminal
start = np.random.choice(starts,p=start_weights )
if start[0] == "S":
p = end_weight_south
else:
p = end_weight
end = np.random.choice(ends,p=p)
stop = np.random.choice(stops[terminal])
trip = Element('trip')
trip.set('id', column + '_TNC_' + str(count))
trip.set('type', 'passenger')
trip.set('color', "#bb0000")
trip.set('depart',str(time))
trip.set('from',start)
trip.set('to',end)
trip.set('departSpeed', "max")
trip.set('departLane', "best")
count+=1
routes.append(trip)
duration = str(np.random.choice(stop_duration))
ET.SubElement(trip,"stop",busStop=stop,duration=duration,parking='true')
routes[:] = sorted(routes, key=lambda child: (child.tag,float(child.get('depart'))))
file_name = Date + "." + level+ ".TNC.curb.xml"
folder = "../Example_Files/TempDemandXML"
print("Saving to xml: ", file_name)
with open(os.path.join(folder,file_name), 'wb') as f:
f.write(minidom.parseString(ET.tostring(routes)).toprettyxml(encoding="utf-8"))
create_sumo_demand_TNC_curbside(people,level)
|
Congestion_Policies/Generate_TNC_Trips.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from keras import layers
from keras import models
#network design with 4 conv layer and 2 dense(fully connected layer)
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(150, 150, 3))) #using 32nos of 3x3 kernels (each kernel has depth of size 3 just like the i/p image)
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(128, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
#flattening the o/p after conv layers, since the dense layer accept i/p as 1D vector not as a matrix
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid')) #sigmoid activation gives probability to which of the classes the i/p image belong
#model configuration
from keras import optimizers
model.compile(loss='binary_crossentropy',
optimizer=optimizers.RMSprop(lr=1e-4),
metrics=['acc'])
# +
#data preprocessing - done with ImageDataGenerator class of Keras
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale=1./255) #argument rescale of method ImageDataGen rescales the pixel values of the image (which is btwn 0 to 255 i.e, 8bit) to [0,1]
test_datagen = ImageDataGenerator(rescale=1./255) #by dividing it by 255
train_generator = train_datagen.flow_from_directory( #using flow_from_directory method of ImageDataGen class to generate images with labels of cats/dogs
train_dir, #directory from which the data is being generated
target_size=(150, 150), #resizes the input image to 150x150
batch_size=20, #the data will be generated in batches of 20 samples each
class_mode='binary') #the binary class_mode sets the 2 sub classes in sub directories of train_data as the labels of i/p data
validation_generator = test_datagen.flow_from_directory( #generates validation data from validation directory like above
validation_dir,
target_size=(150, 150),
batch_size=20,
class_mode='binary')
for data_batch, labels_batch in train_generator:
print('data batch shape:', data_batch.shape)
print('labels batch shape:', labels_batch.shape)
break
|
DeepLearningWithPython/06_CatsVsDogs/.ipynb_checkpoints/CatsVsDogsmain-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Releases Visualisation
import requests
import pandas as pd
import re
import json
from IPython.core.display import display, HTML
from modules.Network import *
display(HTML("<style>.container { width:70% !important; }</style>"))
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
# We get the json file with the information from
# https://wmwaredata.s3.us-east-2.amazonaws.com/releases.json
url = 'https://wmwaredata.s3.us-east-2.amazonaws.com/releases.json'
r = requests.get(url, allow_redirects=True)
open('data/releases.json', 'wb').write(r.content)
# +
df = pd.read_json('data/releases.json')
descriptions = []
for i in range(len(df)):
content = re.sub('<.*?>', '', df['release_name'].iloc[i])
des = wrap_by_word(content, n=5)
des = wrap_by_char(des, n=45)
descriptions.append(des)
df['description'] = descriptions
df = df.drop('release_name', axis=1)
df = df.rename(columns={'repo_repo': 'repo_name'})
df = df[['repo_name', 'repo_url', 'release_tag', 'release_date', 'description']]
df
# -
# ## Repo's Releases
# +
# CREATE JSON FILE WITH NODES AND EDGES
network = Network()
network.groups = ['Repo', 'Tag', 'Date', 'Description']
network.groupSettings = group_settings_R
for repo in set(list(df['repo_name'])):
# REPO
network.add_node(label=repo, group='Repo')
df1 = df[df['repo_name'] == repo].reset_index()
for i in range(len(df1)):
# TAG
tag = 'Tag: ' + df1['release_tag'].iloc[i]
network.add_node(label=tag, group='Tag', repeat_nodes=True)
network.add_edge(network.nodesIds[repo][0], network.nodesIds[tag][-1])
df2 = df1[df1['release_tag'] == tag].reset_index()
# DATE
d = df1['release_date'].iloc[i]
if d is not None:
d = d.split('T')
date = 'Date: '+ d[0] + '\nTime: ' + d[1][:-1]
else:
date = ''
network.add_node(label=date, group='Date', repeat_nodes=True)
network.add_edge(network.nodesIds[tag][-1], network.nodesIds[date][-1])
# DESCRIPTION
des = df1['description'].iloc[i]
if des != '':
network.add_node(label=des, group='Description', repeat_nodes=True)
network.add_edge(network.nodesIds[date][-1], network.nodesIds[des][-1])
network.save_to_json('data/releases_nodes_edges.json')
# + language="html"
# <div id="mynetwork"></div>
# + language="javascript"
# requirejs.config({
# paths: {
# vis: 'vis'
# }
# });
#
#
# require(['vis'], function(vis){
#
# var json = $.getJSON("data/releases_nodes_edges.json")
# .done(function(data){
# var data = {
# nodes: data.nodes,
# edges: data.edges
# };
# var network = new vis.Network(container, data, options);
# });
#
# var options = {
# width: '1000px',
# height: '800px',
# locale: 'en',
# physics: true,
# interaction: {
# hover:true,
# tooltipDelay: 300
# },
# layout: {
# randomSeed: 1,
# improvedLayout: true,
# }
# };
#
# var container = document.getElementById("mynetwork");
# });
|
upwork-devs/Pappaterra-Lucia/Releases Visualisation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wasiqrumaney/privacy/blob/master/notebooks/pate2017single.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="GovDUv_4rmZ0" colab_type="text"
# ###Creating directories
# + id="qeXV2eZVcKdC" colab_type="code" colab={}
# !mkdir data
# !mkdir models
# + [markdown] id="MiV7xWvEryi1" colab_type="text"
# ### Importing libraries
# + id="JHCqrf1Hr3_F" colab_type="code" colab={}
# %matplotlib inline
# # !pip install -q tf-nightly-2.0-preview
import gzip
import math
import os
import sys
import tarfile
import numpy as np
from scipy.io import loadmat as loadmat
from six.moves import cPickle as pickle
from six.moves import urllib
from six.moves import xrange
import tensorflow as tf
from datetime import datetime as dt
import time
import seaborn as sns
import matplotlib.pyplot as plt
# + [markdown] id="PNGjnBY0zd0r" colab_type="text"
# ### Flags
# + id="gnpMJ1E-zcyd" colab_type="code" colab={}
dataset = 'mnist'
epochs_per_decay = 350
nb_teachers = 10
# teacher_id = 0
dropout_seed = 123
batch_size = 128
nb_labels = 10
deeper = False
batch_size = 128
max_steps = 3000
log_device_placement = False
learning_rate = 5
MOVING_AVERAGE_DECAY = 0.9999 # The decay to use for the moving average.
LEARNING_RATE_DECAY_FACTOR = 0.1 # Learning rate decay factor.
train_dir = '/content/models'
data_dir = '/content/data'
stdnt_share = 1000
lap_scale = 10
teachers_dir = '/content/models'
teachers_max_steps = 3000
# + [markdown] id="IKjgCRnb07F2" colab_type="text"
# ### Code from utils.py
# + id="7Qa51ppr041i" colab_type="code" colab={}
def batch_indices(batch_nb, data_length, batch_size):
"""
This helper function computes a batch start and end index
:param batch_nb: the batch number
:param data_length: the total length of the data being parsed by batches
:param batch_size: the number of inputs in each batch
:return: pair of (start, end) indices
"""
# Batch start and end index
start = int(batch_nb * batch_size)
end = int((batch_nb + 1) * batch_size)
# When there are not enough inputs left, we reuse some to complete the batch
if end > data_length:
shift = end - data_length
start -= shift
end -= shift
return start, end
# + id="DCttCPwxDpS6" colab_type="code" colab={}
def accuracy(logits, labels):
"""
Return accuracy of the array of logits (or label predictions) wrt the labels
:param logits: this can either be logits, probabilities, or a single label
:param labels: the correct labels to match against
:return: the accuracy as a float
"""
assert len(logits) == len(labels)
if len(np.shape(logits)) > 1:
# Predicted labels are the argmax over axis 1
predicted_labels = np.argmax(logits, axis=1)
else:
# Input was already labels
assert len(np.shape(logits)) == 1
predicted_labels = logits
# Check against correct labels to compute correct guesses
correct = np.sum(predicted_labels == labels.reshape(len(labels)))
# Divide by number of labels to obtain accuracy
accuracy = float(correct) / len(labels)
# Return float value
return accuracy
# + [markdown] id="3uMyjaLHsbcR" colab_type="text"
# ### Downloading dataset
# + id="kH5AOMxzqY2L" colab_type="code" colab={}
file_urls = ['http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/train-labels-idx1-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-images-idx3-ubyte.gz',
'http://yann.lecun.com/exdb/mnist/t10k-labels-idx1-ubyte.gz',
]
# + id="1U0IkLGjsGNM" colab_type="code" colab={}
def maybe_download(file_urls, directory):
"""Download a set of files in temporary local folder."""
# This list will include all URLS of the local copy of downloaded files
result = []
# For each file of the dataset
for file_url in file_urls:
# Extract filename
filename = file_url.split('/')[-1]
# If downloading from GitHub, remove suffix ?raw=True from local filename
if filename.endswith("?raw=true"):
filename = filename[:-9]
# Deduce local file url
#filepath = os.path.join(directory, filename)
filepath = directory + '/' + filename
# Add to result list
result.append(filepath)
# Test if file already exists
if not tf.gfile.Exists(filepath):
def _progress(count, block_size, total_size):
sys.stdout.write('\r>> Downloading %s %.1f%%' % (filename,
float(count * block_size) / float(total_size) * 100.0))
sys.stdout.flush()
filepath, _ = urllib.request.urlretrieve(file_url, filepath, _progress)
print()
statinfo = os.stat(filepath)
print('Successfully downloaded', filename, statinfo.st_size, 'bytes.')
return result
# + id="3NEOBwWbstRS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="f98e9bf9-90ab-48cb-ef12-84368e44ae77"
local_urls = maybe_download(file_urls, '/content/data')
# + id="zgV_kSpHtqpm" colab_type="code" colab={}
def extract_mnist_data(filename, num_images, image_size, pixel_depth):
"""
Extract the images into a 4D tensor [image index, y, x, channels].
Values are rescaled from [0, 255] down to [-0.5, 0.5].
"""
if not tf.gfile.Exists(filename+'.npy'):
with gzip.open(filename) as bytestream:
bytestream.read(16)
buf = bytestream.read(image_size * image_size * num_images)
data = np.frombuffer(buf, dtype=np.uint8).astype(np.float32)
data = (data - (pixel_depth / 2.0)) / pixel_depth
data = data.reshape(num_images, image_size, image_size, 1)
np.save(filename, data)
return data
else:
with tf.gfile.Open(filename+'.npy', mode='rb') as file_obj:
return np.load(file_obj)
def extract_mnist_labels(filename, num_images):
"""
Extract the labels into a vector of int64 label IDs.
"""
if not tf.gfile.Exists(filename+'.npy'):
with gzip.open(filename) as bytestream:
bytestream.read(8)
buf = bytestream.read(1 * num_images)
labels = np.frombuffer(buf, dtype=np.uint8).astype(np.int32)
np.save(filename, labels)
return labels
else:
with tf.gfile.Open(filename+'.npy', mode='rb') as file_obj:
return np.load(file_obj)
# + id="wzuF-jCSt8PV" colab_type="code" colab={}
# Extract it into np arrays.
train_data = extract_mnist_data(local_urls[0], 60000, 28, 1)
train_labels = extract_mnist_labels(local_urls[1], 60000)
test_data = extract_mnist_data(local_urls[2], 10000, 28, 1)
test_labels = extract_mnist_labels(local_urls[3], 10000)
# + id="hprJ-5oNuAmm" colab_type="code" colab={}
def partition_dataset(data, labels, nb_teachers, teacher_id):
"""
Simple partitioning algorithm that returns the right portion of the data
needed by a given teacher out of a certain nb of teachers
Args:
data: input data to be partitioned
labels: output data to be partitioned
nb_teachers: number of teachers in the ensemble (affects size of each
partition)
teacher_id: id of partition to retrieve
"""
# Sanity check
assert len(data) == len(labels)
assert int(teacher_id) < int(nb_teachers)
# This will floor the possible number of batches
batch_len = int(len(data) / nb_teachers)
# Compute start, end indices of partition
start = teacher_id * batch_len
end = (teacher_id+1) * batch_len
# Slice partition off
partition_data = data[start:end]
partition_labels = labels[start:end]
return partition_data, partition_labels
# + [markdown] id="V42M5EKZONjX" colab_type="text"
# ### Tensorboard
# + id="C1DQ6Z7YOdxC" colab_type="code" colab={}
# Delete any old logs.... be smart while using this
% rm -rf /content/logs/
# + id="NuMs_zF_yjV8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0c82b3b0-072a-4bf4-ddd0-11ee680c826b"
# Install latest Tensorflow build
# !pip install -q tf-nightly-2.0-preview
import tensorflow as tf
# %load_ext tensorboard
# + id="v5yhxAkiOleF" colab_type="code" colab={}
% mkdir -p '/content/logs/tensorboard/teacher/'
# + id="sqjdT2udO_2M" colab_type="code" colab={}
% mkdir -p '/content/logs/tensorboard/student/'
# + id="v29Gv0nuOzPP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 215} outputId="31f30973-3635-47cf-ef71-f1f4904f7c71"
import datetime
current_time = str(dt.now().timestamp())
teacher_log_dir = '/content/logs/tensorboard/teacher/' + current_time
student_log_dir = '/content/logs/tensorboard/student/' + current_time
teacher_summary_writer = summary.FileWriter(teacher_log_dir)
student_summary_writer = summary.FileWriter(student_log_dir)
# + id="ZEfTvzKAPRwa" colab_type="code" colab={}
# + [markdown] id="MIulZ9vrOSTB" colab_type="text"
# ### DeepCNN
# + id="rE5xycGy_haw" colab_type="code" colab={}
def _variable_on_cpu(name, shape, initializer):
"""Helper to create a Variable stored on CPU memory.
Args:
name: name of the variable
shape: list of ints
initializer: initializer for Variable
Returns:
Variable Tensor
"""
with tf.device('/cpu:0'):
var = tf.get_variable(name, shape, initializer=initializer)
return var
# + id="HEnBdvWXy2at" colab_type="code" colab={}
def _variable_with_weight_decay(name, shape, stddev, wd):
"""Helper to create an initialized Variable with weight decay.
Note that the Variable is initialized with a truncated normal distribution.
A weight decay is added only if one is specified.
Args:
name: name of the variable
shape: list of ints
stddev: standard deviation of a truncated Gaussian
wd: add L2Loss weight decay multiplied by this float. If None, weight
decay is not added for this Variable.
Returns:
Variable Tensor
"""
var = _variable_on_cpu(name, shape,
tf.truncated_normal_initializer(stddev=stddev))
if wd is not None:
weight_decay = tf.multiply(tf.nn.l2_loss(var), wd, name='weight_loss')
tf.add_to_collection('losses', weight_decay)
return var
# + id="ZV68edrZB5Rl" colab_type="code" colab={}
def inference(images, dropout=False):
"""Build the CNN model.
Args:
images: Images returned from distorted_inputs() or inputs().
dropout: Boolean controlling whether to use dropout or not
Returns:
Logits
"""
first_conv_shape = [5, 5, 1, 64]
# conv1
with tf.variable_scope('conv1') as scope:
kernel = _variable_with_weight_decay('weights',
shape=first_conv_shape,
stddev=1e-4,
wd=0.0)
conv = tf.nn.conv2d(images, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [64], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv1 = tf.nn.relu(bias, name=scope.name)
if dropout:
conv1 = tf.nn.dropout(conv1, 0.3, seed=dropout_seed)
# pool1
pool1 = tf.nn.max_pool(conv1,
ksize=[1, 3, 3, 1],
strides=[1, 2, 2, 1],
padding='SAME',
name='pool1')
# norm1
norm1 = tf.nn.lrn(pool1,
4,
bias=1.0,
alpha=0.001 / 9.0,
beta=0.75,
name='norm1')
# conv2
with tf.variable_scope('conv2') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[5, 5, 64, 128],
stddev=1e-4,
wd=0.0)
conv = tf.nn.conv2d(norm1, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [128], tf.constant_initializer(0.1))
bias = tf.nn.bias_add(conv, biases)
conv2 = tf.nn.relu(bias, name=scope.name)
if dropout:
conv2 = tf.nn.dropout(conv2, 0.3, seed=dropout_seed)
# norm2
norm2 = tf.nn.lrn(conv2,
4,
bias=1.0,
alpha=0.001 / 9.0,
beta=0.75,
name='norm2')
# pool2
pool2 = tf.nn.max_pool(norm2,
ksize=[1, 3, 3, 1],
strides=[1, 2, 2, 1],
padding='SAME',
name='pool2')
# local3
with tf.variable_scope('local3') as scope:
# Move everything into depth so we can perform a single matrix multiply.
reshape = tf.reshape(pool2, [batch_size, -1])
dim = reshape.get_shape()[1].value
weights = _variable_with_weight_decay('weights',
shape=[dim, 384],
stddev=0.04,
wd=0.004)
biases = _variable_on_cpu('biases', [384], tf.constant_initializer(0.1))
local3 = tf.nn.relu(tf.matmul(reshape, weights) + biases, name=scope.name)
if dropout:
local3 = tf.nn.dropout(local3, 0.5, seed=dropout_seed)
# local4
with tf.variable_scope('local4') as scope:
weights = _variable_with_weight_decay('weights',
shape=[384, 192],
stddev=0.04,
wd=0.004)
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.1))
local4 = tf.nn.relu(tf.matmul(local3, weights) + biases, name=scope.name)
if dropout:
local4 = tf.nn.dropout(local4, 0.5, seed=dropout_seed)
# compute logits
with tf.variable_scope('softmax_linear') as scope:
weights = _variable_with_weight_decay('weights',
[192, nb_labels],
stddev=1/192.0,
wd=0.0)
biases = _variable_on_cpu('biases',
[nb_labels],
tf.constant_initializer(0.0))
logits = tf.add(tf.matmul(local4, weights), biases, name=scope.name)
return logits
# + id="TafRCATMzE1f" colab_type="code" colab={}
def inference_deeper(images, dropout=False):
"""Build a deeper CNN model.
Args:
images: Images returned from distorted_inputs() or inputs().
dropout: Boolean controlling whether to use dropout or not
Returns:
Logits
"""
if dataset == 'mnist':
first_conv_shape = [3, 3, 1, 96]
else:
first_conv_shape = [3, 3, 3, 96]
# conv1
with tf.variable_scope('conv1') as scope:
kernel = _variable_with_weight_decay('weights',
shape=first_conv_shape,
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(images, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [96], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv1 = tf.nn.relu(bias, name=scope.name)
# conv2
with tf.variable_scope('conv2') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[3, 3, 96, 96],
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(conv1, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [96], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv2 = tf.nn.relu(bias, name=scope.name)
# conv3
with tf.variable_scope('conv3') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[3, 3, 96, 96],
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(conv2, kernel, [1, 2, 2, 1], padding='SAME')
biases = _variable_on_cpu('biases', [96], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv3 = tf.nn.relu(bias, name=scope.name)
if dropout:
conv3 = tf.nn.dropout(conv3, 0.5, seed=dropout_seed)
# conv4
with tf.variable_scope('conv4') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[3, 3, 96, 192],
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(conv3, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv4 = tf.nn.relu(bias, name=scope.name)
# conv5
with tf.variable_scope('conv5') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[3, 3, 192, 192],
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(conv4, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv5 = tf.nn.relu(bias, name=scope.name)
# conv6
with tf.variable_scope('conv6') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[3, 3, 192, 192],
stddev=0.05,
wd=0.0)
conv = tf.nn.conv2d(conv5, kernel, [1, 2, 2, 1], padding='SAME')
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.0))
bias = tf.nn.bias_add(conv, biases)
conv6 = tf.nn.relu(bias, name=scope.name)
if dropout:
conv6 = tf.nn.dropout(conv6, 0.5, seed=dropout_seed)
# conv7
with tf.variable_scope('conv7') as scope:
kernel = _variable_with_weight_decay('weights',
shape=[5, 5, 192, 192],
stddev=1e-4,
wd=0.0)
conv = tf.nn.conv2d(conv6, kernel, [1, 1, 1, 1], padding='SAME')
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.1))
bias = tf.nn.bias_add(conv, biases)
conv7 = tf.nn.relu(bias, name=scope.name)
# local1
with tf.variable_scope('local1') as scope:
# Move everything into depth so we can perform a single matrix multiply.
reshape = tf.reshape(conv7, [batch_size, -1])
dim = reshape.get_shape()[1].value
weights = _variable_with_weight_decay('weights',
shape=[dim, 192],
stddev=0.05,
wd=0)
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.1))
local1 = tf.nn.relu(tf.matmul(reshape, weights) + biases, name=scope.name)
# local2
with tf.variable_scope('local2') as scope:
weights = _variable_with_weight_decay('weights',
shape=[192, 192],
stddev=0.05,
wd=0)
biases = _variable_on_cpu('biases', [192], tf.constant_initializer(0.1))
local2 = tf.nn.relu(tf.matmul(local1, weights) + biases, name=scope.name)
if dropout:
local2 = tf.nn.dropout(local2, 0.5, seed=dropout_seed)
# compute logits
with tf.variable_scope('softmax_linear') as scope:
weights = _variable_with_weight_decay('weights',
[192, nb_labels],
stddev=0.05,
wd=0.0)
biases = _variable_on_cpu('biases',
[nb_labels],
tf.constant_initializer(0.0))
logits = tf.add(tf.matmul(local2, weights), biases, name=scope.name)
return logits
# + id="XrJ9OWe2zL4_" colab_type="code" colab={}
def loss_fun(logits, labels):
"""Add L2Loss to all the trainable variables.
Add summary for "Loss" and "Loss/avg".
Args:
logits: Logits from inference().
labels: Labels from distorted_inputs or inputs(). 1-D tensor
of shape [batch_size]
distillation: if set to True, use probabilities and not class labels to
compute softmax loss
Returns:
Loss tensor of type float.
"""
# Calculate the cross entropy between labels and predictions
labels = tf.cast(labels, tf.int64)
cross_entropy = tf.nn.sparse_softmax_cross_entropy_with_logits(
logits=logits, labels=labels, name='cross_entropy_per_example')
# Calculate the average cross entropy loss across the batch.
cross_entropy_mean = tf.reduce_mean(cross_entropy, name='cross_entropy')
# Add to TF collection for losses
tf.add_to_collection('losses', cross_entropy_mean)
# The total loss is defined as the cross entropy loss plus all of the weight
# decay terms (L2 loss).
return tf.add_n(tf.get_collection('losses'), name='total_loss')
# + id="pc0XzPLI0iN3" colab_type="code" colab={}
def moving_av(total_loss):
"""
Generates moving average for all losses
Args:
total_loss: Total loss from loss().
Returns:
loss_averages_op: op for generating moving averages of losses.
"""
# Compute the moving average of all individual losses and the total loss.
loss_averages = tf.train.ExponentialMovingAverage(0.9, name='avg')
losses = tf.get_collection('losses')
loss_averages_op = loss_averages.apply(losses + [total_loss])
return loss_averages_op
# + id="TBOrj8spBoaW" colab_type="code" colab={}
def train_op_fun(total_loss, global_step):
"""Train model.
Create an optimizer and apply to all trainable variables. Add moving
average for all trainable variables.
Args:
total_loss: Total loss from loss().
global_step: Integer Variable counting the number of training steps
processed.
Returns:
train_op: op for training.
"""
# Variables that affect learning rate.
nb_ex_per_train_epoch = int(60000 / nb_teachers)
num_batches_per_epoch = nb_ex_per_train_epoch / batch_size
decay_steps = int(num_batches_per_epoch * epochs_per_decay)
initial_learning_rate = float(learning_rate) / 100.0
# Decay the learning rate exponentially based on the number of steps.
lr = tf.train.exponential_decay(initial_learning_rate,
global_step,
decay_steps,
LEARNING_RATE_DECAY_FACTOR,
staircase=True)
tf.summary.scalar('learning_rate', lr)
# Generate moving averages of all losses and associated summaries.
loss_averages_op = moving_av(total_loss)
# Compute gradients.
with tf.control_dependencies([loss_averages_op]):
opt = tf.train.GradientDescentOptimizer(lr)
grads = opt.compute_gradients(total_loss)
# Apply gradients.
apply_gradient_op = opt.apply_gradients(grads, global_step=global_step)
# Add histograms for trainable variables.
for var in tf.trainable_variables():
tf.summary.histogram(var.op.name, var)
# Track the moving averages of all trainable variables.
variable_averages = tf.train.ExponentialMovingAverage(
MOVING_AVERAGE_DECAY, global_step)
variables_averages_op = variable_averages.apply(tf.trainable_variables())
with tf.control_dependencies([apply_gradient_op, variables_averages_op]):
train_op = tf.no_op(name='train')
return train_op
# + id="D8nyFB4cBlK5" colab_type="code" colab={}
def _input_placeholder():
"""
This helper function declares a TF placeholder for the graph input data
:return: TF placeholder for the graph input data
"""
image_size = 28
num_channels = 1
# Declare data placeholder
train_node_shape = (batch_size, image_size, image_size, num_channels)
return tf.placeholder(tf.float32, shape=train_node_shape)
# + id="m77TTK_CBbsr" colab_type="code" colab={}
def train(images, labels, ckpt_path, dropout=False):
"""
This function contains the loop that actually trains the model.
:param images: a numpy array with the input data
:param labels: a numpy array with the output labels
:param ckpt_path: a path (including name) where model checkpoints are saved
:param dropout: Boolean, whether to use dropout or not
:return: True if everything went well
"""
# Check training data
assert len(images) == len(labels)
assert images.dtype == np.float32
assert labels.dtype == np.int32
# Set default TF graph
with tf.Graph().as_default():
global_step = tf.Variable(0, trainable=False)
# Declare data placeholder
train_data_node = _input_placeholder()
# Create a placeholder to hold labels
train_labels_shape = (batch_size,)
train_labels_node = tf.placeholder(tf.int32, shape=train_labels_shape)
print("Done Initializing Training Placeholders")
# Build a Graph that computes the logits predictions from the placeholder
if deeper:
logits = inference_deeper(train_data_node, dropout=dropout)
else:
logits = inference(train_data_node, dropout=dropout)
# Calculate loss
loss = loss_fun(logits, train_labels_node)
# loss_scalar = tf.summary.scalar("loss",loss)
# Build a Graph that trains the model with one batch of examples and
# updates the model parameters.
train_op = train_op_fun(loss, global_step)
# Create a saver.
saver = tf.train.Saver(tf.global_variables())
print("Graph constructed and saver created")
# Build an initialization operation to run below.
init = tf.global_variables_initializer()
# Create and init sessions
sess = tf.Session(config=tf.ConfigProto(log_device_placement=log_device_placement)) #NOLINT(long-line)
sess.run(init)
writer = tf.summary.FileWriter('board_beginner') # create writer
writer.add_graph(sess.graph)
print("Session ready, beginning training loop")
# Initialize the number of batches
data_length = len(images)
nb_batches = math.ceil(data_length / batch_size)
for step in xrange(max_steps):
# for debug, save start time
start_time = time.time()
# Current batch number
batch_nb = step % nb_batches
# Current batch start and end indices
start, end = batch_indices(batch_nb, data_length, batch_size)
# Prepare dictionnary to feed the session with
feed_dict = {train_data_node: images[start:end],
train_labels_node: labels[start:end]}
# Run training step
_, loss_value = sess.run([train_op, loss], feed_dict=feed_dict)
loss_scalar = tf.summary.scalar("loss_value",loss_value)
# with teacher_summary_writer.as_default():
# summary.scalar('loss', loss_value, step=step)
# teacher_summary_writer.scalar('loss', loss_value, step=step)
# Compute duration of training step
duration = time.time() - start_time
# Sanity check
assert not np.isnan(loss_value), 'Model diverged with loss = NaN'
# Echo loss once in a while
if step % 100 == 0:
sum1 = sess.run(loss_scalar, feed_dict=feed_dict)
writer.add_summary(sum1,step)
num_examples_per_step = batch_size
examples_per_sec = num_examples_per_step / duration
sec_per_batch = float(duration)
format_str = ('%s: step %d, loss = %.2f (%.1f examples/sec; %.3f '
'sec/batch)')
print (format_str % (dt.now(), step, loss_value,
examples_per_sec, sec_per_batch))
# Save the model checkpoint periodically.
if step % 1000 == 0 or (step + 1) == max_steps:
saver.save(sess, ckpt_path, global_step=step)
saver.save(sess, "model_beginner")
return True
# + id="_3WzkMVl3AKf" colab_type="code" colab={}
def softmax_preds(images, ckpt_path, return_logits=False):
"""
Compute softmax activations (probabilities) with the model saved in the path
specified as an argument
:param images: a np array of images
:param ckpt_path: a TF model checkpoint
:param logits: if set to True, return logits instead of probabilities
:return: probabilities (or logits if logits is set to True)
"""
# Compute nb samples and deduce nb of batches
data_length = len(images)
nb_batches = math.ceil(len(images) / batch_size)
# Declare data placeholder
train_data_node = _input_placeholder()
# Build a Graph that computes the logits predictions from the placeholder
if deeper:
logits = inference_deeper(train_data_node)
else:
logits = inference(train_data_node)
if return_logits:
# We are returning the logits directly (no need to apply softmax)
output = logits
else:
# Add softmax predictions to graph: will return probabilities
output = tf.nn.softmax(logits)
# Restore the moving average version of the learned variables for eval.
variable_averages = tf.train.ExponentialMovingAverage(MOVING_AVERAGE_DECAY)
variables_to_restore = variable_averages.variables_to_restore()
saver = tf.train.Saver(variables_to_restore)
# Will hold the result
preds = np.zeros((data_length, nb_labels), dtype=np.float32)
# Create TF session
with tf.Session() as sess:
# Restore TF session from checkpoint file
saver.restore(sess, ckpt_path)
# Parse data by batch
for batch_nb in xrange(0, int(nb_batches+1)):
# Compute batch start and end indices
start, end = batch_indices(batch_nb, data_length, batch_size)
# Prepare feed dictionary
feed_dict = {train_data_node: images[start:end]}
# Run session ([0] because run returns a batch with len 1st dim == 1)
preds[start:end, :] = sess.run([output], feed_dict=feed_dict)[0]
# Reset graph to allow multiple calls
tf.reset_default_graph()
return preds
# + [markdown] id="jonBN3Zi2Xk1" colab_type="text"
# # Teacher training
# + [markdown] id="CTHP3X4v2e8T" colab_type="text"
# ### Teacher 0
# + id="VokxvHeavfc9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 581} outputId="1317a68b-0aff-42ff-d314-155ca361643a"
teacher_id = 0
tf.summary.FileWriterCache.clear()
import datetime
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
accuracy_scalar = tf.summary.scalar("accuracy",precision)
print('Precision of teacher after training: ' + str(precision))
# + id="JEf2oQ8dLMz0" colab_type="code" colab={}
# %load_ext tensorboard
# %tensorboard --logdir /content/board_beginner
# + [markdown] id="vokjSaOP7vs6" colab_type="text"
# ### Teacher 1
# + id="icAdkyw-7N3k" colab_type="code" colab={}
teacher_id = 1
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="n1ekuWH-74if" colab_type="text"
# ### Teacher 2
# + id="DIkgQYa87NRB" colab_type="code" colab={}
teacher_id = 2
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="hhu4neMR788G" colab_type="text"
# ### Teacher 3
# + id="sqFGomhz7Mmy" colab_type="code" colab={}
teacher_id = 3
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="ZliZISiR8A_Q" colab_type="text"
# ### Teacher 4
# + id="jGMg49pw7L32" colab_type="code" colab={}
teacher_id = 4
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="ibdBqPw68D9L" colab_type="text"
# ### Teacher 5
# + id="2z2zKGFe7LOG" colab_type="code" colab={}
teacher_id = 5
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="zaOjsq3j8G7N" colab_type="text"
# ### Teacher 6
# + id="oC7tpH_p7Kj9" colab_type="code" colab={}
teacher_id = 6
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="Roax8U6k8KF5" colab_type="text"
# ### Teacher 7
# + id="nZGhi1Kz7J8J" colab_type="code" colab={}
teacher_id = 7
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="Vuj4H9La8PVp" colab_type="text"
# ### Teacher 8
# + id="BNgQYu5u7JU1" colab_type="code" colab={}
teacher_id = 8
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="1L9qsyXz8k8F" colab_type="text"
# ### Teacher 9
# + id="sCP9T_QU7IuB" colab_type="code" colab={}
teacher_id = 9
# Retrieve subset of data for this teacher
data, labels = partition_dataset(train_data, train_labels, nb_teachers, teacher_id)
print("Length of training data: " + str(len(labels)))
# Define teacher checkpoint filename and full path
if deeper:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt'
else:
filename = str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt'
ckpt_path = train_dir + '/' + str(dataset) + '_' + filename
# Perform teacher training
assert train(data, labels, ckpt_path)
# Append final step value to checkpoint for evaluation
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Retrieve teacher probability estimates on the test data
teacher_preds = softmax_preds(test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(teacher_preds, test_labels)
print('Precision of teacher after training: ' + str(precision))
# + [markdown] id="9nH4nzlB1WOZ" colab_type="text"
# # Student
# + [markdown] id="BQ4EjePM-xRf" colab_type="text"
# ### Aggregation
# + id="EKleBSeV-wFH" colab_type="code" colab={}
def labels_from_probs(probs):
"""
Helper function: computes argmax along last dimension of array to obtain
labels (max prob or max logit value)
:param probs: numpy array where probabilities or logits are on last dimension
:return: array with same shape as input besides last dimension with shape 1
now containing the labels
"""
# Compute last axis index
last_axis = len(np.shape(probs)) - 1
# Label is argmax over last dimension
labels = np.argmax(probs, axis=last_axis)
# Return as np.int32
return np.asarray(labels, dtype=np.int32)
# + id="k-8Y9oZp_4Eu" colab_type="code" colab={}
def noisy_max(logits, lap_scale, return_clean_votes=False):
"""
This aggregation mechanism takes the softmax/logit output of several models
resulting from inference on identical inputs and computes the noisy-max of
the votes for candidate classes to select a label for each sample: it
adds Laplacian noise to label counts and returns the most frequent label.
:param logits: logits or probabilities for each sample
:param lap_scale: scale of the Laplacian noise to be added to counts
:param return_clean_votes: if set to True, also returns clean votes (without
Laplacian noise). This can be used to perform the
privacy analysis of this aggregation mechanism.
:return: pair of result and (if clean_votes is set to True) the clean counts
for each class per sample and the original labels produced by
the teachers.
"""
# Compute labels from logits/probs and reshape array properly
labels = labels_from_probs(logits)
labels_shape = np.shape(labels)
labels = labels.reshape((labels_shape[0], labels_shape[1]))
# Initialize array to hold final labels
result = np.zeros(int(labels_shape[1]))
if return_clean_votes:
# Initialize array to hold clean votes for each sample
clean_votes = np.zeros((int(labels_shape[1]), 10))
# Parse each sample
for i in xrange(int(labels_shape[1])):
# Count number of votes assigned to each class
label_counts = np.bincount(labels[:, i], minlength=10)
if return_clean_votes:
# Store vote counts for export
clean_votes[i] = label_counts
# Cast in float32 to prepare before addition of Laplacian noise
label_counts = np.asarray(label_counts, dtype=np.float32)
# Sample independent Laplacian noise for each class
for item in xrange(10):
label_counts[item] += np.random.laplace(loc=0.0, scale=float(lap_scale))
# Result is the most frequent label
result[i] = np.argmax(label_counts)
# Cast labels to np.int32 for compatibility with deep_cnn.py feed dictionaries
result = np.asarray(result, dtype=np.int32)
if return_clean_votes:
# Returns several array, which are later saved:
# result: labels obtained from the noisy aggregation
# clean_votes: the number of teacher votes assigned to each sample and class
# labels: the labels assigned by teachers (before the noisy aggregation)
return result, clean_votes, labels
else:
# Only return labels resulting from noisy aggregation
return result
# + id="d_qkrnSc_8yQ" colab_type="code" colab={}
def aggregation_most_frequent(logits):
"""
This aggregation mechanism takes the softmax/logit output of several models
resulting from inference on identical inputs and computes the most frequent
label. It is deterministic (no noise injection like noisy_max() above.
:param logits: logits or probabilities for each sample
:return:
"""
# Compute labels from logits/probs and reshape array properly
labels = labels_from_probs(logits)
labels_shape = np.shape(labels)
labels = labels.reshape((labels_shape[0], labels_shape[1]))
# Initialize array to hold final labels
result = np.zeros(int(labels_shape[1]))
# Parse each sample
for i in xrange(int(labels_shape[1])):
# Count number of votes assigned to each class
label_counts = np.bincount(labels[:, i], minlength=10)
label_counts = np.asarray(label_counts, dtype=np.int32)
# Result is the most frequent label
result[i] = np.argmax(label_counts)
return np.asarray(result, dtype=np.int32)
# + id="CGhvvEM4DOsI" colab_type="code" colab={}
# + [markdown] id="D63DOqVX-178" colab_type="text"
# ### Student training
# + id="c3gBl9PryY-A" colab_type="code" colab={}
def ensemble_preds(dataset, nb_teachers, stdnt_data):
"""
Given a dataset, a number of teachers, and some input data, this helper
function queries each teacher for predictions on the data and returns
all predictions in a single array. (That can then be aggregated into
one single prediction per input using aggregation.py (cf. function
prepare_student_data() below)
:param dataset: string corresponding to mnist, cifar10, or svhn
:param nb_teachers: number of teachers (in the ensemble) to learn from
:param stdnt_data: unlabeled student training data
:return: 3d array (teacher id, sample id, probability per class)
"""
# Compute shape of array that will hold probabilities produced by each
# teacher, for each training point, and each output class
result_shape = (nb_teachers, len(stdnt_data), nb_labels)
# Create array that will hold result
result = np.zeros(result_shape, dtype=np.float32)
# Get predictions from each teacher
for teacher_id in xrange(nb_teachers):
# Compute path of checkpoint file for teacher model with ID teacher_id
if deeper:
ckpt_path = teachers_dir + '/' + str(dataset) + '_' + str(nb_teachers) + '_teachers_' + str(teacher_id) + '_deep.ckpt-' + str(teachers_max_steps - 1) #NOLINT(long-line)
else:
ckpt_path = teachers_dir + '/' + str(dataset) + '_' + str(nb_teachers) + '_teachers_' + str(teacher_id) + '.ckpt-' + str(teachers_max_steps - 1) # NOLINT(long-line)
# Get predictions on our training data and store in result array
result[teacher_id] = softmax_preds(stdnt_data, ckpt_path)
# This can take a while when there are a lot of teachers so output status
print("Computed Teacher " + str(teacher_id) + " softmax predictions")
return result
# + id="GYro3WOf4cwr" colab_type="code" colab={}
def prepare_student_data(dataset, nb_teachers, save=False):
"""
Takes a dataset name and the size of the teacher ensemble and prepares
training data for the student model, according to parameters indicated
in flags above.
:param dataset: string corresponding to mnist, cifar10, or svhn
:param nb_teachers: number of teachers (in the ensemble) to learn from
:param save: if set to True, will dump student training labels predicted by
the ensemble of teachers (with Laplacian noise) as npy files.
It also dumps the clean votes for each class (without noise) and
the labels assigned by teachers
:return: pairs of (data, labels) to be used for student training and testing
"""
# Make sure there is data leftover to be used as a test set
assert stdnt_share < len(test_data)
# Prepare [unlabeled] student training data (subset of test set)
stdnt_data = test_data[:stdnt_share]
# Compute teacher predictions for student training data
teachers_preds = ensemble_preds(dataset, nb_teachers, stdnt_data)
# Aggregate teacher predictions to get student training labels
if not save:
stdnt_labels = noisy_max(teachers_preds, lap_scale)
else:
# Request clean votes and clean labels as well
stdnt_labels, clean_votes, labels_for_dump = noisy_max(teachers_preds, lap_scale, return_clean_votes=True) #NOLINT(long-line)
# Prepare filepath for numpy dump of clean votes
filepath = data_dir + "/" + str(dataset) + '_' + str(nb_teachers) + '_student_clean_votes_lap_' + str(lap_scale) + '.npy' # NOLINT(long-line)
# Prepare filepath for numpy dump of clean labels
filepath_labels = data_dir + "/" + str(dataset) + '_' + str(nb_teachers) + '_teachers_labels_lap_' + str(lap_scale) + '.npy' # NOLINT(long-line)
# Dump clean_votes array
with tf.gfile.Open(filepath, mode='w') as file_obj:
np.save(file_obj, clean_votes)
# Dump labels_for_dump array
with tf.gfile.Open(filepath_labels, mode='w') as file_obj:
np.save(file_obj, labels_for_dump)
# Print accuracy of aggregated labels
ac_ag_labels = accuracy(stdnt_labels, test_labels[:stdnt_share])
print("Accuracy of the aggregated labels: " + str(ac_ag_labels))
# Store unused part of test set for use as a test set after student training
stdnt_test_data = test_data[stdnt_share:]
stdnt_test_labels = test_labels[stdnt_share:]
if save:
# Prepare filepath for numpy dump of labels produced by noisy aggregation
filepath = data_dir + "/" + str(dataset) + '_' + str(nb_teachers) + '_student_labels_lap_' + str(lap_scale) + '.npy' #NOLINT(long-line)
# Dump student noisy labels array
with tf.gfile.Open(filepath, mode='w') as file_obj:
np.save(file_obj, stdnt_labels)
return stdnt_data, stdnt_labels, stdnt_test_data, stdnt_test_labels
# + id="RbBe0m_34klp" colab_type="code" colab={}
# Call helper function to prepare student data using teacher predictions
stdnt_dataset = prepare_student_data(dataset, nb_teachers, save=True)
# Unpack the student dataset
stdnt_data, stdnt_labels, stdnt_test_data, stdnt_test_labels = stdnt_dataset
# Prepare checkpoint filename and path
if deeper:
ckpt_path = train_dir + '/' + str(dataset) + '_' + str(nb_teachers) + '_student_deeper.ckpt' #NOLINT(long-line)
else:
ckpt_path = train_dir + '/' + str(dataset) + '_' + str(nb_teachers) + '_student.ckpt' # NOLINT(long-line)
# Start student training
assert train(stdnt_data, stdnt_labels, ckpt_path)
# Compute final checkpoint name for student (with max number of steps)
ckpt_path_final = ckpt_path + '-' + str(max_steps - 1)
# Compute student label predictions on remaining chunk of test set
student_preds = softmax_preds(stdnt_test_data, ckpt_path_final)
# Compute teacher accuracy
precision = accuracy(student_preds, stdnt_test_labels)
print('Precision of student after training: ' + str(precision))
# + id="Ypl_vpp5Jw92" colab_type="code" colab={}
|
notebooks/pate2017single.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true slideshow={"slide_type": "slide"}
# # Second Project
# + [markdown] hidden=true slideshow={"slide_type": "fragment"}
# In this project, you will implement a debugging reducer able to reduce a Python program containing a specific property down to a syntactically correct Python program that *only* contains said property.
# To do this, you will be given several Python parsers checking for specific properties as well as Python programs containing these properties.
#
# The time frame for this project is **2 weeks**, and the Deadline is **January 15th 23:59**.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "skip"}
import bookutils
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "skip"}
import inspect
import ast
import astor
# + slideshow={"slide_type": "skip"}
from bookutils import show_ast
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "skip"}
from ExpectError import ExpectError
# + slideshow={"slide_type": "fragment"}
# ignore
from typing import Any, Optional, Set
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "slide"}
# ## Description
#
# Imagine we want to create a new compiler for Python programs which transforms the code into a binary executable.
# To this end, at first we need to implement a parser that parses a given program into a set of individual statements which can then be converted to the bytecode.
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### Parsing Python
#
# If we want to parse a programming language like Python, it makes sense to parse the code into an abstract syntax tree (AST) and work with the AST from this point onward.
# The reason we might want to do this, is to preserve the syntactic integrity of our program.
# Let's start with parsing a simple Python function (`foo`) into an AST using `ast.parse()` and printing it.
# + slideshow={"slide_type": "subslide"}
def foo(a, b): # type: ignore
"""
Checks whether a and b are true
"""
if a and b:
return 1
return 0
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# We take a string representation of `foo()`...
# + slideshow={"slide_type": "fragment"}
source = inspect.getsource(foo)
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
print(source)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# ...and convert it to the AST:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
source_ast = ast.parse(source)
show_ast(source_ast)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# We can see how the code is structured by inspecting the AST shown above.
# + [markdown] slideshow={"slide_type": "fragment"}
# To learn the structure of an AST, have a look at the [official Python `ast` reference](http://docs.python.org/3/library/ast) for a list of AST nodes and their attributes.
# This reference is complete, but a bit terse; the ["Green Tree Snakes missing Python AST docs"](https://greentreesnakes.readthedocs.io/en/latest/manipulating.html) provide a good manual on how to modify the tree.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# If we want to inspect certain elements of the source code, we can now use the `ast.NodeVisitor` to visit every node in the AST.
# This can be done by extending this class and implementing *visit-methods* for each type of node in the tree we want to inspect.
# So, let's implement a parser that first parses the source code into an AST and then visits each node one after another.
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class SimpleParser(ast.NodeVisitor):
"""
A simple parser printing all nodes in order
"""
def parse(self, file: str) -> None:
"""
When the parse function is called, we parse the source code
into a tree and then start visiting all nodes.
"""
tree = ast.parse(source=file)
self.visit(tree)
def visit_FunctionDef(self, node: ast.FunctionDef) -> None:
print(node)
self.generic_visit(node)
def visit_arguments(self, node: ast.arguments) -> None:
print(node)
self.generic_visit(node)
def visit_If(self, node: ast.If) -> None:
print(node)
self.generic_visit(node)
def visit_Return(self, node: ast.Return) -> None:
print(node)
self.generic_visit(node)
def visit_arg(self, node: ast.arg) -> None:
print(node)
self.generic_visit(node)
def visit_Num(self, node: ast.Num) -> None:
print(node)
self.generic_visit(node)
def visit_Name(self, node: ast.Name) -> None:
print(node)
self.generic_visit(node)
def visit_BoolOp(self, node: ast.BoolOp) -> None:
print(node)
self.generic_visit(node)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# Using this `SimpleParser`, we can visit each node and print the node object itself.
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
SimpleParser().parse(source)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# Now imagine, instead of just parsing the input and printing each node, we start to process the information present in each node.
# To notify the user if anything breaks, we introduce a `ParserException` that is thrown when we encounter a parsing error.
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
class ParserException(Exception):
pass
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# For instance, let's consider our parser has troubles processing boolean operators.
# We simulate this behavior by extending our parser to throw an exception whenever it encounters a `BoolOp` node.
#
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class SimpleParserPlus(SimpleParser):
"""
A simple parser printing all nodes in order until it reaches a BoolOp
"""
def visit_BoolOp(self, node: ast.BoolOp) -> None:
"""
If a BoolOp is encountered, we throw an exception
"""
raise ParserException("Something went wrong")
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
with ExpectError():
SimpleParserPlus().parse(source)
# + [markdown] slideshow={"slide_type": "subslide"}
# As we can see here, as soon as we encounter a boolean operation, an exception is thrown.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### Reducing Python
#
# Let's get back to our `SimpleParserPlus` example. This parser parses Python code into an AST and then iterates over all nodes. When it reaches a boolean operation, it encounters an error and crashes.
# Of course our example is a very small and artificial one. Furthermore, the input triggering our bug is rather small as well.
# However, in reality a similar situation might present itself to you but with a far less obvious error and a far larger input that causes it.
# In this case, instead of using the entire input, one might want to reduce the input down to only those parts actually causing the error.
# But how do we do that?
# We could, for instance, randomly remove some parts of the program.
# However, this would likely produce syntactically invalid Python code and hence prevent the parser from reaching the desired code location.
# So instead of just removing random parts of the code itself, we will manipulate the AST to preserve the correct syntax.
#
# How do we manipulate an AST produced by `ast.parse`? We can make use of the `ast.NodeTransformer`:
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class NodeReducer(ast.NodeTransformer):
def visit_If(self, node: ast.If) -> ast.AST:
"""
Instead of the `if` node return just its condition
"""
# Apply other transformations to the children first
super().generic_visit(node)
# We create an expression around the node test
new_node = ast.Expr(node.test)
return new_node
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# The `ast.NodeTransformer` again uses the visitor pattern to walk over all nodes.
# This time, however, we change the resulting AST. In our case, if we encounter an *if-statement* we only return the condition.
#
# (Keep in mind, `NodeReducer` changes the ast in place, so if you want to preserve the original tree, you need to copy it with `copy.deepcopy(tree)`)
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
NodeReducer().visit(source_ast)
show_ast(source_ast)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# As you can see, after running the `NodeReducer` there is no longer an *if-statement*, but only its condition.
# A more sophisticated NodeReducer is presented in the [Reducing Failure-Inducing Inputs chapter](https://www.debuggingbook.org/beta/html/DeltaDebugger.html) in the debugging book.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# To learn the structure of an AST, have a look at the [official Python `ast` reference](http://docs.python.org/3/library/ast) for a list of AST nodes and their attributes. This reference is complete, but a bit terse; the ["Green Tree Snakes missing Python AST docs"](https://greentreesnakes.readthedocs.io/en/latest/manipulating.html) provide a good manual on how to modify the tree.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Pro tip**: You can learn the expected structure of any Python fragment by invoking `ast.dump()` on any (parsed) AST.
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
demo_ast = ast.parse('''
if a == b:
c = d
''')
# + slideshow={"slide_type": "fragment"}
ast.dump(demo_ast)
# + [markdown] slideshow={"slide_type": "fragment"}
# The nice thing is that this not only reveals the AST structure, it also tells you how to construct it. If you take the very structure as produced by `ast.dump()` and evaluate it, you obtain the same syntax tree.
# + slideshow={"slide_type": "skip"}
from ast import Module, If, Compare, Load, Eq, Assign, Name, Store
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
ast_from_scratch = Module(body=[If(test=Compare(left=Name(id='a', ctx=Load()), ops=[Eq()], comparators=[Name(id='b', ctx=Load())]), body=[Assign(targets=[Name(id='c', ctx=Store())], value=Name(id='d', ctx=Load()))], orelse=[])])
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
print(astor.to_source(ast_from_scratch))
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# Hence, if you do not know what kind of AST fragment to produce, just parse its text with ``ast.parse()`` and study the expression produced by `ast.dump()`.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### Implementing a Reducing Debugger
#
# Having seen how one can access and transform AST structures using a `NodeReducer` in Python, we can now use that knowledge to implement a `ReducingDebugger`
# capable of reducing the failure inducing Python code down to only that parts of the code that actually triggers the failure.
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class ReducingDebugger:
def __init__(self, parser: SimpleParser) -> None:
"""
We initialize the DebuggingReducer with a parser,
as this is needed to verify whether the failure
is still triggered after the code transformation
"""
self.parser = parser
self.node_reducer = NodeReducer()
def minimize(self, code: str) -> str:
"""
This function takes some Python code as string, reduces
it using the NodeReducer defined earlier and returns the
reduced program.
"""
# Parse the code to a tree
code_ast = ast.parse(source=code)
# Use the visit-method of the NodeReducer to reduce if nodes down to their condition
new_code_ast = self.node_reducer.visit(code_ast)
# After we've updated nodes, we need to fix the tree
ast.fix_missing_locations(new_code_ast)
# Generate code from the reduced tree
new_code = astor.to_source(new_code_ast)
# Test, whether the error is still triggered by the reduced code
try:
self.parser.parse(new_code)
# No exception is thrown. This means the new_code does not
# trigger an error anymore. Therefore, we failed in
# reduction and return the initial code.
return code
except ParserException:
# The error is still triggered. Return the reduced code
return new_code
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# The `ReducingDebugger` implemented above takes some Python code as an input and replaces all if-statements with their conditions.
# Then, it checks whether the newly generated code still triggers the error. If not, it returns the initial code. If the exception is still triggered, it returns the reduced code.
# Let's apply it to the source code of our `foo` function:
# + slideshow={"slide_type": "fragment"}
reducing_debugger = ReducingDebugger(SimpleParserPlus())
new_source = reducing_debugger.minimize(source)
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
print(new_source)
# + pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
with ExpectError():
SimpleParserPlus().parse(new_source)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# As we can see, our `ReducingDebugger` reduces the original function to a smaller input, while still preserving the property of the code that triggers the same error as before.
# However, this reducer is very simple: it applies only one transformation to all `if` nodes at once.
# Instead, one can consider various transformations of the same node as well as transformations of various node types and apply them one by one (or in batches) to gradually reduce the code.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# Keep in mind, that the changes should be made in a way that the code is still accepted by a regular Python interpreter.
# Besides, the goal is to obtain a minimal program that still triggers the error.
# So, your debugging reducer should not simply reduce all possible nodes, but also regularly check whether the reduced code still triggers the failure.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# Therefore, you are required to implement certain modifications that can be done to the AST. In general, these transformations can be one of the following:
#
# 1. Delete a node,
# 2. Substitute a node with the new node (e.g., _pass_ node),
# 3. Substitute a node with all of its children,
# 4. Substitute a node with one of its children.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# More detailed, one can consider, for instance:
#
# 1. Replacing a `BoolOp` node by `True`.
# 2. Replacing a `BoolOp` node by `False`.
# 3. Replacing a `BoolOp` node by its left operand.
# 4. Replacing a `BoolOp` node by its right operand.
# 5. Replacing an `If` node by its "then" body.
# 6. Replacing an `If` node by its condition.
# 7. Replacing an `If` node by its "else" body.
# 8. Replacing all instances of a variable by a constant.
# 9. Replacing expressions by a constant.
# 10. etc.
#
# Please, refer to the [official Python `ast` reference](http://docs.python.org/3/library/ast) for a full list of node types.
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "slide"}
# ## Implementation
#
# To succeed in this project, you need to complete the `MyReducer` class.
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
class MyReducer(ReducingDebugger):
def minimize(self, code: str) -> str:
# TODO: implement this!
return code
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# To this end, you will need to extend the `NodeReducer` with additional transformations and implement a reducer strategy
# which collects all possible transformations and applies them to the AST until it is exhaustively minimized.
# Feel free to make your own implementation or extend the `DeltaDebugger` from the [Reducing Failure-Inducing Inputs chapter](https://www.debuggingbook.org/beta/html/DeltaDebugger.html) with
# the proper strategy.
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "slide"}
# ## Evaluation
#
# We evaluate your project based on public as well as secret tests. In this section, we present **five** different Python parsers as well as **five** Python input programs, which should be minified.
# These parsers check for a specific property in the code and fail the execution if the property exists. The input programs and parsers in this section make up the public test cases.
# If you pass all of those tests **without hardcoding the modifications** you are guaranteed to score 15 points in this project. You can get more points for passing secret tests.
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### Parsers
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
class Parser(ast.NodeVisitor):
"""The base class for a parser"""
def parse_tree(self, tree: ast.AST) -> None:
self.visit(tree)
def parse(self, source: str) -> str:
tree = ast.parse(source=source)
self.parse_tree(tree)
return "The input was successfully parsed."
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# `Parser` is the base class for a parser from which all other parsers are derived. It works the same way as the `SimpleParser` introduced earlier.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# Let's define a couple of parsers which model a failure during processing of a certain code feature.
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Parser1(Parser):
"""
Contains a boolean operation
"""
def visit_BoolOp(self, node: ast.BoolOp) -> None:
raise ParserException(f"Something went wrong")
# + [markdown] slideshow={"slide_type": "fragment"}
# If we feed this parser with an input program which contains a boolean operation, it fails:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
parser1 = Parser1()
input_program = '''
a = True and False'''
with ExpectError():
parser1.parse(input_program)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# In contrast to the above, an input program _without_ a boolean operation is parsed correctly.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
parser1 = Parser1()
input_program = '''
a = 1
b = True
'''
parser1.parse(input_program)
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# The other parsers are
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Parser2(Parser):
"""
Fails if an input program contains `if` statement
"""
def visit_If(self, node: ast.If) -> None:
raise ParserException(f"Something went wrong")
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
parser2 = Parser2()
input_program = '''
if a:
b = 1
else:
b = 2
'''
with ExpectError():
parser2.parse(input_program)
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Parser3(Parser):
"""
Fails if an input program contains a special unicode character
"""
def __init__(self) -> None:
self.assignment = False
self.steps = 0
def check_unicode(self, string: str) -> bool:
return string == u'\u0426'
def generic_visit(self, node: ast.AST) -> None:
self.steps += 1
super().generic_visit(node)
def visit_Assign(self, node: ast.Assign) -> None:
self.assignment = True
self.steps = 0
self.generic_visit(node)
def visit_Str(self, node: ast.Str) -> None:
if self.assignment and self.steps == 3:
if self.check_unicode(node.s):
raise ParserException(f"Something went wrong")
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
parser3 = Parser3()
input_program = '''
a = u'\u0426'
'''
with ExpectError():
parser3.parse(input_program)
# + hidden=true slideshow={"slide_type": "subslide"}
class Parser4(Parser):
"""
Fails if an input program contains a variable which is not defined
"""
def __init__(self) -> None:
self.assignment = False
self.steps = 0
self.variables: Set[str] = set()
def generic_visit(self, node: ast.AST) -> None:
self.steps += 1
super().generic_visit(node)
def visit_Name(self, node: ast.Name) -> None:
if self.assignment and self.steps == 1:
self.variables.add(node.id)
self.assignment = False
self.generic_visit(node)
elif node.id in self.variables:
self.generic_visit(node)
else:
raise ParserException(f"Something went wrong")
def visit_Assign(self, node: ast.Assign) -> None:
self.assignment = True
self.steps = 0
self.generic_visit(node)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
parser4 = Parser4()
input_program = '''
a = 1
b = c
'''
with ExpectError():
parser4.parse(input_program)
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Parser5(Parser):
"""
Fails if an input program contains a list
"""
def visit_List(self, node: ast.List) -> None:
raise ParserException(f"Something went wrong")
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
parser5 = Parser5()
input_program = '''
a = {x * 2 for x in map(int, ['1', '2', '3'])}
'''
with ExpectError():
parser5.parse(input_program)
# + [markdown] slideshow={"slide_type": "subslide"}
# For secret tests we may use other parsers crafted in the same way.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### Input Programs
# + [markdown] slideshow={"slide_type": "fragment"}
# Now we present five public tests.
# Each test case implements a `get_original()` method that returns an input program which your reducer should minify.
# It also has a `parser` field which stores a parser used for a respective input program.
# Besides, `get_minimized()` method provides a reference solution.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Test1:
parser = Parser1()
def get_original(self) -> str:
return '''
def original():
a = True
b = not False
c = 30
for i in range(c):
if i == 15:
if a and b:
return 1
return 0
'''
def get_minimized(self) -> str:
return '''
True and True'''
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Test2:
parser = Parser2()
def get_original(self) -> str:
return '''
def original():
a = True
b = not False
c = 30
for i in range(c):
if i == 15:
if a and b:
return 1
return 0
'''
def get_minimized(self) -> str:
return '''
if True:
return
'''
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Test3:
parser = Parser3()
def get_original(self) -> str:
return '''
def original():
a = 1
b = a
c = a - b
if c < a:
d = ''
while a == b:
d = u'\u0426'
a += 1
return d
return ''
'''
def get_minimized(self) -> str:
return '''
d = u'\u0426'
'''
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Test4:
parser = Parser4()
def get_original(self) -> str:
return '''
def original():
a = 1
b = a
c = a - b
if c < a:
while a == b:
a += 1
return d
return ''
'''
def get_minimized(self) -> str:
return '''
d
'''
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class Test5:
parser = Parser5()
def get_original(self) -> str:
return '''
def original():
a = 1
b = 0
while True:
if a < b:
return [1, 2, 3]
else:
return []
'''
def get_minimized(self) -> str:
return '''
[]
'''
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# For instance, let's take the input program of `Test1`:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
test = Test1()
source = test.get_original()
print(source)
# + [markdown] slideshow={"slide_type": "subslide"}
# And parse it with the provided parser:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
with ExpectError():
test.parser.parse(source)
# + [markdown] slideshow={"slide_type": "subslide"}
# The parser failed.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Testing Infrastructure
#
# The following section introduces the testing infrastructure, which is used to assess your performance in the project. Passing all tests will be enough to complete the project successfully.
# + [markdown] hidden=true pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# We are not going to directly compare a minimized program with the reference solution (as two strings). Instead, we again transform the code into AST and count the number of nodes in both trees.
# If the difference between them is lower than a `THRESHOLD` (and a parser still produces an error), the test is passed.
# To count the number of nodes in the AST, we need a helper class `NodeCounter`.
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class NodeCounter(ast.NodeVisitor):
"""
This node counter is used to assess
the amount of reductions performed by your reducer.
It counts the number of nodes in the AST.
"""
def __init__(self) -> None:
self.num_nodes = 0
def visit(self, node: ast.AST) -> None:
self.num_nodes += 1
self.generic_visit(node)
def count(self, source: str) -> int:
tree = ast.parse(source=source)
self.visit(tree)
return self.num_nodes
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# The AST of the input program is as follows:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
show_ast(ast.parse(source))
print(f"The number of nodes is {NodeCounter().count(source)}")
# + [markdown] slideshow={"slide_type": "fragment"}
# As you can see, the input program of `Test1` has 42 nodes.
# + [markdown] slideshow={"slide_type": "fragment"}
# The testing framework takes a reducer and allows to execute a single test or all tests at once.
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
class TestingFramework:
THRESHOLD = 3
test_cases = {
'test1': Test1(),
'test2': Test2(),
'test3': Test3(),
'test4': Test4(),
'test5': Test5()
}
def __init__(self, reducer: Any) -> None:
self.reducer = reducer
def count_nodes(self, source: Optional[str]) -> int:
if source is None:
return 100000
return NodeCounter().count(source)
def run_test(self, test: Any) -> bool:
"""
run a single test
"""
print(f'Running test {test.__class__.__name__}')
reducer = self.reducer(test.parser)
reduced_code = reducer.minimize(test.get_original())
return (self.has_property(reduced_code, test.parser) and
self.is_minimized(reduced_code, test.get_minimized()))
def run_tests(self) -> None:
"""
run all public tests
"""
passed_tests = 0
for test in self.test_cases.values():
success = self.run_test(test)
if success:
passed_tests += 1
print(f"In total {passed_tests} tests passed")
def has_property(self, source: str, parser: Any) -> bool:
"""returns True if the parser fails to parse the source"""
try:
parser.parse(source)
print(f'HAS PROPERTY: FAIL')
return False
except ParserException:
print(f'HAS PROPERTY: OK')
return True
except Exception as e:
print(f'HAS PROPERTY: FAIL {e}')
return False
def is_minimized(self, reduced: str, reference: str) -> bool:
"""
Returns True if the AST of the reduced code contains
no more then the number of nodes in the reference + a THRESHOLD
"""
count_minimized = self.count_nodes(reduced)
count_reference = self.count_nodes(reference)
if count_minimized <= count_reference + self.THRESHOLD:
print(f'IS MINIMIZED: OK')
return True
else:
print(f'IS MINIMIZED: FAIL')
return False
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# Let's test our `ReducingDebugger` with the following test:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "fragment"}
class Test0:
parser = Parser1()
def get_original(self) -> str:
return '''
if a and b:
c = 1
else:
c = 2
'''
def get_minimized(self) -> str:
return '''
a and b'''
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
tf = TestingFramework(ReducingDebugger)
if tf.run_test(Test0()):
print("Success!")
# + [markdown] slideshow={"slide_type": "fragment"}
# The input program was successfully minimized.
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "fragment"}
# What if we run all public tests?
# + hidden=true pycharm={"name": "#%%\n"} slideshow={"slide_type": "subslide"}
tf.run_tests()
# + [markdown] slideshow={"slide_type": "subslide"}
# Unfortunately, our parser failed to minimize all input programs from the public test suite.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Grading
# + [markdown] slideshow={"slide_type": "fragment"}
# For this project, you can get a total of 30 Points:
#
# * 15 Points will be awarded for passing the public test (**without hardcoding the minimized solution or the sequence of transformations**) presented in this notebook.
# These 15 points mean that you successfully implemented the must-have implementation.
# * 5 Points will be awarded for passing secret tests.
# * 10 Points will be awarded for efficient implementation (see may-have implementation).
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Must-have Features
#
# Implement a reducer which minifies a given program so that its parser still produces an error.
# To this end, collect all possible transformations over the nodes of an AST tree and then apply one change at a time.
# These modifications should be repeated until no further updates can be made without triggering a parser exception.
# If your reducer passes secret test you are awarded additional 5 points.
#
# *Note: Implementing the given modifications should be sufficient to successfully complete this project.*
# + [markdown] pycharm={"name": "#%% md\n"} slideshow={"slide_type": "subslide"}
# ### May-have Features
#
# An implementation only of the must-have features aims for correctness but can be inefficient (if transformations are applied one at a time). So, it can be optimized, for instance, with help of the Delta Debugging approach.
#
# Implement an AST Delta Debugger which efficiently prunes the nodes.
# If your reducer can beat the runtime of our reference implementation (which simply collects all possible transformations and applies them randomly one at a time) you can get up to 10 points (depending on the margin).
#
# Hint: The [Reducing Failure-Inducing Inputs chapter](https://www.debuggingbook.org/beta/html/DeltaDebugger.html) already tries to optimize the reduction of the AST with help of Delta Debugging.
# Hint: For instance, you can find useful _Hierarchical Delta Debugging_ paper from the Background section of **Reducing Failure-Inducing Inputs** Chapter.
|
docs/notebooks/Reducing_Code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computational Assignment 1
# **Assigned Tuesday, 3-17-20.**, **Due Tuesday, 3-22-20.**
# Congratulations on installing the Jupyter Notebook! Welcomne to your first computational assignment!
# Beyond using this as a tool to understand physical chemistry, python and notebooks are actually used widely in scientific analysis. Big data analysis especially uses python notebooks.
# ## Introduction to the notebook
# If you double click on the text above, you will notice the look suddenly changes. Every section of the notebook, including the introductory text, is a technically a code entry. The text is written in a typesetting language called **Markdown**. To learn more see https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet
# To run a code entry in the notebook, select the section you want to run and type
# `shift+enter`
# If you want to make notes on a notebook, you can press the plus sign in the toolbar above to creat a new entry. Then make sure to switch the menu that in the toolbar from **code** to **Markdown**.
# We can also run calculations this way.
# In the entry below, I haved typed
#
# `123+3483`
# Select the entry and type `shift+enter`
123+3483
# Once you run an entry, the output is displayed on the screen, when applicable
# Now try some arithmatic yourself in the blank entry below.
# (Don't forget to hit `shift+enter` to run your calculation!)
12+12
# ## Introduction to programming and python
# Python is a very powerful and intuitive modern programming language. It is easier to learn than many other languages. Because of the wide availability of libraries such **numpy** and **scipy** (among many others), it is very useful for scientific calculations.
# In this section, we will cover some very basic concepts. I assuming that nearly everyone has little or no previous programming experience, which is common for most chemistry and biology students.
# We will slowly build up to the skills we need to run complex calculations!
# ### Our first python code: "Hello World!"
# The first thing we usually learn how to do is print a simple message to the output. Run the following entry.
print("Hello World!")
# Print is function that takes in text as an argument and outputs that text.
# A slightly more complicated example. Run the following entry
# +
# This is a comment in python
# Set a variable
x = 1 + 7
# print the result of the variable
print(x)
# -
# The lines that begin with "#" are comments. They are not read by the notebook and do not affect the code. They are very useful to make your code human readable
#
# This snippet of code set assigned the result of `1+7` to the variable `x` and then used `print` to output that value.
# ## Loops
# One of the benifits of the computer is that it can run a calcualtion many times without having to manually type each line. The way that we do this is to use a **loop**.
# +
# This is an example of a loop
# The colon is required on the first line
for i in (1,2,3):
# This indentation is required for loops
print ("Hello World, iteration",i)
# -
# ### Explanation
#
# 1. The command `for` tells the code that this is a loop
# 2. The variable `i` is the counting variable. Everytime the loop runs, it will sequentially take on a different value from the list
# 3. The `(1,2,3)` is list of values.
# Sometimes we need to run a loop many times or iterate over a large list of numbers. For this, the `range` command is useful
# The command range(a,b) creates a list of numbers from a to b
for i in range(-3,3):
print ("Hello World, iteration",i)
# Note that the `range(a,b)` command makes a list that spans from `a` to `b-1`.
# In the example above `range(-3,3`) makes a list that goes from -3 to 2
# ## Conditional Statements: IF
# Many times we want the computer to do something after analyzing a logical statement. **If this is true, then do that**. These are called conditional statements
# +
# Conditional example
a = 100
if (a>0):
#Like in the loop example, the indentation defines what happens in this
# block of the if statement
print("the number is positive")
elif (a<0):
print("the number is negative")
elif (a==0):
print("the number is zero")
# -
# Now we can try it again with a different value for `a`
# +
# Conditional example again
a = -1234
if (a>0):
print("the number is positive")
elif (a<0):
print("the number is negative")
elif (a==0):
print("the number is zero")
# -
# Once more time
# +
# Conditional example again
a = 0
if (a>0):
print("the number is positive")
elif (a<0):
print("the number is negative")
elif (a==0):
print("the number is zero")
# -
# ## Bringing it all together
# These can all be combined together to perform complicated actions. Note the indentation and colons. They matter.
# ### Combined Example
# +
# A loop with an if statement
for i in range(-1,2):
print("Iteration",i)
if (i==0):
print("zero!")
# -
# # Exercise
# Following the examples above, write a code snippet that uses the `range` command to scan from -10 to 10 and print whether the number is positive, negative, or zero.
#
# **To turn this in, upload the notebook in Github, this will make a committ that I can see.**
# +
# A loop with an if statement
for i in range (-10,10):
print("Iteration",i)
if (i==0):
print("zero!")
# -
|
comp_hw-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Representations:
# * Individual tree
# * Trajectory
# * Radii vs. iteration, with summaries such as log-probability, and the marginal likelihoods of various models
# * Include SMIRKS tags for the individual types
from pickle import load
from glob import glob
fnames = glob('../bayes_implicit_solvent/rjmc_experiments/bugfixed_*gaussian_ll.pkl')
print(fnames)
with open(fnames[0], 'rb') as f:
result = load(f)
result.keys()
traj = result['traj']
log_probs = result['log_probs']
log_acceptance_probabilities = result['log_acceptance_probabilities']
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(log_probs)
plt.xlabel('iteration')
plt.ylabel('log probability')
plt.plot(log_probs[1000:])
plt.xlabel('iteration')
plt.ylabel('log probability')
import numpy as np
def plot_log_prob(t=1000, include_number=True):
plt.xlim(0, len(log_probs))
#min_t = max(0,t - 500)
#plt.ylim(log_probs[0], max(log_probs) + np.abs(max(log_probs) * 0.01))
plt.plot(log_probs[:t])
plt.xlabel('iteration')
plt.ylabel('log probability')
title = 'log posterior probability'
if include_number:
title += ': {}'.format(int(log_probs[t]))
plt.title(title)
plot_log_prob(t=len(log_probs) - 1)
plt.plot(np.diff(log_probs))
np.sum(np.diff(log_probs)<0)
tree = traj[0]
n_types = [tree.number_of_nodes for tree in traj]
plt.plot(n_types)
plt.yticks(list(range(min(n_types), max(n_types) + 1)))
plt.ylabel('# GB types')
plt.xlabel('iteration')
# +
import numpy as np
def plot_running_average_model_evidence(t=1000):
x = list(range(0, max(n_types) + 1))
y = np.bincount(n_types[:t], minlength=len(x))
plt.bar(x, y)
plt.xlabel('# GB-types')
plt.ylabel('time spent')
plt.title('marginal distribution for # GB-types')
plt.xticks(list(range(max(n_types) + 1))[::5])
plt.ylim(0,max(np.bincount(n_types)))
plot_running_average_model_evidence()
# -
plot_running_average_model_evidence(100)
tree = traj[-1]
# +
from simtk import unit
radii = np.array([tree.get_radius(n) / unit.nanometer for n in tree.nodes])
# -
list(zip(tree.nodes, radii))
# +
# to assign colors, let's get all of the types every encountered during the simulation
# then put them in lexicographic order, then make a dictionary that indexes into a nice seaborn color palette
# -
all_types = set()
for tree in traj:
all_types.update(tree.nodes)
all_types = sorted(list(all_types))
all_types
len(all_types)
import seaborn.apionly as sns
color_list = sns.color_palette(palette="husl", n_colors=len(all_types))
color_list = color_list[5:] + color_list[:5]
color_dict = dict(zip(all_types, color_list))
# +
# for scatter plot representation
x = []
y = []
c = []
for i in range(len(traj)):
tree = traj[i]
radii = [tree.get_radius(n) / unit.nanometer for n in tree.nodes]
x += [i] * len(radii)
y += radii
c += [color_dict[t] for t in tree.nodes]
# -
# +
# for line-plot representation
x = np.arange(len(traj))
ys = [np.zeros(len(traj)) * np.nan for _ in all_types]
c = [color_dict[t] for t in all_types]
type_dict = dict(zip(all_types, range(len(all_types))))
for time, tree in enumerate(traj):
for n in tree.nodes:
i = type_dict[n]
ys[i][time] = tree.get_radius(n) / unit.nanometer
# -
print(traj[0])
print(traj[-1])
ys[0] - ys[1]
tree.nodes
tree.get_radius('*'), tree.get_radius('[#1]')
for i in range(len(all_types)):
plt.plot(x, ys[i], c=c[i])
for y, n in zip(ys, all_types):
if np.min(y) < 0:
print(n)
[(n, np.min(np.nan_to_num(y))) for (n, y) in zip(all_types, ys)]
# +
def plot_radii_as_scatter(ax, up_to_iteration=1000):
t = np.argmax(np.array(x) >= up_to_iteration)
ax.scatter(x[:t], y[:t], s=1, color=c[:t])
ax.set_xlim(0, len(traj))
ax.set_ylim(0, max(y) * 1.1)
ax.set_xlabel('iteration')
ax.set_ylabel('radius (Å)')
ax.set_title('Born radii')
def plot_radii(ax, up_to_iteration=1000):
max_y = -np.inf
for i in range(len(all_types)):
y = ys[i][:up_to_iteration] * 10
ax.plot(x[:up_to_iteration], y, c=c[i])
max_y = max(max(y), max_y)
ax.set_xlim(0, len(traj))
ax.set_ylim(0, max_y * 2)
ax.set_xlabel('iteration')
ax.set_ylabel('radius (Å)')
ax.set_title('Born radii')
def prettify_axes(ax):
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
plt.figure(figsize=(8,4))
ax = plt.subplot(1,2,1)
plot_radii(ax, 500)
prettify_axes(ax)
ax = plt.subplot(1,2,2)
plot_radii(ax)
prettify_axes(ax)
# -
node_color = [color_dict[n] for n in tree.G.nodes()]
import networkx as nx
layouts = [
('shell', nx.layout.shell_layout(tree.G)),
('fruchterman_reingold', nx.layout.fruchterman_reingold_layout(tree.G)),
('kamada_kawai', nx.layout.kamada_kawai_layout(tree.G)),
('spectral', nx.layout.spectral_layout(tree.G)),
('circular', nx.layout.circular_layout(tree.G)),
]
for name, pos in layouts:
plt.figure()
plt.title(name)
nx.draw_networkx(tree.G, pos=pos, node_color=node_color, )
# +
# none of these are what I want!
# +
# let's just precompute sensible positions for all of the types
# -
def depth(node):
if node == '*':
return 0
return len(node.split('~'))
depth_dict = dict(zip(all_types, map(depth, all_types)))
depth_dict
height_dict = dict(zip(all_types, 0.3 * np.arange(len(all_types))))
height_dict
max(height_dict.values())
pos_dict = dict(zip(all_types, [(depth_dict[t], height_dict[t]) for t in all_types]))
pos_dict
def draw_typing_tree(tree):
node_color = [color_dict[n] for n in tree.G.nodes()]
nx.draw_networkx(tree.G, pos=pos_dict, node_color=node_color)
plt.xlim(-1,max(height_dict.values()) + 2)
plt.ylim(-1,max(height_dict.values()) + 1)
#plt.xticks([])
#plt.yticks([])
plt.axis('off')
plt.title('GB-typer')
draw_typing_tree(traj[-1])
def plot_summary(t=1000):
plt.figure(figsize=(8,8))
# 1. current tree
ax = plt.subplot(2,2,1)
tree = traj[t]
draw_typing_tree(tree)
# 2. radii
ax = plt.subplot(2,2,2)
plot_radii(ax, t)
prettify_axes(ax)
# 3. log probability
ax = plt.subplot(2,2,3)
plot_log_prob(t)
prettify_axes(ax)
# 4. marginal # GB types
ax = plt.subplot(2,2,4)
plot_running_average_model_evidence(t)
prettify_axes(ax)
# (# types trajectory?)
plt.tight_layout()
def plot_number_of_atom_types(ax, t=1000):
ax.plot(n_types)
ax.set_xlabel('iteration')
ax.set_ylabel('# of types')
ax.set_title('number of GB types in current model')
ax = plt.subplot(1,1,1)
plot_number_of_atom_types(ax)
dtrees = [tuple(t.nodes) for t in traj]
len(set(dtrees))
def plot_three_panels(t=1000):
plt.figure(figsize=(8,8))
## 1. current tree
#ax = plt.subplot(2,2,1)
#tree = traj[t]
#draw_typing_tree(tree)
# 1. radii
ax = plt.subplot(2,1,1)
plot_radii(ax, t)
prettify_axes(ax)
## 2. log probability
#ax = plt.subplot(3,1,2)
#plot_log_prob(t, include_number=False)
#prettify_axes(ax)
# 3. number of atom types
ax = plt.subplot(2,1,2)
plot_number_of_atom_types(ax, t)
plt.yticks(list(range(11, 15)))
prettify_axes(ax)
## 4. marginal # GB types
#ax = plt.subplot(2,2,4)
#plot_running_average_model_evidence(t)
#prettify_axes(ax)
# (# types trajectory?)
plt.tight_layout()
plot_three_panels()
plt.savefig('three_panels_wo_callouts.png', dpi=300)
np.argmax(n_types)
n_types[990:1000]
n_types[992]
# +
inds_to_plot = [0, 500, 800, 992]
plt.figure(figsize=(3*4,3))
for i, t in enumerate(inds_to_plot):
plt.subplot(1,4,i + 1)
draw_typing_tree(traj[t])
plt.title(i + 1)
plt.savefig('callouts.png', dpi=300)
# -
for t in inds_to_plot:
print(traj[t])
print('\n\n')
plot_summary(len(traj) - 1)
len(traj)
plot_summary(2)
from tqdm import tqdm
for t in tqdm(range(2, len(traj))[::50]):
plot_summary(t)
plt.savefig('longer_bugfixed_typing_animation/{:03}.png'.format(t))
plt.close()
# Add subplots for "train" / "test" solvation free energy predictions?
# +
import mdtraj as md
import numpy as np
from pkg_resources import resource_filename
from tqdm import tqdm
from bayes_implicit_solvent.molecule import Molecule
from bayes_implicit_solvent.smarts import atomic_number_dict
from bayes_implicit_solvent.solvation_free_energy import smiles_list
from bayes_implicit_solvent.typers import GBTypingTree
from bayes_implicit_solvent.utils import mdtraj_to_list_of_unitted_snapshots
data_path = '../data/'
np.random.seed(0)
train_mols = []
N_train = 64
smiles_subset = list(smiles_list)
np.random.shuffle(smiles_subset)
train_smiles_subset = smiles_subset[:N_train]
print('looking at only {} entries from FreeSolv'.format(len(train_smiles_subset)))
n_configuration_samples = 10
for smiles in train_smiles_subset:
mol = Molecule(smiles, vacuum_samples=[])
path_to_vacuum_samples = resource_filename('bayes_implicit_solvent',
'vacuum_samples/vacuum_samples_{}.h5'.format(
mol.mol_index_in_smiles_list))
vacuum_traj = md.load(path_to_vacuum_samples)
thinning = int(len(vacuum_traj) / n_configuration_samples)
mol.vacuum_traj = mdtraj_to_list_of_unitted_snapshots(vacuum_traj[::thinning])
#print('thinned vacuum_traj from {} to {}'.format(len(vacuum_traj), len(mol.vacuum_traj)))
train_mols.append(mol)
# -
test_smiles_subset = smiles_subset[N_train:]
print(len(test_smiles_subset))
test_smiles_subset = smiles_subset[N_train:]
test_mols = []
for smiles in test_smiles_subset:
mol = Molecule(smiles, vacuum_samples=[])
path_to_vacuum_samples = resource_filename('bayes_implicit_solvent',
'vacuum_samples/vacuum_samples_{}.h5'.format(
mol.mol_index_in_smiles_list))
vacuum_traj = md.load(path_to_vacuum_samples)
thinning = int(len(vacuum_traj) / n_configuration_samples)
mol.vacuum_traj = mdtraj_to_list_of_unitted_snapshots(vacuum_traj[::thinning])
#print('thinned vacuum_traj from {} to {}'.format(len(vacuum_traj), len(mol.vacuum_traj)))
test_mols.append(mol)
print(len(test_mols))
# +
experiment_train = [mol.experimental_value for mol in train_mols]
experiment_unc_train = [mol.experimental_uncertainty for mol in train_mols]
experiment_test = [mol.experimental_value for mol in test_mols]
experiment_unc_test = [mol.experimental_uncertainty for mol in test_mols]
# -
mol.experimental_value
def get_train_predictions(tree):
preds = []
for mol in train_mols:
r = tree.assign_radii(mol.mol)
s = np.ones(len(r))
preds.append(mol.predict_solvation_free_energy(r, s))
return preds
def get_test_predictions(tree):
preds = []
for mol in test_mols:
r = tree.assign_radii(mol.mol)
s = np.ones(len(r))
preds.append(mol.predict_solvation_free_energy(r, s))
return preds
train_predictions = []
for t in tqdm(range(2, len(traj))[::10]):
tree = traj[t]
tree.ordered_nodes = tree.nodes
train_predictions.append(get_train_predictions(tree))
x_ = np.arange(2, len(traj))[::10]
test_predictions = []
for t in tqdm(range(2, len(traj))[::10]):
tree = traj[t]
test_predictions.append(get_test_predictions(tree))
# +
y_ = [[p[0] for p in pred] for pred in train_predictions]
y_unc = [[p[1] for p in pred] for pred in train_predictions]
y_test = [[p[0] for p in pred] for pred in test_predictions]
y_unc_test = [[p[1] for p in pred] for pred in test_predictions]
# -
np.array(y_test).shape
np.array(y_).shape
np.array(experiment_train).shape
from bayes_implicit_solvent.utils import remove_top_right_spines
# +
all_elements = [1, 6, 7, 8, 9, 15, 16, 17, 35, 53]
element_dict = dict(zip(all_elements, list(range(len(all_elements)))))
# default radii are 0.15
initial_radius_dict = {}
for atomic_number in all_elements:
initial_radius_dict[atomic_number] = 0.15
# a few elements have different radii in the default model
# https://github.com/pandegroup/openmm/blob/master/wrappers/python/simtk/openmm/app/internal/customgbforces.py#L233-L257
initial_radius_dict[1] = 0.12
initial_radius_dict[6] = 0.17
initial_radius_dict[7] = 0.155
initial_radius_dict[14] = 0.21
initial_radius_dict[15] = 0.185
initial_radius_dict[16] = 0.18
initial_radii = np.array([initial_radius_dict[a] for a in all_elements])
# scales
initial_scale_dict = {}
for atomic_number in all_elements:
initial_scale_dict[atomic_number] = 0.8
# a few elements have different radii in the default model
# https://github.com/pandegroup/openmm/blob/master/wrappers/python/simtk/openmm/app/internal/customgbforces.py#L233-L257
initial_scale_dict[1] = 0.85
initial_scale_dict[6] = 0.72
initial_scale_dict[7] = 0.79
initial_scale_dict[8] = 0.85
initial_scale_dict[9] = 0.88
initial_scale_dict[15] = 0.86
initial_scale_dict[16] = 0.96
initial_scales = np.array([initial_scale_dict[a] for a in all_elements])
def pack(radii, scales):
n = len(radii)
theta = np.zeros(2 * n)
theta[:n] = radii
theta[n:2 * n] = scales
return theta
def unpack(theta):
n = int((len(theta)) / 2)
radii, scales = theta[:n], theta[n:2 * n]
return radii, scales
initial_theta = pack(initial_radii, initial_scales)
def construct_array(mol, theta):
radii, scales = unpack(theta)
elements = np.array([a.element.atomic_number for a in mol.top.atoms()])
mol_radii = np.array([radii[element_dict[element]] for element in elements])
mol_scales = np.array([scales[element_dict[element]] for element in elements])
return mol_radii, mol_scales
mol = train_mols[0]
r, s = construct_array(mol, initial_theta)
# -
mol.predict_solvation_free_energy(r, s)
all_mols = train_mols + test_mols
len(all_mols)
# +
from bayes_implicit_solvent.constants import beta
def reduce(value):
"""Input value is in units of kcal/mol, turn it into units of kB T"""
return beta * (value * unit.kilocalorie_per_mole)
def unreduce(value):
"""Input value is in units of kB T, turn it into units of kilocalorie_per_mole"""
return value / (beta * unit.kilocalorie_per_mole)
# +
obc2_all_preds = []
all_expt_values = []
all_expt_uncs = []
problematic_smiles = []
for mol in all_mols:
r, s = construct_array(mol, initial_theta)
pred = unreduce(mol.predict_solvation_free_energy(r, s)[0])
obc2_all_preds.append(pred)
all_expt_values.append(unreduce(mol.experimental_value))
all_expt_uncs.append(unreduce(mol.experimental_uncertainty))
if pred < -30:
problematic_smiles.append(mol.smiles)
# -
c = np.zeros(len(all_expt_uncs))
for i in range(len(all_expt_uncs)):
if np.isclose(all_expt_uncs[i], 1.9):
c[i] = 1
sum(c)
c = np.zeros(len(all_expt_uncs))
for i in range(len(all_expt_uncs)):
if all_expt_uncs[i] >= 1.9:
c[i] = 1
sum(c)
mol = all_mols[0]
subsearch_string = '[#16]~[#17]'
from bayes_implicit_solvent.utils import cached_substructure_matches
for mol in all_mols:
if sum(cached_substructure_matches(mol.mol, subsearch_string)) > 0:
print(mol.smiles)
print(unreduce(mol.experimental_value))
print(unreduce(mol.experimental_uncertainty))
print(problematic_smiles)
diag = np.array(sorted(np.hstack((obc2_all_preds, all_expt_values))))
diag = diag[diag > -30]
min(diag)
# +
def obc2_scatter_no_error_bars(ax, restrict_range=True, color_by_uncertainty=False):
if color_by_uncertainty:
ax.scatter(obc2_all_preds, all_expt_values, s=2, c=all_expt_uncs)
else:
ax.scatter(obc2_all_preds, all_expt_values, s=2)
ax.plot(diag, diag, color='grey')
ax.set_xlabel(r'predicted $\Delta G$ (kcal/mol)')
ax.set_ylabel(r'experimental $\Delta G$ (kcal/mol)')
if restrict_range:
ax.set_xlim(-15,5)
ax.set_ylim(-15,5)
ax.set_xticks(list(range(-15,6,5)))
ax.set_yticks(list(range(-15,6,5)))
ax.set_title('OBC2\n(RMSE: 2.4 kcal/mol)')
remove_top_right_spines(ax)
ax = plt.subplot(1,1,1)
obc2_scatter_no_error_bars(ax)
# -
plt.figure()
ax = plt.subplot(1,1,1)
obc2_scatter_no_error_bars(ax, restrict_range=False, color_by_uncertainty=True)
plt.colorbar()
inds = np.array(obc2_all_preds) >-40
obc2_rmse_excluding_outliers = np.sqrt(np.mean((np.array(obc2_all_preds)[inds] - np.array(all_expt_values)[inds])**2))
obc2_rmse_excluding_outliers
obc2_rmse = np.sqrt(np.mean((np.array(obc2_all_preds) - np.array(all_expt_values))**2))
obc2_rmse
# +
ax = plt.subplot(111)
from bayes_implicit_solvent.utils import remove_top_right_spines
def plot_rmse_train_test(ax):
ax.hlines(obc2_rmse, xmin=x[0], xmax=x[-1], linestyles='--', label='OBC2')
ax.hlines(obc2_rmse_excluding_outliers, xmin=x[0], xmax=x[-1], linestyles='--', color='grey', label='OBC2 -- excluding outliers')
ax.plot(x_, np.sqrt(np.mean((unreduce(np.array(y_) - np.array(experiment_train))**2), axis=1)), label='train')
ax.plot(x_, np.sqrt(np.mean((unreduce(np.array(y_test) - np.array(experiment_test))**2), axis=1)), label='test')
ax.set_ylim(0,)
ax.legend()
ax.set_ylabel(r'RMSE (kcal/mol)')
ax.set_xlabel('RJMC iteration')
ax.set_title('FreeSolv: train and test RMSE')
remove_top_right_spines(ax)
ax = plt.subplot(1,1,1)
plot_rmse_train_test(ax)
plt.savefig('train-and-test-rmse-with-obc2-baseline.png', dpi=300)
# -
from simtk import unit
from bayes_implicit_solvent.constants import kB, temperature
kcalmol_over_kT = (1.0 * unit.kilocalorie_per_mole) / (kB * temperature)
kcalmol_over_kT
np.sqrt(np.mean((unreduce(np.array(y_) - np.array(experiment_train)))**2, axis=1))[-1]
np.sqrt(np.mean((unreduce(np.array(y_test) - np.array(experiment_test)))**2, axis=1))[-1]
obc2_rmse
obc2_rmse_excluding_outliers
# +
def train_scatter(i=0):
plt.errorbar(x=experiment_train, xerr=experiment_unc_train, y=y_[i], yerr=y_unc[i], fmt='none')
plt.plot(sorted(experiment_train), sorted(experiment_train), color='grey')
plt.xlabel('experiment')
plt.ylabel('prediction')
train_scatter(0)
plt.figure()
train_scatter(-1)
# +
def test_scatter(i=0):
plt.errorbar(x=experiment_test, xerr=experiment_unc_test, y=y_test[i], yerr=y_unc_test[i], fmt='none')
plt.plot(sorted(experiment_test), sorted(experiment_test), color='grey')
plt.xlabel('experiment')
plt.ylabel('prediction')
test_scatter(0)
plt.figure()
test_scatter(-1)
# -
def scatter_no_error_bars(ax, i=0):
ax.scatter(y_[i], experiment_train, s=2, label='train')
ax.scatter(y_test[i], experiment_test, s=2, label='test')
plt.legend()
ax.plot(diag, diag, color='grey')
plt.xlim(-15,5)
plt.ylim(-15,5)
ax.set_xticks(list(range(-15,6,5)))
ax.set_yticks(list(range(-15,6,5)))
ax.set_xlabel('predicted $\Delta G$ (kcal/mol)')
ax.set_ylabel(r'experimental $\Delta G$ (kcal/mol)')
ax.set_title('Bayesian\n(train RMSE: 3.3 kcal/mol,\ntest RMSE: 5.5 kcal/mol)')
remove_top_right_spines(ax)
ax = plt.subplot(111)
scatter_no_error_bars(ax, i=-1)
# +
plt.figure(figsize=(4,6))
ax = plt.subplot(2,1,1)
scatter_no_error_bars(ax, i=0)
ax = plt.subplot(2,1,2)
scatter_no_error_bars(ax, i=-1)
plt.tight_layout()
# +
plt.figure(figsize=(5,4))
ax = plt.subplot(2,1,1)
plot_radii(ax, t)
plt.xlabel('')
prettify_axes(ax)
ax = plt.subplot(2,1,2)
plot_rmse_train_test(ax)
plt.tight_layout()
plt.savefig('two-panels-without-callouts.png', dpi=300)
# +
plt.figure(figsize=(3.5,6))
ax = plt.subplot(2,1,1)
obc2_scatter_no_error_bars(ax)
ax = plt.subplot(2,1,2)
scatter_no_error_bars(ax, i=-1)
plt.tight_layout()
plt.savefig('scatter-plots.png', dpi=300)
# -
test_scatter(-1)
# +
def train_scatter(i=0):
plt.errorbar(x=experiment_train, xerr=experiment_unc_train, y=y_[i], yerr=y_unc[i], fmt='none')
plt.plot(sorted(experiment_train), sorted(experiment_train), color='grey')
plt.xlabel('experiment')
plt.ylabel('prediction')
train_scatter(0)
plt.figure()
train_scatter(-1)
# +
plt.errorbar(x=experiment, xerr=experiment_unc, y=y_[-1], yerr=y_unc[-1], fmt='none')
plt.plot(sorted(experiment), sorted(experiment), color='grey')
plt.ylim(min(experiment), max(experiment))
plt.xlabel('experiment')
plt.ylabel('prediction')
# +
plt.errorbar(x=experiment, xerr=experiment_unc, y=y_[0], yerr=y_unc[0], fmt='none')
plt.plot(sorted(experiment), sorted(experiment), color='grey')
#plt.ylim(min(experiment), max(experiment))
plt.xlabel('experiment')
plt.ylabel('prediction')
# -
for pred in predictions[::10]:
plt.figure()
plt.scatter(experiment, pred)
|
notebooks/plot rjmc typing trees.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CHEME 512 Presentation: Case 1.2.3
#
# ### <NAME>
# ### Infinite plate with temperature dependent conductivity
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
capture = mpimg.imread('capture.png')
plt.subplots(figsize=(7, 7))
plt.imshow(capture)
plt.axis('off')
plt.show()
# -
# Starting with the energy conservation equation in the Deen Eq2.4-1
#
# $$\rho C_p\frac{DT}{Dt}=-\nabla q +q'''$$
#
# Using the Fourier's law
#
# $$q=-k\nabla T$$
#
# So
#
# $$\rho C_p\frac{DT}{Dt}=\nabla (k\nabla T)+q'''$$
#
# where $k$ is a function of T.
#
# For this problem,the energy flux is only in x-direction and we assume steady state,then the equation can be simplified
#
# $$\frac{d}{dx}(k\frac{dT}{dx})+q'''=0$$
#
# $$\frac{d}{dx}(k\frac{dT}{dx})=-q'''$$
#
# integrate both sides
#
# $$k\frac{dT}{dx}=-q'''x+C_1$$
#
# where $C_1$= constant
#
# Then bring the $k=k_f+\beta (T-T_f)$
#
# $$(k_f+\beta (T-T_f))\frac{dT}{dx}=-q'''x+C_1$$
#
# Separate variables and integrate,we can get
#
# $$\frac{\beta}{2}T^2+k_fT-\beta T_fT=-\frac{q'''}{2}x^2+C_1x+C_2$$
#
# $$\beta T^2+2k_fT-2\beta T_fT=-q'''x^2+2C_1x+2C_2,(1)$$
#
# where $C_1,C_2$=constant
#
# Then using the Boundary Conditions to solve the constant $C_1,C_2$
#
# for this problem,the Boundary Conditions is easy.
#
# B.C.1 is $$T(x=-L)=T_f$$
#
# B.C.2 is $$T(x=L)=T_f$$
#
# Then we can get two equation:
# $$\beta T_f^2-2\beta T_f^2+2k_fT_f=-q'''L^2-2C_1L+2C_2$$
#
# $$\beta T_f^2-2\beta T_f^2+2k_fT_f=-q'''L^2+2C_1L+2C_2$$
# Solve it and we get
# $$C_1=0$$
# $$C_2=\frac{1}{2}(q'''L^2-\beta T_f^2+2k_fT_f)$$
#
# Bring back the $C_1,C_2$ to the equation (1)
#
# $$\beta T^2+2k_fT-2\beta T_fT=-q'''x^2+q'''L^2-\beta T_f^2+2k_fT_f$$
#
# Then we can rewrite the equation to solve $T(x)$,below is the equation after rewriting
#
# $$\beta T^2-2\beta T_fT+\beta T_f^2+2k_fT-2k_fT_f=q'''(L^2-x^2)$$
#
# $$\beta (T^2-2T_fT+T_f^2)+2k_f(T-T_f)=q'''(L^2-x^2)$$
#
# $$\beta (T-T_f)^2+2k_f(T-T_f)-q'''(L^2-x^2)=0$$
#
# Set $S=T-T_f$ the eqution will be
#
# $$\beta S^2+2k_fS-q'''(L^2-x^2)=0$$
#
# Solve it.We can get
#
# $$S=\frac{-2k_f±\sqrt{4k_f^2+4\beta q'''(L^2-x^2)}}{2\beta}$$
#
# So
#
# $$T-T_f=\frac{-k_f±\sqrt{k_f^2+\beta q'''(L^2-x^2)}}{\beta}$$
#
# Because the solution must satisfy the boundary conditions.So the final solution is
#
# $$T-T_f=\frac{-k_f+\sqrt{k_f^2+\beta q'''(L^2-x^2)}}{\beta}$$
#
# $$T-T_f=\frac{k_f}{\beta}\Bigg[-1+\sqrt{1+\frac{\beta q'''}{k_f^2}(L^2-x^2)}\Bigg]$$
#
# $$T=T_f+\frac{k_f}{\beta}\Bigg[-1+\sqrt{1+\frac{\beta q'''}{k_f^2}(L^2-x^2)}\Bigg]$$
# But from the textbook,the given solution is
#
# $$\frac{(T-T_f)k_f}{q'''L^2}=\frac{2}{B}\Bigg[-1+\sqrt{1+2B(1-x^2)}\Bigg]$$
#
# where $B=\frac{\beta q'''L^2}{2k_{\infty}^2}$,bring it to the solution
#
# $$\frac{(T-T_f)k_f}{q'''L^2}=\frac{4k_{\infty}^2}{\beta q'''L^2}\Bigg[-1+\sqrt{1+2\frac{\beta q'''L^2}{2k_{\infty}^2}(1-x^2)}\Bigg]$$
#
# $$T-T_f=\frac{4k_{\infty}^2}{\beta k_f}\Bigg[-1+\sqrt{1+\frac{\beta q'''L^2}{k_{\infty}^2}(1-x^2)}\Bigg]$$
#
# compared with my solution
#
# $$T-T_f=\frac{k_f}{\beta}\Bigg[-1+\sqrt{1+\frac{\beta q'''}{k_f^2}(L^2-x^2)}\Bigg]$$
import matplotlib.pyplot as plt
import numpy as np
# +
Tf = 50
kf = 10
q = 1
L = 10
x = np.linspace(-L, L, 101)
fig, ax = plt.subplots(figsize=(9,8))
for i,b in enumerate([0.001,1,2,5,10,100,10000]):
T= Tf+kf*(-1+(1+b*q*(L**2-x**2)/(kf**2))**0.5)/b
ax.plot(x, T, label='Beta = {}'.format(b))
plt.xlabel('x')
plt.ylabel('T')
ax.legend()
b = 1
q = 1
fig, ax = plt.subplots(figsize=(9,8))
for i, kf in enumerate([0.001,1,2, 5, 10,100,10000]):
T= Tf+kf*(-1+(1+b*q*(L**2-x**2)/(kf**2))**0.5)/b
ax.plot(x, T, label='kf = {}'.format(kf))
plt.xlabel('x')
plt.ylabel('T')
ax.legend()
# -
|
presentations/10_30_19_Yifeng.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setting Rendering Styles
# This tutorial demonstrates how to instantiate a py3Dmol viewer and set atom styles and colors.
import py3Dmol
# ## Instantiating the py3mol viewer with a PDB ID
# py3Dmol downloads PDB structures using the MMTF file format from https://mmtf.rcsb.org. Prepend the 'pdb:" prefix to the 4-letter PDB ID.
viewer = py3Dmol.view(query='pdb:1STP')
viewer.show()
# ## Setting an Atom Style
# Styles are specified as nested dictionaries. In this example, 'stick' is the AtomStyleSpec [(list of atom styles)](http://3dmol.csb.pitt.edu/doc/types.html#AtomStyleSpec).
viewer.setStyle({'stick': {}})
viewer.show()
# ## Setting a Color Representation
# Colors are specified as a dictionary, e.g., {'color': 'spectrum'} or monochrome colors {'color':'lightgreen'} [(list of colors)](https://github.com/3dmol/3Dmol.js/blob/master/3Dmol/colors.js#L45-L192).
viewer.setStyle({'cartoon': {'color': 'spectrum'}})
viewer.show()
# ## Setting a Color Scheme
# Color schemes can be used to color atoms and residues by properties. A particulary useful color scheme is the <color>Carbon color scheme, for example: 'greenCarbon' [(list of color schemes)](https://github.com/3dmol/3Dmol.js/blob/master/3Dmol/colors.js#L26-L36).
viewer.setStyle({'stick': {'colorscheme': 'greenCarbon'}})
viewer.show()
# Example of coloring by amino acid type
viewer.setStyle({'cartoon': {'colorscheme':'amino'}})
viewer.show()
|
1-3D-visualization/2-Styles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reproject Sentinel-2 data to WGS84
import rasterio
from rasterio.merge import merge
from rasterio.plot import show
from rasterio.warp import calculate_default_transform, reproject, Resampling
import glob
import os
def reproject_raster(in_path, out_path, dst_crs = 'EPSG:4326'):
# reproject raster to project crs
with rasterio.open(in_path) as src:
src_crs = src.crs
transform, width, height = calculate_default_transform(src_crs, dst_crs, src.width, src.height, *src.bounds)
kwargs = src.meta.copy()
kwargs.update({
'crs': dst_crs,
'transform': transform,
'width': width,
'height': height})
with rasterio.open(out_path, 'w', **kwargs) as dst:
for i in range(1, src.count + 1):
reproject(
source=rasterio.band(src, i),
destination=rasterio.band(dst, i),
src_transform=src.transform,
src_crs=src.crs,
dst_transform=transform,
dst_crs=dst_crs,
resampling=Resampling.nearest)
return(out_path)
# File and folder paths
dirpath = r"data\sentinel2"
search_criteria = "*\R10m\TCI.jp2"
q = os.path.join(dirpath, search_criteria)
print(q)
paths = glob.glob(q)
paths
files_to_mosaic = []
for path_in in paths:
folders_split = path_in.split('\\')
filename_split = folders_split[-1].split('.')
filename_split[0] = 'transformed_tci'
filename_out = '.'.join(filename_split)
folders_split[-1] = filename_out
path_out = '\\'.join(folders_split)
reproject_raster(path_in, path_out)
|
jupyter-notebooks/2 - reproject_sentinel2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Words Analysis
#
# This analysis analyses collected text and meta-data from scientific articles about that mention ERPs.
#
# Available information includes the titles, words, and years of all those papers such that this data can be used for further analysis.
# +
# %matplotlib inline
from collections import Counter
import numpy as np
import matplotlib.pyplot as plt
from lisc import Words
from lisc.data import ArticlesAll
from lisc.data.utils import threshold_counter, combine_lists
from lisc.utils import SCDB, load_object
from lisc.utils.io import load_txt_file
from lisc.plts.words import plot_years, plot_wordcloud
from lisc.analysis.words import get_all_counts, get_all_values, get_attribute_counts
# -
import seaborn as sns
sns.set_context('talk')
# Import custom project code
import sys
sys.path.append('../code')
from plts import plot_count_hist, plot_attrs_by_year
# ## Setup
# Settings
SAVE_FIG = False
# Set the file locations
term_dir = '../terms/'
figs_dir = '../data/figures/words'
db = SCDB('../data/')
# Set the name of the file to load
name = 'erps' # 'erps', 'test'
# Load the words object
words = load_object('words_' + name, directory=db, reload_results=True)
# Load exclusions for analysis
exclusions = load_txt_file('analysis_exclusions.txt', term_dir, split_elements=False)
# ### Check Database Information
#
# Check the metadata about the data collection, including checking the database data were collected from.
# Check database scrape info
words.meta_data.db_info
# Check requester object info
words.meta_data.requester
# ## Data Organization
# Process data to collapse across all articles
words.process_combined_results(exclusions=exclusions)
# ### Check Number of Articles
# Check the number of ERPs
print('Number of ERPs: \t {}'.format(words.n_terms))
# Check the total number of articles in the collected dataset
n_articles = [data.n_articles for data in words]
total_article_count = sum(n_articles)
print('Number of articles: \t', total_article_count)
# Plot the histogram of the number of ERP papers collected
plot_count_hist(n_articles, log=True, bins=10,
xlabel='Number of articles', ylabel='Count of ERP Components',
save_fig=SAVE_FIG, file_name='erp_hist.pdf', directory=figs_dir)
# Collect all the DOIs across all papers
dois = get_all_values(words, 'dois', unique=True)
# Check the number of unique papers, based on DOIs
print('Number of unique DOIs: \t', len(set(dois)))
# Check how many articles were found per ERP
for data in words.results:
print(data.label, '\t', data.n_articles)
# ## Check Attribute Counts
# Define a list of attributes to check
attrs = ['words', 'keywords', 'authors', 'journals', 'dois', 'titles', 'years']
# Check the proportion of papers, for each attribute, that has data
print('Attribute counts (total # articles: {}):'.format(total_article_count))
for attr in attrs:
count = get_attribute_counts(words, attr)
print(' {:5d} articles ({:6.2f} %) have {}.'.format(\
count, count / total_article_count * 100, attr))
# ### Check Years of Publication
# Get global count of authors
years = set(get_all_values(words, 'years'))
min_year, max_year = min(years), max(years)
# Print out summary of the year range of publications
print('Collected papers were published between the years: {}-{}'.format(min_year, max_year))
# Get the count of articles per year, and drop 2021 (incomplete year)
year_counts = get_all_counts(words, 'years', combine=True)
year_counts.pop(2021);
# Plot the total number of articles, across components, per year
plot_years(year_counts, figsize=(6, 5),
save_fig=SAVE_FIG, file_name='words_year_counts.pdf', directory=figs_dir)
# ### Check number of distinct components studies per year
# Collect the set of years each component was studied in
comp_years_all = {}
for label, res in zip(words.labels, words.results):
comp_years_all[label] = set(res.years)
# Collect the number of unique components studied per year
comp_years = {}
for year in range(min(years), max(years)):
count = 0
for temp in comp_years_all.values():
count += 1 if year in temp else 0
comp_years[year] = count
# Plot the number of unique components studied per year
plot_years(comp_years, ylabel='Unique Components', figsize=(6, 5),
save_fig=SAVE_FIG, file_name='words_year_components.pdf', directory=figs_dir)
# ### Check Authors across all papers
# Get global count of authors
author_counts = get_all_counts(words, 'authors', combine=True)
# Check the number of unique authors
print('Total number of authors :\t {:5d}'.format(len(author_counts)))
print('Number of authors > 10 :\t {:5d}'.format(len(threshold_counter(author_counts, 10))))
print('Number of authors > 100 :\t {:5d}'.format(len(threshold_counter(author_counts, 100))))
# Check the most common authors
print('Most common authors:')
for (author, count) in author_counts.most_common(10):
print(' {:15s} \t {}'.format(' '.join(author), count))
# ### Check Journals across all papers
# Get global count of journals
journals_counts = get_all_counts(words, 'journals', combine=True)
# Check the number of unique authors
print('Total number of journals :\t {:5d}'.format(len(journals_counts)))
print('Number of journals > 10 :\t {:5d}'.format(len(threshold_counter(journals_counts, 10))))
print('Number of journals > 100 :\t {:5d}'.format(len(threshold_counter(journals_counts, 100))))
# Check the most common journals
print('Most common journals:')
for (journal, count) in journals_counts.most_common(10):
print(' {:55s} \t {}'.format(journal.split(':')[0], count))
# ### Keywords
# Get global count of authors
kw_counts = get_all_counts(words, 'keywords', combine=True)
# Check the number of unique keywords
print('Total number of keywords :\t {:5d}'.format(len(kw_counts)))
print('Number of keywords > 10 :\t {:5d}'.format(len(threshold_counter(kw_counts, 10))))
print('Number of keywords > 100 :\t {:5d}'.format(len(threshold_counter(kw_counts, 100))))
# Check the most common keywords across all components
print('Most common keywords:')
for (kw, count) in kw_counts.most_common(10):
print(' {:20s} \t {}'.format(kw, count))
# Check most common keywords per component
for data in words.combined_results:
data.check_frequencies('keywords', n_check=5)
# ### Abstract Words
# Get global count of words
word_counts = get_all_counts(words, 'words', combine=True)
# Check the number of unique words
print('Total number of words :\t {:5d}'.format(len(word_counts)))
print('Number of words > 100 :\t {:5d}'.format(len(threshold_counter(word_counts, 100))))
print('Number of words > 1000 :\t {:5d}'.format(len(threshold_counter(word_counts, 1000))))
# Check the most common keywords across all components
print('Most common words:')
for (word, count) in word_counts.most_common(10):
print(' {:20s} \t {}'.format(word, count))
# Check most common words
for data in words.combined_results:
data.check_frequencies('words', n_check=5)
# ## Check Values by Year
# Extract lists of all values for attributes of interest
all_years = get_all_values(words, 'years')
all_journals = get_all_values(words, 'journals')
all_authors = get_all_values(words, 'authors')
# Collect the number of unique journals & authors by year
journals_by_year = {}
authors_by_year = {}
for year in range(min_year, max_year):
inds = np.where(np.array(all_years) == year)[0]
journals_by_year[year] = len(set(np.array(all_journals)[inds]))
authors_by_year[year] = len(set(combine_lists([all_authors[ind] for ind in inds])))
# Plot unique journals & authors by year
plot_attrs_by_year(journals_by_year, authors_by_year, figsize=(6, 5),
save_fig=SAVE_FIG, file_name='words_attr_year.pdf', directory=figs_dir)
# ## Example Component Summaries
# Set indices for some example components
inds = [15, 51]
print(words.labels[inds[0]], words.labels[inds[1]])
# Check some example summaries
for ind in inds:
words.combined_results[ind].create_summary()
words.combined_results[ind].print_summary()
# ### WordCloud Plots
# Create wordclouds
_, axes = plt.subplots(1, 2, figsize=(15, 6))
plot_wordcloud(words.combined_results[ind].words, inds[0], ax=axes[0])
axes[0].set_title(words.labels[inds[0]]);
plot_wordcloud(words.combined_results[ind].words, inds[1], ax=axes[1])
axes[1].set_title(words.labels[inds[1]]);
# ### History Plots
# Check an example history plot
_, axes = plt.subplots(1, 2, figsize=(15, 6))
plot_years(words.combined_results[inds[0]].years, [None, 2020], ax=axes[0])
axes[0].set_title(words.labels[inds[0]]);
plot_years(words.combined_results[inds[1]].years, [None, 2020], ax=axes[1])
axes[1].set_title(words.labels[inds[1]]);
|
notebooks/02-WordsAnalyses.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Removendo dados em estruturas existentes
# - declaração DELETE remove uma ou mais linhas de uma tabela ou view;
# - DELETE é diferente de fazer um DROP (que remove a tabela em si). No DELETE você só esá apagando uma das linhas da sua tabela.
# - o parâmetro table_sources pode ser usado para especificar tabelas ou views adicionais que podem ser usadas na cláusula WHERE;
# - Muita gente não remove linha, são práticas novas, pois antigamente espaço de disco era muito caro, atualmente não. Como o espaço de disco ficou barato, muita gente não usa o comando DELETE. As pessoas preferem criar um comando a mais, uma flag e dizer: "tupla utilizada ou não". Quando for remover, apenas põe como flag "tupla não utilizada". Assim, aquela entidade se mantém integra, você ainda consegue acessar aquela entidade que foi removida, mas você ainda não a removeu do seu banco de dados. A vantagem é que o comando DELETE, assim como o DROP, você não tem como voltar atrás. Uma vez que você deleta uma linha ela some do seu banco de dados. O mesmo vale para o DROP, se você dropar uma tabela ou coluna, ela não existe mais. Por isso muita gente prefere adicionar uma coluna a mais e manter a linha no banco de dados do que remover.
# - Mas há casos em que precisamos remover o dado, por isso que esses comandos ainda existem no banco de dados e na linguagem SQL.
# - Comando DELETE é similar ao INSERT. Leia assim: delete dado/tabela/visão da tabela x, aonde você tem uma condição.
# __Sintaxe do DELETE__
# `DELETE table_or_view -- DELETE nome_da_tabela/visao_q_vc_quer_remover
# FROM table_sources
# WHERE search_condition`
# __DELETE sem a cláusula WHERE__
# `DELETE FROM SomeTable;`
# Ex.: DELETE FROM Sales.SalesPerson; -> removo o dado da tabela x. Não estou informando quais dados estou removendo. A cláusula está bem limpa, neste caso ele vai apagar todos os dados da minha tabela. Apago a tabela sem apagar a estrutura da minah tabela. A diferença desse comando para o DROP table é que eu estou apagando a tabela e ai eu teria que criá-la de novo. Com o DELETE FROM e o nome da tabela em si eu estou apagando os dados da tabela, mas ainda mantenho a estrutura da tabela.
#
# __DELETE usando TOP__
# Do mesmo modo como há o comando TOP para INSERT, eu posso apagar algumas linhas iniciais e não todos os dados da minha tabela.
# `DELETE TOP (1)
# FROM SomeTable;`
# Ex.:
# DELETE TOP (2.5) PERCENT
# FROM Production.ProductInventory;
# Veja que apago as linhas principais, utilizo um percentual, apago o top 2.5% da minha tabela. Independente do que for o conteúdo, ele apaga as 2,5% de linhas que aparecem. Esse comando é útil quando você está trabalhando com teste, com algum exemplo em que o dado não importa, você pode apagar para ter mais espaço em disco ou algum outro motivo.
#
# __DELETE usando uma Subquery__
# Também posso fazer subquery, que nem na inserção de dados. Posso fazer uma consulta, a qual me dá um resultado e este resultado que eu estou apagando. No exemplo abaixo apago um dado através de uma query de consulta. Apago o dado da tabela x, aonde o ID se encontra em um SELECT. Por ex. quem fez menos vendas eu apago. Ou delete da tabela x quem começa com o nome Pedro, ou quem tem o ID x... Eu posso fazer esse tipo de construção no meu comando DELETE.
# `DELETE FROM SomeTable
# WHERE SomeColumn IN
# (Subquery Definition)`
#
# `DELETE FROM
# Sales.SalesPersonQuotaHistory
# (SELECT SalesSalesPersonID
# FROM Sales.SalesPerson
# WHERE SalesYTD > 2500000.00;`
#
# A vantagem é que vo0cê pode querer apagar só quem tem por ex. o CPF entre 1000 e 2.00.000, ou quem nasceu entre 70 e 80, já que não quero apagar o resto dos dados. Através das opções do WHERE eu consigo construir quais situações eu estou querendo apagar. Quais linhas eu estou selecionando.
# - Também consigo truncar a tabela. Truncar a tabela é eu conseguir quebrar a tabela em partes.
# - A declaração TRUNCATE é mais rápida que o DELETE, mas não há como restringir as linhas que serão removidas através da cláusula WHERE, não consigo utilizar o comando WHERE (essa é uma desvantagem). Já no DELETE eu consigo usar o WHERE, porém é mais lento.
# __Sintaxe do TRUNCATE TABLE__
# `TRUNCATE TABLE
# [ { database_name.[ schema_name ]. | schema_name . } ]
# table_name
# [ ; ]`
#
# Ex.:
# `TRUNCATE TABLE Cliente;`
# `TRUNCATE TABLE Sales.SalesPerson;`
# A boa prática ao usar o comando DELETE é aplicar um SELECT antes, para verificar se os dados retornados são os que queremos eliminar. Dessa forma teremos certeza de que estamos criando o campo WHERE corretamente.
# `SELECT name FROM Cliente
# WHERE name like 'marcalo%';`
# Caso o retorno seja o que queremos eliminar, substituimos o SELECT pelo DELETE. Essa prática não consome muit tempo e vai lhe dar segurança oa usar o comando DELETE sem medo de apagar um dado, sem apagar algo que não tem mais volta (a não se rque você tenha um backup do banco ou da tabela).
# `DELETE FROM Cliente
# WHERE name like 'marcelo%';`
#
# P.s.: vamos aprender a fazer consultas mais adiante.
#
# # 2. Atualizando dados em estruturas existentes
# - A declaração UPDATE altera valores dos dados de uma ou mais linhas de uma tabela. No caso você já tem um valor inserido;
# - Por ex. você nasceu em 01/01/2000, porém você errou a data e nasceu em 01/01/2007. Você não precisa apagar toda essa linha e inseri-la de novo. Você consegue alterar o valor através do comando UPDATE.
# - Uma declaração UPDATE referenciando uma table or view pode alterar os dados somente em uma tabela ao mesmo tempo;
# - UPDATE tem 3 claúsulas principais:
# - SET – Lista de campos, separados por vírgula, que serão alterados.
# - FROM – Fornece objetos fonte para a cláusula SET.
# - WHERE – Especifica a condição de procura para aplicar as alterações com a cláusula SET.
# __Sintaxe do UPDATE__
# `UPDATE table_or_view
# SET column_name = expression
# FROM table_sources
# WHERE search_condition`
#
# Leia assim: UPDATE tabela x, SET qual o conjunto que você está mudando (ex. set coluna_placa pelo valor x), FROM nome_da_tabela, e o WHERE você dá a condição. Você pode ou não usar o WHERE, se você não usar o WHERE você está alterando todas as colunas, todos os valores que estão naquela coluna (informada no SET). Por isso é recomendável usar o WHERE para garantir que você não está cometendo nenhum equívoco e está fazendo a mudança correta.
# Ex.:
#
# __Declaração simples do UPDATE__
# `UPDATE SomeTable
# SET Column = Value`
#
# UPDATE tabela venda, eu mudo o Bonus para 6000 independente de qualquer cláusula. Todos os bônus estão sendo alterados para 6000.
# `UPDATE Sales.SalesPerson
# SET Bonus = 6000;`
#
# Veja que posso fazer operações matemáticas através do UPDATE, por ex.: pego o Bonus e multiplico por 2.
# `UPDATE Sales.SalesPerson
# SET Bonus = Bonus * 2;`
#
# __UPDATE COM A CLÁUSULA WHERE__
# `UPDATE SomeTable
# SET Column = Value
# WHERE SearchExpression`
#
# `UPDATE Production.Product
# SET Color = N’Metallic Red’
# WHERE Name LIKE ’Road-250%’
# AND Color = ’Red’;`
# Leia assim: update a tabela Product, onde eu estou mudando Color que vai ser Metallic Red; aonde na tabela name aonde Road-250% e cor é red. Ou seja, altero a cor onde é red e onde o campo é Road-250%. Dou regras, em que conforme o DELETE você tem que rodar um SELECT antes para ver se está condizente, se realmente é aquele campo que você quer apagar, no UPDATE também é bom fazer isso. Ignora a parte do SET e ao invés disso faz um SELECT * FROM Product WHERE... para ver se os campos estão coerentes. Se é o que realmente você quer mudar. É um tempo a mais, mas que vai lhe dar uma tranquilidade.
#
# __UPDATE usando uma Subquery__
# Também posso fazer que nem o DELETE e o INSERT, posso fazer uma subquery, em que eu estou executando um UPDATE aonde o meu WHERE tem uma subqeury. Parte do meu WHERE é o reusltado de alguma busca.
# `UPDATE SomeTable
# SET Column = Value
# FROM SomeSubquery
# `
#
# `UPDATE Sales.SalesPerson
# SET SalesYTD = SalesYTD + SubTotal
# FROM Sales.SalesPerson AS sp
# JOIN Sales.SalesOrderHeader AS so
# ON sp.BusinessEntityID = so.SalesPersonID
# AND so.OrderDate = (SELECT MAX(OrderDate)
# FROM Sales.SalesOrderHeader
# WHERE SalesPersonID = sp.BusinessEntityID);`
#
#
# __Exemplo:__
# `
# Insert into Aluno (Nome) VALUES (‘Matheus’ )
# Insert into Prova ( Matricula, Nota ) VALUES (503, 10 )
# Delete from prova where Matricula = 500
# Delete from aluno where Matricula = 500`
# - Veja que eu faço o Delete duas vezes, pois não é porque eu apaguei de uma tabela, que ele apaga de outra tabela. Tenho que apagar de todas as tabelas, a não ser que tenha alguma CONSTRAINT que me dá alguma ordem ou regra específica.
# - Vou alterar a matricula da Prova, de 502 para 504:
# `Insert into Aluno (Nome) VALUES (‘Felipe’ ) --inseri Felipe
# Update prova set Matricula = 504 where matricula = 502
# `
#
# 
# 
#
#
#
# # 3. Exercícios
# - Vide no sqlfiddle: http://sqlfiddle.com/#!9/de747e/2
|
SQL/.ipynb_checkpoints/5. DML - Data Manipulation Language - parte 2 - UPDATE e DELETE-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import gzip
# Plotting gender stuff
df = pd.read_csv(gzip.open('data_cleaned/cleanedBigGun_gender.csv.gz'))
df.head()
df.columns
data = pd.DataFrame(df.groupby('year').sum()['# of Male Victims'])
data['# of Female Victims'] = df.groupby('year').sum()['# of Female Victims']
data['# of Male Suspects'] = df.groupby('year').sum()['# of Male Suspects']
data['# of Female Suspects'] = df.groupby('year').sum()['# of Female Suspects']
data.reset_index(inplace = True)
data.rename(columns={"year": "Year"}, inplace = True)
data
# +
from bokeh.core.properties import value
from bokeh.io import show, output_file
from bokeh.models import ColumnDataSource, HoverTool
from bokeh.plotting import figure
from bokeh.transform import dodge
from bokeh.models.widgets import DataTable, TableColumn
from bokeh.layouts import layout
output_file('genderBarCharts.html')
width_of_each_bar = .2
data.Year = data.Year.astype(str)
data['# of Male Victims'] = data['# of Male Victims'].astype(str)
data['# of Female Victims'] = data['# of Female Victims'].astype(str)
df = data.to_dict(orient='list')
years = df['Year']
data_max = data[['# of Female Suspects','# of Female Victims','# of Male Suspects','# of Male Victims']].apply(pd.to_numeric).values.max()
source = ColumnDataSource(data=df)
p = figure(x_range=years, y_range=(0, data_max + 3000),
plot_height=600, plot_width=650, title="Gender Count of Victims and Suspects by Year",
toolbar_location=None)
renderers1 = p.vbar(x=dodge('Year', -0.3, range=p.x_range), top='# of Male Victims', width=width_of_each_bar, source=source,
color="blue", legend=value("# of Male Victims"))
hover1 = HoverTool(tooltips=[
("Year", "@Year"),
("# of Male Victims", "@{# of Male Victims}{0,0[.]00}")
], renderers=[renderers1])
p.add_tools(hover1)
renderers2 = p.vbar(x=dodge('Year', -0.1, range=p.x_range), top='# of Female Victims', width=width_of_each_bar, source=source,
color="lightskyblue", legend=value("# of Female Victims"))
hover2 = HoverTool(tooltips=[
("Year", "@Year"),
("# of Female Victims", "@{# of Female Victims}{0,0[.]00}")
], renderers=[renderers2])
p.add_tools(hover2)
renderers3 = p.vbar(x=dodge('Year', 0.1, range=p.x_range), top='# of Male Suspects', width=width_of_each_bar, source=source,
color="red", legend=value("# of Male Suspects"))
hover3 = HoverTool(tooltips=[
("Year", "@Year"),
("# of Male Suspects", "@{# of Male Suspects}{0,0[.]00}")
], renderers=[renderers3])
p.add_tools(hover3)
renderers4 = p.vbar(x=dodge('Year', 0.3, range=p.x_range), top='# of Female Suspects', width=width_of_each_bar, source=source,
color="pink", legend=value("# of Female Suspects"))
hover4 = HoverTool(tooltips=[
("Year", "@Year"),
("# of Female Suspects", "@{# of Female Suspects}{0,0[.]00}")
], renderers=[renderers4])
p.add_tools(hover4)
#p.x_range.range_padding = 0.2
p.xgrid.grid_line_color = None
p.legend.location = "top_center"
p.legend.orientation = "horizontal"
columns = [
TableColumn(field="Year", title="Year"),
TableColumn(field="# of Male Victims", title="# of Male Victims"),
TableColumn(field="# of Female Victims", title="# of Female Victims"),
TableColumn(field="# of Male Suspects", title="# of Male Suspects"),
TableColumn(field="# of Female Suspects", title="# of Female Suspects")
]
p_data = DataTable(source=source, columns=columns, width=600, height=600, selectable = True)
# show the results
show(layout([p, p_data]))
|
.ipynb_checkpoints/genderGraphing-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# https://github.com/wesm/pydata-book/tree/1st-edition
# +
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# used for example for random
from numpy import *
# for matplot
# %matplotlib inline
# -
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# # <font color='green'>FEATURES</font> #
#
# <font color="#000000">
# <br><br>
# <ol type="1">
# <li>Arrays enable you to perform mathematical operations on whole blocks of data using similar syntax to the equivalent
# operations between scalar elements
# <br><br>
# <li>Arrays are important because they enable you to express batch operations on data
# without writing any for loops. This is usually called <font color="red">**vectorization**.</font>
#
# </font>
# +
data = np.array([[0.9526, -0.246 , -0.8856],
[ 0.5639, 0.2379, 0.9104]])
data
data*10
data+data
# -
# An ndarray is a generic multidimensional container for homogeneous data; that is, **all
# of the elements must be the same type**.
# +
type(data)
data.shape
data.dtype
# -
# # <font color='green'>CREATION</font>
# array creation routines
# <br>https://docs.scipy.org/doc/numpy-1.13.0/reference/routines.array-creation.html
# +
# creating tuple, list and than converting to ndarrray
data1 = [6, 7.5, 8, 0, 1]
type(data1)
data1
arr1 = np.array(data1)
type(arr1)
arr1
arr1.dtype
# Additional excercises:
# convert to tuple
# check for int
# -
#
# <ol type="1">
# <li><font color='blue'>Number of array dimensions.</font>
#
# ```python
# ndarray.ndim
# ```
# </li>
# <li>
# <font color='blue'>Length of one array element in bytes.</font>
#
# ```python
# ndarray.itemsize
# ```
#
# </li>
# <li>
# <font color='blue'>Number of elements in the array.<br>
#
# Equivalent to np.prod(a.shape), i.e., the product of the array’s dimensions.</font>
# ```python
# ndarray.size
# ```
# </li>
# <li>
# <font color='blue'>Tuple of array dimensions.</font>
# ```python
# ndarray.shape
# ```
# </li>
#
# +
# 2-dimensional array
data2 = [[1, 2, 3, 5], [5, 6, 7, 8]]
arr2 = np.array(data2)
arr2
arr2.ndim
arr2.shape
arr2.itemsize
arr2.size
# multiplication of all elements in array
np.prod(data2, axis = 0)
# -
# **<font color='green'>CREATION of dummy</font>**
# +
# arrays of 0's
np.zeros(10)
# arrays of 1's
np.ones((2, 8))
# arrays of garbage values empty(#of arrays, rows, columns)
empty_ex = np.empty((2, 3, 2))
empty_ex
empty_ex.shape
#Return a new array with the same shape and type as a given array.
arr2
np.empty_like(arr2)
#identity matrix
np.identity(3)
#Return a 2-D array with ones on the diagonal and zeros elsewhere.
np.eye(3)
#Return a new array of given shape and type, filled with fill_value.
np.full((2, 2), 12)
# -
#arange is an array-valued version of the built-in Python range function:
np.arange(15)
# **<font color='green'>DTYPE</font>**
#
# supported dtypes:
# <img src="IMG\data_type1.png">
# <img src="IMG\data_type2.png">
# +
arr1 = np.array([1, 2, 3], dtype=np.float64)
arr1.dtype
arr2 = np.array([1, 2, 3], dtype=np.int32)
arr2.dtype
# +
arr = np.array([3.7, -1.2, -2.6, 0.5, 12.9, 10.1])
arr
#change from float to int result in trunc of numbers
arr.astype(np.int32)
#changing form string to float
numeric_strings = np.array(['1.25', '-9.6', '42'], dtype=np.string_)
numeric_strings
numeric_strings = numeric_strings.astype(float)
numeric_strings
numeric_strings.dtype
#!! Calling astype always creates a new ab (a copy of the data), even if the new dtype is the same as the old dtype.
# -
# ================================================================================================================================
# <font color='green'>Matrix calculations element-wise</font>
#
# +
arr = np.array([[1., 2., 3.], [4., 5., 6.]])
print "arr matrix"
arr
arr.shape
print "arrt - transposed matrix"
arrt = arr.transpose()
arrt
print "============================================="
print "Matrix operations element-wise"
arr * arr
arr - arr
arr + arr
arr*2
1/arr
a = 0.5
arr ** a
# -
# # <font color='green'>SLICE and DICE</font>
#
# <font color="#000000">
# <ol type="1">
# An important first distinction
# from lists is that array slices are views on the original array. This means that
# <font color="red">the data is not copied, and any modifications to the **view** will be reflected in the source array</font>
#
# As NumPy has
# been designed with large data use cases in mind, you could imagine performance and
# memory problems if NumPy insisted on copying data left and right.
# +
#1-dimensional arrays
arr = np.arange(10)
arr
arr_slice = arr[5:8]
print "change of 1st element of arr_slice result change in arr 6th element"
arr_slice[1] = 12345
arr
arr_slice[:] = 64
arr
print "if you want to copy array instead of work on view you can use .copy() function"
arr_copy = arr[5:8].copy()
arr_copy
arr_copy[1] = 12345
arr
arr_copy
# -
# <font color='green'>arrays indexing </font>
#
# Matrix multiplicaton
# If **A** is an **n × m** matrix and **B** is an **m × p** matrix,
# <img src="np_arrays.PNG">
# +
#m-dimensional-arrays
arr2d = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
arr2d
arr2d[2]
#[row,column]
arr2d[0][2]
arr2d[0, 2]
# +
names = np.array(['Bob', 'Joe', 'Will', 'Bob', 'Will', 'Joe', 'Joe'])
# assigning random values
data = np.random.randn(7, 4)
names
data
print "=============================================================================================="
names == 'Bob'
data[names == 'Bob']
data[names == 'Bob', 2:]
data[names == 'Bob', 3]
#!! Selecting data from an array by boolean indexing always creates a copy of the data, even if the returned array is unchanged.
# +
names = np.array(['Bob', 'Joe', 'Will', 'Bob', 'Will', 'Joe', 'Joe'])
names != 'Bob'
data[-(names == 'Bob')]
print "Selecting two of the three names to combine multiple boolean conditions, use boolean arithmetic operators like & (and) and | (or):"
mask = (names == 'Bob') | (names == 'Will')
mask
data[mask]
# +
data
data[data < 0] = 0
data
data[names != 'Joe'] = 7
data
# +
arr = np.empty((8, 4))
for i in range(4):
arr[i] = i
arr
arr[1, 2:]
#To select out a subset of the rows in a particular order, you can simply pass a list or ndarray of integers specifying the desired order:
arr[[4, 3, 0, 6]]
# -
# <font color='green'>**FANCY INDEXING** </font>
# +
arr = np.arange(32)
arr
arr_reshape = arr.reshape((8, 4))
arr_reshape
print" Passing multiple index arrays does something slightly different; it selects a 1D array of elements corresponding to each tuple of indices:\
in below example we are taking pairs [1,0] [5,3] [7,1] [2,2]"
arr_reshape[[1, 5, 7, 2], [0, 3, 1, 2]]
#incorrect!!!
arr_reshape[[1,0],[5,3],[7,1],[2,2]]
###!!! Keep in mind that fancy indexing, unlike slicing, always copies the data into a new array.
# -
# <img src="fancy_slice.PNG">
# +
#use to chnage the order of matrix np.ix_([rows], [columns])
arr_reshape[np.ix_([1, 5, 7, 2], [0, 3, 1, 2])]
# -
# # <font color='green'>TRANSPOSING ARRAYS AND SWAPPInG AXES</font>
# Transposing is a special form of reshaping which similarly returns a view on the underlying
# data without copying anything. Arrays have the transpose method and also
# the special T attribute
#
# ```python
# arr.t
# arr.transpose()
# ```
# <br>
# computing
# the inner matrix product XTX using np.dot:
# ```python
# np.dot(arr.T, arr)
# ```
# +
arr = np.array([[1., 2., 3.], [4., 5., 6.]])
print "XtX"
arr
arr.T
arr.T * arr # element-wise matrix multiplication
np.dot(arr.T, arr) # normal matrix multiplication
arr.transpose()
# +
arr = np.arange(16).reshape((2, 2, 4))
arr
#to sa permutacje, ja nie rouzmiem permutacji :/
#https://pl.wikipedia.org/wiki/Permutacja
arr.transpose((1, 0, 2))
arr.transpose((2, 0, 1))
# -
# ## swaping axes - > reurn view not copy
# arr = np.arange(16).reshape((2, 2, 4))
# arr
# arr.swapaxes(1, b1)
arr = np.arange(16).reshape((2, 2, 4))
arr
arr.swapaxes(1, 2)
# # <font color='green'>Universal Functions: Fast Element-wise Array Functions</font>
# <img src="IMG\unary_ufuncs.PNG">
#
# <img src="IMG\binary_fun.PNG">
# +
arr = np.arange(10)
arr
print"========================================="
print"SQRT"
np.sqrt(arr)
arr ** 0.5
# -
#take 2 arrays(thus, binary ufuncs) and return a single array as the result:
x = np.arange(10)
x[5] = 20
y = np.arange(3,13)
z = np.array(np.random.rand(10))*10
x
y
z
np.max(x)
np.max(y)
np.maximum(x, z) # element-wise maximum
np.maximum(x, y) # element-wise maximum
# +
#modf Return the fractional(poprzecinku) and integral(naturalne) parts of an array, element-wise.
arr = np.arange(3,13)*0.75
arr
np.modf(arr)
# +
#arange(start, stop, distance)
points = np.arange(-5, 5, 0.01)
points1 = np.arange(0, 5)
points2 = np.arange(5,10)
points1
points2
points[:10]
points.shape
xs, ys = np.meshgrid(points1, points2)
ys
xs
#import matplotlib.pyplot as plt
z = np.sqrt(xs ** 2 + ys ** 2)
plt.imshow(z, cmap=plt.cm.gray); plt.colorbar()
plt.title("Image plot of $\sqrt{x^2 + y^2}$ for a grid of values")
# -
# # <font color='green'>Expressing Conditional Logic as Array Operations</font>
#
# A typical use of where in data analysis is to produce a new array of values based on another array.
#
# ```python
# np.where
# ```
#
# <img src="np_where.PNG">
#
# https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.where.html
#
#
# +
xarr = np.array([1.1, 1.2, 1.3, 1.4, 1.5])
yarr = np.array([2.1, 2.2, 2.3, 2.4, 2.5])
cond = np.array([True, False, True, True, False])
xarr
cond
yarr
#a list function (not effective as it's working only with 1d array and have poor performance)
result = [(x if c else y)
for x, y, c in zip(xarr, yarr, cond)]
result
# numpy is easier :)
result = np.where(cond, xarr, yarr)
result
print"The second and third arguments to np.where don’t need to be arrays; one or both of\
them can be scalars"
result = np.where(cond, xarr, 0)
result
# +
#Suppose you had a matrix of randomly generated data and you wanted to replace \
#all positive values with 2 and all negative values with -2.\
#This is very easy to do with np.where:\
arr = np.random.randn(4, 4)
arr
np.where(arr > 0, 2, -2)
np.where(arr > 0, 2, arr) # set only positive values to 2
# -
# <font color='green'>**NESTED np.where** </font>
# +
cond1 = np.array([True, False, False, True, False, True])
cond2 = np.array([False, True, False, True, False, True])
np.where(cond1 & cond2, 0,
np.where(cond1, 1,
np.where(cond2, 2, 3)))
# -
# # <font color='green'>Mathematical and Statistical Methods</font>
#
# <img src="IMG\bas_stat.PNG">
#
# +
arr = np.arange(12).reshape((3, 4))
arr
arr.mean()
np.mean(arr)
arr.sum()
#Functions like mean and sum take an optional axis argument which computes the statistic
#over the given axis, resulting in an array with one fewer dimension:
arr.mean(axis=0) # by columns
arr.mean(axis=1) # by rows
arr.sum(0) #by columns
# -
#Other methods like cumsum and cumprod do not aggregate, instead producing an array of the intermediate results:
arr = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
arr
arr.cumsum(0)
arr.cumprod(1)
# <font color='green'>**Methods for Boolean Arrays** </font><br><br>
# Boolean values are coerced to 1 (True) and 0 (False) in the above methods. Thus, sum
# is often used as a means of counting True values in a boolean array:
#
# ```python
# np.any()
# np.all()
# ```
#
# Methods any() and all() are useful especially for boolean arrays.
# any tests whether one or more values in an array is True, while all checks if every value
# is True
# +
arr = np.array([[0, 3, 8], [1, 4, 2], [6, 3, 8]])
arr
(arr > 3).sum() # Number of positive values
bools = np.array([False, False, True, False])
bools2 = np.array([True, True, True, True])
bools3 = np.array([False, False, False, False])
bools.any()
bools3.any()
bools.all()
bools2.all()
# -
# ================================================================================================================================
# # <font color='green'>SORTING & SET OPERATIONS</font>
#
# <img src="IMG\set_ops.PNG">
#
# +
arr = np.array([[5, 0, 8], [1, 4, 2], [6, 3, 8]])
arr
arr.sort() #by default rows
arr
arr2 = np.array([[5, 0, 8], [1, 4, 2], [6, 3, 8]])
arr2.sort(0) #by columns
arr2
arr3 = np.array([[5, 0, 8], [1, 4, 2], [6, 3, 8]])
arr3.sort(1) #by rows
arr3
# -
#The top level method np.sort returns a sorted copy of an array instead of modifying
#the array in place. A quick-and-dirty way to compute the quantiles of an array is to sort
#it and select the value at a particular rank:
large_arr = np.random.randn(1000)
type(large_arr)
large_arr.sort()
large_arr[int(0.05 * len(large_arr)):] # 5% quantile
names = np.array(['Bob', 'Joe', 'Will', 'Bob', 'Will', 'Joe', 'Joe'])
np.unique(names)
ints = np.array([3, 3, 3, 2, 2, 1, 1, 4, 4])
np.unique(ints)
# function,
#
# ```python
# np.in1d
# ```
# tests membership of the values in one array in another,
# returning a boolean array:
values = np.array([6, 0, 0, 3, 2, 5, 6])
np.in1d(values, [2, 3, 6])
# ================================================================================================================================
# # <font color='green'>File Input and Output with Arrays</font>
#
# ```python
# np.save
# np.load
# ```
# are the two workhorse functions for efficiently saving and loading
# array data on disk. Arrays are saved by default in an uncompressed raw binary format
# with file extension **__.npy.__**
#
#
#
#
arr = np.arange(10)
np.save('some_array', arr)
# <img src="save.PNG">
bla = np.load('some_array.npy')
bla
# You save multiple arrays in a zip archive using np.savez and passing the arrays as keyword
# arguments:
arr2 = ([1,5,6,8,4])
np.savez('array_archive.npz', a=arr, b=arr2)
# <img src="IMG\archive.PNG">
#When loading an .npz file, you get back a dict-like object which loads the individual
#arrays lazily:
arch = np.load('array_archive.npz')
arch['b']
arch['a']
# <font color='green'>**Saving and Loading Text Files** </font><br><br>
arr = np.loadtxt('array_ex.txt', delimiter=',')
arr
# np.savetxt performs the inverse operation: writing an array to a delimited text file.
# genfromtxt is similar to loadtxt but is geared for structured arrays and missing data
# handling; see Chapter 12 for more on structured arrays.
# +
arr = np.array([[0.580052, 0.18673 , 1.040717, 1.134411],
[ 0.194163, -0.636917, -0.938659, 0.124094],
[-0.12641 , 0.268607, 0, 0.047428],
[-1.484413, 0.004176, -0.744203, 0.005487],
[ 2.302869, 0.200131, NaN, -1.88109 ],
[-0.19323 , 1.047233, 0.482803, 0.960334]])
np.savetxt('test.txt', arr)
arr1 = np.genfromtxt('test.txt', delimiter=' ')
arr1
arr2 = np.loadtxt('test.txt', delimiter=' ')
arr2
# -
# ================================================================================================================================
# # <font color='green'>LINEAR ALGEBRA</font>
# Linear algebra, like matrix multiplication, decompositions, determinants, and other
# square matrix math, is an important part of any array library. Unlike some languages
# like MATLAB, multiplying two two-dimensional arrays with _*_ is an element-wise
# product instead of a matrix dot product. As such, there is a function dot, both an array
# method, and a function in the _numpy_ namespace, for matrix multiplication:
#
#
# If **A** is an **n × m** matrix and **B** is an **m × p** matrix,
# <img src="matrix_multiplication.PNG">
#
# +
x = np.array([[1., 2., 3.], [4., 5., 6.]])
y = np.array([[6., 23.], [-1, 7], [8, 9]])
x
x.shape
y
y.shape
x.dot(y) # equivalently np.dot(x, y)
y.dot(x)
z = np.array([[1,2], [8,9], [9,10], [1,6]])
y.dot(z)
# -
# <img src="IMG\linalg.PNG">
# **Rozkład QR** – w algebrze liniowej rozkład macierzy A do postaci iloczynu dwóch macierzy A = QR, gdzie Q jest macierzą ortogonalną ( QTQ = I ) i R jest macierzą trójkątną górną [1]. Na bazie rozkładu QR możliwa jest realizacja metody najmniejszych kwadratów[2] oraz metod rozwiązywania układów równań liniowych[1].
#
# matrix inverse: https://www.mathsisfun.com/algebra/matrix-inverse.html
# macierze: http://matematykawwakacje.pl.tl/Macierze.htm#Macierz_ortogonalna
#
# ```python
# numpy.linalg
# ```
#
# has a standard set of matrix decompositions and things like inverse and
# determinant. These are implemented under the hood using the same industry-standard
# Fortran libraries used in other languages like MATLAB and R, such as like BLAS, LAPACK,
# or possibly (depending on your NumPy build) the Intel MKL:
# +
from numpy.linalg import inv, qr
X = np.arange(1,26).reshape(5,5)
print"x"
X
print "mat"
mat = X.T.dot(X)
mat
print "inv(mat)"
inv(mat)
print "mat.dot(inv(mat))"
mat.dot(inv(mat))
np.dot(mat, inv(mat))
# -
# ========================================================================================================================
# # <font color='green'>Random Number Generation</font>
# <img src="random.PNG">
#you can get a 4 by 4 array of samples from the standard normal distribution using normal:
samples = np.random.normal(size=(4, 4))
samples
# +
#comparision on Python built-in function random and numpy function
from random import normalvariate
N = 1000000
# %timeit samples = [normalvariate(0, 1) for _ in xrange(N)]
# %timeit np.random.normal(size=N)
# -
# Example: Random Walks<br>
# _"Python for data analysis"_ page 108
|
Python_for_data_analysis_NUMPY.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="20zzE1qqeDjU" colab_type="text"
# #**Module-02**
# 1. 1\. python Introduction
# 1. 1\. Two
# 1. 1\. Three
#
#
# + [markdown] id="z4FB0GuvjN90" colab_type="text"
# How to checks python version
# + id="CuYwLUT3bP0I" colab_type="code" outputId="19a92f5a-581d-4645-8c78-b7eb2d41bddf" executionInfo={"status": "ok", "timestamp": 1584298668917, "user_tz": 420, "elapsed": 3901, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !python -V
# + [markdown] id="vxK35D8IkL7C" colab_type="text"
# Check which python you atre using write now
# + id="HrUHT-Gqjn9C" colab_type="code" outputId="cfd47f61-689b-4c82-f1c4-b269053a903e" executionInfo={"status": "ok", "timestamp": 1584298821444, "user_tz": 420, "elapsed": 3635, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !which python
# + [markdown] id="AGGtkp_aklOq" colab_type="text"
# ##Write first python program
#
#
#
# 1. Open a new File
# 2. Write a program
# 3. save as a .py extension
# 4. Before running a program check current working directory of file is same as program file directory.
# 5. python3 filename.py
#
# + id="AfXJ8hZKj2Xy" colab_type="code" outputId="9f399041-7098-458a-d38a-9a29cd66b1de" executionInfo={"status": "ok", "timestamp": 1584299573720, "user_tz": 420, "elapsed": 3745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !pwd
# + id="Jiqq0YjzkjIQ" colab_type="code" outputId="2d820810-1492-46bc-84ec-8242a638a378" executionInfo={"status": "ok", "timestamp": 1584299595590, "user_tz": 420, "elapsed": 3832, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + id="qWoCkQNInUSy" colab_type="code" outputId="9fcb6fb1-aa4e-4fce-af13-57e7af0624a7" executionInfo={"status": "ok", "timestamp": 1584300989928, "user_tz": 420, "elapsed": 3628, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls
# + id="-jXLFoV3so9y" colab_type="code" outputId="884a8ddd-e434-4741-8074-47108ca4b9a1" executionInfo={"status": "ok", "timestamp": 1584301012904, "user_tz": 420, "elapsed": 3569, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# !python3 test.py
# + [markdown] id="OEg1u-Fpt7Nq" colab_type="text"
# ##Comments in python
#
#
# 1. Single line Comment
# 2. Multi line Comment
#
#
# > Single line Comment
#
#
# + id="_S9Qkw05sumA" colab_type="code" colab={}
#My first single line comment
# + [markdown] id="5nHKIiWyvks6" colab_type="text"
#
#
# > Multy line Comment
#
# + id="SH6RvwUvtiCx" colab_type="code" outputId="35b238bb-fa76-4b64-ae44-e61aa741704f" executionInfo={"status": "ok", "timestamp": 1584302354159, "user_tz": 420, "elapsed": 1360, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
""" multi lne comment
print(""hello")
print("how are you")
"""
# + id="IAijHYY-wto1" colab_type="code" outputId="94848a7f-ae13-4cd6-f6f2-8ad8a526e303" executionInfo={"status": "ok", "timestamp": 1584302357404, "user_tz": 420, "elapsed": 1318, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
''' multi lne comment
print(""hello")
print("how are you")
'''
# + id="bQoBTVs8w2a4" colab_type="code" colab={}
# !python3 test.py
# + [markdown] id="M0g-hCai1eFx" colab_type="text"
# #Multi line statement
# line continuation character ( \\ )
# + id="iYgc3Q_SxCXI" colab_type="code" outputId="cc628c60-2b4e-4ddc-bc46-0bdeebef4fad" executionInfo={"status": "ok", "timestamp": 1584304502902, "user_tz": 420, "elapsed": 3512, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
# !python3 multiline.py
# + [markdown] id="SXNdgqd16xag" colab_type="text"
# If statement contain in this brackets (below), then no need for line continuation character
#
# * ( )
# * { }
# * [ ]
#
#
# + id="BRdVnmOO3LFq" colab_type="code" outputId="79a182e5-a1f2-4bc7-b7ac-30ca3b3e0b2f" executionInfo={"status": "ok", "timestamp": 1584313146639, "user_tz": 420, "elapsed": 3868, "user": {"displayName": "Md Meraz", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
# !python3 multiline2.py
# + [markdown] id="M6IIlpwwbX7e" colab_type="text"
# ##MultiLine Print
# + id="eOnbgkkB8SrB" colab_type="code" outputId="a661f9a0-3c6c-4390-9115-27f68723bcfe" executionInfo={"status": "ok", "timestamp": 1584313458096, "user_tz": 420, "elapsed": 3586, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 306}
# !python3 multiline_print.py
# + [markdown] id="Wf0Z5ocKc-qm" colab_type="text"
# ##Python Quotation
#
#
# * Single quote '-------'
# * Double quote "-------"
# * Triple qupte """--------""
# ---
# >Single and double line comment are use for single line string
#
# >Triple quotes is only use in multiline string only
#
#
# + id="TymI3d-YcM9w" colab_type="code" outputId="39ec619e-25eb-422c-e69b-e99537d0ea99" executionInfo={"status": "ok", "timestamp": 1584315224872, "user_tz": 420, "elapsed": 1217, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiCDqeiThy6Uz7npH4ERYGnqSgnpBR6l78yLv1B=s64", "userId": "16501587712343191977"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
print ('India')
print(" Jai Hind")
print("""Bharat
mata ki
Jai """)
# + [markdown] id="fPEwi18hoK0t" colab_type="text" language="html"
# # <img src=">
# #
|
mod_02_python_intro/mod_02_01_about_python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3.8
# ---
# # Tutorial: Bring your own data (Part 3 of 3)
#
# ## Introduction
#
# In the previous [Tutorial: Train a model in the cloud](2.train-model.ipynb) article, the CIFAR10 data was downloaded using the builtin `torchvision.datasets.CIFAR10` method in the PyTorch API. However, in many cases you are going to want to use your own data in a remote training run. This article focuses on the workflow you can leverage such that you can work with your own data in Azure Machine Learning.
#
# By the end of this tutorial you would have a better understanding of:
#
# - How to upload your data to Azure
# - Best practices for working with cloud data in Azure Machine Learning
# - Working with command-line arguments
#
# ---
#
# ## Your machine learning code
#
# By now you have your training script running in Azure Machine Learning, and can monitor the model performance. Let's _parametrize_ the training script by introducing
# arguments. Using arguments will allow you to easily compare different hyperparmeters.
#
# Presently our training script is set to download the CIFAR10 dataset on each run. The python code in [train-with-cloud-data-and-logging.py](../../code/models/pytorch/cifar10-cnn/train-with-cloud-data-and-logging.py) now uses **`argparse` to parametize the script.**
# ### Understanding your machine learning code changes
#
# The script `train-with-cloud-data-and-logging.py` has leveraged the `argparse` library to set up the `--data-path`, `--learning-rate`, `--momentum`, and `--epochs` arguments:
#
# ```python
# import argparse
# ...
# parser = argparse.ArgumentParser()
# parser.add_argument("--data-path", type=str, help="Path to the training data")
# parser.add_argument("--learning-rate", type=float, default=0.001, help="Learning rate for SGD")
# parser.add_argument("--momentum", type=float, default=0.9, help="Momentum for SGD")
# parser.add_argument("--epochs", type=int, default=2, help="Number of epochs to train")
# args = parser.parse_args()
# ```
#
# The script was adapted to update the optimizer to use the user-defined parameters:
#
# ```python
# optimizer = optim.SGD(
# net.parameters(),
# lr=args.learning_rate, # get learning rate from command-line argument
# momentum=args.momentum, # get momentum from command-line argument
# )
# ```
#
# Similarly the training loop was adapted to update the number of epochs to train to use the user-defined parameters:
# ```python
# for epoch in range(args.epochs):
# ```
#
# ## Upload your data to Azure
#
# In order to run this script in Azure Machine Learning, you need to make your training data available in Azure. Your Azure Machine Learning workspace comes equipped with a _default_ **Datastore** - an Azure Blob storage account - that you can use to store your training data.
#
# > <span style="color:purple; font-weight:bold">! NOTE <br>
# > Azure Machine Learning allows you to connect other cloud-based datastores that store your data. For more details, see [datastores documentation](./concept-data.md).</span>
#
# !pip install --upgrade torchvision
# +
from azureml.core import Workspace, Dataset
from torchvision import datasets
ws = Workspace.from_config()
datasets.CIFAR10(".", download=True)
ds = ws.get_default_datastore()
ds.upload(
src_dir="cifar-10-batches-py",
target_path="datasets/cifar10",
overwrite=False,
)
# +
import os
import shutil
os.remove("cifar-10-python.tar.gz")
shutil.rmtree("cifar-10-batches-py")
# -
# The `target_path` specifies the path on the datastore where the CIFAR10 data will be uploaded.
#
# ## Submit your machine learning code to Azure Machine Learning
#
# As you have done previously, create a new Python control script:
# +
import git
from pathlib import Path
prefix = Path(git.Repo(".", search_parent_directories=True).working_tree_dir)
prefix
# + tags=["remote run", "batchai", "configure run", "use notebook widget", "get metrics", "use datastore"]
from azureml.core import (
Workspace,
Experiment,
Environment,
ScriptRunConfig,
Dataset,
)
from azureml.widgets import RunDetails
ws = Workspace.from_config()
ds = Dataset.File.from_files(
path=(ws.get_default_datastore(), "datasets/cifar10")
)
env = Environment.from_conda_specification(
name="pytorch-env-tutorial",
file_path=prefix.joinpath("environments", "pytorch-example.yml"),
)
exp = Experiment(
workspace=ws, name="getting-started-train-model-cloud-data-tutorial"
)
src = ScriptRunConfig(
source_directory=prefix.joinpath(
"code", "models", "pytorch", "cifar10-cnn"
),
script="train-with-cloud-data-and-logging.py",
compute_target="cpu-cluster",
environment=env,
arguments=[
"--data-path",
ds.as_mount(),
"--learning-rate",
0.003,
"--momentum",
0.92,
"--epochs",
2,
],
)
run = exp.submit(src)
RunDetails(run).show()
# -
# ### Understand the control code
#
# The above control code has the following additional code compared to the control code written in [previous tutorial](03-train-model.ipynb)
#
# **`ds = Dataset.File.from_files(path=(datastore, 'datasets/cifar10'))`**: A Dataset is used to reference the data you uploaded to the Azure Blob Store. Datasets are an abstraction layer on top of your data that are designed to improve reliability and trustworthiness.
#
#
# **`src = ScriptRunConfig(...)`**: We modified the `ScriptRunConfig` to include a list of arguments that will be passed into training script. We also specified `ds.as_mount()`, which means the directory specified will be _mounted_ to the compute target.
#
# ## Inspect the 70_driver_log log file
#
# In the navigate to the 70_driver_log.txt file - you should see the following output:
#
# ```
# Processing 'input'.
# Processing dataset FileDataset
# {
# "source": [
# "('workspaceblobstore', 'datasets/cifar10')"
# ],
# "definition": [
# "GetDatastoreFiles"
# ],
# "registration": {
# "id": "XXXXX",
# "name": null,
# "version": null,
# "workspace": "Workspace.create(name='XXXX', subscription_id='XXXX', resource_group='X')"
# }
# }
# Mounting input to /tmp/tmp9kituvp3.
# Mounted input to /tmp/tmp9kituvp3 as folder.
# Exit __enter__ of DatasetContextManager
# Entering Run History Context Manager.
# Current directory: /mnt/batch/tasks/shared/LS_root/jobs/dsvm-aml/azureml/tutorial-session-3_1600171983_763c5381/mounts/workspaceblobstore/azureml/tutorial-session-3_1600171983_763c5381
# Preparing to call script [ train.py ] with arguments: ['--data_path', '$input', '--learning_rate', '0.003', '--momentum', '0.92']
# After variable expansion, calling script [ train.py ] with arguments: ['--data_path', '/tmp/tmp9kituvp3', '--learning_rate', '0.003', '--momentum', '0.92']
#
# Script type = None
# ===== DATA =====
# DATA PATH: /tmp/tmp9kituvp3
# LIST FILES IN DATA PATH...
# ['cifar-10-batches-py', 'cifar-10-python.tar.gz']
# ```
#
# Notice:
#
# 1. Azure Machine Learning has mounted the blob store to the compute cluster automatically for you
# 2. The ``ds.as_mount()`` used in the control script resolves to the mount point
# 3. In the machine learning code we include a line to list the directorys under the data directory - you can see the list above
# If you're not going to use what you've created here, delete the resources you just created with this quickstart so you don't incur any charges for storage. In the Azure portal, select and delete your resource group.
|
tutorials/getting-started-train/3.pytorch-model-cloud-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # All imports
# +
import numpy as np
import functools
import itertools
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# -
# # Mathsy Aux Functions
# +
# Returns the probability of n theta bernoulli variables getting r positives multiplied by
def bernoulli_prob(n,r,theta):
theta *= 1.
return (theta**r) * (1-theta)**(n-r)
# Scales a vector to have L1 length equal to 1
def normalize_vector(vect):
return np.divide(vect,sum(vect))
# Given a pair of values and probabilities, it returns E[f(x)]
def expectation(values,probabilities,function):
values = list(map(function,values))
return np.average(values, weights = probabilities)
# Given distribution A and distribution B on np.linspace(0,1,len(distribution))
# It returns the expected value
def expectations(distribution_A,distribution_B,function):
a = zip(distribution_A,np.linspace(0,1,len(distribution_A)))
b = zip(distribution_B,np.linspace(0,1,len(distribution_B)))
c = itertools.product(a,b)
def f(x):
return (x[0][0]*x[1][0],function(x[0][1],x[1][1]))
d = map(f,c)
return list(d)
# -
# # pdfs Aux Functions
# +
# Returns a pdf from a number of observations and number of transactions, assuming a initial uniform pdf
# TODO: If n is too big the probabilities get set to zero due to floating point errors
# TODO: Need to start using BigInt types or find another way around this
def generate_pdf(n,r,bins=10**3):
# n : number of observations
# r : number of positives
# bins : number of points in the x coordinate to be generated
x_coordinate = np.linspace(0,1,bins+1)
f = functools.partial(bernoulli_prob,n,r)
y_coordinate = list(map(f,x_coordinate))
# TODO: Find a better way to normalize a vector
y_coordinate = np.multiply(y_coordinate,(bins*1./sum(y_coordinate)))
return y_coordinate
# Updates the confidence interval based on the original pdf
def bayesian_update(original_pdf,update_factor, bins):
assert len(original_pdf) == bins
assert len(update_factor) == bins
updated_pdf = np.multiply(original_pdf,update_factor)
# TODO: Find a better way to normalize a vector
updated_pdf = np.multiply(updated_pdf,bins*1./sum(updated_pdf))
return updated_pdf
# -
# # Plot Aux Functions
# +
def plot_function(distribution,fig_width=20,fig_height=3):
distribution.sort(key=lambda t: t[1])
plt.rcParams["figure.figsize"] = [fig_width,fig_height]
x,y = list(zip(*distribution))
plt.plot(x,y)
plt.ylabel('Expected value')
plt.show()
# Compares the plots given arrays of equal length
def plot_pdfs(distributions,bins=1000,fig_width=20,fig_height=3):
plt.rcParams["figure.figsize"] = [fig_width,fig_height]
patches = []
for distribution,color,name in distributions:
y_coordinate = distribution
x_coordinate = np.linspace(0,1,len(y_coordinate))
plt.plot(x_coordinate,y_coordinate,color)
patches.append(mpatches.Patch(color=color, label=name))
plt.legend(handles=patches)
plt.ylabel('probabiity density function')
plt.show()
# -
# # Main
obs = [(10,2,'g','alternative_1'),(100,3,'b','alternative_2'),(100,4,'r','control')]
def f(a):
return (generate_pdf(a[0],a[1]),a[2],a[3])
obs = list(map(f,obs))
plot_pdfs(obs)
def f(a,b):
return a-b
obs[0][0]
obs[1][0]
vect = expectations(obs[0][0],obs[1][0],f)
vect[:10]
plot_function(vect)
vect
x,y = list(zip(*vect))
vect.sort(key=lambda t: t[1])
expectations([.1,.2,.7],[.2,.4,.4],lambda a,b:a-b)
|
WIP/Multi Armed Bandit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import netCDF4
import numpy as np
import datetime
# +
f = netCDF4.Dataset()
# +
# -
# %matplotlib inline
# +
# plot with the rainrate
import matplotlib.pyplot as plt
import matplotlib
# +
# Some adjustments to the axis labels, ticks and fonts
def load_xy_style(ax, xlabel='Time [UTC]', ylabel='Height [m]'):
"""
Method that alters the apperance of labels on the x and y axis in place.
Note:
If xlabel == 'Time [UTC]', the x axis ticks have major
ticks every 3 hours and minor ticks every 30 minutes.
Args:
ax (matplotlib.axis) :: axis that gets adjusted
**xlabel (string) :: name of the x axis label
**ylabel (string) :: name of the y axis label
"""
ax.set_xlabel(xlabel, fontweight='semibold', fontsize=15)
ax.set_ylabel(ylabel, fontweight='semibold', fontsize=15)
if xlabel == 'Time [UTC]':
ax.xaxis.set_major_formatter(matplotlib.dates.DateFormatter('%H:%M')
ax.xaxis.set_major_locator(matplotlib.dates.HourLocator(interval=3))
ax.xaxis.set_minor_locator(matplotlib.dates.MinuteLocator(byminute=[0,30]))
ax.yaxis.set_minor_locator(matplotlib.ticker.MultipleLocator(500))
ax.tick_params(axis='both', which='major', labelsize=14, width=3, length=4)
ax.tick_params(axis='both', which='minor', width=2, length=3)
def load_cbar_style(cbar, cbar_label=''):
"""
Method that alters the apperance of labels on the color bar axis in place.
Args:
ax (matplotlib.axis) :: axis that gets adjusted
**cbar_label (string) :: name of the cbar axis label, Defaults to empty string.
"""
cbar.ax.set_ylabel(cbar_label, fontweight='semibold', fontsize=15)
cbar.ax.tick_params(axis='both', which='major', labelsize=14, width=2, length=4)
# +
# reflectivity plot
# +
# velocity plot with custom colorplot
import Colormaps
# similar for velocity
# +
# 2d histogram
# +
# lidar beta and linear depolarization in one figure with two subplots
|
notebooks/part4_netcdf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Space Invaders - missiles
# [Vidéo d'accompagnement](https://vimeo.com/502720408)
# Maintenant que nous savons gérer le mouvement d'un simple rectangle avec le clavier ou automatiquement, appliquons nos connaissances pour modéliser les projectiles du jeu.
# ## Objectif 2: Gérer les missiles
# Un missile est basiquement un rectangle...
#
# Il est donc temps de faire le point sur notre module `rectangle.py` pour **réutiliser** au maximum ce que nous avons déjà fait et/ou l'**adapter** à notre nouvelle problématique.
#
# **module** *rectangle.py*:
#
# 1. objet rectangle (dictionnaire) a pour attributs (clés): `id`, `largeur`, `hauteur`, `position`, `vitesse`
#
# 2. fonction «constructeur» car sert à produire un objet rectangle:
#
# `initialiser_rect`:renvoie un objet rectangle
#
# 3. fonctions ayant un objet rectangle comme premier paramètre:
#
# - `gauche` et `droite`: déplace le rectangle vers la gauche ou la droite de 5 pixels
# - `reagir`: associe un gestionnaire d'événement au rectangle
# - `set_vitesse`: pour régler l'attribut vitesse du rectangle
# - `lancer`: pour déclencher son mouvement automatique dans la scène (gauche-droite)
# Après avoir soigneusement étudié ce plan, quel sera celui du **module** *missile.py*?
# 1. objet missile (dictionnaire) a pour attributs (clés): _____
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# `id`, `largeur`, `hauteur`, `position`, `vitesse`; rien à ajouter ou retrancher
# -
# 2. fonction «constructeur» pour un objet missile: ____
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# `initialiser_missile`: renvoie un objet missile ayant la même structure qu'un objet rectangle; penser à modifier les valeurs par défaut de largeur et hauteur.
# -
# 3. fonctions ayant un objet missile comme premier paramètre:
# - ...
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# on **élimine** `gauche`, `droite` et `reagir`
#
# on **conserve**: `set_vitesse` (identique), `lancer` (code à adapter)
#
# on **ajoute**: `supprimer`: qui sert à éliminer le missile lorsqu'il sort de la scene.
# -
# ## «Schéma» du module `missile.py`
# Suivant notre analyse, notre module doit ressembler à ce qui suit; `...` indique des zones où le code est le même que pour le module `rectangle.py`:
# ```python
# # fichier space_invaders/missile.py
# pass
#
# def initialiser_missile(x, y, largeur=5, hauteur=15, couleur="white"):
# pass
#
# def set_vitesse(fig, v):
# pass
#
# def lancer(fig):
# pass
#
# def supprimer(fig):
# pass
#
# if __name__ == "__main__":
# # code pour tester le module
# pass
# ```
# ## Tout à un début et ... une fin!
# `initialiser...` sert clairement à donner naissance à notre «missile»: **allocation** de ressource graphique et d'un dictionnaire.
#
# `supprimer` devra servir à le faire «mourir» c'est-à-dire à **désallouer** les ressources graphiques (chercher dans la documentation du Canvas).
# ```python
# def supprimer(fig):
# # désallouer ressouce graphique
# scene.______
# ```
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# ```python
# def supprimer(fig):
# # désallouer ressouce graphique
# scene.delete(fig["id"])
# ```
# -
# ## `lancer`
# Souvenez-vous que cette fonction est, normalement, rappelée automatiquement par `fen.after(...)` sauf si le missile sort de la scene.
# Dans *space invaders* les missiles se déplacent verticalement (soit du haut vers le bas soit le contraire selon le lanceur)
# Enfin, un missile qui sort de l'écran est un missile ... mort...
#
# **Note**: nous gérerons les collisions plus tard et donc n'en tenons pas compte ici.
# ```python
# def lancer(fig):
# ...
#
# # déplacement pour 50ms
# ____ = v / 20
#
# ...
# largeur, hauteur = ___
#
# # si le missile est sorti de la scene (entièrement) le détruire
# # conseil: faire un dessin!
# if ____ or ____ or ____ or ____:
# ____
#
# # à présent, on peut agir ...
# ___
#
# ...
# ```
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# ```python
# def lancer(fig):
# v = fig["vitesse"]
# # si la vitesse est nulle, inutile de continuer
# if v == 0: return
#
# # déplacement pour 100ms
# dy = v / 20
#
# x, y = fig["position"]
# largeur, hauteur = fig["largeur"], fig["hauteur"]
#
# # si le missile est sorti de la scene (entièrement) le détruire
# if x > LARGEUR or x + largeur < 0 or y > HAUTEUR or y + hauteur < 0:
# supprimer(fig)
# # attention à utiliser return pour mettre fin à la fonction! sinon...
# return
#
# # à présent, on peut agir ...
# scene.move(fig["id"], 0, dy)
# # ne pas oublier de mettre à jour
# fig["position"] = x, y + dy
#
# # et on recommence
# fen.after(50, lambda: lancer(fig))
# ```
# -
# ## `set_vitesse`
# ```python
# def set_vitesse(fig, v):
# fig["vitesse"] = v
# ```
# ## Tester que tout fonctionne bien
# Commençons par un **test basique** pour corriger au maximum les erreurs éventuelles
# ```python
# if __name__ == "__main__":
# m = ___(LARGEUR // 2, HAUTEUR - 30)
# ____(m, -50)
# ____
# fen.mainloop()
# ```
# **Solution** - n'ouvrir qu'après avoir suffisamment cherché!
# + [markdown] jupyter={"source_hidden": true}
# ```python
# if __name__ == "__main__":
# m = initialiser_missile(LARGEUR // 2, HAUTEUR - 30)
# set_vitesse(m, -50)
# lancer(m)
# fen.mainloop()
# ```
# -
# Puis mettons nos missiles à l'épreuve des balles: réalisons un «**stress test**»
# **VERSION NSI si la syntaxe des listes en compréhensions a été abordée (sinon voir ISN)**
# ```python
# if __name__ == "__main__":
# from random import randint
# missiles = [ ____(randint(0, LARGEUR), HAUTEUR - 30) for _ in range(30) ]
# for m in ____:
# set_vitesse( ____, -randint(50, 300) )
# ____.bind( '<space>', lambda evt: lancer( missiles.pop() ) )
# fen.mainloop()
# ```
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# ```python
# if __name__ == "__main__":
# from random import randint
# missiles = [ initialiser_missile(randint(0, LARGEUR), HAUTEUR - 30) for _ in range(30) ]
# for m in missiles:
# # pourquoi pas une liste en compréhension ...???
# set_vitesse( m, -randint(50, 300) )
# fen.bind( '<space>', lambda evt: lancer( missiles.pop() ) )
# fen.mainloop()
# ```
# -
# **VERSION ISN**
# ```python
# if __name__ == "__main__":
# from random import randint
# missiles = []
# for _ in range(30):
# m = ____(randint(0, LARGEUR), HAUTEUR - 30)
# ____.append(____)
# for m in ____:
# set_vitesse( ____, -randint(50, 300) )
# ____.bind( '<space>', lambda evt: lancer( missiles.pop() ) )
# fen.mainloop()
# ```
# **Solution** - n'ouvrir qu'après avoir suffisemment cherché!
# + [markdown] jupyter={"source_hidden": true}
# ```python
# if __name__ == "__main__":
# from random import randint
# missiles = []
# for _ in range(30):
# m = initialiser_missile(randint(0, LARGEUR), HAUTEUR - 30)
# missiles.append(m)
# for m in missiles:
# set_vitesse( m, -randint(50, 300) )
# fen.bind( '<space>', lambda evt: lancer( missiles.pop() ) )
# fen.mainloop()
# ```
# -
# ## Solution complète
# #### Fichier `missile.py`
# + [markdown] editable=false jupyter={"source_hidden": true}
# ```python
# # module space_invaders/missile.py
# from scene import fen, scene, LARGEUR, HAUTEUR
#
# def initialiser_missile(x, y, largeur=5, hauteur=15, couleur="white"):
# _id = scene.create_rectangle(x, y, x+largeur, y+hauteur, fill=couleur)
# return {
# "id": _id,
# "largeur": largeur,
# "hauteur": hauteur,
# "position": (x, y),
# "vitesse": 0 # en pixels par seconde
# }
#
# def set_vitesse(fig, v):
# fig["vitesse"] = v
#
# def lancer(fig):
# v = fig["vitesse"]
# # si la vitesse est nulle, inutile de continuer
# if v == 0: return
#
# # déplacement pour 100ms
# dy = v / 20
#
# x, y = fig["position"]
# largeur, hauteur = fig["largeur"], fig["hauteur"]
#
# # si le missile est sorti de la scene (entièrement) le détruire
# if x > LARGEUR or x + largeur < 0 or y > HAUTEUR or y + hauteur < 0:
# supprimer(fig)
# # attention à utiliser return pour mettre fin à la fonction! sinon...
# return
#
# # à présent, on peut agir ...
# scene.move(fig["id"], 0, dy)
# # ne pas oublier de mettre à jour
# fig["position"] = x, y + dy
#
# # et on recommence
# fen.after(50, lambda: lancer(fig))
#
# def supprimer(fig):
# # désallouer ressouce graphique
# scene.delete(fig["id"])
# # désallouer le dictionnaire
# del fig
#
# if __name__ == "__main__":
# from random import randint
# missiles = []
# for _ in range(30):
# m = initialiser_missile(randint(0, LARGEUR), HAUTEUR - 30)
# missiles.append(m)
# for m in missiles:
# set_vitesse( m, -randint(50, 300) )
# fen.bind( '<space>', lambda evt: lancer( missiles.pop() ) )
# fen.mainloop()
# ```
|
02_interface_graphique/space_invaders/01_missiles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 Neuron
#
# * Input (x1, ... xn)
# * Weights (w1, ... wn)
# * Single Neuron
# * Activation Function (relu, tanh, sigmoid etc.)
# * Output
#
# 
#
#
# # 2 Neuronales Netz
#
# * Input Layer
# * Hidden Layer
# * Output Layer
#
# 
#
# # 3 More about Deep Learning...
#
# * Forward Propagation
# * Backpropagation
# * Batches
# * Epoches
#
# # 4 Softmax
#
# 
# x = [1,5,3]
# 
#
# # 5 One Hot
#
# # 6 Regularization
|
3_neural_network/1_intro_neural_network.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 0.0. Business Understanding
# #### Stack Overflow Survey
# ##### Data:
# - What: dataset with the results of a survey of developers in 2021
# - Granularity: per respondent
#
# ##### Goal:
# - Provide information about the Data Science Developers subset;
# - Provide insights: 3 insights about the Data Scientists;
# - Provide a Machine Learning Model to predict the salary of Data Scientists.
# # 1.0. Imports
# ## 1.1. Frameworks
# +
import pandas as pd
import numpy as np
import math
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error
# -
# ## 1.2. Config
# %matplotlib inline
plt.rcParams['figure.figsize'] = [25, 12]
plt.rcParams['font.size'] = 24
sns.set()
# ## 1.3. Data
df = pd.read_csv(r'./data/2021_survey_results.csv', sep=',') # import data
# # 2.0. Data Understanding
df2 = df.copy()
# ## 2.1. Columns / Rows
print(f'Rows: {df2.shape[0]}')
print(f'Columns: {df2.shape[1]}')
# ## 2.2. Dtypes
df2.dtypes
# ## 2.3. Null Values
df2.isnull().sum()
# # 3.0. Filter Data
df3 = df2.copy()
# ## 3.1. Select Rows
# +
# drop na values
df3 = df3.dropna(axis=0, subset=['DevType'])
# select only data scientists
df3 = df3.loc[df3['DevType'] == 'Data scientist or machine learning specialist', :]
# -
# ## 3.2. Select Columns
df3 = df3.loc[:, ['ResponseId', 'Employment', 'YearsCode',
'LanguageHaveWorkedWith', 'DatabaseHaveWorkedWith',
'PlatformHaveWorkedWith', 'ConvertedCompYearly']]
# # 4.0. Exploratory Data Analysis
df4 = df3.copy()
def count_list_series(df, column, sep=';', only_values=False):
"""
Input:
df - pandas dataframe containing the column to be counted;
column - columns with the list entries that must have the values counted;
sep - the type of separator used in the list.
describe - if True, return count and percentage.
Output:
df - a new dataframe containing:
Value: the name of the variable to be counted;
Count: count of the variable;
Percentage: the percentage of the value in relation to the number of lines.
"""
final_list = []
df_aux = df[column].dropna(axis=0) # droping na values to analysis. selecting only entries with a list
for index, value in enumerate(df_aux):
list_languages = value.split(sep)
for language in list_languages:
final_list.append(language)
if only_values == False:
value, counts = np.unique(final_list, return_counts=True)
df_final = pd.DataFrame(counts, value).sort_values(by=0, ascending=False).reset_index().rename({'index': 'Value', 0: 'Count'}, axis=1)
df_final['Percentage'] = round((100 * df_final['Count'] / len(df)), 2)
return df_final
else:
return set(final_list)
# ## Q1 - What are the most used languages and tools by Data Scientists or machine learning especialists
# ### Top 3 Languages:
# - **Python**: 92.72%
# - **SQL**: 49.62%
# - **R**: 31.11%
#
# ### Top 3 Databases:
# - **PostgreSQL**: 33.69%
# - **MySQL**: 33.38%
# - **SQLite**: 21.40%
#
# ### Top 3 Platforms:
# - **AWS**: 44.01%
# - **Google Cloud**: 30.80%
# - **Microsoft Azure**: 22.61%
# +
df_language = count_list_series(df=df4, column='LanguageHaveWorkedWith')
sns.barplot(x='Percentage', y='Value', data=df_language[:10], palette=sns.color_palette('Blues', 1))
# Graph configuration
plt.title('Most Used Languages', fontsize=21)
plt.xlabel('Percentage Users', fontsize=15)
plt.ylabel('Language', fontsize=15)
plt.savefig('./images/languages')
# -
df_language.head(10)
# +
df_database = count_list_series(df=df4, column='DatabaseHaveWorkedWith')
sns.barplot(x='Percentage', y='Value', data=df_database[:10], palette=sns.color_palette('Blues', 1))
# Graph configuration
plt.title('Most Used Databases', fontsize=21)
plt.xlabel('Percentage Users', fontsize=15)
plt.ylabel('Database', fontsize=15)
plt.savefig('./images/databases')
# -
df_database.head(10)
# +
df_platform = count_list_series(df=df4, column='PlatformHaveWorkedWith')
sns.barplot(x='Percentage', y='Value', data=df_platform[:10], palette=sns.color_palette('Blues', 1))
# Graph configuration
plt.title('Most Used Platforms', fontsize=21)
plt.xlabel('Percentage Users', fontsize=15)
plt.ylabel('Platform', fontsize=15)
plt.savefig('./images/platforms')
# -
df_platform.head(10)
# ## Q2 - How long have data scientists been programming
# ### Experience:
# - **0 -> 9**: 51.84%
# - **10 -> 18.0**: 35.23
# - **19 -> 27.0**: 7.84%
# - **28 -> 36.0**: 3.00%
# - **37 -> 45.0**: 2.00%
df4.loc[df4['YearsCode'] == 'Less than 1 year', 'YearsCode'] = 0
df_years_code = df4['YearsCode'].dropna().astype('int64')
# +
sns.distplot(df_years_code)
# Graph configuration
plt.title('Distribution of Years Code', fontsize=21)
plt.xlabel('Years Code', fontsize=15)
plt.ylabel('Density', fontsize=15)
plt.savefig('./images/years_code')
# -
sns.boxplot(x = df_years_code)
df_group_years = pd.cut(x=df_years_code, bins=5).reset_index().groupby('YearsCode').count()
df_group_years['Percentage'] = df_group_years['index'] / np.sum(df_group_years['index'])
df_group_years.columns = ['Count', 'Percentage']
df_group_years
# ## Q3 - What's the average salary for Data Scientists or machine learning especialists
# ### Salary:
# - **Employed part-time**: 35586.45
# - **Employed full-time**: 112979.35
# - **Independent contractor, freelancer, or self-employed**: 619168.66
df_salary = df4[['Employment', 'ConvertedCompYearly']].groupby('Employment').mean().reset_index().sort_values(by='ConvertedCompYearly')
df_salary.dropna(inplace=True)
# +
sns.barplot(x='Employment', y='ConvertedCompYearly', data=df_salary,
palette=sns.color_palette('Blues', 1))
# Graph configuration
plt.title('Salary x Employment Type', fontsize=21)
plt.xlabel('Employment Type', fontsize=15)
plt.ylabel('$ Salary', fontsize=15)
plt.savefig('./images/salary')
# -
df_salary
# # 5.0. Prepare Data
df5 = df4.copy()
# ## 5.1. NaN Values
df5.isnull().mean()
# +
# ConvertedCompYearly -> drop
df5.dropna(subset=['ConvertedCompYearly'], axis=0, inplace=True)
# YearsCode -> mean.
df5['YearsCode'] = df5['YearsCode'].fillna(df5['YearsCode'].astype('float64').mean()).astype('int64')
# LanguageHaveWorkedWith -> no_response
df5['LanguageHaveWorkedWith'] = df5['LanguageHaveWorkedWith'].fillna('no_response')
# DatabaseHaveWorkedWith -> no_response
df5['DatabaseHaveWorkedWith'] = df5['LanguageHaveWorkedWith'].fillna('no_response')
# PlatformHaveWorkedWith -> no_response
df5['PlatformHaveWorkedWith'] = df5['LanguageHaveWorkedWith'].fillna('no_response')
# -
# ## 5.2. Encode
df5.isnull().sum()
# +
# LanguageHaveWorkedWith, DatabaseHaveWorkedWith, PlatformHaveWorkedWith
# One hot Encoding
list_columns = ['LanguageHaveWorkedWith', 'DatabaseHaveWorkedWith', 'PlatformHaveWorkedWith']
for column in list_columns:
set_tool = count_list_series(df5, column, only_values=True)
for tool in set_tool:
df5[tool] = df5[column].apply(lambda x: 1 if tool in x else 0)
# delete columns
df5 = df5.drop(axis=1, columns=list_columns)
# +
# create dummies
dummies = pd.get_dummies(df5['Employment'])
# merging dummies and exclude the initial column
df5 = pd.merge(left=df5, right=dummies, left_on = df5.index, right_on=dummies.index).drop('Employment', axis=1)
# -
# ## 5.3. Filter
# +
# drop key_0 em RespondeId columns
df5 = df5.drop(['key_0', 'ResponseId'], axis=1)
# +
# drop outliers
df5 = df5.loc[df5['ConvertedCompYearly'] <= 1000000, :]
# -
# ## 5.4. Separate Variables
X = df5.drop('ConvertedCompYearly', axis=1)
y = df5.loc[:, 'ConvertedCompYearly']
# ## 5.5. Train and Test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# # 6.0. Data Modelling
from sklearn.ensemble import RandomForestRegressor
# ## 6.1. Baseline - mean
# create a baseline
y_hat_baseline = [y_train.mean() for x in range(0, len(y_test))]
# Y_real x baseline
plt.plot(y_hat_baseline)
plt.plot(y_real)
# ## 6.2. RandomForestRegressor
# instantiate linear regression
ln_reg = RandomForestRegressor()
# train linear regression
ln_reg.fit(X_train, y_train)
# +
# y predict
y_hat = ln_reg.predict(X_test)
# y real
y_real = y_test.values
# -
# y_real x model
plt.plot(y_hat)
plt.plot(y_real)
# # 7.0. Evaluate the Results
# Metric: MAE - Mean Absolute Error - The average absolute error.
#
# The model is no better than a baseline. We must create features, increase the amount of data and test new models to improve performance.
# Mean Absolute Percentage Erro - Baseline
mean_absolute_error(y_real, y_hat_baseline)
# Mean Absolute Percentage Error - Model
mean_absolute_error(y_real, y_hat)
|
notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] qat-category="1-Circuit generation in Python" qat-summary="1-Creation of an EPR pair using two qubits"
# # Creation of an EPR pair using two qubits
#
#
#
# Let us start with a simple circuit: the creation of an EPR pair using two qubits.
#
#
# First, we need to import relevant objects from the python AQASM module:
# -
from qat.lang.AQASM import Program, H, CNOT
# ## Creation of the quantum program
#
# Then, we can declare a new object ``Program``. Let us give it an explicit name:
epr_prog = Program()
# To create our EPR pair, we need to manipulate two qbits.
# Qbits are manipulated through qbit registers only (to keep things structured).
# Registers are allocated as follows:
qbits = epr_prog.qalloc(2)
# Now, we can access our qbits using the register "qbits".
#
# Registers behave like python list/arrays.
#
# Here our qbits will be refered to using qbits[0] and qbits[1].
#
# To create our EPR pair, we simply implement the appropriate 2-qbit rotation using a Hadamard gate (H) on the first qbit, followed by a controlled NOT gate (CNOT) on both qbits:
epr_prog.apply(H, qbits[0])
epr_prog.apply(CNOT, qbits)
# Notice that since the CNOT is applied on both qbits (it is a 2-qbit gate), we can pass the whole register as argument to the ``.apply`` method.
# The corresponding circuit object can be extracted directly from the Program object as follows:
circuit = epr_prog.to_circ()
print(circuit.ops)
# %qatdisplay circuit
# ## Simulation of the execution of the circuit
# Now that we have a proper circuit, we can try and simulate it:
# +
#Let us import some qpu connected to a classical linear algebra simulator
from qat.qpus import LinAlg
qpu = LinAlg()
job = circuit.to_job()
result = qpu.submit(job)
for sample in result:
print("State", sample.state, "with amplitude", sample.amplitude)
# -
# ## Export to Atos Quantum Assembly Language (AQASM) format
# We can also export our circuit in the AQASM format:
epr_prog.export("helloworld.aqasm")
# The generated file *helloworld.aqasm* should look like this:
# !cat helloworld.aqasm
# and can be compiled to circ format as follows:
# !aqasm2circ helloworld.aqasm
|
misc/notebooks/tutorials/basic/epr_pair.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# ###############################
# 1D Tensor Networks & Algorithms
# ###############################
#
# Although ``quimb.tensor`` aims to be an interactive and general
# base for arbitrary tensor networks, it also has fast
# implementations of the following:
#
# Static:
#
# * 1-site :class:`~quimb.tensor.tensor_dmrg.DMRG1` (OBC and PBC)
# * 2-site :class:`~quimb.tensor.tensor_dmrg.DMRG2` (OBC and PBC)
# * 1-site :class:`~quimb.tensor.tensor_dmrg.DMRGX`
#
# Time Evolving:
#
# * :class:`~quimb.tensor.tensor_1d_tebd.TEBD`
#
# Two site DMRGX and TDVP slot into the same framework and should be
# easy to implement. All of these are based on 1D tensor networks, the
# primary representation of which is the *matrix product state*.
#
#
# Matrix Product States
# ---------------------
#
# The basic constructor for MPS is :class:`~quimb.tensor.tensor_1d.MatrixProductState`.
# This is a subclass of :class:`~quimb.tensor.tensor_core.TensorNetwork`,
# with a special tagging scheme (``MPS.site_tag_id``) and special index
# naming sceme (``MPS.site_ind_id``).
# It is also possible to instantiate a MPS directly from a dense vector using
# :meth:`~quimb.tensor.tensor_1d.MatrixProductState.from_dense`, though
# this is obviously not efficient for many sites.
#
# In the following, we just generate a random MPS, and demonstrate some basic functionality.
# -
# %config InlineBackend.figure_formats = ['svg']
from quimb.tensor import *
p = MPS_rand_state(L=20, bond_dim=50)
print(f"Site tags: '{p.site_tag_id}', site inds: '{p.site_ind_id}'")
print(p) # shows the full list of constituent tensors
p.show() # 1D tensor networks also have a ascii ``show`` method
# We can then __canonicalize__ the MPS:
p.left_canonize()
p.show()
# And we can compute the inner product as:
p.H @ p
# This relies on them sharing the same physical indices, ``site_ind_id``,
# which the conjugated copy ``p.H`` naturally does.
#
# Like any TN, we can __graph__ the overlap for example, and make use of the
# site tags to color it:
(p.H & p).draw(color=[f'I{i}' for i in range(20)])
# + raw_mimetype="text/restructuredtext" active=""
# I.e. we used the fact that 1D tensor networks are tagged with the structure ``"I{}"``
# denoting their sites. See the :ref:`examples` for how to fix the positions of tensors when drawing them.
#
# We can also **add** MPS, and **multiply/divide** them by scalars:
# -
p2 = (p + p) / 2
p2.show()
# Which doubles the bond dimension, as expected, but should still be normalized:
p2.H @ p2
# Because the MPS is the addition of two identical states, it should also __compress__ right back down:
p2.compress(form=10)
p2.show()
# Where we have also set the __orthogonality center__ at the site 10.
#
# When tensor networks are imbued with a ``structure``, they
# can be indexed with integers and slices, which automatically get
# converted using ``TN.site_tag_id``:
p2[10] # get the tensor(s) with tag 'I10'.
# Note the tensor has matching physical index ``'k10'``.
#
# This tensor is the orthogonality center so:
#
# ─>─>─●─<─<─ ╭─●─╮
# ... │ │ │ │ │ ... = │ │ │
# ─>─>─●─<─<─ ╰─●─╯
# i=10 i=10
#
# should compute the normalization of the *whole* state:
p2[10].H @ p2[10] # all indices match -> inner product
# Or equivalently:
p2[10].norm()
# If two tensor networks with the same ``structure`` are combined, it is propagated.
# For example ``(p2.H & p2)`` can still be sliced.
#
# Since the MPS is in canonical form, left and right pieces of the overlap
# should form the identity. The following forms a TN of the inner product,
# selects the *2* tensors corresponding to the last site (``-1``), contracts them,
# then gets the underlying data:
((p2.H & p2).select(-1) ^ all).data.round(12) # should be close to the identity
# + raw_mimetype="text/restructuredtext" active=""
# Various builtin quantities are available to compute too:
#
# - :meth:`~quimb.tensor.tensor_1d.MatrixProductState.entropy`
# - :meth:`~quimb.tensor.tensor_1d.MatrixProductState.schmidt_gap`
# - :meth:`~quimb.tensor.tensor_1d.MatrixProductState.magnetization`
# - :meth:`~quimb.tensor.tensor_1d.TensorNetwork1DVector.correlation`
# - :meth:`~quimb.tensor.tensor_1d.MatrixProductState.logneg_subsys`
#
# and other non-trivial quantities such as the mutual information
# can be easily calculated using a combination of -
# :meth:`~quimb.tensor.tensor_1d.MatrixProductState.partial_trace_compress`
# and :func:`~quimb.approx_spectral_function` (see :ref:`Examples`).
# Finally, many quantities can be computed using local 'gates' see the section
# :ref:`gating`.
# + raw_mimetype="text/restructuredtext" active=""
# Matrix Product Operators
# ------------------------
#
# The raw MPO class is :class:`~quimb.tensor.tensor_1d.MatrixProductOperator`,
# which shares many features with :class:`~quimb.tensor.tensor_1d.MatrixProductState`,
# but has both a ``MPO.upper_ind_id`` and a ``MPO.lower_ind_id``.
#
# Here we generate a random hermitian MPO and form a 'overlap' network with our MPS:
# +
A = MPO_rand_herm(20, bond_dim=7, tags=['HAM'])
pH = p.H
# This inplace modifies the indices of each to form overlap
p.align_(A, pH)
(pH & A & p).draw(color='HAM', iterations=20, initial_layout='kamada_kawai')
# -
# Compute the actual contraction (``...`` means contract everything, but use the structure if possible):
(pH & A & p) ^ ...
# + raw_mimetype="text/restructuredtext" active=""
# Building Hamiltonians
# ---------------------
#
# There a few built-in MPO hamiltoanians:
#
# * :class:`~quimb.tensor.tensor_gen.MPO_ham_heis`
# * :class:`~quimb.tensor.tensor_gen.MPO_ham_ising`
# * :class:`~quimb.tensor.tensor_gen.MPO_ham_XY`
# * :class:`~quimb.tensor.tensor_gen.MPO_ham_mbl`
#
# These all accept a ``cyclic`` argument to enable periodic boundary
# conditions (PBC), and a ``S`` argument to set the size of spin.
#
# For generating other spin Hamiltonians see
# :class:`~quimb.tensor.tensor_gen.SpinHam1D`, or consider using the raw
# constructor of :class:`~quimb.tensor.tensor_1d.MatrixProductOperator`.
#
#
# Quick DMRG2 Intro
# -----------------
#
# First we build a Hamiltonian term by term (though we could just use :class:`~quimb.tensor.tensor_gen.MPO_ham_heis`):
# -
builder = SpinHam1D(S=1)
builder += 1/2, '+', '-'
builder += 1/2, '-', '+'
builder += 1, 'Z', 'Z'
H = builder.build_mpo(L=100)
# + raw_mimetype="text/restructuredtext" active=""
# Then we construct the 2-site DMRG object (:class:`~quimb.tensor.tensor_dmrg.DMRG2`), with the Hamiltonian MPO and a default sequence of maximum bond dimensions and a bond compression cutoff:
# -
dmrg = DMRG2(H, bond_dims=[10, 20, 100, 100, 200], cutoffs=1e-10)
# The ``DMRG`` object will automatically detect OBC/PBC. Now we can solve to a certain absolute energy tolerance, showing progress and a schematic of the final state:
dmrg.solve(tol=1e-6, verbosity=1)
dmrg.state.show(max_width=80)
# + raw_mimetype="text/restructuredtext" active=""
# There are many options stored in the dict ``DMRG.opts`` - an explanation of each of
# these is given in :func:`~quimb.tensor.tensor_dmrg.get_default_opts`, and it may be
# necessary to tweak these to achieve the best performance/accuracy, especially for
# PBC (see :ref:`examples`).
#
# .. note:: Performance Tips
#
# 1. Make sure numpy is linked to a fast BLAS (e.g. MKL version that comes with conda).
# 2. Install slepc4py, to use as the iterative eigensolver, it's faster than scipy.
# 3. If the hamiltonian is real, compile and use a real version of SLEPC
# (set the environment variable PETSC_ARCH before launch).
# 4. Periodic systems are in some ways easier to solve if *longer*, since this
# reduces correlations the 'long way round'.
#
#
# Quick TEBD Intro
# ----------------
#
# Time Evolving Block Decimation (:class:`~quimb.tensor.tensor_tebd.TEBD`) requires not a
# MPO but a specficiation of the local, interacting term(s) of a Hamiltonian.
# This is encapsulated in the :class:`~quimb.tensor.tensor_tebd.LocalHam1D` object, which is
# initialized with the sum of two site terms ``H2`` and one-site terms (if any), ``H1``.
#
# :class:`~quimb.tensor.tensor_tebd.LocalHam1D` objects can also be built directly
# from a :class:`~quimb.tensor.tensor_gen.SpinHam1D` instance
# using the :meth:`~quimb.tensor.tensor_gen.SpinHam1D.build_local_ham` method.
# There are also the following built-in LocalHam1D Hamiltonians:
#
# * :class:`~quimb.tensor.tensor_gen.ham1d_heis`
# * :class:`~quimb.tensor.tensor_gen.ham1d_ising`
# * :class:`~quimb.tensor.tensor_gen.ham1d_XY`
#
# Here we build a :class:`~quimb.tensor.tensor_tebd.LocalHam1D` using a
# ``SpinHam1D``:
# +
builder = SpinHam1D(S=1 / 2)
builder.add_term(1.0, 'Z', 'Z')
builder.add_term(0.9, 'Y', 'Y')
builder.add_term(0.8, 'X', 'X')
builder.add_term(0.6, 'Z')
H = ham_1d_heis(20, bz=0.1)
# check the two site term
H.terms[0, 1]
# + raw_mimetype="text/restructuredtext" active=""
# Then we set up an initial state and the :class:`~quimb.tensor.tensor_tebd.TEBD` object itself -
# which mimics the general api of :class:`quimb.Evolution`:
# -
psi0 = MPS_neel_state(20)
tebd = TEBD(psi0, H)
# Now we are ready to evolve. By setting a ``tol``, the required timestep ``dt`` is computed for us:
tebd.update_to(T=3, tol=1e-3)
# After the evolution we can see that entanglement has been generated throughout the chain:
tebd.pt.show()
# + raw_mimetype="text/restructuredtext" active=""
# A more complete demonstration can be found in the :ref:`examples`.
# + raw_mimetype="text/restructuredtext" active=""
# .. _gating:
#
# Gates: compute local quantities and simulate circuits
# -----------------------------------------------------
#
# On top of the builtin methods mentioned earlier
# (:meth:`~quimb.tensor.tensor_1d.MatrixProductState.entropy`,
# :meth:`~quimb.tensor.tensor_1d.MatrixProductState.schmidt_gap`,
# :meth:`~quimb.tensor.tensor_1d.MatrixProductState.magnetization`,
# :meth:`~quimb.tensor.tensor_1d.TensorNetwork1DVector.correlation`,
# :meth:`~quimb.tensor.tensor_1d.MatrixProductState.logneg_subsys`, etc.),
# many other quantities are encapsulated by the
# :meth:`~quimb.tensor.tensor_1d.TensorNetwork1DVector.gate` method,
# which works on any 1D tensor network vector (MPS, MERA, etc.).
# This 'applies' a given operator to 1 or more sites, whilst maintaining
# the 'physical', outer indices.
# This not only directly allows quantum circuit style computation simulation
# but also makes local quantities (i.e. non-MPO) easy to compute:
# +
import quimb as qu
Z = qu.pauli('Z')
# compute <psi0|Z_i|psi0> for neel state above
[
psi0.gate(Z, i).H @ psi0
for i in range(10)
]
# + raw_mimetype="text/restructuredtext" active=""
# There are four ways in which a gate can be applied:
#
# * Lazily (``contract=False``) - the gate is added to the tensor network
# but nothing is contracted. This is the default.
# * Lazily with split (``contract='split-gate'``) - the gate is split before
# it is added to the network.
# * Eagerly (``contract=True``) - the gate is contracted into the tensor
# network. If the gate acts on more than one site this will produce
# larger tensors.
# * Swap and Split (``contract='swap+split'``) - sites will be swapped
# until adjacent, the gate will be applied and the resulting
# tensor split, then the sites swapped back into their original positions.
# This explicitly maintains the exact structure of an MPS (at the cost of
# increasing bond dimension), unlike the other two methods.
#
# Here's a quantum computation style demonstration of the lazy method:
# +
import quimb as qu
# some operators to apply
H = qu.hadamard()
CNOT = qu.controlled('not')
# setup an intitial register of qubits
n = 10
psi0 = MPS_computational_state('0' * n, tags='PSI0')
# apply hadamard to each site
for i in range(n):
psi0.gate_(H, i, tags='H')
# apply CNOT to even pairs
for i in range(0, n, 2):
psi0.gate_(CNOT, (i, i + 1), tags='CNOT')
# apply CNOT to odd pairs
for i in range(1, n - 1, 2):
psi0.gate_(CNOT, (i, i + 1), tags='CNOT')
# -
# Note we have used the inplace ``gate_`` (with a trailing
# underscore) which modifies the original ``psi0`` object.
# However ``psi0`` has its physical site indices mantained
# such that it overall looks like the same object:
sorted(psi0.outer_inds())
(psi0.H & psi0) ^ all
# But the network now contains the gates as additional tensors:
psi0.draw(color=['PSI0', 'H', 'CNOT'], show_inds=True)
# With the swap and split method MPS form is always maintained, which
# allows a canonical form and thus optimal trimming of singular values:
# +
n = 10
psi0 = MPS_computational_state('0' * n)
for i in range(n):
# 'swap+split' will be ignore to one-site gates
psi0.gate_(H, i, contract='swap+split')
# use Z-phase to create entanglement
Rz = qu.phase_gate(0.42)
for i in range(n):
psi0.gate_(Rz, i, contract='swap+split')
for i in range(0, n, 2):
psi0.gate_(CNOT, (i, i + 1), contract='swap+split')
for i in range(1, n - 1, 2):
psi0.gate_(CNOT, (i, i + 1), contract='swap+split')
# act with one long-range CNOT
psi0.gate_(CNOT, (2, n - 2), contract='swap+split')
# -
# We now still have an MPS, but with increased bond dimension:
psi0.show()
# Finally, the eager (``contract=True``) method works fairly simply:
psi0_CNOT = psi0.gate(CNOT, (1, n -2 ), contract=True)
psi0_CNOT.draw(color=[psi0.site_tag(i) for i in range(n)])
# + raw_mimetype="text/restructuredtext" active=""
# Where we can see that the gate, site 1, and site 8 have been combined
# into a new rank-6 tensor.
#
# A much more detailed run-through of quantum circuit simulation using
# tensor networks and the :class:`~quimb.tensor.circuit.Circuit` object
# can be found in the example :ref:`quantum-circuit-example`.
|
docs/tensor-1d.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(start = -10, stop = 10, num = 100)
fx = 1/(2*np.pi) / (np.cosh(x/2))
gx = 1/(np.sqrt(2*np.pi))*np.exp(-x**2/2)
plt.plot(x,gx, lw=7, label = "Gaussian Dist")
plt.plot(x,fx, lw=7, label="HSD")
plt.legend()
plt.savefig("GD_HSD.png")
plt.show()
|
hypsecant_related/.ipynb_checkpoints/Untitled-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''keras_2'': conda)'
# language: python
# name: python36964bitkeras2conda59be9c9ec18e4cfd8a6b710b6bd0170c
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/UN-GCPDS/python-gcpds.EEG_Tensorflow_models/blob/main/Experimental/DW_LCAM/%5B1%5D_Main_EEG_representation_Giga.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="nWUE2feMKN6p" colab={"base_uri": "https://localhost:8080/"} outputId="426fb2a8-5b6a-4f0b-d665-52ac451fa427"
from google.colab import drive
drive.mount('/content/drive')
# + id="OKXbjSorJ6f3" colab={"base_uri": "https://localhost:8080/"} outputId="9fe405aa-22c2-4607-ca82-da98785589a1"
# Supporting modules
#-------------------------------------------------------------------------------
import numpy as np
import scipy.io as sio
import pywt
import pandas as pd
import pickle
import os
import matplotlib.pyplot as plt
import cv2
import warnings
import shutil
from scipy.signal import butter, lfilter, lfilter_zi, filtfilt
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.model_selection import StratifiedKFold,train_test_split,ShuffleSplit
warnings.filterwarnings("ignore")
#-------------------------------------------------------------------------------
# !pip install mne==0.19
import mne
from mne.decoding import CSP
from mne.io import read_raw_gdf
#-------------------------------------------------------------------------------
# + id="VAmdfwyKJ6f7"
# Definitions-------------------------------------------------------------------
def leer_GIGA_data(path_filename,ch,vt,sbj_id):
#--- info ------------------------------------------------------------------
# 2 ---> sample rate
# 7 ---> imaginary_left
# 8 ---> imaginary_right
# 11 ---> imaginary_event
# 14 ---> bad_trials
# class1: left
# class2: right
#---------------------------------------------------------------------------
raw = sio.loadmat(path_filename)
eeg_raw = raw['eeg']
sfreq = np.float(eeg_raw[0][0][2])
id_MI = np.where(eeg_raw[0][0][11]==1)
id_MI = id_MI[1]
raw_c1 = []
raw_c2 = []
y_c1 = []
y_c2 = []
for i in range(len(id_MI)):
l_thr = id_MI[i]-(sfreq*2-1)
h_thr = id_MI[i]+(sfreq*5)
tmp_c1 = eeg_raw[0][0][7][ch,np.int(l_thr):np.int(h_thr)]
tmp_c2 = eeg_raw[0][0][8][ch,np.int(l_thr):np.int(h_thr)]
raw_c1.append(tmp_c1[:,np.int(vt[0]*sfreq):np.int(vt[1]*sfreq)])
raw_c2.append(tmp_c2[:,np.int(vt[0]*sfreq):np.int(vt[1]*sfreq)])
y_c1.append(1.0)
y_c2.append(2.0)
# remove bad trials---------------------------------------------------------
id_bad_tr_voltage_c1 = eeg_raw[0][0][14][0][0][0][0][0]
id_bad_tr_voltage_c2 = eeg_raw[0][0][14][0][0][0][0][1]
id_bad_tr_mi_c1 = eeg_raw[0][0][14][0][0][1][0][0]
id_bad_tr_mi_c2 = eeg_raw[0][0][14][0][0][1][0][1]
ref_axis_c1 = 1
ref_axis_c2 = 1
if id_bad_tr_mi_c1.shape[0]>id_bad_tr_mi_c1.shape[1]:
id_bad_tr_mi_c1 = id_bad_tr_mi_c1.T
if id_bad_tr_mi_c2.shape[0]>id_bad_tr_mi_c2.shape[1]:
id_bad_tr_mi_c2 = id_bad_tr_mi_c2.T
if id_bad_tr_voltage_c1.shape[1] == 0:
id_bad_tr_voltage_c1 = np.reshape(id_bad_tr_voltage_c1, (id_bad_tr_voltage_c1.shape[0], id_bad_tr_mi_c1.shape[1]))
if id_bad_tr_voltage_c2.shape[1] == 0:
id_bad_tr_voltage_c2 = np.reshape(id_bad_tr_voltage_c2, (id_bad_tr_voltage_c2.shape[0], id_bad_tr_mi_c2.shape[1]))
if (id_bad_tr_voltage_c1.shape[1] > id_bad_tr_mi_c1.shape[1]):
if id_bad_tr_mi_c1.shape[0] == 0:
id_bad_tr_mi_c1 = np.reshape(id_bad_tr_mi_c1, (id_bad_tr_mi_c1.shape[0],id_bad_tr_voltage_c1.shape[1]))
ref_axis_c1 = 0
if (id_bad_tr_voltage_c2.shape[1] > id_bad_tr_mi_c2.shape[1]):
if id_bad_tr_mi_c2.shape[0] == 0:
id_bad_tr_mi_c2 = np.reshape(id_bad_tr_mi_c2, (id_bad_tr_mi_c2.shape[0],id_bad_tr_voltage_c2.shape[1]))
ref_axis_c2 = 0
if (id_bad_tr_mi_c1.shape[0] > id_bad_tr_voltage_c1.shape[0]):
ref_axis_c1 = 0
if (id_bad_tr_mi_c2.shape[0] > id_bad_tr_voltage_c2.shape[0]):
ref_axis_c2 = 0
if (id_bad_tr_voltage_c1.shape[0] > id_bad_tr_mi_c1.shape[0]):
ref_axis_c1 = 0
if (id_bad_tr_voltage_c2.shape[0] > id_bad_tr_mi_c2.shape[0]):
ref_axis_c2 = 0
id_bad_tr_c1 = np.concatenate((id_bad_tr_voltage_c1,id_bad_tr_mi_c1),axis=ref_axis_c1)
id_bad_tr_c1 = id_bad_tr_c1.ravel()-1
for ele in sorted(id_bad_tr_c1, reverse = True):
del raw_c1[ele]
del y_c1[ele]
id_bad_tr_c2 = np.concatenate((id_bad_tr_voltage_c2,id_bad_tr_mi_c2),axis=ref_axis_c2)
id_bad_tr_c2= id_bad_tr_c2.ravel()-1
for ele in sorted(id_bad_tr_c2, reverse = True):
del raw_c2[ele]
del y_c2[ele]
Xraw = np.array(raw_c1 + raw_c2)
y = np.array(y_c1 + y_c2)
return Xraw, y, sfreq
#-------------------------------------------------------------------------------
def bank_filter_epochsEEG(Xraw, fs, f_frec):
nf,ff = f_frec.shape
epochs,channels,T = Xraw.shape
Xraw_f = np.zeros((epochs,channels,T,nf))
for f in range(nf):
lfc = f_frec[f,0]
hfc = f_frec[f,1]
b,a = butter_bandpass(lfc, hfc, fs)
zi = lfilter_zi(b, a)
for n in range(epochs):
for c in range(channels):
zi = lfilter_zi(b, a)
Xraw_f[n,c,:,f] = lfilter(b, a, Xraw[n,c,:],zi = zi*Xraw[n,c,0])[0]
return Xraw_f
#-------------------------------------------------------------------------------
def butter_bandpass(lowcut, highcut, fs, order=5):
nyq = 0.5 * fs
low = lowcut / nyq
high = highcut / nyq
b, a = butter(order, [low, high], btype='band')
return b, a
#-------------------------------------------------------------------------------
def calculo_cwt(x,fs):
wname = 'cmor'
delta = 1/fs
coef,freq = pywt.cwt(x.T,np.arange(1,32),wname,delta)
return coef, freq
#-------------------------------------------------------------------------------
def cwt_feat_extraction(X,fs,freq_ref):
X_cwt = np.zeros((X.shape[0],X.shape[1],2))
for tr in range(X.shape[0]):#loop across trials
for ch in range(X.shape[1]):#loop across channels
coef, freq = calculo_cwt(np.squeeze(X[tr,ch,:,0]),fs)
coef = np.abs(coef)
fb_valavg = []
for fb in range(freq_ref.shape[0]):#loop across filter bands
coef_mat = coef[np.where((freq > freq_ref[fb,0]) & (freq <freq_ref[fb,1])),:]
coef_mat = np.squeeze(coef_mat[0,:,:])
X_cwt[tr,ch,fb] = np.mean(coef_mat.flatten())
return X_cwt
#-------------------------------------------------------------------------------
from sklearn.base import BaseEstimator, TransformerMixin
class CSP_epochs_filter_extractor(TransformerMixin,BaseEstimator):
def __init__(self, fs,f_frec=[4,30], ncomp=4,reg='empirical'):
self.reg = reg
self.fs = fs
self.f_frec = f_frec
self.ncomp = ncomp
def _averagingEEG(self,X):
epochs,channels,T = X.shape
Xc = np.zeros((epochs,channels,T))
for i in range(epochs):
Xc[i,:,:] = X[i,:,:] - np.mean(X[i,:,:])
return Xc
def _bank_filter_epochsEEG(self,X):
nf,ff = self.f_frec.shape
epochs,channels,T = X.shape
X_f = np.zeros((epochs,channels,T,nf))
for f in range(nf):
lfc = self.f_frec[f,0]
hfc = self.f_frec[f,1]
b,a = butter_bandpass(lfc, hfc, self.fs)
X_f[:,:,:,f] = filtfilt(b,a,X,axis=2)
return X_f
def _CSP_epochsEEG(self,Xraw, y,*_):
ncomp = self.ncomp
mne.set_log_level('WARNING')
epochs,channels,T,nf = Xraw.shape
Xcsp = np.zeros((epochs,self.ncomp,nf))
self.filters =np.zeros((self.ncomp,channels,nf))
csp_l = []
for f in range(nf):
csp_l+= [CSP(n_components=ncomp, reg=self.reg, log=False,transform_into='average_power').fit(Xraw[:,:,:,f],y)]
Xcsp[:,:,f] = csp_l[f].transform(Xraw[:,:,:,f])
self.filters[:,:,f] = csp_l[f].filters_[:self.ncomp]
return csp_l, Xcsp
def fit(self,Xraw,y, *_):
Xraw = self._averagingEEG(Xraw)
Xraw_f = self._bank_filter_epochsEEG(Xraw)
self.csp_l, self.Xcsp = self._CSP_epochsEEG(Xraw_f, y)
return self
def transform(self, Xraw, *_):
Xraw = self._averagingEEG(Xraw)
Xraw_f = self._bank_filter_epochsEEG(Xraw)
epochs,channels,T,nf = Xraw_f.shape
ncomp = self.ncomp
result = np.zeros((epochs,ncomp,nf))
for f in range(nf):
result[:,:,f] = self.csp_l[f].transform(Xraw_f[:,:,:,f])
return result
#-------------------------------------------------------------------------------
def csp_feat_extraction(Xtrain,ytrain,Xtest,fs,f_frec):
# Y = W.T * X
# A*Y = X ---- A= pinv(W.T)
XT_train = np.zeros((Xtrain.shape[0],Xtrain.shape[1],2))
XT_test = np.zeros((Xtest.shape[0],Xtest.shape[1],2))
ncomp = 6# Biclass (4-6) -- Multiclass (8-12)
csp_c = CSP_epochs_filter_extractor(fs=fs,f_frec=f_frec, ncomp=ncomp)
XT = csp_c.fit_transform(Xtrain,ytrain)
Filt_ = csp_c.filters
# train/test
for tr in range(Xtrain.shape[0]):#loop across train trials
for fb in range(len(f_frec)):#loop across filter bands
Xpr_tr = []
Xpr_tr = np.dot(Filt_[:,:,fb],Xtrain[tr,:,:])
Xfr_tr = []
Xfr_tr = np.dot(np.linalg.pinv(Filt_[:,:,fb]),Xpr_tr)
XT_train[tr,:,fb] = np.mean(np.abs(Xfr_tr),axis=1)
for tr in range(Xtest.shape[0]):#loop across test trials
for fb in range(len(f_frec)):#loop across filter bands
Xpr_ts = []
Xpr_ts = np.dot(Filt_[:,:,fb],Xtest[tr,:,:])
Xfr_ts = []
Xfr_ts = np.dot(np.linalg.pinv(Filt_[:,:,fb]),Xpr_ts)
XT_test[tr,:,fb] = np.mean(np.abs(Xfr_ts),axis=1)
return XT_train, XT_test
#-------------------------------------------------------------------------------
def topomap_generation(types,time_inf,time_sup,id_sbj,info):
cmap = 'gray'
newX = 40
newY = 40
for itm in range(len(types)): #len(types)
#-----------------------------------------------------------------------
path = '/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/X_'+types[itm]+'_sbj_'+str(id_sbj)+'_Tw_'+str(time_inf)+'s_'+str(time_sup)+'s.pickle'
with open(path, 'rb') as f:
XT_train, XT_test, y_train, y_test = pickle.load(f)
#-----------------------------------------------------------------------
try:
os.mkdir('figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm])
except OSError:
print('Folder exists!')
#-----------------------------------------------------------------------
# train
X = XT_train.copy()
#-----------------------------------------------------------------------
try:
os.mkdir('figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/train')
except OSError:
print('Folder exists!')
#-----------------------------------------------------------------------
X_train_reshape = np.zeros((X.shape[0],X.shape[2],int(newX),int(newY)))
#-----------------------------------------------------------------------
fig_mu = plt.figure(figsize=(10,10))
for tr in range(X.shape[0]):
fig_mu.clear()
image_mu = []
img_mu = []
rho_mu = []
rho_mu = (X[tr,:,0]-np.min(X[tr,:,0]))/(np.max(X[tr,:,0])-np.min(X[tr,:,0]))
mne.viz.plot_topomap(rho_mu, info, sensors=False, show=False, cmap=cmap, contours=0)
path_mu = 'figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/train/sbj_'+str(id_sbj)+'_tr_'+str(tr+1)+'_fb_mu.png'
fig_mu.savefig(fname=path_mu,dpi=40,format='png',facecolor='w')
image_mu = cv2.imread(path_mu,0)
img_mu = cv2.resize(image_mu,(int(newX),int(newY)))
X_train_reshape[tr,0,:,:] = img_mu
#-----------------------------------------------------------------------
fig_beta = plt.figure(figsize=(10,10))
for tr in range(X.shape[0]):#
fig_beta.clear()
image_beta = []
img_beta = []
rho_beta = []
rho_beta = (X[tr,:,1]-np.min(X[tr,:,1]))/(np.max(X[tr,:,1])-np.min(X[tr,:,1]))
mne.viz.plot_topomap(rho_beta, info, sensors=False, show=False, cmap=cmap, contours=0)
path_beta = 'figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/train/sbj_'+str(id_sbj)+'_tr_'+str(tr+1)+'_fb_beta.png'
fig_beta.savefig(fname=path_beta,dpi=40,format='png',facecolor='w')
image_beta = cv2.imread(path_beta,0)
img_beta = cv2.resize(image_beta,(int(newX),int(newY)))
X_train_reshape[tr,1,:,:] = img_beta
#-----------------------------------------------------------------------
X = X_train_reshape.copy()
#-----------------------------------------------------------------------
# test
X1 = XT_test.copy()
#-----------------------------------------------------------------------
try:
os.mkdir('figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/test')
except OSError:
print('Folder exists!')
#-----------------------------------------------------------------------
X_test_reshape = np.zeros((X1.shape[0],X1.shape[2],int(newX),int(newY)))
#-----------------------------------------------------------------------
fig_mu = plt.figure(figsize=(10,10))
for tr in range(X1.shape[0]):
fig_mu.clear()
image_mu = []
img_mu = []
rho_mu = []
rho_mu = (X1[tr,:,0]-np.min(X1[tr,:,0]))/(np.max(X1[tr,:,0])-np.min(X1[tr,:,0]))
mne.viz.plot_topomap(rho_mu, info, sensors=False, show=False, cmap=cmap, contours=0)
path_mu = 'figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/test/sbj_'+str(id_sbj)+'_tr_'+str(tr+1)+'_fb_mu.png'
fig_mu.savefig(fname=path_mu,dpi=40,format='png',facecolor='w')
image_mu = cv2.imread(path_mu,0)
img_mu = cv2.resize(image_mu,(int(newX),int(newY)))
X_test_reshape[tr,0,:,:] = img_mu
#-----------------------------------------------------------------------
fig_beta = plt.figure(figsize=(10,10))
for tr in range(X1.shape[0]):
fig_beta.clear()
image_beta = []
img_beta = []
rho_beta = []
rho_beta = (X1[tr,:,1]-np.min(X1[tr,:,1]))/(np.max(X1[tr,:,1])-np.min(X1[tr,:,1]))
mne.viz.plot_topomap(rho_beta, info, sensors=False, show=False, cmap=cmap, contours=0)
path_beta = 'figures/'+str(time_inf)+'s-'+str(time_sup)+'s/'+types[itm]+'/test/sbj_'+str(id_sbj)+'_tr_'+str(tr+1)+'_fb_beta.png'
fig_beta.savefig(fname=path_beta,dpi=40,format='png',facecolor='w')
image_beta = cv2.imread(path_beta,0)
img_beta = cv2.resize(image_beta,(int(newX),int(newY)))
X_test_reshape[tr,1,:,:] = img_beta
#-----------------------------------------------------------------------
X1 = X_test_reshape.copy()
#-----------------------------------------------------------------------
Xtr = X
Xts = X1
with open('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/CWT_CSP_data_mubeta_8_30_Tw_'+str(time_inf)+'s_'+str(time_sup)+'s_subject'+str(id_sbj)+'_'+types[itm]+'_resized_10.pickle', 'wb') as f:
pickle.dump([Xtr, Xts, y_train, y_test], f)
#-----------------------------------------------------------------------
#-------------------------------------------------------------------------------
# + [markdown] id="mYwzOWMfJ6gE"
# ### CWT and CSP feature extraction
# + id="XB4Ggr3sJ6gG"
,# Experiment information--------------------------------------------------------
th = np.array([[0.5, 2.5],[1.5, 3.5],[2.5, 4.5],[3.5, 5.5],[4.5, 6.5]])
th_name = np.array([[-1.5, 0.5],[-0.5, 1.5],[0.5, 2.5],[1.5, 3.5],[2.5, 4.5]])
freq_ref = np.array([[8,12],[12,30]])
Nsbj = [1]
#-------------------------------------------------------------------------------
for sbj in range(len(Nsbj)):#loop across subjects
for i in range(th_name.shape[0]):#loop across time windows #
#-----------------------------------------------------------------------
print('Subject - '+str(Nsbj[sbj])+' - Time window '+str(i+1)+' of '+str(th_name.shape[0]))
#-----------------------------------------------------------------------
# load EEG signals------------------------------------------------------
name = '/content/drive/MyDrive/Universidad-2020/CNN_GIGA/GIGAdata/s' + str(Nsbj[sbj])
filename_train = name+'.mat'
ch = np.arange(0,64)
vt = [th[i,0],th[i,1]]
Xraw,y,sfreq = leer_GIGA_data(filename_train,ch,vt,Nsbj[sbj])
fs = sfreq
#-----------------------------------------------------------------------
# Filtering-------------------------------------------------------------
f_frec = np.transpose(np.array([[8],[30]]))
Xraw_filt = bank_filter_epochsEEG(Xraw, fs, f_frec)
#-----------------------------------------------------------------------
# Split in train/test subsets-------------------------------------------
rs = ShuffleSplit(n_splits=1, train_size=0.9, test_size=0.1, random_state=0)
for train_index, test_index in rs.split(y):
X_train, y_train = Xraw_filt[train_index], y[train_index]
X_test, y_test = Xraw_filt[test_index], y[test_index]
#-----------------------------------------------------------------------
if i==0:
with open('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/idxs_train_test_'+str(Nsbj[sbj])+'.pickle', 'wb') as f:
pickle.dump([train_index, test_index], f)
#-----------------------------------------------------------------------
# Compute CWT feature extraction----------------------------------------
X_cwt_train = cwt_feat_extraction(X_train,fs,freq_ref)
X_cwt_test = cwt_feat_extraction(X_test,fs,freq_ref)
#-----------------------------------------------------------------------
# Compute CSP feature extraction----------------------------------------
X_csp_train,X_csp_test = csp_feat_extraction(np.squeeze(X_train),y_train,np.squeeze(X_test),fs,freq_ref)
#-----------------------------------------------------------------------
# Save extracted features-----------------------------------------------
with open('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/X_cwt_sbj_'+str(Nsbj[sbj])+'_Tw_'+str(th_name[i,0])+'s_'+str(th_name[i,1])+'s.pickle', 'wb') as f:
pickle.dump([X_cwt_train, X_cwt_test, y_train, y_test], f)
with open('/content/drive/MyDrive/Colab Notebooks/GradCam_Paper/GigaData/data/X_csp_sbj_'+str(Nsbj[sbj])+'_Tw_'+str(th_name[i,0])+'s_'+str(th_name[i,1])+'s.pickle', 'wb') as f:
pickle.dump([X_csp_train, X_csp_test, y_train, y_test], f)
#-----------------------------------------------------------------------
print('Feature Extraction Done!!!\n')
# + [markdown] id="GLCWkBZuJ6gI"
# ### Topographic map montage
# + id="z9HZiWklJ6gJ"
# set EEG montage using standard 10-20 system-----------------------------------
channels_names = ['FP1','AF7','AF3','F1','F3','F5','F7','FT7','FC5','FC3','FC1','C1',
'C3','C5','T7','TP7','CP5','CP3','CP1','P1','P3','P5','P7','P9','PO7',
'PO3','O1','Iz','Oz','POz','Pz','CPz','FPz','FP2','AF8','AF4','AFz',
'Fz','F2','F4','F6','F8','FT8','FC6','FC4','FC2','FCz','Cz','C2','C4',
'C6','T8','TP8','CP6','CP4','CP2','P2','P4','P6','P8','P10','PO8',
'PO4','O2']
montage = mne.channels.read_montage('standard_1020', channels_names)
info = mne.create_info(channels_names, sfreq=512, ch_types="eeg",
montage=montage)
f,ax = plt.subplots(1,1,figsize=(3,3))
mne.viz.plot_sensors(info, show_names=True,axes=ax)
#-------------------------------------------------------------------------------
# + [markdown] id="4Lm3bQWmJ6gK"
# ### Topoplot generation
# + id="b8gK-YnRJ6gK"
# Load feat data----------------------------------------------------------------
th_name = np.array([[-1.5, 0.5],[-0.5, 1.5],[0.5, 2.5],[1.5, 3.5],[2.5, 4.5]])
types = ['cwt','csp']
Nsbj = [30]
#-------------------------------------------------------------------------------
for sbj in range(len(Nsbj)):#loop across subjects
try:
os.mkdir('figures')
except OSError:
print('Folder exists!')
for i in range(th_name.shape[0]):#loop across time windows
print('Subject - '+str(Nsbj[sbj])+' - Time window '+str(i+1)+' of '+str(th_name.shape[0]))
try:
os.mkdir('figures/'+str(th_name[i,0])+'s-'+str(th_name[i,1])+'s')
except OSError:
print('Folder exists!')
topomap_generation(types,th_name[i,0],th_name[i,1],Nsbj[sbj],info)
shutil.rmtree('figures', ignore_errors=True)
print('Topoplot generation Done!!!\n')
#-------------------------------------------------------------------------------
|
Experimental/DW_LCAM/[1]_Main_EEG_representation_Giga.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
import numpy as np
import pandas as pd
import xarray as xr
import fsspec
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import cartopy.io.shapereader as shpreader
import matplotlib.pyplot as plt
#scientific colormaps
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import LinearSegmentedColormap
cm_data = np.loadtxt("./ScientificColormaps6/vik/vik.txt")
vik_map = LinearSegmentedColormap.from_list("vik", cm_data)
cm_data = np.loadtxt("./ScientificColormaps6/roma/roma.txt")
roma_map = LinearSegmentedColormap.from_list("roma", cm_data)
roma_map2 = LinearSegmentedColormap.from_list("roma", cm_data[-1::-1])
warnings.simplefilter('ignore') # filter some warning messages
xr.set_options(display_style="html") #display dataset nicely
# -
sst_anomaly_monthlyask_kubernetes import KubeCluster
from dask.distributed import Client
cluster = KubeCluster()
cluster.adapt(minimum=1, maximum=200, interval='2s', wait_count=3)
client = Client(cluster)
cluster
# +
# %%time
file_location = 's3://mur-sst/zarr'
ds_sst = xr.open_zarr(fsspec.get_mapper(file_location, anon=True),consolidated=True)
ds_sst
# -
# # subset data to region of interest
subset = ds_sst.sel(lat=slice(32,48),lon=slice(-132.27,-117))
# # calculate climatology using 2003-2019
# +
sst_monthly = subset.resample(time='1MS').mean('time',keep_attrs=True,skipna=False)
climatology_mean_monthly = sst_monthly.sel(time=slice('2003-01-01','2019-01-01')).groupby('time.month').mean('time',keep_attrs=True,skipna=False)
sst_anomaly_monthly = sst_monthly.groupby('time.month')-climatology_mean_monthly #take out annual mean to remove trends
sst_anomaly_monthly = sst_anomaly_monthly.persist()
sst_anomaly_monthly
# -
mask = sst_monthly.mask.isel(time=0).persist()
#axis([-132.27 -117 32.88 48.25]);
for itime in range(-100,-30):
test = sst_anomaly_monthly.isel(time=itime)#.load()
tem = test.where(mask<2)
tstr = str(test.time.data)[0:10]
plt.figure(figsize=(10, 8))
ax = plt.axes(projection=ccrs.PlateCarree())
plt.pcolormesh(tem.lon,tem.lat,tem.analysed_sst,transform=ccrs.PlateCarree(),cmap=vik_map,vmin=-2,vmax=2)
ax.coastlines(resolution='50m', color='black', linewidth=1)
ax.add_feature(cfeature.LAND)
ax.add_feature(cfeature.STATES.with_scale('10m'))
ax.set_extent([-132.27,-117,32,48])
plt.colorbar(ax=ax,label='SST Anomaly (K)')
tt=plt.text(-122,47,tstr,fontsize=16)
plt.savefig('./figs/anom'+tstr+'.png')
|
sst_science/monthly_california_sst_anomalies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Script to scrape load data from the website of the TSOC
#
# This script uses selenium and geckodriver to download automatically the electricity consumption data available at the website of the TSOC. Normally, the data are available in 15-day tranches, so this script makes the process of getting multi-month or multi-year data easier.
#
# The way it works is that we generate a list of start dates with 15 days difference of each other, covering the range of dates we want to download. Then, we formulate the URL and use Firefox in headless mode, to access the website and get the data.
#
# Of course, if the TSOC changes their website, things will fail.
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from datetime import date, timedelta
import time
import os
# +
# Get list of data I already have. This way we don't need to re-download everything again
# and reduces stress on the website. Be kind.
mypath = './power_data/'
datafiles = [f for f in os.listdir(mypath) if os.path.isfile(os.path.join(mypath, f))]
# Usually, the last file downloaded included days with NaN values. If for example we ask for 15-days
# but there are 10 days available, it will fill in the rest with NaN. So, we remove the last
# file to get an updated one.
lastfile = datafiles.pop()
os.remove(mypath+lastfile)
# Extract the start dates from the file names
datafilesclean = []
for f in datafiles:
datafilesclean.append(f[18:28])
# -
# Generate the dates I'll be requesting. It's from the start date to today with jumps of 15 days
start_date = date(2019, 1, 1)
end_date = date.today()
delta = timedelta(days=15)
datelist = []
while start_date <= end_date:
if start_date.strftime("%Y-%m-%d") not in datafilesclean:
datelist.append(start_date.strftime("%d-%m-%Y"))
start_date += delta
print(datelist)
# +
# This is the XPATH of the button to click to get the excel file
# I got this by "inspecting" the website
myxpath = "/html/body/div[2]/div/div/div/div/article/div/div[11]/div[1]/button"
# This is the path to geckodriver.exe and the download directory
# You need to have firefox installed as well for this to work
geckopath = r'C:\Users\p3tri\geckodriver.exe'
download_dir = 'C:\\Users\\p3tri\\OneDrive - Cyprus University of Technology\\Research projects\\2020\\EAC timeseries\\data\\'
# Tell to Firefox where to download the excel files automatically and where to put them
profile = webdriver.FirefoxProfile()
profile.set_preference("browser.download.folderList", 2)
profile.set_preference("browser.download.manager.showWhenStarting", False)
profile.set_preference("browser.download.dir", download_dir )
profile.set_preference("browser.helperApps.neverAsk.saveToDisk", "application/vnd.openxmlformats-officedocument.spreadsheetml.sheet")
options = webdriver.FirefoxOptions()
options.add_argument('-headless')
# Initialise the firefox browser
driver = webdriver.Firefox(executable_path=geckopath,firefox_profile=profile, options=options)
# -
# Loop over the dates and download
for dt in datelist:
# Generate the link to fetch the data
url="https://tsoc.org.cy/archive-total-daily-system-generation-on-the-transmission-system/?startdt="+dt+"&enddt=%2B15days"
# print(dt)
driver.get(url)
try:
btn = WebDriverWait(driver, 20).until(EC.element_to_be_clickable((By.XPATH, myxpath)))
except TimeoutException:
print("Loading took too much time: "+dt)
time.sleep(5)
btn.click()
time.sleep(30)
# Close everything
driver.close()
driver.quit()
|
power_data_scrap.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,.md//md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # `nbev3devsim` Setup Examples
#
# This notebook provides a suite of examples demonstrating different views over the `nbev3devsim` simulator, as well as ways of automating code execution within the simulator.
#
# The notebook also serves to act as informal documentation and an informal test suite.
# ## Load in the the simulator
#
# Running the follow cell should:
#
# - style the notebook with a two column like display, with the notebook shifted to the left column and the pop-up widget (a JQuery dialog widget) floating over the empty space to the right;
# - load in simulator widget.
#
# The widget should be draggable and resizable, with widget sizing controls available in its top bar.
#
# The notebook column should be resizable: draggable left and right to change the width.
#
# Known issues:
#
# - if the widget is larger than the display port, the scrolling and layout of the notebook breaks. The fix is to resize the widget to something that fits in the browser view, then scroll the notebook. The notebook is also drag resizable in such cases by clicking and dragging the bottom right hand corner of the notebook column.
# - sometimes the simulator view inside the widget gets detached from the widget sides and fails to fill the widget panel effectively. The fix is to click the widget *maximize* button and then the *restore* button.
# +
from nbev3devsim.load_nbev3devwidget import roboSim, eds
# %load_ext nbev3devsim
# -
# ## Overview of the user interface
#
# The user interface is based around a series of separate display panels. It is designed to try to provide ways of minimising how much of the screen is taken up by controls rather than the actual simulator.
#
# Each panel may contain display elements and/or additional display control.
#
# When a panel is displayed, clicking on the panel title will toggle the display of contents within the panel to *collapse* or *reveal* its content.
# ### *UI controls*
#
# The (unlabelled) *UI controls* area at the top of the widget provides display toggle button controls for displaying display and simulator control panels.
#
# When a display toggle button is pressed, it will show a dark background and the corresponding panel will be displayed. When the display toggle button has a white background, the corresponding panel will be hidden.
# + [markdown] activity=true
# Click individual *UI controls* buttons to toggle the display of various other control panels on and off.
# -
# The *UI controls* are always displayed.
#
# The and *Simulator world* panel are also generally displayed by default.
#
# Within many of the panels are one or more toggle display switches. The emphasised element is the one that is selected:
#
# - if the toggle button is to the left and shaded grey, the corresponding panel is *hidden*;
# - if toggle button is to the right and showing green, the corresponding panel is *shown*.
# + [markdown] activity=true
# Use the *Hide/Show* controls in the *UI controls* panel to show/hide the *Display controls* and *Simulator controls* panels.
#
# Click on the panel labels to collapse/reveal the contents of the corresponding panel.
# -
# ### *Display controls* panel
#
# ```
# Controlled via *UI controls* panel
# ```
#
# The *Display controls* panel contains controls for showing and hiding a wide range of panels:
#
# - *Settings*: various robot and simulator configuration settings;
# - *Output*: logging console for displaying print and error message from simulator program execution;
# - *Noise controls*: controls for applying noise to sensors and motors;
# - *Instrumentation*: display sensor and motor values;
# - *Sensor arrays*: display image sensor array views;
# - *Chart*: display realtime chart of instrumented values.
# ### *Simulator controls* panel
#
# ```
# Controlled via *UI controls* panel
#
# Keyboard shortcut: H
# ```
#
# The *Simulator controls* panel displays various controls for working directly with the simulator:
#
# - display world;
# - display positioning;
# - display code;
# - pen up / down;
# - run simulator;
# - clear trace.
# ### *Simulator world* panel
#
# ```
# Controlled via *Simulator* panel
#
# Keyboard shortcut: W
# ```
#
# The *World* panel is the simulator world view where the simulated robot performs its actions.
#
# *Known issues: the styling of this is broken. A fix to the layout in `EV3devSim.js` may also be required.*
# ### *Positioning* panel
#
# ```
# Accessed via *Simulator* panel
# Magics: --positioning / -X
# Keyboard shortcut: X
# ```
#
# The *Positioning* panel provides controls for displaying and positioning the simulated robot within the world view.
# ### *Code display* panel
#
# ```
# Accessed via *Simulator* panel
# Magics: --code / -D
# Keyboard shortcut: D
# ```
#
# The *Code display* panel will show the program currently downloaded into the simulator.
#
# *Known issues: if you download a new program to the simulator when the `Code display` is open the code is not updated. Closing and opening the display should display the updated program.*
# ### *Settings* panel
#
# ```
# Magics: --settings / -Z
# Keyboard shortcut: Z
# ```
#
# The *Settings* panel includes controls for setting:
#
# - robot configuration;
# - simulator background (map);
# - obstacles configuration;
# - collaborative mode (*experimental*).
# ### *Output* panel
#
# ```
# Magics: --output / -O
# Keyboard shortcut: O
# ```
#
# The *Output* panel is a terminal display window for viewing printed output and error messages when a program is run.
# ### *Noise controls* panel
#
# ```
# Magics: --noisecontrols / -z
# Keyboard shortcut: z
# ```
#
# The *Noise controls* panel displays controls for managing motor and sensor noise that is applied to the robot.
# ### *Instrumentation* panel
#
# ```
# Magics: --instrumentation / -i
# Keyboard shortcut: i
# ```
#
# The *Instrumentation* panel displays sensor and motor readings.
#
# *Known issue: I think this is continually updated. It would make sense to only update it if it is visible.*
# ### *Sensor arrays* panel
#
# ```
# Magics: --array / -A
# Keyboard shortcut: A
# ```
#
# The *Sensor arrays* panel displays the view of the sensor image arrays.
#
# *Known issue: I think this is continually updated. It would make sense to only update it if it is visible.*
# ### *Chart* panel
#
# ```
# Magics: --chart / -c
# Keyboard shortcut: c
# ```
#
# The *Chart* panel displays a realtime updated chart.
#
# *Known issues: need to check that this is only updated if it is displayed.*
# ### *Robot configurator* panel
#
# ```
# Accessed via *Settings* panel
# ```
#
# The *Robot configurator* panel allows you to view, edit, save and load robot configuration files.
#
# ### *Obstacles configurator* panel
#
# ```
# Accessed via *Settings* panel
# ```
#
# The *Obstacles configurator* panel allows you to:
#
# - edit, save and load obstacle configuration files;
# - control display of the walls around the simulated world;
# - control display of ultrasound rays.
#
# *Know issues: I don't think the walls are sized/ rendered correctly. Also, the walls don't seem to block robot progress?*
# ## Magic controls
#
# The notebook code cells are use to pass code to, and control the behaviour of, the widget using several magics:
#
# ```python
# # %sim_magic / %%sim_magic
# # %%sim_magic_imports
# # %%sim_magic_preloaded
# ```
#
# The `%sim_magic` line magic can be used to configure the simulator and display various help messages.
#
# When operating as a cell magic, the magic:
# - inspects and act on magic switches;
# - downloads code from the code cell into the simulator.
#
# The `%%sim_magic_imports` and `%%sim_magic_preloaded` magics generally operate as cell magics, with the exception of running as a line magic to display the code that they use to automatically prefix any code downloaded to the simulator.
#
# For example, a full list of switches can be displayed by passing the `--help / -h` flag:
# %sim_magic --help
# + [markdown] activity=true
# Run the follow code cells to preview the boilerplate code prepended by the corresponding magic.
# -
# %sim_magic_imports --preview
# %sim_magic_preloaded --preview
# Successfully downloading a program to the simulator is rewarded with an audible alert. To suppress the audible alert, pass the `--quiet / -q` flag in the magic command.
# + [markdown] activity=true
# Download a dummy program with some boilerplate code automatically prepended to it by running the following code cell. Then preview the code downloaded to the simulator via the *UI controls — Simulator controls — Show code* display button.
# -
# %%sim_magic_imports
pass
# *Known issues: the `--stop / -s` flag is currently broken.*
# One really handy switch is the `--background / -b` switch which lets us load in one of the pre-bundled backgrounds. We *could * selecting these from the settings menu:
# %sim_magic --settings
# Or instead we can select and load a background via a magic switch:
# %sim_magic -b MNIST_Digits
# ## Keyboard shortcuts
#
# The long term aim is to duplicate the boolean magic commands using keyboard shortcuts using the same single letter command.
#
# Simulator keyboard shortcuts are only enabled when the mouse cursor is within the bounds of the simulator widget.
#
# At the moment, the following keyboard shortcuts are supported:
#
# - `R` : run the currently loaded program in the simulator;
# - `S` : stop the currently running program in the simulator;
# - `p` : toggle pen up / down;
# - `X` : toggle display of positioning controls.
# - `A` : toggle display of sensor image array panel;
# - `O` : toggle display of output panel;
# - `c` : toggle display of chart panel;
# - `i` : toggle display of instrumentation panel;
# - `W` : toggle display of simulator world panel;
# - `z` : toggle display of simulator noise controls;
# - `Z` : toggle display of simulator configuration controls;
# - `D` : toggle display of code panel;
# - `H` : toggle display of simulator run controls.
#
# *Known issues: the `-S` operation to stop the currently running simulator program does not work.*
#
# For example, in the magic, by default the pen is up but we can set the pen down mode:
# %sim_magic --pendown
# If you move the mouse cursor over the simulator widget, you should also be able to toggle the pen up / pen down mode by pressing the *p* key.
#
# You should be able to toggle the various panel displays by pressing the appropriate keyboard shortcut key whilst the mouse cursor is over the widget.
# ## Controlling the simulator user interface configuration
#
# If you inspect the simulator interface, you will see it contains a range of controls for hiding and revealing various parts of the user interface (the *Hide/Show* toggle buttons).
#
# By default, the *Simulator controls* and *Simulator World* are displayed:
# %sim_magic
# Several magic switches are defined that control the display of simulator display panels, typically using the same single character shortcut as the keyboard shortcut controls:
#
# - `--output / -O`: Show output;
# - `--chart / -c`: Show chart;
# - `--instrumentation / -V`: Show sensor and motor values;
# - `--array / -A`: Show sensor array;
# - `--noisecontrols / -z`: Show sensor and motor noise controls;
# - `--positioning / -X`: Show positioning controls
# - `--worldcontrols / -Z`: Show world controls
# - `--hide / -H`: Hide simulator controls
# - `--world / -W`: Hide world
#
# So for example, we can configure the simulator to show only the world display among the optional display elements by suppressing the display of the simulator controls (all other panels are hidden by default):
#
# *Known issues: if the obstacles or robot config panels were opened manually, they will remain open.*
# %sim_magic -H
# We can hide the world display (at the bottom of the widget), by passing just the `-W` flag:
# %sim_magic -W
# We can pass multiple flags separately (`-W -H`) or compounded (`-WH`):
# %sim_magic -WHZ
# The display switches thus provide us with a means of scripting how the simulator widget controls are displayed for any given activity.
#
# For example, we might want to configure the simulator to show the sensor image array and the noise controls, but hide most of the other show/hide controls to free up screen real estate (note that the world is displayed unless we explicitly hide it):
# %sim_magic -Az
# Run the following code cell to toggle through the various cell displays:
# +
import time
for i in "HOcVAzZW":
# %sim_magic -$i
time.sleep(2)
# -
# We can also script values for the various numerical sliders that appear in the simulator user interface:
#
# - `--xpos, -x`: x co-ord config;
# - `--ypos, -y`: y co-ord config;
# - `--angle, -a`: Angle config;
# - `--sensornoise, -N`: Sensor noise, 0..128;
# - `--motornoise, -M`: Motor noise, 0..500.
#
# We can set just a single co-ordinate value:
# %sim_magic --positioning -y 450
# Or multiple values:
# %sim_magic -X -x 200 -y 700 -a 150
# ## Downloading programs to the simulator
#
# We can download a program to the simulator by prefixing the code cell with one of the simulator magics. The different magics prepend various bits of boilerplate code to the program before downloading it.
#
# When a program is downloaded, there is an audible alert:
# %%sim_magic
pass
# We can download and run a program automatically (`--autorun / -R`). Note that the run status indicator changes colour from red (not running) to green (running) as the program runs and then back to red when the program completes.
# %%sim_magic -R
import time
time.sleep(3)
# We can view the downloaded code by opening the *Code display* panel:
# %sim_magic -D
# Trying to run a broken program in the simulator will give an audible warning:
#
# *Known issue: there is no way to disable this at the moment.*
# +
# %%sim_magic -R -q -O
broken
# -
# We can print to the *Output* display panel.
#
# For example, let's hide the *Simulator controls* and *Simulator world* panels but display the *Output* window and then download and run a program that writes to it:
# %%sim_magic -OHW -R
import time
for i in range(3):
print("Hello number {}".format(i))
time.sleep(1)
# We can get a downloaded program to talk...
# +
# %%sim_magic_preloaded -R
say("hello")
# -
# When driving the robot, we can enable a "pen down" feature to leave a trace showing the path followed by the robot.
#
# For example, drive forward a short way without the pen down:
# +
# %%sim_magic_preloaded -x 100 -y 800 -R
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# -
# Now drive forward with the pen down (`--pendown / -p`):
# +
# %%sim_magic_preloaded -x 100 -y 820 -R -p
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# -
# If we drive forward again, the trace will remain in place.
#
# For example, change the pen colour ( `--pencolor / -P`) and drive forwards again:
# +
# %%sim_magic_preloaded -x 100 -y 840 -R -p -P green
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# -
# *Known issue: the pen colour selector widget is not enabled if the pen down toggle is selected from the magic.*
# Alternatively, we can clear the trace (`--clear / -C`) and then drive forward, again with the pen down and a selected color:
# +
# %%sim_magic_preloaded -x 100 -y 800 -R -p -C -P orange
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# -
# You can suppress the audible download alert by passing the `--quiet, -q` switch:
# %%sim_magic -q
pass
# ### Using the *Simulator controls*
#
# ```
# Magics: --hide / -H
# Keyboard shortcut: H
# ```
#
# The *simulator controls* panel provides manual user interface controls for controlling the simulator and managing the display of several panels related to simulator operation:
#
# - toggle display of the simulator world;
# - toggle display of the positioning controls;
# - toggle display of the code panel showing the currently downloaded program;
# - toggle the pen up / pen down state;
# - select the pen color;
# - clear pen traces from the current simulator world view;
# - start (*Run*) / stop the execution of the program currently downloaded to the simulator in the simulator; a status light (*red* on stop, *green* when running) displays the current run state;
# %sim_magic -W
# ### Viewing sensor and motor *Instrumentation*
#
# ```
# Magics: --instrumentation / -i
# Keyboard shortcut: i
# ```
#
# The *Instrumentation* panel provides information regarding sensor and motor readings.
#
# - `LeftMotor` / `RightMotor` report tachometer counts for each motor;
# - `Sensor1` / `Sensor2` report left and right light sensor values as follows: reflected light percentage for the red component range 0..100; average reflected light percentage over all three RGB components range 0..100; 3-tuple of RGB values, each in range 0..255;
# - `Ultrasonic`: distance reading to obstacle;
# - `Gyro`: the *angle* in degrees since the sensor was initialised and the *rate* at which the sensor is rotating, in degrees/second.
# %sim_magic -i -WH
# Run the following code cell to drive the robot over the *Testcard* background and see how the sensor values update as the robot moves.
# +
# %%sim_magic_preloaded -i -HR -b Testcard
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# -
# ### Using *Sensor arrays*
#
# ```
# Magics: --array / -A
# Keyboard shortcut: A
# ```
#
# The sensor image arrays provide a view of what the sensor can see at the pixel level.
#
# Displaying the sensor arrays and hiding the world provides opportunities for exploration style robot programming challenges.
# %sim_magic -A -WH -b MNIST_Digits
# ### Using *Noise controls*
#
# ```
# Magics:
# --noisecontrols / -z
# --sensornoise / -N [0..128]
# --motornoise / -M [0..500]
#
# Keyboard shortcut: z
# ```
#
# The *Noise controls* provide a numerical slider to set the noise level across both motors (range 0..500) and both light sensors (range 0..128).
# %sim_magic -z -WR
# Noise levels can be set on the light sensors via the `--sensornoise / -N` switch:
# %sim_magic -zWR --sensornoise 100
# %sim_magic -zWR --motornoise 100
# + [markdown] activity=true
# Drag and drop the robot around the plain white background, observing the light sensor instrumentation values and the sensor array view for different levels of light sensor noise.
# -
# %%sim_magic_preloaded -iAHR -x 100 -b Empty_Map -N 100
# + [markdown] activity=true
# Run the following code cell to download a program that will drive the robot forward a short way:
# +
# %%sim_magic_preloaded -HR -R -x 100 -y 700 -M 0
tank_drive.on_for_seconds(SpeedPercent(50),
SpeedPercent(50), 2)
# + [markdown] activity=true
# Now run the program several times in the presence of sensor noise:
# +
import time
# %sim_magic -RC -x 100 -y 700 -M 100 -p -P red
time.sleep(3)
# %sim_magic -R -x 100 -y 700 -M 200 -p -P orange
time.sleep(3)
# %sim_magic -R -x 100 -y 700 -M 300 -p -P green
# -
# ### Using the Live *Chart*
#
# ```
# Magics: --chart / c
# Keyboard shortcut: c
# ```
#
# The live *Chart* panel displays a dynamic line chart that can be configured to display selected instrumentation data traces logged to the output in a specific way.
#
# Supported chart traces:
#
# - *colour*;
# - *Left light* and *Right light* sensor values;
# - *Ultrasonic* distance;
# - *Gyro* angle;
# - *Left wheel* and *Right wheel* tacho counts.
# %sim_magic -c -WH
#
|
content/99. Technical Appendices/99.2 nbev3devsim user interface.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AEBS - AOBxFAC_2F product - long term teporal development
#
# Demonstration of `AOBxFAC_2F` product.
#
# Feel free to change the `SPACECRAFT`, `START_TIME`, and `END_TIME` paramteres and re-run the notebook.
#
# The notebook retrieves and displays data for the selected satellite and time range.
# +
#------------------------------------------------------------------------------
# User input:
SPACECRAFT = 'A'
START_TIME='2015-05-01T00:00:00Z'
END_TIME='2015-07-31T00:00:00Z'
SERVER_URL = 'https://staging.viresdisc.vires.services/ows'
#------------------------------------------------------------------------------
from viresclient import SwarmRequest
PI_START = +1
PI_STOP = -1
SwarmRequest.COLLECTIONS.update({
'AOB_FAC': ['SW_OPER_AOB{}FAC_2F'.format(x) for x in 'ABC'],
})
SwarmRequest.PRODUCT_VARIABLES.update({
'AOB_FAC': [
'Latitude_QD', 'Longitude_QD', 'MLT_QD',
'Boundary_Flag', 'Quality', 'Pair_Indicator'
],
})
try:
request = SwarmRequest(SERVER_URL)
request.set_collection('SW_OPER_AOB%sFAC_2F' % SPACECRAFT)
request.set_products(
measurements=[
'Latitude_QD', 'Longitude_QD', 'MLT_QD', 'Boundary_Flag',
'Quality', 'Pair_Indicator',
],
auxiliaries=['QDOrbitDirection', 'OrbitNumber', 'Kp'],
)
response = request.get_between(
start_time=START_TIME,
end_time=END_TIME,
)
print('Sources:\n\t%s' % '\n\t'.join(response.sources))
boundaries = response.as_xarray()
except Exception as error:
print('ERROR: ', error)
else:
print(boundaries)
# +
from numpy import stack
from matplotlib.pyplot import figure, subplot, show
# %matplotlib inline
def _orbit_lat(latitude, orbit_direction):
orbit_latitude = latitude.copy()
mask = orbit_direction == -1
orbit_latitude[mask] = 180 - latitude[mask]
orbit_latitude[orbit_latitude > 180] -= 360
return orbit_latitude
def _get_pair_indices(time, pair_indicator):
idx_start = (pair_indicator == PI_START).nonzero()[0]
idx_stop = (pair_indicator == PI_STOP).nonzero()[0]
if idx_start.size > 0 and idx_stop.size > 0:
if time[idx_start[0]] > time[idx_stop[0]]:
idx_stop = idx_stop[1:]
if time[idx_start[-1]] > time[idx_stop[-1]]:
idx_start = idx_start[:-1]
assert idx_start.size == idx_start.size
return stack((idx_start, idx_stop), axis=1)
def plot_aob(ax, is_north=True):
b_time = boundaries['Timestamp'].values
b_pair_indicator = boundaries['Pair_Indicator'].values
b_lat_qd = _orbit_lat(
boundaries['Latitude_QD'].values,
boundaries['QDOrbitDirection'].values
)
idx = _get_pair_indices(b_time, b_pair_indicator)
l_ao = ax.plot(b_time[idx].transpose(), b_lat_qd[idx].transpose(), '-', c='tab:blue', ms=3)
l_aob = ax.plot(b_time, b_lat_qd, '+', c='tab:red', ms=3)
if is_north:
ylim = [45, 135]
yticks = [45, 60, 75, 90, 105, 120, 135]
ylabels = ['+45\u00B0\u2191', '+60\u00B0\u2191', '+75\u00B0\u2191', '+90\u00B0', '+75\u00B0\u2193', '+60\u00B0\u2193', '+45\u00B0\u2193']
else:
ylim = [-135, -45]
yticks = [-135, -120, -105, -90, -75, -60, -45]
ylabels = ['\u221245\u00B0\u2193', '\u221260\u00B0\u2193', '\u221275\u00B0\u2193', '\u221290\u00B0', '\u221275\u00B0\u2191', '\u221260\u00B0\u2191', '\u221245\u00B0\u2191']
ax.legend((l_ao[0], l_aob[0]), ('Auroral Oval', 'Auroral Boundary'))
ax.set_ylim(ylim)
ax.set_yticks(yticks)
ax.set_yticklabels(ylabels)
ax.grid()
ax.set_ylabel('QD-latitude')
def plot_kp(ax, is_north=True):
b_time = boundaries['Timestamp'].values
b_kp = boundaries['Kp'].values
ax.plot(b_time, b_kp)
ax.grid()
ax.set_yticks(range(0, 10))
ax.set_ylim([-0.5, 9.5])
ax.set_ylabel('Kp index')
fig = figure(figsize=(18, 12), dpi=100)
plot_aob(subplot(311), is_north=True)
plot_aob(subplot(312), is_north=False)
plot_kp(subplot(313))
|
AEBS/AEBS_AOB_FAC_longterm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content-dl/blob/main/tutorials/W3D2_BasicReinforcementLearning/student/W3D2_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# # Tutorial 1: Introduction to Reinforcement Learning
# **Week 3, Day 2: Basic Reinforcement Learning (RL)**
#
# **By Neuromatch Academy**
#
# __Content creators:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# __Content editors:__ <NAME>
#
# __Production editors:__ <NAME>
#
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# ---
# # Tutorial Objectives
#
# By the end of the tutorial, you should be able to:
#
# 1. Within the RL framework, be able to identify the different components: environment, agent, states, and actions.
# 2. Understand the Bellman equation and components involved.
# 3. Implement tabular value-based model-free learning (Q-learning and SARSA).
# 4. Run a DQN agent and experiment with different hyperparameters.
# 5. Have a high-level understanding of other (nonvalue-based) RL methods.
# 6. Discuss real-world applications and ethical issues of RL.
#
# **Note:** There is an issue with some images not showing up if you're using a Safari browser. Please switch to Chrome if this is the case.
# + cellView="form"
# @title Tutorial slides
# @markdown These are the slides for the videos in this tutorial
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/m3kqy/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
# -
# ---
#
# # Setup
#
# Run the following 5 cells in order to set up needed functions. Don't worry about the code for now!
# + cellView="form"
# @title Install requirements
from IPython.display import clear_output
# @markdown we install the acme library, see [here](https://github.com/deepmind/acme) for more info
# @markdown WARNING: There may be errors and warnings reported during the installation.
# @markdown However, they should be ignored.
# !apt-get install -y xvfb ffmpeg --quiet
# !pip install --upgrade pip --quiet
# !pip install imageio --quiet
# !pip install imageio-ffmpeg
# !pip install gym --quiet
# !pip install enum34 --quiet
# !pip install dm-env --quiet
# !pip install pandas --quiet
# !pip install keras-nightly==2.5.0.dev2021020510 --quiet
# !pip install grpcio==1.34.0 --quiet
# !pip install tensorflow --quiet
# !pip install typing --quiet
# !pip install einops --quiet
# !pip install dm-acme --quiet
# !pip install dm-acme[reverb] --quiet
# !pip install dm-acme[tf] --quiet
# !pip install dm-acme[envs] --quiet
# !pip install dm-env --quiet
clear_output()
# +
# Import modules
import gym
import enum
import copy
import time
import acme
import torch
import base64
import dm_env
import IPython
import imageio
import warnings
import itertools
import collections
import numpy as np
import pandas as pd
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import matplotlib as mpl
import matplotlib.pyplot as plt
import tensorflow.compat.v2 as tf
from acme import specs
from acme import wrappers
from acme.utils import tree_utils
from acme.utils import loggers
from torch.autograd import Variable
from torch.distributions import Categorical
from typing import Callable, Sequence
tf.enable_v2_behavior()
warnings.filterwarnings('ignore')
np.set_printoptions(precision=3, suppress=1)
# + cellView="form"
# @title Figure settings
import ipywidgets as widgets # interactive display
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
mpl.rc('image', cmap='Blues')
# + cellView="form"
# @title Helper Functions
# @markdown Implement helpers for value visualisation
map_from_action_to_subplot = lambda a: (2, 6, 8, 4)[a]
map_from_action_to_name = lambda a: ("up", "right", "down", "left")[a]
def plot_values(values, colormap='pink', vmin=-1, vmax=10):
plt.imshow(values, interpolation="nearest",
cmap=colormap, vmin=vmin, vmax=vmax)
plt.yticks([])
plt.xticks([])
plt.colorbar(ticks=[vmin, vmax])
def plot_state_value(action_values, epsilon=0.1):
q = action_values
fig = plt.figure(figsize=(4, 4))
vmin = np.min(action_values)
vmax = np.max(action_values)
v = (1 - epsilon) * np.max(q, axis=-1) + epsilon * np.mean(q, axis=-1)
plot_values(v, colormap='summer', vmin=vmin, vmax=vmax)
plt.title("$v(s)$")
def plot_action_values(action_values, epsilon=0.1):
q = action_values
fig = plt.figure(figsize=(8, 8))
fig.subplots_adjust(wspace=0.3, hspace=0.3)
vmin = np.min(action_values)
vmax = np.max(action_values)
dif = vmax - vmin
for a in [0, 1, 2, 3]:
plt.subplot(3, 3, map_from_action_to_subplot(a))
plot_values(q[..., a], vmin=vmin - 0.05*dif, vmax=vmax + 0.05*dif)
action_name = map_from_action_to_name(a)
plt.title(r"$q(s, \mathrm{" + action_name + r"})$")
plt.subplot(3, 3, 5)
v = (1 - epsilon) * np.max(q, axis=-1) + epsilon * np.mean(q, axis=-1)
plot_values(v, colormap='summer', vmin=vmin, vmax=vmax)
plt.title("$v(s)$")
def plot_stats(stats, window=10):
plt.figure(figsize=(16,4))
plt.subplot(121)
xline = range(0, len(stats.episode_lengths), window)
plt.plot(xline, smooth(stats.episode_lengths, window=window))
plt.ylabel('Episode Length')
plt.xlabel('Episode Count')
plt.subplot(122)
plt.plot(xline, smooth(stats.episode_rewards, window=window))
plt.ylabel('Episode Return')
plt.xlabel('Episode Count')
# + cellView="form"
# @title Helper functions
def smooth(x, window=10):
return x[:window*(len(x)//window)].reshape(len(x)//window, window).mean(axis=1)
# + cellView="form"
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# for DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
# + cellView="form"
# @title Set device (GPU or CPU). Execute `set_device()`
# especially if torch modules used.
# inform the user if the notebook uses GPU or CPU.
def set_device():
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("WARNING: For this notebook to perform best, "
"if possible, in the menu under `Runtime` -> "
"`Change runtime type.` select `GPU` ")
else:
print("GPU is enabled in this notebook.")
return device
# -
SEED = 2021
set_seed(seed=SEED)
DEVICE = set_device()
# ---
# # Section 1: Introduction to Reinforcement Learning
# + cellView="form"
# @title Video 1: Introduction to RL
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV18V411p7iK", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"BWz3scQN50M", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Acme: a research framework for reinforcement learning
#
# **Acme** is a library of reinforcement learning (RL) agents and agent building blocks by Google DeepMind. Acme strives to expose simple, efficient, and readable agents, that serve both as reference implementations of popular algorithms and as strong baselines, while still providing enough flexibility to do novel research. The design of Acme also attempts to provide multiple points of entry to the RL problem at differing levels of complexity.
#
# For more information see [github repository](https://github.com/deepmind/acme).
# ---
# # Section 2: General Formulation of RL Problems and Gridworlds
#
# + cellView="form"
# @title Video 2: General Formulation and MDPs
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1k54y1E7Zn", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"h6TxAALY5Fc", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# The agent interacts with the environment in a loop corresponding to the following diagram. The environment defines a set of <font color='blue'>**actions**</font> that an agent can take. The agent takes an action informed by the <font color='redorange'>**observations**</font> it receives, and will get a <font color='green'>**reward**</font> from the environment after each action. The goal in RL is to find an agent whose actions maximize the total accumulation of rewards obtained from the environment.
#
#
# <center><img src="https://drive.google.com/uc?id=1KktLm5mdWx1ORotxeYCq1WcQHkXzRT4F" width="500" /></center>
#
#
#
# ## Section 2.1: The Environment
#
#
# For this practical session we will focus on a **simple grid world** environment,which consists of a 9 x 10 grid of either wall or empty cells, depicted in black and white, respectively. The smiling agent starts from an initial location and needs to navigate to reach the goal square.
#
# <center>
# <img src="https://drive.google.com/uc?id=163QdCqrPybJVVO0NhDxpun5O0YZmCnsI" width="500" />
# </center>
#
# Below you will find an implementation of this Gridworld as a ```dm_env.Environment```.
#
# There is no coding in this section, but if you want, you can look over the provided code so that you can familiarize yourself with an example of how to set up a **grid world** environment.
#
#
#
# + cellView="form"
# @title Implement GridWorld { form-width: "30%" }
# @markdown *Double-click* to inspect the contents of this cell.
class ObservationType(enum.IntEnum):
STATE_INDEX = enum.auto()
AGENT_ONEHOT = enum.auto()
GRID = enum.auto()
AGENT_GOAL_POS = enum.auto()
class GridWorld(dm_env.Environment):
def __init__(self,
layout,
start_state,
goal_state=None,
observation_type=ObservationType.STATE_INDEX,
discount=0.9,
penalty_for_walls=-5,
reward_goal=10,
max_episode_length=None,
randomize_goals=False):
"""Build a grid environment.
Simple gridworld defined by a map layout, a start and a goal state.
Layout should be a NxN grid, containing:
* 0: empty
* -1: wall
* Any other positive value: value indicates reward; episode will terminate
Args:
layout: NxN array of numbers, indicating the layout of the environment.
start_state: Tuple (y, x) of starting location.
goal_state: Optional tuple (y, x) of goal location. Will be randomly
sampled once if None.
observation_type: Enum observation type to use. One of:
* ObservationType.STATE_INDEX: int32 index of agent occupied tile.
* ObservationType.AGENT_ONEHOT: NxN float32 grid, with a 1 where the
agent is and 0 elsewhere.
* ObservationType.GRID: NxNx3 float32 grid of feature channels.
First channel contains walls (1 if wall, 0 otherwise), second the
agent position (1 if agent, 0 otherwise) and third goal position
(1 if goal, 0 otherwise)
* ObservationType.AGENT_GOAL_POS: float32 tuple with
(agent_y, agent_x, goal_y, goal_x)
discount: Discounting factor included in all Timesteps.
penalty_for_walls: Reward added when hitting a wall (should be negative).
reward_goal: Reward added when finding the goal (should be positive).
max_episode_length: If set, will terminate an episode after this many
steps.
randomize_goals: If true, randomize goal at every episode.
"""
if observation_type not in ObservationType:
raise ValueError('observation_type should be a ObservationType instace.')
self._layout = np.array(layout)
self._start_state = start_state
self._state = self._start_state
self._number_of_states = np.prod(np.shape(self._layout))
self._discount = discount
self._penalty_for_walls = penalty_for_walls
self._reward_goal = reward_goal
self._observation_type = observation_type
self._layout_dims = self._layout.shape
self._max_episode_length = max_episode_length
self._num_episode_steps = 0
self._randomize_goals = randomize_goals
if goal_state is None:
# Randomly sample goal_state if not provided
goal_state = self._sample_goal()
self.goal_state = goal_state
def _sample_goal(self):
"""Randomly sample reachable non-starting state."""
# Sample a new goal
n = 0
max_tries = 1e5
while n < max_tries:
goal_state = tuple(np.random.randint(d) for d in self._layout_dims)
if goal_state != self._state and self._layout[goal_state] == 0:
# Reachable state found!
return goal_state
n += 1
raise ValueError('Failed to sample a goal state.')
@property
def layout(self):
return self._layout
@property
def number_of_states(self):
return self._number_of_states
@property
def goal_state(self):
return self._goal_state
@property
def start_state(self):
return self._start_state
@property
def state(self):
return self._state
def set_state(self, x, y):
self._state = (y, x)
@goal_state.setter
def goal_state(self, new_goal):
if new_goal == self._state or self._layout[new_goal] < 0:
raise ValueError('This is not a valid goal!')
# Zero out any other goal
self._layout[self._layout > 0] = 0
# Setup new goal location
self._layout[new_goal] = self._reward_goal
self._goal_state = new_goal
def observation_spec(self):
if self._observation_type is ObservationType.AGENT_ONEHOT:
return specs.Array(
shape=self._layout_dims,
dtype=np.float32,
name='observation_agent_onehot')
elif self._observation_type is ObservationType.GRID:
return specs.Array(
shape=self._layout_dims + (3,),
dtype=np.float32,
name='observation_grid')
elif self._observation_type is ObservationType.AGENT_GOAL_POS:
return specs.Array(
shape=(4,), dtype=np.float32, name='observation_agent_goal_pos')
elif self._observation_type is ObservationType.STATE_INDEX:
return specs.DiscreteArray(
self._number_of_states, dtype=int, name='observation_state_index')
def action_spec(self):
return specs.DiscreteArray(4, dtype=int, name='action')
def get_obs(self):
if self._observation_type is ObservationType.AGENT_ONEHOT:
obs = np.zeros(self._layout.shape, dtype=np.float32)
# Place agent
obs[self._state] = 1
return obs
elif self._observation_type is ObservationType.GRID:
obs = np.zeros(self._layout.shape + (3,), dtype=np.float32)
obs[..., 0] = self._layout < 0
obs[self._state[0], self._state[1], 1] = 1
obs[self._goal_state[0], self._goal_state[1], 2] = 1
return obs
elif self._observation_type is ObservationType.AGENT_GOAL_POS:
return np.array(self._state + self._goal_state, dtype=np.float32)
elif self._observation_type is ObservationType.STATE_INDEX:
y, x = self._state
return y * self._layout.shape[1] + x
def reset(self):
self._state = self._start_state
self._num_episode_steps = 0
if self._randomize_goals:
self.goal_state = self._sample_goal()
return dm_env.TimeStep(
step_type=dm_env.StepType.FIRST,
reward=None,
discount=None,
observation=self.get_obs())
def step(self, action):
y, x = self._state
if action == 0: # up
new_state = (y - 1, x)
elif action == 1: # right
new_state = (y, x + 1)
elif action == 2: # down
new_state = (y + 1, x)
elif action == 3: # left
new_state = (y, x - 1)
else:
raise ValueError(
'Invalid action: {} is not 0, 1, 2, or 3.'.format(action))
new_y, new_x = new_state
step_type = dm_env.StepType.MID
if self._layout[new_y, new_x] == -1: # wall
reward = self._penalty_for_walls
discount = self._discount
new_state = (y, x)
elif self._layout[new_y, new_x] == 0: # empty cell
reward = 0.
discount = self._discount
else: # a goal
reward = self._layout[new_y, new_x]
discount = 0.
new_state = self._start_state
step_type = dm_env.StepType.LAST
self._state = new_state
self._num_episode_steps += 1
if (self._max_episode_length is not None and
self._num_episode_steps >= self._max_episode_length):
step_type = dm_env.StepType.LAST
return dm_env.TimeStep(
step_type=step_type,
reward=np.float32(reward),
discount=discount,
observation=self.get_obs())
def plot_grid(self, add_start=True):
plt.figure(figsize=(4, 4))
plt.imshow(self._layout <= -1, interpolation='nearest')
ax = plt.gca()
ax.grid(0)
plt.xticks([])
plt.yticks([])
# Add start/goal
if add_start:
plt.text(
self._start_state[1],
self._start_state[0],
r'$\mathbf{S}$',
fontsize=16,
ha='center',
va='center')
plt.text(
self._goal_state[1],
self._goal_state[0],
r'$\mathbf{G}$',
fontsize=16,
ha='center',
va='center')
h, w = self._layout.shape
for y in range(h - 1):
plt.plot([-0.5, w - 0.5], [y + 0.5, y + 0.5], '-w', lw=2)
for x in range(w - 1):
plt.plot([x + 0.5, x + 0.5], [-0.5, h - 0.5], '-w', lw=2)
def plot_state(self, return_rgb=False):
self.plot_grid(add_start=False)
# Add the agent location
plt.text(
self._state[1],
self._state[0],
u'😃',
# fontname='symbola',
fontsize=18,
ha='center',
va='center',
)
if return_rgb:
fig = plt.gcf()
plt.axis('tight')
plt.subplots_adjust(0, 0, 1, 1, 0, 0)
fig.canvas.draw()
data = np.fromstring(fig.canvas.tostring_rgb(), dtype=np.uint8, sep='')
w, h = fig.canvas.get_width_height()
data = data.reshape((h, w, 3))
plt.close(fig)
return data
def plot_policy(self, policy):
action_names = [
r'$\uparrow$', r'$\rightarrow$', r'$\downarrow$', r'$\leftarrow$'
]
self.plot_grid()
plt.title('Policy Visualization')
h, w = self._layout.shape
for y in range(h):
for x in range(w):
# if ((y, x) != self._start_state) and ((y, x) != self._goal_state):
if (y, x) != self._goal_state:
action_name = action_names[policy[y, x]]
plt.text(x, y, action_name, ha='center', va='center')
def plot_greedy_policy(self, q):
greedy_actions = np.argmax(q, axis=2)
self.plot_policy(greedy_actions)
def build_gridworld_task(task,
discount=0.9,
penalty_for_walls=-5,
observation_type=ObservationType.STATE_INDEX,
max_episode_length=200):
"""Construct a particular Gridworld layout with start/goal states.
Args:
task: string name of the task to use. One of {'simple', 'obstacle',
'random_goal'}.
discount: Discounting factor included in all Timesteps.
penalty_for_walls: Reward added when hitting a wall (should be negative).
observation_type: Enum observation type to use. One of:
* ObservationType.STATE_INDEX: int32 index of agent occupied tile.
* ObservationType.AGENT_ONEHOT: NxN float32 grid, with a 1 where the
agent is and 0 elsewhere.
* ObservationType.GRID: NxNx3 float32 grid of feature channels.
First channel contains walls (1 if wall, 0 otherwise), second the
agent position (1 if agent, 0 otherwise) and third goal position
(1 if goal, 0 otherwise)
* ObservationType.AGENT_GOAL_POS: float32 tuple with
(agent_y, agent_x, goal_y, goal_x).
max_episode_length: If set, will terminate an episode after this many
steps.
"""
tasks_specifications = {
'simple': {
'layout': [
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
],
'start_state': (2, 2),
'goal_state': (7, 2)
},
'obstacle': {
'layout': [
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
[-1, 0, 0, 0, 0, 0, -1, 0, 0, -1],
[-1, 0, 0, 0, -1, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
],
'start_state': (2, 2),
'goal_state': (2, 8)
},
'random_goal': {
'layout': [
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, -1, -1, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, 0, 0, 0, 0, 0, 0, 0, 0, -1],
[-1, -1, -1, -1, -1, -1, -1, -1, -1, -1],
],
'start_state': (2, 2),
# 'randomize_goals': True
},
}
return GridWorld(
discount=discount,
penalty_for_walls=penalty_for_walls,
observation_type=observation_type,
max_episode_length=max_episode_length,
**tasks_specifications[task])
def setup_environment(environment):
"""Returns the environment and its spec."""
# Make sure the environment outputs single-precision floats.
environment = wrappers.SinglePrecisionWrapper(environment)
# Grab the spec of the environment.
environment_spec = specs.make_environment_spec(environment)
return environment, environment_spec
# -
#
# We will use two distinct tabular GridWorlds:
# * `simple` where the goal is at the bottom left of the grid, little navigation required.
# * `obstacle` where the goal is behind an obstacle the agent must avoid.
#
# You can visualize the grid worlds by running the cell below.
#
# Note that **S** indicates the start state and **G** indicates the goal.
#
# +
# Visualise GridWorlds
# Instantiate two tabular environments, a simple task, and one that involves
# the avoidance of an obstacle.
simple_grid = build_gridworld_task(
task='simple', observation_type=ObservationType.GRID)
obstacle_grid = build_gridworld_task(
task='obstacle', observation_type=ObservationType.GRID)
# Plot them.
simple_grid.plot_grid()
plt.title('Simple')
obstacle_grid.plot_grid()
plt.title('Obstacle')
# -
# In this environment, the agent has four possible <font color='blue'>**actions**</font>: `up`, `right`, `down`, and `left`. The <font color='green'>**reward**</font> is `-5` for bumping into a wall, `+10` for reaching the goal, and `0` otherwise. The episode ends when the agent reaches the goal, and otherwise continues. The **discount** on continuing steps, is $\gamma = 0.9$.
#
# Before we start building an agent to interact with this environment, let's first look at the types of objects the environment either returns (e.g., <font color='redorange'>**observations**</font>) or consumes (e.g., <font color='blue'>**actions**</font>). The `environment_spec` will show you the form of the <font color='redorange'>**observations**</font>, <font color='green'>**rewards**</font> and **discounts** that the environment exposes and the form of the <font color='blue'>**actions**</font> that can be taken.
#
# + cellView="form"
# @title Look at environment_spec { form-width: "30%" }
# Note: setup_environment is implemented in the same cell as GridWorld.
environment, environment_spec = setup_environment(simple_grid)
print('actions:\n', environment_spec.actions, '\n')
print('observations:\n', environment_spec.observations, '\n')
print('rewards:\n', environment_spec.rewards, '\n')
print('discounts:\n', environment_spec.discounts, '\n')
# -
#
# We first set the environment to its initial state by calling the `reset()` method which returns the first observation and resets the agent to the starting location.
#
environment.reset()
environment.plot_state()
# Now we want to take an action to interact with the environment. We do this by passing a valid action to the `dm_env.Environment.step()` method which returns a `dm_env.TimeStep` namedtuple with fields `(step_type, reward, discount, observation)`.
#
# Let's take an action and visualise the resulting state of the grid-world. (You'll need to rerun the cell if you pick a new action.)
# **Note for kaggle users:** As kaggle does not render the forms automatically the students should be careful to notice the various instructions and manually play around with the values for the variables
# + cellView="form"
# @title Pick an action and see the state changing
action = "left" #@param ["up", "right", "down", "left"] {type:"string"}
action_int = {'up': 0,
'right': 1,
'down': 2,
'left':3 }
action = int(action_int[action])
timestep = environment.step(action) # pytype: dm_env.TimeStep
environment.plot_state()
# + cellView="form"
# @title Run loop { form-width: "30%" }
# @markdown This function runs an agent in the environment for a number of
# @markdown episodes, allowing it to learn.
# @markdown *Double-click* to inspect the `run_loop` function.
def run_loop(environment,
agent,
num_episodes=None,
num_steps=None,
logger_time_delta=1.,
label='training_loop',
log_loss=False,
):
"""Perform the run loop.
We are following the Acme run loop.
Run the environment loop for `num_episodes` episodes. Each episode is itself
a loop which interacts first with the environment to get an observation and
then give that observation to the agent in order to retrieve an action. Upon
termination of an episode a new episode will be started. If the number of
episodes is not given then this will interact with the environment
infinitely.
Args:
environment: dm_env used to generate trajectories.
agent: acme.Actor for selecting actions in the run loop.
num_steps: number of steps to run the loop for. If `None` (default), runs
without limit.
num_episodes: number of episodes to run the loop for. If `None` (default),
runs without limit.
logger_time_delta: time interval (in seconds) between consecutive logging
steps.
label: optional label used at logging steps.
"""
logger = loggers.TerminalLogger(label=label, time_delta=logger_time_delta)
iterator = range(num_episodes) if num_episodes else itertools.count()
all_returns = []
num_total_steps = 0
for episode in iterator:
# Reset any counts and start the environment.
start_time = time.time()
episode_steps = 0
episode_return = 0
episode_loss = 0
timestep = environment.reset()
# Make the first observation.
agent.observe_first(timestep)
# Run an episode.
while not timestep.last():
# Generate an action from the agent's policy and step the environment.
action = agent.select_action(timestep.observation)
timestep = environment.step(action)
# Have the agent observe the timestep and let the agent update itself.
agent.observe(action, next_timestep=timestep)
agent.update()
# Book-keeping.
episode_steps += 1
num_total_steps += 1
episode_return += timestep.reward
if log_loss:
episode_loss += agent.last_loss
if num_steps is not None and num_total_steps >= num_steps:
break
# Collect the results and combine with counts.
steps_per_second = episode_steps / (time.time() - start_time)
result = {
'episode': episode,
'episode_length': episode_steps,
'episode_return': episode_return,
}
if log_loss:
result['loss_avg'] = episode_loss/episode_steps
all_returns.append(episode_return)
# Log the given results.
logger.write(result)
if num_steps is not None and num_total_steps >= num_steps:
break
return all_returns
# + cellView="form"
# @title Implement the evaluation loop { form-width: "30%" }
# @markdown This function runs the agent in the environment for a number of
# @markdown episodes, without allowing it to learn, in order to evaluate it.
# @markdown *Double-click* to inspect the `evaluate` function.
def evaluate(environment: dm_env.Environment,
agent: acme.Actor,
evaluation_episodes: int):
frames = []
for episode in range(evaluation_episodes):
timestep = environment.reset()
episode_return = 0
steps = 0
while not timestep.last():
frames.append(environment.plot_state(return_rgb=True))
action = agent.select_action(timestep.observation)
timestep = environment.step(action)
steps += 1
episode_return += timestep.reward
print(
f'Episode {episode} ended with reward {episode_return} in {steps} steps'
)
return frames
def display_video(frames: Sequence[np.ndarray],
filename: str = 'temp.mp4',
frame_rate: int = 12):
"""Save and display video."""
# Write the frames to a video.
with imageio.get_writer(filename, fps=frame_rate) as video:
for frame in frames:
video.append_data(frame)
# Read video and display the video.
video = open(filename, 'rb').read()
b64_video = base64.b64encode(video)
video_tag = ('<video width="320" height="240" controls alt="test" '
'src="data:video/mp4;base64,{0}">').format(b64_video.decode())
return IPython.display.HTML(video_tag)
# -
# ## Section 2.2: The Agent
#
# We will be implementing Tabular & Function Approximation agents. Tabular agents are purely in Python.
#
# All agents will share the same interface from the Acme `Actor`. Here we borrow a figure from Acme to show how this interaction occurs:
#
# ### Agent interface
#
#
# <center><img src="https://drive.google.com/uc?id=1T7FTpA9RgDYFkciDFZK4brNyURZN_ZGp" width="500" /></center>
#
# Each agent implements the following functions:
#
# ```python
# class Agent(acme.Actor):
# def __init__(self, number_of_actions, number_of_states, ...):
# """Provides the agent the number of actions and number of states."""
#
# def select_action(self, observation):
# """Generates actions from observations."""
#
# def observe_first(self, timestep):
# """Records the initial timestep in a trajectory."""
#
# def observe(self, action, next_timestep):
# """Records the transition which occurred from taking an action."""
#
# def update(self):
# """Updates the agent's internals to potentially change its behavior."""
# ```
#
# Remarks on the `observe()` function:
#
# 1. In the last method, the `next_timestep` provides the `reward`, `discount`, and `observation` that resulted from selecting `action`.
#
# 2. The `next_timestep.step_type` will be either `MID` or `LAST` and should be used to determine whether this is the last observation in the episode.
#
# 3. The `next_timestep.step_type` cannot be `FIRST`; such a timestep should only ever be given to `observe_first()`.
#
# ### Coding Exercise 2.1: Random Agent
#
# Below is a partially complete implemention of an agent that follows a random (non-learning) policy. Fill in the ```select_action``` method.
#
# The ```select_action``` method should return a random **integer** between 0 and ```self._num_actions``` (not a tensor or an array!)
class RandomAgent(acme.Actor):
def __init__(self, environment_spec):
"""Gets the number of available actions from the environment spec."""
self._num_actions = environment_spec.actions.num_values
def select_action(self, observation):
"""Selects an action uniformly at random."""
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete the select action method")
#################################################
# TODO return a random integer beween 0 and self._num_actions.
# HINT: see the reference for how to sample a random integer in numpy:
# https://numpy.org/doc/1.16/reference/routines.random.html
action = ...
return action
def observe_first(self, timestep):
"""Does not record as the RandomAgent has no use for data."""
pass
def observe(self, action, next_timestep):
"""Does not record as the RandomAgent has no use for data."""
pass
def update(self):
"""Does not update as the RandomAgent does not learn from data."""
pass
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_7eaa84d6.py)
#
#
# + cellView="form"
# @title Visualisation of a random agent in GridWorld { form-width: "30%" }
# Create the agent by giving it the action space specification.
agent = RandomAgent(environment_spec)
# Run the agent in the evaluation loop, which returns the frames.
frames = evaluate(environment, agent, evaluation_episodes=1)
# Visualize the random agent's episode.
display_video(frames)
# -
# ---
# # Section 3: The Bellman Equation
# + cellView="form"
# @title Video 3: The Bellman Equation
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Lv411E7CB", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"cLCoNBmYUns", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
#
# In this tutorial we focus mainly on <font color='green'>**value based methods**</font>, where agents maintain a value for all state-action pairs and use those estimates to choose actions that maximize that <font color='green'>**value**</font> (instead of maintaining a policy directly, like in <font color='blue'>**policy gradient methods**</font>).
#
# We represent the <font color='green'>**action-value function**</font> (otherwise known as $\color{green}Q$-function associated with following/employing a policy $\pi$ in a given MDP as:
#
# \begin{equation}
# \color{green}Q^{\color{blue}{\pi}}(\color{red}{s},\color{blue}{a}) = \mathbb{E}_{\tau \sim P^{\color{blue}{\pi}}} \left[ \sum_t \gamma^t \color{green}{r_t}| s_0=\color{red}s,a_0=\color{blue}{a} \right]
# \end{equation}
#
# where $\tau = \{\color{red}{s_0}, \color{blue}{a_0}, \color{green}{r_0}, \color{red}{s_1}, \color{blue}{a_1}, \color{green}{r_1}, \cdots \}$
#
#
# Recall that efficient value estimations are based on the famous **_Bellman Expectation Equation_**:
#
# \begin{equation}
# \color{green}Q^\color{blue}{\pi}(\color{red}{s},\color{blue}{a}) = \sum_{\color{red}{s'}\in \color{red}{\mathcal{S}}}
# \color{purple}P(\color{red}{s'} |\color{red}{s},\color{blue}{a})
# \left(
# \color{green}{R}(\color{red}{s},\color{blue}{a}, \color{red}{s'})
# + \gamma \color{green}V^\color{blue}{\pi}(\color{red}{s'})
# \right)
# \end{equation}
#
# where $\color{green}V^\color{blue}{\pi}$ is the expected $\color{green}Q^\color{blue}{\pi}$ value for a particular state, i.e. $\color{green}V^\color{blue}{\pi}(\color{red}{s}) = \sum_{\color{blue}{a} \in \color{blue}{\mathcal{A}}} \color{blue}{\pi}(\color{blue}{a} |\color{red}{s}) \color{green}Q^\color{blue}{\pi}(\color{red}{s},\color{blue}{a})$.
#
# ---
# # Section 4: Policy Evaluation
# + cellView="form"
# @title Video 4: Policy Evaluation
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV15f4y157zA", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"HAxR4SuaZs4", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ### Lecture footnotes:
#
# **Episodic vs non-episodic environments:** Up until now, we've mainly been talking about episodic environments, or environments that terminate and reset (resampled) after a finite number of steps. However, there are also *non-episodic* environments, in which an agent cannot count on the environment resetting. Thus, they are forced to learn in a *continual* fashion.
#
# **Policy iteration vs value iteration:** Compare the two equations below, noting that the only difference is that in value iteration, the second sum is replaced by a max.
#
# *Policy iteration (using Bellman expectation equation)*
# \begin{equation}
# \color{green}Q_\color{green}{k}(\color{red}{s},\color{blue}{a}) \leftarrow \color{green}{R}(\color{red}{s},\color{blue}{a}) +\gamma \sum_{\color{red}{s'}\in \color{red}{\mathcal{S}}}
# \color{purple}P(\color{red}{s'} |\color{red}{s},\color{blue}{a})
# \sum_{\color{blue}{a'} \in \color{blue}{\mathcal{A}}} \color{blue}{\pi_{k-1}}(\color{blue}{a'} |\color{red}{s'}) \color{green}{Q_{k-1}}(\color{red}{s'},\color{blue}{a'})
# \end{equation}
#
# *Value iteration (using Bellman optimality equation)*
# \begin{equation}
# \color{green}Q_\color{green}{k}(\color{red}{s},\color{blue}{a}) \leftarrow \color{green}{R}(\color{red}{s},\color{blue}{a}) +\gamma \sum_{\color{red}{s'}\in \color{red}{\mathcal{S}}}
# \color{purple}P(\color{red}{s'} |\color{red}{s},\color{blue}{a})
# \max_{\color{blue}{a'}} \color{green}{Q_{k-1}}(\color{red}{s'},\color{blue}{a'})
# \end{equation}
# ### Coding Exercise 4.1 Policy Evaluation Agent
#
# Tabular agents implement a function `q_values()` returning a matrix of Q values
# of shape: (`number_of_states`, `number_of_actions`)
#
# In this section, we will implement a `PolicyEvalAgent` as an ACME actor: given an `evaluation_policy` $\pi_e$ and a `behaviour_policy` $\pi_b$, it will use the `behaviour_policy` to choose actions, and it will use the corresponding trajectory data to evaluate the `evaluation_policy` (i.e. compute the Q-values as if you were following the `evaluation_policy`).
#
# Algorithm:
#
# **Initialize** $Q(\color{red}{s}, \color{blue}{a})$ for all $\color{red}{s}$ ∈ $\mathcal{\color{red}S}$ and $\color{blue}a$ ∈ $\mathcal{\color{blue}A}(\color{red}s)$
#
# **Loop forever**:
#
# 1. $\color{red}{s} \gets{}$current (nonterminal) state
#
# 2. $\color{blue}{a} \gets{} \text{behaviour_policy }\pi_b(\color{red}s)$
#
# 3. Take action $\color{blue}{a}$; observe resulting reward $\color{green}{r}$, discount $\gamma$, and state, $\color{red}{s'}$
#
# 4. Compute TD-error: $\delta = \color{green}R + \gamma Q(\color{red}{s'}, \underbrace{\pi_e(\color{red}{s'}}_{\color{blue}{a'}})) − Q(\color{red}s, \color{blue}a)$
#
# 4. Update Q-value with a small $\alpha$ step: $Q(\color{red}s, \color{blue}a) \gets Q(\color{red}s, \color{blue}a) + \alpha \delta$
#
#
# We will use a uniform `random policy` as our `evaluation policy` here, but you could replace this with any policy you want, such as a greedy one.
# Uniform random policy
def random_policy(q):
return np.random.randint(4)
class PolicyEvalAgent(acme.Actor):
def __init__(self, environment_spec, evaluated_policy,
behaviour_policy=random_policy, step_size=0.1):
self._state = None
# Get number of states and actions from the environment spec.
self._number_of_states = environment_spec.observations.num_values
self._number_of_actions = environment_spec.actions.num_values
self._step_size = step_size
self._behaviour_policy = behaviour_policy
self._evaluated_policy = evaluated_policy
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Initialize your Q-values!")
#################################################
# TODO Initialize the Q-values to be all zeros.
# (Note: can also be random, but we use zeros here for reproducibility)
# HINT: This is a table of state and action pairs, so needs to be a 2-D
# array. See the reference for how to create this in numpy:
# https://numpy.org/doc/stable/reference/generated/numpy.zeros.html
self._q = ...
self._action = None
self._next_state = None
@property
def q_values(self):
# return the Q values
return self._q
def select_action(self, observation):
# Select an action
return self._behaviour_policy(self._q[observation])
def observe_first(self, timestep):
self._state = timestep.observation
def observe(self, action, next_timestep):
s = self._state
a = action
r = next_timestep.reward
g = next_timestep.discount
next_s = next_timestep.observation
# Compute TD-Error.
self._action = a
self._next_state = next_s
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Need to select the next action")
#################################################
# TODO Select the next action from the evaluation policy
# HINT: Refer to step 4 of the algorithm above.
next_a = ...
self._td_error = r + g * self._q[next_s, next_a] - self._q[s, a]
def update(self):
# Updates
s = self._state
a = self._action
# Q-value table update.
self._q[s, a] += self._step_size * self._td_error
# Update the state
self._state = self._next_state
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_7b3f830c.py)
#
#
# + cellView="form"
# @title Perform policy evaluation { form-width: "30%" }
# @markdown Here you can visualize the state value and action-value functions for the "simple" task.
num_steps = 1e3
# Create the environment
grid = build_gridworld_task(task='simple')
environment, environment_spec = setup_environment(grid)
# Create the policy evaluation agent to evaluate a random policy.
agent = PolicyEvalAgent(environment_spec, evaluated_policy=random_policy)
# run experiment and get the value functions from agent
returns = run_loop(environment=environment, agent=agent, num_steps=int(num_steps))
# get the q-values
q = agent.q_values.reshape(grid._layout.shape + (4, ))
# visualize value functions
print('AFTER {} STEPS ...'.format(num_steps))
plot_action_values(q, epsilon=1.)
# -
# ---
# # Section 5: Tabular Value-Based Model-Free Learning
# + cellView="form"
# @title Video 5: Model-Free Learning
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1iU4y1n7M6", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"Y4TweUYnexU", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ### Lecture footnotes:
#
# **On-policy (SARSA) vs off-policy (Q-learning) TD control:** Compare the two equations below and see that the only difference is that for Q-learning, the update is performed assuming that a greedy policy is followed, which is not the one used to collect the data, hence the name *off-policy*.
#
# *SARSA*
# \begin{equation}
# \color{green}Q(\color{red}{s},\color{blue}{a}) \leftarrow \color{green}Q(\color{red}{s},\color{blue}{a}) +\alpha(\color{green}{r} + \gamma\color{green}{Q}(\color{red}{s'},\color{blue}{a'}) - \color{green}{Q}(\color{red}{s},\color{blue}{a}))
# \end{equation}
#
# *Q-learning*
# \begin{equation}
# \color{green}Q(\color{red}{s},\color{blue}{a}) \leftarrow \color{green}Q(\color{red}{s},\color{blue}{a}) +\alpha(\color{green}{r} + \gamma\max_{\color{blue}{a'}} \color{green}{Q}(\color{red}{s'},\color{blue}{a'}) - \color{green}{Q}(\color{red}{s},\color{blue}{a}))
# \end{equation}
# ## Section 5.1: On-policy control: SARSA Agent
# In this section, we are focusing on control RL algorithms, which perform the **evaluation** and **improvement** of the policy synchronously. That is, the policy that is being evaluated improves as the agent is using it to interact with the environent.
#
#
# The first algorithm we are going to be looking at is SARSA. This is an **on-policy algorithm** -- i.e: the data collection is done by leveraging the policy we're trying to optimize.
#
# As discussed during lectures, a greedy policy with respect to a given $\color{Green}Q$ fails to explore the environment as needed; we will use instead an $\epsilon$-greedy policy with respect to $\color{Green}Q$.
#
# ### SARSA Algorithm
#
# **Input:**
# - $\epsilon \in (0, 1)$ the probability of taking a random action , and
# - $\alpha > 0$ the step size, also known as learning rate.
#
# **Initialize:** $\color{green}Q(\color{red}{s}, \color{blue}{a})$ for all $\color{red}{s}$ ∈ $\mathcal{\color{red}S}$ and $\color{blue}a$ ∈ $\mathcal{\color{blue}A}$
#
# **Loop forever:**
#
# 1. Get $\color{red}s \gets{}$current (non-terminal) state
#
# 2. Select $\color{blue}a \gets{} \text{epsilon_greedy}(\color{green}Q(\color{red}s, \cdot))$
#
# 3. Step in the environment by passing the selected action $\color{blue}a$
#
# 4. Observe resulting reward $\color{green}r$, discount $\gamma$, and state $\color{red}{s'}$
#
# 5. Compute TD error: $\Delta \color{green}Q \gets
# \color{green}r + \gamma \color{green}Q(\color{red}{s'}, \color{blue}{a'}) − \color{green}Q(\color{red}s, \color{blue}a)$, <br> where $\color{blue}{a'} \gets \text{epsilon_greedy}(\color{green}Q(\color{red}{s'}, \cdot))$
#
# 5. Update $\color{green}Q(\color{red}s, \color{blue}a) \gets \color{green}Q(\color{red}s, \color{blue}a) + \alpha \Delta \color{green}Q$
#
# ### Coding Exercise 5.1: Implement $\epsilon$-greedy
# Below you will find incomplete code for sampling from an $\epsilon$-greedy policy, to be used later when we implement an agent that learns values according to the SARSA algorithm.
#
#
#
def epsilon_greedy(
q_values_at_s: np.ndarray, # Q-values in state s: Q(s, a).
epsilon: float = 0.1 # Probability of taking a random action.
):
"""Return an epsilon-greedy action sample."""
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete epsilon greedy policy function")
#################################################
# TODO generate a uniform random number and compare it to epsilon to decide if
# the action should be greedy or not
# HINT: Use np.random.random() to generate a random float from 0 to 1.
if ...:
#TODO Greedy: Pick action with the largest Q-value.
action = ...
else:
# Get the number of actions from the size of the given vector of Q-values.
num_actions = np.array(q_values_at_s).shape[-1]
# TODO else return a random action
# HINT: Use np.random.randint() to generate a random integer.
action = ...
return action
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_524ce08f.py)
#
#
# + cellView="form"
# @title Sample action from $\epsilon$-greedy { form-width: "30%" }
# @markdown With $\epsilon=0.5$, you should see that about half the time, you will get back the optimal
# @markdown action 3, but half the time, it will be random.
# Create fake q-values
q_values = np.array([0, 0, 0, 1])
# Set epsilon = 0.5
epsilon = 0.5
action = epsilon_greedy(q_values, epsilon=epsilon)
print(action)
# -
# ### Coding Exercise 5.2: Run your SARSA agent on the `obstacle` environment
#
# This environment is similar to the Cliff-walking example from [Sutton & Barto](http://incompleteideas.net/book/RLbook2018.pdf) and allows us to see the different policies learned by on-policy vs off-policy methods. Try varying the number of steps.
class SarsaAgent(acme.Actor):
def __init__(self,
environment_spec: specs.EnvironmentSpec,
epsilon: float,
step_size: float = 0.1
):
# Get number of states and actions from the environment spec.
self._num_states = environment_spec.observations.num_values
self._num_actions = environment_spec.actions.num_values
# Create the table of Q-values, all initialized at zero.
self._q = np.zeros((self._num_states, self._num_actions))
# Store algorithm hyper-parameters.
self._step_size = step_size
self._epsilon = epsilon
# Containers you may find useful.
self._state = None
self._action = None
self._next_state = None
@property
def q_values(self):
return self._q
def select_action(self, observation):
return epsilon_greedy(self._q[observation], self._epsilon)
def observe_first(self, timestep):
# Set current state.
self._state = timestep.observation
def observe(self, action, next_timestep):
# Unpacking the timestep to lighten notation.
s = self._state
a = action
r = next_timestep.reward
g = next_timestep.discount
next_s = next_timestep.observation
# Compute the action that would be taken from the next state.
next_a = self.select_action(next_s)
# Compute the on-policy Q-value update.
self._action = a
self._next_state = next_s
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete the on-policy Q-value update")
#################################################
# TODO complete the line below to compute the temporal difference error
# HINT: see step 5 in the pseudocode above.
self._td_error = ...
def update(self):
# Optional unpacking to lighten notation.
s = self._state
a = self._action
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete value update")
#################################################
# Update the Q-value table value at (s, a).
# TODO: Update the Q-value table value at (s, a).
# HINT: see step 6 in the pseudocode above, remember that alpha = step_size!
self._q[s, a] += ...
# Update the current state.
self._state = self._next_state
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_4f341a18.py)
#
#
# + cellView="form"
# @title Run SARSA agent and visualize value function
num_steps = 1e5 # @param {type:"number"}
num_steps = int(num_steps)
# Create the environment.
grid = build_gridworld_task(task='obstacle')
environment, environment_spec = setup_environment(grid)
# Create the agent.
agent = SarsaAgent(environment_spec, epsilon=0.1, step_size=0.1)
# Run the experiment and get the value functions from agent
returns = run_loop(environment=environment, agent=agent, num_steps=num_steps)
print('AFTER {0:,} STEPS ...'.format(num_steps))
# Get the Q-values and reshape them to recover grid-like structure of states.
q_values = agent.q_values
grid_shape = grid.layout.shape
q_values = q_values.reshape([*grid_shape, -1])
# Visualize the value and Q-value tables.
plot_action_values(q_values, epsilon=1.)
# Visualize the greedy policy.
environment.plot_greedy_policy(q_values)
# -
# ##Section 5.2 Off-policy control: Q-learning Agent
# Reminder: $\color{green}Q$-learning is a very powerful and general algorithm, that enables control (figuring out the optimal policy/value function) both on and off-policy.
#
# **Initialize** $\color{green}Q(\color{red}{s}, \color{blue}{a})$ for all $\color{red}{s} \in \color{red}{\mathcal{S}}$ and $\color{blue}{a} \in \color{blue}{\mathcal{A}}$
#
# **Loop forever**:
#
# 1. Get $\color{red}{s} \gets{}$current (non-terminal) state
#
# 2. Select $\color{blue}{a} \gets{} \text{behaviour_policy}(\color{red}{s})$
#
# 3. Step in the environment by passing the selected action $\color{blue}{a}$
#
# 4. Observe resulting reward $\color{green}{r}$, discount $\gamma$, and state, $\color{red}{s'}$
#
# 5. Compute the TD error: $\Delta \color{green}Q \gets \color{green}{r} + \gamma \color{green}Q(\color{red}{s'}, \color{blue}{a'}) − \color{green}Q(\color{red}{s}, \color{blue}{a})$, <br>
# where $\color{blue}{a'} \gets \arg\max_{\color{blue}{\mathcal A}} \color{green}Q(\color{red}{s'}, \cdot)$
#
# 6. Update $\color{green}Q(\color{red}{s}, \color{blue}{a}) \gets \color{green}Q(\color{red}{s}, \color{blue}{a}) + \alpha \Delta \color{green}Q$
#
# Notice that the actions $\color{blue}{a}$ and $\color{blue}{a'}$ are not selected using the same policy, hence this algorithm being **off-policy**.
# ### Coding Exercise 5.3: Implement Q-Learning
#
# +
QValues = np.ndarray
Action = int
# A value-based policy takes the Q-values at a state and returns an action.
ValueBasedPolicy = Callable[[QValues], Action]
class QLearningAgent(acme.Actor):
def __init__(self,
environment_spec: specs.EnvironmentSpec,
behaviour_policy: ValueBasedPolicy,
step_size: float = 0.1):
# Get number of states and actions from the environment spec.
self._num_states = environment_spec.observations.num_values
self._num_actions = environment_spec.actions.num_values
# Create the table of Q-values, all initialized at zero.
self._q = np.zeros((self._num_states, self._num_actions))
# Store algorithm hyper-parameters.
self._step_size = step_size
# Store behavior policy.
self._behaviour_policy = behaviour_policy
# Containers you may find useful.
self._state = None
self._action = None
self._next_state = None
@property
def q_values(self):
return self._q
def select_action(self, observation):
return self._behaviour_policy(self._q[observation])
def observe_first(self, timestep):
# Set current state.
self._state = timestep.observation
def observe(self, action, next_timestep):
# Unpacking the timestep to lighten notation.
s = self._state
a = action
r = next_timestep.reward
g = next_timestep.discount
next_s = next_timestep.observation
# Compute the TD error.
self._action = a
self._next_state = next_s
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete the off-policy Q-value update")
#################################################
# TODO complete the line below to compute the temporal difference error
# HINT: This is very similar to what we did for SARSA, except keep in mind
# that we're now taking a max over the q-values (see lecture footnotes above).
# You will find the function np.max() useful.
self._td_error = ...
def update(self):
# Optional unpacking to lighten notation.
s = self._state
a = self._action
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete value update")
#################################################
# Update the Q-value table value at (s, a).
# TODO: Update the Q-value table value at (s, a).
# HINT: see step 6 in the pseudocode above, remember that alpha = step_size!
self._q[...] += ...
# Update the current state.
self._state = self._next_state
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_0f0ff9d8.py)
#
#
# -
# ### Run your Q-learning agent on the `obstacle` environment
#
# + cellView="form"
# @title Run your Q-learning
epsilon = 1. # @param {type:"number"}
num_steps = 1e5 # @param {type:"number"}
num_steps = int(num_steps)
# environment
grid = build_gridworld_task(task='obstacle')
environment, environment_spec = setup_environment(grid)
# behavior policy
behavior_policy = lambda qval: epsilon_greedy(qval, epsilon=epsilon)
# agent
agent = QLearningAgent(environment_spec, behavior_policy, step_size=0.1)
# run experiment and get the value functions from agent
returns = run_loop(environment=environment, agent=agent, num_steps=num_steps)
# get the q-values
q = agent.q_values.reshape(grid.layout.shape + (4,))
# visualize value functions
print('AFTER {:,} STEPS ...'.format(num_steps))
plot_action_values(q, epsilon=0)
# visualise the greedy policy
grid.plot_greedy_policy(q)
plt.show()
# -
# ### Experiment with different levels of greediness
#
# * The default was $\epsilon=1.$, what does this correspond to?
# * Try also $\epsilon =0.1, 0.5$. What do you observe? Does the behaviour policy affect the training in any way?
# + cellView="form"
# @title Run the cell
epsilon = 0.1 # @param {type:"number"}
num_steps = 1e5 # @param {type:"number"}
num_steps = int(num_steps)
# environment
grid = build_gridworld_task(task='obstacle')
environment, environment_spec = setup_environment(grid)
# behavior policy
behavior_policy = lambda qval: epsilon_greedy(qval, epsilon=epsilon)
# agent
agent = QLearningAgent(environment_spec, behavior_policy, step_size=0.1)
# run experiment and get the value functions from agent
returns = run_loop(environment=environment, agent=agent, num_steps=num_steps)
# get the q-values
q = agent.q_values.reshape(grid.layout.shape + (4,))
# visualize value functions
print('AFTER {:,} STEPS ...'.format(num_steps))
plot_action_values(q, epsilon=epsilon)
# visualise the greedy policy
grid.plot_greedy_policy(q)
plt.show()
# -
# ---
# # Section 6: Function Approximation
# + cellView="form"
# @title Video 6: Function approximation
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1sg411M7cn", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"7_MYePyYhrM", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# <center>
# <img src="https://drive.google.com/uc?id=1XIj68U3eB1bKYfIEHAcVbfwobmMYQQ4X" width="500" />
# </center>
#
# So far we only considered look-up tables for value-functions. In all previous cases every state and action pair $(\color{red}{s}, \color{blue}{a})$, had an entry in our $\color{green}Q$-table. Again, this is possible in this environment as the number of states is equal to the number of cells in the grid. But this is not scalable to situations where, say, the goal location changes or the obstacles are in different locations at every episode (consider how big the table could be in this situation?).
#
# An example (not covered in this tutorial) is ATARI from pixels, where the number of possible frames an agent can see is exponential in the number of pixels on the screen.
#
# <center><img width="200" alt="portfolio_view" src="https://miro.medium.com/max/1760/1*XyIpmXXAjbXerDzmGQL1yA.gif"></center>
#
# But what we **really** want is just to be able to *compute* the Q-value, when fed with a particular $(\color{red}{s}, \color{blue}{a})$ pair. So if we had a way to get a function to do this work instead of keeping a big table, we'd get around this problem.
#
# To address this, we can use **function approximation** as a way to generalize Q-values over some representation of the very large state space, and **train** them to output the values they should. In this section, we will explore $\color{green}Q$-learning with function approximation, which (although it has been theoretically proven to diverge for some degenerate MDPs) can yield impressive results in very large environments. In particular, we will look at [Neural Fitted Q (NFQ) Iteration](http://ml.informatik.uni-freiburg.de/former/_media/publications/rieecml05.pdf) and [Deep Q-Networks (DQN)](https://www.cs.toronto.edu/~vmnih/docs/dqn.pdf).
#
#
# ## Section 6.1 Replay Buffers
# An important property of off-policy methods like $\color{green}Q$-learning is that they involve two policies: one for exploration and one that is being optimized (via the $\color{green}Q$-function updates). This means that we can generate data from the **behavior** policy and insert that data into some form of data storage---usually referred to as **replay**.
#
# In order to optimize the $\color{green}Q$-function we can then sample data from the replay <font color='purple'>**dataset**</font> and use that data to perform an update. An illustration of this learning loop is shown below.
#
# <center><img src="https://drive.google.com/uc?id=1ivTQBHWkYi_J9vWwXFd2sSWg5f2TB5T-" width="400" /></center>
#
# In the next cell we will show how to implement a simple replay buffer. This can be as simple as a python list containing transition data. In more complicated scenarios we might want to have a more performance-tuned variant, we might have to be more concerned about how large replay is and what to do when its full, and we might want to sample from replay in different ways. But a simple python list can go a surprisingly long way.
# +
# Simple replay buffer
# Create a convenient container for the SARS tuples required by deep RL agents.
Transitions = collections.namedtuple(
'Transitions', ['state', 'action', 'reward', 'discount', 'next_state'])
class ReplayBuffer(object):
"""A simple Python replay buffer."""
def __init__(self, capacity: int = None):
self.buffer = collections.deque(maxlen=capacity)
self._prev_state = None
def add_first(self, initial_timestep: dm_env.TimeStep):
self._prev_state = initial_timestep.observation
def add(self, action: int, timestep: dm_env.TimeStep):
transition = Transitions(
state=self._prev_state,
action=action,
reward=timestep.reward,
discount=timestep.discount,
next_state=timestep.observation,
)
self.buffer.append(transition)
self._prev_state = timestep.observation
def sample(self, batch_size: int) -> Transitions:
# Sample a random batch of Transitions as a list.
batch_as_list = random.sample(self.buffer, batch_size)
# Convert the list of `batch_size` Transitions into a single Transitions
# object where each field has `batch_size` stacked fields.
return tree_utils.stack_sequence_fields(batch_as_list)
def flush(self) -> Transitions:
entire_buffer = tree_utils.stack_sequence_fields(self.buffer)
self.buffer.clear()
return entire_buffer
def is_ready(self, batch_size: int) -> bool:
return batch_size <= len(self.buffer)
# -
# ## Section 6.2: NFQ Agent
# [Neural Fitted Q Iteration](http://ml.informatik.uni-freiburg.de/former/_media/publications/rieecml05.pdf) was one of the first papers to demonstrate how to leverage recent advances in Deep Learning to approximate the Q-value by a neural network.$^1$
# In other words, the value $\color{green}Q(\color{red}{s}, \color{blue}{a})$ are approximated by the output of a neural network $\color{green}{Q_w}(\color{red}{s}, \color{blue}{a})$ for each possible action $\color{blue}{a} \in \color{blue}{\mathcal{A}}$.$^2$
#
# When introducing function approximations, and neural networks in particular, we need to have a loss to optimize. But looking back at the tabular setting above, you can see that we already have some notion of error: the **TD error**.
#
# By training our neural network to output values such that the *TD error is minimized*, we will also satisfy the Bellman Optimality Equation, which is a good sufficient condition to enforce, to obtain an optimal policy.
# Thanks to automatic differentiation, we can just write the TD error as a loss, e.g., with an $\ell^2$ loss, but others would work too:
#
# \begin{equation}
# L(\color{green}w) = \mathbb{E}\left[ \left( \color{green}{r} + \gamma \max_\color{blue}{a'} \color{green}{Q_w}(\color{red}{s'}, \color{blue}{a'}) − \color{green}{Q_w}(\color{red}{s}, \color{blue}{a}) \right)^2\right].
# \end{equation}
#
# Then we can compute the gradient with respect to the parameters of the neural network and improve our Q-value approximation incrementally.
#
# NFQ builds on $\color{green}Q$-learning, but if one were to update the Q-values online directly, the training can be unstable and very slow.
# Instead, NFQ uses a replay buffer, similar to what we see implemented above (Section 6.1), to update the Q-value in a batched setting.
#
# When it was introduced, it also was entirely off-policy using a uniformly random policy to collect data, which was prone to instability when applied to more complex environments (e.g. when the input are pixels or the tasks are longer and more complicated).
# But it is a good stepping stone to the more complex agents used today. Here, we will look at a slightly different and modernised implementation of NFQ.
#
# Below you will find an incomplete NFQ agent that takes in observations from a gridworld. Instead of receiving a tabular state, it receives an observation in the form of its (x,y) coordinates in the gridworld, and the (x,y) coordinates of the goal.
# <br />
#
# The goal of this coding exercise is to complete this agent by implementing the loss, using mean squared error.
#
# ---
#
# <sub>$^1$ If you read the NFQ paper, they use a "control" notation, where there is a "cost to minimize", instead of "rewards to maximize", so don't be surprised if signs/max/min do not correspond.</sub>
#
# <sub>$^2$ We could feed it $\color{blue}{a}$ as well and ask $Q_w$ for a single scalar value, but given we have a fixed number of actions and we usually need to take an $argmax$ over them, it's easiest to just output them all in one pass.</sub>
#
# ### Coding Exercise 6.1: Implement NFQ
# +
# Create a convenient container for the SARS tuples required by NFQ.
Transitions = collections.namedtuple(
'Transitions', ['state', 'action', 'reward', 'discount', 'next_state'])
class NeuralFittedQAgent(acme.Actor):
def __init__(self,
environment_spec: specs.EnvironmentSpec,
q_network: nn.Module,
replay_capacity: int = 100_000,
epsilon: float = 0.1,
batch_size: int = 1,
learning_rate: float = 3e-4):
# Store agent hyperparameters and network.
self._num_actions = environment_spec.actions.num_values
self._epsilon = epsilon
self._batch_size = batch_size
self._q_network = q_network
# Container for the computed loss (see run_loop implementation above).
self.last_loss = 0.0
# Create the replay buffer.
self._replay_buffer = ReplayBuffer(replay_capacity)
# Setup optimizer that will train the network to minimize the loss.
self._optimizer = torch.optim.Adam(self._q_network.parameters(),lr = learning_rate)
self._loss_fn = nn.MSELoss()
def select_action(self, observation):
# Compute Q-values.
q_values = self._q_network(torch.tensor(observation).unsqueeze(0)) # Adds batch dimension.
q_values = q_values.squeeze(0) # Removes batch dimension
# Select epsilon-greedy action.
if self._epsilon < torch.rand(1):
action = q_values.argmax(axis=-1)
else:
action = torch.randint(low=0, high=self._num_actions , size=(1,), dtype=torch.int64)
return action
def q_values(self, observation):
q_values = self._q_network(torch.tensor(observation).unsqueeze(0))
return q_values.squeeze(0).detach()
def update(self):
if not self._replay_buffer.is_ready(self._batch_size):
# If the replay buffer is not ready to sample from, do nothing.
return
# Sample a minibatch of transitions from experience replay.
transitions = self._replay_buffer.sample(self._batch_size)
# Note: each of these tensors will be of shape [batch_size, ...].
s = torch.tensor(transitions.state)
a = torch.tensor(transitions.action,dtype=torch.int64)
r = torch.tensor(transitions.reward)
d = torch.tensor(transitions.discount)
next_s = torch.tensor(transitions.next_state)
# Compute the Q-values at next states in the transitions.
with torch.no_grad():
q_next_s = self._q_network(next_s) # Shape [batch_size, num_actions].
max_q_next_s = q_next_s.max(axis=-1)[0]
# Compute the TD error and then the losses.
target_q_value = r + d * max_q_next_s
# Compute the Q-values at original state.
q_s = self._q_network(s)
# Gather the Q-value corresponding to each action in the batch.
q_s_a = q_s.gather(1, a.view(-1,1)).squeeze(0)
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete the NFQ Agent")
#################################################
# TODO Average the squared TD errors over the entire batch using
# self._loss_fn, which is defined above as nn.MSELoss()
# HINT: Take a look at the reference for nn.MSELoss here:
# https://pytorch.org/docs/stable/generated/torch.nn.MSELoss.html
# What should you put for the input and the target?
loss = ...
# Compute the gradients of the loss with respect to the q_network variables.
self._optimizer.zero_grad()
loss.backward()
# Apply the gradient update.
self._optimizer.step()
# Store the loss for logging purposes (see run_loop implementation above).
self.last_loss = loss.detach().numpy()
def observe_first(self, timestep: dm_env.TimeStep):
self._replay_buffer.add_first(timestep)
def observe(self, action: int, next_timestep: dm_env.TimeStep):
self._replay_buffer.add(action, next_timestep)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_f42d1415.py)
#
#
# -
# ### Train and Evaluate the NFQ Agent
#
# + cellView="form"
# @title Training the NFQ Agent
epsilon = 0.4 # @param {type:"number"}
max_episode_length = 200
# Create the environment.
grid = build_gridworld_task(
task='simple',
observation_type=ObservationType.AGENT_GOAL_POS,
max_episode_length=max_episode_length)
environment, environment_spec = setup_environment(grid)
# Define the neural function approximator (aka Q network).
q_network = nn.Sequential(nn.Linear(4, 50),
nn.ReLU(),
nn.Linear(50, 50),
nn.ReLU(),
nn.Linear(50, environment_spec.actions.num_values))
# Build the trainable Q-learning agent
agent = NeuralFittedQAgent(
environment_spec,
q_network,
epsilon=epsilon,
replay_capacity=100_000,
batch_size=10,
learning_rate=1e-3)
returns = run_loop(
environment=environment,
agent=agent,
num_episodes=500,
logger_time_delta=1.,
log_loss=True)
# + cellView="form"
# @title Evaluating the agent (set $\epsilon=0$)
# Temporarily change epsilon to be more greedy; remember to change it back.
agent._epsilon = 0.0
# Record a few episodes.
frames = evaluate(environment, agent, evaluation_episodes=5)
# Change epsilon back.
agent._epsilon = epsilon
# Display the video of the episodes.
display_video(frames, frame_rate=6)
# + cellView="form"
# @title Visualise the learned $Q$ values
# Evaluate the policy for every state, similar to tabular agents above.
environment.reset()
pi = np.zeros(grid._layout_dims, dtype=np.int32)
q = np.zeros(grid._layout_dims + (4, ))
for y in range(grid._layout_dims[0]):
for x in range(grid._layout_dims[1]):
# Hack observation to see what the Q-network would output at that point.
environment.set_state(x, y)
obs = environment.get_obs()
q[y, x] = np.asarray(agent.q_values(obs))
pi[y, x] = np.asarray(agent.select_action(obs))
plot_action_values(q)
# -
# Compare the Q-values approximated with the neural network with the tabular case in **Section 5.3**. Notice how the neural network is generalizing from the visited states to the unvisited similar states, while in the tabular case we updated the value of each state only when we visited that state.
# ### Compare the greedy and behaviour ($\epsilon$-greedy) policies
# + cellView="form"
# @title Compare the greedy policy with the agent's policy
# @markdown Notice that the agent's behavior policy has a lot more randomness,
# @markdown due to the high $\epsilon$. However, the greedy policy that's learned
# @markdown is optimal.
environment.plot_greedy_policy(q)
plt.figtext(-.08, .95, 'Greedy policy using the learnt Q-values')
plt.title('')
plt.show()
environment.plot_policy(pi)
plt.figtext(-.08, .95, "Policy using the agent's behavior policy")
plt.title('')
plt.show()
# -
# ---
# # Section 7: DQN
# + cellView="form"
#@title Video 7: Deep Q-Networks (DQN)
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Mo4y1Q7yD", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"HEDoNtV1y-w", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
#
# <!-- <center><img src="https://drive.google.com/uc?id=1ivTQBHWkYi_J9vWwXFd2sSWg5f2TB5T-" width="500" /></center> -->
#
# <center><img src="https://media.springernature.com/full/springer-static/image/art%3A10.1038%2Fnature14236/MediaObjects/41586_2015_Article_BFnature14236_Fig1_HTML.jpg" width="500" /></center>
#
# In this section, we will look at an advanced deep RL Agent based on the following publication, [Playing Atari with Deep Reinforcement Learning](https://deepmind.com/research/publications/playing-atari-deep-reinforcement-learning), which introduced the first deep learning model to successfully learn control policies directly from high-dimensional pixel inputs using RL.
#
# Here the agent will act directly on a pixel representation of the gridworld. You can find an incomplete implementation below.
#
#
# ### Coding Exercise 7.1: Run a DQN Agent
#
# +
class DQN(acme.Actor):
def __init__(self,
environment_spec: specs.EnvironmentSpec,
network: nn.Module,
replay_capacity: int = 100_000,
epsilon: float = 0.1,
batch_size: int = 1,
learning_rate: float = 5e-4,
target_update_frequency: int = 10):
# Store agent hyperparameters and network.
self._num_actions = environment_spec.actions.num_values
self._epsilon = epsilon
self._batch_size = batch_size
self._q_network = q_network
# create a second q net with the same structure and initial values, which
# we'll be updating separately from the learned q-network.
self._target_network = copy.deepcopy(self._q_network)
# Container for the computed loss (see run_loop implementation above).
self.last_loss = 0.0
# Create the replay buffer.
self._replay_buffer = ReplayBuffer(replay_capacity)
# Keep an internal tracker of steps
self._current_step = 0
# How often to update the target network
self._target_update_frequency = target_update_frequency
# Setup optimizer that will train the network to minimize the loss.
self._optimizer = torch.optim.Adam(self._q_network.parameters(), lr=learning_rate)
self._loss_fn = nn.MSELoss()
def select_action(self, observation):
# Compute Q-values.
# Sonnet requires a batch dimension, which we squeeze out right after.
q_values = self._q_network(torch.tensor(observation).unsqueeze(0)) # Adds batch dimension.
q_values = q_values.squeeze(0) # Removes batch dimension
# Select epsilon-greedy action.
if self._epsilon < torch.rand(1):
action = q_values.argmax(axis=-1)
else:
action = torch.randint(low=0, high=self._num_actions , size=(1,), dtype=torch.int64)
return action
def q_values(self, observation):
q_values = self._q_network(torch.tensor(observation).unsqueeze(0))
return q_values.squeeze(0).detach()
def update(self):
self._current_step += 1
if not self._replay_buffer.is_ready(self._batch_size):
# If the replay buffer is not ready to sample from, do nothing.
return
# Sample a minibatch of transitions from experience replay.
transitions = self._replay_buffer.sample(self._batch_size)
# Optionally unpack the transitions to lighten notation.
# Note: each of these tensors will be of shape [batch_size, ...].
s = torch.tensor(transitions.state)
a = torch.tensor(transitions.action,dtype=torch.int64)
r = torch.tensor(transitions.reward)
d = torch.tensor(transitions.discount)
next_s = torch.tensor(transitions.next_state)
# Compute the Q-values at next states in the transitions.
with torch.no_grad():
#################################################
# Fill in missing code below (...),
# then remove or comment the line below to test your implementation
raise NotImplementedError("Student exercise: complete the DQN Agent")
#################################################
#TODO get the value of the next states evaluated by the target network
#HINT: use self._target_network, defined above.
q_next_s = ... # Shape [batch_size, num_actions].
max_q_next_s = q_next_s.max(axis=-1)[0]
# Compute the TD error and then the losses.
target_q_value = r + d * max_q_next_s
# Compute the Q-values at original state.
q_s = self._q_network(s)
# Gather the Q-value corresponding to each action in the batch.
q_s_a = q_s.gather(1, a.view(-1,1)).squeeze(0)
# Average the squared TD errors over the entire batch
loss = self._loss_fn(target_q_value, q_s_a)
# Compute the gradients of the loss with respect to the q_network variables.
self._optimizer.zero_grad()
loss.backward()
# Apply the gradient update.
self._optimizer.step()
if self._current_step % self._target_update_frequency == 0:
self._target_network.load_state_dict(self._q_network.state_dict())
# Store the loss for logging purposes (see run_loop implementation above).
self.last_loss = loss.detach().numpy()
def observe_first(self, timestep: dm_env.TimeStep):
self._replay_buffer.add_first(timestep)
def observe(self, action: int, next_timestep: dm_env.TimeStep):
self._replay_buffer.add(action, next_timestep)
# Create a convenient container for the SARS tuples required by NFQ.
Transitions = collections.namedtuple(
'Transitions', ['state', 'action', 'reward', 'discount', 'next_state'])
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_d6d1b1d0.py)
#
#
# + cellView="form"
# @title Train and evaluate the DQN agent
epsilon = 0.25 # @param {type: "number"}
num_episodes = 1000 # @param {type: "integer"}
grid = build_gridworld_task(
task='simple',
observation_type=ObservationType.GRID,
max_episode_length=200)
environment, environment_spec = setup_environment(grid)
# Build the agent's network.
class Permute(nn.Module):
def __init__(self, order: list):
super(Permute,self).__init__()
self.order = order
def forward(self, x):
return x.permute(self.order)
q_network = nn.Sequential(Permute([0, 3, 1, 2]),
nn.Conv2d(3, 32, kernel_size=4,
stride=2,padding=1),
nn.ReLU(),
nn.Conv2d(32, 64, kernel_size=3,
stride=1, padding=1),
nn.ReLU(),
nn.MaxPool2d(3, 1),
nn.Flatten(),
nn.Linear(384, 50),
nn.ReLU(),
nn.Linear(50, environment_spec.actions.num_values)
)
agent = DQN(
environment_spec=environment_spec,
network=q_network,
batch_size=10,
epsilon=epsilon,
target_update_frequency=25)
returns = run_loop(
environment=environment,
agent=agent,
num_episodes=num_episodes,
num_steps=100000)
# + cellView="form"
# @title Visualise the learned $Q$ values
# Evaluate the policy for every state, similar to tabular agents above.
pi = np.zeros(grid._layout_dims, dtype=np.int32)
q = np.zeros(grid._layout_dims + (4,))
for y in range(grid._layout_dims[0]):
for x in range(grid._layout_dims[1]):
# Hack observation to see what the Q-network would output at that point.
environment.set_state(x, y)
obs = environment.get_obs()
q[y, x] = np.asarray(agent.q_values(obs))
pi[y, x] = np.asarray(agent.select_action(obs))
plot_action_values(q)
# + cellView="form"
# @title Compare the greedy policy with the agent's policy
environment.plot_greedy_policy(q)
plt.figtext(-.08, .95, "Greedy policy using the learnt Q-values")
plt.title('')
plt.show()
environment.plot_policy(pi)
plt.figtext(-.08, .95, "Policy using the agent's epsilon-greedy policy")
plt.title('')
plt.show()
# -
# ---
# # Section 8: Beyond Value Based Model-Free Methods
# + cellView="form"
# @title Video 8: Other RL Methods
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV14w411977Y", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"1N4Jm9loJx4", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Cartpole task
#
# Here we switch to training on a different kind of task, which has a continuous action space: Cartpole in [Gym](https://gym.openai.com/). As you recall from the video, policy-based methods are particularly well-suited for these kinds of tasks. We will be exploring two of those methods below.
#
#
# <center><img src="https://user-images.githubusercontent.com/10624937/42135683-dde5c6f0-7d13-11e8-90b1-8770df3e40cf.gif" height="250" /></center>
# + cellView="form"
# @title Make a CartPole environment, `gym.make('CartPole-v1')`
env = gym.make('CartPole-v1')
# Set seeds
env.seed(SEED)
set_seed(SEED)
# -
# ## Section 8.1: Policy gradient
#
# Now we will turn to policy gradient methods. Rather than defining the policy in terms of a value function, i.e. $\color{blue}\pi(\color{red}s) = \arg\max_{\color{blue}a}\color{green}Q(\color{red}s, \color{blue}a)$, we will directly parameterize the policy and write it as the distribution
#
# \begin{equation}
# \color{blue}a_t \sim \color{blue}\pi_{\theta}(\color{blue}a_t|\color{red}s_t).
# \end{equation}
#
# Here $\theta$ represent the parameters of the policy. We will update the policy parameters using gradient ascent to **maximize** expected future reward.
#
# One convenient way to represent the conditional distribution above is as a function that takes a state $\color{red}s$ and returns a distribution over actions $\color{blue}a$.
#
# Defined below is an agent which implements the REINFORCE algorithm.
# REINFORCE (Williams 1992) is the simplest model-free general reinforcement learning technique.
#
# The **basic idea** is to use probabilistic action choice. If the reward at the end turns out to be high, we make **all** actions in this sequence **more likely** (otherwise, we do the opposite).
#
# This strategy could reinforce "bad" actions as well, however they will turn out to be part of trajectories with low reward and will likely not get accentuated.
#
# From the lectures, we know that we need to compute
#
# \begin{equation}
# \nabla J(\theta)
# = \mathbb{E}
# \left[
# \sum_{t=0}^T \color{green} G_t
# \nabla\log\color{blue}\pi_\theta(\color{red}{s_t})
# \right]
# \end{equation}
#
# where $\color{green} G_t$ is the sum over future rewards from time $t$, defined as
#
# \begin{equation}
# \color{green} G_t
# = \sum_{n=t}^T \gamma^{n-t}
# \color{green} R(\color{red}{s_t}, \color{blue}{a_t}, \color{red}{s_{t+1}}).
# \end{equation}
#
# The algorithm below will collect the state, action, and reward data in its buffer until it reaches a full trajectory. It will then update its policy given the above gradient (and the Adam optimizer).
#
# A policy gradient trains an agent without explicitly mapping the value for every state-action pair in an environment by taking small steps and updating the policy based on the reward associated with that step. In this section, we will build a small network that trains using policy gradient using PyTorch.
#
# The agent can receive a reward immediately for an action or it can receive the award at a later time such as the end of the episode.
#
# The policy function our agent will try to learn is $\pi_\theta(a,s)$, where $\theta$ is the parameter vector, $s$ is a particular state, and $a$ is an action.
#
# Monte-Carlo Policy Gradient approach will be used, which means the agent will run through an entire episode and then update policy based on the rewards obtained.
# + cellView="form"
# @title Set the hyperparameters for Policy Gradient
num_steps = 300
learning_rate = 0.01 # @param {type:"number"}
gamma = 0.99 # @param {type:"number"}
dropout = 0.6 # @param {type:"number"}
# @markdown Only used in Policy Gradient Method:
hidden_neurons = 128 # @param {type:"integer"}
# -
# ### Coding Exercise 8.1: Creating a simple neural network
#
# Below you will find some incomplete code. Fill in the missing code to construct the specified neural network.
#
# Let us define a simple feed forward neural network with one hidden layer of 128 neurons and a dropout of 0.6. Let's use Adam as our optimizer and a learning rate of 0.01. Use the hyperparameters already defined rather than using explicit values.
#
# Using dropout will significantly improve the performance of the policy. Do compare your results with and without dropout and experiment with other hyper-parameter values as well.
class PolicyGradientNet(nn.Module):
def __init__(self):
super(PolicyGradientNet, self).__init__()
self.state_space = env.observation_space.shape[0]
self.action_space = env.action_space.n
#################################################
## TODO for students: Define two linear layers
## from the first expression
raise NotImplementedError("Student exercise: Create FF neural network.")
#################################################
# HINT: you can construct linear layers using nn.Linear(); what are the
# sizes of the inputs and outputs of each of the layers? Also remember
# that you need to use hidden_neurons (see hyperparameters section above).
# https://pytorch.org/docs/stable/generated/torch.nn.Linear.html
self.l1 = ...
self.l2 = ...
self.gamma = ...
# Episode policy and past rewards
self.past_policy = Variable(torch.Tensor())
self.reward_episode = []
# Overall reward and past loss
self.past_reward = []
self.past_loss = []
def forward(self, x):
model = torch.nn.Sequential(
self.l1,
nn.Dropout(p=dropout),
nn.ReLU(),
self.l2,
nn.Softmax(dim=-1)
)
return model(x)
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_9aaf4a83.py)
#
#
# -
# Now let's create an instance of the network we have defined and use Adam as the optimizer using the learning_rate as hyperparameter already defined above.
policy = PolicyGradientNet()
pg_optimizer = optim.Adam(policy.parameters(), lr=learning_rate)
# ### Select Action
#
# The `select_action()` function chooses an action based on our policy probability distribution using the PyTorch distributions package. Our policy returns a probability for each possible action in our action space (move left or move right) as an array of length two such as [0.7, 0.3]. We then choose an action based on these probabilities, record our history, and return our action.
def select_action(state):
#Select an action (0 or 1) by running policy model and choosing based on the probabilities in state
state = torch.from_numpy(state).type(torch.FloatTensor)
state = policy(Variable(state))
c = Categorical(state)
action = c.sample()
# Add log probability of chosen action
if policy.past_policy.dim() != 0:
policy.past_policy = torch.cat([policy.past_policy, c.log_prob(action).reshape(1)])
else:
policy.past_policy = (c.log_prob(action).reshape(1))
return action
# ### Update policy
#
# This function updates the policy.
# #### Reward $G_t$
# We update our policy by taking a sample of the action value function $Q^{\pi_\theta} (s_t,a_t)$ by playing through episodes of the game. $Q^{\pi_\theta} (s_t,a_t)$ is defined as the expected return by taking action $a$ in state $s$ following policy $\pi$.
#
# We know that for every step the simulation continues we receive a reward of 1. We can use this to calculate the policy gradient at each time step, where $r$ is the reward for a particular state-action pair. Rather than using the instantaneous reward, $r$, we instead use a long term reward $ v_{t} $ where $v_t$ is the discounted sum of all future rewards for the length of the episode. $v_{t}$ is then,
#
# \begin{equation}
# \color{green} G_t
# = \sum_{n=t}^T \gamma^{n-t}
# \color{green} R(\color{red}{s_t}, \color{blue}{a_t}, \color{red}{s_{t+1}}).
# \end{equation}
#
# where $\gamma$ is the discount factor (0.99). For example, if an episode lasts 5 steps, the reward for each step will be [4.90, 3.94, 2.97, 1.99, 1].
# Next we scale our reward vector by substracting the mean from each element and scaling to unit variance by dividing by the standard deviation. This practice is common for machine learning applications and the same operation as Scikit Learn's __[StandardScaler](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)__. It also has the effect of compensating for future uncertainty.
# #### Update Policy: equation
# After each episode we apply Monte-Carlo Policy Gradient to improve our policy according to the equation:
#
# \begin{equation}
# \Delta\theta_t = \alpha\nabla_\theta \, \log \pi_\theta (s_t,a_t)G_t
# \end{equation}
#
# We will then feed our policy history multiplied by our rewards to our optimizer and update the weights of our neural network using stochastic gradient **ascent**. This should increase the likelihood of actions that got our agent a larger reward.
# The following function ```update_policy``` updates the network weights and therefore the policy.
#
#
def update_policy():
R = 0
rewards = []
# Discount future rewards back to the present using gamma
for r in policy.reward_episode[::-1]:
R = r + policy.gamma * R
rewards.insert(0, R)
# Scale rewards
rewards = torch.FloatTensor(rewards)
rewards = (rewards - rewards.mean()) / (rewards.std() +
np.finfo(np.float32).eps)
# Calculate loss
pg_loss = (torch.sum(torch.mul(policy.past_policy,
Variable(rewards)).mul(-1), -1))
# Update network weights
# Use zero_grad(), backward() and step() methods of the optimizer instance.
pg_optimizer.zero_grad()
pg_loss.backward()
# Update the weights
for param in policy.parameters():
param.grad.data.clamp_(-1, 1)
pg_optimizer.step()
# Save and intialize episode past counters
policy.past_loss.append(pg_loss.item())
policy.past_reward.append(np.sum(policy.reward_episode))
policy.past_policy = Variable(torch.Tensor())
policy.reward_episode= []
# ### Training
# This is our main policy training loop. For each step in a training episode, we choose an action, take a step through the environment, and record the resulting new state and reward. We call update_policy() at the end of each episode to feed the episode history to our neural network and improve our policy.
def policy_gradient_train(episodes):
running_reward = 10
for episode in range(episodes):
state = env.reset()
done = False
for time in range(1000):
action = select_action(state)
# Step through environment using chosen action
state, reward, done, _ = env.step(action.item())
# Save reward
policy.reward_episode.append(reward)
if done:
break
# Used to determine when the environment is solved.
running_reward = (running_reward * gamma) + (time * (1 - gamma))
update_policy()
if episode % 50 == 0:
print(f"Episode {episode}\tLast length: {time:5.0f}"
f"\tAverage length: {running_reward:.2f}")
if running_reward > env.spec.reward_threshold:
print(f"Solved! Running reward is now {running_reward} "
f"and the last episode runs to {time} time steps!")
break
# ### Run the model
# + cellView="form"
episodes = 500 #@param {type:"integer"}
policy_gradient_train(episodes)
# -
# ### Plot the results
# + cellView="form"
# @title Plot the training performance for policy gradient
def plot_policy_gradient_training():
window = int(episodes / 20)
fig, ((ax1), (ax2)) = plt.subplots(1, 2, sharey=True, figsize=[15, 4]);
rolling_mean = pd.Series(policy.past_reward).rolling(window).mean()
std = pd.Series(policy.past_reward).rolling(window).std()
ax1.plot(rolling_mean)
ax1.fill_between(range(len(policy.past_reward)),
rolling_mean-std, rolling_mean+std,
color='orange', alpha=0.2)
ax1.set_title(f"Episode Length Moving Average ({window}-episode window)")
ax1.set_xlabel('Episode'); ax1.set_ylabel('Episode Length')
ax2.plot(policy.past_reward)
ax2.set_title('Episode Length')
ax2.set_xlabel('Episode')
ax2.set_ylabel('Episode Length')
fig.tight_layout(pad=2)
plt.show()
plot_policy_gradient_training()
# -
# ### Exercise 8.1: Explore different hyperparameters.
#
# Try running the model again, by modifying the hyperparameters and observe the outputs. Be sure to rerun the function definition cells in order to pick up on the updated values.
#
# What do you see when you
#
# 1. increase learning rate
# 2. decrease learning rate
# 3. decrease gamma ($\gamma$)
# 4. increase number of hidden neurons in the network
# ## Section 8.2: Actor-critic
#
# Recall the policy gradient
#
# \begin{equation}
# \nabla J(\theta)
# = \mathbb{E}
# \left[
# \sum_{t=0}^T \color{green} G_t
# \nabla\log\color{blue}\pi_\theta(\color{red}{s_t})
# \right]
# \end{equation}
#
# The policy parameters are updated using Monte Carlo technique and uses random samples. This introduces high variability in log probabilities and cumulative reward values. This leads to noisy gradients and can cause unstable learning.
#
# One way to reduce variance and increase stability is subtracting the cumulative reward by a baseline:
#
# \begin{equation}
# \nabla J(\theta)
# = \mathbb{E}
# \left[
# \sum_{t=0}^T \color{green} (G_t - b)
# \nabla\log\color{blue}\pi_\theta(\color{red}{s_t})
# \right]
# \end{equation}
#
# Intuitively, reducing cumulative reward will make smaller gradients and thus smaller and more stable (hopefully) updates.
#
# From the lecture slides, we know that in Actor Critic Method:
# 1. The “Critic” estimates the value function. This could be the action-value (the Q value) or state-value (the V value).
# 2. The “Actor” updates the policy distribution in the direction suggested by the Critic (such as with policy gradients).
#
# Both the Critic and Actor functions are parameterized with neural networks. The "Critic" network parameterizes the Q-value.
# + cellView="form"
# @title Set the hyperparameters for Actor Critic
learning_rate = 0.01 # @param {type:"number"}
gamma = 0.99 # @param {type:"number"}
dropout = 0.6
# Only used in Actor-Critic Method
hidden_size = 256 # @param {type:"integer"}
num_steps = 300
# -
# ### Actor Critic Network
class ActorCriticNet(nn.Module):
def __init__(self, num_inputs, num_actions, hidden_size, learning_rate=3e-4):
super(ActorCriticNet, self).__init__()
self.num_actions = num_actions
self.critic_linear1 = nn.Linear(num_inputs, hidden_size)
self.critic_linear2 = nn.Linear(hidden_size, 1)
self.actor_linear1 = nn.Linear(num_inputs, hidden_size)
self.actor_linear2 = nn.Linear(hidden_size, num_actions)
self.all_rewards = []
self.all_lengths = []
self.average_lengths = []
def forward(self, state):
state = Variable(torch.from_numpy(state).float().unsqueeze(0))
value = F.relu(self.critic_linear1(state))
value = self.critic_linear2(value)
policy_dist = F.relu(self.actor_linear1(state))
policy_dist = F.softmax(self.actor_linear2(policy_dist), dim=1)
return value, policy_dist
# ### Training
def actor_critic_train(episodes):
all_lengths = []
average_lengths = []
all_rewards = []
entropy_term = 0
for episode in range(episodes):
log_probs = []
values = []
rewards = []
state = env.reset()
for steps in range(num_steps):
value, policy_dist = actor_critic.forward(state)
value = value.detach().numpy()[0, 0]
dist = policy_dist.detach().numpy()
action = np.random.choice(num_outputs, p=np.squeeze(dist))
log_prob = torch.log(policy_dist.squeeze(0)[action])
entropy = -np.sum(np.mean(dist) * np.log(dist))
new_state, reward, done, _ = env.step(action)
rewards.append(reward)
values.append(value)
log_probs.append(log_prob)
entropy_term += entropy
state = new_state
if done or steps == num_steps - 1:
qval, _ = actor_critic.forward(new_state)
qval = qval.detach().numpy()[0, 0]
all_rewards.append(np.sum(rewards))
all_lengths.append(steps)
average_lengths.append(np.mean(all_lengths[-10:]))
if episode % 50 == 0:
print(f"episode: {episode},\treward: {np.sum(rewards)},"
f"\ttotal length: {steps},"
f"\taverage length: {average_lengths[-1]}")
break
# compute Q values
qvals = np.zeros_like(values)
for t in reversed(range(len(rewards))):
qval = rewards[t] + gamma * qval
qvals[t] = qval
#update actor critic
values = torch.FloatTensor(values)
qvals = torch.FloatTensor(qvals)
log_probs = torch.stack(log_probs)
advantage = qvals - values
actor_loss = (-log_probs * advantage).mean()
critic_loss = 0.5 * advantage.pow(2).mean()
ac_loss = actor_loss + critic_loss + 0.001 * entropy_term
ac_optimizer.zero_grad()
ac_loss.backward()
ac_optimizer.step()
# Store results
actor_critic.average_lengths = average_lengths
actor_critic.all_rewards = all_rewards
actor_critic.all_lengths = all_lengths
# ### Run the model
# + cellView="form"
episodes = 500 # @param {type:"integer"}
env.reset()
num_inputs = env.observation_space.shape[0]
num_outputs = env.action_space.n
actor_critic = ActorCriticNet(num_inputs, num_outputs, hidden_size)
ac_optimizer = optim.Adam(actor_critic.parameters())
actor_critic_train(episodes)
# -
# ### Plot the results
# + cellView="form"
# @title Plot the training performance for Actor Critic
def plot_actor_critic_training(actor_critic, episodes):
window = int(episodes / 20)
plt.figure(figsize=(15, 4))
plt.subplot(1, 2, 1)
smoothed_rewards = pd.Series(actor_critic.all_rewards).rolling(window).mean()
std = pd.Series(actor_critic.all_rewards).rolling(window).std()
plt.plot(smoothed_rewards, label='Smoothed rewards')
plt.fill_between(range(len(smoothed_rewards)),
smoothed_rewards - std, smoothed_rewards + std,
color='orange', alpha=0.2)
plt.xlabel('Episode')
plt.ylabel('Reward')
plt.subplot(1, 2, 2)
plt.plot(actor_critic.all_lengths, label='All lengths')
plt.plot(actor_critic.average_lengths, label='Average lengths')
plt.xlabel('Episode')
plt.ylabel('Episode length')
plt.legend()
plt.tight_layout()
plt.show()
plot_actor_critic_training(actor_critic, episodes)
# -
# ### Exercise 8.3: Effect of episodes on performance
#
# Change the episodes from 500 to 3000 and observe the performance impact.
# ### Exercise 8.4: Effect of learning rate on performance
#
# Modify the hyperparameters related to learning_rate and gamma and observe the impact on the performance.
#
# Be sure to rerun the function definition cells in order to pick up on the updated values.
# ---
# # Section 9: RL in the real world
# + cellView="form"
# @title Video 9: Real-world applications and ethics
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1Nq4y1X7AF", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"5kBtiW88QVw", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ## Exercise 9: Group discussion
#
# Form a group of 2-3 and have discussions (roughly 3 minutes each) of the following questions:
#
# 1. **Safety**: what are some safety issues that arise in RL that don’t arise with e.g. supervised learning?
#
# 2. **Generalization**: What happens if your RL agent is presented with data it hasn’t trained on? (“goes out of distribution”)
#
# 3. How important do you think **interpretability** is in the ethical and safe deployment of RL agents in the real world?
#
# + [markdown] colab_type="text"
# [*Click for solution*](https://github.com/NeuromatchAcademy/course-content-dl/tree/main//tutorials/W3D2_BasicReinforcementLearning/solutions/W3D2_Tutorial1_Solution_99944c89.py)
#
#
# -
# ---
# # Section 10: How to learn more
# + cellView="form"
# @title Video 10: How to learn more
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1WM4y1T7G5", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"dKaOpgor5Ek", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# -
# ---
# # Appendix and further reading
#
# Books and lecture notes
# * [Reinforcement Learning: an Introduction by Sutton & Barto](http://incompleteideas.net/book/RLbook2018.pdf)
# * [Algorithms for Reinforcement Learning by <NAME>](https://sites.ualberta.ca/~szepesva/papers/RLAlgsInMDPs.pdf)
#
# Lectures and course
# * [RL Course by <NAME>](https://www.youtube.com/playlist?list=PLzuuYNsE1EZAXYR4FJ75jcJseBmo4KQ9-)
# * [Reinforcement Learning Course | UCL & DeepMind](https://www.youtube.com/playlist?list=PLqYmG7hTraZBKeNJ-JE_eyJHZ7XgBoAyb)
# * [<NAME>skill Stanford RL Course](https://www.youtube.com/playlist?list=PLoROMvodv4rOSOPzutgyCTapiGlY2Nd8u)
# * [RL Course on Coursera by <NAME> & <NAME>](https://www.coursera.org/specializations/reinforcement-learning)
#
# More practical:
# * [Spinning Up in Deep RL by <NAME>](https://spinningup.openai.com/en/latest/)
# * [Acme white paper](https://arxiv.org/abs/2006.00979) & [Colab tutorial](https://github.com/deepmind/acme/blob/master/examples/tutorial.ipynb)
#
# <br>
#
# [Link to the tweet thread with resources recommended by the community](https://twitter.com/FeryalMP/status/1407272291579355136?s=20).
#
# <br>
#
# This Colab is based on the [EEML 2020 RL practical](https://colab.research.google.com/github/eemlcommunity/PracticalSessions2020/blob/master/rl/EEML2020_RL_Tutorial.ipynb) by <NAME> & <NAME>. If you are interested in JAX you should try the colab. If you are interested in Tensorflow, there is also a version of the colab for the [MLSS 2020 RL Tutorial](https://github.com/Feryal/rl_mlss_2020) that you can try :)
#
|
tutorials/W3D2_BasicReinforcementLearning/student/W3D2_Tutorial1.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:core_acc] *
# language: python
# name: conda-env-core_acc-py
# ---
# # Test references created
#
# Here we want to see how well _pilA_ gene aligns against PAO1+phage reference vs PA14+phage reference. This _pilA_ gene has a low sequence identity with ????
import os
import pandas as pd
import numpy as np
from Bio import SeqIO
from core_acc_modules import paths
# + magic_args="-s $paths.PAO1_PHAGE_REF $paths.PAO1_PHAGE_DB_DIR" language="bash"
#
# makeblastdb -in $1 -dbtype nucl -parse_seqids -out $2
# + magic_args="-s $paths.PA14_PHAGE_REF $paths.PA14_PHAGE_DB_DIR" language="bash"
#
# makeblastdb -in $1 -dbtype nucl -parse_seqids -out $2
# + magic_args="-s $paths.PILA_QUERY $paths.PAO1_PILA_BLAST_RESULT $paths.PAO1_PHAGE_DB_DIR" language="bash"
# blastn -query $1 -out $2 -db $3 -outfmt 6
# + magic_args="-s $paths.PILA_QUERY $paths.PA14_PILA_BLAST_RESULT $paths.PA14_PHAGE_DB_DIR" language="bash"
# blastn -query $1 -out $2 -db $3 -outfmt 6
# -
pao1_blast_result = pd.read_csv(paths.PAO1_PILA_BLAST_RESULT, sep="\t", header=None)
pa14_blast_result = pd.read_csv(paths.PA14_PILA_BLAST_RESULT, sep="\t", header=None)
# Add column names described above
col_names = [
"qseqid",
"sseqid",
"pident",
"length",
"mismatch",
"gapopen",
"qstart",
"qend",
"sstart",
"send",
"evalue",
"bitscore",
]
# BLAST results for PAO1
pao1_blast_result.columns = col_names
print(pao1_blast_result.shape)
print(pao1_blast_result["evalue"].max())
pao1_blast_result.head()
# BLAST results for PA14
pa14_blast_result.columns = col_names
print(pa14_blast_result.shape)
print(pa14_blast_result["evalue"].max())
pa14_blast_result.head()
|
archive/test_phage_reference/0_explore_references_created.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process train and test files
# ## About data
# <blockquote><i>application_train/application_test</i>: This is main training and testing data with information about each loan application at Home Credit. Every loan has its own row and is identified by the feature <i>SK_ID_CURR</i>. The training application data comes with the TARGET indicating 0: the loan was repaid or 1: the loan was not repaid. Feature descriptions are as below. These have been taken from file <i>'HomeCredit_columns_description.csv'>/i>. For a more accurate description, please refer to the file.</blockquote>
# ## Feature descriptions
# <blockquote><p style="font-size:13px">
# SK_ID_CURR: ID of loan in our sample<br>
# TARGET: Target variable (1 - client with payment difficulties: he/she had late payment more than X days on at least one of the first Y installments of the loan in our sample, 0 - all other cases)<br>
# NAME_CONTRACT_TYPE: Identification if loan is cash or revolving<br>
# CODE_GENDER: Gender of the client<br>
# FLAG_OWN_CAR: Flag if the client owns a car<br>
# FLAG_OWN_REALTY: Flag if client owns a house or flat<br>
# CNT_CHILDREN: Number of children the client has<br>
# AMT_INCOME_TOTAL: Income of the client<br>
# AMT_CREDIT: Credit amount of the loan<br>
# AMT_ANNUITY: Loan annuity<br>
# AMT_GOODS_PRICE: For consumer loans it is the price of the goods for which the loan is given<br>
# NAME_TYPE_SUITE: Who was accompanying client when he was applying for the loan<br>
# NAME_INCOME_TYPE: Clients income type (businessman, working, maternity leave,…)<br>
# NAME_EDUCATION_TYPE: Level of highest education the client achieved<br>
# NAME_FAMILY_STATUS: Family status of the client<br>
# NAME_HOUSING_TYPE: What is the housing situation of the client (renting, living with parents, ...)<br>
# REGION_POPULATION_RELATIVE: Normalized population of region where client lives (higher number means the client lives in more populated region)<br>
# DAYS_BIRTH: Client's age in days at the time of application<br>
# DAYS_EMPLOYED: How many days before the application the person started current employment<br>
# DAYS_REGISTRATION: How many days before the application did client change his registration<br>
# DAYS_ID_PUBLISH: How many days before the application did client change the identity document with which he applied for the loan<br>
# OWN_CAR_AGE: Age of client's car<br>
# FLAG_MOBIL: Did client provide mobile phone (1=YES, 0=NO)<br>
# FLAG_EMP_PHONE: Did client provide work phone (1=YES, 0=NO)<br>
# FLAG_WORK_PHONE: Did client provide home phone (1=YES, 0=NO)<br>
# FLAG_CONT_MOBILE: Was mobile phone reachable (1=YES, 0=NO)<br>
# FLAG_PHONE: Did client provide home phone (1=YES, 0=NO)<br>
# FLAG_EMAIL: Did client provide email (1=YES, 0=NO)<br>
# OCCUPATION_TYPE: What kind of occupation does the client have<br>
# CNT_FAM_MEMBERS: How many family members does client have<br>
# REGION_RATING_CLIENT: Our rating of the region where client lives (1,2,3)<br>
# REGION_RATING_CLIENT_W_CITY: Our rating of the region where client lives with taking city into account (1,2,3)<br>
# WEEKDAY_APPR_PROCESS_START: On which day of the week did the client apply for the loan<br>
# HOUR_APPR_PROCESS_START: Approximately at what hour did the client apply for the loan<br>
# REG_REGION_NOT_LIVE_REGION: Flag if client's permanent address does not match contact address (1=different, 0=same, at region level)<br>
# REG_REGION_NOT_WORK_REGION: Flag if client's permanent address does not match work address (1=different, 0=same, at region level)<br>
# LIVE_REGION_NOT_WORK_REGION: Flag if client's contact address does not match work address (1=different, 0=same, at region level)<br>
# REG_CITY_NOT_LIVE_CITY: Flag if client's permanent address does not match contact address (1=different, 0=same, at city level)<br>
# REG_CITY_NOT_WORK_CITY: Flag if client's permanent address does not match work address (1=different, 0=same, at city level)<br>
# LIVE_CITY_NOT_WORK_CITY: Flag if client's contact address does not match work address (1=different, 0=same, at city level)<br>
# ORGANIZATION_TYPE: Type of organization where client works<br>
# EXT_SOURCE_1: Normalized score from external data source<br>
# EXT_SOURCE_2: Normalized score from external data source<br>
# EXT_SOURCE_3: Normalized score from external data source<br>
# APARTMENTS_AVG: Normalized information about building where the client lives.<br>
# BASEMENTAREA_AVG: Normalized information about building where the client lives.<br>
# YEARS_BEGINEXPLUATATION_AVG: Normalized information about building where the client lives.<br>
# YEARS_BUILD_AVG: Normalized information about building where the client lives.<br>
# COMMONAREA_AVG: Normalized information about building where the client lives.<br>
# ELEVATORS_AVG: Normalized information about building where the client lives.<br>
# ENTRANCES_AVG: Normalized information about building where the client lives.<br>
# FLOORSMAX_AVG: Normalized information about building where the client lives.<br>
# FLOORSMIN_AVG: Normalized information about building where the client lives.<br>
# LANDAREA_AVG: Normalized information about building where the client lives.<br>
# LIVINGAPARTMENTS_AVG: Normalized information about building where the client lives.<br>
# LIVINGAREA_AVG: Normalized information about building where the client lives.<br>
# NONLIVINGAPARTMENTS_AVG: Normalized information about building where the client lives.<br>
# NONLIVINGAREA_AVG: Normalized information about building where the client lives.<br>
# APARTMENTS_MODE: Normalized information about building where the client lives.<br>
# BASEMENTAREA_MODE: Normalized information about building where the client lives.<br>
# YEARS_BEGINEXPLUATATION_MODE: Normalized information about building where the client lives.<br>
# YEARS_BUILD_MODE: Normalized information about building where the client lives<br>
# COMMONAREA_MODE: Normalized information about building where the client lives<br>
# ELEVATORS_MODE: Normalized information about building where the client lives<br>
# ENTRANCES_MODE: Normalized information about building where the client lives<br>
# FLOORSMAX_MODE: Normalized information about building where the client lives<br>
# FLOORSMIN_MODE: Normalized information about building where the client lives<br>
# LANDAREA_MODE: Normalized information about building where the client lives<br>
# LIVINGAPARTMENTS_MODE: Normalized information about building where the client lives<br>
# LIVINGAREA_MODE: Normalized information about building where the client lives<br>
# NONLIVINGAPARTMENTS_MODE: Normalized information about building where the client lives<br>
# NONLIVINGAREA_MODE: Normalized information about building where the client lives<br>
# APARTMENTS_MEDI: Normalized information about building where the client lives<br>
# BASEMENTAREA_MEDI: Normalized information about building where the client lives<br>
# YEARS_BEGINEXPLUATATION_MEDI: Normalized information about building where the client lives<br>
# YEARS_BUILD_MEDI: Normalized information about building where the client lives<br>
# COMMONAREA_MEDI: Normalized information about building where the client lives<br>
# ELEVATORS_MEDI: Normalized information about building where the client lives<br>
# ENTRANCES_MEDI: Normalized information about building where the client lives<br>
# FLOORSMAX_MEDI: Normalized information about building where the client lives<br>
# FLOORSMIN_MEDI: Normalized information about building where the client lives<br>
# LANDAREA_MEDI: Normalized information about building where the client lives<br>
# LIVINGAPARTMENTS_MEDI: Normalized information about building where the client lives<br>
# LIVINGAREA_MEDI: Normalized information about building where the client lives<br>
# NONLIVINGAPARTMENTS_MEDI: Normalized information about building where the client lives<br>
# NONLIVINGAREA_MEDI: Normalized information about building where the client lives<br>
# FONDKAPREMONT_MODE: Normalized information about building where the client lives<br>
# HOUSETYPE_MODE: Normalized information about building where the client lives<br>
# TOTALAREA_MODE: Normalized information about building where the client lives<br>
# WALLSMATERIAL_MODE: Normalized information about building where the client lives<br>
# EMERGENCYSTATE_MODE: Normalized information about building where the client lives<br>
# OBS_30_CNT_SOCIAL_CIRCLE: How many observation of client's social surroundings with observable 30 DPD (days past due) default<br>
# DEF_30_CNT_SOCIAL_CIRCLE: How many observation of client's social surroundings defaulted on 30 DPD (days past due) <br>
# OBS_60_CNT_SOCIAL_CIRCLE: How many observation of client's social surroundings with observable 60 DPD (days past due) default<br>
# DEF_60_CNT_SOCIAL_CIRCLE: How many observation of client's social surroundings defaulted on 60 (days past due) DPD<br>
# DAYS_LAST_PHONE_CHANGE: How many days before application did client change phone<br>
# FLAG_DOCUMENT_2: Did client provide document 2<br>
# FLAG_DOCUMENT_3: Did client provide document 3<br>
# FLAG_DOCUMENT_4: Did client provide document 4<br>
# FLAG_DOCUMENT_5: Did client provide document 5<br>
# FLAG_DOCUMENT_6: Did client provide document 6<br>
# FLAG_DOCUMENT_7: Did client provide document 7<br>
# FLAG_DOCUMENT_8: Did client provide document 8<br>
# FLAG_DOCUMENT_9: Did client provide document 9<br>
# FLAG_DOCUMENT_10: Did client provide document 10<br>
# FLAG_DOCUMENT_11: Did client provide document 11<br>
# FLAG_DOCUMENT_12: Did client provide document 12<br>
# FLAG_DOCUMENT_13: Did client provide document 13<br>
# FLAG_DOCUMENT_14: Did client provide document 14<br>
# FLAG_DOCUMENT_15: Did client provide document 15<br>
# FLAG_DOCUMENT_16: Did client provide document 16<br>
# FLAG_DOCUMENT_17: Did client provide document 17<br>
# FLAG_DOCUMENT_18: Did client provide document 18<br>
# FLAG_DOCUMENT_19: Did client provide document 19<br>
# FLAG_DOCUMENT_20: Did client provide document 20<br>
# FLAG_DOCUMENT_21: Did client provide document 21<br>
# AMT_REQ_CREDIT_BUREAU_HOUR: Number of enquiries to Credit Bureau about the client one hour before application<br>
# AMT_REQ_CREDIT_BUREAU_DAY: Number of enquiries to Credit Bureau about the client one day before application<br>
# AMT_REQ_CREDIT_BUREAU_WEEK: Number of enquiries to Credit Bureau about the client one week before applicationion)<br>
# AMT_REQ_CREDIT_BUREAU_MON: Number of enquiries to Credit Bureau about the client one month before application <br>
# AMT_REQ_CREDIT_BUREAU_QRT: Number of enquiries to Credit Bureau about the client 3 month before application <br>
# AMT_REQ_CREDIT_BUREAU_YEAR: Number of enquiries to Credit Bureau about the client one day year<br></p></blockquote>
#
# +
# Last amended: 24rd October, 2020
# Myfolder: C:\Users\Administrator\OneDrive\Documents\home_credit_default_risk
# Objective:
# Solving Kaggle problem: Home Credit Default Risk
# Processing application train/test datasets
#
# Data Source: https://www.kaggle.com/c/home-credit-default-risk/data
# Ref: https://www.kaggle.com/jsaguiar/lightgbm-with-simple-features
# +
# 1.0 Libraries
import numpy as np
import pandas as pd
# 1.1 Reduce read data size
# There is a file reducing.py
# in this folder. A class
# in it is used to reduce
# dataframe size
# (Code modified to
# exclude 'category' dtype)
# Refer: https://wkirgsn.github.io/2018/02/10/auto-downsizing-dtypes/
import reducing
# 1.2 Misc
import warnings
import os
warnings.simplefilter(action='ignore', category=FutureWarning)
# -
# 1.3 In view of large dataset, some useful options
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.max_columns', 100)
pd.set_option('display.max_rows', 100)
# 1.4 Display outputs from multiple commands from a jupyter cell
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# +
# 2.0 Onehot encoding (OHE) function. Uses pd.get_dummies()
# i) To transform 'object' columns to dummies.
# ii) Treat NaN as one of the categories
# iii) Returns transformed-data and new-columns created
def one_hot_encoder(df, nan_as_category = True):
original_columns = list(df.columns)
categorical_columns = [col for col in df.columns if df[col].dtype == 'object']
df = pd.get_dummies(df,
columns= categorical_columns,
dummy_na= nan_as_category # Treat NaNs as category
)
new_columns = [c for c in df.columns if c not in original_columns]
return df, new_columns
# -
# 2.1
pathToFolder = "C:\\Users\\Administrator\\OneDrive\\Documents\\home_credit_default_risk"
os.chdir(pathToFolder)
# 2.2 Some constants
num_rows=None # Implies read all rows
nan_as_category = True # While transforming
# 'object' columns to dummies
# +
# 3.0 Read previous application data first
df = pd.read_csv(
'application_train.csv.zip',
nrows = num_rows
)
# 3.0.1 Reduce memory usage by appropriately
# changing data-types per feature:
df = reducing.Reducer().reduce(df)
# +
# 3.0 Read previous application data first
test_df = pd.read_csv(
'application_test.csv.zip',
nrows = num_rows
)
# 3.0.1 Reduce memory usage by appropriately
# changing data-types per feature:
test_df = reducing.Reducer().reduce(test_df)
# -
# 3.1
df.shape # (307511, 122)
df.head()
# 3.1.1 There are 16 object types
df.dtypes.value_counts()
# 3.2
test_df.shape # (48744, 121)
test_df.head()
# 3.3 There are 16 object types
test_df.dtypes.value_counts()
# 3.4 Append test_df to train
df = df.append(test_df).reset_index()
# 3.5 Examine merged data
df.shape # (356255, 123)
df.head()
# 3.6 This gender is rare. So such
# rows can be dropped
df[df['CODE_GENDER'] == 'XNA']
# 3.7 Optional: Remove 4 applications with XNA CODE_GENDER (train set)
df = df[df['CODE_GENDER'] != 'XNA']
# 3.8 Categorical features with Binary encode (0 or 1; two categories)
for bin_feature in ['CODE_GENDER', 'FLAG_OWN_CAR', 'FLAG_OWN_REALTY']:
df[bin_feature], uniques = pd.factorize(df[bin_feature])
# 3.8.1
df.head()
uniques
# 3.8.2
df.dtypes
df.dtypes.value_counts()
df.dtypes['CODE_GENDER']
df.dtypes['FLAG_OWN_CAR']
df.dtypes['FLAG_OWN_REALTY']
# 4.0 Categorical features with One-Hot encode
df, cat_cols = one_hot_encoder(df, nan_as_category)
# 4.1
len(cat_cols) # 146
cat_cols
# 4.2 NaN values for DAYS_EMPLOYED: 365.243 -> nan
df['DAYS_EMPLOYED'].replace(365243, np.nan, inplace= True)
# 4.3 Some simple new features (percentages)
df['DAYS_EMPLOYED_PERC'] = df['DAYS_EMPLOYED'] / df['DAYS_BIRTH']
df['INCOME_CREDIT_PERC'] = df['AMT_INCOME_TOTAL'] / df['AMT_CREDIT']
df['INCOME_PER_PERSON'] = df['AMT_INCOME_TOTAL'] / df['CNT_FAM_MEMBERS']
df['ANNUITY_INCOME_PERC'] = df['AMT_ANNUITY'] / df['AMT_INCOME_TOTAL']
df['PAYMENT_RATE'] = df['AMT_ANNUITY'] / df['AMT_CREDIT']
# 5.0 Save the results for subsequent use:
df.to_csv("processed_df.csv.zip", compression = "zip")
# +
#################
|
application_train_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
"""
A) Series
"""
# pass list, tuple, or array & pandas makes the indeces automatically
future_array1 = [1,2,3,4,5,6] # 1 dimensional array
array1 = np.array(future_array1)
s = pd.Series(array1)
s
"""
B) DataFrame
"""
# DataFrame: object that can contain more than 1 Series
dates = pd.date_range("20180101", periods=10)
data = np.random.random((10,3))
df = pd.DataFrame(data, index=dates, columns=['Column1', 'Column2', 'Column3'])
df
# DataFrames, unlike Arrays, can store elements of different data types!
df['Column4'] = pd.Series([True, False, False, False, True, True, False, True, True, False], index=dates)
df
"""
C) Upload Data
"""
uploaded_data = pd.read_csv("filename.csv", index_col=0)
"""
D) View Data
"""
# View the top of the data set
df.head(3) # <- can leave it (), defaults to 5
# View the bottom of the data set
df.tail(3) # <- can leave it (), defaults to 5
# View each piece of the DataFrame
print (df.index)
print ("")
print (df.columns)
print ("")
print (df.values)
# View description statistics
print (df.describe())
"""
E) Indexing
"""
# Column names
df['Column2']
# Row Indeces
df[0:2] # or df['20180101':'20180102']
# Multi-axis with label
df.loc['20180101':'20180102',['Column1','Column3']]
# Multi-axis with indexing
df.iloc[3:5, 0:2]
df.iloc[4:8, 3:4]
# Boolean
df[df.Column1 > .5]
"""
F) Iterating
"""
for index, row in df.iterrows():
print (index, row[0])
"""
G) Reshaping & Sorting
"""
# Transpose
df.T
# Sort by Index
df.sort_index(axis=0, ascending=False)
# Sort by Values
df.sort_values(by='Column2')
"""
H) Join & Group
"""
# Join
data1 = np.random.random((10,2))
data2 = np.random.random((10,2))
df1 = pd.DataFrame(data1, index=dates, columns=['ColumnA', 'ColumnB'])
df2 = pd.DataFrame(data2, index=dates, columns=['ColumnC', 'ColumnD'])
df1.join(df2) # joins on the index, you can join on other columns with: pd.merge(df1, df2, on="column_name", how='left')
# Group
df.groupby('Column4').corr()
|
practice/intro_Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""Show fenics mesh and displacement solution."""
########################################################### dolfin
from dolfin import *
# Create mesh and define function space
mesh = UnitCubeMesh(12, 12, 12)
V = VectorFunctionSpace(mesh, "Lagrange", 1)
# Mark boundary subdomains
left = CompiledSubDomain("near(x[0], side) && on_boundary", side=0.0)
right = CompiledSubDomain("near(x[0], side) && on_boundary", side=1.0)
# Define Dirichlet boundary (x=0 or x=1)
c = Constant((0.0, 0.0, 0.0))
r = Expression((
"scale*0.0",
"scale*(y0 + (x[1]-y0)*cos(theta) - (x[2]-z0)*sin(theta)-x[1])",
"scale*(z0 + (x[1]-y0)*sin(theta) + (x[2]-z0)*cos(theta)-x[2])",
), scale=0.5, y0=0.5, z0=0.5, theta=pi/4, degree=2 )
bcl = DirichletBC(V, c, left)
bcr = DirichletBC(V, r, right)
w = TrialFunction(V) # Incremental displacement
v = TestFunction(V) # Test function
u = Function(V) # Solution
solve(inner(grad(w), grad(v)) * dx == inner(c, v) * dx, u, [bcl, bcr])
########################################################### vtkplotter
from vtkplotter.dolfin import plot
plot(u, mode='my displaced mesh please!!', shading='flat', cmap='jet')
# -
|
vtkplotter_examples/other/dolfin/notebooks/ex06_elasticity2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import functools
# +
def evaluateOrderOne(vector):
'''
Input: vector
'''
return np.sum(vector)
def __evaluate3Dvector(v):
#print(v)
tmp = np.sum(v)
if tmp == 3:
return 30
elif tmp == 2:
return 0
elif tmp == 0:
return 28
else:
if v[0]==1 :
return 14
elif v[1] == 1:
return 22
elif v[2] == 1:
return 26
def evaluateOrderThree(vector):
'''
Input: 3d vector
'''
try:
if vector.shape[0]>3:
s = 0
# print(int(vector.shape[0]/3),3)
newv = np.reshape(vector,(int(vector.shape[0]/3),3))
# print(newv)
for v in newv:
# print(v)
s += __evaluate3Dvector(v)
return s
else:
return __evaluate3Dvector(vector)
except Exception:
print("vector length invalid",Exception)
# -
evaluateOrderThree(np.array([1,0,0,1,1,0,1,1,0]))
evaluateOrderOne(np.array([1,0,0,1,1,0,1,1,0]))
# +
def bitflip_mutation(bit, probability):
if np.random.rand() < probability:
if bit == 1:
return 0.
else:
return 1.
else:
return bit
def uniform_crossover(i1,i2):
D = i1.shape[0]
u = np.empty((D,))
for i in range(D):
# print(P[i1][i],P[i2][i])
u[i] = np.random.choice(np.array([i1[i],i2[i]]))
return u
# -
def random_init(number,P,D, evaluate_func):
'''
initialize and evaluate the population
number: number of the individuals
P: the list for the population and value
D: dimension
'''
for i in range(number):
vector = np.random.choice(np.array([0,1]),(D,))
P.append((vector,evaluate_func(vector)))
return P
# +
def GEAlgorithm():
'''
the main algorithm
'''
n = 0
t = 1
D = 24
mu = 20
pm = 1/D # mutation rate
lambda_ = 30
evaluate_func = evaluateOrderOne
# Algorithm 2: The initialization procedure of the population
P = list()
P = random_init(mu,P,D,evaluate_func)
# print(P)
terminate = False
x_bsf = (None,-1)
for x,y in P:
if y == evaluate_func(np.ones(D)):
terminate = True
x_bsf = (x,y)
print(terminate,x_bsf)
while(not terminate):
Q = list()
for i in range(lambda_):
# Step1: Mating selection: generate two distinct random number
r = np.arange(len(P))
np.random.shuffle(r)
selected = r[:2]
#print(selected)
# Step2: Variation operator : Crossover
u = uniform_crossover(P[selected[0]][0],P[selected[1]][0])
# print(u)
# Step3: Variation operator2: Mutation
vfunc = np.vectorize(functools.partial(bitflip_mutation, probability=pm))
u = vfunc(u)
# print(u)
# Step4: Evaluate
new_value = evaluate_func(u)
# print("New value:" + str(newvalue))
n += 1
Q.append((u,new_value))
# Step5: Update bsf solution
if(new_value >x_bsf[1]):
x_bsf=(u,new_value)
# Step6: Environment Selection
R = P + Q
sort_result = sorted(R,key=lambda x:x[1],reverse=True)
P = sort_result[:int(len(R)/2)]
t += 1
if(new_value == evaluate_func(np.ones(D))):
terminate = True
print("# of generation:", t)
print("optimal value: %d"%x_bsf[1] )
return (t,n)
if __name__ == "__main__":
count = list()
totaltimes = 1
for i in range(totaltimes):
count.append(GEAlgorithm())
print(count)
# print("Average # of generations %f" % (sum(count)/float(totaltimes)))
|
Intro_to_GA_practice/ex5_mulambda.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: all,-execution,-papermill,-trusted
# formats: ipynb,py//py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown] tags=[]
# # Description
# %% [markdown] tags=[]
# Generates the figure for top cell types for a specified LV (in Settings section below).
# %% [markdown] tags=[]
# # Modules loading
# %% tags=[]
# %load_ext autoreload
# %autoreload 2
# %% tags=[]
import re
from pathlib import Path
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from data.recount2 import LVAnalysis
from utils import chunker
import conf
# %% [markdown] tags=[]
# # Settings
# %% tags=["parameters"]
LV_NAME = "LV246"
# %%
LV_AXIS_THRESHOLD = 3.0
N_TOP_SAMPLES = 400
N_TOP_ATTRS = 15
# %%
OUTPUT_FIGURES_DIR = Path(
conf.MANUSCRIPT["FIGURES_DIR"], "lvs_analysis", f"{LV_NAME.lower()}"
).resolve()
display(OUTPUT_FIGURES_DIR)
OUTPUT_FIGURES_DIR.mkdir(parents=True, exist_ok=True)
# %%
OUTPUT_CELL_TYPE_FILEPATH = OUTPUT_FIGURES_DIR / f"{LV_NAME.lower()}-cell_types.svg"
display(OUTPUT_CELL_TYPE_FILEPATH)
# %% [markdown] tags=[]
# # Load MultiPLIER summary
# %% tags=[]
multiplier_model_summary = pd.read_pickle(conf.MULTIPLIER["MODEL_SUMMARY_FILE"])
# %% tags=[]
multiplier_model_summary.shape
# %% tags=[]
multiplier_model_summary.head()
# %% [markdown] tags=[]
# # Load data
# %% [markdown] tags=[]
# ## Original data
# %% tags=[]
INPUT_SUBSET = "z_score_std"
# %% tags=[]
INPUT_STEM = "projection-smultixcan-efo_partial-mashr-zscores"
# %% tags=[]
input_filepath = Path(
conf.RESULTS["DATA_TRANSFORMATIONS_DIR"],
INPUT_SUBSET,
f"{INPUT_SUBSET}-{INPUT_STEM}.pkl",
).resolve()
display(input_filepath)
assert input_filepath.exists(), "Input file does not exist"
input_filepath_stem = input_filepath.stem
display(input_filepath_stem)
# %% tags=[]
data = pd.read_pickle(input_filepath)
# %% tags=[]
data.shape
# %% tags=[]
data.head()
# %% [markdown]
# ## LV data
# %%
lv_obj = LVAnalysis(LV_NAME, data)
# %%
multiplier_model_summary[
multiplier_model_summary["LV index"].isin((LV_NAME[2:],))
& (
(multiplier_model_summary["FDR"] < 0.05)
| (multiplier_model_summary["AUC"] >= 0.75)
)
]
# %%
lv_data = lv_obj.get_experiments_data()
# %%
lv_data.shape
# %%
lv_data.head()
# %% [markdown]
# # LV cell types analysis
# %% [markdown]
# ## Get top attributes
# %%
lv_attrs = lv_obj.get_attributes_variation_score()
display(lv_attrs.head(20))
# %%
# show those with cell type or tissue in their name
_tmp = pd.Series(lv_attrs.index)
lv_attrs[
_tmp.str.match(
"(?:cell.+type$)|(?:tissue$)|(?:tissue.+type$)",
case=False,
flags=re.IGNORECASE,
).values
].sort_values(ascending=False)
# %%
_tmp = lv_data.loc[
:,
[
"cell type",
"tissue",
"tissue type",
LV_NAME,
],
]
# %%
_tmp_seq = list(chunker(_tmp.sort_values(LV_NAME, ascending=False), 25))
# %%
_tmp_seq[3]
# %%
# what is there in these projects?
lv_data.loc[["SRP007412"]].dropna(how="all", axis=1).sort_values(
LV_NAME, ascending=False
).sort_values(LV_NAME, ascending=False).head(10)
# %% [markdown]
# **No cell types/tissues/cell lines** so not very interesting
# %%
SELECTED_ATTRIBUTE = "cell type"
# %%
# it has to be in the order desired for filling nans in the SELECTED_ATTRIBUTE
SECOND_ATTRIBUTES = ["tissue", "tissue type"]
# %% [markdown]
# ## Get plot data
# %%
plot_data = lv_data.loc[:, [SELECTED_ATTRIBUTE] + SECOND_ATTRIBUTES + [LV_NAME]]
# %%
# if blank/nan, fill cell type column with tissue content
_new_column = plot_data[[SELECTED_ATTRIBUTE] + SECOND_ATTRIBUTES].fillna(
method="backfill", axis=1
)[SELECTED_ATTRIBUTE]
plot_data[SELECTED_ATTRIBUTE] = _new_column
plot_data = plot_data.drop(columns=SECOND_ATTRIBUTES)
plot_data = plot_data.fillna({SELECTED_ATTRIBUTE: "NOT CATEGORIZED"})
# plot_data = plot_data.dropna(subset=[SELECTED_ATTRIBUTE])
# %%
plot_data = plot_data.sort_values(LV_NAME, ascending=False)
# %%
plot_data.head(20)
# %% [markdown]
# ## Customize x-axis values
# %% [markdown]
# When cell type values are not very clear, customize their names by looking at their specific studies to know exactly what the authors meant.
# %%
final_plot_data = plot_data.replace(
{
SELECTED_ATTRIBUTE: {
"normal skin": "Skin",
"liver": "Liver",
"Human Skeletal Muscle Myoblasts (HSMM)": "Skeletal muscle myoblasts",
"astrocytes": "Astrocytes",
"mixture of U87 human glioma cells and WI-38 human lung fibroblast cells": "Glioma cells + lung fibroblast cells",
"functional hepatocytes generated by lineage reprogramming": "Hepatocytes",
"human adipose-derived stem cells": "Adipose-derived stem cells",
"adipose": "Adipose",
"embryonic stem cells": "Embryonic stem cells",
"primary keratinocytes": "Primary keratinocytes",
"fetal liver": "Fetal liver",
"in vitro differentiated erythroid cells": "Erythroid cells",
"WAT": "White adipose tissue",
"BAT": "Brown adipose tissue",
"Uninvolved Breast Tissue Adjacent to ER+ Primary Tumor": "Breast tissue adjacent to ER+ tumor",
"ovarian granulosa cells": "Ovarian granulosa cells",
}
}
)
# %%
# take the top samples only
final_plot_data = final_plot_data.sort_values(LV_NAME, ascending=False)[:N_TOP_SAMPLES]
# %% [markdown]
# ## Threshold LV values
# %%
final_plot_data.loc[
final_plot_data[LV_NAME] > LV_AXIS_THRESHOLD, LV_NAME
] = LV_AXIS_THRESHOLD
# %% [markdown]
# ## Delete samples with no tissue/cell type information
# %%
final_plot_data = final_plot_data[
final_plot_data[SELECTED_ATTRIBUTE] != "NOT CATEGORIZED"
]
# %% [markdown]
# ## Set x-axis order
# %%
attr_order = (
final_plot_data.groupby(SELECTED_ATTRIBUTE)
.max()
.sort_values(LV_NAME, ascending=False)
.index[:N_TOP_ATTRS]
.tolist()
)
# %%
len(attr_order)
# %%
attr_order[:5]
# %% [markdown]
# ## Plot
# %%
with sns.plotting_context("paper", font_scale=2.5), sns.axes_style("whitegrid"):
g = sns.catplot(
data=final_plot_data,
y=LV_NAME,
x=SELECTED_ATTRIBUTE,
order=attr_order,
kind="strip",
height=5,
aspect=2.5,
)
plt.xticks(rotation=45, horizontalalignment="right")
plt.xlabel("")
plt.savefig(
OUTPUT_CELL_TYPE_FILEPATH,
bbox_inches="tight",
facecolor="white",
)
# %% [markdown]
# # Debug
# %%
# with pd.option_context(
# "display.max_rows", None, "display.max_columns", None, "display.max_colwidth", None
# ):
# _tmp = final_plot_data[final_plot_data[SELECTED_ATTRIBUTE].str.contains("ovarian")]
# display(_tmp.head(20))
# %%
# # what is there in these projects?
# lv_data.loc[["SRP056041"]].dropna(how="all", axis=1).sort_values(
# LV_NAME, ascending=False
# ).head(60)
# %%
|
nbs/99_manuscript/lvs/lv246/lv246-cell_types.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### SVM Model For Digit Recognition
from sklearn import svm, metrics #Importing SVM model builder
import numpy as np #to convert into numpy arrays
import os #to change the source directory
import pandas as pd #to upload the data
from sklearn.metrics import accuracy_score # to Calculate accuracy
from pandas_ml import ConfusionMatrix #To Generate Confusion Matrix
from sklearn.metrics import classification_report # To Calculate Precision, Recall
#Importing the data
os.chdir("C:/Users/<NAME>/Desktop/Capstone/Signature Recognition/Data")
label_train = pd.read_csv('train_numbers.csv')
y_train = label_train['Label']
label_test = pd.read_csv('test_numbers.csv')
y_test = label_test['Label']
X_train = label_train.iloc[:,1:1601]
X_test = label_test.iloc[:,1:1601]
# +
#Creating svm Classifier
classifier = svm.LinearSVC()
# Learning Phase
classifier.fit(X_train, y_train)
# Predict Test Set
predicted = classifier.predict(X_test)
#Calculating the accuracy score
score = accuracy_score(y_test, predicted)
print("Accuracy score: %0.2f" % score)
# confusion matrix
ConfusionMatrix(y_test, predicted)
# -
#Calculating the precision and recall
report = classification_report(y_test,predicted)
print(report)
# ### SVM Model for Reversed values of Pixels 0-black and 255 is white
#Importing the dataset
os.chdir("C:/Users/<NAME>/Desktop/Capstone/Signature Recognition/Data")
label_train = pd.read_csv('trainzero.csv')
y_train = label_train['Label']
label_test = pd.read_csv('testzero.csv')
y_test = label_test['Label']
X_train = label_train.iloc[:,1:1601]
X_test = label_test.iloc[:,1:1601]
# +
#Creating SVM Classifier
classifier = svm.LinearSVC()
# Learning Phase
classifier.fit(X_train, y_train)
# Predict Test Set
predicted = classifier.predict(X_test)
score = accuracy_score(y_test, predicted)
print("Accuracy score: %0.2f" % score)
# confusion matrix
ConfusionMatrix(y_test, predicted)
# -
#Calculating Precision and Recall
report = classification_report(y_test,predicted)
print(report)
# ### SVM Model for Dataset with Images of size 100x100 pixels
#Importing the dataset
os.chdir("C:/Users/<NAME>/Desktop/Capstone/Signature Recognition/Data")
label_train = pd.read_csv('train100.csv')
y_train = label_train['Label']
label_test = pd.read_csv('test100.csv')
y_test = label_test['Label']
X_train = label_train.iloc[:,1:10001]
X_test = label_test.iloc[:,1:10001]
# +
#Creating SVM Classifier
classifier = svm.LinearSVC()
# Learning Phase
classifier.fit(X_train, y_train)
# Predict Test Set
predicted = classifier.predict(X_test)
score = accuracy_score(y_test, predicted)
print("Accuracy score: %0.2f" % score)
# confusion matrix
ConfusionMatrix(y_test, predicted)
# -
#Calculating Precision and Recall.
report = classification_report(y_test,predicted)
print(report)
|
codes/Support Vector Machines.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# File to Load
city_data_to_load = "city_data.csv"
ride_data_to_load = "ride_data.csv"
# Read the City and Ride Data
city_data_df = pd.read_csv(city_data_to_load)
ride_data_df = pd.read_csv(ride_data_to_load)
# Combine the data into a single dataset
city_data_df.head()
ride_data_df.head()
city_ride_df = pd.merge(city_data_df, ride_data_df, on="city")
# Display the data table for preview
city_ride_df
# -
# ## Bubble Plot of Ride Sharing Data
# +
# Obtain the x and y coordinates for each of the three city types
urban = city_ride_df[city_ride_df["type"] == "Urban"]
urban_city = urban.groupby(["city"])
urban_ride_count = urban_city.count()["ride_id"]
urban_average_fare = urban_city.mean()["fare"]
urban_driver_count = urban_city.mean()["driver_count"]
suburban = city_ride_df[city_ride_df["type"] == "Suburban"]
suburban_city = suburban.groupby(["city"])
suburban_ride_count = suburban_city.count()["ride_id"]
suburban_average_fare = suburban_city.mean()["fare"]
suburban_driver_count = suburban_city.mean()["driver_count"]
rural = city_ride_df[city_ride_df["type"] == "Rural"]
rural_city = rural.groupby(["city"])
rural_ride_count = rural_city.count()["ride_id"]
rural_average_fare = rural_city.mean()["fare"]
rural_driver_count = rural_city.mean()["driver_count"]
# Build the scatter plots for each city types
urban = plt.scatter(urban_ride_count, urban_average_fare, s=urban_driver_count * 10, c="lightcoral", alpha=0.5, edgecolors="black", label="Urban")
suburban = plt.scatter(suburban_ride_count, suburban_average_fare, s=suburban_driver_count * 10, c="lightskyblue", alpha=0.5, edgecolors="black", label="Suburban")
rural = plt.scatter(rural_ride_count, rural_average_fare, s=rural_driver_count * 10, c="gold", alpha=0.5, edgecolors="black", label="Rural")
# Incorporate the other graph properties
plt.grid()
plt.xlabel("Total Number of Rides (Per City)")
plt.ylabel("Average Fare ($)")
plt.title("Pyber Ride Sharing Data (2016)")
# Create a legend
plt.legend(handles=[urban, suburban, rural], loc="best")
# Incorporate a text label regarding circle size
textstr = "Note: Circle size correlates with driver count per city"
plt.text(45, 30, textstr, fontsize=12)
# Save Figure
plt.show()
# -
# Pyber Ride Sharing Data Analysis
#
# The relationship of number of rides to cost of fare for the data we have on urban areas is reciprocal to that which we have on rural areas. That is to say, there is an indirect relationship between total number of rides and average fare. In other words, urban areas service multiple times more rides than rural areas for half the fare for each ride. Of course, that is to be expected given in urban areas rider destinations are closer, whereas in rural areas rider destinations are farther apart.
# ## Total Fares by City Type
# +
# Calculate Type Percents
urban_total_fare_by_city = urban_city.sum()["fare"]
urban_total_fare = urban_total_fare_by_city.sum()
suburban_total_fare_by_city = suburban_city.sum()["fare"]
suburban_total_fare = suburban_total_fare_by_city.sum()
rural_total_fare_by_city = rural_city.sum()["fare"]
rural_total_fare = rural_total_fare_by_city.sum()
total_fares = urban_total_fare + suburban_total_fare + rural_total_fare
urban_fares_percent = urban_total_fare/total_fares
suburban_fares_percent = suburban_total_fare/total_fares
rural_fares_percent = rural_total_fare/total_fares
percent_total_fares = [urban_fares_percent, suburban_fares_percent, rural_fares_percent]
# Build Pie Chart
labels = ["Urban", "Suburban", "Rural"]
colors = ["lightcoral", "lightskyblue", "gold"]
explode = (0.05, 0, 0)
plt.title("% of Total Fares by City Type")
plt.pie(percent_total_fares, explode=explode, labels=labels, colors = colors, autopct = "%1.2f%%", shadow = True, startangle = 70)
"""
# Save Figure
plt.tight_layout()
plt.savefig("./Images/Total_Fares_by_City_Type.png")
plt.show()
"""
# -
# ## Total Rides by City Type
# +
# Calculate Ride Percents
total_ride_count = sum(urban_ride_count) + sum(suburban_ride_count) + sum(rural_ride_count)
urban_rides_percent = sum(urban_ride_count)/total_ride_count
suburban_rides_percent = sum(suburban_ride_count)/total_ride_count
rural_rides_percent = sum(rural_ride_count)/total_ride_count
percent_total_rides = [urban_rides_percent, suburban_rides_percent, rural_rides_percent]
# Build Pie Chart
labels = ["Urban", "Suburban", "Rural"]
colors = ["lightcoral", "lightskyblue", "gold"]
explode = (0.05, 0, 0)
plt.title("% of Total Rides by City Type")
plt.pie(percent_total_rides, explode=explode, labels=labels, colors = colors, autopct = "%1.2f%%", shadow = True, startangle = 70)
plt.show()
"""
# Save Figure
plt.tight_layout()
plt.savefig("./Images/Total_Rides_by_City_Type.png")
"""
# -
# ## Total Drivers by City Type
# +
# Calculate Driver Percents
total_driver_count = sum(urban_driver_count) + sum(suburban_driver_count) + sum(rural_driver_count)
total_driver_count
urban_drivers_percent = sum(urban_driver_count)/total_driver_count
suburban_drivers_percent = sum(suburban_driver_count)/total_driver_count
rural_drivers_percent = sum(rural_driver_count)/total_driver_count
percent_total_drivers = [urban_drivers_percent, suburban_drivers_percent, rural_drivers_percent]
# Build Pie Charts
labels = ["Urban", "Suburban", "Rural"]
colors = ["lightcoral", "lightskyblue", "gold"]
explode = (0.05, 0, 0)
plt.title("% of Total Drivers by City Type")
plt.pie(percent_total_drivers, explode=explode, labels=labels, colors = colors, autopct = "%1.2f%%", shadow = True, startangle = 70)
plt.show()
# Save Figure
plt.tight_layout()
plt.savefig("./Images/Total_Drivers_by_City_Type.png")
plt.show()
# -
# Percent of Total Drivers by City Type Analysis
#
# The drivers in rural areas are completing more rides per driver then their urban and suburban counterparts. If 68% of all Pyber rides take place in urban areas, but urban drivers comprose 81% of the Pyber drivers, then one could infer:
# 1) many urban drivers who are be employed by Pyber are not picking up rides
# and/or
# 2) the average number of rides per driver is higher in rural areas than in urban areas
|
.ipynb_checkpoints/Pyber-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Displaying multiple images
# In this activity, we will plot images in a grid.
# +
# Import statements
import os
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# -
# Load all four images from the subfolder data.
# Load images
img_filenames = sorted(os.listdir('../../datasets/images'))
imgs = [mpimg.imread(os.path.join('../../datasets/images', img_filename)) for img_filename in img_filenames]
# Visualize the images in a 2x2 grid. Remove the axes and give each image a label.
# Create subplot
fig, axes = plt.subplots(2, 2)
fig.figsize = (6, 6)
fig.dpi = 150
axes = axes.ravel()
# Specify labels
labels = ['coast', 'beach', 'building', 'city at night']
# Plot images
for i in range(len(imgs)):
axes[i].imshow(imgs[i])
axes[i].set_xticks([])
axes[i].set_yticks([])
axes[i].set_xlabel(labels[i])
plt.show()
|
Chapter03/Activity3.07/Activity3.07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Hqp-TQt4vIBw"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/sjchoi86/upstage-basic-deeplearning/blob/main/notebook/optm.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/sjchoi86/upstage-basic-deeplearning/blob/main/notebook/optm.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View Source</a>
# </td>
# </table>
# + [markdown] id="WBr-T2RD6hma"
# # Regression with Different Optimizers
# + id="MAbu9jNhrdOL"
# !pip install matplotlib==3.3.0
# + id="Pgdbxd616YZg"
import numpy as np
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
print ("PyTorch version:[%s]."%(torch.__version__))
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
print ("device:[%s]."%(device))
# + [markdown] id="e6IJu5BU6qJm"
# ### Dataset
# + id="1jldyTcJ6k7X"
n_data = 10000
x_numpy = -3+6*np.random.rand(n_data,1)
y_numpy = np.exp(-(x_numpy**2))*np.cos(10*x_numpy) + 3e-2*np.random.randn(n_data,1)
plt.figure(figsize=(8,5))
plt.plot(x_numpy,y_numpy,'r.',ms=2)
plt.show()
x_torch = torch.Tensor(x_numpy).to(device)
y_torch = torch.Tensor(y_numpy).to(device)
print ("Done.")
# + [markdown] id="J_AHUbKlFy1t"
# ### Define Model
# + id="3tJB4a1NFAd6"
class Model(nn.Module):
def __init__(self,name='mlp',xdim=1,hdims=[16,16],ydim=1):
super(Model, self).__init__()
self.name = name
self.xdim = xdim
self.hdims = hdims
self.ydim = ydim
self.layers = []
prev_hdim = self.xdim
for hdim in self.hdims:
self.layers.append(nn.Linear(
# FILL IN HERE
))
self.layers.append(nn.Tanh()) # activation
prev_hdim = hdim
# Final layer (without activation)
self.layers.append(nn.Linear(prev_hdim,self.ydim,bias=True))
# Concatenate all layers
self.net = nn.Sequential()
for l_idx,layer in enumerate(self.layers):
layer_name = "%s_%02d"%(type(layer).__name__.lower(),l_idx)
self.net.add_module(layer_name,layer)
self.init_param() # initialize parameters
def init_param(self):
for m in self.modules():
if isinstance(m,nn.Conv2d): # init conv
nn.init.kaiming_normal_(m.weight)
nn.init.zeros_(m.bias)
elif isinstance(m,nn.Linear): # lnit dense
nn.init.kaiming_normal_(m.weight)
nn.init.zeros_(m.bias)
def forward(self,x):
return self.net(x)
print ("Done.")
# + id="n-gPx2m3Fvw1"
LEARNING_RATE = 1e-2
# Instantiate models
model_sgd = Model(name='mlp_sgd',xdim=1,hdims=[64,64],ydim=1).to(device)
model_momentum = Model(name='mlp_momentum',xdim=1,hdims=[64,64],ydim=1).to(device)
model_adam = Model(name='mlp_adam',xdim=1,hdims=[64,64],ydim=1).to(device)
# Optimizers
loss = nn.MSELoss()
optm_sgd = optim.SGD(
# FILL IN HERE
)
optm_momentum = optim.SGD(
# FILL IN HERE
)
optm_adam = optim.Adam(
# FILL IN HERE
)
print ("Done.")
# + [markdown] id="b5OO8B6WiQ9V"
# ### Check Parameters
# + id="qu66dEIxFwGc"
np.set_printoptions(precision=3)
n_param = 0
for p_idx,(param_name,param) in enumerate(model_sgd.named_parameters()):
if param.requires_grad:
param_numpy = param.detach().cpu().numpy() # to numpy array
n_param += len(param_numpy.reshape(-1))
print ("[%d] name:[%s] shape:[%s]."%(p_idx,param_name,param_numpy.shape))
print (" val:%s"%(param_numpy.reshape(-1)[:5]))
print ("Total number of parameters:[%s]."%(format(n_param,',d')))
# + [markdown] id="g5PP5rRvi3c6"
# ### Train
# + id="B-uu6x8DFwZ9"
MAX_ITER,BATCH_SIZE,PLOT_EVERY = 1e4,64,500
model_sgd.init_param()
model_momentum.init_param()
model_adam.init_param()
model_sgd.train()
model_momentum.train()
model_adam.train()
for it in range(int(MAX_ITER)):
r_idx = np.random.permutation(n_data)[:BATCH_SIZE]
batch_x,batch_y = x_torch[r_idx],y_torch[r_idx]
# Update with Adam
y_pred_adam = model_adam.forward(batch_x)
loss_adam = loss(y_pred_adam,batch_y)
optm_adam.zero_grad()
loss_adam.backward()
optm_adam.step()
# Update with Momentum
y_pred_momentum = model_momentum.forward(batch_x)
loss_momentum = loss(y_pred_momentum,batch_y)
optm_momentum.zero_grad()
loss_momentum.backward()
optm_momentum.step()
# Update with SGD
y_pred_sgd = model_sgd.forward(batch_x)
loss_sgd = loss(y_pred_sgd,batch_y)
optm_sgd.zero_grad()
loss_sgd.backward()
optm_sgd.step()
# Plot
if ((it%PLOT_EVERY)==0) or (it==0) or (it==(MAX_ITER-1)):
with torch.no_grad():
y_sgd_numpy = model_sgd.forward(x_torch).cpu().detach().numpy()
y_momentum_numpy = model_momentum.forward(x_torch).cpu().detach().numpy()
y_adam_numpy = model_adam.forward(x_torch).cpu().detach().numpy()
plt.figure(figsize=(8,4))
plt.plot(x_numpy,y_numpy,'r.',ms=4,label='GT')
plt.plot(x_numpy,y_sgd_numpy,'g.',ms=2,label='SGD')
plt.plot(x_numpy,y_momentum_numpy,'b.',ms=2,label='Momentum')
plt.plot(x_numpy,y_adam_numpy,'k.',ms=2,label='ADAM')
plt.title("[%d/%d]"%(it,MAX_ITER),fontsize=15)
plt.legend(labelcolor='linecolor',loc='upper right',fontsize=15)
plt.show()
print ("Done.")
# + id="rVf8-DZnrDdz"
|
notebook/optm.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # ** DIscBIO: a user-friendly pipeline for biomarker discovery in single-cell transcriptomics**
# The pipeline consists of four successive steps: data pre-processing, cellular clustering and pseudo-temporal ordering, determining differential expressed genes and identifying biomarkers.
# 
# # CTC Notebook [PART 3]
#
# # Determining differentially expressed genes and biomarkers
# ## Required Packages
library(DIscBIO)
library(partykit)
library(enrichR)
# ## 3. Determining differentially expressed genes (DEGs) For K-means Clustering
# Differentially expressed genes between individual clusters are identified using the significance analysis of sequencing data (SAMseq), which is a new function in significance analysis of microarrays (Li and Tibshirani 2011) in the samr package v2.0 (Tibshirani et all., 2015). SAMseq is a non-parametric statistical function dependent on Wilcoxon rank statistic that equalizes the sizes of the library by a resampling method accounting for the various sequencing depths. The analysis is implemented over the pure raw dataset that has the unnormalized expression read counts after excluding the ERCCs. Furthermore, DEGs in each cluster comparing to all the remaining clusters are determined using binomial differential expression, which is based on binomial counting statistics.
# 
# ## 3.1 Identifying DEGs using SAMseq
# The user can define DEGs between all clusters generated by either K-means or model based clustering by applying the “DEGanalysis” function. Another alternative is to define DEGs between particular clusters generated by K-means clustering by applying the “DEGanalysis2clust” function. The outcome of these two functions is a list of two components:
# - The first component is a data frame showing the Ensembl gage name and the symbole of the detected DEGs
# - The second component is table showing the comparisons, Target cluster, Number of genes and the File name. This component will be used for the downstream analysis.
# +
load("SC.RData") # Loading the "SC" object that has include the data of the k-means clustering
load("Ndata.RData") # Loading the "Ndata" object and stored in the @ndata will be used to plot the expression of genes
load("expdata.RData") # Loading the "expdata" object and stored in the @expdata will be used to plot the expression of genes
sc<-SC # Storing the data of SC in the sc
sc@ndata<-Ndata
sc@expdata<-expdata
########## Removing the unneeded objects
rm(Ndata)
rm(expdata)
rm(SC)
# -
# ## Identifying DEGs using binomial differential expression
# The function ClustDiffGenes identifies differentially regulated genes for each cluster of the K-means clustering in
# comparison to the ensemble of all cells. It returns a list with a data.frame element for each cluster that contains the mean expression across all cells not in the cluster (mean.ncl) and in the cluster (mean.cl), the fold-change in the cluster versus all remaining cells (fc), and the p-value for differential expression between all cells in a cluster and all remaining cells. The p-value is computed based on the overlap of negative binomials fitted to the count distributions within the two groups akin to DESeq.
cdiffBinomial<-ClustDiffGenes(sc,K=4,export = T,fdr=.01,quiet=T) ########## Binomial differential expression analysis
#### To show the result table
head(cdiffBinomial[[1]]) # The first component
head(cdiffBinomial[[2]]) # The second component
# ### Plotting the DEGs
# Volcano plots are used to readily show the DEGs by plotting significance versus fold-change on the y and x axes, respectively.
name<-cdiffBinomial[[2]][1,6] ############ Selecting the DEGs' ############## Down-DEG-cluster1.csv
U<-read.csv(file=paste0(name),head=TRUE,sep=",")
Vplot<-VolcanoPlot(U,value=0.0001,name=name,FS=0.7,fc=0.75)
# ## 4. Identifying biomarkers (decision trees and networking analysis)
# There are several methods to identify biomarkers, among them are decision trees and hub detection through networking analysis. The outcome of STRING analysis is stored in tab separated values (TSV) files. These TSV files served as an input to check both the connectivity degree and the betweenness centrality, which reflects the communication flow in the defined PPI networks
# 
# Decision trees are one of the most efficient classification techniques in biomarkers discovery. Here we use it to predict the sub-population of a target cell based on transcriptomic data. Two types of decision trees can be performed: classification and regression trees (CART) and J48. The decision tree analysis is implemented over a training dataset, which consisted of the DEGs obtained by either SAMseq or the binomial differential expression. The performance of the generated trees can be evaluated for error estimation by ten-fold cross validation assessment using the "J48DTeval" and "RpartEVAL" functions. The decision tree analysis requires the dataset to be class vectored by applying the “ClassVectoringDT” function.
###################### Finding biomarker genes between cluster 1 and cluster 4
First="CL1"
Second="CL4"
load("DATAforDT.RData")
# ### 4.1. J48 Decision Tree
j48dt<-J48DT(DATAforDT) #J48 Decision Tree
summary(j48dt)
rm(j48dt)
# ### 4.2. RPART Decision Tree
rpartDT<-RpartDT(DATAforDT)
rm(rpartDT)
# ### 4.3. Networking Analysis
# To define protein-protein interactions (PPI) over a list of genes, STRING-api is used. The outcome of STRING analysis was stored in tab separated values (TSV) files. These TSV files served as an input to check both the connectivity degree and the betweenness centrality, which reflects the communication flow in the defined PPI networks.
# +
DEGs="All_DEGs"
FileName=paste0(DEGs)
data<-cdiffBinomial[[1]] [1:200,2] # DEGs gene list from Binomial analysis (taking only the firat 200 genes)
ppi<-PPI(data,FileName)
networking<-NetAnalysis(ppi)
networking ##### In case the Examine response components = 200 and an error "linkmat[i, ]" appeared, that means there are no PPI.
# -
data=networking[1:25,1] # plotting the network of the top 25 highly connected genes
network<-Networking(data,FileName,plot_width = 25, plot_height = 10)
# ### 4.4 Gene Enrichment Analysis
dbs <- listEnrichrDbs()
head(dbs)
#print(dbs)
# +
############ Selecting the DEGs' table ##############
DEGs=cdiffBinomial[[2]][1,4] # Down-regulated markers in cluster 1
FileName=paste0(DEGs)
data<-read.csv(file=paste0(DEGs),head=TRUE,sep=",")
data<-as.character(data[,3])
dbs <- c("KEGG_2019_Human","GO_Biological_Process_2015")
enriched <- enrichr(data, dbs)
KEGG_2019_Human<-enriched[[1]][,c(1,2,3,9)]
GO_Biological_Process_2015<-enriched[[2]][,c(1,2,3,9)]
GEA<-rbind(KEGG_2019_Human,GO_Biological_Process_2015)
GEA
# -
|
notebook/DIscBIO-CTCs-Binder-Part3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
print("Pi = {}".format(math.pi))
print("Pi = %s" % (math.pi,))
math.cos(0)
math.sin(math.pi/2.0)
# +
def hello(you):
print("hello {}".format(you))
return
hello("there")
# -
def add(x, y): return x+y
def mul(x, y): return x*y
def div(x, y): return x/y
# +
class Circle(object):
def __init__(self, origin, r):
self.origin = origin
self.r = r
def surface(self):
return math.pi * self.r**2
pass
class Rectangle(object):
def __init__(self, origin, width, height):
self.origin = origin
self.width = width
self.height = height
def surface(self):
return self.height * self.width
pass
class Point(object):
def __init__(self, x, y):
self.x = x
self.y = y
def __str__(self):
return "Point(x={x}, y={y})".format(x=self.x, y=self.y)
pass
# -
p = Point(1,2)
p
print(p)
|
notebooks/intro-python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Batch Normalization in `gluon`
#
# In the preceding section, [we implemented batch normalization ourselves](../chapter04_convolutional-neural-networks/cnn-batch-norm-scratch.ipynb) using NDArray and autograd.
# As with most commonly used neural network layers,
# Gluon has batch normalization predefined,
# so this section is going to be straightforward.
from __future__ import print_function
import mxnet as mx
from mxnet import nd, autograd
from mxnet import gluon
import numpy as np
mx.random.seed(1)
ctx = mx.cpu()
# ## The MNIST dataset
batch_size = 64
num_inputs = 784
num_outputs = 10
def transform(data, label):
return nd.transpose(data.astype(np.float32), (2,0,1))/255, label.astype(np.float32)
train_data = mx.gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=True, transform=transform),
batch_size, shuffle=True)
test_data = mx.gluon.data.DataLoader(mx.gluon.data.vision.MNIST(train=False, transform=transform),
batch_size, shuffle=False)
# ## Define a CNN with Batch Normalization
#
# To add batchnormalization to a ``gluon`` model defined with Sequential,
# we only need to add a few lines.
# Specifically, we just insert `BatchNorm` layers before the applying the ReLU activations.
num_fc = 512
net = gluon.nn.Sequential()
with net.name_scope():
net.add(gluon.nn.Conv2D(channels=20, kernel_size=5))
net.add(gluon.nn.BatchNorm(axis=1, center=True, scale=True))
net.add(gluon.nn.Activation(activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
net.add(gluon.nn.Conv2D(channels=50, kernel_size=5))
net.add(gluon.nn.BatchNorm(axis=1, center=True, scale=True))
net.add(gluon.nn.Activation(activation='relu'))
net.add(gluon.nn.MaxPool2D(pool_size=2, strides=2))
# The Flatten layer collapses all axis, except the first one, into one axis.
net.add(gluon.nn.Flatten())
net.add(gluon.nn.Dense(num_fc))
net.add(gluon.nn.BatchNorm(axis=1, center=True, scale=True))
net.add(gluon.nn.Activation(activation='relu'))
net.add(gluon.nn.Dense(num_outputs))
# ## Parameter initialization
#
net.collect_params().initialize(mx.init.Xavier(magnitude=2.24), ctx=ctx)
# ## Softmax cross-entropy Loss
softmax_cross_entropy = gluon.loss.SoftmaxCrossEntropyLoss()
# ## Optimizer
trainer = gluon.Trainer(net.collect_params(), 'sgd', {'learning_rate': .1})
# ## Write evaluation loop to calculate accuracy
def evaluate_accuracy(data_iterator, net):
acc = mx.metric.Accuracy()
for i, (data, label) in enumerate(data_iterator):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
output = net(data)
predictions = nd.argmax(output, axis=1)
acc.update(preds=predictions, labels=label)
return acc.get()[1]
# ## Training Loop
# +
epochs = 1
smoothing_constant = .01
for e in range(epochs):
for i, (data, label) in enumerate(train_data):
data = data.as_in_context(ctx)
label = label.as_in_context(ctx)
with autograd.record():
output = net(data)
loss = softmax_cross_entropy(output, label)
loss.backward()
trainer.step(data.shape[0])
##########################
# Keep a moving average of the losses
##########################
curr_loss = nd.mean(loss).asscalar()
moving_loss = (curr_loss if ((i == 0) and (e == 0))
else (1 - smoothing_constant) * moving_loss + (smoothing_constant) * curr_loss)
test_accuracy = evaluate_accuracy(test_data, net)
train_accuracy = evaluate_accuracy(train_data, net)
print("Epoch %s. Loss: %s, Train_acc %s, Test_acc %s" % (e, moving_loss, train_accuracy, test_accuracy))
# -
# ## Next
# [Introduction to recurrent neural networks](../chapter05_recurrent-neural-networks/simple-rnn.ipynb)
# For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
|
chapter04_convolutional-neural-networks/cnn-batch-norm-gluon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementation of Logistic Regression
# ## Import Libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import math
titanic_data = pd.read_csv("titanic.csv")
titanic_data.head(10)
print((titanic_data.info()))
print("# Total number of row and column in dataset: %s, %s" % (len(titanic_data.index), len(titanic_data.columns)))
# # Analyzing Data
sns.countplot(x="Survived", data=titanic_data)
# +
# 0 means passenger who had not survived
# -
sns.countplot(x="Survived", hue="Sex", data=titanic_data)
# +
# # O means male, female who had not survived
# -
temp = titanic_data['Pclass'].head(10)
print(temp)
sns.countplot(x="Survived", hue="Pclass", data=titanic_data)
# +
# O means who had not survived from which class
# +
# # Analysis on Age column
# -
titanic_data['Age'].plot.hist()
titanic_data['Fare'].plot.hist()
titanic_data['Fare'].plot.hist(bins=20, figsize=(10,5))
print((titanic_data.info()))
sns.countplot(x="SibSp", data=titanic_data)
# ## Data Wrangling
# +
# Data Wrangling is to remove null or empty data, cleaning data set
# -
print((titanic_data.info()))
titanic_data.isnull()
titanic_data.isnull().sum()
sns.heatmap(titanic_data.isnull(), yticklabels==False)
sns.heatmap(titanic_data.isnull(), cmap="viridis")
sns.boxplot(x="Pclass", y="Age", data=titanic_data)
# # either drop that row or fill some value where value is null
titanic_data.head(5)
titanic_data.drop("Cabin", axis=1, inplace=True)
titanic_data.head(5)
sns.heatmap(titanic_data.isnull(), yticklabels=False, cbar=False)
titanic_data.drop("Age", axis=1, inplace=True)
sns.heatmap(titanic_data.isnull(), yticklabels=False, cbar=False)
titanic_data.isnull().sum()
titanic_data.drop("Embarked", axis=1, inplace=True)
sns.heatmap(titanic_data.isnull(), yticklabels=False, cbar=False)
pd.get_dummies(titanic_data['Embarked'])
titanic_data.isnull().sum()
pd.get_dummies(titanic_data['Sex'])
pd.get_dummies(titanic_data['Sex'], drop_first=True)
sex = pd.get_dummies(titanic_data['Sex'], drop_first=True)
sex.head(5)
embark = pd.get_dummies(titanic_data['Embarked'])
embark.head(5)
embark = pd.get_dummies(titanic_data['Embarked'], drop_first=True)
embark.head(5)
pcl = pd.get_dummies(titanic_data['Pclass'], drop_first=True)
pcl.head(5)
titanic_data = pd.concat([titanic_data,sex, embark, pcl], axis=1)
titanic_data.head(5)
titanic_data.drop(['Sex', 'Embarked', 'Name', 'PassengerId', 'Ticket'], axis=True, inplace=True)
titanic_data.head(5)
titanic_data.drop(['Pclass'], axis=True, inplace=True)
titanic_data.head(5)
# ## Train and Test
X = titanic_data.drop("Survived", axis=1)
y = titanic_data["Survived"]
titanic_data.head(5)
#from sklearn.cross_validation import train_test_split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=1)
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression()
logmodel.fit(X_train, y_train)
predictions = logmodel.predict(X_test)
from sklearn.metrics import classification_report
classification_report(y_test, predictions)
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, predictions)
from sklearn.metrics import accuracy_score
accuracy_score(y_test, predictions)
# ## Collecting Data
# ## Accuracy Check
|
logistic_regression_examples/logistic_regression_ex1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ONLOAD cells are called at beginning of the report
# ## Available global variables:
# * es: elasticsearch connection
# * replacementHT: a dictionary of replacement tags
# * report: the report object
# * params: the parameters (interval are plits in two name_start + name_end
# +
#@ONLOAD
replacementHT["Author"]="<NAME>"
replacementHT["GenerationDate"]=datetime.datetime.now().strftime("%d/%b/%y %H:%M")
replacementHT["Start"]=replacementHT["param2_start"].strftime("%d%b%y")
replacementHT["End"]=replacementHT["param2_end"].strftime("%d%b%y")
# +
#@ONLOAD
replacementHT["Apps"]=es_helper.elastic_to_dataframe(es,"nyx_app").shape[0]
replacementHT["Reports"]=es_helper.elastic_to_dataframe(es,"nyx_reportdef").shape[0]
replacementHT["Privileges"]=es_helper.elastic_to_dataframe(es,"nyx_privilege").shape[0]
replacementHT["Users"]=es_helper.elastic_to_dataframe(es,"nyx_user").shape[0]
# -
# # TEXT
# +
#@PARAGRAPH=TextParagraph
paragraph.text="Fair trade is an institutional arrangement designed to help producers in developing countries achieve better trading conditions. Members of the fair trade movement advocate the payment of higher prices to exporters, as well as improved social and environmental standards."
# +
#@PARAGRAPH=DynamicTable
df_privileges=es_helper.elastic_to_dataframe(es,"nyx_privilege")
df_privileges=df_privileges[["_id","name","value"]]
df_privileges.columns=["ID","Name","Value"]
create_table(paragraph,df_privileges, 'Grid Table 4 Accent 5')
#df_privileges=es_helper.elastic_to_dataframe(es,"nyx_user")
#df_privileges=df_privileges[["_id","name","value"]]
#df_privileges.columns=["ID","Name","Value"]
#create_table(paragraph,df_privileges, 'Grid Table 4 Accent 5')
# -
# # Charts
# +
#@PARAGRAPH=PieChart
# Pie chart, where the slices will be ordered and plotted counter-clockwise:
labels = 'Frogs2', 'Hogs', 'Dogs', 'Logs'
sizes = [15, 30, 45, 10]
explode = (0, 0.1, 0, 0) # only "explode" the 2nd slice (i.e. 'Hogs')
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',shadow=False, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.tight_layout()
plt.savefig("./tmp/piechart.png", dpi = 600)
run = paragraph.add_run()
run.add_picture("./tmp/piechart.png", width=Inches(5.6))
# +
#@PARAGRAPH=BarChart
objects = ('Python', 'C++', 'Java', 'Perl', 'Scala', 'Lisp')
y_pos = np.arange(len(objects))
performance = [10,8,6,4,2,1]
plt.bar(y_pos, performance, align='center', alpha=0.5)
plt.xticks(y_pos, objects)
plt.ylabel('Usage')
plt.title('Programming language usage')
plt.tight_layout()
plt.savefig("./tmp/barchart.png", dpi = 600)
run = paragraph.add_run()
run.add_picture("./tmp/barchart.png", width=Inches(3.6))
# -
# # BROL
# +
from elasticsearch import Elasticsearch as ES, RequestsHttpConnection as RC
import collections
import datetime
from elastic_helper import es_helper
host_params="http://esnodebal:9200"
es = ES(hosts=[host_params])
es.info()
replacementHT={}
|
sources/demoreports/demodocx.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Publications markdown generator for academicpages
#
# Takes a set of bibtex of publications and converts them for use with [vgonzalezd.github.io](vgonzalezd.github.io). This is an interactive Jupyter notebook ([see more info here](http://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/what_is_jupyter.html)).
#
# The core python code is also in `pubsFromBibs.py`.
# Run either from the `markdown_generator` folder after replacing updating the publist dictionary with:
# * bib file names
# * specific venue keys based on your bib file preferences
# * any specific pre-text for specific files
# * Collection Name (future feature)
#
# TODO: Make this work with other databases of citations,
# TODO: Merge this with the existing TSV parsing solution
from pybtex.database.input import bibtex
import pybtex.database.input.bibtex
from time import strptime
import string
import html
import os
import re
#todo: incorporate different collection types rather than a catch all publications, requires other changes to template
publist = {
"proceeding": {
"file" : "proceedings.bib",
"venuekey": "booktitle",
"venue-pretext": "In the proceedings of ",
"collection" : {"name":"publications",
"permalink":"/publication/"}
},
"journal":{
"file": "pubs.bib",
"venuekey" : "journal",
"venue-pretext" : "",
"collection" : {"name":"publications",
"permalink":"/publication/"}
}
}
# +
html_escape_table = {
"&": "&",
'"': """,
"'": "'"
}
def html_escape(text):
"""Produce entities within text."""
return "".join(html_escape_table.get(c,c) for c in text)
# -
for pubsource in publist:
parser = bibtex.Parser()
bibdata = parser.parse_file(publist[pubsource]["file"])
#loop through the individual references in a given bibtex file
for bib_id in bibdata.entries:
#reset default date
pub_year = "1900"
pub_month = "01"
pub_day = "01"
b = bibdata.entries[bib_id].fields
try:
pub_year = f'{b["year"]}'
#todo: this hack for month and day needs some cleanup
if "month" in b.keys():
if(len(b["month"])<3):
pub_month = "0"+b["month"]
pub_month = pub_month[-2:]
elif(b["month"] not in range(12)):
tmnth = strptime(b["month"][:3],'%b').tm_mon
pub_month = "{:02d}".format(tmnth)
else:
pub_month = str(b["month"])
if "day" in b.keys():
pub_day = str(b["day"])
pub_date = pub_year+"-"+pub_month+"-"+pub_day
#strip out {} as needed (some bibtex entries that maintain formatting)
clean_title = b["title"].replace("{", "").replace("}","").replace("\\","").replace(" ","-")
url_slug = re.sub("\\[.*\\]|[^a-zA-Z0-9_-]", "", clean_title)
url_slug = url_slug.replace("--","-")
md_filename = (str(pub_date) + "-" + url_slug + ".md").replace("--","-")
html_filename = (str(pub_date) + "-" + url_slug).replace("--","-")
#Build Citation from text
citation = ""
#citation authors - todo - add highlighting for primary author?
for author in bibdata.entries[bib_id].persons["author"]:
citation = citation+" "+author.first_names[0]+" "+author.last_names[0]+", "
#citation title
citation = citation + "\"" + html_escape(b["title"].replace("{", "").replace("}","").replace("\\","")) + ".\""
#add venue logic depending on citation type
venue = publist[pubsource]["venue-pretext"]+b[publist[pubsource]["venuekey"]].replace("{", "").replace("}","").replace("\\","")
citation = citation + " " + html_escape(venue)
citation = citation + ", " + pub_year + "."
## YAML variables
md = "---\ntitle: \"" + html_escape(b["title"].replace("{", "").replace("}","").replace("\\","")) + '"\n'
md += """collection: """ + publist[pubsource]["collection"]["name"]
md += """\npermalink: """ + publist[pubsource]["collection"]["permalink"] + html_filename
note = False
if "note" in b.keys():
if len(str(b["note"])) > 5:
md += "\nexcerpt: '" + html_escape(b["note"]) + "'"
note = True
md += "\ndate: " + str(pub_date)
md += "\nvenue: '" + html_escape(venue) + "'"
url = False
if "url" in b.keys():
if len(str(b["url"])) > 5:
md += "\npaperurl: '" + b["url"] + "'"
url = True
md += "\ncitation: '" + html_escape(citation) + "'"
md += "\n---"
## Markdown description for individual page
if note:
md += "\n" + html_escape(b["note"]) + "\n"
if url:
md += "\n[Access paper here](" + b["url"] + "){:target=\"_blank\"}\n"
else:
md += "\nUse [Google Scholar](https://scholar.google.com/scholar?q="+html.escape(clean_title.replace("-","+"))+"){:target=\"_blank\"} for full citation"
md_filename = os.path.basename(md_filename)
with open("../_publications/" + md_filename, 'w') as f:
f.write(md)
print(f'SUCESSFULLY PARSED {bib_id}: \"', b["title"][:60],"..."*(len(b['title'])>60),"\"")
# field may not exist for a reference
except KeyError as e:
print(f'WARNING Missing Expected Field {e} from entry {bib_id}: \"', b["title"][:30],"..."*(len(b['title'])>30),"\"")
continue
|
markdown_generator/PubsFromBib.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# We retrieve all the songs for each genre
wav_files = list(df["file_name"]) # listing files
genres=['Hip','Pop','Reggae','Mbalax']
genres_files={'Hip':[],
'Pop':[],
'Reggae':[],
'Mbalax':[]
}
for genre in genres :
for filename in wav_files:
if genre in filename:
genres_files[genre].append(filename)
print("taille de "+ genre+ "= "+str(len(genres_files[genre])))
# -
# retrieve only 20percent of each genre for testing
genres_files_test={'Hip':[],
'Pop':[],
'Reggae':[],
'Mbalax':[]
}
for genre in genres:
i=0
end=round(0.2*len(genres_files[genre]))
for filename in genres_files[genre]:
genres_files_test[genre].append(filename)
i+=1
if i==end :
break
print("taille de "+ genre+ "= "+str(len(genres_files_test[genre])))
# +
# list of test files
test_files=[]
for genre in genres:
test_files+=genres_files_test[genre]
len(test_files)
with open('testset_filenames.txt', 'w') as f:
for file in test_files:
f.write("%s\n" % file)
# +
# Train-test split of files # Same split as used for VGG model
with open('pred_probas/testset_filenames.txt', 'r') as f:
test_files = f.readlines()
test_files = [t.strip() for t in test_files]
indices = []
for t in test_files:
indices.append(df[df.file_name==t].index[0])
df_test = df.iloc[indices,:]
df.drop(labels=indices, inplace=True)
cl_weight = dict(pd.Series(df['label']).value_counts(normalize=True))
x_train = np.asarray(df[df.columns[1:-1]])
x_test = np.asarray(df_test[df.columns[1:-1]])
y_train = np.asarray(df['label'])
y_test = np.asarray(df_test['label'])
# -
|
Drafts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solution {-}
#
# Consider a random variable $X$ with an exponential probability function given as:
# \begin{equation*}
# f_X(x)=
# \begin{cases}
# e^{-x}, &x \geq 0 \\
# 0, &x < 0 \\
# \end{cases}
# \end{equation*}
#
# a) Compute $P(X \geq 2)$:
# +
from sympy import exp, integrate, symbols, oo
x = symbols('x')
PX2 = integrate(exp(-x), (x, 2, oo))
PX2
# -
# b) Compute $P(1 \leq X \leq 2)$:
P1X2 = integrate(exp(-x), (x, 1, 2))
P1X2
# + [markdown] tags=[]
# c) Compute $E(X)$, $E(X^2)$ and $Var(X)$:
# -
EX = integrate(x*exp(-x), (x, 0, oo))
EX
EX2 = integrate(x**2*exp(-x), (x, 0, oo))
EX2
VarX = EX2 - EX**2
VarX
|
Problem 1.22.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# ### Exercise 22: Intro to Matplotlib through a simple scatter plot
people = ['Ann','Brandon','Chen','David','Emily','Farook',
'Gagan','Hamish','Imran','Joseph','Katherine','Lily']
age = [21,12,32,45,37,18,28,52,5,40,48,15]
weight = [55,35,77,68,70,60,72,69,18,65,82,48]
height = [160,135,170,165,173,168,175,159,105,171,155,158]
plt.scatter(age,weight)
plt.show()
plt.figure(figsize=(8,6))
plt.title("Plot of Age vs. Weight (in kgs)",fontsize=20)
plt.xlabel("Age (years)",fontsize=16)
plt.ylabel("Weight (kgs)",fontsize=16)
plt.grid (True)
plt.ylim(0,100)
plt.xticks([i*5 for i in range(12)],fontsize=15)
plt.yticks(fontsize=15)
plt.scatter(x=age,y=weight,c='orange',s=150,edgecolors='k')
plt.text(x=20,y=85,s="Weights are more or less similar \nafter 18-20 years of age",fontsize=15)
plt.vlines(x=20,ymin=0,ymax=80,linestyles='dashed',color='blue',lw=3)
plt.legend(['Weight in kgs'],loc=2,fontsize=12)
plt.show()
# ### Exercise 23: Generating random numbers from a Uniform distribution
x = np.random.randint(1,10)
print(x)
x = np.random.randint(1,10,size=1)
print(x)
x = np.random.randint(1,6,size=10)
print(x)
x = 50+50*np.random.random(size=15)
x= x.round(decimals=2)
print(x)
x = np.random.rand(3,3)
print(x)
# ### Exercise 24: Generating random numbers from a Binomial distribution and Bar plot
x = np.random.binomial(10,0.6,size=8)
print(x)
plt.figure(figsize=(7,4))
plt.title("Number of successes in coin toss",fontsize=16)
plt.bar(left=np.arange(1,9),height=x)
plt.xlabel("Experiment number",fontsize=15)
plt.ylabel("Number of successes",fontsize=15)
plt.show()
# ### Exercise 25: Generating random numbers from Normal distribution and Histogram
x = np.random.normal()
print(x)
heights = np.random.normal(loc=155,scale=10,size=100)
plt.figure(figsize=(7,5))
plt.hist(heights,color='orange',edgecolor='k')
plt.title("Histogram of teen aged students's height",fontsize=18)
plt.xlabel("Height in cm",fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.show()
# ### Exercise 26: Calculation of descriptive statistics from a DataFrame
people_dict={'People':people,'Age':age,'Weight':weight,'Height':height}
people_df=pd.DataFrame(data=people_dict)
people_df
print(people_df['Age'].mean())
print(people_df['Height'].max())
print(people_df['Weight'].std())
np.percentile(people_df['Age'],25)
pcnt_75 = np.percentile(people_df['Age'],75)
pcnt_25 = np.percentile(people_df['Age'],25)
print("Inter-quartile range: ",pcnt_75-pcnt_25)
print(people_df.describe())
# ### Exercise 27: DataFrame even has built-in plotting utilities
people_df['Weight'].hist()
plt.show()
people_df.plot.scatter('Weight','Height',s=150,c='orange',edgecolor='k')
plt.grid(True)
plt.title("Weight vs. Height scatter plot",fontsize=18)
plt.xlabel("Weight (in kg)",fontsize=15)
plt.ylabel("Height (in cm)",fontsize=15)
plt.show()
|
Lesson03/Exercise42-47/Matplotlib and Descriptive Statistics.ipynb
|
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題2.33
;; 基本的なリスト操作のいくつかを集積として定義したものを以下に⽰す。
;; ⽋けている式を埋めて、完成させよ。
;;
;; (define (map p sequence)
;; (accumulate (lambda (x y) ⟨??⟩) nil sequence))
;; (define (append seq1 seq2)
;; (accumulate cons ⟨??⟩ ⟨??⟩))
;; (define (length sequence)
;; (accumulate ⟨??⟩ 0 sequence))
; 集積
(define (accumulate op initial sequence)
(if (null? sequence) initial
(op (car sequence) (accumulate op initial (cdr sequence)))))
;; +
; 元の定義
;(define (map proc items)
; (if (null? items) '()
; (cons (proc (car items)) (map proc (cdr items)))))
; mapの回答
(define (map proc items)
(accumulate (lambda (x y)
(cons (proc x) y)
)
'() items))
;; +
; mapの動作確認
; フィボナッチ数列の偶数のもののリスト
(define (fib n)
(cond ((= n 0) 0)
((= n 1) 1)
(else (+ (fib (- n 1)) (fib (- n 2))))
)
)
(map fib '(0 1 2 3 4 5 6 7 8 9 10))
;; +
; appendの回答
(define (append seq1 seq2)
(accumulate cons seq2 seq1)
)
; 動作確認
(append '(1 2 3 4) '(5 6 7))
;; +
; lengthの回答
(define (length sequence)
(accumulate (lambda (x y)
(+ 1 y)
)
0 sequence))
; 動作確認
(length '(1 2 3 4 5 6 7))
|
exercises/2.33.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd0bd4c67ef142469da7dc4d338a32ac40116904d26076b8e6aa587d80720bc6a2b
# ---
from datetime import datetime
starttime=datetime.strptime("20180301095416","%Y%m%d%H%M%S")
# print(starttime.day)
endtime = datetime.strptime("20180301100225","%Y%m%d%H%M%S")
# starttime, endtime = endtime, starttime
delta = endtime - starttime
min_total = round(delta.seconds / 60)
print(min_total)
# +
from datetime import datetime
import pandas as pd
# filename = "data/Subway_20180301.txt"
filename = "data/Subway_20190301_top100000.txt"
# res = {}
df = pd.read_csv(filename)
# -
def time_group(etime:datetime.time) -> int:
return etime.hour * 6 + (etime.minute // 10) + 1
# + tags=[]
for _, line in df.iterrows():
starttime = datetime.strptime(str(line["UpTime"]), "%Y%m%d%H%M%S")
endtime = datetime.strptime(str(line["DownTime"]), "%Y%m%d%H%M%S")
if starttime.day != 1 or endtime.day != 1:
continue
delta = endtime - starttime
if delta.days < 0:
continue
if delta.seconds > 7200:
continue
if _ > 100:
break
print(starttime,time_group(starttime))
# -
newindex = {}
for i in range(144):
hour = i // 6
dec_min = i % 6
newindex[i] = str(hour).rjust(2,'0') + ":" + str(dec_min * 10).rjust(2,'0') + '-' + str(hour).rjust(2,'0') + ":" + str(dec_min * 10 + 9).rjust(2,'0')
print(newindex)
# +
import pandas as pd
df = pd.read_csv("./out/PeopleInSubwayTime.csv")
data = [df.iloc[:,0].tolist(), df.iloc[:,1].tolist()]
df = pd.read_csv("./out/PeopleInSubwayCount.csv")
data = [df.iloc[:, 0].tolist(), df.iloc[:, 1].tolist()]
print(data)
# -
from mytool import csv2js
csv2js()
|
Visualization2/test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimizing Code: Common Books
# Let's go through an example scenario where we optimize some code to be more efficient. Say we are managing books for a store, and we want to find all the books published within the last two years about code. We have a file that lists all the ids of books published in the last two years, `books_published_last_two_years.txt`, as well as a file for all coding books, `all_coding_books.txt`.
# Here's what the first few lines of each file looks like.
# #### `books_published_last_two_years.txt`
# ```txt
# 1262771
# 9011996
# 2007022
# 9389522
# 8181760
# ...
# ```
# #### `all_coding_books.txt`
# ```txt
# 382944
# 483549
# 103957
# 590274
# 045832
# ...
# ```
# Since we want to find all the coding books published within the last two years, we'd want to find the book ids included in both of these files. Your coworker came up with one approach, and shows you this code to find the books in both files.
import time
import pandas as pd
import numpy as np
# +
with open('../dataset/books-published-last-two-years.txt') as f:
recent_books = f.read().split('\n')
with open('../dataset/all-coding-books.txt') as f:
coding_books = f.read().split('\n')
# +
start = time.time()
recent_coding_books = []
for book in recent_books:
if book in coding_books:
recent_coding_books.append(book)
print(len(recent_coding_books))
print('Duration: {} seconds'.format(time.time() - start))
# -
# Their strategy is to loop through each book in the first file, check if it's contained in the second file, and if so - add it to the final list. This makes sense and is an intuitive first approach. However, there are several things we can do to make this more efficient. Here are some tips.
# ### Tip #1: Use vector operations over loops when possible
# Numpy and pandas are your best friends for this. There are MANY cases in which you can replace loops with Numpy and pandas that use vector operations to make your computations a LOT faster. Sometimes there is a method that does exactly what you need. Other times, you need to be a little creative. This example in particular has a useful method you can use.
#
# Let me show you how I would approach this. No joke, I google: "how to find common elements in two Numpy arrays" and here are the results I get!
#
# In the Jupyter notebook quiz on the next page, use Numpy's `intersect1d` method to get the intersection of the `recent_books` and `coding_books` arrays. I'll give you this same notebook, and I'll put a cell right here with code to record the time it takes to run. Write your line of code in between these time start and time end lines.
# ```python
# start = time.time()
# recent_coding_books =
# print(len(recent_coding_books))
# print('Duration: {} seconds'.format(time.time() - start))
# ```
# ### Tip #2: Know your data structures and which methods are faster
# In addition to relying on Numpy and pandas, it's often good to double check whether there's a data structure or method in Python you can use to accomplish your task more effectively. For example, in this case do you recall a data structure in Python that stores a group of unique elements and can quickly compute intersections and unions of different groups? You can read more about why sets are more efficient than lists for this task in the link on the bottom of this page.
#
# Also, remember how I said I googled everything? Last time, I was googling how to find common elements in specifically Numpy arrays. But you can go more general and google something like "how to find common elements in two lists python" and you'll see posts like [this](https://stackoverflow.com/questions/2864842/common-elements-comparison-between-2-lists) that share and compare different answers. And you can see the set being introduced here.
#
# This seems to have a lot of great explanation and votes, but ultimately we should try different methods and compare their efficiency for our example. Because different methods perform differently in different contexts. So it's smart to always test for yourself. In the next cell of the Jupyter notebook, find out how long it takes to compute the common elements of `recent_books` and `coding_books` using Python's `set.intersection` method. Here again is some code to measure how long this takes.
# ```python
# start = time.time()
# recent_coding_books =
# print(len(recent_coding_books))
# print('Duration: {} seconds'.format(time.time() - start))
# ```
|
Udacity_AWS_ML_Foundations/codes/optimizing-code-common-books-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Bluelord/Kaggle_Courses/blob/main/03%20Data%20Visualization/06%20Choosing%20plot%20Types%20%26%20Styles.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VokHBcuCoAsd"
# # Choosing plot Types & Styles
#
# ---
#
#
#
# ---
#
#
# + [markdown] id="dGj0cFRFoGFi"
# ## Tutorial
#
# ---
# + [markdown] id="rt3ByzRInzke"
# ### What have you learned?
#
# <img src="https://imgur.com/2VmgDnF.png" height="500" width="1000" usemap="#plottingmap" />
# <map name="plottingmap">
# <area shape="rect" coords="262,342,402,476" href="https://www.kaggle.com/alexisbcook/hello-seaborn" title="EXAMPLE: sns.lineplot(data=my_data)">
# <area shape="rect" coords="8,75,154,200" href="https://www.kaggle.com/alexisbcook/bar-charts-and-heatmaps" title="EXAMPLE: sns.swarmplot(x=my_data['Column 1'], y=my_data['Column 2'])">
# <area shape="rect" coords="8,200,154,350" href="https://www.kaggle.com/alexisbcook/bar-charts-and-heatmaps" title="EXAMPLE: sns.regplot(x=my_data['Column 1'], y=my_data['Column 2'])">
# <area shape="rect" coords="8,350,154,500" href="https://www.kaggle.com/alexisbcook/bar-charts-and-heatmaps" title='EXAMPLE: sns.lmplot(x="Column 1", y="Column 2", hue="Column 3", data=my_data)'>
# <area shape="rect" coords="229,10,393,160" href="https://www.kaggle.com/alexisbcook/bar-charts-and-heatmaps" title="EXAMPLE: sns.scatterplot(x=my_data['Column 1'], y=my_data['Column 2'], hue=my_data['Column 3'])">
# <area shape="rect" coords="397,10,566,160" href="https://www.kaggle.com/alexisbcook/line-charts" title="EXAMPLE: sns.heatmap(data=my_data)">
# <area shape="rect" coords="565,10,711,160" href="https://www.kaggle.com/alexisbcook/line-charts" title="EXAMPLE: sns.barplot(x=my_data.index, y=my_data['Column'])">
# <area shape="rect" coords="780,55,940,210" href="https://www.kaggle.com/alexisbcook/scatter-plots" title="EXAMPLE: sns.jointplot(x=my_data['Column 1'], y=my_data['Column 2'], kind='kde')">
# <area shape="rect" coords="780,210,940,350" href="https://www.kaggle.com/alexisbcook/scatter-plots" title="EXAMPLE: sns.kdeplot(data=my_data['Column'], shade=True)">
# <area shape="rect" coords="780,360,1000,500" href="https://www.kaggle.com/alexisbcook/scatter-plots" title="EXAMPLE: sns.distplot(a=my_data['Column'], kde=False)">
# </map>
#
# + [markdown] id="O_6DZi-bolET"
# Dividing the chart types into three broad categories.<br/>
# - **Trends** - A trend is defined as a pattern of change.
# - `sns.lineplot` - **Line charts** are best to show trends over a period of time, and multiple lines can be used to show trends in more than one group.
# - **Relationship** - There are many different chart types that you can use to understand relationships between variables in your data.
# - `sns.barplot` - **Bar charts** are useful for comparing quantities corresponding to different groups.
# - `sns.heatmap` - **Heatmaps** can be used to find color-coded patterns in tables of numbers.
# - `sns.scatterplot` - **Scatter plots** show the relationship between two continuous variables; if color-coded, we can also show the relationship with a third [categorical variable](https://en.wikipedia.org/wiki/Categorical_variable).
# - `sns.regplot` - Including a **regression line** in the scatter plot makes it easier to see any linear relationship between two variables.
# - `sns.lmplot` - This command is useful for drawing multiple regression lines, if the scatter plot contains multiple, color-coded groups.
# - `sns.swarmplot` - **Categorical scatter plots** show the relationship between a continuous variable and a categorical variable.
# - **Distribution** - We visualize distributions to show the possible values that we can expect to see in a variable, along with how likely they are.
# - `sns.distplot` - **Histograms** show the distribution of a single numerical variable.
# - `sns.kdeplot` - **KDE plots** (or **2D KDE plots**) show an estimated, smooth distribution of a single numerical variable (or two numerical variables).
# - `sns.jointplot` - This command is useful for simultaneously displaying a 2D KDE plot with the corresponding KDE plots for each individual variable.
#
# + [markdown] id="PLjXsd_-o08L"
# ### Changing styles with seaborn
# + colab={"base_uri": "https://localhost:8080/"} id="XUDb_dqMpEhA" outputId="0e48ebd7-088e-462a-c6e3-29eb89c467ef"
from google.colab import drive
drive.mount('/content/drive')
# + _kg_hide-input=true _kg_hide-output=true id="lGeyHFCLnzkf"
import pandas as pd
pd.plotting.register_matplotlib_converters()
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="_4NPnS6unzkg" outputId="4efd2f3b-5b69-4e45-bb22-976717eb011a"
# Loadind data
spotify_data = pd.read_csv("/content/drive/MyDrive/Colab Notebooks/Kaggle_Courses/03 Data Visualization/spotify.csv", index_col="Date", parse_dates=True)
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# + [markdown] id="sXRQ5W6snzkh"
# We can quickly change the style of the figure to a different theme with only a single line of code.
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="8oZ2uO1dnzkh" outputId="a8b4bc29-b807-4583-ecc7-711e6df49dde"
# Change the style of the figure to the "dark" theme
sns.set_style("dark")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# + [markdown] id="y8FkqyDcnzkh"
# Seaborn has five different themes: (1)`"darkgrid"`, (2)`"whitegrid"`, (3)`"dark"`, (4)`"white"`, and (5)`"ticks"`, and you need only use a command similar to the one in the code cell above (with the chosen theme filled in) to change it.
# + [markdown] id="D-2zeEIEpvKU"
# ## Exercise
#
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="A_3yH0usqbh7" outputId="f9a904a1-ee91-4b09-ae8a-e954a249afbc"
# Change the style of the figure
sns.set_style("darkgrid")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
# + [markdown] id="Gdk4j6HCrNET"
# Now, try out different themes by amending the first line of code and running the code cell again. Remember the list of available themes:
#
# "darkgrid"<br/>
# "whitegrid"<br/>
# "dark"<br/>
# "white"<br/>
# "ticks"
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="D0rBNorIrMWz" outputId="3ee4d20e-6adc-4db8-842b-820a8b899ef2"
# Change the style of the figure
sns.set_style("ticks")
# Line chart
plt.figure(figsize=(12,6))
sns.lineplot(data=spotify_data)
|
03 Data Visualization/06 Choosing plot Types & Styles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''tfm1'': conda)'
# language: python
# name: python3
# ---
# +
from myutils import Datasets as ds
from myutils import Helpers as h
from myutils import NeuralNetwork as nn
import numpy as np
import tensorflow as tf
import itertools
from sklearn.decomposition import PCA
import os
#Magic Command, so changes in myutils module are reloaded
# %load_ext autoreload
# %autoreload 1
# %aimport myutils
FOLDER = "neuralnetwork"
NUMBER_RUNS = 3
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# -
x_train, y_train, x_test, y_test = ds.ALL_NUMBERS()
#reshape data
x_train = x_train.reshape(x_train.shape[0], 784)
x_test = x_test.reshape(x_test.shape[0], 784)
nn.run(preprocessing="pca",x_train=x_train,y_train=y_train,x_test=x_test,y_test=y_test,run_number=NUMBER_RUNS)
nn.run(preprocessing="raw",x_train=x_train,y_train=y_train,x_test=x_test,y_test=y_test,run_number=NUMBER_RUNS)
# +
title = "pca"
if os.path.exists("results/"+FOLDER+"/"+title+".json"):
os.remove("results/"+FOLDER+"/"+title+".json")
for run_number in range(NUMBER_RUNS):
for subset in itertools.combinations([i for i in range(10)],2):
print(subset)
tf.keras.backend.clear_session()
model = tf.keras.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(4, activation='relu'),
tf.keras.layers.Dense(2, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='adam',
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy'])
x_train_subset = x_train_pca[(y_train == subset[0]) | (y_train == subset[1])]
y_train_subset = y_train[(y_train == subset[0]) | (y_train == subset[1])]
y_train_subset = np.where(y_train_subset == subset[0], 0, y_train_subset)
y_train_subset = np.where(y_train_subset == subset[1], 1, y_train_subset)
x_test_subset = x_test_pca[(y_test == subset[0]) | (y_test == subset[1])]
y_test_subset = y_test[(y_test == subset[0]) | (y_test == subset[1])]
y_test_subset = np.where(y_test_subset == subset[0], 0, y_test_subset)
y_test_subset = np.where(y_test_subset == subset[1], 1, y_test_subset)
hist = model.fit(x_train_subset, y_train_subset, epochs=3, batch_size=32,validation_data=(x_test_subset, y_test_subset),verbose=0)
h.log_results(filename="results/"+FOLDER+"/"+title+".json",result={
str(run_number): {
str(subset):hist.history["val_accuracy"][-1]
}
})
del model
# -
# # Neural Network with Raw 28x28 Images
# +
title = "raw"
if os.path.exists("results/"+FOLDER+"/"+title+".json"):
os.remove("results/"+FOLDER+"/"+title+".json")
for run_number in range(NUMBER_RUNS):
for subset in itertools.combinations([i for i in range(10)],2):
print(run_number, subset)
tf.keras.backend.clear_session()
model = tf.keras.Sequential([
tf.keras.layers.Dense(4, activation='relu'),
tf.keras.layers.Dense(2, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='adam',
loss=tf.keras.losses.binary_crossentropy,
metrics=['accuracy'])
x_train_subset = x_train[(y_train == subset[0]) | (y_train == subset[1])] / 255
y_train_subset = y_train[(y_train == subset[0]) | (y_train == subset[1])]
y_train_subset = np.where(y_train_subset == subset[0], 0, y_train_subset)
y_train_subset = np.where(y_train_subset == subset[1], 1, y_train_subset)
x_test_subset = x_test[(y_test == subset[0]) | (y_test == subset[1])] / 255
y_test_subset = y_test[(y_test == subset[0]) | (y_test == subset[1])]
y_test_subset = np.where(y_test_subset == subset[0], 0, y_test_subset)
y_test_subset = np.where(y_test_subset == subset[1], 1, y_test_subset)
hist = model.fit(x_train_subset, y_train_subset, epochs=3, batch_size=32,validation_data=(x_test_subset, y_test_subset),verbose=0 )
h.log_results(filename="results/"+FOLDER+"/"+title+".json",result={
str(run_number): {
str(subset):hist.history["val_accuracy"][-1]
}
})
del model
# -
|
praktikum/backlog/NeuralNetwork.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Start by importing all the libraries we need
# %matplotlib inline
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
pd.DataFrame
# -
# ## Now, a quick review of what we covered last week
# * we are going to use the library "pandas" to read and manipulate data
# +
# I heard some folks had some errors doing this - lets check around, does this work for everybody?
buildings = pd.read_csv("/Users/jillnaiman1/Downloads/building_inventory.csv",
na_values = {'Year Acquired': 0,
'Year Constructed': 0,
'Square Footage': 0})
# -
# lets also print out what this looks like
buildings
# neat! a cool table!
# how many building entries do we have?
buildings.index
# we can look up a few agency names
# as a way to look at how we access data
buildings.loc[100:110]["Agency Name"]
# there are also ways to print out statistics
# of this data
# last week we used the "summary-like" function
buildings.describe()
# note here of course that things like talking
# about the "mean" congressional district doesn't make
# a whole lot of sense, but the "mean" of total floors
# might be of interest
# we can also do things like see how many unique agency names there are
buildings["Agency Name"].unique()
# we can also create new data objects
# here we use the "group by" subset to
# look at how much square footage each
# agency has
buildings.groupby("Agency Name")["Square Footage"].sum()
# we can use something like numpy to order these as well
sorted_index = np.argsort(buildings.groupby("Agency Name")["Square Footage"].sum())
# lets take a look
buildings.groupby("Agency Name")["Square Footage"].sum()[sorted_index]
# lets reverse sort so we can see top square footage users
buildings.groupby("Agency Name")["Square Footage"].sum()[sorted_index[::-1]]
# so we can see that U of I and the department of corrections are the
# front runners when it comes to square footage
# as we did last time,
# we can also look at how much square
# footage was created in different years
buildings.groupby("Year Acquired")["Square Footage"].sum()
# we can now also look at statistics of our subset
# from different years using "describe" as we've done before
stats = buildings.groupby("Year Acquired")["Square Footage"].describe()
# lets check out some data from the year 2010
stats.loc[2010]
# we can also check out the statistics by
# index, in this case we look at the very first
# entry
stats.iloc[0]
# since we have only the one entry, the STD is meaningless
# +
# alright, so this is in fact a data viz class
# so lets make some plots then!
# lets first start by plotting the maximum square footage
# of any buildings created in a particular year
# (1)
plt.plot(stats["max"], marker='.', linewidth = 1.0)
plt.xlabel("Year")
plt.ylabel("Square Footage")
# (2)
# we can see that there is a large range
# of square footages - from around 0 to
# millions. We'd like to see this better,
# so lets apply a "mutation" to our data
# in the form of how we visualize it
plt.yscale("log") # take the log scale on the yaxis
# now we can see a wider range of values
# from very small to very large on this log-scale
# what this visualization is showing is out of all the
# buildings built in a given year, what is the largest
# building (in square footage) that has been built
# -
# we can make a similar plot for the minimum value
# of the square footage of buildings built in
# a year
plt.plot(stats["min"], marker='.', linewidth = 1.0)
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
# ## Take away
# * so far we have now played a bit with how to read in a dataset, look at different values in nicely formatted pandas tables, or by hand by checking out different values at different indicies
# * We also started making some simple plots: the above plots show two different values from the same dataset - the maximum and minimum values for the square footage built in a year.
# * Now we are going to get into some methods to combine the plots of these two different values (the min & max square footage), and some ways to "fancyify" our plots with things like interactive widgets
# lets start by plotting both datasets on the same axis:
# (1)
plt.plot(stats["max"], marker='.', linewidth = 1.0)
plt.plot(stats["min"], marker='.', linewidth = 1.0)
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
# +
# lets start by plotting both datasets on the same axis:
# (2) ok, but which is which? Lets make a legend!
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# (3) but lets say what I really want to highlight in my visualization
# is to hightlight the difference between the maximum & minimum square
# footage of buildings aquired in a year - this might be a good measure of the
# diversity of buildings thate are built in a year, but there are
# certainly other factors
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#dddddd")
# so with this fill in we can see something interesting - as the
# years progress we are getting a greater diversity of buildings sq footage
# - the bigger things are getting bigger & smaller things getting smaller
# though, this difference seems to peak around 1975 (not sure why)
# we can also see that it seems like more square footage started being built
# in ~1850 - this makes sense since UIUC was founded in 1868
# there were also large expansions around the early 1900's
# Willard airport was completed in 1954 after a period of large expansion after WWII
# -
# ok, this figure is a little small & at low resolution
# lets embiggen it in the next round!
plt.rcParams["figure.dpi"] = 200
# for plotting on my zoomed in screen I'll make it smaller though
plt.rcParams["figure.dpi"] = 100
# +
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#dddddd")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# woohoo!
# -
# now, lets play with something called the "style" of the plot
# if you've used R, you might have used the "ggplot" package
# we can make plots in this style with Python too:
with plt.style.context("ggplot"):
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#aaaaaa")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# now, lets check out which plotting styles might be available
plt.style.available
# but what if we want to see how our plot would look with each
# of these styles? We could just make a bunch of plots
# OR we can play with this interactively with ipywidgets:
import ipywidgets
# +
# lets tell jupyter ipywidgets that we want to
# mess around with the style of the plot
@ipywidgets.interact(style = plt.style.available)
def make_plot(style):
with plt.style.context(style):
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#aaaaaa")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# so now you can see that we get a little dropdown menu that lists
# all the different styles!
# **play with this a bit!!**
# +
# ok, but what did we just do?
# we made something in Jupyter interactive
# we'll have a lot of opportunities to mess around with
# widgets in this class, but I'll just show a few
# quick examples. Read more on the docs:
# https://ipywidgets.readthedocs.io/en/stable/examples/Using%20Interact.html
# basically what ipywidgets.interact does is looks for a function with
# inputs and makes a little interactive option for those inputs,
# so like we did with the "make_plot" function above we can do
# for other things like change the value of a print statement:
@ipywidgets.interact(x=10)
def f(x):
print("my value = " + str(x))
# note this is a little different to the format in the
# docs and you can use what you'd like
# the "@" symbol is a "decorator" and essentially
# its a way to sort of "extend" the interact function
# without modifying it to much.
# At any rate, the take away is that you can call it
# like this, or how they do it in the docs, its up
# to you!
# -
# ok, lets go back to our original plot and look at a few more ways
# to fancify it
with plt.style.context("ggplot"):
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#aaaaaa")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# +
# maybe we want to highlight a particular measurement, lets say
# when UIUC was officially opened
# (1)
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#aaaaaa")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# (2) now, lets highligh our date of interest with a solid black line
plt.plot([1868, 1868], [10, 10**6], color = 'black', linewidth = 2.0)
# -
# so that is all well and good, but
# maybe we don't like the line being stubby
# and we want to extend it to the full length of the window
# we can do this by trying to figure out the min & max y
# for our plot, or we can transform between the window's
# coordinates & the plots with the matplotlib.transforms library
import matplotlib.transforms as mpt
# +
# just redoing plot here
# (1)
plt.plot(stats["max"], marker='.', linewidth = 1.0, label = "Max")
plt.plot(stats["min"], marker='.', linewidth = 1.0, label = "Min")
plt.fill_between(stats.index, stats["min"], stats["max"], color = "#aaaaaa")
plt.xlabel("Year")
plt.ylabel("Square Footage")
plt.yscale("log")
plt.legend()
# (2) now I've been using plt directly instead of
# building a figure with: fig,ax = plt.subplots(figsize=(sizex,sizey))
# but! I can still access the axes object:
ax = plt.gca()
# we'll need this to do our transform
# (3) now, I'm going to store a simple transform from the
# data space to the plot window space:
new_transform = mpt.blended_transform_factory(ax.transData, ax.transAxes)
# and now the call is very much the same but I apply the transformation
# and note that my y-extent is 0-1 which is the bottom of the plot window
# to the top
plt.plot([1868, 1868], [0.0, 1.0], color = 'black', linewidth = 2.0, transform = new_transform)
# we can see now that the line extends the full plot window
# for this example it would have been probably pretty easy to
# just guess what the max & min data axis where, but there
# can be cases where we want to highlight a certain region
# of the plot window for our visualization and in that
# case it is usually much easier to do it in window
# coordinates
# -
# ## Take away
# * We have now made some fancyfied plots. Hurray!
# * We have also seen how to use ipywidgets to make some aspects of our plots interactive
# * Next, we are going to apply these to some histograming options we discussed in lecture
#
# # Interactive binning & histogramming of data! Woo!
# +
# lets apply our ipywidet knowledge to rebinning the mean
# square footage as a function of year aquired from
# our building data
# we talked in the lecture about rebinning things, so lets mess with that!
# lets look at the mean of the square footage
mean_sq = stats["mean"]
mean_sq
# so, for every year this is telling us the mean sq footage
# of aquired buildings
# +
# now lets rebin this into a smaller number of time bins
# there are a few ways to do this, first lets do
# something simple by interpolating the array
# to a smaller year array
# first, lets choose the number of bins - was 171
nbins = 50 # 171/3 = 57
# now lets make an array to store our bin locations
# (1) first, grab all the years
years = buildings["Year Acquired"].unique()
years.sort()
# take out nans
years = years[~pd.isnull(years)]
years
# (2)
# then make new year bins
year_bins = np.linspace(years.min(),years.max(), nbins)
year_bins
# (3) rebin
mean_sq_avg = np.interp(year_bins, years, mean_sq)
mean_sq_avg.shape
# note: this is just doing a linear interpolation between points
# this is close to the (sum(values)/N) as discussed in the lecture
# -
# now, lets plot both!
plt.plot(years,mean_sq)
plt.plot(year_bins,mean_sq_avg)
plt.yscale("log")
plt.xlabel("Years")
plt.ylabel("Mean Square Footage")
# ok, but now lets make the rebinning interactive!
@ipywidgets.interact(nbins=(2,100,2)) # note, this is just saying bins from 2 to 100 in units of 2
def make_plot(nbins):
# then make new year bins
year_bins = np.linspace(years.min(),years.max(), nbins)
# rebin
mean_sq_avg = np.interp(year_bins, years, mean_sq)
# plot
plt.plot(years,mean_sq)
plt.plot(year_bins,mean_sq_avg)
plt.yscale("log")
plt.xlabel("Years")
plt.ylabel("Mean Square Footage")
# +
# we will cover smoothing a bit more in later lectures
# but we can do this too interactively
@ipywidgets.interact(window = (1, 10, 1))
def make_plot(window):
mean_sq_avg = mean_sq.rolling(window=window).mean()
plt.plot(mean_sq, marker = '.', linewidth = 0.5, alpha = 0.5)
plt.plot(mean_sq_avg, marker = '.', linewidth = 1.5)
plt.yscale("log")
# this is now choosing a rolling average over which to do
# an averaging - it takes nearby bins and averages
# this is like (sum(values)/N)
# mean_sq.rolling?
# note in the docs we can also specify a weighing for these
# we'll talk about the types of windows they list here
# (like hamming, triang, etc) later in the course probably
# -
# we can also look at these different windows directly
# here for example the sum over a rolling window of 3
buildings.groupby("Year Acquired")["Square Footage"].sum().rolling(3).sum()
buildings.keys()
|
week03/.ipynb_checkpoints/notebook_notes_week03-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Attack Password with Correlation Power Analysis IV.1 (CPA)
# + hide_input=true
# %run '../util/Metadata.ipynb'
print_metadata()
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Improving-the-code" data-toc-modified-id="Improving-the-code-1"><span class="toc-item-num">1 </span>Improving the code</a></span></li><li><span><a href="#Basic-Setup" data-toc-modified-id="Basic-Setup-2"><span class="toc-item-num">2 </span>Basic Setup</a></span></li><li><span><a href="#Helper-Functions-for-Password-Attack" data-toc-modified-id="Helper-Functions-for-Password-Attack-3"><span class="toc-item-num">3 </span>Helper Functions for Password Attack</a></span></li><li><span><a href="#New-Code---Old-results" data-toc-modified-id="New-Code---Old-results-4"><span class="toc-item-num">4 </span>New Code - Old results</a></span></li><li><span><a href="#Plotting-correlation-vs-traces" data-toc-modified-id="Plotting-correlation-vs-traces-5"><span class="toc-item-num">5 </span>Plotting correlation vs traces</a></span></li><li><span><a href="#Hardening-against-CPA" data-toc-modified-id="Hardening-against-CPA-6"><span class="toc-item-num">6 </span>Hardening against CPA</a></span><ul class="toc-item"><li><span><a href="#Random-start-point" data-toc-modified-id="Random-start-point-6.1"><span class="toc-item-num">6.1 </span>Random start point</a></span></li><li><span><a href="#Dummy-operations" data-toc-modified-id="Dummy-operations-6.2"><span class="toc-item-num">6.2 </span>Dummy operations</a></span></li></ul></li><li><span><a href="#Summary" data-toc-modified-id="Summary-7"><span class="toc-item-num">7 </span>Summary</a></span></li><li><span><a href="#Disconnect" data-toc-modified-id="Disconnect-8"><span class="toc-item-num">8 </span>Disconnect</a></span></li></ul></div>
# -
# In this example we want to discuss possibilities to fight against a CPA attack.
#
# ## Improving the code
#
# Let's first recap the password checking loop from `advanced-passwdcheck-xor`:
# ```c
# passbad = 0;
# for(uint8_t i = 0; i < sizeof(correct_passwd); i++){
# passbad |= correct_passwd[i] ^ passwd[i];
# }
# ```
#
# We revealed in the last example that aboves XOR generates a collision between known and determinable input data and secret data which shall be revealed by an attack.
#
# So, how to lower this correlation?
# ## Basic Setup
# Define Variables
# %run "../util/Init.ipynb"
# Build target and upload
TARGET = 'simpleserial-passwordcheck'
# %store TARGET
# %run "$HELPERSCRIPTS/Prepare.ipynb"
# Import helper functions
# %run "$HELPERSCRIPTS/Setup_Generic.ipynb"
scope.adc.samples = 500
# ## Helper Functions for Password Attack
# +
from bokeh.plotting import figure, show
from bokeh.io import output_notebook
from bokeh.models import CrosshairTool, Label
output_notebook()
# +
import warnings
import random
import tqdm
import numpy as np
password_length = 8
"""Number of bytes of password"""
random_length = 32
"""Number of bytes of random input"""
def capture(command, data):
scope.arm()
target.simpleserial_write(command, data)
ret = scope.capture()
i = 0
while not target.is_done():
i += 1
time.sleep(0.05)
if i > 100:
warnings.warn("Target did not finish operation")
return None
if ret:
warnings.warn("Timeout happened during capture")
return None
return scope.get_last_trace()
def target_set_random(random_input=None):
if random_input is not None:
rand = random_input()
else:
rand = bytes(random.choices(range(0, 256), k=random_length))
target.simpleserial_write('r', rand)
return rand
def target_set_password(password):
target.simpleserial_write('p', password)
return target.simpleserial_read('r', password_length)
def target_check_password(command, password):
target.simpleserial_write(command, password)
return bytes(target.simpleserial_read('r', 1))[0] == 0
def capture_random(command, size=500, random_input=None, attempt_input=None):
"""Collect size number of password attempts with fully random random data."""
traces = []
textins = []
rands = []
for _ in tqdm.tqdm_notebook(range(size)):
rands.append(target_set_random(random_input))
if attempt_input is not None:
pass_guess = attempt_input()
else:
pass_guess = bytes(random.choices(range(0, 256), k=password_length))
traces.append(capture(command, pass_guess))
textins.append(pass_guess)
return np.array(traces), textins, rands
# +
import numpy as np
import warnings
warnings.simplefilter('ignore')
HW = [bin(n).count("1") for n in range(0, 256)]
def hw(n):
return HW[n]
hw_vec = np.vectorize(hw)
def pearson(x: np.array, y: np.array):
x_mean = np.mean(x)
y_mean = np.mean(y)
return sum((x - x_mean) * (y - y_mean)) / np.sqrt(sum((x - x_mean) ** 2) * sum((y - y_mean) ** 2))
def pearson_helper_traces(traces):
return traces_diff, traces_squared
def pearson_pointwise(traces, intermediates):
n = len(intermediates)
d_traces = traces - np.einsum('ij->j', traces, dtype='float64', optimize='optimal') / np.double(n)
d_intermediates = intermediates - np.einsum('i->', intermediates, dtype='float64', optimize='optimal') / np.double(n)
tmp = np.einsum('ij,ij->j', d_traces, d_traces, optimize='optimal')
tmp *= np.einsum('i,i->', d_intermediates, d_intermediates, optimize='optimal')
return np.dot(d_intermediates, d_traces) / np.sqrt(tmp)
def pearson_pointwise_multi(traces, intermediates):
(n, t) = traces.shape
(_, m) = intermediates.shape
d_traces = traces - np.einsum('nt->t', traces, dtype='float64', optimize='optimal') / np.double(n)
d_intermediates = intermediates - np.einsum('nm->m', intermediates, dtype='float64', optimize='optimal') / np.double(n)
tmp1 = np.einsum('nm,nm->m', d_intermediates, d_intermediates, optimize='optimal')
tmp2 = np.einsum('nt,nt->t', d_traces, d_traces, optimize='optimal')
tmp = np.einsum('m,t->mt', tmp1, tmp2, optimize='optimal')
denominator = np.sqrt(tmp)
numerator = np.einsum('nm,nt->mt', d_intermediates, d_traces, optimize='optimal')
return np.nan_to_num(numerator / denominator)
# +
import math
import bokeh.palettes
import bokeh.transform
from bokeh.models import ColumnDataSource
def plot_correlation(correlations, color_palette=bokeh.palettes.Oranges6, **kw):
kw['height'] = kw.get('height', 300)
kw['y_range'] = kw.get('y_range', (-1, 1))
p = figure(sizing_mode='stretch_width', **kw)
p.vbar(
x='points',
top='corr',
width=1,
source=ColumnDataSource(data=dict(
points=range(len(correlations)),
corr=correlations,
abscorr=abs(correlations),
)),
color=bokeh.transform.linear_cmap(
field_name='abscorr',
palette=color_palette,
low=1,
high=0,
),
)
return p
def plot_correlation_vs_traces(
traces,
textins,
password_index=0,
trylist='abcdefghijklmnopqrstuvwxyz0123456789',
plotpoints=500,
):
# Compute data
plotpoints = min(plotpoints, len(traces))
data = np.zeros((len(trylist), plotpoints))
intermediates = np.array([[hw(attempt[password_index] ^ ord(guess)) for guess in trylist] for attempt in textins])
for i in range(0, plotpoints):
j = math.ceil(i / plotpoints * len(traces))
data[:, i] = np.max(np.abs(pearson_pointwise_multi(traces[:j, :], intermediates[:j, :])), axis=1)
source = {
'xs': len(data) * [list(range(0, len(traces), math.ceil(len(traces) / plotpoints)))],
'ys': [corr for corr in data],
'legend': list(trylist),
'color': math.ceil(len(data) / 20) * bokeh.palettes.Category20_20,
}
# Create figure
p = figure(sizing_mode='stretch_width', height=300, tooltips=[('char', '@legend'), ('corrleation', '$y')])
p.multi_line(xs='xs', ys='ys', color='color', source=source)
return p
# -
# ## New Code - Old results
password = b'<PASSWORD>'
target_set_password(password)
target_check_password('1', password)
# +
command = '1'
traces, textins, rands = capture_random(command=command, size=200)
correlations = pearson_pointwise(traces, np.array([hw(textin[0] ^ ord('i')) for textin in textins]))
print('max(correlations) = ', max(abs(correlations)))
show(plot_correlation(correlations))
# -
# We can observe that the aboves plot is exactly the same as in seen in the previous notebook.
#
# ## Plotting correlation vs traces
#
# In this example we took 200 traces and observed a high correlation coefficient. But, wouldn't last less traces?
# Therefore we want to introduce an additional plot which gives an answer to this question. The **correlation vs. traces** plot. It can also be found in the examples of ChipWhisperer.
show(plot_correlation_vs_traces(traces, textins))
# With this graph it is easy to figure out that around 50 - 100 traces are enough to get a reliable result.
#
# ## Hardening against CPA
#
# Let's recap the idea of the CPA: Somewhere in our code the piece of secret information (a character of the real password) *collides* with the some piece of information coming from extern (a character of the password attempt).
#
# Can we avoid this collision? Not really. But we can *hide* it: Realize that it is not necessary that the comparison of the password characters is done in the order of their position inside the password. For instance you could do the whole comparison reversed but still getting the same result.
#
# What would this mean for the CPA attack?
#
# ### Random start point
#
# In the following investigations we will use command `2`. The code looks like this:
# ```c
# uint8_t check_password_xor_randomstart(uint8_t *attempt)
# {
# uint8_t passbad = 0;
#
# uint8_t j = random_buffer[0] % sizeof(password);
#
# trigger_high();
# for (uint8_t i = 0; i < sizeof(password); i++)
# {
# passbad |= password[j] ^ attempt[j];
# j = (j + 1) % sizeof(password);
# }
# trigger_low();
#
# simpleserial_put('r', 1, &passbad);
#
# return passbad;
# }
# ```
# `j` is used as index for the current character and its starting value is given by the `random_buffer`. Usually random is (of course) generated by the microcontroller itself. But in our case it is easier to feed in randomness from outside to have full control.
#
# The following code block shows what happens if we give two possibilities for `j`. To achieve this, an explicit `random_function` is injected into the capturing process. This special function restricts the randomness of the first value to either 0 or 1. Thus, the loop starts either with the first or the second character.
# +
def random_startpoint(startpoint):
return lambda: bytes([random.randint(0, startpoint - 1)] + random.choices(range(256), k=random_length - 1))
command = '2'
target_check_password(command, password)
# -
traces, textins, rands = capture_random(command=command, size=200, random_input=random_startpoint(2))
show(plot_correlation_vs_traces(traces, textins))
# We observe that the after 200 traces the maximal correlation is around 0.5. In contrast to the plain loop where it was around 0.9. This effect can be used more intensively if the increase the possible initial positions.
traces, textins, rands = capture_random(command=command, size=200, random_input=random_startpoint(3))
show(plot_correlation_vs_traces(traces, textins))
# If the first character has 3 possible indices it is impossible after 200 traces to achieve a reliable prediction. Is the loop secure now? Again: No! We just need more traces as the following plot shows.
traces, textins, rands = capture_random(command=command, size=1000, random_input=random_startpoint(3))
show(plot_correlation_vs_traces(traces, textins))
traces, textins, rands = capture_random(command=command, size=20000, random_input=random_startpoint(8))
show(plot_correlation_vs_traces(traces, textins))
# If the starting point of the loop can use all possibilities out of the length of the password it is hard to achieve a reliable prediction after 20000 traces, but it is still possible.
#
# ### Dummy operations
#
# So, do we need a longer password? Not necessarily. We just insert *dummy operations*. There are (in our case) 8 XOR operations which operate on the password and the attempt. By inserting a certain number of XOR operations which do not operate on the password or the attempt we could lower the probability of an index to proceed a certain position even more.
#
# It is a common technique to hide real operations between a (high) number of dummy operations. There are some requirements on these dummy operations:
# 1. In a power trace it must be impossible to distinguish between dummy operations and real ones.
# 2. The computational result of the algorithm must stay the same.
#
# In most cases it is quite hard to write down code with dummy operations satisfying the above conditions. For our password check there is an elegant way to incorporate dummy operations by just extending the buffers to be compared by some random values.
#
# ```c
# uint8_t check_password_xor_randomstart_random_buffer(uint8_t *attempt)
# {
# uint8_t passbad = 0;
# uint8_t buffer1[sizeof(random_buffer)], buffer2[sizeof(random_buffer)];
#
# // Copy random input
# memcpy(buffer1, random_buffer, sizeof(random_buffer));
# memcpy(buffer2, random_buffer, sizeof(random_buffer));
# // Copy password and attempt
# memcpy(buffer1, password, sizeof(password));
# memcpy(buffer2, attempt, sizeof(password));
#
# uint8_t j = random_buffer[0] % sizeof(random_buffer);
#
# trigger_high();
# for (uint8_t i = 0; i < sizeof(random_buffer); i++)
# {
# passbad |= buffer1[j] ^ buffer2[j];
# j = (j + 1) % sizeof(random_buffer);
# }
# trigger_low();
#
# simpleserial_put('r', 1, &passbad);
#
# return passbad;
# }
# ```
command = '3'
target_check_password(command, password)
traces, textins, rands = capture_random(command=command, size=20000)
show(plot_correlation_vs_traces(traces, textins))
# Now every index has a probability of 1/32 to operate on a certain character. This lowers down the correlation so that even after 20000 traces it is not possible to give a reliable prediction.
#
# These examples emphasize the main point of (side-channel) security: Usually nothing is safe against all attacks. It is just a matter of effort to break. And from a security software developers point of view it is an illusion to program anything absolute secure. The target is to achieve the *required amount of security*.
# ## Summary
#
# In this notebook we observed:
#
# * **Randomizing** the execution **order** of critical operations is an excellent way of lowering the strength of CPA.
# * If a random order is not sufficient or not applicable (e.g. AES) it helps to hide the critical operations between a certain number of **dummy operations**.
# ## Disconnect
scope.dis()
target.dis()
|
passwordcheck/Passwordcheck_Attack_4_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import matplotlib
import numpy as np
import pandas as pd
import seaborn as sns
import time
from datetime import date
from matplotlib import pyplot as plt
from pylab import rcParams
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
from tqdm import tqdm_notebook
from xgboost import XGBRegressor
# %matplotlib inline
#### Input params ##################
stk_path = "D:/Power BI/ABIN.csv"
test_size = 0.2 # proportion of dataset to be used as test set
cv_size = 0.2 # proportion of dataset to be used as cross-validation set
N = 3 # for feature at day t, we use lags from t-1, t-2, ..., t-N as features
n_estimators = 100 # Number of boosted trees to fit. default = 100
max_depth = 3 # Maximum tree depth for base learners. default = 3
learning_rate = 0.1 # Boosting learning rate (xgb’s “eta”). default = 0.1
min_child_weight = 1 # Minimum sum of instance weight(hessian) needed in a child. default = 1
subsample = 1 # Subsample ratio of the training instance. default = 1
colsample_bytree = 1 # Subsample ratio of columns when constructing each tree. default = 1
colsample_bylevel = 1 # Subsample ratio of columns for each split, in each level. default = 1
gamma = 0 # Minimum loss reduction required to make a further partition on a leaf node of the tree. default=0
model_seed = 100
fontsize = 14
ticklabelsize = 14
# +
def get_mov_avg_std(df, col, N):
"""
Given a dataframe, get mean and std dev at timestep t using values from t-1, t-2, ..., t-N.
Inputs
df : dataframe. Can be of any length.
col : name of the column you want to calculate mean and std dev
N : get mean and std dev at timestep t using values from t-1, t-2, ..., t-N
Outputs
df_out : same as df but with additional column containing mean and std dev
"""
mean_list = df[col].rolling(window = N, min_periods=1).mean() # len(mean_list) = len(df)
std_list = df[col].rolling(window = N, min_periods=1).std() # first value will be NaN, because normalized by N-1
# Add one timestep to the predictions
mean_list = np.concatenate((np.array([np.nan]), np.array(mean_list[:-1])))
std_list = np.concatenate((np.array([np.nan]), np.array(std_list[:-1])))
# Append mean_list to df
df_out = df.copy()
df_out[col + '_mean'] = mean_list
df_out[col + '_std'] = std_list
return df_out
def scale_row(row, feat_mean, feat_std):
"""
Given a pandas series in row, scale it to have 0 mean and var 1 using feat_mean and feat_std
Inputs
row : pandas series. Need to scale this.
feat_mean: mean
feat_std : standard deviation
Outputs
row_scaled : pandas series with same length as row, but scaled
"""
# If feat_std = 0 (this happens if adj_close doesn't change over N days),
# set it to a small number to avoid division by zero
feat_std = 0.001 if feat_std == 0 else feat_std
row_scaled = (row-feat_mean) / feat_std
return row_scaled
def get_mape(y_true, y_pred):
"""
Compute mean absolute percentage error (MAPE)
"""
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def train_pred_eval_model(X_train_scaled, \
y_train_scaled, \
X_test_scaled, \
y_test, \
col_mean, \
col_std, \
seed=100, \
n_estimators=100, \
max_depth=3, \
learning_rate=0.1, \
min_child_weight=1, \
subsample=1, \
colsample_bytree=1, \
colsample_bylevel=1, \
gamma=0):
'''
Train model, do prediction, scale back to original range and do evaluation
Use XGBoost here.
Inputs
X_train_scaled : features for training. Scaled to have mean 0 and variance 1
y_train_scaled : target for training. Scaled to have mean 0 and variance 1
X_test_scaled : features for test. Each sample is scaled to mean 0 and variance 1
y_test : target for test. Actual values, not scaled.
col_mean : means used to scale each sample of X_test_scaled. Same length as X_test_scaled and y_test
col_std : standard deviations used to scale each sample of X_test_scaled. Same length as X_test_scaled and y_test
seed : model seed
n_estimators : number of boosted trees to fit
max_depth : maximum tree depth for base learners
learning_rate : boosting learning rate (xgb’s “eta”)
min_child_weight : minimum sum of instance weight(hessian) needed in a child
subsample : subsample ratio of the training instance
colsample_bytree : subsample ratio of columns when constructing each tree
colsample_bylevel : subsample ratio of columns for each split, in each level
gamma :
Outputs
rmse : root mean square error of y_test and est
mape : mean absolute percentage error of y_test and est
est : predicted values. Same length as y_test
'''
model = XGBRegressor(seed=model_seed,
n_estimators=n_estimators,
max_depth=max_depth,
learning_rate=learning_rate,
min_child_weight=min_child_weight,
subsample=subsample,
colsample_bytree=colsample_bytree,
colsample_bylevel=colsample_bylevel,
gamma=gamma)
# Train the model
model.fit(X_train_scaled, y_train_scaled)
# Get predicted labels and scale back to original range
est_scaled = model.predict(X_test_scaled)
est = est_scaled * col_std + col_mean
# Calculate RMSE
rmse = math.sqrt(mean_squared_error(y_test, est))
mape = get_mape(y_test, est)
return rmse, mape, est
# +
df = pd.read_csv(stk_path, sep = ",")
# Convert Date column to datetime
df.loc[:, 'Date'] = pd.to_datetime(df['Date'],format='%Y-%m-%d')
# Change all column headings to be lower case, and remove spacing
df.columns = [str(x).lower().replace(' ', '_') for x in df.columns]
# Get month of each sample
df['month'] = df['date'].dt.month
# Sort by datetime
df.sort_values(by='date', inplace=True, ascending=True)
df.head()
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
ax = df.plot(x='date', y='adj_close', style='b-', grid=True)
ax.set_xlabel("Date")
ax.set_ylabel("EUR")
# +
# Get difference between high and low of each day
df['range_hl'] = df['high'] - df['low']
df.drop(['high', 'low'], axis=1, inplace=True)
# Get difference between open and close of each day
df['range_oc'] = df['open'] - df['close']
df.drop(['open', 'close'], axis=1, inplace=True)
df.head()
# +
# Add a column 'order_day' to indicate the order of the rows by date
df['order_day'] = [x for x in list(range(len(df)))]
# merging_keys
merging_keys = ['order_day']
# List of columns that we will use to create lags
lag_cols = ['adj_close', 'range_hl', 'range_oc', 'volume']
lag_cols
# +
shift_range = [x+1 for x in range(N)]
for shift in tqdm_notebook(shift_range):
train_shift = df[merging_keys + lag_cols].copy()
# E.g. order_day of 0 becomes 1, for shift = 1.
# So when this is merged with order_day of 1 in df, this will represent lag of 1.
train_shift['order_day'] = train_shift['order_day'] + shift
foo = lambda x: '{}_lag_{}'.format(x, shift) if x in lag_cols else x
train_shift = train_shift.rename(columns=foo)
df = pd.merge(df, train_shift, on=merging_keys, how='left') #.fillna(0)
del train_shift
# Remove the first N rows which contain NaNs
df = df[N:]
df.head()
# -
df.info()
# +
cols_list = [
"adj_close",
"range_hl",
"range_oc",
"volume"
]
for col in cols_list:
df = get_mov_avg_std(df, col, N)
df.head()
# +
# Get sizes of each of the datasets
num_cv = int(cv_size*len(df))
num_test = int(test_size*len(df))
num_train = len(df) - num_cv - num_test
print("num_train = " + str(num_train))
print("num_cv = " + str(num_cv))
print("num_test = " + str(num_test))
# Split into train, cv, and test
train = df[:num_train]
cv = df[num_train:num_train+num_cv]
train_cv = df[:num_train+num_cv]
test = df[num_train+num_cv:]
print("train.shape = " + str(train.shape))
print("cv.shape = " + str(cv.shape))
print("train_cv.shape = " + str(train_cv.shape))
print("test.shape = " + str(test.shape))
# +
cols_to_scale = [
"adj_close"
]
for i in range(1,N+1):
cols_to_scale.append("adj_close_lag_"+str(i))
cols_to_scale.append("range_hl_lag_"+str(i))
cols_to_scale.append("range_oc_lag_"+str(i))
cols_to_scale.append("volume_lag_"+str(i))
# Do scaling for train set
# Here we only scale the train dataset, and not the entire dataset to prevent information leak
scaler = StandardScaler()
train_scaled = scaler.fit_transform(train[cols_to_scale])
print("scaler.mean_ = " + str(scaler.mean_))
print("scaler.var_ = " + str(scaler.var_))
print("train_scaled.shape = " + str(train_scaled.shape))
# Convert the numpy array back into pandas dataframe
train_scaled = pd.DataFrame(train_scaled, columns=cols_to_scale)
train_scaled[['date', 'month']] = train.reset_index()[['date', 'month']]
print("train_scaled.shape = " + str(train_scaled.shape))
train_scaled.head()
# +
# Do scaling for train+dev set
scaler_train_cv = StandardScaler()
train_cv_scaled = scaler_train_cv.fit_transform(train_cv[cols_to_scale])
print("scaler_train_cv.mean_ = " + str(scaler_train_cv.mean_))
print("scaler_train_cv.var_ = " + str(scaler_train_cv.var_))
print("train_cv_scaled.shape = " + str(train_cv_scaled.shape))
# Convert the numpy array back into pandas dataframe
train_cv_scaled = pd.DataFrame(train_cv_scaled, columns=cols_to_scale)
train_cv_scaled[['date', 'month']] = train_cv.reset_index()[['date', 'month']]
print("train_cv_scaled.shape = " + str(train_cv_scaled.shape))
train_cv_scaled.head()
# +
# Do scaling for dev set
cv_scaled = cv[['date']]
for col in tqdm_notebook(cols_list):
feat_list = [col + '_lag_' + str(shift) for shift in range(1, N+1)]
temp = cv.apply(lambda row: scale_row(row[feat_list], row[col+'_mean'], row[col+'_std']), axis=1)
cv_scaled = pd.concat([cv_scaled, temp], axis=1)
# Now the entire dev set is scaled
cv_scaled.head()
# +
# Do scaling for test set
test_scaled = test[['date']]
for col in tqdm_notebook(cols_list):
feat_list = [col + '_lag_' + str(shift) for shift in range(1, N+1)]
temp = test.apply(lambda row: scale_row(row[feat_list], row[col+'_mean'], row[col+'_std']), axis=1)
test_scaled = pd.concat([test_scaled, temp], axis=1)
# Now the entire test set is scaled
test_scaled.head()
# +
features = []
for i in range(1,N+1):
features.append("adj_close_lag_"+str(i))
features.append("range_hl_lag_"+str(i))
features.append("range_oc_lag_"+str(i))
features.append("volume_lag_"+str(i))
target = "adj_close"
# Split into X and y
X_train = train[features]
y_train = train[target]
X_cv = cv[features]
y_cv = cv[target]
X_train_cv = train_cv[features]
y_train_cv = train_cv[target]
X_sample = test[features]
y_sample = test[target]
print("X_train.shape = " + str(X_train.shape))
print("y_train.shape = " + str(y_train.shape))
print("X_cv.shape = " + str(X_cv.shape))
print("y_cv.shape = " + str(y_cv.shape))
print("X_train_cv.shape = " + str(X_train_cv.shape))
print("y_train_cv.shape = " + str(y_train_cv.shape))
print("X_sample.shape = " + str(X_sample.shape))
print("y_sample.shape = " + str(y_sample.shape))
# -
# Split into X and y
X_train_scaled = train_scaled[features]
y_train_scaled = train_scaled[target]
X_cv_scaled = cv_scaled[features]
X_train_cv_scaled = train_cv_scaled[features]
y_train_cv_scaled = train_cv_scaled[target]
X_sample_scaled = test_scaled[features]
print("X_train_scaled.shape = " + str(X_train_scaled.shape))
print("y_train_scaled.shape = " + str(y_train_scaled.shape))
print("X_cv_scaled.shape = " + str(X_cv_scaled.shape))
print("X_train_cv_scaled.shape = " + str(X_train_cv_scaled.shape))
print("y_train_cv_scaled.shape = " + str(y_train_cv_scaled.shape))
print("X_sample_scaled.shape = " + str(X_sample_scaled.shape))
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
ax = train.plot(x='date', y='adj_close', style='b-', grid=True)
ax = cv.plot(x='date', y='adj_close', style='y-', grid=True, ax=ax)
ax = test.plot(x='date', y='adj_close', style='g-', grid=True, ax=ax)
ax.legend(['train', 'dev', 'test'])
ax.set_xlabel("date")
ax.set_ylabel("USD")
ax.set_title("Without scaling")
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
ax = train_scaled.plot(x='date', y='adj_close', style='b-', grid=True)
ax.legend(['train_scaled'])
ax.set_xlabel("date")
ax.set_ylabel("USD (scaled)")
ax.set_title("With scaling")
# +
# Create the model
model = XGBRegressor(seed=model_seed,
n_estimators=n_estimators,
max_depth=max_depth,
learning_rate=learning_rate,
min_child_weight=min_child_weight,
subsample=subsample,
colsample_bytree=colsample_bytree,
colsample_bylevel=colsample_bylevel,
gamma=gamma)
# Train the regressor
model.fit(X_train_scaled, y_train_scaled)
# +
# Do prediction on train set
est_scaled = model.predict(X_train_scaled)
est = est_scaled * math.sqrt(scaler.var_[0]) + scaler.mean_[0]
# Calculate RMSE
print("RMSE on train set = %0.3f" % math.sqrt(mean_squared_error(y_train, est)))
# Calculate MAPE
print("MAPE on train set = %0.3f%%" % get_mape(y_train, est))
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
est_df = pd.DataFrame({'est': est,
'date': train['date']})
ax = train.plot(x='date', y='adj_close', style='b-', grid=True)
ax = cv.plot(x='date', y='adj_close', style='y-', grid=True, ax=ax)
ax = test.plot(x='date', y='adj_close', style='g-', grid=True, ax=ax)
ax = est_df.plot(x='date', y='est', style='r-', grid=True, ax=ax)
ax.legend(['train', 'dev', 'test', 'predictions'])
ax.set_xlabel("date")
ax.set_ylabel("EUR")
ax.set_title('Without scaling')
# +
# Do prediction on test set
est_scaled = model.predict(X_cv_scaled)
cv['est_scaled'] = est_scaled
cv['est'] = cv['est_scaled'] * cv['adj_close_std'] + cv['adj_close_mean']
# Calculate RMSE
rmse_bef_tuning = math.sqrt(mean_squared_error(y_cv, cv['est']))
print("RMSE on dev set = %0.3f" % rmse_bef_tuning)
# Calculate MAPE
mape_bef_tuning = get_mape(y_cv, cv['est'])
print("MAPE on dev set = %0.3f%%" % mape_bef_tuning)
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
est_df = pd.DataFrame({'est': cv['est'],
'y_cv': y_cv,
'date': cv['date']})
ax = train.plot(x='date', y='adj_close', style='b-', grid=True)
ax = cv.plot(x='date', y='adj_close', style='y-', grid=True, ax=ax)
ax = test.plot(x='date', y='adj_close', style='g-', grid=True, ax=ax)
ax = est_df.plot(x='date', y='est', style='r-', grid=True, ax=ax)
ax.legend(['train', 'dev', 'test', 'predictions'])
ax.set_xlabel("date")
ax.set_ylabel("EUR")
# -
# Plot adjusted close over time, for dev set only
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
ax = train.plot(x='date', y='adj_close', style='b-', grid=True)
ax = cv.plot(x='date', y='adj_close', style='y-', grid=True, ax=ax)
ax = test.plot(x='date', y='adj_close', style='g-', grid=True, ax=ax)
ax = est_df.plot(x='date', y='est', style='r-', grid=True, ax=ax)
ax.legend(['train', 'dev', 'test', 'predictions'])
ax.set_xlabel("date")
ax.set_ylabel("USD")
ax.set_xlim([date(2017, 8, 1), date(2018, 5, 31)])
ax.set_title("Zoom in to dev set")
# View a list of the features and their importance scores
imp = list(zip(train[features], model.feature_importances_))
imp.sort(key=lambda tup: tup[1])
imp[-10:]
d = {'N': [2, 3, 4, 5, 6, 7, 14],
'rmse_dev_set': [1.225, 1.214, 1.231, 1.249, 1.254, 1.251, 1.498],
'mape_pct_dev_set': [0.585, 0.581, 0.590, 0.601, 0.609, 0.612, 0.763]}
pd.DataFrame(d)
# +
param_label = 'n_estimators'
param_list = range(10, 310, 10)
param2_label = 'max_depth'
param2_list = [2, 3, 4, 5, 6, 7, 8, 9]
error_rate = {param_label: [] , param2_label: [], 'rmse': [], 'mape_pct': []}
tic = time.time()
for param in tqdm_notebook(param_list):
# print("param = " + str(param))
for param2 in param2_list:
# Train, predict and eval model
rmse, mape, _ = train_pred_eval_model(X_train_scaled,
y_train_scaled,
X_cv_scaled,
y_cv,
cv['adj_close_mean'],
cv['adj_close_std'],
seed=model_seed,
n_estimators=param,
max_depth=param2,
learning_rate=learning_rate,
min_child_weight=min_child_weight,
subsample=subsample,
colsample_bytree=colsample_bytree,
colsample_bylevel=colsample_bylevel,
gamma=gamma)
# Collect results
error_rate[param_label].append(param)
error_rate[param2_label].append(param2)
error_rate['rmse'].append(rmse)
error_rate['mape_pct'].append(mape)
error_rate = pd.DataFrame(error_rate)
toc = time.time()
print("Minutes taken = " + str((toc-tic)/60.0))
error_rate
# +
# Plot performance versus params
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
temp = error_rate[error_rate[param2_label]==param2_list[0]]
ax = temp.plot(x=param_label, y='rmse', style='bs-', grid=True)
legend_list = [param2_label + '_' + str(param2_list[0])]
color_list = ['r', 'g', 'k', 'y', 'm', 'c', '0.75']
for i in range(1,len(param2_list)):
temp = error_rate[error_rate[param2_label]==param2_list[i]]
ax = temp.plot(x=param_label, y='rmse', color=color_list[i%len(color_list)], marker='s', grid=True, ax=ax)
legend_list.append(param2_label + '_' + str(param2_list[i]))
ax.set_xlabel(param_label)
ax.set_ylabel("RMSE")
matplotlib.rcParams.update({'font.size': 14})
plt.legend(legend_list, loc='center left', bbox_to_anchor=(1.0, 0.5)) # positions legend outside figure
# -
# Get optimum value for param and param2
temp = error_rate[error_rate['rmse'] == error_rate['rmse'].min()]
n_estimators_opt = temp['n_estimators'].values[0]
max_depth_opt = temp['max_depth'].values[0]
print("min RMSE = %0.3f" % error_rate['rmse'].min())
print("optimum params = ")
n_estimators_opt, max_depth_opt
# Get optimum value for param and param2, using MAPE
temp = error_rate[error_rate['mape_pct'] == error_rate['mape_pct'].min()]
print("min MAPE = %0.3f%%" % error_rate['mape_pct'].min())
print("optimum params = ")
temp['n_estimators'].values[0], temp['max_depth'].values[0]
# +
param_label = 'learning_rate'
param_list = [0.001, 0.005, 0.01, 0.05, 0.1, 0.2, 0.3]
param2_label = 'min_child_weight'
param2_list = range(5, 21, 1)
error_rate = {param_label: [] , param2_label: [], 'rmse': [], 'mape_pct': []}
tic = time.time()
for param in tqdm_notebook(param_list):
# print("param = " + str(param))
for param2 in param2_list:
# Train, predict and eval model
rmse, mape, _ = train_pred_eval_model(X_train_scaled,
y_train_scaled,
X_cv_scaled,
y_cv,
cv['adj_close_mean'],
cv['adj_close_std'],
seed=model_seed,
n_estimators=n_estimators_opt,
max_depth=max_depth_opt,
learning_rate=param,
min_child_weight=param2,
subsample=subsample,
colsample_bytree=colsample_bytree,
colsample_bylevel=colsample_bylevel,
gamma=gamma)
# Collect results
error_rate[param_label].append(param)
error_rate[param2_label].append(param2)
error_rate['rmse'].append(rmse)
error_rate['mape_pct'].append(mape)
error_rate = pd.DataFrame(error_rate)
toc = time.time()
print("Minutes taken = " + str((toc-tic)/60.0))
error_rate
# +
# Plot performance versus params
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
temp = error_rate[error_rate[param2_label]==param2_list[0]]
ax = temp.plot(x=param_label, y='rmse', style='bs-', grid=True)
legend_list = [param2_label + '_' + str(param2_list[0])]
color_list = ['r', 'g', 'k', 'y', 'm', 'c', '0.75']
for i in range(1,len(param2_list)):
temp = error_rate[error_rate[param2_label]==param2_list[i]]
ax = temp.plot(x=param_label, y='rmse', color=color_list[i%len(color_list)], marker='s', grid=True, ax=ax)
legend_list.append(param2_label + '_' + str(param2_list[i]))
ax.set_xlabel(param_label)
ax.set_ylabel("RMSE")
matplotlib.rcParams.update({'font.size': 14})
plt.legend(legend_list, loc='center left', bbox_to_anchor=(1.0, 0.5)) # positions legend outside figure
# -
# Get optimum value for param and param2
temp = error_rate[error_rate['rmse'] == error_rate['rmse'].min()]
learning_rate_opt = temp['learning_rate'].values[0]
min_child_weight_opt = temp['min_child_weight'].values[0]
print("min RMSE = %0.3f" % error_rate['rmse'].min())
print("optimum params = ")
learning_rate_opt, min_child_weight_opt
# Get optimum value for param and param2, using MAPE
# We will use RMSE to decide the final optimum params to use
temp = error_rate[error_rate['mape_pct'] == error_rate['mape_pct'].min()]
print("min MAPE = %0.3f%%" % error_rate['mape_pct'].min())
print("optimum params = ")
temp['learning_rate'].values[0], temp['min_child_weight'].values[0]
# +
param_label = 'subsample'
param_list = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1]
param2_label = 'gamma'
param2_list = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9, 1]
error_rate = {param_label: [] , param2_label: [], 'rmse': [], 'mape_pct': []}
tic = time.time()
for param in tqdm_notebook(param_list):
# print("param = " + str(param))
for param2 in param2_list:
# Train, predict and eval model
rmse, mape, _ = train_pred_eval_model(X_train_scaled,
y_train_scaled,
X_cv_scaled,
y_cv,
cv['adj_close_mean'],
cv['adj_close_std'],
seed=model_seed,
n_estimators=n_estimators_opt,
max_depth=max_depth_opt,
learning_rate=learning_rate_opt,
min_child_weight=min_child_weight_opt,
subsample=param,
colsample_bytree=colsample_bytree,
colsample_bylevel=colsample_bylevel,
gamma=param2)
# Collect results
error_rate[param_label].append(param)
error_rate[param2_label].append(param2)
error_rate['rmse'].append(rmse)
error_rate['mape_pct'].append(mape)
error_rate = pd.DataFrame(error_rate)
toc = time.time()
print("Minutes taken = " + str((toc-tic)/60.0))
error_rate
# +
# Plot performance versus params
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
temp = error_rate[error_rate[param2_label]==param2_list[0]]
ax = temp.plot(x=param_label, y='rmse', style='bs-', grid=True)
legend_list = [param2_label + '_' + str(param2_list[0])]
color_list = ['r', 'g', 'k', 'y', 'm', 'c', '0.75']
for i in range(1,len(param2_list)):
temp = error_rate[error_rate[param2_label]==param2_list[i]]
ax = temp.plot(x=param_label, y='rmse', color=color_list[i%len(color_list)], marker='s', grid=True, ax=ax)
legend_list.append(param2_label + '_' + str(param2_list[i]))
ax.set_xlabel(param_label)
ax.set_ylabel("RMSE")
matplotlib.rcParams.update({'font.size': 14})
plt.legend(legend_list, loc='center left', bbox_to_anchor=(1.0, 0.5)) # positions legend outside figure
# -
# Get optimum value for param and param2
temp = error_rate[error_rate['rmse'] == error_rate['rmse'].min()]
subsample_opt = temp['subsample'].values[0]
gamma_opt = temp['gamma'].values[0]
print("min RMSE = %0.3f" % error_rate['rmse'].min())
print("optimum params = ")
subsample_opt, gamma_opt
# Get optimum value for param and param2, using MAPE
# We will use RMSE to decide the final optimum params to use
temp = error_rate[error_rate['mape_pct'] == error_rate['mape_pct'].min()]
print("min MAPE = %0.3f%%" % error_rate['mape_pct'].min())
print("optimum params = ")
temp['subsample'].values[0], temp['gamma'].values[0]
# +
param_label = 'colsample_bytree'
param_list = [0.5, 0.6, 0.7, 0.8, 0.9, 1]
param2_label = 'colsample_bylevel'
param2_list = [0.5, 0.6, 0.7, 0.8, 0.9, 1]
error_rate = {param_label: [] , param2_label: [], 'rmse': [], 'mape_pct': []}
tic = time.time()
for param in tqdm_notebook(param_list):
# print("param = " + str(param))
for param2 in param2_list:
# Train, predict and eval model
rmse, mape, _ = train_pred_eval_model(X_train_scaled,
y_train_scaled,
X_cv_scaled,
y_cv,
cv['adj_close_mean'],
cv['adj_close_std'],
seed=model_seed,
n_estimators=n_estimators_opt,
max_depth=max_depth_opt,
learning_rate=learning_rate_opt,
min_child_weight=min_child_weight_opt,
subsample=subsample_opt,
colsample_bytree=param,
colsample_bylevel=param2,
gamma=gamma_opt)
# Collect results
error_rate[param_label].append(param)
error_rate[param2_label].append(param2)
error_rate['rmse'].append(rmse)
error_rate['mape_pct'].append(mape)
error_rate = pd.DataFrame(error_rate)
toc = time.time()
print("Minutes taken = " + str((toc-tic)/60.0))
error_rate
# +
# Plot performance versus params
rcParams['figure.figsize'] = 10, 8 # width 10, height 8
temp = error_rate[error_rate[param2_label]==param2_list[0]]
ax = temp.plot(x=param_label, y='rmse', style='bs-', grid=True)
legend_list = [param2_label + '_' + str(param2_list[0])]
color_list = ['r', 'g', 'k', 'y', 'm', 'c', '0.75']
for i in range(1,len(param2_list)):
temp = error_rate[error_rate[param2_label]==param2_list[i]]
ax = temp.plot(x=param_label, y='rmse', color=color_list[i%len(color_list)], marker='s', grid=True, ax=ax)
legend_list.append(param2_label + '_' + str(param2_list[i]))
ax.set_xlabel(param_label)
ax.set_ylabel("RMSE")
matplotlib.rcParams.update({'font.size': 14})
plt.legend(legend_list, loc='center left', bbox_to_anchor=(1.0, 0.5)) # positions legend outside figure
# -
# Get optimum value for param and param2
temp = error_rate[error_rate['rmse'] == error_rate['rmse'].min()]
colsample_bytree_opt = temp['colsample_bytree'].values[0]
colsample_bylevel_opt = temp['colsample_bylevel'].values[0]
print("min RMSE = %0.3f" % error_rate['rmse'].min())
print("optimum params = ")
colsample_bytree_opt, colsample_bylevel_opt
# Get optimum value for param and param2, using MAPE
# We will use RMSE to decide the final optimum params to use
temp = error_rate[error_rate['mape_pct'] == error_rate['mape_pct'].min()]
print("min MAPE = %0.3f%%" % error_rate['mape_pct'].min())
print("optimum params = ")
temp['colsample_bytree'].values[0], temp['colsample_bylevel'].values[0]
d = {'param': ['n_estimators', 'max_depth', 'learning_rate', 'min_child_weight', 'subsample', 'colsample_bytree', 'colsample_bylevel', 'gamma', 'rmse', 'mape_pct'],
'original': [n_estimators, max_depth, learning_rate, min_child_weight, subsample, colsample_bytree, colsample_bylevel, gamma, rmse_bef_tuning, mape_bef_tuning],
'after_tuning': [n_estimators_opt, max_depth_opt, learning_rate_opt, min_child_weight_opt, subsample_opt, colsample_bytree_opt, colsample_bylevel_opt, gamma_opt, error_rate['rmse'].min(), error_rate['mape_pct'].min()]}
tuned_params = pd.DataFrame(d)
tuned_params = tuned_params.round(3)
tuned_params
# +
rmse, mape, est = train_pred_eval_model(X_train_cv_scaled,
y_train_cv_scaled,
X_sample_scaled,
y_sample,
test['adj_close_mean'],
test['adj_close_std'],
seed=model_seed,
n_estimators=n_estimators_opt,
max_depth=max_depth_opt,
learning_rate=learning_rate_opt,
min_child_weight=min_child_weight_opt,
subsample=subsample_opt,
colsample_bytree=colsample_bytree_opt,
colsample_bylevel=colsample_bylevel_opt,
gamma=gamma_opt)
# Calculate RMSE
print("RMSE on test set = %0.3f" % rmse)
# Calculate MAPE
print("MAPE on test set = %0.3f%%" % mape)
# +
# Plot adjusted close over time
rcParams['figure.figsize'] = 16, 8 # width 10, height 8
est_df = pd.DataFrame({'est': est,
'y_sample': y_sample,
'date': test['date']})
ax = train.plot(x='date', y='adj_close', style='b-', grid=True)
ax = cv.plot(x='date', y='adj_close', style='y-', grid=True, ax=ax)
ax = test.plot(x='date', y='adj_close', style='g-', grid=True, ax=ax)
ax = est_df.plot(x='date', y='est', style='r-', grid=True, ax=ax)
ax.legend(['train', 'dev', 'test', 'predictions'])
ax.set_xlabel("date")
ax.set_ylabel("EUR")
# -
|
ts_xgboost_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/atlas-github/20190731StarMediaGroup/blob/master/1_Introduction_to_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ZIrGYKSFitp2" colab_type="code" colab={}
print("Hello World")
# + [markdown] id="j5I2EPH4DPEC" colab_type="text"
# # Python Crash Course
#
# Please note, this is not meant to be a comprehensive overview of Python or programming in general. There are recommended sources at the bottom part of this notebook to help you learn more about Python programming.
#
# This notebook will just go through the basic topics in order:
#
# * Data types
# * Numbers
# * Strings
# * Printing
# * Lists
# * Dictionaries
# * Booleans
# * Tuples
# * Sets
# * Comparison Operators
# * if,elif, else Statements
# * for Loops
# * while Loops
# * range()
# + [markdown] id="PNM2SzBfDIwg" colab_type="text"
# # Why Python?
#
# 1. It’s easy to learn
# 2. Now the language of choice for 8 of 10 top US computer science
# programs (Philip Guo, CACM)
# 3. Full featured
# 4. Not just a statistics language, but has full capabilities for data
# acquisition, cleaning, databases, high performance computing, and more
# 5. Strong Data Science Libraries
# 6. The SciPy Ecosystem
#
#
# It's important to note that Python isn't the only way that we could teach data science. We can teach data science in R, in Excel, in any number of software packages. That's because Python is a tool that data scientists use, but the concepts are more important for you to understand.
# + [markdown] id="TP8v7hWHHeMm" colab_type="text"
# ## Data types
#
# ### Numbers
#
# Python can also handle numbers, and basic arithmetic using the following functions.
#
# | Symbol | Task Performed |
# |----|---|
# | + | Addition |
# | - | Subtraction |
# | / | division |
# | % | modulo |
# | * | multiplication |
# | // | floor division |
# | ** | to the power of |
# + id="gz3xuk3vG4I-" colab_type="code" colab={}
1 + 1
# + id="5zbDUjTaHfui" colab_type="code" colab={}
1 * 3
# + id="O2CFuwTfQ3Vk" colab_type="code" colab={}
1 / 2
# + id="yCdWdJ-KQ83R" colab_type="code" colab={}
2 ** 4
# + id="F4w9u3_6Q94W" colab_type="code" colab={}
4 % 2
# + id="3aEo2Kf0Q_L1" colab_type="code" colab={}
5 % 2
# + id="JRmi63QORAb9" colab_type="code" colab={}
(2 + 3) * (5 + 5)
# + [markdown] id="g336z7_fRe2V" colab_type="text"
# ### Variable Assignment
# + id="sdm3h1gjRccN" colab_type="code" colab={}
# Cannot start with number or special characters
name_of_var = 2
# + id="qr8xXW_LRjYn" colab_type="code" colab={}
x = 2
y = 3
# + id="IZsNqSl3Rk7O" colab_type="code" colab={}
x*y
# + id="ZBlV8QIQRpS-" colab_type="code" colab={}
z = x + y
# + id="e5ynGpxCRqYU" colab_type="code" colab={}
z
# + [markdown] id="NpuPnCzAR24W" colab_type="text"
# ### Strings
#
# A string is a set of characters. Strings are used to store data that is _text_.
#
# In Python, Strings are abbreviated "str" and there are many built-in features that allow you to manipulate those strings.
#
# Strings can be enclosed by either double or single quotes, although single quotes are more common.
# + id="uP6-dR-GRzQe" colab_type="code" colab={}
a = "Najib"
# + id="gIOVuFTtR3as" colab_type="code" colab={}
a
# + id="8BntkB-IR8HS" colab_type="code" colab={}
'single quotes'
# + id="2j02mR2oR9hI" colab_type="code" colab={}
"double quotes"
# + [markdown] id="xy21e_5-SBgn" colab_type="text"
# Question: Why would you want to use double quotes on occasion?
# + id="Dlt0ey2PR-nF" colab_type="code" colab={}
" wrap lot's of other quotes"
# + [markdown] id="fByzxk2ISGNv" colab_type="text"
# #### String Operations
#
# s + t -- the concatenation of s and t
#
# n * s -- equivalent to adding s to itself n times
# + id="Lc8WEWYYSCC6" colab_type="code" colab={}
b + " Fries"
# + id="Qku8TwxUSIjU" colab_type="code" colab={}
b = 'French' # Concatenation works quite intuitively
print((b + ' Fries'))
# + id="XkaM-sO1SMdv" colab_type="code" colab={}
c = "yay! "
print(c * 5)
# + [markdown] id="aZiTscm0STVl" colab_type="text"
# #### Slices of String Indices
#
# Characters in a string can be accessed using the standard [ ] syntax, and like Java and C++, Python uses zero-based indexing. This concept will extend much further than strings.
#
# This is easiest to think about if you think of each letter as being in its own box.
#
# s ='hello'
#
# s[1:4] is 'ell' -- chars starting at index 1 and extending up to but not including index 4
#
# s[1:] is 'ello' -- omitting either index defaults to the start or end of the string
#
# s[:] is 'Hello' -- omitting both always gives us a copy of the whole thing (this is the pythonic way to copy a sequence like a string or list)
#
# s[1:100] is 'ello' -- an index that is too big is truncated down to the string length
#
# s[-1] is 'o' -- last char (1st from the end)
#
# s[-4] is 'e' -- 4th from the end
#
# s[:-3] is 'He' -- going up to but not including the last 3 chars.
#
# s[-3:] is 'llo' -- starting with the 3rd char from the end and extending to the end of the string.
#
# h e l l o <br>
# 0 1 2 3 4<br>
# -5 -4 -3 -2 -1<br>
#
#
# s[i] -- ith item of s, origin 0
#
# s[i:j] -- slice of s from i to j
#
# s[i:j:k] -- slice of s from i to j with step k
# + id="sgU685bySOmt" colab_type="code" colab={}
s='hello'
# + id="4s3Z73E4SWMg" colab_type="code" colab={}
s[0:3]
# + id="cEl4V-SuSXMZ" colab_type="code" colab={}
s[1:]
# + [markdown] id="-F4GAb3WSb-9" colab_type="text"
# #### Common built- in methods
#
# min(s) -- smallest item of s
#
# max(s) -- largest item of s
#
# s.count(x) -- total number of occurrences of x in s
#
# s.lower(), s.upper() -- returns the lowercase or uppercase version of the string
#
# s.strip() -- returns a string with whitespace removed from the start and end
#
# s.isalpha()/s.isdigit()/s.isspace()... -- tests if all the string chars are in the various character classes
#
# s.startswith('other'), s.endswith('other') -- tests if the string starts or ends with the given other string
#
# s.replace('old', 'new') -- returns a string where all occurrences of 'old' have been replaced by 'new'
#
# s.split('delim') -- returns a list of substrings separated by the given delimiter. The delimiter is not a regular expression, it's just text. 'aaa,bbb,ccc'.split(',') -> ['aaa', 'bbb', 'ccc']. As a convenient special case s.split() (with no arguments) splits on all whitespace chars.
#
# s.join(list) -- opposite of split(), joins the elements in the given list together using the string as the delimiter. e.g. '---'.join(['aaa', 'bbb', 'ccc']) -> aaa---bbb---ccc
# + id="0qUeOr4cSYdI" colab_type="code" colab={}
st = 'Hello my name is Wyhow'
# + id="nJjvE8RMSeu_" colab_type="code" colab={}
st.upper()
# + id="rzZcEnx-SiGg" colab_type="code" colab={}
st.lower()
# + id="q7m01SqgSjRT" colab_type="code" colab={}
st.split()
# + id="4rreZbU1SnSi" colab_type="code" colab={}
tweets = 'This is a tweets @jackma'
# + id="IiSoCr6ZSoVt" colab_type="code" colab={}
tweets.split('@')[1]
# + [markdown] id="a2qTuNCvSvuM" colab_type="text"
# ### Printing
# + id="1QEew78KSpX_" colab_type="code" colab={}
x = 'hello'
# + id="7aeoFH9LS3vm" colab_type="code" colab={}
x
# + id="qI2pIwFBS5H4" colab_type="code" colab={}
print(x)
# + [markdown] id="OdrBTRC4Teyg" colab_type="text"
# ### Format
#
# The % operator takes a printf-type format string on the left (%d int, %s string, %f/%g floating point), and the matching values in a tuple on the right (a tuple is made of values separated by commas, typically grouped inside parenthesis)
# + id="BhUveOxsS6Kh" colab_type="code" colab={}
num = 'twelve'
name = 'Wyhow'
# + id="zTbFggARTm0b" colab_type="code" colab={}
print('My number is: %d, and my name is: %s' %(num,name))
# + id="BVbYATcjTtlf" colab_type="code" colab={}
print('My number is: {one}, and my name is: {two}'.format(one=num,two=name))
# + id="LPllLk4TTviH" colab_type="code" colab={}
print('My number is: {}, and my name is: {}'.format(num,name))
# + [markdown] id="msR65PPqT47R" colab_type="text"
# ### Lists
#
# Like a string, a **list** is a sequence of values. In a string, the values are characters; in a list, they can be any type. The values in list are called **elements** or sometimes **items**.
#
# List elements are written within square brackets [ ]. Square brackets [ ] access data, with the first element at index 0. Many of the operations defined above for strings also apply to lists. So it can be convenient to think of a string simply as a list of characters.
# + id="485GXvb0TyA_" colab_type="code" colab={}
[1,2,3]
# + id="n-STbUCwT7hd" colab_type="code" colab={}
a = [1, 2,3]
# + id="CJnpPbChT81X" colab_type="code" colab={}
a
# + id="Irgce7RbT9po" colab_type="code" colab={}
['hi',1,[1,2]]
# + id="wVjQ0_1UT_A1" colab_type="code" colab={}
my_list = ['a','b','c']
# + id="Div_zPyHUAYx" colab_type="code" colab={}
my_list.append('d')
# + id="u6HpX3dRUBfa" colab_type="code" colab={}
my_list
# + id="L8PaEma-UCeu" colab_type="code" colab={}
my_list[0]
# + id="I4vsd8DjUDil" colab_type="code" colab={}
my_list[1]
# + id="0XXdZbWJUEWd" colab_type="code" colab={}
my_list[1:]
# + id="WNPp38-FUE6H" colab_type="code" colab={}
my_list[:1]
# + id="AjjuA0zAUFiw" colab_type="code" colab={}
my_list[0] = 'NEW'
# + id="DPZaULVfUKUO" colab_type="code" colab={}
my_list
# + id="GyWIrySkULdW" colab_type="code" colab={}
nest = [1,2,3,[4,5,['target']]]
# + id="US7U_h9mUM37" colab_type="code" colab={}
nest[3]
# + id="eTye9p41UN61" colab_type="code" colab={}
nest[3][2]
# + id="x-Iah_88UPYK" colab_type="code" colab={}
nest[3][2][0]
# + [markdown] id="SX9lwonyURS6" colab_type="text"
# ### List Methods
#
# - list.append(elem) -- adds a single element to the end of the list. Common error: does not return the new list, just modifies the original.
# - list.insert(index, elem) -- inserts the element at the given index, shifting elements to the right.
# - list.extend(list2) adds the elements in list2 to the end of the list. Using + or += on a list is similar to using extend().
# - list.index(elem) -- searches for the given element from the start of the list and returns its index. Throws a ValueError if the element does not appear (use "in" to check without a ValueError).
# - list.remove(elem) -- searches for the first instance of the given element and removes it (throws ValueError if not present)
# - list.sort() -- sorts the list in place (does not return it).
# - sorted(list) -- return sorted list but keeps the original order of the list
# - list.reverse() -- reverses the list in place (does not return it)
# - list.pop(index) -- removes and returns the element at the given index. Returns the rightmost element if index is omitted (roughly the opposite of append()).
# + id="wn7oWwM0UQaX" colab_type="code" colab={}
my_new_list = [5,10,15]
# + id="Mazd9ySjUZ9L" colab_type="code" colab={}
my_new_list.append(20)
# + id="NB7VJCu-Ua9F" colab_type="code" colab={}
my_new_list
# + id="mzGggyRaUcgz" colab_type="code" colab={}
sorted(my_new_list)
# + [markdown] id="rVoQqBtRUipK" colab_type="text"
# ### Dictionaries
#
# Python's efficient key/value hash table structure is called a "dict".
#
# A dictionary is similar to a list, but you access values by looking up a key instead of an index. A key can be any string or number.
#
# The contents of a dict can be written as a series of key:value pairs within braces { }, e.g. dict = {key1:value1, key2:value2, ... }.
#
# The "empty dict" is just an empty pair of curly braces {}.
#
# Looking up or setting a value in a dict uses square brackets, e.g. dict['foo'] looks up the value under the key 'foo'. Strings, numbers, and tuples work as keys, and any type can be a value. Other types may or may not work correctly as keys.
#
# Dictionaries are mutable!
# + id="8wWZE0nHUd2u" colab_type="code" colab={}
d = {'key1':'item1','key2':'item2'}
# + id="6LXL0JJFUg22" colab_type="code" colab={}
d
# + id="dvlu6chTUl2U" colab_type="code" colab={}
d['key1']
# + [markdown] id="zM3l9_mFUo0o" colab_type="text"
# ### Dictionary Methods
#
# - len(dict) -- Gives the total length of the dictionary. This would be equal to the number of items in the dictionary.
# - str(dict) -- Produces a printable string representation of a dictionary
# - type(variable) -- Returns the type of the passed variable. If passed variable is dictionary, then it would return a dictionary type.
# - dict.clear() -- Removes all elements of dictionary dict
# - dict.copy() -- Returns a shallow copy of dictionary dict
# - dict.fromkeys() -- Create a new dictionary with keys from seq and values set to value.
# - dict.get(key, default=None) -- For key key, returns value or default if key not in dictionary
# - dict.items() -- Returns a list of dict's (key, value) tuple pairs
# - dict.keys() -- Returns list of dictionary dict's keys
# - dict.setdefault(key, default=None) -- Similar to get(), but will set dict[key]=default if key is not already in dict
# - dict.update(dict2) -- Adds dictionary dict2's key-values pairs to dict
# - dict.values() -- Returns list of dictionary dict's values
# + id="H34i-ovDUnCc" colab_type="code" colab={}
d.keys()
# + id="qlMUc3LhUrhu" colab_type="code" colab={}
d.items()
# + [markdown] id="mmFfKjugU2rt" colab_type="text"
# ### Booleans
# + id="cjiY6BEhUskS" colab_type="code" colab={}
True & False
# + id="PJANxFSgU1Tg" colab_type="code" colab={}
False
# + [markdown] id="Bji9gzJ4U9uE" colab_type="text"
# ### Tuples
#
# A tuple is a fixed size grouping of elements, such as an (x, y) co-ordinate. Tuples are like lists, except they are **immutable** and do not change size (tuples are not strictly immutable since one of the contained elements could be mutable). Tuples are a convenient way to pass around a little logical, fixed size bundle of values. A function that needs to return multiple values can just return a tuple of the values.
# + id="4VY1uh1-U7y3" colab_type="code" colab={}
t = (1,2,3)
# + id="zp7_OCyPVBOW" colab_type="code" colab={}
t[0]
# + id="erqNfb3qVDor" colab_type="code" colab={}
t[0] = 'NEW'
# + [markdown] id="2r-ilJzEVKMI" colab_type="text"
# ## Comparison Operators
# + id="vQxxHYcmVEev" colab_type="code" colab={}
1 > 2
# + id="3tYQ6Q6TVFpC" colab_type="code" colab={}
1 < 2
# + id="G85bWxreVOip" colab_type="code" colab={}
1 >= 1
# + id="qzV2hdT3VP-a" colab_type="code" colab={}
1 <= 4
# + id="NI6bYIwnVQzY" colab_type="code" colab={}
1 == 1
# + id="PveIqpFZVRy7" colab_type="code" colab={}
'hi' == 'hi'
# + [markdown] id="LTn1TWyOVYDf" colab_type="text"
# ## Logic Operators
# + id="VF1tE17XVStp" colab_type="code" colab={}
(1 > 2) and (2 < 3)
# + id="8zjj052pVble" colab_type="code" colab={}
(1 > 2) or (2 < 3)
# + id="891HyjTYVcrk" colab_type="code" colab={}
(1 == 2) or (2 == 3) or (4 == 4)
# + [markdown] id="v7F_ow3XVfSV" colab_type="text"
# ## if, elif, else Statements
# + id="FRZYbhLKVdtb" colab_type="code" colab={}
if 1 > 2:
print('Yep!')
# + id="vMtJs-u0VjRh" colab_type="code" colab={}
if 1 < 2:
print('yep!')
# + id="MVlcw0bRVkvS" colab_type="code" colab={}
if 1 < 2:
print('first')
else:
print('last')
# + id="B410JJRcVl09" colab_type="code" colab={}
if 1 > 2:
print('first')
else:
print('last')
# + id="ODA7QqH4Vm-y" colab_type="code" colab={}
if 1 == 2:
print('first')
elif 3 == 3:
print('middle')
else:
print('Last')
# + [markdown] id="dUaG8fSSVrux" colab_type="text"
# ## for Loops
#
# Python's *for* and *in* constructs are extremely useful, and the first use of them we'll see is with lists. The *for* construct -- for var in list -- is an easy way to look at each element in a list (or other collection).
# + id="xf867aH7Vphz" colab_type="code" colab={}
seq = ['hi', 'why', 'hello']
# + id="8q0GSe7XV5xj" colab_type="code" colab={}
for item in seq:
print(seq)
# + id="6lZPjYADV9Du" colab_type="code" colab={}
for item in seq:
print('Yep')
# + id="Y3njaFOZV-HT" colab_type="code" colab={}
for spongebob in seq:
print(spongebob+spongebob)
# + [markdown] id="gUEd-XOIWCWb" colab_type="text"
# ## while Loops
# + id="VSyydZ9mV_Sq" colab_type="code" colab={}
i = 2
while i <= 500:
print('i is: {}'.format(i))
i = i**3
# + [markdown] id="reFEPbVcWG-a" colab_type="text"
# ## range()
# + id="RXY--RiyWEzp" colab_type="code" colab={}
range(5)
# + id="ZNl8jABDWIx_" colab_type="code" colab={}
for i in range(5):
print(i)
# + id="LN6sklyJWJ7w" colab_type="code" colab={}
list(range(5))
# + [markdown] id="sVSmBVWKLei6" colab_type="text"
# # Exercises
# + [markdown] id="TOzII68CLl7_" colab_type="text"
# Ask the user for a number. Depending on whether the number is even or odd, print out an appropriate message to the user. Hint: how does an even / odd number react differently when divided by 2?
# + id="CIDCHDPgLuCU" colab_type="code" colab={}
#num = int(input("Enter a number: "))
###INSERT CODE HERE
###Share your code on https://codeshare.io/axEYAR
# + [markdown] id="HHTu1DFRLuek" colab_type="text"
# Extras:
#
# 1. If the number is a multiple of 4, print out a different message.
# 2. Ask the user for two numbers: one number to check (call it num) and one number to divide by (check). If check divides evenly into num, tell that to the user. If not, print a different appropriate message.
# 3. More problems [here](https://www.practicepython.org/).
# + id="JOkQNiyLNNMI" colab_type="code" colab={}
#num = int(input("give me a number to check: "))
###INSERT CODE HERE
###Share your code on https://codeshare.io/axEYAR
# + id="VeXYmf1IPv1e" colab_type="code" colab={}
#num = int(input("give me a number to check: "))
#check = int(input("give me a number to divide by: "))
###INSERT CODE HERE
###Share your code on https://codeshare.io/axEYAR
# + [markdown] id="LffY6KgMWh1d" colab_type="text"
# # Where to go from here
# + [markdown] id="3Qbc0ImtXjZ3" colab_type="text"
# | Link | Image |
# |----|---|
# | [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) | !<img src=https://covers.oreillystatic.com/images/0636920034919/lrg.jpg width="300"> |
# | [Python Crash Course](https://nostarch.com/pythoncrashcourse) | !<img src=https://nostarch.com/images/PythonCrashCourse2e_product.jpg width="300">
|
1_Introduction_to_Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from tqdm import tqdm
from p_tqdm import p_map
import os
import shutil
import networkx as nx
from rdkit import Chem
from rdkit.Chem import AllChem
from rdkit.Chem import rdMolDescriptors
from rdkit.Chem import PandasTools
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem import Draw
from rdkit.Chem import rdDepictor
from rdkit import DataStructs
rdDepictor.SetPreferCoordGen(True)
#from rdkit import RDLogger
#RDLogger.DisableLog('rdApp.warning')
from molvs import standardize_smiles
import pubchempy
# -
# # Repurposed small-molecule drugs for COVID
#
# The following list is from the recent article by Liu et al [1].
#
# 
#
# [1. <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, Research and Development on Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases. ACS Cent. Sci. (2020)](https://doi.org/10.1021/acscentsci.0c00272)
df = pd.read_csv("covid_drugname_cas.csv", delimiter="\t")
df
def name2smiles(name):
"""
Uses pubchempy to fetch the smiles of a molecule, given its name.
"""
pubchem_compounds = pubchempy.get_compounds(name, "name")
try:
return pubchem_compounds[0].isomeric_smiles
except:
return None
df["smiles"] = df["name"].map(name2smiles)
#df.to_csv("mols_w_smiles.csv")
df = pd.read_csv("mols_w_smiles.csv", index_col=0)
PandasTools.AddMoleculeColumnToFrame(df, smilesCol="smiles", molCol="rdmol")
df.rdmol.map(rdDepictor.Compute2DCoords)
img = Draw.MolsToGridImage(df.rdmol,molsPerRow=3,subImgSize=(400,400),legends=list(df.name))
img
img.save("images/liu_article_antivirals.png")
for i,r in df.iterrows():
molname=r["name"]
im = Draw.MolsToGridImage([r.rdmol])
im.save(f"images/{molname}.png")
# # 3D structure generation
#
# - Some of these compounds have quite many rotatable bonds..
# - [Here](https://pubs.acs.org/doi/10.1021/acs.jpcb.5b12272) they studied the conformational energy landscape of ritonavir, the one with most rotbonds
# - Structural to stabilize the bioactive conf? Like in [here](http://dx.doi.org/10.1021/acs.molpharmaceut.9b00437). Probably has been studied, no idea
#
df["nrotbonds"] = df.rdmol.map(rdMolDescriptors.CalcNumRotatableBonds)
df[["name", "nrotbonds"]]
# # Similarity searches
#
# - Downloaded the Pubchem database (107 million molecules)
# - ```wget ftp://ftp.ncbi.nlm.nih.gov/pubchem/Compound/Extras/CID-SMILES.gz```
# - 6GB when uncompressed, can't read it to my memory directly. Split it to 1e5 row chuncks
# - ```split -l 100000 -d CID-SMILES```
#
# - The following (trivially parallelizable) problem would have taken 14 hours with my crappy laptop, sent it to our remote cluster
#
# +
# pubchem_smiles = [f for f in os.listdir() if f.startswith("x")]
# with open("already_looked") as f: already_looked = [i.strip() for i in f.readlines()]
# not_looked = list(set(pubchem_smiles).difference(set(already_looked)))
# fps_antivirals = [Chem.RDKFingerprint(m) for m in df.rdmol]
# def compute_similarities_for_chunk(path_to_cid_smiles):
# def smiles_to_simvec(smiles):
# rdmol=Chem.MolFromSmiles(smiles)
# fp2compare = Chem.RDKFingerprint(rdmol)
# sims = []
# for fp_i in fps_antivirals:
# try:
# sim = DataStructs.FingerprintSimilarity(fp2compare,fp_i)
# except:
# sim = None
# sims.append(sim)
# return sims
# res = []
# with open(path_to_cid_smiles) as f:
# cid_smiles = f.readlines()
# for r in cid_smiles:
# try:
# sims = smiles_to_simvec(r.split()[1])
# res.append(",".join(r.split()) + "," + ",".join([str(i) for i in sims]))
# except:
# continue
# with open(f"sims_{path_to_cid_smiles}", "w") as f:
# f.write("\n".join(res))
# with open(f"already_looked", "a") as f:
# f.write(f"{path_to_cid_smiles}\n")
# p_map(compute_similarities_for_chunk, not_looked, num_cpus=4)
#
# cols = ["cid", "smiles"] + list(df.name)
# dfz = {i : [] for i in df.name}
# with open("already_looked") as f:
# already_looked = f.readlines()
# for i in already_looked:
# df_chunk = pd.read_csv(f"sims_{i}".strip(), names=cols)
# for drugname in df.name:
# sims = df_chunk[df_chunk[drugname] >= 0.7][["cid", "smiles", drugname]].to_numpy().tolist()
# if len(sims)>0:
# dfz[drugname]+=sims
# drugsim_dfs = {k : pd.DataFrame(v, columns="cid smiles similarity".split()).set_index("cid").sort_values("similarity", ascending=False) for k,v in dfz.items()}
# for k,v in drugsim_dfs.items():
# v.to_csv(f"{k}_sims.csv")
# -
# - The molecules that have a Tanimoto similarity > 0.7 (RDKit topological fingerprints) are in ./similar_mols/<molecule>_sims.csv
# - cid, smiles, similarity coefficient
mol_sims_dict = {molname : pd.read_csv(f"similar_mols/{molname}_sims.csv") for molname in df.name}
im = Draw.MolToImage(df[df.name == k].rdmol.values[0])
im
for k, v in mol_sims_dict.items():
plt.figure()
sns.distplot(v["similarity"], bins=20, kde=False)
plt.xlim((0.7,1))
plt.title(f"{k:15} : {v.shape[0]:10} similar compounds")
plt.xlabel("Similarity")
plt.savefig(f"images/{k}_sim.png")
plt.show()
for i in df.name:
url="https://github.com/mrauha/covid19_drugs/blob/master/images/"
string = f' '
print(string)
string
|
covid_drugs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Quick Fix Example 1
#
# Noticed that FNDM1 and FNDM2 were switched session labels. FNDM1 should have ITC tasks, while FNDM2 should have Card/Face/Perfusion. Fix is below
# modules
import flywheel
import pandas as pd
# add the script to the path
import sys
import os
sys.path.append(os.path.abspath("/home/ttapera/bids-on-flywheel/flywheel_bids_tools"))
import query_bids
import upload_bids
from tqdm import tqdm
import math
# query
fw = flywheel.Client()
result = query_bids.query_fw("Reward2018", fw)
# +
# clean query to get a table
view = fw.View(columns='subject')
subject_df = fw.read_view_dataframe(view, result.id)
sessions = []
view = fw.View(columns='acquisition')
pbar = tqdm(total=100)
for ind, row in subject_df.iterrows():
session = fw.read_view_dataframe(view, row["subject.id"])
if(session.shape[0] > 0):
sessions.append(session)
pbar.update(10)
pbar.close()
# -
acquisitions = pd.concat(sessions,ignore_index=True)
acquisitions.head()
# Here are the fndm2 acquisitions; you can see the acquisition labels are incorrect
acquisitions.loc[acquisitions['session.label'] == "fndm2",]
# We assign indeces and swap the session labels for them:
ind2 = acquisitions.loc[acquisitions['session.label'] == "fndm2",].index
ind1 = acquisitions.loc[acquisitions['session.label'] == "fndm1",].index
acquisitions.loc[ind2, 'session.label'] = "fndm1"
acquisitions.loc[ind1, 'session.label'] = "fndm2"
acquisitions.loc[acquisitions['session.label'] == "fndm2",]
# Now that the table is corrected, we upload the changes:
fndm2 = acquisitions.loc[acquisitions['session.label'] == "fndm2",].reset_index()
# get the session object
ses = fw.get(fndm2.loc[0,'session.id'])
# update it's label
ses.update({"label": fndm2.loc[0,'session.label']})
#reload to confirm
ses = ses.reload()
ses.label
# Here's a loop to automate the process:
# +
fndm2 = acquisitions.loc[acquisitions['session.label'] == "fndm2",].reset_index()
fndm2 = fndm2.drop_duplicates('session.id').reset_index()
fndm1 = acquisitions.loc[acquisitions['session.label'] == "fndm1",].reset_index()
fndm1 = fndm1.drop_duplicates('session.id').reset_index()
for ind, row in fndm2.iterrows():
ses = fw.get(row['session.id'])
ses.update({"label": row['session.label']})
for ind, row in fndm1.iterrows():
ses = fw.get(row['session.id'])
ses.update({"label": row['session.label']})
# -
# We re-query flywheel for an updated view:
# +
result = query_bids.query_fw("Reward2018", fw)
view = fw.View(columns='subject')
subject_df = fw.read_view_dataframe(view, result.id)
sessions = []
view = fw.View(columns='acquisition')
pbar = tqdm(total=100)
for ind, row in subject_df.iterrows():
session = fw.read_view_dataframe(view, row["subject.id"])
if(session.shape[0] > 0):
sessions.append(session)
pbar.update(10)
pbar.close()
acquisitions = pd.concat(sessions,ignore_index=True)
# -
# Finally, we see that FNDM1 has card and face tasks...
acquisitions.loc[acquisitions['session.label'] == 'fndm1',].head()
# And FNDM2 has ITC tasks...
acquisitions.loc[acquisitions['session.label'] == 'fndm2',].head()
# Done!
|
notebooks/Fixing Flywheel BIDS Data Using a Table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
# +
import pandas as pd
import pyspark.sql.functions as F
from datetime import datetime
from pyspark.sql.types import *
from pyspark import StorageLevel
import numpy as np
pd.set_option("display.max_rows", 1000)
pd.set_option("display.max_columns", 1000)
pd.set_option("mode.chained_assignment", None)
# +
from pyspark.ml import Pipeline
from pyspark.ml.classification import GBTClassifier
from pyspark.ml.feature import IndexToString, StringIndexer, VectorIndexer
# from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml.feature import OneHotEncoderEstimator, StringIndexer, VectorAssembler
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from pyspark.ml.tuning import CrossValidator, ParamGridBuilder
from pyspark.sql import Row
from pyspark.ml.linalg import Vectors
# +
# # !pip install scikit-plot
# -
import sklearn
import scikitplot as skplt
from sklearn.metrics import classification_report, confusion_matrix, precision_score
# <hr />
# <hr />
# <hr />
result_schema = StructType([
StructField('experiment_filter', StringType(), True),
StructField('undersampling_method', StringType(), True),
StructField('undersampling_column', StringType(), True),
StructField('filename', StringType(), True),
StructField('experiment_id', StringType(), True),
StructField('n_covid', IntegerType(), True),
StructField('n_not_covid', IntegerType(), True),
StructField('model_name', StringType(), True),
StructField('model_seed', StringType(), True),
StructField('model_maxIter', IntegerType(), True),
StructField('model_maxDepth', IntegerType(), True),
StructField('model_maxBins', IntegerType(), True),
StructField('model_minInstancesPerNode', IntegerType(), True),
StructField('model_minInfoGain', FloatType(), True),
StructField('model_featureSubsetStrategy', StringType(), True),
StructField('model_n_estimators', IntegerType(), True),
StructField('model_learning_rate', FloatType(), True),
StructField('model_impurity', StringType(), True),
StructField('model_AUC_ROC', StringType(), True),
StructField('model_AUC_PR', StringType(), True),
StructField('model_covid_precision', StringType(), True),
StructField('model_covid_recall', StringType(), True),
StructField('model_covid_f1', StringType(), True),
StructField('model_not_covid_precision', StringType(), True),
StructField('model_not_covid_recall', StringType(), True),
StructField('model_not_covid_f1', StringType(), True),
StructField('model_avg_precision', StringType(), True),
StructField('model_avg_recall', StringType(), True),
StructField('model_avg_f1', StringType(), True),
StructField('model_avg_acc', StringType(), True),
StructField('model_TP', StringType(), True),
StructField('model_TN', StringType(), True),
StructField('model_FN', StringType(), True),
StructField('model_FP', StringType(), True),
StructField('model_time_exec', StringType(), True),
StructField('model_col_set', StringType(), True)
])
# <hr />
# <hr />
# <hr />
# +
# undersamp_col = ['03-STRSAMP-AG', '04-STRSAMP-EW']
# dfs = ['ds-1', 'ds-2', 'ds-3']
# cols_sets = ['cols_set_1', 'cols_set_2', 'cols_set_3']
undersamp_col = ['02-KMODES']
dfs = ['ds-1']
cols_sets = ['cols_set_3']
# +
# lists of params
model_maxIter = [20, 50, 100]
model_maxDepth = [3, 5, 7]
model_maxBins = [32, 64]
# model_learningRate = [0.01, 0.1, 0.5]
# model_loss = ['logLoss', 'leastSquaresError', 'leastAbsoluteError']
list_of_param_dicts = []
for maxIter in model_maxIter:
for maxDepth in model_maxDepth:
for maxBins in model_maxBins:
params_dict = {}
params_dict['maxIter'] = maxIter
params_dict['maxDepth'] = maxDepth
params_dict['maxBins'] = maxBins
list_of_param_dicts.append(params_dict)
print("There is {} set of params.".format(len(list_of_param_dicts)))
# list_of_param_dicts
# -
prefix = 'gs://ai-covid19-datalake/trusted/experiment_map/'
# <hr />
# <hr />
# <hr />
# +
# filename = 'gs://ai-covid19-datalake/trusted/experiment_map/03-STRSAMP-AG/ds-1/cols_set_1/experiment0.parquet'
# df = spark.read.parquet(filename)
# df.limit(2).toPandas()
# +
# params_dict = {'maxIter': 100,
# 'maxDepth': 3,
# 'maxBins': 32,
# 'learningRate': 0.5,
# 'loss': 'leastAbsoluteError'}
# cols = 'cols_set_1'
# experiment_filter = 'ds-1'
# undersampling_method = '03-STRSAMP-AG',
# experiment_id = 0
# +
# run_gbt(df, params_dict, cols, filename, experiment_filter, undersampling_method, experiment_id)
# -
# <hr />
# <hr />
# <hr />
def run_gbt(exp_df, params_dict, cols, filename, experiment_filter,
undersampling_method, experiment_id):
import time
start_time = time.time()
n_covid = exp_df.filter(F.col('CLASSI_FIN') == 1.0).count()
n_not_covid = exp_df.filter(F.col('CLASSI_FIN') == 0.0).count()
id_cols = ['NU_NOTIFIC', 'CLASSI_FIN']
labelIndexer = StringIndexer(inputCol="CLASSI_FIN", outputCol="indexedLabel").fit(exp_df)
input_cols = [x for x in exp_df.columns if x not in id_cols]
assembler = VectorAssembler(inputCols = input_cols, outputCol= 'features')
exp_df = assembler.transform(exp_df)
# Automatically identify categorical features, and index them.
# Set maxCategories so features with > 4 distinct values are treated as continuous.
featureIndexer = VectorIndexer(inputCol="features", outputCol="indexedFeatures", maxCategories=30).fit(exp_df)
# Split the data into training and test sets (30% held out for testing)
(trainingData, testData) = exp_df.randomSplit([0.7, 0.3])
trainingData = trainingData.persist(StorageLevel.MEMORY_ONLY)
testData = testData.persist(StorageLevel.MEMORY_ONLY)
# Train a RandomForest model.
gbt = GBTClassifier(labelCol = "indexedLabel", featuresCol = "indexedFeatures",
maxIter = params_dict['maxIter'],
maxDepth = params_dict['maxDepth'],
maxBins = params_dict['maxBins'])
# Convert indexed labels back to original labels.
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel",
labels=labelIndexer.labels)
# Chain indexers and forest in a Pipeline
pipeline = Pipeline(stages=[labelIndexer, featureIndexer, gbt, labelConverter])
# Train model. This also runs the indexers.
model = pipeline.fit(trainingData)
# Make predictions.
predictions = model.transform(testData)
pred = predictions.select(['CLASSI_FIN', 'predictedLabel'])\
.withColumn('predictedLabel', F.col('predictedLabel').cast('double'))\
.withColumn('predictedLabel', F.when(F.col('predictedLabel') == 1.0, 'covid').otherwise('n-covid'))\
.withColumn('CLASSI_FIN', F.when(F.col('CLASSI_FIN') == 1.0, 'covid').otherwise('n-covid'))\
.toPandas()
y_true = pred['CLASSI_FIN'].tolist()
y_pred = pred['predictedLabel'].tolist()
report = classification_report(y_true, y_pred, output_dict=True)
evaluator_ROC = BinaryClassificationEvaluator(labelCol="indexedLabel", rawPredictionCol="prediction", metricName="areaUnderROC")
accuracy_ROC = evaluator_ROC.evaluate(predictions)
evaluator_PR = BinaryClassificationEvaluator(labelCol="indexedLabel", rawPredictionCol="prediction", metricName="areaUnderPR")
accuracy_PR = evaluator_PR.evaluate(predictions)
conf_matrix = confusion_matrix(y_true, y_pred)
result_dict = {}
result_dict['experiment_filter'] = experiment_filter
result_dict['undersampling_method'] = undersampling_method
result_dict['filename'] = filename
result_dict['experiment_id'] = experiment_id
result_dict['n_covid'] = n_covid
result_dict['n_not_covid'] = n_not_covid
result_dict['model_name'] = 'GBT'
result_dict['params'] = params_dict
result_dict['model_AUC_ROC'] = accuracy_ROC
result_dict['model_AUC_PR'] = accuracy_PR
result_dict['model_covid_precision'] = report['covid']['precision']
result_dict['model_covid_recall'] = report['covid']['recall']
result_dict['model_covid_f1'] = report['covid']['f1-score']
result_dict['model_not_covid_precision'] = report['n-covid']['precision']
result_dict['model_not_covid_recall'] = report['n-covid']['recall']
result_dict['model_not_covid_f1'] = report['n-covid']['f1-score']
result_dict['model_avg_precision'] = report['macro avg']['precision']
result_dict['model_avg_recall'] = report['macro avg']['recall']
result_dict['model_avg_f1'] = report['macro avg']['f1-score']
result_dict['model_avg_acc'] = report['accuracy']
result_dict['model_TP'] = conf_matrix[0][0]
result_dict['model_TN'] = conf_matrix[1][1]
result_dict['model_FN'] = conf_matrix[0][1]
result_dict['model_FP'] = conf_matrix[1][0]
result_dict['model_time_exec'] = time.time() - start_time
result_dict['model_col_set'] = cols
return result_dict
# <hr />
# <hr />
# <hr />
# # Running GBT on 10 samples for each experiment
# ### 3x col sets -> ['cols_set_1', 'cols_set_2', 'cols_set_3']
# ### 3x model_maxIter -> [100, 200, 300]
# ### 3x model_maxDepth -> [5, 10, 15]
# ### 3x model_maxBins -> [16, 32, 64]
# Total: 10 * 3 * 3 * 3 * 3 = 810
experiments = []
# ### Datasets: strat_samp_lab_agegrp
for uc in undersamp_col:
for ds in dfs:
for col_set in cols_sets:
for params_dict in list_of_param_dicts:
for id_exp in range(50):
filename = prefix + uc + '/' + ds + '/' + col_set + '/' + 'experiment' + str(id_exp) + '.parquet'
exp_dataframe = spark.read.parquet(filename)
# if 'SG_UF_NOT' in exp_dataframe.columns:
# exp_dataframe = exp_dataframe.withColumn('SG_UF_NOT', F.col('SG_UF_NOT').cast('float'))
print('read {}'.format(filename))
undersampling_method = uc
experiment_filter = ds
experiment_id = id_exp
try:
model = run_gbt(exp_dataframe, params_dict, col_set, filename, experiment_filter, undersampling_method, experiment_id)
experiments.append(model)
print("Parameters ==> {}\n Results: \n AUC_PR: {} \n Precision: {} \n Time: {}".format(str(params_dict), str(model['model_AUC_PR']), str(model['model_avg_precision']), str(model['model_time_exec'])))
print('=========================== \n')
except:
print('=========== W A R N I N G =========== \n')
print('Something wrong with the exp: {}, {}, {}'.format(filename, params_dict, col_set))
# <hr />
# <hr />
# <hr />
for i in range(len(experiments)):
for d in list(experiments[i].keys()):
experiments[i][d] = str(experiments[i][d])
# +
# experiments
# -
cols = ['experiment_filter', 'undersampling_method', 'filename', 'experiment_id', 'n_covid', 'n_not_covid', 'model_name', 'params', 'model_AUC_ROC', 'model_AUC_PR', 'model_covid_precision', 'model_covid_recall', 'model_covid_f1', 'model_not_covid_precision', 'model_not_covid_recall', 'model_not_covid_f1', 'model_avg_precision', 'model_avg_recall', 'model_avg_f1', 'model_avg_acc', 'model_TP', 'model_TN', 'model_FN', 'model_FP', 'model_time_exec', 'model_col_set']
intermed_results = spark.createDataFrame(data=experiments).select(cols)
intermed_results.toPandas()
intermed_results.write.parquet('gs://ai-covid19-datalake/trusted/intermed_results/KMODES/GBT_experiments-kmodes-ds1-cs3.parquet', mode='overwrite')
print('finished')
intermed_results.show()
|
05-models/02-experiment-design/07.1-run_gbt_model_kmodes-ds1-cs3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Compute conformer energies for a small molecule
#
# This notebook illustrates reading conformers of a molecule from an SDF file and computation of vacuum conformer energies using a SMIRNOFF force field.
#
# Note that absolute vacuum potential energies can be sensitive to small changes in partial charge, for example due to using OpenEye or AmberTools to generate AM1-BCC charges. However, in our experience, _relative_ conformer energies are fairly consistent between AM1-BCC implementations.
from openff.toolkit.topology import Molecule, Topology
from openff.toolkit.utils import RDKitToolkitWrapper, get_data_file_path
from rdkit.Chem import rdMolAlign
import numpy as np
from simtk import openmm, unit
import shutil
# +
# If using a OFF Toolkit version before 0.7.0, loading SDFs through RDKit and OpenEye may provide
# different behavior in some cases. So, here we force loading through RDKit to ensure the correct behavior
rdktkw = RDKitToolkitWrapper()
# Locate molecule in OpenFF Toolkit package data and copy to local directory
orig_path = get_data_file_path('molecules/ruxolitinib_conformers.sdf')
shutil.copy(orig_path, '.')
# Load in the molecule and its conformers.
# Note that all conformers of the same molecule are loaded as separate Molecule objects
loaded_molecules = Molecule.from_file('ruxolitinib_conformers.sdf', toolkit_registry=rdktkw)
# The logic below only works for lists of molecules, so if a
# single molecule was loaded, cast it to list
if type(loaded_molecules) is not list:
loaded_molecules = [loaded_molecules]
# Collatate all conformers of the same molecule
# NOTE: This isn't necessary if you have already loaded or created multi-conformer molecules;
# it is just needed because our SDF reader does not automatically collapse conformers.
molecules = [loaded_molecules[0]]
for molecule in loaded_molecules[1:]:
if molecule == molecules[-1]:
for conformer in molecule.conformers:
molecules[-1].add_conformer(conformer)
else:
molecules.append(molecule)
n_molecules = len(molecules)
n_conformers = sum([mol.n_conformers for mol in molecules])
print(f'{n_molecules} unique molecule(s) loaded, with {n_conformers} total conformers')
# -
# Load the openff-1.1.0 force field appropriate for vacuum calculations (without constraints)
from openff.toolkit.typing.engines.smirnoff import ForceField
forcefield = ForceField('openff_unconstrained-1.1.0.offxml')
# Loop over molecules and minimize each conformer
for molecule in molecules:
# If the molecule doesn't have a name, set mol.name to be the hill formula
if molecule.name == '':
molecule.name = Topology._networkx_to_hill_formula(molecule.to_networkx())
print('%s : %d conformers' % (molecule.name, molecule.n_conformers))
# Make a temporary copy of the molecule that we can update for each minimization
mol_copy = Molecule(molecule)
# Make an OpenFF Topology so we can parameterize the system
off_top = molecule.to_topology()
print(f"Parametrizing {molecule.name} (may take a moment to calculate charges)")
system = forcefield.create_openmm_system(off_top)
# Use OpenMM to compute initial and minimized energy for all conformers
integrator = openmm.VerletIntegrator(1*unit.femtoseconds)
platform = openmm.Platform.getPlatformByName('Reference')
omm_top = off_top.to_openmm()
simulation = openmm.app.Simulation(omm_top, system, integrator, platform)
# Print text header
print('Conformer Initial PE Minimized PE RMS between initial and minimized conformer')
output = [['Conformer','Initial PE (kcal/mol)','Minimized PE (kcal/mol)','RMS between initial and minimized conformer (Angstrom)']]
for conformer_index, conformer in enumerate(molecule.conformers):
simulation.context.setPositions(conformer)
orig_potential = simulation.context.getState(getEnergy=True).getPotentialEnergy()
simulation.minimizeEnergy()
min_state = simulation.context.getState(getEnergy=True, getPositions=True)
min_potential = min_state.getPotentialEnergy()
# Calculate the RMSD between the initial and minimized conformer
min_coords = min_state.getPositions()
min_coords = np.array([ [atom.x, atom.y, atom.z] for atom in min_coords]) * unit.nanometer
mol_copy._conformers = None
mol_copy.add_conformer(conformer)
mol_copy.add_conformer(min_coords)
rdmol = mol_copy.to_rdkit()
rmslist = []
rdMolAlign.AlignMolConformers(rdmol, RMSlist=rmslist)
minimization_rms = rmslist[0]
# Save the minimized conformer to file
mol_copy._conformers = None
mol_copy.add_conformer(min_coords)
mol_copy.to_file(f'{molecule.name}_conf{conformer_index+1}_minimized.sdf', file_format='sdf')
print('%5d / %5d : %8.3f kcal/mol %8.3f kcal/mol %8.3f Angstroms' % (conformer_index+1, molecule.n_conformers, orig_potential/unit.kilocalories_per_mole, min_potential/unit.kilocalories_per_mole, minimization_rms))
output.append([str(conformer_index+1),
f'{orig_potential/unit.kilocalories_per_mole:.3f}',
f'{min_potential/unit.kilocalories_per_mole:.3f}',
f'{minimization_rms:.3f}'])
# Write the results out to CSV
with open(f'{molecule.name}.csv', 'w') as of:
for line in output:
of.write(','.join(line)+'\n')
# Clean up OpenMM Simulation
del simulation, integrator
|
examples/conformer_energies/conformer_energies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Version du 3 avril 2020, prenant en compte les éléments choice (ajoutés entre-temps)
# +
#Revue le 15 avril. Options pour fichier déjà lemmatisé ajoutées.
def export_tokens_to_csv(chemin_entree, chemin_sortie):
"""
Fonction récupérant les éléments <w> d'un fichier XML-TEI
avec leurs @n pour les exporter dans un fichier CSV à
deux colonnes, l'une pour le numéro, l'autre pour le texte.
Si l'élément <w> comporte des enfants <height> pour les initiales
hautes, ou <choice> pour des modernisations/abréviations,
la fonction compilera une version complète, résolue et modernisée
pour le logiiel Analog.
:param chemin_entree: Le chemin local vers un fichier en XML-TEI
dont on souhaite extraire les tokens pour Analog.
:param chemin_sortie: L'emplacement local dans lequel on souhaite
écrire le fichier CSV à donner à Analog.
"""
import xml.etree.ElementTree as ET
import csv
# Le dictionnaire dans lequel seront stockés les token avant écriture en CSV.
# dico_tokens = {}
# Les colonnes du futur fichier CSV.
# colonnes = ['ID', 'TOKEN']
colonnes = ['ID', 'TOKEN']
# Pour que Python comprenne les éléments dont on parlera,
# il faut lui donner la déclaration TEI, mais comme c'est
# la seule qu'on utilisera, pas besoin de lui donner un préfixe.
# ET.register_namespace('', "http://tei-c.org/ns/1.0")
# On va chercher le fichier XML-TEI et on le lit.
tree = ET.parse(chemin_entree)
root = tree.getroot()
# On ouvre le CSV de sortie en mode "écriture", on y écrit le nom des colonnes.
with open(chemin_sortie, 'w') as csv_file:
csv_contenu = csv.DictWriter(csv_file, fieldnames = colonnes, delimiter=";")
csv_contenu.writeheader()
# On boucle sur les éléments <w> du XML, dans l'ordre du fichier.
for word in root.findall('.//w'):
# On récupère le @n dans la variable "numero"
# et on crée la chaîne "texte", pour l'instant vide.
numero = str(word.get('n'))
texte = ""
# Si <w> n'a pas d'enfant, on récupère le texte tel-quel.
if word.find('.') == None :
texte = str(word.text)
# Sinon, on compile.
else:
# S'il y a du texte avant le premier enfant, on l'ajoute.
if word.text:
texte += str(word.text)
# On boucle sur les enfants du <w> actuel.
for item in word:
# Si l'enfant est un <height>, on récupère son texte.
if item.tag == 'height' or item.tag == 'supplied':
texte += str(item.text)
# S'il y a du texte après la balise fermante et avant
# le prochain enfant ou la balise fermante du <w>,
# on l'ajoute.
if item.tail:
texte += str(item.tail)
elif item.tag == 'lb':
if item.tail:
texte += str(item.tail)
# Si l'enfant est un <choice>, on récupère le texte de son
# second enfant et on vérifie s'il y a du texte après le <choice>.
elif item.tag == 'choice':
texte += str(item[1].text)
if item.tail:
texte += str(item.tail)
elif item.tag == 'c':
texte += item.text
if item.tail:
texte += str(item.tail)
elif item.tag == 'hi':
texte += item.text
if item.tail:
texte += item.tail
elif item.tag == 'add':
# On refait tous les tests.
if item.find('.') == None :
texte = str(item.text)
else:
if item.text:
texte += str(item.text)
for subitem in item:
if subitem.tag == 'lb':
if subitem.tail:
texte += str(subitem.tail)
elif subitem.tag == 'choice':
texte += str(subitem[1].text)
if subitem.tail:
texte += str(subitem.tail)
# Une fois le mot copié ou compilé, on ajoute une unité au dictionnaire de tokens,
# dont la clé est le numéro (le @) et la valeur est la chaîne copiée ou compilée.
# dico_tokens[str(numero)] = str(texte)
# Puis on écrit les valeurs du mot dans les colonnes appropriées.
csv_contenu.writerow(
{
"ID" : numero,
"TOKEN" : str(texte),
}
)
# +
# Pour exécuter la fonction, remplacer les chemins par les chemins actuels,
# d'abord le XML, ensuite le chemin pour créer le CSV.
export_tokens_to_csv('/home/erminea/Documents/CONDE/Morisse-TS-XML/2021-06-07_morisse_w_REnumerotes.xml',
'/home/erminea/Documents/CONDE/Morisse-TS-XML/2021-06-07_morisse_pour_analog.csv')
|
new-corrections/lemmatize-latin-text/extraction_tokens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="z3saB-0g5XwS"
# ## Installs, imports and Boilerplate code
# + colab={} colab_type="code" id="DQb-CoVB4W9Z"
import warnings
warnings.filterwarnings(action='once')
# + colab={"base_uri": "https://localhost:8080/", "height": 83} colab_type="code" executionInfo={"elapsed": 2879, "status": "ok", "timestamp": 1575357200795, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="ijO161Io5AWK" outputId="6702f005-b7bf-4859-ae0a-4203f26b7c48"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import time
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Convolution2D, DepthwiseConv2D, SeparableConv2D
from keras.layers import Dense, Flatten, Activation
from keras.layers import Dropout, BatchNormalization, GlobalAveragePooling2D
from keras.utils import np_utils
from keras import backend as K
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" executionInfo={"elapsed": 5198, "status": "ok", "timestamp": 1575357227048, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="mAHRwUrt7lIM" outputId="b9999437-2d3d-4a4a-e223-c10c1a373229"
# loading and configuring cifar10 dataset
from keras.datasets import cifar10
# x_train/test : train/test features
# y_train/test : train/test labels
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
num_train, img_channels, img_rows, img_cols = x_train.shape
num_test = x_test.shape[0]
num_classes = len(np.unique(y_train))
# + colab={} colab_type="code" id="WLuAGWnr84cK"
class_names = ['airplane', 'automobile', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" executionInfo={"elapsed": 2401, "status": "ok", "timestamp": 1575357232370, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="wiD1KJ-M84FC" outputId="5b75b86d-7fe1-400c-f476-f5202ce49a69"
# https://stackoverflow.com/questions/45965007/multiline-f-string-in-python
print(f"""Train images : {num_train}
Test images : {num_test}
Total classes : {num_classes}
Input image shape : {x_train.shape[1:]}
""")
# + colab={} colab_type="code" id="Xt9JEPEDnbKe"
from keras.optimizers import Adam
from keras.callbacks import LearningRateScheduler
def scheduler(epoch, lr):
return round(0.003 * 1/(1 + 0.319 * epoch), 10)
# + colab={} colab_type="code" id="qtR-7WvHQH9Y"
def plot_model_history(model_history):
fig, ax = plt.subplots(1, 2, figsize=(15, 5))
# summarize history for accuracy
ax[0].plot(range(1, len(model_history.history['acc'])+1), model_history.history['acc'])#, label='Training Acc')
ax[0].plot(range(1, len(model_history.history['val_acc'])+1), model_history.history['val_acc'])#, label='Validation Acc')
ax[0].set_title('Model Accuracy')
ax[0].set_xlabel('Epochs')
ax[0].set_ylabel('Accuracy')
ax[0].set_xticks(np.arange(1, len(model_history.history['acc'])+1), len(model_history.history['acc'])/10)
ax[0].legend(['Train', 'Validation'], loc='best')
# summarize history for loss
ax[1].plot(range(1, len(model_history.history['loss'])+1), model_history.history['loss'])
ax[1].plot(range(1, len(model_history.history['val_loss'])+1), model_history.history['val_loss'])
ax[1].set_title('Model Loss')
ax[1].set_xlabel('Epochs')
ax[1].set_ylabel('Loss')
ax[1].set_xticks(np.arange(1, len(model_history.history['loss'])+1), len(model_history.history['loss'])/10)
ax[1].legend(['Train', 'Validation'], loc='best')
plt.show()
# + colab={} colab_type="code" id="1lGp_6CQUmbT"
def accuracy(test_x, test_y, model):
result = model.predict(test_x)
predicted = np.argmax(result, axis=1)
true = np.argmax(test_y, axis=1)
num_correct = np.sum(predicted == true)
accuracy = float(num_correct)/result.shape[0]
return (accuracy * 100)
# + colab={"base_uri": "https://localhost:8080/", "height": 213} colab_type="code" executionInfo={"elapsed": 2725, "status": "ok", "timestamp": 1575357236893, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="0py3Ab_5-CK-" outputId="284898fa-7531-4d89-83bb-df595cced050"
# Checking images, one for each class
fig = plt.figure(figsize=(8,3))
for i in range(num_classes):
ax = fig.add_subplot(2, 5, 1+i, xticks=[], yticks=[])
idx = np.where(y_train[:]==i)[0] # selecting all images of a class
features = x_train[idx,::]
img_num = np.random.randint(features.shape[0]) # selecting random no. from class idx
im = features[img_num] # selecting features of the image from random no.
ax.set_title(class_names[i])
plt.imshow(im)
plt.show()
# + colab={} colab_type="code" id="p8iKuwFZWsJW"
x_train = x_train.astype('float32')/255
x_test = x_test.astype('float32')/255
# converting class labels to one hot encoded
y_train = np_utils.to_categorical(y_train, num_classes)
y_test = np_utils.to_categorical(y_test, num_classes)
# + [markdown] colab_type="text" id="yIJ4z94omcuU"
# ## Model
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 6304, "status": "ok", "timestamp": 1575357254727, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="wfyDn3tIXTX6" outputId="c15e40d7-d669-4337-d17b-eb1d96ae5008"
# Define the model
model = Sequential()
model.add(DepthwiseConv2D(3, depth_multiplier=1, use_bias=False, padding='same', input_shape=(32, 32, 3))) # 32x3x3x32 RF:3 # padding==border_mode(old version)
model.add(Convolution2D(32, 1, use_bias=False, padding='same')) # 32x3x3x32 RF:
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.05))
model.add(DepthwiseConv2D(3, use_bias=False)) # 30x3x3x64
model.add(Convolution2D(64, 1, use_bias=False)) # 30x3x3x64
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.05))
model.add(DepthwiseConv2D(3, use_bias=False)) # 28x3x3x128
model.add(Convolution2D(128, 1, use_bias=False)) # 28x3x3x128
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.05))
model.add(MaxPooling2D(2)) # 14x3x3x128
model.add(Convolution2D(32, 1, use_bias=False)) # 14x3x3x32
model.add(DepthwiseConv2D(3, use_bias=False)) # 12x3x3x32 RF:3
model.add(Convolution2D(32, 1, use_bias=False)) # 12x3x3x32 RF:
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.1))
model.add(DepthwiseConv2D(3, use_bias=False)) # 10x3x3x64
model.add(Convolution2D(64, 1, use_bias=False)) # 10x3x3x64
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.1))
model.add(DepthwiseConv2D(3, use_bias=False)) # 8x3x3x128
model.add(Convolution2D(128, 1, use_bias=False)) # 8x3x3x128
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.1))
model.add(MaxPooling2D(2)) # 4x3x3x128
model.add(Convolution2D(32, 1, use_bias=False)) # 4x3x3x32
model.add(DepthwiseConv2D(3, use_bias=False)) # 2x3x3x32 RF:3
model.add(Convolution2D(32, 1, use_bias=False)) # 2x3x3x32 RF:
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.15))
model.add(Convolution2D(10, 1, use_bias=False)) # 2x3x3x10 RF:
model.add(GlobalAveragePooling2D()) # 1X10
model.add(Activation('softmax'))
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 110} colab_type="code" executionInfo={"elapsed": 2040, "status": "ok", "timestamp": 1575357384626, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="G6jeD_Vvfp-S" outputId="8f854af8-b4a5-42ad-eb06-51e3c05c2d53"
# Compile the model
model.compile(optimizer=Adam(lr=0.003), loss='categorical_crossentropy', metrics=['accuracy'])
# + colab={} colab_type="code" id="u-cr82_teNuQ"
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1105235, "status": "ok", "timestamp": 1575358494080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="S9uG39-EeapX" outputId="719cef0a-59ab-459b-a978-84f506847496"
# train the model
start = time.time()
# Train the model
model_info = model.fit_generator(datagen.flow(x_train, y_train, batch_size = 128),
samples_per_epoch = x_train.shape[0], nb_epoch = 50,
validation_data = (x_test, y_test),callbacks=[LearningRateScheduler(scheduler, verbose=1)], verbose=1)
end = time.time()
print ("Model took %0.2f seconds to train"%(end - start))
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" executionInfo={"elapsed": 1079977, "status": "ok", "timestamp": 1575358495697, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="ifmyuJy1edf-" outputId="cf8d8745-ca1e-456b-faa4-c978452ee5ef"
# plot model history
plot_model_history(model_info)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" executionInfo={"elapsed": 1080052, "status": "ok", "timestamp": 1575358496032, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mByTu2xLfZj1YR3txrkITcZs9_i74nbaVq6SBdT8Q=s64", "userId": "03151495481760299531"}, "user_tz": -330} id="xe6CIp5zhaGB" outputId="70a73142-9478-422e-80af-0e3aed50bb8b"
# compute test accuracy
print ("Accuracy on test data is: %0.2f"%accuracy(x_test, y_test, model))
# + colab={} colab_type="code" id="XTG-gPfMiCnh"
# + colab={} colab_type="code" id="9wUiDBEVmcd3"
# + colab={} colab_type="code" id="gEa21Y1UrTfg"
|
Week3/Models/S3_model2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:miniconda3-python-tutorial]
# language: python
# name: conda-env-miniconda3-python-tutorial-py
# ---
# # Regrid data
# This code shows as simple way to:
# - Read in a .nc file using xarray
# - regrid data using xesmf
# - calculate gridcell biases & RMSE, averaged over months
# ### -- NOTE: information around coasts is lost with this approach --
#
# +
import xarray as xr
import xesmf as xe
import numpy as np
import pandas as pd
import esmlab
from ctsm_py import utils
import warnings
# some resources for plotting
import matplotlib.pyplot as plt
import cartopy
import cartopy.crs as ccrs
# %matplotlib inline
# -
# supress Runtime warnings that let you know when code isn't too efficiently
warnings.simplefilter("ignore", category=RuntimeWarning)
file = '/gpfs/fs1/work/dlawren/obs_data/WECANN/WECANN_v1.0.metadata.nc'
ds = xr.open_dataset(file, decode_times=True) #time 2007-2015, no coordinates provided
# #### Regrid example from xesmf
# https://xesmf.readthedocs.io/en/latest/notebooks/Rectilinear_grid.html
# #### Modified here to target resolution for CLM
# +
variables = ['GPP']
# --- CLM5 w/ GSWP3 ---
model = 'CLM5_GSWP3'
pattern = '/glade/p/cgd/tss/people/oleson/CLM_LAND_ONLY_RELEASE/CLM5/clm50_r270_1deg_GSWP3V1_iso_newpopd_hist/'\
'lnd/proc/tseries/month_1/clm50_r270_1deg_GSWP3V1_iso_newpopd_hist.clm2.h0.{var}.185001-201412.nc'
file = [pattern.format(var=var) for var in variables]
var = variables
print(file[0])
# Time slices are not identical, get the last 8 years of data from CLM simulation
years = 8
months = years * 12
ds0 = utils.time_set_mid(xr.open_dataset(file[0], decode_times=True), 'time').isel(time=slice(-months,None))
print(ds0.time.shape)
print(ds.year.isel(time=slice(0,months)))
# +
# assign coordinates to time
# also flip longitude 0-360 & sort by ascending values (as with CLM grids)
# I don't think the lat-lon manipulations are really necessary, but it makes life easier
ds = ds.isel(time=slice(0,months))
ds = ds.assign({'time': (['time'], ds0.time),
'lon': (['lon'],xr.where(ds.lon<0, 360+ds.lon, ds.lon))})
ds = ds.sortby('lon', ascending=True)
ds = ds.sortby('lat', ascending=True)
ds['time_bounds'] = ds0.time_bounds #needed for esmlab if we want to calculate climatologies
dr = ds.GPP
dr.isel(time=0).plot(robust=True);
# how does WECAN provide negative GPP values somewhere?
# -
# #### -- Output grid --
ds_out = xr.Dataset({'lat': (['lat'], ds0.lat),
'lon': (['lon'], ds0.lon)})
ds_out
# +
regridder = xe.Regridder(ds, ds_out, 'bilinear')
regridder # print basic regridder information.
# NOTE: [from xesmf], Data arrays should have (lat, lon) on the right.
# If not, use DataArray.transpose or numpy.transpose to preprocess the data.
# Note this is also a pretty quick and dirty job, not sure how coastlines are bing handled?
# -
dr_out = regridder(dr)
dr_out.isel(time=0).plot(robust=True, cmap = "viridis");
ds0.GPP.isel(time=0).plot(robust=True);
## -- looks like we lost the coastline in this regridding...
# check results look similar
dr.sel(lon=260, lat=40, method='nearest').plot() # input data
dr_out.sel(lon=260, lat=40, method='nearest').plot(); # output data
# ### -- Quick statistics, bias and RMSE w/ plots --
ds0['bias'] = xr.DataArray(np.zeros(ds0.GPP.shape), dims=('time','lat', 'lon'))
ds0['bias'][...] = ds0.GPP.values - dr_out.values
# RMSE in setps (1) calculate bias squared (2) take mean over months, (3) square root
mse = xr.ufuncs.square(ds0.bias).groupby('time.month').mean(dim='time').rename('mse')
rmse = xr.ufuncs.sqrt(mse).rename('RMSE (gC/m2/d)')*3600*24 #convert to gC/m2/d
# Quick look at RMSE~month
simple = rmse.plot(x='lon', y='lat', col='month', col_wrap=4)
# Bias plots, here averaged for each month
bias = ds0.bias.groupby('time.month').mean(dim='time').rename('mean bias (gC/m2/d)')*3600*24
bias.plot(x='lon', y='lat', col='month', col_wrap=4, robust=True);
# ### TODO:
# - More useful to look at multiple models
# - Calculate global values, seasonal values or timeseries? How handled in diagnostics or ILAMB?
# - Preserve coastlines, does xesmf have tools to help with this?
|
notebooks/regrid_wecan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A - Using TorchText with Your Own Datasets
#
# In this series we have used the IMDb dataset included as a dataset in TorchText. TorchText has many canonical datasets included for classification, language modelling, sequence tagging, etc. However, frequently you'll be wanting to use your own datasets. Luckily, TorchText has functions to help you to this.
#
# Recall in the series, we:
# - defined the `Field`s
# - loaded the dataset
# - created the splits
#
# As a reminder, the code is shown below:
#
# ```python
# TEXT = data.Field()
# LABEL = data.LabelField()
#
# train_data, test_data = datasets.IMDB.splits(TEXT, LABEL)
#
# train_data, valid_data = train_data.split()
# ```
# There are three data formats TorchText can read: `json`, `tsv` (tab separated values) and`csv` (comma separated values).
#
# **In my opinion, the best formatting for TorchText is `json`, which I'll explain later on.**
#
# ## Reading JSON
#
# Starting with `json`, your data must be in the `json lines` format, i.e. it must be something like:
#
# ```
# {"name": "John", "location": "United Kingdom", "age": 42, "quote": ["i", "love", "the", "united kingdom"]}
# {"name": "Mary", "location": "United States", "age": 36, "quote": ["i", "want", "more", "telescopes"]}
# ```
#
# That is, each line is a `json` object. See `data/train.json` for an example.
#
# We then define the fields:
# +
from torchtext.legacy import data
from torchtext.legacy import datasets
NAME = data.Field()
SAYING = data.Field()
PLACE = data.Field()
# -
# Next, we must tell TorchText which fields apply to which elements of the `json` object.
#
# For `json` data, we must create a dictionary where:
# - the key matches the key of the `json` object
# - the value is a tuple where:
# - the first element becomes the batch object's attribute name
# - the second element is the name of the `Field`
#
# What do we mean when we say "becomes the batch object's attribute name"? Recall in the previous exercises where we accessed the `TEXT` and `LABEL` fields in the train/evaluation loop by using `batch.text` and `batch.label`, this is because TorchText sets the batch object to have a `text` and `label` attribute, each being a tensor containing either the text or the label.
#
# A few notes:
#
# * The order of the keys in the `fields` dictionary does not matter, as long as its keys match the `json` data keys.
#
# - The `Field` name does not have to match the key in the `json` object, e.g. we use `PLACE` for the `"location"` field.
#
# - When dealing with `json` data, not all of the keys have to be used, e.g. we did not use the `"age"` field.
#
# - Also, if the values of `json` field are a string then the `Fields` tokenization is applied (default is to split the string on spaces), however if the values are a list then no tokenization is applied. Usually it is a good idea for the data to already be tokenized into a list, this saves time as you don't have to wait for TorchText to do it.
#
# - The value of the `json` fields do not have to be the same type. Some examples can have their `"quote"` as a string, and some as a list. The tokenization will only get applied to the ones with their `"quote"` as a string.
#
# - If you are using a `json` field, every single example must have an instance of that field, e.g. in this example all examples must have a name, location and quote. However, as we are not using the age field, it does not matter if an example does not have it.
fields = {'name': ('n', NAME), 'location': ('p', PLACE), 'quote': ('s', SAYING)}
# Now, in a training loop we can iterate over the data iterator and access the name via `batch.n`, the location via `batch.p`, and the quote via `batch.s`.
#
# We then create our datasets (`train_data` and `test_data`) with the `TabularDataset.splits` function.
#
# The `path` argument specifices the top level folder common among both datasets, and the `train` and `test` arguments specify the filename of each dataset, e.g. here the train dataset is located at `data/train.json`.
#
# We tell the function we are using `json` data, and pass in our `fields` dictionary defined previously.
train_data, test_data = data.TabularDataset.splits(
path = 'data',
train = 'train.json',
test = 'test.json',
format = 'json',
fields = fields
)
# If you already had a validation dataset, the location of this can be passed as the `validation` argument.
train_data, valid_data, test_data = data.TabularDataset.splits(
path = 'data',
train = 'train.json',
validation = 'valid.json',
test = 'test.json',
format = 'json',
fields = fields
)
# We can then view an example to make sure it has worked correctly.
#
# Notice how the field names (`n`, `p` and `s`) match up with what was defined in the `fields` dictionary.
#
# Also notice how the word `"United Kingdom"` in `p` has been split by the tokenization, whereas the `"united kingdom"` in `s` has not. This is due to what was mentioned previously, where TorchText assumes that any `json` fields that are lists are already tokenized and no further tokenization is applied.
print(vars(train_data[0]))
# We can now use `train_data`, `test_data` and `valid_data` to build a vocabulary and create iterators, as in the other notebooks. We can access all attributes by using `batch.n`, `batch.p` and `batch.s` for the names, places and sayings, respectively.
#
# ## Reading CSV/TSV
#
# `csv` and `tsv` are very similar, except csv has elements separated by commas and tsv by tabs.
#
# Using the same example above, our `tsv` data will be in the form of:
#
# ```
# name location age quote
# John United Kingdom 42 i love the united kingdom
# Mary United States 36 i want more telescopes
# ```
#
# That is, on each row the elements are separated by tabs and we have one example per row. The first row is usually a header (i.e. the name of each of the columns), but your data could have no header.
#
# You cannot have lists within `tsv` or `csv` data.
#
# The way the fields are defined is a bit different to `json`. We now use a list of tuples, where each element is also a tuple. The first element of these inner tuples will become the batch object's attribute name, second element is the `Field` name.
#
# Unlike the `json` data, the tuples have to be in the same order that they are within the `tsv` data. Due to this, when skipping a column of data a tuple of `None`s needs to be used, if not then our `SAYING` field will be applied to the `age` column of the `tsv` data and the `quote` column will not be used.
#
# However, if you only wanted to use the `name` and `age` column, you could just use two tuples as they are the first two columns.
#
# We change our `TabularDataset` to read the correct `.tsv` files, and change the `format` argument to `'tsv'`.
#
# If your data has a header, which ours does, it must be skipped by passing `skip_header = True`. If not, TorchText will think the header is an example. By default, `skip_header` will be `False`.
fields = [('n', NAME), ('p', PLACE), (None, None), ('s', SAYING)]
train_data, valid_data, test_data = data.TabularDataset.splits(
path = 'data',
train = 'train.tsv',
validation = 'valid.tsv',
test = 'test.tsv',
format = 'tsv',
fields = fields,
skip_header = True
)
print(vars(train_data[0]))
# Finally, we'll cover `csv` files.
#
# This is pretty much the exact same as the `tsv` files, expect with the `format` argument set to `'csv'`.
fields = [('n', NAME), ('p', PLACE), (None, None), ('s', SAYING)]
train_data, valid_data, test_data = data.TabularDataset.splits(
path = 'data',
train = 'train.csv',
validation = 'valid.csv',
test = 'test.csv',
format = 'csv',
fields = fields,
skip_header = True
)
print(vars(train_data[0]))
# ## Why JSON over CSV/TSV?
#
# 1. Your `csv` or `tsv` data cannot be stored lists. This means data cannot be already be tokenized, thus everytime you run your Python script that reads this data via TorchText, it has to be tokenized. Using advanced tokenizers, such as the `spaCy` tokenizer, takes a non-negligible amount of time. Thus, it is better to tokenize your datasets and store them in the `json lines` format.
#
# 2. If tabs appear in your `tsv` data, or commas appear in your `csv` data, TorchText will think they are delimiters between columns. This will cause your data to be parsed incorrectly. Worst of all TorchText will not alert you to this as it cannot tell the difference between a tab/comma in a field and a tab/comma as a delimiter. As `json` data is essentially a dictionary, you access the data within the fields via its key, so do not have to worry about "surprise" delimiters.
# ## Iterators
#
# Using any of the above datasets, we can then build the vocab and create the iterators.
NAME.build_vocab(train_data)
SAYING.build_vocab(train_data)
PLACE.build_vocab(train_data)
# Then, we can create the iterators after defining our batch size and device.
#
# By default, the train data is shuffled each epoch, but the validation/test data is sorted. However, TorchText doesn't know what to use to sort our data and it would throw an error if we don't tell it.
#
# There are two ways to handle this, you can either tell the iterator not to sort the validation/test data by passing `sort = False`, or you can tell it how to sort the data by passing a `sort_key`. A sort key is a function that returns a key on which to sort the data on. For example, `lambda x: x.s` will sort the examples by their `s` attribute, i.e their quote. Ideally, you want to use a sort key as the `BucketIterator` will then be able to sort your examples and then minimize the amount of padding within each batch.
#
# We can then iterate over our iterator to get batches of data. Note how by default TorchText has the batch dimension second.
# +
import torch
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
BATCH_SIZE = 1
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
sort = False, #don't sort test/validation data
batch_size=BATCH_SIZE,
device=device)
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
sort_key = lambda x: x.s, #sort by s attribute (quote)
batch_size=BATCH_SIZE,
device=device)
print('Train:')
for batch in train_iterator:
print(batch)
print('Valid:')
for batch in valid_iterator:
print(batch)
print('Test:')
for batch in test_iterator:
print(batch)
|
A - Using TorchText with Your Own Datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# language: python
# name: python3
# ---
# # Merge Bugianen and serve as pmtiles
#
# It's all in the title :)
# Pre-requisite:
# * Bugianen files downloaded in the working directory below (links in https://tartamillo.wordpress.com/bugianen/)
# * s3 object store setup done in aws-cli (for the very end)
# %cd ~/Downloads/dwnmaps/mapdata/mbtiles/Bugianen
import sys
sys.path.append('../../eslope/development/src')
from mbt_util import mbt_merge, mbt_info
mbt_merge('Bugianen 2005 Cervino.mbtiles',
'Bugianen 2005 Cuneese.mbtiles',
'Bugianen 2005 Germanasca.mbtiles',
'Bugianen 2005 Gran Paradiso.mbtiles',
'Bugianen 2005 Lanzo.mbtiles',
'Bugianen 2005 Monregalese.mbtiles',
'Bugianen 2005 Monte Bianco.mbtiles',
'Bugianen 2005 Monte Rosa.mbtiles',
'Bugianen 2005 Monviso.mbtiles',
'Bugianen 2005 Ossola.mbtiles',
'Bugianen 2005 Susa.mbtiles',
'Bugianen 2005 Verbano.mbtiles',
dest='Bugianen.mbtiles',
description='https://tartamillo.wordpress.com/bugianen/')
# Don't keep duplicate indexes
import sqlite3
db = sqlite3.connect('Bugianen.mbtiles')
try:
db.cursor().execute('DROP INDEX IF EXISTS tiles_id')
finally:
db.commit()
db.close()
mbt_info('Bugianen.mbtiles')
# !command -v pmtiles-convert &> /dev/null || pip install pmtiles
# !pmtiles-convert Bugianen.{mb,pm}tiles
# +
# configure your s3
# pip install awscli-plugin-endpoint
# aws configure set plugins.endpoint awscli_plugin_endpoint
# aws configure
# -
# !aws s3 cp --acl public-read ./Bugianen.mbtiles s3://maki/
# !aws s3 cp --acl public-read ./Bugianen.pmtiles s3://maki/
# !aws s3api put-bucket-cors --bucket maki --cors-configuration \
# '{"CORSRules": [{"AllowedOrigins": ["*"], "AllowedHeaders": ["*"], "AllowedMethods": ["GET", "HEAD"], "MaxAgeSeconds": 3000, "ExposeHeaders": ["Etag"]}]}'
|
topo_merge/Bugianen-merge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from glob import glob
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
exps = glob('Compiled/*')
vcs = glob('Inbal/*/value_counts*')
convert = {
'590': '10',
'610': '8',
'620': '7',
'630': '6a',
'640': '6B',
}
# +
vc_dict = {}
for vc in vcs:
key = vc.split('/')[1]
test = pd.read_csv(vc)
vc_dict[key] = test
# +
exp_dict = {}
for exp in exps:
nm = exp.split('_')[1].split(' ')[0]
with open(exp) as fp:
#The first 14 lines of the qTable do not contain spectrum data
for blank in range(0,2):
fp.readline()
wave = []
Q_ext = []
# Q_abs = []
# Q_sca = []
for line in fp:
ary = line.split(",")
wave.append(float(ary[0]))
Q_ext.append(float(ary[1]))
df = pd.DataFrame({'wave': wave, q: Q_ext})
exp_dict[nm] = df
exp_dict[nm] = exp_dict[nm].drop(range(0, 100)).reset_index(drop=True)
# exp_dict[nm] = exp_dict[nm].iloc[::2, :].reset_index(drop=True)
# +
q = 'Q_sca'
x_dict = {}
for x in glob(f'x_*_new_interp_{q}.csv'):
test = pd.read_csv(x)
x_dict[int(x.split('_')[1])] = test
yz_dict = {}
for yz in glob(f'yz_*_new_interp_{q}.csv'):
test = pd.read_csv(yz)
yz_dict[int(yz.split('_')[1])] = test
net_dict = {}
for key in x_dict:
net_dict[key] = x_dict[key] * (1.0/3.0) + yz_dict[key] * (2.0/3.0)
# +
import numpy as np
best_dict = {}
for peak in convert:
print(peak)
exp_temp = exp_dict[peak]
vc_temp = vc_dict[convert[peak]]
vc_temp = vc_temp.reset_index()
sp = 1.15
total_count = 0
test_spectra = exp_dict[peak].copy()
test_spectra[q] = 0.0
for _, row in vc_temp.iterrows():
leng = row['Size']
num = row['Count']
if leng > 35:
break
spdf = net_dict[leng]['%.2f' % sp]
test_spectra[q] += spdf * num
total_count += num
test_spectra[q] = test_spectra[q] / total_count
plt.plot(test_spectra['wave'], test_spectra[q], label='theory')
plt.legend()
plt.title(f'{q}_{sp}nm_spacing')
plt.xlabel(f'Wavelength (nm)')
plt.ylabel(f'Intensity (arbitrary units)')
plt.savefig(f'outimg/{peak}_{q}_sample.png')
plt.close()
df = pd.DataFrame({'wl': test_spectra['wave'], 'computed': test_spectra[q]})
df.to_csv(f'outimg/{peak}_{q}_sample.csv')
# -
|
dda_analysis/q_tables.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# A cheatsheet for Python standard library <br />
# Software License Agreement (MIT License) <br />
# Copyright (c) 2019, <NAME>.
# # "Python Standard Library"
# # math
import math
# +
# constants
print(math.pi)
print(math.e)
print(math.nan)
print(math.inf)
print(-math.inf)
# +
# trigonometry
a1 = math.cos(math.pi / 4) # cos(45) deg
a2 = math.sin(math.pi / 4) # sin(45) deg
print(a1)
print(a2)
# +
# ciel & floor
print(math.ceil(47.3)) # rounds up
print(math.ceil(47.8))
print(math.ceil(48))
print(math.floor(47.3)) # rounds down
print(math.floor(47.8))
print(math.floor(47))
# +
# factorial & square root
print(math.factorial(3))
print(math.sqrt(64))
# +
# GCD : Gratest Common Denominator
print(math.gcd(52,8))
# +
# degrees & radians
print(math.radians(360))
print(math.degrees(math.pi * 2))
# -
# # random
import random
# +
print(random.random()) # float random between 0 & 1
print(random.randrange(2)) # random 0 or 1
print(random.randrange(1,7)) # random between 1 & 6
winners = random.sample(range(100),5) # selects 5 numbers from the range and returns in a list
print(winners)
pets = ['cat', 'dog', 'fish', 'kitten']
print(random.choice(pets))
random.shuffle(pets)
print(pets)
# -
# # statistics
import statistics
# +
data = [10, 15, 10, 11, 12, 10, 10, 13, 14]
print(statistics.mean(data)) # avarage
print(statistics.mode(data)) # most frequent value
print(statistics.median(data)) # mid point of data
print(statistics.variance(data)) # variance - the avarage of squared differences from the mean, tells how varied is the data
print(statistics.stdev(data)) # standard diviation - the square root of devience
# -
# # itertools
import itertools
# infinite count
for x in itertools.count(50,5):
print(x)
if x >= 70: break
# infinite cycle
i = 0
for c in itertools.cycle('RACECAR'):
print(c)
i += 1
if i >= 10: break
# infinite repeat
i = 0
for r in itertools.repeat(True):
print(r)
i += 1
if i >= 10: break
# +
# permutations - all posssible orders of a data
dic1 = {1:'bob' , 2:'john' , 3:'linda'}
for p1 in itertools.permutations(dic1):
print(p1)
for p2 in itertools.permutations(dic1.values()):
print(p2)
# -
# combinations - all posssible orders of a particular number of data
colors = ['red', 'blue', 'green', 'pink']
for c in itertools.combinations(colors, 2):
print(c)
# # command line arguments
# assuming that some arguments are passed as file runs in cmd
import sys
print(sys.argv) # prints the arguments, first argument is file path
sys.argv.remove(sys.argv[0]) # removes the first argument
print(sys.argv)
# # tempfile
import tempfile
tmp = tempfile.TemporaryFile()
tmp.write(b'some data on temp file') # b changes data to byte
tmp.seek(0)
print(tmp.read())
tmp.close()
# # HTML parser
# +
from html.parser import HTMLParser
class HTMLParser(HTMLParser):
def handle_starttag(self, tag, attrs):
print('Start tag: ', tag)
for atr in attrs:
print('attrs:', atr)
def handle_endtag(self, tag):
print('End tag: ', tag)
def handle_comment(self,com):
print('Comment: ', com)
def handle_data(self, data):
print('Data: ', data)
parser = HTMLParser()
parser.feed('<html><head><title>Code</title></head><body><h1><!--hi!-->I am a Coder!</h1></body></html>')
print()
# -
# html data from consule
inhtml = input('Put soem HTML code: ')
parser.feed(inhtml)
print()
# html data from a file
htmlfile = open('sample_html.html', 'r')
s = ''
for line in htmlfile:
s += line
parser.feed(s)
# # text wrap
# +
import textwrap
textdata = ''' This is a text data
for testing text wrapper module
in Python standard libaray.'''
print('No Dedent:') # keeps the beginning tab, does not keep the enters
print(textwrap.fill(textdata))
print('-------------')
print('Dedent: ') # removes the beginning spases and keeps our enters
dedtxt = textwrap.dedent(textdata).strip()
print(dedtxt)
print('-------------')
print('Fill: ')
print(textwrap.fill(dedtxt, width=80)) # sets next line by rhe given width
print('')
print(textwrap.fill(dedtxt, width=10))
print('-------------')
print('Controlled indent: ')
print(textwrap.fill(dedtxt, initial_indent=' ' , subsequent_indent=' '))
print('-------------')
print('Shortening text: ')
shr = textwrap.shorten('Some text data for testing', width=20, placeholder='...')
print(shr)
# -
# # HTTP package, urllib, json
# +
import urllib.request
import json
import textwrap
with urllib.request.urlopen("https://www.googleapis.com/books/v1/volumes?q=isbn:1101904224") as f:
text = f.read()
decodedtext = text.decode('utf-8')
print(textwrap.fill(decodedtext, width=50))
print('------------------------------------------')
obj = json.loads(decodedtext)
print(obj['kind'])
print('------------------------------------------')
print(obj['items'][0]['searchInfo']['textSnippet'])
# -
|
Python_Standard_Llibrary/Python Standard Library .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Install the function using
# >`pip install inequalipy`
# Import the function
import inequalipy as ineq
# Randomly create a distribution:
import numpy as np
a = np.random.normal(5,1,100)
weights = np.ones(len(a), dtype=int)
# weights = np.random.randint(0,100,len(a), dtype=int)
# # Gini Index
# our function
ineq.gini(a)
# our function with weights (of ones)
ineq.gini(a, weights)
ineq.gini
# Pysal's gini coefficient
import inequality as pysal
pysal.gini._gini(a)
# Grasia's gini coefficient
from example import grasia
grasia.gini(a)/100
# # Atkinson
# our function
ineq.atkinson.index(a, 0.5)
from example import ineqpy
ineqpy.atkinson(a,e=0.5)
# # Kolm-Pollak
# our function
ineq.kolmpollak.ede(a, epsilon=0.5)
|
example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from astropy.table import Table
import astropy
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator
import matplotlib
# -
# %matplotlib notebook
print('numpy version: {}'.format(np.__version__))
print('pandas version: {}'.format(pd.__version__))
print('matplotlib version: {}'.format(matplotlib.__version__))
print('astropy version: {}'.format(astropy.__version__))
# # Figure 3
#
# Create Figure 3 (the relative rates of SN types in the ZTF BTS, ASAS-SN, and LOSS) in [Fremling et al. 2020](https://ui.adsabs.harvard.edu/abs/2019arXiv191012973F/abstract).
#
# Data for ASAS-SN are from [Holoien et al. 2019](https://ui.adsabs.harvard.edu/abs/2019MNRAS.484.1899H/abstract), while data for LOSS are from [Li et al. 2011](https://ui.adsabs.harvard.edu/abs/2011MNRAS.412.1441L/abstract).
# BTS data
bts_df = pd.read_hdf('../data/final_rcf_table.h5')
# +
normal_sne = np.where((bts_df.sn_type != 'SLSN-I') &
(bts_df.sn_type != 'SLSN-II') &
(bts_df.sn_type != 'SLSN-R') &
(bts_df.sn_type != 'ambiguous')
)
all_ia = np.where((bts_df.sn_type == 'Ia') |
(bts_df.sn_type == 'Ia-91T') |
(bts_df.sn_type == 'Ia-91bg') |
(bts_df.sn_type == 'Ia-02cx') |
(bts_df.sn_type == 'Ia-csm') |
(bts_df.sn_type == 'Ia-SC')
)
all_ii = np.where((bts_df.sn_type == 'II') |
(bts_df.sn_type == 'IIn') |
(bts_df.sn_type == 'IIb') |
(bts_df.sn_type == 'II-87A')
)
all_se = np.where((bts_df.sn_type == 'Ic') |
(bts_df.sn_type == 'Ib') |
(bts_df.sn_type == 'Ic-BL') |
(bts_df.sn_type == 'Ib/c') |
(bts_df.sn_type == 'Ibn') |
(bts_df.sn_type == 'Ic-pec')
)
slsne = np.where((bts_df.sn_type == 'SLSN-I') |
(bts_df.sn_type == 'SLSN-II') |
(bts_df.sn_type == 'SLSN-R')
)
# -
n_sn = len(bts_df) -3
print('There are {} SNe in the BTS sample'.format(n_sn))
print('{} of these are not superluminous SNe in the BTS sample'.format(len(normal_sne[0])))
print('\t {} ({:.3f}%) are SNe Ia'.format(len(all_ia[0]), 100*len(all_ia[0])/n_sn))
print('\t {} ({:.3f}%) are SNe II'.format(len(all_ii[0]), 100*len(all_ii[0])/n_sn))
print('\t {} ({:.3f}%) are SNe Ib/c'.format(len(all_se[0]), 100*len(all_se[0])/n_sn))
print('\t {} ({:.3f}%) are SLSNe'.format(len(slsne[0]), 100*len(slsne[0])/n_sn))
# #### Read in SN data from ASAS-SN
# +
n_asas_ia = 0
n_asas_91T = 0
n_asas_91bg = 0
n_asas_ii = 0
n_asas_ibc = 0
n_asas_slsn = 0
asas_offset = np.array([])
for release in ['1','2','3','4']:
tab1 = '../data/ASAS_SN/bright_sn_catalog_{}/table1.txt'.format(release)
tab2 = '../data/ASAS_SN/bright_sn_catalog_{}/table2.txt'.format(release)
asassn_tab1 = Table.read(tab1, format='cds')
asassn_tab2 = Table.read(tab2, format='cds')
n_asas_ia += len(np.where( (asassn_tab1['Type'] == 'Ia') |
(asassn_tab1['Type'] == 'Ia-91T') |
(asassn_tab1['Type'] == 'Ia-91bg') |
(asassn_tab1['Type'] == 'Ia+CSM') |
(asassn_tab1['Type'] == 'Ia-pec') |
(asassn_tab1['Type'] == 'Ia-00cx') |
(asassn_tab1['Type'] == 'Ia-06bt') |
(asassn_tab1['Type'] == 'Ia-07if') |
(asassn_tab1['Type'] == 'Ia-09dc') |
(asassn_tab1['Type'] == 'Ia-02cx')
)[0])
n_asas_91T += len(np.where( (asassn_tab1['Type'] == 'Ia-91T') )[0])
n_asas_91bg += len(np.where( (asassn_tab1['Type'] == 'Ia-91bg') )[0])
n_asas_ii += len(np.where( (asassn_tab1['Type'] == 'II') |
(asassn_tab1['Type'] == 'IIP') |
(asassn_tab1['Type'] == 'IIb') |
(asassn_tab1['Type'] == 'II-pec') |
(asassn_tab1['Type'] == 'IIn') |
(asassn_tab1['Type'] == 'IIn-pec') |
(asassn_tab1['Type'] == 'IIn/LBV') |
(asassn_tab1['Type'] == 'IIn-09ip')
)[0])
n_asas_ibc += len(np.where( (asassn_tab1['Type'] == 'Ib') |
(asassn_tab1['Type'] == 'Ib/c') |
(asassn_tab1['Type'] == 'Ibn') |
(asassn_tab1['Type'] == 'Ic') |
(asassn_tab1['Type'] == 'Ic-pec') |
(asassn_tab1['Type'] == 'Ib/c-BL') |
(asassn_tab1['Type'] == 'Ic-BL')
)[0])
n_asas_slsn += len(np.where( (asassn_tab1['Type'] == 'SLSN-II') |
(asassn_tab1['Type'] == 'SLSN-I')
)[0])
n_asas_ia += len(np.where( ( (asassn_tab2['Type'] == 'Ia') |
(asassn_tab2['Type'] == 'Ia-91T') |
(asassn_tab2['Type'] == 'Ia-91bg') |
(asassn_tab2['Type'] == 'Ia+CSM') |
(asassn_tab2['Type'] == 'Ia-pec') |
(asassn_tab2['Type'] == 'Ia-00cx') |
(asassn_tab2['Type'] == 'Ia-06bt') |
(asassn_tab2['Type'] == 'Ia-07if') |
(asassn_tab2['Type'] == 'Ia-09dc') |
(asassn_tab2['Type'] == 'Ia-02cx')
) &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
n_asas_91T += len(np.where( (asassn_tab2['Type'] == 'Ia-91T') &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
n_asas_91bg += len(np.where( (asassn_tab2['Type'] == 'Ia-91bg') &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
n_asas_ii += len(np.where( ( (asassn_tab2['Type'] == 'II') |
(asassn_tab2['Type'] == 'IIP') |
(asassn_tab2['Type'] == 'IIb') |
(asassn_tab2['Type'] == 'II-pec') |
(asassn_tab2['Type'] == 'IIn') |
(asassn_tab2['Type'] == 'IIn-pec') |
(asassn_tab2['Type'] == 'IIn/LBV') |
(asassn_tab2['Type'] == 'IIn-09ip')
) &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
n_asas_ibc += len(np.where( ( (asassn_tab2['Type'] == 'Ib') |
(asassn_tab2['Type'] == 'Ib/c') |
(asassn_tab2['Type'] == 'Ibn') |
(asassn_tab2['Type'] == 'Ic') |
(asassn_tab2['Type'] == 'Ic-pec') |
(asassn_tab2['Type'] == 'Ib/c-BL') |
(asassn_tab2['Type'] == 'Ic-BL')
) &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
n_asas_slsn += len(np.where( ( (asassn_tab2['Type'] == 'SLSN-II') |
(asassn_tab2['Type'] == 'SLSN-I')
) &
(asassn_tab2['Recovered'] == 'Yes')
)[0])
asas_offset = np.append(asas_offset, np.array(asassn_tab1['Offset'][asassn_tab1['HostName'] != 'None'], dtype=float))
asas_offset = np.append(asas_offset,
np.array(asassn_tab2['Offset'][np.where((asassn_tab2['Recovered'] == 'Yes') &
(asassn_tab2['SNName'] != 'PS16dtm'))], dtype=float))
tot_asas = n_asas_ia + n_asas_ii + n_asas_ibc + n_asas_slsn
# +
color_dict = {'blue': '#2C5361',
'orange': '#DB6515',
'yellow': '#CF8E36',
'maroon': '#3B2525',
'purple': '#A588AC',
'beige': '#D2A176'}
fig, ax = plt.subplots()
bar_width = 0.25
epsilon = 0.05
# CI are calculated MultinomCI - available here: https://rdrr.io/cran/DescTools/man/MultinomCI.html
bts_upper = [0.76194918, 0.24740644, 0.08006433, 0.04625050]
bts_lower = [0.67118508, 0.16597629, 0.03415659, 0.01334088]
asassn_upper = [0.78238445, 0.25404044, 0.06727064, 0.01589015]
asassn_lower = [0.6971419929, 0.1744027864, 0.0269571897, 0.0008384526]
# mean
ax.bar(np.arange(4) - bar_width - epsilon,
[len(all_ia[0])/n_sn, len(all_ii[0])/n_sn,
len(all_se[0])/n_sn, len(slsne[0])/n_sn],
bar_width, lw=2, edgecolor=color_dict['orange'], facecolor='None',
hatch='\\', label='ZTF BTS', zorder=10)
# mean + unc
ax.bar(np.arange(4) - bar_width - epsilon,
bts_upper,
bar_width, lw=2, color=color_dict['orange'], edgecolor='None', alpha=0.4)
# mean - unc
ax.bar(np.arange(4) - bar_width - epsilon,
bts_lower,
bar_width, lw=0, color='white', edgecolor='None', zorder=5)
# show Li 11 with uncertainties
ax.bar(np.arange(4),
[0.793, 0.166, 0.041, 0.0],
bar_width, lw=2, edgecolor='0.6', facecolor='None',
label='LOSS', zorder=10)
ax.bar(np.arange(4),
[0.834, 0.216, 0.057, 0.0],
bar_width, lw=2, color='0.6', edgecolor='None', alpha=0.4)
ax.bar(np.arange(4),
[0.737, 0.127, 0.029, 0.0],
bar_width, color='white', zorder=5, lw=0)
# mean
ax.bar(np.arange(4) + bar_width + epsilon,
[n_asas_ia/tot_asas, n_asas_ii/tot_asas,
n_asas_ibc/tot_asas, n_asas_slsn/tot_asas],
bar_width, lw=2, edgecolor=color_dict['blue'], facecolor='None',
hatch='///', label='ASAS-SN', zorder=10)
# mean + unc
ax.bar(np.arange(4) + bar_width + epsilon,
asassn_upper,
bar_width, lw=2, color=color_dict['blue'], edgecolor='None', alpha=0.4)
# # mean - unc
ax.bar(np.arange(4) + bar_width + epsilon,
asassn_lower,
bar_width, lw=0, color='white', edgecolor='None', zorder=5)
ax.tick_params(labelsize=11)
ax.set_xticks([0,1,2,3])
ax.set_xticklabels(['Ia', 'II', 'Ibc', 'SLSN'],fontsize=14)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.yaxis.set_minor_locator(MultipleLocator(0.1))
ax.yaxis.set_major_locator(MultipleLocator(0.2))
ax.set_ylabel('flux-limited fraction',fontsize=14)
leg = ax.legend(fancybox=True, markerscale=20,
labelspacing=1.5, handlelength=4,
fontsize=14, loc=1, borderpad=0.75,
bbox_to_anchor=(0.9,0.8))
for patch in leg.get_patches():
patch.set_height(22)
patch.set_y(-6)
fig.subplots_adjust(left=0.1,bottom=0.07,top=0.99,right=0.99)
fig.savefig('mag_limit_rates.pdf')
|
figures/Figure3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from skimage import data
from skimage import exposure
img = data.moon()
img_eq = exposure.equalize_hist(img)
img_adapthist = exposure.equalize_adapthist(img, clip_limit=0.03)
output = [img, img_eq, img_adapthist]
titles = ['Original', 'Histogram Equalization', 'Adaptive Equalization']
for i in range(3):
plt.subplot(3, 1, i+1)
plt.imshow(output[i], cmap='gray')
plt.title(titles[i])
plt.axis('off')
plt.show()
# -
|
Chapter 11/Section17_03_Histogram_Equalization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:cntk-py34]
# language: python
# name: conda-env-cntk-py34-py
# ---
# + [markdown] nbpresent={"id": "29b9bd1d-766f-4422-ad96-de0accc1ce58"}
# # CNTK 103: Part B - Feed Forward Network with MNIST
#
# ### This notebook was covered in the Live Webinar on 4th of April 2017
# ### The Webinar recording is available [HERE...](https://channel9.msdn.com/blogs/Cloud-and-Enterprise-Premium/Data-Science-Virtual-Machine--A-Walkthrough-of-end-to-end-Analytics-Scenarios)
#
# ## We assume that you have successfully completed CNTK 103 Part A.
#
# In this tutorial we will train a fully connected network on MNIST data. This notebook provides the recipe using Python APIs. If you are looking for this example in BrainScript, please look [here](https://github.com/Microsoft/CNTK/tree/v2.0.beta7.0/Examples/Image/GettingStarted)
#
# ## Introduction
#
# **Problem** (recap from the CNTK 101):
#
# The MNIST data comprises of hand-written digits with little background noise.
#
# **Goal**:
# Our goal is to train a classifier that will identify the digits in the MNIST dataset.
#
# <img src="https://github.com/Azure/DataScienceVM/blob/master/Tutorials/WebinarDocuments-04-04-2017/MiscAssets/DNNdiag1.png?raw=true">
#
# **Approach**:
# The same 5 stages we have used in the previous tutorial are applicable: Data reading, Data preprocessing, Creating a model, Learning the model parameters and Evaluating (a.k.a. testing/prediction) the model.
# - Data reading: We will use the CNTK Text reader
# - Data preprocessing: Covered in part A (suggested extension section).
#
# Rest of the steps are kept identical to CNTK 102.
# + nbpresent={"id": "138d1a78-02e2-4bd6-a20e-07b83f303563"}
# Import the relevant components
from __future__ import print_function
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import sys
import os
from cntk.device import gpu, set_default_device
from cntk import Trainer, StreamConfiguration, text_format_minibatch_source, learning_rate_schedule, UnitType
from cntk.initializer import glorot_uniform
from cntk.learner import sgd
from cntk.ops import *
from IPython.display import Image
# %matplotlib inline
# -
# Specify the target device to be used for computing (this example is showing for GPU usage)
# Note we need to set the device only once during the session.
set_default_device(gpu(0))
# ## Data reading
#
# In this section, we will read the data generated in CNTK 103 Part A.
# +
# Ensure we always get the same amount of randomness
np.random.seed(0)
# Define the data dimensions
input_dim = 784
num_output_classes = 10
# -
# ### Input and Labels
#
# In this tutorial we are using the MNIST data you have downloaded using CNTK_103A_MNIST_DataLoader notebook. The dataset has 60,000 training images and 10,000 test images with each image being 28 x 28 pixels. Thus the number of features is equal to 784 (= 28 x 28 pixels), 1 per pixel. The variable `num_output_classes` is set to 10 corresponding to the number of digits (0-9) in the dataset.
# +
# Ensure the training data is generated and available for this tutorial
train_file = "data/MNIST/Train-28x28_cntk_text.txt"
if os.path.isfile(train_file):
path = train_file
else:
raise ValueError("Please generate the data by completing CNTK 103 Part A")
feature_stream_name = 'features'
labels_stream_name = 'labels'
mb_source = text_format_minibatch_source(path, [
StreamConfiguration(feature_stream_name, input_dim),
StreamConfiguration(labels_stream_name, num_output_classes)])
features_si = mb_source[feature_stream_name]
labels_si = mb_source[labels_stream_name]
print("Training data from file {0} successfully read.".format(path))
# -
# <a id='#Model Creation'></a>
# ## Model Creation
#
# Our feed forward network will be relatively simple with 2 hidden layers (`num_hidden_layers`) with each layer having 200 hidden nodes (`hidden_layers_dim`).
#
# If you are not familiar with the terms *hidden_layer* and *number of hidden layers*, please refer back to CNTK 102 tutorial.
#
# For this tutorial: The number of green nodes (refer to picture above) in each hidden layer is set to 200 and the number of hidden layers (refer to the number of layers of green nodes) is 2. Fill in the following values:
# - num_hidden_layers
# - hidden_layers_dim
#
# Note: In this illustration, we have not shown the bias node (introduced in the logistic regression tutorial). Each hidden layer would have a bias node.
#
#
# <img src="https://github.com/Azure/DataScienceVM/blob/master/Tutorials/WebinarDocuments-04-04-2017/MiscAssets/feedforward_network.jpg?raw=true">
#
num_hidden_layers = 2
hidden_layers_dim = 400
# Network input and output:
# - **input** variable (a key CNTK concept):
# >An **input** variable is a container in which we fill different observations in this case image pixels during model learning (a.k.a.training) and model evaluation (a.k.a. testing). Thus, the shape of the `input_variable` must match the shape of the data that will be provided. For example, when data are images each of height 10 pixels and width 5 pixels, the input feature dimension will be 50 (representing the total number of image pixels). More on data and their dimensions to appear in separate tutorials.
#
#
# **Question** What is the input dimension of your chosen model? This is fundamental to our understanding of variables in a network or model representation in CNTK.
#
# +
# The input variable (representing 1 observation, in our example of age and size) $\bf{x}$ which
# in this case has a dimension of 2.
#
# The label variable has a dimensionality equal to the number of output classes in our case 2.
input = input_variable((input_dim), np.float32)
label = input_variable((num_output_classes), np.float32)
# -
# ## Feed forward network setup
#
# If you are not familiar with the feedforward network, please refer to CNTK 102. In this tutorial we are using the same network.
#
# Let us define the feedforward network one step at a time. The first layer takes an input feature vector ($\bf{x}$) with dimensions (`input_dim`) say $m$) and emits the output a.k.a. *evidence* (first hidden layer $\bf{z_1}$ with dimension (`hidden_layer_dim`) say $n$). Each feature in the input layer is connected with a node in the output later by the weight which is represented by a matrix $\bf{W}$ with dimensions ($m \times n$). The first step is to compute the evidence for the entire feature set. Note: we use **bold** notations to denote matrix / vectors:
#
# $$\bf{z_1} = \bf{W} \cdot \bf{x} + \bf{b}$$
#
# where $\bf{b}$ is a bias vector of dimension $n$.
#
# In the `linear_layer` function, we perform two operations:
# 0. multiply the weights ($\bf{W}$) with the features ($\bf{x}$) and add individual features' contribution,
# 1. add the bias term $\bf{b}$.
#
# The next step is to convert the *evidence* (the output of the linear layer) through a non-linear function a.k.a. *activation functions* of your choice that would squash the evidence to activations using a choice of functions ([found here][]). **Sigmoid** or **Tanh** are historically popular. We will use **sigmoid** function in this tutorial. The output of the sigmoid function often is the input to the next layer or the output of the final layer.
# [found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
#
# **Question**: Try different activation functions by passing different them to `nonlinearity` value and get familiarized with using them.
#
# Now that we have created one hidden layer, we need to iterate through the layers to create a fully connected classifier. Output of the first layer $\bf{h_1}$ becomes the input to the next layer.
#
# In this example we have only 2 layers, hence one could conceivably write the code as:
#
# >`h1 = fully_connected_layer(input, hidden_layer_dim, sigmoid)`
#
# >`h2 = fully_connected_layer(h1, hidden_layer_dim, sigmoid)`
#
# However, this code becomes very quickly difficult to read and update when the number of layers or blocks (in convolutional or recurrent networks) that we will see in later tutorials. CNTK provides a programming construct shown below that greatly eases the burden on the programmer.
#
# >`h = fully_connected_layer(input, hidden_layer_dim, sigmoid)`
#
# >`for i in range(1, num_hidden_layers):`
#
# >>` h = fully_connected_layer(h, hidden_layer_dim, sigmoid)`
#
# This construct is very attractive to write compact representation of large repetitive network components and will be used in many of the subsequent tutorials.
# +
# Define a fully connected feedforward network
def linear_layer(input_var, output_dim):
input_dim = input_var.shape[0]
times_param = parameter(shape=(input_dim, output_dim), init=glorot_uniform())
bias_param = parameter(shape=(output_dim))
t = times(input_var, times_param)
return bias_param + t
def dense_layer(input, output_dim, nonlinearity):
r = linear_layer(input, output_dim)
r = nonlinearity(r)
return r;
def fully_connected_classifier_net(input, num_output_classes, hidden_layer_dim,
num_hidden_layers, nonlinearity):
h = dense_layer(input, hidden_layer_dim, nonlinearity)
for i in range(1, num_hidden_layers):
h = dense_layer(h, hidden_layer_dim, nonlinearity)
r = linear_layer(h, num_output_classes)
return r
# -
# `z` will be used to represent the output of a network.
#
# We introduced sigmoid function in CNTK 102, in this tutorial you should try different activation functions. You may choose to do this right away and take a peek into the performance later in the tutorial or run the preset tutorial and then choose to perform the suggested activity.
#
#
# ** Suggested Activity **
# - Record the training error you get with `sigmoid` as the activation function
# - Now change to `relu` as the activation function and see if you can improve your training error
#
# *Quiz*: Different supported activation functions can be [found here][]. Which activation function gives the least training error?
#
# [found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
# Scale the input to 0-1 range by dividing each pixel by 256.
scaled_input = element_times(constant(1.0 / 256.0), input)
# Create the fully connected classifier.
z = fully_connected_classifier_net(scaled_input, num_output_classes, hidden_layers_dim, num_hidden_layers, relu)
#
# ## Learning model parameters
#
# Now that the network is setup, we would like to learn the parameters $\bf w$ and $\bf b$ for each of the layers in our network. To do so we convert, the computed evidence ($\bf z_{final~layer}$) into a set of predicted probabilities ($\textbf p$) using a `softmax` function.
#
# $$ \textbf{p} = \mathrm{softmax}(\bf{z_{final~layer}})$$
#
# One can see the `softmax` function as an activation function that maps the accumulated evidences to a probability distribution over the classes (Details of the [softmax function][]). Other choices of activation function can be [found here][].
#
# [softmax function]: https://www.cntk.ai/pythondocs/cntk.ops.html?highlight=softmax#cntk.ops.softmax
#
# [found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
#
#
# ## Training
#
# Similar to CNTK 102, we use minimize the cross-entropy between the label and predicted probability by the network. If this terminology sounds strange to you, please refer to the CNTK 102 for a refresher.
#
# If you have already gone through CNTK101, please skip this section and jump to the section titled,
# <a href='#Run the trainer'>Run the trainer'</a>.
#
# The output of the `softmax` is a probability of observations belonging to the respective classes. For training the classifier, we need to determine what behavior the model needs to mimic. In other words, we want the generated probabilities to be as close as possible to the observed labels. This function is called the *cost* or *loss* function and shows what is the difference between the learnt model vs. that generated by the training set.
#
# $$ H(p) = - \sum_{j=1}^C y_j \log (p_j) $$
#
# where $p$ is our predicted probability from `softmax` function and $y$ represents the label. This label provided with the data for training is also called the ground-truth label. In the two-class example, the `label` variable has dimensions of two (equal to the `num_output_classes` or $C$). Generally speaking, if the task in hand requires classification into $C$ different classes, the label variable will have $C$ elements with 0 everywhere except for the class represented by the data point where it will be 1. Understanding the [details][] of this cross-entropy function is highly recommended.
#
# [`cross-entropy`]: http://cntk.ai/pythondocs/cntk.ops.html#cntk.ops.cross_entropy_with_softmax
# [details]: http://colah.github.io/posts/2015-09-Visual-Information/
#
#
#
loss = cross_entropy_with_softmax(z, label)
# #### Evaluation
#
# In order to evaluate the classification, one can compare the output of the network which for each observation emits a vector of evidences (can be converted into probabilities using `softmax` functions) with dimension equal to number of classes.
label_error = classification_error(z, label)
# ### Configure training
#
# The trainer strives to reduce the `loss` function by different optimization approaches, [Stochastic Gradient Descent][] (`sgd`) being one of the most popular one. Typically, one would start with random initialization of the model parameters. The `sgd` optimizer would calculate the `loss` or error between the predicted label against the corresponding ground-truth label and using [gradient-decent][] generate a new set model parameters in a single iteration.
#
# The aforementioned model parameter update using a single observation at a time is attractive since it does not require the entire data set (all observation) to be loaded in memory and also requires gradient computation over fewer datapoints, thus allowing for training on large data sets. However, the updates generated using a single observation sample at a time can vary wildly between iterations. An intermediate ground is to load a small set of observations and use an average of the `loss` or error from that set to update the model parameters. This subset is called a *minibatch*.
#
# With minibatches we often sample observation from the larger training dataset. We repeat the process of model parameters update using different combination of training samples and over a period of time minimize the `loss` (and the error). When the incremental error rates are no longer changing significantly or after a preset number of maximum minibatches to train, we claim that our model is trained.
#
# One of the key parameter for optimization is called the `learning_rate`. For now, we can think of it as a scaling factor that modulates how much we change the parameters in any iteration. We will be covering more details in later tutorial.
# With this information, we are ready to create our trainer.
#
# [optimization]: https://en.wikipedia.org/wiki/Category:Convex_optimization
# [Stochastic Gradient Descent]: https://en.wikipedia.org/wiki/Stochastic_gradient_descent
# [gradient-decent]: http://www.statisticsviews.com/details/feature/5722691/Getting-to-the-Bottom-of-Regression-with-Gradient-Descent.html
# Instantiate the trainer object to drive the model training
learning_rate = 0.2
lr_schedule = learning_rate_schedule(learning_rate, UnitType.minibatch)
learner = sgd(z.parameters, lr_schedule)
trainer = Trainer(z, loss, label_error, [learner])
# First let us create some helper functions that will be needed to visualize different functions associated with training.
# +
from cntk.utils import get_train_eval_criterion, get_train_loss
# Define a utility function to compute the moving average sum.
# A more efficient implementation is possible with np.cumsum() function
def moving_average(a, w=5):
if len(a) < w:
return a[:] # Need to send a copy of the array
return [val if idx < w else sum(a[(idx-w):idx])/w for idx, val in enumerate(a)]
# Defines a utility that prints the training progress
def print_training_progress(trainer, mb, frequency, verbose=1):
training_loss = "NA"
eval_error = "NA"
if mb%frequency == 0:
training_loss = get_train_loss(trainer)
eval_error = get_train_eval_criterion(trainer)
if verbose:
print ("Minibatch: {0}, Loss: {1:.4f}, Error: {2:.2f}%".format(mb, training_loss, eval_error*100))
return mb, training_loss, eval_error
# -
# <a id='#Run the trainer'></a>
# ### Run the trainer
#
# We are now ready to train our fully connected neural net. We want to decide what data we need to feed into the training engine.
#
# In this example, each iteration of the optimizer will work on `minibatch_size` sized samples. We would like to train on all 60000 observations. Additionally we will make multiple passes through the data specified by the variable `num_sweeps_to_train_with`. With these parameters we can proceed with training our simple feed forward network.
# Initialize the parameters for the trainer
minibatch_size = 64
num_samples_per_sweep = 60000
num_sweeps_to_train_with = 10
num_minibatches_to_train = (num_samples_per_sweep * num_sweeps_to_train_with) / minibatch_size
# +
# Run the trainer on and perform model training
training_progress_output_freq = 500
plotdata = {"batchsize":[], "loss":[], "error":[]}
for i in range(0, int(num_minibatches_to_train)):
mb = mb_source.next_minibatch(minibatch_size)
# Specify the input variables mapping in the model to actual minibatch data to be trained
arguments = {input: mb[features_si],
label: mb[labels_si]}
trainer.train_minibatch(arguments)
batchsize, loss, error = print_training_progress(trainer, i, training_progress_output_freq, verbose=1)
if not (loss == "NA" or error =="NA"):
plotdata["batchsize"].append(batchsize)
plotdata["loss"].append(loss)
plotdata["error"].append(error)
# -
# Let us plot the errors over the different training minibatches. Note that as we iterate the training loss decreases though we do see some intermediate bumps.
#
# Hence, we use smaller minibatches and using `sgd` enables us to have a great scalability while being performant for large data sets. There are advanced variants of the optimizer unique to CNTK that enable harnessing computational efficiency for real world data sets and will be introduced in advanced tutorials.
# +
# Compute the moving average loss to smooth out the noise in SGD
plotdata["avgloss"] = moving_average(plotdata["loss"])
plotdata["avgerror"] = moving_average(plotdata["error"])
# Plot the training loss and the training error
import matplotlib.pyplot as plt
plt.figure(1)
plt.subplot(211)
plt.plot(plotdata["batchsize"], plotdata["avgloss"], 'b--')
plt.xlabel('Minibatch number')
plt.ylabel('Loss')
plt.title('Minibatch run vs. Training loss')
plt.show()
plt.subplot(212)
plt.plot(plotdata["batchsize"], plotdata["avgerror"], 'r--')
plt.xlabel('Minibatch number')
plt.ylabel('Label Prediction Error')
plt.title('Minibatch run vs. Label Prediction Error')
plt.show()
# -
# ## Evaluation / Testing
#
# Now that we have trained the network, let us evaluate the trained network on the test data. This is done using `trainer.test_minibatch`.
# +
# Ensure the training data is read and available for this tutorial
test_file = "data/MNIST/Test-28x28_cntk_text.txt"
if os.path.isfile(test_file):
path = test_file
else:
print("Please generate the data by completing CNTK 103 Part A")
feature_stream_name = 'features'
labels_stream_name = 'labels'
test_mb_source = text_format_minibatch_source(path, [
StreamConfiguration(feature_stream_name, input_dim),
StreamConfiguration(labels_stream_name, num_output_classes)])
features_si = mb_source[feature_stream_name]
labels_si = mb_source[labels_stream_name]
print("Test data from file {0} successfully read".format(path))
# +
# Test data for trained model
test_minibatch_size = 512
num_samples = 10000
num_minibatches_to_test = num_samples / test_minibatch_size
test_result = 0.0
for i in range(0, int(num_minibatches_to_test)):
mb = test_mb_source.next_minibatch(test_minibatch_size)
# Specify the mapping of input variables in the model to actual
# minibatch data to be tested with
arguments = {input: mb[features_si],
label: mb[labels_si]}
eval_error = trainer.test_minibatch(arguments)
test_result = test_result + eval_error
# Average of evaluation errors of all test minibatches
print("Average test error: {0:.2f}%".format(test_result*100 / num_minibatches_to_test))
# -
# Note, this error is very comparable to our training error indicating that our model has good "out of sample" error a.k.a. generalization error. This implies that our model can very effectively deal with previously unseen observations (during the training process). This is key to avoid the phenomenon of overfitting.
# We have so far been dealing with aggregate measures of error. Let us now get the probabilities associated with individual data points. For each observation, the `eval` function returns the probability distribution across all the classes. The classifier is trained to recognize digits, hence has 10 classes. First let us route the network output through a `softmax` function. This maps the aggregated activations across the network to probabilities across the 10 classes.
out = softmax(z)
# Let us a small minibatch sample from the test data.
# +
mb = test_mb_source.next_minibatch(test_minibatch_size)
predicted_label_prob = out.eval({input : mb[features_si]})
#orig_label=np.array(mb[labels_si].m_data.data().to_numpy())
orig_label = np.asarray(mb[labels_si].m_data)
# -
# Find the index with the maximum value for both predicted as well as the ground truth
pred = [np.argmax(predicted_label_prob[i,:,:]) for i in range(0,predicted_label_prob.shape[0])]
gtlabel = [np.argmax(orig_label[i,:,:]) for i in range(0, orig_label.shape[0])]
print("Label :", gtlabel[:25])
print("Predicted:", pred[:25])
# Let us visualize some of the results
# +
# Plot a random image
sample_number = 2
#img_data = mb[features_si].m_data.data().to_numpy()
img_data = mb[features_si].value
plt.imshow(img_data[sample_number,:,:].reshape(28,28), cmap="gray_r")
plt.axis('off')
img_gt, img_pred = gtlabel[sample_number], pred[sample_number]
print("Image Label: ", img_pred)
# -
# **Exploration Suggestion**
# - Try exploring how the classifier behaves with different parameters - suggest changing the `minibatch_size` parameter from 25 to say 64 or 128. What happens to the error rate? How does the error compare to the logistic regression classifier?
# - Suggest trying to increase the number of sweeps
# - Can you change the network to reduce the training error rate? When do you see *overfitting* happening?
# #### Code link
#
# If you want to try running the tutorial from python command prompt. Please run the [SimpleMNIST.py](https://github.com/Microsoft/CNTK/tree/v2.0.beta7.0/Examples/Image/Classification/MLP/Python) example.
|
Tutorials/WebinarDocuments-04-04-2017/CNTK_103B_MNIST_FeedForwardNetwork.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# <NAME>
# WSU ID: 10696738
# -
# ## Programming Assignment 2 - Part 1
# ### Cpts 355 - Spring 2016
# ### An Interpreter for a Postscript-like Language
# ### Assigned Feb. 3, 2016
# ### Due Friday, Feb. 12, 2016
# Develop your code in a file named `sps.ipynb`, starting from this notebook file. When you are finished, upload `sps.ipynb` on the course Turnin Page.
#
# The entire interpreter project (Parts 1 and Part 2 together) will count for 10% of your course grade. This first part is worth 20% of that 10%: the intention is to make sure that you are on the right track and have a chance for mid-course correction before completing Part 2. However, note that the work and amount of code involved in Part 1 is a large fraction of the total project, so you need to get going on this part right away.
# ### This assignment is to be your own work. Refer to the course academic integrity statement in the syllabus.
# ## The problem
# In this assignment you will write an interpreter in Python for a small PostScript-like language, concentrating on key computational features of the abstract machine, omitting all PS features related to graphics, and using a somewhat-simplified syntax.
#
# The simplified language, SPS, has the following features of PS
# * integer constants, e.g. `123`: in python3 there is no practical limit on the size of integers
# * boolean constants, `true` and `false` (Note that the boolean constants in python are `True` and `False`)
# * name constants, e.g. `/fact`: start with a `/` and letter followed by an arbitrary sequence of letters and numbers
# * names to be looked up in the dictionary stack, e.g. `fact`: as for name constants, without the `/`
# * code constants: code between matched curly braces `{` ... `}`
# * built-in operators on numbers: `add`, `sub`, `mul`, `div`, `eq`, `lt`, `gt`
# * built-in operators on boolean values: `and`, `or`, `not`; these take boolean operands only. Anything else is an error.
# * built-in sequencing operators: `if`, `ifelse`; make sure that you understand the order of the operands on the stack. Play with ghostscript if necessary to help understand what is happening.
# * stack operators: `dup`, `exch`, `pop`
# * dictionary creation operator: `dict`; takes one operand from the operand stack, ignores it, and creates a new, empty dictionary on the operand stack
# * dictionary stack manipulation operators: `begin`, `end`. `begin` requires one dictionary operand on the operand stack; `end` has no operands.
# * name definition operator: `def`. This requires two operands, a name and a value
# * defining (using `def`) and calling functions
# * stack printing operator (prints contents of stack without changing it): `stack`
# * top-of-stack printing operator (pops the top element of the stack and prints it): `=`
#
# ## Requirements for Part 1 (Due Feb. 12)
# In Part 1 you will build some essential pieces of the interpreter but not yet the full interpreter. The pieces you build will be driven by Python test code rather than actual Postscript programs. The pieces you are going to build first are:
# * The operand stack
# * The dictionary stack
# * The operators that don't involve code arrays: all of the operators except `if`, `ifelse`.
# In Part 2 we will add the implementations for `if`, `ifelse`, calling functions, as well as interpreting input strings in the Postscript language.
# * Looking up names
# ### The operand stack
# The operand stack should be implemented as a Python list. The list will contain **Python** integers, booleans, and strings, and later in Part 2 code arrays. Python integers and booleans on the stack represent Postscript integers and booleans. Python strings on the stack represent names of Postscript variables (see the handling of names and the `def` operator below.
#
# When using a list as a stack one of the decisions you have to make is where the *hot* end of the stack is located. (The *hot* end is where pushing and popping happens). Will the hot end be at position `0`, the head of the list, or at position `-1`, the end of the list? It's your choice.
# ### The dictionary stack
# The dictionary stack is also implemented as a Python list. It will contain **Python** dictionaries which will be the implementation for **Postscript** dictionaries. The dictionary stack needs to support adding and removing dictionaries at the hot end, as well as defining and looking up names.
# ### Operators
# Operators will be implemented as zero-argument Python functions that manipulate the operand and dictionary stacks. For example, the `add` operator could be implemented as the Python function (with comments instead of actual implementations)
# ```
# def add():
# op1 = # pop the top value off the operand stack
# op2 = # pop the top value off the operand stack
# # push (op1 + op2) onto the operand stack
# ```
# You may run into conflicts for some of the names of these functions . For example, the function for the `not` operator can't be named `not` because it is reserved for another use in Python. So you could do something like:
# ```
# def psnot():
# // pop the top value off the operand stack and push its negation onto the operand stack
# ```
# The `begin` and `end` operators are a little different in that they manipulate the dictionary stack in addition to or instead of the operand stack. Remember that the `dict` operator affects *only* the operand stack.
#
# The `def` operator takes two operands from the operand stack: a string (recall that strings in the operand stack represent names of postscript variables) and a value. It changes the dictionary at the hot end of the dictionary stack so that the string is mapped to the value by that dictionary. Notice that `def` does ***not*** change the number of dictionaries on the dictionary stack!
#
# ### Name lookup
#
# Name lookup is implemented by a Python function:
# ```
# def lookup(name):
# # search the dictionaries on the dictionary stack starting at the hot end to find one that contains name
# # return the value associated with name
# ```
# Note that name lookup is ***not*** a Postscript operator, but it ***is*** implemented by a Python function.
# ## Your Code Start Here
# +
# The operand stack: define the operand stack and its operations in this notebook cell
opstack = []
# now define functions to push and pop values on the opstack according to your decision about which
# end should be the hot end. Recall that `pass` in python is a no-op: replace it with your code.
def opPop():
return opstack.pop()
def opPush(value):
opstack.append(value)
# Remember that there is a Postscript operator called "pop" so we choose different names for these functions.
# +
# The dictionary stack: define the dictionary stack and its operations in this cell
dstack = [{}]
# now define functions to push and pop dictionaries on the dictstack, to define name, and to lookup a name
def dictPop():
a = dstack.pop()
if(a == {}):
dictPush(a)
return {}
else:
return a
def dictPush(value):
dstack.append(value)
def define(name, value):
a = opstack.pop()
b = opstack.pop()
x = dstack.pop()
x[b] = a
dstack.append(x)
def lookup(name):
# return the value associated with name
# what is your design decision about what to do when there is no definition for name
# need to determine if name is a number or a name
#if name.isdigit():
# return name
if type(name) == type(1):
return name
else:
# need to lookup the name in the dict
for a in dstack:
if(a == {}):
return None
if name in a.keys():
return a[name]
# if we get here than name doesn't exist
return None
# -
# ## Comment: It should look up from the top to the bottom of dictstack.
# ## -2
#
# +
# Arithmetic operators: define all the arithmetic operators in this cell -- add, sub, mul, div, eq, lt, gt
def EQ():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(a == b):
opstack.append("true")
else:
opstack.append("false")
def Add():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
sum = int(a) + int(b)
opstack.append(sum)
def Mul():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
prod = int(a) * int(b)
opstack.append(prod)
def Div():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(a == 0):
exit()
else:
quote = int(b) / int(a)
opstack.append(quote)
def Lt():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(b < a):
opstack.append("true")
else:
opstack.append("false")
def Gt():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(b > a):
opstack.append("true")
else:
opstack.append("false")
# +
# Boolean operators: define all the boolean operators in this cell -- and, or, not
def And():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(a == 'true' and b == 'true'):
opstack.append('true')
else:
opstack.append('false')
def Not():
a = lookup(opstack.pop())
if(a == 'false'):
opstack.append('true')
else:
opstack.append('false')
def Or():
a = lookup(opstack.pop())
b = lookup(opstack.pop())
if(a == 'true' or b == 'true'):
opstack.append('true')
else:
opstack.append('false')
# +
# Define the stack manipulation operators in this cell: dup, exch, pop
def Dup():
a = opstack.pop()
opstack.append(a)
opstack.append(a)
def Exch():
a = opstack.pop()
b = opstack.pop()
opstack.append(a)
opstack.append(b)
def Pop():
opstack.pop()
# +
# Define the dictionary manipulation operators in this cell: dict, begin, end, def
# name the function for the def operator psDef because def is reserved in Python
def Dict():
# takes one operand from the operand stack, ignores it, and creates a new, empty dictionary on the operand stack
opstack.pop()
dstack.append({})
def Begin(d):
# begin pushes a dictionary on the dictionary stack
dstack.append(d)
def End():
# end removes a dictionary from the dictionary stack
dstack.pop()
def Def():
# def creates or modifies an entry in the top dictionary on the dictionary stack.
a = opstack.pop()
b = opstack.pop()
d = dstack.pop()
d[b] = a
dstack.append(d)
# +
# Define the printing operators in this cell: =, stack
# Pick a good name for the code implementing =
def PrintOP():
print(opstack)
def PrintVal():
a = lookup(opstack.pop())
print (a)
# +
# Define any other operators that I may have forgotten in this cell
# -
# ## Test your code
# With all of that stuff defined, you will be able to test your interpreter using Python code like this:
# +
def testAdd():
opPush(1)
opPush(2)
Add()
if opPop() != 3: return False
return True
def testLookup():
opPush("n1")
opPush(3)
Def()
if lookup("n1") != 3: return False
return True
# go on writing test code for ALL of your code here; think about edge cases, and
# other points where you are likely to make a mistake.
# now an easy way to run all the test cases and make sure that they all return true
# is
testCases = [testAdd, testLookup] # add the names of your test functions to this list
def testAll1():
for test in testCases:
if not test(): return False
return True
# but wouldn't it be nice to run all the tests, instead of stopping on the first failure,
# and see which ones failed
# How about something like:
testCases = [('add', testAdd), ('lookup', testLookup)] # add you test functions to this list along with suitable names
def testAll2():
failedTests = [testName for (testName, testProc) in testCases if not testProc()]
if failedTests:
return ('Some tests failed', failedTests)
else: return ('All tests OK')
# -
testAll2()
# ## Comment:
# ## Excellent Overall!
# ## Final Score = 98
#
|
Postscrpit Interperator/sps(graded).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Vector plot
# +
#from IPython.display import Image
#Image(filename="quiver.jpg")
from pylab import *
# Set limits and number of points in grid
xmax = 2.0
xmin = -xmax
NX = 10
ymax = 2.0
ymin = -ymax
NY = 10
# Make grid and calculate vector components
x = linspace(xmin, xmax, NX)
y = linspace(ymin, ymax, NY)
X, Y = meshgrid(x, y)
S2 = X**2 + Y**2 # This is the radius squared
Bx = -Y/S2
By = +X/S2
figure()
QP = quiver(X,Y,Bx,By)
quiverkey(QP, 0.85, 1.02, 1.0, '1 mT', labelpos='N')
# Set the left, right, bottom, top limits of axes
dx = (xmax - xmin)/(NX - 1) # One less gap than points
dy = (ymax - ymin)/(NY - 1)
axis([xmin-dx, xmax+dx, ymin-dy, ymax+dy])
title('Magnetic Field of a Wire')
xlabel('x (cm)')
ylabel('y (cm)')
show()
# -
# ## Contour Plots
#
#
# The meshgrid command is used to make two grids where X contains the x coordinate for each point and Y contains the y coordinate for each point. A third grid (called V in this example) is filled with values that are functions of the x and y components.
# The lines below make a 400x400 grid.
from pylab import *
xlist = linspace(-2.0, 2.0, 5)
ylist = linspace(-2.0, 2.0, 5)
X, Y = meshgrid(xlist, ylist)
xlist = linspace(-2.0, 2.0, 400)
ylist = linspace(-2.0, 2.0, 400)
X, Y = meshgrid(xlist, ylist)
V= sqrt(X**2 + Y**2)
figure()
CP1 = contour(X, Y, V)
clabel(CP1, fontsize=10)
title('Contour Plot')
xlabel('x (cm)')
ylabel('y (cm)')
show()
figure()
CP2 = contourf(X, Y, V)
colorbar(CP2)
title('Contour Plot')
xlabel('x (cm)')
ylabel('y (cm)')
show()
levels = [0.0, 0.5, 1, 1.5, 2.0, 2.5, 3.0, 3.5, 4.0]
CP3 = contour(X, Y, V, levels, colors='k')
clabel(CP3, colors = 'k', fmt = '%2.1f', fontsize=14)
CP4 = contourf(X, Y, V, levels)
colorbar(CP4)
# ## Histograms
from pylab import *
t = array([4.94,5.98,5.00,6.06,5.94,5.17,5.12,5.06,
2.74,2.91,4.24,6.68,4.89,5.88,5.41,5.53,
3.73,5.80,4.26,5.50,5.73,5.29,7.40,3.55])
figure()
hist(t, facecolor='b', edgecolor='k')
show()
# +
figure()
hist(t, bins=30, edgecolor='k')
show()
# -
bins = linspace(0, 10, 11)
figure()
hist(t, bins, edgecolor='k')
show()
# +
from pylab import *
N = array([4,5,5,6,5,5,5,5,2,2,4,6,4,5,5,5,3,5,4,5,5,5,7,3])
bins = linspace(0, 10, 11)
x = array([2,3,4,5,6,7])
y = array([1,2,3,12,3,2])
figure()
hist(t, bins, edgecolor='b', fill=False,align='left')
scatter(x,y,c='g')
show()
# -
|
Playground/PlotsPython.ipynb
|