code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
from nltk.stem import PorterStemmer
from nltk.corpus import stopwords
paragraph = """I have three visions for India. In 3000 years of our history, people from all over
the world have come and invaded us, captured our lands, conquered our minds.
From Alexander onwards, the Greeks, the Turks, the Moguls, the Portuguese, the British,
the French, the Dutch, all of them came and looted us, took over what was ours.
Yet we have not done this to any other nation. We have not conquered anyone.
We have not grabbed their land, their culture,
their history and tried to enforce our way of life on them.
Why? Because we respect the freedom of others.That is why my
first vision is that of freedom. I believe that India got its first vision of
this in 1857, when we started the War of Independence. It is this freedom that
we must protect and nurture and build on. If we are not free, no one will respect us.
My second vision for India’s development. For fifty years we have been a developing nation.
It is time we see ourselves as a developed nation. We are among the top 5 nations of the world
in terms of GDP. We have a 10 percent growth rate in most areas. Our poverty levels are falling.
Our achievements are being globally recognised today. Yet we lack the self-confidence to
see ourselves as a developed nation, self-reliant and self-assured. Isn’t this incorrect?
I have a third vision. India must stand up to the world. Because I believe that unless India
stands up to the world, no one will respect us. Only strength respects strength. We must be
strong not only as a military power but also as an economic power. Both must go hand-in-hand.
My good fortune was to have worked with three great minds. Dr. <NAME> of the Dept. of
space, Professor <NAME>, who succeeded him and Dr. <NAME>, father of nuclear material.
I was lucky to have worked with all three of them closely and consider this the great opportunity of my life.
I see four milestones in my career"""
# +
sentenc= nltk.sent_tokenize(paragraph)
wordd= nltk.word_tokenize(paragraph)
# -
# NOW HERE I APPLY PORT STREAMER AND STOP WORD
sentences = nltk.sent_tokenize(paragraph)
stemmer = PorterStemmer()
# Stemming Here i perform streming with stop word
for i in range(len(sentences)):
words = nltk.word_tokenize(sentences[i])
words = [stemmer.stem(word) for word in words if word not in set(stopwords.words('english'))]
sentences[i] = ' '.join(words)
print(sentences)
#here i perform lemitization
# Stemming Here i perform lemitizaton with stop word
from nltk.stem import WordNetLemmatizer
# # Here i perform Lemitization
# +
import nltk
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
paragraph = """Thank you all so very much. Thank you to the Academy.
Thank you to all of you in this room. I have to congratulate
the other incredible nominees this year. The Revenant was
the product of the tireless efforts of an unbelievable cast
and crew. First off, to my brother in this endeavor, Mr. Tom
Hardy. Tom, your talent on screen can only be surpassed by
your friendship off screen … thank you for creating a t
ranscendent cinematic experience. Thank you to everybody at
Fox and New Regency … my entire team. I have to thank
everyone from the very onset of my career … To my parents;
none of this would be possible without you. And to my
friends, I love you dearly; you know who you are. And lastly,
I just want to say this: Making The Revenant was about
man's relationship to the natural world. A world that we
collectively felt in 2015 as the hottest year in recorded
history. Our production needed to move to the southern
tip of this planet just to be able to find snow. Climate
change is real, it is happening right now. It is the most
urgent threat facing our entire species, and we need to work
collectively together and stop procrastinating. We need to
support leaders around the world who do not speak for the
big polluters, but who speak for all of humanity, for the
indigenous people of the world, for the billions and
billions of underprivileged people out there who would be
most affected by this. For our children’s children, and
for those people out there whose voices have been drowned
out by the politics of greed. I thank you all for this
amazing award tonight. Let us not take this planet for
granted. I do not take tonight for granted. Thank you so very much."""
# -
sentences = nltk.sent_tokenize(paragraph)
lemmatizer = WordNetLemmatizer()
# Lemmatization
for i in range(len(sentences)):
words = nltk.word_tokenize(sentences[i])
words = [lemmatizer.lemmatize(word) for word in words if word not in set(stopwords.words('english'))]
sentences[i] = ' '.join(words)
print(sentences)
paragraph = """I have three visions for India. In 3000 years of our history, people from all over
the world have come and invaded us, captured our lands, conquered our minds.
From Alexander onwards, the Greeks, the Turks, the Moguls, the Portuguese, the British,
the French, the Dutch, all of them came and looted us, took over what was ours.
Yet we have not done this to any other nation. We have not conquered anyone.
We have not grabbed their land, their culture,
their history and tried to enforce our way of life on them.
Why? Because we respect the freedom of others.That is why my
first vision is that of freedom. I believe that India got its first vision of
this in 1857, when we started the War of Independence. It is this freedom that
we must protect and nurture and build on. If we are not free, no one will respect us.
My second vision for India’s development. For fifty years we have been a developing nation.
It is time we see ourselves as a developed nation. We are among the top 5 nations of the world
in terms of GDP. We have a 10 percent growth rate in most areas. Our poverty levels are falling.
Our achievements are being globally recognised today. Yet we lack the self-confidence to
see ourselves as a developed nation, self-reliant and self-assured. Isn’t this incorrect?
I have a third vision. India must stand up to the world. Because I believe that unless India
stands up to the world, no one will respect us. Only strength respects strength. We must be
strong not only as a military power but also as an economic power. Both must go hand-in-hand.
My good fortune was to have worked with three great minds. Dr. <NAME> of the Dept. of
space, Professor <NAME>, who succeeded him and Dr. <NAME>, father of nuclear material.
I was lucky to have worked with all three of them closely and consider this the great opportunity of my life.
I see four milestones in my career"""
# +
sentences = nltk.sent_tokenize(paragraph)
lemmatizer = WordNetLemmatizer()
# Lemmatization
for i in range(len(sentences)):
words = nltk.word_tokenize(sentences[i])
words = [lemmatizer.lemmatize(word) for word in words if word not in set(stopwords.words('english'))]
sentences[i] = ' '.join(words)
print(sentences)
# -
|
Streaming & Lemitization/NLP with [Streaming ,Lemitization ,Stop word].ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# check installed packages
pip list
# +
from pyspark.sql import SparkSession
# Spark session & context
spark = SparkSession.builder.master('local').getOrCreate()
sc = spark.sparkContext
# Sum of the first 100 whole numbers
rdd = sc.parallelize(range(100 + 1))
rdd.sum()
# 5050
# -
|
src/playbook_101.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## 1. Basic idea
# The algorithm is supposed to generate a list of rules based on PRISM algorithm discussed during the class, and should be able to work with both numerical and categorical data with no empty cells. This project is implemented based on Professor <NAME>'s implementation plan and some code is borrowed from her. I also got direct help from her with regard to implementing the learn_one_rule algorithm.
#
# ## 2. Implementation notes
#
# First, we need a datastructure to store each rule. The data structure is borrowed from Professor <NAME>
#
#
# Each Rule consists of antecedent (Left Hand Side) and consequent (Right Hand Side). The LHS includes multiple conditions joined with AND, and RHS is a class label. The Rule also needs to store its accuracy and coverage.
class Rule:
def __init__(self, class_label):
self.conditions = [] # list of conditions
self.class_label = class_label # rule class
self.accuracy = 0
self.coverage = 0
def addCondition(self, condition):
self.conditions.append(condition)
def setParams(self, accuracy, coverage):
self.accuracy = accuracy
self.coverage = coverage
# Human-readable printing of this Rule
def __repr__(self):
return "If {} then {}. Coverage:{}, accuracy: {}".format(self.conditions, self.class_label,
self.coverage, self.accuracy)
# The list of conditions contains several objects of class _Condition_.
#
# Each condition includes the _attribute name_ and the _value_.
#
# If the _value_ is numeric, then the condition also includes an additional field `true_false` which means the following:
# - *if true_false == True then values are >= value*
# - *if true_false == False then values are < value*
# - If *true_false is None*, then this condition is simply of form *categorical attribute = value*.
class Condition:
def __init__(self, attribute, value, true_false = None):
self.attribute = attribute
self.value = value
self.true_false = true_false
def __repr__(self):
if self.true_false is None:
return "{}={}".format(self.attribute, self.value)
else:
return "{}>={}:{}".format(self.attribute, self.value, self.true_false)
# Next comes the `learn_one_rule` algorithm. The required parameters are the names of the columns, the current subset of data, and the class label (RHS of the rule we are trying to learn). The optional parameters are thresholds `min_coverage` and `min_accuracy`. For normal datasets we can set the coverage value to 30, for example, to prevent creating unreliable rules with insignificant support from data.
#
# The algorithm returns a new rule and maybe a subset of data covered by the Rule or the remaining data (not covered by the rule).
#
# Please note that the algorithm I implemented is prioritizing accuracy, which means that for a 1.0 accuracy rule with 30 coverage, and a 0.9 accuracy rule with 30000 coverage, the first rule would be selected.
#
# The general idea of the implementation is that:
# 1. Create a subset of the original data set that only contains the class label we are interested in
# 2. Find all possible conditions, and for each condition, check if it is int(numerical) or not int(categorical data).
# 3. If it is numerical data, get all possible values and sort the unique values from small to large. For all possible values, test the conditions as a. greater or equal to the value, or b, smaller than the value. Find the greater accuracy. The accuracy is compared with the current best accuracy and coverage.
# 4. If it is categorical, iterate through all possible values to see what is the accuracy and coverage for each category.
# 5. Check the current best result:
# - if I reach coverage 1, that means I can no longer improve and since every new condition would at least keep/reduce my covered subset, I don't want to continue. Thus, it is the best option. I add the current condition to my rule
# - else if I tested all possible conditions, I want to see if I reached a final accpetable result or not. If so, I return the rule with all conditions in it.
# - else if I reached a coverage smaller than min coverage, that means I can no longer subdivide, and have to stop. I check if it is good enough and if so I return the rule without current condition, as the current condition's coverage is too small.
# - else I have to continue, and I remove current attribute from the possible attribute list, and add the condition to the rule, and continue with the iterative loop
import pandas as pd
import numpy as np
def learn_one_rule(column, data, class_label, min_coverage=0, min_accuracy=0.6):
if len(data)==0:
return None
covered_subset = data.copy()
data2 = data.copy()
current_rule = Rule(class_label)
done = False
# filter out data with right labels
classheader = data2.columns[-1]
current_data = data2[data2[classheader] == class_label]
# make sure I am not operating the wrong set of data
columns = column.copy()
columns.pop()
#get the best possible option
while not done:
#reset
current_accuracy = 0
current_coverage = 0
current_condition = Condition(None,None)
index = 0
#go through all possible attributes
for i in range(len(columns)):
current_attribute = columns[i]
#check if it is numerical, get unique values, for each value, there is two rules: greater or equal to, or less than
possible_values = current_data[current_attribute].unique().tolist()
if isinstance(possible_values[0], int) or isinstance(possible_values[0],float):
possible_values.sort()
#test every possible value for the current attribute
for value in possible_values:
#if dealing with numerical values
if isinstance(value, int) or isinstance(value,float):
#get two sets of accuracy and coverage
mycolumn1 = current_data[current_attribute]
correct1 = mycolumn1[mycolumn1 >= value].count()
mycolumn2 = data2[current_attribute]
coverage1 = mycolumn2[mycolumn2 >= value].count()
# this is to avoid 0/0 scenario
if coverage1 == 0:
accuracy1 = 0
else:
accuracy1 = correct1/coverage1
mycolumn3 = current_data[current_attribute]
correct = mycolumn3[mycolumn3 <value].count()
column4 = data2[current_attribute]
coverage = column4[column4 < value].count()
if coverage == 0:
accuracy = 0
else:
accuracy = correct/coverage
#just here to avoid syntax problems, also set flag for later use
GreaterThan = False
# get the best among the two
if accuracy1 > accuracy:
GreaterThan = True
accuracy = accuracy1
coverage = coverage1
elif accuracy1 == accuracy:
if coverage1 > coverage:
GreaterThan = True
coverage = coverage1
Int = True
# if the result is good enough to be recorded, record as condition, and update the
# accuracy/coverage/covered set
if coverage >= min_coverage:
if accuracy > current_accuracy:
index = i
current_coverage = coverage
current_accuracy = accuracy
current_condition= Condition(current_attribute, value, GreaterThan)
if GreaterThan:
covered_subset = data2[data2[current_attribute] >= value]
else:
covered_subset = data2[data2[current_attribute] < value]
# if accuracy is the same, compare coverage
elif accuracy == current_accuracy and coverage > current_coverage:
index = i
current_coverage = coverage
current_accuracy = accuracy
current_condition= Condition(current_attribute, value,GreaterThan)
if GreaterThan:
covered_subset = data2[data2[current_attribute] >= value]
else:
covered_subset = data2[data2[current_attribute] < value]
else:
#compute accuracy
correct = current_data[current_attribute].value_counts()[value]
coverage = data2[current_attribute].value_counts()[value]
accuracy = correct/coverage
#choose the best option based on accuracy
if coverage >= min_coverage:
if accuracy > current_accuracy:
current_coverage = coverage
current_accuracy = accuracy
current_condition= Condition(current_attribute, value)
covered_subset = data2[data2[current_attribute] == value]
index = i
# if accuracy is the same, compare coverage
elif accuracy == current_accuracy and coverage > current_coverage:
current_coverage = coverage
current_accuracy = accuracy
current_condition= Condition(current_attribute, value)
covered_subset = data2[data2[current_attribute] == value]
index = i
#if reached to the end accuracy = 1.0, add to rule and terminate
if current_accuracy == 1.0 and current_coverage > min_coverage:
done = True
current_rule.addCondition(current_condition)
current_rule.accuracy = current_accuracy
current_rule.coverage = current_coverage
#if reached to the end that is acceptable, add to rule and terminate
elif len(columns) == 0:
done = True
if current_rule.accuracy <= min_accuracy or current_rule.coverage < min_coverage:
return (None, None)
#if no rule possible, return none, else, we reached an end and is done
elif current_coverage < min_coverage:
if current_rule.accuracy <= min_accuracy or current_rule.coverage < min_coverage:
return (None, None)
done = True
#default: not done yet, continue
else:
#update possible attributes
columns.pop(index)
current_rule.addCondition(current_condition)
data2 = covered_subset
current_data = data2[data2[classheader] == class_label]
current_rule.accuracy = current_accuracy
current_rule.coverage = current_coverage
#reset
return (current_rule, covered_subset)
# Finally, the main algorithm `learn_rules` takes as parameters list of columns, with the last column representing the class label, and the original data in form of pandas dataframe. Two optional threshold parameters `min_coverage` and `min_accuracy` set up the conditions of rule's validity for a given dataset.
#
# In order to cover the PRISM algorithm, I also added a small part that would run the learn_one_rule algorithm repeatedly to generate the best possible result. Unfortunately, that means the time is at least doubled and I am unable to improve on that.
# +
import pandas as pd
import numpy as np
def learn_rules (columns, data, classes=None,
min_coverage = 30, min_accuracy = 0.6):
# List of final rules
rules = []
# If list of classes of interest is not provided - it is extracted from the last column of data
if classes is not None:
class_labels = classes
else:
class_labels = data[columns[-1]].unique().tolist()
current_data = data.copy()
# This follows the logic of the original PRISM algorithm
# It processes each class in turn.
for class_label in class_labels:
done = False
while len(current_data) > min_coverage and not done:
# Learn one rule
rule, subset = learn_one_rule(columns, current_data, class_label, min_coverage, min_accuracy)
# If the best rule does not pass the coverage threshold - we are done with this class
if rule is None:
break
copylabel = class_labels.copy()
copylabel.remove(class_label)
for mylabel in copylabel:
myrule, mysubset = learn_one_rule(columns, current_data, mylabel, min_coverage, min_accuracy)
if myrule is not None and (myrule.accuracy > rule.accuracy or myrule.coverage > rule.coverage):
rule, subset = myrule, mysubset
# If we get the rule with accuracy and coverage above threshold
if rule.accuracy >= min_accuracy:
rules.append(rule)
#try get the best result.
for id in subset.index:
current_data.drop(index = id, inplace = True)
current_data = current_data.dropna()
else:
done = True
return rules
# -
# ## 3. Correctness test (Borrowed from Professor <NAME>)
# Test your algorithm on the original dataset from the PRISM paper.
#
# The dataset was downloaded from [here](https://archive.ics.uci.edu/ml/datasets/Lenses). The CSV version is included in this repository.
#
# **Attribute Information**:
#
# 3 Classes:
# - __1__ : the patient should be fitted with __hard__ contact lenses,
# - __2__ : the patient should be fitted with __soft__ contact lenses,
# - __3__ : the patient should __not__ be fitted with contact lenses.
#
#
# Attributes:
# 1. age of the patient: (1) young, (2) pre-presbyopic, (3) presbyopic
# 2. spectacle prescription: (1) myope, (2) hypermetrope
# 3. astigmatic: (1) no, (2) yes
# 4. tear production rate: (1) reduced, (2) normal
#
# Presbyopia is physiological insufficiency of accommodation associated with the aging of the eye that results in progressively worsening ability to focus clearly on close objects. So "age=presbiopic" means old.
#
# Hypermetropia: far-sightedness, also known as long-sightedness - cannot see close.
# Myopia: nearsightedness - cannot see at distance.
# +
import pandas as pd
import numpy as np
data_file = "contact_lenses.csv"
data = pd.read_csv(data_file, index_col=['id'])
data.columns
# -
# We can replace numbers with actual values - for clarity.
# +
# classes
conditions = [ data['lenses type'].eq(1), data['lenses type'].eq(2), data['lenses type'].eq(3)]
choices = ["hard","soft","none"]
data['lenses type'] = np.select(conditions, choices)
# age groups
conditions = [ data['age'].eq(1), data['age'].eq(2), data['age'].eq(3)]
choices = ["young","medium","old"]
data['age'] = np.select(conditions, choices)
# spectacles
conditions = [ data['spectacles'].eq(1), data['spectacles'].eq(2)]
choices = ["nearsighted","farsighted"]
data['spectacles'] = np.select(conditions, choices)
# astigmatism
conditions = [ data['astigmatism'].eq(1), data['astigmatism'].eq(2)]
choices = ["no","yes"]
data['astigmatism'] = np.select(conditions, choices)
# tear production rate
conditions = [ data['tear production rate'].eq(1), data['tear production rate'].eq(2)]
choices = ["reduced","normal"]
data['tear production rate'] = np.select(conditions, choices)
print(data)
# -
# The test (do not run it before you finished the implementation of the rule learning algorithm):
# +
column_list = data.columns.to_numpy().tolist()
rules = learn_rules (column_list, data, None, 1, 0.95)
for rule in rules[:20]:
print(rule)
# -
# After manual evaluationi the method seems to work with a better result from professor's method, meaning that it is more accuracy in terms of the PRISM algorithm.
# <NAME>'s results are given below for comparison:
#
# If [tear production rate=reduced] then none. Coverage:12, accuracy: 1.0
#
# If [astigmatism=no, spectacles=farsighted] then soft. Coverage:3, accuracy: 1.0
#
# If [astigmatism=no, age=young] then soft. Coverage:1, accuracy: 1.0
#
# If [astigmatism=no, age=medium] then soft. Coverage:1, accuracy: 1.0
#
# If [age=young] then hard. Coverage:2, accuracy: 1.0
#
# If [spectacles=nearsighted, astigmatism=yes] then hard. Coverage:2, accuracy: 1.0
# Copyright © 2020 Marina Barsky. All rights reserved.
|
Xiaoyi_algorithm_notes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Differential Privacy Jupyter Lab Lesson 1
# Welcome to the Differential Privacy Jupyter Lab Lesson #1.
#
# In this lab, we'll see how the Laplace and the Geometric mechanism can be used in private data analysis.
# +
import numpy
import numpy.random
# TODO: The differential privacy laplace mechanism uses the laplace distribtion
# The original paper used the laplace mechanism because it made the math easier.
# Graph the gaussian & the laplace distribution.
# TODO: Redo this so that we just have a single run
# Then build it to multiple runs.
def dp_laplace(*,private_x, sensitivity, epsilon):
"""This versin of the """
return numpy.random.laplace(private_x, 1.0/epsilon )
# -
"""
Let's assume a hypothetical survey in which there are 100 people who respond.
We want to protect with differential privacy the number of respondents.
The sensitivity is 1 because a person being added or removed will change that
number by 1. Here is such a computation, with an epsilon of 2.0:
"""
dp_laplace(private_x=100, sensitivity=1, epsilon=2.0)
# +
"""
Now we will run this experiment 10 times, to show the range of the protection values:
"""
runs=10
for i in range(runs):
display(dp_laplace(private_x=100, sensitivity=1, epsilon=2.0))
# -
"""
Because our dp_lapace mechanism is built with numpy.random.laplace,
we can repeat that experiment with a single operation.
REMEMBER -- this is just for demonstration purposes. If we were *actually*
using differential privacy, we would just run it once.
We can get integer counts by rounding afterwards, or by using a different
mechanism (the geometeric mechanism gives integers.)
"""
private_data = [100,100,100,100,100,100,100,100,100,100]
display( dp_laplace(private_x = private_data, sensitivity=1.0, epsilon=2.0) )
"""
Here we introduce a nifty tool for displaying tables that's part of the ctools package.
We will re-run the experiment
TODO: redo this with pandas
"""
from ctools.tydoc import jupyter_display_table
private_data = [100] * 10
public_data = dp_laplace(private_x = private_data, sensitivity=1.0, epsilon=2.0)
jupyter_display_table({'epsilon 2.0':public_data}, float_format='{:.4f}')
# +
"""Averaging the 10 draws above with an epsilon of 2.0 is the same a doing a single
draw with an epsilon of 20. Let's compare those two possibilities; they look
pretty simlar (and pretty accurate)"""
import statistics
display("Average of the {} epsilon 2.0 runs: {}".
format(len(public_data), statistics.mean(public_data)))
display("Private query with a single epsilon 20.0 run: {}".format(dp_laplace(private_x = 100.0, sensitivity=1.0, epsilon=20.0)))
# +
"""Here we observe the impact of epsilon by comparing the noise added to a count of 100
for epsilon values of 0.01, 0.1, 1.0, and 2.0."""
def run_experiment(epsilon):
private_data = [100] * 10
return {f"epsilon {epsilon}":
dp_laplace(private_x = private_data, sensitivity=1.0, epsilon=epsilon)}
trials = {"Trial":[f"trial #{i}" for i in range(1,11)]}
jupyter_display_table( {**trials,
**run_experiment(0.01),
**run_experiment(0.1),
**run_experiment(1.0),
**run_experiment(2.0)} )
# +
"""
Instead of protecting 10 independent trial, the approach that we take above
could use used to protect 10 independent measurements of a single population.
Let's protect the population numbers for the District of Columbia from the 2010 Census.
Here we round the counts. That's post-processing, so it's totally okay to do.
We'll be using an epsilon of 0.01 so that we can see some differences.
"""
categories = ["Under 5 years"] + [f"{age} to {age+4} years" for age in range(5,90,5)]+ ["90 years and over"]
true_counts = [32613, 26147, 25041, 39919, 64110, 69649, 55096, 42925, 37734,
38539, 37164, 34274, 29703, 21488, 15481, 11820, 9705, 6496, 3819]
protected_counts = [int(x) for x in
dp_laplace(private_x = true_counts, sensitivity=1.0, epsilon=0.01 )]
jupyter_display_table( {"Age":categories,
"True Counts":true_counts,
"Protected Counts":protected_counts} )
# +
"""By comparing the differences between the counts above and the true counts,
we can see the overall impact of differential privacy for epsilon=0.01.
NOTE --- comparing the protected counts to the true counts is something you cannot
typically do in differential privacy, because that's making a comparision across the noise barrier.
But it's useful for learning DP and understanding how mechanisms work.
"""
diff_counts = [p-t for (p,t) in zip(protected_counts,true_counts)]
jupyter_display_table( {"Age":categories + ['total'],
"True Counts":true_counts + [sum(true_counts)],
"Protected Counts":protected_counts + [sum(protected_counts)],
"Difference":diff_counts + [sum(diff_counts)]
} )
# -
|
dp-lab/DP Lesson 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="VIVfmy_iqvmH" outputId="59e6914b-7eed-4dee-ae32-a66d4d9044ea"
import pandas as pd
import numpy as np
battingdf = pd.read_csv('https://raw.githubusercontent.com/frankData612/data_612/master/baseballdatabank-master/core/Batting.csv')
battingdf.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="F1XxGPEFsZeG" outputId="10fe48fb-2001-4496-e9a3-43925cf5467f"
battingdf.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="Uq44OsRpsjJn" outputId="a5b1605a-dbc8-4fd0-9df8-373ce96f14f9"
battingdf.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="iK5BbQxXtyX4" outputId="81612887-a5dd-498b-9a25-4b57ca9b87e8"
#convert leagueID from string to categorical
battingdf['lgID'] = battingdf['lgID'].astype('category')
print(battingdf['lgID'].dtypes)
# + colab={"base_uri": "https://localhost:8080/"} id="h5c5U967vDLi" outputId="0bdae1fb-d54b-4fcb-9625-701e463c8a2b"
#convert BB from float to string
battingdf['BB'] = battingdf['BB'].astype(str)
print(battingdf['BB'].dtypes)
|
Assignment6/Module7_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from birdy import WPSClient
from birdy.exceptions import ProcessIsNotComplete, ProcessFailed, ProcessCanceled
from IPython.display import Image
cli = WPSClient('http://compute.mips.copernicus-climate.eu/wps') # progress=True
# -
result = cli.cordex_subsetter(year=2000, model='MOHC-HadRM3P', variable='tas', country='France')
try:
output = result.get()
except ProcessIsNotComplete:
print("Please wait ...")
except ProcessFailed:
print("Sorry, somthing went somethrong ...")
except ProcessCanceled:
# TODO: canceled exception is not raised yet
print("Job was canceled.")
else:
print(output)
Image(output.preview)
|
notebooks/examples/c4cds.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 쉘모드 변경
# - Code 쉘 모드 : 파이썬 코드 실행
# - Markdown 쉘 모드 : 문서만들기
#
# ---
#
# * Code 쉘 : esc > y
# * Markdown 쉘 : esc > m
# ### 서울 지하철 평균 이용객 수 분석
#
# #### 가장 이용객수가 많은 지하철은?
# + 강남
# + 홍대
# - 성수
# - 합정
# * 별표 하나는 번호 없는 목록
#
# *별표로 감싸면 기울이기(별표와 띄어쓰기 없이)*
#
# **별표 2개로 감싸면 굵게**
#
# ***별표 3개로 감싸면 굵게이면서 기울이기***
#
# 가로선 --- 이면
#
# ---
#
# 인용문으로 >
# > (실행코드 쓸 때)
# <div align='center'><font color='red'>hello world</font></div>
# <img src='./dataset/puppy.jpg'>
# [링크걸기](https://datascienceschool.net/view-notebook/39569f0132044097a15943bd8f440ca5/)
#
# 이미지링크
# [](https://playdata.io/)
# ## <font color="green">코드결과</font>
# +
import pandas
p1 = pandas.read_csv('./dataset/subway_data1.csv', encoding='utf-8')
# -
p1
# + *맨 좌측의 숫자는 index이다* <br/>
# + **인덱스는 자동으로 설정되지만 사용자가 설정도 가능합니다**<br/>
# + ***value는 데이터를 의미한다***
#
# <table border="1">
# <tr><td>인덱스</td><td>값</td></tr>
# <tr><td></td><td>데이터</td></tr>
# </table>
# <img src="dataset/map_subway.jpg">
#
# 이미지를 클릭하면 지하철홈페이지 링크걸기
# [](http://www.seoulmetro.co.kr/kr/index.do?device=PC)
#
#
|
jupyter/dAnalysis/a_basic_class/Ex01_markdown.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="5rmpybwysXGV"
# ##### Copyright 2018 The TensorFlow Authors
# + cellView="form" colab_type="code" id="m8y3rGtQsYP2" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="hrXv0rU9sIma"
# # 定制化训练:基础
# + [markdown] colab_type="text" id="7S0BwJ_8sLu7"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/beta/tutorials/eager/custom_training"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 Tensorflow.google.cn 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/zh-cn/beta/tutorials/eager/custom_training.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/zh-cn/beta/tutorials/eager/custom_training.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 Github 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/zh-cn/beta/tutorials/eager/custom_training.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="k2o3TTG4TFpt"
# 在之前的教程中,我们讨论了自动差分,一个基本的机器学习模块。在这个教程中,我们将使用在之前介绍的 Tensorflow 基础语句实现简单的机器学习模型。
#
# Tensorflow 也提供了高级神经网络 API(`tf.keras`),可以精简范例代码。我们强烈建议在神经网络方面的工作使用高级的 API。在这篇教程中,我们使用基本规则来训练神经网络,为以后打下牢固基础。
# + [markdown] colab_type="text" id="3LXMVuV0VhDr"
# ## 创建
# + colab_type="code" id="NiolgWMPgpwI" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# !pip install tensorflow==2.0.0-beta1
import tensorflow as tf
# + [markdown] colab_type="text" id="eMAWbDJFVmMk"
# ## 变量
#
# Tensorflow 中的 tensor 是不可变无状态对象。机器学习模型需要可改变状态,比如模型训练和模型预测的代码是相同的,但变量值随着时间而不同(希望尽量小的 loss)。为了应对随着计算而改变的状态,可以利用 Python 的状态可变性。
# + colab_type="code" id="VkJwtLS_Jbn8" colab={}
# 使用 python 状态
x = tf.zeros([10, 10])
# 等价于 x = x + 2, 不改变原本 x 的值
x += 2
print(x)
# + [markdown] colab_type="text" id="wfneTXy7JcUz"
# TensorFlow,拥有内建可变状态操作,比使用底层 Python 状态表示更常见的。比如,表示模型的权重,使用 TensorFlow 变量更方便高效。
#
# 变量是一个对象,这个对象存储着数值,当在 TensorFlow 计算中使用时,会隐式地读取这个存储的数值。有一些操作(`tf.assign_sub`, `tf.scatter_update` 等)会复制 TensorFlow 变量存储的数值。
# + colab_type="code" id="itxmrMil6DQi" colab={}
v = tf.Variable(1.0)
assert v.numpy() == 1.0
# 重新赋值
v.assign(3.0)
assert v.numpy() == 3.0
# 在 TensorFlow 操作中使用 `v`,比如 tf.square() 和重新赋值
v.assign(tf.square(v))
assert v.numpy() == 9.0
# + [markdown] colab_type="text" id="-paSaeq1JzwC"
# 当计算梯度时,会自动跟踪使用变量的计算过程。用变量来表示向量时,TensorFlow 会默认使用稀疏更新,这样可以带来计算和存储高效性。
#
# 使用变量也是一种更快的提醒方式,就是代码的这部分是状态可变的。
# + [markdown] colab_type="text" id="BMiFcDzE7Qu3"
# ## 示例:尝试一个线性模型
#
# 让我们来使用目前为止学到的概念---`Tensor`,`Variable`,和 `GradientTape`---来创建和训练一个简单的模型。一般需要下面这些步骤:
#
# 1. 定义模型
# 2. 定义损失函数
# 3. 获取训练数据
# 4. 通过训练数据运行模型,使用 "optimizer" 来调整变量以满足数据
#
# 在这个教程中,我们使用一个简单线性模型作为示例:`f(x) = x * W + b`,有2个变量- `W` 和 `b`。另外,我们会生成数据让训练好的模型满足 `W = 3.0` 和 `b = 2.0`。
# + [markdown] colab_type="text" id="gFzH64Jn9PIm"
# ### 定义模型
#
# 定义一个简单的类封装变量和计算
# + colab_type="code" id="_WRu7Pze7wk8" colab={}
class Model(object):
def __init__(self):
# 初始化变量值为(5.0, 0.0)
# 实际上,这些变量应该初始化为随机值
self.W = tf.Variable(5.0)
self.b = tf.Variable(0.0)
def __call__(self, x):
return self.W * x + self.b
model = Model()
assert model(3.0).numpy() == 15.0
# + [markdown] colab_type="text" id="xa6j_yXa-j79"
# ### 定义损失函数
#
# 损失函数用来衡量在给定输入的情况下,模型的预测输出与实际输出的偏差。我们这里使用标准 L2 损失函数。
# + colab_type="code" id="Y0ysUFGY924U" colab={}
def loss(predicted_y, desired_y):
return tf.reduce_mean(tf.square(predicted_y - desired_y))
# + [markdown] colab_type="text" id="qutT_fkl_CBc"
# ### 获取训练数据
#
# 我们来生成带噪声的训练数据。
# + colab_type="code" id="gxPTb-kt_N5m" colab={}
TRUE_W = 3.0
TRUE_b = 2.0
NUM_EXAMPLES = 1000
inputs = tf.random.normal(shape=[NUM_EXAMPLES])
noise = tf.random.normal(shape=[NUM_EXAMPLES])
outputs = inputs * TRUE_W + TRUE_b + noise
# + [markdown] colab_type="text" id="-50nq-wPBsAW"
# 在训练模型之前,我们来看看当前的模型表现。我们绘制模型的预测结果和训练数据,预测结果用红色表示,训练数据用蓝色表示。
# + colab_type="code" id="_eb83LtrB4nt" colab={}
import matplotlib.pyplot as plt
plt.scatter(inputs, outputs, c='b')
plt.scatter(inputs, model(inputs), c='r')
plt.show()
print('Current loss: '),
print(loss(model(inputs), outputs).numpy())
# + [markdown] colab_type="text" id="sSDP-yeq_4jE"
# ### 定义训练循环
#
# 我们已经定义了网络模型,并且获得了训练数据。现在对模型进行训练,采用[梯度下降](https://en.wikipedia.org/wiki/Gradient_descent)的方式,通过训练数据更新模型的变量(`W` 和 `b`)使得损失量变小。梯度下降中有很多参数,通过 `tf.train.Optimizer` 实现。我们强烈建议使用这些实现方式,但基于通过基本规则创建模型的精神,在这个特别示例中,我们自己实现基本的数学运算。
# + colab_type="code" id="MBIACgdnA55X" colab={}
def train(model, inputs, outputs, learning_rate):
with tf.GradientTape() as t:
current_loss = loss(model(inputs), outputs)
dW, db = t.gradient(current_loss, [model.W, model.b])
model.W.assign_sub(learning_rate * dW)
model.b.assign_sub(learning_rate * db)
# + [markdown] colab_type="text" id="RwWPaJryD2aN"
# 最后,我们对训练数据重复地训练,观察 `W` 和 `b` 是怎么变化的。
# + colab_type="code" id="XdfkR223D9dW" colab={}
model = Model()
# 收集 W 和 b 的历史数值,用于显示
Ws, bs = [], []
epochs = range(10)
for epoch in epochs:
Ws.append(model.W.numpy())
bs.append(model.b.numpy())
current_loss = loss(model(inputs), outputs)
train(model, inputs, outputs, learning_rate=0.1)
print('Epoch %2d: W=%1.2f b=%1.2f, loss=%2.5f' %
(epoch, Ws[-1], bs[-1], current_loss))
# 显示所有
plt.plot(epochs, Ws, 'r',
epochs, bs, 'b')
plt.plot([TRUE_W] * len(epochs), 'r--',
[TRUE_b] * len(epochs), 'b--')
plt.legend(['W', 'b', 'true W', 'true_b'])
plt.show()
# + [markdown] colab_type="text" id="vPnIVuaSJwWz"
# ## 下一步
#
# 在这个教程中,我们讨论了 `Variable`,而且创建和训练了一个简单的线性模型,使用了在此之前所学习的 TensorFlow 知识点。
#
# 理论上,掌握了 TensorFlow 这些知识点即可用于机器学习研究。实际上,采用高级的 API 比如 `tf.keras` 是更方便的,特别是神经网络,因为它提供了更高级别的内建模块(命名为 "layers"),可以保存和恢复状态,还有配套的损失函数和优化策略等。
#
|
site/zh-cn/beta/tutorials/eager/custom_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8ZelAXba3D05"
# 
#
# [](https://colab.research.google.com/github/JohnSnowLabs/nlu/blob/master/examples/release_notebooks/NLU_3_0_2_release_notebook.ipynb)
#
#
# # Entity Resolution
# **Named entities** are sub-strings in a text that can be classified into catogires. For example, in the String
# `"Tesla is a great stock to invest in "` , the sub-string `"Tesla"` is a named entity, it can be classified with the label `company` by an ML algorithm.
# **Named entities** can easily be extracted by the various pre-trained Deep Learning based NER algorithms provided by NLU.
#
#
#
# After extracting **named entities** an **entity resolution algorithm** can be applied to the extracted named entities. The resolution algorithm classifies each extracted entitiy into a class, which reduces dimensionality of the data and has many useful applications.
# For example :
# - "**Tesla** is a great stock to invest in "
# - "**TSLA** is a great stock to invest in "
# - "**Tesla, Inc** is a great company to invest in"
#
# The sub-strings `Tesla` , `TSLA` and `Tesla, Inc` are all named entities, that are classified with the labeld `company` by the NER algorithm. It tells us, all these 3 sub-strings are of type `company`, but we cannot yet infer that these 3 strings are actually referring to literally the same company.
#
# This exact problem is solved by the resolver algorithms, it would resolve all these 3 entities to a common name, like a company ID. This maps every reference of Tesla, regardless of how the string is represented, to the same ID.
#
# This example can analogusly be expanded to healthcare any any other text problems. In medical documents, the same disease can be referenced in many different ways.
#
# With NLU Healthcare you can leverage state of the art pre-trained NER models to extract **Medical Named Entities** (Diseases, Treatments, Posology, etc..) and **resolve these** to common **healthcare disease codes**.
#
#
# These algorithms are based provided by **Spark NLP for Healthcare's** [SentenceEntitiyResolver](https://nlp.johnsnowlabs.com/docs/en/licensed_annotators#sentenceentityresolver) and [ChunkEntityResolvers](https://nlp.johnsnowlabs.com/docs/en/licensed_annotators#chunkentityresolver)
#
# ## New Entity Resolovers In NLU 3.0.2
#
# | NLU REF | NLP REF |
# |-----------------------------------|-----------------------------------------|
# |[`en.resolve.umls` ](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_major_concepts_en.html)| [`sbiobertresolve_umls_major_concepts`](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_major_concepts_en.html) |
# |[`en.resolve.umls.findings` ](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_findings_en.html)| [`sbiobertresolve_umls_findings`](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_findings_en.html) |
# |[`en.resolve.loinc` ](https://nlp.johnsnowlabs.com/2021/04/29/sbiobertresolve_loinc_en.html)| [`sbiobertresolve_loinc`](https://nlp.johnsnowlabs.com/2021/04/29/sbiobertresolve_loinc_en.html) |
# |[`en.resolve.loinc.biobert` ](https://nlp.johnsnowlabs.com/2021/04/29/sbiobertresolve_loinc_en.html)| [`sbiobertresolve_loinc`](https://nlp.johnsnowlabs.com/2021/04/29/sbiobertresolve_loinc_en.html) |
# |[`en.resolve.loinc.bluebert` ](https://nlp.johnsnowlabs.com/2021/04/29/sbluebertresolve_loinc_en.html)| [`sbluebertresolve_loinc`](https://nlp.johnsnowlabs.com/2021/04/29/sbluebertresolve_loinc_en.html) |
# |[`en.resolve.HPO` ](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_HPO_en.html)| [`sbiobertresolve_HPO`](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_HPO_en.html) |
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="mOC9k0u0Hp2t" outputId="76424895-1343-43d8-cca1-794407cbecc0"
# Upload add your spark_nlp_fo"r_healthcare.json
# !wget http://setup.johnsnowlabs.com/nlu/colab.sh -O - | bash
import nlu
nlu.auth(SPARK_NLP_LICENSE,AWS_ACCESS_KEY_ID,AWS_SECRET_ACCESS_KEY,JSL_SECRET)
# + [markdown] id="2YmPDZxdJ4zO"
# #### [Sentence Entity Resolver for UMLS CUI Codes](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_major_concepts_en.html)
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="pP80GhfAJ2Np" outputId="23d4865a-69ac-41a7-a2d4-66e99a88dfc6"
nlu.load('med_ner.jsl.wip.clinical en.resolve.umls').predict("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting."""
,output_level = 'sentence')
# + id="3uhnHQc3vTa2" colab={"base_uri": "https://localhost:8080/", "height": 485} outputId="de3ebf52-fa8a-4ef8-d03c-251b3547d2b1"
nlu.load('med_ner.jsl.wip.clinical en.resolve.umls').viz("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting.""")
# + [markdown] id="fkoNgzSJNrnf"
# #### [Sentence Entity Resolver for UMLS CUI Codes](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_umls_findings_en.html)
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="4kUf26eFLbdK" outputId="967c52b4-7c78-4759-b26b-bd8c8b328b9b"
nlu.load('med_ner.jsl.wip.clinical en.resolve.umls.findings').predict("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting."""
,output_level = 'sentence')
# + colab={"base_uri": "https://localhost:8080/", "height": 485} id="t5Jx4IupRpI4" outputId="92470bf6-2132-4d5a-e50e-3ddfdb63a16b"
nlu.load('med_ner.jsl.wip.clinical en.resolve.umls.findings').viz("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting."""
)
# + [markdown] id="mPZjWu900n1e"
# #### [Loinc Sentence Entity Resolver](https://nlp.johnsnowlabs.com/2021/04/29/sbiobertresolve_loinc_en.html)
# + colab={"base_uri": "https://localhost:8080/", "height": 457} id="JJXsEYGoz7p-" outputId="78881a56-09dd-413b-dd0c-3c5aa7be59ea"
nlu.load('med_ner.jsl.wip.clinical en.resolve.loinc').predict("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting."""
,output_level = 'sentence')
# + id="p9s1c1NA05MI" colab={"base_uri": "https://localhost:8080/", "height": 485} outputId="75258a04-fb3b-4662-e97e-5bbee4e0aea6"
nlu.load('med_ner.jsl.wip.clinical en.resolve.loinc.biobert').viz("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting.""")
# + [markdown] id="-MKt6PpP3xvC"
# #### [Loinc Sentence Entity Resolver](https://nlp.johnsnowlabs.com/2021/04/29/sbluebertresolve_loinc_en.html)
# + id="SGoizkat1Dmv" colab={"base_uri": "https://localhost:8080/", "height": 457} outputId="e4fb067d-4d04-4581-9dd2-30e85dfc9d24"
nlu.load('med_ner.jsl.wip.clinical en.resolve.loinc.bluebert').predict("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting."""
,output_level = 'sentence')
# + id="UUMlOhdW37Gv" colab={"base_uri": "https://localhost:8080/", "height": 485} outputId="8ad66fee-b301-4e56-c170-dfd68034dddd"
nlu.load('med_ner.jsl.wip.clinical en.resolve.loinc.bluebert').viz("""A 28-year-old female with a history of gestational diabetes mellitus diagnosed eight years prior to presentation and
subsequent type two diabetes mellitus (TSS2DM), one prior episode of HTG-induced pancreatitis three years prior to presentation, associated with an acute
hepatitis, and obesity with a body mass index (BMI) of 33.5 kg/m2, presented with a one-week history of polyuria, polydipsia, poor appetite, and vomiting.""")
# + [markdown] id="ZuHLreVA6nAW"
# #### [Entity Resolver for Human Phenotype Ontology](https://nlp.johnsnowlabs.com/2021/05/16/sbiobertresolve_HPO_en.html)
# + id="R6cdGc-16mrY" colab={"base_uri": "https://localhost:8080/", "height": 440} outputId="484e8b34-e228-4617-9fb0-08bd92e2585b"
nlu.load('med_ner.jsl.wip.clinical en.resolve.HPO').predict("""These disorders include cancer, bipolar disorder, schizophrenia, autism, Cri-du-chat syndrome,
myopia, cortical cataract-linked Alzheimer's disease, and infectious diseases"""
,output_level = 'sentence')
# + id="PYAF55oz4By5" colab={"base_uri": "https://localhost:8080/", "height": 414} outputId="565cfd0a-f51b-4f57-c31a-aa61bba33c8a"
nlu.load('med_ner.jsl.wip.clinical en.resolve.HPO').viz("""These disorders include cancer, bipolar disorder, schizophrenia, autism, Cri-du-chat syndrome,
myopia, cortical cataract-linked Alzheimer's disease, and infectious diseases""")
# + id="vdn-Hj8sj_gK"
|
nlu/release_notebooks/NLU_3_0_2_release_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Permanganant/Shazam_Song_Downloader/blob/main/Shazam_song_downloader.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="exzHLXjuugHs"
# !pip install pytube
# !pip install youtube-search-python
# !pip3 install ShazamAPI
# !pip install ffmpeg-python > /dev/null
# !pip install torchaudio
# !pip install moviepy
# + id="_FSeGRawDt1_"
#Required Libraries
from pytube import YouTube
from pytube import Search
from ShazamAPI import Shazam
import json, requests
from IPython.display import HTML, Audio
from google.colab.output import eval_js
from base64 import b64decode
import numpy as np
import io
import ffmpeg
import tempfile
import pathlib
import torchaudio
import moviepy.editor as mp
# + colab={"base_uri": "https://localhost:8080/", "height": 96} id="kcvFQxbnpmlr" outputId="eb48dd5d-acd2-4e61-bff6-7e884187d542"
#To get audio for Colab this part of the code taken with some little modifications from https://ricardodeazambuja.com/deep_learning/2019/03/09/audio_and_video_google_colab/
AUDIO_HTML = """
<script>
var my_div = document.createElement("DIV");
var my_p = document.createElement("P");
var my_btn = document.createElement("BUTTON");
var t = document.createTextNode("Press to start recording");
my_btn.appendChild(t);
//my_p.appendChild(my_btn);
my_div.appendChild(my_btn);
document.body.appendChild(my_div);
var base64data = 0;
var reader;
var recorder, gumStream;
var recordButton = my_btn;
var handleSuccess = function(stream) {
gumStream = stream;
var options = {
//bitsPerSecond: 8000, //chrome seems to ignore, always 48k
mimeType : 'audio/webm;codecs=opus'
//mimeType : 'audio/webm;codecs=pcm'
};
//recorder = new MediaRecorder(stream, options);
recorder = new MediaRecorder(stream);
recorder.ondataavailable = function(e) {
var url = URL.createObjectURL(e.data);
var preview = document.createElement('audio');
preview.controls = true;
preview.src = url;
document.body.appendChild(preview);
reader = new FileReader();
reader.readAsDataURL(e.data);
reader.onloadend = function() {
base64data = reader.result;
//console.log("Inside FileReader:" + base64data);
}
};
recorder.start();
};
recordButton.innerText = "Recording... press to stop";
navigator.mediaDevices.getUserMedia({audio: true}).then(handleSuccess);
function toggleRecording() {
if (recorder && recorder.state == "recording") {
recorder.stop();
gumStream.getAudioTracks()[0].stop();
recordButton.innerText = "Saving the recording... pls wait!"
}
}
// https://stackoverflow.com/a/951057
function sleep(ms) {
return new Promise(resolve => setTimeout(resolve, ms));
}
var data = new Promise(resolve=>{
//recordButton.addEventListener("click", toggleRecording);
recordButton.onclick = ()=>{
toggleRecording()
sleep(2000).then(() => {
// wait 2000ms for the data to be available...
// ideally this should use something like await...
//console.log("Inside data:" + base64data)
resolve(base64data.toString())
});
}
});
</script>
"""
def get_audio():
display(HTML(AUDIO_HTML))
data = eval_js("data")
binary = b64decode(data.split(',')[1])
process = (ffmpeg
.input('pipe:0')
.output('pipe:1', format='wav')
.run_async(pipe_stdin=True, pipe_stdout=True, pipe_stderr=True, quiet=True, overwrite_output=True)
)
output, err = process.communicate(input=binary)
riff_chunk_size = len(output) - 8
# Break up the chunk size into four bytes, held in b.
q = riff_chunk_size
b = []
for i in range(4):
q, r = divmod(q, 256)
b.append(r)
# Replace bytes 4:8 in proc.stdout with the actual size of the RIFF chunk.
riff = output[:4] + bytes(b) + output[8:]
path = '/content/tmp.wav'
with open(path, 'wb') as f:
f.write(riff)
x, sr = torchaudio.load(path)
return x, sr
audio, sr = get_audio()
# + id="YHN28BRCuuwg"
mp3_file_content_to_recognize = open('tmp.wav', 'rb').read()
shazam = Shazam(mp3_file_content_to_recognize)
recognize_generator = shazam.recognizeSong()
try:
youtube_json = next(recognize_generator)[1]['track']['sections'][2]['youtubeurl']
except ValueError:
print("Oopss..Can't find song, Please Try Again")
# + id="gmVNoDNbVhfs" colab={"base_uri": "https://localhost:8080/"} outputId="a8d2a926-29ab-4e20-b1ce-4ffb46f1f036"
url = requests.get(youtube_json)
text = url.text
data = json.loads(text)
video_uri = data['actions'][0]['uri']
print("Video URI: " ,video_uri)
# + id="QrJCyWRscqj8" colab={"base_uri": "https://localhost:8080/"} outputId="0c006e74-87c7-4c41-cd7d-c08ac3494318"
yt = YouTube(video_uri)
len_query = len(yt.streams)
q = yt.streams
resolution = {'144':False,'240':False,'360':False,'480':False,'720':False,'1080':False,'1440':False,'2160':False}
for i in range (len_query):
q = yt.streams[i]
if q.resolution == '144p':
resolution['144'] = True
elif q.resolution == '240p':
resolution['240'] = True
elif q.resolution == '360p':
resolution['360'] = True
elif q.resolution == '480p':
resolution['480'] = True
elif q.resolution == '720p':
resolution['720'] = True
elif q.resolution == '1080p':
resolution['1080'] = True
elif q.resolution == '1440p':
resolution['1440'] = True
elif q.resolution == '2160p':
resolution['2160'] = True
print("Available resolutions:")
for i in list(resolution.keys()):
if resolution[i] != False:
print(i)
# + id="QCDs48APXsCp"
x = True
y = True
while x == True:
resolution_key = input("Enter a valid Resolution value: ")
for i in range(len(yt.streams)):
if yt.streams[i].resolution == resolution_key+'p':
x = False
while y == True:
key = input("To download as Video click --> v , To download as a mp3 click --> 3 :")
if key == 'v':
try:
yt.streams[i].download()
print("Video downloaded sucessfully")
break
except ValueError:
print("Oopss..Link is not working")
break
y = False
elif key == '3':
try:
song = yt.streams[i].download()
clip = mp.VideoFileClip(song).subclip(0,20)
clip.audio.write_audiofile("theaudio.mp3")
print("MP3 song downloaded sucessfully")
break
except ValueError:
print("Oopss..Link is not working")
break
y = False
break
if x == True:
print("Oopss.. Please enter a valid resolution")
|
Shazam_song_downloader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''base'': conda)'
# name: python3
# ---
# + [markdown] id="SuuFz3YKhsFZ"
# # Tutorial 1: Basics
#
#
# In this tutorial you will learn how to:
# * run LightAutoML training on tabular data
# * obtain feature importances and reports
# * configure resource usage in LightAutoML
#
# Official LightAutoML github repository is [here](https://github.com/sberbank-ai-lab/LightAutoML)
# + [markdown] id="2FzFDY0-hsFb" papermill={"duration": 0.032379, "end_time": "2021-06-22T20:10:29.835505", "exception": false, "start_time": "2021-06-22T20:10:29.803126", "status": "completed"} tags=[]
# <img src="https://github.com/sberbank-ai-lab/LightAutoML/blob/master/imgs/LightAutoML_logo_big.png?raw=1" alt="LightAutoML logo" style="width:100%;"/>
# + [markdown] id="Ocu3h1vQhsFc"
# ## 0. Prerequisites
# + [markdown] id="f1VLa9AlhsFd"
# ### 0.0. install LightAutoML
# + _kg_hide-output=true id="8K1x-sAWhsFd" papermill={"duration": 23.023261, "end_time": "2021-06-22T20:10:52.955691", "exception": false, "start_time": "2021-06-22T20:10:29.932430", "status": "completed"} tags=[]
# # !pip install -U lightautoml
# + [markdown] id="W3jBDmHahsFe" papermill={"duration": 0.066681, "end_time": "2021-06-22T20:10:53.090975", "exception": false, "start_time": "2021-06-22T20:10:53.024294", "status": "completed"} tags=[]
# ### 0.1. Import libraries
#
# Here we will import the libraries we use in this kernel:
# - Standard python libraries for timing, working with OS etc.
# - Essential python DS libraries like numpy, pandas, scikit-learn and torch (the last we will use in the next cell)
# - LightAutoML modules: presets for AutoML, task and report generation module
# + id="1qHj6ybRhsFf" papermill={"duration": 8.32949, "end_time": "2021-06-22T20:11:01.487788", "exception": false, "start_time": "2021-06-22T20:10:53.158298", "status": "completed"} tags=[]
# Standard python libraries
import os
import time
import joblib
# Essential DS libraries
import numpy as np
import pandas as pd
from sklearn.metrics import roc_auc_score
from sklearn.metrics import mean_absolute_error, mean_absolute_percentage_error, r2_score
from sklearn.model_selection import train_test_split
import torch
# LightAutoML presets, task and report generation
from lightautoml.automl.presets.tabular_presets import TabularAutoML, TabularUtilizedAutoML
from lightautoml.tasks import Task
from lightautoml.report.report_deco import ReportDeco
from utils import score_model
# + [markdown] id="I0IlMLt3hsFg" papermill={"duration": 0.064234, "end_time": "2021-06-22T20:11:01.619010", "exception": false, "start_time": "2021-06-22T20:11:01.554776", "status": "completed"} tags=[]
# ### 0.2. Constants
#
# Here we setup the constants to use in the kernel:
# - `N_THREADS` - number of vCPUs for LightAutoML model creation
# - `N_FOLDS` - number of folds in LightAutoML inner CV
# - `RANDOM_STATE` - random seed for better reproducibility
# - `TEST_SIZE` - houldout data part size
# - `TIMEOUT` - limit in seconds for model to train
# - `TARGET_NAME` - target column name in dataset
# + id="bodnajRNhsFh" papermill={"duration": 0.077787, "end_time": "2021-06-22T20:11:01.761030", "exception": false, "start_time": "2021-06-22T20:11:01.683243", "status": "completed"} tags=[]
N_THREADS = 10
N_FOLDS = 5
RANDOM_STATE = 42
TEST_SIZE = 0.2
TIMEOUT = 300
TARGET_NAME = 'DSHORTT1138P2300058'
# + id="HC_Au-z_hsFh"
DATASET_DIR = 'data/'
DATASET_NAME = 'clean_train.csv'
DATASET_FULLNAME = os.path.join(DATASET_DIR, DATASET_NAME)
# DATASET_URL = 'https://raw.githubusercontent.com/sberbank-ai-lab/LightAutoML/master/examples/data/sampled_app_train.csv'
# + [markdown] id="mWdjGDsUhsFi" papermill={"duration": 0.086481, "end_time": "2021-06-22T20:11:01.927314", "exception": false, "start_time": "2021-06-22T20:11:01.840833", "status": "completed"} tags=[]
# ### 0.3. Imported models setup
#
# For better reproducibility fix numpy random seed with max number of threads for Torch (which usually try to use all the threads on server):
# + id="PhlswEnYhsFj" papermill={"duration": 0.087268, "end_time": "2021-06-22T20:11:02.092497", "exception": false, "start_time": "2021-06-22T20:11:02.005229", "status": "completed"} tags=[]
np.random.seed(RANDOM_STATE)
torch.set_num_threads(N_THREADS)
# + [markdown] id="GALb5UnUhsFj" papermill={"duration": 0.072033, "end_time": "2021-06-22T20:11:02.238196", "exception": false, "start_time": "2021-06-22T20:11:02.166163", "status": "completed"} tags=[]
# ### 0.4. Data loading
# Let's check the data we have:
# + id="UX0d5KCYhsFj"
if not os.path.exists(DATASET_FULLNAME):
os.makedirs(DATASET_DIR, exist_ok=True)
dataset = requests.get(DATASET_URL).text
with open(DATASET_FULLNAME, 'w') as output:
output.write(dataset)
# + id="y5meBmpRhsFk" outputId="21640443-7726-4eb8-de78-5138bec39c81" papermill={"duration": 12.710747, "end_time": "2021-06-22T20:11:15.018360", "exception": false, "start_time": "2021-06-22T20:11:02.307613", "status": "completed"} tags=[]
data = pd.read_csv(DATASET_FULLNAME)
data.drop(['Unnamed: 0', 'UUID', 'UNIXDT'], axis=1, inplace=True)
data.head()
# -
data = data.fillna(0)
to_shift = ['T1138P6000096', 'T1138P6000315', 'DMIDT1138P4000064',
'DSHORTT1138P4000064', 'DLONGT1138P4000064', 'DMIDT1138P2600012',
'DSHORTT1138P2600012', 'DLONGT1138P2600012', 'DMIDT1205P2300000',
'DSHORTT1205P2300000', 'DLONGT1205P2300000', 'T1205P2300000',
'T1138P4000064', 'T1138P2600012', 'T1138P600050', 'T1013P500399',
'DSHORTT1138P2300058']
# + id="YaXTGB5ShsFl" outputId="588a21ce-9aca-4211-9076-a107101b4fa3" papermill={"duration": 0.077509, "end_time": "2021-06-22T20:11:15.161419", "exception": false, "start_time": "2021-06-22T20:11:15.083910", "status": "completed"} tags=[]
for var in to_shift:
data[var + '_l1'] = data.sort_values(['WELL_ID', 'DT']).groupby('WELL_ID', group_keys=False)[var].shift()
data[var + '_l2'] = data.sort_values(['WELL_ID', 'DT']).groupby('WELL_ID', group_keys=False)[var].shift(2)
# -
data = data.dropna()
data
data.isnull().sum(axis = 0)
# + [markdown] id="e070gErIhsFm" papermill={"duration": 0.065323, "end_time": "2021-06-22T20:11:21.676311", "exception": false, "start_time": "2021-06-22T20:11:21.610988", "status": "completed"} tags=[]
# ### 0.5. Data splitting for train-holdout
# As we have only one file with target values, we can split it into 80%-20% for holdout usage:
# + id="oIFDDubshsFm" outputId="ebf65f6d-ebac-4dbb-a5da-b6bc0a1dc9b5" papermill={"duration": 0.793619, "end_time": "2021-06-22T20:11:22.537798", "exception": false, "start_time": "2021-06-22T20:11:21.744179", "status": "completed"} tags=[]
# tr_data, te_data = train_test_split(
# data,
# test_size=TEST_SIZE,
# stratify=data['WELL_ID'],
# random_state=RANDOM_STATE
# )
p = 100 * TEST_SIZE
te_data = data.sort_values(['WELL_ID', 'DT'])\
.groupby('WELL_ID', group_keys=False)\
.apply(lambda x: x.tail(int(len(x) * (p / 100))))
tr_data = data.sort_values(['WELL_ID', 'DT'])\
.groupby('WELL_ID', group_keys=False)\
.apply(lambda x: x.head(int(len(x) * (1-TEST_SIZE))))
print(f'Data splitted. Parts sizes: tr_data = {tr_data.shape}, te_data = {te_data.shape}')
tr_data.head()
# + [markdown] id="dsmLK7YRhsFm" papermill={"duration": 0.071526, "end_time": "2021-06-22T20:11:22.853156", "exception": false, "start_time": "2021-06-22T20:11:22.781630", "status": "completed"} tags=[]
# # 1. Task definition
# + [markdown] id="bB7xtQH8hsFn"
# ### 1.1. Task type
#
# On the cell below we create Task object - the class to setup what task LightAutoML model should solve with specific loss and metric if necessary (more info can be found [here](https://lightautoml.readthedocs.io/en/latest/generated/lightautoml.tasks.base.Task.html#lightautoml.tasks.base.Task) in our documentation):
# + id="l7B_h1gRhsFn" papermill={"duration": 0.086442, "end_time": "2021-06-22T20:11:23.010643", "exception": false, "start_time": "2021-06-22T20:11:22.924201", "status": "completed"} tags=[]
task = Task('reg', metric='mae')
# + [markdown] id="l9M-S4ohhsFn" papermill={"duration": 0.070103, "end_time": "2021-06-22T20:11:23.150929", "exception": false, "start_time": "2021-06-22T20:11:23.080826", "status": "completed"} tags=[]
# ### 1.2. Feature roles setup
# + [markdown] id="BXqKJVtghsFo" papermill={"duration": 0.069372, "end_time": "2021-06-22T20:11:23.290153", "exception": false, "start_time": "2021-06-22T20:11:23.220781", "status": "completed"} tags=[]
# To solve the task, we need to setup columns roles. The **only role you must setup is target role**, everything else (drop, numeric, categorical, group, weights etc.) is up to user - LightAutoML models have automatic columns typization inside:
# + id="UNg-F99qhsFo" papermill={"duration": 0.07715, "end_time": "2021-06-22T20:11:23.438830", "exception": false, "start_time": "2021-06-22T20:11:23.361680", "status": "completed"} tags=[]
roles = {
'target': TARGET_NAME,
'drop': ['DT']
}
# + [markdown] id="ohNn5FCmhsFo" papermill={"duration": 0.074284, "end_time": "2021-06-22T20:11:23.582462", "exception": false, "start_time": "2021-06-22T20:11:23.508178", "status": "completed"} tags=[]
# ### 1.3. LightAutoML model creation - TabularAutoML preset
# + [markdown] id="Mnq1ZsnshsFo" papermill={"duration": 0.072649, "end_time": "2021-06-22T20:11:23.726154", "exception": false, "start_time": "2021-06-22T20:11:23.653505", "status": "completed"} tags=[]
# In next the cell we are going to create LightAutoML model with `TabularAutoML` class - preset with default model structure like in the image below:
#
# <img src="https://github.com/sberbank-ai-lab/LightAutoML/blob/master/imgs/tutorial_blackbox_pipeline.png?raw=1" alt="TabularAutoML preset pipeline" style="width:85%;"/>
#
# in just several lines. Let's discuss the params we can setup:
# - `task` - the type of the ML task (the only **must have** parameter)
# - `timeout` - time limit in seconds for model to train
# - `cpu_limit` - vCPU count for model to use
# - `reader_params` - parameter change for Reader object inside preset, which works on the first step of data preparation: automatic feature typization, preliminary almost-constant features, correct CV setup etc. For example, we setup `n_jobs` threads for typization algo, `cv` folds and `random_state` as inside CV seed.
#
# **Important note**: `reader_params` key is one of the YAML config keys, which is used inside `TabularAutoML` preset. [More details](https://github.com/sberbank-ai-lab/LightAutoML/blob/master/lightautoml/automl/presets/tabular_config.yml) on its structure with explanation comments can be found on the link attached. Each key from this config can be modified with user settings during preset object initialization. To get more info about different parameters setting (for example, ML algos which can be used in `general_params->use_algos`) please take a look at our [article on TowardsDataScience](https://towardsdatascience.com/lightautoml-preset-usage-tutorial-2cce7da6f936).
#
# Moreover, to receive the automatic report for our model we will use `ReportDeco` decorator and work with the decorated version in the same way as we do with usual one.
# + id="scfhGQ-zhsFp"
automl = TabularAutoML(
task = task,
timeout = TIMEOUT,
cpu_limit = N_THREADS,
reader_params = {'n_jobs': N_THREADS, 'cv': N_FOLDS, 'random_state': RANDOM_STATE}
)
# + [markdown] id="F4wgkgtdhsFp"
# # 2. AutoML training
# + [markdown] id="ijBDGCgxhsFp"
# To run autoML training use fit_predict method:
# - `train_data` - Dataset to train.
# - `roles` - Roles dict.
# - `verbose` - Controls the verbosity: the higher, the more messages.
# <1 : messages are not displayed;
# >=1 : the computation process for layers is displayed;
# >=2 : the information about folds processing is also displayed;
# >=3 : the hyperparameters optimization process is also displayed;
# >=4 : the training process for every algorithm is displayed;
#
# Note: out-of-fold prediction is calculated during training and returned from the fit_predict method
# + id="wmuNBHeIhsFp" outputId="d7c82260-686a-4d0d-e0a8-2ac6cbfea789"
# %%time
oof_pred = automl.fit_predict(tr_data, roles = roles, verbose = 1)
# + [markdown] id="ijJdL5wRhsFq" papermill={"duration": 0.145098, "end_time": "2021-06-22T20:34:32.530768", "exception": false, "start_time": "2021-06-22T20:34:32.385670", "status": "completed"} tags=[]
# # 3. Prediction on holdout and model evaluation
# -
pd.DataFrame(oof_pred.data[:, 0])[:100].plot()
# + id="TB1nI8SfhsFq" outputId="fe4f1010-a5ae-4ebe-9100-4375468235e9"
# %%time
te_pred = automl.predict(te_data)
print(f'Prediction for te_data:\n{te_pred}\nShape = {te_pred.shape}')
# -
pd.DataFrame(te_data[TARGET_NAME].values)[:100].plot()
te_data[TARGET_NAME].values
oof_pred.data[:, 0]
sum(~np.isnan(oof_pred.data[:, 0]))
# + id="Js7zljjFhsFq" outputId="b3a7a068-e174-48a6-9058-7861972f7237"
print(f'OOF score: {mean_absolute_error(tr_data[TARGET_NAME].values, oof_pred.data[:, 0])}')
print(f'HOLDOUT score: {mean_absolute_error(te_data[TARGET_NAME].values, te_pred.data[:, 0])}')
# -
pd.DataFrame({'true':tr_data[TARGET_NAME].values, 'predict':oof_pred.data[:, 0]})[0:200].plot(figsize=(20,10))
# + id="DW-BJ_mRhsFr"
pd.DataFrame({'true':te_data[TARGET_NAME].values, 'predict':te_pred.data[:, 0]})[0:200].plot(figsize=(20,10))
# + [markdown] id="2_22H2ZOhsFr"
# # 4. Model analysis
# + [markdown] id="esrgT6nQhsFr"
# ## 4.1. Reports
# + [markdown] id="O5Ow6gGXhsFr"
# You can obtain the description of the resulting pipeline:
# + id="rtz-avkChsFr" outputId="cd417de0-e5c2-4fdc-b4a5-7ce80fa45d69"
print(automl.create_model_str_desc())
# + [markdown] id="BetrfmW5hsFs"
# Also for this purposes LightAutoML have ReportDeco, use it to build reports:
# + _kg_hide-output=true id="r4AZQVN-hsFs" papermill={"duration": 1030.159528, "end_time": "2021-06-22T20:28:33.963476", "exception": false, "start_time": "2021-06-22T20:11:23.803948", "status": "completed"} tags=[]
RD = ReportDeco(output_path = 'tabularAutoML_model_report')
automl_rd = RD(
TabularAutoML(
task = task,
timeout = TIMEOUT,
cpu_limit = N_THREADS,
reader_params = {'n_jobs': N_THREADS, 'cv': N_FOLDS, 'random_state': RANDOM_STATE}
)
)
# + id="D9LESAVZhsFs" outputId="fe00b3f2-c320-4247-87da-271cf08a5d74"
# %%time
oof_pred = automl_rd.fit_predict(tr_data, roles = roles, verbose = 1)
# + [markdown] id="Kx2hz5sihsFs"
# So the report is available in tabularAutoML_model_report folder
# + id="p2sDibohhsFt" outputId="e24676d8-29ad-4987-cf6e-81fff27e5689"
# !ls tabularAutoML_model_report
# + id="LduiXBMLhsFt" outputId="f278b287-7f55-453f-e60d-789720b10b48" papermill={"duration": 22.483603, "end_time": "2021-06-22T20:34:55.170931", "exception": false, "start_time": "2021-06-22T20:34:32.687328", "status": "completed"} tags=[]
# %%time
te_pred = automl_rd.predict(te_data)
print(f'Prediction for te_data:\n{te_pred}\nShape = {te_pred.shape}')
# + id="sRm8fz-ehsFt" outputId="5a155d55-8d9b-4934-aab3-73975b1519b9" papermill={"duration": 0.310292, "end_time": "2021-06-22T20:34:55.630539", "exception": false, "start_time": "2021-06-22T20:34:55.320247", "status": "completed"} tags=[]
print(f'OOF score: {mean_absolute_error(tr_data[TARGET_NAME].values, oof_pred.data[:, 0])}')
print(f'HOLDOUT score: {mean_absolute_error(te_data[TARGET_NAME].values, te_pred.data[:, 0])}')
# + id="2IhxQHnthsFt"
# + [markdown] id="jQNos2WKhsFu" papermill={"duration": 0.113545, "end_time": "2021-06-22T20:28:34.191432", "exception": false, "start_time": "2021-06-22T20:28:34.077887", "status": "completed"} tags=[]
# ## 4.2 Feature importances calculation
#
# For feature importances calculation we have 2 different methods in LightAutoML:
# - Fast (`fast`) - this method uses feature importances from feature selector LGBM model inside LightAutoML. It works extremely fast and almost always (almost because of situations, when feature selection is turned off or selector was removed from the final models with all GBM models). no need to use new labelled data.
# - Accurate (`accurate`) - this method calculate *features permutation importances* for the whole LightAutoML model based on the **new labelled data**. It always works but can take a lot of time to finish (depending on the model structure, new labelled dataset size etc.).
#
# In the cell below we will use `automl_rd.model` instead `automl_rd` because we want to take the importances from the model, not from the report. But **be carefull** - everything, which is calculated using `automl_rd.model` will not go to the report.
# + id="vMsvxtsMhsFu" outputId="5b156128-7d4d-4345-ef25-bf63807af838" papermill={"duration": 4.063771, "end_time": "2021-06-22T20:28:38.368139", "exception": false, "start_time": "2021-06-22T20:28:34.304368", "status": "completed"} tags=[]
# %%time
# Fast feature importances calculation
fast_fi = automl_rd.model.get_feature_scores('fast')
fast_fi.set_index('Feature')['Importance'].plot.bar(figsize = (30, 10), grid = True)
# + _kg_hide-output=true id="ZMfyepPdhsFu" outputId="37a6d97c-f96d-4811-ae1e-4ed14af9bfca" papermill={"duration": 349.538712, "end_time": "2021-06-22T20:34:28.353655", "exception": false, "start_time": "2021-06-22T20:28:38.814943", "status": "completed"} tags=[]
# %%time
# Accurate feature importances calculation (Permutation importances) - can take long time to calculate
accurate_fi = automl_rd.model.get_feature_scores('accurate', te_data, silent = False)
# + id="dMfHAgtkhsFu" outputId="42d7cbdc-5c41-49f9-e961-35177250f125" papermill={"duration": 3.739686, "end_time": "2021-06-22T20:34:32.232636", "exception": false, "start_time": "2021-06-22T20:34:28.492950", "status": "completed"} tags=[]
accurate_fi.set_index('Feature')['Importance'].plot.bar(figsize = (30, 10), grid = True)
# + [markdown] id="Q7qUKTYahsFv" papermill={"duration": 0.143136, "end_time": "2021-06-22T20:35:48.498828", "exception": false, "start_time": "2021-06-22T20:35:48.355692", "status": "completed"} tags=[]
# ## Bonus: where is the automatic report?
#
# As we used `automl_rd` in our training and prediction cells, it is already ready in the folder we specified - you can check the output kaggle folder and find the `tabularAutoML_model_report` folder with `lama_interactive_report.html` report inside (or just [click this link](tabularAutoML_model_report/lama_interactive_report.html) for short). It's interactive so you can click the black triangles on the left of the texts to go deeper in selected part.
# + id="1Rp2wd6rhsFv"
# + [markdown] id="wW8l6clhhsFv"
# # 5. Spending more from TIMEOUT - `TabularUtilizedAutoML` usage
#
# Using `TabularAutoML` we spent only 31 second to build the model with setup `TIMEOUT` equal to 5 minutes. To spend (almost) all the `TIMEOUT` we can use `TabularUtilizedAutoML` preset instead of `TabularAutoML`, which has the same API:
# + id="U_cYHJ2uhsFv"
utilized_automl = TabularUtilizedAutoML(
task = task,
timeout = 1000,
cpu_limit = N_THREADS,
reader_params = {'n_jobs': N_THREADS, 'cv': N_FOLDS, 'random_state': RANDOM_STATE},
)
# + id="5WkZpcQQhsFv" outputId="2fee2294-c7c0-4352-fc7e-9dd7b9e8801d"
# %%time
oof_pred = utilized_automl.fit_predict(tr_data, roles = roles, verbose = 1)
# + id="AMFQla-shsFw" outputId="3f11d834-c96d-4206-8630-bed093283634"
print('oof_pred:\n{}\nShape = {}'.format(oof_pred, oof_pred.shape))
# + id="kzQ9rl4zhsFw" outputId="ed9b1de4-bd99-42ea-b512-e4b32cafbdac"
print(utilized_automl.create_model_str_desc())
# + [markdown] id="gm7JW5Y5hsFx"
# Prediction on holdout and metric calculation
# + id="zO8UwnmHhsFx" outputId="be7344d3-6fbf-4645-b354-acc669982de4"
# %%time
te_pred = utilized_automl.predict(te_data)
print(f'Prediction for te_data:\n{te_pred}\nShape = {te_pred.shape}')
# + id="yYf1-yWxhsFx" outputId="acab6aac-9036-4e08-91bd-84de4ced2df4"
print(f'OOF score: {mean_absolute_error(tr_data[TARGET_NAME].values, oof_pred.data[:, 0])}')
print(f'HOLDOUT score: {mean_absolute_error(te_data[TARGET_NAME].values, te_pred.data[:, 0])}')
# -
# # predict on valid 1 and 2
# + id="DLxpDseVhsFx"
def read_valid(link = DATASET_FULLNAME):
data = pd.read_csv(link)
data.drop(['Unnamed: 0', 'UUID', 'UNIXDT'], axis=1, inplace=True)
data.drop(['DLONGT1138P2300058',
'DMIDT1138P2300058',
'LONGUPT1138P2300058',
'MIDUPT1138P2300058',
'SHORTUPT1138P2300058'], axis=1, inplace=True, errors='ignore')
data = data.fillna(0)
to_shift = ['T1138P6000096', 'T1138P6000315', 'DMIDT1138P4000064',
'DSHORTT1138P4000064', 'DLONGT1138P4000064', 'DMIDT1138P2600012',
'DSHORTT1138P2600012', 'DLONGT1138P2600012', 'DMIDT1205P2300000',
'DSHORTT1205P2300000', 'DLONGT1205P2300000', 'T1205P2300000',
'T1138P4000064', 'T1138P2600012', 'T1138P600050', 'T1013P500399',
'DSHORTT1138P2300058']
for var in to_shift:
data[var + '_l1'] = data.sort_values(['WELL_ID', 'DT']).groupby('WELL_ID', group_keys=False)[var].shift()
data[var + '_l2'] = data.sort_values(['WELL_ID', 'DT']).groupby('WELL_ID', group_keys=False)[var].shift(2)
data = data.fillna(0)
data = data.sort_values(['WELL_ID', 'DT'])
return data
# -
valid1 = 'data/cfp_dataset_v1_valid1.csv'
valid2 = 'data/cfp_dataset_v1_valid2.csv'
validdata1 = read_valid(valid1)
validdata2 = read_valid(valid2)
pred_valid1 = utilized_automl.predict(validdata1)
pred_valid2 = utilized_automl.predict(validdata2)
Lama_prediction = {'Valid1':{'true':validdata1[TARGET_NAME].values,
'prediction':pred_valid1.data[:, 0]},
'Valid2':{'true':validdata2[TARGET_NAME].values,
'prediction':pred_valid2.data[:, 0]}}
automl_validation_results = score_model(Lama_prediction)
automl_validation_results.loc['mean_auto_ml_valid'] = automl_validation_results.mean()
automl_validation_results
automl_validation_results.to_csv('automl_validation_results.csv', sep=';', decimal=',', index=True)
pd.DataFrame({'true':validdata1[TARGET_NAME].values, 'predict':pred_valid1.data[:, 0]})[0:200].plot(figsize=(20,10))
pd.DataFrame({'true':validdata2[TARGET_NAME].values, 'predict':pred_valid2.data[:, 0]})[0:200].plot(figsize=(20,10))
joblib.dump(utilized_automl, 'utilized_automl_model.pkl')
utilized_automl=joblib.load('utilized_automl_model.pkl')
# + [markdown] id="do1KErj3hsFy" papermill={"duration": 0.14221, "end_time": "2021-06-22T20:35:48.782561", "exception": false, "start_time": "2021-06-22T20:35:48.640351", "status": "completed"} tags=[]
# # Additional materials
# + [markdown] id="5uGtcmcDhsFy" papermill={"duration": 0.147943, "end_time": "2021-06-22T20:35:49.074531", "exception": false, "start_time": "2021-06-22T20:35:48.926588", "status": "completed"} tags=[]
# - [Official LightAutoML github repo](https://github.com/sberbank-ai-lab/LightAutoML)
# - [LightAutoML documentation](https://lightautoml.readthedocs.io/en/latest)
|
LAMA_basic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unsupervised methods
#
# In this lesson, we'll cover unsupervised computational text anlalysis approaches. The central methods covered are TF-IDF and Topic Modeling. Both of these are common approachs in the social sciences and humanities.
#
# [DTM/TF-IDF](#dtm)<br>
#
# [Topic modeling](#topics)<br>
#
# ### Today you will
# * Understand the DTM and why it's important to text analysis
# * Learn how to create a DTM in Python
# * Learn basic functionality of Python's package scikit-learn
# * Understand tf-idf scores
# * Learn a simple way to identify distinctive words
# * Implement a basic topic modeling algorithm and learn how to tweak it
# * In the process, gain more familiarity and comfort with the Pandas package and manipulating data
#
#
# ### Key Jargon
# * *Document Term Matrix*:
# * a matrix that describes the frequency of terms that occur in a collection of documents. In a document-term matrix, rows correspond to documents in the collection and columns correspond to terms.
# * *TF-IDF Scores*:
# * short for term frequency–inverse document frequency, is a numerical statistic that is intended to reflect how important a word is to a document in a collection or corpus.
# * *Topic Modeling*:
# * A general class of statistical models that uncover abstract topics within a text. It uses the co-occurrence of words within documents, compared to their distribution across documents, to uncover these abstract themes. The output is a list of weighted words, which indicate the subject of each topic, and a weight distribution across topics for each document.
#
# * *LDA*:
# * Latent Dirichlet Allocation. A particular model for topic modeling. It does not take document order into account, unlike other topic modeling algorithms.
# ## DTM/TF-IDF <a id='dtm'></a>
#
# In this lesson we will use Python's scikit-learn package learn to make a document term matrix from a .csv Music Reviews dataset (collected from MetaCritic.com). We will then use the DTM and a word weighting technique called tf-idf (term frequency inverse document frequency) to identify important and discriminating words within this dataset (utilizing the Pandas package). The illustrating question: **what words distinguish reviews of Rap albums, Indie Rock albums, and Jazz albums?**
# +
import os
import numpy as np
import pandas as pd
DATA_DIR = 'data'
music_fname = 'music_reviews.csv'
music_fname = os.path.join(DATA_DIR, music_fname)
# -
# ### First attempt at reading in file
reviews = pd.read_csv(music_fname, sep='\t')
reviews.head()
# Print the text of the first review.
print(reviews['body'][0])
# ### Explore the Data using Pandas
#
# Let's first look at some descriptive statistics about this dataset, to get a feel for what's in it. We'll do this using the Pandas package.
#
# Note: this is always good practice. It serves two purposes. It checks to make sure your data is correct, and there's no major errors. It also keeps you in touch with your data, which will help with interpretation. <3 your data!
#
# First, what genres are in this dataset, and how many reviews in each genre?
#We can count this using the value_counts() function
reviews['genre'].value_counts()
# The first thing most people do is to `describe` their data. (This is the `summary` command in R, or the `sum` command in Stata).
#There's only one numeric column in our data so we only get one column for output.
reviews.describe()
# This only gets us numerical summaries. To get summaries of some of the other columns, we can explicitly ask for it.
reviews.describe(include=['O'])
# Who were the reviewers?
reviews['critic'].value_counts().head(10)
# And the artists?
reviews['artist'].value_counts().head(10)
# We can get the average score as follows:
reviews['score'].mean()
# Now we want to know the average score for each genre? To do this, we use Pandas `groupby` function. You'll want to get very familiar with the `groupby` function. It's quite powerful. (Similar to `collapse` on Stata)
reviews_grouped_by_genre = reviews.groupby("genre")
reviews_grouped_by_genre['score'].mean().sort_values(ascending=False)
# ### Creating the DTM using scikit-learn
#
# Ok, that's the summary of the metadata. Next, we turn to analyzing the text of the reviews. Remember, the text is stored in the 'body' column. First, a preprocessing step to remove numbers.
# +
def remove_digits(comment):
return ''.join([ch for ch in comment if not ch.isdigit()])
reviews['body_without_digits'] = reviews['body'].apply(remove_digits)
reviews
# -
reviews['body_without_digits'].head()
# ### CountVectorizer Function
#
# Our next step is to turn the text into a document term matrix using the scikit-learn function called `CountVectorizer`.
# +
from sklearn.feature_extraction.text import CountVectorizer
countvec = CountVectorizer()
sparse_dtm = countvec.fit_transform(reviews['body_without_digits'])
# -
# Great! We made a DTM! Let's look at it.
sparse_dtm
# This format is called Compressed Sparse Format. It save a lot of memory to store the dtm in this format, but it is difficult to look at for a human. To illustrate the techniques in this lesson we will first convert this matrix back to a Pandas DataFrame, a format we're more familiar with. For larger datasets, you will have to use the Compressed Sparse Format. Putting it into a DataFrame, however, will enable us to get more comfortable with Pandas!
dtm = pd.DataFrame(sparse_dtm.toarray(), columns=countvec.get_feature_names(), index=reviews.index)
dtm.head()
# ### What can we do with a DTM?
#
# We can quickly identify the most frequent words
dtm.sum().sort_values(ascending=False).head(10)
# ### Challenge - SOLUTION
#
# * Print out the most infrequent words rather than the most frequent words. You can look at the [Pandas documentation](http://pandas.pydata.org/pandas-docs/stable/api.html#api-dataframe-stats) for more information.
# * Print the average number of times each word is used in a review.
# * Print this out sorted from highest to lowest.
dtm.sum().sort_values().head()
dtm.mean().sort_values(ascending=False).head()
# ### TF-IDF scores
#
# How to find distinctive words in a corpus is a long-standing question in text analysis. Today, we'll learn one simple approach to this: TF-IDF. The idea behind words scores is to weight words not just by their frequency, but by their frequency in one document compared to their distribution across all documents. Words that are frequent, but are also used in every single document, will not be distinguising. We want to identify words that are unevenly distributed across the corpus.
#
# One of the most popular ways to weight words (beyond frequency counts) is `tf-idf score`. By offsetting the frequency of a word by its document frequency (the number of documents in which it appears) will in theory filter out common terms such as 'the', 'of', and 'and'.
#
# Traditionally, the *inverse document frequency* of word $j$ is calculated as:
#
# $idf_{j} = log\left(\frac{\#docs}{\#docs\,with\,j}\right)$
#
# and the *term freqency - inverse document frequency* is
#
# $tfidf_{ij} = f_{ij}\times{idf_j}$ where $f_{ij}$ is the number of occurences of word $j$ in document $i$.
#
# You can, and often should, normalize the word frequency:
#
# $tfidf_{ij} = \frac{f_{ij}}{\#words\,in\,doc\,i}\times{idf_{j}}$
#
# We can calculate this manually, but scikit-learn has a built-in function to do so. This function also uses log frequencies, so the numbers will not correspond excactly to the calculations above. We'll use the [scikit-learn calculation](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html), but a challenge for you: use Pandas to calculate this manually.
# ### TF-IDFVectorizer Function
#
# To do so, we simply do the same thing we did above with CountVectorizer, but instead we use the function TfidfVectorizer.
# +
from sklearn.feature_extraction.text import TfidfVectorizer
tfidfvec = TfidfVectorizer()
sparse_tfidf = tfidfvec.fit_transform(reviews['body_without_digits'])
sparse_tfidf
# -
tfidf = pd.DataFrame(sparse_tfidf.toarray(), columns=tfidfvec.get_feature_names(), index=reviews.index)
tfidf.head()
# Let's look at the 20 words with highest tf-idf weights.
tfidf.max().sort_values(ascending=False).head(20)
# Ok! We have successfully identified content words, without removing stop words.
# ### Identifying Distinctive Words
#
# What can we do with this? These scores are best used when you want to identify distinctive words for individual documents, or groups of documents, compared to other groups or the corpus as a whole. To illustrate this, let's compare three genres and identify the most distinctive words by genre.
#
# First we add in a column of genre.
tfidf['genre_'] = reviews['genre']
tfidf.head()
# Now lets compare the words with the highest tf-idf weight for each genre.
# +
rap = tfidf[tfidf['genre_']=='Rap']
indie = tfidf[tfidf['genre_']=='Indie']
jazz = tfidf[tfidf['genre_']=='Jazz']
rap.max(numeric_only=True).sort_values(ascending=False).head()
# -
indie.max(numeric_only=True).sort_values(ascending=False).head()
jazz.max(numeric_only=True).sort_values(ascending=False).head()
# There we go! A method of identifying distinctive words.
# ### Challenge - SOLUTION
#
# Instead of outputting the highest weighted words, output the lowest weighted words. How should we interpret these words?
jazz.max(numeric_only=True).sort_values().head()
# # Topic modeling <a id='topics'></a>
#
# The goal of topic models can be twofold: 1/ learning something about the topics themselves, i.e what the the ext is about 2/ reduce the dimensionality of text to represent a document as a weighted average of K topics instead of a vector of token counts over the whole vocabulary. In the latter case, topic modeling a way to treat text as any data in a more tractable way for any subsequent statistical analysis (linear/logistic regression, etc).
#
# There are many topic modeling algorithms, but we'll use LDA. This is a standard model to use. Again, the goal is not to learn everything you need to know about topic modeling. Instead, this will provide you some starter code to run a simple model, with the idea that you can use this base of knowledge to explore this further.
#
# We will run Latent Dirichlet Allocation, the most basic and the oldest version of topic modeling$^1$. We will run this in one big chunk of code. Our challenge: use our knowledge of scikit-learn that we gained above to walk through the code to understand what it is doing. Your challenge: figure out how to modify this code to work on your own data, and/or tweak the parameters to get better output.
#
# First, a bit of theory. LDA is a generative model - a model over the entire data generating process - in which a document is a mixture of topics and topics are probability distributions over tokens in the vocabulary. The (normalized) frequency of word $j$ in document $i$ can be written as:
# $q_{ij} = v_{i1}*\theta_{1j} + v_{i2}*\theta_{2j} + ... + v_{iK}*\theta_{Kj}$
# where K is the total number of topics, $\theta_{kj}$ is the probability that word $j$ shows up in topic $k$ and $v_{ik}$ is the weight assigned to topic $k$ in document $i$. The model treats $v$ and $\theta$ as generated from Dirichlet-distributed priors and can be estimated through Maximum Likelihood or Bayesian methods.
#
# Note: we will be using a different dataset for this technique. The music reviews in the above dataset are often short, one word or one sentence reviews. Topic modeling is not really appropriate for texts that are this short. Instead, we want texts that are longer and are composed of multiple topics each. For this exercise we will use a database of children's literature from the 19th century.
#
# The data were compiled by students in this course: http://english197s2015.pbworks.com/w/page/93127947/FrontPage
# Found here: http://dhresourcesforprojectbuilding.pbworks.com/w/page/69244469/Data%20Collections%20and%20Datasets#demo-corpora
#
# That page has additional corpora, for those interested in exploring text analysis further.
#
# $^1$ Reference: <NAME>., <NAME>, and <NAME> (2003). Latent Dirichlet allocation. Journal of Machine
# Learning Research 3, 993–1022.
# +
literature_fname = os.path.join(DATA_DIR, 'childrens_lit.csv.bz2')
df_lit = pd.read_csv(literature_fname, sep='\t', encoding = 'utf-8', compression = 'bz2', index_col=0)
#drop rows where the text is missing
df_lit = df_lit.dropna(subset=['text'])
df_lit.head()
# -
# Now we're ready to fit the model. This requires the use of CountVectorizer, which we've already used, and the scikit-learn function LatentDirichletAllocation.
#
# See [here](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.LatentDirichletAllocation.html) for more information about this function.
#
# First, we have to import it from sklearn.
from sklearn.decomposition import LatentDirichletAllocation
# In sklearn, the input to LDA is a DTM (with either counts or TF-IDF scores).
tfidf_vectorizer = TfidfVectorizer(max_df=0.80, min_df=50,
stop_words='english')
tfidf = tfidf_vectorizer.fit_transform(df_lit['text'])
tf_vectorizer = CountVectorizer(max_df=0.80, min_df=50,
stop_words='english'
)
tf = tf_vectorizer.fit_transform(df_lit['text'])
# This is where we fit the model.
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
lda = LatentDirichletAllocation(n_topics=10, max_iter=20, random_state=0)
lda = lda.fit(tf)
# This is a function to print out the top words for each topic in a pretty way. Don't worry too much about understanding every line of this code.
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
print("\nTopic #{}:".format(topic_idx))
print(" ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]]))
print()
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, 20)
# ### Challenge
#
# Modify the script above to:
# * increase the number of topics
# * increase the number of printed top words per topic
# * fit the model to the tf-idf matrix instead of the tf one
# ## Topic weights
#
# One thing we may want to do with the output is compare the prevalence of each topic across documents. A simple way to do this (but not memory efficient), is to merge the topic distribution back into the Pandas dataframe.
#
# First get the topic distribution array.
topic_dist = lda.transform(tf)
topic_dist
# Merge back with original dataframe
topic_dist_df = pd.DataFrame(topic_dist)
df_w_topics = topic_dist_df.join(df_lit)
df_w_topics
# Now we can chech the average weight of each topic across gender using `groupby`.
grouped = df_w_topics.groupby('author gender')
grouped[0].mean().sort_values(ascending=False)
# ## LDA as dimensionality reduction
#
# Now that we obtained a distribution of topic weights for each document, we can represent our corpus with a dense document-weight matrix as opposed to our initial sparse DTM. The weights can then replace tokens as features for any subsequent task (classification, prediction, etc). A simple example may consist in measuring cosine similarity between documents. For instance, which book is closest to the first book in our corpus? Let's use pairwise cosine similarity to find out.
#
# NB: cosine similarity measures an angle between two vectors, which provides a measure of distance robust to vectors of different lenghts (total number of tokens)
#
# First, let's turn the DTM into a readable dataframe.
dtm = pd.DataFrame(tf_vectorizer.fit_transform(df_lit['text']).toarray(), columns=tf_vectorizer.get_feature_names(), index = df_lit.index)
# Next let's import the cosine_similarity function from sklearn and print the cosine similarity between the first and second book or the first and third book.
from sklearn.metrics.pairwise import cosine_similarity
print("Cosine similarity between first and second book: " + str(cosine_similarity(dtm.iloc[0,:], dtm.iloc[1,:])))
print("Cosine similarity between first and third book: " + str(cosine_similarity(dtm.iloc[0,:], dtm.iloc[2,:])))
# What if we use the topic weights instead of word frequencies?
# +
dwm = df_w_topics.iloc[:,:10]
print("Cosine similarity between first and second book: " + str(cosine_similarity(dwm.iloc[0,:], dwm.iloc[1,:])))
print("Cosine similarity between first and third book: " + str(cosine_similarity(dwm.iloc[0,:], dwm.iloc[2,:])))
# -
# ### Challenge - SOLUTION
#
# Calculate the cosine similarity between the first book and all other books to identify the most similar one.
sim = cosine_similarity(dwm.iloc[0,:], dwm.iloc[1,:]) #cosine similarity with 2nd book
for i in range(2, len(dwm)):
sim = np.append(sim, cosine_similarity(dwm.iloc[0,:], dwm.iloc[i,:])) #append cosine similarity with i'th book
print("Max similarity: " + str(np.max(sim)))
print("Index of most similar book: " + str(np.argmax(sim)+1))
print("Title of most similar book: " + df_lit['title'][np.argmax(sim)+1])
# ### Further resources
#
# [This blog post](https://de.dariah.eu/tatom/feature_selection.html) goes through finding distinctive words using Python in more detail
#
# Paper: [Fightin’ Words: Lexical Feature Selection and Evaluation for Identifying the Content of Political Conflict](http://languagelog.ldc.upenn.edu/myl/Monroe.pdf), <NAME>, <NAME>, <NAME>
#
# [Topic modeling with Textacy](https://github.com/repmax/topic-model/blob/master/topic-modelling.ipynb)
|
day-2/02-unsupervised-solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch_cyclegan] *
# language: python
# name: conda-env-pytorch_cyclegan-py
# ---
# +
import os
import json
import tqdm
import argparse
from src.util import *
from src.train import *
from src.evaluate import *
# +
parser = argparse.ArgumentParser(description="TestLoop",
formatter_class=argparse.ArgumentDefaultsHelpFormatter)
parser.add_argument("--loop", default="stroll", choices=["stroll", "test"], type=str, dest="loop")
parser.add_argument("--lr", default=2e-4, type=float, dest="lr")
parser.add_argument("--batch_size", default=1, type=int, dest="batch_size")
parser.add_argument("--train_continue", default="on", choices=["on", "off"], type=str, dest="train_continue")
parser.add_argument("--num_epoch", default=100, type=int, dest="num_epoch")
parser.add_argument("--task", default="pose estimation", choices=["pose estimation"], type=str, dest="task")
parser.add_argument("--ny", default=256, type=int, dest="ny")
parser.add_argument("--nx", default=256, type=int, dest="nx")
parser.add_argument("--nch", default=3, type=int, dest="nch")
parser.add_argument("--nker", default=64, type=int, dest="nker")
parser.add_argument("--norm", default='inorm', type=str, dest="norm")
parser.add_argument("--network", default="PoseResNet", choices=["PoseResNet"], type=str, dest="network")
parser.add_argument("--resnet_depth", default=50, choices=[18, 34, 50, 101, 152], type=int, dest="resnet_depth")
parser.add_argument("--joint_weight", default=False, type=bool, dest="joint_weight")
parser.add_argument("--cuda", default="cuda", choices=["cuda", "cuda:0", "cuda:1"], type=str, dest="cuda")
parser.add_argument("--spec", default="all", type=str, dest="spec")
args = parser.parse_args(args=[])
vars = vars(args)
# -
cwd = os.getcwd()
datasets_dir = os.path.join(cwd, "datasets")
test_report_dir = os.path.join(cwd, "test_results")
test_design_list = os.listdir(datasets_dir)
eval_results = {}
for idx_d, design in enumerate(tqdm.tqdm(test_design_list)):
setups_dir = os.path.join(datasets_dir, design)
reports_dir = os.path.join(test_report_dir, design)
setups_list = os.listdir(setups_dir)
for idx_s, setup in enumerate(setups_list):
# Set arguments for evaluation
vars["mode"] = "test"
test_data_dir = os.path.join(setups_dir, setup)
vars["data_dir"] = test_data_dir
with open(os.path.join(test_data_dir, "test", "labels", "mpii_style.json"), "r", encoding="utf-8") as fread:
labels_dict = json.load(fread)
num_mark = len(labels_dict[0]["joints_vis"])
vars["num_mark"] = num_mark
args.ckpt_dir = os.path.join(reports_dir, setup, "checkpoint")
# Evaluate
evals = evaluate(args=args)
eval_results["%s" % design+"-"+setup] = evals
save_dir = os.path.join(test_report_dir, "evaluation_test_dataset.json")
with open(save_dir, "w", encoding = "UTF-8-SIG") as file:
json.dump(eval_results, file, ensure_ascii=False)
avg_acc_array = np.zeros((4, len(evals)))
for i, (key, result) in enumerate(tqdm.tqdm(eval_results.items())):
for j, snip in enumerate(result):
avg_acc = snip["avg_acc"]
avg_acc_array[i, j] = avg_acc
# +
import matplotlib.pyplot as plt
x = np.array(list(range(len(evals))))
unity_mean = np.mean(avg_acc_array[1,:])
gan_mean = np.mean(avg_acc_array[0,:])
plt.figure(figsize=(15, 5))
plt.title("3 labels, test data")
plt.plot(x, avg_acc_array[1,:], color="tab:blue", alpha=0.6, label="Unity")
plt.plot(x, avg_acc_array[0,:], color="tab:orange", alpha=0.6, label="GAN")
plt.axhline(y=unity_mean, color="tab:blue", alpha=1, linestyle="dotted", label=f"mean={unity_mean}")
plt.axhline(y=gan_mean, color="tab:orange", alpha=1, linestyle="dotted", label=f"mean={gan_mean}")
plt.xlabel('Data index')
plt.ylabel('Average accuracy')
plt.legend()
plt.show()
# +
difference = avg_acc_array[1,:] - avg_acc_array[0,:]
mean_diff = np.mean(difference)
plt.figure(figsize=(15, 5))
plt.title("3 labels, test data")
plt.plot(x, difference, color="tab:green", alpha=0.6, label="Unity - GAN")
plt.axhline(y=mean_diff, color="tab:green", alpha=1, linestyle="dotted", label=f"mean={mean_diff}")
plt.xlabel('Data index')
plt.ylabel('Mean accuracy difference')
plt.legend()
plt.show()
# +
unity_mean = np.mean(avg_acc_array[3,:])
gan_mean = np.mean(avg_acc_array[2,:])
plt.figure(figsize=(15, 5))
plt.title("4 labels, test data")
plt.plot(x, avg_acc_array[3,:], color="tab:blue", alpha=0.6, label="Unity")
plt.plot(x, avg_acc_array[2,:], color="tab:orange", alpha=0.6, label="GAN")
plt.axhline(y=unity_mean, color="tab:blue", alpha=1, linestyle="dotted", label=f"mean={unity_mean}")
plt.axhline(y=gan_mean, color="tab:orange", alpha=1, linestyle="dotted", label=f"mean={gan_mean}")
plt.xlabel('Data index')
plt.ylabel('Average accuracy')
plt.legend()
plt.show()
# +
difference = avg_acc_array[3,:] - avg_acc_array[2,:]
mean_diff = np.mean(difference)
plt.figure(figsize=(15, 5))
plt.title("4 labels, test data")
plt.plot(x, difference, color="tab:green", alpha=0.6, label="Unity - GAN")
plt.axhline(y=mean_diff, color="tab:green", alpha=1, linestyle="dotted", label=f"mean={mean_diff}")
plt.xlabel('Data index')
plt.ylabel('Mean accuracy difference')
plt.legend()
plt.show()
# -
|
evaluate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import librosa
import librosa.display
import scipy as sp
import IPython.display as ipd
import matplotlib.pyplot as plt
import numpy as np
# load audio file in the player
audio_path = "0000.wav"
ipd.Audio(audio_path)
# +
# load audio file
signal, sr = librosa.load(audio_path)
print(signal.max())
print(signal.min())
# signal += abs(signal.min())
# signal = (signal/signal.max())*2 -1
# print(signal.shape)
# print(signal.max())
# print(signal.min())
# +
# plot waveform
plt.figure(figsize=(18, 8))
#plt.plot(signal)
librosa.display.waveplot(signal, sr=sr, alpha=0.5)
plt.show()
# -
# derive spectrum using FT
ft = sp.fft.fft(signal)
print(ft.shape)
half_len = int(ft.shape[0]/2)
magnitude = librosa.power_to_db(np.absolute(ft)**2)
frequency = np.linspace(0, sr, len(magnitude))
# plot spectrum
plt.figure(figsize=(18, 8))
plt.plot(frequency[:], magnitude[:]) # magnitude spectrum
plt.xlabel("Frequency (Hz)")
plt.ylabel("Magnitude")
plt.show()
|
2_ftt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Supervised sentiment: overview of the Stanford Sentiment Treebank
# -
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2021"
# + [markdown] slideshow={"slide_type": "-"}
# ## Contents
#
# 1. [Overview of this unit](#Overview-of-this-unit)
# 1. [Set-up](#Set-up)
# 1. [Data readers](#Data-readers)
# 1. [Train split](#Train-split)
# 1. [Root-only formulation](#Root-only-formulation)
# 1. [Including subtrees](#Including-subtrees)
# 1. [Dev and test splits](#Dev-and-test-splits)
# 1. [Tokenization](#Tokenization)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview of this unit
#
# We have a few inter-related goals for this unit:
#
# * Provide a basic introduction to supervised learning in the context of a problem that has long been central to academic research and industry applications: __sentiment analysis__.
#
# * Explore and evaluate a diverse array of methods for modeling sentiment:
# * Hand-built feature functions with (mostly linear) classifiers
# * Dense feature representations derived from VSMs as we built them in the previous unit
# * Recurrent neural networks (RNNs)
#
# * Begin discussing and implementing responsible methods for __hyperparameter optimization__ and __classifier assessment and comparison__.
#
# The unit is built around the [Stanford Sentiment Treebank (SST)](http://nlp.stanford.edu/sentiment/), a widely-used resource for evaluating supervised NLU models, and one that provides rich linguistic representations.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Set-up
#
# * Make sure your environment includes all the requirements for [the cs224u repository](https://github.com/cgpotts/cs224u).
#
# * If you haven't already, download [the course data](http://web.stanford.edu/class/cs224u/data/data.tgz), unpack it, and place it in the directory containing the course repository – the same directory as this notebook. (If you want to put it somewhere else, change `SST_HOME` below.)
# +
from nltk.tokenize.treebank import TreebankWordDetokenizer
from nltk.tokenize.treebank import TreebankWordTokenizer
import os
import pandas as pd
import sst
# -
SST_HOME = os.path.join('data', 'sentiment')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data readers
#
# Our SST distribution is the ternary version of the problem (SST-3). It consists of train/dev/test files with the following columns:
#
# 1. `example_id`: a string with the format 'N-S' where N is the example number and S is the index for the subtree in example N. Both N and S are five-digit numbers with 0-padding.
# 2. `sentence`: a string giving the example sentence.
# 3. `label`: a string giving the label: `'positive'`, `'negative'`, or `'neutral'`. This value is derived from the original SST by mapping labels 0 and 1 to `'negative'`, label 2 to `'neutral'`, and labels 3 and 4 to `'positive'`.
# 4. `is_subtree`: the integer `1` if the example is a (proper) subtree, else `0`. This affects only the train file. Our dev and test splits contain no subtrees – full examples only – and hence `is_subtree` is always `0` for them.
# -
# ### Train split
# When reading in the train split, you have a few options.
# #### Root-only formulation
# The default will include only full examples and retain duplicate examples:
train_df = sst.train_reader(SST_HOME)
train_df.sample(3, random_state=1).to_dict(orient="records")
train_df.shape[0]
# This yields the following label distribution:
train_df.label.value_counts()
# You might want to remove the duplicate examples:
dup_train_df = sst.train_reader(SST_HOME, dedup=True)
dup_train_df.shape[0]
# This removes only ten examples for this setting so it is unlikely to be a significant choice.
# Our CSV-based distribution should make it easy to do basic analysis of the dataset to inform system development.
#
# Here's a look at the distribution of examples by length in characters:
_ = train_df.sentence.str.len().hist().set_ylabel("Length in characters")
# And by word count, assuming a very simple tokenization strategy:
# +
train_df['word_count'] = train_df.sentence.str.split().apply(len)
_ = train_df['word_count'].hist().set_ylabel("Length in words")
# -
_ = train_df.boxplot("word_count", by="label")
# #### Including subtrees
# Much of the special interest of the SST is that it includes labels, not just for full examples, but also for all the constituent words and phrases in those examples. You might also want to try training on this expanded dataset. It's much larger and so experiments will be more costly in terms of time and compute resources, but it could be worth it.
subtree_train_df = sst.train_reader(SST_HOME, include_subtrees=True)
subtree_train_df.shape[0]
subtree_train_df.head()
# +
subtree_train_df['word_count'] = subtree_train_df.sentence.str.split().apply(len)
_ = subtree_train_df['word_count'].hist().set_ylabel("Length in words")
# -
# In this setting, removing duplicates has a large effect, since many subtrees are repeated:
subtree_dedup_train_df = sst.train_reader(SST_HOME, include_subtrees=True, dedup=True)
subtree_dedup_train_df.shape
# Label distribution:
subtree_dedup_train_df.label.value_counts()
# ### Dev and test splits
# For the dev and test splits, we include only the root-level examples, and we do not deduplicate to remain aligned with the original paper. (The dev set has one repeated example, and the test set has none.)
dev_df = sst.dev_reader(SST_HOME)
dev_df.shape
# Label distribution:
dev_df.label.value_counts()
# There is an associated `sst.test_reader(SST_HOME)` with 2,210 (root-only) examples and no duplicates. As always in our field, you should use the test set only at the very end of your system development, and you should never, ever develop a system on the basis of test-set scores.
#
# In a similar vein, you should use the dev set only very sparingly. This will give you a clearer picture of how you will ultimately do on test; over-use of a dev set can lead to over-fitting on that particular dataset with a resulting loss of performance at test time.
#
# In the homework and associated bake-off for this course, we will introduce a second dev/test pair involving sentences about restaurants. The goal there is to have a fresh test set, and to push you to develop a system that works both for the SST movie domain and this new domain.
_ = dev_df.sentence.str.len().hist().set_ylabel("Length in characters")
# +
dev_df['word_count'] = dev_df.sentence.str.split().apply(len)
_ = dev_df['word_count'].hist().set_ylabel("Length in words")
# -
_ = dev_df.boxplot("word_count", by="label")
# ## Tokenization
# The SST began as a collection of sentences from [Rotten Tomatoes](https://www.rottentomatoes.com/) that were released as a corpus by [Pang and Lee 2004](https://doi.org/10.3115/1218955.1218990). The data were parsed as part of the SST project, and we are now releasing them in a flat format similar to what one sees in benchmarks like [GLUE](https://gluebenchmark.com). Along this journey, the sentences have acquired a tokenization scheme that is reminiscent of what one sees in standard [Penn Treenbank](https://catalog.ldc.upenn.edu/docs/LDC95T7/cl93.html) formats, with some additional quirks. This makes the tokens different in sigificant respects from what one sees in most standard English texts:
# +
ex = train_df.iloc[0].sentence
ex
# -
# One can address some of this using the NLTK `TreebankWordDetokenizer`:
detokenizer = TreebankWordDetokenizer()
def detokenize(s):
return detokenizer.detokenize(s.split())
detokenize(ex)
# As you can see, there is additional clean-up one could do, but this is a start.
# Another option would be to go in the reverse – for outside data, one could try to bring it into the SST format:
tokenizer = TreebankWordTokenizer()
def treebank_tokenize(s):
return tokenizer.tokenize(s)
treebank_tokenize("The Rock isn't the new ``Conan'' – he's this generation's Olivier!")
|
sst_01_overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://rhyme.com/assets/img/logo-dark.png" align="center">
# <h2 align="center"> Univariate Linear Regression </h2>
#
# ### Task 2: Load the Data and Libraries
# ---
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# %matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
plt.rcParams['figure.figsize'] = (12, 8)
data = pd.read_csv("food_truck_data.txt")
data.head()
data.info()
#
# ### Task 3: Visualize the Data
# ---
ax = sns.scatterplot(x='Population', y='Profit', data=data)
ax.set_title("Profit in $10000s vs City Population in 10000s");
#
# ### Task 4: Compute the Cost $J(\theta)$
# ---
# The objective of linear regression is to minimize the cost function
#
# $$J(\theta) = \frac{1}{2m} \sum_{i=1}^m (h_\theta(x^{(i)}) - y^{(i)} )^2$$
#
# where $h_{\theta}(x)$ is the hypothesis and given by the linear model
#
# $$h_{\theta}(x) = \theta^Tx = \theta_0 + \theta_1x_1$$
def cost_function(X, y, theta):
m = len(y)
y_pred = X.dot(theta)
error = (y_pred - y) ** 2
return 1 / (2 * m) * np.sum(error)
# +
m = data.Population.values.size
# add another dimension to accomodate the intercept term and set it to all ones
X = np.append(np.ones((m, 1)), data.Population.values.reshape(m, 1), axis=1)
y = data.Profit.values.reshape(m, 1)
theta = np.zeros((2,1))
cost_function(X, y, theta)
# -
#
# ### Task 5: Gradient Descent
# ---
# Minimize the cost function $J(\theta)$ by updating the below equation and repeat unitil convergence
#
# $\theta_j := \theta_j - \alpha \frac{1}{m} \sum_{i=1}^m (h_{\theta}(x^{(i)}) - y^{(i)})x_j^{(i)}$ (simultaneously update $\theta_j$ for all $j$).
def gradient_descent(X, y, theta, alpha, iterations):
m = len(y)
costs = []
for i in range(iterations):
y_pred = X.dot(theta)
error = np.dot(X.transpose(), (y_pred - y))
theta -= alpha * 1/m * error
costs.append(cost_function(X, y, theta))
return theta, costs
# +
theta, costs = gradient_descent(X, y, theta, alpha=0.01, iterations=1000)
print("h(x) = {} + {}x1".format(str(round(theta[0, 0], 2)),
str(round(theta[1, 0], 2))))
# -
costs[999]
# ### Task 6: Visualising the Cost Function $J(\theta)$
# ---
from mpl_toolkits.mplot3d import Axes3D
# +
theta_0 = np.linspace(-10,10,100)
theta_1 = np.linspace(-1,4,100)
cost_values = np.zeros((len(theta_0), len(theta_1)))
for i in range(len(theta_0)):
for j in range(len(theta_1)):
t = np.array([theta_0[i], theta_1[j]])
cost_values[i, j] = cost_function(X, y, t)
# +
fig = plt.figure(figsize = (12, 8))
ax = fig.gca(projection = '3d')
surf = ax.plot_surface(theta_0, theta_1, cost_values, cmap = "viridis", linewidth = 0.2)
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.xlabel("$\Theta_0$")
plt.ylabel("$\Theta_1$")
ax.set_zlabel("$J(\Theta)$")
ax.set_title("Cost Surface")
ax.view_init(30,330)
plt.show()
# -
#
# ### Task 7: Plotting the Convergence
# ---
# Plot $J(\theta)$ against the number of iterations of gradient descent:
plt.plot(costs)
plt.xlabel("Iterations")
plt.ylabel("$J(\Theta)$")
plt.title("Values of Cost Function over iterations of Gradient Descent");
#
# ### Task 8: Training Data with Linear Regression Fit
# ---
theta.shape
theta
# +
theta = np.squeeze(theta)
sns.scatterplot(x = "Population", y= "Profit", data = data)
x_value=[x for x in range(5, 25)]
y_value=[(x * theta[1] + theta[0]) for x in x_value]
sns.lineplot(x_value,y_value)
plt.xlabel("Population in 10000s")
plt.ylabel("Profit in $10,000s")
plt.title("Linear Regression Fit");
# -
# ### Task 9: Inference using the optimized $\theta$ values
# ---
# $h_\theta(x) = \theta^Tx$
def predict(x, theta):
y_pred = np.dot(theta.transpose(), x)
return y_pred
y_pred_1 = predict(np.array([1, 4]),theta) * 10000
print("For a population of 40,000, the model predicts a profit of $" + str(round(y_pred_1, 0)))
y_pred_2 = predict(np.array([1, 8.3]), theta) * 10000
print("For a population of 83,000, the model predicts a profit of $"+str(round(y_pred_2, 0)))
|
Univariate Linear Regression_Completed.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # Demo ARMA node: filtering
using Revise
using ProgressMeter
using LinearAlgebra
using Plots
pyplot();
using ForneyLab
import ForneyLab: unsafeMean, unsafeCov
using ARMA
# ## Data generation
# +
# Parameters
θ_true = [0.7, -.2]
η_true = [1.0, 1.0]
τ_true = 1e4
# Orders
M1 = length(θ_true)
M2 = length(η_true)
M = M1 + M2
# Transient period
tt = 10
# Time horizon
T = 300
# Observation array
output = zeros(T+tt,)
errors = zeros(T+tt,)
# First 2 outputs
output[1] = 0.0
output[2] = 0.0
for k = 1:T+tt
# Errors
errors[k] = sqrt(inv(τ_true))*randn(1)[1]
if k > max(M1,M2)
# Autoregressive moving average function plus noise
output[k] = θ_true'*output[k-1:-1:k-M1] + η_true'*errors[k-1:-1:k-M2] + errors[k]
end
end
# -
plot(tt:T, output[tt:T], color="black", label="", xlabel="time (k)", ylabel="signal", size=(800,300))
# ## Model specification
# +
graph = FactorGraph()
# Observed variables
@RV z_kmin1; placeholder(z_kmin1, :z_kmin1, dims=(M1,))
@RV r_kmin1; placeholder(r_kmin1, :r_kmin1, dims=(M2,))
# Time-invariant parameters
@RV θ ~ GaussianMeanVariance(placeholder(:m_θ, dims=(M,)), placeholder(:v_θ, dims=(M,M)))
@RV τ ~ Gamma(placeholder(:a_τ), placeholder(:b_τ))
# Likelihood
@RV y_k ~ AutoRegressiveMovingAverage(θ, z_kmin1, r_kmin1, τ)
placeholder(y_k, :y_k)
ForneyLab.draw(graph)
# -
q = PosteriorFactorization(θ, τ, ids=[:θ :τ])
algorithm = messagePassingAlgorithm([θ; τ], q)
source_code = algorithmSourceCode(algorithm)
eval(Meta.parse(source_code));
# ## Inference
# +
# Preallocate parameter arrays
params_θ = (zeros(T-tt,M), zeros(T-tt,M,M))
params_τ = (zeros(T-tt,1), zeros(T-tt,1))
# Initialize priors
θ_k = (zeros(M,), 10 .*Matrix{Float64}(I, M,M))
τ_k = (1e4, 1e1)
marginals = Dict(:θ => ProbabilityDistribution(Multivariate, GaussianMeanVariance, m=θ_k[1], v=θ_k[2]),
:τ => ProbabilityDistribution(Univariate, Gamma, a=τ_k[1], b=τ_k[2]))
# Keep track of residuals
predictions = (zeros(T,), zeros(T,))
residuals = zeros(T,)
# Inference
@showprogress for (jj,k) in enumerate(tt+1:T)
# State vector
x_k = [output[k-1:-1:k-M1]; residuals[k-1:-1:k-M2]]
# Posterior predictive
predictions[1][k] = θ_k[1]'*x_k
predictions[2][k] = x_k'*θ_k[2]'*x_k# + inv(τ_k[1]/τ_k[2])
# Compute residual
residuals[k] = output[k] - predictions[1][k]
# Set data
data = Dict(:y_k => output[k],
:z_kmin1 => output[k-1:-1:k-M1],
:r_kmin1 => residuals[k-1:-1:k-M2],
:m_θ => θ_k[1],
:v_θ => θ_k[2],
:a_τ => τ_k[1],
:b_τ => τ_k[2])
# Iterate updates
for n = 1:10
stepθ!(data, marginals)
stepτ!(data, marginals)
end
# Update params
θ_k = (unsafeMean(marginals[:θ]), unsafeCov(marginals[:θ]))
τ_k = (marginals[:τ].params[:a], marginals[:τ].params[:b])
# Store params
params_θ[1][jj,:] = θ_k[1]
params_θ[2][jj,:,:] = θ_k[2]
params_τ[1][jj] = τ_k[1]
params_τ[2][jj] = τ_k[2]
end
# -
# ## Visualization
# Fit to past data
sd_pred = sqrt.(predictions[2])
scatter(tt:T, output[tt:T], color="black", label="data", xlabel="time (k)", ylabel="signal", size=(800,300))
plot!(tt:T, predictions[1][tt:T], ribbon=[sd_pred[tt:T], sd_pred[tt:T]], color="blue", label="filter")
plot(tt+1:T, params_θ[1][:,1], ribbon=[sqrt.(params_θ[2][:,1,1]) sqrt.(params_θ[2][:,1,1])], xlabel="time (k)", label="θ1", size=(800,300))
plot!(tt+1:T, params_θ[1][:,2], ribbon=[sqrt.(params_θ[2][:,2,2]) sqrt.(params_θ[2][:,2,2])], label="θ2")
plot!(tt+1:T, params_θ[1][:,3], ribbon=[sqrt.(params_θ[2][:,3,3]) sqrt.(params_θ[2][:,3,3])], label="θ2")
# +
mτ = params_τ[1] ./ params_τ[2]
vτ = params_τ[1] ./ params_τ[2].^2
plot(tt+1:T, mτ, ribbon=[vτ vτ], color="purple", xlabel="time (k)", label="τ", size=(800,300))
# -
|
demo/demo_filtering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Predict Category of News Using Labeled NYT Data
#
#
#
# - Take out the rows that are in category :- 'Gen Soft', 'Misc', 'Week in Review', 'Magazine', 'Home Desk', 'Cars', 'Living', 'Personal Finance'
# - Take out the rows that are shorter than 150 characters
# - Stratified random sampling 7000 rows for each categories (17 categories)
# - Split 80% for trained dataset and 20% for test dataset
# - Feature extraction (unigram/bigram/trigram) by take out the n-grams that are occurrence less than 0.05% or greater than 50% of trained dataset
# - Fit the model using LinearSVC
import pandas as pd
df = pd.read_csv('../../../nyt_data/nyt_recode_clean.csv.bz2', nrows=10)
select_cols = "categories, OnlineMSoft, NewsDeskSoft, Publication.Date, Publication.Year, Section, Body, Lead.Paragraph, Headline, Online.Headline, Online.Lead.Paragraph, Url, Online.Section, ID"
select_cols = [c.strip() for c in select_cols.split(',')]
for c in select_cols:
if c not in df.columns:
print(c)
df = pd.read_csv('../../../nyt_data/nyt_recode_clean.csv.bz2', usecols=select_cols)
df.dropna(subset=['Body'], inplace=True)
df
df.groupby('categories').size()
# ### Let all other categories ==> Other
df.loc[df.categories.isnull() | df.categories.isin(['Gen Soft', 'Misc', 'Week in Review', 'Magazine', 'Home Desk', 'Cars', 'Living', 'Personal Finance']), 'categories'] = 'Other'
df.groupby('categories').size()
pd.set_option('max_colwidth', 120)
df[df.Body.str.len() < 150][['Body', 'Url']]
# ### Take out the Body text shorter than 150 characters
df = df[df.Body.str.len() > 150]
df.groupby(['categories']).size()
# !pip install nltk
# +
import time
import re
import string
import numpy as np
import nltk
from nltk import word_tokenize
from nltk.stem.porter import PorterStemmer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.metrics import classification_report, accuracy_score, f1_score
from sklearn.model_selection import train_test_split
from sklearn.externals import joblib
stemmer = PorterStemmer()
def stem_tokens(tokens, stemmer):
stemmed = []
for item in tokens:
stemmed.append(stemmer.stem(item))
return stemmed
def tokenize_with_punc(text):
tokens = nltk.word_tokenize(text)
stems = stem_tokens(tokens, stemmer)
return stems
def tokenize(text):
text = "".join([ch for ch in text if ch not in string.punctuation])
tokens = nltk.word_tokenize(text)
stems = stem_tokens(tokens, stemmer)
return stems
with open('../../../roberts_rules/all_text.txt', 'rt') as f:
text = f.read()
text = re.sub(r'\d+', '', text)
vect = CountVectorizer(tokenizer=tokenize, stop_words='english', ngram_range=(2, 3))
vect.fit([text])
roberts_rules = set(vect.get_feature_names())
def most_informative_feature_for_class(vectorizer, classifier, classlabel, n=10):
labelid = list(classifier.classes_).index(classlabel)
feature_names = vectorizer.get_feature_names()
topn = sorted(zip(classifier.coef_[labelid], feature_names))[-n:]
for coef, feat in topn:
print(classlabel, feat, coef)
def most_informative_feature_for_class_svm(vectorizer, classifier, n=10):
labelid = 3 # this is the coef we're interested in.
feature_names = vectorizer.get_feature_names()
svm_coef = classifier.coef_.toarray()
topn = sorted(zip(svm_coef[labelid], feature_names))[-n:]
for coef, feat in topn:
print(feat, coef)
def print_top10(vectorizer, clf, class_labels):
"""Prints features with the highest coefficient values, per class"""
feature_names = vectorizer.get_feature_names()
for i, class_label in enumerate(class_labels):
top10 = np.argsort(clf.coef_[i])[-10:]
print("%s: %s" % (class_label,
" | ".join(feature_names[j] for j in top10)))
def get_top_features(vectorizer, clf, class_labels, n=20):
"""Prints features with the highest coefficient values, per class"""
feature_names = vectorizer.get_feature_names()
top_features = {}
for i, class_label in enumerate(class_labels):
topN = np.argsort(clf.coef_[i])[-n:]
top_features[class_label] = [feature_names[j] for j in topN][::-1]
return top_features
def show_most_informative_features(vectorizer, clf, n=20):
feature_names = vectorizer.get_feature_names()
coefs_with_fns = sorted(zip(clf.coef_[0], feature_names))
top = zip(coefs_with_fns[:n], coefs_with_fns[:-(n + 1):-1])
for (coef_1, fn_1), (coef_2, fn_2) in top:
print("\t%.4f\t%-20s\t\t%.4f\t%-20s" % (coef_1, fn_1, coef_2, fn_2))
def get_most_informative_features(vectorizer, clf, n=20):
feature_names = vectorizer.get_feature_names()
coefs_with_fns = sorted(zip(clf.coef_[0], feature_names))
top_a = coefs_with_fns[:n]
top_b = coefs_with_fns[:-(n + 1):-1]
return top_a, top_b
# -
# ### Stratified Sampling
#SAMPLE_SIZE_PER_CAT = 4000 ==> 0.85
#SAMPLE_SIZE_PER_CAT = 6000 ==> 0.86
SAMPLE_SIZE_PER_CAT = 7000
sdf = pd.DataFrame()
for c in df.categories.unique():
sdf = sdf.append(df[df.categories == c].sample(SAMPLE_SIZE_PER_CAT, random_state=21, replace=True))
sdf.groupby(['categories']).size()
sdf['soft_news'] = 0
sdf.loc[sdf.categories.isin(['Arts', 'Books', 'Classifieds', 'Dining', 'Leisure', 'Obits', 'Other', 'Real Estate', 'Style', 'Travel']), 'soft_news'] = 1
sdf
sdf[['ID', 'Body', 'soft_news']].to_csv('nyt_sample_soft_news_7k.csv', index=False)
# X = sdf.Body
# y = sdf.categories
#
# X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=21, stratify=y)
# +
X = sdf[['ID', 'Body']]
y = sdf.soft_news
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=21, stratify=y)
# -
X_test_data = X_test.copy()
X_train = X_train.Body
X_test = X_test.Body
# +
import re
def custom_tokenizer(doc):
doc = re.sub('\d+', '[NUM]', doc)
return doc.split()
# -
vect = CountVectorizer(ngram_range=(1, 3), min_df=0.0005, max_df=0.5, tokenizer=custom_tokenizer, max_features=20000)
#vect = CountVectorizer(ngram_range=(1, 3), min_df=10, max_df=0.3, tokenizer=custom_tokenizer, max_features=10000)
#vect = CountVectorizer(ngram_range=(1, 3), min_df=200, max_df=0.3, tokenizer=custom_tokenizer)
# %%time
X_train = vect.fit_transform(X_train)
transformer = TfidfTransformer()
X_train = transformer.fit_transform(X_train)
len(vect.vocabulary_)
# %%time
X_test = vect.transform(X_test)
transformer = TfidfTransformer()
X_test = transformer.fit_transform(X_test)
len(vect.vocabulary_)
# +
# %%time
from sklearn.svm import LinearSVC
from sklearn.calibration import CalibratedClassifierCV
est = LinearSVC(penalty='l1', dual=False, tol=1e-3)
# Calibrated with isotonic calibration
clf = CalibratedClassifierCV(est, cv=2, method='isotonic')
t0 = time.time()
clf.fit(X_train, y_train)
t1 = time.time()
y_pred = clf.predict(X_test)
t2 = time.time()
time_clf_train = t1-t0
time_clf_predict = t2-t1
print("Results for classifier")
print("Training time: %fs; Prediction time: %fs" % (time_clf_train, time_clf_predict))
print(classification_report(y_test, y_pred))
# -
f1_score(y_test, y_pred, average='macro')
len(vect.vocabulary_)
clf.classes_
show_most_informative_features(vect, clf.calibrated_classifiers_[0].base_estimator)
get_most_informative_features(vect, clf.calibrated_classifiers_[0].base_estimator)
vect.stop_words_ = None
# %%time
joblib.dump(vect, "../data/us_model/nyt_us_soft_news_vectorizer.joblib", compress=3)
joblib.dump(clf, "../data/us_model/nyt_us_soft_news_classifier.joblib", compress=3)
# +
y_test_df = pd.DataFrame(y_test)
y_test_df.columns = ['true_value']
y_test_df.reset_index(drop=True, inplace=True)
y_test_df['pred_value'] = clf.predict(X_test)
if hasattr(clf, "predict_proba"):
prob = clf.predict_proba(X_test)
else: # use decision function
prob = clf.decision_function(X_test)
prob = \
(prob - prob.min()) / (prob.max() - prob.min())
prob_df = pd.DataFrame(prob)
columns = []
for c in clf.classes_:
columns.append(c)
prob_df.columns = columns
result_df = pd.concat([X_test_data.reset_index(drop=True), y_test_df, prob_df], axis=1)
result_df.to_csv('./tests/us_soft_news_test_prediction_other_calibrated+text.csv', index=False)
result_df
# -
result_df[result_df.pred_value!=result_df.true_value].to_csv('./reports/nyt_soft_news_test_pred_misclass.csv', index=False)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from pandas_confusion import ConfusionMatrix
y_true = y_test.reset_index(drop=True)
confusion_matrix = ConfusionMatrix(y_true, y_pred)
print("Confusion matrix:\n%s" % confusion_matrix)
# -
confusion_matrix.plot()
from sklearn.metrics import confusion_matrix
conf = confusion_matrix(y_true, y_pred)
conf_df = pd.DataFrame(conf)
conf_df.columns = clf.classes_
conf_df.index = clf.classes_
conf_df.to_csv('./reports/us_soft_news_test_confusion_matrix_other_calibrated.csv', index_label="actual \ predicted")
print(conf_df)
|
notnews/models/us_not_news_soft_news.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sofiesoltani/awesome-public-datasets/blob/master/Office_TV_Series.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="iAVb92H8S3tS"
# # 1. Welcome!
# <p><img src="https://assets.datacamp.com/production/project_1170/img/office_cast.jpeg" alt="Markdown">.</p>
# <p><strong>The Office!</strong> What started as a British mockumentary series about office culture in 2001 has since spawned ten other variants across the world, including an Israeli version (2010-13), a Hindi version (2019-), and even a French Canadian variant (2006-2007). Of all these iterations (including the original), the American series has been the longest-running, spanning 201 episodes over nine seasons.</p>
# <p>In this notebook, we will take a look at a dataset of The Office episodes, and try to understand how the popularity and quality of the series varied over time. To do so, we will use the following dataset: <code>datasets/office_episodes.csv</code>, which was downloaded from Kaggle <a href="https://www.kaggle.com/nehaprabhavalkar/the-office-dataset">here</a>.</p>
# <p>This dataset contains information on a variety of characteristics of each episode. In detail, these are:
# <br></p>
# <div style="background-color: #efebe4; color: #05192d; text-align:left; vertical-align: middle; padding: 15px 25px 15px 25px; line-height: 1.6;">
# <div style="font-size:20px"><b>datasets/office_episodes.csv</b></div>
# <ul>
# <li><b>episode_number:</b> Canonical episode number.</li>
# <li><b>season:</b> Season in which the episode appeared.</li>
# <li><b>episode_title:</b> Title of the episode.</li>
# <li><b>description:</b> Description of the episode.</li>
# <li><b>ratings:</b> Average IMDB rating.</li>
# <li><b>votes:</b> Number of votes.</li>
# <li><b>viewership_mil:</b> Number of US viewers in millions.</li>
# <li><b>duration:</b> Duration in number of minutes.</li>
# <li><b>release_date:</b> Airdate.</li>
# <li><b>guest_stars:</b> Guest stars in the episode (if any).</li>
# <li><b>director:</b> Director of the episode.</li>
# <li><b>writers:</b> Writers of the episode.</li>
# <li><b>has_guests:</b> True/False column for whether the episode contained guest stars.</li>
# <li><b>scaled_ratings:</b> The ratings scaled from 0 (worst-reviewed) to 1 (best-reviewed).</li>
# </ul>
# </div>
#
# ---
#
#
# + id="bXxhnDIDlZ9w"
import pandas as pd
data = pd.read_csv('the_office_series.csv', parse_dates=['Date'])
Office_df = data.rename(columns={'Unnamed: 0': 'episode_number'})
Office_df.info()
Office_df['Ratings'].describe()[['25%', '50%', '75%']]
Office_df['GuestStars_has']=Office_df['GuestStars'].notnull()
Office_df.head()
# + id="Z03VRy-rqlaP"
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [11, 7]
# + id="cBW_GWTcAWd1" colab={"base_uri": "https://localhost:8080/"} outputId="06f74575-c70c-4283-ec83-7550894b2063"
cols = []
for ind, row in Office_df.iterrows():
if row['Ratings'] < 7.8:
cols.append('red')
elif row['Ratings'] < 8.2:
cols.append('orange')
elif row['Ratings'] < 8.6:
cols.append('lightgreen')
else:
cols.append('darkgreen')
sizes = []
for ind, row in Office_df.iterrows():
if row['GuestStars_has']==False:
sizes.append(25)
else:
sizes.append(250)
Office_df['colors'] = cols
Office_df['sizes'] = sizes
Office_df.info()
# + id="q9J6bqTxUkvb"
office_df_star=Office_df[Office_df['GuestStars_has']==True]
office_df_nostar=Office_df[Office_df['GuestStars_has']==False]
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="dmdl7FcGxg1R" outputId="1be60b23-ee0e-4afd-bf08-a06fa98ee49d"
fig = plt.figure()
plt.style.use('fivethirtyeight')
plt.scatter(x=office_df_nostar['Date'], y=office_df_nostar['Viewership'], c=office_df_nostar['colors'], s=office_df_nostar['sizes'])
plt.scatter(x=office_df_star['Date'], y=office_df_star['Viewership'], c=office_df_star['colors'], s=office_df_star['sizes'], marker='*')
plt.title("Popularity, Quality, and Guest Appearances on the Office")
plt.xlabel("Release Year")
plt.ylabel("Viewership (Millions)")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="-mzlXEiQXN_Q" outputId="cf13314a-1514-4a7f-c9a4-ea93eab8899f"
Office_df[Office_df['Viewership']==Office_df['Viewership'].max()]['GuestStars']
|
Office_TV_Series.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Linear Magnetic Inversion
# **Objective:**
#
# In this tutorial we will create a simple magnetic problem from scratch using the SimPEG framework.
#
# We are using the integral form of the magnetostatic problem. In the absence of free-currents or changing magnetic field, magnetic material can give rise to a secondary magnetic field according to:
#
# $$\vec b = \frac{\mu_0}{4\pi} \int_{V} \vec M \cdot \nabla \nabla \left(\frac{1}{r}\right) \; dV $$
#
# Where $\mu_0$ is the magnetic permealitity of free-space, $\vec M$ is the magnetization per unit volume and $r$ defines the distance between the observed field $\vec b$ and the magnetized object. Assuming a purely induced response, the strenght of magnetization can be written as:
#
# $$ \vec M = \mu_0 \kappa \vec H_0 $$
#
# where $\vec H$ is an external inducing magnetic field, and $\kappa$ the magnetic susceptibility of matter.
# As derived by Sharma 1966, the integral can be evaluated for rectangular prisms such that:
#
# $$ \vec b(P) = \mathbf{T} \cdot \vec H_0 \; \kappa $$
#
# Where the tensor matrix $\bf{T}$ relates the three components of magnetization $\vec M$ to the components of the field $\vec b$:
#
# $$\mathbf{T} =
# \begin{pmatrix}
# T_{xx} & T_{xy} & T_{xz} \\
# T_{yx} & T_{yy} & T_{yz} \\
# T_{zx} & T_{zy} & T_{zz}
# \end{pmatrix} $$
#
# In general, we discretize the earth into a collection of cells, each contributing to the magnetic data such that:
#
# $$\vec b(P) = \sum_{j=1}^{nc} \mathbf{T}_j \cdot \vec H_0 \; \kappa_j$$
#
# giving rise to a linear problem.
#
# The remaining of this notebook goes through all the important components of a 3D magnetic experiment. From mesh creation, topography, data and inverse problem.
#
# Enjoy.
#
from SimPEG import Mesh
from SimPEG.Utils import mkvc, surface2ind_topo
from SimPEG import Maps
from SimPEG import Regularization
from SimPEG import DataMisfit
from SimPEG import Optimization
from SimPEG import InvProblem
from SimPEG import Directives
from SimPEG import Inversion
from SimPEG import PF
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# First we need to define the direction of the inducing field
# As a simple case, we pick a vertical inducing field of magnitude 50,000nT.
# From old convention, field orientation is given as an azimuth from North
# (positive clockwise) and dip from the horizontal (positive downward).
H0 = (60000.,90.,0.)
# Create a mesh
dx = 5.
hxind = [(dx,5,-1.3), (dx, 10), (dx,5,1.3)]
hyind = [(dx,5,-1.3), (dx, 10), (dx,5,1.3)]
hzind = [(dx,5,-1.3),(dx, 10)]
mesh = Mesh.TensorMesh([hxind, hyind, hzind], 'CCC')
# Get index of the center
midx = int(mesh.nCx/2)
midy = int(mesh.nCy/2)
# Lets create a simple Gaussian topo and set the active cells
[xx,yy] = np.meshgrid(mesh.vectorNx,mesh.vectorNy)
zz = -np.exp( ( xx**2 + yy**2 )/ 75**2 ) + mesh.vectorNz[-1]
topo = np.c_[mkvc(xx),mkvc(yy),mkvc(zz)] # We would usually load a topofile
actv = surface2ind_topo(mesh,topo,'N') # Go from topo to actv cells
actv = np.asarray([inds for inds, elem in enumerate(actv, 1) if elem], dtype = int) - 1
#nC = mesh.nC
#actv = np.asarray(range(mesh.nC))
# Create active map to go from reduce space to full
actvMap = Maps.InjectActiveCells(mesh, actv, -100)
nC = len(actv)
# Create and array of observation points
xr = np.linspace(-20., 20., 20)
yr = np.linspace(-20., 20., 20)
X, Y = np.meshgrid(xr, yr)
# Let just put the observation above the topo
Z = -np.exp( ( X**2 + Y**2 )/ 75**2 ) + mesh.vectorNz[-1] + 5.
#Z = np.ones(shape(X)) * mesh.vectorCCz[-1]
# Create a MAGsurvey
rxLoc = np.c_[mkvc(X.T), mkvc(Y.T), mkvc(Z.T)]
rxLoc = PF.BaseMag.RxObs(rxLoc)
srcField = PF.BaseMag.SrcField([rxLoc],param = H0)
survey = PF.BaseMag.LinearSurvey(srcField)
# -
# Now that we have all our spatial components, we can create our linear system. For a single location and single component of the data, the system would look like this:
#
# $$ b_x =
# \begin{bmatrix}
# T_{xx}^1 &... &T_{xx}^{nc} & T_{xy}^1 & ... & T_{xy}^{nc} & T_{xz}^1 & ... & T_{xz}^{nc}\\
# \end{bmatrix}
# \begin{bmatrix}
# \mathbf{M}_x \\ \mathbf{M}_y \\ \mathbf{M}_z
# \end{bmatrix} \\ $$
#
# where each of $T_{xx},\;T_{xy},\;T_{xz}$ are [nc x 1] long. For the $y$ and $z$ component, we need the two other rows of the tensor $\mathbf{T}$.
# In our simple induced case, the magnetization direction $\mathbf{M_x,\;M_y\;,Mz}$ are known and assumed to be constant everywhere, so we can reduce the size of the system such that:
#
# $$ \vec{\mathbf{d}}_{\text{pred}} = (\mathbf{T\cdot M})\; \kappa$$
#
#
#
# In most geophysical surveys, we are not collecting all three components, but rather the magnitude of the field, or $Total\;Magnetic\;Intensity$ (TMI) data.
# Because the inducing field is really large, we will assume that the anomalous fields are parallel to $H_0$:
#
# $$ d^{TMI} = \hat H_0 \cdot \vec d$$
#
# We then end up with a much smaller system:
#
# $$ d^{TMI} = \mathbf{F\; \kappa}$$
#
# where $\mathbf{F} \in \mathbb{R}^{nd \times nc}$ is our $forward$ operator.
# +
# We can now create a susceptibility model and generate data
# Lets start with a simple block in half-space
model = np.zeros((mesh.nCx,mesh.nCy,mesh.nCz))
model[(midx-2):(midx+2),(midy-2):(midy+2),-6:-2] = 0.02
model = mkvc(model)
model = model[actv]
# Create active map to go from reduce set to full
actvMap = Maps.InjectActiveCells(mesh, actv, -100)
# Creat reduced identity map
idenMap = Maps.IdentityMap(nP = nC)
# Create the forward model operator
prob = PF.Magnetics.MagneticIntegral(mesh, chiMap=idenMap, actInd=actv)
# Pair the survey and problem
survey.pair(prob)
# Compute linear forward operator and compute some data
d = prob.fields(model)
# +
# Plot the model
m_true = actvMap * model
m_true[m_true==-100] = np.nan
plt.figure()
ax = plt.subplot(212)
mesh.plotSlice(m_true, ax = ax, normal = 'Y', ind=midy, grid=True, clim = (0., model.max()), pcolorOpts={'cmap':'viridis'})
plt.title('A simple block model.')
plt.xlabel('x'); plt.ylabel('z')
plt.gca().set_aspect('equal', adjustable='box')
# We can now generate data
data = d + np.random.randn(len(d)) # We add some random Gaussian noise (1nT)
wd = np.ones(len(data))*1. # Assign flat uncertainties
plt.subplot(221)
plt.imshow(d.reshape(X.shape), extent=[xr.min(), xr.max(), yr.min(), yr.max()])
plt.title('True data.')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.subplot(222)
plt.imshow(data.reshape(X.shape), extent=[xr.min(), xr.max(), yr.min(), yr.max()])
plt.title('Data + Noise')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.tight_layout()
# +
# Create distance weights from our linera forward operator
wr = np.sum(prob.G**2.,axis=0)**0.5
wr = ( wr/np.max(wr) )
wr_FULL = actvMap * wr
wr_FULL[wr_FULL==-100] = np.nan
plt.figure()
ax = plt.subplot()
mesh.plotSlice(wr_FULL, ax = ax, normal = 'Y', ind=midx, grid=True, clim = (0, wr.max()),pcolorOpts={'cmap':'viridis'})
plt.title('Distance weighting')
plt.xlabel('x');plt.ylabel('z')
plt.gca().set_aspect('equal', adjustable='box')
# -
# Once we have our problem, we can use the inversion tools in SimPEG to run our inversion:
# +
#survey.makeSyntheticData(data, std=0.01)
survey.dobs= data
survey.std = wd
survey.mtrue = model
# Create a regularization
reg = Regularization.Sparse(mesh, indActive=actv, mapping=idenMap)
reg.cell_weights = wr
reg.norms = [0, 1, 1, 1]
reg.eps_p = 1e-3
reg.eps_1 = 1e-3
dmis = DataMisfit.l2_DataMisfit(survey)
dmis.W = 1/wd
# Add directives to the inversion
opt = Optimization.ProjectedGNCG(maxIter=100 ,lower=0.,upper=1., maxIterLS = 20, maxIterCG= 10, tolCG = 1e-3)
invProb = InvProblem.BaseInvProblem(dmis, reg, opt)
betaest = Directives.BetaEstimate_ByEig()
# Here is where the norms are applied
# Use pick a treshold parameter empirically based on the distribution of model
# parameters (run last cell to see the histogram before and after IRLS)
IRLS = Directives.Update_IRLS(f_min_change = 1e-3, minGNiter=3)
update_Jacobi = Directives.Update_lin_PreCond()
inv = Inversion.BaseInversion(invProb, directiveList=[betaest, IRLS, update_Jacobi])
m0 = np.ones(nC)*1e-4
# -
mrec = inv.run(m0)
# Inversion has converged. We can plot sections through the model.
# +
# Here is the recovered susceptibility model
ypanel = midx
zpanel = -4
m_l2 = actvMap * IRLS.l2model
m_l2[m_l2==-100] = np.nan
m_lp = actvMap * mrec
m_lp[m_lp==-100] = np.nan
m_true = actvMap * model
m_true[m_true==-100] = np.nan
plt.figure()
#Plot L2 model
ax = plt.subplot(231)
mesh.plotSlice(m_l2, ax = ax, normal = 'Z', ind=zpanel, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCy[ypanel],mesh.vectorCCy[ypanel]]),color='w')
plt.title('Plan l2-model.')
plt.gca().set_aspect('equal')
plt.ylabel('y')
ax.xaxis.set_visible(False)
plt.gca().set_aspect('equal', adjustable='box')
# Vertica section
ax = plt.subplot(234)
mesh.plotSlice(m_l2, ax = ax, normal = 'Y', ind=midx, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCz[zpanel],mesh.vectorCCz[zpanel]]),color='w')
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([Z.min(),Z.max()]),color='k')
plt.title('E-W l2-model.')
plt.gca().set_aspect('equal')
plt.xlabel('x')
plt.ylabel('z')
plt.gca().set_aspect('equal', adjustable='box')
#Plot Lp model
ax = plt.subplot(232)
mesh.plotSlice(m_lp, ax = ax, normal = 'Z', ind=zpanel, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCy[ypanel],mesh.vectorCCy[ypanel]]),color='w')
plt.title('Plan lp-model.')
plt.gca().set_aspect('equal')
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
plt.gca().set_aspect('equal', adjustable='box')
# Vertical section
ax = plt.subplot(235)
mesh.plotSlice(m_lp, ax = ax, normal = 'Y', ind=midx, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCz[zpanel],mesh.vectorCCz[zpanel]]),color='w')
plt.title('E-W lp-model.')
plt.gca().set_aspect('equal')
ax.yaxis.set_visible(False)
plt.xlabel('x')
plt.gca().set_aspect('equal', adjustable='box')
#Plot True model
ax = plt.subplot(233)
mesh.plotSlice(m_true, ax = ax, normal = 'Z', ind=zpanel, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCy[ypanel],mesh.vectorCCy[ypanel]]),color='w')
plt.title('Plan true model.')
plt.gca().set_aspect('equal')
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
plt.gca().set_aspect('equal', adjustable='box')
# Vertical section
ax = plt.subplot(236)
mesh.plotSlice(m_true, ax = ax, normal = 'Y', ind=midx, grid=True, clim = (0., model.max()),pcolorOpts={'cmap':'viridis'})
plt.plot(([mesh.vectorCCx[0],mesh.vectorCCx[-1]]),([mesh.vectorCCz[zpanel],mesh.vectorCCz[zpanel]]),color='w')
plt.title('E-W true model.')
plt.gca().set_aspect('equal')
plt.xlabel('x')
ax.yaxis.set_visible(False)
plt.gca().set_aspect('equal', adjustable='box')
# -
# Great, we have a 3D model of susceptibility, but the job is not done yet.
# A VERY important step of the inversion workflow is to look at how well the model can predict the observed data.
# The figure below compares the observed, predicted and normalized residual.
#
# +
# Plot predicted data and residual
plt.figure()
pred = prob.fields(mrec) #: this is matrix multiplication!!
plt.subplot(221)
plt.imshow(data.reshape(X.shape))
plt.title('Observed data.')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.subplot(222)
plt.imshow(pred.reshape(X.shape))
plt.title('Predicted data.')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.subplot(223)
plt.imshow(data.reshape(X.shape) - pred.reshape(X.shape))
plt.title('Residual data.')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.subplot(224)
plt.imshow( (data.reshape(X.shape) - pred.reshape(X.shape)) / wd.reshape(X.shape) )
plt.title('Normalized Residual')
plt.gca().set_aspect('equal', adjustable='box')
plt.colorbar()
plt.tight_layout()
# -
# Good job!
# Hopefully we covered all the important points regarding the inversion of magnetic field data using the integral formulation.
#
# Make sure you visit the notebook for the compact norms regularization.
#
# Cheers!
|
docs/case-studies/PF/Linear_Problem_Mag.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
base=os.path.dirname(os.path.realpath("__file__")).split(os.sep)
os.chdir(os.sep+os.path.join(*base[:-1]))
os.getcwd()
import numpy as np
import normflowpy as nfp
import datasets
import torch
from matplotlib import pyplot as plt
from torch.distributions import MultivariateNormal
from experiments.functions import run_training
# +
dataset_type = datasets.DatasetType.MOONS
n_training_samples = 50000
n_validation_samples = 10000
n_flow_blocks = 3
batch_size = 32
n_epochs = 50
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("Current Working Device is set to:" + str(device))
training_data = datasets.get_dataset(dataset_type, n_training_samples)
training_dataset_loader = torch.utils.data.DataLoader(training_data, batch_size=batch_size,
shuffle=True, num_workers=0)
validation_data = datasets.get_dataset(dataset_type, n_validation_samples)
validation_dataset_loader = torch.utils.data.DataLoader(validation_data, batch_size=batch_size,
shuffle=False, num_workers=0)
# -
# # Create Glow Normalizing Flow Model
dim = training_data.dim() # get data dim
base_distribution = MultivariateNormal(torch.zeros(dim, device=device),
torch.eye(dim, device=device)) # generate a class for base distribution
flows = []
for i in range(n_flow_blocks):
flows.append(
nfp.flows.ActNorm(dim=dim))
flows.append(
nfp.flows.InvertibleFullyConnected(dim=dim))
flows.append(
nfp.flows.AffineCoupling(x_shape=[dim], parity=i % 2, net_class=nfp.base_nets.generate_mlp_class(), nh=32))
flow = nfp.NormalizingFlowModel(base_distribution, flows).to(device)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Set Optimizer and run training
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
optimizer = torch.optim.Adam(flow.parameters(), lr=1e-4)
run_training(n_epochs, training_dataset_loader, validation_dataset_loader, flow, optimizer, device)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Plot probability Map
# + pycharm={"name": "#%%\n"}
def generate_probability_map(n_points, in_x_min, in_x_max, in_y_min, in_y_max, in_flow_model, in_device):
results = []
for x_tag in torch.linspace(in_x_min, in_x_max, n_points):
_results_y = []
for y_tag in torch.linspace(in_y_min, in_y_max, n_points):
d = torch.stack([x_tag, y_tag]).reshape([1, -1]).to(in_device)
_results_y.append(in_flow_model.nll(d).item())
results.append(_results_y)
return np.exp(-np.asarray(results)).T
x_min = -1.4
x_max = 2.1
y_min = -1
y_max = 1.5
res = generate_probability_map(200, x_min, x_max, y_min, y_max, flow, device)
fig, (ax0, ax1) = plt.subplots(2)
x = np.linspace(x_min, x_max, 200)
y = np.linspace(y_min, y_max, 200)
xx, yy = np.meshgrid(x, y)
im = ax0.pcolormesh(xx, yy, res)
ax1.plot(training_data[:, 0], training_data[:, 1], "o")
ax1.grid()
plt.show()
|
examples/moons_glow_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1. Import Libraries
# +
import pandas as pd
import numpy as np
import xgboost as xgb
import tensorflow as tf
import mimic_iv_utils as utils
from functools import reduce
from tqdm import tqdm
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn import metrics
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
from keras.models import Sequential
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
from matplotlib import pyplot as plt
# -
# ## 2. Fetch Data
# +
duration_data = {}
durations = [8, 12, 16, 24, 48, 72, 96, 120, 144]
for d in tqdm(durations):
static = utils.getStaticFeatures()
first_lab = utils.getLabFeatures(duration=d)
last_lab = utils.getLabFeatures(mode='last', duration=d)
first_vitals = utils.getVitalsFeatures(duration=d)
last_vitals = utils.getVitalsFeatures(mode='last', duration=d)
max_vitals = utils.getMinMaxVitalsFeatures(mode='max', duration=d)
min_vitals = utils.getMinMaxVitalsFeatures(duration=d)
avg_vitals = utils.getMinMaxVitalsFeatures(mode='avg', duration=d)
mortality = utils.getInhospitalMortality()
filtered = utils.getFilteredCohort(duration=d)
dfs = [filtered, static, first_lab, last_lab, first_vitals, last_vitals, max_vitals, min_vitals, avg_vitals, mortality]
data = reduce(lambda left, right: pd.merge(left, right, on=['stay_id'], how='inner'), dfs)
data.drop(columns=['subject_id', 'hadm_id', 'stay_id'], inplace=True)
X = data.values
y = X[:,-1]
y = y.astype('int')
X = X[:,0:-1]
X_header = [x for x in data.columns.values]
X_header = X_header[0:-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
duration_data[d] = [X_train, X_test, y_train, y_test]
# -
# ## 3. Deep Learning
results = {}
for k, v in tqdm(duration_data.items()):
X_train, X_test, y_train, y_test = v[0], v[1], v[2], v[3]
X_train_array = np.asarray(X_train).astype('float32')
X_test_array = np.asarray(X_test).astype('float32')
imp_mean = SimpleImputer(missing_values=np.nan, strategy='mean')
imp_mean.fit(X_train_array)
X_train_imputed = imp_mean.transform(X_train_array)
X_test_imputed = imp_mean.transform(X_test_array)
normalizer = preprocessing.Normalization()
normalizer.adapt(X_train_imputed)
X_train_noralized = normalizer(X_train_imputed)
X_test_noralized = normalizer(X_test_imputed)
model = Sequential()
model.add(layers.Dense(X_train.shape[1], activation=tf.nn.relu, kernel_initializer='he_normal', bias_initializer='zeros'))
model.add(layers.Dense(40, activation=tf.nn.relu, kernel_initializer='he_normal', bias_initializer='zeros'))
model.add(layers.Dense(1, activation=tf.nn.sigmoid, kernel_initializer='he_normal', bias_initializer='zeros'))
model.compile(optimizer="adam", loss="binary_crossentropy", metrics=["accuracy"])
model.fit(X_train_noralized, y_train, validation_data=(X_test_noralized, y_test), epochs=5, batch_size=256)
probs = model.predict(X_test_noralized)
results[k] = [y_test, probs]
# ## 4. Results
for k, v in tqdm(results.items()):
y_test = v[0]
probs = v[1]
preds = np.around(probs)
print('--------------------------------------------------------------------------------------------------------------------')
print('-------------------------------------------- * duration ='+ str(k)+' * ----------------------------------------------------')
print('--------------------------------------------------------------------------------------------------------------------')
roc_score = metrics.roc_auc_score(y_test, probs)
print('ROC: ', roc_score)
classification_report = metrics.classification_report(y_test, preds)
print(classification_report)
cm = confusion_matrix(y_test, preds, labels=[0, 1])
disp = ConfusionMatrixDisplay(confusion_matrix=cm, display_labels=['survided', 'not-survived'])
disp.plot()
# +
from sklearn.metrics import roc_curve
plt.figure(figsize=(15,10))
for k, v in tqdm(results.items()):
y_test = v[0]
probs = v[1]
preds = np.around(probs)
fpr, tpr, _ = roc_curve(y_test, preds)
plt.plot(fpr, tpr, marker='.', label='Duration: ' + str(k))
# axis labels
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
# show the legend
plt.legend()
# show the plot
plt.show()
# +
import seaborn as sns
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(15,10))
for k, v in tqdm(results.items()):
y_test = v[0]
probs = v[1]
sns.distplot(probs, hist=True, rug=False, axlabel='Prediction Probability')
fig.legend(labels=results.keys())
plt.show()
|
analysis/machine_learning/mimic_iv_feature_engineering_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="5fCEDCU_qrC0"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# # **Upcoming ESG Regulations**
# + [markdown] colab_type="text" id="GJBs_flRovLc"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# ### **About this Webpage**
# + [markdown] colab_type="text" id="allXugATwQQQ"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# The document you are reading is not a static webpage, but an interactive environment called a **Jupyter notebook** that lets you write, share, and execute code. To learn more about the Jupyter project, see [jupyter.org](https://www.jupyter.org).
#
# Below is a **code cell** with a short Python script that prints a message, does a calculation, and then prints the value of a variable:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="gJr_9dXGpJ05" outputId="9b5524e4-0cf7-4f51-c0af-4ea1da9b27a6"
print('Hello world!')
var1 = 3 * 4
print(var1)
# + [markdown] colab_type="text" id="2fhs6GZ4qFMx"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# **To execute the code in the above cell, select it with a click and then either press the play button to the left of the code, or use the keyboard shortcut "Command/Ctrl + Enter".** To edit the code, just click the cell and start editing.
#
# Variables that you define in one cell can later be used in other cells:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="-gE-Ez1qtyIA" outputId="650f347e-7e2d-4216-f654-86685d27b6c8"
print(var1 + 2)
# + [markdown] colab_type="text" id="lSrWNr3MuFUS"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# Colab notebooks allow you to combine **executable code** and **rich text** in a single document, along with **images**, **HTML**, **LaTeX**, and more. When you create your own Colab notebooks, they are stored in your Google Drive account. You can easily share your Colab notebooks with co-workers or friends, allowing them to comment on your notebooks or even edit them. To learn more, see [Overview of Colab](/notebooks/basic_features_overview.ipynb). To create a new Colab notebook you can use the File menu above, or use the following link: [create a new Colab notebook](http://colab.research.google.com#create=true).
#
# <br><br>
#
# ---
#
#
#
# + [markdown] colab_type="text" id="OwuxHmxllTwN"
# <!---
#
# If you can see this text, you have this cell in "edit mode".
# Click anywhere inside the cell and hit Command/Ctrl + Enter to "render" the cell.
#
# --->
# # **ESG Regulation Timeline**
#
# The following cell is a script written in Python that generates an interactive timeline of the dates at which upcoming ESG Regulations go into force. Scroll down to see the plot. Hover your mouse over a data point for more information.
#
# _Click inside the cell, and then click the play button to execute the code._
# + colab={"base_uri": "https://localhost:8080/", "height": 542} colab_type="code" id="OA3qMV8IUyLu" outputId="726587ed-c5b5-4a64-f879-3e960b4fbd87"
from plotly.offline import iplot
import numpy as np
data = [{'visible': False,
'line': dict(color='#00CED1', width=6),
'name': '𝜈 = '+str(step),
'x': np.arange(0,10,0.01),
'y': np.sin(step*np.arange(0,10,0.01))
} for step in np.arange(0,5,0.1)]
data[10]['visible'] = True
steps = []
for i in range(len(data)):
step = {'method': 'restyle',
'args': ['visible', [False] * len(data)],
}
step['args'][1][i] = True # Toggle i'th trace to "visible"
steps.append(step)
sliders = [dict(
active = 10,
currentvalue = {"prefix": "Frequency: "},
pad = {"t": 50},
steps = steps
)]
layout = dict(sliders=sliders)
fig = dict(data=data, layout=layout)
iplot(fig, filename='Sine Wave Slider')
# + [markdown] colab_type="text" id="m4BiLSfxPG_P"
# # Upcoming ESG Regulations by Jurisdiction
#
# + [markdown] colab_type="text" id="y827J89y-AWi"
# ## European Union
# + [markdown] colab_type="text" id="17747zJbZtZj"
# #### The Disclosure Regulation
# + [markdown] colab_type="text" id="6p5OVD8XQW5t"
# The disclosure Regulation mainly focuses on straaaaaaaange thiiiiiings.
#
# + [markdown] colab_type="text" id="W3CxUXp7ZdP8"
# ### Regulation 2
#
# this one too
# <br>
# lankkkkkkkk
# + [markdown] colab_type="text" id="NhfP0lYXaMwF"
# So I found these docs bleh bleh
# <br>
# linkkkkkkkkkk
# + colab={"base_uri": "https://localhost:8080/", "height": 542} colab_type="code" id="M_4QNJx4-ghh" outputId="bfabfbaf-f4b9-47d4-d453-e8fae523cf94"
from plotly.offline import iplot
import numpy as np
data = [dict(
visible = False,
line=dict(color='#00CED1', width=6),
name = '𝜈 = '+str(step),
x = np.arange(0,10,0.01),
y = np.sin(step*np.arange(0,10,0.01))) for step in np.arange(0,5,0.1)]
data[10]['visible'] = True
steps = []
for i in range(len(data)):
step = dict(
method = 'restyle',
args = ['visible', [False] * len(data)],
)
step['args'][1][i] = True # Toggle i'th trace to "visible"
steps.append(step)
sliders = [dict(
active = 10,
currentvalue = {"prefix": "Frequency: "},
pad = {"t": 50},
steps = steps
)]
layout = dict(sliders=sliders)
fig = dict(data=data, layout=layout)
iplot(fig, filename='Sine Wave Slider')
# + colab={"base_uri": "https://localhost:8080/", "height": 559} colab_type="code" id="C4HZx7Gndbrh" outputId="087c97a3-e56e-42cc-9180-3d4bf61f71b0"
from plotly.offline import iplot
import plotly.graph_objs as go
data = [
go.Contour(
z=[[10, 10.625, 12.5, 15.625, 20],
[5.625, 6.25, 8.125, 11.25, 15.625],
[2.5, 3.125, 5.8, 8.125, 12.5],
[0.625, 1.25, 3.125, 6.25, 10.625],
[0, 0.625, 2.5, 5.625, 10]]
)
]
iplot(data)
print('done')
# + [markdown] colab_type="text" id="-Rh3-Vt9Nev9"
# ## More Resources
#
# ### Working with Notebooks in Colab
# - [Overview of Colaboratory](/notebooks/basic_features_overview.ipynb)
# - [Guide to Markdown](/notebooks/markdown_guide.ipynb)
# - [Importing libraries and installing dependencies](/notebooks/snippets/importing_libraries.ipynb)
# - [Saving and loading notebooks in GitHub](https://colab.research.google.com/github/googlecolab/colabtools/blob/master/notebooks/colab-github-demo.ipynb)
# - [Interactive forms](/notebooks/forms.ipynb)
# - [Interactive widgets](/notebooks/widgets.ipynb)
# - <img src="/img/new.png" height="20px" align="left" hspace="4px" alt="New"></img>
# [TensorFlow 2 in Colab](/notebooks/tensorflow_version.ipynb)
#
# <a name="working-with-data"></a>
# ### Working with Data
# - [Loading data: Drive, Sheets, and Google Cloud Storage](/notebooks/io.ipynb)
# - [Charts: visualizing data](/notebooks/charts.ipynb)
# - [Getting started with BigQuery](/notebooks/bigquery.ipynb)
#
# ### Machine Learning Crash Course
# These are a few of the notebooks from Google's online Machine Learning course. See the [full course website](https://developers.google.com/machine-learning/crash-course/) for more.
# - [Intro to Pandas](/notebooks/mlcc/intro_to_pandas.ipynb)
# - [Tensorflow concepts](/notebooks/mlcc/tensorflow_programming_concepts.ipynb)
# - [First steps with TensorFlow](/notebooks/mlcc/first_steps_with_tensor_flow.ipynb)
# - [Intro to neural nets](/notebooks/mlcc/intro_to_neural_nets.ipynb)
# - [Intro to sparse data and embeddings](/notebooks/mlcc/intro_to_sparse_data_and_embeddings.ipynb)
#
# <a name="using-accelerated-hardware"></a>
# ### Using Accelerated Hardware
# - [TensorFlow with GPUs](/notebooks/gpu.ipynb)
# - [TensorFlow with TPUs](/notebooks/tpu.ipynb)
# + [markdown] colab_type="text" id="ZFFeVI4yO4xi"
# ### Sample
# + colab={"base_uri": "https://localhost:8080/", "height": 542} colab_type="code" id="8YCVGqZkJJxT" outputId="068e57aa-f946-4c31-c36e-54029f878647"
from plotly.offline import iplot
import plotly.graph_objs as go
data = [
go.Contour(
z=[[10, 10.25, 12.5, 15.625, 20],
[5.625, 6.25, 8.125, 11.25, 15.625],
[2.5, 3.125, 5., 8.125, 12.5],
[0.625, 1.25, 3.125, 6.25, 10.625],
[0, 0.625, 2.5, 5.625, 10]]
)
]
iplot(data)
|
Upcoming_ESG_Regulations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from matplotlib import pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
import pickle
with open('logs_bck.pickle', 'rb') as f:
logs = pickle.load(f)
epochs = list(range(1, len(logs[0]['history']['loss'])+1))
# +
loss = pd.DataFrame()
for log_entry in logs:
df_training = pd.DataFrame({
'epoch': epochs,
'loss': log_entry['history']['loss'],
'type': 'training',
'model': log_entry['model'],
'train size': log_entry['train_size']
})
df_validation = pd.DataFrame({
'epoch': epochs,
'loss': log_entry['history']['val_loss'],
'type': 'validation',
'model': log_entry['model'],
'train size': log_entry['train_size']
})
loss = pd.concat([loss, df_training, df_validation])
auc = pd.DataFrame()
for log_entry in logs:
df_training = pd.DataFrame({
'epoch': epochs,
'AUC': log_entry['history']['auc'],
'type': 'training',
'model': log_entry['model'],
'train size': log_entry['train_size']
})
df_validation = pd.DataFrame({
'epoch': epochs,
'AUC': log_entry['history']['val_auc'],
'type': 'validation',
'model': log_entry['model'],
'train size': log_entry['train_size']
})
auc = pd.concat([auc, df_training, df_validation])
# -
sns.set_context("notebook")
g = sns.FacetGrid(loss, col="model", hue='type', row='train size', margin_titles=True)
_ = g.map(sns.lineplot, "epoch", "loss", alpha=.7)
_ = g.add_legend(title='')
_ = g.set_titles(col_template="{col_name}")
_ = g.set(xticks=epochs)
#plt.savefig("loss.svg", format="svg")
sns.set_context("notebook")
g = sns.FacetGrid(auc, col="model", hue='type', row='train size', margin_titles=True) #, col_wrap=3, height=4)
_ = g.map(sns.lineplot, "epoch", "AUC", alpha=.7)
_ = g.add_legend(title='')
_ = g.set_titles(col_template="{col_name}")
_ = g.set(xticks=epochs)
#plt.savefig("auc.svg", format="svg")
# +
evaluation = []
for log_entry in logs:
evaluation.append({
'AUC': log_entry['evaluation'][log_entry['metrics'].index('auc')],
'model': log_entry['model'],
'train size': log_entry['train_size']
})
evaluation = pd.DataFrame(evaluation)
pd.pivot_table(evaluation, values='AUC', index=['model'], columns=['train size'])
# -
sns.set_context("notebook")
g = sns.FacetGrid(evaluation, hue='model', margin_titles=True, height=8, aspect=1.5)
_ = g.map(sns.lineplot, "train size", "AUC", alpha=.7)
_ = g.add_legend(title='')
_ = g.set(xticks=evaluation['train size'].unique())
#plt.savefig("evaluation.svg", format="svg")
|
models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Introducing Keras
#
# Be sure to "pip install keras" first!
#
# Keras is a layer on top of TensorFlow that makes things a lot easier. Not only is it easier to use, it's easier to tune.
#
# Let's set up the same deep neural network we set up with TensorFlow to learn from the MNIST data set.
#
# First we'll import all the stuff we need, which will initialize Keras as a side effect:
# + deletable=true editable=true
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.optimizers import RMSprop
# + [markdown] deletable=true editable=true
# We'll load up the MNIST data set. In Keras, it's a little bit different - there are 60K training samples and 10K test samples. No "validation" samples.
# + deletable=true editable=true
(mnist_train_images, mnist_train_labels), (mnist_test_images, mnist_test_labels) = mnist.load_data()
# + [markdown] deletable=true editable=true
# We need to explicitly convert the data into the format Keras / TensorFlow expects. We divide the image data by 255 in order to normalize it into 0-1 range, after converting it into floating point values.
# + deletable=true editable=true
train_images = mnist_train_images.reshape(60000, 784)
test_images = mnist_test_images.reshape(10000, 784)
train_images = train_images.astype('float32')
test_images = test_images.astype('float32')
train_images /= 255
test_images /= 255
# + [markdown] deletable=true editable=true
# Now we'll convert the 0-9 labels into "one-hot" format, as we did for TensorFlow.
# + deletable=true editable=true
train_labels = keras.utils.to_categorical(mnist_train_labels, 10)
test_labels = keras.utils.to_categorical(mnist_test_labels, 10)
# + [markdown] deletable=true editable=true
# Let's take a peek at one of the training images just to make sure it looks OK:
# + deletable=true editable=true
import matplotlib.pyplot as plt
def display_sample(num):
#Print the one-hot array of this sample's label
print(train_labels[num])
#Print the label converted back to a number
label = train_labels[num].argmax(axis=0)
#Reshape the 768 values to a 28x28 image
image = train_images[num].reshape([28,28])
plt.title('Sample: %d Label: %d' % (num, label))
plt.imshow(image, cmap=plt.get_cmap('gray_r'))
plt.show()
display_sample(1234)
# + [markdown] deletable=true editable=true
# Here's where things get exciting. All that code we wrote in Tensorflow creating placeholders, variables, and defining a bunch of linear algebra for each layer in our neural network? None of that is necessary with Keras!
#
# We can set up the same layers like this. The input layer of 784 features feeds into a ReLU layer of 512 nodes, which then goes into 10 nodes with softmax applied. Couldn't be simpler:
# + deletable=true editable=true
model = Sequential()
model.add(Dense(512, activation='relu', input_shape=(784,)))
model.add(Dense(10, activation='softmax'))
# + [markdown] deletable=true editable=true
# We can even get a nice description of the resulting model:
# + deletable=true editable=true
model.summary()
# + [markdown] deletable=true editable=true
# Setting up our optimizer and loss function is just as simple. We will use the RMSProp optimizer here. Other choices include Adagrad, SGD, Adam, Adamax, and Nadam. See https://keras.io/optimizers/
# + deletable=true editable=true
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
# + [markdown] deletable=true editable=true
# Training our model is also just one line of code with Keras. Here we'll do 10 epochs with a batch size of 100. Keras is slower, and if we're not running on top of a GPU-accelerated Tensorflow this can take a fair amount of time (that's why I've limited it to just 10 epochs.)
# + deletable=true editable=true
history = model.fit(train_images, train_labels,
batch_size=100,
epochs=10,
verbose=2,
validation_data=(test_images, test_labels))
# + [markdown] deletable=true editable=true
# But, even with just 10 epochs, we've outperformed our Tensorflow version considerably!
# + deletable=true editable=true
score = model.evaluate(test_images, test_labels, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# + [markdown] deletable=true editable=true
# As before let's visualize the ones it got wrong. As this model is much better, we'll have to search deeper to find mistakes to look at.
# + deletable=true editable=true
for x in range(1000):
test_image = test_images[x,:].reshape(1,784)
predicted_cat = model.predict(test_image).argmax()
label = test_labels[x].argmax()
if (predicted_cat != label):
plt.title('Prediction: %d Label: %d' % (predicted_cat, label))
plt.imshow(test_image.reshape([28,28]), cmap=plt.get_cmap('gray_r'))
plt.show()
# + [markdown] deletable=true editable=true
# You can see most of the ones it's having trouble with, are images a human would have trouble with as well!
#
# ## Excercise
#
# As before, see if you can improve on the results! Does running more epochs help considerably? How about trying different optimizers?
#
# You can also take advantage of Keras's ease of use to try different topologies quickly. Keras includes a MNIST example, where they add an additional layer, and use Dropout at each step to prevent overfitting, like this:
#
# `
# model = Sequential()
# model.add(Dense(512, activation='relu', input_shape=(784,)))
# model.add(Dropout(0.2))
# model.add(Dense(512, activation='relu'))
# model.add(Dropout(0.2))
# model.add(Dense(10, activation='softmax'))
# `
#
# Try adapting that to our code above and see if it makes a difference or not.
# + deletable=true editable=true
|
7_deep_learning_az/DataScience-Python3/Keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/manashpratim/Algorithms-and-Data-Structures/blob/master/Searching_Algorithms.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xBT76Pyo20TV" colab_type="text"
# ## **Sequential Search**
# + [markdown] id="LlAve3SY3Ig0" colab_type="text"
# **Unordered List**
# + id="tcWTSE1510YR" colab_type="code" colab={}
def seq_search(arr,element):
for i in range(len(arr)):
if arr[i] == element:
return True
return False
# + id="nsQTkkRR35Yi" colab_type="code" colab={}
arr = [1,9,2,8,3,4,7,5,6]
# + id="sgi25LNd3_xM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="86764490-2d1c-408c-a26f-5ea43dcb2ccb"
seq_search(arr,8)
# + id="CeCZ95B94E2t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b7b9e402-2d2d-4b7a-b1d4-c40225c25ec6"
seq_search(arr,2.5)
# + [markdown] id="2O-oA9uc4OFM" colab_type="text"
# **Ordered List**
# + id="gcz4meWK4Q_U" colab_type="code" colab={}
def ordered_seq_search(arr,element):
for i in range(len(arr)):
if arr[i]==element:
return True
break
elif arr[i]>element:
return False
break
# + id="SHDjLQmY5MwM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1ba67241-514b-44fc-ffe5-23e3b9b0e1f8"
ordered_seq_search(sorted(arr),4)
# + id="DEnprFcc5VYc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f73b25b8-dccc-413a-ceca-3811994a58dc"
ordered_seq_search(sorted(arr),4.5)
# + [markdown] id="15P8j3ML5rCg" colab_type="text"
# ## **Binary Search**
# + id="JmGRSZFb7XVr" colab_type="code" colab={}
def binary_search(arr,element):
first=0
last=len(arr)-1
found=False
while first <= last and not found:
mid=(first+last)//2
if arr[mid]==element: #best case
found=True
else:
if element<arr[mid]: #check in the LHS of the mid
last=mid-1
else: #check in RHS of the mid
first=mid+1
return found
# + id="ih7p6Rj98ohG" colab_type="code" colab={}
arr=[1,9,2,8,3,4,7,5,6]
# + id="NTx0BFyL8uot" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="28ba85a2-07db-41f1-bff0-b130722c70a9"
binary_search(sorted(arr),8) #input array must be sorted
# + id="rjS_apnA9Fqf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c286b300-e71d-49cf-db10-706595ef9c96"
binary_search(sorted(arr),4.5)
# + [markdown] id="8s7sdk8T9OTG" colab_type="text"
# **Recursive Implementation**
# + id="6Dvbhg9C9SCa" colab_type="code" colab={}
def binary_search_recursive(arr,element):
if len(arr)==0: #base case
return False
else:
mid=len(arr)//2
if arr[mid]==element:
return True
else:
if element>arr[mid]: #check in the LHS of mid
return binary_search_recursive(arr[mid+1:],element)
else: #check in the RHS of mid
return binary_search_recursive(arr[:mid],element)
# + id="UvOfP668_6ll" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b7bc7ebc-f66c-4ac2-a2ab-81c21cb6c29a"
binary_search_recursive(sorted(arr),8)
# + id="6YKXmLjWAF3x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0154e10d-2e2e-<PASSWORD>"
binary_search_recursive(sorted(arr),4.5)
# + [markdown] id="uumdY0ejPyqu" colab_type="text"
# ## **Hash Table**
# + id="U2sFqUu3QClP" colab_type="code" colab={}
class HashTable(object):
def __init__(self,size):
self.size = size #defining the size of the hash table
self.slots = [None]*self.size #initializing the slots as a list with None values
self.data = [None]*self.size #initializing the data as a list with None values
def hashfunction(self,key,size):
return key%size #the remainder method
def rehashfunction(self,oldhash,size): #for collision resolution
return (oldhash+1)%size
def insert(self,key,data): #function to insert elements into a hash table
if type(key) == str:
s=0
for i in key:
s=s+ord(i)
hashvalue = self.hashfunction(s,len(self.slots))
else:
hashvalue = self.hashfunction(key,len(self.slots))
if self.slots[hashvalue] == None:
self.slots[hashvalue] = key
self.data[hashvalue] = data
else: #incase of collision
if self.slots[hashvalue] == key: #if the key already exists, replace the data
self.data[hashvalue] = data
else:
nextslot = self.rehashfunction(hashvalue,len(self.slots))
while self.slots[nextslot] != None and self.slots[nextslot] !=key: #loop till we find an empty slot or a slot where the key exists
nextslot=self.rehashfunction(nextslot,len(self.slots))
if self.slots[nextslot] == None:
self.slots[nextslot] = key
self.data[nextslot] = data
else: #if the key already exists
self.data[nextslot] = data
def get(self,key): #function to get the data, given a key
if type(key) == str:
s=0
for i in key:
s=s+ord(i)
startslot = self.hashfunction(s,len(self.slots))
else:
startslot = self.hashfunction(key,len(self.slots))
data = None
found = False
stop = False
position = startslot
while self.slots[position]!=None and not found and not stop:
if self.slots[position] == key: #if key is found, retrieve the data
found = True
data = self.data[position]
else:
position=self.rehashfunction(position,len(self.slots))
if position == startslot: #if after searching the whole hashtable we end up in the startslot again, then stop searching
stop = True
if data == None:
print('Invalid Key!')
else:
return data
def delete(self,key): #function to delete the key and data for a given key
if type(key) == str:
s=0
for i in key:
s=s+ord(i)
startslot = self.hashfunction(s,len(self.slots))
else:
startslot = self.hashfunction(key,len(self.slots))
found = False
stop = False
position = startslot
while self.slots[position]!=None and not found and not stop:
if self.slots[position] == key: #if key is found, set the data and key to None
found = True
self.slots[position] = None
self.data[position] = None
else:
position=self.rehashfunction(position,len(self.slots))
if position == startslot: #if after searching the whole hashtable we end up in the startslot again, then stop searching
stop = True
if found == False:
print('Invalid Key!')
else:
print('Key {} deleted!'.format(key))
# + id="5J22BZQGZuJJ" colab_type="code" colab={}
h = HashTable(4) #creating a hashtable of size 4
# + id="o2LRzr_eZ1Ql" colab_type="code" colab={}
h.insert(1,5) #here, the key is 1 and the data at the key is 5
h.insert(2,'two')
# + id="Deea2bNZaFb_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="48a32d79-0290-4107-9d12-2d7b51e88e4f"
h.get(1)
# + id="qx6kdp3lb_s2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0c76b779-a6b7-425d-abf6-421ad61e3619"
h.get(2)
# + id="pnQCRGTYcD3z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2c656791-137b-4d50-f389-df18966e6d57"
h.get(10)
# + id="Yi0n7UgkcH4n" colab_type="code" colab={}
h.insert('apple',5)
h.insert('mango','tasty')
# + id="ivlDbtpQcLYb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dd469010-4902-4953-8047-355bd1ddfedf"
h.get('apple')
# + id="5Q1EjQdsiSxt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fef9676b-42e2-4236-9d56-7c2baeac4c44"
h.get('mango')
# + id="weMbyCJ8hLhO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="99f5ce74-ab65-4b15-b587-fd6667752b9e"
h.delete('apple')
# + id="4bK0UudwhPFK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a7603e3a-25d9-4d63-e56b-359d0b763fa2"
h.get('apple')
# + [markdown] id="pthMzLXChVhy" colab_type="text"
# **That's the end of the hash table implementation.**
#
# **Note:** Python already has a built-in dictionary object that serves as a Hash Table.
|
Searching_Algorithms.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TimeSeries_3.7
# language: python
# name: timeseries_3.7
# ---
# ### Installing the Library
# !pip install MAQTextSDK
# ### Send Request
# +
#Load Text
keyphrase_input = dict()
#Set Text
keyphrase_input["text"] = "Does social capital determine innovation ? To what extent? This paper deals with two questions: Does social capital determine innovation in manufacturing firms? If it is the case, to what extent? To deal with these questions, we review the literature on innovation in order to see how social capital came to be added to the other forms of capital as an explanatory variable of innovation. In doing so, we have been led to follow the dominating view of the literature on social capital and innovation which claims that social capital cannot be captured through a single indicator, but that it actually takes many different forms that must be accounted for. Therefore, to the traditional explanatory variables of innovation, we have added five forms of structural social capital (business network assets, information network assets, research network assets, participation assets, and relational assets) and one form of cognitive social capital (reciprocal trust). In a context where empirical investigations regarding the relations between social capital and innovation are still scanty, this paper makes contributions to the advancement of knowledge in providing new evidence regarding the impact and the extent of social capital on innovation at the two decisionmaking stages considered in this study"
#Top Number Of Keyphrases
keyphrase_input["keyphrases_count"] = 10
#More Score, more different/diverse keyphrases are generated
#Less Score, more duplicate keyphrases are generated
keyphrase_input["diversity_threshold"] = 0.52
#Similarity threshold for Alias/Similar Keyphrase with Top Key Phrase [Similar Column]
#More The Value, More accurate keyphrases are found with top key-phrase [Similar Column]
keyphrase_input["alias_threshold"] = 0.65
# -
#Set API Key
APIKey = 'Valid API Key'
APIEndpoint = "https://maqtextnalyticssdk.azure-api.net/text/"
#Import
import MAQTextSDK.maq_text_analytics_linux as TextSDK
import pandas as pd
# +
#Send Request
textClient = TextSDK.MAQTextAnalyticsLinux(base_url = APIEndpoint)
response = textClient.post_keyphrase_extractor(api_key = APIKey, data_input =keyphrase_input)
response_df = pd.DataFrame(response, columns = ['KeyPhrase','Score','Similar'])
display(response_df)
# -
# ### Send Request using Requests Library
import requests
#Set API Key
APIKey = 'Valid API Key'
APIEndpoint = "https://maqtextnalyticssdk.azure-api.net/text/"
headers = {"APIKey": APIKey}
# +
#Load Text
keyphrase_input = dict()
#Set Text
keyphrase_input["text"] = "Does social capital determine innovation ? To what extent? This paper deals with two questions: Does social capital determine innovation in manufacturing firms? If it is the case, to what extent? To deal with these questions, we review the literature on innovation in order to see how social capital came to be added to the other forms of capital as an explanatory variable of innovation. In doing so, we have been led to follow the dominating view of the literature on social capital and innovation which claims that social capital cannot be captured through a single indicator, but that it actually takes many different forms that must be accounted for. Therefore, to the traditional explanatory variables of innovation, we have added five forms of structural social capital (business network assets, information network assets, research network assets, participation assets, and relational assets) and one form of cognitive social capital (reciprocal trust). In a context where empirical investigations regarding the relations between social capital and innovation are still scanty, this paper makes contributions to the advancement of knowledge in providing new evidence regarding the impact and the extent of social capital on innovation at the two decisionmaking stages considered in this study"
#Top Number Of Keyphrases
keyphrase_input["keyphrases_count"] = 10
#More Score, more different/diverse keyphrases are generated
#Less Score, more duplicate keyphrases are generated
keyphrase_input["diversity_threshold"] = 0.52
#Similarity threshold for Alias/Similar Keyphrase with Top Key Phrase
#More The Value, More accurate similar keyphrases are found with top key-phrase
keyphrase_input["alias_threshold"] = 0.65
# -
response = requests.post(APIEndpoint + "/KeyPhrase", headers=headers, json=keyphrase_input)
# +
response_df = pd.DataFrame(response.json(), columns = ['KeyPhrase','Score','Similar'])
display(response_df)
# -
# ### Error Handling
# #### Invalid or Expired API Key
#Set API Key
APIKey = 'Invalid/Expired API Key'
APIEndpoint = "https://maqtextnalyticssdk.azure-api.net/text/"
# +
#Send Request
textClient = TextSDK.MAQTextAnalyticsLinux(base_url = APIEndpoint)
response = textClient.post_keyphrase_extractor(api_key = APIKey, data_input = keyphrase_input)
print(response)
|
Samples/KeyPhrasesDemo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
import numpy as np
import random
import keras
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
import tensorflow as tf
from tensorflow.keras import models
from tensorflow.keras import layers
import matplotlib.pyplot as plt
# %matplotlib inline
tf.__version__
# +
# number of characters in a word.
# for instance abccba has nb_chars = 6
nb_chars = 5
# number of possible characters used during the encoding.
# for instance abcde leads to 01234 has nb_letters = 5
nb_letters = 26
# number of words samples to be generated
nb_words = 10000
# percentage of words that will be used for validation
percentage_split = 0.60
# number of epochs for fitting the model training step
nb_epochs = 200
# -
# total number of combinations
nb_letters**nb_chars
# +
def create_inputs(nb_words, nb_chars, nb_letters):
'''Create a numpy array of nb_words rows with nb_chars columns each element
being a random letter of nb_letters (a, b...)'''
words = np.zeros((nb_words, nb_chars), dtype=int)
for w in range(nb_words):
optim_tentative = False
if optim_tentative == True and w%10 != 0:
i = random.randint(0, nb_letters-1)
for c in range(nb_chars):
words[w, c] = ord('a') + i
else:
for c in range(nb_chars):
i = random.randint(0, nb_letters-1)
words[w, c] = ord('a') + i
return words
def encrypt(words, nb_words, nb_chars):
'''Encrypt each element of a numpy array of nb_words rows with nb_chars
columns each item with a secret algorithm'''
encrypted_words = words.copy()
encrypted_words_probs = np.zeros((nb_words, nb_chars, nb_chars))
#val_max = -1
for w in range(nb_words):
for c in range(nb_chars): # 0,1,2,3,4
encrypted_words[w,c] = int(words[w,c]) - 49
val = encrypted_words[w,c] - 48
#if val > val_max:
# val_max = val
# add entropy (i.e. mistakes in the encryption)
#epsilon = random.randint(0, 100)
#if epsilon == 5 and val != val_max:
#val +=1
#print('w:',w,', c:',c,', [wc]:', val)
#encrypted_words_probs[w, c, val ] = 1.0
encrypted_words[w,c] = val
return encrypted_words
# -
def build_model(nb_chars, nb_letters):
# This returns a tensor
inputs = layers.Input(shape=(nb_chars,), dtype='float32', name='main_input')
#original_inputs = tf.keras.Input(shape=(original_dim,), name='encoder_input')
# a layer instance is callable on a tensor, and returns a tensor
x = layers.Dense(4096, activation='relu', name='hl_1')(inputs)
#x = layers.Dense(2048, activation='relu', name='hl_1')(inputs)
#x = layers.Dense(64, activation='relu', name='hl_2')(x)
outputs = []
losses = {}
for o in range(nb_chars):
name_i = 'output_'+str(o)
output_i = layers.Dense(nb_letters, activation='softmax', dtype='float32', name=name_i)(x)
outputs.append(output_i)
losses[name_i] = 'categorical_crossentropy'
model = models.Model(inputs=inputs, outputs=outputs)
rmsprop = tf.keras.optimizers.RMSprop(lr=0.01)
model.compile(optimizer=rmsprop,
loss=losses,
metrics=['accuracy'])
return model
def print_readable_inputs(x):
words = []
for w in x:
word = ''
for c in w:
word += chr(c)
words.append(word)
print(words)
def print_readable_outputs_(outputs, nb_words, nb_chars):
# outputs are listed : first, per char, second by sample, third by letter probability
words = [''] * nb_words
c_i = 0
for char in outputs:
s_i = 0
for sample in char:
l_i = 0
best_value = -float('inf')
best_letter = -1
for letter_probs in sample:
if letter_probs > best_value:
best_value = letter_probs
best_letter = l_i
l_i += 1
words[s_i] += str(best_letter)
if c_i != nb_chars - 1:
words[s_i] += ' '
s_i += 1
c_i += 1
print(words)
def print_readable_outputs(outputs, nb_words, nb_chars):
# outputs are listed : first, per char, second by sample, third by letter probability
words = [''] * nb_words
c_i = 0
for char in outputs:
s_i = 0
for sample in char:
best_letter = np.argmax(sample)
words[s_i] += str(best_letter)
if c_i != nb_chars - 1:
words[s_i] += ' '
s_i += 1
c_i += 1
print(words)
# +
x = create_inputs(nb_words, nb_chars, nb_letters)
print('x: (as readable inputs)')
first_n_samples = 4
print_readable_inputs(x[:first_n_samples])
print('x (partial):\n', x[:first_n_samples], 'out of ',len(x))
print()
# process the x data as useful ANN input data
scaler = StandardScaler()
#scaler = MinMaxScaler()
x_train = scaler.fit_transform(x)
print('x_train:\n', x_train[:first_n_samples], 'out of ',len(x_train))
print()
# create output data for training
y = encrypt(x, nb_words, nb_chars)
print('y (readable):\n', y)
print()
# process the y data as useful ANN output data
y_train0 = keras.utils.to_categorical(y, nb_letters)
print('y (less readable):\n', y_train0[:first_n_samples], 'out of ',len(y_train0))
print('')
# process the y data as useful ANN multiple-outputs data
y_train = []
for c in range(nb_chars):
# extract each 'char' colomn from the global y_train0 tensor
# in order to have multiplue yi_train outputs tensors
yi_train = y_train0[:,c,:]
y_train.append(yi_train)
# Not really displayable, henced commented
#print('y_train):')
#print(y_train[:first_n_samples])
# -
coding_model = build_model(nb_chars, nb_letters)
print(coding_model.summary())
history = coding_model.fit(x_train, y_train, validation_split=percentage_split, batch_size=32, epochs=nb_epochs, verbose=1)
history_dict = history.history
print(history_dict.keys())
# +
# Plot training & validation accuracy values (of first char only)
plt.plot(history.history['val_output_0_accuracy'])
plt.plot(history.history['val_output_0_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values (of first char only)
plt.plot(history.history['output_0_loss'])
plt.plot(history.history['val_output_0_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# +
nb_words_to_test = 100000
x_test = create_inputs(nb_words_to_test, nb_chars, nb_letters)
x_test_scaled = scaler.transform(x_test)
y_test_raw = encrypt(x_test, nb_words_to_test, nb_chars)
y_test_raw_cate = keras.utils.to_categorical(y_test_raw, nb_letters)
# process the y data as useful ANN multiple-outputs data
y_test = []
for c in range(nb_chars):
# extract each 'char' colomn from the global y_train0 tensor
# in order to have multiplue yi_train outputs tensors
yi_test = y_test_raw_cate[:,c,:]
y_test.append(yi_test)
print('\n# Evaluate on test data')
results = coding_model.evaluate(x_test_scaled, y_test, batch_size=128)
for r in range(len(results)):
print(coding_model.metrics_names[r],':',results[r])
# +
nb_words_to_test = 3
x_test = create_inputs(nb_words_to_test, nb_chars, nb_letters)
print_readable_inputs(x_test)
print("x_test=\n", x_test)
x_test_scaled = scaler.transform(x_test)
print("x_test_scaled=\n", x_test_scaled)
print('-->')
prediction = coding_model.predict(x_test_scaled)
#print(prediction)
print('prediction')
print_readable_outputs(prediction, nb_words_to_test, nb_chars)
print('check prediction')
y_test = encrypt(x_test, nb_words_to_test, nb_chars)
print("y_test=\n", y_test)
|
multiple_outputs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
# Read School and Student Data File and store into Pandas DataFrames
school_data = pd.read_csv(school_data_to_load)
student_data = pd.read_csv(student_data_to_load)
# Combine the data into a single dataset.
school_data_complete = pd.merge(student_data, school_data, how="left", on=["school_name", "school_name"])
# -
school_data_complete.head(3)
# ## District Summary
#
# * Calculate the total number of schools
#
# * Calculate the total number of students
#
# * Calculate the total budget
#
# * Calculate the average math score
#
# * Calculate the average reading score
#
# * Calculate the percentage of students with a passing math score (70 or greater)
#
# * Calculate the percentage of students with a passing reading score (70 or greater)
#
# * Calculate the percentage of students who passed math **and** reading (% Overall Passing)
#
# * Create a dataframe to hold the above results
#
# * Optional: give the displayed data cleaner formatting
tot_num_schools = school_data_complete["School ID"].nunique()
tot_num_students = school_data_complete["Student ID"].count()
tot_budget = sum(school_data_complete["budget"].unique())
avg_math_score = school_data_complete["math_score"].mean()
avg_read_score = school_data_complete["reading_score"].mean()
perc_math_pass = sum(school_data_complete.math_score >= 70)/tot_num_students
perc_read_pass = sum(school_data_complete.reading_score >= 70)/tot_num_students
overall_pass = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)]['student_name'].count()/tot_num_students
district_summary_df = pd.DataFrame({"Total Schools": [tot_num_schools],
"Total Students": [tot_num_students],
"Total Budget": [tot_budget],
"Average Math Score": [avg_math_score],
"Average Reading Score": [avg_read_score],
"% Passing Math": [perc_math_pass],
"% Passing Reading": [perc_read_pass],
"% Overall Passing": [overall_pass]})
district_summary_df = district_summary_df.style.format({"Total Budget": "${:,.2f}",
"Total Students": "{:,.0f}",
"Average Reading Score": "{:.2f}",
"Average Math Score": "{:.2f}",
"% Passing Math": "{:.2%}",
"% Passing Reading": "{:.2%}",
"% Overall Passing": "{:.2%}"})
district_summary_df
# ## School Summary
# * Create an overview table that summarizes key metrics about each school, including:
# * School Name
# * School Type
# * Total Students
# * Total School Budget
# * Per Student Budget
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * % Overall Passing (The percentage of students that passed math **and** reading.)
#
# * Create a dataframe to hold the above results
school_summary = school_data_complete.groupby(["school_name"])
school_type = school_summary["type"].unique()
students_per_school = school_summary["Student ID"].count()
school_budget = school_summary["budget"].unique()
per_student_budget = school_budget / students_per_school
avg_math_school = school_summary["math_score"].mean()
avg_read_school = school_summary["reading_score"].mean()
math_pass_school = school_data_complete[school_data_complete['math_score'] >= 70].groupby('school_name')['Student ID'].count()/students_per_school
read_pass_school = school_data_complete[school_data_complete['reading_score'] >= 70].groupby('school_name')['Student ID'].count()/students_per_school
overall_pass_school = school_data_complete[(school_data_complete['math_score'] >= 70) & (school_data_complete['reading_score'] >= 70)].groupby('school_name')['Student ID'].count()/students_per_school
school_summary_df = pd.DataFrame({"School Type": school_type,
"Total Students": students_per_school,
"Total School Budget": school_budget,
"Per Student Budget": per_student_budget,
"Average Math Score": avg_math_school,
"Average Reading Score": avg_read_school,
"% Passing Math": math_pass_school,
"% Passing Reading": read_pass_school,
"% Overall Passing": overall_pass_school})
school_summary_df.index.name = None
school_style_df = school_summary_df.copy().head(5)
school_style_df['School Type'] = school_style_df['School Type'].astype(str).str[1:-1]
school_style_df['School Type'] = school_style_df['School Type'].astype(str).str[1:-1]
school_style_df['Total School Budget'] = school_style_df['Total School Budget'].astype(str).str[1:-1]
school_style_df['Per Student Budget'] = school_style_df['Per Student Budget'].astype(str).str[1:-1]
school_style_df.loc[:,"Total School Budget"] = school_style_df["Total School Budget"].astype(float).map("${:,.2f}".format)
school_style_df.loc[:,"Per Student Budget"] = school_style_df["Per Student Budget"].astype(float).map("${:,.2f}".format)
school_style_df = school_style_df.style.format({"Average Math Score": "{:.2f}",
"Average Reading Score": "{:.2f}",
"% Passing Math": "{:.2%}",
"% Passing Reading": "{:.2%}",
"% Overall Passing": "{:.2%}"})
school_style_df
# ### Another approach
school_data_df = school_data_complete.copy()
school_data_df["passing_math"] = school_data_df["math_score"] >= 70
school_data_df["passing_reading"] = school_data_df["reading_score"] >= 70
school_group = school_data_df.groupby(["school_name"]).mean()
school_group["Per Student Budget"] = school_group["budget"]/school_group["size"]
school_group["% Passing Math"] = round(school_group["passing_math"]*100,2)
school_group["% Passing Reading"] = round(school_group["passing_reading"]*100,2)
school_group["% Overall Passing"] = round(school_data_df[(school_data_df['math_score'] >= 70) & (school_data_df['reading_score'] >= 70)].groupby('school_name')['Student ID'].count()/school_group["size"]*100,2)
school_data_summary = pd.merge(school_group, school_data, how="left", on=["school_name", "school_name"])
school_summary_dataframe = pd.DataFrame({"School Name": school_data_summary["school_name"],
"School Type": school_data_summary["type"],
"Total Students":school_data_summary["size_x"],
"Total School Budget": school_data_summary["budget_x"],
"Per Student Budget":school_data_summary["Per Student Budget"],
"Average Math Score":round(school_data_summary["math_score"],2),
"Average Reading Score":round(school_data_summary["reading_score"],2),
"% Passing Math": school_data_summary["% Passing Math"],
"% Passing Reading": school_data_summary["% Passing Reading"],
"% Overall Passing": school_data_summary["% Overall Passing"]})
school_summary_df_formatted = school_summary_dataframe.copy()
school_summary_df_formatted["Total Students"] = school_summary_df_formatted["Total Students"].map("{:,.0f}".format)
school_summary_df_formatted["Total School Budget"] = school_summary_df_formatted["Total School Budget"].map("${:,.2f}".format)
school_summary_df_formatted["Per Student Budget"] = school_summary_df_formatted["Per Student Budget"].map("${:,.2f}".format)
#Display
school_summary_df_formatted.head()
# ## Top Performing Schools (By % Overall Passing)
# * Sort and display the top five performing schools by % overall passing.
school_summary_df.sort_values(by='% Overall Passing', ascending=False, inplace=True)
top_schools = school_summary_df.copy().head(5)
top_schools['School Type'] = top_schools['School Type'].astype(str).str[1:-1]
top_schools['School Type'] = top_schools['School Type'].astype(str).str[1:-1]
top_schools['Total School Budget'] = top_schools['Total School Budget'].astype(str).str[1:-1]
top_schools['Per Student Budget'] = top_schools['Per Student Budget'].astype(str).str[1:-1]
top_schools.loc[:,"Total School Budget"] = top_schools["Total School Budget"].astype(float).map("${:,.2f}".format)
top_schools.loc[:,"Per Student Budget"] = top_schools["Per Student Budget"].astype(float).map("${:,.2f}".format)
top_schools = top_schools.style.format({"Average Math Score": "{:.2f}",
"Average Reading Score": "{:.2f}",
"% Passing Math": "{:.2%}",
"% Passing Reading": "{:.2%}",
"% Overall Passing": "{:.2%}"})
top_schools
# + [markdown] tags=[]
# ### Another approach
# -
top_schools = school_summary_dataframe.sort_values(["% Overall Passing"], ascending=False)
top_schools.head()
# ## Bottom Performing Schools (By % Overall Passing)
# * Sort and display the five worst-performing schools by % overall passing.
school_summary_df.sort_values(by='% Overall Passing', ascending=True, inplace=True)
bottom_schools = school_summary_df.copy().head(5)
# + tags=[]
bottom_schools['School Type'] = bottom_schools['School Type'].astype(str).str[1:-1]
bottom_schools['School Type'] = bottom_schools['School Type'].astype(str).str[1:-1]
bottom_schools['Total School Budget'] = bottom_schools['Total School Budget'].astype(str).str[1:-1]
bottom_schools['Per Student Budget'] = bottom_schools['Per Student Budget'].astype(str).str[1:-1]
bottom_schools.loc[:,"Total School Budget"] = bottom_schools["Total School Budget"].astype(float).map("${:,.2f}".format)
bottom_schools.loc[:,"Per Student Budget"] = bottom_schools["Per Student Budget"].astype(float).map("${:,.2f}".format)
bottom_schools = bottom_schools.style.format({"Average Math Score": "{:.2f}",
"Average Reading Score": "{:.2f}",
"% Passing Math": "{:.2%}",
"% Passing Reading": "{:.2%}",
"% Overall Passing": "{:.2%}"})
bottom_schools
# + [markdown] tags=[]
# ### Another approach
# -
bottom_schools = school_summary_dataframe.sort_values(["% Overall Passing"], ascending=True)
bottom_schools.head()
# ## Math Scores by Grade
# * Create a table that lists the average Math Score for students of each grade level (9th, 10th, 11th, 12th) at each school.
#
# * Create a pandas series for each grade. Hint: use a conditional statement.
#
# * Group each series by school
#
# * Combine the series into a dataframe
#
# * Optional: give the displayed data cleaner formatting
nine_grade_math = school_data_complete[school_data_complete["grade"] == "9th"].groupby("school_name").mean()["math_score"]
ten_grade_math = school_data_complete[school_data_complete["grade"] == "10th"].groupby("school_name").mean()["math_score"]
eleven_grade_math = school_data_complete[school_data_complete["grade"] == "11th"].groupby("school_name").mean()["math_score"]
twelve_grade_math = school_data_complete[school_data_complete["grade"] == "12th"].groupby("school_name").mean()["math_score"]
math_by_school_df = round(pd.DataFrame({"9th":nine_grade_math, "10th":ten_grade_math, "11th":eleven_grade_math, "12th":twelve_grade_math}), 2)
math_by_school_df.index.name = None
math_by_school_df.head(5)
# ## Reading Score by Grade
# * Perform the same operations as above for reading scores
nine_grade_read = school_data_complete[school_data_complete["grade"] == "9th"].groupby("school_name").mean()["reading_score"]
ten_grade_read = school_data_complete[school_data_complete["grade"] == "10th"].groupby("school_name").mean()["reading_score"]
eleven_grade_read = school_data_complete[school_data_complete["grade"] == "11th"].groupby("school_name").mean()["reading_score"]
twelve_grade_read = school_data_complete[school_data_complete["grade"] == "12th"].groupby("school_name").mean()["reading_score"]
read_by_school_df = round(pd.DataFrame({"9th":nine_grade_read, "10th":ten_grade_read, "11th":eleven_grade_read, "12th":twelve_grade_read}), 2)
read_by_school_df.index.name = None
read_by_school_df.head(5)
# ## Scores by School Spending
# * Create a table that breaks down school performances based on average Spending Ranges (Per Student). Use 4 reasonable bins to group school spending. Include in the table each of the following:
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * Overall Passing Rate (Average of the above two)
spending_df = school_summary_dataframe.copy()
spending_df.head(2)
spending_bins = [0, 583.99, 628.99, 643.99, 675]
spending_bin_names = ["<$584", "$585-629", "$630-644", "$645-675"]
spending_df["Spending Ranges (Per Student)"] = pd.cut(spending_df["Per Student Budget"],spending_bins, labels=spending_bin_names)
spending_df_grouped = spending_df.groupby("Spending Ranges (Per Student)").mean().round(2)
spending_df_grouped.drop(['Total Students', 'Total School Budget', 'Per Student Budget'], axis=1, inplace=True)
spending_df_grouped
# ## Scores by School Size
# * Perform the same operations as above, based on school size.
size_df = school_summary_dataframe.copy()
size_df["size"] = school_data_summary["size_x"]
size_df.head(2)
size_bins = [0, 999.99, 1999.99, 5000]
size_bin_names = ["Small (<1000)", "Medium (1000-2000)", "Large (2000-5000)"]
size_df["School Size"] = pd.cut(size_df["size"],size_bins, labels=size_bin_names)
size_df_grouped = size_df.groupby("School Size").mean().round(2)
size_df_grouped.drop(['size', 'Total Students', 'Total School Budget', 'Per Student Budget'], axis=1, inplace=True)
size_df_grouped
# ## Scores by School Type
# * Perform the same operations as above, based on school type
type_df = school_summary_dataframe.copy()
type_df["School Type"] = school_data_summary["type"]
type_df.head(2)
type_df_grouped = type_df.groupby("School Type").mean().round(2)
type_df_grouped.drop(['Total Students', 'Total School Budget', 'Per Student Budget'], axis=1, inplace=True)
type_df_grouped
|
PyCitySchools/PyCitySchools_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
import requests
from io import TextIOWrapper
from zipfile import ZipFile
import io
from matplotlib.colors import LinearSegmentedColormap
from numpy import array
from numpy import argmax
from sklearn import tree
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import precision_score, recall_score, f1_score
from sklearn.metrics import confusion_matrix
# Files were previously uploaded from data cleaning onto Github repository.
# Just need to pull them up with the following:
feat_url = "https://raw.githubusercontent.com/jasmultani5391/Census-Data/master/featDF.csv"
download1 = requests.get(feat_url).content
complete_url = "https://raw.githubusercontent.com/jasmultani5391/Census-Data/master/completeDF.csv"
download2 = requests.get(complete_url).content
# Read the downloaded content and turn it into a pandas dataframe
featDF = pd.read_csv(io.StringIO(download1.decode('utf-8')))
completeDF = pd.read_csv(io.StringIO(download2.decode('utf-8')))
#print(featDF.head(4))
#print(completeDF.head(4))
# +
# First, we must find the best K index that gives us the highest
# accuracy. We previously created a class called NearestK that contains
# the method to search for the best K value to use. We'll use the featDF
# over the completeDF because the former contains all the one-hot encoded
# columns that converted qualitative to quantitative data.
labelDF = featDF['salary_label']
featDF = featDF.drop(['salary_label'],
axis=1
)
# +
trndfdata, tstdfdata, trndflbl, tstdflbl = train_test_split(featDF,
labelDF,
random_state=0)
rf_classifier = RandomForestClassifier(min_samples_leaf=70,
n_estimators=150,
bootstrap=True,
oob_score=True, #use out-of-bag samples
n_jobs=-1,
random_state=0,
max_features='sqrt'
)
# Fit classifier on train dataset.
rf_classifier.fit(trndfdata, np.ravel(trndflbl))
# Score based on train dataset.
train_df_score = rf_classifier.score(trndfdata, trndflbl)
print('Train Accuracy Score %: ' + str(round(train_df_score*100, 4)))
predict_trainlabels = rf_classifier.predict(trndfdata)
trainprecision = precision_score(trndflbl, predict_trainlabels, average='binary')
print('Train Precision %: ' + str(round(trainprecision*100, 4)))
trainrecall = recall_score(trndflbl, predict_trainlabels, average='binary')
print('Train Recall %: ' + str(round(trainrecall*100, 4)))
trainf1 = f1_score(trndflbl, predict_trainlabels, average='binary')
print('Train F1 %: ' + str(round(trainf1*100, 4)))
print('\n\n')
# Score based on test dataset.
test_df_score = rf_classifier.score(tstdfdata, tstdflbl)
print('Test Accuracy Score %: ' + str(round(test_df_score*100, 4)))
predict_testlabels = rf_classifier.predict(tstdfdata)
testprecision = precision_score(tstdflbl, predict_testlabels, average='binary')
print('Test Precision %: ' + str(round(testprecision*100, 4)))
testrecall = recall_score(tstdflbl, predict_testlabels, average='binary')
print('Test Recall %: ' + str(round(testrecall*100, 4)))
testf1 = f1_score(tstdflbl, predict_testlabels, average='binary')
print('Test F1 %: ' + str(round(testf1*100, 4)))
# Most important coefficients
important_coefficients = rf_classifier.feature_importances_
feature_columns = list(featDF.head(0))
importantcoef_columns = dict(zip(feature_columns, important_coefficients))
importantcoef_series = pd.Series(importantcoef_columns)
importantcoef_series = importantcoef_series.sort_values(ascending=False)
importantcoef = dict(importantcoef_series)
print(importantcoef_series)
# Confirm we have the total number of features included, which should be 45.
print(len(importantcoef))
# -
|
Decision Forest Classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Detect and Delete outliers with Optimus
# An outlier is an observation that lies an abnormal distance from other values in a random sample from a population. In a sense, this definition leaves it up to the analyst (or a consensus process) to decide what will be considered abnormal. Before abnormal observations can be singled out, it is necessary to characterize normal observations.
#
# You have to be careful when studying outliers because how do you know if an outlier is the result of a data glitch, or a real data point -- indeed maybe not an outlier.
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("..")
from optimus import Optimus
# Create optimus
op = Optimus()
# +
from pyspark.sql.types import *
df = op.create.df(
[
("words", "str", True),
("num", "int", True),
("animals", "str", True),
("thing", StringType(), True),
("two strings", StringType(), True),
("filter", StringType(), True),
("num 2", "int", True),
("date", "string", True),
("num 3", "str", True)
],[
(" I like fish ", 1, "dog", "&^%$#housé", "cat-car", "a",1, "20150510", '3'),
(" zombies", 2, "cat", "tv", "dog-tv", "b",2, "20160510", '3'),
("simpsons cat lady", 2, "frog", "table","eagle-tv-plus","1",3, "20170510", '4'),
(None, 3, "eagle", "glass", "lion-pc", "c",4, "20180510", '5'),
(None, 5, "eagle", "glass", "lion-pc", "c",4, "20180510", '5'),
(None, 6, "eagle", "glass", "lion-pc", "c",4, "20180510", '5'),
(None, 7, "eagle", "glass", "lion-pc", "c",4, "20180510", '5'),
(None, 20, "eagle", "glass", "lion-pc", "c",4, "20180510", '5')
]
)
df.table()
# -
# From a quick inspection of the dataframe we can guess that the 1000 in the column `num` can be an outlier. You can perform a very intense search to see if it is actually and outlier, if you need something like that please check out [these articles and tutorials](http://www.datasciencecentral.com/profiles/blogs/11-articles-and-tutorials-about-outliers)
# With optimus you can perform several analysis too to check if a value is an outlier. First lets run some visual analysis. Remember to check the [Main Example](https://github.com/ironmussa/Optimus/blob/master/examples/Optimus_Example.ipynb) for more.
# ## Outlier detection
# One of the commonest ways of finding outliers in one-dimensional data is to mark as a potential outlier any point that is more than two standard deviations, say, from the mean (I am referring to sample means and standard deviations here and in what follows). But the presence of outliers is likely to have a strong effect on the mean and the standard deviation, making this technique unreliable.
# That's why we have programmed in Optimus the median absolute deviation from median, commonly shortened to the median absolute deviation (MAD). It is the median of the set comprising the absolute values of the differences between the median and each data point. If you want more information on the subject please read the amazing article by Leys et al. about dtecting outliers [here](http://www.sciencedirect.com/science/article/pii/S0022103113000668)
from optimus.outliers.outliers import OutlierDetector
od = OutlierDetector()
# ### Zscore
od.z_score(df, "num", threshold=1).table()
od.z_score(df, ["num", "num 2"],1).table()
# ### IQR
od.iqr(df, "num").table()
# ### MAD
od.mad(df, "num", 1).table()
# ### Modified Zscore
od.modified_z_score(df, "num", 1).table()
|
examples/new-api-outliers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iAy5RvwucBoD"
# Imports
import torch
import torchvision # torch package for vision related things
import torch.nn.functional as F # Parameterless functions, like (some) activation functions
import torchvision.datasets as datasets # Standard datasets
import torchvision.transforms as transforms # Transformations we can perform on our dataset for augmentation
from torch import optim # For optimizers like SGD, Adam, etc.
from torch import nn # All neural network modules
from torchsummary import summary
from torch import Tensor
from torch.utils.data import DataLoader # Gives easier dataset managment by creating mini batches etc.
from tqdm import tqdm # For nice progress bar!
import os
import numpy as np
import matplotlib.pyplot as plt
import time
from typing import Union
# + colab={"base_uri": "https://localhost:8080/", "height": 343} id="QPyCoRdSfF-p" outputId="2fdea71e-babe-4999-e643-929a1bf30262"
################################################# datasets #################################################
# Path and Hyperparameters
#os.listdir('./drive/MyDrive/21-2-ML/')
storage = './drive/MyDrive/21-2-ML/CIFAR100/'
in_channels = 3
num_class = 100
BS = 64
# Apply Transformation :
img_transformation = transforms.Compose([
#transforms.RandomAffine(degrees=(-10,10), translate=(0, 0.05)),
transforms.RandomHorizontalFlip(p = 0.5),
transforms.ToTensor()
])
# Prepare the Dataset :
train_data = datasets.CIFAR100(root = storage, train = True, transform = img_transformation, download = True)
test_data = datasets.CIFAR100(root = storage, train = False, transform = img_transformation, download = True)
train_loader = DataLoader(dataset=train_data, batch_size=BS, shuffle=True)
test_loader = DataLoader(dataset=test_data, batch_size=BS, shuffle=True) # Shuffle for every epoch
# Check the prepared dataset : iter through the training loader
dataiter = iter(train_loader)
img, lab = dataiter.next()
def imshow(img):
img = img / 2 + 0.5 # unnormalize. Dataset images are already normalized!!
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# show images
imshow(torchvision.utils.make_grid(img))
# print labels
print(" ".join('%5s' % lab[j] for j in range(BS)))
# + colab={"base_uri": "https://localhost:8080/"} id="OZqjLaL5nALG" outputId="77b7c3e4-c1c9-4ac4-fe18-42080521e59c"
print(lab.unique()) # Okay. labels are wetted properly.
print(img[0].shape)
# + colab={"base_uri": "https://localhost:8080/"} id="3vxKPkSlk-nD" outputId="0ffc21d5-c21e-4919-bf2a-33bba8e024fc"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Conv3x3 block
def conv3x3(in_ch, out_ch, stride = 1) ->nn.Conv2d:
return nn.Conv2d(in_ch, out_ch, kernel_size=3, stride=stride, padding=1, bias=False)
# Conv1x1 block
def conv1x1(in_ch, out_ch, stride = 1) ->nn.Conv2d:
return nn.Conv2d(in_ch, out_ch, kernel_size=1, stride=stride, bias=False)
# Basic Residual block for ResNet 18 / 34
class ResBlock(nn.Module):
expansion = 1
def __init__(self, in_ch, out_ch, stride=1, downsample = None):
super(ResBlock, self).__init__()
# Sequential(residual fcn)
self.residualBlock = nn.Sequential(
conv3x3(in_ch, out_ch, stride=stride),
nn.BatchNorm2d(out_ch),
nn.LeakyReLU(negative_slope=0.01, inplace=True),
conv3x3(out_ch, out_ch),
nn.BatchNorm2d(out_ch)
)
self.downsample = downsample
def forward(self, x):
identity = x
out = self.residualBlock(x)
if self.downsample is not None:
identity = self.downsample(x)
out += identity
out = nn.ReLU(inplace=True)(out)
return out
# Bottleneck block for ResNet 50 / 101 / 152
class BottleNeck(nn.Module):
expansion = 4
def __init__(self, in_ch, out_ch, stride = 1, downsample = None):
super(BottleNeck, self).__init__()
self.residualBlock = nn.Sequential(
conv1x1(in_ch, out_ch),
nn.BatchNorm2d(out_ch),
nn.LeakyReLU(negative_slope=0.01, inplace=True),
conv3x3(out_ch, out_ch, stride=stride),
nn.BatchNorm2d(out_ch),
nn.LeakyReLU(negative_slope=0.01, inplace=True),
conv1x1(out_ch, out_ch*BottleNeck.expansion),
nn.BatchNorm2d(out_ch*BottleNeck.expansion)
)
self.downsample = downsample
def forward(self, x):
identity = x
out = self.residualBlock(x)
if self.downsample is not None:
identity = self.downsample(x)
out += identity
out = nn.ReLU(inplace=True)(out)
return out
class ResNet(nn.Module):
def __init__(self, block, n_layers, n_class = 100):
super(ResNet, self).__init__()
# The initial Conv layer
self.in_ch = 16
self.conv = conv3x3(3,16)
self.BN = nn.BatchNorm2d(16)
self.relu = nn.ReLU(inplace=True)
# Blocks.
self.layer1 = self.make_layer(block, 32, n_layers[0])
self.layer2 = self.make_layer(block, 64, n_layers[1], stride=2)
self.layer3 = self.make_layer(block, 128, n_layers[2], stride=2)
self.layer4 = self.make_layer(block, 256, n_layers[3], stride=2)
# Output layer elements
self.pool = nn.AdaptiveAvgPool2d((1,1)) # The output size.
self.fc1 = nn.Linear(256*block.expansion, n_class)
self.drop = nn.Dropout(p=0.5)
def make_layer(self, block, out_ch, n_block, stride = 1):
'''
:param block: block type, ResBlock or BottleNeck
:param out_ch: numbers of output channels
:param n_block: number of blocks that will be stacked in the layer
:param stride: stride of the initial block of the layer.
:return: The entire ResNet model.
'''
downsample = None
# Usage : for example, output of layer 1 shape = ( , 16, 32, 32)
# 1) Here self.in_ch = 16
# 2) out_ch of layer 2 = 32, hence downsample will be
# conv3x3(in = 16, out = 32, stride = 2) and added in the first block of the layer.
if (stride!=1) or (self.in_ch!=out_ch):
downsample = nn.Sequential(
conv3x3(self.in_ch, out_ch*block.expansion, stride=stride),
nn.BatchNorm2d(out_ch*block.expansion)
)
layers = []
layers.append(block(self.in_ch, out_ch, stride, downsample)) # Add the block with downsample layer!
self.in_ch = out_ch*block.expansion
# Repeat blocks with the given list of numbers :
for i in range(n_block):
layers.append(block(self.in_ch, out_ch))
self.in_ch = out_ch*block.expansion
return nn.Sequential(*layers)
def forward(self, x):
out = self.conv(x)
out = self.BN(out)
out = self.relu(out)
# Body
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
# Output
out = self.pool(out)
out = out.view(out.size(0), -1)
out = self.drop(out)
out = self.fc1(out)
return out
def ResNet18():
return ResNet(ResBlock, [2,2,2,2])
def ResNet34():
return ResNet(ResBlock, [3,4,6,3])
def ResNet50():
return ResNet(BottleNeck, [3,4,6,3])
def ResNet101():
return ResNet(BottleNeck, [3,4,23,3])
def ResNet152():
return ResNet(BottleNeck, [3,8,36,3])
test_X = torch.randn(12, 3, 32, 32).to(device)
model = ResNet50()
model = model.to(device)
out = model(test_X)
print(out.shape)
summary(model, (3,32,32), device = device.type)
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="rmVj5tnI4_wr" outputId="49b6dead-b100-4ba7-81a8-859736a962c0"
'''model = ResNet(ResBlock, [3,3,3]).to(device)
X = torch.randn(16, 3, 32, 32).to(device)
output = model(X)
print(output.size(),'\n')
summary(model, (3,32,32), device=device.type)'''
# + id="0TQOjPe6orgB"
lr_init = 0.0009
############################ Loss function and optimization #############################
loss_func = nn.CrossEntropyLoss(reduction = 'sum') # Sum-up the CE loss in the minibatch
optimizer = torch.optim.Adam(model.parameters(), lr = lr_init)
##################################### Lr scheduling #####################################
# print(optimizer.param_groups) All the layer weights.. and Hyperparameters!
from torch.optim.lr_scheduler import ReduceLROnPlateau
# patience : n epoch동안 loss reduction 없으면
# factor : factor만큼 lr에 multplication함
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.3, patience=5)
######################### Function that calculates lr and loss ##########################
def get_lr(opt : optimizer) -> float:
for param_group in opt.param_groups:
return param_group['lr']
def batch_metric(prediction, target): # Returns correct number of data in a minibatch.
onehot_pred = prediction.argmax(1, keepdim = True) # Extracts maximum prob. index
corrects = onehot_pred.eq(target.view_as(onehot_pred)).sum().item()
# .sum() -> Tensor(scalar) --> .item()->scalar!
return corrects # Therefore metric == accuracy, and passed trough scheduler
def batch_loss(loss_func, prediction, target, optim = None):
loss = loss_func(prediction, target)
corrects = batch_metric(prediction, target)
if optim is not None: # In the training step
optim.zero_grad() # zero-initialize gradients(since pytorch sums-up gradients)
loss.backward() # BP
optim.step() # https://tutorials.pytorch.kr/beginner/pytorch_with_examples.html
return loss.item(), corrects # loss.item() returns a scarlar value.
# + id="CMASSAbgmyZ8"
###################### Get loss per epoch and return metric #######################
def loss_in_epoch(model, loss_func, train_dataloader, optimizer = None) -> int:
running_loss = 0.0
running_corrects = 0.0
len_data = len(train_dataloader.dataset) # .dataset = attibutes of the whole dataset.
for X_batch, Y_batch in train_dataloader:
X_batch = X_batch.to(device)
Y_batch = Y_batch.to(device)
Y_pred = model(X_batch)
BatchLoss_train, corrects = batch_loss(loss_func, Y_pred, Y_batch, optimizer)
running_loss += BatchLoss_train
#print(BatchLoss_train)
if corrects is not None:
running_corrects += corrects
loss = running_loss / len_data # Aggregate all losses in epoch -> divide to len(dataset)
metric = running_corrects / len_data # Aggregate all corrects in '' -> divide to ''
return loss, metric
################################ Training Function ################################
def train_model(model, params : dict) -> Union[model, dict, dict] : # Multiple output annotation : Union[]
# Assert parameters.
n_epoch = params['n_epoch']
optimizer = params['optimizer']
loss_func = params['loss_func']
train_dl = params['train_dl']
val_dl = params['val_dl']
lr_scheduler= params['lr_scheduler']
# To store losses and accuracy:
loss_history = {'train' : [], 'val' : []}
metric_history = {'train' : [], 'val' : []}
# Record the best loss:
best_loss = float('inf')
start_time = time.time()
for epoch in range(n_epoch):
current_lr = get_lr(optimizer)
print(f"Epoch [{epoch+1}/{n_epoch}], current lr = {current_lr:.8f}")
model.train()
train_loss, train_metric = loss_in_epoch(model, loss_func, train_dl, optimizer)
loss_history['train'].append(train_loss)
metric_history['train'].append(train_metric)
with torch.no_grad(): # Off all layers, i.e. stop update
model.eval() # Since Dropout/norm.layers added, we turn off them in the eval.step.
val_loss, val_metric = loss_in_epoch(model, loss_func, val_dl)
loss_history['val'].append(val_loss)
metric_history['val'].append(val_metric)
#if val_loss < best_loss :
# Here we aim to implement a storing function of the best model weights
# pass
lr_scheduler.step(val_loss) # Update lr_scheduler with metric = Accuracy
print(f'training loss = {train_loss:.6f}, validation loss = {val_loss:.6f}, \
accuracy = {100*val_metric:.4f}, time = {(time.time() - start_time):.3f} sec' )
print('--'*10)
return model, loss_history, metric_history
# + colab={"base_uri": "https://localhost:8080/", "height": 71} id="f3C2ym5hzM_K" outputId="3fdd5491-26c3-4907-ba0c-aff5ccd23bc5"
'''testlr = []
test_scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, gamma = 0.95)
for i in range(100):
optimizer.step()
testlr.append(optimizer.param_groups[0]['lr'])
#print(len(optimizer.param_groups))
test_scheduler.step()
plt.plot(range(100), testlr)'''
# + colab={"base_uri": "https://localhost:8080/"} id="8u0fILtck84u" outputId="1d3e7589-eba5-4bed-b589-304f3efe6979"
params_train = {'n_epoch' : 50,
'optimizer' : optimizer, # Adam(model.parameters(), lr = lr_init)
'loss_func' : loss_func, # CrossEntropyLoss(reduction = 'sum')
'train_dl' : train_loader,
'val_dl' : test_loader, # Here i use validation set as a test set. In practice, NEVER DO THIS!
'lr_scheduler' : scheduler #
}
model, loss_hist, metric_hist = train_model(model, params_train)
# + id="6u1Iko8tQEJy" colab={"base_uri": "https://localhost:8080/"} outputId="8d4e82fd-bfdc-40ce-f5cb-3e26bd993613"
PATH = './drive/MyDrive/21-2-ML/CIFAR100/Res18_CIFAR100_ckpt'
torch.save(model.state_dict(), PATH) # Save the entire model
test_model = ResNet50().to(device)
test_model.load_state_dict(torch.load(PATH))
test_model.eval()
# + id="r1n2UeUeXP3t" colab={"base_uri": "https://localhost:8080/"} outputId="de1596fc-69f0-45af-d940-7a31d2405deb"
for i in range(20):
n_accurate = 0
n_total = 0
# Sum of the accurate samples / Total test samples*100
for X, Y in test_loader:
n_total += len(X)
n_accurate += batch_metric(test_model(X.to(device)), Y.to(device))
print(n_accurate/n_total*100)
|
CIFAR10 modulization/PyTorch_ResNet_Full.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parameter identification example
#
# Here is a simple toy model that we use to demonstrate the working of the inference package
#
# $\emptyset \xrightarrow[]{k_1} X \; \; \; \; X \xrightarrow[]{d_1} \emptyset$
#
# ### Run the MCMC algorithm to identify parameters from the experimental data
#
# In this demonstration, we will try to use multiple trajectories of data taken under multiple initial conditions and different length of time points?
# +
# %matplotlib inline
# %config InlineBackend.figure_format = "retina"
from matplotlib import rcParams
rcParams["savefig.dpi"] = 100
rcParams["figure.dpi"] = 100
rcParams["font.size"] = 20
# -
# ## Using Gaussian prior for `k1`
# +
# %matplotlib inline
import bioscrape as bs
from bioscrape.types import Model
from bioscrape.inference import py_inference
import numpy as np
import pylab as plt
import pandas as pd
# Import a bioscrape/SBML model
M = Model(sbml_filename = 'toy_sbml_model.xml')
# Import data from CSV
# Import a CSV file for each experiment run
df = pd.read_csv('test_data.csv', delimiter = '\t', names = ['X','time'], skiprows = 1)
M.set_species({'X':df['X'][0]})
# Create prior for parameters
prior = {'d1' : ['gaussian', 0.2, 200]}
sampler, pid = py_inference(Model = M, exp_data = df, measurements = ['X'], time_column = ['time'],
nwalkers = 5, init_seed = 0.15, nsteps = 1500, sim_type = 'deterministic',
params_to_estimate = ['d1'], prior = prior)
# -
# # Using uniform priors and estimating both `k1` and `d1`
# ## and use the pid => parameter inference object directly.
# +
# %matplotlib inline
import bioscrape as bs
from bioscrape.types import Model
from bioscrape.inference import py_inference
import numpy as np
import pylab as plt
import pandas as pd
# Import a bioscrape/SBML model
M = Model(sbml_filename = 'toy_sbml_model.xml')
# Import data from CSV
# Import a CSV file for each experiment run
df = pd.read_csv('test_data.csv', delimiter = '\t', names = ['X','time'], skiprows = 1)
M.set_species({'X':df['X'][0]})
prior = {'d1' : ['uniform', 0, 10], 'k1' : ['uniform', 0, 100]}
sampler, pid = py_inference(Model = M, exp_data = df, measurements = ['X'], time_column = ['time'],
nwalkers = 20, init_seed = 0.15, nsteps = 5500, sim_type = 'deterministic',
params_to_estimate = ['d1', 'k1'], prior = prior)
# -
# ### Check mcmc_results.csv for the results of the MCMC procedure and perform your own analysis.
#
# ### You can also plot the results as follows
# +
from bioscrape.simulator import py_simulate_model
M_fit = Model(sbml_filename = 'toy_sbml_model.xml')
M_fit.set_species({'X':df['X'][0]})
timepoints = pid.timepoints
flat_samples = sampler.get_chain(discard=200, thin=15, flat=True)
inds = np.random.randint(len(flat_samples), size=200)
for ind in inds:
sample = flat_samples[ind]
for pi, pi_val in zip(pid.params_to_estimate, sample):
M_fit.set_parameter(pi, pi_val)
plt.plot(timepoints, py_simulate_model(timepoints, Model= M_fit)['X'], "C1", alpha=0.1)
# plt.errorbar(, y, yerr=yerr, fmt=".k", capsize=0)
# plt.plot(timepoints, list(pid.exp_data['X']), label = 'data')
plt.plot(timepoints, py_simulate_model(timepoints, Model = M)['X'], "k", label="original model")
plt.legend(fontsize=14)
plt.xlabel("Time")
plt.ylabel("[X]");
# -
flat_samples = sampler.get_chain(discard = 200, thin = 15,flat = True)
flat_samples
# ## Alll methods above have other advanced options that you can use. Refer to Parameter Identification Tools and Advanced Examples notebook for more details. There are many other tools available such as for multiple initial conditions and timepoints for each trajectory, options for the estimator etc.
|
inference examples/Gaussian prior example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
tf.set_random_seed(222)
# Predicting animal type based on various features
xy = np.loadtxt('data-04-zoo.csv', delimiter=',', dtype=np.float32)
x_data = xy[:, 0:-1]
y_data = xy[:, [-1]]
print(x_data.shape, y_data.shape)
nb_classes = 7
X = tf.placeholder(tf.float32, [None, 16])
Y = tf.placeholder(tf.int32, [None, 1]) #0~6
Y_one_hot = tf.one_hot(Y, nb_classes)
print("one hot: ",Y_one_hot)
Y_one_hot = tf.reshape(Y_one_hot, [-1, nb_classes])
print("reshape", Y_one_hot)
W = tf.Variable(tf.random_normal([16, nb_classes]), name= 'weight')
b = tf.Variable(tf.random_normal([nb_classes]), name = 'bias')
# +
# tf.nn.softmax computes softmax activations
# softmax = exp(logits) / reduce_sum(exp(logits), dim)
logits = tf.matmul(X,W) + b
hypothesis = tf.nn.softmax(logits)
# +
# Cross entropy cost/loss
cost_ini = tf.nn.softmax_cross_entropy_with_logits(logits = logits, labels = Y_one_hot)
cost = tf.reduce_mean(cost_ini)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(cost)
# -
prediction = tf.argmax(hypothesis, 1)
correct_prediction = tf.equal(prediction, tf.argmax(Y_one_hot, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for step in range(2000):
sess.run(optimizer, feed_dict = {X: x_data, Y: y_data})
if step % 100 == 0:
loss, acc = sess.run([cost, accuracy], feed_dict = {X: x_data, Y: y_data})
print("Step: {:5}\tLoss: {:.3f}\tAcc: {:.2%}".format(step, loss, acc))
#predict output
pred = sess.run(prediction, feed_dict = {X:x_data})
# y_data: (N,1) = flatten => (N, ) matches pred.shape
for p, y in zip(pred, y_data.flatten()):
print("[{}] Prediction: {} True Y: {}".format(p == int(y), p, int(y)))
|
Lab6_2. Zoo_Softmax_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import slug
import os
import glob
import h5py
import numpy as np
import matplotlib.pyplot as plt
from kungpao import io
from kungpao.display import display_single, IMG_CMAP, SEG_CMAP
from astropy.io import fits
from astropy import wcs
from astropy.table import Table, Column
from IPython.display import clear_output
# -
ra_cen = 129.603676 # ra of object
dec_cen = -1.606419 # dec of object
redshift = 0.052043
fits_data = fits.open('./Images/cut_Dragonfly_r.fits')
# +
if not os.path.isdir('HDF5'):
os.mkdir('HDF5')
f = h5py.File('./HDF5/Dragonfly_test.h5','w')
dt = h5py.special_dtype(vlen=str)
info = f.create_dataset('info', (10,2), dtype='S20')
info[0] = 'edition', 'Dragonfly'
info[1] = 'ra', ra_cen
info[2] = 'dec', dec_cen
info[3] = 'size (pix)', str([fits_data[0].shape[0], fits_data[0].shape[1]])
info[4] = 'redshift', redshift
g1 = f.create_group('Image')
g1.create_dataset('image', data=fits_data[0].data)
g1.create_dataset('image_header', data=fits_data[0].header.tostring(), dtype=dt)
f.close()
# -
f = h5py.File('./HDF5/Dragonfly_test.h5', 'r')
print(f['info'][:])
f.close()
# ### 1-D profile
# +
prefix = 'Dragonfly_test'
f = h5py.File('./HDF5/Dragonfly_test.h5', 'r+')
img = f['Image']['image'].value
w = wcs.WCS(f['Image']['image_header'].value)
g4 = f.create_group('Mask')
g5 = f.create_group('MaskedImage')
# phys_size
redshift = float(f['info'][4,1])
phys_size = slug.phys_size(redshift)
# extract obj
data = img
data = data.byteswap().newbyteorder()
objects, segmap = slug.extract_obj(
data,
b=15,
f=3,
sigma=3,
pixel_scale=slug.Dragonfly_pixel_scale,
deblend_cont=0.0001,
deblend_nthresh=128,
show_fig=False)
# make mask
seg_mask = slug.make_binary_mask(data, w, segmap, radius=1.0, show_fig=False, threshold=0.01, gaia=True)
# evaluate_sky
bkg_global = slug.evaluate_sky_dragonfly(data, b=15, f=3, sigma=1.5, radius=1.0,
threshold=0.005, show_fig=False, show_hist=False)
f['info'][5] = 'global bkg', bkg_global.globalback
f['info'][6] = 'global rms', bkg_global.globalrms
g4.create_dataset('mask', data=seg_mask)
HSC_mask = fits.open('./Data/msk_rebin_HSC_for_Dragonfly.fits')[0].data
g4.create_dataset('HSCmask', data=HSC_mask)
g5.create_dataset('maskedimage', data=(data-bkg_global.globalback)*(~seg_mask))
g5.create_dataset('HSCmaskedimage', data=(data-bkg_global.globalback)*(~HSC_mask))
# Save image and mask
if not os.path.isdir('Images'):
os.mkdir('Images')
if not os.path.isdir('Masks'):
os.mkdir('Masks')
img_fits = './Images/' + prefix + '_img.fits'
msk_fits = './Masks/' + prefix + '_msk.fits'
io.save_to_fits((data-bkg_global.globalback), img_fits, wcs=w)
io.save_to_fits(seg_mask.astype('uint8'), msk_fits, wcs=w)
display_single((data-bkg_global.globalback)*(~seg_mask))
plt.show(block=False)
# +
# Run ELLIPSE
phys_size = slug.phys_size(redshift)
iraf_path = '/Users/jiaxuanli/Research/slug/slug/iraf/macosx/'
ell_free, ell_fix = slug.run_SBP(
img_fits,
'./Data/msk_rebin_HSC_for_Dragonfly.fits',
slug.Dragonfly_pixel_scale,
phys_size,
iraf_path,
step=0.2,
sma_ini=5.0,
sma_max=80.0,
n_clip=1,
low_clip=3.0,
upp_clip=3.0,
outPre=prefix)
f['info'][7] = 'mean_e', ell_fix['ell'][10]
f['info'][8] = 'mean_pa', ell_fix['pa'][10]
f.create_dataset('ell_fix', data=ell_fix)
f.create_dataset('ell_free', data=ell_free)
f.close()
# -
f = h5py.File('./HDF5/Dragonfly_test.h5', 'r')
slug.h5_print_attrs(f)
print(f['info'].value)
f.close()
# +
f = h5py.File('./HDF5/Dragonfly_test.h5', 'r')
bkgval_header = float(fits.open('./Images/coadd_SloanR.fits')[0].header['BACKVAL'])
off_set = bkgval_header - float(f['info'].value[5][1])
# +
ellipse_fix = f['ell_fix'].value
img = f['Image']['image'].value
masked_img = f['MaskedImage']['maskedimage'].value
fig = plt.figure(figsize=(28, 8))
grid = plt.GridSpec(1, 14, wspace=0.1, hspace=0.1)
ax1 = fig.add_subplot(grid[0, 0:4])
ax1 = slug.display_isophote(
img,
ellipse_fix,
slug.Dragonfly_pixel_scale,
text='Dragonfly\ Image',
ax=ax1)
ax2 = fig.add_subplot(grid[0, 4:8])
ax2 = slug.display_isophote(
masked_img,
ellipse_fix,
slug.Dragonfly_pixel_scale,
text='Binary\ Masked',
ax=ax2,
circle=60)
ax3 = fig.add_subplot(grid[0, 9:])
ax3.tick_params(direction='in')
slug.SBP_single(ellipse_fix, redshift, slug.Dragonfly_pixel_scale, slug.Dragonfly_zeropoint_r, ax=ax3,
physical_unit=True, x_max=(200**0.25), vertical_line=False, show_dots=False, linecolor='firebrick',
linestyle='-', label="Dragonfly")
slug.SBP_single(ellipse_fix, redshift, slug.Dragonfly_pixel_scale, slug.Dragonfly_zeropoint_r, ax=ax3, offset=-off_set,
physical_unit=True, x_max=(200**0.25), vertical_line=False, show_dots=False, linecolor='gray',
linestyle='-.', label="Dragonfly\ BACKVAL")
plt.ylim(15.5, 33.5)
#plt.vlines((100 * 0.168)**0.25, 15.5, 32.5, linestyle='-.', label='60 arcsec')
ax3.invert_yaxis()
plt.subplots_adjust(hspace=0.)
#f.close()
#plt.savefig('./Figures/' + prefix + '.png', dpi=100, bbox_inches='tight')
# -
|
demo/Dragonfly-1D-profile-HDF5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 14
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### Code from previous chapters
def make_system(beta, gamma):
"""Make a system object for the SIR model.
beta: contact rate in days
gamma: recovery rate in days
returns: System object
"""
init = State(S=89, I=1, R=0)
init /= np.sum(init)
t0 = 0
t_end = 7 * 14
return System(init=init, t0=t0, t_end=t_end,
beta=beta, gamma=gamma)
def update_func(state, t, system):
"""Update the SIR model.
state: State (s, i, r)
t: time
system: System object
returns: State (sir)
"""
s, i, r = state
infected = system.beta * i * s
recovered = system.gamma * i
s -= infected
i += infected - recovered
r += recovered
return State(S=s, I=i, R=r)
def run_simulation(system, update_func):
"""Runs a simulation of the system.
system: System object
update_func: function that updates state
returns: TimeFrame
"""
unpack(system)
frame = TimeFrame(columns=init.index)
frame.row[t0] = init
for t in linrange(t0, t_end):
frame.row[t+1] = update_func(frame.row[t], t, system)
return frame
def calc_total_infected(results):
"""Fraction of population infected during the simulation.
results: DataFrame with columns S, I, R
returns: fraction of population
"""
return get_first_value(results.S) - get_last_value(results.S)
def sweep_beta(beta_array, gamma):
"""Sweep a range of values for beta.
beta_array: array of beta values
gamma: recovery rate
returns: SweepSeries that maps from beta to total infected
"""
sweep = SweepSeries()
for beta in beta_array:
system = make_system(beta, gamma)
results = run_simulation(system, update_func)
sweep[system.beta] = calc_total_infected(results)
return sweep
# ## SweepFrame
#
# The following sweeps two parameters and stores the results in a `SweepFrame`
def sweep_parameters(beta_array, gamma_array):
"""Sweep a range of values for beta and gamma.
beta_array: array of infection rates
gamma_array: array of recovery rates
returns: SweepFrame with one row for each beta
and one column for each gamma
"""
frame = SweepFrame(columns=gamma_array)
for gamma in gamma_array:
frame[gamma] = sweep_beta(beta_array, gamma)
return frame
# Here's what the results look like.
beta_array = linspace(0.1, 0.9, 11)
gamma_array = linspace(0.1, 0.7, 4)
frame = sweep_parameters(beta_array, gamma_array)
frame.head()
# And here's how we can plot the results.
# +
for gamma in gamma_array:
label = 'gamma = ' + str(gamma)
plot(frame[gamma], label=label)
decorate(xlabel='Contacts per day (beta)',
ylabel='Fraction infected',
loc='upper left')
# -
# It's often useful to separate the code that generates results from the code that plots the results, so we can run the simulations once, save the results, and then use them for different analysis, visualization, etc.
# ### Contact number
# After running `sweep_parameters`, we have a `SweepFrame` with one row for each value of `beta` and one column for each value of `gamma`.
frame.shape
# The following loop shows how we can loop through the columns and rows of the `SweepFrame`. With 11 rows and 4 columns, there are 44 elements.
for gamma in frame.columns:
series = frame[gamma]
for beta in series.index:
frac_infected = series[beta]
print(beta, gamma, frac_infected)
# Now we can wrap that loop in a function and plot the results. For each element of the `SweepFrame`, we have `beta`, `gamma`, and `frac_infected`, and we plot `beta/gamma` on the x-axis and `frac_infected` on the y-axis.
def plot_sweep_frame(frame):
"""Plot the values from a SweepFrame.
For each (beta, gamma), compute the contact number,
beta/gamma
frame: SweepFrame with one row per beta, one column per gamma
"""
for gamma in frame.columns:
series = frame[gamma]
for beta in series.index:
frac_infected = series[beta]
plot(beta/gamma, frac_infected, 'ro')
# Here's what it looks like:
# +
plot_sweep_frame(frame)
decorate(xlabel='Contact number (beta/gamma)',
ylabel='Fraction infected',
legend=False)
savefig('figs/chap06-fig03.pdf')
# -
# It turns out that the ratio `beta/gamma`, called the "contact number" is sufficient to predict the total number of infections; we don't have to know `beta` and `gamma` separately.
#
# We can see that in the previous plot: when we plot the fraction infected versus the contact number, the results fall close to a curve.
# ### Analysis
# In the book we figured out the relationship between $c$ and $s_{\infty}$ analytically. Now we can compute it for a range of values:
s_inf_array = linspace(0.0001, 0.9999, 101);
c_array = log(s_inf_array) / (s_inf_array - 1);
# `total_infected` is the change in $s$ from the beginning to the end.
frac_infected = 1 - s_inf_array
frac_infected_series = Series(frac_infected, index=c_array);
# Now we can plot the analytic results and compare them to the simulations.
# +
plot_sweep_frame(frame)
plot(frac_infected_series, label='Analysis')
decorate(xlabel='Contact number (c)',
ylabel='Fraction infected')
savefig('figs/chap06-fig04.pdf')
# -
# The agreement is generally good, except for values of `c` less than 1.
# ## Exercises
# **Exercise:** If we didn't know about contact numbers, we might have explored other possibilities, like the difference between `beta` and `gamma`, rather than their ratio.
#
# Write a version of `plot_sweep_frame`, called `plot_sweep_frame_difference`, that plots the fraction infected versus the difference `beta-gamma`.
#
# What do the results look like, and what does that imply?
beta_array = linspace(0.1, 0.9, 11)
gamma_array = linspace(0.1, 0.7, 4)
frame = sweep_parameters(beta_array, gamma_array)
frame.head()
def plot_sweep_frame_difference(frame):
"""Plot the values from a SweepFrame.
For each (beta, gamma), compute the contact number,
beta/gamma
frame: SweepFrame with one row per beta, one column per gamma
"""
for gamma in frame.columns:
series = frame[gamma]
for beta in series.index:
frac_infected = series[beta]
plot(beta-gamma, frac_infected, 'ro')
# +
plot_sweep_frame_difference(frame)
# plot(frac_infected_series, label='Analysis')
decorate(xlabel='beta - gamma',
ylabel='Fraction infected')
# -
# **Exercise:** Suppose you run a survey at the end of the semester and find that 26% of students had the Freshman Plague at some point.
#
# What is your best estimate of `c`?
#
# Hint: if you print `frac_infected_series`, you can read off the answer.
print(frac_infected_series)
print("This implies that c = 1.158132")
# +
# Alternative solution
"""We can use `np.interp` to look up `s_inf` and
estimate the corresponding value of `c`, but it only
works if the index of the series is sorted in ascending
order. So we have to use `sort_index` first.
"""
frac_infected_series.sort_index(inplace=True)
np.interp(0.26, frac_infected_series, frac_infected_series.index)
# -
|
code/chap14mine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Predição do alfabeto de sinais
# ## Libs utilizadas no desenvolvimento
# +
import csv
import string
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.metrics import confusion_matrix,accuracy_score, classification_report
# -
# ## Leitura e Modelagem dos Dados
def get_data(filename):
with open(filename) as training_file:
training_reader = csv.reader(training_file, delimiter=',')
image = []
labels = []
line_count = 0
for row in training_reader:
if line_count == 0:
line_count +=1
else:
labels.append(row[0])
temp_image = row[1:785]
image_data_as_array = np.array_split(temp_image, 28)
image.append(image_data_as_array)
line_count += 1
images = np.array(image).astype('float32')
labels = np.array(labels).astype('float32')
print(f'Processed {line_count} lines.')
return images, labels
# +
train_imgs, train_labels = get_data("../datasets/sign_mnist_train.csv")
test_imgs, test_labels = get_data("../datasets/sign_mnist_test.csv")
print("Total Training images", train_imgs.shape)
print("Total Training labels",train_labels.shape)
print("Total Testing images",test_imgs.shape)
print("Total Testing labels",test_labels.shape)
# +
# Mapeamento das letras do alfabeto de acordo com a posição dela
alphabets = string.ascii_lowercase
map_letter = {}
for i, letter in enumerate(alphabets):
map_letter[letter] = i
map_letter = {v:k for k, v in map_letter.items()}
train_labels_s = pd.Series(train_labels).map(map_letter)
test_labels_s = pd.Series(test_labels).map(map_letter)
# -
# ## Vizualização Geral dos Dados
# +
fig, axes = plt.subplots(nrows=4, ncols=4, figsize=(12,12), subplot_kw={'xticks': [], 'yticks': []})
for i, ax in enumerate(axes.flat):
img = train_imgs[i].reshape(28,28)
ax.imshow(img, cmap='gray')
title = train_labels_s[i]
ax.set_title(title, fontsize=15)
plt.tight_layout()
plt.show()
# -
plt.figure(figsize=(15,6))
sns.histplot(sorted(train_labels_s), color='purple')
plt.title("Número de fotos por categoria", fontsize=15)
plt.xticks(fontsize=15)
plt.show()
# ## Data Augmentation
# +
train_imgs_s = np.expand_dims(train_imgs, axis=3)
test_imgs_s = np.expand_dims(test_imgs, axis=3)
print(train_imgs_s.shape)
print(test_imgs_s.shape)
# +
train_datagen = ImageDataGenerator(rescale=1.0/255,
height_shift_range=0.1,
width_shift_range=0.1,
zoom_range=0.1,
shear_range=0.1,
rotation_range=10,
fill_mode='nearest',
horizontal_flip=True)
validation_datagen = ImageDataGenerator(rescale=1.0/255)
train_datagenerator = train_datagen.flow(train_imgs_s, train_labels, batch_size=32)
validation_datagenerator = validation_datagen.flow(test_imgs_s, test_labels, batch_size=32)
# -
# ## Definição de um Callback personalizado
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if (logs.get('accuracy')>0.99):
print("\nCancelando treinamento devido a acurácia de 99%!")
self.model.stop_training = True
# ## Construção do Modelo
# ### Arquitetura do Modelo
# +
model = keras.Sequential()
model.add(layers.Conv2D(32, (3,3), activation='relu', input_shape=(28,28,1)))
model.add(layers.MaxPooling2D(2,2))
model.add(layers.Conv2D(128, (3,3), activation='relu'))
model.add(layers.MaxPooling2D(2,2))
model.add(layers.Conv2D(512, (3,3), activation='relu'))
model.add(layers.MaxPooling2D(2,2))
model.add(layers.Flatten())
model.add(layers.Dense(1024, activation='relu'))
model.add(layers.Dropout(0.2))
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dropout(0.2))
model.add(layers.Dense(25, activation='softmax'))
# -
# ### Sumário do Modelo
model.summary()
# ### Treinamento e Parâmetros do Modelo
# +
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
learning_rate_reduction = keras.callbacks.ReduceLROnPlateau(monitor='val_accuracy',
patience = 2,
verbose=1,factor=0.25,
min_lr=0.0001)
callbacks = myCallback()
history = model.fit(train_datagenerator,
validation_data = validation_datagenerator,
steps_per_epoch = len(train_labels)//32,
epochs = 100,
validation_steps = len(test_labels)//32,
callbacks = [callbacks, learning_rate_reduction])
model.save('../models/model.h5')
# -
# ### Gráficos do Treinamento
# +
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(15,6))
axes[0].plot(epochs, acc, 'r', label='Training accuracy')
axes[0].plot(epochs, val_acc, 'b', label='Validation accuracy')
axes[0].set_title('Training and validation accuracy')
axes[0].legend()
axes[1].plot(epochs, loss, 'r', label='Training Loss')
axes[1].plot(epochs, val_loss, 'b', label='Validation Loss')
axes[1].set_title('Training and validation loss')
axes[1].legend()
plt.show()
# -
# ### Carregando o Modelo
model_load = keras.models.load_model('../models/model.h5')
model_load.evaluate(test_imgs_s, test_labels, return_dict=True)
# ## Avaliando as Métricas do Modelo
# ### <NAME>
# +
pred = model.predict(test_imgs_s)
pred = np.argmax(pred,axis=1)
# Get the accuracy score
acc = accuracy_score(test_labels,pred)
# Display the results
print(f'## {acc*100:.2f}% accuracy on the test set')
# -
# ### Classification Report
# +
y_test_letters = [map_letter[x] for x in test_labels]
pred_letters = [map_letter[x] for x in pred]
print(classification_report(y_test_letters, pred_letters))
# -
# ### Matrix de Confusão
cf_matrix = confusion_matrix(y_test_letters, pred_letters, normalize='true')
plt.figure(figsize = (20,15))
sns.heatmap(cf_matrix, annot=True, fmt='.2f', xticklabels = sorted(set(y_test_letters)), yticklabels = sorted(set(y_test_letters)),cbar=False)
plt.title('Normalized Confusion Matrix\n', fontsize = 23)
plt.xlabel("Pred Class",fontsize=15)
plt.ylabel("True Label",fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15,rotation=0)
plt.show()
# ### Casos de Acerto
correct = np.nonzero(pred == test_labels)[0]
plt.figure(figsize=(12, 6))
i = 0
for c in correct[:8]:
plt.subplot(2,4,i+1)
plt.imshow(test_imgs[c].reshape(28,28), cmap="gray", interpolation='none')
plt.title("Pred:{}, True:{}".format(pred_letters[c], y_test_letters[c]), size=15)
plt.tight_layout()
plt.axis('off')
i += 1
wrong = np.nonzero(pred != test_labels)[0]
plt.figure(figsize=(12, 6))
i = 0
for c in wrong[:8]:
plt.subplot(2,4,i+1)
plt.imshow(test_imgs[c].reshape(28,28), cmap="gray", interpolation='none')
plt.title("Pred:{}, True:{}".format(pred_letters[c], y_test_letters[c]), size=15)
plt.tight_layout()
plt.axis('off')
i += 1
|
notebook/model_construct.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
You can run either this notebook locally (if you have all the dependencies and a GPU) or on Google Colab.
Instructions for setting up Colab are as follows:
1. Open a new Python 3 notebook.
2. Import this notebook from GitHub (File -> Upload Notebook -> "GITHUB" tab -> copy/paste GitHub URL)
3. Connect to an instance with a GPU (Runtime -> Change runtime type -> select "GPU" for hardware accelerator)
4. Run this cell to set up dependencies.
"""
# If you're using Google Colab and not running locally, run this cell.
# !pip install wget
# !apt-get install sox libsndfile1 ffmpeg
# !pip install nemo_toolkit[asr]==0.10.0b10
# !pip install unidecode
# !mkdir configs
# !wget -P configs/ https://raw.githubusercontent.com/NVIDIA/NeMo/master/examples/asr/notebooks/configs/jasper_an4.yaml
# -
# # Introduction to End-To-End Automatic Speech Recognition
#
# This notebook contains a basic tutorial of Automatic Speech Recognition (ASR) concepts, introduced with code snippets using the [NeMo framework](https://github.com/NVIDIA/NeMo).
# We will first introduce the basics of the main concepts behind speech recognition, then explore concrete examples of what the data looks like and walk through putting together a simple end-to-end ASR pipeline.
#
# We assume that you are familiar with general machine learning concepts and can follow Python code, and we'll be using the [AN4 dataset from CMU](http://www.speech.cs.cmu.edu/databases/an4/) (with processing using `sox`).
# ## Conceptual Overview: What is ASR?
#
# ASR, or **Automatic Speech Recognition**, refers to the problem of getting a program to automatically transcribe spoken language (speech-to-text). Our goal is usually to have a model that minimizes the **Word Error Rate (WER)** metric when transcribing speech input. In other words, given some audio file (e.g. a WAV file) containing speech, how do we transform this into the corresponding text with as few errors as possible?
#
# Traditional speech recognition takes a generative approach, modeling the full pipeline of how speech sounds are produced: from a **language model** that encapsulates likely orderings of words (e.g. an n-gram model), to a **pronunciation model** for each word in the vocabulary (e.g. a pronunciation table), to an **acoustic model** that translates the pronunciations to audio waveforms (e.g. a Gaussian Mixture Model), and so on.
#
# Then, if we receive some spoken input, our goal would be to find the most likely sequence of text that would result in the given audio according to our pipeline of models. Overall, with traditional speech recognition, we try to model `Pr(audio|transcript)*Pr(transcript)`, and take the argmax of this over possible transcripts.
#
# Over time, neural nets advanced to the point where each component of the traditional speech recognition model could be replaced by a neural model that had better performance and that had a greater potential for generalization. For example, we could replace an n-gram model with a neural language model, and replace a pronunciation table with a neural pronunciation model, and so on. However, each of these neural models need to be trained individually on different tasks, and errors in any model in the pipeline could throw off the whole prediction.
#
# Thus, we can see the appeal of **end-to-end ASR architectures**--discriminative models that simply take an audio input and give a textual output, and in which all components are trained together towards the same goal. A much easier pipeline to handle!
# ### End-To-End ASR
#
# With an end-to-end model, we want to directly learn `Pr(transcript|audio)` in order to predict the transcripts from the original audio. Since we are dealing with sequential information--audio data over time that corresponds to a sequence of letters--RNNs are the obvious choice. But now we have a pressing problem to deal with: since our input sequence (number of audio timesteps) is not the same length as our desired output (transcript length), how do we match each time step from the audio data to the correct output characters?
#
# Earlier speech recognition approaches relied on **temporally-aligned data**, in which each segment of time in an audio file was matched up to a corresponding speech sound such as a phoneme or word. However, if we would like to have the flexibility to predict letter-by-letter to prevent OOV (out of vocabulary) issues, then each time step in the data would have to be labeled with the letter sound that the speaker is making at that point in the audio file. With that information, it seems like we should simply be able to try to predict the correct letter for each time step and then collapse the repeated letters (e.g. the prediction output `LLLAAAAPPTOOOPPPP` would become `LAPTOP`). It turns out that this idea has some problems: not only does alignment make the dataset incredibly labor-intensive to label, but also, what do we do with words like "book" that contain consecutive repeated letters? Simply squashing repeated letters together would not work in that case!
#
# 
#
# Modern end-to-end approaches get around this using methods that don't require manual alignment at all, so that the input-output pairs are really just the raw audio and the transcript--no extra data or labeling required. Let's briefly go over two popular approaches that allow us to do this, Connectionist Temporal Classification (CTC) and sequence-to-sequence models with attention.
#
# #### Connectionist Temporal Classification (CTC)
#
# In normal speech recognition prediction output, we would expect to have characters such as the letters from A through Z, numbers 0 through 9, spaces ("\_"), and so on. CTC introduces a new intermediate output token called the **blank token** ("-") that is useful for getting around the alignment issue.
#
# With CTC, we still predict one token per time segment of speech, but we use the blank token to figure out where we can and can't collapse the predictions. The appearance of a blank token helps separate repeating letters that should not be collapsed. For instance, with an audio snippet segmented into `T=11` time steps, we could get predictions that look like `BOO-OOO--KK`, which would then collapse to `"BO-O-K"`, and then we would remove the blank tokens to get our final output, `BOOK`.
#
# Now, we can predict one output token per time step, then collapse and clean to get sensible output without any fear of ambiguity from repeating letters! A simple way of getting predictions like this would be to apply a bidirectional RNN to the audio input, apply softmax over each time step's output, and then take the token with the highest probability. The method of always taking the best token at each time step is called **greedy decoding, or max decoding**.
#
# To calculate our loss for backprop, we would like to know the log probability of the model producing the correct transcript, `log(Pr(transcript|audio))`. We can get the log probability of a single intermediate output sequence (e.g. `BOO-OOO--KK`) by summing over the log probabilities we get from each token's softmax value, but note that the resulting sum is different from the log probability of the transcript itself (`BOOK`). This is because there are multiple possible output sequences of the same length that can be collapsed to get the same transcript (e.g. `BBO--OO-KKK` also results in `BOOK`), and so we need to **marginalize over every valid sequence of length `T` that collapses to the transcript**.
#
# Therefore, to get our transcript's log probability given our audio input, we must sum the log probabilities of every sequence of length `T` that collapses to the transcript (e.g. `log(Pr(output: "BOOK"|audio)) = log(Pr(BOO-OOO--KK|audio)) + log(Pr(BBO--OO-KKK|audio)) + ...`). In practice, we can use a dynamic programming approach to calculate this, accumulating our log probabilities over different "paths" through the softmax outputs at each time step.
#
# If you would like a more in-depth explanation of how CTC works, or how we can improve our results by using a modified beam search algorithm, feel free to check out the Further Reading section at the end of this notebook for more resources.
#
# #### Sequence-to-Sequence with Attention
#
# One problem with CTC is that predictions at different time steps are conditionally independent, which is an issue because the words in a continuous utterance tend to be related to each other in some sensible way. With this conditional independence assumption, we can't learn a language model that can represent such dependencies, though we can add a language model on top of the CTC output to mitigate this to some degree.
#
# A popular alternative is to use a sequence-to-sequence model with attention. A typical seq2seq model for ASR consists of some sort of **bidirectional RNN encoder** that consumes the audio sequence timestep-by-timestep, and where the outputs are then passed to an **attention-based decoder**. Each prediction from the decoder is based on attending to some parts of the entire encoded input, as well as the previously outputted tokens.
#
# The outputs of the decoder can be anything from word pieces to phonemes to letters, and since predictions are not directly tied to time steps of the input, we can just continue producing tokens one-by-one until an end token is given (or we reach a specified max output length). This way, we do not need to deal with audio alignment, and our predicted transcript is just the sequence of outputs given by our decoder.
#
# Now that we have an idea of what some popular end-to-end ASR models look like, let's take a look at the audio data we'll be working with for our example.
# ## Taking a Look at Our Data (AN4)
#
# The AN4 dataset, also known as the Alphanumeric dataset, was collected and published by Carnegie Mellon University. It consists of recordings of people spelling out addresses, names, telephone numbers, etc., one letter or number at a time, as well as their corresponding transcripts. We choose to use AN4 for this tutorial because it is relatively small, with 948 training and 130 test utterances, and so it trains quickly.
#
# Before we get started, let's download and prepare the dataset. The utterances are available as `.sph` files, so we will need to convert them to `.wav` for later processing. Please make sure you have [Sox](http://sox.sourceforge.net/) installed for this step (see the "Downloads" section of the main page).
# This is where the an4/ directory will be placed.
# Change this if you don't want the data to be extracted in the current directory.
data_dir = '.'
# +
import glob
import os
import subprocess
import tarfile
import wget
# Download the dataset. This will take a few moments...
print("******")
if not os.path.exists(data_dir + '/an4_sphere.tar.gz'):
an4_url = 'http://www.speech.cs.cmu.edu/databases/an4/an4_sphere.tar.gz'
an4_path = wget.download(an4_url, data_dir)
print(f"Dataset downloaded at: {an4_path}")
else:
print("Tarfile already exists.")
an4_path = data_dir + '/an4_sphere.tar.gz'
if not os.path.exists(data_dir + '/an4/'):
# Untar and convert .sph to .wav (using sox)
tar = tarfile.open(an4_path)
tar.extractall(path=data_dir)
print("Converting .sph to .wav...")
sph_list = glob.glob(data_dir + '/an4/**/*.sph', recursive=True)
for sph_path in sph_list:
wav_path = sph_path[:-4] + '.wav'
cmd = ["sox", sph_path, wav_path]
subprocess.run(cmd)
print("Finished conversion.\n******")
# -
# You should now have a folder called `an4` that contains `etc/an4_train.transcription`, `etc/an4_test.transcription`, audio files in `wav/an4_clstk` and `wav/an4test_clstk`, along with some other files we will not need.
#
# Now we can load and take a look at the data. As an example, file `cen2-mgah-b.wav` is a 2.6 second-long audio recording of a man saying the letters "G L E N N" one-by-one (feel free to check this out by listening to `./an4/wav/an4_clstk/mgah/cen2-mgah-b.wav`). In an ASR task, the WAV file would be our input, and "G L E N N" would be our desired output.
#
# Let's plot the waveform, which is simply a line plot of the sequence of values that we read from the file. This is a format of viewing audio that you are likely to be familiar with seeing in many audio editors and visualizers:
# +
# %matplotlib inline
import librosa
import librosa.display
import matplotlib.pyplot as plt
import numpy as np
# Plot our example audio file's waveform
example_file = data_dir + '/an4/wav/an4_clstk/mgah/cen2-mgah-b.wav'
audio, sample_rate = librosa.load(example_file)
plt.rcParams['figure.figsize'] = (15,7)
plt.title('Waveform of Audio Example')
plt.ylabel('Amplitude')
_ = librosa.display.waveplot(audio)
# -
# We can see the activity in the waveform that corresponds to each letter in the audio, as our speaker here enunciates quite clearly!
# You can kind of tell that each spoken letter has a different "shape," and it's interesting to note that last two blobs look relatively similar, which is expected because they are both the letter "N."
#
# ### Spectrograms and Mel Spectrograms
#
# However, since audio information is more useful in the context of frequencies of sound over time, we can get a better representation than this raw sequence of 57,330 values.
# We can apply a [Fourier Transform](https://en.wikipedia.org/wiki/Fourier_transform) on our audio signal to get something more useful: a **spectrogram**, which is a representation of the energy levels (i.e. amplitude, or "loudness") of each frequency (i.e. pitch) of the signal over the duration of the file.
# A spectrogram (which can be viewed as a heat map) is a good way of seeing how the *strengths of various frequencies in the audio vary over time*, and is obtained by breaking up the signal into smaller, usually overlapping chunks and performing a Short-Time Fourier Transform (STFT) on each.
#
# Let's examine what the spectrogram of our sample looks like.
# +
# Get spectrogram using Librosa's Short-Time Fourier Transform (stft)
spec = np.abs(librosa.stft(audio))
spec_db = librosa.amplitude_to_db(spec, ref=np.max) # Decibels
# Use log scale to view frequencies
librosa.display.specshow(spec_db, y_axis='log', x_axis='time')
plt.colorbar()
plt.title('Audio Spectrogram');
# -
# Again, we are able to see each letter being pronounced, and that the last two blobs that correspond to the "N"s are pretty similar-looking. But how do we interpret these shapes and colors? Just as in the waveform plot before, we see time passing on the x-axis (all 2.6s of audio). But now, the y-axis represents different frequencies (on a log scale), and *the color on the plot shows the strength of a frequency at a particular point in time*.
#
# We're still not done yet, as we can make one more potentially useful tweak: using the **Mel Spectrogram** instead of the normal spectrogram. This is simply a change in the frequency scale that we use from linear (or logarithmic) to the mel scale, which is "a perceptual scale of pitches judged by listeners to be equal in distance from one another" (from [Wikipedia](https://en.wikipedia.org/wiki/Mel_scale)).
#
# In other words, it's a transformation of the frequencies to be more aligned to what humans perceive; a change of +1000Hz from 2000Hz->3000Hz sounds like a larger difference to us than 9000Hz->10000Hz does, so the mel scale normalizes this such that equal distances sound like equal differences to the human ear. Intuitively, we use the mel spectrogram because in this case we are processing and transcribing human speech, such that transforming the scale to better match what we hear is a useful procedure.
# +
# Plot the mel spectrogram of our sample
mel_spec = librosa.feature.melspectrogram(audio, sr=sample_rate)
mel_spec_db = librosa.power_to_db(mel_spec, ref=np.max)
librosa.display.specshow(
mel_spec_db, x_axis='time', y_axis='mel')
plt.colorbar()
plt.title('Mel Spectrogram');
# -
# ## Building a Simple ASR Pipeline in NeMo
#
# Now that we have an idea of what the audio data looks like, we can start building our end-to-end ASR pipeline!
#
# We'll be using the **Neural Modules (NeMo) toolkit** for this part, so if you haven't already, you should download and install NeMo and its dependencies. To do so, just follow the directions on the [GitHub page](https://github.com/NVIDIA/NeMo), or in the [documentation](https://nvidia.github.io/NeMo/).
#
# NeMo lets us easily hook together the components (modules) of our model, such as the data layer, intermediate layers, and various losses, without worrying too much about implementation details of individual parts or connections between modules. If you're curious, you can read more about NeMo and how it works in the documentation pages linked above.
# NeMo's "core" package
import nemo
# NeMo's ASR collection
import nemo.collections.asr as nemo_asr
# ### Creating Data Manifests
#
# The first thing we need to do now is to create manifests for our training and evaluation data, which will contain the metadata of our audio files. NeMo data layers take in a standardized manifest format where each line corresponds to one sample of audio, such that the number of lines in a manifest is equal to the number of samples that are represented by that manifest. A line must contain the path to an audio file, the corresponding transcript (or path to a transcript file), and the duration of the audio sample.
#
# Here's an example of what one line in a NeMo-compatible manifest might look like:
# ```
# {"audio_filepath": "path/to/audio.wav", "duration": 3.45, "text": "this is a nemo tutorial"}
# ```
#
# We can build our training and evaluation manifests using `an4/etc/an4_train.transcription` and `an4/etc/an4_test.transcription`, which have lines containing transcripts and their corresponding audio file IDs:
# ```
# ...
# <s> P I T T S B U R G H </s> (cen5-fash-b)
# <s> TWO SIX EIGHT FOUR FOUR ONE EIGHT </s> (cen7-fash-b)
# ...
# ```
# +
# --- Building Manifest Files --- #
import json
# Function to build a manifest
def build_manifest(transcripts_path, manifest_path, wav_path):
with open(transcripts_path, 'r') as fin:
with open(manifest_path, 'w') as fout:
for line in fin:
# Lines look like this:
# <s> transcript </s> (fileID)
transcript = line[: line.find('(')-1].lower()
transcript = transcript.replace('<s>', '').replace('</s>', '')
transcript = transcript.strip()
file_id = line[line.find('(')+1 : -2] # e.g. "cen4-fash-b"
audio_path = os.path.join(
data_dir, wav_path,
file_id[file_id.find('-')+1 : file_id.rfind('-')],
file_id + '.wav')
duration = librosa.core.get_duration(filename=audio_path)
# Write the metadata to the manifest
metadata = {
"audio_filepath": audio_path,
"duration": duration,
"text": transcript
}
json.dump(metadata, fout)
fout.write('\n')
# Building Manifests
print("******")
train_transcripts = data_dir + '/an4/etc/an4_train.transcription'
train_manifest = data_dir + '/an4/train_manifest.json'
if not os.path.isfile(train_manifest):
build_manifest(train_transcripts, train_manifest, 'an4/wav/an4_clstk')
print("Training manifest created.")
test_transcripts = data_dir + '/an4/etc/an4_test.transcription'
test_manifest = data_dir + '/an4/test_manifest.json'
if not os.path.isfile(test_manifest):
build_manifest(test_transcripts, test_manifest, 'an4/wav/an4test_clstk')
print("Test manifest created.")
print("******")
# -
# ### Building Training and Evaluation DAGs
#
# Let's take a look at the model that we will be building, and how we specify its parameters.
#
# #### The Jasper Model
#
# We will be putting together a small [Jasper (Just Another SPeech Recognizer) model](https://arxiv.org/abs/1904.03288).
# In brief, Jasper architectures consist of a repeated block structure that utilizes 1D convolutions.
# In a Jasper_KxR model, `R` sub-blocks (consisting of a 1D convolution, batch norm, ReLU, and dropout) are grouped into a single block, which is then repeated `K` times.
# We also have a one extra block at the beginning and a few more at the end that are invariant of `K` and `R`, and we use CTC loss.
#
# A Jasper model looks like roughly this:
#
# 
#
# #### Specifying Our Model with a YAML Config File
#
# For this tutorial, we'll build a *Jasper_4x1 model*, with `K=4` blocks of single (`R=1`) sub-blocks and a *greedy CTC decoder*, using the configuration found in `./configs/jasper_an4.yaml`.
#
# If we open up this config file, we find that there is an entry labeled `JasperEncoder`, with a field called `jasper` that contains a list with multiple entries. Each of the members in this list specifies one block in our model, and looks something like this:
# ```
# - filters: 256
# repeat: 1
# kernel: [11]
# stride: [2]
# dilation: [1]
# dropout: 0.2
# residual: false
# ```
# The first member of the list corresponds to the first block in the Jasper architecture diagram, which appears regardless of `K` and `R`.
# Next, we have four entries that correspond to the `K=4` blocks, and each has `repeat: 1` since we are using `R=1`.
# These are followed by two more entries for the blocks that appear at the end of our Jasper model before the CTC loss.
#
# There are also some entries at the top of the file that specify that we should be shuffling our training data but not our evaluation data (see `AudioToTextDataLayer_train` and `AudioToTextDataLayer_eval`), and some specifications for preprocessing and converting the audio data (in `AudioToMelSpectrogramPreprocessor`).
#
# Using a YAML config such as this is helpful for getting a quick and human-readable overview of what your architecture looks like, and allows you to swap out model and run configurations easily without needing to change your code.
#
# #### Building Training and Evaluation DAGs with NeMo
#
# Building a model using NeMo consists of (1) instantiating the neural modules we need and (2) specifying the DAG by linking them together.
# In NeMo, **the training and inference pipelines are managed by a `NeuralModuleFactory`**, which takes care of checkpointing, callbacks, and logs, along with other details in training and inference. We set its `log_dir` argument to specify where our model logs and outputs will be written, and can set other training and inference settings in its constructor. For instance, if we were **resuming training from a checkpoint**, we would set the argument `checkpoint_dir=<path_to_checkpoint>`.
#
# Along with logs in NeMo, you can optionally view the tensorboard logs with the `create_tb_writer=True` argument to the `NeuralModuleFactory`. By default all the tensorboard log files will be stored in `{log_dir}/tensorboard`, but you can change this with the `tensorboard_dir` argument. One can load tensorboard logs through tensorboard by running `tensorboard --logdir=<path_to_tensorboard dir>` in the terminal.
# +
# Create our NeuralModuleFactory, which will oversee the neural modules.
neural_factory = nemo.core.NeuralModuleFactory(
log_dir=data_dir+'/an4_tutorial/',
create_tb_writer=True)
logger = nemo.logging
# -
# Note that if you would like your pipeline to support running on CPU, you should
# instead add these imports and then add an additional 'placement' argument:
#
# ```python
# import torch
# from nemo.core import DeviceType
#
# neural_factory = nemo.core.NeuralModuleFactory(
# log_dir=data_dir+'/an4_tutorial/',
# placement=(DeviceType.GPU if torch.cuda.is_available() else DeviceType.CPU)
# )
# ```
#
# Now that we have our neural module factory, we can **specify our neural modules and instantiate them**. Here, we load the parameters for each module from the configuration file using `import_from_config`. For the module parameters that we can't read directly from the config file or want to overwrite, we can use the `overwrite_params` argument.
# +
# --- Config Information ---#
from ruamel.yaml import YAML
config_path = './configs/jasper_an4.yaml'
yaml = YAML(typ='safe')
with open(config_path) as f:
params = yaml.load(f)
labels = params['labels'] # Vocab
# --- Instantiate Neural Modules --- #
# Create training and test data layers (which load data) and data preprocessor
data_layer_train = nemo_asr.AudioToTextDataLayer.import_from_config(
config_path,
"AudioToTextDataLayer_train",
overwrite_params={"manifest_filepath": train_manifest}
) # Training datalayer
data_layer_test = nemo_asr.AudioToTextDataLayer.import_from_config(
config_path,
"AudioToTextDataLayer_eval",
overwrite_params={"manifest_filepath": test_manifest}
) # Eval datalayer
data_preprocessor = nemo_asr.AudioToMelSpectrogramPreprocessor.import_from_config(
config_path, "AudioToMelSpectrogramPreprocessor"
)
# Create the Jasper_4x1 encoder as specified, and a CTC decoder
encoder = nemo_asr.JasperEncoder.import_from_config(
config_path, "JasperEncoder"
)
decoder = nemo_asr.JasperDecoderForCTC.import_from_config(
config_path, "JasperDecoderForCTC",
overwrite_params={"num_classes": len(labels)}
)
ctc_loss = nemo_asr.CTCLossNM(num_classes=len(labels))
greedy_decoder = nemo_asr.GreedyCTCDecoder()
# -
# The next step is to assemble our training DAG by specifying the inputs to each neural module.
# +
# --- Assemble Training DAG --- #
audio_signal, audio_signal_len, transcript, transcript_len = data_layer_train()
processed_signal, processed_signal_len = data_preprocessor(
input_signal=audio_signal,
length=audio_signal_len)
encoded, encoded_len = encoder(
audio_signal=processed_signal,
length=processed_signal_len)
log_probs = decoder(encoder_output=encoded)
preds = greedy_decoder(log_probs=log_probs) # Training predictions
loss = ctc_loss(
log_probs=log_probs,
targets=transcript,
input_length=encoded_len,
target_length=transcript_len)
# -
# We would like to be able to evaluate our model on the test set, as well, so let's set up the evaluation DAG.
#
# Our **evaluation DAG will reuse most of the parts of the training DAG with the exception of the data layer**, since we are loading the evaluation data from a different file but evaluating on the same model. Note that if we were using data augmentation in training, we would also leave that out in the evaluation DAG.
# +
# --- Assemble Validation DAG --- #
(audio_signal_test, audio_len_test,
transcript_test, transcript_len_test) = data_layer_test()
processed_signal_test, processed_len_test = data_preprocessor(
input_signal=audio_signal_test,
length=audio_len_test)
encoded_test, encoded_len_test = encoder(
audio_signal=processed_signal_test,
length=processed_len_test)
log_probs_test = decoder(encoder_output=encoded_test)
preds_test = greedy_decoder(log_probs=log_probs_test) # Test predictions
loss_test = ctc_loss(
log_probs=log_probs_test,
targets=transcript_test,
input_length=encoded_len_test,
target_length=transcript_len_test)
# -
# ### Running the Model
#
# We would like to be able to monitor our model while it's training, so we use **callbacks**. In general, *callbacks are functions that are called at specific intervals over the course of training or inference*, such as at the start or end of every *n* iterations, epochs, etc. The callbacks we'll be using for this are the `SimpleLossLoggerCallback`, which reports the training loss (or another metric of your choosing, such as WER for ASR tasks), and the `EvaluatorCallback`, which regularly evaluates the model on the test set. Both of these callbacks require you to pass in the tensors to be evaluated--these would be the final outputs of the training and eval DAGs above.
#
# Another useful callback is the `CheckpointCallback`, for saving checkpoints at set intervals. We create one here just to demonstrate how it works.
# +
# --- Create Callbacks --- #
# We use these imports to pass to callbacks more complex functions to perform.
from nemo.collections.asr.helpers import monitor_asr_train_progress, \
process_evaluation_batch, process_evaluation_epoch
from functools import partial
train_callback = nemo.core.SimpleLossLoggerCallback(
# Notice that we pass in loss, predictions, and the transcript info.
# Of course we would like to see our training loss, but we need the
# other arguments to calculate the WER.
tensors=[loss, preds, transcript, transcript_len],
# The print_func defines what gets printed.
print_func=partial(
monitor_asr_train_progress,
labels=labels),
tb_writer=neural_factory.tb_writer
)
# We can create as many evaluation DAGs and callbacks as we want,
# which is useful in the case of having more than one evaluation dataset.
# In this case, we only have one.
eval_callback = nemo.core.EvaluatorCallback(
eval_tensors=[loss_test, preds_test, transcript_test, transcript_len_test],
user_iter_callback=partial(
process_evaluation_batch, labels=labels),
user_epochs_done_callback=process_evaluation_epoch,
eval_step=500, # How often we evaluate the model on the test set
tb_writer=neural_factory.tb_writer
)
checkpoint_saver_callback = nemo.core.CheckpointCallback(
folder=data_dir+'/an4_checkpoints',
step_freq=1000 # How often checkpoints are saved
)
if not os.path.exists(data_dir+'/an4_checkpoints'):
os.makedirs(data_dir+'/an4_checkpoints')
# -
# Now that we have our model and callbacks set up, **how do we run it**?
#
# Once we create our neural factory and the callbacks for the information that we want to see, we can **start training** by simply calling the train function on the tensors we want to optimize and our callbacks!
# +
# --- Start Training! --- #
neural_factory.train(
tensors_to_optimize=[loss],
callbacks=[train_callback, eval_callback, checkpoint_saver_callback],
optimizer='novograd',
optimization_params={
"num_epochs": 100, "lr": 0.01, "weight_decay": 1e-4
})
# Training for 100 epochs will take a little while, depending on your machine.
# It should take about 20 minutes on Google Colab.
# At the end of 100 epochs, your evaluation WER should be around 20-25%.
# -
# There we go! We've put together a full training pipeline for the model and trained it for 100 epochs.
#
# ### Inference
#
# What if we have a trained model that we **just want to run inference** on?
#
# In that case, we just need to instantiate and link up the modules needed for the evaluation DAG (same procedure as before), and then run `infer` to get the results. Let's see what performing inference with our last checkpoint would look like.
# +
# --- Inference Only --- #
# We've already built the inference DAG above, so all we need is to call infer().
evaluated_tensors = neural_factory.infer(
# These are the tensors we want to get from the model.
tensors=[loss_test, preds_test, transcript_test, transcript_len_test],
# checkpoint_dir specifies where the model params are loaded from.
checkpoint_dir=(data_dir+'/an4_checkpoints')
)
# Process the results to get WER
from nemo.collections.asr.helpers import word_error_rate, \
post_process_predictions, post_process_transcripts
greedy_hypotheses = post_process_predictions(
evaluated_tensors[1], labels)
references = post_process_transcripts(
evaluated_tensors[2], evaluated_tensors[3], labels)
wer = word_error_rate(hypotheses=greedy_hypotheses, references=references)
print("*** Greedy WER: {:.2f} ***".format(wer * 100))
# -
# And that's it!
# ## Model Improvements
#
# You already have all you need to create your own ASR model in NeMo, but there are a few more tricks that you can employ if you so desire. In this section, we'll briefly cover a few possibilities for improving an ASR model.
#
# ### Data Augmentation
#
# There exist several ASR data augmentation methods that can increase the size of our training set.
#
# For example, we can perform augmentation on the spectrograms by zeroing out specific frequency segments ("frequency masking") or time segments ("time masking") as described by [SpecAugment](https://arxiv.org/abs/1904.08779), or zero out rectangles on the spectrogram as in [Cutout](https://arxiv.org/pdf/1708.04552.pdf). In NeMo, we can do all three of these by simply adding in a `SpectrogramAugmentation` neural module. (As of now, it does not perform the time warping from the SpecAugment paper.)
# +
# Create a SpectrogramAugmentation module
spectrogram_aug = nemo_asr.SpectrogramAugmentation(
rect_masks=5, rect_time=120, rect_freq=50)
# Rearrange training DAG to use augmentation.
# The following code is mostly copy/pasted from the "Assemble Training DAG"
# section, with only one line added!
audio_signal, audio_signal_len, transcript, transcript_len = data_layer_train()
processed_signal, processed_signal_len = data_preprocessor(
input_signal=audio_signal,
length=audio_signal_len)
############## This is the only part that's changed! ##############
processed_signal_aug = spectrogram_aug(input_spec=processed_signal)
encoded, encoded_len = encoder(
audio_signal=processed_signal_aug, # Change this argument too
length=processed_signal_len)
###################################################################
log_probs = decoder(encoder_output=encoded)
preds = greedy_decoder(log_probs=log_probs) # Training predictions
loss = ctc_loss(
log_probs=log_probs,
targets=transcript,
input_length=encoded_len,
target_length=transcript_len)
# And then you can train as usual.
# If you want to try it out in this notebook,
# be sure to run neural_factory.reset_trainer() before training again!
# -
# Another popular method of ASR data augmentation is speed perturbation, where the audio is sped up or slowed down slightly (e.g. 10% faster or slower). See the `SpeedPerturbation` class in the ASR collection for more details.
#
# ### Using a Language Model
#
# Though a language model (LM) may not be especially suited to a task like AN4 where we have a bunch of letters being read in sequence, adding a language model for inference can the improve WER in most other ASR tasks, where the speech more closely matches normal patterns. We can use the probability distribution that a language model gives us to better match our predictions to sequences of words we would be more likely to see in the real world, such as correcting "keyboard and house" to "keyboard and mouse."
#
# If you have a language model that you'd like to use with a NeMo model, you can add a `BeamSearchDecoderWithLM` module to your DAG to get beam search predictions that use your language model file.
#
# For the sake of example, even though an LM won't help much for this dataset, we'll go through how to set this up.
# First, if you're on your own machine, you'll want to run the script `NeMo/scripts/install_decoders.sh` (or `NeMo/scripts/install_decoders_MacOS.sh`, if appropriate).
#
# **Only run the following code block if you are using Google Colab.**
# +
# If you are using Google Colab, run this cell.
# The following is mostly copied from `NeMo/scripts/install_decoders.sh`.
# This will take a little while.
# !apt-get install swig
# !git clone https://github.com/PaddlePaddle/DeepSpeech
# !cd DeepSpeech; git checkout b3c728d
# !mv DeepSpeech/decoders/swig_wrapper.py DeepSpeech/decoders/swig/ctc_decoders.py
# !mv DeepSpeech/decoders/swig ./decoders
# !cd decoders; sed -i "s/\.decode('utf-8')//g" ctc_decoders.py; \
# sed -i 's/\.decode("utf-8")//g' ctc_decoders.py; \
# sed -i "s/name='swig_decoders'/name='ctc_decoders'/g" setup.py; \
# sed -i "s/-space_prefixes\[i\]->approx_ctc/space_prefixes\[i\]->score/g" decoder_utils.cpp; \
# sed -i "s/py_modules=\['swig_decoders'\]/py_modules=\['ctc_decoders', 'swig_decoders'\]/g" setup.py; \
# chmod +x setup.sh; \
# ./setup.sh
# The following is a bit of a hack to get the import to work.
# If the path is wrong, check the last few lines of installer output for the correct path.
# (There should be a line like: "Installed <path>".)
os.sys.path.append('/usr/local/lib/python3.6/dist-packages/ctc_decoders-1.1-py3.6-linux-x86_64.egg')
# -
# Next, we'll download and unzip a 3-gram language model built from the LibriSpeech corpus, courtesy of OpenSLR. (See more [here](http://www.openslr.org/11).) We also convert it to lowercase, because the decoder we use expects all lowercase.
#
# The download will take several minutes, so don't panic!
# +
import gzip
import shutil
lm_gzip_path = os.path.join(data_dir, '3-gram.pruned.1e-7.arpa.gz')
if not os.path.exists(lm_gzip_path):
print("Downloading pruned 3-gram model.")
lm_url = 'http://www.openslr.org/resources/11/3-gram.pruned.1e-7.arpa.gz'
lm_gzip_path = wget.download(lm_url, data_dir)
print("Downloaded the 3-gram language model.")
else:
print("Pruned .arpa.gz already exists.")
uppercase_lm_path = os.path.join(data_dir, '3-gram.pruned.1e-7.arpa')
if not os.path.exists(uppercase_lm_path):
with gzip.open(lm_gzip_path, 'rb') as f_zipped:
with open(uppercase_lm_path, 'wb') as f_unzipped:
shutil.copyfileobj(f_zipped, f_unzipped)
print("Unzipped the 3-gram language model.")
else:
print("Unzipped .arpa already exists.")
lm_path = os.path.join(data_dir, 'lowercase_3-gram.pruned.1e-7.arpa')
if not os.path.exists(lm_path):
with open(uppercase_lm_path, 'r') as f_upper:
with open(lm_path, 'w') as f_lower:
for line in f_upper:
f_lower.write(line.lower())
print("Converted language model file to lowercase.")
# -
# Now we're set up to create the `BeamSearchDecoderWithLM` module.
#
# You can tune the hyperparameters of `beam_width`, `alpha`, and `beta` to your liking, or even perform a search over hyperparameters. For now, we'll just use a measly `beam_width` of 16, and some arbitrary `alpha` and `beta`.
### Instantiating the module ###
beam_search_lm = nemo_asr.BeamSearchDecoderWithLM(
vocab=labels,
beam_width=16,
alpha=2, beta=1.5,
lm_path=lm_path,
num_cpus=max(os.cpu_count(), 1),
input_tensor=False # We will be inputting numpy values rather than PT tensors.
)
# Once we've done that, we want to perform inference again. Notice the slightly different set of tensors we evaluate--this time we'll need the log probabilities and not the loss, as well as the encoded length.
#
# We then want to get the log probabilities of the output of the encoder, then pass them through the `forward()` function of the beam search decoder module to get our predictions.
# +
# Infer again to get the info we need!
evaluated_tensors = neural_factory.infer(
tensors=[log_probs_test, preds_test, transcript_test, transcript_len_test, encoded_len_test],
checkpoint_dir=(data_dir+'/an4_checkpoints')
)
eval_log_probs, eval_preds, eval_transcript, eval_transcript_len, eval_encoded_len = evaluated_tensors
# Convert our log probs from inference to a list of numpy arrays for beam search with LM
np_log_probs = []
for i, batch in enumerate(eval_log_probs): # Iterate through batches
for j in range(batch.shape[0]): # Iterate through each batch entry
# Get the log-probs for each entry, but mask off data longer than the entry
np_log_probs.append(batch[j][: eval_encoded_len[i][j], :].cpu().numpy())
# Exponentiate -- the BeamSearchDecoderWithLM class assumes we've done it already if we pass in numpy values.
np_log_probs_exp = [np.exp(p) for p in np_log_probs]
# Get predictions!
beam_predictions = beam_search_lm(
log_probs=np_log_probs_exp,
log_probs_length=None,
force_pt=True)
# -
# Now that we have beam search predictions, we want to get the best hypothesis for each evaluation sample, so we iterate through each sample and grab the top prediction.
#
# Once we have that, we process our reference transcripts and can then use them to calculate the WERs of our top predictions using beam search based on our language model!
# +
# Get the top beam search hypothesis for each sample
beam_hypotheses = []
for mini_batch in beam_predictions:
for sample_hypotheses in mini_batch:
# sample_hypotheses is a set of (probability, prediction) pairs
beam_hypotheses.append(sample_hypotheses[0][1]) # Take top prediction
# Process our reference transcripts
references = post_process_transcripts(eval_transcript, labels=labels,
transcript_len_list=eval_transcript_len)
# Calculate top beam search prediction WERs!
wer = word_error_rate(hypotheses=beam_hypotheses, references=references)
print("BEAM WER {:.2f}".format(wer*100))
# -
# ### Fast Training
#
# Last but not least, we could simply speed up training our model! If you have the resources, you can speed up training by splitting the workload across multiple GPUs. Otherwise (or in addition), there's always mixed precision training, which allows you to increase your batch size.
#
# You can read more about both mixed precision and multi-GPU training using NeMo on [this page of the documentation](https://nvidia.github.io/NeMo/training.html).
# ## Further reading/watching:
#
# That's all for now! If you'd like to learn more about the topics covered in this tutorial, here are some resources that may interest you:
# - [Stanford Lecture on ASR](https://www.youtube.com/watch?v=3MjIkWxXigM)
# - ["An Intuitive Explanation of Connectionist Temporal Classification"](https://towardsdatascience.com/intuitively-understanding-connectionist-temporal-classification-3797e43a86c)
# - [Explanation of CTC with Prefix Beam Search](https://medium.com/corti-ai/ctc-networks-and-language-models-prefix-beam-search-explained-c11d1ee23306)
# - [Listen Attend and Spell Paper (seq2seq ASR model)](https://arxiv.org/abs/1508.01211)
# - [Explanation of the mel spectrogram in more depth](https://towardsdatascience.com/getting-to-know-the-mel-spectrogram-31bca3e2d9d0)
# - [Jasper Paper](https://arxiv.org/abs/1904.03288)
# - [SpecAugment Paper](https://arxiv.org/abs/1904.08779)
# - [Explanation and visualization of SpecAugment](https://towardsdatascience.com/state-of-the-art-audio-data-augmentation-with-google-brains-specaugment-and-pytorch-d3d1a3ce291e)
# - [Cutout Paper](https://arxiv.org/pdf/1708.04552.pdf)
|
examples/asr/notebooks/1_ASR_tutorial_using_NeMo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Reservoir of Izhikevich neuron models
# In this script a reservoir of neurons models with the differential equations proposed by Izhikevich is defined.
# +
# %matplotlib inline
import pyNN.nest as p
from pyNN.random import NumpyRNG, RandomDistribution
from pyNN.utility import Timer
import matplotlib.pyplot as plt
import numpy as np
timer = Timer()
p.setup(timestep=0.1) # 0.1ms
# -
# ## Definition of Inputs
# The input can be:
# - the joint position of the robot arm (rate coded or temporal coded)
poisson_input = p.SpikeSourcePoisson(rate = 10, start = 20.)
#input_neuron = p.Population(2, p.SpikeSourcePoisson, {'rate': 0.7}, label='input')
input_neuron = p.Population(2, poisson_input, label='input')
# ## Definition of neural populations
#
# Izhikevich spiking model with a quadratic non-linearity:
#
# dv/dt = 0.04*v^2 + 5*v + 140 - u + I
#
# du/dt = a*(b*v - u)
# +
n = 1500 # number of cells
exc_ratio = 0.8 # ratio of excitatory neurons
n_exc = int(round(n*0.8))
n_inh = n-n_exc
print n_exc, n_inh
celltype = p.Izhikevich()
# default_parameters = {'a': 0.02, 'c': -65.0, 'd': 2.0, 'b': 0.2, 'i_offset': 0.0}¶
# default_initial_values = {'v': -70.0, 'u': -14.0}¶
exc_cells = p.Population(n_exc, celltype, label="Excitatory_Cells")
inh_cells = p.Population(n_inh, celltype, label="Inhibitory_Cells")
# initialize with a uniform random distributin
# use seeding for reproducability
rngseed = 98766987
parallel_safe = True
rng = NumpyRNG(seed=rngseed, parallel_safe=parallel_safe)
unifDistr = RandomDistribution('uniform', (-75,-65), rng=rng)
exc_cells.initialize(v=unifDistr)
inh_cells.initialize(v=unifDistr)
# -
# ## Definition of readout neurons
# Decide:
# - 2 readout neurons: representing in which direction to move the joint
# - 1 readout neuron: representing the desired goal position of the joint
readout_neurons = p.Population(2, celltype, label="readout_neuron")
# ## Define the connections between the neurons
# +
inp_conn = p.AllToAllConnector()
rout_conn = p.AllToAllConnector()
w_exc = 20. # later add unit
w_inh = 51. # later add unit
delay_exc = 1 # defines how long (ms) the synapse takes for transmission
delay_inh = 1
stat_syn_exc = p.StaticSynapse(weight =w_exc, delay=delay_exc)
stat_syn_inh = p.StaticSynapse(weight =w_inh, delay=delay_inh)
weight_distr_exc = RandomDistribution('normal', [w_exc, 1e-3], rng=rng)
weight_distr_inh = RandomDistribution('normal', [w_inh, 1e-3], rng=rng)
exc_synapse = p.TsodyksMarkramSynapse(U=0.04, tau_rec=100.0, tau_facil=1000.0,
weight=weight_distr_exc, delay=lambda d: 0.1+d/100.0)
inh_synapse = p.TsodyksMarkramSynapse(U=0.04, tau_rec=100.0, tau_facil=1000.0,
weight=weight_distr_inh, delay=lambda d: 0.1+d/100.0)
# tau_rec: depression time constant (ms)
# tau_facil: facilitation time constant (ms)
pconn = 0.01 # sparse connection probability
exc_conn = p.FixedProbabilityConnector(pconn, rng=rng)
inh_conn = p.FixedProbabilityConnector(pconn, rng=rng)
connections = {}
connections['e2e'] = p.Projection(exc_cells, exc_cells, exc_conn,
synapse_type=stat_syn_exc, receptor_type='excitatory')
connections['e2i'] = p.Projection(exc_cells, inh_cells, exc_conn,
synapse_type=stat_syn_exc,receptor_type='excitatory')
connections['i2e'] = p.Projection(inh_cells, exc_cells, inh_conn,
synapse_type=stat_syn_inh,receptor_type='inhibitory')
connections['i2i'] = p.Projection(inh_cells, inh_cells, inh_conn,
synapse_type=stat_syn_inh,receptor_type='inhibitory')
connections['inp2e'] = p.Projection(input_neuron, exc_cells, inp_conn,
synapse_type=stat_syn_exc,receptor_type='excitatory')
connections['inp2i'] = p.Projection(input_neuron, inh_cells, inp_conn,
synapse_type=stat_syn_exc,receptor_type='excitatory')
connections['e2rout'] = p.Projection(exc_cells, readout_neurons, rout_conn,
synapse_type=stat_syn_exc,receptor_type='excitatory')
connections['i2rout'] = p.Projection(inh_cells, readout_neurons, rout_conn,
synapse_type=stat_syn_inh,receptor_type='inhibitory')
# -
# ## Setup recording and run the simulation
readout_neurons.record(['v','spikes'])
exc_cells.record(['v','spikes'])
p.run(1000)
# ## Plotting the Results
# +
p.end()
data_rout = readout_neurons.get_data()
data_exc = exc_cells.get_data()
# -
fig_settings = {
'lines.linewidth': 0.5,
'axes.linewidth': 0.5,
'axes.labelsize': 'small',
'legend.fontsize': 'small',
'font.size': 8
}
plt.rcParams.update(fig_settings)
plt.figure(1, figsize=(6,8))
def plot_spiketrains(segment):
for spiketrain in segment.spiketrains:
y = np.ones_like(spiketrain) * spiketrain.annotations['source_id']
plt.plot(spiketrain, y, '.')
plt.ylabel(segment.name)
plt.setp(plt.gca().get_xticklabels(), visible=False)
def plot_signal(signal, index, colour='b'):
label = "Neuron %d" % signal.annotations['source_ids'][index]
plt.plot(signal.times, signal[:, index], colour, label=label)
plt.ylabel("%s (%s)" % (signal.name, signal.units._dimensionality.string))
plt.setp(plt.gca().get_xticklabels(), visible=False)
plt.legend()
# Plot readout neurons
# +
n_panels = sum(a.shape[1] for a in data_rout.segments[0].analogsignalarrays) + 2
plt.subplot(n_panels, 1, 1)
plot_spiketrains(data_rout.segments[0])
panel = 3
for array in data_rout.segments[0].analogsignalarrays:
for i in range(array.shape[1]):
plt.subplot(n_panels, 1, panel)
plot_signal(array, i, colour='bg'[panel%2])
panel += 1
plt.xlabel("time (%s)" % array.times.units._dimensionality.string)
plt.setp(plt.gca().get_xticklabels(), visible=True)
plt.savefig("neo_example.png")
# -
# Plot excitatory cells
# +
n_panels = sum(a.shape[1] for a in data_exc.segments[0].analogsignalarrays) + 2
plt.subplot(n_panels, 1, 1)
plot_spiketrains(data_exc.segments[0])
panel = 3
for array in data_exc.segments[0].analogsignalarrays:
for i in range(array.shape[1]):
plt.subplot(n_panels, 1, panel)
plot_signal(array, i, colour='bg'[panel%2])
panel += 1
plt.xlabel("time (%s)" % array.times.units._dimensionality.string)
plt.setp(plt.gca().get_xticklabels(), visible=True)
plt.savefig("neo_example.png")
# -
|
src/experimental_code/.ipynb_checkpoints/Izh_reservoir_stdp-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 6 Strings
# ### 6.1 A string is a sequence
fruit = 'banana'
letter = fruit[1]
print(letter)
letter = fruit[0]
print(letter)
# ### 6.2 Getting the length of a string
length = len(fruit)
print(length)
last = fruit[length-1]
print(last)
# ### 6.3 Traversal through a string with a loop
index = 0
while index < len(fruit):
letter = fruit[index]
print(letter)
index = index + 1
for char in fruit:
print(char)
# Exercise 1
index = len(fruit) - 1
while index >= 0:
letter = fruit[index]
print(letter)
index = index - 1
# ### 6.4 String slices
s = 'Monty Python'
print(s[0:5])
print(s[6:12])
fruit = 'banana'
fruit[:3]
fruit[3:]
fruit[3:3]
# Exercise 2
fruit[:]
fruit
# ### 6.5 Strings are immutable
greeting = 'Hello world!'
greeting[0] = 'J'
new_greeting = 'J' + greeting[1:]
print(new_greeting)
# ### 6.6 Looping and counting
word = 'banana'
count = 0
for letter in word:
if letter == 'a':
count = count + 1
print(count)
# Exercise 3
def charcount(s, c):
count = 0
for letter in s:
if letter == c:
count = count + 1
return count
charcount('banana', 'a')
# ### 6.7 The IN operator
'a' in 'banana'
'seed' in 'banana'
# ### 6.8 String comparison
# word is assigned above
if word == 'banana':
print('All right, bananas.')
word = 'pineapple'
if word < 'banana':
print('Your word, ' + word + ', comes before banana.')
elif word > 'banana':
print('Your word, ' + word + ', comes after banana.')
else:
print('All right, bananas.')
# ### 6.9 String methods (capitalize, upper, lower, find, strip, startswith)
# ##### Also see https://docs.python.org/library/stdtypes.html#string-methods
stuff = 'Hello world'
type(stuff)
# list available string methods (functions)
dir(stuff)
help(str.capitalize)
name = 'lisa'
name.capitalize()
name.upper()
name = 'ELIZA'
name.lower()
name.lower().capitalize()
word = 'banana'
index = word.find('a')
print(index)
word.find('na')
# find 'na' - start looking at character with index 3
word.find('na', 3)
# remove leading and trailing spaces
line = ' Here we go! '
line.strip()
line = 'Have a nice day!'
line.startswith('Have')
line.startswith('h')
line.lower().startswith('h')
# ### 6.10 Parsing strings
data = 'From <EMAIL> Sat Jan 5 09:14:16 2008'
atpos = data .find('@')
print(atpos)
sppos = data.find(' ', atpos)
print(sppos)
host = data[atpos+1:sppos]
print(host)
# ### 6.11 Format operator (%d formats integers; %g formats floats, %s formats strings)
# ##### Also see https://docs.python.org/library/stdtypes.html#printf-style-string-formatting
camels = 42
'%d' % camels
'I have spotted %d camels.' % camels
'In %d years, I have spotted %g %s.' % (3, 0.1, 'camels')
'%d %d %d' % (1, 2)
'%d' % 'dollars'
|
06-strings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # VGP 245 Assignment 1
# # Python Basics
# ### Question 1 - Object Types and Data Structures
#
# Give briefly describe and give an example for each of the following types
#
# Number:
#
# Strings:
#
# Lists:
#
# Dictionaries:
#
# Tuples:
#
# ### Question 2 - Numbers
#
# Write an equation that can calculate the total amount earned in a week if a person worked 4 hours a day, 5 days a week, and
# $25.20 per hour.
#
# ### Question 3 - Numbers
# What is the result of the following?
#
# * 4 * (6 + 5)
# * 4 * 6 + 6
# * 4 + 6 * 5
#
# ### Question 4 - Numbers
#
# Write an example of getting the square of 87 and the square root of 2304
import math
print(87**2)
print(math.sqrt(2304))
# ### Question 5 - Strings
# Given the following string 'Master Chef', output the sub string 'Chef' and then reverse it
name = '<NAME>'
# output the second character of the string above
print(name[1])
# output Chef from above into a new variable and print it out
name2 = name[7:11]
print(name2)
# output the reverse of the variable above and print it out
print(name2[::-1])
# ### Question 6 - List
# Build the list ['Billy', 'Jason', 'Tommy'] in 2 different ways
#
# Method 1
list1 = ["Billy", "Jason", "Tommy"]
# Method 2
list2 = ("Billy", "Jason", "Tommy")
# Add an 'Zack' and 'Kimberly' into the list above and remove 'Tommy'
# Add Zack and Kimberly to the list you made in Question 6
list1.append("Zack")
list1.append("Kimberly")
# +
# Remove Tommy from the list
del list1[2]
# -
# ### Question 7 - List
# Sort the list below
#
my_numbers = [41, 32, 5, 37, 27, 14, 24, 13, 6]
# hint there's a built-in function for that
# ### Question 8 - Dictionaries
#
# Using the dictionary below, find the value from a given key.
# +
rangers = {'Jason': 'Red', 'Tommy': 'Green', 'Billy': 'Blue', 'Zack': 'Black'}
# find out which color is Tommy from the dictionary above and print it out
print(rangers['Tommy'])
# Tommy became the 'White' ranger and is no longer the green ranger
# Update the Tommy to reflect the changes and print it out
rangers['Tommy'] = 'White'
print(rangers['Tommy'])
# It turns out there are other rangers, put them together into a list of dictionaries
zeo_ranger = {'Tommy': 'Red', 'Adam': 'Black', 'Rocky': 'Blue', 'Kat': 'Pink', 'Jason': 'Gold'}
turbo_ranger = {'Tommy': 'Red', 'Adam': 'Green', 'Justin': 'Blue', 'Kat': 'Pink'}
space_ranger = {'Andros': 'Red', 'Carlos': 'Black', 'TJ': 'Blue', 'Cassie': 'Pink'}
all_the_rangers = {'Tommy': 'Red', 'Adam': 'Black', 'Rocky': 'Blue', 'Kat': 'Pink', 'Jason': 'Gold',
'Tommy': 'Red', 'Adam': 'Green', 'Justin': 'Blue', 'Kat': 'Pink',
'Andros': 'Red', 'Carlos': 'Black', 'TJ': 'Blue', 'Cassie': 'Pink'}
# Find out the different how many rangers are there total
# hint you will need a nested for loop
# -
# ### Quesiton 9 - Tuple
# What is the difference between a Tuple and a list?
# +
tuple_of_fruits = ('apple', 'berries', 'cherries')
list_of_fruits = ['apple', 'berries', 'cherries']
# list 2 things and show an example
# 1.tuples and lists are enclosed differently tuples use () while lists use []
list1 = [1, 2, 3]
tuple1 = (7, 8, 9)
# 2.unlike list once a tuple is made it cannot be changed
del tuple1[1]
# -
# ### Question 10 Boolean
# Using the dictionary in question 8, do the following:
#
# using the dictionary `rangers` check if the black ranger is 'Adam', and print the result
if rangers['Zack'] == 'Black':
print("Adam is the black ranger")
else:
print("Adam is not the black ranger")
# Check if the blue ranger is not Bobby
if rangers['Billy'] == 'bobby':
print('true')
else:
print('false')
# What does the 2.0 == 2 return?
if 2.0 == 2:
print('true')
else:
print('false')
# What does 2 > 5 return?
if 2 > 5:
print('true')
else:
print('false')
# What does 2 <= 1 return?
if 2 <= 1:
print('true')
else:
print('false')
# That's it. You can complete this and add it into your GitHub repo, and send me a message on slack or submit it on omnivox. See you in class.
|
Class 1/homework1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <a id="top"></a>
# # MIRI MRS Spectroscopy of a Late M Star
# -
# **Use case:** Extract spatial-spectral features from IFU cube and measure their attributes.<br>
# **Data:** KMOS datacube of point sources in the LMC from Jones et al. (in prep).<br>
# **Tools:** specutils, spectral_cube, photutils, astropy, aplpy, scipy.<br>
# **Cross-intrument:** MIRI<br>
# **Documentation:** This notebook is part of a STScI's larger [post-pipeline Data Analysis Tools Ecosystem](https://jwst-docs.stsci.edu/jwst-post-pipeline-data-analysis).<br>
#
# **Note**: Ultimately, this notebook will include MIRI simulated data cubes obtained using MIRISim (https://wiki.miricle.org//bin/view/Public/MIRISim_Public)
# and run through the JWST pipeline (https://jwst-pipeline.readthedocs.io/en/latest/) of
# point sources with spectra representative of late M type stars.
#
# ## Introduction
#
# This notebook analyzes one star represented by a dusty SED corresponding to the ISO SWS spectrum of
# W Per from Kraemer et al. (2002) and Sloan et al. (2003) to cover the MRS spectral range 5-28 microns. Analysis of JWST spectral cubes requires extracting spatial-spectral features of interest and measuring their attributes.
#
# The first part of the notebook will process the datacube and automatically detect and extract spectra (summed over its spatial region) for all point sources in the cube. Then it will read in a datacube generated at Stage 3 of the JWST pipeline or use near-IR data from KMOS as a representative example of an IR data cube. The analysis will use `photutils` to automatically detect sources in the continuum image and use an aperture mask generated with `spectral-cube` to extract the spectra of each point source in the data cube.
#
# The second part of the notebook will perform data analysis using `specutils`. Specifically, it will fit a model photosphere/blackbody to the spectra. Then it will calculate the centroids, line integrated flux and equivalent width for each dust and molecular feature.
#
# ## To Do:
# - Replace KMOS data cube with JWST/MIRI simulation of an M star ran through JWST piplieline.
# - Make function to extract spectra from datacube using an apeture.
# - Replace blackbody fit to the photosphere part of the spectra with a stellar photosphere model.
# - Make sure errors have been propagated correctly in the caculation of centroids, line integrated flux and
# equivalent widths.
# - Make simple function within the `specutils` framework to fit a continium and measure centroids, line integrated flux and
# equivalent widths of broad solid state and molecular features.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Imports
# + slideshow={"slide_type": "fragment"}
# Import useful python packages
import numpy as np
# Import packages to display images inline in the notebook
import matplotlib.pyplot as plt
# %matplotlib inline
# Set general plotting options
params={'legend.fontsize':'18','axes.labelsize':'18',
'axes.titlesize':'18','xtick.labelsize':'18',
'ytick.labelsize':'18','lines.linewidth':2,'axes.linewidth':2,'animation.html': 'html5'}
plt.rcParams.update(params)
plt.rcParams.update({'figure.max_open_warning': 0})
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
# Import astropy packages
from astropy import units as u
from astropy.io import ascii
from astropy.wcs import WCS
from astropy.table import Table, vstack
from astropy.stats import sigma_clipped_stats
from astropy.nddata import StdDevUncertainty
# Import packages to deal with spectralcubes
from spectral_cube import SpectralCube
# To find stars in the MRS spectralcubes and do aperture photometry
from photutils import DAOStarFinder, CircularAperture
# To deal with 1D spectrum
from specutils import Spectrum1D
from specutils.fitting import fit_generic_continuum
from specutils.manipulation import box_smooth, extract_region, SplineInterpolatedResampler
from specutils.analysis import line_flux, centroid, equivalent_width
from specutils.spectra import SpectralRegion
# To make nice plots with WCS axis
import aplpy
# To fit a curve to the data
from scipy.optimize import curve_fit
# -
# ## Set paths to the Data and Outputs
#
# For now use KMOS data cube of YSOs in the LMC from Jones et al in prep.
#
# TODO: Update with MIRISim JWST pipeline processed data in future itterations.
# +
# Setup an input directory where relevant data is located
data_in_path = "https://data.science.stsci.edu/redirect/JWST/jwst-data_analysis_tools/MRS_Spectroscopy_Late_M_Star/"
data_cube_file = data_in_path + "NGC346_K_2c_COMBINED_CUBE_Y551.fits"
# Path to output directory
data_out_path = "./"
# Setup an output directory to save the extracted 1D spectra
outdir_spectra = data_out_path + '/spectra/'
# -
# Some housekeeping if using the KMOS data rather than simulated JWST/MRS data
# Define good wavelength ranges for each grating from which to make the data cube
YJgrating = [1.02, 1.358] # microns
Hgrating = [1.44, 1.85] # microns
Kgrating = [2.1, 2.42] # microns
# + [markdown] slideshow={"slide_type": "slide"}
# ## Load and Display the Data cube
#
# **Developer note** Note the `SpectralCube` package is designed for sub-mm/radio data it expects a beam!
# This is preferred to other packages available due to much of its functionality and ease of use.
# JWST NIRSpec and MIRI both have instruments that give data cubes (with two positional dimensions and one spectral
# dimension) as the final pipeline product, as do many ground based telescopes, which do not have a beam.
#
#
# https://spectral-cube.readthedocs.io/en/stable/index.html
# -
cube = SpectralCube.read(data_cube_file, hdu=1)
print(cube)
# +
# Cube dimensions and trimming
# Data order in cube is (n_spectral, n_y, n_x)
# Trim the ends of the cube where the data quality is poor
subcube = cube.spectral_slab(Kgrating[0] * u.micron, Kgrating[1] * u.micron)
# Rename subcube to equal cube - done in case step above is not necessary
cube = subcube
# Chop out the NaN borders
cmin = cube.minimal_subcube()
# +
# Make a continuum image (Sum/average over Wavelength)
# Note: many mathematical options are available median is preferred
cont_img = cmin.median(axis = 0)
# Extract the target name
name_long = cont_img.header["OBJECT"]
name, _ = name_long.split("/")
# -
# Quick plot the continuum image now the NaN borders removed
plt.imshow(cont_img.value)
plt.tight_layout()
plt.show()
#Plot the continuum in WCS
F = aplpy.FITSFigure(cont_img.hdu, north = True)
F.show_colorscale()
F.add_label(0.1, 0.9, name, relative = True, size = 22, weight = 'bold')
F.axis_labels.set_font(size = 22)
F.tick_labels.set_font(size = 18, stretch = 'condensed')
# ## Now to detect the point source in the datacube and extract and plot the spectra for each source
#
# **Developer note** Finding a way to streamline the process of detecting sources within a data cube and extracting their
# spectra would be extremely valuable.
#
# For data cubes like the JWST/MIRI MRS information on the point sources in the FOV and also obtaining a source subtracted
# data cube will be necessary (See the `PampelMuse` software for an example on how spectral extraction is implemented for
# near-IR data cubes like MUSE).
#
# Note these backgrounds of diffuse emission can be quite complex.
#
# On these source extracted data cubes (see `SUBTRES` in `PampelMuse`) I would like to produce moment maps
# (https://casa.nrao.edu/Release3.4.0/docs/UserMan/UserManse41.html) and Position-Velocity (PV) diagrams
# (https://casa.nrao.edu/Release4.1.0/doc/UserMan/UserManse42.html).
#
# ### 1) Use `Photutils` to detect stars/point sources in the continuum image
#
# The first step of the analysis is to identify those sources for which it is feasible to extract spectra from the IFU
# data. Ideally we can estimate the signal-to-noise ratio (S/N) for all sources in the cube, do a number of checks to
# determine the status of every source and loop through these (brightest first) to extract the spectra.
#
# ### 2) Extract the spectra from the datacube using `SpectralCube`
#
# **Note** There are multiple ways of extracting spectra from datacubes. The simplest is slice the cube along a single
# pixel but this is not ideal for point sources which should cover multiple pixels.
# Here I use *Aperture Extraction*.
#
# - The flux from each point source was obtained via a circular aperture. This requires you to mask the data, and make a
# circular mask and a maskedsubcube.
#
# - A background measured using a square/rectangular aperture sliced in pixel coordinates to produce a sub-cube.
#
# - A annulus surrounding the point source to measure the local background.
#
# - Using predefined regions from DS9 etc. to create a mask [`Not used here`].
#
# *If have a small number of data cubes selecting the source extraction region and background region manually using
# `cubeviz` would be useful here.*
#
# Mathematical operation e.g. `max, mean, sum, median, std` should then be applied to the region in the aperture.
#
# Below I show a few different options from the simple to the complex, which takes into account the background emission
# within the data cube. Taking into account the background may not always be the preferred method but the option should
# always be available when using an aperture extraction.
#
# #### Steps to find the background
#
# 1) Define a background region either as an annulus or as a rectangle away from the source
#
# 2) Find the median of all the background pixels to account for variations
#
# 3) Find number of pixels in background and number of pixels in the point source aperture
#
# 4) Find the sum of all the pixels in the point source aperture
#
# 5) Correct for background using the sum star flux minus median of background * pixels in star aperture
#
#
#
# **Advanced Developer Note** Using Aperture Extraction to obtain the spectra for each source in the data cube is still
# very simplistic. It should be noted that the MIRI aperture changes as a function of wavelength, the steps above do not
# account for this.
# A good example of software that looks at extracting point sources from datacubes is: `PampelMuse`, by <NAME>.
# https://gitlab.gwdg.de/skamann/pampelmuse; https://ui.adsabs.harvard.edu/abs/2013A%26A...549A..71K/abstract
#
# An `optimal spectrum extraction` procedure would take into account the varying PSF through the cube, to produce an
# accurate spectra with the maximum possible signal-to-noise ratio. This weights the extracted data by the S/N of each
# pixel (Horne 1986) and would be ideal for when there is a complex background or for extracting spatially-blended source.
# For small cubes its best to fit a PSF profile to all resolved sources simultaneously, but this might not be possible in
# larger data sets.
#
# **Advanced Developer Note 2** In dense fields like globular clusters, with a significant number of unresolved sources or
# in embedded star-forming clusters, a more advanced treatment of the background would be necessary. For instance using a
# homogeneous grid across the field of view with parameters controlling the bin size would be ideal. If a variable
# background is not accounted for in a PSF extraction systematic residuals in the data would be present where background
# is over or underestimated.
#
#
# ## Detect, extract and plot 1D spectrum of each source in the cube
#
# ### First automatically identify all the point sources in the cube using `photutils`
# Make an array to store results of the source detection within the data cube
name_val = []
source_val = []
ra_val =[]
dec_val =[]
# +
# Crop out Edges and take absolute value of the continuum image
cont_img = cont_img[1:13, 1:13]
# Find the background in the collapsed datacube
mean, median, std = sigma_clipped_stats(cont_img.value, sigma = 2.0)
# Get a list of sources using a dedicated source detection algorithm
# Find sources at least 3* background (typically)
daofind = DAOStarFinder(fwhm = 2.0, threshold = 3. * std)
sources = daofind(cont_img.value - median)
print("\n Number of sources in field: ", len(sources))
# -
# ### If point sources are present in the cube extract and plot the spectrum of each source
#
# #### In the cell below we:
#
# 1) Extract a spectra for each detected object using aperture photometry, and a circular masked region.
#
# 2) Make an estimate of the background in the datacube using both: an annulus around each source and a box region away
# from the source - this box and annulus is hard coded and not ideal for other datasets or multiple cubes.
#
# 3) Generate a background corrected spectrum.
#
# 4) Plots the spectra and its various background corrected versions.
#
# 5) Convert the spectra into Jy.
#
# 6) Write each of the spectra to a file. (They could be put into a `specutils` `Spectrum1D` object at this stage but I
# have not done this here.) This file is loaded by all other routines to do analysis on the data.
# +
if len(sources) > 0:
print()
for col in sources.colnames:
sources[col].info.format = '%.8g' # for consistent table output
print(sources)
# From the list of sources in the field get their RA and DEC (ICRS)
print()
# Positions in pixels
positions = Table([sources['xcentroid'], sources['ycentroid']])
# Instantiate WCS object
w = WCS(cont_img.header)
# Convert to RA & Dec (ICRS)
radec_lst = w.pixel_to_world(sources['xcentroid'], sources['ycentroid'])
#-----------------------------------------------------
# We are now entering a loop which does multiple processing steps on each
# point source detected in the cube
for count_s, _ in enumerate(sources):
print(radec_lst[count_s].to_string('hmsdms'))
name_val.append(name)
source_val.append(count_s)
ra_val.append(radec_lst[count_s].ra.deg)
dec_val.append(radec_lst[count_s].dec.deg)
#-----------------------------------------------------
# Aperture Extract spectrum of point source - using a circular aperture
# Size of frame
ysize_pix = cmin.shape[1]
xsize_pix = cmin.shape[2]
# Set up some centroid pixel for the source
ycent_pix = sources['ycentroid'][count_s]
xcent_pix = sources['xcentroid'][count_s]
# Make an aperture radius for source
# If made into a function this value should not be hardcoded
aperture_rad_pix = 2
# Make a masked array for the aperture
yy, xx = np.indices([ysize_pix,xsize_pix], dtype = 'float')
radius = ((yy-ycent_pix)**2 + (xx-xcent_pix)**2)**0.5
# Select pixels within the aperture radius
mask = radius <= aperture_rad_pix
# Make a masked cube
maskedcube = cmin.with_mask(mask)
# Pixels in aperture
pix_in_ap = np.count_nonzero(mask == 1)
# Extract the spectrum from only the circular aperture - use sum
spectrum = maskedcube.sum(axis = (1,2))
# Extract the noise spectrum for the source
noisespectrum = maskedcube.std(axis = (1,2))
# Measure a spectrum from the background - Use an annulus around the source
# NOTE: Hardcoded values in for annulus size - improve
# Select pixels within an annulus
an_mask = (radius > aperture_rad_pix + 1) & (radius <= aperture_rad_pix + 2)
# Make a masked cube
an_maskedcube = cmin.with_mask(an_mask)
# Extract the background spectrum from only the annulus
bkg_spectrum = an_maskedcube.median(axis = (1,2))
# Background corrected spectrum - annulus
corr_sp = spectrum - (bkg_spectrum * pix_in_ap)
# Try measuring a spectrum from the background -> Use a box away from source.
# NOTE: Hardcoded values in for box region - improve
bkgcube = cmin[: , 1:3, 10:13]
bkgbox_spectrum = bkgcube.median(axis = (1,2))
bkg_img = bkgcube.median(axis = 0)
# Background corrected spectrum - box
corr_sp_box = spectrum - (bkgbox_spectrum * pix_in_ap)
#-----------------------------------------------------
# Plot the spectrum extracted from circular aperture via: a sum extraction
plt.figure(figsize = (10,5))
plt.plot(maskedcube.spectral_axis.value, spectrum.value,
label = 'Source')
plt.plot(maskedcube.spectral_axis.value, corr_sp.value,
label = 'Bkg Corr')
plt.plot(maskedcube.spectral_axis.value, corr_sp_box.value,
label = 'Bkg Corr box')
plt.xlabel('Wavelength (microns)')
plt.ylabel(spectrum.unit)
plt.gcf().text(0.5, 0.85, name, fontsize = 14, ha = 'center')
plt.gcf().text(0.5, 0.80, radec_lst[count_s].to_string('decimal'),
ha = 'center', fontsize=14)
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
#-----------------------------------------------------
# Convert flux from erg / (Angstrom cm2 s) to Jy
spectrumJy = spectrum.to(
u.Jy, equivalencies = u.spectral_density(maskedcube.spectral_axis))
corr_sp_Jy = corr_sp.to(
u.Jy, equivalencies = u.spectral_density(maskedcube.spectral_axis))
corr_sp_box_Jy = corr_sp_box.to(
u.Jy, equivalencies= u.spectral_density(maskedcube.spectral_axis))
noiseSp_Jy = noisespectrum.to(
u.Jy, equivalencies = u.spectral_density(maskedcube.spectral_axis))
#-----------------------------------------------------
# Save each extracted spectrum to a file
# Set an output name
spec_outname = name + "_" + str(count_s) + "_" + "spec"
# Make output table
specdata_tab = Table([maskedcube.spectral_axis, corr_sp_Jy, noiseSp_Jy,
spectrumJy, corr_sp_box_Jy],
names=['wave_mum', 'cspec_Jy', 'err_fl_Jy',
'spec_Jy', 'cSpec_box_Jy'])
# Write the file
# ascii.write(specdata_tab, outdir_spectra + spec_outname +".csv",
# format = 'csv', overwrite = True)
#-----------------------------------------------------
# Do aperture photometry on the sources - Only if using sum of image
# Take list of star positions from DAOFIND use this to define an aperture
if len(sources) == 2: # To overcome in array order
sources = vstack([sources, sources])
positions_pix = (sources['xcentroid'], sources['ycentroid'])
else:
positions_pix = (sources['xcentroid'], sources['ycentroid'])
apertures = CircularAperture(positions_pix, r = 2.) # Aperture radius = 2 pixels
#-----------------------------------------------------
# As a check to make sure all obvious point sources have been identified
# plot the cube with the NaN borders removed and overplot the apertures
# for the extracted sources
plt.figure()
plt.subplot(1, 2, 1)
plt.imshow(cont_img.value, cmap='Greys', origin='lower')
apertures.plot(color='blue', lw=1.5, alpha=0.5)
plt.subplot(1, 2, 2)
plt.imshow(cont_img.value, origin='lower')
plt.tight_layout()
plt.show()
plt.close()
else:
# Plot the cube with the NaN borders removed
plt.figure()
plt.imshow(cont_img.value, origin='lower')
plt.tight_layout()
plt.show()
plt.close()
# -
# Make table of extracted sources
source_extspec_tab = Table([name_val, source_val, ra_val, dec_val],
names = ("name", "source_no", "ra", "dec"))
print(source_extspec_tab)
# ## Data analysis - on the extracted spectra using `specutils`
# With the present lack of JWST flight data, we instead use the SWS spectra of an dusty AGB star, a cool M-star.
# +
# Set the paths to the spectral data extracted from the datacube above
dusty_AGB_spec_file = data_in_path + '63702662.txt'
spectra_file = dusty_AGB_spec_file
# +
# Read in the spectra - as saved as text files & do some housekeeping
data = ascii.read(spectra_file)
if data.colnames[0] == 'col1':
data['col1'].name = 'wave_mum'
data['col2'].name = 'cspec_Jy'
data['col3'].name = 'err_fl_Jy'
wav = data['wave_mum'] * u.micron # Wavelength: microns
fl = data['cspec_Jy'] * u.Jy # Fnu: Jy
efl = data['err_fl_Jy'] * u.Jy # Error flux: Jy
# Make a 1D spectrum object
spec = Spectrum1D(spectral_axis = wav, flux = fl, uncertainty = StdDevUncertainty(efl))
# -
# **Note** Reading in a spectra comprised of multiple spectral components this file may have a spectral order column. In
# many instances these orders are not correctly stitched together due to issues with background and flux calibration. A
# spectral file with an order column that can read into the `Spectrum1D` is necessary to do corrections and scaling on
# each segment individually to fix the jumps between the spectra.
# +
# Apply a 5 pixel boxcar smoothing to the spectrum
spec_bsmooth = box_smooth(spec, width = 5)
# Plot the spectrum & smoothed spectrum to inspect features
plt.figure(figsize = (8,4))
plt.plot(spec.spectral_axis, spec.flux, label = 'Source')
plt.plot(spec.spectral_axis, spec_bsmooth.flux, label = 'Smoothed')
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
# -
# ### Fit a continuum - find the best-fitting template (stellar photosphere model or blackbody)
#
# **Note** - Would idealy like to fit the photosphere with a set of Phoenix Models - but cant get that to work.
# I think `template_comparison` may be a good function here to work with the Phoenix Models which have been setup to
# interface with `pysynphot`.
#
# For now switching to a blackbody.
#
# - For AGB stars with a photosphere component fit a stellar photosphere model or a blackbody to short wavelength end of
# the spectra
def blackbody_Fnu(lam, T, A):
""" Blackbody as a function of wavelength (um) and temperature (K).
Function returns the Planck function in f_nu units
# [Y Jy] = 1.0E+23 * [X erg/cm^2/s/Hz] = 10E+26 [X Watts/m^2/Hz]
"""
from scipy.constants import h, k, c
lam = 1e-6 * lam # convert to metres
bb_nu = 2*h*c / (lam**3 * (np.exp(h*c / (lam*k*T)) - 1)) # units of W/m^2/Hz/Steradian ; f_nu units
return A * bb_nu
# +
# Only want to fit to a small wavelength range at the start of the spectra
phot_fit_region = [3.0, 9.4] # Microns
# Trim the specrum to the region showing a stellar photosphere
sub_region_phot = SpectralRegion([(phot_fit_region[0], phot_fit_region[1])] * u.micron)
sub_spectrum_phot = extract_region(spec, sub_region_phot)
# +
# fit BB to the data
def phot_fn(wa, T1, A):
return blackbody_Fnu(wa, T1, A)
popt, pcov = curve_fit(phot_fn, sub_spectrum_phot.spectral_axis.value,
sub_spectrum_phot.flux.value, p0=(3000, 10000),
sigma=sub_spectrum_phot.uncertainty.quantity)
# Get the best fitting parameter value and their 1 sigma errors
best_t1, best_a1 = popt
sigma_t1, sigma_a1 = np.sqrt(np.diag(pcov))
ybest = blackbody_Fnu(spec.spectral_axis.value, best_t1, best_a1)
print ('Parameters of best-fitting model:')
print ('T1 = %.2f +/- %.2f' % (best_t1, sigma_t1))
degrees_of_freedom = len(sub_spectrum_phot.spectral_axis.value) - 2
resid = (sub_spectrum_phot.flux.value - phot_fn(sub_spectrum_phot.spectral_axis.value, *popt)) \
/ sub_spectrum_phot.uncertainty.quantity
chisq = np.dot(resid, resid)
print ('nchi2 %.2f' % (chisq.value / degrees_of_freedom))
# +
# Plot the spectrum & the model fit to the short wavelength region of the data.
plt.figure(figsize = (8,4))
plt.plot(spec.spectral_axis, spec.flux, label = 'Source')
plt.plot(spec.spectral_axis, ybest, label = 'BB')
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("Spectrum with blackbody fit")
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
# Now subtract the BB and plot the underlying dust continuum
plt.figure(figsize = (8,4))
plt.plot(spec.spectral_axis, spec.flux.value - ybest, label = 'Dust spectra')
plt.axhline(0, color='r', linestyle = 'dashdot', alpha=0.5)
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("Continuum-subtracted spectrum")
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
# -
# ### Now have the dust continuum want to look for features and measure their properties.
#
# Want to find:
# - Equivalent width
# - Equivalent flux
# - Optical depth
# - Centroids = wavelength with half the flux on either side
#
# #### As an example lets focus on the amorphous silicate 10 micron region.
#
# **Method - used repeatedly**
#
# - Fit a spline to the photosphere continuum subtracted spectra excluding the feature in this fit.
# - Trim the spectra to that wavelength region as the spline is now a different size to the full wavelength range of the
# spectra.
# - Make a continuum subtracted and and continuum normalised spectra.
# - Convert the units of the flux from Jy to W/m^2/wavelength for nice units post line integration.
# - Determine the feature line flux in units of W/m^2 and the feature centroid. Use continuum subtracted spectra.
# - Determine the feature equivalent width. Use continuum normalised spectra.
# - Make sure errors have been propagated correctly.
# - Store these results in a table
# - Several molecular and dust features are normally present in the spectra. Repeat for each feature.
#
# **Note**
# This seems like a long winded way to do this. Is there a simpler approach?
#
# > For instance a tool that takes four wavelengths, fits a line using the data from lam0 to lam1 and lam2 to lam3, then
# >passes the continuum-subtracted spectrum for line integration from lam1 to lam2 with error propagation is needed
# >several times for dust features. But with the current spectra1d framework this takes many steps to write manually and
# >is beyond tedious after doing this for 2 features let alone 20+. Similar framework is also needed for the integrated
# >line centroid with uncertainty, and the extracted equivalent width.
# +
# Fit a spline to the 10 micron feature to isolate it.
bbsub_spectra = spec - ybest # continuum subtracted spectra - Dust only
# Fit a local continuum between the flux densities at: 8.0 - 8.1 & 14.9 - 15.0 microns
# (i.e. excluding the line itself)
sw_region = 8.0 #lam0
sw_line = 8.1 #lam1
lw_line = 14.9 #lam2
lw_region = 15.0 #lam3
# Zoom in on the line complex & extract
line_reg_10 = SpectralRegion([(sw_region*u.um, lw_region*u.um)])
line_spec = extract_region(bbsub_spectra, line_reg_10)
# Fit a local continuum - exclude the actual dust feature when doing the fit
lgl_fit = fit_generic_continuum(line_spec,
exclude_regions = SpectralRegion([(sw_line*u.um,
lw_line*u.um)]))
# Determine Y values of the line continuum
line_y_continuum = lgl_fit(line_spec.spectral_axis)
#-----------------------------------------------------------------
# Generate a continuum subtracted and continuum normalised spectra
line_spec_norm =line_spec / line_y_continuum
line_spec_consub = line_spec - line_y_continuum
#-----------------------------------------------------------------
# Plot the dust feature & continuum fit to the region
plt.figure(figsize = (8, 4))
plt.plot(line_spec.spectral_axis, line_spec.flux.value,
label = 'Dust spectra 10 micron region')
plt.plot(line_spec.spectral_axis, line_y_continuum, label = 'Local continuum')
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("10$\mu$m feature plus local continuum")
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
#-----------------------------------------------------------------
# Plot the continuum subtracted 10 micron feature
plt.figure(figsize = (8,4))
plt.plot(line_spec_consub.spectral_axis, line_spec_consub.flux,
label = 'continuum subtracted')
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("Continuum subtracted 10$\mu$m feature")
plt.tight_layout()
plt.show()
plt.close()
# +
# Calculate the Line flux; Line Centroid; Equivalent width
# NOTE: Where are errors computed with these functions?
line_centroid = centroid(line_spec_consub, SpectralRegion(sw_line*u.um, lw_line*u.um))
line_flux_val = line_flux(line_spec_consub, SpectralRegion(sw_line*u.um, lw_line*u.um))
equivalent_width_val = equivalent_width(line_spec_norm)
# Hack to convert the line flux value into more conventional units
# Necessary as spectra has mixed units: f_nu+lambda
line_flux_val = (line_flux_val * u.micron).to(u.W * u.m**-2 * u.micron,
u.spectral_density(line_centroid)) / u.micron
print("Line_centroid: {:.6} ".format(line_centroid))
print("Integrated line_flux: {:.6} ".format(line_flux_val))
print("Equivalent width: {:.6} ".format(equivalent_width_val))
# -
# **Developer note** The hack in the cell above is necessary, as the line flux computed by `specutils` would return
# units of Jy micron and it is hard to convert this into conventional units within the current `specutils` framework.
# Line flux units should be in units of in W/m^2. Implementing a simple way to convert the flux and associate error to
# other units when dealing with a 1d spectal object with "mixed" spectral x and y axis units seems necessary.
# +
# Compute the optical depth of the 10 micron feature
tau = -(np.log(line_spec.flux.value / line_y_continuum.value))
optdepth_spec = Spectrum1D(spectral_axis = line_spec.spectral_axis,
flux = tau*(u.Jy/u.Jy))
# -
# **Developer note** Trying to put optical depth into a Spectrum1D object results in an error as no units.
# But the optical depth is unit-less - using (u.Jy/u.Jy) as work arround.
# Plot the optical depth of the 10 micron region vs wavelength
plt.figure(figsize = (10,6))
plt.plot(optdepth_spec.spectral_axis, optdepth_spec.flux)
plt.xlabel("Wavelength ({:latex})".format(spec.spectral_axis.unit))
plt.ylabel('Tau')
plt.tight_layout()
plt.show()
plt.close()
# **Note** At this point repeat *all* the steps above to isolate solid-state features e.g. for the forsterite feature at
# at approx 13.3 microns.
# #### Now try looking for low crystalline silicate features at 23, 28, 33 microns in the spectra.
# +
bbsub_spectra = spec - ybest # photosphere continuum subtracted spectra
spline_points = [20.0, 21.3, 22.0, 24.4, 25.5, 33.8, 35.9] * u.micron
fluxc_resample = SplineInterpolatedResampler()
# Generate a spline fit to the dust continuum
spline_spec = fluxc_resample(bbsub_spectra, spline_points)
# +
# Plot the underlying dust continuum and spline fit
plt.figure(figsize = (8,4))
plt.plot(bbsub_spectra.spectral_axis, bbsub_spectra.flux.value, label = 'Dust spectra')
plt.plot(spline_spec.spectral_axis, spline_spec.flux.value, label = 'Spline spectra')
plt.axhline(0, color='r', linestyle='dashdot', alpha=0.5)
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("Continuum-subtracted spectrum with spline")
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
# Plot the underlying dust continuum and spline fit
plt.figure(figsize = (8,4))
plt.plot(bbsub_spectra.spectral_axis, bbsub_spectra.flux.value, label = 'Dust spectra')
plt.plot(spline_spec.spectral_axis, spline_spec.flux.value, label = 'Spline spectra')
plt.xlim(spline_points[0].value, spline_points[-1].value)
plt.xlabel('Wavelength (microns)')
plt.ylabel("Flux ({:latex})".format(spec.flux.unit))
plt.title("Zoom of continuum-subtracted spectrum with spline")
plt.legend(frameon = False, fontsize = 'medium')
plt.tight_layout()
plt.show()
plt.close()
# -
# **Developer note** By fitting a spline to a sub region the spectral shapes are no longer the same.
# ` bbsub_spectra.flux.value - spline_spec.flux.value` now brakes. Would need to trim the spectrum to the spline size to
# start looking closely for low contrast dust features and again measure their properties (see above). Some wrapper to
# stop repeating the same steps over and over would be nice.
# ## Additional Resources
#
# - [PampelMuse](https://gitlab.gwdg.de/skamann/pampelmuse)
# - [CASA](https://casa.nrao.edu/Release3.4.0/docs/UserMan/UserManse41.html)
# + [markdown] slideshow={"slide_type": "slide"}
# ## About this notebook
# **Author:** <NAME>, Project Scientist, UK ATC.
# **Updated On:** 2020-08-11
# -
# ***
# [Top of Page](#top)
|
_notebooks/MRS_Mstar_analysis/JWST_Mstar_dataAnalysis_usecase.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python3
# ---
# <h1> Logistic Regression <h1>
# <h2> ROMÂNĂ <h2>
# <blockquote><p>În final, o să observăm dacă Google PlayStore a avut destule date pentru a putea prezice popularitatea unei aplicații de trading sau pentru topul jocurilor plătite.
# Lucrul acesta se va face prin împărțirea descărcărilor în 2 variabile dummy. Cu mai mult de 1.000.000 pentru variabila 1 și cu mai puțin de 1.000.000 pentru variabila 0, pentru
# aplicațiile de Trading și pentru jocurile plătite cu mai mult de 670.545 de descărcari pentru variabila 1 iar 0 corespunde celorlalte aplicații.</p></blockquote>
# <h2>ENGLISH<h2>
#
# <blockquote><p>Lastly, we shall see if Google PlayStore had enough data in order to predict the popularity of a trading app or for the top paid games of the store .
# This will be done by dividing the downloads into 2 dummy variables. With more than 1,000,000 for variable 1 and less than 1,000,000 for variable 0, for Trading applications and for paid games with more than 670,545 downloads for variable 1 and 0 corresponding to the other applications.</p></blockquote>
#
# <h3>Now we shall create a logistic regression model using a 80/20 ratio between the training sample and the testing sample<h3>
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
def Log_reg(x,y):
model = LogisticRegression(solver='liblinear',C=10, random_state=0).fit(x,y)
print("Model accuracy",model.score(x,y))
cm = confusion_matrix(y, model.predict(x))
fig, ax = plt.subplots(figsize=(8, 8))
ax.imshow(cm)
ax.grid(False)
ax.xaxis.set(ticks=(0, 1), ticklabels=('Predicted 0s', 'Predicted 1s'))
ax.yaxis.set(ticks=(0, 1), ticklabels=('Actual 0s', 'Actual 1s'))
ax.set_ylim(1.5, -0.5)
for i in range(2):
for j in range(2):
ax.text(j, i, cm[i, j], ha='center', va='center', color='black')
plt.title('Confusion Matrix')
plt.show()
print(classification_report(y, model.predict(x)))
scores = cross_val_score(model, x,y, cv=10)
print('Cross-Validation Accuracy Scores', scores)
scores = pd.Series(scores)
print("Mean Accuracy: ",scores.mean())
# +
import pandas as pd
import numpy as np
#path = "D:\Java\VS-CodPitonul\\GAME.xlsx"
#df = pd.read_excel (path, sheet_name='Sheet1')
path = "D:\Java\VS-CodPitonul\\Trading_Apps.xlsx"
df = pd.read_excel (path, sheet_name='Results')
'''
RO: Folosește dropna daca ai valori lipsă, altfel îți va da eroare
ENG: Use dropna only if you have missing values else you will recive an error message
'''
#df = df.dropna()
# +
#Log_reg(df[['Score','Ratings','Reviews','Months_From_Release','Price']],df['Instalari_Bin']) #For GAME.xlsx
Log_reg(df[['Score','Ratings','Reviews','Months_From_Release']],df['Instalari_Bin']) #For Trading_Apps.xlsx
# -
|
Code/LogisticRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import division
import numpy as np
from numpy import linalg as LA
#np.seterr(divide='ignore') # these warnings are usually harmless for this code
from matplotlib import pyplot as plt
import matplotlib
# %matplotlib inline
import os
import scipy.stats as stats
import pyhsmm
from pyhsmm.util.text import progprint_xrange
import pyhsmm.basic.distributions as distributions
import scipy.io as sio
import csv
import copy
import time
import pickle
from sqlalchemy.orm import sessionmaker
from sqlalchemy import Table, MetaData, Column, Integer, String
from sqlalchemy import create_engine
from sqlalchemy.ext.declarative import declarative_base
from sklearn import preprocessing
filename = 'data_devices_trip.sav'
data_devices_trip = pickle.load(open(filename, 'rb'))
#EEFECTS: return new data in form: data = {} and data[device]={"trip":[]}
def dataTransform(data_devices):
data = {}
for i, devi in enumerate(data_devices):
#print(i, devi)
data[devi] = {}
for ii in range(data_devices[devi].shape[0]):
data_temp = data_devices[devi][ii]
trip = int(data_temp[0])
speed = data_temp[1]
acc = data_temp[2]
try:
data[devi][trip].append([speed,acc])
except:
data[devi][trip] = []
data[devi][trip].append([speed,acc])
return data
# get data_devices_trip = {} and data_devices_trip[device]={"trip":[]}
filename = 'data_devices.sav'
data_devices = pickle.load(open(filename, 'rb'))
data_devices_trip = dataTransform(data_devices)
#another way to get data_devices_trip, but this way is a little bit slow
#filename = 'data_devices_trip.sav'
#data_devices_trip = pickle.load(open(filename, 'rb'))
# +
posteriormodels = {}
i = 0
for devi, value1 in data_devices_trip.items() :
#for i, devi in enumerate(data_devices):
print('devi', devi)
if(len(data_devices_trip[devi]) == 0):
print('oops, this is a none set')
continue
else:
posteriormodels[devi]={}
for trip,value2 in data_devices_trip[devi].items():
print('trip',trip)
data_trip = np.array(data_devices_trip[devi][trip])
data_scaled = preprocessing.scale(data_trip)#implement data normalization
Nmax = 200 # preset the maximum states
# and some hyperparameters
obs_dim = data_scaled.shape[1] # data dimensions
obs_hypparams = {'mu_0':np.zeros(np.int(obs_dim)),
'sigma_0':np.eye(np.int(obs_dim)),
'kappa_0':0.25, # 0.2 5
'nu_0':obs_dim+2}
# Define the observation distribution
obs_distns = [pyhsmm.distributions.Gaussian(**obs_hypparams) for state in range(Nmax)]
# Define the posterior inference model
posteriormodels[devi][trip] = pyhsmm.models.WeakLimitStickyHDPHMM(
kappa=6.,alpha=1.,gamma=1.,init_state_concentration=1.,
obs_distns=obs_distns)
# Sampling process, for 100 round
Sampling_step = 100
Sampling_xaxis = range(1,Sampling_step+1)
# Add the data to the model and train
posteriormodels[devi][trip].add_data(data_scaled)
Meth2_LLH = np.zeros((Sampling_step,1))
# Sampling process, for 100 around
for idx in progprint_xrange(Sampling_step):
posteriormodels[devi][trip].resample_model()
#Meth2_LLH[idx] = posteriormodel.log_likelihood()
i = i + 1
if i == 6:
break
# save the model to disk
filename = 'posterior_models_test.sav'
pickle.dump(posteriormodels, open(filename, 'wb'))
# -
posteriormodels = {}
i = 0
for devi, value1 in data_devices_trip.items() :
#for i, devi in enumerate(data_devices):
print('devi', devi)
if(len(data_devices_trip[devi]) == 0):
print('oops, this is a none set')
continue
else:
posteriormodels[devi]={}
i = i + 1
if i == 6:
break
|
2. Posterior_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from kinase_binding.learning.data_analysis import biolab_indexing, get_distance_matrix, get_morgan_fingerprints
from kinase_binding.learning.splts import BiolabSplitter, create_test_set_from_folds
from rdkit import Chem
import dill
import pandas as pd
# -
# # Split
base_path = '../data/p38'
data_fpath = base_path+'/data.csv'
df=pd.read_csv(data_fpath)
if 'Unnamed: 0' in df.columns:
df = df.drop(columns=['Unnamed: 0'])
df=biolab_indexing(df, 'rdkit')
df.to_csv(data_fpath)
dm=get_distance_matrix(get_morgan_fingerprints(df['rdkit']))
# +
spl = BiolabSplitter(dm, 7)
folds = spl.create_folds()
train_val_folds, train_test = create_test_set_from_folds(folds, 0)
# +
with open(base_path+'/train_val_folds.pkl', "wb") as out_f:
dill.dump(train_val_folds, out_f)
with open(base_path+'/train_test_folds.pkl', "wb") as out_f:
dill.dump(train_test, out_f)
|
learning/.ipynb_checkpoints/spitting-checkpoint.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Topic Modeling
# Eyeballing parliamentary minutes for two election periods using topic modelling is performed in this notebook. The initial creation of n-grams takes long (~1h.
# - Language model from spaCy
# - n-grams models are large in size and quick to compute given n-gram prepared text
# - Corpus and LDA with gensim
# - Visualization with pyLDAvis
# %% [markdown]
# ## Setup
# %%
VERBOSE = False
# %%
import codecs
from pathlib import Path
import _pickle as pickle
import pyLDAvis
import pyLDAvis.gensim_models
import spacy
from gensim.corpora import Dictionary, MmCorpus
from gensim.models import Phrases
from gensim.models.ldamulticore import LdaMulticore
from gensim.models.word2vec import LineSentence
# %%
# Create subfolder 'tm' and 'out' in 'data'
Path("data/tm").mkdir(parents=True, exist_ok=True)
Path("data/out").mkdir(parents=True, exist_ok=True)
# %% [markdown] tags=[]
# #### Configuration
# - Models are large in size and quick to compute -> save only when re-running the notebook frequently
# %%
SAVE_MODELS = True
# language_model = "de_dep_news_trf"
language_model = "de_core_news_lg"
gerNLP = spacy.load(language_model)
speeches_txt_filepath = "data/plpr_alltext.txt"
# %% [markdown]
# ### Helper Functions
# The helper functions help with text preprocessing, the creation of n-grams, and to show results. The creation of the lemmatized sentence corpuse is memory intense. Reduce batch size and the number of parallel processes if needed.
# %%
def preview(filepath, N=1000):
"""Previews N characters of file."""
with open(filepath) as temp:
head = next(temp)
head = head[:N]
temp.close()
return head
def preview_lines(filepath, N=5):
"""Previews N lines of file."""
with open(filepath) as temp:
head = [next(temp) for i in range(N)]
temp.close()
return head
def punct_space(token):
"""Removes punctuation and whitespace."""
return token.is_punct or token.is_space
def line_speech(filename):
"""Reads lines and ignores lines breaks"""
with codecs.open(filename, encoding="utf_8") as f:
for speech in f:
yield speech.replace("\\n", "\n")
def lemmatized_sentence_corpus_to_file(input_file, output_file):
"""Parses speeches with spaCy, writes lemmatized sentences to file."""
with codecs.open(output_file, "w", encoding="utf_8") as f:
for parsed_speech in gerNLP.pipe(
line_speech(input_file), batch_size=100, n_process=8
):
for sent in parsed_speech.sents:
parsed_sent = " ".join(
[token.lemma_ for token in sent if not punct_space(token)]
)
f.write(parsed_sent + "\n")
# %% [markdown]
# ## Unigrams
# The text file is a text-only extract of the speeches of the plenary protocols dataframe in `nb_02`. To create unigrams, all text is cleaned by stripping junk such as stop words or meaningless filter words, conjugated words are reversed to their base form.
# %%
preview(speeches_txt_filepath)
# %%
# %%time
# long running
unigram_sentences_filepath = "data/tm/unigram_sent_all.txt"
if Path(unigram_sentences_filepath).exists():
print(f"Unigram sentences available at {unigram_sentences_filepath}")
else:
print(f"Unigram sentences not available. Now creating {unigram_sentences_filepath}")
lemmatized_sentence_corpus_to_file(
input_file=speeches_txt_filepath, output_file=unigram_sentences_filepath
)
unigram_sentences = LineSentence(unigram_sentences_filepath)
# preview_lines(unigram_sentences_filepath)
# %% [markdown]
# ## Bigrams
# Bigrams (or any larger structure of n-grams) represent word pairs (or triplets, quadruples, etc.) of words commonly appearing together. "Renewable" and "energy" used independetly do not convey the same meaning as "renewable_energy".
# %%
# %%time
bigram_model_filepath = "data/tm/bigram_model_all"
if Path(bigram_model_filepath).exists():
print(f"Bigram model available at {bigram_model_filepath}")
bigram_model = Phrases.load(bigram_model_filepath)
else:
print(f"Bigram model not available. Now creating {bigram_model_filepath}")
bigram_model = Phrases(unigram_sentences)
if SAVE_MODELS:
bigram_model.save(bigram_model_filepath)
# %%
bigram_sentences_filepath = "data/tm/bigram_sent_all.txt"
if Path(bigram_sentences_filepath).exists():
print(f"Bigram sentences available at {bigram_sentences_filepath}")
else:
print(f"Bigram sentences not available. Now creating {bigram_sentences_filepath}")
with codecs.open(bigram_sentences_filepath, "w", encoding="utf_8") as f:
for unigram_sentence in unigram_sentences:
bigram_sentence = " ".join(bigram_model[unigram_sentence])
f.write(bigram_sentence + "\n")
bigram_sentences = LineSentence(bigram_sentences_filepath)
preview_lines(bigram_sentences_filepath)
# %% [markdown]
# ## Trigrams
# As bigrams are used to create trigrams, there is the chance of two bigrams being combined which would be a 4-gram.
# %%
# %%time
trigram_model_filepath = "data/tm/trigram_model_all"
if Path(trigram_model_filepath).exists():
print(f"Trigram model available at {trigram_model_filepath}")
trigram_model = Phrases.load(trigram_model_filepath)
else:
print(f"Trigram model not available. Now creating {trigram_model_filepath}")
trigram_model = Phrases(bigram_sentences)
if SAVE_MODELS:
trigram_model.save(trigram_model_filepath)
# %%
# short running
trigram_sentences_filepath = "data/tm/trigram_sent_all.txt"
if Path(trigram_sentences_filepath).exists():
print(f"Trigram sentences available at {trigram_sentences_filepath}")
else:
print(f"Trigram sentences not available. Now creating {trigram_sentences_filepath}")
with codecs.open(trigram_sentences_filepath, "w", encoding="utf_8") as f:
for bigram_sentence in bigram_sentences:
trigram_sentence = " ".join(trigram_model[bigram_sentence])
f.write(trigram_sentence + "\n")
trigram_sentences = LineSentence(trigram_sentences_filepath)
preview_lines(trigram_sentences_filepath)
# %%
# %%time
# long running
trigram_speeches_filepath = "data/tm/trigram_transformed_speeches_all.txt"
if Path(trigram_speeches_filepath).exists():
print(f"Trigram speeches available at {trigram_speeches_filepath}")
else:
print(f"Trigram speeches not available. Now creating {trigram_speeches_filepath}")
with codecs.open(trigram_speeches_filepath, "w", encoding="utf_8") as f:
for parsed_speech in gerNLP.pipe(
line_speech(speeches_txt_filepath), batch_size=100, n_process=15
):
# lemmatize the text, removing punctuation and whitespace
unigram_speech = [
token.lemma_ for token in parsed_speech if not punct_space(token)
]
# apply the first-order and second-order phrase models
bigram_speech = bigram_model[unigram_speech]
trigram_speech = trigram_model[bigram_speech]
# remove any remaining stopwords
trigram_speech = [
term
for term in trigram_speech
if term.lower() not in spacy.lang.de.STOP_WORDS
]
# stop words found here: https://github.com/explosion/spaCy/blob/master/spacy/lang/de/stop_words.py
# write the transformed speech as a line in the new file
trigram_speech = " ".join(trigram_speech)
f.write(trigram_speech + "\n")
preview_lines(trigram_speeches_filepath, N=2)
# %%
print(f"""
File {trigram_speeches_filepath} contains {sum(1 for line in open(trigram_speeches_filepath))} documents
"""
)
# %% [markdown]
# ## Latent Dirichlet Allocation
# In this section, the text is transformed into a corpus, which is the collection of documents over which topics are discovered using LDA. As a first intermediate step, the speech documents are represented with a dictionary, where n-grams are keys and occurances counts within speech documents are the respective values.
#
# The parameters `THRES_BELOW` AND `THRES_ABOVE` define, which keywords can define a topic. `THRES_BELOW` is the minimum number of documents, in which a keyword needs to occur to be able to define a topic. `THRES_ABOVE` is a relative value, it defines the maximum fraction of documents, which may contain a keyword for the keyword to be able to define topics. Accordingly, keywords which are too common, cannot define a topic, and special terminology of a single speech does not, either.
# %%
THRESH_BELOW = 10
THRESH_ABOVE = 0.05
thres_suffix = f"TB{str(THRESH_BELOW)}_TA{str(THRESH_ABOVE)}".replace(".", "")
trigram_dictionary_filepath = f"data/tm/trigram_dict_all_{thres_suffix}.dict"
if Path(trigram_dictionary_filepath).exists():
print(f"Trigram dictionary available at {trigram_dictionary_filepath}")
trigram_dictionary = Dictionary.load(trigram_dictionary_filepath)
else:
print(
f"Trigram dictionary not available. Now creating {trigram_dictionary_filepath}"
)
trigram_speeches = LineSentence(trigram_speeches_filepath)
trigram_dictionary = Dictionary(trigram_speeches)
# filter tokens that are very rare or too common from
# the dictionary (filter_extremes) and reassign integer ids (compactify)
trigram_dictionary.filter_extremes(no_below=THRESH_BELOW, no_above=THRESH_ABOVE)
trigram_dictionary.compactify()
trigram_dictionary.save(trigram_dictionary_filepath)
# %%
def trigram_bow_generator(filepath):
"""
generator function to read speeches from a file
and yield a bag-of-words representation
"""
for speech in LineSentence(filepath):
yield trigram_dictionary.doc2bow(speech)
# %%
trigram_bow_filepath = f"data/tm/trigram_bow_corpus_all_{thres_suffix}.mm"
if Path(trigram_bow_filepath).exists():
print(f"Trigram bag-of-words available at {trigram_bow_filepath}")
else:
print(f"Trigram bag-of-words not available. Now creating {trigram_bow_filepath}")
# generate bag-of-words representations for
# all speeches and save them as a matrix
MmCorpus.serialize(
trigram_bow_filepath, trigram_bow_generator(trigram_speeches_filepath)
)
# load the finished bag-of-words corpus from disk
trigram_bow_corpus = MmCorpus(trigram_bow_filepath)
# %% [markdown]
# ## Topic Models & Visuals
# Latent topics are finally within the corpus are finally derived. "Latent" means that topic belongingness may not be obvious at first sight for a document. The output is by no means finite and requires manual review and validation.
# %%
# %%time
# medium-long running
topics = [50, 250, 500]
for number_of_topics in topics:
# topic model
lda_model_filepath = (
f"data/tm/lda_model_{thres_suffix}_{str(number_of_topics)}"
)
if Path(lda_model_filepath).exists():
print(f"Trigram bag-of-words available at {lda_model_filepath}")
# load the finished LDA model from disk
lda = LdaMulticore.load(lda_model_filepath)
else:
print(f"Trigram bag-of-words not available. Now creating {lda_model_filepath}")
lda = LdaMulticore(
trigram_bow_corpus,
num_topics=number_of_topics,
id2word=trigram_dictionary,
workers=8,
)
lda.save(lda_model_filepath)
# topic model visual
LDAvis_data_filepath = (
f"data/tm/ldavis_prepared_{thres_suffix}_{str(number_of_topics)}"
)
if Path(LDAvis_data_filepath).exists():
print(f"LDA Visualization available at {LDAvis_data_filepath}")
with open(LDAvis_data_filepath, "rb") as f:
LDAvis_prepared = pickle.load(f)
else:
print(f"LDA visualization not available. Now creating {LDAvis_data_filepath}")
LDAvis_prepared = pyLDAvis.gensim_models.prepare(
lda, trigram_bow_corpus, trigram_dictionary
)
with open(LDAvis_data_filepath, "wb") as f:
pickle.dump(LDAvis_prepared, f)
# topic model html visual
LDAvis_html_filepath = (
f"data/out/lda_viz_{thres_suffix}_{str(number_of_topics)}.html"
)
if Path(LDAvis_html_filepath).exists():
print(f"LDA Visualization available at {LDAvis_html_filepath}")
else:
print(
f"LDA html visualization not available. Now creating {LDAvis_html_filepath}"
)
pyLDAvis.save_html(LDAvis_prepared, LDAvis_html_filepath)
# %% [markdown]
# ## Single Model Review
# %%
DEFAULT_NO_TOPICS = 250
DEFAULT_THRESHS = "TB{}_TA{}".format(str(THRESH_BELOW), str(THRESH_ABOVE).replace(".",""))
lda_model_filepath = (
f"data/tm/lda_model_{DEFAULT_THRESHS}_{str(DEFAULT_NO_TOPICS)}"
)
# load the finished LDA model from disk
lda = LdaMulticore.load(lda_model_filepath)
LDAvis_data_filepath = (
f"data/tm/ldavis_prepared_{thres_suffix}_{str(DEFAULT_NO_TOPICS)}"
)
with open(LDAvis_data_filepath, "rb") as f:
LDAvis_prepared = pickle.load(f)
print(f"Model in use: {LDAvis_data_filepath}")
# %% [markdown]
# In verbose mode, the next cell displays a visualization of topic models, disabling the Jupyter menu bar. Delete the output to see the menu bar again.
# %%
if VERBOSE:
pyLDAvis.display(LDAvis_prepared)
# %% [markdown]
# View the notebook [here](https://nbviewer.jupyter.org/github/sebas-seck/bundestag_nlp/blob/main/nb_03_topic_modelling.ipynb#topic=0&lambda=1&term=) with Jupyter's nbviewer as the interactive visualizations are not rendered with the static display of notebooks on Github. Alternatively, paste the link to the notebook on Github [here](https://nbviewer.jupyter.org/).
#
# The definition of the number of latent topics to uncover has no set definition. Given the unsupervised nature of Topic modeling, I expect numbers of varying magnitude to result in differing broadness of topics.
# %%
def lda_description(review_text, min_topic_freq=0.05):
"""
accept the original text of a review and (1) parse it with spaCy,
(2) apply text pre-proccessing steps, (3) create a bag-of-words
representation, (4) create an LDA representation, and
(5) print a sorted list of the top topics in the LDA representation
"""
# parse the review text with spaCy
parsed_review = gerNLP(review_text)
# lemmatize the text and remove punctuation and whitespace
unigram_review = [token.lemma_ for token in parsed_review if not punct_space(token)]
# apply the first-order and secord-order phrase models
bigram_review = bigram_model[unigram_review]
trigram_review = trigram_model[bigram_review]
# remove any remaining stopwords
trigram_review = [
term for term in trigram_review if not term in spacy.lang.de.STOP_WORDS
]
# create a bag-of-words representation
review_bow = trigram_dictionary.doc2bow(trigram_review)
# create an LDA representation
review_lda = lda[review_bow]
# sort with the most highly related topics first
review_lda = sorted(review_lda, key=lambda topic_number_freq: -topic_number_freq[1])
for topic_number, freq in review_lda:
if freq < min_topic_freq:
break
# print the most highly related topic names and frequencies
print("{:25} {}".format(topic_number, round(freq, 3)))
# %% [markdown]
# ## Speech Review
# %%
review_text1 = """<NAME>, möglicherweise ist das ein Anlass, um über andere Strukturen nachzudenken. Im Land Brandenburg, aus dem ich komme,
gibt es im Süden einen Bestand von 60 000 Schweinen an einem Standort. Stellen wir uns vor, dass dieser Standort wegen der Afrikanischen
Schweinepest auf einmal in einer Restriktionszone liegt. Dann werden wir wahrscheinlich nicht umhinkommen, den gesamten Bestand zu töten.
Ist es nicht an der Zeit, einmal ernsthaft darüber nachzudenken, ob solche Megaställe nicht der Vergangenheit angehören sollten und ob unter
Aspekten der Tierseuchenbekämpfung nicht Regionen mit sehr dichtem Tierbestand als auch solche riesengroßen Bestände vermieden werden sollten?
Das ist einfach sehr schwierig in einer Tierseuchensituation zu händeln. Ich glaube zudem, dass die in Rede stehenden Maßnahmen ethisch nicht
mehr vertretbar sind. Deswegen lautet meine Frage: Müssen wir nicht auch über Strukturen bei den Tierbeständen nachdenken?"""
review_text2 = """Vielen Dank, <NAME>. – <NAME>, gestern fand eine informelle Tagung der Entwicklungsminister der
Europäischen Union statt. Auf der Tagesordnung stand unter anderem der mehrjährige Finanzrahmen der Europäischen Union. Die Kommission bereitet
die Debatte vor. Das Europäische Parlament wie auch der Ministerrat in allen seinen Formationen wird sich zu der Frage positionieren müssen,
wie der Haushalt der Europäischen Union im Zeitrahmen des nächsten mehrjährigen Finanzrahmens aufzustellen ist. In diesem Zusammenhang hat
Entwicklungsminister Dr. <NAME> dazu aufgerufen, die internationalen Aufgaben der Europäischen Union, insbesondere mit Blick auf Afrika,
deutlich zu stärken."""
review_text3 = """Welche konkreten rechtlichen Überlegungen haben die Ostbeauftragte <NAME> und das Bundeswirtschaftsministerium dazu
veranlasst, für eine Studie des Göttinger Instituts für Demokratieforschung zum Thema 'Rechtsextremismus und Fremdenfeindlichkeit in
Ostdeutschland', die nach eigenen Angaben von <NAME> selbst nach Nacherfüllungsmöglichkeit eine 'schlicht nicht hinnehmbare Schlamperei'
darstellt, von der sie sich öffentlich distanziert hat und die für sie 'jeden Wert ... verloren' hatte, nicht nur die Rückforderung von
bereits ausgezahlten Geldern zu unterlassen, sondern auch noch zu einem Zeitpunkt, als die Unbrauchbarkeit der Studie bereits bekannt war,
einen bis dahin noch nicht ausgezahlten Betrag hierfür zu zahlen, wie unter anderem die Zeitung 'Die Welt' am 12. Februar 2018 berichtet hat,
und wie hoch war der Betrag, der erst nach Bekanntwerden der Mangelhaftigkeit der Studie an das Göttinger Institut für Demokratieforschung bzw.
die Georg-August-Universität Göttingen ausgezahlt wurde?"""
# %%
lda_description(review_text1)
# %%
lda_description(review_text2)
# %%
lda_description(review_text3)
# %% [markdown]
# ## Energy Politics Topics
# %% [markdown]
# Which keywords make up the energy politics topic? Looking at a few speeches from three plenary debates, topics can be identified which can be reviewed manually for further, relevant terms to handcraft topic models later.
#
# - [17/96](https://dip21.bundestag.de/dip21/btp/17/17096.pdf): March 17th 2011, second meeting after the catastrophe, first technical debates about nuclear power
# - [17/117](https://dip21.bundestag.de/dip21/btp/17/17117.pdf): Discussions on changing the Atomic Energy Act
# - [17/229](https://dip21.bundestag.de/dip21/btp/17/17229.pdf): March 15th 2013, shortly after the second anniversary of the catastrophe
# %%
speeches = {
11636: "<NAME>",
14197: "<NAME>",
14198: "<NAME>",
14199: "<NAME>",
14200: "<NAME>",
29594: "<NAME>",
29595: "<NAME>",
29596: "<NAME>",
}
# %%
with open(speeches_txt_filepath) as f:
d = f.readlines()
# %%
for k, v in speeches.items():
print(v)
text = d[k-1]
print(lda_description(text))
# %%
|
nb_03_topic_modelling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#export
from local.test import *
from local.basics import *
from local.vision.core import *
from local.vision.data import *
from local.vision.augment import *
from local.vision import models
# +
#default_exp vision.learner
# -
from local.notebook.showdoc import *
# # Learner for the vision applications
#
# > All the functions necessary to build `Learner` suitable for transfer learning in computer vision
# ## Cut a pretrained model
# export
def _is_pool_type(l): return re.search(r'Pool[123]d$', l.__class__.__name__)
m = nn.Sequential(nn.AdaptiveAvgPool2d(5), nn.Linear(2,3), nn.Conv2d(2,3,1), nn.MaxPool3d(5))
test_eq([bool(_is_pool_type(m_)) for m_ in m.children()], [True,False,False,True])
# export
def has_pool_type(m):
"Return `True` if `m` is a pooling layer or has one in its children"
if _is_pool_type(m): return True
for l in m.children():
if has_pool_type(l): return True
return False
m = nn.Sequential(nn.AdaptiveAvgPool2d(5), nn.Linear(2,3), nn.Conv2d(2,3,1), nn.MaxPool3d(5))
assert has_pool_type(m)
test_eq([has_pool_type(m_) for m_ in m.children()], [True,False,False,True])
#export
def create_body(arch, pretrained=True, cut=None):
"Cut off the body of a typically pretrained `arch` as determined by `cut`"
model = arch(pretrained=pretrained)
#cut = ifnone(cut, cnn_config(arch)['cut'])
if cut is None:
ll = list(enumerate(model.children()))
cut = next(i for i,o in reversed(ll) if has_pool_type(o))
if isinstance(cut, int): return nn.Sequential(*list(model.children())[:cut])
elif callable(cut): return cut(model)
else: raise NamedError("cut must be either integer or a function")
# `cut` can either be an integer, in which case we cut the model at the coresponding layer, or a function, in which case, this funciton returns `cut(model)`. It defaults to `cnn_config(arch)['cut']` if `arch` is in `cnn_config`, otherwise to the first layer that contains some pooling.
# +
tst = lambda pretrained : nn.Sequential(nn.Conv2d(4,5,3), nn.BatchNorm2d(5), nn.AvgPool2d(1), nn.Linear(3,4))
m = create_body(tst)
test_eq(len(m), 2)
m = create_body(tst, cut=3)
test_eq(len(m), 3)
m = create_body(tst, cut=noop)
test_eq(len(m), 4)
# -
# ## Head and model
#export
def create_head(nf, nc, lin_ftrs=None, ps=0.5, concat_pool=True, bn_final=False, lin_first=False):
"Model head that takes `nf` features, runs through `lin_ftrs`, and out `nc` classes."
lin_ftrs = [nf, 512, nc] if lin_ftrs is None else [nf] + lin_ftrs + [nc]
ps = L(ps)
if len(ps) == 1: ps = [ps[0]/2] * (len(lin_ftrs)-2) + ps
actns = [nn.ReLU(inplace=True)] * (len(lin_ftrs)-2) + [None]
pool = AdaptiveConcatPool2d() if concat_pool else nn.AdaptiveAvgPool2d(1)
layers = [pool, Flatten()]
if lin_first: layers.append(nn.Dropout(ps.pop(0)))
for ni,no,p,actn in zip(lin_ftrs[:-1], lin_ftrs[1:], ps, actns):
layers += LinBnDrop(ni, no, bn=True, p=p, act=actn, lin_first=lin_first)
if lin_first: layers.append(nn.Linear(lin_ftrs[-2], nc))
if bn_final: layers.append(nn.BatchNorm1d(lin_ftrs[-1], momentum=0.01))
return nn.Sequential(*layers)
tst = create_head(5, 10)
tst
# +
#hide
mods = list(tst.children())
test_eq(len(mods), 9)
assert isinstance(mods[2], nn.BatchNorm1d)
assert isinstance(mods[-1], nn.Linear)
tst = create_head(5, 10, lin_first=True)
mods = list(tst.children())
test_eq(len(mods), 8)
assert isinstance(mods[2], nn.Dropout)
# -
#export
from local.callback.hook import num_features_model
#export
def create_cnn_model(arch, nc, cut, pretrained, lin_ftrs=None, ps=0.5, custom_head=None,
bn_final=False, concat_pool=True, init=nn.init.kaiming_normal_):
"Create custom convnet architecture using `base_arch`"
body = create_body(arch, pretrained, cut)
if custom_head is None:
nf = num_features_model(nn.Sequential(*body.children())) * (2 if concat_pool else 1)
head = create_head(nf, nc, lin_ftrs, ps=ps, concat_pool=concat_pool, bn_final=bn_final)
else: head = custom_head
model = nn.Sequential(body, head)
if init is not None: apply_init(model[1], init)
return model
tst = create_cnn_model(models.resnet18, 10, None, True)
#export
@delegates(create_cnn_model)
def cnn_config(**kwargs):
"Convenienc function to easily create a config for `create_cnn_model`"
return kwargs
# +
pets = DataBlock(blocks=(ImageBlock, CategoryBlock),
get_items=get_image_files,
splitter=RandomSplitter(),
get_y=RegexLabeller(pat = r'/([^/]+)_\d+.jpg$'))
dbunch = pets.databunch(untar_data(URLs.PETS)/"images", item_tfms=RandomResizedCrop(300, min_scale=0.5), bs=64,
batch_tfms=[*aug_transforms(size=224), Normalize(*imagenet_stats)])
# +
#TODO: refactor, i.e. something like this?
# class ModelSplitter():
# def __init__(self, idx): self.idx = idx
# def split(self, m): return L(m[:self.idx], m[self.idx:]).map(params)
# def __call__(self,): return {'cut':self.idx, 'split':self.split}
# -
#export
def default_split(m:nn.Module): return L(m[0], m[1:]).map(params)
# +
#export
def _xresnet_split(m): return L(m[0][:3], m[0][3:], m[1:]).map(params)
def _resnet_split(m): return L(m[0][:6], m[0][6:], m[1:]).map(params)
def _squeezenet_split(m:nn.Module): return L(m[0][0][:5], m[0][0][5:], m[1:]).map(params)
def _densenet_split(m:nn.Module): return L(m[0][0][:7],m[0][0][7:], m[1:]).map(params)
def _vgg_split(m:nn.Module): return L(m[0][0][:22], m[0][0][22:], m[1:]).map(params)
def _alexnet_split(m:nn.Module): return L(m[0][0][:6], m[0][0][6:], m[1:]).map(params)
_default_meta = {'cut':None, 'split':default_split}
_xresnet_meta = {'cut':-3, 'split':_xresnet_split }
_resnet_meta = {'cut':-2, 'split':_resnet_split }
_squeezenet_meta = {'cut':-1, 'split': _squeezenet_split}
_densenet_meta = {'cut':-1, 'split':_densenet_split}
_vgg_meta = {'cut':-2, 'split':_vgg_split}
_alexnet_meta = {'cut':-2, 'split':_alexnet_split}
# -
#export
model_meta = {
models.xresnet.xresnet18 :{**_xresnet_meta}, models.xresnet.xresnet34: {**_xresnet_meta},
models.xresnet.xresnet50 :{**_xresnet_meta}, models.xresnet.xresnet101:{**_xresnet_meta},
models.xresnet.xresnet152:{**_xresnet_meta},
models.resnet18 :{**_resnet_meta}, models.resnet34: {**_resnet_meta},
models.resnet50 :{**_resnet_meta}, models.resnet101:{**_resnet_meta},
models.resnet152:{**_resnet_meta},
models.squeezenet1_0:{**_squeezenet_meta},
models.squeezenet1_1:{**_squeezenet_meta},
models.densenet121:{**_densenet_meta}, models.densenet169:{**_densenet_meta},
models.densenet201:{**_densenet_meta}, models.densenet161:{**_densenet_meta},
models.vgg11_bn:{**_vgg_meta}, models.vgg13_bn:{**_vgg_meta}, models.vgg16_bn:{**_vgg_meta}, models.vgg19_bn:{**_vgg_meta},
models.alexnet:{**_alexnet_meta}}
# ## `Learner` convenience functions
#export
@delegates(Learner.__init__)
def cnn_learner(dbunch, arch, loss_func=None, pretrained=True, cut=None, splitter=None, config=None, **kwargs):
"Build a convnet style learner"
if config is None: config = {}
meta = model_meta.get(arch, _default_meta)
model = create_cnn_model(arch, get_c(dbunch), ifnone(cut, meta['cut']), pretrained, **config)
learn = Learner(dbunch, model, loss_func=loss_func, splitter=ifnone(splitter, meta['split']), **kwargs)
if pretrained: learn.freeze()
return learn
# The model is built from `arch` using the number of final activation inferred from `dbunch` by `get_c`. It might be `pretrained` and the architecture is cut and split using the default metadata of the model architecture (this can be customized by passing a `cut` or a `splitter`). To customize the model creation, use `cnn_config` and pass the result to the `config` argument.
learn = cnn_learner(dbunch, models.resnet34, loss_func=CrossEntropyLossFlat(), config=cnn_config(ps=0.25))
#export
@delegates(models.unet.DynamicUnet.__init__)
def unet_config(**kwargs):
"Convenience function to easily create a config for `DynamicUnet`"
return kwargs
#export
@delegates(Learner.__init__)
def unet_learner(dbunch, arch, loss_func=None, pretrained=True, cut=None, splitter=None, config=None, **kwargs):
"Build a unet learner from `dbunch` and `arch`"
if config is None: config = {}
meta = model_meta.get(arch, _default_meta)
body = create_body(arch, pretrained, ifnone(cut, meta['cut']))
try: size = dbunch.train_ds[0][0].size
except: size = dbunch.one_batch()[0].shape[-2:]
model = models.unet.DynamicUnet(body, get_c(dbunch), size, **config)
learn = Learner(dbunch, model, loss_func=loss_func, splitter=ifnone(splitter, meta['split']), **kwargs)
if pretrained: learn.freeze()
return learn
# +
camvid = DataBlock(blocks=(ImageBlock, MaskBlock),
get_items=get_image_files,
splitter=RandomSplitter(),
get_y=lambda o: untar_data(URLs.CAMVID_TINY)/'labels'/f'{o.stem}_P{o.suffix}')
dbunch = camvid.databunch(untar_data(URLs.CAMVID_TINY)/"images", batch_tfms=aug_transforms())
dbunch.show_batch(max_n=9, vmin=1, vmax=30)
# -
#TODO: Find a way to pass the classes properly
dbunch.vocab = np.loadtxt(untar_data(URLs.CAMVID_TINY)/'codes.txt', dtype=str)
learn = unet_learner(dbunch, models.resnet34, loss_func=CrossEntropyLossFlat(axis=1), config=unet_config())
# ## Show functions
#export
@typedispatch
def show_results(x:TensorImage, y, samples, outs, ctxs=None, max_n=10, rows=None, cols=None, figsize=None, **kwargs):
if ctxs is None: ctxs = get_grid(min(len(samples), max_n), rows=rows, cols=cols, add_vert=1, figsize=figsize)
ctxs = show_results[object](x, y, samples, outs, ctxs=ctxs, max_n=max_n, **kwargs)
return ctxs
#export
@typedispatch
def show_results(x:TensorImage, y:TensorCategory, samples, outs, ctxs=None, max_n=10, rows=None, cols=None, figsize=None, **kwargs):
if ctxs is None: ctxs = get_grid(min(len(samples), max_n), rows=rows, cols=cols, add_vert=1, figsize=figsize)
for i in range(2):
ctxs = [b.show(ctx=c, **kwargs) for b,c,_ in zip(samples.itemgot(i),ctxs,range(max_n))]
ctxs = [r.show(ctx=c, color='green' if b==r else 'red', **kwargs)
for b,r,c,_ in zip(samples.itemgot(1),outs.itemgot(0),ctxs,range(max_n))]
return ctxs
#export
@typedispatch
def show_results(x:TensorImage, y:(TensorImageBase, TensorPoint, TensorBBox), samples, outs, ctxs=None, max_n=10, rows=None, cols=None, figsize=None, **kwargs):
if ctxs is None: ctxs = get_grid(min(len(samples), max_n), rows=rows, cols=cols, add_vert=1, figsize=figsize, double=True)
for i in range(2):
ctxs[::2] = [b.show(ctx=c, **kwargs) for b,c,_ in zip(samples.itemgot(i),ctxs[::2],range(max_n))]
for x in [samples,outs]:
ctxs[1::2] = [b.show(ctx=c, **kwargs) for b,c,_ in zip(x.itemgot(0),ctxs[1::2],range(max_n))]
return ctxs
#export
@typedispatch
def plot_top_losses(x: TensorImage, y:TensorCategory, samples, outs, raws, losses, rows=None, cols=None, figsize=None, **kwargs):
axs = get_grid(len(samples), rows=rows, cols=cols, add_vert=1, figsize=figsize, title='Prediction/Actual/Loss/Probability')
for ax,s,o,r,l in zip(axs, samples, outs, raws, losses):
s[0].show(ctx=ax, **kwargs)
ax.set_title(f'{o[0]}/{s[1]} / {l.item():.2f} / {r.max().item():.2f}')
#export
@typedispatch
def plot_top_losses(x: TensorImage, y:TensorMultiCategory, samples, outs, raws, losses, rows=None, cols=None, figsize=None, **kwargs):
axs = get_grid(len(samples), rows=rows, cols=cols, add_vert=1, figsize=figsize)
for i,(ax,s) in enumerate(zip(axs, samples)): s[0].show(ctx=ax, title=f'Image {i}', **kwargs)
rows = get_empty_df(len(samples))
outs = L(s[1:] + o + (Str(r), Float(l.item())) for s,o,r,l in zip(samples, outs, raws, losses))
for i,l in enumerate(["target", "predicted", "probabilities", "loss"]):
rows = [b.show(ctx=r, label=l, **kwargs) for b,r in zip(outs.itemgot(i),rows)]
display_df(pd.DataFrame(rows))
# ## Export -
#hide
from local.notebook.export import notebook2script
notebook2script(all_fs=True)
|
dev/21_vision_learner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="z_YqsRpmaMFF" colab_type="code" colab={}
#Description:This program detects breast cancer,based off of data.
# + id="28_gMgriawn5" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + id="zNSTE5K_bH6z" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 436} outputId="dce74010-f0c0-4f5d-de2b-8122261736cc"
from google.colab import files
uploaded=files.upload()
df=pd.read_csv("cancer.csv")
df.head(10)
# + id="Sm3_CSuNdG4b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c8331685-c76d-42a7-98dc-e03248f76bb4"
df.shape
# + id="iRrVxwyKe65E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 588} outputId="c21b9cae-e27b-43d4-8571-0f9185cf57f2"
df.isna().sum()
# + id="0vkhiaHDfdFd" colab_type="code" colab={}
df=df.dropna(axis=1)
# + id="N0MIsBR4fxi3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8b9a9b1a-2d1b-49b0-8f98-9224ded544a9"
df.shape
# + id="8Dvsvlygf3NT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="d9f58be5-b730-4190-8b8d-f7d1ce5dfaf0"
#count of the number of malignant (M) or benign (b) cells
df['diagnosis'].value_counts()
# + id="niqMm3RcgXDY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="7fe05d4c-9307-4562-c7a2-5ac895bdb7c4"
sns.countplot(df['diagnosis'],label='count')
# + id="dZ5Mwt3sglLl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 571} outputId="aae4dc59-14b1-48a8-e847-e2d7ec7b976e"
#all datatypes that need to be encoded
df.dtypes
# + id="TkcXJUjQhZeW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 454} outputId="33d5828b-9adf-4e3d-dee8-25521800b79a"
#encode categorical values
from sklearn.preprocessing import *
labelencoder_y=LabelEncoder()
labelencoder_y.fit_transform(df.iloc[:,1].values)
# + id="sEeT5CHOirNX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 218} outputId="4750bc9b-74ba-4368-bb37-621cf009396e"
df.iloc[:,1]=labelencoder_y.fit_transform(df.iloc[:,1].values)
df.iloc[:,1]
# + id="yznui4qCi9KL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 764} outputId="63dd13b0-2d60-4fbe-b025-03854d01d703"
sns.pairplot(df.iloc[:,1:6],hue='diagnosis')
# + id="Z74Q_3NHjqpu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="6a81cd67-2b3a-4f5b-d16e-83e4eab007cb"
df.head(5)
# + id="gKhbc8w1j56s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 410} outputId="0aad0cfe-dd47-4fce-93b7-19679142ecf0"
#colleration of column
y=df.iloc[:,1:12].corr()
y
# + id="5K7LcApLkuFj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 725} outputId="71695e98-e1bb-4d97-c860-2ed282b1ce32"
plt.figure(figsize=(10,10))
sns.heatmap(y,annot=True, fmt='.0%')
# + id="5ylKd_4PsaRi" colab_type="code" colab={}
#split the datset into independent(x) and dependent(Y)
X=df.iloc[:,2:31].values
Y=df.iloc[:,1].values
# + id="26b0JUpQQIAl" colab_type="code" colab={}
#75% training and 25% testing spliting
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test=train_test_split(X, Y, test_size = 0.25 , random_state=0)
# + id="u9pNyFu2V6yG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="17b2dc19-fcb6-452c-b767-12d5523df070"
sc=StandardScaler()
X_train=sc.fit_transform(X_train)
X_test=sc.fit_transform(X_test)
X_train
# + id="swTYD30kYbDX" colab_type="code" colab={}
def models(X_train, Y_train):
#logistic regression
from sklearn.linear_model import LogisticRegression
log=LogisticRegression(random_state=0)
log.fit(X_train,Y_train)
#decision Tree
from sklearn.tree import DecisionTreeClassifier
tree =DecisionTreeClassifier(criterion='entropy',random_state=0)
tree.fit(X_train,Y_train)
#random forest classifier
from sklearn.ensemble import RandomForestClassifier
forest=RandomForestClassifier(n_estimators =10,criterion ="entropy",random_state=0)
forest.fit(X_train,Y_train)
print("[0]Logistic Regression Training Accuracy:",log.score(X_train,Y_train))
print("[1]Decision Tree Classifier Training Accuracy",tree.score(X_train,Y_train))
print("[2]Random Forest Classifier Training Accuracy:",forest.score(X_train,Y_train))
return log,tree,forest
# + id="bJO7abGofS0-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="a9e281b3-da13-4d14-89c6-89f91cc14f1a"
model= models(X_train,Y_train)
# + id="D0U9zBeOfqKp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="76f29081-b71e-4128-bb0c-865b68e0bbe1"
#test data on confusion matrix and accuracy
from sklearn.metrics import confusion_matrix
cm=confusion_matrix(Y_test,model[0].predict(X_test))
print(cm)
# + id="_iVkmjqdgSe-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c9f9531a-0ea6-4451-f14a-1e35092aa9b9"
TP=cm[0][0]
TN=cm[1][1]
FN=cm[1][0]
FP=cm[0][1]
print("Testing accuracy=",(TP+TN)/(TP+TN+FN+FP))
# + id="DFJUSAWLgz0i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 319} outputId="8612cd53-f89c-412f-f83e-6c9e72f5e6ee"
for i in range(len(model)):
print("Model",i)
cm=confusion_matrix(Y_test,model[i].predict(X_test))
TP=cm[0][0]
TN=cm[1][1]
FN=cm[1][0]
FP=cm[0][1]
print(cm)
if(i==0):
print("Logistic Regression model")
print("Testing accuracy=",(TP+TN)/(TP+TN+FN+FP))
if(i==1):
print("Decision Tree classifier model")
print("Testing accuracy=",(TP+TN)/(TP+TN+FN+FP))
if(i==2):
print("Random Forest classifier model")
print("Testing accuracy=",(TP+TN)/(TP+TN+FN+FP))
print()
# + id="_tObCdNeilzw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="64e4c8ec-1278-47d3-b97b-ff535426c539"
#other matrix model
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
print(classification_report(Y_test,model[0].predict(X_test)))
print(accuracy_score(Y_test,model[0].predict(X_test)))
# + id="inF25SlIxXwH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 672} outputId="43a62433-e123-4c34-e785-b460227fa2c0"
for i in range(len(model)):
print('Model',i)
if(i==0):
print("Logistic Regression model")
print(classification_report(Y_test,model[i].predict(X_test)))
print(accuracy_score(Y_test,model[i].predict(X_test)))
if(i==1):
print("Decision Tree classifier model")
print(classification_report(Y_test,model[i].predict(X_test)))
print(accuracy_score(Y_test,model[i].predict(X_test)))
if(i==2):
print("Random Forest classifier model")
print(classification_report(Y_test,model[i].predict(X_test)))
print(accuracy_score(Y_test,model[i].predict(X_test)))
print()
# + id="uvSdrTDdzxV3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 168} outputId="cc20b8cf-1ea4-4204-e5a1-30bc2e333ccf"
#prediction in the forest classifier model
pred=model[2].predict(X_test)
print(pred)
print()
print(Y_test)
|
MLostraining.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table style="width:100%">
# <tr>
# <td style="background-color:#EBF5FB; border: 1px solid #CFCFCF">
# <b>National generation capacity: Processing notebook</b>
# <ul>
# <li><a href="main.ipynb">Main notebook</a></li>
# <li>Processing notebook (this)</li>
# <li><a href="tests.ipynb">Check notebook</a></li>
# </ul>
# <br>This Notebook is part of the <a href="http://data.open-power-system-data.org/national_generation_capacity">National Generation Capacity Datapackage</a> of <a href="http://open-power-system-data.org">Open Power System Data</a>.
# </td>
# </tr>
# </table>
# # Table of Contents
# * [1. Introductory notes](#1.-Introductory-notes)
# * [2. Script setup](#2.-Script-setup)
# * [3. Data download and processing](#3.-Data-download-and-processing)
# * [3.1 Manually compiled dataset](#3.1-Manually-compiled-dataset)
# * [3.2 EUROSTAT data](#3.2-EUROSTAT-data)
# * [3.3 ENTSO-E data](#3.3-ENTSO-E-data)
# * [3.3.1 ENTSO-E statistical data](#3.3.1-ENTSO-E-statistical-data)
# * [3.3.2 ENTSO-E SO&AF data](#3.3.2-ENTSO-E-SO&AF-data)
# * [3.3.3 ENTSO-E Transparency Plaftform](#3.3.1-ENTSO-E-Transparency-Platform)
# * [3.3.4 ENTSO-E Power Statistics](#3.3.4-ENTSO-E-Power-Statistics)
# * [3.4 Merge data sources](#3.4-Merge-data-sources)
# * [4. Convert stacked data to crosstable format](#4.-Convert-stacked-data-to-crosstable-format)
# * [5. Output](#5.-Output)
# * [5.1 Write results to file](#5.1-Write-results-to-file)
# * [5.2 Formatting of Excel tables](#5.2-Formatting-of-Excel-tables)
# * [5.3 Write checksums](#5.3-Write-checksums)
# * [6. Documentation of the data package](#6.-Documentation-of-the-data-package)
# # 1. Introductory notes
# The script processes the compiled nationally aggregated generation capacity for European countries. Due to varying formats and data specifications of references for national generation capacity, the script firstly focuses on rearranging the manually compiled data. Thus, the script itself does not collect, select, download or manage data from original sources. Secondly, international data sources, such as EUROSTAT and ENTSO-E, are directly downloaded from original web sources and complement the initial data set.
# # 2. Script setup
# +
# some functions and classes that are defined in seperate files
import functions.helper_functions as func
import functions.soaf as soaf
# core packages
import os
import pandas as pd
import numpy as np
# packages to copy files, write sqllite data bases and manipulate excel files
import shutil
import sqlite3
import openpyxl
from openpyxl.styles import PatternFill, colors, Font, Alignment
from openpyxl.utils import get_column_letter
import yaml
import json
# -
# # 3. Data download and processing
# We compile data from different national and international sources. Firstly, national data sources are manually compiled due to varying data formats and specifications. Secondly, international sources are compiled directly and appended to the compiled data set. The international data sources comprise:
# - [EUROSTAT](http://ec.europa.eu/eurostat/product?code=nrg_113a&mode=view)
# - [ENTSO-E Statistical data](https://www.entsoe.eu/data/data-portal/miscellaneous/Pages/default.aspx)
# - [ENTSO-E System Outlook and Adequacy Forecast](https://www.entsoe.eu/outlooks/maf/Pages/default.aspx)
# - [ENTSO-E Transparency Platform](https://transparency.entsoe.eu/)
# - [ENTSO-E Power Statistics](https://www.entsoe.eu/data/power-stats/)
#
# In the following section, the data sets are downloaded and uploaded to Python.
# ## 3.1 Manually compiled dataset
# The manually compiled dataset is imported and rearranged to a DataFrame for further processing. The dataset comprises for each European country and specified generation technology different data entries, which are based on different sources. As these sources differ by country and year, information on the corresponding reference are directly given with the data entry.
# +
data_file = 'National_Generation_Capacities.xlsx'
filepath = os.path.join('input', data_file)
# Read data into pandas
data_raw = pd.read_excel(filepath,
sheet_name='Summary',
header=None,
na_values=['-'],
skiprows=0)
# Deal with merged cells from Excel: fill first three rows with information
data_raw.iloc[0:2] = data_raw.iloc[0:2].fillna(method='ffill', axis=1)
# Set index for rows
data_raw = data_raw.set_index([0])
data_raw.index.name = 'technology'
# Extract energylevels from raw data for later use
energylevels_raw = data_raw.iloc[:, 0:5]
energylevels_raw.head()
# +
# Delete definition of energy levels from raw data
data_raw.drop(data_raw.columns[[0, 1, 2, 3, 4, 5]], axis=1, inplace=True)
level_names = ['country', 'type', 'year', 'source',
'source_type', 'weblink', 'capacity_definition']
# Set multiindex column names
data_raw.columns = pd.MultiIndex.from_arrays(data_raw[:7].values,
names=level_names)
# Remove 3 rows which are already used as column names
data_raw = data_raw[pd.notnull(data_raw.index)]
# Extract the ordering of technologies
technology_order = data_raw.index.str.replace('- ', '').values.tolist()
data_raw.head()
# -
data_raw.columns[data_raw.columns.duplicated()]
data_raw[data_raw.index.duplicated()]
# +
# Reshape dataframe to list
data_opsd = pd.DataFrame(data_raw.stack(level=level_names))
# Reset index for dataframe
data_opsd.reset_index(inplace=True)
data_opsd['technology'] = data_opsd['technology'].str.replace('- ', '')
data_opsd.rename(columns={0: 'capacity'}, inplace=True)
data_opsd['capacity'] = pd.to_numeric(data_opsd['capacity'], errors='coerce')
# For some source, permission to publish data
banlist = ['ELIA', 'BMWi', 'Mavir']
davail = 'data available, but cannot be provided'
data_opsd.loc[data_opsd['source'].isin(banlist), 'comment'] = davail
data_opsd.head()
# -
# ## 3.2 EUROSTAT data
# EUROSTAT publishes annual structural data on national electricity generation capacities for European countries. The dataset is available in the EUROSTAT database within the category 'Environment and Energy' ([nrg_113a](http://ec.europa.eu/eurostat/product?code=nrg_113a&mode=view)).
# +
url_eurostat = 'Link unavailable'
filepath_eurostat = os.path.join('input', 'Eurostat', 'Eurostat.tsv.gz')
data_eurostat = pd.read_csv(filepath_eurostat,
compression='gzip',
sep='\t|,',
engine='python'
)
data_eurostat.head()
# +
id_vars = ['unit', 'product','indic_nrg', 'geo\\time']
data_eurostat = pd.melt(data_eurostat, id_vars=id_vars,
var_name='year', value_name='value')
data_eurostat.head()
# +
data_definition = pd.read_csv(os.path.join('input', 'definition_EUROSTAT_indic.txt'),
header=None,
names=['indic', 'description',
'energy source'],
sep='\t')
data_eurostat = data_eurostat.merge(data_definition,
how='left',
left_on='indic_nrg',
right_on='indic')
# -
# The classification of generation capacities in the EUROSTAT dataset is specified in [Regulation (EC) No 1099/2008](http://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32008R1099&from=EN) (Annex B, 3.3). The available EUROSTAT dataset [nrg_113a](http://ec.europa.eu/eurostat/product?code=nrg_113a&mode=view) covers the following indicators:
#
# |indic_nrg | Description | Technology in OPSD |
# |---|---|---|
# |**12_1176011**| **Electrical capacity, main activity producers - Combustible Fuels**| Fossil fuels & bioenergy|
# |**12_1176012**| **Electrical capacity, autoproducers - Combustible Fuels**| Fossil fuels & bioenergy|
# |*12_1176061*| *Electrical capacity, main activity producers - Mixed plants*| |
# |*12_1176101*| *Electrical capacity, main activity producers - Other Sources*| |
# |*12_1176102*| *Electrical capacity, autoproducers - Other Sources*| |
# |*12_1176111*| *Electrical capacity, main activity producers - Steam*||
# |*12_1176112*| *Electrical capacity, autoproducers - Steam*||
# |*12_1176121*| *Electrical capacity, main activity producers - Gas Turbine*||
# |*12_1176122*| *Electrical capacity, autoproducers - Gas Turbine*||
# |*12_1176131*| *Electrical capacity, main activity producers - Combined Cycle*||
# |*12_1176132*| *Electrical capacity, autoproducers - Combined Cycle*||
# |*12_1176141*| *Electrical capacity, main activity producers - Internal Combustion*||
# |*12_1176142*| *Electrical capacity, autoproducers - Internal Combustion*||
# |*12_1176401*| *Electrical capacity, main activity producers - Other Type of Generation*| |
# |*12_1176402*| *Electrical capacity, autoproducers - Other Type of Generation*| |
# |12_1176253| Net maximum capacity - Municipal Wastes| Non-renewable waste|
# |12_1176263| Net maximum capacity - Wood/Wood Wastes/Other Solid Wastes| Other bioenergy and renewable waste|
# |12_1176273| Net maximum capacity - Biogases| Biomass and biogas|
# |12_1176283| Net maximum capacity - Industrial Wastes (non-renewable)| Non-renewable waste|
# |12_1176343| Net maximum capacity - Liquid Biofuels| Biomass and biogas|
# |**12_1176031**| **Electrical capacity, main activity producers - Nuclear**| Nuclear|
# |**12_1176032**| **Electrical capacity, autoproducers - Nuclear**| Nuclear|
# |**12_1176051**| **Electrical capacity, main activity producers - Hydro**| Hydro|
# |**12_1176052**| **Electrical capacity, autoproducers - Hydro**| Hydro|
# |12_1176071| Net electrical capacity, main activity producers - Pure Pumped Hydro| Pumped storage|
# |12_1176072| Net electrical capacity, autoproducers - Pure Pumped Hydro| Pumped storage|
# |*12_117615*| *Net maximum capacity - Hydro <1 MW*| |
# |*12_117616*| *Net maximum capacity - Hydro >= 1 MW and <= 10 MW*| |
# |*12_117617*| *Net maximum capacity - Hydro 10 MW and over*| |
# |**12_1176301**| **Electrical capacity, main activity producers - Tide, wave and ocean**| Marine|
# |**12_1176302**| **Electrical capacity, autoproducers - Tide, wave and ocean**| Marine|
# |*12_1176303*| *Net maximum capacity - Tide, Wave, Ocean*||
# |**12_1176081**| **Electrical capacity, main activity producers - Geothermal**| Geothermal|
# |**12_1176082**| **Electrical capacity, autoproducers - Geothermal**| Geothermal|
# |*12_1176083*| *Net maximum capacity - Geothermal*| |
# |**12_1176091**| **Electrical capacity, main activity producers - Wind**| Wind|
# |**12_1176092**| **Electrical capacity, autoproducers - Wind**| Wind|
# |**12_1176233**| **Net maximum capacity - Solar Photovoltaic**| Photovoltaics|
# |**12_1176243**| **Net maximum capacity - Solar Thermal Electric**| Concentrated solar power|
#
# **Bold** rows indicate top level classes within the EUROSTAT classification, whereas normal and *italic* rows cover different kinds of subclassifications. Especially within the top level 'Combustible fuels' different kinds of subcategorizations based on fuel or technology are available. Simarily, 'Hydro' is differentiated by type (e.g. pumped-hydro storage) or capacity classes. *Italic* rows are not further considered within the OPSD dataset due to the mismatch with existing technology classes.
# +
data_eurostat = data_eurostat[data_eurostat['energy source'].isnull() == False]
values_as_string = data_eurostat['value'].astype(str)
string_values = values_as_string.str.split(' ', 1).str[0]
string_values.replace(':', np.nan, inplace=True)
subset_nan = string_values.isnull()
data_eurostat['value'] = string_values
data_eurostat['year'] = data_eurostat['year'].astype(int)
data_eurostat['value'] = data_eurostat['value'].astype(float)
data_eurostat.head()
# +
data_eurostat = data_eurostat.drop(['unit', 'product', 'indic_nrg',
'indic', 'description'], axis=1)
data_eurostat = data_eurostat.rename(columns={'geo\\time': 'country',
'energy source': 'technology',
'value': 'capacity'})
# +
data_eurostat['country'].replace({'UK': 'GB', 'EL': 'GR'}, inplace=True)
drop_list = data_eurostat[data_eurostat['country'].isin(['EU28','EA19'])].index
data_eurostat.drop(drop_list, inplace=True)
by_columns = ['technology', 'year', 'country']
data_eurostat = pd.DataFrame(data_eurostat.groupby(by_columns)['capacity'].sum())
data_eurostat_isnull = data_eurostat['capacity'].isnull() == True
data_eurostat.reset_index(inplace=True)
data_eurostat.head()
# +
eurostat_pivot = data_eurostat.pivot_table(values='capacity',
index=['country','year'],
columns='technology')
eurostat_pivot.head()
# +
eurostat_pivot['Differently categorized solar'] = 0
eurostat_pivot['Solar'] = eurostat_pivot[['Photovoltaics', 'Concentrated solar power']].sum(axis=1)
eurostat_pivot['Differently categorized wind'] = eurostat_pivot['Wind']
bio_arr = ['Biomass and biogas', 'Other bioenergy and renewable waste']
eurostat_pivot['Bioenergy and renewable waste'] = eurostat_pivot[bio_arr].sum(axis=1)
res_arr = ['Hydro', 'Wind', 'Solar', 'Geothermal', 'Marine', 'Bioenergy and renewable waste']
eurostat_pivot['Renewable energy sources'] = eurostat_pivot[res_arr].sum(axis=1)
eurostat_pivot['Fossil fuels'] = eurostat_pivot['Fossil fuels'] - eurostat_pivot['Bioenergy and renewable waste']
eurostat_pivot['Differently categorized fossil fuels'] = eurostat_pivot['Fossil fuels']\
- eurostat_pivot['Non-renewable waste']
total_arr = ['Fossil fuels','Nuclear','Renewable energy sources']
eurostat_pivot['Total'] = eurostat_pivot[total_arr].sum(axis=1)
eurostat_pivot.head()
# +
data_eurostat = eurostat_pivot.stack().reset_index().rename(columns={0: 'capacity'})
data_eurostat['source'] = 'EUROSTAT'
data_eurostat['source_type'] = 'Statistical Office'
data_eurostat['capacity_definition'] = 'Unknown'
data_eurostat['type'] = 'Installed capacity in MW'
data_eurostat['weblink'] = url_eurostat
data_eurostat.head()
# -
# ## 3.3 ENTSO-E data
# The ENTSO-E publishes annual data on national generation capacites in different specifications and formats. We use two relevant data sources from the ENTSOE-E, which comprises firstly statistical data within the [Data Portal (up to 2015)](https://www.entsoe.eu/data/data-portal/Pages/default.aspx) or [ENTSO-E Transparency Platform](https://transparency.entsoe.eu/), and secondly datasets compiled within the [ENTSO-E System Outlook & Adequacy Forecast (SO&AF)](https://www.entsoe.eu/outlooks/maf/Pages/default.aspx). The ENTSO-E Transparency Platform is currently not implemented as a data source for national generation capacities.
#
# The advantage of the ENTSO-E SO&AF is the higher granularity of the data with respect to the main fuel or technology. However, as the SO&AF provides a forecast on future system conditions in particular peak hours, the dataset also accounts for expected capacity changes throughout the years. Therefore, we only consider years which are closest to the publication year of the respective SO&AF.
# ### 3.3.1 ENTSO-E Statistical Data
# In the following, we use the statistical data available in the [Data Portal (up to 2015)](https://www.entsoe.eu/data/data-portal/Pages/default.aspx).
# +
url_entsoe = 'https://docstore.entsoe.eu/Documents/Publications/Statistics/NGC_2010-2015.xlsx'
filepath_entsoe = func.downloadandcache(url_entsoe, 'Statistics.xls',
os.path.join('ENTSO-E','Data Portal 2010-2015')
)
data_entsoe_raw = pd.read_excel(filepath_entsoe)
data_entsoe_raw.head()
# +
dict_energy_source = {'hydro': 'Hydro',
'of which storage': 'Reservoir',
'of which run of river': 'Run-of-river',
'of which pumped storage': 'Pumped storage',
'nuclear': 'Nuclear',
'of which wind': 'Wind',
'of which solar': 'Solar',
'of which biomass': 'Biomass and biogas',
'fossil_fuels': 'Fossil fuels',
'other': 'Other or unspecified energy sources',
"Country": "country",
'fossil_fueals': 'Fossil fuels'}
data_entsoe_raw.rename(columns=dict_energy_source,
inplace=True)
data_entsoe_raw.drop(columns='representativity', inplace=True)
data_entsoe_raw.head()
# +
data_entsoe_raw['Differently categorized solar'] = data_entsoe_raw['Solar']
data_entsoe_raw['Differently categorized wind'] = data_entsoe_raw['Wind']
data_entsoe_raw['Bioenergy and renewable waste'] = data_entsoe_raw['Biomass and biogas']
data_entsoe_raw['Differently categorized fossil fuels'] = data_entsoe_raw['Fossil fuels']
data_entsoe_raw['Differently categorized hydro'] = (
data_entsoe_raw['Hydro']
- data_entsoe_raw['Run-of-river']
- data_entsoe_raw['Reservoir']
- data_entsoe_raw['Pumped storage'])
data_entsoe_raw['Differently categorized renewable energy sources'] = (
data_entsoe_raw['renewable']
- data_entsoe_raw['Wind']
- data_entsoe_raw['Solar']
- data_entsoe_raw['Biomass and biogas'])
data_entsoe_raw.drop(columns='renewable', inplace=True)
data_entsoe_raw['Renewable energy sources'] = (
data_entsoe_raw['Hydro']
+ data_entsoe_raw['Wind']
+ data_entsoe_raw['Solar']
+ data_entsoe_raw['Bioenergy and renewable waste']
+ data_entsoe_raw['Differently categorized renewable energy sources'])
data_entsoe_raw['Total'] = (
data_entsoe_raw['Renewable energy sources']
+ data_entsoe_raw['Nuclear']
+ data_entsoe_raw['Fossil fuels']
+ data_entsoe_raw['Other or unspecified energy sources'])
data_entsoe = pd.melt(data_entsoe_raw,
id_vars=['country', 'year'],
var_name='technology',
value_name='capacity')
data_entsoe.head()
# +
data_entsoe['country'].replace('NI', 'GB', inplace=True)
# set negative capacities to zero
data_entsoe.loc[data_entsoe['capacity'] < 0, 'capacity'] = 0
data_entsoe['source'] = 'ENTSO-E Data Portal'
data_entsoe['source_type'] = 'Other association'
data_entsoe['capacity_definition'] = 'Net capacity'
data_entsoe['type'] = 'Installed capacity in MW'
data_entsoe.head()
# -
# ### 3.3.2 ENTSO-E SO&AF data
# +
soafs = [soaf.SoafDataRaw('https://www.entsoe.eu/fileadmin/user_upload/_library/SDC/SOAF/SO_AF_2011_-_2025_.zip',
'SO_AF_2011_-_2025_.zip',
'SO&AF 2011 - 2025 Scenario B.xls',
2011),
soaf.SoafDataRaw('https://www.entsoe.eu/fileadmin/user_upload/_library/SDC/SOAF/120705_SOAF_2012_Dataset.zip',
'120705_SOAF_2012_Dataset.zip',
'SOAF 2012 Scenario B.xls',
2012),
soaf.SoafDataRaw('https://www.entsoe.eu/fileadmin/user_upload/_library/publications/entsoe/So_AF_2013-2030/130403_SOAF_2013-2030_dataset.zip',
'130403_SOAF_2013-2030_dataset.zip',
'ScB.xls',
2013),
soaf.SoafDataRaw('https://www.entsoe.eu/Documents/SDC%20documents/SOAF/140602_SOAF%202014_dataset.zip',
'140602_SOAF%202014_dataset.zip',
'ScB.xlsx',
2014),
soaf.SoafDataRaw('https://www.entsoe.eu/Documents/Publications/SDC/data/SO_AF_2015_dataset.zip',
'SO_AF_2015_dataset.zip',
os.path.join('SO&AF 2015 dataset', 'ScB_publication.xlsx'),
2016)]
data_soaf = pd.concat([s.transformed_df for s in soafs])
# Correct that in the Soaf2015 datatset the year column is 2016 instead of 2015
data_soaf['year'].replace({2016 : 2015}, inplace=True)
data_soaf.head()
# +
soaf_unstacked = func.unstackData(data_soaf)
soaf_unstacked['Differently categorized solar'] = soaf_unstacked['Solar']
soaf_unstacked['Differently categorized wind'] = soaf_unstacked['Wind']\
- soaf_unstacked['Offshore']\
- soaf_unstacked['Onshore']
soaf_unstacked['Differently categorized hydro'] = soaf_unstacked['Hydro']\
- soaf_unstacked['Run-of-river']\
- soaf_unstacked['Reservoir including pumped storage']
soaf_unstacked['Bioenergy and renewable waste'] = soaf_unstacked['Biomass and biogas']
soaf_unstacked['Differently categorized renewable energy sources'] = (
soaf_unstacked['renewable']
- soaf_unstacked['Wind']
- soaf_unstacked['Solar']
- soaf_unstacked['Biomass and biogas'])
soaf_unstacked.drop(columns='renewable', inplace=True)
subtract_fossils_arr = ['Lignite','Hard coal','Oil','Natural gas','Mixed fossil fuels']
soaf_unstacked['Differently categorized fossil fuels'] = soaf_unstacked['Fossil fuels']\
- soaf_unstacked[subtract_fossils_arr].sum(axis=1)
res_arr = ['Solar','Wind','Bioenergy and renewable waste','Hydro',
'Differently categorized renewable energy sources']
soaf_unstacked['Renewable energy sources'] = soaf_unstacked[res_arr].sum(axis=1)
total_arr = ['Renewable energy sources','Fossil fuels','Nuclear',
'Other or unspecified energy sources']
soaf_unstacked['Total'] = soaf_unstacked[total_arr].sum(axis=1)
soaf_unstacked.head()
# +
data_soaf = func.restackData(soaf_unstacked)
data_soaf.loc[data_soaf['capacity'] < 0, 'capacity'] = 0
data_soaf['source'] = 'ENTSO-E SOAF'
data_soaf['type'] = 'Installed capacity in MW'
data_soaf['capacity_definition'] = 'Net capacity'
data_soaf['source_type'] = 'Other association'
data_soaf['weblink'] = url_entsoe
data_soaf.head()
# -
# ### 3.3.3 ENTSO-E Transparency Platform
# +
# file pattern for the single years
filenamepattern = '_1_InstalledGenerationCapacityAggregated.csv'
list_of_data_tables = [] # list to append
# iterate over the years from 2015 to 2020
for i in range(2015,2021):
filepath = os.path.join('input',
'ENTSO-E',
'Transparency',
'InstalledGenerationCapacityAggregated',
str(i) + filenamepattern)
list_of_data_tables.append(pd.read_csv(filepath, delimiter="\t", encoding = "UTF-16"))
# merge the datasets of the single of files into one pandas dataframe
data_transparency = pd.concat(list_of_data_tables, ignore_index=True)
data_transparency.head()
# +
# rename columns according to the opsd standards
data_transparency.rename(columns={'ProductionType': 'technology',
'AggregatedInstalledCapacity': 'capacity',
'MapCode': 'country',
'Year': 'year'}, inplace=True)
# drop non relevant columns
data_transparency = data_transparency.filter(items=['technology','capacity','country','year'], axis=1)
# drop countries that are not part of opsd
data_transparency = data_transparency[data_transparency['country'].isin(data_opsd.country.unique())]
data_transparency.head()
# +
# adapt energy source notation
dict_energy_source = {'Biomass': 'Biomass and biogas',
'Fossil Brown coal/Lignite': 'Lignite',
'Fossil Coal-derived gas': 'Mixed fossil fuels',
'Fossil Gas': 'Natural gas',
'Fossil Hard coal': 'Hard coal',
'Fossil Oil': 'Oil',
'Fossil Oil shale': 'Oil',
'Fossil Peat': 'Other fossil fuels',
'Hydro Pumped Storage': 'Pumped storage',
'Hydro Run-of-river and poundage': 'Run-of-river',
'Hydro Water Reservoir': 'Reservoir',
'Other': 'Other or unspecified energy sources',
'Other renewable': 'Differently categorized renewable energy sources',
'Waste': 'Other bioenergy and renewable waste',
'Wind Offshore': 'Offshore',
'Wind Onshore': 'Onshore',
' ': np.nan}
data_transparency['technology'].replace(dict_energy_source, inplace=True)
data_transparency.head()
# +
# add missing categories
transparency_pivot = data_transparency.pivot_table(values='capacity',
index=['country','year'],
columns='technology')
# technology level
transparency_pivot['Differently categorized solar'] = transparency_pivot['Solar']
transparency_pivot['Differently categorized natural gas'] = transparency_pivot['Natural gas']
transparency_pivot['Non-renewable waste'] = 0
transparency_pivot['Differently categorized fossil fuels'] = 0
# level 3
hydro_arr = ['Pumped storage', 'Reservoir', 'Run-of-river']
transparency_pivot['Hydro'] = transparency_pivot[hydro_arr].sum(axis=1)
wind_arr = ['Onshore', 'Offshore']
transparency_pivot['Wind'] = transparency_pivot[wind_arr].sum(axis=1)
# level 2
bio_arr = ['Biomass and biogas', 'Other bioenergy and renewable waste']
transparency_pivot['Bioenergy and renewable waste'] = transparency_pivot[bio_arr].sum(axis=1)
#level 1
res_arr = ['Hydro', 'Wind', 'Solar', 'Geothermal', 'Marine',
'Bioenergy and renewable waste', 'Differently categorized renewable energy sources']
transparency_pivot['Renewable energy sources'] = transparency_pivot[res_arr].sum(axis=1)
fossil_arr = ['Lignite', 'Hard coal', 'Oil', 'Natural gas', 'Mixed fossil fuels', 'Other fossil fuels']
transparency_pivot['Fossil fuels'] = transparency_pivot[fossil_arr].sum(axis=1)
# level 0
total_arr = ['Fossil fuels','Nuclear','Renewable energy sources', 'Other or unspecified energy sources']
transparency_pivot['Total'] = transparency_pivot[total_arr].sum(axis=1)
transparency_pivot.reset_index(inplace=True)
transparency_pivot.head()
# +
data_transparency = pd.melt(transparency_pivot,
id_vars=['country', 'year'],
var_name='technology',
value_name='capacity')
data_transparency = data_transparency.loc[data_transparency["year"] < 2020, :]
data_transparency['source'] = 'ENTSO-E Transparency Platform'
data_transparency['source_type'] = 'Other association'
data_transparency['capacity_definition'] = 'Net capacity'
data_transparency['type'] = 'Installed capacity in MW'
data_transparency['weblink'] = ('https://transparency.entsoe.eu/generation'
'/r2/installedGenerationCapacityAggregation/show')
data_transparency.head()
# -
# ### 3.3.4 ENTSO-E Power Statistics
# +
row_of_year = {2014: 9,
2015: 53,
2016: 97,
2017: 141,
2018: 185}
dataframes = []
for year, row in row_of_year.items():
# read the dataframe for each year
power_statistics_raw = pd.read_excel(os.path.join('input',
'ENTSO-E',
'Power Statistics',
'NGC.xlsx'),
header=[0,1],
sheet_name='NGC',
skiprows=row,
nrows=42)
# drop non relevant columns
power_statistics_raw.drop(columns='Coverage ratio in %', level=1, inplace=True)
power_statistics_raw.drop(columns=['Unnamed: 1_level_1','Unnamed: 2_level_1','Unnamed: 3_level_1','Unnamed: 4_level_1'], level=1, inplace=True)
# get rid of multi index
df = power_statistics_raw.set_index(year).stack().reset_index().drop('level_1', axis=1)
# remove leftovers of multi index in the index column
df["technology"] = df[year].apply(lambda x: x[0])
df.drop(columns=year, inplace=True)
# stack df to the opsd standard format
stacked_df = df.melt(id_vars='technology', var_name='country', value_name='capacity')
# add information about the year
stacked_df['year'] = year
# append to the main list of dataframes
dataframes.append(stacked_df)
power_statistics = pd.concat(dataframes)
power_statistics.head()
# +
# drop countries that are not covered in opsd
opsd_countries = data_opsd.country.unique()
drop_list_country = power_statistics.loc[~power_statistics['country'].isin(opsd_countries)].index.to_list()
power_statistics.drop(drop_list_country, inplace=True)
# technology classes to be dropped
tech_to_drop = ['Non-Renewable', 'Fossil fuels', 'Renewable','Non-renwable hydro',
'Total Waste', 'Bio', 'Renewable Hydro', 'Comments', 'Total NGC']
drop_list_tech = power_statistics.loc[power_statistics['technology'].isin(tech_to_drop)].index.to_list()
power_statistics.drop(drop_list_tech, inplace=True)
# replace string with values that can be used in math operations
power_statistics['capacity'].replace(to_replace='Not Expected', value=0, inplace=True)
power_statistics['capacity'].replace(to_replace='Not Available', value=np.nan, inplace=True)
power_statistics.head()
# +
# Not included because already categorized in OPSD standard:
# Nuclear, Solar, Geothermal, Wind
dict_energy_source = {'Of which hydro pure pumped storage':'Pumped storage',
'Of which Hydro mixed pumped storage (non renewable part)':'Pumped storage',
'Of which Fossil Brown coal/Lignite':'Lignite',
'Of which Fossil Coal-derived gas':'Differently categorized fossil fuels',
'Of which Fossil Gas':'Natural gas',
'Of which Fossil Hard coal':'Hard coal',
'Of which Fossil Oil':'Oil',
'Of which Fossil Oil shale':'Oil',
'Of which Fossil Peat':'Differently categorized fossil fuels',
'Of which Mixed fuels':'Mixed fossil fuels',
'Of which Other fossil fuels':'Other fossil fuels',
'Non-renewable Waste':'Non-renewable waste',
'Other non-renewable':'Differently categorized fossil fuels',
'Of which Wind offshore':'Offshore',
'Of which Wind onshore':'Onshore',
'Of which Solar PV':'Photovoltaics',
'Of which Solar Thermal':'Differently categorized solar',
'Of which Biomass':'Biomass and biogas',
'Of which Biogas':'Biomass and biogas',
'Renewable Waste':'Other bioenergy and renewable waste',
'Of which Hydro Pure storage':'Reservoir',
'Of which Hydro Run-of-river and pondage':'Run-of-river',
'Of which Hydro mixed pumped storage (renewable part)':'Pumped storage',
'Of which Hydro Marine (tidal/wave)':'Marine',
'Other renewable (not listed)':'Differently categorized renewable energy sources',
'Non identified (other not listed)':'Other or unspecified energy sources',
'Total Hydro':'Hydro'}
power_statistics["technology"].replace(dict_energy_source, inplace=True)
power_statistics.head()
# -
powerstats_pivot = power_statistics.pivot_table(values='capacity',
index=['country','year'],
columns='technology').reset_index()
powerstats_pivot.head()
# +
# technology level
powerstats_pivot['Differently categorized natural gas'] = powerstats_pivot['Natural gas']
# level 2
powerstats_pivot['Bioenergy and renewable waste'] = (
powerstats_pivot['Biomass and biogas'] +
powerstats_pivot['Other bioenergy and renewable waste'])
#level 1
fossil_techs = ['Lignite', 'Hard coal', 'Oil', 'Natural gas', 'Non-renewable waste',
'Mixed fossil fuels', 'Other fossil fuels', 'Differently categorized fossil fuels']
powerstats_pivot['Fossil fuels'] = powerstats_pivot[fossil_techs].sum(axis=1)
res_tech = ['Hydro', 'Wind', 'Solar', 'Geothermal', 'Marine', 'Bioenergy and renewable waste',
'Differently categorized renewable energy sources']
powerstats_pivot['Renewable energy sources'] = powerstats_pivot[res_tech].sum(axis=1)
total_arr = ['Fossil fuels','Nuclear','Renewable energy sources']
powerstats_pivot['Total'] = powerstats_pivot[total_arr].sum(axis=1)
powerstats_pivot.head()
# +
data_power_statistics = powerstats_pivot.melt(id_vars=['country', 'year'],
var_name='technology',
value_name='capacity')
data_power_statistics['source'] = 'ENTSO-E Power Statistics'
data_power_statistics['source_type'] = 'Other association'
data_power_statistics['capacity_definition'] = 'Net capacity'
data_power_statistics['type'] = 'Installed capacity in MW'
data_power_statistics['weblink'] = ('https://www.entsoe.eu/data/'
'power-stats/net-gen-capacity/')
data_power_statistics.head()
# -
# ## 3.4 Merge data sources
# +
dataframes = [data_opsd, data_eurostat, data_soaf, data_entsoe, data_transparency, data_power_statistics]
data = pd.concat(dataframes, sort=False)
data['comment'] = data['comment'].fillna('').astype(str)
col_order = ['technology', 'source', 'source_type', 'weblink', 'year', 'type',
'country', 'capacity_definition', 'capacity', 'comment']
data = data[col_order]
energy_source_mapping = pd.read_csv(os.path.join('input','energy_source_mapping.csv'),
index_col ='name')
energy_source_mapping.replace({0: False, 1: True}, inplace=True)
data = data.merge(energy_source_mapping,
left_on='technology',
right_index=True,
how='left')
new_level_names = {"Level 0": "energy_source_level_0",
"Level 1": "energy_source_level_1",
"Level 2": "energy_source_level_2",
"Level 3": "energy_source_level_3",
"Technology level": "technology_level"}
data.rename(columns=new_level_names, inplace=True)
data.head()
# -
# # 4. Convert stacked data to crosstable format
# +
cols = ['technology', 'source', 'source_type', 'weblink','year',
'type', 'country', 'capacity_definition', 'capacity']
data_crosstable = pd.pivot_table(data[cols],
index=['technology'],
columns=['country', 'type', 'year',
'source', 'source_type',
'weblink', 'capacity_definition'],
values='capacity')
# Apply initial ordering of technologies
data_crosstable = data_crosstable.reindex(technology_order)
# Delete index naming
data_crosstable.index.name = None
data_crosstable.columns.names = ('Country (ISO code)',
'Type of data', 'Year',
'Source', 'Type of source',
'Weblink',
'Capacity definition (net, gross, unknown)')
data_crosstable.head()
# +
energylevels_table = energylevels_raw[6:]
energylevels_table.columns = pd.MultiIndex.from_arrays(energylevels_raw[:6].values,
names=['country', 'type', 'year',
'source', 'source_type',
'capacity_definition'
])
energylevels_table = energylevels_table.reset_index()
energylevels_table['technology'] = energylevels_table['technology'].str.replace('- ', '')
energylevels_table = energylevels_table.set_index('technology')
# Delete index naming
energylevels_table.index.name = None
energylevels_table.columns.names = ('Country (ISO code)',
'Description', None,
None, None,
'Level')
energylevels_table.head()
# -
# # 5. Output
# Delete downloaded zip files
for root, dirs, files in os.walk("download"):
for file in files:
item = os.path.join(root, file)
if item.endswith(".zip"):
os.remove(item)
print("Deleted: " + item)
# Copy input files
orig_data_path = os.path.join('output', 'original_data')
shutil.rmtree(orig_data_path, ignore_errors=True)
func.copydir(os.path.join('input'), orig_data_path)
func.copydir(os.path.join('download'), orig_data_path)
# ## 5.1 Write results to file
# Write stacked data to formats: csv, xls and sql.
# +
# Write the result to file
data.to_csv(os.path.join('output', 'national_generation_capacity_stacked.csv'),
encoding='utf-8', index_label='ID')
# Write the results to excel file
data.to_excel(os.path.join('output', 'national_generation_capacity_stacked.xlsx'),
sheet_name='output', index_label='ID')
# Write the results to sql database
data.to_sql('national_generation_capacity_stacked',
sqlite3.connect(os.path.join('output',
'national_generation_capacity.sqlite')),
if_exists="replace", index_label='ID')
# -
# Write data in human readable form to excel.
# Write crosstable data to excel file
writer = pd.ExcelWriter(os.path.join('output', 'national_generation_capacity.xlsx'))
data_crosstable.to_excel(writer, sheet_name='output')
energylevels_table.to_excel(writer, sheet_name='technology levels')
writer.save()
# ## 5.2 Formatting of Excel tables
# +
outputxls = openpyxl.load_workbook(os.path.join('output',
'national_generation_capacity.xlsx'))
ws1 = outputxls['output']
ws2 = outputxls['technology levels']
ws1_rows, ws1_cols = data_crosstable.shape
amount_cols = ws1_cols + 1 # correct 0 index
ws1.column_dimensions['A'].width = 50
ws2.column_dimensions['A'].width = 50
# +
blackfont = Font(color=colors.BLACK, italic=False, bold=False)
blackfontitalic = Font(color=colors.BLACK, italic=True, bold=False)
blackfontbold = Font(color=colors.BLACK, italic=False, bold=True)
align0 = Alignment(horizontal='left', indent=0)
align1 = Alignment(horizontal='left', indent=1)
align2 = Alignment(horizontal='left', indent=2)
# darkest grey
colour = "{0:02X}{1:02X}{2:02X}".format(166, 166, 166)
grey166 = PatternFill(fgColor=colour, bgColor=colour, patternType="solid")
# darker grey
colour = "{0:02X}{1:02X}{2:02X}".format(191, 191, 191)
grey191 = PatternFill(fgColor=colour, bgColor=colour, patternType="solid")
# lighter grey
colour = "{0:02X}{1:02X}{2:02X}".format(217, 217, 217)
grey217 = PatternFill(fgColor=colour, bgColor=colour, patternType="solid")
# lightest grey
colour = "{0:02X}{1:02X}{2:02X}".format(242, 242, 242)
grey242 = PatternFill(fgColor=colour, bgColor=colour, patternType="solid")
# +
for col in range(2, amount_cols+1):
colname = openpyxl.utils.cell.get_column_letter(col)
ws1.column_dimensions[colname].width = 16
for col in range(1, amount_cols+1):
# format column name block
for row in range(2,8):
ws1.cell(row=row, column=col).font = blackfont
# format cells that contain the values
for row in range(9, 48):
ws1.cell(row=row, column=col).fill = grey242
ws1.cell(row=row, column=1).font = blackfontitalic
ws1.cell(row=row, column=1).alignment = align2
# format row 'Total' with dark grey
ws1.cell(row=47, column=col).fill = grey166
ws1.cell(row=47, column=col).font = blackfontbold
# format level 1
for row in [9, 22, 23, 46]:
ws1.cell(row=row, column=col).fill = grey191
ws1.cell(row=row, column=col).font = blackfontbold
ws1.cell(row=row, column=1).alignment = align0
# format level 2
for row in [10, 11, 12, 13, 18, 19, 20, 21, 24, 31, 35, 39, 40, 41, 45]:
ws1.cell(row=row, column=col).fill = grey217
ws1.cell(row=row, column=1).alignment = align1
ws1.cell(row=47, column=1).alignment = align0
ws1.freeze_panes = ws1['B8'] #freeze first column and header rows
# +
# do the same for the second worksheet 'technology levels'
for col in range(1, 7):
colname = get_column_letter(col + 1)
ws2.column_dimensions[colname].width = 25
for row in range(2, 8):
ws2.cell(row=row, column=col).font = blackfont
for row in range(9, 48):
ws2.cell(row=row, column=col).fill = grey242
ws2.cell(row=row, column=1).font = blackfontitalic
ws2.cell(row=row, column=1).alignment = align2
# format row 'Total' with dark grey
ws2.cell(row=46, column=col).fill = grey166
# format level 1
for row in [8, 21, 22, 45]:
ws2.cell(row=row, column=col).fill = grey191
ws2.cell(row=row, column=col).font = blackfontbold
ws2.cell(row=row, column=1).font = blackfontbold
ws2.cell(row=row, column=1).alignment = align0
# format level 2
for row in [9, 10, 11, 12, 17, 18, 19, 20, 23, 30, 34, 38, 39, 40, 44]:
ws2.cell(row=row, column=col).fill = grey217
ws2.cell(row=row, column=1).alignment = align1
ws2.cell(row=46, column=1).alignment = align0
# -
additional_notes = openpyxl.load_workbook(os.path.join('input',
'National_Generation_Capacities.xlsx'))['Additional notes']
# # copy additional notes to output file
for col in range(1, 3):
for row in range(1, 10):
add_notes_value = additional_notes.cell(row=row, column=col).value
ws1.cell(row=row + 50, column=col).value = add_notes_value
ws1.cell(row=51, column=1).font = blackfontbold
ws1.cell(row=row + 51, column=1).font = blackfontitalic
outputxls.save(os.path.join('output', 'national_generation_capacity.xlsx'))
# ## 5.3 Write checksums
# +
files = ['national_generation_capacity.xlsx',
'national_generation_capacity_stacked.csv',
'national_generation_capacity_stacked.xlsx',
'national_generation_capacity.sqlite']
hash_dict = {}
filesize_dict = {}
with open('checksums.txt', 'w') as f:
for file_name in files:
path = os.path.join('output', file_name)
file_hash = func.get_sha_hash(path)
hash_dict[file_name] = file_hash
filesize_dict[file_name] = os.path.getsize(path)
f.write('{},{}\n'.format(file_name, file_hash))
# -
# # 6. Documentation of the data package
# We document the data packages meta data in the specific format JSON as proposed by the Open Knowledge Foundation. See the Frictionless Data project by OKFN (http://data.okfn.org/) and the Data Package specifications (http://dataprotocols.org/data-packages/) for more details.
#
# In order to keep the notebook more readable, we first formulate the metadata in the human-readable YAML format using a multi-line string. We then parse the string into a Python dictionary and save that to disk as a JSON file.
# +
with open(os.path.join('input', 'metadata.yml'), 'r') as f:
metadata = yaml.load(f.read(), Loader=yaml.BaseLoader)
metadata['resources'][0]['hash'] = hash_dict['national_generation_capacity.xlsx']
metadata['resources'][1]['hash'] = hash_dict['national_generation_capacity_stacked.csv']
metadata['resources'][0]['bytes'] = filesize_dict['national_generation_capacity.xlsx']
metadata['resources'][1]['bytes'] = filesize_dict['national_generation_capacity_stacked.csv']
# +
datapackage_json = json.dumps(metadata, indent=4, separators=(',', ': '))
# Write the information of the metadata
with open(os.path.join('output', 'datapackage.json'), 'w') as f:
f.write(datapackage_json)
# -
# End of script.
|
processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An Introduction To `aima-python`
#
# The [aima-python](https://github.com/aimacode/aima-python) repository implements, in Python code, the algorithms in the textbook *[Artificial Intelligence: A Modern Approach](http://aima.cs.berkeley.edu)*. A typical module in the repository has the code for a single chapter in the book, but some modules combine several chapters. See [the index](https://github.com/aimacode/aima-python#index-of-code) if you can't find the algorithm you want. The code in this repository attempts to mirror the pseudocode in the textbook as closely as possible and to stress readability foremost; if you are looking for high-performance code with advanced features, there are other repositories for you. For each module, there are three files, for example:
#
# - [**`logic.py`**](https://github.com/aimacode/aima-python/blob/master/logic.py): Source code with data types and algorithms for fealing with logic; functions have docstrings explaining their use.
# - [**`logic.ipynb`**](https://github.com/aimacode/aima-python/blob/master/logic.ipynb): A notebook like this one; gives more detailed examples and explanations of use.
# - [**`tests/test_logic.py`**](https://github.com/aimacode/aima-python/blob/master/tests/test_logic.py): Test cases, used to verify the code is correct, and also useful to see examples of use.
#
# There is also an [aima-java](https://github.com/aimacode/aima-java) repository, if you prefer Java.
#
# ## What version of Python?
#
# The code is tested in Python [3.4](https://www.python.org/download/releases/3.4.3/) and [3.5](https://www.python.org/downloads/release/python-351/). If you try a different version of Python 3 and find a problem, please report it as an [Issue](https://github.com/aimacode/aima-python/issues). There is an incomplete [legacy branch](https://github.com/aimacode/aima-python/tree/aima3python2) for those who must run in Python 2.
#
# We recommend the [Anaconda](https://www.continuum.io/downloads) distribution of Python 3.5. It comes with additional tools like the powerful IPython interpreter, the Jupyter Notebook and many helpful packages for scientific computing. After installing Anaconda, you will be good to go to run all the code and all the IPython notebooks.
#
# ## IPython notebooks
#
# The IPython notebooks in this repository explain how to use the modules, and give examples of usage.
# You can use them in two ways:
#
# 1. View static HTML pages. (Just browse to the [repository](https://github.com/aimacode/aima-python) and click on a `.ipynb` file link.)
# 2. Run, modify, and re-run code, live. (Download the repository (by [zip file](https://github.com/aimacode/aima-python/archive/master.zip) or by `git` commands), start a Jupyter notebook server with the shell command "`jupyter notebook`" (issued from the directory where the files are), and click on the notebook you want to interact with.)
#
#
# You can [read about notebooks](https://jupyter-notebook-beginner-guide.readthedocs.org/en/latest/) and then [get started](https://nbviewer.jupyter.org/github/jupyter/notebook/blob/master/docs/source/examples/Notebook/Running%20Code.ipynb).
# # Helpful Tips
#
# Most of these notebooks start by importing all the symbols in a module:
from logic import *
# From there, the notebook alternates explanations with examples of use. You can run the examples as they are, and you can modify the code cells (or add new cells) and run your own examples. If you have some really good examples to add, you can make a github pull request.
#
# If you want to see the source code of a function, you can open a browser or editor and see it in another window, or from within the notebook you can use the IPython magic funtion `%psource` (for "print source"):
# %psource WalkSAT
# Or see an abbreviated description of an object with a trainling question mark:
# +
# WalkSAT?
# -
# # Authors
#
# This notebook by [<NAME>](https://github.com/chiragvartak) and [<NAME>](https://github.com/norvig).
|
intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This notebook explores the binary arithmetic error in Python and potential solutions
import sys
import numpy
import pandas
from decimal import *
# creating dummy values and weights
values = 1e-10 * numpy.ones(5)
weights = values/values.sum()
weights.sum()
# try using Decimal. Following this website "https://docs.python.org/2/library/decimal.html",
# I ought to be able to set the precision using getcontext...but that isn't working
weights_list = []
getcontext().prec = 5
Decimal(values[0])
getcontext().prec = 5
Decimal(1)/Decimal(7)
Decimal(1)
# try using Decimal
weights_list = []
values = 1e-10 * numpy.ones(5)
sum_values = values.sum()
for value in values:
getcontext().prec = 5
weight = Decimal(value)/Decimal(sum_values)
weights_list.append(weight)
weights = numpy.array(weights_list)
numpy.float(weights.sum())
|
notebooks/monte_carlo_dev/binary_arithmetic_error.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import numpy.linalg as la
datasets = ['CIFAR', 'MNIST']
net_names = ['ConvBig', 'ConvMed']
perturbations = ['Haze', 'ContrastVariation', 'Rotation']
# +
import re
class Interval:
def __init__(self, interval_str):
m = re.match(r'\[(.*),(.*)\]', interval_str)
self.lower, self.upper = float(m.group(1)), float(m.group(2))
def get_abs_max(self):
return max(abs(self.lower), abs(self.upper))
# +
from statistics import median
NUM_IMAGES = 100
all_data = {}
for dataset in datasets:
all_data[dataset] = {}
for net in net_names:
if dataset == 'MNIST' and net == 'ConvMed':
continue
all_data[dataset][net] = {}
for perturbation in perturbations:
all_data[dataset][net][perturbation] = {}
filename = f'original/results/results_nosplit/{net}_{dataset}_{perturbation}_nosplit.txt'
with open(filename) as f:
content = f.readlines()
content = [x.strip() for x in content]
data = []
for header, arr in zip(content[::2], content[1::2]):
items = header.split(',')
interval_size = float(items[4])
time = float(items[6])
jacobians = np.array(list(map(lambda x: Interval(x).get_abs_max(), arr.split(';')[:-1]))).reshape(NUM_IMAGES, 10)
avg_norm = 0
for jacobi in jacobians:
avg_norm += la.norm(jacobi, np.inf)
avg_norm /= NUM_IMAGES
data.append((interval_size, time, avg_norm))
all_data[dataset][net][perturbation] = data
# +
from statistics import median
NUM_IMAGES = 100
all_data_sound = {}
for dataset in datasets:
all_data_sound[dataset] = {}
for net in net_names:
if dataset == 'MNIST' and net == 'ConvMed':
continue
all_data_sound[dataset][net] = {}
for perturbation in perturbations:
all_data_sound[dataset][net][perturbation] = {}
filename = f'sound/results/results_nosplit/{net}_{dataset}_{perturbation}_nosplit.txt'
with open(filename) as f:
content = f.readlines()
content = [x.strip() for x in content]
data = []
for header, arr in zip(content[::2], content[1::2]):
items = header.split(',')
interval_size = float(items[4])
time = float(items[6])
jacobians = np.array(list(map(lambda x: Interval(x).get_abs_max(), arr.split(';')[:-1]))).reshape(NUM_IMAGES, 10)
avg_norm = 0
for jacobi in jacobians:
avg_norm += la.norm(jacobi, np.inf)
avg_norm /= NUM_IMAGES
data.append((interval_size, time, avg_norm))
all_data_sound[dataset][net][perturbation] = data
# +
perturbations = ['HazeThenRotation', 'ContrastVariationThenRotation', 'ContrastVariationThenHaze']
interval_sizes = np.array([10**(-0.25*k) for k in range(4, 20, 3)]) * 2
from statistics import median
NUM_IMAGES = 10
for dataset in datasets:
for net in net_names:
if net == 'ConvMed' and dataset == 'MNIST':
continue
for perturbation in perturbations:
filename = f'original/results_compose/results_compose_nosplit/{net}_{dataset}_{perturbation}_nosplit.txt'
with open(filename) as f:
content = f.readlines()
content = [x.strip() for x in content]
data = []
for header, arr in zip(content[::2], content[1::2]):
items = header.split(',')
interval_size = float(items[4])
time = float(items[7])
jacobians = np.array(list(map(lambda x: Interval(x).get_abs_max(), arr.split(';')[:-1]))).reshape(NUM_IMAGES, 20)
avg_norm = 0
for jacobi in jacobians:
jacobi = jacobi.reshape(2, 10).T
avg_norm += la.norm(jacobi, np.inf)
avg_norm /= NUM_IMAGES
add = 0
for isize in interval_sizes:
if np.isclose(isize, interval_size):
add = 1
break
if add:
data.append((interval_size, time, avg_norm))
all_data[dataset][net][perturbation] = data
# +
from statistics import median
NUM_IMAGES = 10
for dataset in datasets:
for net in net_names:
if net == 'ConvMed' and dataset == 'MNIST':
continue
for perturbation in perturbations:
filename = f'sound/results_compose/results_compose_nosplit/{net}_{dataset}_{perturbation}_nosplit.txt'
with open(filename) as f:
content = f.readlines()
content = [x.strip() for x in content]
data = []
for header, arr in zip(content[::2], content[1::2]):
items = header.split(',')
interval_size = float(items[4])
time = float(items[7])
jacobians = np.array(list(map(lambda x: Interval(x).get_abs_max(), arr.split(';')[:-1]))).reshape(NUM_IMAGES, 20)
avg_norm = 0
for jacobi in jacobians:
jacobi = jacobi.reshape(2, 10).T
avg_norm += la.norm(jacobi, np.inf)
avg_norm /= NUM_IMAGES
add = 0
for isize in interval_sizes:
if np.isclose(isize, interval_size):
add = 1
break
if add:
data.append((interval_size, time, avg_norm))
all_data_sound[dataset][net][perturbation] = data
# +
from statistics import median as med
table = r"""
\begin{tabular}{c|c|c|c|c|c|c}
& Haze & Contrast & Rotation & Haze-Rotation & Contrast-Rotation & Contrast-Haze \\
\hline
\multirow{2}{*}{MNIST ConvBig} & 0 & 0 & 0 & 0 & 0 & 0 \\
& 0 & 0 & 0 & 0 & 0 & 0 \\
\hline
\multirow{2}{*}{CIFAR ConvMed} & 0 & 0 & 0 & 0 & 0 & 0 \\
& 0 & 0 & 0 & 0 & 0 & 0 \\
\hline
\multirow{2}{*}{CIFAR ConvBig} & 0 & 0 & 0 & 0 & 0 & 0 \\
& 0 & 0 & 0 & 0 & 0 & 0
\end{tabular}"""
table = table.replace('0', 'holder')
perturbations = ['Haze', 'ContrastVariation', 'Rotation', 'HazeThenRotation', 'ContrastVariationThenRotation', 'ContrastVariationThenHaze']
for dataset, net in [('MNIST','ConvBig'), ('CIFAR','ConvMed'), ('CIFAR','ConvBig')]:
# relative error
for perturbation in perturbations:
errors_norm = []
time_overhead = []
for orig, sound in zip(all_data[dataset][net][perturbation], all_data_sound[dataset][net][perturbation]):
errors_norm.append(abs(orig[2] - sound[2]) / abs(orig[2]))
time_overhead.append(round(sound[1] / orig[1], 3))
table = table.replace('holder', f'{max(errors_norm):.2e}', 1)
# time overhead
for perturbation in perturbations:
errors_norm = []
time_overhead = []
for orig, sound in zip(all_data[dataset][net][perturbation], all_data_sound[dataset][net][perturbation]):
errors_norm.append(abs(orig[2] - sound[2]) / abs(orig[2]))
time_overhead.append(round(sound[1] / orig[1], 3))
table = table.replace('holder', f'{round(max(time_overhead), 2)}', 1)
print(table)
# -
|
eval/fp_soundness/.ipynb_checkpoints/Analyze_Soundness-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-rbig]
# language: python
# name: conda-env-.conda-rbig-py
# ---
# # Information Theory Measures w/ RBIG
# +
import sys, os
from pyprojroot import here
# spyder up to find the root
root = here(project_files=[".here"])
# append to path
sys.path.append(str(root))
# %load_ext autoreload
# %autoreload 2
# +
import sys
import numpy as np
from sklearn.utils import check_random_state
import matplotlib.pyplot as plt
import corner
# %load_ext autoreload
# %autoreload 2
# -
# ---
# ## Total Correlation
# +
#Parameters
n_samples = 10000
d_dimensions = 10
seed = 123
rng = check_random_state(seed)
# -
# #### Sample Data
# +
# Generate random normal data
data_original = rng.randn(n_samples, d_dimensions)
# Generate random Data
A = rng.rand(d_dimensions, d_dimensions)
data = data_original @ A
# covariance matrix
C = A.T @ A
vv = np.diag(C)
# -
# #### Calculate Total Correlation
# +
tc_original = np.log(np.sqrt(vv)).sum() - 0.5 * np.log(np.linalg.det(C))
print(f"TC: {tc_original:.4f}")
# -
# ### RBIG - TC
from rbig._src.total_corr import rbig_total_corr
tc_rbig = rbig_total_corr(data, zero_tolerance=30)
print(f"TC (RBIG): {tc_rbig:.4f}")
print(f"TC: {tc_original:.4f}")
# ---
#
# ## Entropy
# #### Sample Data
# +
#Parameters
n_samples = 5000
d_dimensions = 10
seed = 123
rng = check_random_state(seed)
# Generate random normal data
data_original = rng.randn(n_samples, d_dimensions)
# Generate random Data
A = rng.rand(d_dimensions, d_dimensions)
data = data_original @ A
# -
# #### Calculate Entropy
from rbig._src.entropy import entropy_marginal
# +
Hx = entropy_marginal(data)
H_original = Hx.sum() + np.linalg.slogdet(A)[1]
print(f"H: {H_original:.4f}")
# -
print(f"H: {H_original * np.log(2):.4f}")
# ### Entropy RBIG
from rbig._src.entropy import entropy_rbig
H_rbig = entropy_rbig(data, zero_tolerance=30)
print(f"Entropy (RBIG): {H_rbig:.4f}")
print(f"Entropy: {H_original:.4f}")
# ---
# ## Mutual Information
# #### Sample Data
# +
#Parameters
n_samples = 10000
d_dimensions = 10
seed = 123
rng = check_random_state(seed)
# Generate random Data
A = rng.rand(2 * d_dimensions, 2 * d_dimensions)
# Covariance Matrix
C = A @ A.T
mu = np.zeros((2 * d_dimensions))
dat_all = rng.multivariate_normal(mu, C, n_samples)
CX = C[:d_dimensions, :d_dimensions]
CY = C[d_dimensions:, d_dimensions:]
X = dat_all[:, :d_dimensions]
Y = dat_all[:, d_dimensions:]
# -
# #### Calculate Mutual Information
# +
H_X = 0.5 * np.log(2 * np.pi * np.exp(1) * np.abs(np.linalg.det(CX)))
H_Y = 0.5 * np.log(2 * np.pi * np.exp(1) * np.abs(np.linalg.det(CY)))
H = 0.5 * np.log(2 * np.pi * np.exp(1) * np.abs(np.linalg.det(C)))
mi_original = H_X + H_Y - H
mi_original #*= np.log(2)
print(f"MI: {mi_original:.4f}")
# -
# ### RBIG - Mutual Information
from rbig._src.mutual_info import MutualInfoRBIG
# +
# Initialize RBIG class
rbig_model = MutualInfoRBIG()
# fit model to the data
rbig_model.fit(X, Y);
# +
H_rbig = rbig_model.mutual_info()
print(f"MI (RBIG): {H_rbig:.4f}")
print(f"MI: {mi_original:.4f}")
# -
print(f"TC (X) (RBIG): {rbig_model.rbig_model_X.total_correlation():.4f}")
print(f"TC (Y) (RBIG): {rbig_model.rbig_model_Y.total_correlation():.4f}")
print(f"TC (XY_t) (RBIG): {rbig_model.rbig_model_XY.total_correlation():.4f}")
# ---
# ## Kullback-Leibler Divergence (KLD)
# #### Sample Data
# +
#Parameters
n_samples = 10000
d_dimensions = 10
mu = 0.4 # how different the distributions are
seed = 123
rng = check_random_state(seed)
# Generate random Data
A = rng.rand(d_dimensions, d_dimensions)
# covariance matrix
cov = A @ A.T
# Normalize cov mat
cov = A / A.max()
# create covariance matrices for x and y
cov_x = np.eye(d_dimensions)
cov_y = cov_x.copy()
mu_x = np.zeros(d_dimensions) + mu
mu_y = np.zeros(d_dimensions)
# generate multivariate gaussian data
X = rng.multivariate_normal(mu_x, cov_x, n_samples)
Y = rng.multivariate_normal(mu_y, cov_y, n_samples)
# -
# #### Calculate KLD
# +
kld_original = 0.5 * ((mu_y - mu_x) @ np.linalg.inv(cov_y) @ (mu_y - mu_x).T +
np.trace(np.linalg.inv(cov_y) @ cov_x) -
np.log(np.linalg.det(cov_x) / np.linalg.det(cov_y)) - d_dimensions)
print(f'KLD: {kld_original:.4f}')
# -
# ### RBIG - KLD
X.min(), X.max()
Y.min(), Y.max()
# +
# %%time
n_layers = 100000
rotation_type = 'PCA'
random_state = 0
zero_tolerance = 60
tolerance = None
pdf_extension = 10
pdf_resolution = None
verbose = 0
# Initialize RBIG class
kld_rbig_model = RBIGKLD(n_layers=n_layers,
rotation_type=rotation_type,
random_state=random_state,
zero_tolerance=zero_tolerance,
tolerance=tolerance,
pdf_resolution=pdf_resolution,
pdf_extension=pdf_extension,
verbose=verbose)
# fit model to the data
kld_rbig_model.fit(X, Y);
# +
# Save KLD value to data structure
kld_rbig= kld_rbig_model.kld*np.log(2)
print(f'KLD (RBIG): {kld_rbig:.4f}')
print(f'KLD: {kld_original:.4f}')
# -
|
notebooks/information_theory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering NIST headlines and descriptions
#
# adapted from https://github.com/star-is-here/open_data_day_dc
# ## Introduction:
# In this workshop we show you an example of a workflow in data science from initial data ingestion, cleaning, modeling, and ultimately clustering. In this example we scrape the news feed of the National Institute of Standards and Technology ([NIST](www.nist.gov)). For those not in the know, NIST is comprised of multiple research centers which include:
# * Center for Nanoscale Science and Technology (CNST)
# * Engineering Laboratory (EL)
# * Information Technology Laboratory (ITL)
# * NIST Center for Neutron Research (NCNR)
# * Material Measurement Laboratory (MML)
# * Physical Measurement Laboratory (PML)
#
# This makes it an easy target for topic modeling, a way of identifying patterns in a corpus that uses __clustering__.
from lxml import html
import requests
from __future__ import print_function
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cluster import KMeans, MiniBatchKMeans
from time import time
# ## Get the Data
# ### Building the list of headlines and descriptions
#
# We request NIST news based on the following URL, 'http://www.nist.gov/allnews.cfm?s=01-01-2014&e=12-31-2014'. For this workshop, we look at only 2014 news articles posted on the NIST website.
#
# We then pass that retrieved content to our HTML parser and search for a specific div class, "select_portal_module_wrapper" which is assigned to every headline and every description (headlines receive a strong tag and descriptions receive a p tag).
#
# We then merge both the headline and description into one entry in the list because we don't need to differentiate between title and description.
# +
print("Retrieving data from NIST...")
# Retrieve the data from the web page.
page = requests.get('https://www.nist.gov/news-events/news/search?combine=&field_campus_tid=All&term_node_tid_depth_1=All&date_filter%5Bmin%5D%5Bdate%5D=January+01%2C+2016&date_filter%5Bmax%5D%5Bdate%5D=June+30%2C+2016&items_per_page=200')
# Use html module to parse it out and store in tree.
tree = html.fromstring(page.content)
# Create list of news headlines and descriptions.
# This required obtaining the xpath of the elements by examining the web page.
list_of_headlines = tree.xpath('//h3[@class="nist-teaser__title"]/a/text()')
list_of_descriptions = tree.xpath('//div[@class="field-body field--body nist-body nist-teaser__content"]/text()')
#Combine each headline and description into one value in a list
news=[]
for each_headline in list_of_headlines:
for each_description in list_of_descriptions:
news.append(each_headline+each_description)
print("Last item in list retrieved: %s" % news[-1])
# -
# ## Term Frequency-Inverse Document Frequency
#
# - The weight of a term that occurs in a document is proportional to the term frequency.
# - Term frequency is the number of times a term occurs in a document.
# - Inverse document frequency diminishes the weight of terms that occur very frequently in the document set and increases the weight of terms that occur rarely.
#
# 
#
#
# ### Convert collection of documents to TF-IDF matrix
# We now call a TF-IDF vectorizer to create a sparse matrix with term frequency-inverse document frequency weights:
# +
print("Extracting features from the training dataset using a sparse vectorizer")
t0 = time()
# Create a sparse word occurrence frequency matrix of the most frequent words
# with the following parameters:
# Maximum document frequency = half the total documents
# Minimum document frequency = two documents
# Toss out common English stop words.
vectorizer = TfidfVectorizer(input=news, max_df=0.5, min_df=2, stop_words='english')
# This calculates the counts
X = vectorizer.fit_transform(news)
print("done in %fs" % (time() - t0))
print("n_samples: %d, n_features: %d" % X.shape)
print()
# -
# ## Let's do some clustering!
#
# 
#
# I happen to know there are 15 [subject areas](http://www.nist.gov/subject_areas.cfm) at NIST:
# - Bioscience & Health
# - Building and Fire Research
# - Chemistry
# - Electronics & Telecommunications
# - Energy
# - Environment/Climate
# - Information Technology
# - Manufacturing
# - Materials Science
# - Math
# - Nanotechnology
# - Physics
# - Public Safety & Security
# - Quality
# - Transportation
#
# So, why don't we cheat and set the number of clusters to 15?
#
# Then we call the KMeans clustering model from sklearn and set an upper bound to the number of iterations for fitting the data to the model.
#
# Finally we list out each centroid and the top 10 terms associated with each centroid.
# +
# Set the number of clusters to 15
k = 15
# Initialize the kMeans cluster model.
km = KMeans(n_clusters=k, init='k-means++', max_iter=100)
print("Clustering sparse data with %s" % km)
t0 = time()
# Pass the model our sparse matrix with the TF-IDF counts.
km.fit(X)
print("done in %0.3fs" % (time() - t0))
print()
print("Top terms per cluster:")
order_centroids = km.cluster_centers_.argsort()[:, ::-1]
terms = vectorizer.get_feature_names()
for i in range(k):
print("Cluster %d:" % (i+1), end='')
for ind in order_centroids[i, :10]:
print(' %s' % terms[ind], end='')
print()
# -
# ## Questions
# 1. How do the results compare to NIST's listed [subject areas](http://www.nist.gov/subject_areas.cfm)?
#
# 2. How would you operationalize this model?
|
notebook/nist_clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 01-Introduction to Python for general usage
#
# #### This notebook demonstrates the basic programming required for general usage if the chosen language is Python.
# ### Table of Contents
# * [01 - Variables and Datatypes](#variables)
# * [02 - List and Array (denoted by [....])](#lists)
# * [03 - Dictionary (Hash table) (Denoted by {...})](#dict)
# * [04 - Arithmetic Operations](#arithmetic)
# * [05 - Logical operations](#logical-operations)
# * [06 - Conditional Statements](#conditional)
# * [07 - Loop Statements](#loops)
# * [08 - Error Handling using try and except blocks](#error-handling)
# * [09 - Functions](#functions)
#
# ### 01 - Variables, datatypes <a class="anchor" id="variables"></a>
x = 1 # assign value 1 to variable `x`
y = "hello"
z = 3.12
x, y, z = 1, "hello", 3.12 # same as above
type(x), type(y), type(z) # print datatype
# ### 02 - List and Array (denoted by [....]) <a class="anchor" id="lists"></a>
a1 = [1,2,3,4,5] # Homogenous list of Integers. Also called Array. Array is Homogenous.
a2 = [1,2,"hello",2.3456,True] # Heterogenous list. Mix of Integer, String, Float and Boolean
print(a1)
print(a2)
a3 = [i for i in range(10)] # generate a list of int values over a given range
a3
# ##### Get help on any Python function documentation
# get help on any python function by prefixing with `?`. In this example `range`. This only works in Jupyter notebook.
# ?range
a2[2] # access the value in 2nd index of list `a2`
# ### 03 - Dictionary (Hash table) (Denoted by {...}) <a class="anchor" id="dict"></a>
d1 = {"key1": "value1", "key2":"value2"} # dictionary is nothing but 1 or more key:value pairs. In JSON format.
print(d1)
print(type(d1))
d2 = {"k1":"v1","k2":"v2","k3":"v3","k4":"v4","k5":"v5","k6":"v6"}
d2["k3"] # print the value associated with key `k3`
# ### 04 - Arithmetic Operations <a class="anchor" id="arithmetic"></a>
# ##### Arithmentic operations on basic datatypes
a, b = 5, 10 # value assignment
print("Addition:", a+b)
print("Multiplication:", a*b)
print("Subtraction:", a-b)
print("Division:", a/b)
# ##### Arithmetic operations on Lists
a1 = [1,2,3,4,5,"hello"]
a2=[6,7,8,9,10]
print(a1+a2) # add 2 lists
print(a1-a2) # List subtract does not work
# +
# List subtract works as follows
a1 = [1,2,3,4,5]
a2 = [2,3]
a3 = set(a1) - set(a2) # a3 is stored as `set` type. `set` type does not allow duplicates.
print("List a3: ", a3, "Type: ", type(a3))
a3 = list(a3) # Convert set to list type.
print("List a3: ", a3, "Type: ", type(a3))
# -
a1 = [2,5,1,7,2,8]
sorted(a1, reverse=False) # Sort the list in ascending order. To sort in descending, set `reverse=True`.
# ### 05 - Logical operations <a class="anchor" id="logical-operations"></a>
a, b = 5, 10
print(a == b) # check if a is equal to b
print(a != b) # check if a is not equal to b
print(a > b) # check if a is greater than b
print(a >= b) # check if a is greater than or equal to b
print(a < b) # check if a is less than b
# ### 06 - Conditional Statements <a class="anchor" id="conditional"></a>
a, b = 5, 10
if a == b:
print(f"{a} == {b}") # Python f strings: f"...{variable}..."
else:
print(f"{a} != {b}")
print(f"{a} == {b}") if a == b else print(f"{a} != {b}") # same as above cell
# Use logical operations in conditional statements
if a < b:
print(f"{a} is less than {b}")
elif a > b:
print(f"{a} is greater than {b}")
else:
print(f"{a} is equal to {b}")
# ### 07 - Loops <a class="anchor" id="loops"></a>
for i in range(5): # print value of `i` 5 times. Ranging from 0...4
print(i)
l1 = [i for i in range(10)] # prepare a list of values using for loop
l1
# +
a2 = [1,2,"hello",2.3456,True]
# iterate over a list and print all items
for item in a2:
print(item)
# +
d2 = {"k1":"v1","k2":"v2","k3":"v3","k4":"v4","k5":"v5","k6":"v6"}
# iterate over key and value pairs in a dictionary
for key in d2:
print(key, ":", d2[key])
# -
# ### 08 - Error Handling using try and except blocks <a class="anchor" id="error-handling"></a>
print(1/0) # Example of an error such as Divide by Zero
try: # try block. Place potential faulty code in here which is prone to generating errors.
print(1/0)
except Exception as e: # Catch the exception occurred in the try block and treat it appropriately.
print(f"The error is caught. The error is: {e}")
# ### 09 - Functions <a class="anchor" id="functions"></a>
# #### Example function block structure
# def function_name(arg1, arg2, ..., argn, *args <optional>, *kwargs <optional>):
# # write code below
# <code block>
# return # if no return value
# return ret_val1, ret_val2, ..., ret_valn # if n return values
#
# #### Function calls
# Function calls can be made from any location in the code as shown below.
#
# function_name(arg1, arg2, ..., argn)
#
#
# +
# Example of a basic function
def basic_calculator(op_type, operand1, operand2):
if op_type == "add":
result = operand1 + operand2
elif op_type == "sub":
result = operand1 - operand2
elif op_type == "mul":
result = operand1 * operand2
elif op_type == "div":
result = operand1 / operand2
elif op_type == "mod":
result = operand1 % operand2
elif op_type == None or op_type == "":
raise ValueError("op_type is not specified.")
return result
# +
op1, op2 = 10, 20
val = basic_calculator("add", op1, op2) # function call
print(f"Addition of {op1} + {op2} = {val}")
val = basic_calculator(op_type="sub", operand1=op1, operand2=op2) # function call same as above. Can rearrange the argument order if specified arg_name=value.
print(f"Subtraction of {op1} - {op2} = {val}")
val = basic_calculator(operand2=op2, operand1=op1, op_type="mul") # function call. Arguments order rearranged.
print(f"Multiplication of {op1} * {op2} = {val}")
val = basic_calculator("div", op1, op2) # function call
print(f"Division of {op1} / {op2} = {val}")
# -
val = basic_calculator("", op1, op2)
try:
val = basic_calculator("", op1, op2)
except Exception as e:
print(e)
|
python_introduction/01-python-for-data-science.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # ORB feature detector and binary descriptor
#
# This example demonstrates the ORB feature detection and binary description
# algorithm. It uses an oriented FAST detection method and the rotated BRIEF
# descriptors.
#
# Unlike BRIEF, ORB is comparatively scale and rotation invariant while still
# employing the very efficient Hamming distance metric for matching. As such, it
# is preferred for real-time applications.
#
# +
from skimage import data
from skimage import transform
from skimage.feature import (match_descriptors, corner_harris,
corner_peaks, ORB, plot_matches)
from skimage.color import rgb2gray
import matplotlib.pyplot as plt
img1 = rgb2gray(data.astronaut())
img2 = transform.rotate(img1, 180)
tform = transform.AffineTransform(scale=(1.3, 1.1), rotation=0.5,
translation=(0, -200))
img3 = transform.warp(img1, tform)
descriptor_extractor = ORB(n_keypoints=200)
descriptor_extractor.detect_and_extract(img1)
keypoints1 = descriptor_extractor.keypoints
descriptors1 = descriptor_extractor.descriptors
descriptor_extractor.detect_and_extract(img2)
keypoints2 = descriptor_extractor.keypoints
descriptors2 = descriptor_extractor.descriptors
descriptor_extractor.detect_and_extract(img3)
keypoints3 = descriptor_extractor.keypoints
descriptors3 = descriptor_extractor.descriptors
matches12 = match_descriptors(descriptors1, descriptors2, cross_check=True)
matches13 = match_descriptors(descriptors1, descriptors3, cross_check=True)
fig, ax = plt.subplots(nrows=2, ncols=1)
plt.gray()
plot_matches(ax[0], img1, img2, keypoints1, keypoints2, matches12)
ax[0].axis('off')
ax[0].set_title("Original Image vs. Transformed Image")
plot_matches(ax[1], img1, img3, keypoints1, keypoints3, matches13)
ax[1].axis('off')
ax[1].set_title("Original Image vs. Transformed Image")
plt.show()
|
digital-image-processing/notebooks/features_detection/plot_orb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This script is used to make the "WHAM meta files" from the umbrella files.
# The Eric Theide python implementation of EMUS seems to work best with this format
import sys, os, os.path
import glob
import scipy as sp
import numpy as np
from emus import usutils as uu
from emus import emus, avar
import matplotlib
import matplotlib.pyplot as pp
from mpl_toolkits.mplot3d import Axes3D
import yt
from yt.frontends.boxlib.data_structures import AMReXDataset
from tempfile import TemporaryFile
# %pylab inline
# +
# After phi_0 and \kappa data has been extracted using "Extract_Data.ipynb", load data from the folder
# with umbrella files that have had this data removed.
# the block below computes the number of samples in the umbrella files ( assuming they all have the same amount)
location='./umb_files' #data file location. A copy of the data should be used.
list = sorted(os.listdir("./umb_files"))
Number_of_Umbrellas=len(list)
dat = np.loadtxt('./umb_files/umbrella00000000.txt',usecols=[0],unpack=True) #any of the files should work
Number_of_Samples_in_Umb=dat.shape[0]
#load umbrella parameters
centers = np.loadtxt('phi0_centers.txt')
fks = np.loadtxt('spring_constants.txt')
# Additonal EMUS parameters should be set here
period=None
dim=1
T=0.01
k_B=1
# +
#hold umbrella data in 2D array with column index corresponding to umbrella index.
dat_array=numpy.zeros(shape=(Number_of_Samples_in_Umb,Number_of_Umbrellas),dtype=float64)
H_data=numpy.zeros(shape=(Number_of_Samples_in_Umb,Number_of_Umbrellas),dtype=float64)
No_umb_H_data=numpy.zeros(shape=(Number_of_Samples_in_Umb,Number_of_Umbrellas),dtype=float64)
i=0
for filename in list:
a=os.path.join(location, filename)
Temp=np.loadtxt(a)
dat_array[:,i]=Temp[:,0]
H_data[:,i]=Temp[:,2]
No_umb_H_data[:,i]=Temp[:,1]
i=i+1
# Data is then reformatted that it is made compatible with EMUS struvture
# This follows the format of their example in their github repo
# Essentially, every first index should correspond to all the data in that umbrella
cv_data=numpy.zeros(shape=(Number_of_Umbrellas,Number_of_Samples_in_Umb,),dtype=float64)
Hamiltonian=numpy.zeros(shape=(Number_of_Umbrellas,Number_of_Samples_in_Umb,),dtype=float64)
Ham_no_umb=numpy.zeros(shape=(Number_of_Umbrellas,Number_of_Samples_in_Umb,),dtype=float64)
for i in range (0,Number_of_Umbrellas):
cv_data[i]=dat_array[0:Number_of_Samples_in_Umb,i]
Hamiltonian[i]=H_data[0:Number_of_Samples_in_Umb,i]
Ham_no_umb[i]=No_umb_H_data[0:Number_of_Samples_in_Umb,i]
list2=[0]*(len(list))
i=0
for filename in list:
list2[i]=os.path.join(location, filename)
i=i+1
cv_traj=(cv_data)
list2=np.asarray(list2)
# +
# A folder in the "WHAM meta file" format will be made
# This is a format that is standard for chemistry people
# EMUS is easier to work with in this format
# See WHAM documentation of grossfield for a precise description
# Switch location to a new copy of original data file with title denoting the WHAM format
location='./umb_files_Wham_Format' #data file location
list = sorted(os.listdir("./umb_files_Wham_Format"))
list3=[0]*(len(list))
i=0
for filename in list:
list3[i]=os.path.join(location, filename)
i=i+1
## code below is for removing the last two THREE of the data files(none phi average part).
## ONLY RUN ONCE on the NON-original data
# temp=np.zeros(Number_of_Samples_in_Umb)
# for filename in list3:
# with open(filename, 'r') as fin:
# data = fin.read().splitlines(True)
# for j in range (0,Number_of_Samples_in_Umb):
# temp[j]=float(data[j][0:20])
# with open(filename, 'w') as fout:
# np.savetxt(fout, temp, fmt="%10.14f",delimiter='\t')
# +
## Add time column to trimmed data set.
##*****RUN ONCE******#
# DT=2.4414e-5 # time step used in run
# DT_vec=np.arange(Number_of_Samples_in_Umb)+1
# DT_vec=DT_vec*DT # create a vector of these timesteps (i.e dt,2*dt,3*dt,...)
# # Add time step column to data as a column before the phi averages column
# for filename in list3:
# with open(filename, 'r') as fin:
# data = fin.read().splitlines(True)
# data=np.asarray(data)
# Place_holder = np.zeros(data.size, dtype=[('var1', float64), ('var2', float64)])
# Place_holder['var1']=DT_vec
# Place_holder['var2']=data
# np.savetxt(filename, Place_holder, fmt="%10.12f %10.12f",delimiter='\t')
# +
# Now we find the "middle" of the data where our loop is made.
# The location prints to screen
# Depending on the where the "turning point" of the average data is, find the max or min
A=np.where(centers == centers.min()) # find min since we go from 1.0 to 0.74 to 1.0
# np.where(centers == centers.min())
A[0]
# -
# SET middle location index manually here
middle=171
# +
# Overwrite copy of umbrella file with time step and phi average with WHAM format of data
#In this case we consider data with 2 pars. The 1.0 to 0.74 trip, and the 0.74 to 1.0 trip.
# We make two meta files for this
#part 1 (1.0 to 0.74)
list4=np.asarray(list3[0:middle+1])
MAKE_WHAM_META_ARR = np.zeros(list4.size, dtype=[('var1', 'U60'), ('var2', float64), ('var3', float64)])
MAKE_WHAM_META_ARR['var1']=list4[0:middle+1]
MAKE_WHAM_META_ARR['var2']=centers[0:middle+1]
MAKE_WHAM_META_ARR['var3']=fks[0:middle+1]
np.savetxt('ONE_TO_074_META.txt', MAKE_WHAM_META_ARR, fmt="%60s %10.8f %10.8f",delimiter='\t')
# +
#part 2 (0.74 to 1.0)
list5=np.asarray(list3[middle+1:])
MAKE_WHAM_META_ARR_2 = np.zeros(list5.size, dtype=[('var1', 'U60'), ('var2', float64), ('var3', float64)])
MAKE_WHAM_META_ARR_2['var1']=list5
MAKE_WHAM_META_ARR_2['var2']=centers[middle+1:]
MAKE_WHAM_META_ARR_2['var3']=fks[middle+1:]
np.savetxt('074_TO_ONE_META.txt', MAKE_WHAM_META_ARR_2, fmt="%60s %10.8f %10.8f",delimiter='\t')
# +
meta_file = 'ONE_TO_074_META.txt' # Path to Meta File
psis, cv_trajs, neighbors = uu.data_from_meta(
meta_file, dim, T=T, k_B=k_B, period=period)
meta_file = '074_TO_ONE_META.txt' # Path to Meta File
psis, cv_trajs, neighbors = uu.data_from_meta(
meta_file, dim, T=T, k_B=k_B, period=period)
|
unmaintained/_GL_alt/Python_notebooks_KL_new/EMUS_scripts/Make_meta_files(step2).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#define function that scrapes one page at a time and returns a panda dataframe
def scrape_page(page_num):
import requests # b_soup_1.py
import re
from bs4 import BeautifulSoup
#spoof the user agent to make page scrapable
headers = {'User-Agent': 'Mozilla/5.0 (Macintosh; Intel Mac OS X 10_10_1) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/39.0.2171.95 Safari/537.36'}
#define position search term (one word) and location
position='analyst'
location='Pittsburgh,+PA'
page_str=str(page_num)
#url for search query
url='https://www.indeed.com/jobs?q='+position+'&l='+location+'&start='+page_str
search_page=requests.get(url,headers=headers)
search_page
soup=BeautifulSoup(search_page.content,'html.parser')
all_cards=soup.find_all(class_='jobsearch-SerpJobCard row result')
#empty list to store job posting urls
job_urls=[]
#it seems to be easier to pull posting dates and locations from search page, so do that here
posting_dates=[]
locations=[]
#iterate through all_cards, extract job posting urls, posting dates, and locations
for card in all_cards:
#urls
job_id=card['data-jk']
job_url='https://www.indeed.com/viewjob?jk='+job_id
job_urls.append(job_url)
#posting dates
#some jobs don't have posting dates so try them
try:
job_post_date=card.find(class_='date').get_text()
except:
job_post_date=' '
posting_dates.append(job_post_date)
#locations
job_location=card.find(class_='location').get_text()
locations.append(job_location)
#locations
#lets make this dataframe
#declare the empty lists that will become columns
position_name=[]
employer=[]
full_description=[]
#in the future, we can develop a list of keywords that we'll
#search postings for, but for now, just return all capitalized
#words besides common words
capital_words=[]
for job_url in job_urls:
#retrieve job page
job_page=requests.get(job_url,headers=headers)
#make job page into BeautifulSoup object
job_soup=BeautifulSoup(job_page.content,'html.parser')
#retrieve the job title
job_position=job_soup.find(class_='icl-u-xs-mb--xs').get_text()
#retrieve company name from job rating line on page
job_rating_line=job_soup.find(class_='jobsearch-InlineCompanyRating')
job_employer=job_rating_line.find(class_='icl-u-lg-mr--sm').get_text()
#retrieve full job description
job_descrip=job_soup.find(class_='jobsearch-JobComponent-description').get_text()
#replace line breaks in description with spaces
job_descrip.replace('\n',' ')
#find capitalized words in job description
capitals=re.findall('([A-Z][a-z]+)', job_descrip)
#replace commonly capitalized words
for x in ['The','Our','A','As','On','Using','We', 'In', 'Some']:
try:
capitals[:]=(value for value in capitals if value!=x)
except:
capitals
#make capitals into a set instead of a list
capitals=set(capitals)
capital_string=', '.join(capitals)
#append all of the items created to the empty lists from last step
position_name.append(job_position)
employer.append(job_employer)
full_description.append(job_descrip)
capital_words.append(capitals)
#store results in dictionary, then data frame
import pandas as pd
result_dict={'Job Title':position_name,
'Employer':employer,
'Location':locations,
'Posting Date':posting_dates,
'Posting Url':job_urls,
'Full Job Description':full_description,
'Keywords':capital_words
}
result_frame=pd.DataFrame(result_dict)
return(result_frame)
# -
|
Prototype 2_5/.ipynb_checkpoints/scrape page as function-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
data = []
data.append({'name':'Coleoptera','trank':'order','lineage':'Animalia|Arthropoda|Insecta|Coleoptera'})
data.append({'name':'Curculionidae','trank':'family','lineage':'Animalia|Arthropoda|Insecta|Coleoptera|Curculionidae'})
data.append({'name':'Trigonops','trank':'genus','lineage':'Animalia|Arthropoda|Insecta|Coleoptera|Curculionidae|Trigonops'})
data.append({'name':'Chrysomelidae','trank':'family','lineage':'Animalia|Arthropoda|Insecta|Coleoptera|Chrysomelidae'})
data.append({'name':'Phytorus','trank':'genus','lineage':'Animalia|Arthropoda|Insecta|Coleoptera|Chrysomelidae|Phytorus'})
data
level_dict = {'kingdom': 1, 'phylum': 2, 'class': 3, 'order': 3, 'family': 4, 'genus': 5, 'species': 6}
# +
for d in data:
# -
target = {t: {'name': 'Coleoptera', 'trank': 'order'} }
# + active=""
# $('#using_json_2').jstree({ 'core' : {
# 'data' : [
# { "id" : "ajson1", "parent" : "#", "text" : "Simple root node" },
# { "id" : "ajson2", "parent" : "#", "text" : "Root node 2" },
# { "id" : "ajson3", "parent" : "ajson2", "text" : "Child 1" },
# { "id" : "ajson4", "parent" : "ajson2", "text" : "Child 2" },
# ]
# } });
|
Untitled1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import random
2000/120
random.expovariate(1/40)
# +
epochs = []
for i in range(2000):
epochs.append(random.expovariate(2000/120))
# -
epochs
sum(epochs)/2000
plt.plot(epochs)
plt.show()
def next_time(rate):
return
# +
interarrival=[]
while sum(interarrival) < 120:
interarrival.append(random.expovariate(55.64166))
print(len(interarrival))
# -
random.expovariate(55.641)
|
PoissonProcesses.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
#
# <a href="https://www.quantrocket.com/disclaimer/">Disclaimer</a>
# # End-of-Day Analysis with Alphalens
#
# Alphalens is an open-source performance analysis library which pairs well with the Pipeline API. In this notebook we will use Alphalens to analyze whether our momentum factor is predictive of forward returns.
#
# > Using Alphalens makes sense when you believe your end-of-day Pipeline rules have alpha. In contrast, if your Pipeline rules simply perform a basic screen and the alpha is entirely provided by your intraday trading rules, it might make more sense to omit this step.
# Let's re-run our pipeline from the previous notebook:
# +
from zipline.pipeline import Pipeline
from zipline.pipeline.factors import AverageDollarVolume, Returns
from zipline.research import run_pipeline
pipeline = Pipeline(
columns={
"1y_returns": Returns(window_length=252),
},
screen=AverageDollarVolume(window_length=30) > 10e6
)
factors = run_pipeline(pipeline, start_date="2017-01-01", end_date="2019-01-01")
# -
# To see if our momentum factor is predictive of forward returns, we use the factor data to request forward returns for the corresponding assets and dates, then format the factor and returns data for use with Alphalens:
# +
from zipline.research import get_forward_returns
import alphalens as al
# Get forward returns (this provides forward 1-day returns by default)
forward_returns = get_forward_returns(factors)
# format the data for Alphalens
al_data = al.utils.get_clean_factor(
factors["1y_returns"],
forward_returns,
quantiles=2 # For a very small sample universe, you might only want 2 quantiles
)
# -
# Then we create a tear sheet to look at the factor. For a predictive factor, the higher quantiles should perform better than the lower quantiles.
from alphalens.tears import create_full_tear_sheet
create_full_tear_sheet(al_data)
# ***
#
# ## *Next Up*
#
# Part 4: [Intraday Trading Rules](Part4-Intraday-Trading-Rules.ipynb)
|
intro_zipline/Part3-End-of-Day-Analysis-With-Alphalens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/logo.jpg" style="display: block; margin-left: auto; margin-right: auto;" alt="לוגו של מיזם לימוד הפייתון. נחש מצויר בצבעי צהוב וכחול, הנע בין האותיות של שם הקורס: לומדים פייתון. הסלוגן המופיע מעל לשם הקורס הוא מיזם חינמי ללימוד תכנות בעברית.">
# # <span style="text-align: right; direction: rtl; float: right;">Unpacking</span>
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">הקדמה</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הנה שורה שתיראה לנו קצת שונה תחבירית מכל מה שהכרנו עד עכשיו:
# </p>
country, population = ('Israel', 8712000)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בצד ימין יש tuple שמכיל שני איברים, ואנחנו מבצעים השמה לשני משתנים ששמותיהם <var>country</var> ו־<var>population</var>.<br>
# אומנם לא הייתי מעניק לפיסת הקוד הזו את תואר פיסת הקוד הנאה בעולם, אבל נראה שפייתון יודעת להריץ אותה:<br>
# </p>
print(f"There are {population} people living in {country}.")
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# חזינו כאן במעין השמה כפולה שפייתון מאפשרת לנו לבצע:<br>
# את האיבר הראשון בצד ימין היא הכניסה לאיבר הראשון בצד שמאל, ואת האיבר השני בצד ימין היא הכניסה לאיבר השני בצד שמאל.<br>
# </p>
# <img src="images/unpacking.svg?v=3" style="display: block; margin-left: auto; margin-right: auto;" alt="ארבע ריבועים, שניים בימין בצבע אפור ושניים בשמאל בצבע כתום. בין הריבועים האפורים לכתומים מפריד סימן שווה. בתוך הריבוע השמאלי ביותר כתוב country, ואליו יוצא חץ מהריבוע האפור, השלישי משמאל, שבו כתוב Israel. אל הריבוע הכתום השני, שממוקם כריבוע השני משמאל ובו כתוב population, יוצא חץ מהריבוע האפור, הממוקם כרביעי משמאל, שבו כתוב 8712000. בין הריבועים האפורים יש פסיק, וכך גם בין הריבועים הכתומים.">
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# התחביר שמאפשר לפייתון לפרק מבנה ולבצע השמה של האיברים שבו לכמה שמות משתנים בו־זמנית, קיבל את השם <dfn>unpacking</dfn>.
# </p>
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">Unpacking לתוך משתנים</span>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">המקרה הקלאסי</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מתכנתים רבים מנצלים את הנוחות שב־unpacking, ולכן תמצאו שהוא מופיע פעמים רבות בקוד "בעולם האמיתי".<br>
# אפשר לבצע השמה לכמה משתנים שנרצה בו־זמנית:
# </p>
a, b, c, d, e = (1, 2, 3, 4, 5)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הטכניקה הזו עובדת גם עבור טיפוסים אחרים שמוגדרים כ־iterable:
# </p>
a, b = [1, 2]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ואפילו בלי סוגריים (זה לא קסם – בצד ימין נוצר בפועל tuple):
# </p>
a, b = 1, 2
print(type(a))
print(b)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# או כ"חילוץ איברים" מתוך משתנה קיים לתוך כמה משתנים נפרדים:
# </p>
point_on_map = (36.672011, 65.807761)
x, y = point_on_map
print(f"The treasure should be in ({x}, {y}).")
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בדוגמה האחרונה יצרנו בשורה הראשונה משתנה שמצביע ל־tuple. ב־tuple ישנם שני מספרים מסוג float.<br>
# בשורה השנייה פירקנו את ה־tuple – הערך הראשון שלו הוכנס לתוך המשתנה <var>x</var> והערך השני שלו הוכנס לתוך המשתנה <var>y</var>.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">unpacking בלולאות for</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שימוש מקובל מאוד ל־unpacking, שאותו כבר הספקתם לראות בעבר, מתרחש בלולאות for.<br>
# ניצור רשימה של tuple־ים, שבה כל tuple ייצג מדינה ואת מספר האנשים החיים בה:
# </p>
countries_with_population = [
('Cyprus', 1198575),
('Eswatini', 1148130),
('Djibouti', 973560),
('Fiji', 889953),
]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בעולם ללא unpacking, היינו צריכים לכתוב כך:
# </p>
for item in countries_with_population:
country = item[0]
population = item[1]
print(f"There are {population:9,} people in {country}.")
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אבל כיוון שאנחנו חיים בעולם טוב יותר, פייתון מאפשרת לנו לעשות את הטריק הבא:
# </p>
for country, population in countries_with_population:
print(f"There are {population:9,} people in {country}.")
# <table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
# <caption style="direction: rtl; text-align: center;">תצוגה של המשתנה <var>countries_with_population</var> ושל צורת הפירוק שלו</caption>
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">3</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"Cyprus"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">1198575</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"Eswatini"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">1148130</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"Djibouti"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">973560</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid;">
# <table style="font-size: 1.5rem; border: 0px solid black; border-spacing: 0px;">
# <tr>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-bottom: 1px solid;">0</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: left; border-left: 1px solid #555555; border-bottom: 1px solid;">1</td>
# </tr>
# <tbody>
# <tr>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #FF8578;">"Fiji"</td>
# <td style="padding-top: 8px; padding-bottom: 8px; padding-left: 10px; padding-right: 10px; vertical-align: bottom; border: 2px solid; background-color: #98FB98;">889953</td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# </td>
# </tr>
# <tr style="background: #f5f5f5;">
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right;">-4</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-3</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-2</td>
# <td style="padding-left: 4px; padding-top: 2px; padding-bottom: 3px; font-size: 1.3rem; color: #777; text-align: right; border-left: 1px solid #555555;">-1</td>
# </tr>
# </tbody>
# </table>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# באיור למעלה, התאים האדומים מפורקים לתוך המשתנה <var>country</var> והתאים הירוקים לתוך המשתנה <var>population</var>.<br>
# כל איטרציה של הלולאה תגרום לפירוק של צמד אחד מתוך הרשימה, ולהשמתו בהתאמה לתוך צמד המשתנים שבראש לולאת ה־for.
# </p>
# #### <span style="text-align: right; direction: rtl; float: right; clear: both;">במילונים</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הרעיון של unpacking נעשה שימושי מאוד כשאנחנו רוצים לעבור בו־זמנית על המפתח ועל הערך של מילון.<br>
# כל שנצטרך לעשות כדי לקבל את המפתח לצד הערך בכל איטרציה, זה להשתמש בפעולה <code>items</code> על המילון:
# </p>
countries_with_population = {
'Cyprus': 1198575,
'Eswatini': 1148130,
'Djibouti': 973560,
'Fiji': 889953,
}
for country, population in countries_with_population.items():
print(f"There are {population:9,} people in {country}.")
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">unpacking לערכי חזרה מפונקציה</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# המקרה הזה לא שונה מהמקרים הקודמים שראינו, אבל חשבתי שיהיה נחמד לראות אותו מפורשות.<br>
# הפעם ניקח פונקציה שמחזירה tuple, ונשתמש ב־unpacking כדי לפרק את הערכים שהיא מחזירה למשתנים.<br>
# </p>
def division_and_modulo(number, divisor):
division = number // divisor
modulo = number % divisor
return (division, modulo) # אפשר גם בלי הסוגריים
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נשתמש בפונקציה:
# </p>
division_and_modulo(5, 2)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# והרי שאם מוחזר לנו tuple, אפשר לעשות לו unpacking:
# </p>
div, mod = division_and_modulo(5, 2)
print(f"division: {div}, modulo: {mod}")
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">Unpacking לארגומנטים</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נבחן את הקוד הפשוט הבא:<br>
# </p>
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
treasure_location = (36.671111, 65.808056)
print_treasure_location(treasure_location[0], treasure_location[1])
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הגדרנו פונקציה שמדפיסה לנו יפה מיקומים לפי x ו־y שהיא מקבלת.<br>
# המימוש הגיוני, אבל אוי א בראך! השימוש לא מאוד נוח!<br>
# בכל פעם אנחנו צריכים לפרק את ה־tuple שמכיל את המיקום ל־2 איברים, ולשלוח כל אחד מהם בנפרד.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">unpacking לארגומנטים לפי מיקום</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כפתרון למצב, פייתון מאפשרת לנו לעשות את הקסם הנפלא הבא:
# </p>
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
treasure_location = (36.671111, 65.808056)
print_treasure_location(*treasure_location)
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נראה מוזר? זו לא טעות, זהו באמת תחביר שעדיין לא ראינו!<br>
# הכוכבית מפרקת את ה־tuple שהגדרנו, <var>treasure_location</var>, ושולחת לארגומנט <var>x</var> את הערך הראשון ולארגומנט <var>y</var> את הערך השני.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם היו לנו ערכים רבים, היינו יכולים להשתמש באותו טריק בלולאה:
# </p>
treasure_locations = [
(36.671111, 65.808056),
(53.759748, -2.648121),
(52.333333, 1.183333),
(52.655278, -1.906667),
]
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
for treasure_location in treasure_locations:
print_treasure_location(*treasure_location)
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אבל נזכור שאנחנו יכולים להשתמש גם בתעלול שלמדנו על unpacking בתוך for:
# </p>
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
for treasure_x, treasure_y in treasure_locations:
print_treasure_location(treasure_x, treasure_y)
# -
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">unpacking לארגומנטים לפי שם הארגומנט</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מצוידים בפונקציה להדפסת המיקום, אנחנו מקבלים את רשימת המילונים הבאה:
# </p>
treasure_maps = [
{'x': 36.671111, 'y': 65.808056},
{'x': 53.759748, 'y': -2.648121},
{'x': 52.333333, 'y': 1.183333},
{'x': 52.655278, 'y': -1.906667}
]
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אין מנוס, דנו אותנו לחיי עבדות של פירוק מילונים ולשליחת ערכיהם לפונקציות! 😢
# </p>
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
for location in treasure_maps:
print_treasure_location(location.get('x'), location.get('y'))
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# או שאולי לא?
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# במקרה המיוחד מאוד של מילון, אפשר לשלוח לפונקציה את הפרמטרים בעזרת unpacking עם שתי כוכביות.<br>
# המפתחות של המילון צריכים לציין את שם הפרמטרים של הפונקציה, והערכים במילון יהיו הערך שיועבר לאותם פרמטרים:
# </p>
# +
treasure_maps = [
{'x': 36.671111, 'y': 65.808056},
{'x': 53.759748, 'y': -2.648121},
{'x': 52.333333, 'y': 1.183333},
{'x': 52.655278, 'y': -1.906667}
]
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
for location in treasure_maps:
print_treasure_location(**location)
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מה קרה פה?<br>
# באיטרציה הראשונה <var>location</var> היה <code>{'x': 36.671111, 'y': 65.808056}</code>.<br>
# הפונקציה <code>print_treasure_locations</code> מחכה שיעבירו לה ערך לפרמטר <var>x</var> ולפרמטר <var>y</var>.<br>
# ה־unpacking שעשינו בעזרת שתי הכוכביות העביר את הערך של המפתח <code>'x'</code> במילון לפרמטר <var>x</var>, ואת הערך של המפתח <code>'y'</code> במילון לפרמטר <var>y</var>.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נראה דוגמה נוספת לפונקציה שמקבלת שנה, חודש ויום ומחזירה לנו תאריך כמחרוזת:
# </p>
# +
def stringify_date(year, month, day):
return f'{year}-{month}-{day}'
date = {'year': 1815, 'month': 12, 'day': 10}
print(stringify_date(**date))
# -
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">שגיאות</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אנחנו רגילים שפייתון די סובלנית, אבל על unpacking תקין היא לא מוותרת.<br>
# בדוגמה הבאה אנחנו מנסים לחלץ שני איברים לתוך שלושה משתנים, וזה לא נגמר טוב:
# </p>
a, b, c = (1, 2)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נקבל שגיאה דומה, אך שונה, כשננסה לחלץ מספר לא נכון של ארגומנטים לתוך פונקציה:
# </p>
# +
def print_treasure_location(x, y):
print(f"{x}°N, {y}°E")
location_3d = (36.671111, 65.808056, 63.124592)
print_treasure_location(*location_3d)
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם ננסה לעשות unpacking לאיבר שאינו iterable, תתקבל השגיאה הבאה:
# </p>
a, b = 5
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">קוד לדוגמה</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# סדרת פיבונאצ'י היא סדרה שמתחילה באיברים 1 ו־1, וכל איבר בה הוא סכום שני האיברים הקודמים לו.<br>
# האיברים הראשונים בסדרה, הם (מימין לשמאל) 1, 1, 2, 3, 5, 8, וכך הסדרה ממשיכה.<br>
# במימוש פונקציה שמקבלת מספר ומחזירה את סכום כל איברי הסדרה עד אותו מספר, נוכל להשתמש ב־unpacking כדי לשפר את הקריאות של הפונקציה:
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הפונקציה בלי unpacking:
# </p>
def fibonacci_no_unpacking(number):
a = 1
b = 1
total = 0
while a <= number:
total = total + a
temp = a
a = b
b = temp + b
return total
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הפונקציה עם unpacking:
# </p>
# +
def fibonacci_sum(number):
a, b = 1, 1
total = 0
while a <= number:
total = total + a
a, b = b, a + b
return total
fibonacci_sum(8)
# -
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">תרגילים</span>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">אליבי לרוצחים</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לפניכם tuple המכיל כמה מילונים, כאשר כל מילון מייצג דמות חשודה ברצח.<br>
# בתוך כל אחד מהמילונים, תחת המפתח <em>evidences</em>, ישנו tuple שבו שני איברים.<br>
# האיבר הראשון הוא הנשק שתפסה המשטרה, והאיבר השני הוא המיקום המרכזי שבו הייתה הדמות באותו היום.<br>
# בהינתן שהרוצח השתמש באקדח דרינגר (derringer) ב־Petersen House, הדפיסו רק את שמות האנשים שעדיין חשודים ברצח.<br>
# השתדלו להשתמש ב־unpacking לפחות פעמיים.
# </p>
suspects = (
{'name': 'Anne', 'evidences': ('derringer', 'Caesarea')},
{'name': 'Taotao', 'evidences': ('derringer', 'Petersen House')},
{'name': 'Pilpelet', 'evidences': ('Master Sword', 'Hyrule')},
)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לנוחיותכם, הנה פונקציה שמקבלת כלי נשק ומיקום, ובודקת אם הראיות תואמות:
# </p>
def check_evidences(weapon, location):
return weapon.lower() == 'derringer' and location.lower() == 'petersen house'
# +
def print_suspects(suspects):
for suspect in suspects:
if check_evidences(*suspect['evidences']):
print (suspect['name'])
print_suspects(suspects)
|
week4/4_Unpacking.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WnAa-QR1oeL4" colab_type="text"
# # Imports
# + id="d6NRfj76ojE8" colab_type="code" colab={}
import numpy as np
import tensorflow as tf
import keras
from keras.preprocessing.image import img_to_array, load_img
from keras.applications.inception_resnet_v2 import InceptionResNetV2, decode_predictions, preprocess_input
from IPython.core.display import display
from keras.applications.vgg19 import VGG19
from keras.applications.vgg19 import preprocess_input as preprocess_input_vgg19
from keras.applications.vgg19 import decode_predictions as decode_vgg19
# + [markdown] id="hinQ_rUorEsO" colab_type="text"
# # Constants
# + id="hC6dpJAhpSta" colab_type="code" colab={}
FILE_1 = '01 Umbrella.jpg'
FILE_2 = '02 Couple.jpg'
FILE_3 = '03 Ocean.jpg'
# + [markdown] id="sEuNV70OrSyD" colab_type="text"
# # Preprocessing Images
# + id="l_1UMxiYoEcq" colab_type="code" outputId="ae0e39ce-1903-4c22-de55-f436c9f3ad1b" colab={"base_uri": "https://localhost:8080/", "height": 316}
pic = load_img(FILE_1, target_size=(299, 299))
display(pic)
# + id="U3UxVTLCug_5" colab_type="code" outputId="d8d30f15-f784-4d78-bece-836f50f85b8d" colab={"base_uri": "https://localhost:8080/", "height": 34}
pic_array = img_to_array(pic)
pic_array.shape
# + id="KRpYb_1zc89Z" colab_type="code" outputId="9cc32c30-9636-4d21-dcb6-d0f857ec1fac" colab={"base_uri": "https://localhost:8080/", "height": 34}
expanded = np.expand_dims(pic_array, axis=0)
expanded.shape
# + id="iAoyAFSPdhkO" colab_type="code" colab={}
preprocessed = preprocess_input(expanded)
# + [markdown] id="m2gD7R4JhWSO" colab_type="text"
# **Challenge:** Create a function called ```format_img_inceptionresnet()``` that takes a filename as an argument. The function needs to load the image in the default resolution for InceptionResNetv2, convert the image to an array and return the preprocessed image for the InceptionResNetv2 model.
# + id="v5rRYOWjjFrS" colab_type="code" colab={}
def format_img_inceptionresnet(filename):
pic = load_img(filename, target_size=(299,299))
pic_arr = img_to_array(pic)
expanded = np.expand_dims(pic_arr, axis=0)
return preprocess_input(expanded)
# + id="CkYkVjFtqpIG" colab_type="code" colab={}
def format_img_vgg19(filename):
pic = load_img(filename, target_size=(224,224))
pic_arr = img_to_array(pic)
# expanded = np.expand_dims(pic_arr, axis=0)
expanded = pic_arr.reshape(1, pic_arr.shape[0], pic_arr.shape[1], pic_arr.shape[2])
return preprocess_input_vgg19(expanded)
# + [markdown] id="hILeQtmTzyWv" colab_type="text"
# # Load InceptionResNet
# + id="qrug1XPDuuNC" colab_type="code" outputId="439b7875-fac5-4237-b4a9-e27d43840e7b" colab={"base_uri": "https://localhost:8080/", "height": 154}
# %%time
inception_model = InceptionResNetV2(weights='imagenet')
# + id="1vY-iNec0Ncg" colab_type="code" colab={}
inception_model.graph = tf.get_default_graph()
# + [markdown] id="SPiAWaQ1bXWT" colab_type="text"
# # Making Predictions
# + id="8ck3IK6T161p" colab_type="code" outputId="d56aff33-0b04-42a8-e7a1-282e4ab67d84" colab={"base_uri": "https://localhost:8080/", "height": 101}
prediction = inception_model.predict(preprocessed)
decode_predictions(prediction)
# + id="nPL0u9c1b79H" colab_type="code" outputId="1df4349a-9080-4e57-e931-db2e0958b341" colab={"base_uri": "https://localhost:8080/", "height": 400}
data = format_img_inceptionresnet('04 Horse.jpg')
prediction = inception_model.predict(data)
display(load_img('04 Horse.jpg'))
decode_predictions(prediction)
# + [markdown] id="iuLK-fUCpN8Y" colab_type="text"
# # Testing the VGG19 Model
# + [markdown] id="Zbdn5KVKpf0u" colab_type="text"
# **Challenge:** Use the VGG19 Model from Keras with the ImageNet weights to make a prediction on several of the sample images. Load the model into the notebook. Process the data for VGG19. Then make a prediction. Look at the documentation for hints.
# + id="wfUQ5j1Xpe0c" colab_type="code" outputId="2c4dab92-d3b1-46da-f3a2-a7ca32b54f19" colab={"base_uri": "https://localhost:8080/", "height": 70}
vgg19_model = VGG19()
# + id="q2FPrzeMkSJk" colab_type="code" outputId="7a31dadd-372f-467e-e490-d2ef48d5e95e" colab={"base_uri": "https://localhost:8080/", "height": 357}
data = format_img_vgg19(FILE_3)
pred = vgg19_model.predict(data)
display(load_img(FILE_3))
decode_vgg19(pred)
# + id="WRZsFAn-wYwM" colab_type="code" outputId="f90e6e3c-4a44-4267-826b-0f58a0ab2666" colab={"base_uri": "https://localhost:8080/", "height": 400}
data = format_img_vgg19('04 Horse.jpg')
pred = vgg19_model.predict(data)
display(load_img('04 Horse.jpg'))
decode_vgg19(pred)
# + id="BNzr8tiwxS9b" colab_type="code" colab={}
|
src/09_Section_9_Introduction_to_Neural_Networks_and_How_to_Use_Pre-Trained_Models/02_Download_the_Complete_Notebook_Here/09 Neural Nets Pretrained Image Classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generate C2 correction from anti-Stokes and Stokes Raman intensity ratios.
#
# Example based on vibrational transitions in CCl4, C6H6 and C6H12
#
# ---
#
# In this scheme, the known temperature is used to compute the true(or reference) intensity ratio of the anti-Stokes and Stokes bands.
# Experimental band areas are loaded as numpy arrays. Initial coefs are used to model the wavelength-dependent sensitivity (which is unity at zero at 0 cm-1). Coefs of the polynomial are optimized in least-squares scheme to obtain smallest difference in the true and experimental intensity ratios.
import numpy as np
import genC2_antiStokes_Stokes
print(genC2_antiStokes_Stokes.__doc__)
init_coef_linear = np.zeros(1)
init_coef_linear[0] = 0.0
genC2_antiStokes_Stokes.residual_linear(init_coef_linear)
genC2_antiStokes_Stokes.run_fit_linear ( 0.95 )
# ---
#
init_coef_quadratic = np.zeros(2)
init_coef_quadratic[0] = 0.5
init_coef_quadratic[1] = 0.25
genC2_antiStokes_Stokes.run_fit_quadratic ( 1.0, -0.0025 )
|
PythonModule/determine_C2/vibrationalRaman_liquids/antiStokes_Stokes_ratios/example/example_antiStokes_Stokes_Raman_intensities.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# # **强化学习——Deep Deterministic Policy Gradient (DDPG)**
# **作者:**:[EastSmith](https://github.com/EastSmith)
#
# **日期:** 2021.11
#
# **AI Studio项目**:[点击体验](https://aistudio.baidu.com/aistudio/projectdetail/1702021)
#
# ## **一、介绍**
#
# ### 深度确定策略梯度(Deep Deterministic Policy Gradient,DDPG)
# * 它是一种学习连续动作的无模型策略算法。
# * 它结合了DPG(确定性策略梯度)和DQN(深度Q网络)的思想。它利用DQN中的经验重放和延迟更新的目标网络,并基于DPG,可以在连续的动作空间上运行。
#
# ### 要解决的问题
# * 你正试图解决经典的倒立摆控制问题。在此设置中,你只能执行两个操作:向左摆动或向右摆动。
# * 对于Q-学习算法来说,这个问题的挑战在于动作是连续的而不是离散的。也就是说,你必须从-2到+2的无限操作中进行选择,而不是使用像-1或+1这样的两个离散操作。
#
# ### 快速理论
#
# * 就像**演员-评论家**的方法一样,它有两个网络:
#
# **演员**-它提出一个给定状态的动作。
#
# **评论家**-它预测给定的状态和动作是好(正值)还是坏(负值)。
#
# * DDPG使用的另外2种技术:
# **首先,它使用两个目标网络。**
#
# 为什么?因为它增加了训练的稳定性。简言之,它是从估计的目标和目标网络学习更新,从而保持它估计的目标稳定。
#
# 从概念上讲,这就像是说,“我有一个如何玩这个好主意,我要尝试一下,直到我找到更好的东西”,而不是说“我要在每一个动作之后重新学习如何玩好整个游戏”。
#
# **第二,使用经验回放。**
#
# 它存储元组列表(状态、动作、奖励、下一个状态),而不是仅仅从最近的经验中学习,而是从取样中学习到迄今为止积累的所有经验。
#
# ### 现在,看看它是如何实现的。
# ## **二、环境配置**
# 本教程基于Paddle 2.2.0 编写,如果你的环境不是本版本,请先参考官网[安装](https://www.paddlepaddle.org.cn/install/quick) Paddle 2.2.0。
import gym
import paddle
import paddle.nn as nn
from itertools import count
from paddle.distribution import Normal
import numpy as np
from collections import deque
import random
import paddle.nn.functional as F
from visualdl import LogWriter
# ## **三、实施深度确定策略梯度网络(Deep Deterministic Policy Gradient,DDPG)**
# * **这里定义了演员和评论家网络。这些都是具有ReLU激活的基本全连接模型。**
# **注意**:你需要为Actor的最后一层使用tanh激活,将值映射到-1到1之间。
# * **Memory类定义了经验回放池。**
#
# 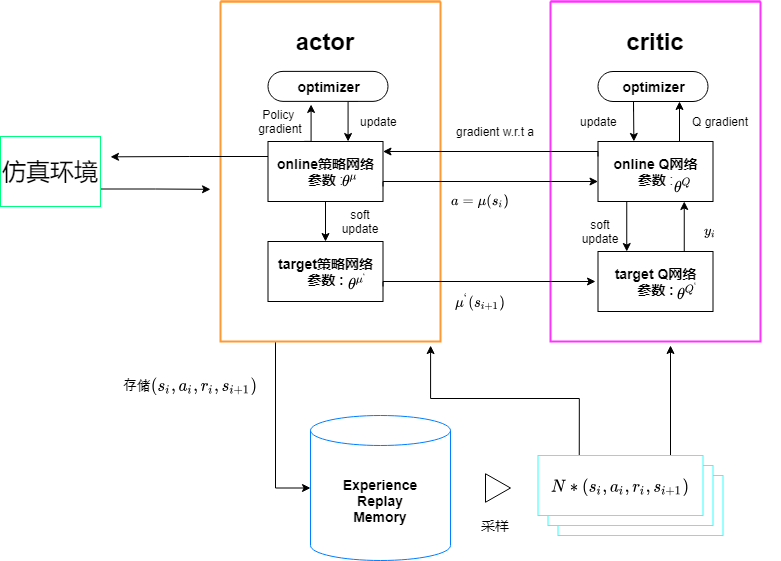
#
#
# +
# 定义评论家网络结构
# DDPG这种方法与Q学习紧密相关,可以看作是连续动作空间的深度Q学习。
class Critic(nn.Layer):
def __init__(self):
super(Critic, self).__init__()
self.fc1 = nn.Linear(3, 256)
self.fc2 = nn.Linear(256 + 1, 128)
self.fc3 = nn.Linear(128, 1)
self.relu = nn.ReLU()
def forward(self, x, a):
x = self.relu(self.fc1(x))
x = paddle.concat((x, a), axis=1)
x = self.relu(self.fc2(x))
x = self.fc3(x)
return x
# 定义演员网络结构
# 为了使DDPG策略更好地进行探索,在训练时对其行为增加了干扰。 原始DDPG论文的作者建议使用时间相关的 OU噪声 ,
# 但最近的结果表明,不相关的均值零高斯噪声效果很好。 由于后者更简单,因此是首选。
class Actor(nn.Layer):
def __init__(self, is_train=True):
super(Actor, self).__init__()
self.fc1 = nn.Linear(3, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 1)
self.relu = nn.ReLU()
self.tanh = nn.Tanh()
self.noisy = Normal(0, 0.2)
self.is_train = is_train
def forward(self, x):
x = self.relu(self.fc1(x))
x = self.relu(self.fc2(x))
x = self.tanh(self.fc3(x))
return x
def select_action(self, epsilon, state):
state = paddle.to_tensor(state,dtype="float32").unsqueeze(0)
with paddle.no_grad():
action = self.forward(state).squeeze() + self.is_train * epsilon * self.noisy.sample([1]).squeeze(0)
return 2 * paddle.clip(action, -1, 1).numpy()
# 重播缓冲区:这是智能体以前的经验, 为了使算法具有稳定的行为,重播缓冲区应该足够大以包含广泛的体验。
# 如果仅使用最新数据,则可能会过分拟合,如果使用过多的经验,则可能会减慢模型的学习速度。 这可能需要一些调整才能正确。
class Memory(object):
def __init__(self, memory_size: int) -> None:
self.memory_size = memory_size
self.buffer = deque(maxlen=self.memory_size)
def add(self, experience) -> None:
self.buffer.append(experience)
def size(self):
return len(self.buffer)
def sample(self, batch_size: int, continuous: bool = True):
if batch_size > len(self.buffer):
batch_size = len(self.buffer)
if continuous:
rand = random.randint(0, len(self.buffer) - batch_size)
return [self.buffer[i] for i in range(rand, rand + batch_size)]
else:
indexes = np.random.choice(np.arange(len(self.buffer)), size=batch_size, replace=False)
return [self.buffer[i] for i in indexes]
def clear(self):
self.buffer.clear()
# -
# ## **四、训练模型**
# ### 算法伪代码
#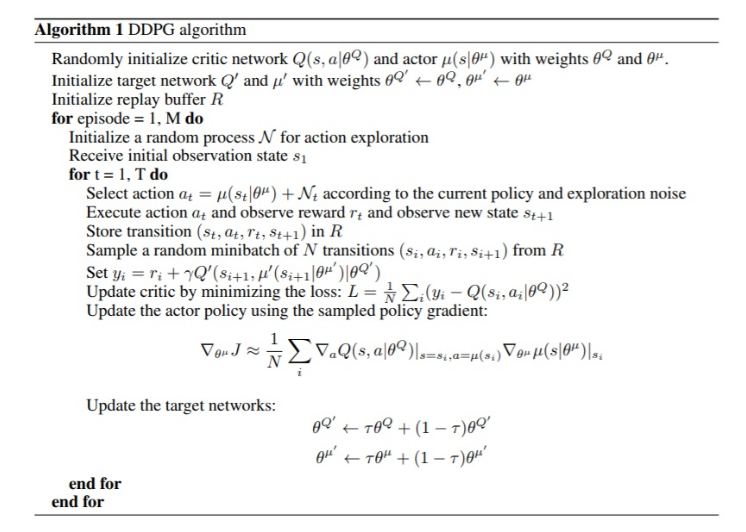 
#
#
#
# +
# 定义软更新的函数
def soft_update(target, source, tau):
for target_param, param in zip(target.parameters(), source.parameters()):
target_param.set_value( target_param * (1.0 - tau) + param * tau)
# 定义环境、实例化模型
env = gym.make('Pendulum-v0')
actor = Actor()
critic = Critic()
actor_target = Actor()
critic_target = Critic()
# 定义优化器
critic_optim = paddle.optimizer.Adam(parameters=critic.parameters(), learning_rate=3e-5)
actor_optim = paddle.optimizer.Adam(parameters=actor.parameters(), learning_rate=1e-5)
# 定义超参数
explore = 50000
epsilon = 1
gamma = 0.99
tau = 0.001
memory_replay = Memory(50000)
begin_train = False
batch_size = 32
learn_steps = 0
epochs = 250
writer = LogWriter('logs')
# 训练循环
for epoch in range(0, epochs):
state = env.reset()
episode_reward = 0
for time_step in range(200):
action = actor.select_action(epsilon, state)
next_state, reward, done, _ = env.step([action])
episode_reward += reward
reward = (reward + 8.1) / 8.1
memory_replay.add((state, next_state, action, reward))
if memory_replay.size() > 1280:
learn_steps += 1
if not begin_train:
print('train begin!')
begin_train = True
experiences = memory_replay.sample(batch_size, False)
batch_state, batch_next_state, batch_action, batch_reward = zip(*experiences)
batch_state = paddle.to_tensor(batch_state,dtype="float32")
batch_next_state = paddle.to_tensor(batch_next_state,dtype="float32")
batch_action = paddle.to_tensor(batch_action,dtype="float32").unsqueeze(1)
batch_reward = paddle.to_tensor(batch_reward,dtype="float32").unsqueeze(1)
# 均方误差 y - Q(s, a) , y是目标网络所看到的预期收益, 而 Q(s, a)是Critic网络预测的操作值。
# y是一个移动的目标,评论者模型试图实现的目标;这个目标通过缓慢的更新目标模型来保持稳定。
with paddle.no_grad():
Q_next = critic_target(batch_next_state, actor_target(batch_next_state))
Q_target = batch_reward + gamma * Q_next
critic_loss = F.mse_loss(critic(batch_state, batch_action), Q_target)
critic_optim.clear_grad()
critic_loss.backward()
critic_optim.step()
writer.add_scalar('critic loss', critic_loss.numpy(), learn_steps)
# 使用Critic网络给定值的平均值来评价Actor网络采取的行动。 我们力求使这一数值最大化。
# 因此,我们更新了Actor网络,对于一个给定状态,它产生的动作尽量让Critic网络给出高的评分。
critic.eval()
actor_loss = - critic(batch_state, actor(batch_state))
# print(actor_loss.shape)
actor_loss = actor_loss.mean()
actor_optim.clear_grad()
actor_loss.backward()
actor_optim.step()
critic.train()
writer.add_scalar('actor loss', actor_loss.numpy(), learn_steps)
soft_update(actor_target, actor, tau)
soft_update(critic_target, critic, tau)
if epsilon > 0:
epsilon -= 1 / explore
state = next_state
writer.add_scalar('episode reward', episode_reward, epoch)
if epoch % 50 == 0:
print('Epoch:{}, episode reward is {}'.format(epoch, episode_reward))
if epoch % 200 == 0:
paddle.save(actor.state_dict(), 'model/ddpg-actor' + str(epoch) + '.para')
paddle.save(critic.state_dict(), 'model/ddpg-critic' + str(epoch) + '.para')
print('model saved!')
# -
# 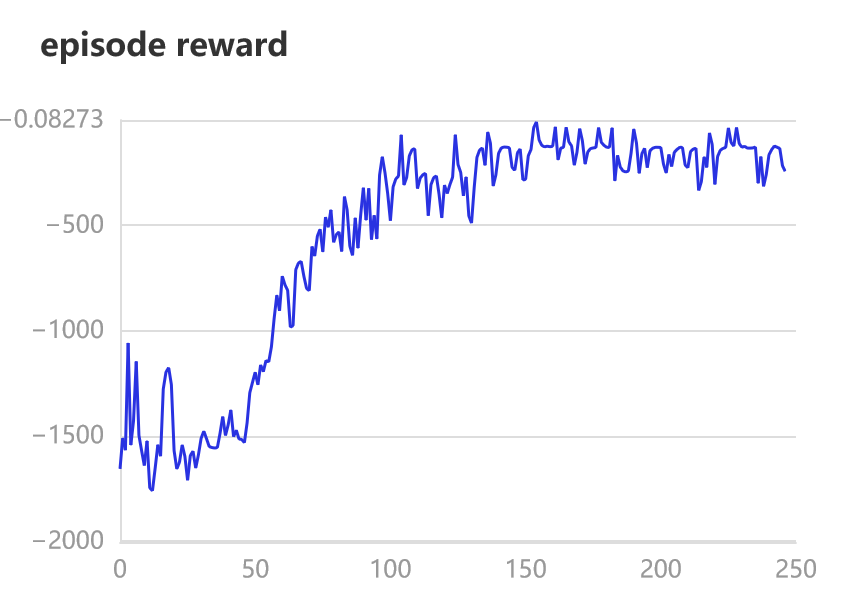
#
#
# ## **五、效果展示**
# 在训练的早期
#
# 
#
#
# 在训练的后期
#
# 
#
# ## **六、总结和建议**
# * DDPG中同时用到了“基于价值”与“基于策略”这两种思想。
# * experience replay memory的使用:actor同环境交互时,产生的transition数据序列是在时间上高度关联的,如果这些数据序列直接用于训练,会导致神经网络的过拟合,不易收敛。
# DDPG的actor将transition数据先存入经验缓冲池, 然后在训练时,从经验缓冲池中随机采样mini-batch数据,这样采样得到的数据可以认为是无关联的。
# * target 网络和online 网络的使用, 使得学习过程更加稳定,收敛更有保障。
# * 如果训练进行的正确,平均奖励将随着时间的推移而增加。请随意尝试演员和评论家网络的不同学习率、tau值和架构。
# * 倒立摆问题的复杂度较低,但DDPG在许多其它问题上都有很好的应用。另一个很好的环境是LunarLandingContinuous-v2,但是需要更多的训练才能获得好的效果。
|
docs/practices/reinforcement_learning/deep_deterministic_policy_gradient.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solution to Kaggle Problem 'Digit Recognizer'
# ***
#
# **Name:AI-23**
#
# **Submission Date:16-01-2018**
#
# ***
# # Abstract
# In this project, .csv file of binary image of digit 0 to 9 are given. The problem is the predict the label of the test samples.
# The main purpose of solving this problem are:
# * Taking input of binary image file and preprocessing
# * Using Keras Deep model like CNN to classify image file
# # Convolutional Neural Network Approach
# Convnet of Convolution Neural Network is used to extract the necessary features nedded to cassify an image. In this problem, CNN (Convolutional Neural Network) is used to solve the problem. In this types of problem where trainining data is small, image augmentation (Creating variation in the training samples) can be very useful. In the [first approach](#f) only cnn network is used without any image augmentation. In the [second approach](#f) cnn with input data from ImageDataGenerator for image augmentation is used. In the [third approach](#t) BatchNormalization layer is added to improve the clasification accuracy.
# <a id="f"></a>
# ### First Approach
# In this approach, CNN without any image augmentation is used. First all the libray file is added.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
# %matplotlib inline
np.random.seed(2)
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
sns.set(style='white', context='notebook', palette='deep')
# -
# Loading data and extracting labels
# +
# Load the data
train = pd.read_csv("../mnistdata/train.csv")
test = pd.read_csv("../mnistdata/test.csv")
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# free some space
del train
g = sns.countplot(Y_train)
Y_train.value_counts()
# -
# #### Preprocessing
# Normalizing the data because doing so helps to converge the model more quickly.
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0
# Reshaping the data to make suitable for CNN input
# Reshape image in 3 dimensions (height = 28px, width = 28px , canal = 1)
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
# Making label suitable for multiclass classification
# Encode labels to one hot vectors (ex : 2 -> [0,0,1,0,0,0,0,0,0,0])
Y_train = to_categorical(Y_train, num_classes = 10)
# Set the random seed
random_seed = 2
# Split the train and the validation set for the fitting
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=random_seed)
# Some examples
g = plt.imshow(X_train[0][:,:,0])
# #### Defining the CNN model architechture
# +
# Set the CNN model
# my CNN architechture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 -> Flatten -> Dense -> Dropout -> Out
model = Sequential()
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(10, activation = "softmax"))
# -
# Define the optimizer
optimizer = RMSprop(lr=0.001, rho=0.9, epsilon=1e-08, decay=0.0)
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
# Set a learning rate annealer
learning_rate_reduction = ReduceLROnPlateau(monitor='val_acc',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
epochs = 1 # Turn epochs to 30 to get 0.9967 accuracy
batch_size = 86
# Without data augmentation i obtained an accuracy of 0.98114
cnn = model.fit(X_train, Y_train, batch_size = batch_size, epochs = epochs,
validation_data = (X_val, Y_val), verbose = 2)
# +
# predict results
results = model.predict(test)
# select the indix with the maximum probability
results = np.argmax(results,axis = 1)
results = pd.Series(results,name="Label")
# +
submission = pd.concat([pd.Series(range(1,28001),name = "ImageId"),results],axis = 1)
submission.to_csv("cnn_mnist_datagen_1st_approach.csv",index=False)
# -
# #### Kaggle Submission Score
# The kaggle score (categorization accuracy) for this approach is 0.98099
# 
# <a id="s"></a>
# ### Second Approach
# In this approach image augmentation is used with the help of keras libraru module ImageDataGenerator. Data preprocessing is done as same as 1st approach.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
# %matplotlib inline
np.random.seed(2)
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
sns.set(style='white', context='notebook', palette='deep')
# +
# Load the data
train = pd.read_csv("../mnistdata/train.csv")
test = pd.read_csv("../mnistdata/test.csv")
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# free some space
del train
g = sns.countplot(Y_train)
Y_train.value_counts()
# -
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0
# Reshape image in 3 dimensions (height = 28px, width = 28px , canal = 1)
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
# Encode labels to one hot vectors (ex : 2 -> [0,0,1,0,0,0,0,0,0,0])
Y_train = to_categorical(Y_train, num_classes = 10)
# Set the random seed
random_seed = 2
# Split the train and the validation set for the fitting
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=random_seed)
# Some examples
g = plt.imshow(X_train[0][:,:,0])
# +
# Set the CNN model
# my CNN architechture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 -> Flatten -> Dense -> Dropout -> Out
model = Sequential()
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(10, activation = "softmax"))
# -
# Define the optimizer
optimizer = RMSprop(lr=0.001, rho=0.9, epsilon=1e-08, decay=0.0)
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
# Set a learning rate annealer
learning_rate_reduction = ReduceLROnPlateau(monitor='val_acc',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
epochs = 30 # Turn epochs to 30 to get 0.9967 accuracy
batch_size = 86
# +
# With data augmentation to prevent overfitting (accuracy 0.99286)
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, # randomly shift images vertically (fraction of total height)
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
# -
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val),
verbose = 2, steps_per_epoch=X_train.shape[0] // batch_size
, callbacks=[learning_rate_reduction])
# +
# Plot the loss and accuracy curves for training and validation
fig, ax = plt.subplots(2,1)
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['acc'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_acc'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
# +
# predict results
results = model.predict(test)
# select the indix with the maximum probability
results = np.argmax(results,axis = 1)
results = pd.Series(results,name="Label")
# +
submission = pd.concat([pd.Series(range(1,28001),name = "ImageId"),results],axis = 1)
submission.to_csv("with_augmentation_cnn_mnist_datagen_2nd_approach.csv",index=False)
# -
# #### Kaggle Score
# The kaggle score (categorization accuracy) for this approach is 0.99614 which is better than the first approach
# 
# <a id="t"></a>
# ### Third Approach
# In this approach, Dropout rate is increased and BatchNormalization layer is added to the fully connected neural network of CNN to further increase the predition accuracy.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
# %matplotlib inline
np.random.seed(2)
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, BatchNormalization
from keras.optimizers import RMSprop
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
sns.set(style='white', context='notebook', palette='deep')
# -
# Load the data
train = pd.read_csv("../mnistdata/train.csv")
test = pd.read_csv("../mnistdata/test.csv")
# +
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# free some space
del train
g = sns.countplot(Y_train)
Y_train.value_counts()
# -
# Check the data
X_train.isnull().any().describe()
test.isnull().any().describe()
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0# Normalize the data
# Reshape image in 3 dimensions (height = 28px, width = 28px , canal = 1)
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
# Encode labels to one hot vectors (ex : 2 -> [0,0,1,0,0,0,0,0,0,0])
Y_train = to_categorical(Y_train, num_classes = 10)
# Set the random seed
random_seed = 2
# Split the train and the validation set for the fitting
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=random_seed)
# Some examples
g = plt.imshow(X_train[0][:,:,0])
# +
# Set the CNN model
# my CNN architechture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 -> Flatten -> Dense -> Dropout -> BatchNormalization -> Out
model = Sequential()
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(Conv2D(filters = 32, kernel_size = (5,5),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(Conv2D(filters = 64, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(BatchNormalization())
model.add(Dropout(0.8))
model.add(Dense(10, activation = "softmax"))
# -
# Define the optimizer
optimizer = RMSprop(lr=0.001, rho=0.9, epsilon=1e-08, decay=0.0)
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
# Set a learning rate annealer
learning_rate_reduction = ReduceLROnPlateau(monitor='val_acc',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
epochs = 30 # Turn epochs to 30 to get 0.9967 accuracy
batch_size = 86
# +
# With data augmentation to prevent overfitting (accuracy 0.99286)
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, # randomly shift images vertically (fraction of total height)
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
# -
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val),
verbose = 2, steps_per_epoch=X_train.shape[0] // batch_size
, callbacks=[learning_rate_reduction])
# +
# Plot the loss and accuracy curves for training and validation
fig, ax = plt.subplots(2,1)
ax[0].plot(history.history['loss'], color='b', label="Training loss")
ax[0].plot(history.history['val_loss'], color='r', label="validation loss",axes =ax[0])
legend = ax[0].legend(loc='best', shadow=True)
ax[1].plot(history.history['acc'], color='b', label="Training accuracy")
ax[1].plot(history.history['val_acc'], color='r',label="Validation accuracy")
legend = ax[1].legend(loc='best', shadow=True)
# +
# Display some error results
# Errors are difference between predicted labels and true labels
errors = (Y_pred_classes - Y_true != 0)
Y_pred_classes_errors = Y_pred_classes[errors]
Y_pred_errors = Y_pred[errors]
Y_true_errors = Y_true[errors]
X_val_errors = X_val[errors]
def display_errors(errors_index,img_errors,pred_errors, obs_errors):
""" This function shows 6 images with their predicted and real labels"""
n = 0
nrows = 2
ncols = 3
fig, ax = plt.subplots(nrows,ncols,sharex=True,sharey=True)
for row in range(nrows):
for col in range(ncols):
error = errors_index[n]
ax[row,col].imshow((img_errors[error]).reshape((28,28)))
ax[row,col].set_title("Predicted label :{}\nTrue label :{}".format(pred_errors[error],obs_errors[error]))
n += 1
# Probabilities of the wrong predicted numbers
Y_pred_errors_prob = np.max(Y_pred_errors,axis = 1)
# Predicted probabilities of the true values in the error set
true_prob_errors = np.diagonal(np.take(Y_pred_errors, Y_true_errors, axis=1))
# Difference between the probability of the predicted label and the true label
delta_pred_true_errors = Y_pred_errors_prob - true_prob_errors
# Sorted list of the delta prob errors
sorted_dela_errors = np.argsort(delta_pred_true_errors)
# Top 6 errors
most_important_errors = sorted_dela_errors[-6:]
# Show the top 6 errors
display_errors(most_important_errors, X_val_errors, Y_pred_classes_errors, Y_true_errors)
# +
# predict results
results = model.predict(test)
# select the indix with the maximum probability
results = np.argmax(results,axis = 1)
results = pd.Series(results,name="Label")
# +
submission = pd.concat([pd.Series(range(1,28001),name = "ImageId"),results],axis = 1)
submission.to_csv("08with_augmentation_batchnorm_cnn_mnist_datagen.csv",index=False)
# -
# ### Conclusion
# The best classification accuracy is obtained for this problem using image augmentation, larger drop out rate and BatchNormalization layer.
|
.ipynb_checkpoints/3rdapproach-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas - Reorder Columns
# Swap two columns, or change the order of columns
import pandas as pd
df = pd.read_csv('iris.data', names=['A','B','C','D','Label'])
df
# 1) get a list of the column names.
titles = list(df.columns)
titles
# 2) Swap or move whatever columns you want in the list.
titles[1], titles[2] = titles[2], titles[1]
titles
# 3) Reassign the columns in the DataFrame.
df = df[titles]
df
|
Pandas/Pandas Reorder Columns in DataFrame.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pachterlab/BLCSBGLKP_2020/blob/master/notebooks/memtime.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="O8d3P2jYCAMa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0841bfb7-b0b2-4f95-d954-d7437dc5d887"
# !date
# + id="O2YLya2XCAMi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="4f186959-f67a-4dad-8929-476549cf9956"
# !git clone https://github.com/pachterlab/BLCSBGLKP_2020.git
# + [markdown] id="veLpmUcECAMm" colab_type="text"
# # Memory and time
# + id="-ureJ3dBCAMn" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import string
from collections import defaultdict
from collections import OrderedDict
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib as mpl
import matplotlib.patches as mpatches
def nd(arr):
return np.asarray(arr).reshape(-1)
def yex(ax):
lims = [
np.min([ax.get_xlim(), ax.get_ylim()]), # min of both axes
np.max([ax.get_xlim(), ax.get_ylim()]), # max of both axes
]
# now plot both limits against eachother
ax.plot(lims, lims, 'k-', alpha=0.75, zorder=0)
ax.set_aspect('equal')
ax.set_xlim(lims)
ax.set_ylim(lims)
return ax
cm = {1:"#D43F3A", 0:"#3182bd"}
fsize=20
plt.rcParams.update({'font.size': fsize})
# %config InlineBackend.figure_format = 'retina'
# + id="BM_50JEgCAMt" colab_type="code" colab={}
df = pd.read_csv("BLCSBGLKP_2020/data/kb/memtime.txt", sep="\t")
# + id="x8KvraNeCAM7" colab_type="code" colab={}
df["wall_time_s"] = df["wall_time"].apply(lambda x: int(x.split(":")[0])*60 + float(x.split(":")[1]))
# + id="WvZ6vZjmCANE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="56aaba77-126f-4121-aff0-d1cd5ceb54de"
df
# + id="5J9G-BdcCANI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="c3108405-e805-4384-b156-5932295157a5"
fig, ax = plt.subplots(figsize=(5,5))
x = df["n_reads"].values.astype(int)
xind = np.arange(0, len(x)*2, 2)
y = df["wall_time_s"].values.astype(float)/60
yy = df["max_mem_bytes"]/1e9
rt = ax.bar(xind-0.5, y, width=0.8, label="Runtime")
# ax.bar(xind+0.5, yy, width=0.8)
ax2 = ax.twinx()
mem = ax2.bar(xind+0.5, yy, width=0.8, color="orange", label="Memory")
ax2.set_ylim(0, 8)
ax2.set_ylabel("Maxmimum RAM [Gb]")
ax.set_xticks(xind)
fmt = lambda x: "{:,.0f}".format(x)
ax.set_xticklabels([fmt(i) for i in x], ha="right", rotation=45)
ax.set_xlabel("Number of reads")
ax.set_ylabel("Time [min]")
ax.set_ylim(0, 8)
ax.legend(handles=[rt, mem])
#plt.savefig("./figs/memtime.png",bbox_inches='tight', dpi=300)
plt.show()
# + id="ezuRvkoSCANV" colab_type="code" colab={}
|
notebooks/memtime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.10 64-bit
# language: python
# name: python361064bit3b840f9918f246278fc4b65bf6247be2
# ---
# Introduction
# --------------
#
# In this section, we consider the very important problem of resolving two nearby frequencies using the DFT. This spectral analysis problem is one of the cornerstone problems in signal processing and we therefore highlight some nuances. We also investigate the circular convolution as a tool to uncover the mechanics of frequency resolution as the uncertainty principle emerges again.
# + jupyter={"outputs_hidden": false}
import numpy as np
import matplotlib.pyplot as plt
Nf = 64 # N- DFT size
fs = 64 # sampling frequency
f = 10 # one signal
t = np.arange(0,1,1/fs) # time-domain samples
deltaf = 1/2. # second nearby frequency
fig,ax = plt.subplots(2,1,sharex=True,sharey=True)
fig.set_size_inches((8,3))
x=np.cos(2*np.pi*f*t) + np.cos(2*np.pi*(f+2)*t) # 2 Hz frequency difference
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[0].plot(np.linspace(0,fs,Nf),abs(X),'-o')
ax[0].set_title(r'$\delta f = 2$',fontsize=18)
ax[0].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[0].grid()
x=np.cos(2*np.pi*f*t) + np.cos(2*np.pi*(f+deltaf)*t) # delta_f frequency difference
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[1].plot(np.linspace(0,fs,Nf),abs(X),'-o')
ax[1].set_title(r'$\delta f = 1/2$',fontsize=14)
ax[1].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[1].set_xlabel('Frequency (Hz)',fontsize=18)
ax[1].set_xlim(xmax = fs/2)
ax[1].set_ylim(ymax=6)
ax[1].grid()
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# Seeking Better Frequency Resolution with Longer DFT
# ----------------------------------------------------
#
# The top plot above shows the magnitude of the DFT for an input that is the sum of two frequencies separated by 2 Hz. Using the parameters we have chosen for the DFT, we can easily see there are two distinct frequencies in the input signal. The bottom plot shows the same thing except that here the frequencies are only separated by 0.5 Hz and, in this case, the two frequencies are not so easy to separate. From this figure, it would be difficult to conclude how many frequencies are present and at what magnitude.
#
# At this point, the usual next step is to increase the size of the DFT since the frequency resolution is $f_s/N$. Thus, the idea is to increase this resolution until the two frequencies separate. This is shown in the next figure.
# + jupyter={"outputs_hidden": false}
Nf = 64*2
fig,ax = plt.subplots(2,1,sharex=True,sharey=True)
fig.set_size_inches((8,4))
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[0].plot(np.linspace(0,fs,len(X)),abs(X),'-o',ms=3.)
ax[0].set_title(r'$N=%d$'%Nf,fontsize=18)
ax[0].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[0].grid()
Nf = 64*4
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[1].plot(np.linspace(0,fs,len(X)),abs(X),'-o',ms=3.)
ax[1].set_title(r'$N=%d$'%Nf,fontsize=18)
ax[1].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[1].set_xlabel('Frequency (Hz)',fontsize=18)
ax[1].set_xlim(xmax = fs/2)
ax[1].set_ylim(ymax=6)
ax[1].grid()
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# As the figure above shows, increasing the size of the DFT did not help matters much. Why is this? Didn't we increase the frequency resolution using a larger DFT? Why can't we separate frequencies now?
#
# The Uncertainty Principle Strikes Back
# --------------------------------------------
#
# The problem here is a manifestation the uncertainty principle we [previously discussed](http://python-for-signal-processing.blogspot.com/2012/09/investigating-sampling-theorem-in-this.html). Remember that taking a larger DFT doesn't add anything new; it just picks off more discrete frequencies on the unit circle. Note that we want to analyze a particular signal $x(t)$, but we have only a *finite section* of that signal. In other words, what we really have are samples of the product of $x(t),t\in \mathbb{R}$ and a rectangular time-window, $r(t)$, that is zero except $r(t)=1 \Leftrightarrow t\in[0,1]$. This means that the DFT is structured according to the rectangular window, which explains the `sinc` shapes we have seen here.
#
# The following figure shows the updated DFT using a longer duration rectangular window.
# + jupyter={"outputs_hidden": false}
t = np.arange(0,2,1/fs)
x=np.cos(2*np.pi*f*t) + np.cos(2*np.pi*(f+deltaf)*t)
Nf = 64*2
fig,ax = plt.subplots(2,1,sharex=True,sharey=True)
fig.set_size_inches((8,4))
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[0].plot(np.linspace(0,fs,len(X)),abs(X),'-o',ms=3.)
ax[0].set_title(r'$N=%d$'%Nf,fontsize=18)
ax[0].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[0].grid()
Nf = 64*8
X = np.fft.fft(x,Nf)/np.sqrt(Nf)
ax[1].plot(np.linspace(0,fs,len(X)),abs(X),'-o',ms=3.)
ax[1].set_title(r'$N=%d$'%Nf,fontsize=18)
ax[1].set_ylabel(r'$|X(k)|$',fontsize=18)
ax[1].set_xlabel('Frequency (Hz)',fontsize=18)
ax[1].set_xlim(xmax = fs/2)
ax[1].set_ylim(ymax=6)
ax[1].grid()
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# The top plot in the figure above shows the DFT of the longer duration signal with $N=128$. The bottom plot shows the same signal with larger DFT length of $N=512$ and a clear separation between the two frequencies. Thus, as opposed to the previous case, a longer DFT *did* resolve the nearby frequencies, but it needed a longer duration signal to do it. Why is this? Consider the DFT of the rectangular windows of length $N_s$,
#
# $$ X[k] = \frac{1}{\sqrt N}\sum_{n=0}^{N_s-1} \exp\left( \frac{2\pi}{N} k n \right) $$
#
# after some re-arrangement, this reduces to
#
#
# $$ |X[k]|=\frac{ 1}{\sqrt N}\left|\frac{\sin \left( N_s \frac{2\pi}{N} k\right)}{\sin \left( \frac{2\pi}{N} k \right)}\right|$$
#
# which bears a strong resemblence to our [original](http://python-for-signal-processing.blogspot.com/2012/09/investigating-sampling-theorem-in-this.html) `sinc` function. The following figure is a plot of this function
# + jupyter={"outputs_hidden": false}
def abs_sinc(k=None,N=64,Ns=32):
if k is None: k = np.arange(0,N-1)
y = np.where(k == 0, 1.0e-20, k)
return abs(np.sin( Ns*2*np.pi/N*y)/np.sin(2*np.pi*y/N))/np.sqrt(N)
fig,ax=plt.subplots()
fig.set_size_inches((8,3))
ax.plot(abs_sinc(N=512,Ns=10),label='duration=10')
ax.plot(abs_sinc(N=512,Ns=20),label='duration=20')
ax.set_xlabel('DFT Index',fontsize=18)
ax.set_ylabel(r'$|X(\Omega_k)|$',fontsize=18)
ax.set_title('Rectangular Windows DFTs',fontsize=18)
ax.grid()
ax.legend(loc=0);
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# Note that the DFT grows taller and narrower as the sampling duration increases (i.e. longer rectangular window). The amplitude growth occurs because the longer window accumulates more "energy" than the shorter window. The length of the DFT is the same for both lines shown so only the length of the rectangular window varies. The point is that taking a longer duration rectangular window improves the frequency resolution! This fact is just the uncertainty principle at work. Looking at the `sinc` formula, the null-to-null width of the main lobe in frequency terms is the following
#
# $$ \delta f = 2\frac{N}{2 N_s} \frac{f_s}{N} =\frac{f_s}{N_s} $$
#
# Thus, two frequencies that differ by at least this amount should be resolvable in these plots.
#
# Thus, in our last example, we had $f_s= 64,N_s = 128 \Rightarrow \delta f = 1/2$ Hz and we were trying to separate two frequencies 0.5 Hz apart so we were right on the edge in this case. I invite you to download this IPython notebook and try longer or shorter signal durations to see how these plots change. Incidentally, this is where some define the notion of *frequency bin* as the DFT resolution ($ f_s/N $) divided by this minimal resolution, $ f_s/N_s $ which gives $ N_s/N $. In other words, the DFT measures frequency in discrete *bins* of minimal resolution, $ N_s/N $.
#
# However, sampling over a longer duration only helps when the signal frequencies are *stable* over the longer duration. If these frequencies drift during the longer sampling interval or otherwise become contaminated with other signals, then advanced techniques become necessary.
#
# Let's consider in detail how the DFT of the rectangular window affects resolution by considering the circular convolution.
#
# ## Circular Convolution
#
# Suppose we want to compute the DFT of a product $z_n=x_n y_n$ as shown below,
#
# $$ Z_k = \frac{1}{\sqrt N}\sum_{n=0}^{N-1} (x_n y_n) W_N^{n k} $$
#
# in terms of the respective DFTs of $x_n$ and $y_n$, $X_k$ and $Y_k$, respectively, where
#
# $$ x_n = \frac{1}{\sqrt N}\sum_{p=0}^{N-1} X_p W_N^{-n p} $$
#
# and
#
# $$ y_n = \frac{1}{\sqrt N}\sum_{m=0}^{N-1} Y_m W_N^{-n m} $$
#
# Then, substituting back in gives,
#
# $$Z_k = \frac{1}{\sqrt N} \frac{1}{N} \sum_{p=0}^{N-1} X_p \sum_{m=0}^{N-1} Y_m \sum_{n=0}^{N-1} W_N^{n k -n p - n m}$$
#
#
# The last term evaluates to
#
# $$ \sum_{n=0}^{N-1} W_N^{n k -n p - n m} = \frac{1-W_N^{N(k-p-m)}}{1-W_N^{k-p-m}} \hspace{2em} = \frac{1-e^{j2\pi(k-p-m)}}{1-e^{j 2\pi (k-p-m)/N}}$$
#
# This is zero everywhere except where $k-p-m= q N$ ($q\in \mathbb{Z}$) in which case it is $N$. Substituting all this back into our expression gives the *circular convolution* usually denoted as
#
# $$ Z_k = \frac{1}{\sqrt N} \sum_{p=0}^{N-1} X_p Y_{((k-p))_N} = X_k \otimes_N Y_k $$
#
# where the double subscripted parenthesis emphasizes the periodic nature of the index. The circular convolution tells us to compute the DFT $Z_k$ directly from the corresponding DFTs $X_k$ and $Y_k$.
#
# Let's work through an example to see this in action.
# + jupyter={"outputs_hidden": false}
def dftmatrix(Nfft=32,N=None):
'construct DFT matrix'
k= np.arange(Nfft)
if N is None: N = Nfft
n = np.arange(N)
U = np.matrix(np.exp(1j* 2*np.pi/Nfft *k*n[:,None])) # use numpy broadcasting to create matrix
return U/np.sqrt(Nfft)
Nf = 32 # DFT size
U = dftmatrix(Nf,Nf)
x = U[:,12].real # input signal
X = U.H*x # DFT of input
rect = np.ones((int(Nf/2), 1)) # short rectangular window
z = x[:int(Nf/2)] # product of rectangular window and x (i.e. chopped version of x)
R = dftmatrix(Nf,Nf/2).H*rect # DFT of rectangular window
Z = dftmatrix(Nf,Nf/2).H*z # DFT of product of x_n and r_n
# + jupyter={"outputs_hidden": false}
idx=np.arange(Nf)-np.arange(Nf)[:,None] # use numpy broadcasting to setup summand's indices
idx[idx<0]+=Nf # add periodic Nf to negative indices for wraparound
a = np.arange(Nf) # k^th frequency index
fig,ax = plt.subplots(4,8,sharex=True,sharey=True)
fig.set_size_inches((12,5))
for i,j in enumerate(ax.flat):
#markerline, stemlines, baseline = j.stem(np.arange(Nf),abs(R[idx[:,i],0])/np.sqrt(Nf))
#setp(markerline, 'markersize', 3.)
j.fill_between(np.arange(Nf),1/np.sqrt(Nf)*abs(R[idx[:,i],0]).flat,0,alpha=0.3)
markerline, stemlines, baseline =j.stem(np.arange(Nf),abs(X))
plt.setp(markerline, 'markersize', 4.)
plt.setp(markerline,'markerfacecolor','r')
plt.setp(stemlines,'color','r')
j.axis('off')
j.set_title('k=%d'%i,fontsize=8)
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# The figure above shows the <font color="blue">rectangular window DFT in blue, $R_k$</font> against the sinusoid <font color="red">input signal in red, $X_k$</font>, for each value of $k$ as the two terms slide past each other from left to right, top to bottom. In other words, the $k^{th}$ term in $Z_k$, the DFT of the product $x_n r_n $, can be thought of as the inner-product of the red and blue lines. This is not exactly true because we are just plotting magnitudes and not the real/imaginary parts, but it's enough to understand the mechanics of the circular convolution.
#
# A good way to think about the rectangular window's `sinc` shape as it slides past the input signal is as a *probe* with a resolution defined by its mainlobe width. For example, in frame $k=12$, we see that the peak of the rectangular window coincides with the peak of the input frequency so we should expect a large value for $Z_{k=12}$ which is shown below. However, if the rectangular window were shorter, corresponding to a wider mainlobe width, then two nearby frequencies could be draped in the same mainlobe and would then be indistinguishable in the resulting DFT because the DFT for that value of $k$ is the inner-product (i.e. a complex number) of the two overlapping graphs.
#
# The figure below shows the direct computation of the DFT of $Z_k$ matches the circular convolution method using $X_k$ and $R_k$.
# + jupyter={"outputs_hidden": false}
fig,ax=plt.subplots()
fig.set_size_inches((7,3))
ax.plot(a,abs(R[idx,0]*X)/np.sqrt(Nf), label=r'$|Z_k|$ = $X_k\otimes_N R_k$')
ax.plot(a, abs(Z),'o',label=r'$|Z_k|$ by DFT')
ax.set_xlabel('DFT index,k',fontsize=18)
ax.set_ylabel(r'$|Z_k|$',fontsize=18)
ax.set_xticks(np.arange(ax.get_xticks().max()))
ax.tick_params(labelsize=8)
ax.legend(loc=0)
ax.grid()
# fig.savefig('figure_00@.png', bbox_inches='tight', dpi=300)
# -
# ## Summary
# In this section, we unpacked the issues involved in resolving two nearby frequencies using the DFT and once again confronted the uncertainty principle in action. We realized that longer DFTs cannot distinguish nearby frequencies unless the signal is sampled over a sufficient duration. Additionally, we developed the circular convolution as a tool to visualize the exactly how a longer sampling duration helps resolve frequencies.
#
# As usual, the original corresponding IPython notebook for this post is available for download [here](https://github.com/unpingco/Python-for-Signal-Processing/blob/master/Frequency_Resolution.ipynb).
#
# Comments and corrections welcome!
# References
# ---------------
#
# * <NAME>., and <NAME>. "Signals and Systems." Prentice-Hall, (1997).
# * <NAME>. "Digital signal processing: principles algorithms and applications". Pearson Education India, 2001.
|
notebook/Frequency_Resolution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Benchmark FRESA.CAD BSWIMS final Script
#
# This algorithm implementation uses R code and a Python library (rpy2) to connect with it, in order to run the following it is necesary to have installed both on your computer:
#
# - R (you can download in https://www.r-project.org/) <br>
# - install rpy2 by <code> pip install rpy2 </code>
import numpy as np
import pandas as pd
import sys
from pathlib import Path
import tadpole_algorithms
from tadpole_algorithms.models import BenchmarkSVM_R
from tadpole_algorithms.preprocessing.split import split_test_train_tadpole
#rpy2 libs and funcs
import rpy2.robjects.packages as rpackages
from rpy2.robjects.vectors import StrVector
from rpy2.robjects import r, pandas2ri
from rpy2 import robjects
# +
# Load D1_D2 train and possible test data set
data_path_train_test = Path("data/TADPOLE_D1_D2.csv")
data_df_train_test = pd.read_csv(data_path_train_test)
# Load D4 evaluation data set
data_path_eval = Path("data/TADPOLE_D4_corr.csv")
data_df_eval = pd.read_csv(data_path_eval)
# Split data in test, train and evaluation data
train_df, test_df, eval_df = split_test_train_tadpole(data_df_train_test, data_df_eval)
#instanciate the model to get the functions
model = BenchmarkSVM_R()
#set the flag to true to use a preprocessed data
USE_PREPROC = True
#preprocess the data
AdjustedTrainFrame,testingFrame,Train_Imputed,Test_Imputed = model.preproc_tadpole_D1_D2(train_df,USE_PREPROC)
#train and predit
Forecast_D2 = model.Forecast_D2_HLCM_EM(AdjustedTrainFrame,testingFrame,Train_Imputed,Test_Imputed,False)
# -
AdjustedTrainFrame,testingFrame,Train_Imputed,Test_Imputed = model.preproc_tadpole_D3(train_df,USE_PREPROC)
#train and predit
Forecast_D3 = model.Forecast_D3_HLCM_EM(AdjustedTrainFrame,testingFrame,Train_Imputed,Test_Imputed,False)
# +
from tadpole_algorithms.evaluation import evaluate_forecast
from tadpole_algorithms.evaluation import print_metrics
# Evaluate the model
dictionary = evaluate_forecast(eval_df,Forecast_D3 )
# Print metrics
print_metrics(dictionary)
# +
from tadpole_algorithms.evaluation import evaluate_forecast
from tadpole_algorithms.evaluation import print_metrics
# Evaluate the model
dictionary = evaluate_forecast(eval_df, Forecast_D2)
# Print metrics
print_metrics(dictionary)
|
benchmarking_FRESA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2. Tree cover widget
#
# The default view for the tree cover widget should be for 'All Region'.
# We provide a mapping between the text to select in the Location drop-down and the dataset IDs that need to be called in conjunction with a specific selection.
#
#
# For the default "All Region" location, we will need to show 3 slices in the donut chart:
# * Tree plantations
# * Natural forest (tree cover 2010 - tree plantations)
# * Non-forest (total area - tree cover 2010)
#
# We show how to calculate this below.
#
# **BUT**
#
# If other Locations are selected, (e.g. Protected Areas), we need a different donut chart. One with less data (only tree cover, and non-forest).
#
# * Tree cover
# * Non-forest
#
#
#
# *Notes: below this line are extra notes not needed for Front-end dev*
#
# - adm0 = BRA, adm1 = 4 is Amazonas
# - adm0 = BRA, adm1 = 4, adm2 = 141 is Amaturá (many forests)
# - adm0 = BRA, adm1 = 12, adm2 = 1434 is Mato Grosso, Cáceres
# - adm0 = BRA, adm1 = 14, adm2 = 2404 is Para, Altamira
# - adm0 = BRA, adm1 = 16, adm2 = 3135 - largest area of plantations (Turning this on seems to reveal a bug)
#
# +
#Import Global Metadata etc
# %run '0.Importable_Globals.ipynb'
# -
# # GLOBAL UPDATE
#
# To adapt for global we need to allow the SQL to accept adm0 = None.
# +
# VARIABLES FOR TREE COVER WIDGET
ds = '499682b1-3174-493f-ba1a-368b4636708e'
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
adm0 = None
adm1 = None
adm2 = None
threshold = 30
location = 'All Region'
extent_year = 2000 #2000 or 2010
tags = ["land_cover", "conservation", "people", "land_use"]
selectable_polynames = ['gadm28', "mining", "wdpa", 'landmark']
# -
def global_extent_queries(p_name, year, adm0, adm1=None, adm2 = None, threshold=30):
if adm2:
print('Request for adm2 area')
sql = (f"SELECT SUM({year}) as value, "
f"SUM(area_gadm28) as total_area "
f"FROM data "
f"WHERE iso = '{adm0}' "
f"AND adm1 = {adm1} "
f"AND adm2 = {adm2} "
"AND thresh = {threshold} "
f"AND polyname = '{p_name}'")
return sql
elif adm1:
print('Request for adm1 area')
sql = (f"SELECT SUM({year}) as value, "
f"SUM(area_gadm28) as total_area "
f"FROM data "
f"WHERE iso = '{adm0}' "
f"AND adm1 = {adm1} "
f"AND thresh = {threshold} "
f"AND polyname = '{p_name}'")
return sql
if adm0:
print('Request for adm0 area')
sql = (f"SELECT SUM({year}) as value, "
f"SUM(area_gadm28) as total_area "
f"FROM data "
f"WHERE iso = '{adm0}' "
f"AND thresh = {threshold} "
f"AND polyname = '{p_name}'")
else:
print('Request for Global area')
sql = (f"SELECT SUM({year}) as value, "
f"SUM(area_gadm28) as total_area "
f"FROM data "
f"WHERE thresh = {threshold} "
f"AND polyname = '{p_name}'")
return sql
# +
# Get location extent and area
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
sql = global_extent_queries(p_name=polynames[location], year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
tree_cover_extent_2010 = r.json().get('data')[0].get('value')
except:
tree_cover_extent_2010 = 0.0
print(f"\n{adm0} {adm1} {adm2} Gadm28 Tree cover extent = {tree_cover_extent_2010} ha")
try:
total_area = r.json().get('data')[0].get('total_area')
except:
total_area = None
print(f"total area = {total_area} ha")
# +
# For some locations we will also need to retrieve an area for plantations
# This is the area of UMD forest cover intersecting tree plantations at admin2 level
if location in ['All Region']:
sql = global_extent_queries(p_name=polynames['Plantations'], year=extent_year_dict[extent_year], adm0=adm0,
adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
plantations = r.json().get('data')[0].get('value')
except:
plantations = 0.0
print(f"\n{adm0} {adm1} {adm2} plantation area = {plantations} ha")
else:
print(f"No plantation data for '{location}'")
plantations = None
# +
# Pie chart, where the slices will be ordered and plotted counter-clockwise:
if adm0 and not adm1 and not adm2:
dynamic_sentence = (f"Tree cover for {location.lower()} of {iso_to_countries[adm0]}, "
f"with tree canopy of \u2265{threshold}%")
elif adm0 and adm1 and not adm2:
dynamic_sentence = (f"Tree cover for {location.lower()} of {areaId_to_name[adm1]}, "
f"with tree canopy of \u2265{threshold}%")
elif adm0 and adm1 and adm2:
dynamic_sentence = (f"Tree cover for {location.lower()} of {areaId_to_name[adm2]}, "
f"with tree canopy of \u2265{threshold}%")
else:
dynamic_sentence = (f"Global Tree cover "
f"with tree canopy of \u2265{threshold}%")
if location in ['All Region']:
labels = ['Tree plantations', 'Natural Forest', 'Non-forest']
sizes = [plantations, tree_cover_extent_2010 - plantations, total_area - tree_cover_extent_2010]
colors = ['orange','green','#E2CF96']
else:
labels = ['Tree cover', 'Non-forest']
sizes = [tree_cover_extent_2010, total_area - tree_cover_extent_2010]
colors = ['green','grey']
fig1, ax1 = plt.subplots()
ax1.pie(sizes, labels=labels, autopct='%1.1f%%',
shadow=False, startangle=90, colors=colors)
ax1.axis('equal')
centre_circle = plt.Circle((0,0),0.75,color='black', fc='white',linewidth=0.5)
fig1 = plt.gcf()
fig1.gca().add_artist(centre_circle)
plt.suptitle('Tree cover extent')
plt.title(dynamic_sentence)
plt.show()
# -
print(f"Globally there is {int(tree_cover_extent_2010)}Ha ", end="")
print(f"of forest, covering around ", end="")
print(f"{round(100 * (tree_cover_extent_2010) / total_area, 1)}% of the Earth's land. ", end="")
print(f"Planted forests account for {int(plantations)}Ha of this. ", end="")
# # IFL ver
# +
# VARIABLES FOR TREE COVER WIDGET
ds = '499682b1-3174-493f-ba1a-368b4636708e'
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
adm0 = None
adm1 = None
adm2 = None
threshold = 30
location = 'Intact Forest Landscapes'
extent_year = 2000
tags = ["land_cover"]
selectable_polynames = ['ifl_2013', 'ifl_2013__mining','ifl_2013__wdpa']
# +
# Get Intact Forest Landscapes extent and region area
if location in ['Intact Forest Landscapes',
'Protected Areas in Intact Forest Landscapes',
'Mining in Intact Forest Landscapes']:
sql = global_extent_queries(p_name=polynames['Intact Forest Landscapes'], year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
intact_forest = r.json().get('data')[0].get('value')
except:
intact_forest = 0.0
print(f"\n{adm0} {adm1} {adm2} intact forst area = {intact_forest} ha")
else:
print(f"No data, for '{intact_forest}'")
intact_forest = None
# +
# Get specific extent and area values for intersections (mining in intact forest etc)
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
if location in ['Intact Forest Landscapes']:
poly = polynames['All Region']
elif location in ['Protected Areas in Intact Forest Landscapes']:
poly = polynames['Protected Areas']
elif location in ['Mining in Intact Forest Landscapes']:
poly = polynames['Mining']
sql = global_extent_queries(p_name=poly, year=extent_year_dict[extent_year], adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
tree_cover_extent_2010 = r.json().get('data')[0].get('value')
except:
tree_cover_extent_2010 = 0.0
print(f"\n{adm0} {adm1} {adm2} Gadm28 Tree cover extent = {tree_cover_extent_2010} ha")
try:
total_area = r.json().get('data')[0].get('total_area')
except:
total_area = None
print(f"total area = {total_area} ha")
# +
# For some locations we will also need to retrieve an area for plantations
# If user is interested on subregion (i.e mining) then get plantations within that subregion
if location in ['Protected Areas in Intact Forest Landscapes']:
plantations_poly = polynames['Protected areas in Plantations']
elif location in ['Mining in Intact Forest Landscapes']:
plantations_poly = polynames['Mining in Plantation Areas']
else:
plantations_poly = polynames['Plantations']
sql = global_extent_queries(p_name=plantations_poly, year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
plantations = r.json().get('data')[0].get('value')
except:
plantations = 0.0
print(f"\n{adm0} {adm1} {adm2} plantation area = {plantations} ha")
# +
# Pie chart, where the slices will be ordered and plotted counter-clockwise:
if location in ['Protected Areas in Intact Forest Landscapes', 'Mining in Intact Forest Landscapes']:
labels = ['Intact Forest', 'Degraded Forest', 'Non-forest']
sizes = [intact_forest, total_area - intact_forest, total_area - tree_cover_extent_2010]
colors = ['green', 'yellow','#E2CF96']
labels.append('Plantations')
sizes.append(plantations)
colors.append('orange')
sizes[1] = sizes[1] - plantations
elif location in ['Intact Forest Landscapes']:
labels = ['Intact Forest', 'Degraded Forest', 'Non-forest']
sizes = [intact_forest, tree_cover_extent_2010 - intact_forest, total_area - tree_cover_extent_2010]
colors = ['green', 'yellow','#E2CF96']
labels.append('Plantations')
sizes.append(plantations)
colors.append('orange')
sizes[1] = sizes[1] - plantations
fig1, ax1 = plt.subplots()
ax1.pie(sizes, labels=labels, autopct='%1.1f%%',
shadow=False, startangle=90, colors=colors)
ax1.axis('equal')
centre_circle = plt.Circle((0,0),0.75,color='black', fc='white',linewidth=0.5)
fig1 = plt.gcf()
fig1.gca().add_artist(centre_circle)
plt.suptitle('Tree cover extent')
plt.title(dynamic_sentence)
plt.show()
# -
print(f"Globally there is {int(intact_forest)}Ha ", end="")
print(f"of intact forest, which is around ", end="")
print(f"{round(100 * (intact_forest) / tree_cover_extent_2010, 1)}% of the Earth's total tree cover. ", end="")
# # Primary Forest
# +
# VARIABLES FOR TREE COVER WIDGET
ds = '499682b1-3174-493f-ba1a-368b4636708e'
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
adm0 = None
adm1 = None
adm2 = None
threshold = 30
location = 'Primary Forests'
year = 2000
tags = ["land_cover"]
selectable_polynames = ['gadm28', 'primary_forest', 'primary_forest__mining',
'primary_forest__wdpa','primary_forest__landmark']
# +
# Get extent and area of region
url = f"https://production-api.globalforestwatch.org/v1/query/{ds}"
if location in ['Primary Forests']:
poly = polynames['All Region']
elif location in ['Protected Areas in Primary Forests']:
poly = polynames['Protected Areas']
elif location in ['Mining in Primary Forests']:
poly = polynames['Mining']
elif location in ['Indigenous Lands in Primary Forests']:
poly = polynames['Indigenous Lands']
sql = global_extent_queries(p_name=poly, year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
tree_cover_extent_2010 = r.json().get('data')[0].get('value')
except:
tree_cover_extent_2010 = 0.0
print(f"\n{adm0} {adm1} {adm2} Gadm28 Tree cover extent = {tree_cover_extent_2010} ha")
try:
total_area = r.json().get('data')[0].get('total_area')
except:
total_area = None
print(f"total area = {total_area} ha")
# +
# get primary forests extent and area
if location in ['Primary Forests', 'Mining in Primary Forests',
'Protected Areas in Primary Forests','Indigenous Lands in Primary Forests']:
sql = global_extent_queries(p_name=polynames['Primary Forests'], year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
primary_forest = r.json().get('data')[0].get('value')
except:
primary_forest = 0.0
print(f"\n{adm0} {adm1} {adm2} plantation area = {primary_forest} ha")
else:
print(f"No data, for '{primary_forest}'")
primary_forest = None
# +
# Get plantation extent within the region
if location in ['Protected Areas in Primary Forests']:
plantations_poly = polynames['Protected areas in Plantations']
elif location in ['Indigenous Lands in Primary Forests']:
plantations_poly = polynames['Indigenous Lands in Plantations']
elif location in ['Mining in Primary Forests']:
plantations_poly = polynames['Mining in Plantation Areas']
else:
plantations_poly = polynames['Plantations']
sql = global_extent_queries(p_name=plantations_poly, year=extent_year_dict[extent_year],
adm0=adm0, adm1=adm1, adm2=adm2, threshold=threshold)
r = requests.get(url, params = {"sql": sql})
print(r.url)
print(f'Status: {r.status_code}')
pprint(r.json())
try:
plantations = r.json().get('data')[0].get('value')
except:
plantations = 0.0
print(f"\n{adm0} {adm1} {adm2} plantation area = {plantations} ha")
# +
if location in ['Mining in Primary Forests',
'Protected Areas in Primary Forests','Indigenous Lands in Primary Forests']:
labels = ['Primary Forest', 'Other Forest', 'Non-forest']
sizes = [primary_forest, total_area - primary_forest, total_area - super_location]
colors = ['green', 'yellow','#E2CF96']
labels.append('Plantations')
sizes.append(plantations)
colors.append('orange')
sizes[1] = sizes[1] - plantations
elif location in ['Primary Forests']:
labels = ['Primary Forest', 'Other Forest', 'Non-forest']
sizes = [primary_forest, tree_cover_extent_2010 - primary_forest, total_area - tree_cover_extent_2010]
colors = ['green', 'yellow','#E2CF96']
labels.append('Plantations')
sizes.append(plantations)
colors.append('orange')
sizes[1] = sizes[1] - plantations
fig1, ax1 = plt.subplots()
ax1.pie(sizes, labels=labels, autopct='%1.1f%%',
shadow=False, startangle=90, colors=colors)
ax1.axis('equal')
centre_circle = plt.Circle((0,0),0.75,color='black', fc='white',linewidth=0.5)
fig1 = plt.gcf()
fig1.gca().add_artist(centre_circle)
plt.suptitle('Tree cover extent')
plt.title(dynamic_sentence)
plt.show()
# -
print(f"Globally there is {int(primary_forest)}Ha ", end="")
print(f"of primary forest, which is around ", end="")
print(f"{round(100 * (primary_forest) / tree_cover_extent_2010, 1)}% of the Earth's total tree cover. ", end="")
|
Global_Widgets/Global_Tree_Cover_Widgets.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # Jacobi Example
# dependencies
using LFAToolkit
using LinearAlgebra
using Pkg
Pkg.activate("./")
Pkg.instantiate()
using Plots
# ## Spectrum of Symbol, p=2
# + code_folding=[7]
# setup
p = 2
dimension = 1
mesh = Mesh1D(1.0)
# operator
diffusion = GalleryOperator("diffusion", p+1, p+1, mesh)
# Jacobi smoother
jacobi = Jacobi(diffusion)
# + code_folding=[14, 16, 25]
# full operator symbols
numbersteps = 250
maxeigenvalue = 0
θ_min = -π/2
θ_max = 3π/2
θ_step = 2π/(numbersteps-1)
θ_range = θ_min:θ_step:θ_max
# compute and plot smoothing factor
# setup
ω = [1.00]
eigenvalues = zeros(numbersteps, p)
# compute
for i in 1:numbersteps
θ = [θ_range[i]]
if abs(θ[1]) > π/512
A = computesymbols(jacobi, ω, θ)
currenteigenvalues = [real(val) for val in eigvals(I-A)]
eigenvalues[i, :] = currenteigenvalues
end
end
# plot
xrange = θ_range/π
plot(
xrange,
xlabel="θ/π",
xtickfont=font(12, "Courier"),
eigenvalues,
ytickfont=font(12, "Courier"),
ylabel="λ",
linewidth=3,
legend=:none,
title="Spectrum of Jacobi Symbol",
palette=palette(:tab10)
)
ylims!(min(0.0, eigenvalues...) * 1.1, max(eigenvalues...) * 1.1)
# -
savefig("jacobi_spectrum_2")
# ## Spectrum of Symbol, p=4
# +
# setup
p = 4
dimension = 1
mesh = Mesh1D(1.0)
# operator
diffusion = GalleryOperator("diffusion", p+1, p+1, mesh)
# Jacobi smoother
jacobi = Jacobi(diffusion)
# + code_folding=[14, 16, 25]
# full operator symbols
numbersteps = 250
maxeigenvalue = 0
θ_min = -π/2
θ_max = 3π/2
θ_step = 2π/(numbersteps-1)
θ_range = θ_min:θ_step:θ_max
# compute and plot smoothing factor
# setup
ω = [1.00]
eigenvalues = zeros(numbersteps, p)
# compute
for i in 1:numbersteps
θ = [θ_range[i]]
if abs(θ[1]) > π/512
A = computesymbols(jacobi, ω, θ)
currenteigenvalues = [real(val) for val in eigvals(I-A)]
eigenvalues[i, :] = currenteigenvalues
end
end
# plot
xrange = θ_range/π
plot(
xrange,
xlabel="θ/π",
xtickfont=font(12, "Courier"),
eigenvalues,
ytickfont=font(12, "Courier"),
ylabel="λ",
linewidth=3,
legend=:none,
title="Spectrum of Jacobi Symbol",
palette=palette(:tab10)
)
ylims!(min(0.0, eigenvalues...) * 1.1, max(eigenvalues...) * 1.1)
# -
savefig("jacobi_spectrum_4")
|
papers/copper-mountain-2021/jupyter/1d_jacobi.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0
# language: julia
# name: julia-1.3
# ---
# +
using Printf
using Statistics
using Flux
using DifferentialEquations
using DiffEqFlux
using JLD2
using Plots
using Flux: @epochs
# -
file = jldopen("../data/ocean_convection_profiles.jld2");
# +
Is = keys(file["timeseries/t"])
Nz = file["grid/Nz"]
Nt = length(Is)
t = zeros(Nt)
T = T_data = zeros(Nt, Nz)
for (i, I) in enumerate(Is)
t[i] = file["timeseries/t/$I"]
T[i, :] = file["timeseries/T/$I"][1, 1, 2:Nz+1]
end
# +
z = file["grid/zC"]
anim = @gif for n=1:10:Nt
t_str = @sprintf("%.2f", t[n] / 86400)
plot(T[n, :], z, linewidth=2,
xlim=(19, 20), ylim=(-100, 0), label="",
xlabel="Temperature (C)", ylabel="Depth (z)",
title="Free convection: $t_str days", show=false)
end
display(anim)
# -
function coarse_grain(data, resolution)
@assert length(data) % resolution == 0
s = length(data) / resolution
data_cs = zeros(resolution)
for i in 1:resolution
t = data[Int((i-1)*s+1):Int(i*s)]
data_cs[i] = mean(t)
end
return data_cs
end
# +
coarse_resolution = cr = 32
T_cs = zeros(Nt, coarse_resolution)
for n=1:Nt
T_cs[n, :] = coarse_grain(T[n, :], coarse_resolution)
end
T_cs = transpose(T_cs) |> Array;
# +
dTdt_NN = Chain(Dense(cr, 2cr, tanh),
Dense(2cr, cr))
ps = Flux.params(dTdt_NN)
T₀ = T_cs[:, 1]
n_train = round(Int, Nt/2)
t_train = t[1:n_train] ./ 86400
tspan_train = (t_train[1], t_train[end])
neural_pde_prediction(T₀) = neural_ode(dTdt_NN, T₀, tspan_train, Tsit5(), saveat=t_train, reltol=1e-7, abstol=1e-9)
# +
opt = ADAM(0.1)
data = [(T₀, T_cs[:, 1:n_train])]
loss_function(T₀, T_data) = sum(abs2, T_data .- neural_pde_prediction(T₀))
# -
# Callback function to observe training.
cb = function ()
loss = loss_function(T₀, T_cs[:, 1:n_train]) # Not very generalizable...
println("loss = $loss")
end
for _ in 1:10
Flux.train!(loss_function, ps, data, opt, cb = cb)
end
# +
tspan = (t[1], t[end]) ./ 86400
nn_pred = neural_ode(dTdt_NN, T₀, tspan, Tsit5(), saveat=t ./86400, reltol=1e-7, abstol=1e-9) |> Flux.data
z_cs = coarse_grain(z, cr)
anim = @gif for n=1:10:Nt
t_str = @sprintf("%.2f", t[n] / 86400)
plot(T_cs[:, n], z_cs, linewidth=2,
xlim=(19, 20), ylim=(-100, 0), label="Data",
xlabel="Temperature (C)", ylabel="Depth (z)",
title="Free convection: $t_str days",
legend=:bottomright, show=false)
if n <= n_train
plot!(nn_pred[:, n], z_cs, linewidth=2, label="Neural ODE (train)", show=false)
else
plot!(nn_pred[:, n], z_cs, linewidth=2, linestyle=:dash, label="Neural ODE (test)", show=false)
end
end
# -
|
notebooks/Neural free convection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nes-lter-ims-dev]
# language: python
# name: conda-env-nes-lter-ims-dev-py
# ---
# +
# parse dates and times using Pandas' "to_datetime" function
import pandas as pd
pd.to_datetime('2019-07-22')
# +
# pandas uses the terms "datetime" and "timestamp" to mean
# pretty much the same thing
# -
# pandas will infer the format if it's not ambiguous
pd.to_datetime('July 22, 2019 12:34:56')
pd.to_datetime('7/22/2019')
pd.to_datetime('22 Jul 2019 1:30pm')
# you should probably be using the UTC timezone for everything
# to do this, pass utc=True to to_datetime
# note how a timezone offset is shown
pd.to_datetime('22 July 2019 6:30pm', utc=True)
# represent relative times
pd.to_timedelta('2d')
# +
# do date arithmetic
from datetime import datetime
current_time = pd.to_datetime(datetime.utcnow(), utc=True)
current_time + pd.to_timedelta('12h')
# -
current_time - pd.to_datetime('July 22 2019 16:00', utc=True)
# +
# timestamps in pandas are either "timezone-aware" or not.
# temporal operations won't work between timezone-aware and
# non-timezone-aware timestamps
# here's a non-timezone-aware timestamp
time_a = pd.to_datetime('2019-07-22 12:34:56')
time_a
# -
# if your string has a timezone notation in it, Pandas will parse it
time_b = pd.to_datetime('2019-07-22 12:34:56 +0000')
time_b
# if it doesn't, use utc=True when parsing
time_c = pd.to_datetime('2019-07-22 12:34:56', utc=True)
time_c
# +
# time operations like comparison do not work unless
# either both timezones are non-timezone-aware or
# they are both timezone-aware
try:
time_a > time_b
except TypeError as e:
print(e)
# +
# ^ this comes up in Pandas applications all the time
# because a lot of tabular data uses timestamps with no timezone notation
# always use utc=True, unless you're using a different timezone
# +
# note that to_datetime works for iterables including arrays
# and Pandas Series
time_strings = [
'1969-10-27',
'1971-03-14',
'2009-07-04'
]
timestamps = pd.to_datetime(time_strings, utc=True)
timestamps
# +
# and operations on them are vectorized
a_day = pd.to_timedelta('1d')
timestamps + a_day
# -
timestamps > pd.to_datetime('2000-01-01', utc=True)
|
06 pandas time utilities.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mean-Variance Optimization
# MPT solves for the optimal portfolio weights to minimize volatility for a given expected return, or maximize returns for a given level of volatility. The key requisite input are expected asset returns, standard deviations, and the covariance matrix.
# Diversification works because the variance of portfolio returns depends on the covariance of the assets and can be reduced below the weighted average of the asset variances by including assets with less than perfect correlation. In particular, given a vector, ω, of portfolio weights and the covariance matrix, $\Sigma$, the portfolio variance, $\sigma_{\text{PF}}$ is defined as:
# $$\sigma_{\text{PF}}=\omega^T\Sigma\omega$$
# Markowitz showed that the problem of maximizing the expected portfolio return subject to a target risk has an equivalent dual representation of minimizing portfolio risk subject to a target expected return level, $μ_{PF}$. Hence, the optimization problem becomes:
# $$
# \begin{align}
# \min_\omega & \quad\quad\sigma^2_{\text{PF}}= \omega^T\Sigma\omega\\
# \text{s.t.} &\quad\quad \mu_{\text{PF}}= \omega^T\mu\\
# &\quad\quad \lVert\omega\rVert =1
# \end{align}
# $$
# We can calculate an efficient frontier using `scipy.optimize.minimize` and the historical estimates for asset returns, standard deviations, and the covariance matrix.
# ## Imports & Settings
import warnings
warnings.filterwarnings('ignore')
# +
# %matplotlib inline
import pandas as pd
import numpy as np
from numpy.random import random, uniform, dirichlet, choice
from numpy.linalg import inv
from scipy.optimize import minimize
import pandas_datareader.data as web
import matplotlib.pyplot as plt
from matplotlib.ticker import FuncFormatter
import seaborn as sns
# -
sns.set_style('whitegrid')
np.random.seed(42)
cmap = sns.diverging_palette(10, 240, n=9, as_cmap=True)
# ## Prepare Data
# We select historical data for tickers included in the S&P500 (according to Wikipedia) from 1998-2017.
with pd.HDFStore('../data/assets.h5') as store:
sp500_stocks = store['sp500/stocks']
sp500_stocks.head()
with pd.HDFStore('../data/assets.h5') as store:
prices = (store['quandl/wiki/prices']
.adj_close
.unstack('ticker')
.filter(sp500_stocks.index)
.sample(n=30, axis=1))
# ## Compute Inputs
# ### Compute Returns
start = 2008
end = 2017
# Create month-end monthly returns and drop dates that have no observations:
weekly_returns = prices.loc[f'{start}':f'{end}'].resample('W').last().pct_change().dropna(how='all')
weekly_returns = weekly_returns.dropna(axis=1)
weekly_returns.info()
# ### Set Parameters
stocks = weekly_returns.columns
n_obs, n_assets = weekly_returns.shape
n_assets, n_obs
NUM_PF = 100000 # no of portfolios to simulate
x0 = uniform(0, 1, n_assets)
x0 /= np.sum(np.abs(x0))
# ### Annualization Factor
periods_per_year = round(weekly_returns.resample('A').size().mean())
periods_per_year
# ### Compute Mean Returns, Covariance and Precision Matrix
mean_returns = weekly_returns.mean()
cov_matrix = weekly_returns.cov()
# The precision matrix is the inverse of the covariance matrix:
precision_matrix = pd.DataFrame(inv(cov_matrix), index=stocks, columns=stocks)
# ### Risk-Free Rate
# Load historical 10-year Treasury rate:
treasury_10yr_monthly = (web.DataReader('DGS10', 'fred', start, end)
.resample('M')
.last()
.div(periods_per_year)
.div(100)
.squeeze())
rf_rate = treasury_10yr_monthly.mean()
# ## Simulate Random Portfolios
# The simulation generates random weights using the Dirichlet distribution, and computes the mean, standard deviation, and SR for each sample portfolio using the historical return data:
def simulate_portfolios(mean_ret, cov, rf_rate=rf_rate, short=True):
alpha = np.full(shape=n_assets, fill_value=.05)
weights = dirichlet(alpha=alpha, size=NUM_PF)
if short:
weights *= choice([-1, 1], size=weights.shape)
returns = weights @ mean_ret.values + 1
returns = returns ** periods_per_year - 1
std = (weights @ weekly_returns.T).std(1)
std *= np.sqrt(periods_per_year)
sharpe = (returns - rf_rate) / std
return pd.DataFrame({'Annualized Standard Deviation': std,
'Annualized Returns': returns,
'Sharpe Ratio': sharpe}), weights
simul_perf, simul_wt = simulate_portfolios(mean_returns, cov_matrix, short=False)
df = pd.DataFrame(simul_wt)
df.describe()
# ### Plot Simulated Portfolios
# +
ax = simul_perf.plot.scatter(x=0, y=1, c=2, cmap='Blues',
alpha=0.5, figsize=(14, 9), colorbar=True,
title=f'{NUM_PF:,d} Simulated Portfolios')
max_sharpe_idx = simul_perf.iloc[:, 2].idxmax()
sd, r = simul_perf.iloc[max_sharpe_idx, :2].values
print(f'Max Sharpe: {sd:.2%}, {r:.2%}')
ax.scatter(sd, r, marker='*', color='darkblue', s=500, label='Max. Sharpe Ratio')
min_vol_idx = simul_perf.iloc[:, 0].idxmin()
sd, r = simul_perf.iloc[min_vol_idx, :2].values
ax.scatter(sd, r, marker='*', color='green', s=500, label='Min Volatility')
plt.legend(labelspacing=1, loc='upper left')
plt.tight_layout()
# -
# ## Compute Annualize PF Performance
# Now we'll set up the quadratic optimization problem to solve for the minimum standard deviation for a given return or the maximum SR.
#
# To this end, define the functions that measure the key metrics:
def portfolio_std(wt, rt=None, cov=None):
"""Annualized PF standard deviation"""
return np.sqrt(wt @ cov @ wt * periods_per_year)
def portfolio_returns(wt, rt=None, cov=None):
"""Annualized PF returns"""
return (wt @ rt + 1) ** periods_per_year - 1
def portfolio_performance(wt, rt, cov):
"""Annualized PF returns & standard deviation"""
r = portfolio_returns(wt, rt=rt)
sd = portfolio_std(wt, cov=cov)
return r, sd
# ## Max Sharpe PF
# Define a target function that represents the negative SR for scipy's minimize function to optimize, given the constraints that the weights are bounded by [-1, 1], if short trading is permitted, and [0, 1] otherwise, and sum to one in absolute terms.
def neg_sharpe_ratio(weights, mean_ret, cov):
r, sd = portfolio_performance(weights, mean_ret, cov)
return -(r - rf_rate) / sd
weight_constraint = {'type': 'eq',
'fun': lambda x: np.sum(np.abs(x))-1}
def max_sharpe_ratio(mean_ret, cov, short=False):
return minimize(fun=neg_sharpe_ratio,
x0=x0,
args=(mean_ret, cov),
method='SLSQP',
bounds=((-1 if short else 0, 1),) * n_assets,
constraints=weight_constraint,
options={'tol':1e-10, 'maxiter':1e4})
# ## Compute Efficient Frontier
# The solution requires iterating over ranges of acceptable values to identify optimal risk-return combinations
def min_vol_target(mean_ret, cov, target, short=False):
def ret_(wt):
return portfolio_returns(wt, mean_ret)
constraints = [{'type': 'eq',
'fun': lambda x: ret_(x) - target},
weight_constraint]
bounds = ((-1 if short else 0, 1),) * n_assets
return minimize(portfolio_std,
x0=x0,
args=(mean_ret, cov),
method='SLSQP',
bounds=bounds,
constraints=constraints,
options={'tol': 1e-10, 'maxiter': 1e4})
# The mean-variance frontier relies on in-sample, backward-looking optimization. In practice, portfolio optimization requires forward-looking input. Unfortunately, expected returns are notoriously difficult to estimate accurately.
#
# The covariance matrix can be estimated somewhat more reliably, which has given rise to several alternative approaches. However, covariance matrices with correlated assets pose computational challenges since the optimization problem requires inverting the matrix. The high condition number induces numerical instability, which in turn gives rise to Markovitz curse: the more diversification is required (by correlated investment opportunities), the more unreliable the weights produced by the algorithm.
# ## Min Volatility Portfolio
def min_vol(mean_ret, cov, short=False):
bounds = ((-1 if short else 0, 1),) * n_assets
return minimize(fun=portfolio_std,
x0=x0,
args=(mean_ret, cov),
method='SLSQP',
bounds=bounds,
constraints=weight_constraint,
options={'tol': 1e-10, 'maxiter': 1e4})
def efficient_frontier(mean_ret, cov, ret_range, short=False):
return [min_vol_target(mean_ret, cov, ret) for ret in ret_range]
# ## Run Calculation
# ### Get random PF
simul_perf, simul_wt = simulate_portfolios(mean_returns, cov_matrix, short=False)
print(simul_perf.describe())
simul_max_sharpe = simul_perf.iloc[:, 2].idxmax()
simul_perf.iloc[simul_max_sharpe]
# ### Get Max Sharpe PF
max_sharpe_pf = max_sharpe_ratio(mean_returns, cov_matrix, short=False)
max_sharpe_perf = portfolio_performance(max_sharpe_pf.x, mean_returns, cov_matrix)
r, sd = max_sharpe_perf
pd.Series({'ret': r, 'sd': sd, 'sr': (r-rf_rate)/sd})
# From simulated pf data
# ### Get Min Vol PF
min_vol_pf = min_vol(mean_returns, cov_matrix, short=False)
min_vol_perf = portfolio_performance(min_vol_pf.x, mean_returns, cov_matrix)
# ### Get Efficent PFs
ret_range = np.linspace(simul_perf.iloc[:, 1].min(), simul_perf.iloc[:, 1].max(), 50)
eff_pf = efficient_frontier(mean_returns, cov_matrix, ret_range, short=True)
eff_pf = pd.Series(dict(zip([p['fun'] for p in eff_pf], ret_range)))
# ### Plot Result
# The simulation yields a subset of the feasible portfolios, and the efficient frontier identifies the optimal in-sample return-risk combinations that were achievable given historic data.
#
# The below figure shows the result including the minimum variance portfolio and the portfolio that maximizes the SR and several portfolios produce by alternative optimization strategies. The efficient frontier
# +
fig, ax = plt.subplots()
simul_perf.plot.scatter(x=0, y=1, c=2, ax=ax, cmap='Blues',alpha=0.25,
figsize=(14, 9), colorbar=True)
eff_pf[eff_pf.index.min():].plot(linestyle='--', lw=2, ax=ax, c='k',
label='Efficient Frontier')
r, sd = max_sharpe_perf
ax.scatter(sd, r, marker='*', color='k', s=500, label='Max Sharpe Ratio PF')
r, sd = min_vol_perf
ax.scatter(sd, r, marker='v', color='k', s=200, label='Min Volatility PF')
kelly_wt = precision_matrix.dot(mean_returns).clip(lower=0).values
kelly_wt /= np.sum(np.abs(kelly_wt))
r, sd = portfolio_performance(kelly_wt, mean_returns, cov_matrix)
ax.scatter(sd, r, marker='D', color='k', s=150, label='Kelly PF')
std = weekly_returns.std()
std /= std.sum()
r, sd = portfolio_performance(std, mean_returns, cov_matrix)
ax.scatter(sd, r, marker='X', color='k', s=250, label='Risk Parity PF')
r, sd = portfolio_performance(np.full(n_assets, 1/n_assets), mean_returns, cov_matrix)
ax.scatter(sd, r, marker='o', color='k', s=200, label='1/n PF')
ax.legend(labelspacing=0.8)
ax.set_xlim(0, eff_pf.max()+.4)
ax.set_title('Mean-Variance Efficient Frontier', fontsize=16)
ax.yaxis.set_major_formatter(FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
ax.xaxis.set_major_formatter(FuncFormatter(lambda y, _: '{:.0%}'.format(y)))
sns.despine()
fig.tight_layout();
|
ml4trading-2ed/05_strategy_evaluation/04_mean_variance_optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Transfer Learning
#
# A Convolutional Neural Network (CNN) for image classification is made up of multiple layers that extract features, such as edges, corners, etc; and then use a final fully-connected layer to classify objects based on these features. You can visualize this like this:
#
# <table>
# <tr><td rowspan=2 style='border: 1px solid black;'>⇒</td><td style='border: 1px solid black;'>Convolutional Layer</td><td style='border: 1px solid black;'>Pooling Layer</td><td style='border: 1px solid black;'>Convolutional Layer</td><td style='border: 1px solid black;'>Pooling Layer</td><td style='border: 1px solid black;'>Fully Connected Layer</td><td rowspan=2 style='border: 1px solid black;'>⇒</td></tr>
# <tr><td colspan=4 style='border: 1px solid black; text-align:center;'>Feature Extraction</td><td style='border: 1px solid black; text-align:center;'>Classification</td></tr>
# </table>
#
# *Transfer Learning* is a technique where you can take an existing trained model and re-use its feature extraction layers, replacing its final classification layer with a fully-connected layer trained on your own custom images. With this technique, your model benefits from the feature extraction training that was performed on the base model (which may have been based on a larger training dataset than you have access to) to build a classification model for your own specific set of object classes.
#
# How does this help? Well, think of it this way. Suppose you take a professional tennis player and a complete beginner, and try to teach them both how to play raquetball. It's reasonable to assume that the professional tennis player will be easier to train, because many of the underlying skills involved in raquetball are already learned. Similarly, a pre-trained CNN model may be easier to train to classify specific set of objects because it's already learned how to identify the features of common objects, such as edges and corners. Fundamentally, a pre-trained model can be a great way to produce an effective classifier even when you have limited data with which to train it.
#
# In this notebook, we'll see how to implement transfer learning for a classification model using TensorFlow.
# ## Install and import TensorFlow libraries
#
# Let's start by ensuring that we have the latest version of the **TensorFlow** package installed and importing the Tensorflow libraries we're going to use.
# !pip install --upgrade tensorflow
# + tags=[]
import tensorflow
from tensorflow import keras
print('TensorFlow version:',tensorflow.__version__)
print('Keras version:',keras.__version__)
# -
# ## Prepare the base model
#
# To use transfer learning, we need a base model from which we can use the trained feature extraction layers. The ***resnet*** model is an CNN-based image classifier that has been pre-trained using a huge dataset of 3-color channel images of 224x224 pixels. Let's create an instance of it with some pretrained weights, excluding its final (top) prediction layer.
# + tags=["outputPrepend"]
base_model = keras.applications.resnet.ResNet50(weights='imagenet', include_top=False, input_shape=(224,224,3))
print(base_model.summary())
# -
# ## Prepare the image data
#
# The pretrained model has many layers, starting with a convolutional layer that starts the feature extraction process from image data.
#
# For feature extraction to work with our own images, we need to ensure that the image data we use to train our prediction layer has the same number of features (pixel values) as the images originally used to train the feature extraction layers, so we need data loaders for color images that are 224x224 pixels in size.
#
# Tensorflow includes functions for loading and transforming data. We'll use these to create a generator for training data, and a second generator for test data (which we'll use to validate the trained model). The loaders will transform the image data to match the format used to train the original resnet CNN model and normalize them.
#
# Run the following cell to define the data generators and list the classes for our images.
# + tags=[]
from tensorflow.keras.preprocessing.image import ImageDataGenerator
data_folder = 'data/shapes'
pretrained_size = (224,224)
batch_size = 30
print("Getting Data...")
datagen = ImageDataGenerator(rescale=1./255, # normalize pixel values
validation_split=0.3) # hold back 30% of the images for validation
print("Preparing training dataset...")
train_generator = datagen.flow_from_directory(
data_folder,
target_size=pretrained_size, # resize to match model expected input
batch_size=batch_size,
class_mode='categorical',
subset='training') # set as training data
print("Preparing validation dataset...")
validation_generator = datagen.flow_from_directory(
data_folder,
target_size=pretrained_size, # resize to match model expected input
batch_size=batch_size,
class_mode='categorical',
subset='validation') # set as validation data
classnames = list(train_generator.class_indices.keys())
print("class names: ", classnames)
# -
# ## Create a prediction layer
#
# We downloaded the complete *resnet* model excluding its final prediction layer, so need to combine these layers with a fully-connected (*dense*) layer that takes the flattened outputs from the feature extraction layers and generates a prediction for each of our image classes.
#
# We also need to freeze the feature extraction layers to retain the trained weights. Then when we train the model using our images, only the final prediction layer will learn new weight and bias values - the pre-trained weights already learned for feature extraction will remain the same.
# + tags=["outputPrepend"]
from tensorflow.keras import applications
from tensorflow.keras import Model
from tensorflow.keras.layers import Flatten, Dense
# Freeze the already-trained layers in the base model
for layer in base_model.layers:
layer.trainable = False
# Create prediction layer for classification of our images
x = base_model.output
x = Flatten()(x)
prediction_layer = Dense(len(classnames), activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=prediction_layer)
# Compile the model
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# Now print the full model, which will include the layers of the base model plus the dense layer we added
print(model.summary())
# -
# ## Train the Model
#
# With the layers of the CNN defined, we're ready to train it using our image data. The weights used in the feature extraction layers from the base resnet model will not be changed by training, only the final dense layer that maps the features to our shape classes will be trained.
# + tags=[]
# Train the model over 3 epochs
num_epochs = 3
history = model.fit(
train_generator,
steps_per_epoch = train_generator.samples // batch_size,
validation_data = validation_generator,
validation_steps = validation_generator.samples // batch_size,
epochs = num_epochs)
# -
# ## View the loss history
#
# We tracked average training and validation loss for each epoch. We can plot these to verify that the loss reduced over the training process and to detect *over-fitting* (which is indicated by a continued drop in training loss after validation loss has levelled out or started to increase).
# +
# %matplotlib inline
from matplotlib import pyplot as plt
epoch_nums = range(1,num_epochs+1)
training_loss = history.history["loss"]
validation_loss = history.history["val_loss"]
plt.plot(epoch_nums, training_loss)
plt.plot(epoch_nums, validation_loss)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['training', 'validation'], loc='upper right')
plt.show()
# -
# ## Evaluate model performance
#
# We can see the final accuracy based on the test data, but typically we'll want to explore performance metrics in a little more depth. Let's plot a confusion matrix to see how well the model is predicting each class.
# + tags=[]
# Tensorflow doesn't have a built-in confusion matrix metric, so we'll use SciKit-Learn
import numpy as np
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
# %matplotlib inline
print("Generating predictions from validation data...")
# Get the image and label arrays for the first batch of validation data
x_test = validation_generator[0][0]
y_test = validation_generator[0][1]
# Use the model to predict the class
class_probabilities = model.predict(x_test)
# The model returns a probability value for each class
# The one with the highest probability is the predicted class
predictions = np.argmax(class_probabilities, axis=1)
# The actual labels are hot encoded (e.g. [0 1 0], so get the one with the value 1
true_labels = np.argmax(y_test, axis=1)
# Plot the confusion matrix
cm = confusion_matrix(true_labels, predictions)
plt.imshow(cm, interpolation="nearest", cmap=plt.cm.Blues)
plt.colorbar()
tick_marks = np.arange(len(classnames))
plt.xticks(tick_marks, classnames, rotation=85)
plt.yticks(tick_marks, classnames)
plt.xlabel("Predicted Shape")
plt.ylabel("Actual Shape")
plt.show()
# -
# ## Use the trained model
#
# Now that we've trained the model, we can use it to predict the class of an image.
# + tags=[]
from tensorflow.keras import models
import numpy as np
from random import randint
import os
# %matplotlib inline
# Function to predict the class of an image
def predict_image(classifier, image):
from tensorflow import convert_to_tensor
# The model expects a batch of images as input, so we'll create an array of 1 image
imgfeatures = img.reshape(1, img.shape[0], img.shape[1], img.shape[2])
# We need to format the input to match the training data
# The generator loaded the values as floating point numbers
# and normalized the pixel values, so...
imgfeatures = imgfeatures.astype('float32')
imgfeatures /= 255
# Use the model to predict the image class
class_probabilities = classifier.predict(imgfeatures)
# Find the class predictions with the highest predicted probability
index = int(np.argmax(class_probabilities, axis=1)[0])
return index
# Function to create a random image (of a square, circle, or triangle)
def create_image (size, shape):
from random import randint
import numpy as np
from PIL import Image, ImageDraw
xy1 = randint(10,40)
xy2 = randint(60,100)
col = (randint(0,200), randint(0,200), randint(0,200))
img = Image.new("RGB", size, (255, 255, 255))
draw = ImageDraw.Draw(img)
if shape == 'circle':
draw.ellipse([(xy1,xy1), (xy2,xy2)], fill=col)
elif shape == 'triangle':
draw.polygon([(xy1,xy1), (xy2,xy2), (xy2,xy1)], fill=col)
else: # square
draw.rectangle([(xy1,xy1), (xy2,xy2)], fill=col)
del draw
return np.array(img)
# Create a random test image
classnames = os.listdir(os.path.join('data', 'shapes'))
classnames.sort()
img = create_image ((224,224), classnames[randint(0, len(classnames)-1)])
plt.axis('off')
plt.imshow(img)
# Use the classifier to predict the class
class_idx = predict_image(model, img)
print (classnames[class_idx])
# -
# ## Learn More
#
# * [Tensorflow Documentation](https://www.tensorflow.org/tutorials/images/transfer_learning)
|
05c - Transfer Learning (Tensorflow).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import langdetect
# +
#Load CSVs in dataframes
csv1 = pd.read_csv('1.csv', delimiter = ',',encoding = "ISO-8859-1")
csv2 = pd.read_csv('2.csv', delimiter = ',',encoding = "ISO-8859-1")
# -
#Merge Dataframes
df = pd.concat([csv1,csv2])
df.reset_index(inplace=True, drop=True)
#Filter by 'From: (Address)'
df_filtered_from = df[df['From: (Name)'].str.contains('***NAME OF THE PERSON TO FILTER***')]
#Filter NaN
df_filtered_nan = df_filtered_from[df_filtered_from['Body'].str.contains('NaN')==False]
df_filtered_nan = df_filtered_nan[df_filtered_nan['Body'].str.match('\r\n')==False]
df_filtered_nan = df_filtered_nan[df_filtered_nan['Body'].str.match(' \r\n\r\n')==False]
df_filtered_nan.reset_index(inplace=True, drop=True)
#Create a list with languages for emails
language_list = []
for index, row in df_filtered_nan.iterrows():
language_list.append(langdetect.detect(row['Body']))
#Add column to dataframe with languages
df_filtered_nan["Language"] = language_list
print(df_filtered_nan.shape)
#Filter emails based on language
df_filtered_language = df_filtered_nan[df_filtered_nan['Language'].str.match('en')]
for index, row in df_filtered_language.iterrows():
df_filtered_language.at[index, 'Body'] = row['Body'].split("****URL THAT USUALLY MEANS AN EMAIL END****")[0]
for index, row in df_filtered_language.iterrows():
df_filtered_language.at[index, 'Body'] = row['Body'].split("_")[0]
for index, row in df_filtered_language.iterrows():
df_filtered_language.at[index, 'Body'] = row['Body'].split("**** NAME OF THE PERSONA THAT MEANS EMAIL SIGNATURE *****")[0]
# +
#Concatenate Body values in list
result = df_filtered_language['Body'].str.cat(sep=' ')
result = result.replace("\r","")
result = result.replace("\n","")
result = result.replace("_","")
result = result.replace(".","")
result = result.replace('****URL THAT USUALLY MEANS AN EMAIL END****',"")
# -
#Save string to txt
with open("Output.txt", "w", encoding="utf-8") as text_file:
text_file.write(result)
#Write dataframe to csv for debugging purposes
df_filtered_language.to_csv("result.csv", sep=',')
|
Language Model Pre Processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Working with Compute
#
# When you run a script as an Azure Machine Learning experiment, you need to define the execution context for the experiment run. The execution context is made up of:
#
# * The Python environment for the script, which must include all Python packages used in the script.
# * The compute target on which the script will be run. This could be the local workstation from which the experiment run is initiated, or a remote compute target such as a training cluster that is provisioned on-demand.
#
# In this lab, you'll explore *environments* and *compute targets* for experiments.
#
# ## Connect to Your Workspace
#
# The first thing you need to do is to connect to your workspace using the Azure ML SDK.
#
# > **Note**: If the authenticated session with your Azure subscription has expired since you completed the previous exercise, you'll be prompted to reauthenticate.
# +
import azureml.core
from azureml.core import Workspace
# Load the workspace from the saved config file
ws = Workspace.from_config()
print('Ready to use Azure ML {} to work with {}'.format(azureml.core.VERSION, ws.name))
# -
# ## Prepare Data
#
# In this lab, you'll use a dataset containing details of diabetes patients. Run the cell below to create this dataset (if you already created it in a previous lab, the code will find the existing version.)
# +
from azureml.core import Dataset
default_ds = ws.get_default_datastore()
if 'diabetes dataset' not in ws.datasets:
default_ds.upload_files(files=['./data/diabetes.csv', './data/diabetes2.csv'], # Upload the diabetes csv files in /data
target_path='diabetes-data/', # Put it in a folder path in the datastore
overwrite=True, # Replace existing files of the same name
show_progress=True)
#Create a tabular dataset from the path on the datastore (this may take a short while)
tab_data_set = Dataset.Tabular.from_delimited_files(path=(default_ds, 'diabetes-data/*.csv'))
# Register the tabular dataset
try:
tab_data_set = tab_data_set.register(workspace=ws,
name='diabetes dataset',
description='diabetes data',
tags = {'format':'CSV'},
create_new_version=True)
print('Dataset registered.')
except Exception as ex:
print(ex)
else:
print('Dataset already registered.')
# -
# ## Create a Training Script
#
# Run the following two cells to create:
# 1. A folder for a new experiment
# 2. An training script file that uses **scikit-learn** to train a model and **matplotlib** to plot a ROC curve.
# +
import os
# Create a folder for the experiment files
experiment_folder = 'diabetes_training_logistic'
os.makedirs(experiment_folder, exist_ok=True)
print(experiment_folder, 'folder created')
# +
# %%writefile $experiment_folder/diabetes_training.py
# Import libraries
import os
import argparse
from azureml.core import Run
import pandas as pd
import numpy as np
import joblib
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import roc_auc_score
# Set regularization hyperparameter (passed as an argument to the script)
parser = argparse.ArgumentParser()
parser.add_argument('--regularization', type=float, dest='reg_rate', default=0.01, help='regularization rate')
args = parser.parse_args()
reg = args.reg_rate
# Get the experiment run context
run = Run.get_context()
# load the diabetes data (passed as an input dataset)
print("Loading Data...")
diabetes = run.input_datasets['diabetes'].to_pandas_dataframe()
# Separate features and labels
X, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values
# Split data into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# Train a logistic regression model
print('Training a logistic regression model with regularization rate of', reg)
run.log('Regularization Rate', np.float(reg))
model = LogisticRegression(C=1/reg, solver="liblinear").fit(X_train, y_train)
# calculate accuracy
y_hat = model.predict(X_test)
acc = np.average(y_hat == y_test)
print('Accuracy:', acc)
run.log('Accuracy', np.float(acc))
# calculate AUC
y_scores = model.predict_proba(X_test)
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
run.log('AUC', np.float(auc))
os.makedirs('outputs', exist_ok=True)
# note file saved in the outputs folder is automatically uploaded into experiment record
joblib.dump(value=model, filename='outputs/diabetes_model.pkl')
run.complete()
# -
# ## Define an Environment
#
# When you run a Python script as an experiment in Azure Machine Learning, a Conda environment is created to define the execution context for the script. Azure Machine Learning provides a default environment that includes many common packages; including the **azureml-defaults** package that contains the libraries necessary for working with an experiment run, as well as popular packages like **pandas** and **numpy**.
#
# You can also define your own environment and add packages by using **conda** or **pip**, to ensure your experiment has access to all the libraries it requires.
#
# Run the following cell to create an environment for the diabetes experiment.
# +
from azureml.core import Environment
from azureml.core.conda_dependencies import CondaDependencies
# Create a Python environment for the experiment
diabetes_env = Environment("diabetes-experiment-env")
diabetes_env.python.user_managed_dependencies = False # Let Azure ML manage dependencies
diabetes_env.docker.enabled = True # Use a docker container
# Create a set of package dependencies (conda or pip as required)
diabetes_packages = CondaDependencies.create(conda_packages=['scikit-learn'],
pip_packages=['azureml-defaults', 'azureml-dataprep[pandas]'])
# Add the dependencies to the environment
diabetes_env.python.conda_dependencies = diabetes_packages
print(diabetes_env.name, 'defined.')
# -
# Now you can use the environment for the experiment by assigning it to an Estimator (or RunConfig).
#
# The following code assigns the environment you created to a generic estimator, and submits an experiment. As the experiment runs, observe the run details in the widget and in the **azureml_logs/60_control_log.txt** output log, you'll see the conda environment being built.
# +
from azureml.train.estimator import Estimator
from azureml.core import Experiment
from azureml.widgets import RunDetails
# Set the script parameters
script_params = {
'--regularization': 0.1
}
# Get the training dataset
diabetes_ds = ws.datasets.get("diabetes dataset")
# Create an estimator
estimator = Estimator(source_directory=experiment_folder,
inputs=[diabetes_ds.as_named_input('diabetes')],
script_params=script_params,
compute_target = 'local',
environment_definition = diabetes_env,
entry_script='diabetes_training.py')
# Create an experiment
experiment = Experiment(workspace = ws, name = 'diabetes-training')
# Run the experiment
run = experiment.submit(config=estimator)
# Show the run details while running
RunDetails(run).show()
run.wait_for_completion()
# -
# The experiment successfully used the environment, which included all of the packages it required.
# Having gone to the trouble of defining an environment with the packages you need, you can register it in the workspace.
# Register the environment
diabetes_env.register(workspace=ws)
# ## Run an Experiment on a Remote Compute Target
#
# In many cases, your local compute resources may not be sufficient to process a complex or long-running experiment that needs to process a large volume of data; and you may want to take advantage of the ability to dynamically create and use compute resources in the cloud.
#
# Azure ML supports a range of compute targets, which you can define in your workpace and use to run experiments; paying for the resources only when using them. In this case, we'll run the diabetes training experiment on a compute cluster with a unique name of your choosing, so let's verify that exists (and if not, create it) so we can use it to run training experiments.
#
# > **Important**: Change *your-compute-cluster* to a unique name for your compute cluster in the code below before running it! Cluster names must be globally unique names between 2 to 16 characters in length. Valid characters are letters, digits, and the - character.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
cluster_name = "nikhilvmcluster"
try:
# Check for existing compute target
training_cluster = ComputeTarget(workspace=ws, name=cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
# If it doesn't already exist, create it
try:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS11_V2', max_nodes=2)
training_cluster = ComputeTarget.create(ws, cluster_name, compute_config)
training_cluster.wait_for_completion(show_output=True)
except Exception as ex:
print(ex)
# -
# Now you're ready to run the experiment on the compute you created. You can do this by specifying the **compute_target** parameter in the estimator (you can set this to either the name of the compute target, or a **ComputeTarget** object.)
#
# You'll also reuse the environment you registered previously.
# +
from azureml.train.estimator import Estimator
from azureml.core import Environment, Experiment
from azureml.widgets import RunDetails
# Get the environment
registered_env = Environment.get(ws, 'diabetes-experiment-env')
# Set the script parameters
script_params = {
'--regularization': 0.1
}
# Get the training dataset
diabetes_ds = ws.datasets.get("diabetes dataset")
# Create an estimator
estimator = Estimator(source_directory=experiment_folder,
inputs=[diabetes_ds.as_named_input('diabetes')],
script_params=script_params,
compute_target = cluster_name, # Run the experiment on the remote compute target
environment_definition = registered_env,
entry_script='diabetes_training.py')
# Create an experiment
experiment = Experiment(workspace = ws, name = 'diabetes-training')
# Run the experiment
run = experiment.submit(config=estimator)
# Show the run details while running
RunDetails(run).show()
run.wait_for_completion()
# -
# The experiment will take quite a lot longer because a container image must be built with the conda environment, and then the cluster nodes must be started and the image deployed before the script can be run. For a simple experiment like the diabetes training script, this may seem inefficient; but imagine you needed to run a more complex experiment with a large volume of data that would take several hours on your local workstation - dynamically creating more scalable compute may reduce the overall time significantly.
#
# While you're waiting for the experiment to run, you can check on the status of the compute in the widget above or in [Azure Machine Learning studio](https://ml.azure.com).
#
# > **Note**: After some time, the widget may stop updating. You'll be able to tell the experiment run has completed by the information displayed immediately below the widget and by the fact that the kernel indicator at the top right of the notebook window has changed from **⚫** (indicating the kernel is running code) to **◯** (indicating the kernel is idle).
#
# After the experiment has finished, you can get the metrics and files generated by the experiment run. The files will include logs for building the image and managing the compute.
# Get logged metrics
metrics = run.get_metrics()
for key in metrics.keys():
print(key, metrics.get(key))
print('\n')
for file in run.get_file_names():
print(file)
# **More Information**:
#
# - For more information about environments in Azure Machine Learning, see [Reuse environments for training and deployment by using Azure Machine Learning](https://docs.microsoft.com/azure/machine-learning/how-to-use-environments).
# - For more information about compute targets in Azure Machine Learning, see [What are compute targets in Azure Machine Learning?](https://docs.microsoft.com/azure/machine-learning/concept-compute-target).
|
mslearn-aml-labs/04-Working_with_Compute.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Create a python module (a file with extension ‘.py’) with the following functions:
#
# 1. (1 points) Write a python reads creates a dataframe from a URL that points to a CSV file such as the pronto data or CSVs in data.gov.
#
# 1. (6 points) Create the function test_create_dataframe that takes as input: (a) a pandas DataFrame and (b) a list of column names. The function returns True if the following conditions hold:
#
# - The DataFrame contains only the columns that you specified as the second argument.
# - The values in each column have the same python type
# - There are at least 10 rows in the DataFrame.
#
# read_url is the function read data from URL
#
# test_create_dataframe is the function testing the coditions
#
# ### Sample case
import hw2
url='https://data.seattle.gov/api/views/65db-xm6k/rows.csv?accessType=DOWNLOAD'
df = hw2.read_url(url)
df.head()
# ### Sample case when all conditions hold
colName = list(df.columns)
print(colName)
type_list = []
for i in range(len(df.columns)):
columns = df.iloc[:,i]
type0 = type(columns[0])
type_list.append(all(isinstance(columns[j],type0) for j in range(len(df))))
type_list
hw2.test_create_dataframe(df,colName)
# ### Test False Case
df.head()
df.iloc[1,1] = 'not number'
# ### Function inside hw2 module
import pandas as pd
def read_url(url):
return pd.read_csv(url)
def test_create_dataframe(df,colName):
# return true if all conditions satisfied
# The DataFrame contains only the columns that you specified as the second argument.
# The values in each column have the same python type
# There are at least 10 rows in the DataFrame.
if (list(df.columns.values) == colName) & len(set(df.dtypes.values.tolist())) == 1 & (len(df) >=10):
return True
i = 0
columns = df.iloc[:,i]
type0 = type(columns[0])
all(isinstance(columns[j],type0) for j in range(len(df)))
len(df.columns)
type(df.iloc[:,1][0])
# +
type_list = []
for i in range(len(df.columns)):
columns = df.iloc[:,i]
type0 = type(columns[0])
type_list.append(all(isinstance(columns[j],type0) for j in range(len(df))))
# -
all(type_list)
all([False,True])
|
hw2_test_case_demo_weikun.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="YyeVlSYkhahF" cellView="form"
#@title Imports, initial setup (Ctrl+F9 to run all)
import os
import re
import pandas as pd
import matplotlib.pyplot as plt
from scipy.signal import find_peaks
import copy
try:
import gamry_parser
except:
subprocess.run(
["pip", "install", "gamry-parser"],
encoding="utf-8",
shell=False)
finally:
import gamry_parser
gp = gamry_parser.CyclicVoltammetry()
print('Done.')
# + id="ZGoqracvk9q2" cellView="form"
"""
### SCRIPT CONFIGURATION SETTINGS ###
"""
#@markdown **Experimental Setup**
#@markdown Where should the notebook search for DTA files? Examples (using google colab):
#@markdown - Mounted google drive folder: `/content/drive/`
#@markdown - If uploading files manually, : `/content/`).
data_path = "/content/" #@param {type:"string"}
#@markdown Filter which files we want to analyze
file_pattern = "Search-For-Text" #@param {type:"string"}
#@markdown Extract trace labels from file name (e.g. `[17:].lower()` => drop the first 17 characters from the filename and convert to lowercase). The trace labels are used for category labeling (and plot legends)
file_label_xform = "[51:]" #@param {type:"string"}
# create a "results" dataframe to contain the values we care about
data_df = pandas.DataFrame()
settings_df = pandas.DataFrame()
peaks_df = pandas.DataFrame()
# identify files to process
files = [f for f in os.listdir(data_path) if
os.path.splitext(f)[1].lower() == ".dta" and
len(re.findall(file_pattern.upper(), f.upper())) > 0
]
# + cellView="form" id="8MFNF2Qz6lef"
#@markdown **Process Data and Detect Peaks**
#@markdown Which CV curves (cycle number) should be sampled? (`0` would select the first CV curve from each file)
curves_to_sample = "0" #@param {type:"string"}
curves_to_sample = [int(item.strip()) for item in curves_to_sample.split(",")]
#@markdown Peak Detection: specify the peak detection parameters
peak_width_mV = 75 #@param {type:"integer"}
peak_height_nA = 25 #@param {type:"integer"}
peak_thresh_max_mV = 800 #@param {type:"integer"}
peak_thresh_min_mV = -100 #@param {type:"integer"}
# this method finds the row that has an index value closest to the desired time elapsed
def duration_lookup(df, elapsed):
return df.index.get_loc(elapsed, method='nearest')
# iterate through each DTA file
for index, file in enumerate(files):
print("Checking File {}".format(file))
label, ext = os.path.splitext(file)
my_label = "-".join(eval("label{}".format(file_label_xform)).strip().split())
# load the dta file using gamry parser
gp.load(filename=os.path.join(data_path, file))
is_cv = gp.get_header().get("TAG") == "CV"
if not is_cv:
# if the DTA file is a different experiment type, skip it and move to the next file.
print("File `{}` is not a CV experiment. Skipping".format(file))
del files[index] # remove invalid file from list
continue
# for each CV file, let's extract the relevant information
cv = gamry_parser.CyclicVoltammetry(filename=os.path.join(data_path, file))
cv.load()
for curve_num in curves_to_sample:
print("\tProcessing Curve #{}".format(curve_num))
v1, v2 = cv.get_v_range()
settings = pandas.DataFrame({
"label": my_label,
"curves": cv.get_curve_count(),
"v1_mV": v1*1000,
"v2_mV": v2*1000,
"rate_mV": cv.get_scan_rate(),
}, index=[0])
settings_df = settings_df.append(settings)
data = copy.deepcopy(cv.get_curve_data(curve=curve_num))
data.Im = data.Im*1e9
data.Vf = data.Vf*1e3
data["label"] = my_label #"{:03d}-{}".format(index, curve_num)
data_df = data_df.append(data)
# find peaks in the data
dV = cv.get_scan_rate() # in mV
peak_width = int(peak_width_mV/dV)
peaks_pos, props_pos = find_peaks(
data.Im,
width=peak_width,
distance=2*peak_width,
height=peak_height_nA
)
peaks_neg, props_neg = find_peaks(
-data.Im,
width=peak_width,
distance=2*peak_width,
height=peak_height_nA
)
peaks = list(peaks_pos) + list(peaks_neg)
# remove peaks that are out of min/max range
peaks = [peak
for peak in peaks
if data.Vf.iloc[peak] >= peak_thresh_min_mV and data.Vf.iloc[peak] <= peak_thresh_max_mV]
# add detected peaks to aggregated peak dataframe
peaks = data.iloc[peaks].sort_values(by="Vf")
peaks["index"] = peaks.index
peaks.reset_index(level=0, inplace=True)
peaks_df = peaks_df.append(peaks)
peaks_df = peaks_df[["label", "index", "Vf", "Im"]]
# print("\tdetected peaks (mV)", [int(peak) for peak in data.iloc[peaks].Vf.sort_values().tolist()])
print("\nFile Metadata")
print(settings_df.to_string(index=False))
print("\nPeaks Detected")
print(peaks_df.to_string(index=False))
# + id="Ulne80RrpBrW" cellView="form"
#@markdown **I-V plot**: Overlay the loaded CyclicVoltammetry Curves
from plotly.subplots import make_subplots
import plotly.graph_objects as go
from plotly.colors import DEFAULT_PLOTLY_COLORS
fig = make_subplots(rows=1, cols=1, shared_xaxes=True, vertical_spacing=0.02)
for (index, exp_id) in enumerate(data_df.label.unique()):
data = data_df.loc[data_df.label == exp_id]
newTrace = go.Scatter(
x=data.Vf,
y=data.Im,
mode='lines',
name=exp_id,
legendgroup=files[index],
line=dict(color=DEFAULT_PLOTLY_COLORS[index]),
)
fig.add_trace(newTrace, row=1, col=1)
peak = peaks_df.loc[peaks_df.label == exp_id]
newTrace = go.Scatter(
x=peak.Vf, y=peak.Im,
mode="markers",
showlegend=False,
marker=dict(size=12,
color=DEFAULT_PLOTLY_COLORS[index],
)
)
fig.add_trace(newTrace, row=1, col=1)
layout = {
'title': {'text': 'Cyclic Voltammetry Overlay',
'yanchor': 'top',
'y': 0.95,
'x': 0.5 },
'xaxis': {
'anchor': 'x',
'title': 'voltage, mV'
},
'yaxis': {
'title': 'current, nA',
'type': 'linear'
''
},
'width': 1200,
'height': 500,
'margin': dict(l=30, r=20, t=60, b=20),
}
fig.update_layout(layout)
config={
'displaylogo': False,
'modeBarButtonsToRemove': ['select2d', 'lasso2d', 'hoverClosestCartesian', 'toggleSpikelines','hoverCompareCartesian']
}
fig.show(config=config)
|
demo/notebook_cyclicvoltammetry_peakdetect.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 其他的迭代环境
# ### 1. for 循环使用了迭代协议;
# ### 2. 任何从左到右扫描对象的工具都使用迭代协议;
for line in open('untitled3.py'):
print(line.upper(), end = '')
# ### 列表解析,in的子集测试,内置函数map, sorted, zip 都会使用迭代协议
uppers = [line.upper() for line in open('untitled3.py')] #列表解析
uppers
'@AUTHOR: NICKC\n' in open('untitled3.py').read().upper() # in的子集测试
map(str.upper, open('untitled3.py')) # map内置函数
list(map(str.upper, open('untitled3.py')))
f = open('untitled3.py')
enumerate(map(f.readline, open('untitled3.py'))) # map的function选项取决于后面作用对象的类型
open('untitled3.py').readline()
[line for line in open('untitled3.py')]
def function(x):
return x+' \n'+ x
list(map(function, open('untitled3.py')))
from sympy import tanh
def function(x):
return tanh(x).evalf()
list(map(function,list(range(1,101))))
# ### map很像列表解析,但是没有列表解析强大,map需要函数而列表解析可以是任意的表达式
S = """Python includes various additional built-ins that process iterables, too: sorted sorts
items in an iterable, zip combines items from iterables, enumerate pairs items in an
iterable with relative positions, filter selects items for which a function is true, and
reduce runs pairs of items in an iterable through a function. All of these accept iterables,
and zip, enumerate, and filter also return an iterable in Python 3.0, like map. Here they
are in action running the file’s iterator automatically to scan line by line:""" # S 是个字符串
L = S.split()
print(L, end='')
print(sorted(L), end = '') # 字符串首先是从1-100,从A到Z, 再从a到z
L1 = []
from numpy import random
for i in range(100):
L1.append(random.randint(100))
print(L1, end = '' )
print(sorted(L), end= '')
L2 = []
for i in range(100,200):
L2.append(random.randint(100,200))
print(L2, end='')
print(list(zip(L1, L2)), end= '')
print(list(enumerate(S.split(sep='.'))))
print(list(filter(str.isalpha, S.split()))) # 当str.isalpha是正确的时候,元素就会过滤起来
sorted(open('untitled3.py'))
list(zip(open("untitled3.py"), open("untitled3.py")))
list(enumerate(open("untitled3.py")))
bool('0'), bool(0), bool('1'), bool('')
list(filter(bool, open('untitled3.py')))
bool('0'), bool(0), bool('1'), bool('')
if 0:
print("Hello world")
if 1:
print("Hello world")
if bool('0'):
print("Hello world")
if bool(1):
print("Hello world")
bool(""), bool(''), bool("""""")
import functools, operator
functools.reduce(operator.add, open('untitled3.py'))
# +
# functools.reduce?
# -
S = 'import sys\n' + 'print(sys.path)'
S
# ### from sympy import tanh # tanh 就不是内置函数
L = list(range(100))
from numpy import random
L1 = []
L2 = []
for i in range(100):
L1.append(random.randint(1,101)) # int 就是integer的缩写, integer是整数的意思
for j in range(100):
L2.append(random.randint(1,101))
print(L1, end=''); print('\n'); print(L2, end='')
type(L1.sort()); print('\n');print(sorted(L2), end='')
# ### L1.sort() 对对象本身是即时生效的
print(L1) # L1.sort() 对对象本身是即时生效的
# ### 不是即时生效的例子
a = L1
print( 'a = '+ str(a), end='')
print('\n')
b = a # b是等于列表L1, 而不是a这个名字
print('b = ' + str(b), end ='')
print('\n')
a = a + L2
print('a = ' + str(a), end= "")
print('\n')
print(b, end='')
sum([i for i in range(101)])
D = dict(enumerate(L1))
type(D), type(L1), type(tuple(L1))
print(D, end ='')
sum(D) == (0+99)*100/2
# ### any函数所作用的对象中只要有一项的布尔值为True就返回True
any(['',''])
all(['spam','','ni'])
# +
# any?
# -
S = 'python'
L = []
for i in S:
L.append(i)
L
max(L)
min(L)
for i in L:
print('%s = ' % i + str(ord(i)))
max(['A','a'])
print(ord('A'), ord('a'))
# ### 52个英文字母(包含大小写) + 键盘上的其他按键都对应一个ASCII码
bin(65), hex(65), oct(65)
print(ord('x'), ord('='), ord('2'), ord('\n'),ord('x')+ord('=')+ord('2')+ord('\n') )
S = 'import sys\n'
list((i for i in S))
print([ord(i) for i in list((i for i in S))])
f = open('script1.py', mode='w')
f.write('import sys\n')
f.write('print(sys.path)\n')
f.write('x = 2\n')
f.write('print(2**33)\n')
f.close()
f = open('script1.py', 'r')
f.read()
f.close()
# # %load script1.py
import sys
print(sys.path)
x = 2
print(2**33)
max(open('script1.py'))
ord('i'), ord('p'), ord('x')
L1 = list(range(1,101,1))
L2 = []
for i in L1:
L2.append(str(i))
S = '+'.join(L2)
type(L1[0])
eval(S)
S
import functools, operator
functools.reduce(operator.add, L1)
print(L1, end='')
print('\n')
print(L2, end='')
# ### Sympy 完全是python写的,并不像看起来的那样上流
from sympy import *
init_session()
Eq(Sum(n,(n,1,100)), Sum(n,(n,1,100)).doit())
Eq(Product(n, (n,1,100)), Product(n, (n,1,100)).doit())
factorial(100)
L1 = list(range(1,101,1))
functools.reduce(operator.add, L1)
L1 = list(range(1,101,1))
I = functools.reduce(operator.mul, L1)
I
# # %load script1.py
import sys
print(sys.path)
x = 2
print(2**33)
list(open('script1.py'))
tuple(open('script1.py'))
a, b, c, d= open('script1.py')
a, d
a, *b = open('script1.py')
type(a), type(b)
print(a); print('\n'); print(b)
set(open('script1.py'))
# +
# set?
# -
set()
L1
print(set(L2)) #对于字符串来说,它返回的是无序的
print(set(L1)) #对于整数来说,它返回的是有序的
# ### 创造的是一个无序的元素独一无二的集合,而且作用于不同的对象的话,得到的结果也是不一样的
set('asafgjasa') # 字符串
D = dict(enumerate(L2))
set(D) # set它把字典当中的关键词提取出来,创造新的集合
D2 = {i : i for i in L2}
print(set(D2),end='')
print(set(set(D2))) # set为什么返回的结果与预想不一样
print(set(tuple(L2))) # 元组
print(set("""Strictly speaking, the max and min functions can be applied to files as well—they automatically use the iteration protocol to scan the file and pick out the lines with the highest
and lowest string values, respectively though I’ll leave valid use cases to your
imagination"""), end='')
print(set("""石正丽等中国科学家发现:进化的“军备竞赛”(arms race)塑造了病毒及其受体的多样性。
鉴定涉及种间传播的关键残基对于预测潜在的病原体、了解病毒如何从野生动物向人类跃迁,非常重要。
以前,研究者已经在中华菊头蝠中鉴定出具有不同遗传特征的SARS相关冠状病毒(SARSr-CoV)。而这份最新研究还展现了中华菊头蝠种群中蝙蝠受体ACE2(血管紧张素转化酶2)的高度多样性。这些ACE2变体支持SARS病毒和SARS相关冠状病毒的感染,但对不同刺突蛋白具有不同的结合亲和力。
SARS相关冠状病毒刺突蛋白对人ACE2拥有更高结合亲和力,显示这些病毒具有向人类跃迁传染的能力。ACE2和SARS相关冠状病毒刺突蛋白之间的界面处残基的正向选择,表明它们之间存在长期和持续的协同进化动力学。因此,持续监视蝙蝠中的这一组病毒对于预防下一个SARS样疾病非常必要。
以上研究来自中科院武汉病毒所石正丽团队与福建师范大学生命科学学院欧阳松应教授在预印本平台 bioRxiv 上发表的论文:Evolutionary arms race between virus and host drives genetic diversity in bat SARS related coronavirus spike genes。
中华菊头蝠是SARS病毒的宿主,其体内还携带多种SARS相关冠状病毒。这些病毒具有高度的遗传多样性,尤其是病毒的刺突蛋白基因。尽管有着不同程度的变异,一些蝙蝠SARS相关冠状病毒仍可以利用人类受体ACE2进入人体细胞。研究者推测,蝙蝠的ACE2受体和SARS相关冠状病毒刺突蛋白之间,有着相互作用,而这驱动了SARS相关冠状病毒的遗传多样性。
研究者鉴定出了一系列中华菊头蝠ACE2变异体,这些变异体中有一些与SARS-CoV刺突蛋白有相互作用的多态位点。携带不同刺突蛋白的伪病毒或SARS相关冠状病毒,在表达了蝙蝠ACE2变体的细胞中有着不同的瞬时感染效率。通过测定SARS病毒、SARS相关冠状病毒刺突蛋白与蝙蝠受体、人类受体分子之间的结合亲和力,能观察到相关的结果。
所有被测试的蝙蝠SARS相关冠状病毒刺突蛋白与人ACE2的结合亲和力,均高于其对蝙蝠ACE2的结合亲和力。不过SARS相关冠状病毒刺突蛋白与人ACE2的结合亲和力,比SARS-CoV刺突蛋白与人ACE的亲和力低10倍。
结构建模表明,刺突和ACE2之间的结合亲和力差异可能是由于这两个分子界面中某些关键残基的改变而引起。分子进化分析表明,这些残基处于强的正选择。
这些结果表明SARS新冠病毒刺突蛋白和蝙蝠ACE2可能随着时间的推移而互相进化,并经历彼此的选择压力,从而触发了进化的“军备竞赛”动力学。这进一步证明了,中华菊头蝠是SARS相关冠状病毒的天然宿主。
冠状病毒是包膜病毒,包含单股正链RNA。该亚科有四个属,即α、β、γ和δ。α冠状病毒和β冠状病毒起源于蝙蝠或啮齿动物,而γ冠状病毒和δ冠状病毒起源于鸟类。自21世纪初以来,三种β型冠状病毒已引起人类严重肺炎暴发。分别是SARS-CoV,MERS-CoV和SARS-CoV-2。
SARS-CoV-2引发的疫情使人们回想起17年前发生的SARS疫情。SARS是一种人畜共患病,在接下来的几年中,科学家从中国和欧洲不同地区的蝙蝠中检测或分离出了具有不同遗传特征的75种SARS相关冠状病毒(SARSr-CoV)。
蝙蝠SARS相关冠状病毒与人类和果子狸的SARS-CoVs有96%的核苷酸序列相似度,其中可变区最多的是刺突蛋白(S)和辅助蛋白ORF3和ORF8。此外,研究者已经确定了不同蝙蝠SARS相关冠状病毒基因组中能找到SARS-CoV的所有基因构建基块,这表明SARS病毒的祖先是通过蝙蝠SARS相关冠状病毒基因组的重组而来,其起源于蝙蝠。
病毒感染的第一步是识别细胞受体,这也是必不可少的步骤。冠状病毒的进入是由病毒刺突蛋白(Spike,S)和细胞表面受体之间的特异性相互作用介导,然后病毒与宿主膜之间发生融合。冠状病毒刺突蛋白在功能上分为两个亚基:细胞附着亚基(S1)和膜融合亚基(S2)。 S1区域包含N端结构域(NTD)和C端结构域(CTD);两者均可用于冠状病毒受体结合(RBD)。
对于SARS-CoV,其S1-CTD作为RBD与细胞的受体即血管紧张素转换酶2(ACE2)结合。冷冻电镜和晶体结构分析,确定了SARS病毒的S-RBD与人ACE2之间界面中的一些关键残基。
根据S蛋白的大小,蝙蝠SARS相关冠状病毒可以分为两个不同的进化枝。进化枝1包含病毒具有与SARS病毒大小相同的刺突蛋白。而由于5、12或13个氨基酸缺失,属于进化枝2的病毒其刺突蛋白则比SARS病毒的小。
尽管RBD有所不同,所有进化枝1毒株都可以使用ACE2进入细胞,而进化枝2毒株则由于上述缺失无法直接进入。这些结果表明,就基因组相似性和ACE2的使用而言,进化枝1的成员很可能是SARS病毒的直接来源。
ACE2在功能上分为两个结构域:N末端结构域参与SARS-CoV结合,C末端结构域参与心功能的调节。先前的结果表明,不同来源的ACE2的C末端结构域相对保守,而N末端结构域在物种间显示出更多的多样性。此前已证明SARS病毒可以利用水鼠耳蝠的ACE2和中华菊头蝠的ACE2。RBD结合位点中的微小突变,可将ACE2从对SARS-CoV结合不易感转变为易感。由于属于进化枝1的所有SARS相关冠状病毒都可从中华菊头蝠体内提取出来,而且也都可以利用ACE2,因此研究者提出问题:中华菊头蝠ACE2中的变异是否可能有导致了蝙蝠SARS相关冠状病毒的多样性。
研究团队研究了中华菊头蝠ACE2基因的多态性,并通过分子进化分析,蛋白质亲和力测定和病毒感染测定相结合,评估了它们对不同蝙蝠SARS相关冠状病毒刺突蛋白的敏感性和结合亲和力。
结果表明,SARS相关冠状病毒的刺突蛋白多样性可能会受到中华菊头蝠ACE2变体的自然选择压力; 在长期共存期间,SARSr-CoV刺突蛋白可能会被中华菊头蝠的ACE2选择,以维持自身遗传多样性并适合中华菊头蝠的种群。
ACE2基因在中华菊头蝠种群中表现出高度多态性
根据蝙蝠SARS相关冠状病毒的流行情况以及样品组织的可用性和质量,研究者使用来自三个省(湖北,广东和云南)的样品进行ACE2扩增。
除了团队先前测序过的蝙蝠ACE2(分别从湖北,广西和云南收集的样本ID 832、411和3357)和其他蝙蝠ACE2(GenBank登记号ACT66275,这是从香港收集的样本)外,研究者从21只中华菊头蝠蝠个体中获得了ACE2基因序列:湖北有5个,广东有9个,云南有7个。这些蝙蝠ACE2序列在其物种内显示98-100%的氨基酸同一性,与人ACE2的显示80-81%的氨基酸同一性。
这些蝙蝠ACE2在N端区域观察到了主要变化,包括一些先前已确定与SARS病毒的 S-RBD接触的残基。根据非同义SNP分析鉴定出8个残基,包括24、27、31、34、35、38.41和42。这8个残基的组合产生了8个等位基因,包括RIESEDYK,LIEFENYQ,RTESENYQ,RIKSEDYQ,QIKSEDYQ, RMTSEDYQ,EMKT KDHQ和EIKT EIKTKDHQ,分别命名为等位基因1-8。
除了先前研究(等位基因4、7和8)中的ACE2基因型数据外,研究者在中华菊头蝠种群中还鉴定出5个新的等位基因。“等位基因2”在两个省的样本中有发现,“等位基因4”在3个省中有发现,而其他等位基因似乎在地理上受到限制。总之,在广东发现了3个等位基因(4、6和8),云南发现了4个等位基因(1、2、4和7),在湖北发现了3个等位基因(2、4和5),在广西和香港分别找到了1个等位基因。在发现SARS病毒直接祖先的云南一蝙蝠洞中,研究者发现了4个等位基因共存。
综上所述,这些数据表明ACE2变异体已经在不同地区的中华菊头蝠种群中长期存在。与SARS病毒的S-RBD直接接触的位点的取代,表明它们在SARS病毒的进化和传播过程中可能具有重要功能。"""), end ='')
{ line for line in open('script1.py')}
ord('i'), ord('p'), ord('x')
# # %load script1.py
import sys
print(sys.path)
x = 2
print(2**33)
ord('7'), ord('2')
{ix: line for ix, line in enumerate(open('script1.py'))}
{line for line in open('script1.py') if line[0] == 'p'}
{ix: line for (ix, line) in enumerate(open('script1.py'))}
ord('s') # 函数
def f(a, b,c, d):print(a,b,c,d,sep=',')
f(1,2,3,4)
f(*[1,2,3,45]) # a, b, c, d是单个参数
a, *b = open('script1.py')
b
def f(a,b,c,d): print(a*2, b*3, c*4, d*5, end='\n' )
f(*open('script1.py'))
# # %load script1.py
import sys
print(sys.path)
x = 2
print(2**33)
X = (1,2)
Y = (3,4)
zip(X,Y)
tuple(zip(X,Y)); list(zip(X,Y))
zip(zip(X,Y))
A, B = zip(*zip(X,Y))
print(A, end='\n'); print(B, end='\n')
C, D = zip(X,Y) # X = (1,2) Y = (3,4)
print(C, end='\n'); print(D, end='\n')
A, B = zip(C,D)
A, B = zip(zip(X,Y)) #
print(A, end='\n'); print(B, end='\n')
A, B = zip(X,Y)
C = A, B
C
E, F= zip(C,)
print(E, end='\n'); print(F, end='\n')
# ##### 什么叫字典视图对象?
# ### Python3.0中的新的可迭代对象
# Python 3 的版本比Python2.x 更加强调迭代
zip('abc','xyz') # 在Python3.x中返回的是可迭代的对象,而在2.x版本中返回是一个列表
list(zip('abc','xyz'))
# ### range迭代器
R = range(10)
R
I = iter(R)
next(I)
next(I), next(I), next(I), next(I), next(I), next(I), next(I)
# ### range 对象只支持迭代,索引,长度函数,它们不支持其他的序列操作
len(R), R[0], R[-1], next(I), I.__next__()
# #### 版本偏差,xrange, file.xreadlines() 只适用于python 2.x; range和open在python 3.x取而代之
range(10), map(list(range(10)), list(range(10,20))), zip(list(range(10)),list(range(10,20))), filter(bool, open('script1.py'))
f = open('script1.py')
f is iter(f)
R = range(3); M = map(abs, list(range(3,6))); Z = zip(list(range(3)),list(range(3,6))); F = filter(bool, open('script1.py'))
R is iter(R), M is iter(M), Z is iter(Z), F is iter(F)
# #### 除了range, map和zip,以及filter都是自身的迭代器(它们在一次遍历之后,就不能在进行访问了)
f.read()
f.read() # 一次迭代完全后,它们将会迭代完全
list(map(abs,(-1,0,1)))
Z = zip((1,2,3),(10,20,30))
print('%s != ' % str(next(Z)), next(Z))
next(Z), next(Z), next(Z)
# ### map,以及zip还有filter都是一次性迭代器,像文件迭代器一样,如果一次性遍历完之后,就会给出停止迭代的错误,
for pair in Z: print(pair)
Z = zip((1,2,3),(10,20,30))
for pair in Z: print(pair)
for pair in Z: print(pair)
# +
def f(x):
if bool(x) == False:
return x
else:
pass
list(filter(f,['spam','','ni']))
# -
type(False)
list('')
list(filter(bool,['spam','','ni']))
# #### filter, map, zip它们既可以处理可迭代对象,也可以产生一个可迭代对象
# #### range, 字典视图对象不能处理可迭代对象,但是可以产生一个可迭代对象
range(1,101,2)
list(range(1,101,2))
# ### 多个迭代器 vs 单个迭代器
R = range(3)
R is iter(R)
Z = zip((1,2,3),(10,12,13))
I1 = iter(Z)
I2 = iter(Z)
Z is iter(Z), I1 is Z, I2 is Z
next(I1), next(I1), next(I2)
M = map(abs, (-1,0,1))
I1 = iter(M); I2 = iter(M)
print(next(I1),next(I2),next(I1))
# next(I2)
R = range(3)
I1, I2 = iter(R), iter(R)
R is iter(R), I1 is R, I2 is R
[next(I1), next(I2), next(I1)]
class Text():
"""
This is just an experiment performed for iterations related to classes
"""
def PLUS(x,y):
return str(x)+str(y)
"""
PLUS can be used to convert two items to strings and add them together, and return the result.
"""
def SUBSTR(x,y):
return x[:y]
Text.PLUS(1,2)
Text.SUBSTR('asahfjasfhajfjah',4)
Text.PLUS(5,6789)
# ### 文件,字典以及字典的方法(keys,values,items) 都和range,map,zip,filter这些函数一样都会返回可迭代对象
S = """As we saw briefly in Chapter 8, in Python 3.0 the dictionary keys, values, and items
methods return iterable view objects that generate result items one at a time, instead
of producing result lists all at once in memory. View items maintain the same physical
ordering as that of the dictionary and reflect changes made to the underlying dictionary.
Now that we know more about iterators, here’s the rest of the story:"""
D = dict(enumerate(S))
print(list(D.keys()), end='')
I = iter(D)
next(I), next(I), next(I), next(I), next(I)
for key in D.keys(): print(key,end=' ')
print('\n')
for key in D: print(key, end=' ')
import numpy as np
D = dict(a=1,c=3,b=3)
print(D)
D = {'a': 1, 'c': 3, 'b': 2}
print(D)
D = { s : int(i) for s, i in zip('acb', '132')}
print(D)
D = {}
for i in [1,2,3]:
for s in 'abc':
D[s] = i
D
for k in D: print(k, D[k], end=' ') #并不按照ASCII的顺序
for k in sorted(D.keys()): print(k,D[k], end=' ')
for k in sorted(D): print(k, D[k], end= ' ')
help(Text)
|
Chapter 14 Iterations and Comprehensions (2).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # COMP 534 - Applied Artificial Intelligence
# ## CA1 - Binary classification
# This notebook was produced as a deliverable for a group project for the above module, as part of the 2021-2022 Data Science and Artificial Intelligence MSc course at the University of Liverpool. It comprises a performance analysis of three supervised machine learning methods for solving a binary classification problem.
# ### Preparation
# #### Setup
# Import required libraries, tools and classifiers.
# +
# Data handling and mathematical tools
import pandas as pd
import numpy as np
# Creating plots
import matplotlib.pyplot as plt
import seaborn as sns
# Various tools from sklearn
# Data preparation
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
# Machine learning algorithms
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
# Hyperparameter optimisation
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.model_selection import GridSearchCV
# Performance evaluation
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
# -
# #### Data
# The performance evaluation is based on analysis of the Pima Indian Diabetes Dataset, from the American National Institute of Diabetes and Digestive and Kidney Diseases. The aim of the analyses is to predict whether or not an individual has diabetes based on a variety of measurements. The dataset is available for download on Kaggle.
# https://www.kaggle.com/uciml/pima-indians-diabetes-database (last accessed 12/03/22)
# Read data from csv
df = pd.read_csv('diabetes.csv')
# Assign feature and target column headers for handling
features = list(df.columns[:-1])
target = df.columns[-1]
# We can visualise the distribution of features in the dataset and begin to understand their relative importance in predicting the presence of diabetes by plotting as below.
# Create 8 subplots, 1 for each feature
fig, axs = plt.subplots(1, 8, figsize=(12, 4))
# Plot distribution of each feature as a violin plot broken down by class label
for i, ax in enumerate(axs):
sns.violinplot(ax=ax, data=df, x='Outcome', y=features[i])
fig.tight_layout()
# There are differences between the distributions of the two classes for each feature, suggesting all features will be useful in predicting the presence of diabetes. The distinction between each class's distribution is more marked for some features than others, specifically pregnancies, glucose and age.
#
# It is also apparent that a number of features have a significant proportion of records with zero values, particularly skin thickness and insulin. Pregnancies, glucose, blood pressure, skin thickness, insulin and BMI all contain at least one zero value. This is valid for number of pregnancies but thought to be invalid otherwise. As a result a data cleaning step will be performed to account for these values.
# #### Cleaning
# For feature values of zero in glucose, blood pressure, skin thickness, insulin and BMI, the value will be replaced by the mean of that feature of the subset of records belonging to the same class and excluding zero values.
# Helper function to work with data cleaning lambda
# For passed feature, updates 0 values to mean of feature for objects of same class
# Takes:
# - row: row of dataframe
# - feature: df column name
# - df: dataframe
def dataCleaningHelper(row, feature, df):
# Check if feature value at row = 0 and class label of row = 0
if row[feature] == 0 and row['Outcome'] == 0:
# Update feature value to mean of feature for class label 0
return df[(df['Outcome'] == 0) & (df[feature] != 0)][feature].mean()
# Check if feature value at row = 0 and class label of row = 1
elif row[feature] == 0 and row['Outcome'] == 1:
# Update feature value to mean of feature for class label 1
return df[(df['Outcome'] == 1) & (df[feature] != 0)][feature].mean()
# Otherwise no update
else:
return row[feature]
# Iterate over selected column headers
for feature in ['Glucose', 'BloodPressure', 'SkinThickness', 'Insulin', 'BMI']:
# Replace feature values of 0 with mean feature value of objects in same class
df[feature] = df.apply(lambda row: dataCleaningHelper(row, feature, df), axis=1)
# ### Models and Analysis
# The dataset is now ready to be used for development of our machine learning methods. The same procedure will be repeated for each method to enable our evaluation.
# #### Setup
# Hyperparameter tuning will done using grid search cross validation over a defined hyperparameter search space. Several evaluation metrics will be computed from the final test results for each method.
# Function to optimise algorithm hyperparameters
# Performs grid search with cross validation to evaluate hyperparameter combinations
# Takes:
# - algorithm: machine learning model object
# - searchSpace: dictionary containing hyperparameter names (keys) and list of values to search through (values)
# - X_train: training data feature values
# - y_train: training data class labels
def hyperparameterTuner(algorithm, searchSpace, X_train, y_train):
# Initialise cross validator
# Repeated to account for variance across samples
# Stratified to ensure constant proportion of each classes in each fold
crossValidator = RepeatedStratifiedKFold(n_splits=5, random_state=1, n_repeats=3)
# Initialise search
# Grid search to perform exhaustive search of all hyperparameter combinations in searchSpace
hyperparameterSearch = GridSearchCV(algorithm, searchSpace, scoring='accuracy', n_jobs=-1, cv=crossValidator)
# Fit search with training data
# Performs search over searchSpace
hyperparameterSearch.fit(X_train, y_train)
return hyperparameterSearch
# Helper function to compute and store evaluation metrics
# Takes:
# - confusion matrix: 2x2 array
# - name of algorithm used: str
def getEvaluationMetrics(confusionMatrix, algorithmName):
# Extract counts of each result category from confusion matrix
[tp, fn], [fp, tn] = confusionMatrix
# Compute evaluation metrics
accuracy = (tp+tn)/(tp+tn+fp+fn)
precision = tp/(tp+fp)
recall = tp/(tp+fn)
fScore = (2*precision*recall)/(precision+recall)
# Return metrics as dictionary
return {'Algorithm':algorithmName, 'Accuracy':accuracy, 'Precision':precision, 'Recall':recall, 'F-Score':fScore}
# Initialise empty dataframe to store model evaluation metrics
evaluationDf = pd.DataFrame(columns=['Algorithm', 'Accuracy', 'Precision', 'Recall', 'F-Score'])
# #### K-Nearest Neighbours Classifier
# ##### Data preparation
# +
# Split data into 80% train and 20% test sets
# Data stratified by class due to imbalance in classes (65% negative - 35% positive)
# Data shuffled to reduce any bias in the order of the dataset
# Define random state to ensure the same train test split can be used for each method
X_train, X_test, y_train, y_test = train_test_split(df[features],
df[target],
test_size=0.20,
stratify=df[target],
shuffle=True,
random_state=123,
)
# KNN method detrimentally impacted by differences between the units of features
# Normalise to account for this - updated features in range [0, 1]
scaler = MinMaxScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# -
# ##### Baseline model using default hyperparameters
# Create KNN classifier using default hyperparameters
knnBaselineModel = KNeighborsClassifier()
# Train model
knnBaselineModel.fit(X_train, y_train)
# Test model
knnBaselinePredictions = knnBaselineModel.predict(X_test)
# ##### Hyperparameter search space
# +
# Define KNN hyperparameter search space
# Default values for parameters are shown in comments
searchSpace = {
# 'algorithm': 'auto',
# 'leaf_size': 30,
# 'metric': 'minkowski',
# 'metric_params': None,
# 'n_jobs': None,
'n_neighbors': list(range(3,25,2)), # 'n_neighbors': 5,
'p': [1, 2, 3], # 'p': 2,
'weights': ['uniform', 'distance'] # 'weights': 'uniform'}
}
# Store dictionary of default hyperparameters used in search for plotting later
defaultParameters = {
'n_neighbors': 5,
'p': 2,
'weights': 'uniform'
}
# -
# ##### Perform search
# Create KNN classifier for hyperparameter search
knnSearchModel = KNeighborsClassifier()
# Perform search based on search space
search = hyperparameterTuner(knnSearchModel, searchSpace, X_train, y_train)
# Store best parameters found
optimumParameters = search.best_params_
# Store all results in new dataframe
results = pd.DataFrame(search.cv_results_)
# ##### Plot search results
# +
# Create figure for results
fig, ax = plt.subplots()
# Lineplot to display trend for each hyperparameter investigated
sns.lineplot(
data=results,
y='mean_test_score',
x='param_n_neighbors',
hue='param_p',
style='param_weights',
palette="Set1",
ax=ax,
)
# Add point to highlight best accuracy achieved
ax.scatter(optimumParameters['n_neighbors'], results[results['params'] == optimumParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Optimised',
marker='o'
)
# Add point to show accuracy achieved using default hyperparameters
ax.scatter(defaultParameters['n_neighbors'], results[results['params'] == defaultParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Default',
marker='^'
)
# Move legend outside figure
ax.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
# Fix x axis markers to whole numbers
ax.xaxis.get_major_locator().set_params(integer=True)
# Add title
fig.suptitle('KNN accuracies across hyperparameter search space', fontsize=12)
fig.savefig('kNN_hyperparameterSearchResults', bbox_inches='tight')
plt.show()
# -
# ##### Test best hyperparameters
# Create KNN classifier using best parameters
knnBestModel = KNeighborsClassifier(
n_neighbors=optimumParameters['n_neighbors'],
p=optimumParameters['p'],
weights=optimumParameters['weights']
)
# Train model
knnBestModel.fit(X_train, y_train)
# Test model
knnBestPredictions = knnBestModel.predict(X_test)
# ##### Compare baseline and optimised accuracies
print(f'Baseline model test accuracy: {100*((knnBaselinePredictions==y_test).sum()/y_test.shape[0]):.2f}%')
print(f'Optimised model test accuracy: {100*((knnBestPredictions==y_test).sum()/y_test.shape[0]):.2f}%')
# ##### Evaluation metrics
# Create confusion matrix for KNN optimised hyperparameter test results
confusionMatrixKNN = confusion_matrix(y_test, knnBestPredictions, labels=[1, 0])
# Store evaluation metrics in dataframe
evaluationDf = evaluationDf.append(getEvaluationMetrics(confusionMatrixKNN, 'k-Nearest Neighbours'), ignore_index=True)
# #### Random Forest
# ##### Data preparation
# Repeat data splitting process
# Use same random state to ensure the same train test split
X_train, X_test, y_train, y_test = train_test_split(df[features],
df[target],
test_size=0.20,
stratify=df[target],
shuffle=True,
random_state=123,
)
# ##### Baseline model using default hyperparameters
# Create random forest classifier using default hyperparameters
rfBaselineModel = RandomForestClassifier()
# Train model
rfBaselineModel.fit(X_train, y_train)
# Test model
rfBaselinePredictions = rfBaselineModel.predict(X_test)
# ##### Hyperparameter search space
# +
# Define Random Forest hyperparameter search space
# Default values for parameters are shown in comments
searchSpace = {
# 'bootstrap': True,
# 'ccp_alpha': 0.0,
# 'class_weight': None,
'criterion': ['gini', 'entropy'], # 'criterion': 'gini',
# 'max_depth': None,
'max_features': [3, 4, 5, 6, 7, 8], # 'max_features': 'auto' (= square root of number of features)
# 'max_leaf_nodes': None,
# 'max_samples': None,
# 'min_impurity_decrease': 0.0,
# 'min_samples_leaf': 1,
# 'min_samples_split': 2,
# 'min_weight_fraction_leaf': 0.0,
'n_estimators': [10, 100, 1000], # 'n_estimators': 100,
# 'n_jobs': None,
# 'oob_score': False,
# 'random_state': None,
# 'verbose': 0,
# 'warm_start': False
}
# Store dictionary of default hyperparameters used in search for plotting later
defaultParameters = {
'criterion': 'gini',
'max_features': 3, # sqrt(8) ~ 3
'n_estimators': 100,
}
# -
# ##### Perform search
# Create random forest classifier for hyperparameter search
rfSearchModel = RandomForestClassifier()
# Perform search based on search space
search = hyperparameterTuner(rfSearchModel, searchSpace, X_train, y_train)
# Store best parameters found
optimumParameters = search.best_params_
# Store all results in new dataframe
results = pd.DataFrame(search.cv_results_)
# ##### Plot search results
# +
# Create figure for results
fig, ax = plt.subplots()
# Lineplot to display trend for each hyperparameter investigated
sns.lineplot(
data=results,
y='mean_test_score',
x='param_max_features',
hue='param_n_estimators',
style='param_criterion',
palette="Set1",
ax=ax,
)
# Add point to highlight best accuracy achieved
ax.scatter(optimumParameters['max_features'], results[results['params'] == optimumParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Optimised',
marker='o'
)
# # Add point to show accuracy achieved using default hyperparameters
ax.scatter(defaultParameters['max_features'], results[results['params'] == defaultParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Default',
marker='^'
)
# Move legend outside figure
ax.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
# Fix x axis markers to whole numbers
ax.xaxis.get_major_locator().set_params(integer=True)
# Add title
fig.suptitle('Random Forest accuracies across hyperparameter search space', fontsize=12)
fig.savefig('RF_hyperparameterSearchResults', bbox_inches='tight')
plt.show()
# -
# ##### Test best hyperparameters
# Create Random Forest classifier using best parameters
rfBestModel = RandomForestClassifier(
criterion=optimumParameters['criterion'],
max_features=optimumParameters['max_features'],
n_estimators=optimumParameters['n_estimators'],
)
# Train model
rfBestModel.fit(X_train, y_train)
# Test model
rfBestPredictions = rfBestModel.predict(X_test)
# ##### Compare baseline and optimised accuracies
print(f'Baseline model test accuracy: {100*((rfBaselinePredictions==y_test).sum()/y_test.shape[0]):.2f}%')
print(f'Optimised model test accuracy: {100*((rfBestPredictions==y_test).sum()/y_test.shape[0]):.2f}%')
# ##### Evaluation metrics
# Create confusion matrix for random forest optimised hyperparameter test results
confusionMatrixRF = confusion_matrix(y_test, rfBestPredictions, labels=[1, 0])
# Store evaluation metrics in dataframe
evaluationDf = evaluationDf.append(getEvaluationMetrics(confusionMatrixRF, 'Random Forest'), ignore_index=True)
# #### Support Vector Machine Classifier
# ##### Data preparation
# +
# Split data into 80% train and 20% test sets
# Data stratified by class due to imbalance in classes (65% negative - 35% positive)
# Data shuffled to reduce any bias in the order of the dataset
# Define random state to ensure the same train test split can be used for each method
X_train, X_test, y_train, y_test = train_test_split(df[features],
df[target],
test_size=0.20,
stratify=df[target],
shuffle=True,
random_state=123,
)
# Standardise features to negate the impact of different feature units
# Updated features have mean 0 and standard deviation 1
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# -
# ##### Baseline model using default hyperparameters
# Create support vector classifier using default hyperparameters
svcBaselineModel = SVC()
# Train model
svcBaselineModel.fit(X_train, y_train)
# Test model
svcBaselinePredictions = svcBaselineModel.predict(X_test)
# ##### Hyperparameter search space
# +
# Define SVC hyperparameter search space
# Default values for parameters are shown in comments
searchSpace = {
'C': np.logspace(-2, 3, num=6), # 'C': 1.0,
# 'break_ties': False,
# 'cache_size': 200,
# 'class_weight': None,
# 'coef0': 0.0,
# 'decision_function_shape': 'ovr',
'degree': [2, 3, 4], # 'degree': 3, (only used for polynomial kernel)
# 'gamma': 'scale',
'kernel': ['linear', 'poly', 'rbf', 'sigmoid'], # 'kernel': 'rbf',
# 'max_iter': -1,
# 'probability': False,
# 'random_state': None,
# 'shrinking': True,
# 'tol': 0.001,
# 'verbose': False
}
# Store dictionary of default hyperparameters used in search for plotting later
defaultParameters = {
'C': 1.0,
'degree': 3,
'kernel': 'rbf',
}
# -
# ##### Perform search
# Create SVC classifier for hyperparameter search
svcSearchModel = SVC()
# Perform search based on search space
search = hyperparameterTuner(svcSearchModel, searchSpace, X_train, y_train)
# Store best parameters found
optimumParameters = search.best_params_
# Store all results in new dataframe
results = pd.DataFrame(search.cv_results_)
# ##### Plot search results
# +
# Create figure for results
fig, ax = plt.subplots()
# Lineplot to display trend for each hyperparameter investigated
sns.lineplot(
data=results,
y='mean_test_score',
x='param_C',
hue='param_kernel',
style='param_degree',
palette="Set1",
ax=ax,
)
# Add point to highlight best accuracy achieved
ax.scatter(optimumParameters['C'], results[results['params'] == optimumParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Optimised',
marker='o'
)
# Add point to show accuracy achieved using default hyperparameters
ax.scatter(defaultParameters['C'], results[results['params'] == defaultParameters]['mean_test_score'],
facecolors='none',
edgecolors='k',
linewidth=2,
s=100,
label='Default',
marker='^'
)
# Set logarithmic x axis
ax.set_xscale('log')
# Move legend outside figure
ax.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
# Add title
fig.suptitle('Support Vector Classifier accuracies across hyperparameter search space', fontsize=12)
fig.savefig('SVC_hyperparameterSearchResults', bbox_inches='tight')
plt.show()
# -
# ##### Test best hyperparameters
# Create SVC using best parameters
svcBestModel = SVC(
C=optimumParameters['C'],
kernel=optimumParameters['kernel'],
degree=optimumParameters['degree'],
)
# Train model
svcBestModel.fit(X_train, y_train)
# Test model
svcBestPredictions = svcBestModel.predict(X_test)
# ##### Compare baseline and optimised accuracies
print(f'Baseline model test accuracy: {100*((svcBaselinePredictions==y_test).sum()/y_test.shape[0]):.2f}%')
print(f'Optimised model test accuracy: {100*((svcBestPredictions==y_test).sum()/y_test.shape[0]):.2f}%')
# ##### Evaluation metrics
# Create confusion matrix for SVC optimised hyperparameter test results
confusionMatrixSVC = confusion_matrix(y_test, svcBestPredictions, labels=[1, 0])
# Store evaluation metrics in dataframe
evaluationDf = evaluationDf.append(getEvaluationMetrics(confusionMatrixSVC, 'Support Vector Classifier'), ignore_index=True)
# ### Evaluation
# The performance of the three methods will be evaluation using their confusion matrices and the associated metrics. These will be discussed further in the accompanying report.
# +
# Create figure for confusion matrices
fig, axs = plt.subplots(1, 3, figsize=(12,4))
# Create confusion matrix display for knn, add to figure and provide title
disp_knn = ConfusionMatrixDisplay(confusion_matrix=confusionMatrixKNN, display_labels=[1,0])
disp_knn.plot(ax=axs[0], colorbar=False, cmap='cividis')
axs[0].set_title('k-Nearest Neighbours')
# Create confusion matrix display for rf, add to figure and provide title
disp_rf = ConfusionMatrixDisplay(confusion_matrix=confusionMatrixRF, display_labels=[1,0])
disp_rf.plot(ax=axs[1], colorbar=False, cmap='cividis')
axs[1].set_title('Random Forest')
# Create confusion matrix display for svc, add to figure and provide title
disp_svc = ConfusionMatrixDisplay(confusion_matrix=confusionMatrixSVC, display_labels=[1,0])
disp_svc.plot(ax=axs[2], colorbar=False, cmap='cividis')
axs[2].set_title('Support Vector Classifier')
# Add title
fig.suptitle('Confusion matrices comparison', fontsize=12)
fig.savefig('confusionMatrices', bbox_inches='tight')
plt.show()
# -
# Display evaluation metrics dataframe
evaluationDf.set_index('Algorithm')
|
COMP534-AppliedArtificialIntelligence/COMP534-CA1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # CME 193 - Scientific Python
# ### Lecture 6 (4/26)
# Spring 2016, Stanford University
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Last time
# * Object Oriented Programming!
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Today
# * Dealing with data
# * Building some predictive models
# * Some buzzwords 4 u
# + [markdown] slideshow={"slide_type": "slide"}
# # Intro to Pandas
# + [markdown] slideshow={"slide_type": "fragment"}
# ### (and a bit of Seaborn)
# + [markdown] slideshow={"slide_type": "slide"}
# ## What is Pandas?
# + [markdown] slideshow={"slide_type": "fragment"}
# * Open source, well maintained library for the most fundamental portion of science / research / data science
# * Data structures, data analysis tools
# * Makes the barriers to entry from R / SAS / Stata as small as possible...
# * Some nice plotting wrappers
# + [markdown] slideshow={"slide_type": "fragment"}
# If you like this kind of stuff, definitely check out the Pandas cookbook!
# https://github.com/jvns/pandas-cookbook
# + [markdown] slideshow={"slide_type": "subslide"}
# ## What is Seaborn?
# + [markdown] slideshow={"slide_type": "fragment"}
# Seaborn is a context and style manager for matplotlib, the standard plotting package for Python.
# + [markdown] slideshow={"slide_type": "fragment"}
# TL;DR -- makes stuff pretty
# + slideshow={"slide_type": "subslide"}
# %matplotlib inline
# + slideshow={"slide_type": "fragment"}
import numpy as np
import pandas as pd
# + slideshow={"slide_type": "fragment"}
import matplotlib.pyplot as plt
import seaborn as sns
# + slideshow={"slide_type": "fragment"}
sns.set_style("dark")
sns.set_context("talk")
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's start looking at some data!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Iris Data
# + [markdown] slideshow={"slide_type": "fragment"}
# We'll use the standard Iris data set. Let's load data **directly** from an online source!
# + slideshow={"slide_type": "fragment"}
iris = pd.read_csv(
'https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data',
header=None,
names=['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
)
# + [markdown] slideshow={"slide_type": "fragment"}
# What's going on here?
# + slideshow={"slide_type": "subslide"}
type(iris)
# + slideshow={"slide_type": "fragment"}
iris.head()
# + [markdown] slideshow={"slide_type": "slide"}
# # DATA SCIENCE!
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's do some exploratory data science with Pandas + Iris!
# + [markdown] slideshow={"slide_type": "fragment"}
# If you use data in what you do, you probably aren't doing enough of this!
# + slideshow={"slide_type": "fragment"}
iris.head(n=2)
# + slideshow={"slide_type": "subslide"}
sns.pairplot(iris, hue='species')
# + slideshow={"slide_type": "subslide"}
sns.corrplot(iris.drop('species', 1))
# + [markdown] slideshow={"slide_type": "subslide"}
# Can index using dot notation!
# + slideshow={"slide_type": "fragment"}
iris.sepal_length.head()
# + [markdown] slideshow={"slide_type": "fragment"}
# Calculating summary statistics is easy
# + slideshow={"slide_type": "fragment"}
print 'The mean sepal length is {}'.format(iris.sepal_length.mean())
# + slideshow={"slide_type": "fragment"}
iris.sepal_length.ix[0]
# + [markdown] slideshow={"slide_type": "subslide"}
# It's easy to select columns
# + slideshow={"slide_type": "fragment"}
iris[['sepal_length', 'sepal_width']].head()
# + slideshow={"slide_type": "subslide"}
x = iris[['sepal_length', 'sepal_width']][:3]
# + slideshow={"slide_type": "fragment"}
x
# + slideshow={"slide_type": "fragment"}
X = x.values
# + slideshow={"slide_type": "fragment"}
X
# + [markdown] slideshow={"slide_type": "slide"}
# # Data Exploration from **scratch**
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's do some data science in numpy!
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's get principal components analysis working (PCA)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### What is PCA?
# + [markdown] slideshow={"slide_type": "fragment"}
# * Transform a bunch of data into a set of linearly uncorrelated (orthogonal) features
# * Think of it as a decomposition -- we're able to figure out how much variance per feature
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Some Math...
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's say we have data $X$, and each feature (variable) has zero mean, unit variance.
# + [markdown] slideshow={"slide_type": "fragment"}
# We talked about the SVD before.
# + [markdown] slideshow={"slide_type": "fragment"}
# We said that for *any* $X\in\mathbb{C}$, we can find:
#
# $$
# X = U\Sigma V^T
# $$
#
# Where $U$ and $V$ are unitary, and $\Sigma$ is diagonal.
# + [markdown] slideshow={"slide_type": "subslide"}
# Suppose we wanted to find a linear mapping, call it $W$, that transforms my data $X$ into it Principal Component, i.e., $T = XW$ are my principal components.
# + [markdown] slideshow={"slide_type": "fragment"}
# I claim that setting $W = V$ (from SVD) satisfies this. Furthermore, the uncorrelated version of my data $X$ is simply:
#
# $$
# XW = U\Sigma V^T W = U\Sigma W^T W = U\Sigma
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# So, if I wanted to transform my data, all I need to do is:
#
# * Center / normalize $X$
# * Find $X = U\Sigma V^T$
# * My principal components are $T = U\Sigma$
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's do this with Iris data!
# + [markdown] slideshow={"slide_type": "subslide"}
# First, let's convert Iris to a numpy array
# + slideshow={"slide_type": "fragment"}
X = iris.drop('species', 1).values
# + slideshow={"slide_type": "fragment"}
X.shape
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's center our matrix! We'll use **broadcasting**...
# + slideshow={"slide_type": "fragment"}
print 'Mean Values: {}'.format(X.mean(axis=0))
print 'Mean Shape: {}'.format(X.mean(axis=0).shape)
print 'Stdev Values: {}'.format(X.std(axis=0))
print 'Stdev Shape: {}'.format(X.std(axis=0).shape)
# + slideshow={"slide_type": "subslide"}
X_centered = (X - X.mean(axis=0)) / X.std(axis=0)
# + [markdown] slideshow={"slide_type": "fragment"}
# shapes: `((150, 4) - (4, )) / (4, )`
# + slideshow={"slide_type": "fragment"}
print 'Mean Values: {}'.format(X_centered.mean(axis=0))
print 'Stdev Values: {}'.format(X_centered.std(axis=0))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Let's take our SVD and get to work!
# + slideshow={"slide_type": "subslide"}
U, S, V = np.linalg.svd(X_centered)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's form $\Sigma$...
# + [markdown] slideshow={"slide_type": "fragment"}
# `S` isnt in the right form yet
# + slideshow={"slide_type": "fragment"}
S
# + slideshow={"slide_type": "subslide"}
# sigma needs to be the same shape as X
sigma = np.zeros(X.shape)
# + slideshow={"slide_type": "fragment"}
# we need the upper 4x4 square to be the diagonal matrix we expect
sigma[:X.shape[1]] = np.diag(S)
# + slideshow={"slide_type": "fragment"}
sigma
# + slideshow={"slide_type": "subslide"}
# let's verify our shortcut, and grab our PCs
T = np.dot(U, sigma)
# + slideshow={"slide_type": "fragment"}
# and treating V as a mapping...let's also grab PCs
Tprime = np.dot(X_centered, V.T)
# + slideshow={"slide_type": "fragment"}
np.allclose(T, Tprime)
# + slideshow={"slide_type": "fragment"}
T.shape
# + slideshow={"slide_type": "subslide"}
cols = ['pc_%s' % i for i in range(T.shape[-1])]
print cols
# + slideshow={"slide_type": "fragment"}
pc = pd.DataFrame(T, columns=cols)
# + slideshow={"slide_type": "fragment"}
pc['species'] = iris.species
# + slideshow={"slide_type": "subslide"}
pc.head()
# + slideshow={"slide_type": "subslide"}
COLORS = ['red', 'blue', 'green']
for i, (key, group) in enumerate(pc.groupby('species')):
plt.hist(group['pc_0'].values, histtype='step', label=key, color=COLORS[i])
plt.title('Iris, first principal component')
plt.legend()
# + slideshow={"slide_type": "subslide"}
sns.lmplot('pc_0', 'pc_1', data=pc, hue='species', fit_reg=False)
# + [markdown] slideshow={"slide_type": "fragment"}
# This seems to be picking up most variation in two components
# + [markdown] slideshow={"slide_type": "fragment"}
# How can we verify this fact?
# + [markdown] slideshow={"slide_type": "subslide"}
# Well, let's look at $U\Sigma V^T$, and call $\sigma_i = \Sigma_{ii}$.
# + [markdown] slideshow={"slide_type": "fragment"}
# I claim that the percent variance explained by the $i$th Principal Component is
#
# $$
# \frac{\sigma_i}{\sum_{j=1}^{p}\sigma_j}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's look at this with our data!
# + slideshow={"slide_type": "subslide"}
print S
# + slideshow={"slide_type": "fragment"}
v = (100.) * (S / S.sum())
# + slideshow={"slide_type": "fragment"}
for i, pc in enumerate(v):
print 'Principal Component #%i accounts for %.2f%% of the variance' % (i, pc)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's go one final step further! Let's look at cumulative variance.
# + slideshow={"slide_type": "fragment"}
v_tot = v.cumsum()
plt.plot(range(1, 5), v_tot, color='red')
plt.xticks(range(1, 5))
plt.grid(True)
plt.xlabel('# PC included')
plt.ylabel('Cumulative % variance explained')
# + [markdown] slideshow={"slide_type": "slide"}
# # Thanks
# + [markdown] slideshow={"slide_type": "fragment"}
# Office hours will be from 5-6:30 in Y2E2 105
|
nb/2016_spring/lecture-6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Support Vector Machine (SVM) is a supervised machine learning algorithm which can be used for both classification or regression challenges. However, it is mostly used in classification problems. In this algorithm, we plot each data item as a point in n-dimensional space (where n is number of features you have) with the value of each feature being the value of a particular coordinate. Then, we perform classification by finding the hyper-plane that differentiate the two classes very well
# The important parameters to tune in SVM are gamma, c and Kernel
#
# * Kernel: linear or radial
# * gamma: the free parameter of the Gaussian radial basis function. A small gamma means a Gaussian with a large variance so the influence of x_j is more, i.e. if x_j is a support vector, a small gamma implies the class of this support vector will have influence on deciding the class of the vector x_i even if the distance between them is large. If gamma is large, then variance is small implying the support vector does not have wide-spread influence. Technically speaking, large gamma leads to high bias and low variance models, and vice-versa.
# * c: the parameter for the soft margin cost function, which controls the influence of each individual support vector; this process involves trading error penalty for stability. It also controls the trade off between smooth decision boundary and classifying the training points correctly.
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import svm, datasets
# we only take the first two features. We could avoid this ugly slicing by using a two-dim dataset
## we can also plot it on 2d plane if we deal with 2d dataset
iris = datasets.load_iris()
y = iris.target
X = iris.data[:, :2]
X.shape,y.shape
# we create an instance of SVM and fit out data. We do not scale our
# data since we want to plot the support vectors
C = 1.0 # SVM regularization parameter
svc = svm.SVC(kernel='linear', C=1,gamma='auto').fit(X, y)
# create a mesh to plot in
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
h = (x_max / x_min)/100
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z.shape
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx,yy,Z,cmap=plt.cm.Paired, alpha=0.8)
##red is for class 1 brown for class2 and blue for class 0
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y+100, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with linear kernel')
# using radial
svc = svm.SVC(kernel='rbf', C=1,gamma='auto').fit(X, y)
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with Radial kernel')
##change value of gamma to 10
svc = svm.SVC(kernel='rbf', C=1,gamma=10.0).fit(X, y)
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with Radial kernel and gamma=10, it has lower variance but higher bias then gamma=0')
##change value of gamma to 100
svc = svm.SVC(kernel='rbf', C=1,gamma=100.0).fit(X, y)
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with Radial kernel and gamma=100, it has lower variance but higher bias then gamma=10')
##changing the value c
svc = svm.SVC(kernel='rbf', C=100,gamma='auto').fit(X, y)
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with Radial kernel and c=100, it effects the smoothness of decision boundry')
##changing the value c
svc = svm.SVC(kernel='rbf', C=1000,gamma='auto').fit(X, y)
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with Radial kernel and c=100, it effects the smoothness of decision boundry')
# ### Pros and Cons associated with SVM
# #### Pros:
# * It works really well with clear margin of separation
# * It is effective in high dimensional spaces.
# * It is effective in cases where number of dimensions is greater than the number of samples.
# * It uses a subset of training points in the decision function (called support vectors), so it is also memory efficient.
# #### Cons:
# * It doesn’t perform well, when we have large data set because the required training time is higher
# * It also doesn’t perform very well, when the data set has more noise i.e. target classes are overlapping
# * SVM doesn’t directly provide probability estimates, these are calculated using an expensive five-fold cross-validation. It is related SVC method of Python scikit-learn library.
|
Using SVM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Comparing Python vs TensorFlow Performance
#
# To justify why we use TensorFlow over normal python, we can run some benchmarks for simple operations and compare the two implementations.
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import time as time
# ### Matrix Multiplication
#
# We compare the time takes for Python and TensorFlow to evaluate the product of two matrices of varying sizes.
n_replicate = 15
matrix_sizes = [2,8,32,128,512,2048,8192,16384]
# #### In Python
for n in matrix_sizes:
run_time = []
for i in range(n_replicate):
start = time.time()
a = np.random.uniform(size=(n,n))
b = np.random.uniform(size=(n,n))
c = np.matmul(a,b)
end = time.time()
run_time.append(end-start)
mean = np.mean(run_time)
sd = np.std(run_time)
print("For a {}x{} Matrix, Runtime = {:0.4f} +/- {:0.4f} secs".format(n,n,mean,sd))
# #### In TensorFlow
for n in matrix_sizes:
run_time = []
for i in range(n_replicate):
start = time.time()
a = tf.random_uniform([n,n])
b = tf.random_uniform([n,n])
c = tf.matmul(a,b)
with tf.Session() as sess:
c = sess.run(c)
end = time.time()
run_time.append(end-start)
mean = np.mean(run_time)
sd = np.std(run_time)
print("For a {}x{} Matrix, Runtime = {:0.4f} +/- {:0.4f} secs".format(n,n,mean,sd))
# ### RK4 Integration
#
# We compare the time takes for Python and TensorFlow to evaluate the integrate varying numbers of the differential equations of the form $\dot x = \sin{xt}$ .
n_replicate = 15
equation_sizes = [1,10,100,1000,10000,100000,1000000]
t = np.arange(0,5,0.01)
# #### In Python
# +
def python_check_type(y,t): # Ensure Input is Correct
return y.dtype == np.floating and t.dtype == np.floating
class python_Integrator():
def integrate(self,func,y0,t):
time_delta_grid = t[1:] - t[:-1]
y = np.zeros((y0.shape[0],t.shape[0]))
y[:,0] = y0
for i in range(time_delta_grid.shape[0]):
k1 = func(y[:,i], t[i]) # RK4 Integration Steps
half_step = t[i] + time_delta_grid[i] / 2
k2 = func(y[:,i] + time_delta_grid[i] * k1 / 2, half_step)
k3 = func(y[:,i] + time_delta_grid[i] * k2 / 2, half_step)
k4 = func(y[:,i] + time_delta_grid[i] * k3, t[i] + time_delta_grid[i])
y[:,i+1]= (k1 + 2 * k2 + 2 * k3 + k4) * (time_delta_grid[i] / 6) + y[:,i]
return y
def odeint_python(func,y0,t):
y0 = np.array(y0)
t = np.array(t)
if python_check_type(y0,t):
return python_Integrator().integrate(func,y0,t)
else:
print("error encountered")
def f(X,t):
return np.sin(X*t)
for n in equation_sizes:
run_time = []
for i in range(n_replicate):
start = time.time()
solution = odeint_python(f,[0.]*n,t)
end = time.time()
run_time.append(end-start)
mean = np.mean(run_time)
sd = np.std(run_time)
print("For a {} Equations, Runtime = {:0.4f} +/- {:0.4f} secs".format(n,mean,sd))
# -
# #### In TensorFlow
# +
def tf_check_type(t, y0): # Ensure Input is Correct
if not (y0.dtype.is_floating and t.dtype.is_floating):
raise TypeError('Error in Datatype')
class Tf_Integrator():
def integrate(self, func, y0, t):
time_delta_grid = t[1:] - t[:-1]
def scan_func(y, t_dt):
t, dt = t_dt
dy = self._step_func(func,t,dt,y)
return y + dy
y = tf.scan(scan_func, (t[:-1], time_delta_grid),y0)
return tf.concat([[y0], y], axis=0)
def _step_func(self, func, t, dt, y):
k1 = func(y, t)
half_step = t + dt / 2
dt_cast = tf.cast(dt, y.dtype) # Failsafe
k2 = func(y + dt_cast * k1 / 2, half_step)
k3 = func(y + dt_cast * k2 / 2, half_step)
k4 = func(y + dt_cast * k3, t + dt)
return tf.add_n([k1, 2 * k2, 2 * k3, k4]) * (dt_cast / 6)
def odeint_tf(func, y0, t):
t = tf.convert_to_tensor(t, preferred_dtype=tf.float64, name='t')
y0 = tf.convert_to_tensor(y0, name='y0')
tf_check_type(y0,t)
return Tf_Integrator().integrate(func,y0,t)
def f(X,t):
return tf.sin(X*t)
for n in equation_sizes:
run_time = []
for i in range(n_replicate):
start = time.time()
state = odeint_tf(f,tf.constant([0.]*n,dtype=tf.float64),t)
with tf.Session() as sess:
state = sess.run(state)
end = time.time()
run_time.append(end-start)
mean = np.mean(run_time)
sd = np.std(run_time)
print("For a {} Equations, Runtime = {:0.4f} +/- {:0.4f} secs".format(n,mean,sd))
|
Tutorial/Supplementary: Jupyter Notebooks/Supplementary: Benchmark/Benchmark.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Car Number Plate Detection using OpenCV
# This code template is for car number plate detection using OpenCV
# ## Required Packages
import cv2
import numpy as np
import matplotlib.pyplot as plt
# ## Importing our Classifier
#
# **Link:** [classifier used in this template](https://drive.google.com/file/d/15o6VctOvVgySRoQMigPIRly9oBnU4Q1L/view)
cascade = cv2.CascadeClassifier('')
# ## Reading our image
# OpenCV-Python is a library of Python bindings designed to solve computer vision problems.
#
# We use the the **"cv2.imread()"** method to load an image from the specified file path.
cars = cv2.imread('')
# ## Converting Image to Gray
gray = cv2.cvtColor(cars, cv2.COLOR_BGR2GRAY)
# ## Converting image to colored image
def converToRGB(image):
return cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
# ## Drawing bounding boxes
cars_detected = cascade.detectMultiScale(
gray,
scaleFactor=1.2,
minNeighbors=5,
minSize=(20,20))
print('Found Numbers', len(cars_detected))
for (x, y, w, h) in cars_detected:
cv2.rectangle(cars, (x,y), (x+w,y+h),(145,60,255), 5)
# ## Final Image
plt.figure(figsize=(20,20))
plt.imshow(converToRGB(cars));
# ## Creator: <NAME>, Github: [Profile](https://github.com/abhishek-252)
#
|
Audio Visual/Problems/CarNumberPlateDetection_OpenCV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementing logistic regression from scratch
#
# The goal of this notebook is to implement your own logistic regression classifier. You will:
#
# * Extract features from Amazon product reviews.
# * Convert an SFrame into a NumPy array.
# * Implement the link function for logistic regression.
# * Write a function to compute the derivative of the log likelihood function with respect to a single coefficient.
# * Implement gradient ascent.
# * Given a set of coefficients, predict sentiments.
# * Compute classification accuracy for the logistic regression model.
#
# Let's get started!
#
# ## Fire up Turi Create
#
# Make sure you have the latest version of Turi Create.
import turicreate
# ## Load review dataset
# For this assignment, we will use a subset of the Amazon product review dataset. The subset was chosen to contain similar numbers of positive and negative reviews, as the original dataset consisted primarily of positive reviews.
products = turicreate.SFrame('amazon_baby_subset.sframe/')
# One column of this dataset is 'sentiment', corresponding to the class label with +1 indicating a review with positive sentiment and -1 indicating one with negative sentiment.
products['sentiment']
# Let us quickly explore more of this dataset. The 'name' column indicates the name of the product. Here we list the first 10 products in the dataset. We then count the number of positive and negative reviews.
products.head(10)['name']
print('# of positive reviews =', len(products[products['sentiment']==1]))
print('# of negative reviews =', len(products[products['sentiment']==-1]))
# **Note:** For this assignment, we eliminated class imbalance by choosing
# a subset of the data with a similar number of positive and negative reviews.
#
# ## Apply text cleaning on the review data
#
# In this section, we will perform some simple feature cleaning using **SFrames**. The last assignment used all words in building bag-of-words features, but here we limit ourselves to 193 words (for simplicity). We compiled a list of 193 most frequent words into a JSON file.
#
# Now, we will load these words from this JSON file:
import json
with open('important_words.json', 'r') as f: # Reads the list of most frequent words
important_words = json.load(f)
important_words = [str(s) for s in important_words]
print(important_words)
# Now, we will perform 2 simple data transformations:
#
# 1. Remove punctuation using [Python's built-in](https://docs.python.org/2/library/string.html) string functionality.
# 2. Compute word counts (only for **important_words**)
#
# We start with *Step 1* which can be done as follows:
# +
import string
def remove_punctuation(text):
try: # python 2.x
text = text.translate(None, string.punctuation)
except: # python 3.x
translator = text.maketrans('', '', string.punctuation)
text = text.translate(translator)
return text
products['review_clean'] = products['review'].apply(remove_punctuation)
# -
# Now we proceed with *Step 2*. For each word in **important_words**, we compute a count for the number of times the word occurs in the review. We will store this count in a separate column (one for each word). The result of this feature processing is a single column for each word in **important_words** which keeps a count of the number of times the respective word occurs in the review text.
#
#
# **Note:** There are several ways of doing this. In this assignment, we use the built-in *count* function for Python lists. Each review string is first split into individual words and the number of occurances of a given word is counted.
for word in important_words:
products[word] = products['review_clean'].apply(lambda s : s.split().count(word))
# The SFrame **products** now contains one column for each of the 193 **important_words**. As an example, the column **perfect** contains a count of the number of times the word **perfect** occurs in each of the reviews.
products['perfect']
# Now, write some code to compute the number of product reviews that contain the word **perfect**.
#
# **Hint**:
# * First create a column called `contains_perfect` which is set to 1 if the count of the word **perfect** (stored in column **perfect**) is >= 1.
# * Sum the number of 1s in the column `contains_perfect`.
counter = 0
for state in products['perfect']:
if state > 0:
counter += 1
counter
# **Quiz Question**. How many reviews contain the word **perfect**?
# ## Convert SFrame to NumPy array
#
# As you have seen previously, NumPy is a powerful library for doing matrix manipulation. Let us convert our data to matrices and then implement our algorithms with matrices.
#
# First, make sure you can perform the following import. If it doesn't work, you need to go back to the terminal and run
#
# `pip install numpy`.
import numpy as np
# We now provide you with a function that extracts columns from an SFrame and converts them into a NumPy array. Two arrays are returned: one representing features and another representing class labels. Note that the feature matrix includes an additional column 'intercept' to take account of the intercept term.
def get_numpy_data(data_sframe, features, label):
data_sframe['intercept'] = 1
features = ['intercept'] + features
features_sframe = data_sframe[features]
feature_matrix = features_sframe.to_numpy()
label_sarray = data_sframe[label]
label_array = label_sarray.to_numpy()
return(feature_matrix, label_array)
# Let us convert the data into NumPy arrays.
# Warning: This may take a few minutes...
feature_matrix, sentiment = get_numpy_data(products, important_words, 'sentiment')
# **Are you running this notebook on an Amazon EC2 t2.micro instance?** (If you are using your own machine, please skip this section)
#
# It has been reported that t2.micro instances do not provide sufficient power to complete the conversion in acceptable amount of time. For interest of time, please refrain from running `get_numpy_data` function. Instead, download the [binary file](https://s3.amazonaws.com/static.dato.com/files/coursera/course-3/numpy-arrays/module-3-assignment-numpy-arrays.npz) containing the four NumPy arrays you'll need for the assignment. To load the arrays, run the following commands:
# ```
# arrays = np.load('module-3-assignment-numpy-arrays.npz')
# feature_matrix, sentiment = arrays['feature_matrix'], arrays['sentiment']
# ```
feature_matrix.shape
# **Quiz Question:** How many features are there in the **feature_matrix**?
#
# **Quiz Question:** Assuming that the intercept is present, how does the number of features in **feature_matrix** relate to the number of features in the logistic regression model?
# Now, let us see what the **sentiment** column looks like:
sentiment
# ## Estimating conditional probability with link function
# Recall from lecture that the link function is given by:
# $$
# P(y_i = +1 | \mathbf{x}_i,\mathbf{w}) = \frac{1}{1 + \exp(-\mathbf{w}^T h(\mathbf{x}_i))},
# $$
#
# where the feature vector $h(\mathbf{x}_i)$ represents the word counts of **important_words** in the review $\mathbf{x}_i$. Complete the following function that implements the link function:
'''
produces probablistic estimate for P(y_i = +1 | x_i, w).
estimate ranges between 0 and 1.
'''
def predict_probability(feature_matrix, coefficients):
# Take dot product of feature_matrix and coefficients
# YOUR CODE HERE
scores = np.dot(feature_matrix, coefficients)
# Compute P(y_i = +1 | x_i, w) using the link function
# YOUR CODE HERE
predictions = 1 / (1 + np.exp(-scores))
# return predictions
return predictions
# **Aside**. How the link function works with matrix algebra
#
# Since the word counts are stored as columns in **feature_matrix**, each $i$-th row of the matrix corresponds to the feature vector $h(\mathbf{x}_i)$:
# $$
# [\text{feature_matrix}] =
# \left[
# \begin{array}{c}
# h(\mathbf{x}_1)^T \\
# h(\mathbf{x}_2)^T \\
# \vdots \\
# h(\mathbf{x}_N)^T
# \end{array}
# \right] =
# \left[
# \begin{array}{cccc}
# h_0(\mathbf{x}_1) & h_1(\mathbf{x}_1) & \cdots & h_D(\mathbf{x}_1) \\
# h_0(\mathbf{x}_2) & h_1(\mathbf{x}_2) & \cdots & h_D(\mathbf{x}_2) \\
# \vdots & \vdots & \ddots & \vdots \\
# h_0(\mathbf{x}_N) & h_1(\mathbf{x}_N) & \cdots & h_D(\mathbf{x}_N)
# \end{array}
# \right]
# $$
#
# By the rules of matrix multiplication, the score vector containing elements $\mathbf{w}^T h(\mathbf{x}_i)$ is obtained by multiplying **feature_matrix** and the coefficient vector $\mathbf{w}$.
# $$
# [\text{score}] =
# [\text{feature_matrix}]\mathbf{w} =
# \left[
# \begin{array}{c}
# h(\mathbf{x}_1)^T \\
# h(\mathbf{x}_2)^T \\
# \vdots \\
# h(\mathbf{x}_N)^T
# \end{array}
# \right]
# \mathbf{w}
# = \left[
# \begin{array}{c}
# h(\mathbf{x}_1)^T\mathbf{w} \\
# h(\mathbf{x}_2)^T\mathbf{w} \\
# \vdots \\
# h(\mathbf{x}_N)^T\mathbf{w}
# \end{array}
# \right]
# = \left[
# \begin{array}{c}
# \mathbf{w}^T h(\mathbf{x}_1) \\
# \mathbf{w}^T h(\mathbf{x}_2) \\
# \vdots \\
# \mathbf{w}^T h(\mathbf{x}_N)
# \end{array}
# \right]
# $$
# **Checkpoint**
#
# Just to make sure you are on the right track, we have provided a few examples. If your `predict_probability` function is implemented correctly, then the outputs will match:
# +
dummy_feature_matrix = np.array([[1.,2.,3.], [1.,-1.,-1]])
dummy_coefficients = np.array([1., 3., -1.])
correct_scores = np.array( [ 1.*1. + 2.*3. + 3.*(-1.), 1.*1. + (-1.)*3. + (-1.)*(-1.) ] )
correct_predictions = np.array( [ 1./(1+np.exp(-correct_scores[0])), 1./(1+np.exp(-correct_scores[1])) ] )
print('The following outputs must match ')
print('------------------------------------------------')
print('correct_predictions =', correct_predictions)
print('output of predict_probability =', predict_probability(dummy_feature_matrix, dummy_coefficients))
# -
# ## Compute derivative of log likelihood with respect to a single coefficient
#
# Recall from lecture:
# $$
# \frac{\partial\ell}{\partial w_j} = \sum_{i=1}^N h_j(\mathbf{x}_i)\left(\mathbf{1}[y_i = +1] - P(y_i = +1 | \mathbf{x}_i, \mathbf{w})\right)
# $$
#
# We will now write a function that computes the derivative of log likelihood with respect to a single coefficient $w_j$. The function accepts two arguments:
# * `errors` vector containing $\mathbf{1}[y_i = +1] - P(y_i = +1 | \mathbf{x}_i, \mathbf{w})$ for all $i$.
# * `feature` vector containing $h_j(\mathbf{x}_i)$ for all $i$.
#
# Complete the following code block:
def feature_derivative(errors, feature):
# Compute the dot product of errors and feature
derivative = np.dot(errors, feature)
# Return the derivative
return derivative
# In the main lecture, our focus was on the likelihood. In the advanced optional video, however, we introduced a transformation of this likelihood---called the log likelihood---that simplifies the derivation of the gradient and is more numerically stable. Due to its numerical stability, we will use the log likelihood instead of the likelihood to assess the algorithm.
#
# The log likelihood is computed using the following formula (see the advanced optional video if you are curious about the derivation of this equation):
#
# $$\ell\ell(\mathbf{w}) = \sum_{i=1}^N \Big( (\mathbf{1}[y_i = +1] - 1)\mathbf{w}^T h(\mathbf{x}_i) - \ln\left(1 + \exp(-\mathbf{w}^T h(\mathbf{x}_i))\right) \Big) $$
#
# We provide a function to compute the log likelihood for the entire dataset.
def compute_log_likelihood(feature_matrix, sentiment, coefficients):
indicator = (sentiment==+1)
scores = np.dot(feature_matrix, coefficients)
logexp = np.log(1. + np.exp(-scores))
# Simple check to prevent overflow
mask = np.isinf(logexp)
logexp[mask] = -scores[mask]
lp = np.sum((indicator-1)*scores - logexp)
return lp
# **Checkpoint**
#
# Just to make sure we are on the same page, run the following code block and check that the outputs match.
# +
dummy_feature_matrix = np.array([[1.,2.,3.], [1.,-1.,-1]])
dummy_coefficients = np.array([1., 3., -1.])
dummy_sentiment = np.array([-1, 1])
correct_indicators = np.array( [ -1==+1, 1==+1 ] )
correct_scores = np.array( [ 1.*1. + 2.*3. + 3.*(-1.), 1.*1. + (-1.)*3. + (-1.)*(-1.) ] )
correct_first_term = np.array( [ (correct_indicators[0]-1)*correct_scores[0], (correct_indicators[1]-1)*correct_scores[1] ] )
correct_second_term = np.array( [ np.log(1. + np.exp(-correct_scores[0])), np.log(1. + np.exp(-correct_scores[1])) ] )
correct_ll = sum( [ correct_first_term[0]-correct_second_term[0], correct_first_term[1]-correct_second_term[1] ] )
print('The following outputs must match ')
print('------------------------------------------------')
print('correct_log_likelihood =', correct_ll)
print('output of compute_log_likelihood =', compute_log_likelihood(dummy_feature_matrix, dummy_sentiment, dummy_coefficients))
# -
# ## Taking gradient steps
# Now we are ready to implement our own logistic regression. All we have to do is to write a gradient ascent function that takes gradient steps towards the optimum.
#
# Complete the following function to solve the logistic regression model using gradient ascent:
# +
from math import sqrt
def logistic_regression(feature_matrix, sentiment, initial_coefficients, step_size, max_iter):
coefficients = np.array(initial_coefficients) # make sure it's a numpy array
for itr in range(max_iter):
# Predict P(y_i = +1|x_i,w) using your predict_probability() function
# YOUR CODE HERE
predictions = predict_probability(feature_matrix, coefficients)
# Compute indicator value for (y_i = +1)
indicator = (sentiment==+1)
# Compute the errors as indicator - predictions
errors = indicator - predictions
for j in range(len(coefficients)): # loop over each coefficient
# Recall that feature_matrix[:,j] is the feature column associated with coefficients[j].
# Compute the derivative for coefficients[j]. Save it in a variable called derivative
# YOUR CODE HERE
derivative = feature_derivative(errors, feature_matrix[:, j])
# add the step size times the derivative to the current coefficient
## YOUR CODE HERE
coefficients[j] += step_size * derivative
# Checking whether log likelihood is increasing
if itr <= 15 or (itr <= 100 and itr % 10 == 0) or (itr <= 1000 and itr % 100 == 0) \
or (itr <= 10000 and itr % 1000 == 0) or itr % 10000 == 0:
lp = compute_log_likelihood(feature_matrix, sentiment, coefficients)
print('iteration %*d: log likelihood of observed labels = %.8f' % \
(int(np.ceil(np.log10(max_iter))), itr, lp))
return coefficients
# -
# Now, let us run the logistic regression solver.
coefficients = logistic_regression(feature_matrix, sentiment, initial_coefficients=np.zeros(194),
step_size=1e-7, max_iter=301)
# **Quiz Question:** As each iteration of gradient ascent passes, does the log likelihood increase or decrease?
# ## Predicting sentiments
# Recall from lecture that class predictions for a data point $\mathbf{x}$ can be computed from the coefficients $\mathbf{w}$ using the following formula:
# $$
# \hat{y}_i =
# \left\{
# \begin{array}{ll}
# +1 & \mathbf{x}_i^T\mathbf{w} > 0 \\
# -1 & \mathbf{x}_i^T\mathbf{w} \leq 0 \\
# \end{array}
# \right.
# $$
#
# Now, we will write some code to compute class predictions. We will do this in two steps:
# * **Step 1**: First compute the **scores** using **feature_matrix** and **coefficients** using a dot product.
# * **Step 2**: Using the formula above, compute the class predictions from the scores.
#
# Step 1 can be implemented as follows:
# Compute the scores as a dot product between feature_matrix and coefficients.
scores = np.dot(feature_matrix, coefficients)
# Now, complete the following code block for **Step 2** to compute the class predictions using the **scores** obtained above:
class_predictions = np.array(turicreate.SArray(scores).apply(lambda x: 1 if x > 0 else -1))
print (class_predictions)
# **Quiz Question:** How many reviews were predicted to have positive sentiment?
unique, counts = np.unique(class_predictions, return_counts=True)
print (unique, counts)
# ## Measuring accuracy
#
# We will now measure the classification accuracy of the model. Recall from the lecture that the classification accuracy can be computed as follows:
#
# $$
# \mbox{accuracy} = \frac{\mbox{# correctly classified data points}}{\mbox{# total data points}}
# $$
#
# Complete the following code block to compute the accuracy of the model.
num_mistakes = (class_predictions != sentiment).sum() # YOUR CODE HERE
accuracy = len(sentiment) # YOUR CODE HERE
print("-----------------------------------------------------")
print('# Reviews correctly classified =', len(products) - num_mistakes)
print('# Reviews incorrectly classified =', num_mistakes)
print('# Reviews total =', len(products))
print("-----------------------------------------------------")
print('Accuracy = %.2f' % accuracy)
# **Quiz Question**: What is the accuracy of the model on predictions made above? (round to 2 digits of accuracy)
# ## Which words contribute most to positive & negative sentiments?
# Recall that in Module 2 assignment, we were able to compute the "**most positive words**". These are words that correspond most strongly with positive reviews. In order to do this, we will first do the following:
# * Treat each coefficient as a tuple, i.e. (**word**, **coefficient_value**).
# * Sort all the (**word**, **coefficient_value**) tuples by **coefficient_value** in descending order.
coefficients = list(coefficients[1:]) # exclude intercept
word_coefficient_tuples = [(word, coefficient) for word, coefficient in zip(important_words, coefficients)]
word_coefficient_tuples = sorted(word_coefficient_tuples, key=lambda x:x[1], reverse=True)
# Now, **word_coefficient_tuples** contains a sorted list of (**word**, **coefficient_value**) tuples. The first 10 elements in this list correspond to the words that are most positive.
# ### Ten "most positive" words
#
# Now, we compute the 10 words that have the most positive coefficient values. These words are associated with positive sentiment.
word_coefficient_tuples[0:10]
# **Quiz Question:** Which word is **not** present in the top 10 "most positive" words?
#
# - love
# - easy
# - great
# - perfect
# - cheap
# ### Ten "most negative" words
#
# Next, we repeat this exercise on the 10 most negative words. That is, we compute the 10 words that have the most negative coefficient values. These words are associated with negative sentiment.
word_coefficient_tuples[-10:]
# **Quiz Question:** Which word is **not** present in the top 10 "most negative" words?
#
# - need
# - work
# - disappointed
# - even
# - return
|
Courses/Machine Learning Classification/Implementing logistic regression from scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import json
data_path_1 = '../../2020_MARCH (2).json'
data_path_2 = '../../2020_FEBRUARY (2).json'
def location_history_processing_pipeline(raw_json_file_locations):
raw_json = concat_src_files(raw_json_file_locations)
places_df, location_df = extract_location_information_from_raw(raw_json)
return location_df
def concat_src_files(raw_json_file_locations):
all_data = []
for file in raw_json_file_locations:
print(file)
json_data = json.load(open(file, 'rb'))
all_data += json_data['timelineObjects']
return {'timelineObjects': all_data}
def extract_location_information_from_raw(raw_json):
places_data = []
location_data = []
for json_dict in raw_json['timelineObjects']:
if "placeVisit" in json_dict:
places_data.append(json_dict['placeVisit'])
location_dict = json_dict['placeVisit']['location']
location_dict['visit_start_time'] = json_dict['placeVisit']['duration']['startTimestampMs']
location_dict['visit_end_time'] = json_dict['placeVisit']['duration']['endTimestampMs']
location_data.append(location_dict)
return pd.DataFrame(places_data), pd.DataFrame(location_data)
df1 = location_history_processing_pipeline([data_path_1, data_path_2])
df1[df1['name'] == '<NAME>'].visit_start_time.apply(lambda x: datetime.fromtimestamp(float(x) / 1000))
cases = pd.read_csv("covid-cases-coordinates.csv")
# +
E7 = 10000000
#cases['Latitude'] = cases['Latitude'].apply(lambda x: int(x * E7))
#cases['Longitude'] = cases['Longitude'].apply(lambda x: int(x * E7))
# -
cases['Time'] = pd.to_datetime(cases['Time'])
cases
# # Risk heuristics
# +
DANGEROUS_ZONE_RANGE = 0.1
CRITICAL_OVERFLOW_TIMES = 5
def diminishing_importance(x_raw, x_limit, speed):
x = (x_limit - min(x_limit, x_raw)) * 1.0 / x_limit
#print(x)
#if x >= 1 - DANGEROUS_ZONE_RANGE:
# return 1 - DANGEROUS_ZONE_RANGE + min(x_limit / x_raw * (DANGEROUS_ZONE_RANGE / CRITICAL_OVERFLOW_TIMES), DANGEROUS_ZONE_RANGE)
return max(0, np.exp(speed * (x - 1)))
# -
# !pip install geopy
# +
import geopy.distance
from datetime import datetime
TIMESTAMP_PREC = 1000.0
MAX_TIME_PASSED = 10000
def distance_between_points(latitude1, longitude1, latitude2, longitude2):
return geopy.distance.geodesic((latitude1,longitude1),(latitude2,longitude2)).m
def is_in_date_range(date_x, date_start, date_end):
return date_x >= date_start and date_x <= date_end
def get_top3_cases(place, cases):
cases_extended = cases.copy()
cases_extended['distance'] = cases_extended.apply(lambda x: distance_between_points(x.Latitude, x.Longitude, place.latitudeE7 * 1.0 / E7, place.longitudeE7 * 1.0 / E7), axis=1)
cases_extended['time_passed'] = cases_extended.apply(lambda x: 0 if is_in_date_range(x.Time, datetime.fromtimestamp(float(place.visit_start_time) / TIMESTAMP_PREC), datetime.fromtimestamp(float(place.visit_end_time) / TIMESTAMP_PREC))\
else (datetime.fromtimestamp(int(place["visit_start_time"]) / TIMESTAMP_PREC) - x.Time).total_seconds() // 60, axis = 1)
cases_extended['time_passed'] = cases_extended.apply(lambda x: MAX_TIME_PASSED if x['time_passed'] < 0 else x['time_passed'], axis = 1)
return cases_extended.sort_values(by=["distance","time_passed"], ascending=True)[:3]
# -
get_top3_cases(df1.iloc[3], cases)
# +
DURATION_START_LIMIT = 15
DURATION_END_LIMIT = 180
MAX_DISTANCE = 1000 # meters
MAX_TIME = 7200 # 15 days in minutes
TIME_DIMINISHING_SPEED = 10
DISTANCE_DIMINISHING_SPEED = 6
# +
def calculate_harmonic_mean(x, y):
return 2 * x * y / (x + y)
def get_duration_multiplier(duration):
return min(1 + max(duration - DURATION_START_LIMIT, 0) / DURATION_END_LIMIT, 2)
places_by_time = df1.sort_values(by="visit_start_time",ascending=False)
def duration_between_timestamps(timestamp1, timestamp2):
return abs((datetime.fromtimestamp(float(timestamp2) / TIMESTAMP_PREC) - datetime.fromtimestamp(float(timestamp1) / TIMESTAMP_PREC)).seconds // 60)
components_values = []
for i in range(places_by_time.shape[0]):
place = places_by_time.iloc[i]
duration = duration_between_timestamps(place.visit_start_time, place.visit_end_time)
duration_multiplier = get_duration_multiplier(duration)
top3_cases = get_top3_cases(place, cases)
print(top3_cases)
for j in range(3):
top_case = top3_cases.iloc[j]
distance_factor = diminishing_importance(top_case.distance, MAX_DISTANCE, DISTANCE_DIMINISHING_SPEED)
time_factor = diminishing_importance(top_case.time_passed, MAX_TIME, TIME_DIMINISHING_SPEED)
print(place)
print(top_case)
print('--')
score = (duration_multiplier / 2) * calculate_harmonic_mean(distance_factor, time_factor)
components_values.append(top_case.values.tolist() + [place.address, duration, duration_multiplier, distance_factor, time_factor, place.visit_start_time, score])
components = pd.DataFrame(components_values, columns=["No", "Address", "Name", "City", "Time", "Latitude", "Longitude", "distance", "time_passed", "address", "duration","duration_multiplier","distance_factor","time_factor", "visit_start_time", "score"])
sorted_components = components.sort_values(by="score", ascending=False)
# -
diminishing_importance(5 * 600, MAX_TIME, TIME_DIMINISHING_SPEED)
diminishing_importance(550, MAX_DISTANCE, DISTANCE_DIMINISHING_SPEED)
# +
W1 = 0.7
W2 = 0.2
W3 = 0.1
risk = W1 * sorted_components.iloc[0].score + W2 * sorted_components.iloc[1].score + W3 * sorted_components.iloc[2].score
risk
# -
sorted_components
(datetime.fromtimestamp(int(sorted_components.iloc[0].visit_start_time) / 1000) - sorted_components.iloc[0].Time).total_seconds() // 60#.seconds // 60
datetime.fromtimestamp(int(sorted_components.iloc[0].visit_start_time) / 1000)
datetime.fromtimestamp(int(1584113900482) / 1000)
sorted_components = sorted_components[["Address", "Name", "City", "Time", "distance", "time_passed", "address", "score"]]
sorted_components = sorted_components.rename(columns={"Address": "Case Address", "address": "Your visit"})
sorted_components[:3]
df1.shape
185 * 3
|
risk-evaluation/1. location history preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime as dt
# +
#skip this cell
#
#OUTDATED
#OUTDATED
#OUTDATED
#OUTDATED
#
#
#get 4 closest points
#
#do change
folder_llpair = r'/Users/Pat/Documents/Harvard/'
filename_llpair = r'Nam12_Latitude_Longitude_Pairs.csv'
folder = r'/Users/Pat/Documents/Harvard/Interpolation/2016-05-18_1633/'
csv_name_loc = 'ws_path_2016-05-18_1633.csv'
csv_name_room = 'grid_closest_to_path_2016-05-18_1633.csv'
#do not change
#read in nam12 grid
df = pd.read_csv(folder_llpair+filename_llpair, delim_whitespace = 1)
#read in locations of windsond (delta 100m)
df_loc = pd.read_table(folder+csv_name_loc,sep=',')
#Output Dataframe
df_room = pd.DataFrame(columns = ['x1','x2','x3','x4','y1','y2','y3','y4','d1','d2','d3','d4','lat','lon','alt'])
lat = df_loc['lat'].tolist()
lon = df_loc['lon'].tolist()
alt = df_loc['alt'].tolist()
for i in range(len(lat)):
#calculate distance of points
df['Delta'] = (np.sqrt(np.square((abs(df.LAT - lat[i])) + np.square(abs(df.LON - lon[i])))))
#get the 4 closest points
x,y = (df.I[df.Delta.sort_values()[:4].index].tolist(), df.J[df.Delta.sort_values()[:4].index].tolist())
#prepare data for Dataframe
room = x + y + df.Delta.sort_values()[:4].tolist()
room.append(lat[i])
room.append(lon[i])
room.append(alt[i])
#append data to DataFrame
df_room.loc[len(df_room)] = [room[n] for n in range(15)]
#to csv
df_room.to_csv(path_or_buf = folder+csv_name_room, sep = ',',index = False)
df_room
# +
#Fetch 4 points around
#
#
#
#
#do change
folder_llpair = r'/Users/Pat/Documents/Harvard/'
filename_llpair = r'Nam12_Latitude_Longitude_Pairs.csv'
folder = r'/Users/Pat/Documents/Harvard/Interpolation/2016-05-18_1633/'
csv_name_loc = 'ws_path_2016-05-18_1633.csv'
csv_name_room = 'grid_around_path_2016-05-18_1633.csv'
#windsond info
t_peak = dt.timedelta(hours = 21,minutes = 32, seconds = 30) #UTC!!
alt_peak = 5211 #MSL
a_rize = 1.6 #m/s
#do not change
#read in nam12 grid
df = pd.read_csv(folder_llpair+filename_llpair, delim_whitespace = 1)
#read in locations of windsond (delta 100m)
df_loc = pd.read_table(folder+csv_name_loc,sep=',')
#Output Dataframe
df_room = pd.DataFrame(columns = ['x1','y1','x2','y2','x3','y3','x4','y4','d1','d2','d3','d4','lat','lon','alt','time_est'])
lat = df_loc['lat'].tolist()
lon = df_loc['lon'].tolist()
alt = df_loc['alt'].tolist()
for i in range(len(lat)):
#calculate distance of points
df['Delta'] = (np.sqrt(np.square((abs(df.LAT - lat[i])) + np.square(abs(df.LON - lon[i])))))
#x,y
x,y = df.loc[df['Delta'] == df['Delta'].min().item()].I.item(), df.loc[df['Delta'] == df['Delta'].min().item()].J.item()
#x2,y2
if df.loc[(df['I'] == x+1) & (df['J'] == y)].Delta.item() < df.loc[(df['I'] == x-1) & (df['J'] == y)].Delta.item():
x2, y2 = df.loc[(df['I'] == x+1) & (df['J'] == y)].I.item(), df.loc[(df['I'] == x+1) & (df['J'] == y)].J.item()
else:
x2, y2 = df.loc[(df['I'] == x-1) & (df['J'] == y)].I.item(), df.loc[(df['I'] == x-1) & (df['J'] == y)].J.item()
#x3.y3
if df.loc[(df['I'] == x) & (df['J'] == y+1)].Delta.item() < df.loc[(df['I'] == x) & (df['J'] == y-1)].Delta.item():
x3, y3 = df.loc[(df['I'] == x) & (df['J'] == y+1)].I.item(), df.loc[(df['I'] == x) & (df['J'] == y+1)].J.item()
else:
x3, y3 = df.loc[(df['I'] == x) & (df['J'] == y-1)].I.item(), df.loc[(df['I'] == x) & (df['J'] == y-1)].J.item()
#x4,y4
x4,y4 = x2,y3
#d1-d4
d1 = df.loc[(df['I'] == x) & (df['J'] == y)].Delta.item()
d2 = df.loc[(df['I'] == x2) & (df['J'] == y2)].Delta.item()
d3 = df.loc[(df['I'] == x3) & (df['J'] == y3)].Delta.item()
d4 = df.loc[(df['I'] == x4) & (df['J'] == y4)].Delta.item()
#print x,y,x2,y2,x3,y3,x4,y4
grid_list = [x,y,x2,y2,x3,y3,x4,y4,d1,d2,d3,d4]
#prepare data for Dataframe
room = grid_list
room.append(lat[i])
room.append(lon[i])
room.append(alt[i])
#add time row
room.append(t_peak - dt.timedelta(seconds = (alt_peak - int(alt[i]))/a_rize))
#append data to DataFrame
df_room.loc[len(df_room)] = [room[n] for n in range(16)]
#to csv
df_room.to_csv(path_or_buf = folder+csv_name_room, sep = ',',index = False)
df_room
# +
#test grid visually
n = 35
df_indy = df.loc[(df['LAT'] < 40.20) & (df['LAT'] > 39.40) & (df['LON'] > -86.70) & (df['LON'] < -85.60)]
df_point = df.loc[(df['I'] == df_room.x1[n]) & (df['J'] == df_room.y1[n])]
df_point2 = df.loc[(df['I'] == df_room.x2[n]) & (df['J'] == df_room.y2[n])]
df_point3 = df.loc[(df['I'] == df_room.x3[n]) & (df['J'] == df_room.y3[n])]
df_point4 = df.loc[(df['I'] == df_room.x4[n]) & (df['J'] == df_room.y4[n])]
plt.clf()
plt.plot(df_indy.LON, df_indy.LAT,'o')
plt.plot(df_room.lon[n],df_room.lat[n],'ro')
plt.plot(df_point.LON,df_point.LAT,'go')
plt.plot(df_point2.LON,df_point2.LAT,'go')
plt.plot(df_point3.LON,df_point3.LAT,'go')
plt.plot(df_point4.LON,df_point4.LAT,'go')
plt.show()
# +
#get all unique combinations
x1 = df_room.x1.tolist()
x2 = df_room.x2.tolist()
x3 = df_room.x3.tolist()
x4 = df_room.x4.tolist()
y1 = df_room.y1.tolist()
y2 = df_room.y2.tolist()
y3 = df_room.y3.tolist()
y4 = df_room.y4.tolist()
unique_grid_points = []
for n in range(len(x1)):
tup = (x1[n],y1[n])
if tup not in unique_grid_points:
unique_grid_points.append(tup)
tup = (x2[n],y2[n])
if tup not in unique_grid_points:
unique_grid_points.append(tup)
tup = (x3[n],y3[n])
if tup not in unique_grid_points:
unique_grid_points.append(tup)
tup = (x4[n],y4[n])
if tup not in unique_grid_points:
unique_grid_points.append(tup)
unique_grid_points.sort()
print unique_grid_points
# +
import subprocess
import time
#do change
folder_nam12 = '/Users/Pat/Documents/Harvard/Nam12/hysplit/'
#valid times are 0,3,6,9,12,15,18,21
folder_time_pair = [('20160518_nam12',18),('20160518_nam12',21)]
#fetch all needed profiles
for n,info in enumerate(folder_time_pair):
for entry in unique_grid_points:
x,y = entry
lat, lon = df.loc[(df['I'] == x) & (df['J'] == y)].LAT.item(), df.loc[(df['I'] == x) & (df['J'] == y)].LON.item()
print info[0],info[1],lat,lon
#First time point is AXXX-YYY, second is BXXX-YYY, ... (maxium suffix is len is 8)
ident = chr(n + ord('A'))
suffix = '%s%i-%i' %(ident,int(x),int(y))
subprocess.call("/Users/Pat/Hysplit4/exec/profile "
"-d%s "
"-f%s "
"-y%s -x%s "
"-o%s "
"-p%s".replace('\xe2','').replace('\x80','').replace('\xa8','')
%(folder_nam12, info[0], str(lat), str(lon), info[1], suffix), shell=True)
print 'Got profile: %i/%i' %(int(x),int(y))
print 'Done'
# +
#read in all 4 profiles
#do change
folder_profile = folder
profile_identifiers = ['A','B']
#windsond info
t_peak = dt.timedelta(hours = 17,minutes = 32, seconds = 30)
alt_peak = 5211 #MSL
a_rize = 1.6 #m/s
#do not change
df_8pts_profile = pd.DataFrame(columns = ['temp','wnd_spd','wnd_dir','relh','alt','time_ident'])
def bilinear_interpolation(x, y, points):
'''Interpolate (x,y) from values associated with four points.
The four points are a list of four triplets: (x, y, value).
The four points can be in any order. They should form a rectangle.
>>> bilinear_interpolation(12, 5.5,
... [(10, 4, 100),
... (20, 4, 200),
... (10, 6, 150),
... (20, 6, 300)])
165.0
'''
# See formula at: http://en.wikipedia.org/wiki/Bilinear_interpolation
points = sorted(points) # order points by x, then by y
(x1, y1, q11), (_x1, y2, q12), (x2, _y1, q21), (_x2, _y2, q22) = points
if x1 != _x1 or x2 != _x2 or y1 != _y1 or y2 != _y2:
raise ValueError('points do not form a rectangle')
if not x1 <= x <= x2 or not y1 <= y <= y2:
#print "ValueError('(x, y) not within the rectangle')"
return np.nan
return (q11 * (x2 - x) * (y2 - y) +
q21 * (x - x1) * (y2 - y) +
q12 * (x2 - x) * (y - y1) +
q22 * (x - x1) * (y - y1)) / ((x2 - x1) * (y2 - y1) + 0.0)
for n in range(len(df_room)):
#this flag will trigger, if there is not upper or lower level
skip_flag = 0
for t in profile_identifiers:
#read in the 4 profiles of each line in df_room
#profile1
name_profile = 'profile_%s%i-%i.txt' %(t,int(df_room.x1[n]),int(df_room.y1[n]))
df_prf1 = pd.read_table(folder_profile+ name_profile,sep=r'\s*', header = [11,12], engine='python')
lat1, lon1 = df.loc[(df['I'] == int(df_room.x1[n])) & (df['J'] == int(df_room.y1[n]))].LAT.item(), df.loc[(df['I'] == int(df_room.x1[n])) & (df['J'] == int(df_room.y1[n]))].LON.item()
#get the values for the level 1 lower than the alt
df_prf1_mod = df_prf1.loc[lambda x: df_prf1.HGTS.m<df_room.alt[n],:]
if len(df_prf1_mod) > 0:
#print df_prf1_mod.iloc[-1,:]
uwnd_dir1_low = df_prf1_mod.iloc[-1,:]['UWND']['W->E']
vwnd_dir1_low = df_prf1_mod.iloc[-1,:]['VWND']['S->N']
wnd_dir1_low = 270 - (np.arctan2(vwnd_dir1_low, uwnd_dir1_low) * (180 / np.pi))
if wnd_dir1_low > 360:
wnd_dir1_low = wnd_dir1_low - 360
hgts1_low = df_prf1_mod.iloc[-1,:].HGTS.m
temp1_low = df_prf1_mod.iloc[-1,:].TEMP.oC
relh1_low = df_prf1_mod.iloc[-1,:]['RELH']['%']
uwnd_spd1_low = df_prf1_mod.iloc[-1,:]['UWND']['m/s']
vwnd_spd1_low = df_prf1_mod.iloc[-1,:]['VWND']['m/s']
wnd_spd1_low = np.sqrt(np.square(uwnd_spd1_low)+np.square(vwnd_spd1_low))
#print wnd_dir1_low, hgts1_low, temp1_low, relh1_low, wnd_spd1_low
else:
skip_flag = 1
#get the values for the level 1 higher than the alt
df_prf1_mod = df_prf1.loc[lambda x: df_prf1.HGTS.m>df_room.alt[n],:]
if len(df_prf1_mod) > 0:
#print df_prf1_mod.iloc[0,:]
uwnd_dir1_hi = df_prf1_mod.iloc[0,:]['UWND']['W->E']
vwnd_dir1_hi = df_prf1_mod.iloc[0,:]['VWND']['S->N']
wnd_dir1_hi = 270 - (np.arctan2(vwnd_dir1_hi, uwnd_dir1_hi) * (180 / np.pi))
if wnd_dir1_hi > 360:
wnd_dir1_hi = wnd_dir1_hi - 360
hgts1_hi = df_prf1_mod.iloc[0,:].HGTS.m
temp1_hi = df_prf1_mod.iloc[0,:].TEMP.oC
relh1_hi = df_prf1_mod.iloc[0,:]['RELH']['%']
uwnd_spd1_hi = df_prf1_mod.iloc[0,:]['UWND']['m/s']
vwnd_spd1_hi = df_prf1_mod.iloc[0,:]['VWND']['m/s']
wnd_spd1_hi = np.sqrt(np.square(uwnd_spd1_hi)+np.square(vwnd_spd1_hi))
#print wnd_dir1_hi, hgts1_hi, temp1_hi, relh1_hi, wnd_spd1_hi
else:
skip_flag = 1
#profile2
name_profile = 'profile_%s%i-%i.txt' %(t,int(df_room.x2[n]),int(df_room.y2[n]))
df_prf2 = pd.read_table(folder_profile+ name_profile,sep=r'\s*', header = [11,12], engine='python')
lat2, lon2 = df.loc[(df['I'] == int(df_room.x2[n])) & (df['J'] == int(df_room.y2[n]))].LAT.item(), df.loc[(df['I'] == int(df_room.x2[n])) & (df['J'] == int(df_room.y2[n]))].LON.item()
#get the values for the level 1 lower than the alt
df_prf2_mod = df_prf2.loc[lambda x: df_prf2.HGTS.m<df_room.alt[n],:]
if len(df_prf2_mod) > 0:
#print df_prf2_mod.iloc[-1,:]
uwnd_dir2_low = df_prf2_mod.iloc[-1,:]['UWND']['W->E']
vwnd_dir2_low = df_prf2_mod.iloc[-1,:]['VWND']['S->N']
wnd_dir2_low = 270 - (np.arctan2(vwnd_dir2_low, uwnd_dir2_low) * (180 / np.pi))
if wnd_dir2_low > 360:
wnd_dir2_low = wnd_dir2_low - 360
hgts2_low = df_prf2_mod.iloc[-1,:].HGTS.m
temp2_low = df_prf2_mod.iloc[-1,:].TEMP.oC
relh2_low = df_prf2_mod.iloc[-1,:]['RELH']['%']
uwnd_spd2_low = df_prf2_mod.iloc[-1,:]['UWND']['m/s']
vwnd_spd2_low = df_prf2_mod.iloc[-1,:]['VWND']['m/s']
wnd_spd2_low = np.sqrt(np.square(uwnd_spd2_low)+np.square(vwnd_spd2_low))
#print wnd_dir2_low, hgts2_low, temp2_low, relh2_low, wnd_spd2_low
else:
skip_flag = 1
#get the values for the level 1 higher than the alt
df_prf2_mod = df_prf2.loc[lambda x: df_prf2.HGTS.m>df_room.alt[n],:]
if len(df_prf2_mod) > 0:
#print df_prf2_mod.iloc[0,:]
uwnd_dir2_hi = df_prf2_mod.iloc[0,:]['UWND']['W->E']
vwnd_dir2_hi = df_prf2_mod.iloc[0,:]['VWND']['S->N']
wnd_dir2_hi = 270 - (np.arctan2(vwnd_dir2_hi, uwnd_dir2_hi) * (180 / np.pi))
if wnd_dir2_hi > 360:
wnd_dir2_hi = wnd_dir2_hi - 360
hgts2_hi = df_prf2_mod.iloc[0,:].HGTS.m
temp2_hi = df_prf2_mod.iloc[0,:].TEMP.oC
relh2_hi = df_prf2_mod.iloc[0,:]['RELH']['%']
uwnd_spd2_hi = df_prf2_mod.iloc[0,:]['UWND']['m/s']
vwnd_spd2_hi = df_prf2_mod.iloc[0,:]['VWND']['m/s']
wnd_spd2_hi = np.sqrt(np.square(uwnd_spd2_hi)+np.square(vwnd_spd2_hi))
#print wnd_dir2_hi, hgts2_hi, temp2_hi, relh2_hi, wnd_spd2_hi
else:
skip_flag = 1
#profile3
name_profile = 'profile_%s%i-%i.txt' %(t,int(df_room.x3[n]),int(df_room.y3[n]))
df_prf3 = pd.read_table(folder_profile+ name_profile,sep=r'\s*', header = [11,12], engine='python')
lat3, lon3 = df.loc[(df['I'] == int(df_room.x3[n])) & (df['J'] == int(df_room.y3[n]))].LAT.item(), df.loc[(df['I'] == int(df_room.x3[n])) & (df['J'] == int(df_room.y3[n]))].LON.item()
#get the values for the level 1 lower than the alt
df_prf3_mod = df_prf3.loc[lambda x: df_prf3.HGTS.m<df_room.alt[n],:]
if len(df_prf3_mod) > 0:
#print df_prf3_mod.iloc[-1,:]
uwnd_dir3_low = df_prf3_mod.iloc[-1,:]['UWND']['W->E']
vwnd_dir3_low = df_prf3_mod.iloc[-1,:]['VWND']['S->N']
wnd_dir3_low = 270 - (np.arctan2(vwnd_dir3_low, uwnd_dir3_low) * (180 / np.pi))
if wnd_dir3_low > 360:
wnd_dir3_low = wnd_dir3_low - 360
hgts3_low = df_prf3_mod.iloc[-1,:].HGTS.m
temp3_low = df_prf3_mod.iloc[-1,:].TEMP.oC
relh3_low = df_prf3_mod.iloc[-1,:]['RELH']['%']
uwnd_spd3_low = df_prf3_mod.iloc[-1,:]['UWND']['m/s']
vwnd_spd3_low = df_prf3_mod.iloc[-1,:]['VWND']['m/s']
wnd_spd3_low = np.sqrt(np.square(uwnd_spd3_low)+np.square(vwnd_spd3_low))
#print wnd_dir3_low, hgts3_low, temp3_low, relh3_low, wnd_spd3_low
else:
skip_flag = 1
#get the values for the level 1 higher than the alt
df_prf3_mod = df_prf3.loc[lambda x: df_prf3.HGTS.m>df_room.alt[n],:]
if len(df_prf3_mod) > 0:
#print df_prf3_mod.iloc[0,:]
uwnd_dir3_hi = df_prf3_mod.iloc[0,:]['UWND']['W->E']
vwnd_dir3_hi = df_prf3_mod.iloc[0,:]['VWND']['S->N']
wnd_dir3_hi = 270 - (np.arctan2(vwnd_dir3_hi, uwnd_dir3_hi) * (180 / np.pi))
if wnd_dir3_hi > 360:
wnd_dir3_hi = wnd_dir3_hi - 360
hgts3_hi = df_prf3_mod.iloc[0,:].HGTS.m
temp3_hi = df_prf3_mod.iloc[0,:].TEMP.oC
relh3_hi = df_prf3_mod.iloc[0,:]['RELH']['%']
uwnd_spd3_hi = df_prf3_mod.iloc[0,:]['UWND']['m/s']
vwnd_spd3_hi = df_prf3_mod.iloc[0,:]['VWND']['m/s']
wnd_spd3_hi = np.sqrt(np.square(uwnd_spd3_hi)+np.square(vwnd_spd3_hi))
#print wnd_dir3_hi, hgts3_hi, temp3_hi, relh3_hi, wnd_spd3_hi
else:
skip_flag = 1
#profile4
name_profile = 'profile_%s%i-%i.txt' %(t,int(df_room.x4[n]),int(df_room.y4[n]))
df_prf4 = pd.read_table(folder_profile+ name_profile,sep=r'\s*', header = [11,12], engine='python')
lat4, lon4 = df.loc[(df['I'] == int(df_room.x4[n])) & (df['J'] == int(df_room.y4[n]))].LAT.item(), df.loc[(df['I'] == int(df_room.x4[n])) & (df['J'] == int(df_room.y4[n]))].LON.item()
#get the values for the level 1 lower than the alt
df_prf4_mod = df_prf4.loc[lambda x: df_prf4.HGTS.m<df_room.alt[n],:]
if len(df_prf4_mod) > 0:
#print df_prf4_mod.iloc[-1,:]
uwnd_dir4_low = df_prf4_mod.iloc[-1,:]['UWND']['W->E']
vwnd_dir4_low = df_prf4_mod.iloc[-1,:]['VWND']['S->N']
wnd_dir4_low = 270 - (np.arctan2(vwnd_dir4_low, uwnd_dir4_low) * (180 / np.pi))
if wnd_dir4_low > 360:
wnd_dir4_low = wnd_dir4_low - 360
hgts4_low = df_prf4_mod.iloc[-1,:].HGTS.m
temp4_low = df_prf4_mod.iloc[-1,:].TEMP.oC
relh4_low = df_prf4_mod.iloc[-1,:]['RELH']['%']
uwnd_spd4_low = df_prf4_mod.iloc[-1,:]['UWND']['m/s']
vwnd_spd4_low = df_prf4_mod.iloc[-1,:]['VWND']['m/s']
wnd_spd4_low = np.sqrt(np.square(uwnd_spd4_low)+np.square(vwnd_spd4_low))
#print wnd_dir4_low, hgts4_low, temp4_low, relh4_low, wnd_spd4_low
else:
skip_flag = 1
#get the values for the level 1 higher than the alt
df_prf4_mod = df_prf4.loc[lambda x: df_prf4.HGTS.m>df_room.alt[n],:]
if len(df_prf4_mod) > 0:
#print df_prf1_mod.iloc[0,:]
uwnd_dir4_hi = df_prf4_mod.iloc[0,:]['UWND']['W->E']
vwnd_dir4_hi = df_prf4_mod.iloc[0,:]['VWND']['S->N']
wnd_dir4_hi = 270 - (np.arctan2(vwnd_dir4_hi, uwnd_dir4_hi) * (180 / np.pi))
if wnd_dir4_hi > 360:
wnd_dir4_hi = wnd_dir4_hi - 360
hgts4_hi = df_prf4_mod.iloc[0,:].HGTS.m
temp4_hi = df_prf4_mod.iloc[0,:].TEMP.oC
relh4_hi = df_prf4_mod.iloc[0,:]['RELH']['%']
uwnd_spd4_hi = df_prf4_mod.iloc[0,:]['UWND']['m/s']
vwnd_spd4_hi = df_prf4_mod.iloc[0,:]['VWND']['m/s']
wnd_spd4_hi = np.sqrt(np.square(uwnd_spd4_hi)+np.square(vwnd_spd4_hi))
#print wnd_dir4_hi, hgts4_hi, temp4_hi, relh4_hi, wnd_spd4_hi
else:
skip_flag = 1
if skip_flag == 0:
#do the 8 point interpolation (first bilinear (4point) then linear (2xbilinear))
#linearize the grid for bilinear interpolation
bilin_x1 = (lat1+lat2)/2
bilin_x2 = (lat3+lat4)/2
bilin_y1 = (lon1+lon3)/2
bilin_y2 = (lon2+lon4)/2
#print bilin_x1,bilin_y1,bilin_x2,bilin_y2,df_room.lat[n],df_room.lon[n]
#hgts
par_input = [hgts1_low,hgts2_low,hgts3_low,hgts4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
hgts_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [hgts1_hi,hgts2_hi,hgts3_hi,hgts4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
hgts_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
#temp
par_input = [temp1_low,temp2_low,temp3_low,temp4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
temp_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [temp1_hi,temp2_hi,temp3_hi,temp4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
temp_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
fp = [temp_low, temp_hi]
temp = np.interp(df_room.alt[n], [hgts_low,hgts_hi], fp)
#wnd_spd
par_input = [wnd_spd1_low,wnd_spd2_low,wnd_spd3_low,wnd_spd4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_spd_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [wnd_spd1_hi,wnd_spd2_hi,wnd_spd3_hi,wnd_spd4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_spd_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
fp = [wnd_spd_low, wnd_spd_hi]
wnd_spd = np.interp(df_room.alt[n], [hgts_low,hgts_hi], fp)
#wnd_dir
par_input = [wnd_dir1_low,wnd_dir2_low,wnd_dir3_low,wnd_dir4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_dir_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [wnd_dir1_hi,wnd_dir2_hi,wnd_dir3_hi,wnd_dir4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_dir_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
fp = [wnd_dir_low, wnd_dir_hi]
wnd_dir = np.interp(df_room.alt[n], [hgts_low,hgts_hi], fp)
#wnd_spd
par_input = [wnd_spd1_low,wnd_spd2_low,wnd_spd3_low,wnd_spd4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_spd_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [wnd_spd1_hi,wnd_spd2_hi,wnd_spd3_hi,wnd_spd4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
wnd_spd_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
fp = [wnd_spd_low, wnd_spd_hi]
wnd_spd = np.interp(df_room.alt[n], [hgts_low,hgts_hi], fp)
#relh
par_input = [relh1_low,relh2_low,relh3_low,relh4_low]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
relh_low = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
par_input = [relh1_hi,relh2_hi,relh3_hi,relh4_hi]
bilin_grid = [(bilin_x1, bilin_y1, par_input[0]),(bilin_x1, bilin_y2, par_input[1]),
(bilin_x2, bilin_y1, par_input[2]),(bilin_x2, bilin_y2, par_input[3])]
relh_hi = bilinear_interpolation(df_room.lat[n], df_room.lon[n], bilin_grid)
fp = [relh_low, relh_hi]
relh = np.interp(df_room.alt[n], [hgts_low,hgts_hi], fp)
if t == 'A':
vec_A = [temp, wnd_spd, wnd_dir, relh, df_room.alt[n],'A']
#df_8pts_profile.loc[len(df_8pts_profile)] = [vec_A[i] for i in range(len(vec_A))]
elif t == 'B':
vec_B = [temp, wnd_spd, wnd_dir, relh, df_room.alt[n],'B']
#df_8pts_profile.loc[len(df_8pts_profile)] = [vec_B[i] for i in range(len(vec_B))]
#elif t == 'C':
# vec_C = [temp, wnd_spd, wnd_dir, relh, df_room.alt[n],'C']
# df_8pts_profile.loc[len(df_8pts_profile)] = [vec_C[i] for i in range(len(vec_C))]
#interpolation in time
vec_final = []
for m in range(4):
profile_times = [(dt.timedelta(hours = 18) + dt.timedelta(hours = 1,minutes = 30)).total_seconds(),
(dt.timedelta(hours = 21) + dt.timedelta(hours = 1,minutes = 30)).total_seconds()]
profile_values = [vec_A[m],vec_B[m]]
vec_final.append(np.interp(df_room.time_est[n].total_seconds(), profile_times, profile_values))
vec_final.append(df_room.alt[n])
vec_final.append('Final')
df_8pts_profile.loc[len(df_8pts_profile)] = [vec_final[i] for i in range(len(vec_final))]
print 'Finished level: %i, %sm' %(n, df_room.alt[n])
df_8pts_profile
# +
from matplotlib.backends.backend_pdf import PdfPages
#do change
nam12_h_offset = ' 16pt_interp'
#nam12_h_offset_B = '+21h'
#windsond csv
folder_ws = folder
filename_ws = 'ws_profile_2016-05-18_1633.txt'
ground_height = 213
#nam12profiles
df_8pts_profile_A = df_8pts_profile.loc[df_8pts_profile.time_ident == 'Final']
#df_8pts_profile_B = df_8pts_profile.loc[df_8pts_profile.time_ident == 'B']
pp = PdfPages(folder + '8pt_+1821.pdf')
plot_title = 'fallCreek_18052016_1633'
#dataframe for windsond data
df_ws = pd.read_csv(folder_ws+filename_ws, header = [0,1], delim_whitespace = 1)
#x = temp, y = height
plt.clf()
plt.figure(1)
plt.plot(df_8pts_profile_A.temp,df_8pts_profile_A.alt,'b',label = 'nam12'+nam12_h_offset)
#plt.plot(df_8pts_profile_B.temp,df_8pts_profile_B.alt,'g',label = 'nam12'+nam12_h_offset_B)
plt.plot(df_ws['temp'],df_ws['alt']+ground_height,'r', label = 'windsond')
plt.ylabel('Height [m]')
plt.xlabel('Temp [C]')
plt.title('TEMP: '+plot_title)
plt.grid(True)
plt.legend(loc='best')
plt.savefig(pp, format='pdf')
#plt.show()
#x = temp, y = windspeed
plt.figure(2)
plt.plot(df_8pts_profile_A.wnd_spd,df_8pts_profile_A.alt,'b',label = 'nam12'+nam12_h_offset)
#plt.plot(df_8pts_profile_B.wnd_spd,df_8pts_profile_B.alt,'g',label = 'nam12'+nam12_h_offset_B)
plt.plot(df_ws['spd'],df_ws['alt']+ground_height,'r', label = 'windsond')
plt.ylabel('Height [m]')
plt.xlabel('Windspeed [m/s]')
plt.title('WINDSPEED: '+plot_title)
plt.grid(True)
plt.legend(loc='best')
pp.savefig()
#plt.show()
#x = hum, y = height
plt.figure(3)
plt.plot(df_8pts_profile_A.relh,df_8pts_profile_A.alt,'b',label = 'nam12'+nam12_h_offset)
#plt.plot(df_8pts_profile_B.relh,df_8pts_profile_B.alt,'g',label = 'nam12'+nam12_h_offset_B)
plt.plot(df_ws['hum'],df_ws['alt']+ground_height,'r', label = 'windsond')
plt.ylabel('Height [m]')
plt.xlabel('Humidity [%]')
plt.title('HUM: '+plot_title)
plt.grid(True)
plt.legend(loc='best')
pp.savefig()
#plt.show()
#x = wind-dir, y = height
plt.figure(4)
plt.plot(df_8pts_profile_A.wnd_dir,df_8pts_profile_A.alt,'b',label = 'nam12'+nam12_h_offset)
#plt.plot(df_8pts_profile_B.wnd_dir,df_8pts_profile_B.alt,'g',label = 'nam12'+nam12_h_offset_B)
plt.plot(df_ws['wind-dir'],df_ws['alt']+ground_height,'r', label = 'windsond')
plt.ylabel('Height [m]')
plt.xlabel('Wind direction [o]')
plt.title('WIND-DIR: '+plot_title)
plt.grid(True)
plt.legend(loc='best')
pp.savefig()
pp.close()
plt.show()
# -
df_room.time_est[0].total_seconds()
|
2016-05-18_1633/.ipynb_checkpoints/Nam12-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:lab_conda]
# language: python
# name: conda-env-lab_conda-py
# ---
# +
# Lab 12 Character Sequence RNN
import torch
import torch.nn as nn
from torch.autograd import Variable
torch.manual_seed(777) # reproducibility
sentence = ("if you want to build a ship, don't drum up people together to "
"collect wood and don't assign them tasks and work, but rather "
"teach them to long for the endless immensity of the sea.")
char_set = list(set(sentence))
char_dic = {w: i for i, w in enumerate(char_set)}
# -
char_dic
# +
# hyperparameters
learning_rate = 0.1
num_epochs = 500
input_size = len(char_set) # RNN input size (one hot size)
hidden_size = len(char_set) # RNN output size
num_classes = len(char_set) # final output size (RNN or softmax, etc.)
sequence_length = 10 # any arbitrary number
num_layers = 2 # number of layers in RNN
dataX = []
dataY = []
for i in range(0, len(sentence) - sequence_length):
x_str = sentence[i:i + sequence_length]
y_str = sentence[i + 1: i + sequence_length + 1]
print(i, x_str, '->', y_str)
x = [char_dic[c] for c in x_str] # x str to index
y = [char_dic[c] for c in y_str] # y str to index
dataX.append(x)
dataY.append(y)
batch_size = len(dataX)
x_data = torch.Tensor(dataX)
y_data = torch.LongTensor(dataY)
# +
# one hot encoding
def one_hot(x, num_classes):
idx = x.long()
idx = idx.view(-1, 1)
x_one_hot = torch.zeros(x.size()[0] * x.size()[1], num_classes)
x_one_hot.scatter_(1, idx, 1)
x_one_hot = x_one_hot.view(x.size()[0], x.size()[1], num_classes)
return x_one_hot
x_one_hot = one_hot(x_data, num_classes)
inputs = Variable(x_one_hot)
labels = Variable(y_data)
# -
x_one_hot
# +
class LSTM(nn.Module):
def __init__(self, num_classes, input_size, hidden_size, num_layers):
super(LSTM, self).__init__()
self.num_classes = num_classes
self.num_layers = num_layers
self.input_size = input_size
self.hidden_size = hidden_size
self.sequence_length = sequence_length
# Set parameters for RNN block
# Note: batch_first=False by default.
# When true, inputs are (batch_size, sequence_length, input_dimension)
# instead of (sequence_length, batch_size, input_dimension)
self.lstm = nn.LSTM(input_size=input_size, hidden_size=hidden_size,
num_layers=num_layers,batch_first=True)
# Fully connected layer
self.fc = nn.Linear(hidden_size, num_classes)
def forward(self, x):
# Initialize hidden and cell states
h_0 = Variable(torch.zeros(
self.num_layers, x.size(0), self.hidden_size))
c_0 = Variable(torch.zeros(
self.num_layers, x.size(0), self.hidden_size))
# h_0 = Variable(torch.zeros(
# self.num_layers, x.size(0), self.hidden_size))
# c_0 = Variable(torch.zeros(
# self.num_layers, x.size(0), self.hidden_size))
# Propagate input through LSTM
# Input: (batch, seq_len, input_size)
out, _ = self.lstm(x, (h_0, c_0))
# Note: the output tensor of LSTM in this case is a block with holes
# > add .contiguous() to apply view()
out = out.contiguous().view(-1, self.hidden_size)
# Return outputs applied to fully connected layer
out = self.fc(out)
return out
# +
# Instantiate RNN model
lstm = LSTM(num_classes, input_size, hidden_size, num_layers)
# Set loss and optimizer function
criterion = torch.nn.CrossEntropyLoss() # Softmax is internally computed.
optimizer = torch.optim.Adam(lstm.parameters(), lr=learning_rate)
# +
# Train the model
for epoch in range(num_epochs):
outputs = lstm(inputs)
optimizer.zero_grad()
# obtain the loss function
# flatten target labels to match output
loss = criterion(outputs, labels.view(-1))
loss.backward()
optimizer.step()
# obtain the predicted indices of the next character
_, idx = outputs.max(1)
idx = idx.data.numpy()
idx = idx.reshape(-1, sequence_length) # (170,10)
# display the prediction of the last sequence
result_str = [char_set[c] for c in idx[-2]]
print("epoch: %d, loss: %1.3f" % (epoch + 1, loss.data[0]))
print("Predicted string: ", ''.join(result_str))
print("Learning finished!")
# -
|
Practice5/15_Long_CHar_RNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#02_end_to_end_machine_learning_project 练习#
# +
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "end_to_end_project"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
# -
|
Untitled1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="uQ62eZUEvexJ"
# # Exercício de Programação 2: PageRank
#
# <font color="red">**Prazo de submissão: 23:55 do dia 18/01/2021** </font>
#
# 2020.2 Álgebra Linear Computacional - DCC - UFMG
#
# <NAME> Fabricio
#
# Instruções:
# * Antes de submeter suas soluções, certifique-se de que tudo roda como esperado. Primeiro, **reinicie o kernel** no menu, selecione Kernel$\rightarrow$Restart e então execute **todas as células** (no menu, Cell$\rightarrow$Run All)
# * **Apenas o arquivo .py deve ser submetido**. Você deve salvar o seu notebook em Python script (no menu, File $\rightarrow$ Download .py) e enviar o script Python no Ambiente de Programação Virtual.
# * **Preste bastante atenção nos nomes das variáveis e métodos** (irá estar em negrito), se elas estiverem diferentes do que foi pedido no exercício, *sua resposta será considerada incorreta pelo corretor automático*.
# * Os exercícios desse EP utilizam os conceitos do PageRank vistos em aula, inclusive o de *matrizes de transição*, porém, neste notebook em alguns exercícios as matrizes podem estar num formato **transposto** do que vimos em aula, ou seja, com os links de saída da página $i$ na linha $i$, ao invés da coluna (mais detalhes sobre isso no notebook).
# * Não deixe de preencher seu nome e número de matrícula na célula a seguir
# + [markdown] id="RPKeHzw6vexK"
# **Nome:** <NAME>
#
# **Matrícula:** 2020054293
#
# * Todo material consultado na Internet deve ser referenciado (incluir URL).
#
# Este trabalho está dividido em três partes:
# * **Parte 0**: Esta parte não vale nota, mas é fundamental para entender o que se pede a seguir.
# * **Parte 1**: Pagerank sem saltos aleatórios em grafo pequeno
# * **Parte 2**: Pagerank (com saltos aleatórios) em grafo pequeno
# + [markdown] id="Oz9PxS-YvexL"
# ## Parte 0: Revisão de conceitos
#
# I. O **primeiro autovetor** (isto é, o autovetor associado ao maior autovalor em módulo) pode ser calculado rapidamente através do método da potência, desde que o *gap* entre o maior e o segundo maior autovalor (em módulo) seja grande. Uma implementação simples do método da potência é mostrada a seguir.
# + id="_HiRBalhvexL"
import numpy as np
def powerMethod(A, niter=10):
n = len(A)
w = np.ones((n,1))/n
for i in range(niter):
w = A.dot(w)
return w
# + [markdown] id="h_W0jcC62IWZ"
#
# II. Dado um grafo $G=(V, E)$ com n vértices, podemos obter **uma matriz de transição** $A$ de tamanho $n \times n$ em que cada elemento $ij$ na matriz representa uma aresta direcionada do vértice $i$ para o vértice $j$. Por exemplo, para o seguinte grafo direcionado:
#
# <img src='https://www.dropbox.com/s/wmk8v8worinoqk0/grafo-simples-2.PNG?raw=1'>
#
# a matriz de transição seria:
#
# $$
# A =
# \begin{bmatrix}
# 0 & 1 & 0 & 0 & 0 \\
# 0 & 0 & 1 & 0 & 0 \\
# 1 & 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 0 & 1 \\
# 0 & 1 & 0 & 0 & 0
# \end{bmatrix}
# $$
#
# **Essa notação é um pouco diferente da que vimos em aula**, já que no vídeo e os slides as saídas de cada página eram dadas nas colunas, e não nas linhas. Mesmo com essa diferença, podemos realizar as mesmas operações que vimos em aula. Por exemplo:
#
# - Para multiplicar a matriz $A$ por um vetor, podemos usar por exemplo: $A^\top \textbf{v}$. Ou alternativamente podemos usar $\textbf{v}^\top A$.
# - Para obter a quantidade de links de saída, precisamos somar ao longo das linhas, ao invés das colunas.
# + [markdown] id="QI7kGQZZvexQ"
# III. Dado um grafo $G=(V,E)$, podemos obter uma **matriz de probabilidade de transição** $P$ dividindo-se cada linha de $A$ pela soma dos elementos da linha. Seja $D$ a matriz diagonal contendo a soma das linhas de $A$. Temos que
#
# $$
# P = D^{-1} \times A.
# $$
# + [markdown] id="b2cJ5hesvexR"
# IV. A matriz de probabilidade de transição $P$ de certos grafos direcionados satisfaz
#
# $$
# v^\top P = v^\top \textrm{ou $P^\top v = v$},
# $$
#
# onde $v$ é o primeiro autovetor de $P^\top$. A equação da direita é mais fácil de ser utilizada, pois ela tem a forma canônica $Ax=b$. Já a equação da esquerda é mais fácil de ser interpretada. Para todo $j=1,\ldots,n$,
#
# $$
# \sum_{i=1} v_i P_{ij} = v_j \\
# \Rightarrow \sum_{i=1} v_i \frac{A_{ij}}{D_{ii}} = v_j \\
# \Rightarrow \sum_{i:(i,j) \in E} v_i \frac{1}{D_{ii}} = v_j
# $$
#
# em que nesse último somatório, a soma se dá apenas aos vértices $i$ que possuem link para $j$, ou seja, a aresta $(i,j)$ pertence ao conjunto $E$ de arestas $i:(i,j) \in E$.
# + [markdown] id="oW-klTh6vexR"
# V. Assuma que $v$ seja normalizado de forma que $\sum_j v_j = 1$. O PageRank (sem saltos) de um vértice $j$ é dado por $v_j$, onde $v$ é o primeiro autovetor de $P^\top$. Esta é uma maneira de medir sua relevância. A intuição da Equação $\sum_{i:(i,j) \in E} v_i /D_{ii} = v_j$ é que a relevância de $j$ é a soma das relevâncias dos vértices $i$ que apontam para $j$ normalizados pelos seus respectivos graus de saída.
# + [markdown] id="znlbzFthvexS"
# ## Parte 1: Pagerank sem saltos aleatórios em grafo pequeno
#
# Considere o grafo a seguir composto por $n=4$ vértices e $m=8$ arestas.
# <img src='https://www.dropbox.com/s/oxibt5daw1g4dw3/directedgraph.png?raw=1'>
#
# Certifique-se de que encontrou as $m=8$ arestas.
# + [markdown] id="crXuarjrvexS"
# **1.1** Crie um numpy array chamado <b> A </b>, contendo a matriz de adjacência.
# + id="MtCaXDS3vexT"
#Insira seu código para a questão 1.1 aqui
A = np.array([
[0,1,1,0],
[0,0,1,1],
[0,0,0,1],
[1,1,1,0],
])
# + [markdown] id="jxIdLmlyvexZ"
# **1.2** Escreva uma função chamada <b>matrizDeTransicao</b> que recebe como entrada uma matriz $n \times n$ e retorna a matriz de probabilidade de transição desta matriz. Aplique a função em <b>A</b> e armazene o resultado na variável <b>P</b>, e depois imprima <b>P</b>.
# + id="KYbc3agDvexZ"
#Insira seu código para a questão 1.2 aqui
def matrizDeTransicao(A):
d = np.sum(A, axis=1)
D = np.diag(d)
P = np.matmul(np.linalg.inv(D), A)
return P
P = matrizDeTransicao(A)
print(P)
# + [markdown] id="kssQbOAavexe"
# **1.3** Use a função <i>np.linalg.eig</i> para calcular o primeiro autovetor de $P^\top$. Normalize o autovetor pela sua soma em uma variável chamada <b>autovec</b> e imprima o resultado. (Observação: os elementos do autovetor serão retornados como números complexos, mas a parte imaginária será nula e pode ser ignorada.)
# + id="f16Ed-KPvexe"
#Insira seu código para a questão 1.3 aqui
autoval,autovec = np.linalg.eig(P.T)
autovec = (autovec[:,0])/np.sum(autovec[:,0].real)
autovec = autovec.real
print(autovec)
# + [markdown] id="4jKEgUnEvexh"
# **1.4** Verifique que o método da potência aplicado a $P^\top$ retorna uma aproximação para o primeiro autovetor. Atribua o resultado retornado pelo método na variável <b> result_pm </b> e imprima-o.
# + id="2HAkj9fOvexi"
#Insira seu código para a questão 1.4 aqui
result_pm = powerMethod(np.transpose(P))
print(result_pm)
# + [markdown] id="pyAZd73fvexn"
# **1.5** Implemente uma função <b>powerMethodEps(A, epsilon)</b> que executa o método da potência até que a condição de convergência $\|w_{t} - w_{t-1}\| < \epsilon$ seja atingida. Para a matriz $P^\top$ com $\epsilon=10^{-5}$, armazene o resultado do método da potência na variável <b>result_pm_eps</b> *(1.5.1)*, e o número de iterações na variável <b>nb_iters</b> *(1.5.2)*.
#
# Imprima o resultado das duas variáveis.
# + id="6dk2a5i7vexo"
#Insira seu código para a questão 1.5 aqui
def powerMethodEps(A, epsilon=1e-5):
nb_iters = 1
n = len(A)
w = np.ones((n,1))/n
w_new = A @ w
while np.linalg.norm(w - w_new) > epsilon:
w = w_new
w_new = A @ w
nb_iters += 1
return w, nb_iters
result_pm_eps, nb_iters = powerMethodEps(P.T)
print(result_pm_eps, nb_iters)
# + [markdown] id="MQwHkfXvvexr"
# ## Parte II: Pagerank (com saltos aleatórios) em grafo pequeno
#
# Agora iremos modificar a matriz A de forma a:
# * adicionar um novo vértice 4, e
# * adicionar uma aresta de 3 para 4.
#
# Obviamente a matriz de probabilidade de transição não está definida para a nova matriz $A$. Vértices que não possuem arestas de saída (como o vértice 4) são conhecidos como *dangling nodes*. Para resolver este e outros problemas, incluiremos a possibilidade de realizar saltos aleatórios de um vértice para qualquer outro vértice.
#
# Em particular, assume-se que com probabilidade $\alpha$, seguimos uma das arestas de saída em $A$ e, com probabilidade $1-\alpha$ realizamos um salto aleatório, isto é, transicionamos do vértice $v$ para um dos $n$ vértices do grafo (incluindo $v$) escolhido uniformemente. Quando não existem *dangling nodes*, a nova matriz de probabilidade de transição é dada por
#
# $$
# P = \alpha D^{-1} A + (1-\alpha) \frac{\mathbf{1}\mathbf{1}^\top}{n}
# $$
#
# Quando existem *dangling nodes*, a única possibilidade a partir desses nós é fazer saltos aleatórios. Mais precisamente, se $i$ é um vértice sem arestas de saída, desejamos que a $i$-ésima linha de $P$ seja o vetor $[1/n,\ldots,1/n]$. Uma forma de satisfazer essa definição é preencher com 1's as linhas de $A$ que correspondem aos *dangling nodes*. Uma desvantagem desta estratégia é que faz com que $A$ fique mais densa (mais elementos não-nulos).
#
# Um valor típico usado para $\alpha$ é $0.85$.
# + [markdown] id="KuYrPgEUvexr"
# **2.1** Crie um novo numpy array chamado <b> A_new </b> contendo o vértice 4 e a aresta (3,4).
# + id="YcLYaqbHvexs"
#Insira seu código para a questão 2.1 aqui
A_new = np.array([
[0,1,1,0,0],
[0,0,1,1,0],
[0,0,0,1,0],
[1,1,1,0,1],
[0,0,0,0,0],
])
print(A_new)
# + [markdown] id="oce_KjnYvexv"
# **2.2** Crie uma função **fixDangling(M)** que retorna uma cópia modificada da matriz de adjacência **M** onde cada *dangling node* do grafo original possui arestas para todos os vértices do grafo. *Dica:* Você pode criar um vetor $d$ com os graus de saída e acessar as linhas de $M$ correpondentes aos *dangling nodes* por $M[d==0,:]$. Imprima uma nova matriz chamada **A_fixed** retornada após chamar *fixDangling* para **A_new**.
# + id="0JN8E2Flvexw"
#Insira seu código para a questão 2.2 aqui
def fixDangling(M):
d = np.sum(M, axis=1)
A_fixed = np.copy(M)
A_fixed[d==0] = 1
return A_fixed
A_fixed = fixDangling(A_new)
print(A_fixed)
# + [markdown] id="DLlA7cscvexz"
# **2.3** Crie uma função **matrizDeTransicao(M, alpha)** que receba como parâmetro também a probabilidade *alpha* de não fazermos um salto aleatório. Você pode assumir que **M** foi retornada por *fixDanglig*, logo, não possui *dangling nodes*. Imprima as matrizes:
# * *(2.3.1)* **P_2** obtida ao chamar *matrizDeTransicao* para os parâmetros **A** e **alpha**=$0.85$;
# * *(2.3.2)* **P_new** obtida ao chamar matrizDeTransicao para os parâmetros **A_fixed** e **alpha**=$0.85$.
# + id="F5wdtLIVvex0"
# Insira seu código para a questão 2.3 aqui
def matrizDeTransicao(M, alpha=1.0):
n = A.shape[0]
d = A.sum(axis=1)
P = (alpha*A)/d[:,None]+(1-alpha)*(np.ones((n,n))/n)
return P
P_2 = matrizDeTransicao(A, 0.85)
P_new = matrizDeTransicao(A_fixed, 0.85)
print(P_2)
print(P_new)
# + [markdown] id="fSXsxfT_vex5"
# **2.4** Armazene, respectivamente, o resultado do método da potência com:
# * *(2.4.1)* $P_2^\top$ e $\epsilon=10^{-5}$
# * *(2.4.2)* $P_\textrm{new}^\top$ e $\epsilon=10^{-5}$.
#
# nas variáveis **pm_eps_P2** e **pm_eps_Pnew**;
# + id="gH-t5-OUvex5"
#Insira seu código para a questão 2.4 aqui
pm_eps_P2, iters = powerMethodEps(P_2.T,epsilon=1e-5)
pm_eps_Pnew, iters = powerMethodEps(P_new.T,epsilon=1e-5)
print(pm_eps_P2)
print(pm_eps_Pnew)
# + [markdown] id="qMgo37N1vex8"
# **2.5** Sejam $i_\max$ e $i_\min$ os índices dos vértices com maior e menor PageRank de **A_fixed**. Vamos verificar como a adição de um novo link pode ajudar a promover uma página web (vértice). Adicione uma aresta do vértice $i_\max$ para o vértice $i_\min$ (se já houver aresta, aumente de 1 para 2 o elemento da matriz de adjacência). Salve o valor do novo pagerank na variável **new_pagerank**. Qual é o novo pagerank de $i_\min$?
# + id="9TLsIp_3vex8"
#Insira seu código para a questão 2.5 aqui
ind_sorted = np.argsort(np.squeeze(pm_eps_Pnew))
imax = ind_sorted[ 1]
imax = ind_sorted[-1]
imin = ind_sorted[0]
# imin = np.argmin(pm_eps_Pnew)
# imin = np.argmin(pm_eps_Pnew)
print('imin: {} - imax: {}'.format(imin, imax))
A_fixed2 = A_fixed.copy()
A_fixed2[imax,imin] += 1
print(A_fixed2, type(A_fixed2))
P_fixed2 = matrizDeTransicao(A_fixed2,0.85)
print(P_fixed2, type(P_fixed2))
new_pagerank, iters = powerMethodEps(P_fixed2.T,epsilon=1e-4)
print(new_pagerank)
|
src/eps/exercicios/EP2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Project 3: Implement a Planning Search
# ### SandBox
#
# <sub><NAME>. August 28, 2017<sub>
#
# #### Abstract
#
# _In this project, I will solve deterministic logistics planning problems for an Air Cargo transport system using a planning search agent. I will start by defining a group of air cargo domain problems in classical PDDL (Planning Domain Definition Language) fashion, and then I will set up the problems for search, experimenting with various automatically generated heuristics to solve the problems._
# -
# ## 1. Introduction
#
# In this section, I will give some background about the problem addressed and the goal of the project....
import sys
sys.path.append("../") # include the root directory as the main
import aind.eda as eda
import pandas as pd
import numpy as np
|
notebooks/sandbox.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ME7: Neural Networks
#
# We will have serveral exercises of neural networks using Sklearn neural network multi-layer perceptron (MLP) packages. Although keras with tensorflow framework is more commonly used in ML, this will be a good start to know how NN can be used for classification and regression problems. Keras/tensorflow will be explored in the next assignment.
#
# Please read the document on Scikit Learn:
#
# https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPClassifier.html
#
# https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPRegressor.html
# Write your name and your collaborators if any.
#
# -
# -
# ### Set up
# +
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import pandas as pd
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
from sklearn.model_selection import train_test_split
from matplotlib.colors import ListedColormap
from sklearn.datasets import make_classification, make_blobs
cmap_bold = ListedColormap(['#FFFF00', '#00FF00', '#0000FF','#000000'])
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "NN"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
# -
# ### Utility functions
# +
def plot_class_regions_for_classifier_subplot(clf, X, y, X_test, y_test, title, subplot, target_names = None, plot_decision_regions = True):
numClasses = np.amax(y) + 1
color_list_light = ['#FFFFAA', '#EFEFEF', '#AAFFAA', '#AAAAFF']
color_list_bold = ['#EEEE00', '#000000', '#00CC00', '#0000CC']
cmap_light = ListedColormap(color_list_light[0:numClasses])
cmap_bold = ListedColormap(color_list_bold[0:numClasses])
h = 0.03
k = 0.5
x_plot_adjust = 0.1
y_plot_adjust = 0.1
plot_symbol_size = 50
x_min = X[:, 0].min()
x_max = X[:, 0].max()
y_min = X[:, 1].min()
y_max = X[:, 1].max()
x2, y2 = np.meshgrid(np.arange(x_min-k, x_max+k, h), np.arange(y_min-k, y_max+k, h))
P = clf.predict(np.c_[x2.ravel(), y2.ravel()])
P = P.reshape(x2.shape)
if plot_decision_regions:
subplot.contourf(x2, y2, P, cmap=cmap_light, alpha = 0.8)
subplot.scatter(X[:, 0], X[:, 1], c=y, cmap=cmap_bold, s=plot_symbol_size, edgecolor = 'black')
subplot.set_xlim(x_min - x_plot_adjust, x_max + x_plot_adjust)
subplot.set_ylim(y_min - y_plot_adjust, y_max + y_plot_adjust)
if (X_test is not None):
subplot.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cmap_bold, s=plot_symbol_size, marker='^', edgecolor = 'black')
train_score = clf.score(X, y)
test_score = clf.score(X_test, y_test)
title = title + "\nTrain score = {:.2f}, Test score = {:.2f}".format(train_score, test_score)
subplot.set_title(title)
if (target_names is not None):
legend_handles = []
for i in range(0, len(target_names)):
patch = mpatches.Patch(color=color_list_bold[i], label=target_names[i])
legend_handles.append(patch)
subplot.legend(loc=0, handles=legend_handles)
def plot_class_regions_for_classifier(clf, X, y, X_test=None, y_test=None, title=None, target_names = None, plot_decision_regions = True):
numClasses = np.amax(y) + 1
color_list_light = ['#FFFFAA', '#EFEFEF', '#AAFFAA', '#AAAAFF']
color_list_bold = ['#EEEE00', '#000000', '#00CC00', '#0000CC']
cmap_light = ListedColormap(color_list_light[0:numClasses])
cmap_bold = ListedColormap(color_list_bold[0:numClasses])
h = 0.03
k = 0.5
x_plot_adjust = 0.1
y_plot_adjust = 0.1
plot_symbol_size = 50
x_min = X[:, 0].min()
x_max = X[:, 0].max()
y_min = X[:, 1].min()
y_max = X[:, 1].max()
x2, y2 = np.meshgrid(np.arange(x_min-k, x_max+k, h), np.arange(y_min-k, y_max+k, h))
P = clf.predict(np.c_[x2.ravel(), y2.ravel()])
P = P.reshape(x2.shape)
plt.figure()
if plot_decision_regions:
plt.contourf(x2, y2, P, cmap=cmap_light, alpha = 0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cmap_bold, s=plot_symbol_size, edgecolor = 'black')
plt.xlim(x_min - x_plot_adjust, x_max + x_plot_adjust)
plt.ylim(y_min - y_plot_adjust, y_max + y_plot_adjust)
if (X_test is not None):
plt.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cmap_bold, s=plot_symbol_size, marker='^', edgecolor = 'black')
train_score = clf.score(X, y)
test_score = clf.score(X_test, y_test)
title = title + "\nTrain score = {:.2f}, Test score = {:.2f}".format(train_score, test_score)
if (target_names is not None):
legend_handles = []
for i in range(0, len(target_names)):
patch = mpatches.Patch(color=color_list_bold[i], label=target_names[i])
legend_handles.append(patch)
plt.legend(loc=0, handles=legend_handles)
if (title is not None):
plt.title(title)
plt.show()
# -
# ### Activation functions
#
# - You should explore these activation functions to find the best function for your dataset(s).
# +
xrange = np.linspace(-2, 2, 200)
plt.figure(figsize=(7,6))
plt.plot(xrange, np.maximum(xrange, 0), label = 'relu')
plt.plot(xrange, np.tanh(xrange), label = 'tanh')
plt.plot(xrange, 1 / (1 + np.exp(-xrange)), label = 'logistic')
plt.legend()
plt.title('Neural network activation functions')
plt.xlabel('Input value (x)')
plt.ylabel('Activation function output')
plt.show()
# -
# ## Part 0
#
# - The examples in Part 0 build classifiers and regressors using neural networks.
#
# - We will use synthetic datasets to demonstrate the NN modeling process.
#
# - Classifier examples demonstrate the model performance through the class boundary with model score (overall accuracy).
#
#
# ### Neural networks on Classification
# - Read and run each cell of the given examples and understand the results
#
# - Tasks: you might have warnings related to data normalization or/and number of iterations.
# - <span style="color:red"> Fix the issues and remove warnings (if possible). </span>
#
#
# #### SkLearn Neural networks for classification
# - Please also read the document on Scikit Learn
# - https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPClassifier.html
#
# ### (1) Synthetic dataset1: Binary classification
#
# - A synthetic dataset contains two features (x1 and x2)
# - We use single hidden layer NN.
# +
# synthetic dataset for classification (binary)
plt.figure()
plt.title('Sample binary classification problem with two informative features')
X_C2, y_C2 = make_classification(n_samples = 100, n_features=2,
n_redundant=0, n_informative=2,
n_clusters_per_class=1, flip_y = 0.1,
class_sep = 0.5, random_state=0)
plt.scatter(X_C2[:, 0], X_C2[:, 1], marker= 'o', c=y_C2, s=50, cmap=cmap_bold)
plt.show()
# -
# #### NN modeling on synthetic dataset 1
#
# - We may need to increase the number of iterations or scale the data (normalization).
# +
from sklearn.neural_network import MLPClassifier
fig, subaxes = plt.subplots(3, 1, figsize=(6,18))
X_train, X_test, y_train, y_test = train_test_split(X_C2, y_C2, random_state=0)
for units, axis in zip([1, 10, 100], subaxes):
# create a model and training it
# we may need to increase the number of iterations or scale the data (normalization)
nnclf = MLPClassifier(hidden_layer_sizes = [units], solver='lbfgs',
random_state = 0).fit(X_train, y_train)
title = 'Dataset 1: Neural net classifier, 1 layer, {} units'.format(units)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
# -
# ### (2) Synthetic dataset 2: binary classification
#
# - More difficult synthetic dataset for classification (binary) with classes that are not linearly separable.
# - We apply single hidden layer NN.
# generate a synthetic dataset
X_D2, y_D2 = make_blobs(n_samples = 100, n_features = 2,
centers = 8, cluster_std = 1.3,
random_state = 4)
y_D2 = y_D2 % 2
plt.figure()
plt.title('Sample binary classification problem with non-linearly separable classes')
plt.scatter(X_D2[:,0], X_D2[:,1], c=y_D2,
marker= 'o', s=50, cmap=cmap_bold)
plt.show()
# #### NN modeling on synthetic dataset 2
#
# - We may need to increase the number of iterations or scale the data (normalization).
# +
from sklearn.neural_network import MLPClassifier
fig, subaxes = plt.subplots(3, 1, figsize=(6,18))
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
for units, axis in zip([1, 10, 100], subaxes):
# create a model and training it
# we may need to increase the number of iterations or scale the data (normalization)
nnclf = MLPClassifier(hidden_layer_sizes = [units], solver='lbfgs',
random_state = 0).fit(X_train, y_train)
title = 'Dataset 1: Neural net classifier, 1 layer, {} units'.format(units)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
# -
# #### Apply two hidden layer NN on synthetic dataset 2
# +
#from adspy_shared_utilities import plot_class_regions_for_classifier
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
# model training with two hidden layers
nnclf = MLPClassifier(hidden_layer_sizes = [10, 10], solver='lbfgs',
random_state = 0).fit(X_train, y_train)
plot_class_regions_for_classifier(nnclf, X_train, y_train, X_test, y_test,
'Dataset 1: Neural net classifier, 2 layers, 10/10 units')
# -
# #### NN on synthetic dataset 2: Regularization parameter: alpha
# +
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
fig, subaxes = plt.subplots(4, 1, figsize=(6, 23))
for this_alpha, axis in zip([0.01, 0.1, 1.0, 5.0], subaxes):
nnclf = MLPClassifier(solver='lbfgs', activation = 'tanh',
alpha = this_alpha,
hidden_layer_sizes = [100, 100],
random_state = 0).fit(X_train, y_train)
title = 'Dataset 2: NN classifier, alpha = {:.3f} '.format(this_alpha)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
# -
# #### NN on synthetic dataset2: the effect of different choices of activation function
# +
X_train, X_test, y_train, y_test = train_test_split(X_D2, y_D2, random_state=0)
fig, subaxes = plt.subplots(3, 1, figsize=(6,18))
for this_activation, axis in zip(['logistic', 'tanh', 'relu'], subaxes):
nnclf = MLPClassifier(max_iter = 500, solver='lbfgs', activation = this_activation,
alpha = 0.01, hidden_layer_sizes = [100, 100],
random_state = 0).fit(X_train, y_train)
title = 'Dataset 2: NN classifier, 2 layers 10/10, {} \
activation function'.format(this_activation)
plot_class_regions_for_classifier_subplot(nnclf, X_train, y_train,
X_test, y_test, title, axis)
plt.tight_layout()
# -
# ### Neural networks on Regression
#
# - NN can be also applied for regression problems.
#
# #### sklearn Neural Networks for regression
#
# https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPRegressor.html
# #### A synthetic dataset with one feature
#
# - A simple example that can be visualized the regression.
# +
# synthetic dataset for simple regression
from sklearn.datasets import make_regression
plt.figure()
plt.title('Sample regression problem with one input variable')
X_R1, y_R1 = make_regression(n_samples = 100, n_features=1,
n_informative=1, bias = 150.0,
noise = 30, random_state=0)
plt.scatter(X_R1, y_R1, marker= 'o', s=50)
plt.show()
# -
# ##### A NN regressor modeling
# +
from sklearn.neural_network import MLPRegressor
fig, subaxes = plt.subplots(2, 3, figsize=(11,8), dpi=70)
X_predict_input = np.linspace(-3, 3, 50).reshape(-1,1)
X_train, X_test, y_train, y_test = train_test_split(X_R1[0::5], y_R1[0::5], random_state = 0)
for thisaxisrow, thisactivation in zip(subaxes, ['tanh', 'relu', 'logistic']):
for thisalpha, thisaxis in zip([0.0001, 0.1, 1.0, 100], thisaxisrow):
# create a model
mlpreg = MLPRegressor(hidden_layer_sizes = [100,100],
activation = thisactivation,
alpha = thisalpha,
solver = 'lbfgs').fit(X_train, y_train)
y_predict_output = mlpreg.predict(X_predict_input)
thisaxis.set_xlim([-2.5, 0.75])
thisaxis.plot(X_predict_input, y_predict_output,
'^', markersize = 10)
thisaxis.plot(X_train, y_train, 'o')
thisaxis.set_xlabel('Input feature')
thisaxis.set_ylabel('Target value')
thisaxis.set_title('MLP regression\nalpha={}, activation={})'
.format(thisalpha, thisactivation))
plt.tight_layout()
# -
# ## Part 1
#
# ### NN Application to real-world dataset for classification
#
# - You will be working on two datasets: (1) Breast cancer dataset, and (2) fruit dataset
#
# - Before you start NN modeling, prepare the datasets (a) a dataset without normalization, (b) a dataset with normalization.
#
# - Please make sure that training data and test data must be the same scale. This means that the normalization process should be done before you split the data into training data and test data.
#
# - For each dataset, conduct classificaton modeling and evalute the model performance. We would suggest you use evaluation metrics for classification (accuracy, precision, recall, f1-score, etc.)
#
# - 1. Apply neural network with 2 hidden layers with varying number of units (e.g., 10, 20, 50, 100) for each layer.
# - You may want to use different number of units for the two hidden layers.
#
# - 2. Find out the optimal alpha parameter value for regularization.
# - alpha = [0.01, 0.05, 0.1, 0.5, 1.0, 5.0]
#
# - 3. Apply three different activation functions and show the effect.
# - activation = ['tanh', relu', 'logistic']
#
# - 4. (extra) Build a NN model with 3 hidden layers and check if the model is improvled.
#
# - Compare the results without normalization and with normalization.
#
# ### 1. Breast Cancer dataset
# +
# Breast cancer dataset for classification
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
(X_cancer, y_cancer) = load_breast_cancer(return_X_y = True)
print(X_cancer.shape) # 30 attributes
# -
# #### Add cells for your modeling below.
# #### Let's first choose the first two attributes and display the data
#
# - This is simply for visualization purpose. You can include most (or all) attributes for X.
# +
# You should choose some attributes for X
# You also should check X's shpae and y_cancer's shape
# We show how to do below as an example,
# but you should be able to do this process on your own.
X = X_cancer[:, :2]
print(X.shape)
print(y_cancer.shape)
# -
plt.figure()
plt.title('Sample binary classification problem with non-linearly separable classes')
plt.scatter(X[:,0], X[:,1], c=y_cancer,
marker= 'o', s=50, cmap=cmap_bold)
plt.show()
# #### Apply neural network with 2 hidden layers with varying number of units (e.g., 10, 20, 50, 100) on non-normalized data
#
# - We are using all 30 attribues of X
# - X is not normalized.
# #### Data normalization
#
# - We are using all 30 attribues of X
# - Please keep in mind that we do not normalize y values (it is class label!)
# #### Apply neural network with 2 hidden layers with varying number of units (e.g., 10, 20, 50, 100) on normalized data
#
# - We are using all 30 attribues of X
# - X is normalized.
# #### Find the optimal alpha parameter value for regularization
# #### Apply different activation functions (logistic, tanh, relu) and show the effect.
# ### (2) Fruit dataset
#
# - We show how to read txt format files using Pandas read_csv() function.
# - Make sure you check shape of X and shape of y
# +
# fruits dataset
fruits = pd.read_csv('./data/fruit_data_with_colors.txt', sep='\t', engine='python')
feature_names_fruits = ['height', 'width', 'mass', 'color_score']
X_fruits = fruits[feature_names_fruits]
y_fruits = fruits['fruit_label']
target_names_fruits = ['apple', 'mandarin', 'orange', 'lemon']
X_fruits_2d = fruits[['height', 'width']]
y_fruits_2d = fruits['fruit_label']
print(X_fruits_2d.shape)
print(y_fruits_2d.shape)
#print(X_fruits_2d)
# -
# #### Apply neural network with 2 hidden layers with varying number of units (e.g., 10, 20, 50, 100) on not normalized data
# #### Apply varying regularization parameter alpha on non-normalized data
# #### Apply three different activation functions on non-normalized data¶
# ### Normalize data
# #### Apply neural network with 2 hidden layers with varying number of units (e.g., 10, 20, 50, 100) on normalized data
# #### Find the optimal alpha parameter value for regularization
# #### Apply different activation functions (logistic, tanh, relu) and show the effect.
# ## Part 2
#
# - Write a short summary of your analysis result of neural networks (submitted on Canvas).
#
# - Provide a link to the notebook on Github.
|
ME7_NN/.ipynb_checkpoints/NN_basic-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small>
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
# </small>
#
#
#
# # Deep Learning From Basics to Practice
# ## by <NAME>, https://dlbasics.com, http://glassner.com
# ------
# ## Chapter 15: Scikit-Learn
# ### Notebook 8: Datasets
import numpy as np
import math
import matplotlib.pyplot as plt
from sklearn.datasets import make_moons
from sklearn.datasets import make_circles
from sklearn.datasets import make_blobs
from sklearn.model_selection import train_test_split
import seaborn as sns ; sns.set()
# +
# Make a File_Helper for saving and loading files.
save_files = True
import os, sys, inspect
current_dir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
sys.path.insert(0, os.path.dirname(current_dir)) # path to parent dir
from DLBasics_Utilities import File_Helper
file_helper = File_Helper(save_files)
# -
clr_list = ['#00CEAB', '#FFE53F', '#F26363']
# +
np.random.seed(42)
plt.figure(figsize=(8,12))
plt.subplot(3, 2, 1)
(moons_xy, moons_labels) = make_moons(n_samples=800)
clrs = [clr_list[v] for v in moons_labels]
plt.scatter(moons_xy[:,0], moons_xy[:,1], c=clrs, s=15)
plt.title('make_moons(), 0 noise')
plt.subplot(3, 2, 2)
(moons_xy, moons_labels) = make_moons(n_samples=800, noise=0.08)
clrs = [clr_list[v] for v in moons_labels]
plt.scatter(moons_xy[:,0], moons_xy[:,1], c=clrs, s=15)
plt.title('make_moons(), 0.08 noise')
plt.subplot(3, 2, 3)
(circle_xy, circle_labels) = make_circles(n_samples=200)
clrs = [clr_list[v] for v in circle_labels]
plt.scatter(circle_xy[:,0], circle_xy[:,1], c=clrs, s=15)
plt.title('make_circles(), 0 noise')
plt.subplot(3, 2, 4)
(circle_xy, circle_labels) = make_circles(n_samples=200, noise=.08)
clrs = [clr_list[v] for v in circle_labels]
plt.scatter(circle_xy[:,0], circle_xy[:,1], c=clrs, s=15)
plt.title('make_circles(), .08 noise')
plt.subplot(3, 2, 5)
(blob_xy, blob_labels) = make_blobs(n_samples=800, n_features=2,
centers=[[-10,-10], [-2.5, 2.5], [10,10]])
clrs = [clr_list[v] for v in blob_labels]
plt.scatter(blob_xy[:,0], blob_xy[:,1], c=clrs, s=25)
plt.title('make_blobs()')
plt.subplot(3, 2, 6)
(blob_xy, blob_labels) = make_blobs(n_samples=800, n_features=2,
centers=[[-10,-10], [-2.5, 2.5], [10,10]], cluster_std=[1,3,5])
clrs = [clr_list[v] for v in blob_labels]
plt.scatter(blob_xy[:,0], blob_xy[:,1], c=clrs, s=25)
plt.title('make_blobs(), std=(1,3,5)')
file_helper.save_figure('synthetic-datasets')
plt.show()
|
Chapter15-Scikit-Learn/Scikit-Learn-Notebook-8-Datasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Immutability
name = "Sam"
name[0] = 'P'
# +
# name[0] = 'P'
# -
name[1:]
last_letters = name[1:]
'P' + last_letters
x = 'Hello World'
x = x + " Python is great!"
x
letter = 'z'
letter * 10
2 + 3
'2' + '4'
x = 'Hello World'
x.upper()
x
x.upper
x.lower()
x.split()
x = 'His this is a string'
x.split()
x.split('i')
print('hello')
|
Section 3: Python Object and Data Structure Basics/String Properties and Methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
PIXIV_USERNAME = "userbay"
PIXIV_PASSWORD = "<PASSWORD>"
# +
from pixivpy3 import *
api = AppPixivAPI()
# api = ByPassSniApi() # bypass the GFW
# api.require_appapi_hosts()
api.set_accept_language('zh-cn') # tags翻译成中文
token = api.login(PIXIV_USERNAME, PIXIV_PASSWORD)
user_id = token.response.user.id
print(token.response.user)
# -
# # PixivCrawler (with pixivpy)
# +
import os
import json
import time
import random
import numpy as np
import pandas as pd
import sqlite3 as lite
from sqlalchemy import create_engine
try:
from tqdm.notebook import tqdm # new tqdm
except:
from tqdm import tqdm_notebook as tqdm
class PixivCrawler(object):
def __init__(self, api, illust_db='pixiv_illusts.db'):
self.api = api
self.illust_db = illust_db
self.user_info = None
def randSleep(self, base=0.1, rand=0.5):
"休眠随机的时间"
time.sleep(base + rand*random.random())
def GetUserDetail(self, user_id):
"查询指定用户的基本信息"
self.last_user = self.api.user_detail(user_id)
return self.last_user
def GetUserBookmarks(self, user_id, restrict='public'):
"获取指定用户的收藏列表"
df_list = []
next_qs = {'user_id': user_id, 'restrict': restrict}
user = self.GetUserDetail(user_id)
self.randSleep(0.1)
with tqdm(total=user.profile.total_illust_bookmarks_public,
desc="api.user_bookmarks_illust") as pbar:
while next_qs != None:
json_result = self.api.user_bookmarks_illust(**next_qs)
tmp_df = pd.DataFrame.from_dict(json_result.illusts)
df_list.append(tmp_df)
pbar.update(tmp_df.shape[0])
next_qs = self.api.parse_qs(json_result.next_url)
self.randSleep(0.1)
df = pd.concat(df_list).rename(columns={'id': 'illust_id'})
df['user_id'] = df.user.apply(lambda d: d['id'])
return df.set_index('illust_id')
def GetUserIllusts(self, user_id, type='illust'):
"获取指定用户的作品列表(illusts/manga)"
df_list = []
next_qs = {'user_id': user_id, 'type': type, 'filter': 'for_ios'}
user = self.GetUserDetail(user_id)
if type == 'illust':
total = user.profile.total_illusts
elif type == 'manga':
total = user.profile.total_manga
else:
raise Exception("Unsupported type=%d" % type)
self.randSleep(0.1)
with tqdm(total=total, desc="api.user_illusts") as pbar:
while next_qs != None:
json_result = self.api.user_illusts(**next_qs)
tmp_df = pd.DataFrame.from_dict(json_result.illusts)
df_list.append(tmp_df)
pbar.update(tmp_df.shape[0])
next_qs = self.api.parse_qs(json_result.next_url)
self.randSleep(0.1)
df = pd.concat(df_list).rename(columns={'id': 'illust_id'})
df['user_id'] = df.user.apply(lambda d: d['id'])
return df.set_index('illust_id')
def GetIllustRanking(self, mode, date, total=100):
"获取作品排行榜"
df_list = []
next_qs = {'mode': mode, 'date': date, 'filter': 'for_ios'}
with tqdm(total=total, desc="api.illust_ranking") as pbar:
while next_qs != None:
json_result = self.api.illust_ranking(**next_qs)
tmp_df = pd.DataFrame.from_dict(json_result.illusts)
df_list.append(tmp_df)
pbar.update(tmp_df.shape[0])
next_qs = self.api.parse_qs(json_result.next_url)
self.randSleep(0.3)
df = pd.concat(df_list).rename(columns={'id': 'illust_id'})
df['user_id'] = df.user.apply(lambda d: d['id'])
return df.set_index('illust_id')
def GetFollowingUsers(self, user_id, restrict='public'):
"获取指定用户跟踪的用户列表,返回user_ids"
user_ids = []
next_qs = {'user_id': user_id, 'restrict': restrict}
user = self.GetUserDetail(user_id)
with tqdm(total=user.profile.total_follow_users,
desc="api.user_following") as pbar:
while next_qs != None:
json_result = self.api.user_following(**next_qs)
for one_user in json_result.user_previews:
user_ids.append(one_user.user.id)
pbar.update(len(json_result.user_previews))
next_qs = self.api.parse_qs(json_result.next_url)
self.randSleep(0.3, 0.8)
return np.array(user_ids)
def UpdateIllusts(self, df_illusts):
sql_df = df_illusts.copy()
# 数组类字段转json
sql_df['image_urls'] = sql_df.image_urls.apply(json.dumps)
sql_df['meta_pages'] = sql_df.meta_pages.apply(json.dumps)
sql_df['meta_single_page'] = sql_df.meta_single_page.apply(json.dumps)
sql_df['series'] = sql_df.series.apply(json.dumps)
sql_df['tags'] = sql_df.tags.apply(json.dumps)
sql_df['tools'] = sql_df.tools.apply(json.dumps)
sql_df['user'] = sql_df.user.apply(json.dumps)
# 先读取文件里的illusts存储,并用新的数据代替key相同的内容
if os.path.isfile(self.illust_db):
# 读取文件的数据并丢弃同样的illust_id (保留新的illust_id)
db_df = self.DBIllusts(ensure_json=False)
db_df = db_df[~db_df.index.isin(sql_df.index)]
merged_df = pd.concat([sql_df, db_df], sort=False)
else:
merged_df = sql_df
# 合并后df写入文件(replace方式)
engine = create_engine('sqlite:///' + self.illust_db, echo=False)
merged_df.to_sql('illusts', con=engine, if_exists='replace')
return merged_df
def DBIllusts(self, sql="SELECT * FROM illusts WHERE illust_id > 0", ensure_json=True):
with lite.connect(self.illust_db) as conn:
sql_df = pd.read_sql_query(sql, conn, index_col='illust_id')
# 还原json字段
if ensure_json:
sql_df['image_urls'] = sql_df.image_urls.apply(json.loads)
sql_df['meta_pages'] = sql_df.meta_pages.apply(json.loads)
sql_df['meta_single_page'] = sql_df.meta_single_page.apply(
json.loads)
sql_df['series'] = sql_df.series.apply(json.loads)
sql_df['tags'] = sql_df.tags.apply(json.loads)
sql_df['tools'] = sql_df.tools.apply(json.loads)
sql_df['user'] = sql_df.user.apply(json.loads)
return sql_df
crawl = PixivCrawler(api)
# -
# ## GetUserBookmarks(public)
df_bookmarks = crawl.GetUserBookmarks(user_id)
_ = crawl.UpdateIllusts(df_bookmarks)
# ## GetFollowingUsers(public)
user_ids = crawl.GetFollowingUsers(user_id)
random.shuffle(user_ids)
for uid in tqdm(user_ids, desc="GetFollowingUsers"):
df = crawl.GetUserIllusts(uid)
_ = crawl.UpdateIllusts(df)
crawl.randSleep(1.1, 5.0)
# ## GetIllustRanking
# mode: [day, week, month, day_male, day_female, week_original, week_rookie, day_manga]
# date: '2016-08-01'
# mode (Past): [day, week, month, day_male, day_female, week_original, week_rookie,
# day_r18, day_male_r18, day_female_r18, week_r18, week_r18g]
df_ranking = crawl.GetIllustRanking('week', '2019-11-01')
_ = crawl.UpdateIllusts(df_ranking)
|
notebooks/pixivCrawler.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="zXWF5wD-8PIs"
from sklearn import datasets
cancer = datasets.load_breast_cancer()
# + colab={"base_uri": "https://localhost:8080/"} id="QtNpIDHySsDc" outputId="7ea436aa-cf44-4618-b529-fc75b308ff24"
print("Features: ", cancer.feature_names)
# + colab={"base_uri": "https://localhost:8080/"} id="Y6fVKuwZSyrI" outputId="6acd7792-e73a-494c-f78a-e74c6da48661"
print("Labels: ", cancer.target_names)
# + colab={"base_uri": "https://localhost:8080/"} id="6gq6nH86S2l7" outputId="756bf6d7-20d0-4ee4-e63b-3b98752b1eff"
cancer.data.shape
# + colab={"base_uri": "https://localhost:8080/"} id="OKa1Pom1S8Kk" outputId="d6303a88-2524-49bf-aedb-68fff2a213d6"
print(cancer.data[0:5])
# + colab={"base_uri": "https://localhost:8080/"} id="vUD2iD5tTRjq" outputId="c3caaabe-3a20-4686-b1fa-35552a183d21"
# print the cancer labels (0:malignant, 1:benign)
print(cancer.target)
# + id="zSx9FNbjTZqf"
# Import train_test_split function
from sklearn.model_selection import train_test_split
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(cancer.data, cancer.target, test_size=0.3,random_state=109) # 70% training and 30% test
# + id="GNy2XlEcTlNA"
#Import svm model
from sklearn import svm
#Create a svm Classifier
clf = svm.SVC(kernel='linear') # Linear Kernel
#Train the model using the training sets
clf.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = clf.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="G0F0LLNbT2Km" outputId="ee044423-109a-453e-f592-914694060932"
import matplotlib.pyplot as plt
plt.plot(X_train,y_train,"b.")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="Q3Q_PeWaUCqd" outputId="54517789-e38b-4da2-b579-394b6512ef71"
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
# Model Accuracy: how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="fjRoEeOTUsra" outputId="9e9793d6-d8e3-4c0e-de3d-5745722d6df5"
# Model Precision: what percentage of positive tuples are labeled as such?
print("Precision:",metrics.precision_score(y_test, y_pred))
# Model Recall: what percentage of positive tuples are labelled as such?
print("Recall:",metrics.recall_score(y_test, y_pred))
|
src/third_month/SupportVectorMachine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import random
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"] = "1"
os.environ["TF_FORCE_GPU_ALLOW_GROWTH"] = "true"
# +
import numpy as np
import pandas as pd
import tensorflow as tf
from evaluator import evaluate
from data_loader import load_kdd_cup_urc, load_yahoo_A1, load_yahoo_A2, load_yahoo_A3, load_yahoo_A4, load_power_demand # Univariate Datasets
from data_loader import load_nasa, load_ecg, load_gesture, load_smd # Multivariate Datasets
from tensorflow import keras
from tensorflow.keras import layers
from tqdm.notebook import tqdm
# THESE LINES ARE FOR REPRODUCIBILITY
random.seed(0)
np.random.seed(0)
tf.random.set_seed(0)
# -
class Sampling(layers.Layer):
"""Uses (z_mean, z_log_var) to sample z, the vector encoding a digit."""
def call(self, inputs):
z_mean, z_log_var = inputs
batch = tf.shape(z_mean)[0]
dim = tf.shape(z_mean)[1]
epsilon = tf.keras.backend.random_normal(shape=(batch, dim))
return z_mean + tf.exp(0.5 * z_log_var) * epsilon
class VAE(keras.Model):
def __init__(self, encoder, decoder, **kwargs):
super(VAE, self).__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
self.total_loss_tracker = keras.metrics.Mean(name="total_loss")
self.reconstruction_loss_tracker = keras.metrics.Mean(
name="reconstruction_loss"
)
self.kl_loss_tracker = keras.metrics.Mean(name="kl_loss")
@property
def metrics(self):
return [
self.total_loss_tracker,
self.reconstruction_loss_tracker,
self.kl_loss_tracker,
]
def train_step(self, data):
with tf.GradientTape() as tape:
z_mean, z_log_var, z = self.encoder(data)
reconstruction = self.decoder(z)
reconstruction_loss = tf.reduce_mean(
tf.reduce_sum(
keras.losses.binary_crossentropy(data, reconstruction), axis=1
)
)
kl_loss = -0.5 * (1 + z_log_var - tf.square(z_mean) - tf.exp(z_log_var))
kl_loss = tf.reduce_mean(tf.reduce_sum(kl_loss, axis=1))
total_loss = reconstruction_loss + kl_loss
grads = tape.gradient(total_loss, self.trainable_weights)
self.optimizer.apply_gradients(zip(grads, self.trainable_weights))
self.total_loss_tracker.update_state(total_loss)
self.reconstruction_loss_tracker.update_state(reconstruction_loss)
self.kl_loss_tracker.update_state(kl_loss)
return {
"loss": self.total_loss_tracker.result(),
"reconstruction_loss": self.reconstruction_loss_tracker.result(),
"kl_loss": self.kl_loss_tracker.result(),
}
def CNN_VAE(X_train):
latent_dim = 16
encoder_inputs = keras.Input(shape=(X_train.shape[1], X_train.shape[2]))
x = layers.Conv1D(32, 3, activation="relu", strides=2, padding="same")(encoder_inputs)
x = layers.Conv1D(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Flatten()(x)
x = layers.Dense(128, activation="relu")(x)
z_mean = layers.Dense(latent_dim, name="z_mean")(x)
z_log_var = layers.Dense(latent_dim, name="z_log_var")(x)
z = Sampling()([z_mean, z_log_var])
encoder = keras.Model(encoder_inputs, [z_mean, z_log_var, z], name="encoder")
latent_inputs = keras.Input(shape=(latent_dim,))
x = layers.Dense(32* 64, activation="relu")(latent_inputs)
x = layers.Reshape((32, 64))(x)
x = layers.Conv1DTranspose(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Conv1DTranspose(32, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Flatten()(x)
x = layers.Dense(X_train.shape[1] * X_train.shape[2])(x)
decoder_outputs = layers.Reshape([X_train.shape[1], X_train.shape[2]])(x)
decoder = keras.Model(latent_inputs, decoder_outputs, name="decoder")
model = VAE(encoder, decoder)
model.compile(optimizer=keras.optimizers.Adam(learning_rate=0.001))
history = model.fit(X_train, epochs=50, batch_size=128, verbose=0)
return model
# ### Yahoo S5
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
# + tags=[]
for loader in [load_yahoo_A1, load_yahoo_A2, load_yahoo_A3, load_yahoo_A4]:
datasets = loader(64, 1)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(loader.__name__.replace('load_', ''))
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(loader.__name__.replace('load_', ''), np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
# -
yahoo_results = pd.DataFrame(total_scores)
yahoo_results.groupby('dataset').mean()
# ### NASA
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
for loader in [load_nasa]:
datasets = loader(100, 100)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(f'D{i+1}')
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(f'D{i+1}', np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
nasa_results = pd.DataFrame(total_scores)
nasa_results.groupby('dataset').mean()
# ### SMD
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
for loader in [load_smd]:
datasets = loader(64, 1)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(loader.__name__.replace('load_', ''))
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(loader.__name__.replace('load_', ''), np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
smd_results = pd.DataFrame(total_scores)
smd_results.groupby('dataset').mean()
# ### ECG
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
for loader in [load_ecg]:
datasets = loader(32, 16)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(f'D{i+1}')
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(f'D{i+1}', np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
ecg_results = pd.DataFrame(total_scores)
ecg_results.groupby('dataset').mean()
# ### Power Demand
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
for loader in [load_power_demand]:
datasets = loader(64, 1)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(loader.__name__.replace('load_', ''))
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(loader.__name__.replace('load_', ''), np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
power_results = pd.DataFrame(total_scores)
power_results.groupby('dataset').mean()
# ### 2D Gesture
total_scores = {'dataset': [], 'f1': [], 'pr_auc': [], 'roc_auc': []}
for loader in [load_gesture]:
datasets = loader(64, 1)
x_trains, x_tests, y_tests = datasets['x_train'], datasets['x_test'], datasets['y_test']
for i in tqdm(range(len(x_trains))):
tf.keras.backend.clear_session()
X_train = x_trains[i]
X_test = x_tests[i]
model = CNN_VAE(X_train)
X_test_rec = model.decoder.predict(model.encoder.predict(X_test)[-1])
scores = evaluate(X_test, X_test_rec, y_tests[i], is_reconstructed=True)
total_scores['dataset'].append(loader.__name__.replace('load_', ''))
total_scores['f1'].append(np.max(scores['f1']))
total_scores['pr_auc'].append(scores['pr_auc'])
total_scores['roc_auc'].append(scores['roc_auc'])
print(loader.__name__.replace('load_', ''), np.max(scores['f1']), scores['pr_auc'], scores['roc_auc'])
gesture_results = pd.DataFrame(total_scores)
gesture_results.groupby('dataset').mean()
|
Baseline - CNN-VAE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
bitsl = [3.63, 7.12, 10.61, 13.91, 16.69, 19.97, 25.55, 29.13,
32.07, 34.08, 36.65, 42.95, 43.38, 49.12, 52.22,
53.47, 59.06, 60.12, 63.26]
norm = [11.75, 23.51, 36.51, 47.09, 58.85, 72.29, 82.45, 94.07,
105.82, 117.59, 131.64, 141.17, 154.37, 166.89,
177.64, 188.30, 201.75, 213.63, 224.61]
iters = [i*100 for i in range(1,20)]
df = pd.DataFrame([{"iters": k, "norm": i, "bitsl":j} for i,j,k in zip(bitsl, norm, iters)])
# +
X1 = df.iloc[:, 0].values.reshape(-1, 1)
Y1 = df.iloc[:, 1].values.reshape(-1, 1)
linear_regressor1 = LinearRegression()
linear_regressor1.fit(X1, Y1)
Y_pred1 = linear_regressor1.predict(X1)
X2 = df.iloc[:, 0].values.reshape(-1, 1)
Y2 = df.iloc[:, 2].values.reshape(-1, 1)
linear_regressor2 = LinearRegression()
linear_regressor2.fit(X2, Y2)
Y_pred2 = linear_regressor2.predict(X2)
plt.figure(figsize=(20,10))
ax = plt.axes()
ax.set_facecolor('#e5ecf6')
plt.grid(True, linewidth=1.0, color='white', linestyle='-', alpha=0.7)
plt.scatter(iters, bitsl, color='blue', alpha=0.8, label='with bitslicing')
plt.scatter(iters, norm, color='green', alpha=0.8, label='without bitslicing')
plt.plot(X1, Y_pred1, color='blue', alpha=0.2)
plt.plot(X1, Y_pred2, color='green', alpha=0.2)
plt.legend(loc="upper right")
plt.title("With bitslicing vs without")
plt.xlabel("Number of iterations")
plt.ylabel("Time used in millisecond")
plt.show()
# +
# smaller image
X1 = df.iloc[:, 0].values.reshape(-1, 1)
Y1 = df.iloc[:, 1].values.reshape(-1, 1)
linear_regressor1 = LinearRegression()
linear_regressor1.fit(X1, Y1)
Y_pred1 = linear_regressor1.predict(X1)
X2 = df.iloc[:, 0].values.reshape(-1, 1)
Y2 = df.iloc[:, 2].values.reshape(-1, 1)
linear_regressor2 = LinearRegression()
linear_regressor2.fit(X2, Y2)
Y_pred2 = linear_regressor2.predict(X2)
plt.figure(figsize=(13,8))
ax = plt.axes()
ax.set_facecolor('#e5ecf6')
plt.grid(True, linewidth=1.0, color='white', linestyle='-', alpha=0.7)
plt.scatter(iters, bitsl, color='blue', alpha=0.8, label='with bitslicing')
plt.scatter(iters, norm, color='green', alpha=0.8, label='without bitslicing')
plt.plot(X1, Y_pred1, color='blue', alpha=0.2)
plt.plot(X1, Y_pred2, color='green', alpha=0.2)
plt.legend(loc="upper right")
plt.title("With bitslicing vs without")
plt.xlabel("Number of iterations")
plt.ylabel("Time used in millisecond")
plt.show()
# -
#get stats
df.describe()
# time per iteration
df['pern'] = df['norm'] / df['iters']
df['perb'] = df['bitsl'] / df['iters']
df.describe()
df
|
implementation/timings/graphs.ipynb
|