code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### tuple and default operations
# +
tuples1 = (1,8,33,5,3,15,9,4,10)
tuples2 = (7,2,4,2,1,6,4,7,9)
print(tuples)
## count function
print("Count of the elements in tuple : ",tuples.count(5))
## index function
print('Index of the element : ', tuples.index(15))
## length function
print('length of the tuple : ',len(tuples))
## minimum function
print('minimum of the tuple : ',min(tuples))
## Concatenation function
tuple3 = tuples1 + tuples2
print("Concatenation tuple : ",tuple3)
# -
|
tuple_and_operations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from shapely import geometry
from pulp import *
import pandas as pd
import geopandas as gpd
import numpy as np
from geopy.distance import geodesic
import matplotlib.pyplot as plt
from matplotlib.collections import LineCollection
import json
from area import area
import pickle
from vincenty import *
from shapely.strtree import STRtree
import networkx as nx
# -
ne = gpd.read_file('./../../../data/ne_10m_countries.gpkg')
ne.plot()
mp = ne.unary_union
len(mp)
import pickle
pickle.dump(mp,open('./../../../data/world_mp.pkl','wb'))
pulp.pulpTestAll()
dist = np.arange(0,10,0.1)
mw = np.array([ii/10 for ii in range(10,0,-1)] + [ii/10 for ii in range(11)])
mw
dist2d = np.stack([dist]*mw.shape[0])
mw2d = np.stack([mw]*dist.shape[0])
Z = dist2d**2
mw2d
alpha=0.5
# case: B=1
E_mw = mw2d
# ## Toy Model
pts_A = np.random.rand(10*2).reshape(10,2)*10
pts_B = np.random.rand(3*2).reshape(3,2)*10
pts_A = [geometry.Point(pt[0],pt[1]) for pt in pts_A.tolist()]
pts_B = [geometry.Point(pt[0],pt[1]) for pt in pts_B.tolist()]
for ii_A,pt in enumerate(pts_A):
pt.MW = np.random.rand(1)*5
pt.name='A_'+str(ii_A)
for ii_B,pt in enumerate(pts_B):
pt.MW = np.random.rand(1)*40
pt.name='B_'+str(ii_B)
def milp_geodesic_network_satisficing(pts_A, pts_B, alpha,v=False):
pts_A_dict = {pt.name:pt for pt in pts_A}
pts_B_dict = {pt.name:pt for pt in pts_B}
A_names = [pt.name for pt in pts_A]
B_names = [pt.name for pt in pts_B]
Z = {pt.name:{} for pt in pts_A}
MW_A = {pt.name:pt.MW for pt in pts_A}
MW_B = {pt.name:pt.MW for pt in pts_B}
if v:
print ('generating Z..')
for pt_A in pts_A:
for pt_B in pts_B:
Z[pt_A.name][pt_B.name]=(geodesic([pt_A.y,pt_A.x], [pt_B.y,pt_B.x]).kilometers)**2
sum_Z = sum([Z[A_name][B_name] for A_name in A_names for B_name in B_names])
### declare model
model = LpProblem("Network Satisficing Problem",LpMinimize)
### Declare variables
# B -> Bipartite Network
B = LpVariable.dicts("Bipartite",(A_names,B_names),0,1,LpInteger)
# abs_diffs -> absolute value forcing variable
abs_diffs = LpVariable.dicts("abs_diffs",B_names,cat='Continuous')
### Declare constraints
# Contstraint - abs diffs edges
for B_name in B_names:
model += abs_diffs[B_name] >= (MW_B[B_name] - lpSum([MW_A[A_name]*B[A_name][B_name] for A_name in A_names]))/MW_B[B_name],"abs forcing pos {}".format(B_name)
model += abs_diffs[B_name] >= -1 * (MW_B[B_name] - lpSum([MW_A[A_name]*B[A_name][B_name] for A_name in A_names]))/MW_B[B_name], "abs forcing neg {}".format(B_name)
# Constraint - bipartite edges
for A_name in A_names:
model += lpSum([B[A_name][B_name] for B_name in B_names]) <= 1,"Bipartite Edges {}".format(A_name)
### Affine equations
# Impedence error
E_z = sum([Z[A_name][B_name]*B[A_name][B_name] for A_name in A_names for B_name in B_names])/sum_Z
# mw error
E_mw = sum([abs_diffs[B_name] for B_name in B_names])/len(B_names)
### Objective function
model += E_z*alpha + (1-alpha)*E_mw, "Loss"
if v:
print ('solving model...')
model.solve(pulp.GUROBI_CMD())
if v:
print(pulp.LpStatus[model.status])
return model, B, E_z, E_mw, Z
model, B, E_z, E_mw, Z = milp_geodesic_network_satisficing(pts_A, pts_B, 0.5, v=True)
pts_A_dict = {pt.name:pt for pt in pts_A}
pts_B_dict = {pt.name:pt for pt in pts_B}
E_z.value()
for e in E_mw:
print (e.name, e.value())
for kk, vv in B.items():
for kk2, vv2 in vv.items():
print (kk,kk2,vv2.value())
# +
### visualise
fig, axs = plt.subplots(1,1,figsize=(12,12))
for pt in pts_A:
axs.scatter(pt.x, pt.y, s=pt.MW*10,c='g')
for pt in pts_B:
axs.scatter(pt.x,pt.y,s=pt.MW*10,c='r')
lines = []
for k1, v in B.items():
for k2,v2 in v.items():
if v2.varValue>0:
lines.append([[pts_A_dict[k1].x,pts_A_dict[k1].y],[pts_B_dict[k2].x,pts_B_dict[k2].y]])
lc = LineCollection(lines, color='grey')
axs.add_collection(lc)
plt.show()
# -
# ## To data
# ### wrangle data
fts = json.load(open('./results_fcs_v1.3/ABCD_simplified.geojson','r'))
GB = json.load(open('GB_v1.3.geojson','r'))
wri = pd.read_csv('global_power_plant_database.csv')
ws = pd.read_csv('wri_mixin.csv')
GB_pts = []
for ii_f,ft in enumerate(GB['features']):
poly = geometry.shape(ft['geometry'])
pp = poly.representative_point()
pp.MW = area(geometry.mapping(poly))*54/1000/1000
if ft['properties']['SPOT_ids_0']=='':
pp.name=str(ii_f)+'_'+str(ft['properties']['S2_ids_0'])
else:
pp.name=str(ii_f)+'_'+str(ft['properties']['SPOT_ids_0'])
GB_pts.append(pp)
wri.fuel1.unique()
wri_records = wri[(wri.country=='GBR')&(wri.fuel1=='Solar')][['capacity_mw', 'name','latitude','longitude']].to_dict('records')
for ii_r,r in enumerate(wri_records):
r['MW'] = r['capacity_mw']
r['pt'] = geometry.Point(r['longitude'],r['latitude'])
r['name'] = str(ii_r)+'_'+str(r['name'])
ws_records = ws[ws.iso2=='GB'][['WS Coords','MW','WS Name']].to_dict('records')
for r in ws_records:
r['WS Coords'] = r['WS Coords'].replace(' E','')
r['pt'] = geometry.Point(float(r['WS Coords'].split(',')[-1]),float(r['WS Coords'].split(',')[0]))
r['name'] = r['WS Name']
records = wri_records + ws_records
country_shps = json.load(open('/home/lucaskruitwagen/DPHIL/DPHIL_CLASSIFICATION/Solar_PV/NE/ne_10m_admin_0_countries.geojson','r'))
GB_shp = [ft for ft in country_shps['features'] if ft['properties']['ISO_A2']=='GB']
R_pts = []
for r in records:
pp = r['pt']
pp.MW =r['MW']
pp.name=r['name']
R_pts.append(pp)
# ### Get components with threshold
def get_matches_treenx(A_polys,P_polys):
### return a list of matches. both polys need a property which is primary_id
P_tree = STRtree(P_polys)
G = nx.Graph()
for pp_A in A_polys:
t_result = P_tree.query(pp_A)
t_result = [pp for pp in t_result if pp.intersects(pp_A)]
if len(t_result)>0:
for r in t_result:
G.add_edge(pp_A.name,r.name)
return list(nx.connected_components(G))
buffer_dist = 1500 #m
# +
GB_polys = []
for pt in GB_pts:
pp = pt.buffer(V_dir([pt.y,pt.x],buffer_dist,0)[0][0]-pt.y)
pp.name = pt.name
GB_polys.append(pp)
R_polys = []
for pt in R_pts:
pp = pt.buffer(V_dir([pt.y,pt.x],buffer_dist,0)[0][0]-pt.y)
pp.name = pt.name
R_polys.append(pp)
# -
components = get_matches_treenx(GB_polys,R_polys)
len(components)
# +
### visualise components
fig, axs = plt.subplots(1,1,figsize=(16,16))
for shp_ft in GB_shp[1:2]:
geom = geometry.shape(shp_ft['geometry'])
if geom.type=='Polygon':
xs,ys = geom.exterior.xy
axs.plot(xs,ys,c='grey')
if geom.type=='MultiPolygon':
for subgeom in geom:
xs,ys = subgeom.exterior.xy
axs.plot(xs,ys,c='grey')
for c in components:
pts = [pt for pt in GB_pts if pt.name in c]
pts += [pt for pt in R_pts if pt.name in c]
rnd_col = np.random.rand(3,)
axs.scatter([pt.x for pt in pts], [pt.y for pt in pts],c=[rnd_col]*len(pts))
plt.show()
# -
alpha = 0.15
B_dict = {}
# +
E_z_all = 0
E_mw_all = 0
for ii_c, component in enumerate(components):
if ii_c % 100==0:
print (ii_c)
pts_A = [pt for pt in GB_pts if pt.name in component]
pts_B = [pt for pt in R_pts if pt.name in component]
#print (len(pts_A),len(pts_B))
model, B, E_z, E_mw, Z = milp_geodesic_network_satisficing(pts_A, pts_B, alpha)
E_z_all += E_z.value()
E_mw_all += E_mw.value()
B_dict[ii_c] = B
#print (Z)
#for k1, v in B.items():
# for k2,v2 in v.items():
# if v2.varValue>0:
# print (k1,k2,v2,11)
print ('E_z', E_z_all, 'E_mw',E_mw_all)
# +
### visualise results
fig, axs = plt.subplots(1,1,figsize=(72,72))
for shp_ft in GB_shp[1:2]:
geom = geometry.shape(shp_ft['geometry'])
if geom.type=='Polygon':
xs,ys = geom.exterior.xy
axs.plot(xs,ys,c='grey')
if geom.type=='MultiPolygon':
for subgeom in geom:
xs,ys = subgeom.exterior.xy
axs.plot(xs,ys,c='grey')
### cyan and purple - unmatched outside of components
for ii_c, component in enumerate(components):
pts_A = [pt for pt in GB_pts if pt.name in component]
pts_B = [pt for pt in R_pts if pt.name in component]
pts_A_dict = {pt.name:pt for pt in pts_A}
pts_B_dict = {pt.name:pt for pt in pts_B}
lines = []
pts_recs = []
pts_obs = []
done_As = []
done_Bs = []
#print (B_dict[ii_c])
for k1, v in B_dict[ii_c].items():
#k1 - cluster
for k2,v2 in v.items():
#k2 -
#print (k1,k2,v2, v2.varValue)
if v2.varValue>0:
#matches observations
pts_obs.append([pts_A_dict[k1].x,pts_A_dict[k1].y,pts_A_dict[k1].MW])
done_As.append(k1)
#mathes dataset
pts_recs.append([pts_B_dict[k2].x,pts_B_dict[k2].y,pts_B_dict[k2].MW])
done_Bs.append(k2)
#match lines
lines.append([[pts_A_dict[k1].x,pts_A_dict[k1].y],[pts_B_dict[k2].x,pts_B_dict[k2].y]])
#print (ii_c,len(component),len(pts_A),len(pts_B),len(lines))
lc = LineCollection(lines, color='grey')
### pink and green for matched
axs.scatter([pt[0] for pt in pts_obs],[pt[1] for pt in pts_obs],s=[pt[2] for pt in pts_obs],c='green')
axs.scatter([pt[0] for pt in pts_recs],[pt[1] for pt in pts_recs],s=[pt[2] for pt in pts_recs],c='pink')
for name,pt in pts_A_dict.items():
if name not in done_As:
axs.scatter(pt.x,pt.y,s=pt.MW,c='b')
for name,pt in pts_B_dict.items():
if name not in done_Bs:
axs.scatter(pt.x,pt.y,s=pt.MW,c='r')
axs.add_collection(lc)
axs.set_xlim([-6,-4])
axs.set_ylim([49,51])
plt.show()
# -
# ### get impedence network
pts_R_dict = {pt.name:pt for pt in R_pts}
pts_GB_dict = {pt.name:pt for pt in GB_pts}
R_names = [pt.name for pt in R_pts]
GB_names = [pt.name for pt in GB_pts]
Z = {pt.name:{} for pt in GB_pts}
MW_GB = {pt.name:pt.MW for pt in GB_pts}
MW_R = {pt.name:pt.MW for pt in R_pts}
len(GB_pts), len(R_pts)
for ii_A,pt_A in enumerate(GB_pts):
if ii_A % 100 ==0:
print ('ii_A',ii_A)
for pt_B in R_pts:
#print (pt_A.name, pt_B.name)
Z[pt_A.name][pt_B.name]=geodesic([pt_A.x,pt_A.y], [pt_B.x,pt_B.y]).kilometers
pickle.dump(Z,open('Z.pickle','wb'))
Z= pickle.load(open('Z.pickle','rb'))
# ### Set up problem
# #### Define problem
model = LpProblem("Network Satisficing Problem",LpMinimize)
# #### Define Variables
# B -> bipartite
B = LpVariable.dicts("Bipartite",(GB_names,R_names),0,1,LpInteger)
# Differences between MW and MW in (needs variable to ensure absolute value)
abs_diffs = LpVariable.dicts("abs_diffs",R_names,cat='Continuous')
# #### Define Constraints
# Contstraint - abs diffs edges
for R_name in R_names:
model += abs_diffs[R_name] >= (MW_R[R_name] - lpSum([MW_GB[GB_name]*B[GB_name][R_name] for GB_name in GB_names]))/MW_R[R_name],"abs forcing pos {}".format(R_name)
model += abs_diffs[R_name] >= -1 * (MW_R[R_name] - lpSum([MW_GB[GB_name]*B[GB_name][R_name] for GB_name in GB_names]))/MW_R[R_name], "abs forcing neg {}".format(R_name)
# Contstraint - bipartite edges
for GB_name in GB_names:
model += lpSum([B[GB_name][R_name] for R_name in R_names]) <= 1,"Bipartite Edges {}".format(GB_name)
# #### Define constants and affine equations
alpha = 0.5
sum_Z = sum([Z[GB_name][R_name] for GB_name in GB_names for R_name in R_names])
E_z = sum([Z[GB_name][R_name]*B[GB_name][R_name] for GB_name in GB_names for R_name in R_names])/sum_Z
E_mw = sum([abs_diffs[R_name] for R_name in R_names])/len(R_names)
# #### Defin Objective Function
# Objective function
model += E_z*alpha + (1-alpha)*E_mw, "Loss"
model.solve()
pulp.LpStatus[model.status]
E_z.value()
for e in E_mw:
print (e.name, e.value())
# +
### visualise
fig, axs = plt.subplots(1,1,figsize=(12,12))
for pt in pts_A:
axs.scatter(pt.x, pt.y, s=pt.MW*10,c='g')
for pt in pts_B:
axs.scatter(pt.x,pt.y,s=pt.MW*10,c='r')
lines = []
for k1, v in B.items():
for k2,v2 in v.items():
if v2.varValue>0:
lines.append([[pts_A_dict[k1].x,pts_A_dict[k1].y],[pts_B_dict[k2].x,pts_B_dict[k2].y]])
lc = LineCollection(lines, color='grey')
axs.add_collection(lc)
plt.show()
# -
# ## Demo
model = LpProblem("Profit maximising problem", pulp.LpMaximize)
A = LpVariable('A', lowBound=0, cat='Integer')
B = LpVariable('B', lowBound=0, cat='Integer')
profit = 3000 * A + 45000 * B
# +
# Objective function
model += profit, "Profit"
# Constraints
model += 3 * A + 4 * B <= 30
model += 5 * A + 6 * B <= 60
model += 1.5 * A + 3 * B <= 21
# -
model.solve()
pulp.LpStatus[model.status]
print ("Production of Car A = {}".format(A.varValue))
print ("Production of Car B = {}".format(B.varValue))
# ## Archive
E_z_1 = LpVariable.dicts('E_z_1', (A_names,B_names),lowBound=0, cat='Continuous')
E_z_0 = LpVariable.dicts('E_z_0',A_names, lowBound=0, cat='Continuous')
E_z_t = LpVariable('E_z_t',lowBound = 0, cat='Continuous')
for A_name in A_names:
for B_name in B_names:
model+= E_z_1 == bipartite[A_name][B_name] * Z[A_name][B_name]
for A_name in A_names:
model += E_z_0[A_name] == lpSum([E_z_1[A_name][B_name] for B_name in B_names])
|
solarpv/analysis/matching/MILP_WRI-matching_stripped.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Here I have some useful python codes.
# Import library
from __future__ import division, print_function, absolute_import
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
print('Import library')
# # How to sort a Python dict by value
# +
# How to sort a Python dict by value
# (== get a representation sorted by value)
xs = {'a': 4, 'b': 3, 'c': 2, 'd': 1}
sorted(xs.items(), key=lambda x: x[1])
# -
# # Left pad with zero
# Left pad with zero
n = '4'
n.zfill(3)
# # Subplots example
# +
import numpy as np
import matplotlib.pyplot as plt
# Data for plotting
t = np.arange(0.01, 20.0, 0.01)
# Create figure
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2, 2)
# log y axis
ax1.semilogy(t, np.exp(-t / 5.0))
ax1.set(title='semilogy')
ax1.grid()
# log x axis
ax2.semilogx(t, np.sin(2 * np.pi * t))
ax2.set(title='semilogx')
ax2.grid()
# log x and y axis
ax3.loglog(t, 20 * np.exp(-t / 10.0), basex=2)
ax3.set(title='loglog base 2 on x')
ax3.grid()
# With errorbars: clip non-positive values
# Use new data for plotting
x = 10.0**np.linspace(0.0, 2.0, 20)
y = x**2.0
ax4.set_xscale("log", nonposx='clip')
ax4.set_yscale("log", nonposy='clip')
ax4.set(title='Errorbars go negative')
ax4.errorbar(x, y, xerr=0.1 * x, yerr=5.0 + 0.75 * y)
# ylim must be set after errorbar to allow errorbar to autoscale limits
ax4.set_ylim(bottom=0.1)
fig.tight_layout()
plt.show()
# -
# # Namedtuple
# +
# Using namedtuple is way shorter than defining a class manually.
# Namedtuple is immutable
from collections import namedtuple
Car = namedtuple('Car', 'company name color mileage data')
my_car = Car('Toyota', 'Sienna LE', 'gray', [3812.4], [i for i in range(5)])
print(my_car)
my_car.mileage[0] = 1000
my_car.data.sort(reverse=True)
print(my_car)
# +
import collections
# collections.deque?
# -
|
notebooks/.ipynb_checkpoints/Useful_codes-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../../Pierian_Data_Logo.png' /></a>
# ___
# # Matplotlib Overview Lecture
# ## Introduction
# Matplotlib is the "grandfather" library of data visualization with Python. It was created by <NAME>. He created it to try to replicate MatLab's (another programming language) plotting capabilities in Python. So if you happen to be familiar with matlab, matplotlib will feel natural to you.
#
# It is an excellent 2D and 3D graphics library for generating scientific figures.
#
# Some of the major Pros of Matplotlib are:
#
# * Generally easy to get started for simple plots
# * Support for custom labels and texts
# * Great control of every element in a figure
# * High-quality output in many formats
# * Very customizable in general
#
# Matplotlib allows you to create reproducible figures programmatically. Let's learn how to use it! Before continuing this lecture, I encourage you just to explore the official Matplotlib web page: http://matplotlib.org/
#
# ## Installation
#
# You'll need to install matplotlib first with either:
#
# conda install matplotlib
# or
# pip install matplotlib
#
# ## Importing
# Import the `matplotlib.pyplot` module under the name `plt` (the tidy way):
import matplotlib.pyplot as plt
# You'll also need to use this line to see plots in the notebook:
# %matplotlib inline
# That line is only for jupyter notebooks, if you are using another editor, you'll use: **plt.show()** at the end of all your plotting commands to have the figure pop up in another window.
# # Basic Example
#
# Let's walk through a very simple example using two numpy arrays:
# ### Example
#
# Let's walk through a very simple example using two numpy arrays. You can also use lists, but most likely you'll be passing numpy arrays or pandas columns (which essentially also behave like arrays).
#
# ** The data we want to plot:**
import numpy as np
x = np.linspace(0, 5, 11)
y = x ** 2
x
y
# ## Functinal method per usare matplotlib
# Lo vediamo brevemente perché è meglio utilizzare l'altro modo (object oriented method)
#
#
# ### Basic Matplotlib Commands
#
# We can create a very simple line plot using the following ( I encourage you to pause and use Shift+Tab along the way to check out the document strings for the functions we are using).
plt.plot(x,y, 'r*')
plt.plot(x, y, 'r') # 'r' is the color red
plt.xlabel('X Axis Title Here')
plt.ylabel('Y Axis Title Here')
plt.title('String Title Here')
plt.show()
# ## Creating Multiplots on Same Canvas
# plt.subplot(nrows, ncols, plot_number)
plt.subplot(1,2,1)
plt.plot(x, y, 'r--') # More on color options later
plt.subplot(1,2,2)
plt.plot(y, x, 'g*-');
# ___
# # Matplotlib Object Oriented Method
# Now that we've seen the basics, let's break it all down with a more formal introduction of Matplotlib's Object Oriented API. This means we will instantiate figure objects and then call methods or attributes from that object.
# ## Introduction to the Object Oriented Method
# The main idea in using the more formal Object Oriented method is to create figure objects and then just call methods or attributes off of that object. This approach is nicer when dealing with a canvas that has multiple plots on it.
#
# To begin we create a figure instance. Then we can add axes to that figure:
fig = plt.figure()
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8])
# +
# Create Figure (empty canvas)
fig = plt.figure() # un canvas vuoto che può contenere cose
# Add set of axes to figure
# % da left, % da bottom, width, height (range 0 to 1)
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8])
# Plot on that set of axes
axes.plot(x, y, 'b')
axes.set_xlabel('Set X Label') # Notice the use of set_ to begin methods
axes.set_ylabel('Set y Label')
axes.set_title('Set Title')
# -
# Code is a little more complicated, but the advantage is that we now have full control of where the plot axes are placed, and we can easily add more than one axis to the figure:
# +
# Creates blank canvas
fig = plt.figure()
axes1 = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # main axes
axes2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
# Larger Figure Axes 1
axes1.plot(x, y, 'b')
axes1.set_xlabel('X_label_axes2')
axes1.set_ylabel('Y_label_axes2')
axes1.set_title('Axes 2 Title')
# Insert Figure Axes 2
axes2.plot(y, x, 'r')
axes2.set_xlabel('X_label_axes2')
axes2.set_ylabel('Y_label_axes2')
axes2.set_title('Axes 2 Title');
# -
# ## subplots()
#
# The plt.subplots() object will act as a more automatic axis manager.
#
# Basic use cases:
# +
# Use similar to plt.figure() except use tuple unpacking to grab fig and axes
fig, axes = plt.subplots()
# Now use the axes object to add stuff to plot
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
# -
# Then you can specify the number of rows and columns when creating the subplots() object:
# Empty canvas of 1 by 2 subplots
fig, axes = plt.subplots(nrows=2, ncols=3)
plt.tight_layout() # per eliminare l'overlapping tra i plot
# Empty canvas of 1 by 2 subplots
fig, axes = plt.subplots(nrows=1, ncols=2)
# Axes is an array of axes to plot on
axes
# We can iterate through this array:
# +
for ax in axes:
ax.plot(x, y, 'b')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
# Display the figure object
fig
# -
fig, axes = plt.subplots(nrows=1, ncols=2)
axes[0].plot(x,y)
axes[1].plot(y,x)
# A common issue with matplolib is overlapping subplots or figures. We ca use **fig.tight_layout()** or **plt.tight_layout()** method, which automatically adjusts the positions of the axes on the figure canvas so that there is no overlapping content:
# +
fig, axes = plt.subplots(nrows=1, ncols=2)
for ax in axes:
ax.plot(x, y, 'g')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
fig
plt.tight_layout()
# -
# ### Figure size, aspect ratio and DPI
# Matplotlib allows the aspect ratio, DPI and figure size to be specified when the Figure object is created. You can use the `figsize` and `dpi` keyword arguments.
# * `figsize` is a tuple of the width and height of the figure in inches
# * `dpi` is the dots-per-inch (pixel per inch).
#
# For example:
fig = plt.figure(figsize=(8,4), dpi=100)
# The same arguments can also be passed to layout managers, such as the `subplots` function:
# +
fig, axes = plt.subplots(figsize=(12,3))
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
# -
# ## Saving figures
# Matplotlib can generate high-quality output in a number formats, including PNG, JPG, EPS, SVG, PGF and PDF.
# To save a figure to a file we can use the `savefig` method in the `Figure` class:
fig.savefig("filename.png")
# Here we can also optionally specify the DPI and choose between different output formats:
fig.savefig("filename.png", dpi=200)
# ____
# ## Legends, labels and titles
# Now that we have covered the basics of how to create a figure canvas and add axes instances to the canvas, let's look at how decorate a figure with titles, axis labels, and legends.
# **Figure titles**
#
# A title can be added to each axis instance in a figure. To set the title, use the `set_title` method in the axes instance:
ax.set_title("title");
# **Axis labels**
#
# Similarly, with the methods `set_xlabel` and `set_ylabel`, we can set the labels of the X and Y axes:
ax.set_xlabel("x")
ax.set_ylabel("y");
# ### Legends
# You can use the **label="label text"** keyword argument when plots or other objects are added to the figure, and then using the **legend** method without arguments to add the legend to the figure:
# +
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.plot(x, x**2, label="x**2")
ax.plot(x, x**3, label="x**3")
ax.legend()
# -
# Notice how are legend overlaps some of the actual plot!
#
# The **legend** function takes an optional keyword argument **loc** that can be used to specify where in the figure the legend is to be drawn. The allowed values of **loc** are numerical codes for the various places the legend can be drawn. See the [documentation page](http://matplotlib.org/users/legend_guide.html#legend-location) for details. Some of the most common **loc** values are:
# +
# Lots of options....
ax.legend(loc=1) # upper right corner
ax.legend(loc=2) # upper left corner
ax.legend(loc=3) # lower left corner
ax.legend(loc=4) # lower right corner
# .. many more options are available
# Most common to choose
ax.legend(loc=0) # let matplotlib decide the optimal location
fig
# -
# ## Setting colors, linewidths, linetypes
#
# Matplotlib gives you *a lot* of options for customizing colors, linewidths, and linetypes.
#
# There is the basic MATLAB like syntax (which I would suggest you avoid using for more clairty sake:
# ### Colors with MatLab like syntax
# With matplotlib, we can define the colors of lines and other graphical elements in a number of ways. First of all, we can use the MATLAB-like syntax where `'b'` means blue, `'g'` means green, etc. The MATLAB API for selecting line styles are also supported: where, for example, 'b.-' means a blue line with dots:
# MATLAB style line color and style
fig, ax = plt.subplots()
ax.plot(x, x**2, 'b.-') # blue line with dots
ax.plot(x, x**3, 'g--') # green dashed line
# ### Colors with the color= parameter
# We can also define colors by their names or RGB hex codes and optionally provide an alpha value using the `color` and `alpha` keyword arguments. Alpha indicates opacity.
# +
fig, ax = plt.subplots()
ax.plot(x, x+1, color="blue", alpha=0.5) # half-transparant
ax.plot(x, x+2, color="#8B008B") # RGB hex code
ax.plot(x, x+3, color="#FF8C00") # RGB hex code
# -
# ### Line and marker styles
# To change the line width, we can use the `linewidth` or `lw` keyword argument. The line style can be selected using the `linestyle` or `ls` keyword arguments:
# +
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(x, x+1, color="red", linewidth=0.25)
ax.plot(x, x+2, color="red", linewidth=0.50)
ax.plot(x, x+3, color="red", linewidth=1.00)
ax.plot(x, x+4, color="red", linewidth=2.00)
# possible linestype options ‘-‘, ‘–’, ‘-.’, ‘:’, ‘steps’
ax.plot(x, x+5, color="green", lw=3, linestyle='-')
ax.plot(x, x+6, color="green", lw=3, ls='-.')
ax.plot(x, x+7, color="green", lw=3, ls=':')
# custom dash
line, = ax.plot(x, x+8, color="black", lw=1.50)
line.set_dashes([5, 10, 15, 10]) # format: line length, space length, ...
# possible marker symbols: marker = '+', 'o', '*', 's', ',', '.', '1', '2', '3', '4', ...
ax.plot(x, x+9, color="blue", lw=3, ls='-', marker='+')
ax.plot(x, x+10, color="blue", lw=3, ls='--', marker='o')
ax.plot(x, x+11, color="blue", lw=3, ls='-', marker='s')
ax.plot(x, x+12, color="blue", lw=3, ls='--', marker='1')
# marker size and color
ax.plot(x, x+13, color="purple", lw=1, ls='-', marker='o', markersize=2)
ax.plot(x, x+14, color="purple", lw=1, ls='-', marker='o', markersize=4)
ax.plot(x, x+15, color="purple", lw=1, ls='-', marker='o', markersize=8, markerfacecolor="red")
ax.plot(x, x+16, color="purple", lw=1, ls='-', marker='s', markersize=8,
markerfacecolor="yellow", markeredgewidth=3, markeredgecolor="green");
# -
# ### Control over axis appearance
# In this section we will look at controlling axis sizing properties in a matplotlib figure.
# ## Plot range
# We can configure the ranges of the axes using the `set_ylim` and `set_xlim` methods in the axis object, or `axis('tight')` for automatically getting "tightly fitted" axes ranges:
# +
fig, axes = plt.subplots(1, 3, figsize=(12, 4))
axes[0].plot(x, x**2, x, x**3)
axes[0].set_title("default axes ranges")
axes[1].plot(x, x**2, x, x**3)
axes[1].axis('tight')
axes[1].set_title("tight axes")
axes[2].plot(x, x**2, x, x**3)
axes[2].set_ylim([0, 60])
axes[2].set_xlim([2, 5])
axes[2].set_title("custom axes range");
# -
# # Special Plot Types
#
# There are many specialized plots we can create, such as barplots, histograms, scatter plots, and much more. Most of these type of plots we will actually create using pandas. But here are a few examples of these type of plots:
plt.scatter(x,y)
from random import sample
data = sample(range(1, 1000), 100)
plt.hist(data)
# +
data = [np.random.normal(0, std, 100) for std in range(1, 4)]
# rectangular box plot
plt.boxplot(data,vert=True,patch_artist=True);
# -
# ## Further reading
# * http://www.matplotlib.org - The project web page for matplotlib.
# * https://github.com/matplotlib/matplotlib - The source code for matplotlib.
# * http://matplotlib.org/gallery.html - A large gallery showcaseing various types of plots matplotlib can create. Highly recommended!
# * http://www.loria.fr/~rougier/teaching/matplotlib - A good matplotlib tutorial.
# * http://scipy-lectures.github.io/matplotlib/matplotlib.html - Another good matplotlib reference.
#
|
04-Visualization-Matplotlib-Pandas/04-01-Matplotlib/Matplotlib Concepts Lecture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mapuertag/Datascience300/blob/main/Clase1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LPPQqGrfqWq4"
# **Introducción**
#
#
# ---
# Para manejar un código en Python, vamos a seguir las siguientes recomendaciones:
# 1. Usar siempre minúsculas
# 2. Se le puede añadir números
# 3. Aparecerá error si no estan seguidos de la declaración de la variable con el número asignado: es decir luisa1(correcto) luisa 1 (incorrecto)
#
# En python hay varios tipos de datos compuestos y estan disponibles por defecto en los interpretes:
#
# 1. Númericos
# 2. Secuencias
# 3. Mapeos
# 4. Conjuntos usados para agrupar otros valores.
#
# **Diferencia entre constantes y variables**: En mátematicas llamamos constante a una magnitud que no cambia con el paso del tiempo. Ejemplo: Contabilidad: Gastos Fijos_ Muebles de oficina Ejemplo 2: 6
#
# Por otro lado, el concepto de variable, como la cantidad que es susceptible a tomar distintos valores númericos. Ejemplo:
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="wTuP4jG8V_fQ" outputId="89bd80d0-7f56-4bbe-b710-70781df7214b"
x=80
print(x)
# + [markdown] id="ENLbdOjfWaMG"
# **Tipos de datos**:
#
# **Enteros**:
#
# Son los números que no tienes decimales Pueden ser positivos y negativos
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="SD3x7e9hWuor" outputId="38576b5f-431b-42df-cd31-8233fcda520f"
a=int(3.964)
print(a,type(a))
# + [markdown] id="OhgM65sGXFw0"
# El comando int es para hacer alusión a número enteros. A parte el comando float, permite valores con decimales.
# + colab={"base_uri": "https://localhost:8080/"} id="kZPLw-dmXKev" outputId="17c5f7ee-da4a-41ff-da5a-5f6928cb4ed6"
b=float(8.23)
print(b,type (b))
# + [markdown] id="1MGsfJ7SXfen"
# **Tipo cadena**:
# Las cadenas son texto encerrado entre comillas (Simples o dobles) y se pueden conformar de diferentes caracteres (Númericos, Albéticos, Especiales #$+*%). Tener en cuenta, las cadenas admiten operadores como la suma y la resta. (Variable String)
#
# + colab={"base_uri": "https://localhost:8080/"} id="LFatv-voX1C8" outputId="1f5a4e70-9cbf-42ee-a707-31d89d246489"
val2="<NAME>"
print(val2)
val1="Luisa"
print(val1)
# + colab={"base_uri": "https://localhost:8080/"} id="P6BpQGFyYETX" outputId="ce1df153-8115-4c5d-f254-f4e6ba851052"
n="Aprender"
z="Python"
w="Aprender Python"
nz=n+" "+z
print(w)
q="5"
p="2"
b=int(q)
c=int(p)
print(b-c)
# + [markdown] id="OPp2YJPIYLYO"
# **Tipos Booleanos**: Este tipo de variale solo tendrá un valor de Verdadero o Falso. Nota: Son valores muy usados en condiciones y bucles.
# + colab={"base_uri": "https://localhost:8080/"} id="ishYgxs9YWXb" outputId="36bf60b1-1116-4a02-fbe6-8ca367b125c8"
lola=True
print("El valor es verdadero:",lola,", el cual es de tipo", type(lola))
# + id="DxHCyf_pYeCe"
Python cuenta con tipos de datos que admiten colecciones:
1. Listas
2. Tuplas
3. Diccionarios
# + [markdown] id="_0MSXw7j483j"
# **Tipos de conjuntos**: Son una colección de datos sin elementos que repiten
# + colab={"base_uri": "https://localhost:8080/"} id="yc86cO6Y5MQO" outputId="db1fd2ed-efcd-4590-8d6b-3b2ee009b611"
fru="pera","manzana","naranja"
color= "morado","blanco","azul"
print(fru,color)
# + [markdown] id="8rxr4rmT6G8i"
# **Tipo de listas**
# Las almacenan vectores, siempre empiezan a nombrarse desde el elemento cero(0). Apertir de allí empieza el conteo. Se usan corchetes
# Ejemplo de listas de Python:
#
# + colab={"base_uri": "https://localhost:8080/"} id="20Q6z4647AqB" outputId="e751c1be-770e-417f-b730-8a331006aed4"
ines=["5","uva","lila","perro","celular","micro","botella","tetero","10"]
print(Ines)
fe=ines[5:7]
print(fe)
# + [markdown] id="iVg3iNiG8snj"
# **Tipo Tuplas**
# Es una lista que no se puede modificar después de la creación de esta: Tuplas anidadas. Agrupación (Tuplas). Generalmente se usa parentesis
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="U-SvV4iy9Sm-" outputId="605064a0-1760-4004-827f-cda2125ade07"
tupla=23,28,"hello"
print(tupla)
otra=tupla,(1,2,3,4)
print(otra)
# + [markdown] id="5VMqkWDg9ojH"
# **Tipo Diccionarios**
# Define los datos uno a uno entre un campo (ID-Identificador-Clave) y un valor
#
# + id="ywee2Pm298F3" colab={"base_uri": "https://localhost:8080/"} outputId="942b1226-7def-451c-8a69-041421d5a556"
datos_b={
"nombres":"Diana",
"apellidos":"Perea",
"cedula":"2345671",
"est_civil":"Viuda",
"lugar_nacimiento":"Neiva",
"fecha_nacimiento":"24/12/1980",
}
print("ID del diccionario", datos_b.keys())
print("ID del diccionario", datos_b.values())
print("ID del diccionario", datos_b.items())
print("Fecha de Nacimiento de Diana:", datos_b['fecha_nacimiento'])
# + [markdown] id="fOh1cDjBZYf8"
# **Ejemplo práctico**:
# + id="iM6zPXIhCBAx" colab={"base_uri": "https://localhost:8080/"} outputId="4c13f8ee-bd4f-449a-8eee-46f18223fa52"
zapatos_0={}
zapatos_0["talla"]="grande"
zapatos_0["color"]="rojo"
zapatos_0["material"]="cuero"
zapatos_0["tacón"]="alto"
zapatos_0["precio"]=90
zapatos_1={}
zapatos_1["talla"]="medio"
zapatos_1["color"]="negro"
zapatos_1["material"]="sintetico"
zapatos_1["tacón"]="plataforma"
zapatos_1["precio"]=60
zapatos_2={}
zapatos_2["talla"]="pequeño"
zapatos_2["color"]="blanco"
zapatos_2["material"]="tela"
zapatos_2["tacón"]="plano"
zapatos_2["precio"]=30
print(zapatos_0)
print(zapatos_1)
print(zapatos_2)
compra0=zapatos_0["precio"]
compra1=zapatos_1["precio"]
compra2=zapatos_2["precio"]
compratotal=compra0+compra1+compra2
print("la compra total fue de:",compratotal)
print("la compra total fue de:"+" "+str(compratotal))
|
Clase1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pybot
import string
import time
import numpy as np
import bot_utils
from collections import defaultdict
from IPython.display import SVG
# Keras imports
import tensorflow as tf
import tensorflow.keras.preprocessing.text as tftext
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers import Dense, Embedding, LSTM, Input, Bidirectional, Concatenate
from tensorflow.keras.utils import model_to_dot
from attention import AttentionLayer
# Gensim and Spelling
import gensim.downloader as api
glove_vectors = api.load("glove-wiki-gigaword-100")
from spellchecker import SpellChecker
import concurrent.futures
# Telegram
import logging
from threading import Timer
from telegram import ReplyKeyboardMarkup, ReplyKeyboardRemove, Update, ChatAction
from telegram.ext import (
Updater,
CommandHandler,
MessageHandler,
Filters,
ConversationHandler,
CallbackContext,
)
HYPER_PARAMS = {
'ENABLE_GPU' : True,
'ENABLE_SPELLING_SUGGESTION' : False,
'MESSAGE_TYPING_DELAY' : 4.0, # Seconds to mimic bot typing
'DATA_BATCH_SIZE' : 128,
'DATA_BUFFER_SIZE' : 10000,
'EMBEDDING_DIMENSION' : 100, # Dimension of the GloVe embedding vector
'MAX_SENT_LENGTH' : 11, # Maximum length of sentences
'MAX_SAMPLES' : 2000000, # Maximum samples to consider (useful for laptop memory) 200000
'MIN_WORD_OCCURENCE' : 30, # Minimum word count. If condition not met word replaced with <UNK>
'MODEL_LAYER_DIMS' : 500,
'MODEL_LEARN_RATE' : 1e-3,
'MODEL_LEARN_EPOCHS' : 10,
'MODEL_TRAINING' : True # If False - Model weights are loaded from file
}
if HYPER_PARAMS['ENABLE_GPU']:
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], enable=True)
# ### Importing the data set
# https://www.cs.cornell.edu/~cristian/Cornell_Movie-Dialogs_Corpus.html
#
# movie_lines:
# - lineID
# - characterID (who uttered this phrase)
# - movieID
# - character name
# - text of the utterance
#
# movie_conversations:
# - characterID of the first character involved in the conversation
# - characterID of the second character involved in the conversation
# - movieID of the movie in which the conversation occurred
# - list of the utterances that make the conversation, in chronological
# order: ['lineID1','lineID2',É,'lineIDN']
# has to be matched with movie_lines.txt to reconstruct the actual content
#
lines = open('data/movie_lines.txt', encoding = 'utf-8', errors = 'ignore').read().split('\n')
conversations = open('data/movie_conversations.txt', encoding = 'utf-8', errors = 'ignore').read().split('\n')
# ### Pre-processing the data
# - Perform cleaning
# - Extract Q&As
# - Tokenize
# - Add padding
# +
def clean_text_to_lower(text):
"""Performs cleaning and lowers text"""
text = text.lower()
# Expand contractions to words, and remove characters
for word in text.split():
if word in bot_utils.CONTRACTIONS:
text = text.replace(word, bot_utils.CONTRACTIONS[word])
# Remove punctuation
text = text.translate(str.maketrans('', '', string.punctuation))
return text
def get_questions_answers(lines, conversations):
"""Builds question and answers from data sets"""
# Map id to text of the utterance
id_to_line = {}
for line in lines:
line_ = line.split(' +++$+++ ')
if len(line_) == 5:
id_to_line[line_[0]] = line_[4]
questions, answers = [], []
for line in conversations:
# Extract conversation turns
conversation = line.split(' +++$+++ ')
if len(conversation) == 4:
conversation_turns = conversation[3][1:-1].split(', ')
turns_list = [turn[1:-1] for turn in conversation_turns]
# Split to Q & A
for i_turn in range(len(conversation_turns) - 1):
questions.append(clean_text_to_lower(id_to_line[conversation_turns[i_turn][1:-1]]))
answers.append(clean_text_to_lower(id_to_line[conversation_turns[i_turn + 1][1:-1]]))
if len(questions) >= HYPER_PARAMS['MAX_SAMPLES']:
return questions, answers
return questions, answers
def tokenize(lines, conversations):
"""Tokenizes sets to sequences of integers, and adds special tokens.
Also reduces the vocabulary by replacing low frequency words with an unknown token.
Returns:
tokenizer: tokenizer, which might be used later to reverse integers to words,
tokenized_questions: list, questions as sequence of integers
tokenized_answers: list, answer as sequence of integers
size_vocab: int, unique number of words in vocabulary.
special_tokens: dict, mappings for special tokens"""
questions, answers = get_questions_answers(lines, conversations)
tokenizer = tftext.Tokenizer(oov_token='<UNK>')
tokenizer.fit_on_texts(questions + answers)
# Make UNK tokens, reindex tokenizer dicts
sorted_by_word_count = sorted(tokenizer.word_counts.items(), key=lambda kv: kv[1], reverse=True)
tokenizer.word_index = {}
tokenizer.index_word = {}
index = 1
for word, count in sorted_by_word_count:
if count >= HYPER_PARAMS['MIN_WORD_OCCURENCE']:
tokenizer.word_index[word] = index
tokenizer.index_word[index] = word
index += 1
# Add special tokens
special_tokens = {}
special_tokens['<PAD>'] = 0
special_tokens['<UNK>'] = len(tokenizer.word_index)
special_tokens['<SOS>'] = special_tokens['<UNK>'] + 1
special_tokens['<EOS>'] = special_tokens['<SOS>'] + 1
for special_token, index_value in special_tokens.items():
tokenizer.word_index[special_token] = index_value
tokenizer.index_word[index_value] = special_token
# Tokenize to integer sequences
tokenized_questions = tokenizer.texts_to_sequences(questions)
tokenized_answers = tokenizer.texts_to_sequences(answers)
# Size is equal to the last token's index
size_vocab = special_tokens['<EOS>'] + 1
# Add sentence position tokens
tokenized_questions = [[special_tokens['<SOS>']] + tokenized_question + [special_tokens['<EOS>']]
for tokenized_question in tokenized_questions]
tokenized_answers = [[special_tokens['<SOS>']] + tokenized_answer + [special_tokens['<EOS>']]
for tokenized_answer in tokenized_answers]
# Add padding at end so we can use a static input size to the model
tokenized_questions = pad_sequences(tokenized_questions,
maxlen=HYPER_PARAMS['MAX_SENT_LENGTH'],
padding='post')
tokenized_answers = pad_sequences(tokenized_answers,
maxlen=HYPER_PARAMS['MAX_SENT_LENGTH'],
padding='post')
return tokenizer, tokenized_questions, tokenized_answers, size_vocab, special_tokens
# -
tokenizer, tokenized_questions, tokenized_answers,\
size_vocab, special_tokens = tokenize(lines, conversations)
size_vocab
# **Create tf.data.Dataset** <br>
# Allows caching and prefetching to speed up training.
# +
dataset = tf.data.Dataset.from_tensor_slices((
{
'encoder_inputs': tokenized_questions[:, 1:], # Skip <SOS> token
'decoder_inputs': tokenized_answers[:, :-1] # Skip <EOS> token
},
{
'outputs': tokenized_answers[:, 1:] # Skip <SOS> token
},
))
dataset = dataset.cache()
dataset = dataset.shuffle(HYPER_PARAMS['DATA_BUFFER_SIZE'])
dataset = dataset.batch(HYPER_PARAMS['DATA_BATCH_SIZE'])
dataset = dataset.prefetch(tf.data.experimental.AUTOTUNE)
# -
# ### Create contextual embedding from GloVe
# Create embedding layer with out context, from a pre-trained Word2Vec Model from Glove:<br>
# https://nlp.stanford.edu/projects/glove/
# +
def load_glove_weights(dimension_embedding):
""" Load GloVe pre-trained model"""
path='glove/glove.6B.' + str(dimension_embedding) + 'd.txt'
word_to_vec = {}
with open(path, encoding='utf-8') as file:
for line in file:
values = line.split()
word_to_vec[values[0]] = np.asarray(values[1:], dtype='float32')
file.close()
return word_to_vec
def load_embedding_weights(word_dictionary, size_vocab, dimension_embedding):
"""Loads embedding weights from GloVe based on our context"""
word_to_vec = load_glove_weights(dimension_embedding)
embedding_matrix = np.zeros((size_vocab,
HYPER_PARAMS['EMBEDDING_DIMENSION']))
for word, index in word_dictionary.items():
embedding_vector = word_to_vec.get(word)
# Word is within GloVe dictionary
if embedding_vector is not None:
embedding_matrix[index] = embedding_vector
return embedding_matrix, word_to_vec
def build_embedding_layer(word_dictionary, size_vocab, dimension_embedding, length_input):
"""Builds a non-trainable embedding layer from GloVe
pre-trained model, based on our context"""
embedding_matrix, word_to_vec = load_embedding_weights(word_dictionary, size_vocab, dimension_embedding)
size_vocab = size_vocab
embedding_layer = Embedding(size_vocab,
dimension_embedding,
input_length=length_input,
weights=[embedding_matrix],
trainable=False)
return embedding_matrix, word_to_vec, embedding_layer
# -
embedding_matrix, word_to_vec, embedding_layer = build_embedding_layer(tokenizer.word_index,
size_vocab,
HYPER_PARAMS['EMBEDDING_DIMENSION'],
HYPER_PARAMS['MAX_SENT_LENGTH'])
# ### Seq2Seq Model
def seq2seq(embedding_layer, length_input, size_vocab, layer_dims):
"""LSTM Seq2Seq model with attention"""
length_input = length_input - 1
#
# Encoder,
# Biderectional (RNNSearch) as explained in https://arxiv.org/pdf/1409.0473.pdf
#
encoder_inputs = Input(shape=(length_input, ), name='encoder_inputs')
encoder_embedding = embedding_layer(encoder_inputs)
ecoder_lstm = Bidirectional(LSTM(layer_dims,
return_state=True,
return_sequences=True,
dropout=0.05,
recurrent_initializer='glorot_uniform',
name='encoder_lstm'),
name='encoder_bidirectional')
encoder_outputs, forward_h, forward_c, backward_h, backward_c = ecoder_lstm(encoder_embedding)
# For annotating sequences, we concatenate forward hidden state with backward one as explained in top paper
state_h = Concatenate(name='encoder_hidden_state')([forward_h, backward_h])
state_c = Concatenate(name='encoder_cell_state')([forward_c, backward_c])
encoder_states = [state_h, state_c]
#
# Decoder,
# Unidirectional
#
decoder_inputs = Input(shape=(length_input, ), name='decoder_inputs')
decoder_embedding = embedding_layer(decoder_inputs)
decoder_lstm = LSTM(layer_dims * 2, # to match bidirectional size
return_state=True,
return_sequences=True,
dropout=0.05,
recurrent_initializer='glorot_uniform',
name='decoder_lstm')
# Set encoder to use the encoder state as initial states
decoder_output,_ , _, = decoder_lstm(decoder_embedding,
initial_state=encoder_states)
# Attention
attention_layer = AttentionLayer(name='attention_layer')
attention_output, attention_state = attention_layer([encoder_outputs, decoder_output])
decoder_concat = Concatenate(axis=-1)([decoder_output, attention_output])
# Output layer
outputs = Dense(size_vocab, name='outputs', activation='softmax')(decoder_concat)
return Model([encoder_inputs, decoder_inputs], outputs), decoder_embedding
# +
model, decoder_embedding = seq2seq(embedding_layer,
HYPER_PARAMS['MAX_SENT_LENGTH'],
size_vocab,
HYPER_PARAMS['MODEL_LAYER_DIMS'])
optimizer = Adam(learning_rate=HYPER_PARAMS['MODEL_LEARN_RATE'])
model.compile(optimizer=optimizer, loss='sparse_categorical_crossentropy', metrics=['acc'])
SVG(model_to_dot(model, show_shapes=True, show_layer_names=False,
rankdir='TB', dpi=65).create(prog='dot', format='svg'))
# -
# ### Training
# Optionally load weights
if HYPER_PARAMS['MODEL_TRAINING']:
model.fit(dataset, epochs=HYPER_PARAMS['MODEL_LEARN_EPOCHS'], batch_size=HYPER_PARAMS['DATA_BATCH_SIZE'])
model.save('backup_{0}.h5'.format(HYPER_PARAMS['MODEL_LEARN_EPOCHS']))
else:
model.load_weights('backup_{0}.h5'.format(HYPER_PARAMS['MODEL_LEARN_EPOCHS']))
# ### Predictions processing
# - Follow the same tokeninzing approach as before
# - For words that are not in our original context, we use the GloVe embedding to find the most similar word within our vocabulary. If it is still a weird word, it will be replaced by an unknown token
# +
def get_known_words(sentence_words, glove_word_dictionary, local_word_dictionary, topn=20):
"""Pre-process the input text, corrects spelling,
get similar words from gensim if input word is out of context
or replace with <UNK> for bad words."""
result = []
for word in sentence_words:
# Correct any spelling mistakes
spell = SpellChecker()
word = spell.correction(word)
if word in local_word_dictionary:
result.append(word)
else:
# Determine if top match is within our local context
found_sim_word = False
if word in glove_word_dictionary:
similar_words = glove_vectors.similar_by_word(word, topn=topn)
for (sim_word, measure) in similar_words:
if sim_word in local_word_dictionary.keys():
found_sim_word = True
result.append(sim_word)
break
# Funny word
if not found_sim_word:
result.append('<UNK>')
return result
def pre_process_new_questions(text, tokenizer, glove_vectors, special_tokens):
""" Process text to words within our context,
and tokenize."""
# Pre-process
text_ = clean_text_to_lower(text)
split_text = text_.split(' ')
split_text = get_known_words(split_text, glove_vectors, tokenizer.word_index)
processed_question = " ".join(split_text)
# Tokenize to sequence of ints
# using original tokenizer
tokenized_question = tokenizer.texts_to_sequences([processed_question])
tokenized_question = [[special_tokens['<SOS>']] +
tokenized_question[0] +
[special_tokens['<EOS>']]]
# Padding
tokenized_question = pad_sequences(tokenized_question,
maxlen=HYPER_PARAMS['MAX_SENT_LENGTH'] - 1,
padding='post')
return processed_question, tokenized_question[0]
def post_process_new_answers(text_sequence, tokenizer, glove_vectors):
answer = []
# Build string from text sequence
for text_id in text_sequence:
if text_id != tokenizer.word_index['<EOS>'] and\
text_id != tokenizer.word_index['<PAD>']:
word = tokenizer.index_word[text_id]
if len(answer) == 0:
word = word.capitalize()
answer.append(word)
answer = " ".join(answer)
return answer
# -
# **Example of correction** <br>
# Vanish is not within our model's context, but dissapear is
processed_question, _ = pre_process_new_questions('i wouzld like tso vansish', tokenizer, glove_vectors, special_tokens)
processed_question
# ### Predictions
# +
def get_encoder_decoder(model):
""" Get the encoder and decoder models,
used to do inference of text sequences"""
# Encoder model
encoder_model = Model(model.get_layer('encoder_inputs').output,
[model.get_layer('encoder_bidirectional').output[0],
[model.get_layer('encoder_hidden_state').output,
model.get_layer('encoder_cell_state').output]])
# Decoder model
decoder_state_input_h = Input(shape=(HYPER_PARAMS['MODEL_LAYER_DIMS'] * 2, ))
decoder_state_input_c = Input(shape=(HYPER_PARAMS['MODEL_LAYER_DIMS'] * 2, ))
decoder_states_inputs = [decoder_state_input_h, decoder_state_input_c]
decoder_outputs, state_h, state_c = model.get_layer('decoder_lstm')(decoder_embedding,
initial_state=decoder_states_inputs)
decoder_states = [state_h, state_c]
decoder_model = Model([model.get_layer('decoder_inputs').output,
decoder_states_inputs],
[decoder_outputs] + decoder_states)
return encoder_model, decoder_model
def infer_answer_sentence(model, encoder_model, decoder_model, raw_question, tokenizer, glove_vectors, special_tokens):
processed_question, tokenized_question = pre_process_new_questions(raw_question,
tokenizer,
glove_vectors,
special_tokens)
tokenized_question = tf.expand_dims(tokenized_question, axis=0)
# Encode input sentence
encoder_outputs, states = encoder_model.predict(tokenized_question)
# Create starting input with only <SOS> token
decoder_input = tf.expand_dims(special_tokens['<SOS>'], 0)
decoded_answer_sequence = []
while len(decoded_answer_sequence) < HYPER_PARAMS['MAX_SENT_LENGTH']:
decoder_outputs , decoder_hidden_states , decoder_cell_states = decoder_model.predict([decoder_input] + states)
# Apply attention to decoder output
attention_layer = AttentionLayer()
attention_outputs, attention_states = attention_layer([encoder_outputs, decoder_outputs])
decoder_concatenated = Concatenate(axis=-1)([decoder_outputs, attention_outputs])
dense_output = model.get_layer('outputs')(decoder_concatenated)
# Get argmax of output
sampled_word = np.argmax(dense_output[0, -1, :])
# Finished output sentence
if sampled_word == special_tokens['<EOS>']:
break
# Store prediction, and use it as the decoder's new input
decoded_answer_sequence.append(sampled_word)
decoder_input = tf.expand_dims(sampled_word, 0)
# Update internal states each timestep
states = [decoder_hidden_states, decoder_cell_states]
answer = post_process_new_answers(decoded_answer_sequence, tokenizer, glove_vectors)
return answer
# -
encoder_model, decoder_model = get_encoder_decoder(model)
# ## Interface with Telegram
# +
# Enable logging
logging.basicConfig(
format='%(asctime)s - %(name)s - %(levelname)s - %(message)s', level=logging.INFO
)
updater = Updater("1396342801:AAElqHfd2RGJI-lhdbVevFYSIW4HolMjS7E", use_context=True)
# Get the dispatcher to register handlers
dispatcher = updater.dispatcher
spell = SpellChecker()
# +
WELCOMED_IDS = []
MARKUP_MESSAGES = defaultdict(int)
def welcome_message(chat_id, context):
context.bot.send_message(chat_id=chat_id, text=
'Hi there! I am NLTChatBot. \n' +
'I have studied the transcripts of 617 movies to learn to speak.' +
' Ask me anything! ' + u'\U0001F603' )
WELCOMED_IDS.append(chat_id)
def check_grammar(message):
_message = message.replace("?", "").split(' ')
processed_message = []
highlited_grammar = []
flag = False
for word in _message:
corrected_word = spell.correction(word)
processed_message.append(corrected_word)
if corrected_word != word:
# Bold correction
highlited_grammar.append('<b>'+ word + '</b>')
flag = True
else:
highlited_grammar.append(word)
return ' '.join(highlited_grammar), ' '.join(processed_message), flag
def handle_message(update, context):
# Delete old markup option if user didn't select it
if MARKUP_MESSAGES[update.effective_chat.id] != 0:
try:
context.bot.deleteMessage(chat_id = update.effective_chat.id,
message_id = MARKUP_MESSAGES[update.effective_chat.id])
# Chat possibly closed
except:
MARKUP_MESSAGES[update.effective_chat.id] = 0
MARKUP_MESSAGES[update.effective_chat.id] = 0
# New user?
if update.effective_chat.id not in WELCOMED_IDS:
welcome_message(update.effective_chat.id, context)
return
if HYPER_PARAMS['ENABLE_SPELLING_SUGGESTION']:
# Check grammar
highlited_grammar, processed_message, flag = check_grammar(update.message.text)
if flag:
# Spelling suggestions,
reply_keyboard = [[processed_message]]
message = update.message.reply_text('I am not sure what you meant with - ' + highlited_grammar,
reply_markup=ReplyKeyboardMarkup(reply_keyboard,
one_time_keyboard=True),
parse_mode='HTML')
MARKUP_MESSAGES[update.effective_chat.id] = message.message_id
return
else:
processed_message = update.message.text
# Reply to user
answer = infer_answer_sentence(model,
encoder_model,
decoder_model,
processed_message,
tokenizer,
glove_vectors,
special_tokens)
context.bot.send_message(chat_id=update.effective_chat.id, text=answer)
def received_message(update, context):
"""Mimics the typing on keybord,
before calling the message handling"""
if HYPER_PARAMS['MESSAGE_TYPING_DELAY'] > 0:
context.bot.send_chat_action(chat_id=update.effective_chat.id, action=ChatAction.TYPING)
timed_response = Timer(HYPER_PARAMS['MESSAGE_TYPING_DELAY'], handle_message, [update, context])
timed_response.start()
else:
handle_message(update, context)
# -
message_handler = MessageHandler(Filters.text & (~Filters.command), received_message)
dispatcher.add_handler(message_handler)
updater.start_polling()
|
NLPChatBot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _nb_visualization:
# -
# # Visualization
# + raw_mimetype="text/restructuredtext" active=""
# .. toctree::
# :maxdepth: 1
# :hidden:
#
# scatter.ipynb
# pcp.ipynb
# heatmap.ipynb
# petal.ipynb
# radar.ipynb
# radviz.ipynb
# star.ipynb
# video.ipynb
#
# -
# Different visualization techniques are available. Each of them has different purposes and is suitable for less or higher dimensional objective spaces.
#
# The following visualizations can be used:
#
# |Name|Class|Convenience|
# |---|---|---|
# |[Scatter Plots (2D/3D/ND)](scatter.ipynb)|Scatter|"scatter"|
# |[Parallel Coordinate Plots (PCP)](pcp.ipynb)|ParallelCoordinatePlot|"pcp"|
# |[Heatmap](heatmap.ipynb)|Heatmap|"heat"|
# |[Petal Diagram](petal.ipynb)|Petal|"petal"|
# |[Radar](radar.ipynb)|Radar|"radar"|
# |[Radviz](radviz.ipynb)|Radviz|"radviz"|
# |[Star Coordinates](star.ipynb)|StarCoordinate|"star"|
# |[Video](video.ipynb)|Video|-|
#
# Each of them is implemented in a class which can be used directly. However, it might
# be more comfortable to either use the factory function in some cases.
# For example for scatter plots the following initiates the same object:
# +
# directly using the class
from pymoo.visualization.scatter import Scatter
plot = Scatter()
# the global factory method
from pymoo.factory import get_visualization
plot = get_visualization("scatter")
# -
# The advantages of the convenience function is that just by changing the string a different visualization
# can be chosen (without changing any other import). Moreover, we desire to keep the global interface in the factory the same, whereas the implementation details, such as class names might change.
# Please note, that the visualization implementations are just a wrapper around [matplotlib](https://matplotlib.org) and all keyword arguments are still useable.
# For instance, if two different set of points should be plotted in different colors with different markers in a scatter plot:
#
# +
import numpy as np
A = np.random.random((20,2))
B = np.random.random((20,2))
from pymoo.factory import get_visualization
plot = get_visualization("scatter")
plot.add(A, color="green", marker="x")
plot.add(B, color="red", marker="*")
plot.show()
# -
# This holds for all our visualizations. However, depending on the visualization the matplotlib function that is used and the corresponding keyword arguments might change. For example, in for the PetalWidth Plot polygons are drawn which has different keywords than the plot of matplotlib.
# Furthermore, the plots have some default arguments to be used to set them during initialization:
# +
from pymoo.visualization.petal import Petal
from pymoo.visualization.util import default_number_to_text
np.random.seed(5)
A = np.random.random((1,6))
plot = Petal(
# change the overall figure size (does not work for all plots)
figsize=(8, 6),
# directly provide the title (str or tuple for options)
title=("My Plot", {"pad" : 30}),
# plot a legend (tuple for options)
legend=False,
# make the layout tight before returning
tight_layout=True,
# the boundaries for normalization purposes (does not apply for every plot
# either 2d array [[min1,..minN],[max1,...,maxN]] or just two numbers [min,max]
bounds=[0,1],
# if normalized, the reverse can be potted (1-values)
reverse=False,
# the color map to be used
cmap="tab10",
# modification of the axis style
axis_style=None,
# function to be used to plot numbers
func_number_to_text=default_number_to_text,
# change the axis labels - could be a list just the prefix
axis_labels=["Objective %s" % i for i in range(1,7)],
)
plot.add(A, label="A")
plot.show()
# -
# For each visualization a documentation is provided.
|
source/visualization/index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/victordibia/taxi/blob/main/notebooks/taxi_model_training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Kwkhb5bOpjRq"
pip install scikit-learn==0.23.2 pandas==1.1.3
# + id="ssdEOTGs_M6u"
import pandas as pd
import numpy as np
from tqdm.notebook import tqdm
# + id="3R_JRetrInvM"
df = pd.read_csv("taxi/data/training.csv")
# + id="tS2N5tAnJIOl"
categorical_features = ["month","week","hour","isweekday","holiday", "dayofweek", "PULocationID", "DOLocationID"] #["PULocationID", "DOLocationID"]
feature_list = ["passenger_count","trip_time","fare_amount"]
numeric_features = ["passenger_count"]
dfa = df
dfa[categorical_features] = dfa[categorical_features].astype("category")
df_data = dfa[feature_list + categorical_features]
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="udNeyu8po_fn" outputId="b0405e04-2356-4a2c-d193-5d02eb1caf56"
train_sample_size = 300000
df_draw = df_data.sample(train_sample_size, random_state=42)
both_labels = df_draw[["fare_amount", "trip_time"]]
df_draw.drop(["trip_time","fare_amount"], axis =1, inplace=True)
df_draw.head(2)
# + colab={"base_uri": "https://localhost:8080/"} id="gKsrhJybov6w" outputId="e4a1e9a0-2501-408b-b01f-f636c8fdff53"
categorical_idx = df_draw.columns.get_indexer(categorical_features)
numeric_idx = df_draw.columns.get_indexer(numeric_features)
categorical_idx, numeric_idx
# + id="WCJAW34yJNTL"
import time
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.neural_network import MLPRegressor
from sklearn.svm import SVR
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder, StandardScaler, LabelEncoder
from sklearn.metrics import classification_report, mean_squared_error, mean_absolute_error, mean_squared_log_error
from sklearn.compose import ColumnTransformer
from sklearn.compose import make_column_selector as selector
num_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
cat_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
preprocessor = ColumnTransformer(
transformers=[
# ('num', num_transformer, selector(dtype_include='float64')),
('num', num_transformer, numeric_idx),
('cat', cat_transformer, categorical_idx)
])
# keep track of all details for models we train
def train_model(model, data, labels):
X = data.values
y = labels.values
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# pipe = Pipeline([('scaler', StandardScaler()),('clf', model["clf"])])
pipe = Pipeline(steps=[
('preprocessor', preprocessor),
('clf', model["clf"])])
start_time = time.time()
# with joblib.parallel_backend('dask'):
# pipe.fit(X_train, y_train)
pipe.fit(X_train, y_train)
train_time = time.time() - start_time
train_preds = pipe.predict(X_train)
test_preds = pipe.predict(X_test)
np.histogram(train_preds)
np.histogram(test_preds)
train_rmse = np.sqrt(mean_squared_error(y_train, train_preds))
test_rmse = np.sqrt(mean_squared_error(y_test, test_preds))
train_mae = mean_absolute_error(y_train, train_preds)
test_mae = mean_absolute_error(y_test, test_preds)
train_accuracy = pipe.score(X_train, y_train)
test_accuracy = pipe.score(X_test, y_test)
model_details = {"name": model["name"],
"train_rmse":train_rmse,
"test_rmse":test_rmse,
"train_mae":train_mae,
"test_mae":test_mae,
"train_time": train_time,
"model": pipe,
"y_test": y_test,
"test_preds": test_preds
}
return model_details
models = [
{"name": "Random Forest", "clf": RandomForestRegressor(n_estimators=100)},
# {"name": "Gradient Boosting", "clf": GradientBoostingRegressor(n_estimators=100)},
{"name": "MLP Classifier", "clf": MLPRegressor(solver='adam', alpha=1e-1, hidden_layer_sizes=(20,20,10,5,2), max_iter=500, random_state=42)}
]
# + colab={"base_uri": "https://localhost:8080/", "height": 100, "referenced_widgets": ["d1bd0549db2e438e8ade9327c701d874", "c581476b7e1f4b0ba89d18917444b637", "743c9ce76d904d6b9952977f0addefba", "5dc28368ceff48d4a4411ec9dc4212a6", "2a572e37d003423f84594fe95488da41", "ede8d0bacc9e498e891b3ee33385087a", "b26049d81ea5465cab3aed38a4fea362", "90fd406027f24f608e096a486da83a10"]} id="xqSOQ6VMJWVz" outputId="cd038a4b-5bfb-4dde-b26d-7ddc337d0918"
def plot_models(trained_models,outcome_type):
# visualize accuracy and run time
setup_plot()
model_df = pd.DataFrame(trained_models)
model_df.sort_values("test_rmse", inplace=True)
ax = model_df[["train_rmse","test_rmse", "name"]].plot(kind="line", x="name", figsize=(19,5), title="Classifier Performance Sorted by Test Accuracy - " + outcome_type)
ax.legend(["Train RMSE", "Test RMSE"])
for p in ax.patches:
ax.annotate( str( round(p.get_height(),3) ), (p.get_x() * 1.005, p.get_height() * 1.005))
ax.title.set_size(20)
plt.box(False)
model_df.sort_values("train_time", inplace=True)
ax= model_df[["train_time","name"]].plot(kind="line", x="name", figsize=(19,5), grid=True, title="Classifier Training Time (seconds)" + outcome_type)
ax.title.set_size(20)
ax.legend(["Train Time"])
plt.box(False)
def train_models(models, data, labels, outcome_type):
trained_models = []
for model in tqdm(models):
model_details = train_model(model, data, labels)
model_details["label"] = outcome_type
trained_models.append(model_details)
return trained_models
# %time multi_output_model = train_models(models, df_draw, both_labels, "Trip Time")
# + id="8WWG2YQ2Jizo"
def plot_trained_models(trained_model, title, window_size=100):
plt.figure(figsize=(14,5))
fare = [x[0] for x in trained_model["y_test"][:window_size]]
triptime = [x[1] for x in trained_model["y_test"][:window_size]]
fare_preds = [x[0] for x in trained_model["test_preds"][:window_size]]
triptime_preds = [x[1] for x in trained_model["test_preds"][:window_size]]
plt.plot(fare , label="Ground Truth")
plt.plot(fare_preds , label="Predicitions" )
plt.title("Model Predictions vs Ground Truth | " + trained_model["name"] + " | " + " Fare ")
plt.legend(loc="upper right")
plt.figure(figsize=(14,5))
plt.plot(triptime , label="Ground Truth")
plt.plot(triptime_preds , label="Predicitions" )
plt.title("Model Predictions vs Ground Truth | " + trained_model["name"] + " | " + " Time ")
plt.legend(loc="upper right")
print("RMSE: Train",trained_model["train_rmse"], "Test",trained_model["test_rmse"] )
print("MAE: Train",trained_model["train_mae"], "Test",trained_model["test_mae"] )
# + colab={"base_uri": "https://localhost:8080/"} id="fvMSV8e6p3ib" outputId="9a3ac15f-5db8-44e9-bacb-3f59f3882ebe"
multi_output_model[0]["test_preds"].shape, multi_output_model[0]["y_test"].shape
# + id="axJdMzb2wfPh" colab={"base_uri": "https://localhost:8080/", "height": 689} outputId="a2ce76e6-d28a-4467-f23e-701c4c61a26f"
plot_trained_models(multi_output_model[0], "Trip Time")
# + id="V9u9DfFrLWBA" colab={"base_uri": "https://localhost:8080/", "height": 689} outputId="6ab1baf2-29e0-4338-f401-2ed7d9b0b704"
plot_trained_models(multi_output_model[1], "Trip Time")
# + id="rm9JTSY8KXd9"
def result_to_df(trained_model, title):
print("Results for ", title)
model_df = pd.DataFrame(trained_model)
return model_df[["name","train_rmse","test_rmse", "train_mae","test_mae"]]
# + id="8uuXfnhAKcJB" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="d561a8f6-a391-44c5-ca16-6fb3ef0f739a"
result_to_df(multi_output_model, "Trip Fare")
# + [markdown] id="P6b-xqk9x_nX"
# ## Export Model
#
# - Write models to joblib file which can then be used to setup a Cloud AI Platform Model Endpoint.
# + id="_JWOjiSBLIFk"
# !mkdir models
# !mkdir models/mlp
# !mkdir models/randomforest
joblib.dump(multi_output_model[0]["model"],"models/randomforest/model.joblib")
joblib.dump(multi_output_model[1]["model"],"models/mlp/model.joblib")
# !gsutil -m cp -r models gs://taximodel
# + id="DUwrWtkKpd5-"
|
notebooks/taxi_model_training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Basic RNNs in Tensorflow
# - 1 Layer of 5 recurrent neurons
# - Outputs of a layer of recurrent neurons for all instances in a mini-batch
# $$ \textbf{Y}_{(t)} = \phi \big( \textbf{X}_{(t)} . \textbf{W}_x + \textbf{Y}_{(t-1)}^T . \textbf{W}_y + b \big) $$
# $$ = \phi \big( \big[ \textbf{X}_{(t)} \textbf{Y}_{(t-1)} \big] . \textbf{W} + b \big)$$
#
# where $ \textbf{W} = {\textbf{W}_x\brack \textbf{W}_y} $
#
# #### Notations in vectorized form:
# - $m =$ number of instances in mini-batch
# - $n_{neurons}$ number of neurons
# - $n_{inputs}$ number of input features
# - $dim(x)$ a function that determines shape or dimension of element $x$.
# - $\textbf{Y}_{(t)}$ is the layer output at time $t$ for each instance of the mini-batch:
# $$ dim(\textbf{Y}_{(t)}) = m\times n_{neurons}$$
# - $\textbf{X}_{(t)}$ is a matrix containing the inputs for all instances:
# $$ dim(\textbf{X}_{(t)}) = m \times n_{inputs}$$
# - $\textbf{W}_{x}$ is a matrix containing the connections weights for the inputs of the **current** time step:
# $$ dim(\textbf{W}_{x}) = n_{inputs} \times n_{neurons} $$
# - $\textbf{W}_{y}$ is a matrix containing the connections werights for the outputs of the **previous** time step:
# $$ dim(\textbf{W}_{y}) = n_{nuerons} \times n_{neurons} $$
# - Weight matrices $\textbf{W}_x$ and $\textbf{W}_y$ are often concatenated into a single matrix $\textbf{W}$ of shape $(n_{inputs} + n_{nuerons}) \times n_{neurons}$
# - Bias term $b$ is just a 1-dimensional vector of size $1 \times n_{neurons}$
# #### TODO: Put image of network we are building here
# +
import tensorflow as tf
from tensorflow.contrib.layers import fully_connected
import os
import numpy as np
tf.set_random_seed(1) # seed to obtain similar outputs
os.environ['CUDA_VISIBLE_DEVICES'] = '' # avoids using GPU for this session
# -
# ### Single Recurrent Neuron
# 
# +
# Implementation of RNN with Single Neuron
N_INPUTS = 4
N_NEURONS = 1
class SingleRNN(object):
def __init__(self, n_inputs, n_neurons):
self.X0 = tf.placeholder(tf.float32, [None, n_inputs])
self.X1 = tf.placeholder(tf.float32, [None, n_inputs])
self.Wx = tf.Variable(tf.random_normal(shape=[n_inputs, n_neurons], dtype=tf.float32))
self.Wy = tf.Variable(tf.random_normal(shape=[n_neurons, n_neurons], dtype=tf.float32))
b = tf.Variable(tf.zeros([1, n_neurons], dtype=tf.float32))
self.Y0 = tf.tanh(tf.matmul(self.X0, self.Wx) + b)
self.Y1 = tf.tanh(tf.matmul(self.Y0, self.Wy) + tf.matmul(self.X1, self.Wx) + b)
# +
# Now we feed input at both time steps
# Generate mini-batch with 4 instances (i.e., each instance has an input sequence of exactly two inputs)
# instance1 instance2 instance3 instance4
X0_batch = np.array([[0,1,2,0], [3,4,5,0], [6,7,8,0], [9,0,1,0]]) # t = 0
X1_batch = np.array([[9,8,7,0], [0,0,0,0], [6,5,4,0], [3,2,1,0]]) # t = 1
model = SingleRNN(N_INPUTS, N_NEURONS)
with tf.Session() as sess:
# initialize and run all variables so that we can use their values directly
init = tf.global_variables_initializer()
sess.run(init)
Y0_val, Y1_val, Wx, Wy = sess.run([model.Y0, model.Y1, model.Wx, model.Wy], feed_dict={model.X0: X0_batch, model.X1: X1_batch})
# -
print(Y0_val)
print(Wx)
# ---
# ### Basic Layer of Recurrent Neurons
# 
# +
# RNN unrolled through two time steps
N_INPUTS = 3 # number of features in input
N_NEURONS = 5
class BasicRNN(object):
def __init__(self, n_inputs, n_neurons):
self.X0 = tf.placeholder(tf.float32, [None, n_inputs])
self.X1 = tf.placeholder(tf.float32, [None, n_inputs])
Wx = tf.Variable(tf.random_normal(shape=[n_inputs, n_neurons], dtype=tf.float32))
Wy = tf.Variable(tf.random_normal(shape=[n_neurons, n_neurons], dtype=tf.float32))
b = tf.Variable(tf.zeros([1, n_neurons], dtype=tf.float32))
self.Y0 = tf.tanh(tf.matmul(self.X0, Wx) + b)
self.Y1 = tf.tanh(tf.matmul(self.Y0, Wy) + tf.matmul(self.X1, Wx) + b)
# +
# Now we feed input at both time steps
# Generate mini-batch with 4 instances (i.e., each instance has an input sequence of exactly two inputs)
# instance1 instance2 instance3 instance4
X0_batch = np.array([[0,1,2], [3,4,5], [6,7,8], [9,0,1]]) # t = 0
X1_batch = np.array([[9,8,7], [0,0,0], [6,5,4], [3,2,1]]) # t = 1
model = BasicRNN(N_INPUTS, N_NEURONS)
with tf.Session() as sess:
# initialize and run all variables so that we can use their values directly
init = tf.global_variables_initializer()
sess.run(init)
Y0_val, Y1_val = sess.run([model.Y0, model.Y1], feed_dict={model.X0: X0_batch, model.X1: X1_batch})
# -
print(Y0_val) # output at t = 0 with 4 X 5 dimensions (m X n_neurons)
print(Y1_val) # output at t = 1
# ---
# ### RNNs using Static Unrolling Through Time
#
# 
#
# Deals with situations when dealing with extremely large inputs and outputs
class StaticRNN(object):
def __init__(self, n_steps, n_inputs, n_neurons):
self.X = tf.placeholder(tf.float32, [None, n_steps, n_inputs])
X_seqs = tf.unstack(tf.transpose(self.X, perm =[1,0,2]))
basic_cell = tf.contrib.rnn.BasicRNNCell(num_units=n_neurons) # build copies of cell for each time step (unrolling).
output_seqs, states = tf.contrib.rnn.static_rnn(basic_cell, X_seqs, dtype=tf.float32) # does chaining for each input with the cells
self.outputs = tf.transpose(tf.stack(output_seqs), perm=[1,0,2])
# must create 3D tensor as this is what Tensorflow RNN Cell requires
X_batch = np.array([
# t = 0 t = 1
[[0,1,2], [9,8,7]], # instance 0
[[3,4,5], [0,0,0]], # instance 1
[[6,7,8], [6,5,4]], # instance 2
[[9,0,1], [3,2,1]], # instnace 3
])
X_batch.shape
# +
tf.reset_default_graph() # resetting graph everything we run the code to avoid TF error
N_STEPS = 2
model = StaticRNN(N_STEPS, N_INPUTS, N_NEURONS)
with tf.Session() as sess:
# initialize and run all variables so that we can use their values directly
init = tf.global_variables_initializer()
sess.run(init)
output_vals = model.outputs.eval(feed_dict={model.X: X_batch})
# -
print(output_vals)
# There is still a problem with Static RNN version because it builds one cell per time step, which can amplify if we had a very large time steps.
# ---
# ### Dynamic Unrolling Through Time
# Basically runs a while loop over a cell the appropriate number of times. Definitely a much cleaner and computationally effecient way of building RNNs on tensorflow. Also, no need to unstack, stack, or transpose.
#
# Here we also handle variable length input sequences.
class DynamicRNN(object):
def __init__(self, n_steps, n_inputs, n_neurons):
# denotes the size of the sequence (helpful for supporting varying sizes of input sequences, e.g., sentences)
self.seq_length = tf.placeholder(tf.int32, [None])
self.X = tf.placeholder(tf.float32, [None, n_steps, n_inputs]) # 3D tensor
basic_cell = tf.contrib.rnn.BasicRNNCell(num_units=n_neurons)
self.outputs, self.states = tf.nn.dynamic_rnn(basic_cell, self.X, dtype=tf.float32, sequence_length=self.seq_length)
# +
# must create 3D tensor as this is what Tensorflow RNN Cell requires
X_batch = np.array([
# t = 0 t = 1
[[0,1,2], [9,8,7]], # instance 0
[[3,4,5], [0,0,0]], # instance 1 (padded with a zero vector)
[[6,7,8], [6,5,4]], # instance 2
[[9,0,1], [3,2,1]], # instnace 4
])
seq_length_batch = np.array([2,1,2,2])
# +
tf.reset_default_graph() # resetting graph everything we run the code to avoid TF error
N_STEPS = 2
model = DynamicRNN(N_STEPS, N_INPUTS, N_NEURONS)
with tf.Session() as sess:
# initialize and run all variables so that we can use their values directly
init = tf.global_variables_initializer()
sess.run(init)
outputs_val, states_val = sess.run([model.outputs, model.states],
feed_dict={model.X: X_batch, model.seq_length: seq_length_batch})
# -
print(outputs_val) # pairs t = 0 , t = 1 and outputs them as multidimensional array
print(states_val) # notice that states contain the final state (i.e, t = 1)
# ---
# ### Training RNN Classifier on MNIST
#
# 
#
# Note that even though image classification can be done more effectively using CNN, RNNs will still perform well since the sequence is also important in the process of drawing digits.
class ImageRNN(object):
def __init__(self, n_steps, n_inputs, n_neurons, n_outputs):
self.X = tf.placeholder(tf.float32, [None, n_steps, n_inputs])
self.y = tf.placeholder(tf.int32, [None])
basic_cell = tf.contrib.rnn.BasicRNNCell(num_units=n_neurons)
outputs, states = tf.nn.dynamic_rnn(basic_cell, self.X, dtype=tf.float32)
# computes loss
logits = fully_connected(states, n_outputs, activation_fn=None) # log probabilities
self.loss = tf.reduce_mean(tf.nn.sparse_softmax_cross_entropy_with_logits(labels=self.y, logits=logits))
# evaluation (accuracy)
correct = tf.nn.in_top_k(logits, self.y, 1) # tf.equal(tf.argmax(logits, 1), tf.argmax(self.y, 1))
self.accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
# parameters
N_STEPS = 28
N_INPUTS = 28
N_NEURONS = 150
N_OUTPUTS = 10
N_EPHOCS = 10
BATCH_SIZE = 150
DISPLAY_STEP = 1
LEARNING_RATE = 0.001
# +
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/")
X_test = mnist.test.images.reshape((-1, N_STEPS, N_INPUTS))
y_test = mnist.test.labels
# +
tf.reset_default_graph()
# build model
model = ImageRNN(N_STEPS, N_INPUTS, N_NEURONS, N_OUTPUTS)
# training procedures (backpropagation)
optimizer = tf.train.AdamOptimizer(learning_rate=LEARNING_RATE)
training_op = optimizer.minimize(model.loss)
with tf.Session() as sess:
# initialize and run all variables
init = tf.global_variables_initializer()
sess.run(init)
for epoch in range(N_EPHOCS):
avg_cost = 0. # average loss
total_batch = mnist.train.num_examples // BATCH_SIZE
for iteration in range(total_batch): # note iterations are depended on batch size (increase for faster computation)
X_batch, y_batch = mnist.train.next_batch(BATCH_SIZE)
X_batch = X_batch.reshape((-1, N_STEPS, N_INPUTS))
_, cost = sess.run([training_op, model.loss], feed_dict={model.X: X_batch, model.y: y_batch})
avg_cost += cost / total_batch
acc_train = model.accuracy.eval(feed_dict={model.X: X_batch, model.y: y_batch})
acc_test = model.accuracy.eval(feed_dict={model.X: X_test, model.y: y_test})
# Display cost, accuracy (test, train) based on display step
if (epoch+1) % DISPLAY_STEP == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", "{:.9f}".format(avg_cost))
print("Train Accuracy: ", acc_train, " ", "Test Accuracy: ", acc_test)
print("Training Finished!!!")
# -
# ---
# ### References:
# - [All sorts of Text Classificaiton Deep Learning models](https://github.com/brightmart/text_classification)
# - [NTHU Machine Learning](https://nthu-datalab.github.io/ml/labs/13_Sentiment_Analysis_and_Neural_Machine_Translation/13_Sentiment_Analysis_and_Neural_Machine_Translation.html)
# - [Introduction to RNN](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/)
|
1.1_Intro_RNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
import seaborn
# import mpld3
import numpy as np
seaborn.set()
np.set_printoptions(precision=4, suppress=True)
# mpld3.enable_notebook()
# -
mirex_data = json.load(open("mirex_runs.json"))
years = [yr for yr in sorted(mirex_data.keys()) if mirex_data[yr]]
counts = [mirex_data[yr][0] for yr in years]
years, counts
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
idx = np.arange(len(counts))
ax.bar(idx - 0.5, counts, width=1.0, fc='green', ec='k')
ax.set_xlim(-0.5, len(counts) - 0.5)
ax.set_xticks(idx)
ax.set_xticklabels(years, rotation=30)
ax.set_ylabel("Algorithm Runs")
x_margin = 0.25
fig, ax = plt.subplots(1, 1, figsize=(8, 4))
idx = np.arange(len(counts))
ax.plot(idx, counts, 'k', marker='o')
ax.set_xlim(-x_margin, len(counts)-1+x_margin)
ax.set_xticks(idx)
ax.set_xticklabels(years, rotation=30)
ax.set_ylabel("Algorithm Runs")
ax.set_xlabel("MIREX Year")
plt.tight_layout()
fig.savefig("../paper/figs/mirex_runs.pdf")
|
data/mirex-viz.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysing merge sort and modified merge sort - CZ2001 Lab4
#
# ### Aim:
# 1. Generate datasets of varying sizes (1,000 - 1 million)
# 2. Count key comparisons and CPU times on the data generated, and compare the two variations of mergesort on these parameters
# 3. Study the performance of the algorithm as `S` varies
# 4. Determine the value of `S` by trial and error
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
import random, csv, copy
import mergeAndInsertion
# ## Getting comparison metrics for original mergesort
n=1000
data = [random.randint(1, 20000) for _ in range(n)] # below fine if you run it the second time?!
# testing mergesort
m=mergeAndInsertion.mergeAndInsertion(copy.deepcopy(data))
# print('time taken:',m.timeTaken)
m.testMerge()
mergeTime = m.timeTaken
mergeComparisons = m.keyCmpCounter
print('Mergesort:')
print(f'\ttime taken : {mergeTime}ns \n\tkeyComparisons : {mergeComparisons}')
# ## Analyzing modified mergesort
# letting 'S' vary from 0 to n
tableList =[]
for S in range(0,n):
m = mergeAndInsertion.mergeAndInsertion(copy.deepcopy(data));
m.testMergeModified(S)
time = m.timeTaken
comparisons = m.keyCmpCounter
tableList.append([S, time, comparisons])
tableDf = pd.DataFrame(tableList, columns = ['S', 'time', 'keyCmp'])
tableDf.head()
# +
x = tableDf.iloc[:,0]
y = tableDf.iloc[:,1]
z = pd.Series([mergeTime]*n)
fig,ax = plt.subplots()
ax.plot(x,y,z)
ax.set(xlabel = 'S', ylabel = 'time')
x = tableDf.iloc[:,0]
y = tableDf.iloc[:,2]
z = pd.Series([mergeComparisons]*n) # y = mergeComparisons
fig2,ax2 = plt.subplots()
ax2.plot(x,y,z)
# ax.plot([0,n], [mergeComparisons,mergeComparisons])
ax2.set(xlabel = 'S', ylabel = 'keycomparison')
plt.show()
# -
x = tableDf.iloc[:,0]
y = tableDf.iloc[:,2]
z = pd.Series([mergeComparisons]*n) # y = mergeComparisons
fig,ax = plt.subplots()
ax.plot(x,y,z)
# ax.plot([0,n], [mergeComparisons,mergeComparisons])
ax.set(xlabel = 'S', ylabel = 'keycomparison')
plt.show()
# # Function with which we vary 'n'
def compareOriginalAndModified(n, ascending = False, descending = False, step = 1):
data = [random.randint(1, 20000) for _ in range(n)]
if(ascending):
data = sorted(data)
if(descending):
data = sorted(data, reverse = True)
# Mergesort
m=mergeAndInsertion.mergeAndInsertion(copy.deepcopy(data))
m.testMerge()
mergeTime = m.timeTaken
mergeComparisons = m.keyCmpCounter
print('Mergesort:')
print(f'\ttime taken : {mergeTime}ns \n\tkeyComparisons : {mergeComparisons}')
# Mergesort modifed :
# letting 'S' vary from 0 to n
tableList =[]
for S in range(0,n, step):
m = mergeAndInsertion.mergeAndInsertion(copy.deepcopy(data));
m.testMergeModified(S)
time = m.timeTaken
comparisons = m.keyCmpCounter
tableList.append([S, time, comparisons])
tableDf = pd.DataFrame(tableList, columns = ['S', 'time', 'keyCmp'])
x = tableDf.iloc[:,0]
y = tableDf.iloc[:,1]
z = pd.Series([mergeTime]*n)
fig,ax = plt.subplots()
ax.plot(x,y,z)
ax.set(xlabel = 'S', ylabel = 'time')
x = tableDf.iloc[:,0]
y = tableDf.iloc[:,2]
z = pd.Series([mergeComparisons]*n) # y = mergeComparisons
fig2,ax2 = plt.subplots()
ax2.plot(x,y,z)
# ax.plot([0,n], [mergeComparisons,mergeComparisons])
ax2.set(xlabel = 'S', ylabel = 'keycomparison')
plt.show()
print('------------------------------------------------------------')
# # Generating plots for different types of data:
n=1000
# ### Random data, size n
compareOriginalAndModified(n)
# ### Random data, size n, in ascending order
compareOriginalAndModified(n, ascending = True)
# ### Random data, size n, in descending order
compareOriginalAndModified(n, descending = True)
# ## Generating plots for different values of 'n'
# ### For n = 1000
compareOriginalAndModified(1000)
# ### For n = 10,000
compareOriginalAndModified(10000, step = 250)
# ### For n = 100,000
compareOriginalAndModified(100000, step = 2500)
|
lab3/modifiedMergesortAnalysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies and Setup
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# Hide warning messages in notebook
import warnings
warnings.filterwarnings('ignore')
# File to Load (Remember to Change These)
mouse_drug_data_to_load = "../Pymaceuticals/mouse_drug_data.csv"
clinical_trial_data_to_load = "../Pymaceuticals/clinicaltrial_data.csv"
# -
# Read the Mouse Data
mouse_drug_pd = pd.read_csv(mouse_drug_data_to_load)
# ## Tumor Response to Treatment
#Read the Clinical Trial Data
clinical_trial_pd = pd.read_csv(clinical_trial_data_to_load)
# Combine the data into a single dataset
# Display the data table for preview
drug_clinical_data = pd.merge(mouse_drug_pd, clinical_trial_pd, on='Mouse ID', how="right")
drug_clinical_data.columns
# +
# Store the Mean Tumor Volume Data Grouped by Drug and Timepoint
mean_tumor_volume_by_drug_timepoint_groupby = drug_clinical_data.groupby(['Drug', 'Timepoint']).mean()['Tumor Volume (mm3)']
#Reset index
mean_tumor_volume_by_drug_timepoint = mean_tumor_volume_by_drug_timepoint_groupby.reset_index()
# Convert to DataFrame
mean_tumor_volume_by_drug_timepoint_df = pd.DataFrame(mean_tumor_volume_by_drug_timepoint)
# -
# Preview DataFrame
mean_tumor_volume_by_drug_timepoint_df.head()
# +
# Store the Standard Error of Tumor Volumes Grouped by Drug and Timepoint
standard_error_tumor_volume_grouped_by_drug_timepoint = drug_clinical_data.groupby(['Drug', 'Timepoint']).sem()['Tumor Volume (mm3)']
# Convert to DataFrame
standard_error_tumor_volume_grouped_by_drug_timepoint_df = pd.DataFrame(standard_error_tumor_volume_grouped_by_drug_timepoint)
# Reset Index
reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df = standard_error_tumor_volume_grouped_by_drug_timepoint_df.reset_index()
reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.head()
# -
# Minor Data Munging to Re-Format the Data Frames(Mean Tumor Volume)
data_munged_mean_tumor_volume_by_drug_timepoint_groupby = mean_tumor_volume_by_drug_timepoint.pivot(columns='Drug',index='Timepoint')['Tumor Volume (mm3)']
data_munged_mean_tumor_volume_by_drug_timepoint_groupby.head()
# Minor Data Munging to Re-Format the Data Frames(Standard Error)
data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df = reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.pivot(columns='Drug',index='Timepoint')['Tumor Volume (mm3)']
data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.head()
# +
# Generate the Plot (with Error Bars)
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Capomulin'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Capomulin'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Infubinol'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Infubinol'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ketapril'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Ketapril'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Naftisol'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Naftisol'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Placebo'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Placebo'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Propriva'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Propriva'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ramicane'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Ramicane'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Stelasyn'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Stelasyn'])
plt.errorbar(data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df.index,data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Zoniferol'], marker="o", yerr=data_munged_reset_index_standard_error_tumor_volume_grouped_by_drug_timepoint_df['Zoniferol'])
plt.title("Mean Tumor Volume")
plt.ylabel("Timepoint")
plt.xlabel('Drug')
plt.grid(True)
plt.show()
# -
# Show the Figure
plt.show()
# 
# ## Metastatic Response to Treatment
# +
# Store the Mean Met. Site Data Grouped by Drug and Timepoint
mean_tumor_volume_by_metsite_groupby = drug_clinical_data.groupby(['Drug', 'Timepoint']).mean()['Metastatic Sites']
#Reset index
metsite_by_drug_timepoint = mean_tumor_volume_by_metsite_groupby.reset_index()
# Convert to DataFrame
metsite_by_drug_timepoint_df = pd.DataFrame(metsite_by_drug_timepoint)
# Preview DataFrame
metsite_by_drug_timepoint_df.head()
# +
# Store the Standard Error associated with Met. Sites Grouped by Drug and Timepoint
standard_error_metsite_grouped_by_drug_timepoint = drug_clinical_data.groupby(['Drug', 'Timepoint']).sem()['Metastatic Sites']
# Convert to DataFrame
standard_error_metsite_grouped_by_drug_timepoint_df = pd.DataFrame(standard_error_metsite_grouped_by_drug_timepoint)
# Reset Index
reset_index_standard_error_metsite_grouped_by_drug_timepoint_df = standard_error_metsite_grouped_by_drug_timepoint_df.reset_index()
# Preview DataFrame
reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.head()
# +
# Minor Data Munging to Re-Format the Data Frames
data_munged_mean_metsite_by_drug_timepoint_groupby = metsite_by_drug_timepoint_df.pivot(columns='Drug',index='Timepoint')['Metastatic Sites']
# Preview that Reformatting worked
data_munged_mean_metsite_by_drug_timepoint_groupby.head()
# -
# Minor Data Munging to Re-Format the Data Frames(Standard Error)
data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df = reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.pivot(columns='Drug',index='Timepoint')['Metastatic Sites']
data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.head()
# +
# Generate the Plot (with Error Bars)
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Capomulin'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Capomulin'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Ceftamin'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Ceftamin'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Infubinol'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Infubinol'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Ketapril'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Ketapril'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Naftisol'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Naftisol'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Placebo'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Placebo'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Propriva'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Propriva'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Ramicane'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Ramicane'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Stelasyn'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Stelasyn'])
plt.errorbar(data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df.index,data_munged_mean_metsite_by_drug_timepoint_groupby['Zoniferol'], marker="o", yerr=data_munged_reset_index_standard_error_metsite_grouped_by_drug_timepoint_df['Zoniferol'])
plt.title('Metastatic Sites')
plt.ylabel('Timepoint')
plt.xlabel('Drug')
plt.grid(True)
# Save the Figure
plt.savefig("../Pymaceuticals/matplotlib-challenge-metsites.png")
# Show the Figure
plt.show()
# -
# 
# ## Survival Rates
# Store the Count of Mice Grouped by Drug and Timepoint (W can pass any metric)
mice_count_by_drug_timepoint_groupby = drug_clinical_data.groupby(['Drug', 'Timepoint']).count()['Mouse ID']
# +
# Reset Index
reset_index_mouseid_grouped_by_drug_timepoint_df = mice_count_by_drug_timepoint_groupby.reset_index()
# Preview Reset Index
reset_index_mouseid_grouped_by_drug_timepoint_df.head()
# +
# Convert to DataFrame
mice_count_by_drug_timepoint_groupby_df = pd.DataFrame(reset_index_mouseid_grouped_by_drug_timepoint_df)
# Preview DataFrame
mice_count_by_drug_timepoint_groupby_df.columns
# -
# Minor Data Munging to Re-Format the Data Frames
data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df = mice_count_by_drug_timepoint_groupby_df.pivot(columns='Drug',index='Timepoint')['Mouse ID']
# Preview the Data Frame
data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df.head()
# +
# Store the Standard Error associated with Met. Sites Grouped by Drug and Timepoint
standard_error_mice_grouped_by_drug_timepoint = drug_clinical_data.groupby(['Drug','Timepoint']).sem()['Mouse ID']
# Convert to DataFrame
standard_error_mice_grouped_by_drug_timepoint_df = pd.DataFrame(standard_error_mice_grouped_by_drug_timepoint)
# Reset Index
reset_index_standard_error_mice_grouped_by_drug_timepoint_df = standard_error_mice_grouped_by_drug_timepoint_df.reset_index()
# Preview DataFrame
reset_index_standard_error_mice_grouped_by_drug_timepoint_df.head()
# +
# Minor Data Munging to Re-Format the Data Frames (Standard Error)
data_munged_mice_by_drug_timepoint_groupby = reset_index_standard_error_mice_grouped_by_drug_timepoint_df.pivot(columns='Drug',index='Timepoint')['Mouse ID']
# Preview that Reformatting worked
data_munged_mice_by_drug_timepoint_groupby.head()
# +
# Generate the Scatter Plot (Accounting for percentages)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Capomulin']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Capomulin']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Ceftamin']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Ceftamin']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Infubinol']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Infubinol']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Ketapril']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Ketapril']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Naftisol']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Naftisol']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Placebo']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Placebo']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Propriva']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Propriva']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Ramicane']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Ramicane']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Stelasyn']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Stelasyn']/25*100)
plt.errorbar(data_munged_mice_by_drug_timepoint_groupby.index,data_munged_reset_index_mouseid_grouped_by_drug_timepoint_df['Zoniferol']/25*100, marker="o", yerr=data_munged_mice_by_drug_timepoint_groupby['Zoniferol']/25*100)
plt.title('Mouse ID')
plt.ylabel('Timepoint')
plt.xlabel('Drug')
plt.grid(True)
# Save the Figure
plt.savefig("../Pymaceuticals/matplotlib-challenge-survival-rate.png")
plt.show()
# -
# 
# ## Summary Bar Graph
# +
# Calculate the percent changes for each drug
capomulin_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Capomulin'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Capomulin'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Capomulin'].iloc[0]*100
# Display the data to confirm
capomulin_drug_percent
# +
# Calculate the percent changes for each drug
ceftamin_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ceftamin'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ceftamin'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ceftamin'].iloc[0]*100
# Display the data to confirm
ceftamin_drug_percent
# +
# Calculate the percent changes for each drug
infubinol_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Infubinol'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Infubinol'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Infubinol'].iloc[0]*100
# Display the data to confirm
infubinol_drug_percent
# +
# Calculate the percent changes for each drug
ketapril_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ketapril'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ketapril'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ketapril'].iloc[0]*100
# Display the data to confirm
ketapril_drug_percent
# +
# Calculate the percent changes for each drug
naftisol_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Naftisol'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Naftisol'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Naftisol'].iloc[0]*100
# Display the data to confirm
naftisol_drug_percent
# +
# Calculate the percent changes for each drug
placebo_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Placebo'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Placebo'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Placebo'].iloc[0]*100
# Display the data to confirm
placebo_drug_percent
# +
# Calculate the percent changes for each drug
propriva_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Propriva'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Propriva'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Propriva'].iloc[0]*100
# Display the data to confirm
propriva_drug_percent
# +
# Calculate the percent changes for each drug
ramicane_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ramicane'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ramicane'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Ramicane'].iloc[0]*100
# Display the data to confirm
ramicane_drug_percent
# +
# Calculate the percent changes for each drug
stelasyn_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Stelasyn'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Stelasyn'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Stelasyn'].iloc[0]*100
# Display the data to confirm
stelasyn_drug_percent
# +
# Calculate the percent changes for each drug
zoniferol_drug_percent = (data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Zoniferol'].iloc[9]-data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Zoniferol'].iloc[0])/data_munged_mean_tumor_volume_by_drug_timepoint_groupby['Zoniferol'].iloc[0]*100
# Display the data to confirm
zoniferol_drug_percent
# -
# Store all Relevant Percent Changes into a Tuple
drug_percent_change_tuple = {'Capomulin': capomulin_drug_percent,'Ceftamin': ceftamin_drug_percent, 'Infubinol': infubinol_drug_percent, 'Ketapril': ketapril_drug_percent, 'Placebo': placebo_drug_percent, 'Propriva': propriva_drug_percent, 'Ramicane': ramicane_drug_percent, 'Stelasyn': stelasyn_drug_percent, 'Zoniferol': zoniferol_drug_percent}
total_drug_percent_change = pd.Series(drug_percent_change_tuple)
total_drug_percent_change
#Convert to a Data Frame
total_drug_percent_change_df =pd.DataFrame(total_drug_percent_change)
total_drug_percent_change_df
reset_index = total_drug_percent_change_df.reset_index()
# Splice the data between passing and failing drugs
drug_percentage_change = {'Capomulin': [capomulin_drug_percent],'Ceftamin': [ceftamin_drug_percent], 'Infubinol': [infubinol_drug_percent], 'Ketapril': [ketapril_drug_percent], 'Placebo': [placebo_drug_percent], 'Propriva': [propriva_drug_percent], 'Ramicane': [ramicane_drug_percent], 'Stelasyn': [stelasyn_drug_percent], 'Zoniferol': [zoniferol_drug_percent]}
drug_percentage_change
df = pd.DataFrame(drug_percentage_change)
df.head()
bin_range = [-60,0,60]
passing_failing_drugs_groups = ['Passing Drugs','Failing Drugs']
colors = {'Passing Drugs': 'blue',
'Failing Drugs': 'red'}
# +
x = pd.cut(list(drug_percent_change_tuple.values()), bin_range, labels=passing_failing_drugs_groups)
total_drug_percent_change_df["Pass or Fail"] = x
total_drug_percent_change_df.head()
# -
# 
# +
# Add labels, tick marks, etc.
plt.title('Pass or Fail')
plt.ylabel('Timepoint')
plt.xlabel('Drug')
plt.grid(True)
plt.bar(x,'Pass or Fail',facecolor=red,alpha=0.75)
plt.xticks('Drug','Timepoint')
# Save the Figure
plt.savefig("../Pymaceuticals/matplotlib-challenge-summary-bar-graph.png")
# Show the Figure
plt.show
# -
|
homework.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Mo7sY0FSBrVU"
import numpy as np
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="7ddr8Tn8D31l" outputId="b3222eb2-e7cc-46fc-d306-ad283761bb0f"
df= pd.read_csv('stock.csv')
df
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="zWml3Ro9EJ6x" outputId="aaa14c18-2e71-4075-f765-e68a62680ebb"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="-QDTzidXEL_W" outputId="d5d39740-5ec8-42d1-f446-46e288941942"
df.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="o98bexmSENzi" outputId="0208f3e8-213a-4c45-f527-c3311af4a285"
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="LUzhIl3bESJP" outputId="83dc89f3-2ec2-4199-f3e9-52d3d2bf38bc"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="4Z4864deEWk6" outputId="86c621cd-9e57-4f5c-b82b-5f27b1b27ca2"
df.columns.values
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="f4q1smm6Ec3I" outputId="cc5f29a7-2b41-472b-e5ef-66fe5d7c11b8"
df.corr()
# + colab={"base_uri": "https://localhost:8080/", "height": 447} id="kYK3bW99EhkH" outputId="f5121caf-0571-4076-a8aa-842db94391a0"
df['Date']= pd.to_datetime(df.Date, format= '%Y-%m-%d')
df.index = df['Date']
df
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="ohY2Zi-8W2bV" outputId="39e276c2-8462-45ae-e8f4-3d3dcb643b6c"
import matplotlib.pyplot as plt
plt.figure(figsize=(16,8))
plt.plot(df['Close'],label='CloseHistory')
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="nNA3CEVsH8yL" outputId="5203ce12-123a-465c-aa65-ca7c637ca4e8"
data= df.sort_index(ascending=True, axis=0)
new_data = pd.DataFrame(index=range(0,len(df)), columns=['Date','Close'])
new_data
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="6sGuW6TlIpou" outputId="608d75c8-9bee-40ee-b8e0-1866a7e53388"
for i in range(0,len(data)):
new_data['Date'][i]= data['Date'][i]
new_data['Close'][i]= data['Close'] [i]
new_data
# + id="skZB6ICod-8i"
new_data.index = new_data.Date
new_data.drop('Date', axis=1, inplace=True)
# + id="BMYxYU84X7AF"
dataset = new_data.values
train = dataset[0:987,:]
valid = dataset[987:,:]
# + id="TwRH04pQ-zSd"
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense, Dropout, LSTM
# + id="LPHbvErkZTCz"
scaler = MinMaxScaler(feature_range=(0, 1))
scaled_data = scaler.fit_transform(dataset)
# + id="l574v9nl-9yV"
x_train, y_train = [], []
for i in range(60,len(train)):
x_train.append(scaled_data[i-60:i,0])
y_train.append(scaled_data[i,0])
x_train, y_train = np.array(x_train), np.array(y_train)
x_train = np.reshape(x_train, (x_train.shape[0],x_train.shape[1],1))
# + colab={"base_uri": "https://localhost:8080/"} id="1zyX6Ozu_AFo" outputId="f51f25c1-4c7d-4cb9-ab0c-35050952851d"
model = Sequential()
model.add(LSTM(units=50, return_sequences=True, input_shape=(x_train.shape[1],1)))
model.add(LSTM(units=50))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
model.fit(x_train, y_train, epochs=1, batch_size=1, verbose=2)
# + id="xJBCo2ZD_A6R"
inputs = new_data[len(new_data) - len(valid) - 60:].values
inputs = inputs.reshape(-1,1)
inputs = scaler.transform(inputs)
# + id="ILF_1nvD_M4Z"
X_test = []
for i in range(60,inputs.shape[0]):
X_test.append(inputs[i-60:i,0])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0],X_test.shape[1],1))
closing_price = model.predict(X_test)
closing_price = scaler.inverse_transform(closing_price)
# + colab={"base_uri": "https://localhost:8080/", "height": 399} id="2m4LY0Bb_R8b" outputId="0c4c0dea-28a8-4b1f-920d-3b847235c0df"
train = new_data[:987]
valid = new_data[987:]
valid['Predictions'] = closing_price
plt.plot(train['Close'])
plt.plot(valid[['Close','Predictions']])
|
Machine Learning Experiment 6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # OT for image color adaptation with mapping estimation
#
# OT for domain adaptation with image color adaptation [6] with mapping
# estimation [8].
#
# [6] <NAME>., <NAME>., <NAME>., & <NAME>. (2014). Regularized
# discrete optimal transport. SIAM Journal on Imaging Sciences, 7(3), 1853-1882.
#
# [8] <NAME>, <NAME>, <NAME>, <NAME>, "Mapping estimation for
# discrete optimal transport", Neural Information Processing Systems (NIPS), 2016.
#
# +
# Authors: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: MIT License
# sphinx_gallery_thumbnail_number = 3
import numpy as np
import matplotlib.pylab as pl
import ot
r = np.random.RandomState(42)
def im2mat(img):
"""Converts and image to matrix (one pixel per line)"""
return img.reshape((img.shape[0] * img.shape[1], img.shape[2]))
def mat2im(X, shape):
"""Converts back a matrix to an image"""
return X.reshape(shape)
def minmax(img):
return np.clip(img, 0, 1)
# -
# ## Generate data
#
#
# +
# Loading images
I1 = pl.imread('../../data/ocean_day.jpg').astype(np.float64) / 256
I2 = pl.imread('../../data/ocean_sunset.jpg').astype(np.float64) / 256
X1 = im2mat(I1)
X2 = im2mat(I2)
# training samples
nb = 1000
idx1 = r.randint(X1.shape[0], size=(nb,))
idx2 = r.randint(X2.shape[0], size=(nb,))
Xs = X1[idx1, :]
Xt = X2[idx2, :]
# -
# ## Domain adaptation for pixel distribution transfer
#
#
# +
# EMDTransport
ot_emd = ot.da.EMDTransport()
ot_emd.fit(Xs=Xs, Xt=Xt)
transp_Xs_emd = ot_emd.transform(Xs=X1)
Image_emd = minmax(mat2im(transp_Xs_emd, I1.shape))
# SinkhornTransport
ot_sinkhorn = ot.da.SinkhornTransport(reg_e=1e-1)
ot_sinkhorn.fit(Xs=Xs, Xt=Xt)
transp_Xs_sinkhorn = ot_sinkhorn.transform(Xs=X1)
Image_sinkhorn = minmax(mat2im(transp_Xs_sinkhorn, I1.shape))
ot_mapping_linear = ot.da.MappingTransport(
mu=1e0, eta=1e-8, bias=True, max_iter=20, verbose=True)
ot_mapping_linear.fit(Xs=Xs, Xt=Xt)
X1tl = ot_mapping_linear.transform(Xs=X1)
Image_mapping_linear = minmax(mat2im(X1tl, I1.shape))
ot_mapping_gaussian = ot.da.MappingTransport(
mu=1e0, eta=1e-2, sigma=1, bias=False, max_iter=10, verbose=True)
ot_mapping_gaussian.fit(Xs=Xs, Xt=Xt)
X1tn = ot_mapping_gaussian.transform(Xs=X1) # use the estimated mapping
Image_mapping_gaussian = minmax(mat2im(X1tn, I1.shape))
# -
# ## Plot original images
#
#
# +
pl.figure(1, figsize=(6.4, 3))
pl.subplot(1, 2, 1)
pl.imshow(I1)
pl.axis('off')
pl.title('Image 1')
pl.subplot(1, 2, 2)
pl.imshow(I2)
pl.axis('off')
pl.title('Image 2')
pl.tight_layout()
# -
# ## Plot pixel values distribution
#
#
# +
pl.figure(2, figsize=(6.4, 5))
pl.subplot(1, 2, 1)
pl.scatter(Xs[:, 0], Xs[:, 2], c=Xs)
pl.axis([0, 1, 0, 1])
pl.xlabel('Red')
pl.ylabel('Blue')
pl.title('Image 1')
pl.subplot(1, 2, 2)
pl.scatter(Xt[:, 0], Xt[:, 2], c=Xt)
pl.axis([0, 1, 0, 1])
pl.xlabel('Red')
pl.ylabel('Blue')
pl.title('Image 2')
pl.tight_layout()
# -
# ## Plot transformed images
#
#
# +
pl.figure(2, figsize=(10, 5))
pl.subplot(2, 3, 1)
pl.imshow(I1)
pl.axis('off')
pl.title('Im. 1')
pl.subplot(2, 3, 4)
pl.imshow(I2)
pl.axis('off')
pl.title('Im. 2')
pl.subplot(2, 3, 2)
pl.imshow(Image_emd)
pl.axis('off')
pl.title('EmdTransport')
pl.subplot(2, 3, 5)
pl.imshow(Image_sinkhorn)
pl.axis('off')
pl.title('SinkhornTransport')
pl.subplot(2, 3, 3)
pl.imshow(Image_mapping_linear)
pl.axis('off')
pl.title('MappingTransport (linear)')
pl.subplot(2, 3, 6)
pl.imshow(Image_mapping_gaussian)
pl.axis('off')
pl.title('MappingTransport (gaussian)')
pl.tight_layout()
pl.show()
|
_downloads/acdb5c8d9a410d04b44379453a1620f0/plot_otda_mapping_colors_images.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
from itertools import cycle
import numpy as np
import pandas as pd
# setup plot details
colors = cycle(["navy", "turquoise", "darkorange", "cornflowerblue", "teal"])
from sklearn.multiclass import OneVsRestClassifier
from sklearn.preprocessing import label_binarize
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import LinearSVC
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import average_precision_score
from sklearn.metrics import (precision_recall_curve,PrecisionRecallDisplay)
from sklearn.svm import SVC
# +
_, ax = plt.subplots(figsize=(7, 8))
f_scores = np.linspace(0.2, 0.8, num=4)
lines, labels = [], []
for f_score in f_scores:
x = np.linspace(0.01, 1)
y = f_score * x / (2 * x - f_score)
(l,) = plt.plot(x[y >= 0], y[y >= 0], color="gray", alpha=0.2)
plt.annotate("f1={0:0.1f}".format(f_score), xy=(0.9, y[45] + 0.02))
display = PrecisionRecallDisplay(
recall=recall["micro"],
precision=precision["micro"],
average_precision=average_precision["micro"],
)
display.plot(ax=ax, name="Micro-average precision-recall", color="gold")
for i in range(n_classes):
plt.plot(fpr[i], tpr[i], label='ROC curve of class {0} (area = {1:0.2f})'.format(i, roc_auc[i]))
display.plot(ax=ax, name=f"Precision-recall for class {i}", color=color)
# add the legend for the iso-f1 curves
handles, labels = display.ax_.get_legend_handles_labels()
handles.extend([l])
labels.extend(["iso-f1 curves"])
# set the legend and the axes
ax.set_xlim([0.0, 1.0])
ax.set_ylim([0.0, 1.05])
ax.legend(handles=handles, labels=labels, loc="best")
ax.set_title("Extension of Precision-Recall curve to multi-class")
plt.show()
# -
# ### 1)Data Set Validation
# +
# roc curve and auc
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from matplotlib import pyplot
# generate 2 class dataset
X, y = make_classification(n_samples=1000, n_classes=2, random_state=1)
# split into train/test sets
trainX, testX, trainy, testy = train_test_split(X, y, test_size=0.5, random_state=2)
# generate a no skill prediction (majority class)
ns_probs = [0 for _ in range(len(testy))]
# -
# ### Logistic Regression Modeling
# +
# fit a model
model = LogisticRegression(solver='lbfgs')
model.fit(trainX, trainy)
# predict probabilities
lr_probs = model.predict_proba(testX)
# keep probabilities for the positive outcome only
lr_probs = lr_probs[:, 1]
# -
# ### Calculating ROC Accuracy Score
# +
# calculate scores
ns_auc = roc_auc_score(testy, ns_probs)
lr_auc = roc_auc_score(testy, lr_probs)
# summarize scores
print('No Skill: ROC AUC=%.3f' % (ns_auc))
print('Logistic: ROC AUC=%.3f' % (lr_auc))
# -
# ### Plotting ROC
# +
# calculate roc curves
ns_fpr, ns_tpr, _ = roc_curve(testy, ns_probs)
lr_fpr, lr_tpr, _ = roc_curve(testy, lr_probs)
# plot the roc curve for the model
pyplot.plot(ns_fpr, ns_tpr, linestyle='--', label='No Skill')
pyplot.plot(lr_fpr, lr_tpr, marker='.', label='Logistic')
# axis labels
pyplot.xlabel('False Positive Rate')
pyplot.ylabel('True Positive Rate')
# show the legend
pyplot.legend()
# show the plot
pyplot.show()
# -
# ## 2) Library Import
# +
# precision-recall curve and f1
from sklearn.datasets import make_classification
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import f1_score
from sklearn.metrics import auc
from matplotlib import pyplot
# generate 2 class dataset
X, y = make_classification(n_samples=1000, n_classes=2, random_state=1)
# split into train/test sets
trainX, testX, trainy, testy = train_test_split(X, y, test_size=0.5, random_state=2)
# -
# ## Logistic Regression
# +
# fit a model
model = LogisticRegression(solver='lbfgs')
model.fit(trainX, trainy)
# predict probabilities
lr_probs = model.predict_proba(testX)
# keep probabilities for the positive outcome only
lr_probs = lr_probs[:, 1]
# -
# ## F1 Scores
# predict class values
yhat = model.predict(testX)
lr_precision, lr_recall, _ = precision_recall_curve(testy, lr_probs)
lr_f1, lr_auc = f1_score(testy, yhat), auc(lr_recall, lr_precision)
# summarize scores
print('Logistic: f1=%.3f auc=%.3f' % (lr_f1, lr_auc))
# ### PLotting the Model
# +
# plot the precision-recall curves
no_skill = len(testy[testy==1]) / len(testy)
pyplot.plot([0, 1], [no_skill, no_skill], linestyle='--', label='No Skill')
pyplot.plot(lr_recall, lr_precision, marker='.', label='Logistic')
# axis labels
pyplot.xlabel('Recall')
pyplot.ylabel('Precision')
# show the legend
pyplot.legend()
# show the plot
pyplot.show()
# -
|
Supervised Learning/Model Validation/Precision-Recall With Scikit Learn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DAT210x - Programming with Python for DS
# ## Module5- Lab6
# +
import random, math
import pandas as pd
import numpy as np
import scipy.io
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
matplotlib.style.use('ggplot') # Look Pretty
# Leave this alone until indicated:
Test_PCA = False
# -
# ### A Convenience Function
# This method is for your visualization convenience only. You aren't expected to know how to put this together yourself, although you should be able to follow the code by now:
def Plot2DBoundary(DTrain, LTrain, DTest, LTest):
# The dots are training samples (img not drawn), and the pics are testing samples (images drawn)
# Play around with the K values. This is very controlled dataset so it should be able to get perfect classification on testing entries
# Play with the K for isomap, play with the K for neighbors.
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_title('Transformed Boundary, Image Space -> 2D')
padding = 0.1 # Zoom out
resolution = 1 # Don't get too detailed; smaller values (finer rez) will take longer to compute
colors = ['blue','green','orange','red']
# ------
# Calculate the boundaries of the mesh grid. The mesh grid is
# a standard grid (think graph paper), where each point will be
# sent to the classifier (KNeighbors) to predict what class it
# belongs to. This is why KNeighbors has to be trained against
# 2D data, so we can produce this countour. Once we have the
# label for each point on the grid, we can color it appropriately
# and plot it.
x_min, x_max = DTrain[:, 0].min(), DTrain[:, 0].max()
y_min, y_max = DTrain[:, 1].min(), DTrain[:, 1].max()
x_range = x_max - x_min
y_range = y_max - y_min
x_min -= x_range * padding
y_min -= y_range * padding
x_max += x_range * padding
y_max += y_range * padding
# Using the boundaries, actually make the 2D Grid Matrix:
xx, yy = np.meshgrid(np.arange(x_min, x_max, resolution),
np.arange(y_min, y_max, resolution))
# What class does the classifier say about each spot on the chart?
# The values stored in the matrix are the predictions of the model
# at said location:
Z = model.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
# Plot the mesh grid as a filled contour plot:
plt.contourf(xx, yy, Z, cmap=plt.cm.terrain, z=-100)
# ------
# When plotting the testing images, used to validate if the algorithm
# is functioning correctly, size them as 5% of the overall chart size
x_size = x_range * 0.05
y_size = y_range * 0.05
# First, plot the images in your TEST dataset
img_num = 0
for index in LTest.index:
# DTest is a regular NDArray, so you'll iterate over that 1 at a time.
x0, y0 = DTest[img_num,0]-x_size/2., DTest[img_num,1]-y_size/2.
x1, y1 = DTest[img_num,0]+x_size/2., DTest[img_num,1]+y_size/2.
# DTest = our images isomap-transformed into 2D. But we still want
# to plot the original image, so we look to the original, untouched
# dataset (at index) to get the pixels:
img = df.iloc[index,:].reshape(num_pixels, num_pixels)
ax.imshow(img,
aspect='auto',
cmap=plt.cm.gray,
interpolation='nearest',
zorder=100000,
extent=(x0, x1, y0, y1),
alpha=0.8)
img_num += 1
# Plot your TRAINING points as well... as points rather than as images
for label in range(len(np.unique(LTrain))):
indices = np.where(LTrain == label)
ax.scatter(DTrain[indices, 0], DTrain[indices, 1], c=colors[label], alpha=0.8, marker='o')
# Plot
plt.show()
# ### The Assignment
# Use the same code from Module4/assignment4.ipynb to load up the `face_data.mat` file into a dataframe called `df`. Be sure to calculate the `num_pixels` value, and to rotate the images to being right-side-up instead of sideways. This was demonstrated in the [Lab Assignment 4](https://github.com/authman/DAT210x/blob/master/Module4/assignment4.ipynb) code.
mat = scipy.io.loadmat('Datasets/face_data.mat')
df = pd.DataFrame(mat['images']).T
num_images, num_pixels = df.shape
num_pixels = int(math.sqrt(num_pixels))
for i in range(num_images):
df.loc[i,:] = df.loc[i,:].reshape(num_pixels, num_pixels).T.reshape(-1)
# Load up your face_labels dataset. It only has a single column, and you're only interested in that single column. You will have to slice the column out so that you have access to it as a "Series" rather than as a "Dataframe". This was discussed in the the "Slicin'" lecture of the "Manipulating Data" reading on the course website. Use an appropriate indexer to take care of that. Be sure to print out the labels and compare what you see to the raw `face_labels.csv` so you know you loaded it correctly.
label = pd.read_csv('Datasets/face_labels.csv')
label = label.iloc[:, 0]
# Do `train_test_split`. Use the same code as on the EdX platform in the reading material, but set the random_state=7 for reproducibility, and the test_size to 0.15 (150%). Your labels are actually passed in as a series (instead of as an NDArray) so that you can access their underlying indices later on. This is necessary so you can find your samples in the original dataframe. The convenience methods we've written for you that handle drawing expect this, so that they can plot your testing data as images rather than as points:
# +
# .. your code here ..
# -
# ### Dimensionality Reduction
# +
if Test_PCA:
# INFO: PCA is used *before* KNeighbors to simplify your high dimensionality
# image samples down to just 2 principal components! A lot of information
# (variance) is lost during the process, as I'm sure you can imagine. But
# you have to drop the dimension down to two, otherwise you wouldn't be able
# to visualize a 2D decision surface / boundary. In the wild, you'd probably
# leave in a lot more dimensions, which is better for higher accuracy, but
# worse for visualizing the decision boundary;
#
# Your model should only be trained (fit) against the training data (data_train)
# Once you've done this, you need use the model to transform both data_train
# and data_test from their original high-D image feature space, down to 2D
# TODO: Implement PCA here. ONLY train against your training data, but
# transform both your training + test data, storing the results back into
# data_train, and data_test.
# .. your code here ..
else:
# INFO: Isomap is used *before* KNeighbors to simplify your high dimensionality
# image samples down to just 2 components! A lot of information has been is
# lost during the process, as I'm sure you can imagine. But if you have
# non-linear data that can be represented on a 2D manifold, you probably will
# be left with a far superior dataset to use for classification. Plus by
# having the images in 2D space, you can plot them as well as visualize a 2D
# decision surface / boundary. In the wild, you'd probably leave in a lot more
# dimensions, which is better for higher accuracy, but worse for visualizing the
# decision boundary;
# Your model should only be trained (fit) against the training data (data_train)
# Once you've done this, you need use the model to transform both data_train
# and data_test from their original high-D image feature space, down to 2D
# TODO: Implement Isomap here. ONLY train against your training data, but
# transform both your training + test data, storing the results back into
# data_train, and data_test.
# .. your code here ..
# -
# Implement `KNeighborsClassifier` here. You can use any K value from 1 through 20, so play around with it and attempt to get good accuracy. Fit the classifier against your training data and labels.
# +
# .. your code here ..
# -
# Calculate and display the accuracy of the testing set (data_test and label_test):
# +
# .. your code here ..
# -
# Let's chart the combined decision boundary, the training data as 2D plots, and the testing data as small images so we can visually validate performance:
Plot2DBoundary(data_train, label_train, data_test, label_test)
# After submitting your answers, experiment with using using PCA instead of ISOMap. Are the results what you expected? Also try tinkering around with the test/train split percentage from 10-20%. Notice anything?
# +
# .. your code changes above ..
# -
|
Module5/.ipynb_checkpoints/Module5 - Lab6-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# ## Traininig of the Inclusive classifier
#
# **4.3 Inclusive classifier** This trains the Inclusive classifier, a combination of the Particle-sequence classifier with the High Level Features classifier.
#
# To run this notebook we used the following configuration:
# * *Software stack*: Spark 2.4.3, analytics-zoo 0.5.1
# * *Platform*: CentOS 7, Python 3.6
# * *Spark cluster*: Analytix
# + deletable=true editable=true
# pip install pyspark or use your favorite way to set Spark Home, here we use findspark
import findspark
findspark.init('/home/luca/Spark/spark-2.4.3-bin-hadoop2.7') #set path to SPARK_HOME
# + deletable=true editable=true
# Configure according to your environment
from pyspark.sql import SparkSession
pyspark_python = "<path to python>/bin/python"
analytics_zoo_jar = "<path>/analytics-zoo-bigdl_0.8.0-spark_2.4.3-0.5.1-jar-with-dependencies.jar"
analytics_zoo_python_api = "<path>/analytics-zoo-bigdl_0.8.0-spark_2.4.3-0.5.1-python-api.zip"
spark = SparkSession.builder \
.appName("4.3-Training-InclusiveClassifier") \
.master("yarn") \
.config("spark.driver.memory","8g") \
.config("spark.executor.memory","14g") \
.config("spark.executor.cores","6") \
.config("spark.executor.instances","70") \
.config("spark.dynamicAllocation.enabled","false") \
.config("spark.shuffle.reduceLocality.enabled","false") \
.config("spark.shuffle.blockTransferService","nio") \
.config("spark.scheduler.minRegisteredResourcesRatio","1.0") \
.config("spark.speculation","false") \
.config("spark.eventLog.enabled","false") \
.config("spark.jars",analytics_zoo_jar) \
.config("spark.submit.pyFiles",analytics_zoo_python_api) \
.config("spark.pyspark.python",pyspark_python) \
.getOrCreate()
# + deletable=true editable=true
# Check if Spark Session has been created correctly
spark
# + [markdown] deletable=true editable=true
# ## Load train and test dataset
# + deletable=true editable=true
PATH = "hdfs://analytix/Training/Spark/TopologyClassifier/"
trainDF = spark.read.format('parquet')\
.load(PATH + 'trainUndersampled.parquet')\
.select(['GRU_input', 'HLF_input', 'encoded_label'])
testDF = spark.read.format('parquet')\
.load(PATH + 'testUndersampled.parquet')\
.select(['GRU_input', 'HLF_input', 'encoded_label'])
# + deletable=true editable=true
trainDF.printSchema()
# + deletable=true editable=true
print("Number of events in the test dataset:", testDF.count())
print("Number of events in the training dataset:", trainDF.count())
# + [markdown] deletable=true editable=true
# ## Create the model
# + deletable=true editable=true
# Init analytics zoo
from zoo.common.nncontext import *
sc = init_nncontext("Inclusive Classifier")
# + deletable=true editable=true
from zoo.pipeline.api.keras.optimizers import Adam
from zoo.pipeline.api.keras.models import Sequential
from zoo.pipeline.api.keras.layers.core import *
from zoo.pipeline.api.keras.layers.recurrent import GRU
from zoo.pipeline.api.keras.engine.topology import Merge
## GRU branch
gruBranch = Sequential() \
.add(Masking(0.0, input_shape=(801, 19))) \
.add(GRU(
output_dim=50,
activation='tanh'
)) \
.add(Dropout(0.2)) \
## HLF branch
hlfBranch = Sequential() \
.add(Dropout(0.2, input_shape=(14,)))
## Concatenate the branches
branches = Merge(layers=[gruBranch, hlfBranch], mode='concat')
## Create the model
model = Sequential() \
.add(branches) \
.add(Dense(25, activation='relu')) \
.add(Dense(3, activation='softmax'))
# + [markdown] deletable=true editable=true
# ## Create train and validation RDD
#
# We need to create an RDD of `Sample`, a tuple of the form (`features`, `label`). The two elements of this touple should be `numpy arrays`.
# + deletable=true editable=true
from bigdl.util.common import Sample
import numpy as np
trainRDD = trainDF.rdd.map(lambda row: Sample.from_ndarray(
[np.array(row.GRU_input), np.array(row.HLF_input)],
np.array(row.encoded_label)
))
testRDD = testDF.rdd.map(lambda row: Sample.from_ndarray(
[np.array(row.GRU_input), np.array(row.HLF_input)],
np.array(row.encoded_label)
))
# + deletable=true editable=true
# Let's have a look at one element of trainRDD
trainRDD.take(1)
# + [markdown] deletable=true editable=true
# We can see that `Sample.feature` is now composed by the list of 801 particles with 19 features each (`shape=[801 19]`) plus the HLF (`shape=[14]`) and the encoded label (`shape=[3]`).
# + [markdown] deletable=true editable=true
# ## Optimizer setup and training
# + deletable=true editable=true
# Set of hyperparameters
numEpochs = 50
# The batch used by BDL must be a multiple of numExecutors * executorCores
# Because data will be equally distibuted inside each executor
workerBatch = 32
numExecutors = int(spark.conf.get('spark.executor.instances'))
executorCores = int(spark.conf.get('spark.executor.cores'))
BDLbatch = workerBatch * numExecutors * executorCores
# + deletable=true editable=true
from bigdl.optim.optimizer import *
from bigdl.nn.criterion import CategoricalCrossEntropy
# optim_method = Adam(learningrate=0.002, learningrate_decay=0.0002, epsilon=9e-8)
optim_method = Adam()
model.compile(optimizer=optim_method, loss=CategoricalCrossEntropy(), metrics=[Loss(CategoricalCrossEntropy())])
# + [markdown] deletable=true editable=true
# Let's define a directory to store logs (i.e. train and validation losses) and save models
# + deletable=true editable=true
# name of our application
appName = "InclusiveClassifier"
# Change it!
logDir = "/tmp"
# Check if there is already an application with the same name
# and remove it, otherwise logs will be appended to that app
import os
try:
os.system('rm -rf '+logDir+'/'+appName)
except:
pass
print("Saving logs to {}".format(logDir+'/'+appName))
# + deletable=true editable=true
model.set_tensorboard(logDir, appName)
# + [markdown] deletable=true editable=true
# We are now ready to launch the training.
#
# Warning relevant for CERN SWAN service users: During the training it would be better to shutdown the Toggle Spark Monitoring Display because each iteration is seen as a Spark job, therefore the toggle will try to display everything causing problem to the browser.
# + deletable=true editable=true
# %time model.fit(x=trainRDD, batch_size=BDLbatch, nb_epoch=numEpochs, validation_data=testRDD, distributed=True)
# + [markdown] deletable=true editable=true
# ## Plot loss
# + deletable=true editable=true
# %matplotlib notebook
import matplotlib.pyplot as plt
plt.style.use('seaborn-darkgrid')
trainSummary = TrainSummary(log_dir=logDir,app_name=appName)
loss = np.array(trainSummary.read_scalar("Loss"))
valSummary = ValidationSummary(log_dir=logDir,app_name=appName)
val_loss = np.array(valSummary.read_scalar("Loss"))
plt.plot(loss[:,0], loss[:,1], label="Training loss")
plt.plot(val_loss[:,0], val_loss[:,1], label="Validation loss", color='crimson', alpha=0.8)
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.legend()
plt.title("Inclusive classifier loss")
plt.show()
# + [markdown] deletable=true editable=true
# ## Save the model
# + deletable=true editable=true
modelDir = logDir + '/models'
model.saveModel(
modelPath = modelDir + '/' + appName + '.bigdl',
weightPath = modelDir + '/' + appName + '.bin',
over_write = True
)
# + [markdown] deletable=true editable=true
# It is possible to load the model in the following way:
# ```Python
# model = Model.loadModel(modelPath=modelPath+'.bigdl', weightPath=modelPath+'.bin')
# ```
# + [markdown] deletable=true editable=true
# ## Prediction
# + deletable=true editable=true
pred = model.predict(testRDD)
# + deletable=true editable=true
y_pred = np.asarray(pred.collect())
y_true = np.asarray(testDF.select('encoded_label').rdd\
.map(lambda row: np.asarray(row.encoded_label)).collect())
# + deletable=true editable=true
from sklearn.metrics import roc_curve, auc
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(3):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# + deletable=true editable=true
plt.figure()
plt.plot(fpr[0], tpr[0], lw=2,
label='Inclusive classifier (AUC) = %0.4f' % roc_auc[0])
plt.plot([0, 1], [0, 1], linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('Background Contamination (FPR)')
plt.ylabel('Signal Efficiency (TPR)')
plt.title('$tt$ selector')
plt.legend(loc="lower right")
plt.show()
# + [markdown] deletable=true editable=true
# ## Confusion Matrix
# + deletable=true editable=true
from sklearn.metrics import accuracy_score
print('Accuracy of the Inclusive classifier: {:.4f}'.format(
accuracy_score(np.argmax(y_true, axis=1),np.argmax(y_pred, axis=1))))
# + deletable=true editable=true
import seaborn as sns
from sklearn.metrics import confusion_matrix
labels_name = ['qcd', 'tt', 'wjets']
labels = [0,1,2]
cm = confusion_matrix(np.argmax(y_true, axis=1), np.argmax(y_pred, axis=1), labels=labels)
## Normalize CM
cm = cm / cm.astype(np.float).sum(axis=1)
fig, ax = plt.subplots()
ax = sns.heatmap(cm, annot=True, fmt='g')
ax.xaxis.set_ticklabels(labels_name)
ax.yaxis.set_ticklabels(labels_name)
plt.title('Confusion matrix - Inclusive classifier')
plt.xlabel('True labels')
plt.ylabel('Predicted labels')
plt.show()
# + deletable=true editable=true
spark.stop()
# + deletable=true editable=true
|
Training_BigDL_Zoo/4.3-Training-InclusiveClassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="H12ohaovEpt7"
# # Improving Remote Poverty Detection with Multiview Learning — TESTING
# This notebook explores the data and tests the methods using only a subset of the data (Ethiopia) to more efficeintly narrow the search space for methods and hyperparameters.
# ## SETUP
# + colab={"base_uri": "https://localhost:8080/"} id="2rvMNx26EGaB" executionInfo={"status": "ok", "timestamp": 1622583268553, "user_tz": 240, "elapsed": 29863, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="346c9ce2-ce96-4ca0-b74f-f9ac04454c08"
from google.colab import files, drive
drive.mount('/content/drive')
# ! pip install geoio
# + colab={"base_uri": "https://localhost:8080/"} id="M8FvxJqd6pM0" executionInfo={"status": "ok", "timestamp": 1622583320942, "user_tz": 240, "elapsed": 2543, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="ae6f8fbf-3413-46ac-c638-ba5d0f564c63"
# !ls drive/MyDrive/detecting-poverty/data/landsat
# + id="8s3itW6xGXAA" executionInfo={"status": "ok", "timestamp": 1622583322873, "user_tz": 240, "elapsed": 1934, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}}
# ! cp drive/MyDrive/detecting-poverty/modules/* ./
# + id="zYciIYnSGceI" executionInfo={"status": "ok", "timestamp": 1622583354974, "user_tz": 240, "elapsed": 5246, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}}
import numpy as np
import torch
import pandas as pd
from data_loaders import LandsatViirs, LandsatTransform, ViirsTransform
from conv_ved import ConvVED, ResnetVAE
import utils
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Lasso
import geoio
import matplotlib.pyplot as plt
# %matplotlib inline
from utils import create_space
import os
from PIL import Image
import torchvision.transforms.functional as TF
# to handle truncated images
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
import random
torch.random.manual_seed(31220)
random.seed(31220)
# for better traceback with CUDA errors
os.environ['CUDA_LAUNCH_BLOCKING'] = "1"
# + [markdown] id="3KLMgM23mKRI"
# ## The DATA
# ### Survey and Reference Data
# + colab={"base_uri": "https://localhost:8080/", "height": 159} id="sDD7RK-vHy2x" executionInfo={"status": "ok", "timestamp": 1622583357973, "user_tz": 240, "elapsed": 1287, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="1ed3e0f2-2357-43c8-b787-64beb67e7493"
full_reference_data = pd.read_csv('drive/MyDrive/detecting-poverty/data/image_download_actual.csv')
print(full_reference_data.shape)
full_reference_data.head(3)
# + id="Q9SuQO4IFGsa" executionInfo={"status": "ok", "timestamp": 1622583360611, "user_tz": 240, "elapsed": 1843, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}}
NIGHTLIGHTS_DIRS = ['drive/MyDrive/detecting-poverty/data/viirs/viirs_2015_' + tif_name for tif_name in ['00N060W.tif', '75N060W.tif']]
viirs_tifs = [geoio.GeoImage(ndir) for ndir in NIGHTLIGHTS_DIRS]
# + [markdown] id="1RRul0HzmcFk"
# ## Convolutional Variation Resnet-Encoder Convolutional-Decoder
# ### Data Pipeline
# + id="w4OfAPIgMU8t" executionInfo={"status": "ok", "timestamp": 1622583390478, "user_tz": 240, "elapsed": 24886, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}}
# create dataset objects
ref_features = ['image_lat', 'image_lon', 'image_name', 'country']
target = 'cons_pc'
traindev_ref_data, test_ref_data, Ytraindev, Ytest = train_test_split(
full_reference_data[ref_features],
full_reference_data[target],
train_size=0.9
)
train_ref_data, dev_ref_data, Ytrain, Ydev = train_test_split(
traindev_ref_data,
Ytraindev,
train_size=0.9
)
viirs_transform = ViirsTransform(viirs_tifs)
landsat_transform = LandsatTransform('drive/MyDrive/detecting-poverty/data/landsat', width=224, height=224)
# training_loader = LandsatViirs(
# df=train_ref_data,
# viirs_transform=viirs_transform,
# landsat_transform=landsat_transform
# )
# dev_loader = LandsatViirs(
# df=dev_ref_data,
# viirs_transform=viirs_transform,
# landsat_transform=landsat_transform
# )
# + [markdown] id="GBxcg2-tmjZU"
# ### Model Testing
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["a39adc121fe74087b075a218fa6e9850", "5475895c4ff445d3b11af2f3d020059a", "3e7f977c90164c0da097c4fcaa2596c1", "9ed4057d8bfe49a781c1ff93b463c3c1", "85b3bf2fe1a3480385db64261b675d51", "1cde0f21235341529792a19dd2c4a568", "27fb835fa4f24dae856ccad05fe6a8e2", "c3082fd1e13c40d992d518f74c4ed2f5"]} id="oyevRRSxa7TG" executionInfo={"status": "ok", "timestamp": 1622584230821, "user_tz": 240, "elapsed": 835885, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="53126999-5ac5-42e4-f2d7-713921981a96"
# TESTING MODULES WITH ONLY ETHIOPIA DATA
# init model
conv_ved = ConvVED(
n_components=64,
net=ResnetVAE,
image_in_channels=3,
image_out_channels=1,
lr=2e-3,
batch_size=32,
kkl=1,
kv=1,
path='resnet_ved.pth',
# cuda=False
)
# data loaders
eth_training_loader = LandsatViirs(
df=train_ref_data[train_ref_data.country == 'eth'],
viirs_transform=viirs_transform,
landsat_transform=landsat_transform
)
eth_dev_loader = LandsatViirs(
df=dev_ref_data[dev_ref_data.country == 'eth'],
viirs_transform=viirs_transform,
landsat_transform=landsat_transform
)
# train
conv_ved.fit(
eth_training_loader, Xd=eth_dev_loader, epochs=5
)
# + [markdown] id="gNLGvhiSacZA"
# ### Consumption Prediction
# + colab={"base_uri": "https://localhost:8080/"} id="PYBGnaJwjboN" executionInfo={"status": "ok", "timestamp": 1622584323075, "user_tz": 240, "elapsed": 42194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="6c686b19-f018-4638-f222-4f54484143de"
eth_train_features = conv_ved.transform(eth_training_loader)
eth_dev_features = conv_ved.transform(eth_dev_loader)
# + colab={"base_uri": "https://localhost:8080/"} id="FlW-aBOWkgF5" executionInfo={"status": "ok", "timestamp": 1622584480706, "user_tz": 240, "elapsed": 210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="0fb81291-285d-43c8-c8d8-17ee72c66be4"
eth_dev_features
# + colab={"base_uri": "https://localhost:8080/"} id="7gmH__-4NYEF" executionInfo={"status": "ok", "timestamp": 1622584599723, "user_tz": 240, "elapsed": 1264, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="7ed13cfd-a598-4932-943e-6ad4f5a7478e"
from predictors import elastic_net, logistic
results, downstream_model = elastic_net(
Xtrain=eth_train_features,
Xdev=eth_dev_features,
Ytrain=Ytrain[train_ref_data.country == 'eth'],
Ydev=Ydev[dev_ref_data.country == 'eth'],
verbose=True,
# scoring='f1'
)
# + [markdown] id="m-q2hLoTrEpJ"
# ## Long Training
# + colab={"base_uri": "https://localhost:8080/"} id="lkZkECCaq6FA" executionInfo={"status": "ok", "timestamp": 1622587054864, "user_tz": 240, "elapsed": 633356, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="11e737a8-e776-4f4f-fd76-c2aaa51a0aa5"
# TESTING MODULES WITH ONLY ETHIOPIA DATA
# init model
conv_ved = ConvVED(
n_components=64,
net=ResnetVAE,
image_in_channels=3,
image_out_channels=1,
lr=2e-3,
batch_size=32,
kkl=1,
kv=1,
path='resnet_ved.pth',
# cuda=False
)
# train
conv_ved.fit(
eth_training_loader, Xd=eth_dev_loader, epochs=20
)
# + colab={"base_uri": "https://localhost:8080/"} id="dNPhWgEhuZQn" executionInfo={"status": "ok", "timestamp": 1622587122648, "user_tz": 240, "elapsed": 41908, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="f02b1f89-b31e-47da-d2f1-b421688ececf"
eth_train_features = conv_ved.transform(eth_training_loader)
eth_dev_features = conv_ved.transform(eth_dev_loader)
# + colab={"base_uri": "https://localhost:8080/"} id="a9EXV6h_uaGu" executionInfo={"status": "ok", "timestamp": 1622587149372, "user_tz": 240, "elapsed": 1179, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiGHdh6BtwcVBOlBTfpiwHfHij6pb9p8XlIjneTpg=s64", "userId": "11790179745156165663"}} outputId="0dd048bf-bd7d-447f-d94c-b65a7bf956fa"
results, downstream_model = elastic_net(
Xtrain=eth_train_features,
Xdev=eth_dev_features,
Ytrain=Ytrain[train_ref_data.country == 'eth'],
Ydev=Ydev[dev_ref_data.country == 'eth'],
verbose=True,
# scoring='f1'
)
|
notebooks/resnetVAE-subset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Lab 3: Quantum Phase Estimation
# In this lab, you will implement a quantum program to determine the global phase applied by a unitary operator on its eigenstate. In order to do this, you will write `Qiskit` code for the quantum phase estimation algorithm following the material presented in lectures 7 to 9.
#
# You might find the following chapters of the Qiskit Textbook useful:
# - **Quantum phase estimation**: https://qiskit.org/textbook/ch-algorithms/quantum-phase-estimation.html
# - **Quantum Fourier transform**: https://qiskit.org/textbook/ch-algorithms/quantum-fourier-transform.html
#
# Remember, to run a cell in Jupyter notebooks, you press `Shift` + `Return/Enter` on your keyboard.
# ### Installing necessary packages
# Before we begin, you will need to install some prerequisites into your environment. Run the cell below to complete these installations. At the end, the cell outputs will be cleared.
# +
# !pip install -U -r resources/requirements.txt
from IPython.display import clear_output
clear_output()
# -
# # Review of Quantum Phase Estimation
# 
# You might recall from lectures 7 to 9 that the goal of quantum phase estimation is to determine the phase $\theta$ applied by a unitary operator $U$ on its eigenstate $\vert\psi\rangle$ such that
#
# $$U\vert\psi\rangle = e^{2\pi i\theta}\vert\psi\rangle$$
#
# This is done in four main steps.
#
# 1. First, we begin by creating a superposition of all $2^n$ computational basis states on the $n$ measurement qubits by applying a Hadamard ($H$) gate on each qubit starting off in the state $\vert0\rangle^{\otimes n}$. We also initialize the target qubits (in this case only one) into an eigenstate $\vert\psi\rangle$ of the unitary operator $U$. Here, the exponent $\otimes n$ means that we have a tensor product of the states of $n$ qubits.
#
# 2. Second, we apply the unitary operator $U$ with various powers onto the target qubits (in this case only one) by controlling it with each of the different measurement qubits. The schematic above shows the ordering and respective powers.
#
# 3. Third, we apply an inverse quantum Fourier transform on the $n$ measurement qubits.
#
# 4. Finally, we measure the $n$ qubits and read out $2^n\theta$.
#
#
# # Graded Exercise 1: Implementing Quantum Phase Estimation
#
# In this lab, we will implement the unitary operator $U$ of a single qubit given by
#
# $$U = \begin{bmatrix}1 & 0\\ 0 & e^{2\pi i\theta}\end{bmatrix}$$
#
# for which an eigenstate is the single-qubit state $\vert1\rangle$. The operator applies a phase
#
# $$U\vert1\rangle = e^{2\pi i\theta}\vert1\rangle$$
#
# Our objective is to determine theta using quantum phase estimation. We will use $\theta=0.5$ and $n = 5$ measurement qubits.
# ### 1. Initializing the qubits
#
# We will need to initialize our qubits as described above by applying a Hadamard gate on each of the $n$ measurement qubits. We will also set the target qubit to $\vert1\rangle$, since that is the eigenstate onto which the unitary operator $U$ will be applied.
#
# We have created a function below called `initialize_qubits` which takes in three arguments. The first argument is the quantum circuit onto which the gates will be applied. The second argument, `measurement_qubits`, is the list of measurement qubits. The third argument, `target_qubit`, is the target qubit for the unitary operator.
def initialize_qubits(given_circuit, measurement_qubits, target_qubit):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.h(measurement_qubits)
given_circuit.x(target_qubit)
### WRITE YOUR CODE BETWEEN THESE LINES - END
# ### 2. Implementing the unitary operator
#
# We have created a function below called `unitary_operator` which takes in three arguments. The first argument is the quantum circuit onto which the operator will be applied. The second argument, `control_qubit`, is the control qubit for the unitary operator. The third argument, `target_qubit`, is the target qubit for the unitary operator. Finally, the fourth argument, `theta`, sets the value of $\theta$.
#
# The function implements the unitary operator described above by using `Qiskit`'s controlled-$u_1$ gate. The matrix for the $u_1$ gate is
#
# $$u_1 = \begin{bmatrix}1 & 0\\ 0 & e^{i\theta}\end{bmatrix}$$
#
# **Note that the phase from the $u_1$ gate differs from that of the $U$ gate by a factor of $2\pi$. You will need to account for this difference in your work.**
import numpy as np
pi = np.pi
def unitary_operator(given_circuit, control_qubit, target_qubit, theta):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.cu1(2*pi*theta, control_qubit, target_qubit)
### WRITE YOUR CODE BETWEEN THESE LINES - END
# You will also need to apply different powers of the unitary operator for the quantum phase estimation algorithm. In order to do this, you can either create a loop that applies that operator several times, or take advantage of the fact that the matrix for $u_1$ is diagonal, and simply multiply the phase by the power.
#
# We have created a function below called `unitary_operator_exponent` which takes in four arguments. The first argument is the quantum circuit onto which the operator will be applied. The second argument, `control_qubit`, is the control qubit for the unitary operator. The third argument, `target_qubit`, is the target qubit for the unitary operator. Finally, the fourth argument, `theta`, sets the value of $\theta$. The fourth argument, `exponent` is the number of times that the unitary operator needs to be applied.
def unitary_operator_exponent(given_circuit, control_qubit, target_qubit, theta, exponent):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.cu1(2*pi*theta*exponent, control_qubit, target_qubit)
### WRITE YOUR CODE BETWEEN THESE LINES - END
# ### 3. Implementing an inverse quantum Fourier transform
#
# You will also need to implement an inverse quantum Fourier transform as part of the quantum phase estimation algorithm. You can do this using two methods.
#
# -- Method 1 (easier) is to use `Qiskit`'s circuit library to give you a box that implements the inverse quantum fourier transform. You can do this using `qiskit.circuit.library.qft(num_qubits).inverse()`. The documentation for this is here: https://qiskit.org/documentation/stubs/qiskit.circuit.library.QFT.html
#
# -- Method 2 (harder) is to implement the gates of the inverse quantum Fourier transform by hand. We strongly recommend following the detailed discussion in the `Qiskit` textbook for examples.
#
# We have created a function below called `apply_iqft` which takes in three arguments. The first argument is the quantum circuit onto which the operator will be applied. The second argument, `measurement_qubits`, is the set of qubits onto which the inverse quantum Fourier transform will be applied. The third argument, `n`, is the number of measurement qubits for which the inverse quantum Fourier transform needs to be created.
from qiskit.circuit.library import QFT
def apply_iqft(given_circuit, measurement_qubits, n):
### WRITE YOUR CODE BETWEEN THESE LINES - START
given_circuit.append(QFT(n).inverse(), measurement_qubits)
### WRITE YOUR CODE BETWEEN THESE LINES - END
# ### 4. Putting it all together
#
# Finally, we combine the functions to construct the quantum program that implements the quantum phase estimation algorithm.
#
# The next lines of code put everything together. **You do not need to modify anything below, but you will need to run the cell to submit your solution.**
from qiskit import QuantumCircuit
# +
def qpe_program(n, theta):
# Create a quantum circuit on n+1 qubits (n measurement, 1 target)
qc = QuantumCircuit(n+1, n)
# Initialize the qubits
initialize_qubits(qc, range(n), n)
# Apply the controlled unitary operators in sequence
for x in range(n):
exponent = 2**(n-x-1)
unitary_operator_exponent(qc, x, n, theta, exponent)
# Apply the inverse quantum Fourier transform
apply_iqft(qc, range(n), n)
# Measure all qubits
qc.measure(range(n), range(n))
return qc
n = 5; theta = 0.5
mycircuit = qpe_program(n, theta)
mycircuit.draw(output='text')
# -
# That's it! You might find it useful to run your quantum circuit and see the measurement outcomes, as well as visualize the statevector at the end.
#
# In order to run your quantum circuit and get the measurement outcomes, you simply need to run `Qiskit`'s `execute` function as follows.
from qiskit import Aer, execute
simulator = Aer.get_backend('qasm_simulator')
counts = execute(mycircuit, backend=simulator, shots=1000).result().get_counts(mycircuit)
from qiskit.visualization import plot_histogram
plot_histogram(counts)
# You can use the measured counts to determine $\theta$ using the following lines of code. Here, we are looking for the outcome with the largest probability, and dividing by $2^n$ since the quantum phase estimation algorithm outputs $2^n\theta$. Note also that we are reversing the order of the bits in the outcome, since `Qiskit` uses the top-most outcome bit to come from the top-most qubit.
import operator
highest_probability_outcome = max(counts.items(), key=operator.itemgetter(1))[0][::-1]
measured_theta = int(highest_probability_outcome, 2)/2**n
print("Using %d qubits with theta = %.2f, measured_theta = %.2f." % (n, theta, measured_theta))
# # Additional reading
#
# - On pi day of 2020 (March 14, 2020), we added a chapter to the `Qiskit` textbook showing how to estimate the value of $\pi$ using the quantum phase estimation algorithm. You can find that implementation here: https://qiskit.org/textbook/ch-demos/piday-code.html
|
Labs/introqcqh-lab-3/lab-3-solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pickle_file = '/home/sulabh/Documents/work/dmgdetection/pytorch/model/model_final.pkl'
import pickle
a = pickle.loads(, 'rb')
with open('/home/sulabh/Documents/work/dmgdetection/pytorch/model/model_final.pkl', 'rb') as f:
d = pickle.load(f, encoding='latin1')
d.keys()
import torch
original = torch.load('/home/sulabh/Documents/work/dmgdetection/pytorch/model/model_final.pkl', map_location='cpu')
# +
from maskrcnn_benchmark.config import cfg
from maskrcnn_benchmark.utils.c2_model_loading import load_c2_format
cfg.merge_from_file("../configs/caffe2/e2e_mask_rcnn_X_101_32x8d_FPN_1x_caffe2.yaml")
path = '/home/sulabh/Documents/work/dmgdetection/pytorch/model/model_final.pkl'
_d = load_c2_format(cfg, path)
keys = [k for k in _d['model'].keys()]
print(sorted(keys))
# -
def _transfer_pretrained_weights(model, pretrained_model_pth):
pretrained_weights = torch.load(pretrained_model_pth)['model']
new_dict = {k.replace('module.',''):v for k, v in pretrained_weights.items()
if 'cls_score' not in k and 'bbox_pred' not in k}
this_state = model.state_dict()
this_state.update(new_dict)
model.load_state_dict(this_state)
return model
|
demo/loading pickle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact, FloatSlider
t = np.linspace(0.0, 5.0)
t
A = 0.5
y = A*np.sin(t)
y
# %matplotlib widget
fig, ax = plt.subplots()
lines = ax.plot(t, y)
lines
type(lines)
line = lines[0]
line
[1, 2, 3]
def update_line(A):
y = A*np.sin(t)
return y
update_line(2.0)
def update_line(A=0.5):
y = A*np.sin(t)
return y
update_line(A=1.0)
update_line()
def update_line(A=0.5):
y = A*np.sin(t)
line.set_ydata(y)
update_line(A=0.2)
widget = interact(update_line, A=FloatSlider(value=0.5, min=0.0, max=5.0))
|
content/materials/notebooks/2020/l02_plotting_widget_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
X = np.array([2.5, 5.1, 3.2, 8.5, 3.5, 1.5, 9.2, 5.5, 8.3, 2.7, 7.7, 5.9, 4.5,
3.3, 1.1, 8.9, 2.5, 1.9, 6.1, 7.4, 2.7, 4.8, 3.8, 6.9, 7.8], dtype='float64')
y = np.array([21, 47, 27, 75, 30, 20, 88, 60, 81, 25, 85, 62, 41, 42, 17, 95, 30,
24, 67, 69, 30, 54, 35, 76, 86], dtype='int64')
import matplotlib.pyplot as plt
plt.scatter(X,y, color = 'red')
X = X.reshape(25,1)
y = y.reshape(25,1)
# # Model satup
from sklearn.linear_model import LinearRegression
model=LinearRegression()
# # Training
model.fit(X,y)
# # Predicting
y_pred= model.predict(X)
y_pred
# # Visualisation
plt.figure(figsize=(12,8))
plt.scatter(X, y, color = 'blue')
plt.plot(X, y_pred, color = 'green')
plt.title('Linear regression plot')
plt.xlabel('Hours')
plt.ylabel('Score')
plt.show()
# # Equation
model.coef_
model.intercept_
# # Evaluating Model
from sklearn.metrics import r2_score
r2_score(y, y_pred)
# # Predicting
model.predict([[2.67],[3.5]])
|
Marks_predt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: drone_racing
# language: python
# name: drone_racing
# ---
from scipy.interpolate import CubicSpline, CubicHermiteSpline
import argparse
import airsimneurips as airsim
import cvxpy as cp
import numpy as np
import time
import airsimneurips as airsim
import gtp
import baseline_racer
import baseline_racer_gtp
import hum_drum_racer
# %matplotlib inline
import matplotlib.pyplot as plt
import track_defs
gate_pose_dicts = track_defs.soccer_medium_gate_pose_dicts
gate_inner_dims_dict= { 'x_val': 1.6,
'y_val': 0.2,
'z_val': 1.6}
gate_outer_dims_dict= { 'x_val': 2.1333333333333333,
'y_val': 0.2,
'z_val': 2.1333333333333333}
# +
# airsim.Quaternionr(x_val,y_val,z_val,w_val) # quaternion ordering
gate_poses = [
airsim.Pose(
airsim.Vector3r(
d['position']['x_val'],
d['position']['y_val'],
d['position']['z_val']
),
airsim.Quaternionr(
d['orientation']['x_val'],
d['orientation']['y_val'],
d['orientation']['z_val'],
d['orientation']['w_val']
)
) for d in gate_pose_dicts
]
# gate_directions = [gtp.rotate_vector(g.orientation, airsim.Vector3r(1,0,0)) for g in gate_poses]
gate_directions = [gtp.rotate_vector(g.orientation, airsim.Vector3r(0,1,0)) for g in gate_poses]
# try rearranging the dimensions here:
gate_inner_dims = airsim.Vector3r(
gate_inner_dims_dict['x_val'],
gate_inner_dims_dict['y_val'],
gate_inner_dims_dict['z_val'],
)
gate_outer_dims = airsim.Vector3r(
gate_outer_dims_dict['x_val'],
gate_outer_dims_dict['y_val'],
gate_outer_dims_dict['z_val'],
)
drone_names = ["drone_1", "drone_2"]
drone_params = [
{"r_safe": 0.4,
"r_coll": 0.3,
"v_max": 20.0,
"a_max": 15.0},
{"r_safe": 0.4,
"r_coll": 0.3,
"v_max": 20.0,
"a_max": 15.0}]
parser = argparse.ArgumentParser(description='')
parser.add_argument('--dt', type=float, default=0.05)
parser.add_argument('--dt_min', type=float, default=0.05)
parser.add_argument('--r_safe', type=float, default=0.0)
parser.add_argument('--v_max', type=float, default=80.0)
parser.add_argument('--a_max', type=float, default=40.0)
parser.add_argument('--n', type=int, default=14)
parser.add_argument('--blocking_behavior', dest='blocking', action='store_true', default=False)
parser.add_argument('--vel_constraints', dest='vel_constraints', action='store_true', default=False)
parser.add_argument('--horizon', type=int, default=10)
parser.add_argument('--no_resample', dest='resample', action='store_false', default=True)
parser.add_argument('--replan_from_lookahead', dest='replan_from_lookahead', action='store_true', default=False)
parser.add_argument('--plot_gtp', dest='plot_gtp', action='store_true', default=False)
parser.add_argument('--level_name', type=str, choices=["Soccer_Field_Easy", "Soccer_Field_Medium", "ZhangJiaJie_Medium", "Building99_Hard",
"Qualifier_Tier_1", "Qualifier_Tier_2", "Qualifier_Tier_3"], default="ZhangJiaJie_Medium")
parser.add_argument('--enable_viz_traj', dest='viz_traj', action='store_true', default=False)
parser.add_argument('--race_tier', type=int, choices=[1,2,3], default=1)
traj_params = parser.parse_known_args()[0]
start_state = airsim.MultirotorState()
start_state.kinematics_estimated.position = airsim.Vector3r(6.373129367828369, 81.43741607666016, -42.88162612915039)
# -
start_state.kinematics_estimated.position.z_val += 3
start_state.kinematics_estimated.position
controller = gtp.IBRController(traj_params,drone_params,gate_poses)
# +
fig, axs = plt.subplots(2, 2,figsize=(15,15))
axs[0,0].plot([c[0] for c in controller.track.track_centers],[c[1] for c in controller.track.track_centers])
axs[0,0].scatter([g.position.x_val for g in gate_poses],[g.position.y_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[0,0].plot([g.position.x_val, g.position.x_val+d.x_val],[g.position.y_val, g.position.y_val+d.y_val],c="red")
axs[0,0].axis('equal')
axs[1,0].plot([c[0] for c in controller.track.track_centers],[c[2] for c in controller.track.track_centers])
axs[1,0].scatter([g.position.x_val for g in gate_poses],[g.position.z_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[1,0].plot([g.position.x_val, g.position.x_val+d.x_val],[g.position.z_val, g.position.z_val+d.z_val],c="red")
axs[1,0].axis('equal')
axs[0,1].plot([c[2] for c in controller.track.track_centers],[c[1] for c in controller.track.track_centers])
axs[0,1].scatter([g.position.z_val for g in gate_poses],[g.position.y_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[0,1].plot([g.position.z_val, g.position.z_val+d.z_val],[g.position.y_val, g.position.y_val+d.y_val],c="red")
axs[0,1].axis('equal')
axs[0,1].set_ylim(ymin=4,ymax=89)
# axs[0,1].set_ylim(ymin=axs[0,0].get_ylim()[0],ymax=axs[0,0].get_ylim()[1])
plt.show()
# -
# # visualize GTP trajectories
# +
# start_pos = start_state.kinematics_estimated.position
# p0 = [start_pos.x_val, start_pos.y_val, start_pos.z_val]
# joint_p0 = np.concatenate([p0-np.array([[2,0,0]]),p0+np.array([[2,0,0]])])
# # ego_id = 1; opp_id = 0;
# ego_id = 0; opp_id = 1;
# base_traj = controller.init_trajectory(ego_id,joint_p0[ego_id,:])
# # ego_traj = controller.iterative_br(ego_id,joint_p0,n_game_iterations=5)
# # opp_traj = controller.iterative_br(opp_id,joint_p0,n_game_iterations=5)
# # # ego_traj = controller.best_response(ego_id,joint_p0,[ego_traj,opp_traj])
# +
# base_traj[1,:]
# +
# plt.figure(figsize=(12,8))
# plt.scatter([g.position.x_val for g in gate_poses],[g.position.y_val for g in gate_poses])
# plt.scatter([g.position.x_val + d.x_val for (g,d) in zip(gate_poses,gate_directions)],
# [g.position.y_val + d.y_val for (g,d) in zip(gate_poses,gate_directions)])
# plt.plot(base_traj[:,0],base_traj[:,1],"black")
# # plt.plot(opp_traj[:,0],opp_traj[:,1],"red")
# # plt.plot(ego_traj[:,0],ego_traj[:,1],"green")
# plt.axis('equal')
# plt.show()
# -
# # visualize global trajectories
optimizer = hum_drum_racer.GlobalTrajectoryOptimizer(traj_params,drone_params[1],gate_poses,gate_inner_dims,gate_outer_dims)
optimizer.traj_params.r_safe = 0.2
optimizer.traj_params.dt = 0.1
traj = optimizer.compute_global_optimal_trajectory(start_state)
pos = traj.pos
vel = traj.vel
accel = traj.accel
t_vec = traj.t_vec
# +
fig, axs = plt.subplots(2, 2,figsize=(15,15))
axs[0,0].plot([p[0] for p in pos],[p[1] for p in pos])
axs[0,0].scatter([g.position.x_val for g in gate_poses],[g.position.y_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[0,0].plot([g.position.x_val, g.position.x_val+d.x_val],[g.position.y_val, g.position.y_val+d.y_val],c="red")
axs[0,0].axis('equal')
axs[1,0].plot([p[0] for p in pos],[p[2] for p in pos])
axs[1,0].scatter([g.position.x_val for g in gate_poses],[g.position.z_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[1,0].plot([g.position.x_val, g.position.x_val+d.x_val],[g.position.z_val, g.position.z_val+d.z_val],c="red")
axs[1,0].axis('equal')
axs[0,1].plot([p[2] for p in pos],[p[1] for p in pos])
axs[0,1].scatter([g.position.z_val for g in gate_poses],[g.position.y_val for g in gate_poses])
for (g,d) in zip(gate_poses,gate_directions):
axs[0,1].plot([g.position.z_val, g.position.z_val+d.z_val],[g.position.y_val, g.position.y_val+d.y_val],c="red")
axs[0,1].axis('equal')
axs[0,1].set_ylim(ymin=4,ymax=89)
# axs[0,1].set_ylim(ymin=axs[0,0].get_ylim()[0],ymax=axs[0,0].get_ylim()[1])
plt.show()
# -
fig, axs = plt.subplots(1,2,figsize=(15,4))
axs[0].plot(t_vec,vel)
axs[0].plot(t_vec,[np.linalg.norm(v) for v in vel])
axs[1].plot(t_vec,accel)
plt.show()
|
baselines/HumDrumExperiments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
from fastai import *
from fastai.vision import *
from fastai.vision.models.wrn import wrn_22
from fastai.docs import *
torch.backends.cudnn.benchmark = True
# -
untar_data(CIFAR_PATH)
ds_tfms = ([*rand_pad(4, 32), flip_lr(p=0.5)], [])
data = image_data_from_folder(CIFAR_PATH, valid='test', ds_tfms=ds_tfms, tfms=cifar_norm, bs=512)
learn = Learner(data, wrn_22(), metrics=accuracy).to_fp16()
learn.fit_one_cycle(30, 3e-3, wd=0.4, div_factor=10, pct_start=0.5)
# With mixup
learn = Learner(data, wrn_22(), metrics=accuracy).to_fp16().mixup()
learn.fit_one_cycle(24, 3e-3, wd=0.2, div_factor=10, pct_start=0.5)
|
examples/cifar.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison with Stata
#
# For potential users coming from [Stata](https://en.wikipedia.org/wiki/Stata)
# this page is meant to demonstrate how different Stata operations would be
# performed in pandas.
#
# If you’re new to pandas, you might want to first read through [10 Minutes to pandas](../04_user_guide/10min.ipynb#min)
# to familiarize yourself with the library.
#
# As is customary, we import pandas and NumPy as follows:
# + hide-output=false
import pandas as pd
import numpy as np
# -
# ## Data structures
# ### General terminology translation
#
# |pandas|Stata|
# |:------------------:|:------------------:|
# |DataFrame|data set|
# |column|variable|
# |row|observation|
# |groupby|bysort|
# |NaN|.|
# ### `DataFrame`
#
# A `DataFrame` in pandas is analogous to a Stata data set – a two-dimensional
# data source with labeled columns that can be of different types. As will be
# shown in this document, almost any operation that can be applied to a data set
# in Stata can also be accomplished in pandas.
# ### `Series`
#
# A `Series` is the data structure that represents one column of a
# `DataFrame`. Stata doesn’t have a separate data structure for a single column,
# but in general, working with a `Series` is analogous to referencing a column
# of a data set in Stata.
# ### `Index`
#
# Every `DataFrame` and `Series` has an `Index` – labels on the
# *rows* of the data. Stata does not have an exactly analogous concept. In Stata, a data set’s
# rows are essentially unlabeled, other than an implicit integer index that can be
# accessed with `_n`.
#
# In pandas, if no index is specified, an integer index is also used by default
# (first row = 0, second row = 1, and so on). While using a labeled `Index` or
# `MultiIndex` can enable sophisticated analyses and is ultimately an important
# part of pandas to understand, for this comparison we will essentially ignore the
# `Index` and just treat the `DataFrame` as a collection of columns. Please
# see the indexing documentation for much more on how to use an
# `Index` effectively.
# ### Copies vs. in place operations
#
# Most pandas operations return copies of the `Series`/`DataFrame`. To make the changes “stick”,
# you’ll need to either assign to a new variable:
#
# > sorted_df = df.sort_values(“col1”)
#
#
# or overwrite the original one:
# + [markdown] hide-output=false
# >df = df.sort_values("col1")
# -
# >**Note**
# >
# >You will see an `inplace=True` keyword argument available for some methods:
# + [markdown] hide-output=false
# >df.sort_values("col1", inplace=True)
#
# Its use is discouraged. :ref:`More information. <indexing.view_versus_copy>`
# -
# ## Data input / output
# ### Constructing a DataFrame from values
#
# A Stata data set can be built from specified values by
# placing the data after an `input` statement and
# specifying the column names.
# + [markdown] hide-output=false
# ```stata
# input x y
# 1 2
# 3 4
# 5 6
# end
# ```
#
# -
# A pandas `DataFrame` can be constructed in many different ways,
# but for a small number of values, it is often convenient to specify it as
# a Python dictionary, where the keys are the column names
# and the values are the data.
# + hide-output=false
df = pd.DataFrame({"x": [1, 3, 5], "y": [2, 4, 6]})
df
# -
# ### Reading external data
#
# Like Stata, pandas provides utilities for reading in data from
# many formats. The `tips` data set, found within the pandas
# tests ([csv](https://raw.github.com/pandas-dev/pandas/master/pandas/tests/io/data/csv/tips.csv))
# will be used in many of the following examples.
#
# Stata provides `import delimited` to read csv data into a data set in memory.
# If the `tips.csv` file is in the current working directory, we can import it as follows.
# + [markdown] hide-output=false
# ```stata
# import delimited tips.csv
# ```
#
# -
# The pandas method is `read_csv()`, which works similarly. Additionally, it will automatically download
# the data set if presented with a url.
# + hide-output=false
url = (
"https://raw.github.com/pandas-dev"
"/pandas/master/pandas/tests/io/data/csv/tips.csv"
)
tips = pd.read_csv(url)
tips
# -
# Like `import delimited`, `read_csv()` can take a number of parameters to specify
# how the data should be parsed. For example, if the data were instead tab delimited,
# did not have column names, and existed in the current working directory,
# the pandas command would be:
# + [markdown] hide-output=false
# ```python
# tips = pd.read_csv("tips.csv", sep="\t", header=None)
#
# # alternatively, read_table is an alias to read_csv with tab delimiter
# tips = pd.read_table("tips.csv", header=None)
# ```
#
# -
# pandas can also read Stata data sets in `.dta` format with the `read_stata()` function.
# + [markdown] hide-output=false
# ```python
# df = pd.read_stata("data.dta")
# ```
#
# -
# In addition to text/csv and Stata files, pandas supports a variety of other data formats
# such as Excel, SAS, HDF5, Parquet, and SQL databases. These are all read via a `pd.read_*`
# function. See the IO documentation for more details.
# ### Limiting output
#
# By default, pandas will truncate output of large `DataFrame`s to show the first and last rows.
# This can be overridden by changing the pandas options, or using
# `DataFrame.head()` or `DataFrame.tail()`.
# + hide-output=false
tips.head(5)
# -
# The equivalent in Stata would be:
# + [markdown] hide-output=false
# ```stata
# list in 1/5
# ```
#
# -
# ### Exporting data
#
# The inverse of `import delimited` in Stata is `export delimited`
# + [markdown] hide-output=false
# ```stata
# export delimited tips2.csv
# ```
#
# -
# Similarly in pandas, the opposite of `read_csv` is `DataFrame.to_csv()`.
# + [markdown] hide-output=false
# ```python
# tips.to_csv("tips2.csv")
# ```
#
# -
# pandas can also export to Stata file format with the `DataFrame.to_stata()` method.
# + [markdown] hide-output=false
# ```python
# tips.to_stata("tips2.dta")
# ```
#
# -
# ## Data operations
# ### Operations on columns
#
# In Stata, arbitrary math expressions can be used with the `generate` and
# `replace` commands on new or existing columns. The `drop` command drops
# the column from the data set.
# + [markdown] hide-output=false
# ```stata
# replace total_bill = total_bill - 2
# generate new_bill = total_bill / 2
# drop new_bill
# ```
#
# -
# pandas provides vectorized operations by specifying the individual `Series` in the
# `DataFrame`. New columns can be assigned in the same way. The `DataFrame.drop()` method drops
# a column from the `DataFrame`.
# + hide-output=false
tips["total_bill"] = tips["total_bill"] - 2
tips["new_bill"] = tips["total_bill"] / 2
tips
tips = tips.drop("new_bill", axis=1)
# -
# ### Filtering
#
# Filtering in Stata is done with an `if` clause on one or more columns.
# + [markdown] hide-output=false
# ```stata
# list if total_bill > 10
# ```
#
# -
# DataFrames can be filtered in multiple ways; the most intuitive of which is using
# boolean indexing.
# + hide-output=false
tips[tips["total_bill"] > 10]
# -
# The above statement is simply passing a `Series` of `True`/`False` objects to the DataFrame,
# returning all rows with `True`.
# + hide-output=false
is_dinner = tips["time"] == "Dinner"
is_dinner
is_dinner.value_counts()
tips[is_dinner]
# -
# ### If/then logic
#
# In Stata, an `if` clause can also be used to create new columns.
# + [markdown] hide-output=false
# ```stata
# generate bucket = "low" if total_bill < 10
# replace bucket = "high" if total_bill >= 10
# ```
#
# -
# The same operation in pandas can be accomplished using
# the `where` method from `numpy`.
# + hide-output=false
tips["bucket"] = np.where(tips["total_bill"] < 10, "low", "high")
tips
# -
#
# <dl style='margin: 20px 0;'>
# <dt>::</dt>
# <dd>
#
# <dl style='margin: 20px 0;'>
# <dt>suppress</dt>
# <dd>
# </dd>
#
# </dl>
#
# tips = tips.drop(“bucket”, axis=1)
#
# </dd>
#
# </dl>
# ### Date functionality
#
# Stata provides a variety of functions to do operations on
# date/datetime columns.
# + [markdown] hide-output=false
# ```stata
# generate date1 = mdy(1, 15, 2013)
# generate date2 = date("Feb152015", "MDY")
#
# generate date1_year = year(date1)
# generate date2_month = month(date2)
#
# * shift date to beginning of next month
# generate date1_next = mdy(month(date1) + 1, 1, year(date1)) if month(date1) != 12
# replace date1_next = mdy(1, 1, year(date1) + 1) if month(date1) == 12
# generate months_between = mofd(date2) - mofd(date1)
#
# list date1 date2 date1_year date2_month date1_next months_between
# ```
#
# -
# The equivalent pandas operations are shown below. In addition to these
# functions, pandas supports other Time Series features
# not available in Stata (such as time zone handling and custom offsets) –
# see the timeseries documentation for more details.
# + hide-output=false
tips["date1"] = pd.Timestamp("2013-01-15")
tips["date2"] = pd.Timestamp("2015-02-15")
tips["date1_year"] = tips["date1"].dt.year
tips["date2_month"] = tips["date2"].dt.month
tips["date1_next"] = tips["date1"] + pd.offsets.MonthBegin()
tips["months_between"] = tips["date2"].dt.to_period("M") - tips[
"date1"
].dt.to_period("M")
tips[
["date1", "date2", "date1_year", "date2_month", "date1_next", "months_between"]
]
# -
# cleanup
tips = tips.drop(["date1", "date2", "date1_year", "date2_month", "date1_next", "months_between"],
axis=1,
)
# ### Selection of columns
#
# Stata provides keywords to select, drop, and rename columns.
# + [markdown] hide-output=false
# ```stata
# keep sex total_bill tip
#
# drop sex
#
# rename total_bill total_bill_2
# ```
#
# -
# The same operations are expressed in pandas below.
# #### Keep certain columns
# + hide-output=false
tips[["sex", "total_bill", "tip"]]
# -
# #### Drop a column
# + hide-output=false
tips.drop("sex", axis=1)
# -
# #### Rename a column
# + hide-output=false
tips.rename(columns={"total_bill": "total_bill_2"})
# -
# ### Sorting by values
#
# Sorting in Stata is accomplished via `sort`
# + [markdown] hide-output=false
# ```stata
# sort sex total_bill
# ```
#
# -
# pandas has a `DataFrame.sort_values()` method, which takes a list of columns to sort by.
# + hide-output=false
tips = tips.sort_values(["sex", "total_bill"])
tips
# -
# ## String processing
# ### Finding length of string
#
# Stata determines the length of a character string with the `strlen()` and
# `ustrlen()` functions for ASCII and Unicode strings, respectively.
# + [markdown] hide-output=false
# ```stata
# generate strlen_time = strlen(time)
# generate ustrlen_time = ustrlen(time)
# ```
#
# -
# You can find the length of a character string with `Series.str.len()`.
# In Python 3, all strings are Unicode strings. `len` includes trailing blanks.
# Use `len` and `rstrip` to exclude trailing blanks.
# + hide-output=false
tips["time"].str.len()
tips["time"].str.rstrip().str.len()
# -
# ### Finding position of substring
#
# Stata determines the position of a character in a string with the `strpos()` function.
# This takes the string defined by the first argument and searches for the
# first position of the substring you supply as the second argument.
# + [markdown] hide-output=false
# ```stata
# generate str_position = strpos(sex, "ale")
# ```
#
# -
# You can find the position of a character in a column of strings with the `Series.str.find()`
# method. `find` searches for the first position of the substring. If the substring is found, the
# method returns its position. If not found, it returns `-1`. Keep in mind that Python indexes are
# zero-based.
# + hide-output=false
tips["sex"].str.find("ale")
# -
# ### Extracting substring by position
#
# Stata extracts a substring from a string based on its position with the `substr()` function.
# + [markdown] hide-output=false
# ```stata
# generate short_sex = substr(sex, 1, 1)
# ```
#
# -
# With pandas you can use `[]` notation to extract a substring
# from a string by position locations. Keep in mind that Python
# indexes are zero-based.
# + hide-output=false
tips["sex"].str[0:1]
# -
# ### Extracting nth word
#
# The Stata `word()` function returns the nth word from a string.
# The first argument is the string you want to parse and the
# second argument specifies which word you want to extract.
# + [markdown] hide-output=false
# ```stata
# clear
# input str20 string
# "<NAME>"
# "<NAME>"
# end
#
# generate first_name = word(name, 1)
# generate last_name = word(name, -1)
# ```
#
# -
# The simplest way to extract words in pandas is to split the strings by spaces, then reference the
# word by index. Note there are more powerful approaches should you need them.
# + hide-output=false
firstlast = pd.DataFrame({"String": ["<NAME>", "<NAME>"]})
firstlast["First_Name"] = firstlast["String"].str.split(" ", expand=True)[0]
firstlast["Last_Name"] = firstlast["String"].str.rsplit(" ", expand=True)[0]
firstlast
# -
# ### Changing case
#
# The Stata `strupper()`, `strlower()`, `strproper()`,
# `ustrupper()`, `ustrlower()`, and `ustrtitle()` functions
# change the case of ASCII and Unicode strings, respectively.
# + [markdown] hide-output=false
# ```stata
# clear
# input str20 string
# "<NAME>"
# "<NAME>"
# end
#
# generate upper = strupper(string)
# generate lower = strlower(string)
# generate title = strproper(string)
# list
# ```
#
# -
# The equivalent pandas methods are `Series.str.upper()`, `Series.str.lower()`, and
# `Series.str.title()`.
# + hide-output=false
firstlast = pd.DataFrame({"string": ["<NAME>", "<NAME>"]})
firstlast["upper"] = firstlast["string"].str.upper()
firstlast["lower"] = firstlast["string"].str.lower()
firstlast["title"] = firstlast["string"].str.title()
firstlast
# -
# ## Merging
#
# The following tables will be used in the merge examples:
# + hide-output=false
df1 = pd.DataFrame({"key": ["A", "B", "C", "D"], "value": np.random.randn(4)})
df1
df2 = pd.DataFrame({"key": ["B", "D", "D", "E"], "value": np.random.randn(4)})
df2
# -
# In Stata, to perform a merge, one data set must be in memory
# and the other must be referenced as a file name on disk. In
# contrast, Python must have both `DataFrames` already in memory.
#
# By default, Stata performs an outer join, where all observations
# from both data sets are left in memory after the merge. One can
# keep only observations from the initial data set, the merged data set,
# or the intersection of the two by using the values created in the
# `_merge` variable.
# + [markdown] hide-output=false
# ```stata
# * First create df2 and save to disk
# clear
# input str1 key
# B
# D
# D
# E
# end
# generate value = rnormal()
# save df2.dta
#
# * Now create df1 in memory
# clear
# input str1 key
# A
# B
# C
# D
# end
# generate value = rnormal()
#
# preserve
#
# * Left join
# merge 1:n key using df2.dta
# keep if _merge == 1
#
# * Right join
# restore, preserve
# merge 1:n key using df2.dta
# keep if _merge == 2
#
# * Inner join
# restore, preserve
# merge 1:n key using df2.dta
# keep if _merge == 3
#
# * Outer join
# restore
# merge 1:n key using df2.dta
# ```
#
# -
# pandas DataFrames have a `merge()` method, which provides similar functionality. The
# data does not have to be sorted ahead of time, and different join types are accomplished via the
# `how` keyword.
# + hide-output=false
inner_join = df1.merge(df2, on=["key"], how="inner")
inner_join
left_join = df1.merge(df2, on=["key"], how="left")
left_join
right_join = df1.merge(df2, on=["key"], how="right")
right_join
outer_join = df1.merge(df2, on=["key"], how="outer")
outer_join
# -
# ## Missing data
#
# Both pandas and Stata have a representation for missing data.
#
# pandas represents missing data with the special float value `NaN` (not a number). Many of the
# semantics are the same; for example missing data propagates through numeric operations, and is
# ignored by default for aggregations.
# + hide-output=false
outer_join
outer_join["value_x"] + outer_join["value_y"]
outer_join["value_x"].sum()
# -
# One difference is that missing data cannot be compared to its sentinel value.
# For example, in Stata you could do this to filter missing values.
# + [markdown] hide-output=false
# ```stata
# * Keep missing values
# list if value_x == .
# * Keep non-missing values
# list if value_x != .
# ```
#
# -
# In pandas, `Series.isna()` and `Series.notna()` can be used to filter the rows.
# + hide-output=false
outer_join[outer_join["value_x"].isna()]
outer_join[outer_join["value_x"].notna()]
# -
# pandas provides a variety of methods to work with missing data. Here are some examples:
# ### Drop rows with missing values
# + hide-output=false
outer_join.dropna()
# -
# ### Forward fill from previous rows
# + hide-output=false
outer_join.fillna(method="ffill")
# -
# ### Replace missing values with a specified value
#
# Using the mean:
# + hide-output=false
outer_join["value_x"].fillna(outer_join["value_x"].mean())
# -
# ## GroupBy
# ### Aggregation
#
# Stata’s `collapse` can be used to group by one or
# more key variables and compute aggregations on
# numeric columns.
# + [markdown] hide-output=false
# ```stata
# collapse (sum) total_bill tip, by(sex smoker)
# ```
#
# -
# pandas provides a flexible `groupby` mechanism that allows similar aggregations. See the
# groupby documentation for more details and examples.
# + hide-output=false
tips_summed = tips.groupby(["sex", "smoker"])[["total_bill", "tip"]].sum()
tips_summed
# -
# ### Transformation
#
# In Stata, if the group aggregations need to be used with the
# original data set, one would usually use `bysort` with `egen()`.
# For example, to subtract the mean for each observation by smoker group.
# + [markdown] hide-output=false
# ```stata
# bysort sex smoker: egen group_bill = mean(total_bill)
# generate adj_total_bill = total_bill - group_bill
# ```
#
# -
# pandas provides a [Transformation](../04_user_guide/36_groupby.ipynb#groupby-transform) mechanism that allows these type of operations to be
# succinctly expressed in one operation.
# + hide-output=false
gb = tips.groupby("smoker")["total_bill"]
tips["adj_total_bill"] = tips["total_bill"] - gb.transform("mean")
tips
# -
# ### By group processing
#
# In addition to aggregation, pandas `groupby` can be used to
# replicate most other `bysort` processing from Stata. For example,
# the following example lists the first observation in the current
# sort order by sex/smoker group.
# + [markdown] hide-output=false
# ```stata
# bysort sex smoker: list if _n == 1
# ```
#
# -
# In pandas this would be written as:
# + hide-output=false
tips.groupby(["sex", "smoker"]).first()
# -
# ## Other considerations
# ### Disk vs memory
#
# pandas and Stata both operate exclusively in memory. This means that the size of
# data able to be loaded in pandas is limited by your machine’s memory.
# If out of core processing is needed, one possibility is the
# [dask.dataframe](https://dask.pydata.org/en/latest/dataframe.html)
# library, which provides a subset of pandas functionality for an
# on-disk `DataFrame`.
|
01_comparison/comparison_with_stata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.stats as st
import scipy.optimize as opt
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d # for 3d plots
# # Fit data with function $y = f(x)$ by minimizing sum or squared errors.
# +
time_sec = np.linspace(0, 10, 1000)
current_pA = 100 * np.exp(-time_sec / 2) + 20 * (np.random.rand(len(time_sec)) - 0.5)
plt.plot(time_sec, current_pA)
plt.xlabel('Time (sec)')
plt.ylabel('Current (pA)');
# +
def exponential_decay(time, amp, tau):
return amp * np.exp(-time / tau)
result = opt.curve_fit(exponential_decay, time_sec, current_pA)
print(result)
# +
amp, tau = result[0]
print(amp, tau)
# +
result = opt.curve_fit(exponential_decay, time_sec, current_pA, p0=[90, 4], bounds=([80, 1], [120, 10]))
amp, tau = result[0]
print(amp, tau)
# -
plt.plot(time_sec, current_pA)
plt.plot(time_sec, exponential_decay(time_sec, amp, tau), lw=3)
plt.xlabel('Time (sec)')
plt.ylabel('Current (pA)');
# # Find optimal parameters that minimize an objective (or cost) function.
# ### Find $x$ that minimizes $(|x| - 5)^2 + x$
# +
# cost function to be minimized
def cost_func(x):
return (abs(x) - 5)**2 + x
# plot cost function
x = np.linspace(-10, 10, 100)
plt.plot(x, cost_func(x))
plt.xlabel('$x$')
plt.ylabel('$(|x| - 5)^2 + x$');
# +
x0 = [-10] # initial guess
# find value of x that minimizes cost function
result = opt.minimize(cost_func, x0)
print(result)
print()
print(result.x)
# -
x0 = [10]
result = opt.minimize(cost_func, x0)
print(result)
x0 = [10]
result = opt.minimize(cost_func, x0, method='slsqp')
print(result)
x0 = [-10] # initial guess
bnds = opt.Bounds([0], [10]) # lower bounds, upper bounds
result = opt.minimize(cost_func, x0, method='slsqp', bounds=bnds)
print(result)
# ### Find $x$ and $y$ that minimizes $10 (x - 5)^2 + 9 (y + 3)^2$
params = [1, 3]
x, y = params
print(x, y)
# +
# cost function to be minimized
def cost_func(params):
x, y = params
return 10 * (x - 5)**2 + 9 * (y + 3)**2
# plot cost function
x = np.linspace(-10, 10, 100)
y = np.linspace(-10, 10, 100)
cost = np.zeros((len(y), len(x)))
for row in range(len(y)):
for col in range(len(x)):
cost[row, col] = cost_func([x[col], y[row]])
plt.imshow(cost, aspect='auto', cmap=plt.cm.nipy_spectral_r)
plt.xticks([0, 24, 49, 74, 99], [-10, -5, 0, 5, 10])
plt.yticks([0, 24, 49, 74, 99], [-10, -5, 0, 5, 10])
plt.xlabel('x')
plt.ylabel('y');
plt.title('cost')
plt.colorbar();
# -
x0 = -10
y0 = 10
guess = [x0, y0] # initial guess
bnds = opt.Bounds([-10, -10], [10, 10]) # lower bounds, upper bounds
result = opt.minimize(cost_func, guess, method='slsqp', bounds=bnds)
print(result)
# ### Find $x$ and $y$ that minimizes $10 (x - 5)^2 + 9 (y + 3)^2$ under the constraint $x = y$
#
# You will need to cast all constraints in one of these two forms:
#
# 1. equality_constraint(params) = 0
# 2. inequality_constraint(params) >= 0
# +
def constraint_that_x_equals_y(params):
x, y = params
return x - y
con_eq = {'type': 'eq', 'fun': constraint_that_x_equals_y}
guess = [-10, 10] # initial guess
bnds = opt.Bounds([-10, -10], [10, 10]) # lower bounds, upper bounds
result = opt.minimize(cost_func, guess, method='slsqp', bounds=bnds, constraints=con_eq)
print(result)
# +
plt.imshow(cost, aspect='auto', cmap=plt.cm.nipy_spectral_r)
plt.xticks([0, 24, 49, 74, 99], [-10, -5, 0, 5, 10])
plt.yticks([0, 24, 49, 74, 99], [-10, -5, 0, 5, 10])
plt.xlabel('x')
plt.ylabel('y');
plt.title('cost')
plt.colorbar();
plt.plot([0, 99], [0, 99])
x1, y1 = result.x # optimal values
col = np.argmin(np.abs(x - x1))
row = np.argmin(np.abs(y - y1))
plt.plot(col, row, 'o');
# -
# ### Find $x$ and $y$ that minimizes $10 (x - 5)^2 + 9 (y + 3)^2$ under the constraint $x <= y + 2$
# +
def constraint_that_x_less_than_or_equal_to_y(params):
x, y = params
return y + 2 - x
con_ineq = {'type': 'ineq', 'fun': constraint_that_x_less_than_or_equal_to_y}
guess = [-10, 10] # initial guess
bnds = opt.Bounds([-10, -10], [10, 10]) # lower bounds, upper bounds
result = opt.minimize(cost_func, guess, method='slsqp', bounds=bnds, constraints=con_ineq)
print(result)
# -
# ### You place a mouse in a chamber with two paths, one of which leads to a reward and one that doesn't, and observe whether it chooses the path to the reward. You repeat the experiment 10 times per mouse for 100 mice (randomizing where the reward is each time). Based on your data (see below), what is the maximum likelihood probability that a mouse will select the path to the reward? What is the 95% confidence interval for your estimate?
num_rewards_out_of_10 = np.array([ 6, 5, 7, 4, 8, 4, 8, 6, 3, 4, 6, 10, 7, 7, 7, 7, 7,
9, 7, 7, 5, 6, 8, 8, 8, 5, 8, 4, 6, 6, 7, 6, 8, 9,
5, 4, 5, 8, 8, 6, 9, 8, 7, 4, 8, 7, 9, 9, 6, 8, 7,
4, 6, 8, 8, 8, 6, 8, 8, 6, 6, 6, 7, 8, 9, 7, 9, 9,
9, 7, 6, 5, 5, 9, 6, 9, 7, 7, 4, 9, 8, 9, 8, 6, 4,
8, 8, 8, 8, 8, 6, 8, 6, 8, 6, 8, 5, 9, 9, 5], dtype=int)
# +
def binomNegLoglikelihood(k, n, p):
return -st.binom.logpmf(k, n, p).sum()
def dataNegLoglikelihood(p):
return binomNegLoglikelihood(num_rewards_out_of_10, 10, p)
p = opt.minimize_scalar(dataNegLoglikelihood, bounds=(0, 1), method='bounded')
print(p)
# -
# ### You record the number of times each of 50 neurons spike within 1 minute after application of a novel compound. What is the maximum likelihood estimate of these neuron's spike rate in response to the compound? What is the 95% confidence interval for your estimate?
observed_num_spikes_in_a_minute_for_all_neurons = np.array([32, 35, 32, 37, 32, 26, 30, 35, 33, 37, 34, 31, 28, 29, 33, 29, 30,
25, 32, 29, 34, 41, 33, 29, 30, 18, 30, 29, 30, 42, 22, 30, 21, 38,
36, 33, 33, 38, 32, 39, 32, 38, 36, 42, 25, 29, 23, 28, 43, 49], dtype=int)
# +
def poissonNegLoglikelihood(k, mu):
return -st.poisson.logpmf(k, mu).sum()
def dataNegLoglikelihood(mu):
return poissonNegLoglikelihood(observed_num_spikes_in_a_minute_for_all_neurons, mu)
lb = observed_num_spikes_in_a_minute_for_all_neurons.min()
ub = observed_num_spikes_in_a_minute_for_all_neurons.max()
p = opt.minimize_scalar(dataNegLoglikelihood, bounds=(lb, ub), method='bounded')
print(p)
print()
print("MLE of spike rate in reponse to compound is", p.x, "spikes per minute")
# -
# ### Repeat above for mice in chamber with reward and non-reward paths. However, this time you lost your records for how many times you tested each mouse! Get a maximum likelihood estimate for both the probability that a mouse selects the reward path and the number of times each mouse was placed in the chamber and 95% confidence intervals for each estimate.
# +
num_rewards_out_of_who_knows = num_rewards_out_of_10
def binomNegLoglikelihood(k, n, p):
return -st.binom.logpmf(k, n, p).sum()
def dataNegLoglikelihood(params):
n, p = params
return binomNegLoglikelihood(num_rewards_out_of_who_knows, n, p)
n0 = 20
p0 = 0.25
guess = [n0, p0] # initial guess
bnds = opt.Bounds([0, 0], [20, 1]) # lower bounds, upper bounds
result = opt.minimize(dataNegLoglikelihood, guess, method='slsqp', bounds=bnds)
print(result)
# +
n, p = result.x
print("MLE probability of choosing reward path is", p, "on each of", int(n), "attempts")
# -
# # Example exercises
# ### 1. Fit the following data to a theoretical function.
time_sec = np.linspace(0, 1, 100)
voltage_mV = np.array([-19.33535068, -19.81593 , -19.38068935, -19.01444258,
-19.01279965, -18.08972043, -19.05443624, -16.92871871,
-17.41921874, -17.13110999, -16.23081011, -15.93055289,
-16.6963649 , -16.02562308, -16.08446757, -15.70107301,
-16.14943005, -16.24260514, -16.2479143 , -15.26316983,
-14.14899883, -14.2671532 , -13.85340797, -15.38637959,
-13.62949667, -14.09838187, -14.5240729 , -14.52185707,
-13.02644386, -12.98712829, -14.37612648, -14.06115682,
-13.58528874, -13.74801745, -13.31550763, -13.82579987,
-13.37824516, -13.02067885, -12.20748865, -13.12844023,
-12.80277207, -12.85018719, -12.76829176, -12.65668693,
-11.97756266, -11.46451702, -11.67070899, -12.32710194,
-12.93915408, -12.80260755, -11.62231014, -12.64038186,
-11.91782401, -10.87750533, -11.82992523, -11.67502684,
-11.82323243, -10.76213803, -12.05737906, -11.3578388 ,
-10.44744459, -11.76906988, -11.36599376, -12.1965993 ,
-10.16171931, -11.60630276, -12.0049248 , -11.61201511,
-10.11810532, -10.02273279, -10.38912268, -11.72630602,
-10.08001563, -10.58014772, -10.03496926, -11.50078207,
-10.01827333, -11.1164367 , -10.86874543, -10.15982785,
-9.72053465, -10.04952839, -10.31409222, -11.28338912,
-10.65026096, -11.20603978, -9.6280181 , -9.73437495,
-10.38121048, -9.64101998, -11.08428724, -9.69146819,
-9.6996263 , -9.6253655 , -10.97046052, -11.28989399,
-9.63686909, -9.87094353, -10.55579744, -9.6242286 ])
# +
plt.plot(time_sec, voltage_mV);
def exp_rise(t, amp, tau, constant):
return amp * (1 - np.exp(-t / tau)) + constant
result = opt.curve_fit(exp_rise, time_sec, voltage_mV, p0=[10, 0.2, -20], bounds=([1, 0.01, -100], [40, 10, 100]))
amp, tau, constant = result[0]
print(amp, tau, constant)
plt.plot(time_sec, exp_rise(time_sec, amp, tau, constant), lw=3)
# -
# ### 2. Using electron microscopy you count the number of docked vesicles per synapse. What is the maximum likelihood estimate for the average number of docked vesicles per synapse given you data below? Also provide 95% confidence limits on this estimate.
# each entry is # of docked vesicles at a particular synapse
num_docked_vesicles = np.array([ 8, 3, 7, 7, 5, 5, 3, 7, 7, 7, 6, 3, 7, 5, 3, 8, 5,
8, 5, 5, 3, 3, 6, 7, 8, 13, 7, 7, 7, 5, 4, 9, 13, 3,
5, 7, 6, 5, 4, 4, 3, 5, 13, 7, 8, 5, 4, 9, 4, 8, 3,
5, 3, 5, 3, 5, 6, 7, 7, 6, 8, 8, 7, 5, 9, 3, 11, 9,
8, 5, 2, 9, 8, 8, 2, 8, 8, 5, 8, 4, 6, 8, 5, 6, 3,
3, 7, 7, 2, 4, 6, 7, 7, 4, 4, 2, 7, 3, 9, 7, 7, 4,
5, 9, 6, 6, 5, 2, 7, 6, 4, 6, 4, 7, 7, 8, 8, 8, 5,
7, 5, 6, 4, 2, 7, 8, 4, 2, 6, 3, 5, 5, 5, 8, 7, 2,
4, 8, 9, 10, 5, 6, 5, 5, 5, 4, 3, 2, 3, 5, 7, 2, 5,
7, 5, 10, 8, 5, 3, 4, 6, 7, 8, 3, 5, 7, 9, 7, 4, 6,
3, 6, 7, 5, 9, 4, 5, 13, 9, 10, 6, 7, 6, 5, 9, 8, 8,
3, 5, 3, 7, 6, 8, 6, 3, 5, 8, 6, 6, 3], dtype=int)
# +
def poissonNegLoglikelihood(k, mu):
return -st.poisson.logpmf(k, mu).sum()
def cost_function(mu):
return poissonNegLoglikelihood(num_docked_vesicles, mu)
p = opt.minimize_scalar(cost_function, bounds=(0, 100), method='bounded')
print(p)
# -
# ### 3. You record the time-dependent current through a single ion channel at +50 mV in symmetrical ionic solutions (time series data is below). Based on the data, determine the channel's conductance states and their 95% confidence intervals.
# Single channel current record (pA). Sample interval is 100 microseconds.
current_pA = np.array([ 0.00000000e+00, -2.35394235e-01, 6.07833286e-01, -8.70306907e-01,
-1.25841170e+00, -1.74297695e-02, -5.58284641e-01, 2.78609759e+00,
4.78248770e+00, 4.69373968e+00, 5.67881976e+00, 4.99705998e+00,
6.30184317e+00, 4.83487181e+00, 3.03183825e+00, 5.83206241e+00,
5.35819319e+00, -4.60627502e-01, 1.07486132e+00, 2.91376156e+00,
2.10070182e+00, 1.18350752e+00, 1.91661039e+00, 2.27939756e+00,
1.29232847e+00, 1.48268820e+00, 4.61369862e+00, 5.00355371e+00,
-1.44819161e+00, -5.51972368e-01, 2.98400128e-01, 7.46190742e-01,
-6.42705287e-01, 4.58047840e-01, -7.41532194e-01, -6.73262703e-01,
5.91905023e-01, 2.41612119e-01, 2.99097843e+00, 1.68857562e+00,
2.54164978e+00, 2.73297315e+00, 2.44545466e+00, 1.17925091e+00,
2.75972609e+00, 1.37661225e+00, 1.98199173e+00, 1.76662384e+00,
2.51339495e+00, 1.97739440e+00, 2.52634436e+00, 8.51353231e-01,
-5.73178723e-01, 1.63084881e+00, 3.21066731e+00, 6.50098026e+00,
3.82993941e+00, -3.40622218e-01, -6.44956162e-01, -5.14403137e-01,
-2.07115509e-01, -9.05523326e-01, 3.59862599e-01, 1.35010587e-01,
-4.15770501e-03, -2.82345061e-01, 2.81339139e-01, -2.87053959e-01,
2.39041366e+00, 4.35900735e-01, 1.66756888e+00, 1.91837972e+00,
1.11969643e+00, 1.31477977e+00, 1.45233161e+00, 1.70625427e+00,
2.18176920e+00, 2.23218499e+00, 1.66333979e+00, 1.74158175e+00,
3.86019668e+00, 1.50568450e+00, 1.25141608e+00, 2.20152044e+00,
4.69589336e+00, 4.25880606e+00, 4.61861707e+00, -5.94591437e-01,
-3.65028789e-01, 3.23722523e-01, 6.19410844e+00, 5.58166939e+00,
3.85071042e+00, 4.40009238e+00, 8.12198881e-01, 4.42617977e-01,
5.95656399e-01, -1.10237496e+00, -8.98242376e-01, 7.94843118e-02,
5.02012149e+00, 5.47512233e+00, 5.25582988e+00, 5.49404794e+00,
4.28887172e+00, 4.61565686e+00, 3.83608122e+00, 2.35337233e+00,
3.67022237e+00, -1.11334635e+00, -3.18741077e-01, -3.99539006e-01,
8.52420828e-02, -3.93682375e-01, -2.68777379e-02, -5.10853746e-02,
-1.15787989e-01, -3.52036704e-01, -5.32425938e-01, -1.24672576e+00,
3.78593323e-01, 1.15892914e+00, -5.83994444e-01, 2.78768157e-01,
1.39543987e-01, -5.79143158e-01, 3.39523161e+00, 5.22938269e+00,
5.12957734e+00, 5.99947723e+00, 3.47899706e+00, 4.54859360e+00,
4.43192113e+00, 6.18596821e+00, 4.80092205e+00, 3.84851588e+00,
5.52286481e+00, 4.22197068e+00, 5.25556506e+00, 3.39726765e+00,
3.95806880e+00, 5.19732271e-02, 3.82649645e-01, -2.81867318e-01,
1.54658894e-01, 7.41042105e-01, 1.19612111e-01, 3.08308106e-01,
1.30592584e-01, -4.90093303e-01, 1.76953535e-02, 3.70764975e-01,
6.93714048e-01, 1.34719391e-03, -1.32212088e-01, -5.78001048e-01,
1.75952612e-01, -3.85325860e-01, 2.97573118e+00, 4.78669205e+00,
4.61278529e+00, 2.76800451e+00, 4.16105050e+00, 5.33284168e+00,
-6.91454966e-01, 7.44278510e-01, -1.78087687e-01, -1.07690707e-02,
-5.15949200e-01, -2.12352159e-02, 1.43678730e-02, 4.10445502e-01,
1.02651487e-01, -6.21051441e-02, 9.73525935e-02, 3.51205459e-01,
-1.53660529e-01, -4.67862784e-01, 4.76170250e-01, 1.48609704e-01,
6.97893890e-01, 5.81011874e+00, 1.17941897e+00, -3.81008936e-01,
1.12724230e+00, -6.24757237e-01, 7.51028577e-01, 2.66629092e-01,
5.50412454e+00, 4.19133725e+00, 4.09773610e+00, 4.91765871e+00,
4.48516386e+00, 3.42712419e+00, 5.04727020e+00, 5.76303846e+00,
5.64527893e+00, 5.31863192e+00, 6.84720646e+00, 5.10201998e+00,
6.13293645e+00, 5.17184085e+00, 3.28588438e+00, 6.21459053e+00,
8.86636021e-01, 4.30073067e-01, -1.82835345e-01, 6.23432272e+00,
4.89739508e+00, 4.95640821e+00, 5.86488527e+00, 4.76614324e+00,
5.56635125e+00, 5.58515436e+00, 5.32609459e+00, 4.44922633e+00,
4.68761265e+00, 5.72018314e+00, 4.53798775e+00, 5.35117506e+00,
6.55592233e+00, 7.31036662e-01, 1.00165977e-02, 1.00127817e+00,
-5.12566393e-01, 6.64430341e-02, 4.94824937e-02, -2.52122454e-01,
-1.66718603e-03, 4.01108487e+00, 5.38854432e+00, 4.40596752e+00,
4.31745088e+00, 4.09093292e+00, 5.49939608e+00, 4.38892305e+00,
7.06768792e+00, 4.83096902e+00, 4.67238235e+00, 4.64054073e+00,
5.97315744e+00, 3.64447394e+00, 6.52360451e+00, 1.67080047e-01,
1.58743815e-01, -4.97994325e-01, 2.77442586e-01, 8.40816780e-01,
-2.76123328e-01, 2.96957222e-01, 8.67329185e-01, -2.15183359e-01,
1.97688390e+00, 2.50587484e+00, 5.33830774e+00, 6.35012627e+00,
4.72571451e+00, 4.12477664e+00, 4.98959977e+00, 4.99791463e+00,
4.31788380e+00, 4.82419428e+00, 6.24065982e+00, 4.71929306e+00,
6.47166049e+00, 4.64263725e+00, 4.25388140e+00, 4.97108273e+00,
5.36356874e+00, 6.09683618e+00, 4.70881307e+00, 3.90790662e+00,
4.60344649e+00, 2.96657248e+00, 4.91172918e+00, 4.86004171e+00,
-1.03664236e-01, -1.28981910e+00, 5.65662422e+00, 4.84480518e+00,
4.63513934e+00, 3.88702111e+00, 5.40899961e+00, 6.58228256e+00,
4.22522761e+00, 4.98799388e+00, 3.82113599e+00, 5.34481406e+00,
3.48372344e+00, 2.76332305e+00, 5.07734330e+00, 4.19223767e+00,
5.44372783e+00, 5.74934559e+00, 4.93328862e+00, 6.47041815e+00,
4.80420346e+00, 4.76978276e+00, 5.05511742e+00, 4.95050316e+00,
3.94901753e+00, 3.92472554e+00, 4.31938430e+00, 4.65210990e+00,
6.15180663e+00, 5.07877829e+00, 6.56012409e+00, 4.20723605e+00,
3.81550402e+00, 4.47061072e+00, -5.69054016e-01, -7.84616662e-01,
-1.98468796e-01, 5.38468108e-01, 5.73989511e-01, 1.50372009e-02,
1.96245180e-01, -7.09689441e-01, 7.76448894e-01, 5.03919904e-01,
-1.69266354e-01, -7.83270455e-01, -8.94048883e-01, -1.17435501e+00,
-6.64616930e-01, 3.99032164e-01, 7.07776055e-01, -5.98138336e-01,
2.16213375e-01, 3.04162160e-01, -1.24266851e+00, -1.42416970e-01,
6.19975070e-01, 4.84911325e-01, 1.95967750e-01, -6.37770775e-01,
-5.83109741e-01, -9.62725747e-01, -5.18823095e-01, -5.76940857e-01,
4.62372911e-01, 8.69445149e-01, 5.12538010e-01, 3.17208541e-01,
4.72208582e-01, 2.27873851e+00, 2.57947901e+00, 2.55378886e+00,
-6.21728442e-01, -8.04110171e-01, 6.13548342e-01, -2.34156894e-01,
5.60106130e+00, 4.58944257e+00, 4.52930781e+00, 6.89404816e+00,
5.18856013e+00, 3.70978746e+00, 5.64345571e+00, 4.17924589e+00,
5.97330067e+00, 1.53794649e-01, 2.05903817e-01, 1.61260938e-01,
-3.25067734e-01, 1.50956982e-01, 4.09703821e-01, -2.67476991e-01,
5.50912650e+00, -3.84203946e-01, -1.09823328e-01, -9.11846052e-03,
-6.60663999e-02, 8.09315308e-01, 5.69463865e-01, -4.69163402e-01,
-1.25743717e-01, 3.17528871e-01, -8.31831349e-01, 5.14788905e-01,
-4.50043642e-01, -2.09737709e-01, -5.43059326e-02, -5.14679082e-01,
-1.84233722e-01, 2.92250622e+00, 6.04219983e+00, 5.89374422e+00,
5.08316147e+00, 5.28852812e+00, 4.55730956e+00, 3.40790975e+00,
4.26451066e+00, 4.39505582e+00, 3.01505874e+00, 5.22432918e+00,
4.12567297e+00, 3.85602698e+00, 5.32965183e+00, 4.14539342e+00,
3.68378241e+00, 4.47342468e+00, 5.05517480e+00, 6.59834411e+00,
5.55650193e+00, 6.91460435e+00, 4.21584833e+00, 6.02797597e+00,
6.21484966e+00, 5.78263423e+00, 6.02205711e+00, 4.73538849e+00,
6.78430844e+00, 5.31165875e+00, 4.16615106e+00, 6.24564358e+00,
6.12474151e+00, 4.78900502e+00, 5.87334550e+00, 6.09462959e+00,
5.44633275e+00, 5.04946494e+00, 3.55625112e+00, 1.97616094e+00,
1.27371899e+00, 5.15156729e-02, 6.96208827e-01, 6.69849259e-01,
4.45580988e+00, 4.02572944e+00, 2.17927668e+00, 6.55604770e+00,
3.33227557e+00, 4.82226224e+00, 4.89847403e+00, 2.84086030e+00,
-5.86029968e-01, 2.36353907e-01, 2.39716393e-01, -7.07255090e-01,
-2.81219915e-01, 2.51116339e-01, -1.26565191e-01, 3.62540753e+00,
3.32995387e+00, 5.66744265e-01, -3.40645496e-02, -5.84484719e-01,
5.78613493e-02, 3.23157318e-01, 9.60771854e-02, -4.16136912e-01,
-7.04181139e-02, 4.23666253e-01, -5.74010844e-01, -1.71738551e-01,
-2.42243438e-01, 2.18816668e-01, 3.07139554e-01, 3.49351347e-01,
8.56725469e-01, 1.23632321e+00, 1.88519013e-01, -2.98144414e-01,
4.98438837e-02, 1.14320966e+00, -1.09747355e-01, 3.19141747e+00,
1.67241140e+00, 3.66922162e+00, 1.46583340e+00, 2.85149140e+00,
2.74238434e+00, 2.32424380e+00, 2.86499143e+00, 2.38703208e+00,
2.05881964e+00, 1.75789261e+00, 1.77086193e+00, 1.70326953e+00,
2.52617867e+00, 2.07655081e+00, 1.07607905e+00, 2.43453813e+00,
2.26353454e+00, 1.90824367e+00, 1.01588795e+00, 1.93368475e+00,
1.99684441e+00, 5.11396615e+00, -4.58835008e-01, 8.90157108e-01,
3.88577889e-01, 1.80190657e+00, 9.10633663e-01, 2.89357535e+00,
5.66504206e+00, 5.49020086e+00, 3.08693614e+00, 4.67819246e+00,
5.19606891e+00, 4.82684580e+00, 3.88366081e+00, 4.49152794e+00,
4.34359416e+00, 4.35466445e+00, 3.30521825e+00, 6.25648225e+00,
5.02653493e+00, -5.70298789e-02, 1.19753305e+00, 3.42595810e-01,
1.67493637e+00, 2.26428932e+00, 1.47106154e+00, 1.50521973e+00,
2.09963688e+00, 1.95238363e+00, 1.71197507e+00, 2.18808090e+00,
3.20780417e+00, 1.65151195e+00, 1.61326552e+00, 3.09809517e+00,
1.88794987e+00, 2.18740681e+00, 7.74253556e-01, 2.77897882e+00,
1.72264326e+00, 2.41937107e+00, 1.44320491e+00, 5.17589501e-01,
2.00088254e+00, 2.41455666e+00, 2.51899870e+00, 1.64813695e+00,
4.68627910e+00, 5.67713834e+00, 3.52135890e+00, 4.89507513e+00,
7.18450996e+00, 4.11871362e+00, 5.17891649e+00, 6.31167964e+00,
5.46112205e+00, 4.03201460e+00, 3.92611121e+00, 5.84590943e+00,
5.36973422e+00, 5.59472581e+00, 4.28178480e+00, 4.37445322e+00,
5.05222825e+00, 5.52243570e+00, 3.23795458e+00, 5.16305168e+00,
5.26622084e+00, 6.01434228e+00, 4.12277624e+00, 6.21006910e+00,
6.24229037e+00, 4.23761650e+00, 5.09941210e+00, 5.35582130e+00,
4.09610898e+00, 5.42990944e+00, 4.46057780e+00, 5.57508105e+00,
5.01842699e+00, 5.01288934e+00, -1.52280443e+00, -2.33384021e-02,
-6.34707019e-02, -1.00099157e+00, 2.68210925e-01, 4.22525634e+00,
5.33389294e+00, 3.42543670e+00, 5.47599883e+00, 3.93018892e+00,
5.06096559e+00, 5.11148086e+00, 5.71579167e+00, 6.34809632e+00,
6.11894498e+00, 4.82520063e+00, 5.33995915e+00, 4.76015774e+00,
5.74386634e+00, 4.74162452e+00, 5.14921295e+00, 2.42957509e+00,
5.22449423e+00, 4.61466962e+00, 3.45230630e+00, 4.42505258e+00,
5.23595054e+00, 3.13562129e+00, 3.38412124e+00, 5.54940694e+00,
5.24741588e+00, 5.20281489e+00, 5.78871816e+00, 4.73847482e+00,
5.15666998e+00, 5.68131499e+00, 4.55421657e+00, 6.45650213e+00,
6.32737988e+00, 4.53161324e+00, 4.03978912e+00, 5.69315915e+00,
5.10470017e+00, 4.50330832e+00, 4.57634074e+00, 4.98727683e+00,
5.10453786e+00, 4.60708248e+00, 2.53164025e-01, -4.43106675e-01,
2.65099292e+00, 6.55287395e+00, 5.51439659e+00, 5.16165839e+00,
3.58207227e+00, 1.00171123e+00, -1.34743915e-01, 1.07353283e+00,
9.43654937e-01, 3.75702439e+00, 4.80809556e+00, 5.44679085e+00,
3.54428702e-02, 4.57189642e-01, -6.22028786e-03, 4.92928767e-01,
3.09224777e-01, 3.89566279e-01, 3.19268180e-01, 3.54326036e-01,
-9.72095846e-01, -6.78172273e-01, -1.68570029e-01, -3.36378664e-01,
-4.51651633e-01, -6.34563358e-01, -3.33411742e-01, 4.06929719e-01,
-2.63869480e-01, -2.27360242e-01, -5.50530063e-01, -8.00015938e-02,
-5.05033354e-01, 2.56639866e-01, -6.71518215e-02, -5.53206933e-01,
-5.25135475e-01, -2.53552619e-01, -1.06549885e+00, -4.94171583e-01,
3.42475039e-01, 8.41688979e-01, 3.32803148e-01, -5.20871100e-01,
5.87397550e-02, 1.96749995e+00, 2.29304688e+00, 2.29672192e+00,
7.35592817e-02, 1.75138038e+00, 1.75796971e+00, 2.69261781e+00,
3.44781270e+00, 1.71453366e+00, 2.29723326e+00, 1.60633201e+00,
3.06150336e+00, 1.32374036e+00, 1.94679548e+00, 1.22769026e+00,
5.42947746e+00, 5.89559369e+00, 5.63004935e+00, 5.05891805e+00,
5.28104571e+00, 2.72631153e+00, 5.49943719e+00, 4.67899259e+00,
6.20384231e+00, 5.01494446e+00, 4.22665514e+00, 4.81657598e+00,
2.38426134e+00, 5.38265912e+00, 6.42894192e+00, 3.06540277e+00,
5.92706574e+00, 6.67872976e-01, 2.78216391e+00, 8.03945493e-01,
2.72272307e+00, 2.52631306e+00, 1.86918398e+00, 2.46094378e+00,
2.88882336e+00, 2.22845569e+00, 1.71159936e+00, 1.27204212e+00,
3.00080403e+00, 1.63441038e+00, 1.89847505e+00, 2.92439964e+00,
2.28317064e+00, 2.30928990e+00, 1.56514084e+00, 1.80904833e+00,
4.15168874e+00, 4.41225582e+00, 5.36599510e+00, 5.73157588e+00,
4.52129243e+00, 5.06654091e+00, 4.18619254e+00, 7.13478843e+00,
1.92832261e+00, 1.39923528e+00, 1.55280253e+00, 2.41123612e+00,
2.58210760e+00, 1.13366612e+00, 2.98031988e+00, 1.72448109e+00,
2.16135015e+00, 1.79579051e+00, 2.45928505e+00, 9.65086076e-01,
3.39874514e+00, 2.53253911e+00, 2.29074849e+00, 8.51206219e-01,
1.90698624e+00, 2.27474697e+00, 2.36703413e+00, 1.11421027e+00,
3.06094100e+00, 2.49187196e+00, 2.55557142e+00, 1.73560141e+00,
1.32353615e+00, 1.20405983e+00, 8.01954351e-01, 3.63372157e+00,
4.06946668e+00, 5.35883815e+00, 4.53815453e+00, 4.50734149e+00,
4.42884467e+00, 5.15082273e+00, 5.38048767e+00, -4.83060337e-01,
5.86324781e-01, -7.07635657e-01, -3.96769767e-02, -1.71928536e-01,
1.59561715e-01, 5.72425704e-01, 5.03455513e+00, 2.47486072e+00,
2.86333190e+00, 6.23822818e+00, 3.44193571e+00, 5.86772308e+00,
5.91485831e+00, 5.63492174e+00, 1.98388811e-02, 1.91886033e-01,
-2.86882974e-01, 3.39431116e-01, -5.11399980e-01, 7.32068641e-01,
3.35959486e+00, 4.35632324e+00, -5.98912740e-01, 1.04535587e-01,
8.27681537e-01, -1.23641570e+00, 2.56551076e+00, 5.59692672e+00,
5.87656682e+00, 6.04474640e+00, 6.75038499e+00, 3.76111285e+00,
6.15217880e+00, 5.22876038e+00, 5.38097539e-01, 2.49936229e-01,
7.66582790e-01, 1.13116318e-01, 4.69509486e-01, -2.40432983e-01,
3.64384014e-01, -3.93192343e-02, -4.67088178e-01, 2.37359997e+00,
2.02762815e+00, 5.28426900e+00, 6.49650666e+00, 6.88385717e-01,
-5.66918501e-01, 7.14217986e-01, 6.34703268e-04, 6.58382771e-01,
-9.68954276e-01, 3.69113024e-01, -3.42755629e-02, 3.61400383e-01,
1.68605611e-01, 1.32412039e+00, 4.99299216e+00, 5.85098743e+00,
4.43981428e+00, 4.34171615e+00, 6.75843133e+00, 5.48285066e+00,
4.72453113e+00, 6.26784060e+00, 3.79397173e+00, 4.46706639e+00,
-5.16051490e-01, 1.80721884e+00, 1.27800963e+00, 2.65462601e+00,
1.73931061e+00, 2.36206879e+00, 1.78028518e+00, 2.64296002e+00,
2.53924418e+00, 2.68635313e+00, 1.52218848e+00, 1.80082431e+00,
9.89247193e-01, 1.83857348e+00, 2.56870733e+00, 1.79388177e+00,
6.43174868e+00, 4.05676529e+00, 5.16035089e+00, 8.10403268e+00,
6.33395956e+00, 5.36969250e+00, 4.11021517e+00, 5.57863174e+00,
2.39729125e+00, 4.87265882e+00, 1.53999248e+00, 3.16692176e-01,
-9.68394495e-02, -5.48466391e-02, 5.03758778e-01, -3.86349026e-01,
3.90149996e-01, -2.59305907e-01, -2.75176353e-01, 6.07524663e-01,
1.51878435e-01, 4.89789530e-01, -3.63861950e-02, -3.59848445e-01,
5.49975461e+00, 5.24749798e+00, 4.51709104e+00, 4.51697794e+00,
3.26617802e+00, 5.41773177e+00, 5.08079949e+00, 4.47700465e+00,
5.15032511e+00, 5.44431222e+00, 6.26265551e+00, 5.22118916e-02,
-7.29572053e-01, 1.80990868e-01, 3.87541970e-01, -2.62406441e-01,
2.25509443e-01, 2.21612096e-01, -9.27519392e-02, 7.82146116e-01,
1.26083109e-01, -1.20822439e+00, -6.45699877e-01, -5.98345610e-01,
-5.80377986e-01, 6.27625432e-03, 3.26924906e-01, -1.16486764e-01,
1.07788700e-01, -3.23001830e-01, 4.69522684e-01, 1.15845476e+00,
-5.73753815e-01, 5.94855932e+00, 4.81268300e+00, 3.59854874e+00,
3.31126973e+00, 2.14634417e+00, 8.76389478e-01, 1.02613886e+00,
1.82915419e+00, 2.94004634e+00, 3.85671064e+00, 5.27870687e+00,
7.02120176e+00, 6.31854655e+00, 7.83279294e+00, 8.44696228e-01,
6.06551408e-01, 2.44036705e+00, 2.14885036e+00, 2.42615539e+00,
1.24812405e+00, 2.51986227e+00, 1.55529541e+00, 5.88929664e+00,
5.27394828e+00, 4.31596058e+00, 3.17104408e+00, 5.57176325e+00,
5.68050660e+00, 3.85081684e+00, 6.14503970e+00, 3.54355996e+00,
4.97140261e+00, 4.45069669e+00, 4.93222238e+00, 8.95799483e-02,
-3.77974444e-01, 5.62340607e-01, 1.50443813e-01, -3.38391311e-01,
-1.16628334e+00, -8.43633004e-01, -5.06260355e-02, -1.57859465e-01,
-1.38528486e-01, -1.30226510e-01, -5.15944750e-01, 2.69338674e-01,
-2.11566463e-02, -2.13466381e-02, 8.10999760e-01, 7.15971437e-02,
6.59688135e+00, 2.64405256e+00, 1.51465420e+00, 2.22868548e+00,
2.82653800e+00, 1.72987041e+00, 1.92337588e+00, 2.99460875e+00,
1.64045313e+00, 2.00815223e+00, 8.26479943e-01, 1.26351041e+00,
2.18910372e+00, 1.07603690e+00, 1.15761229e+00, 2.12761980e+00,
2.90168507e+00, 2.32714947e+00, 2.99920749e+00, 2.73666409e+00,
1.52107281e+00, 1.49327412e+00, 2.04311653e+00, 1.46003820e+00,
2.25981785e+00, 1.87948603e+00, 1.96461422e+00, 2.99696402e+00,
1.48325323e+00, 1.34831203e+00, 1.75584295e+00, 2.85157391e+00,
2.38644556e+00, 1.41198351e+00, 2.74144257e+00, 5.08657668e+00,
4.25039843e+00, 5.30212904e+00, 4.96883289e+00, 5.76473842e+00,
5.67735333e+00, 5.69871463e+00, 5.94869568e+00, 4.01429796e+00,
4.95619286e+00, 5.67372506e+00, 5.71424239e+00, 3.73974484e+00])
fig = plt.figure(figsize=(18, 6))
plt.plot(current_pA)
plt.hist(current_pA, 30);
|
lecture_08_optimization/optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Environment setting
# move to source directory
# %cd ../
# %pwd
# %ls
# import major modules
import pandas as pd
import numpy as np
import sys
from pathlib import Path
import seaborn as sns
import matplotlib.pyplot as plt
sys.stdout.flush()
from util.easydict import EasyDict
import json
# +
version = '0012'
use_small_data = False
dsize = '.small' if use_small_data is True else ''
# import config
r = json.load(open(f'config/result_{version}{dsize}.json'))
r = EasyDict(r)
r
# -
# # Read data
log = pd.read_csv(r.paths.train_log_path, delimiter='\t')
print(log.shape)
log.head()
sns.lineplot(x="iteration", y="train_auc", data=log)
sns.lineplot(x="iteration", y="val_auc", data=log)
plt.ylabel('auc')
importance = pd.read_csv(r.paths.importance_path)
importance.columns = ['feature', 'importance']
importance.head()
data = importance.sort_values(by="importance", ascending=False)[0:20]
data.head()
sns.barplot(x="importance", y="feature", data=data);
|
notebook/0012-xgb-magic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 12) Intro to NumPy (prodounced Num-Pie), Numerical Python
# ## Libraries commonly used for Data Science
#
# numpy, scipy, matplotlib, pandas, and scikit-learn will all be at least briefly covered in this course. The two we have not used yet are the last two; you can import them with:
# ```
# $ conda install pandas scikit-learn
# ```
#
# We'll go over each one, starting with NumPy, which we've already used several times.
#
# Related references:
#
# - https://jakevdp.github.io/WhirlwindTourOfPython/15-preview-of-data-science-tools.html
# - https://jakevdp.github.io/PythonDataScienceHandbook/02.00-introduction-to-numpy.html
# ## The Case for NumPy
#
# NumPy provides an efficient way to store and manipulate multi-dimensional dense arrays in Python.
# The important features of NumPy are:
#
# - It provides an ``ndarray`` structure, which allows efficient storage and manipulation of vectors, matrices, and higher-dimensional datasets.
# - It provides a readable and efficient syntax for operating on this data, from simple element-wise arithmetic to more complicated linear algebraic operations. (As we've previously discussed: do not write your own loops; use libraries as much as possible to speed up programming and your programs)
#
# In the simplest case, NumPy arrays look a lot like Python lists.
# For example, here is an array containing the range of numbers 1 to 9 (compare this with Python's built-in ``range()``):
# +
import numpy as np
x = np.arange(1, 10)
print(type(x), x)
y = list(range(1, 10))
print(type(y), y)
# -
# Key differences include:
#
# - We can directly do math on an ndarray, versus needing a loop (= slow) for lists
# - Lists are always 1D (although you can have lists of lists) while arrays can have any number of dimensions
x_squared = x**2
print(x_squared)
y_squared = [val ** 2 for val in y]
print(y_squared)
m = x.reshape((3,3)) # This reshape command will only work if the total size remains the same
print("matrix:")
print(m)
# Furthermore, NumPy knows how to do lots of math, including linear algebra.
#
# What is a matrix? Who remembers what the transpose of a matrix is?
#
# For next class, would you like a review of linear algebra basics?
print(m.T)
# The multiple ways to make ndarrays include making them from lists or lists of lists.
# an array from a list
a = np.array([3.14, 4, 2, 3])
print(a, a.shape)
m * 2.5
# nested lists result in multi-dimensional arrays
list_of_lists = [list(range(i, i + 3)) for i in [2, 4, 6, 8]]
print(list_of_lists)
b = np.array(list_of_lists)
print(b, b.shape)
# so do lists of lists
c = np.array([[1, 2], [3, 4], [5, 6]])
print(c, c.shape)
# If you don't have a specific set of values you want to use, it is more efficient to directly generate ndarrays.
# Create a length-10 integer array filled with zeros
np.zeros(10, dtype=int)
# Create a 3x5 floating-point (the default type) array filled with ones
np.ones((3, 5))
# Create a 3x5 array filled with 3.14
np.full((3, 5), np.nan)
# Create an array filled with a linear sequence
# Starting at 0, ending at 20, stepping by 2
# (this is similar to the built-in range() function)
np.arange(0, 20, 2)
np.arange(0, 1, 5)
# Create an array of five values evenly spaced between 0 and 1
np.linspace(0, 1, 5)
# Create a 3x3 array of uniformly distributed
# random values between 0 and 1
np.random.random((3, 3))
# Create a 3x3 array of normally distributed random values
# with mean 0 and standard deviation 1
np.random.normal(0, 1, (3, 3))
# Create a 3x3 array of random integers in the interval [0, 10)
np.random.randint(0, 10, (3, 3))
# Create a 4x4 identity matrix
np.eye(4)
# Create an uninitialized array of three integers
# The values will be whatever happens to already exist at that memory location
np.empty(3)
# Other data types you can use in numpy arrays include booleans and complex numbers.
# The follow two commands are equivalent
np.ones((3, 5), dtype=bool)
np.full((3, 5), True)
np.zeros((3, 5), dtype=complex)
# ## Some Useful NumPy Array Attributes
#
# First let's discuss some useful array attributes. We'll start by defining three random arrays, a one-dimensional, two-dimensional, and three-dimensional array. We'll use NumPy's random number generator, which we will seed with a set value in order to ensure that the same random arrays are generated each time this code is run:
# +
np.random.seed(0) # seed for reproducibility
x1 = np.random.randint(10, size=6) # One-dimensional array
x2 = np.random.randint(10, size=(3, 4)) # Two-dimensional array
x3 = np.random.randint(10, size=(3, 4, 5)) # Three-dimensional array
# -
print("x3 shape:", x3.shape) # shape
print("x3 ndim: ", x3.ndim) # number of dimensions
print("x3 size: ", x3.size) # total size of the array
print("dtype:", x3.dtype) # data type
# ## Some Ways ndarrays Are Like Python Lists: Slicing and indexing
#
# What we previously learned about list slicing and indexing applies here, too:
x1
print(x1)
print(type(x1))
x1[4]
x1[0]
x1[-2]
# Let's compare 2D indexing
list2 = [[3, 5, 2, 4], [7, 6, 8, 8], [1, 6, 7, 7]]
print(list2)
print(x2)
print(list2[2][0])
print(x2[2, 0])
# And modification
list2[0][1] = 12
x2[0, 1] = 12
print(list2)
print(x2)
# ### The general format for slicing is:
# ~~~
# x[start:stop:step]
# ~~~
z = np.arange(10)
d = list(range(10))
print(z)
print(d)
print(z[:5])
print(d[:5])
print(z[4:7])
print(d[4:7])
print(z[::2]) # every other element
print(d[::2])
print(z[3::2]) # every other element, starting at 3
print(d[3::2])
print(z[::-1]) # all elements, reversed
print(d[::-1])
# ## And some differences
#
# Lists can be heterogeneous. Arrays cannot.
list2[0][1] = 8.4
x2[0, 1] = 8.4
print(list2)
print(x2)
x2[0] = x2[1] * 1.1
print(x2)
# For ndarrays, slicing works similarly for higher dimensional arrays. Things are more complicated for lists of lists.
print(list2)
print(x2)
print(list2[:2][:1])
print(list2[:2])
print(x2[:2, :3])
print(list2[2][:2])
print(x2[0]) # equivalent to x2[0, :], prints the first row
print(list2[0])
# ### Subparts of an array will change the parent array
print(x2)
x2_part = x2[:2, :2]
print(x2_part)
x2_part[0,1] = 0
print(x2)
# We can get around this similarly to how we worked with this problem in lists--make a copy!
x2_part_copy = x2[:2, :2].copy()
x2_part_copy[0, 0] = 42
print(x2)
# ## Array Concatenation and Splitting
# There are three main ways that arrays can be joined: `np.concatenate`, `np.vstack`, and `np.hstack`. In each case, the arrays to be joined must be of compatible dimensions.
#
# [np.concatenate](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.concatenate.html) tacks the second array onto the first. You can specify the axis along which it is to be joined (default is `axis=0`).
x = np.array([1, 2, 3])
y = np.array([3, 2, 1])
z = np.concatenate([x, y])
print(z)
a = np.array([[1, 2], [3, 4]])
b = np.array([[5, 6]])
c = np.concatenate([a, b])
print(c)
# multiple arrays can be concatenated at once
d = np.array([[7, 8]])
e = np.concatenate([a, b.T, d.T], axis=1)
print(e)
# [np.vstack](https://docs.scipy.org/doc/numpy/reference/generated/numpy.vstack.html) and [np.hstack](https://docs.scipy.org/doc/numpy/reference/generated/numpy.hstack.html) work like concatenate, without your having to remember which axis is vertical and which is horizontal.
print(a)
print(b)
f = np.vstack([a, b])
print(f)
g = np.hstack([a, b.T])
print(g)
# The opposite of stacking is [splitting](https://docs.scipy.org/doc/numpy/reference/generated/numpy.split.html).
x = [1, 2, 3, 99, 99, 3, 2, 1]
x1, x2, x3, x4 = np.split(x, [3, 5, 6]) # the second argument gives the split points
print(x1, x2, x3, x4)
grid = np.arange(25).reshape((5, 5))
print(grid)
upper, lower = np.vsplit(grid, [2])
print(upper)
print(lower)
left, right = np.hsplit(grid, [2])
print(left)
print(right)
# Similarly, [np.dsplit](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dsplit.html#numpy.dsplit) will split arrays along the third axis (depth).
# Next up: computations with ndarrays!
|
source/notebooks/lecture12_numpy_attributes_methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Note: The original CelebA dataset is of 11 GB which we were unable to load it on Google Drive (space constraints). So, we have decided to work on Aligned CelebA dataset.
# + [markdown] id="Po4ZUf5yXXBt"
# # Import Libraries
# + executionInfo={"elapsed": 27533, "status": "ok", "timestamp": 1617775659984, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}, "user_tz": -330} id="Mc9Qpz2YXXB0"
from utilities import *
import cv2
import numpy as np
import matplotlib.pyplot as plt
from glob import glob
from tqdm import tqdm
# + [markdown] id="WKkdtSFgXXB1"
# # Initializations
# + executionInfo={"elapsed": 27534, "status": "ok", "timestamp": 1617775659990, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}, "user_tz": -330} id="nMlasld7XXB1"
# initialize_bboxes_file(file_path = '../../CelebA/metadata/list_bbox_celeba.txt')
initialize_dlib_shape_predictor("../../Face Landmark Detection Pre-trained models/shape_predictor_81_face_landmarks.dat")
# + [markdown] id="tIuHgVoKXXB2"
# # Low level features
# + executionInfo={"elapsed": 28054, "status": "ok", "timestamp": 1617775660516, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}, "user_tz": -330} id="b4jAzBp_tK1Z"
output_file_path = "../../data/low level/celeba/"
final_features = [None] * (11*4) # Why 11? As 11 face parts are being returned. Why 4? As each face part is associated with 4 histograms.
for i in range(len(final_features)):
final_features[i] = np.empty(shape = (0, HISTOGRAM_BINS))
file_exception = open(output_file_path + "images_under_error.txt", "w")
# + id="iG6d7ox2XXB2" tags=[]
for file_name in tqdm(sorted(glob("../../../CelebA/img_align_celeba/img_align_celeba/*"))[:20000]):
try:
# Face extraction
# img_face_extracted = face_extraction_lfw(cv2.cvtColor(cv2.imread(file_name), cv2.COLOR_BGR2RGB))
# Face alignment
img_face_aligned = cv2.cvtColor(cv2.imread(file_name), cv2.COLOR_BGR2RGB)
shape = face_alignment_dlib(img_face_aligned, display_intermediate_results = False, do_not_align = True)
## Uncomment below to see the aligned eyes
# img_face_aligned_copy = img_face_aligned.copy()
# for j in range(len(shape)):
# cv2.circle(img_face_aligned_copy, tuple(shape[j]), 2, (255, 0, 0), -1)
# left_eye_center, right_eye_center = get_center_eyes_dlib(shape)
# cv2.line(img_face_aligned_copy, tuple(left_eye_center), tuple(right_eye_center), (0, 0, 255), 1)
# cv2.circle(img_face_aligned_copy, tuple(left_eye_center), 2, (0, 255, 0), -1)
# cv2.circle(img_face_aligned_copy, tuple(right_eye_center), 2, (0, 255, 0), -1)
# plt.imshow(img_face_aligned_copy)
# plt.axis('off')
# plt.show()
# Low level features extraction
"""
features =
rgb_left_eye, hsv_left_eye, grad_mag_left_eye, grad_orien_left_eye,
rgb_right_eye, hsv_right_eye, grad_mag_right_eye, grad_orien_right_eye,
rgb_nose, hsv_nose, grad_mag_nose, grad_orien_nose,
rgb_mouth, hsv_mouth, grad_mag_mouth, grad_orien_mouth,
rgb_chin, hsv_chin, grad_mag_chin, grad_orien_chin,
rgb_moustache, hsv_moustache, grad_mag_moustache, grad_orien_moustache,
rgb_left_cheek, hsv_left_cheek, grad_mag_left_cheek, grad_orien_left_cheek,
rgb_right_cheek, hsv_right_cheek, grad_mag_right_cheek, grad_orien_right_cheek,
rgb_forehead, hsv_forehead, grad_mag_forehead, grad_orien_forehead,
rgb_full_face, hsv_full_face, grad_mag_full_face, grad_orien_full_face,
rgb_hair, hsv_hair, grad_mag_hair, grad_orien_hair
"""
features = extract_low_level_features_research_paper(img_face_aligned, shape, display_intermediate_results = False)
for i in range(len(features)):
final_features[i] = np.append(final_features[i], features[i].reshape(1, HISTOGRAM_BINS), axis = 0)
except Exception as e:
file_exception.write(file_name + " " + str(e) + "\n")
print("Completed")
# + [markdown] id="HmaOv5w5X1GO"
# # Save histogram data as .npy
# + id="Slik1lSCtPNs"
features_title = ['rgb_left_eye', 'hsv_left_eye', 'grad_mag_left_eye', 'grad_orien_left_eye', \
'rgb_right_eye', 'hsv_right_eye', 'grad_mag_right_eye', 'grad_orien_right_eye', \
'rgb_nose', 'hsv_nose', 'grad_mag_nose', 'grad_orien_nose', \
'rgb_mouth', 'hsv_mouth', 'grad_mag_mouth', 'grad_orien_mouth', \
'rgb_chin', 'hsv_chin', 'grad_mag_chin', 'grad_orien_chin', \
'rgb_moustache', 'hsv_moustache', 'grad_mag_moustache', 'grad_orien_moustache', \
'rgb_left_cheek', 'hsv_left_cheek', 'grad_mag_left_cheek', 'grad_orien_left_cheek', \
'rgb_right_cheek', 'hsv_right_cheek', 'grad_mag_right_cheek', 'grad_orien_right_cheek', \
'rgb_forehead', 'hsv_forehead', 'grad_mag_forehead', 'grad_orien_forehead', \
'rgb_full_face', 'hsv_full_face', 'grad_mag_full_face', 'grad_orien_full_face', \
'rgb_hair', 'hsv_hair', 'grad_mag_hair', 'grad_orien_hair']
for i in range(len(features_title)):
np.save(output_file_path + features_title[i], final_features[i])
# + [markdown] id="VO5O73WtXXB3"
# # Clear resources
# + id="LAD_jq1bXXB6"
# close_bboxes_file()
file_exception.close()
|
src/low level/LowLevelAlignedCelebA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # intro to matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
heart_disease = pd.read_csv('heart-disease.csv')
heart_disease
over_50 = heart_disease[heart_disease['age']>50]
over_50.head()
plt.style.use('seaborn-whitegrid')
# +
fig,(ax0,ax1) = plt.subplots(nrows=2,
ncols=1,
figsize=(10,10))
fig.suptitle('Heart Disease Analysis',fontweight='bold',fontsize=30)
ax0.set(title="chol vs age",xlabel="age",ylabel="chol")
ax1.set(title="thalach vs age",xlabel="age",ylabel="thalach")
ax0.set_xlim([50,80])
ax1.set_xlim([50,80])
ax1.set_ylim([60,200])
ax0.axhline(over_50['chol'].mean(),linestyle="--",color='red')
ax1.axhline(over_50['thalach'].mean(),linestyle="--",color='red')
scatter1 = ax0.scatter(x=over_50['age'],y=over_50['chol'],c=over_50['target'],cmap='winter')
scatter2 = ax1.scatter(x=over_50['age'],y=over_50['thalach'],c=over_50['target'],cmap='summer')
ax0.legend(*scatter1.legend_elements(),title='target')
ax1.legend(*scatter2.legend_elements(),title='target');
# -
|
intro_to_matplotlib.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''phuse-python-sde-sep2021-nm3hVWFa'': pipenv)'
# name: python3
# ---
# # The CDISC Library API
#
# There's no other way to put it - PDF is a terrible medium for data sharing. There is a standard, but implementations of it are wide and varied. Issues around handling tables, paragraphs, etc have made it a very human (and thereby accident prone) approach.
#
# The CDISC Library exposes the core metadata via a RESTful Web API. Access is available for members with fair use provisions and also for Open Source Contributors if you are eligible. To request an account go to [CDISC Library](https://www.cdisc.org/cdisc-library) and click on `Request an account`. Once you are setup you will be able to generate a token (a character string) that you can use to make requests against the API (this is primarily used to avoid people abusing the service)
# +
# as before, we import our requests library
import requests
# We use the python-dotenv library to merge in the CDISC_LIBRARY_API_TOKEN
# - this reads from a formatted file and adds the values to the environment
from dotenv import load_dotenv
# this loads in the values from the .env file
load_dotenv()
import os
# prove it worked
print(f"Loaded token {os.environ['CDISC_LIBRARY_API_TOKEN'][:5]}...")
# Supply this globally
CDISC_API_URL = "https://library.cdisc.org/api"
# +
# the token we loaded needs to be added to the headers using the name 'api-key'
# if we don't pass the token, what happens?
unauth = requests.get(f"{CDISC_API_URL}/mdr/products")
print(f"Requesting {unauth.url} got status {unauth.status_code}")
# +
# we pass the `api-key` in the headers
response = requests.get(f"{CDISC_API_URL}/mdr/products", headers={'api-key': os.environ['CDISC_LIBRARY_API_TOKEN']})
print(f"Requesting {response.url} got status {response.status_code}")
# +
# The request we made above was to the webservice that returns the list of products
# as we know the response is JSON we can access the model as a dictionary using the `json` method
results = response.json()
# What elements are in the result set?
print("Keys", results.keys())
# +
# Inspection of this object helps us find what we are looking for
links = results.get('_links')
print("Links:", links.keys())
# +
# the _links collection is a common pattern for representing where you can 'go' from a location
# self represents the current resource
print("self -> ", links['self'])
# this particular resource has sets of related resources (organised by product group)
for product_group in links.keys():
if product_group == "self":
continue
for product, details in links.get(product_group).get('_links').items():
print(f"{product_group} -> {product} -> {details}")
# +
# the href attribute is a value you can pick to resolve the entity
href = links["terminology"]["_links"]["self"]["href"]
response = requests.get(f"{CDISC_API_URL}{href}", headers={'api-key': os.environ['CDISC_LIBRARY_API_TOKEN']})
print(response.json())
# +
# you can reuse the client authentication details by creating a Session
from requests import Session
client = Session()
client.headers['api-key'] = os.getenv("CDISC_LIBRARY_API_TOKEN")
terminology = client.get(f"{CDISC_API_URL}{href}")
print(f"Packages: {terminology.json().get('_links').get('packages')}")
# +
# let's request a particular standard (say SDTMIG version 3-2)
sdtm_ig_32 = client.get(f"{CDISC_API_URL}/mdr/sdtmig/3-2")
# +
# decode the response
ig = sdtm_ig_32.json()
# inspect the response
print(ig.keys())
# this API call returns a class oriented structure
for ig_class in ig.get('classes'):
# note, General Observations has no direct datasets, so we need to guard
if "datasets" in ig_class:
# iterate over the datasets
for dataset in ig_class.get('datasets'):
# each dataset has a set of variables
for dataset_variable in dataset.get('datasetVariables'):
print(f"{ig_class.get('name')} -> {dataset.get('name')} -> [{dataset_variable.get('ordinal')}] {dataset_variable.get('name')}")
# +
# can go directly to the datasets
datasets = client.get(f"{CDISC_API_URL}/mdr/sdtmig/3-2/datasets").json()
# +
dm = None
# the href attribute is a link that you can reuse with the BASE URI
for dataset_link in datasets.get('_links').get('datasets'):
if dataset_link.get('title') == 'Demographics':
dm = client.get(f"{CDISC_API_URL}{dataset_link.get('href')}").json()
# +
print(dm.get('name'))
print(dm.get('description'))
print(dm.get('_links'))
# pull the variables for the dataset
for dv in dm.get('datasetVariables'):
print(f"{dm.get('name')} -> {dv.get('name')}")
# +
import pprint
# The dataset variables also have accessible attributes
# pull the variables for the dataset
for dv in dm.get('datasetVariables'):
if dv.get('name') == 'SEX':
print(pprint.pprint(dv))
# -
# # API Specifications
#
# The APIs themselves are sensible in terms of how they are laid out; we go from context (eg SDTM IG) to version (eg 3-3) to dataset (eg DM) to variable (eg SEX). The API is documented using a standard called OpenAPI (previously known as Swagger) - this is a format for publishing information about an API; information includes:
# * Routes (where to get the data)
# * Methods (how to access the data, usually using HTTP Verbs)
# * Formats (what parameters are required)
# * Structures (how the data looks)
# * Responses (what outcomes can you expect)
#
# The use of standard representations makes the developer view ([here](https://www.cdisc.org/cdisc-library/api-documentation))
#
# Using a standard OpenAPI based webservice makes it easier for developers to access and understand what the API exposes. You can use tools to automatically generate clients so code like the following is possible (this was built using [openapi-python-client](https://github.com/openapi-generators/openapi-python-client)):
#
# ```python
# # create a module and then use the module
# from cdisc_library_api_client.api.sdtm.ig.sdtmig_get_dataset import sync_detailed
#
# # URL construction and authentication are handled transparently
# client = AuthenticatedClient(base_url="https://library.cdisc.org/api", token=token)
#
# token = os.getenv("CDISC_LIBRARY_TOKEN")
#
# # Specifications
# version = "3-3"
# domain = "DM"
#
# # make the call
# mm = await sync_detailed(client=client, version=version, dataset=domain)
# dataset = mm.parsed
#
# # dataset is an instance of an SDTMigDataset
# print("Dataset: {dataset.name} ({dataset.label})")
# ```
#
#
|
workbooks/03-The-CDISC-Library-API.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Make Predictions for Upcoming IPO
# - Retrieves upcoming IPO from NASDAQ
# - Lists probabilities for upcoming peformance
# - 1D, 1W, 1M and 3M outcome prediction
# +
import numpy as np
import pandas as pd
import nasdaq
import ml
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import VotingClassifier
from pandas_datareader import data as pdr
import fix_yahoo_finance as yf
yf.pdr_override()
# %matplotlib inline
import matplotlib.pyplot as plt
import time
import datetime
# -
#params
today = datetime.datetime.today()
start_date = str(datetime.datetime(today.year, today.month-1, 1).date())
print('predictions for', start_date)
# # Create NASDAQ IPO List
df_symbols = nasdaq.get_ipo_list(start_date)
print('symbols', df_symbols.shape[0])
df_symbols.head()
# # Train Models (RF/LR - Train/Test Split)
#load most up to date data
df = pd.read_csv('../Datasets/4 keywords.csv', index_col='Symbol')
# #quarter and month
# months = df.loc[:, 'Q1':'MKT12'].columns
# classes = df.loc[:, '1D':'3M'].columns
# others = df.columns.difference(months).difference(classes)
#
# #standardize
# df = pd.concat([ml.standardize(df[others]), df[months], df[classes]], axis=1)
#feature importance
indices = ml.show_feature_importance(df, '3M')
#feature selection
df1 = ml.select_features(indices.shape[0], indices, df)
#run ML flow
ml.run_ml_flow(df1)
# # Train Model (Ensemble - Entire Data)
# +
#prepare test set
targets = ['1D', '1W', '1M', '3M']
df_symbols = df.loc[df_symbols.index]
#drop targets
df_symbols.drop(targets, axis=1, inplace=True)
#drop columns with nanas
df_symbols.dropna(axis=1, inplace=True)
df_symbols.head()
# -
# align train and test frames
df = df.dropna()
df = df[list(df_symbols.columns) + targets]
# +
#train & predict for all targets
prediction = pd.DataFrame(columns=targets, index=df_symbols.index)
X_test = df_symbols.values
for target in targets:
#split
X_train = df.values[:,:-4]
y_train = df[target].map(lambda x: 1 if x > 0 else 0).values
#classifier
clf = VotingClassifier(estimators=[('lr', LogisticRegression(random_state=1)), ('rf', RandomForestClassifier(n_estimators=50, max_depth=5, random_state=1))], voting='soft')
#fit
clf.fit(X_train, y_train)
#predict
probas = clf.predict_proba(X_test)
prediction[target] = probas[:,1]
# -
# # Show Predictions
prediction.plot(figsize=(15,5), title='Probabilities')
prediction.tail(10)
|
Notebooks/Predictor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GEM Tutorial with COBRApy
#
# This exercise will walk through some of the basic operations in working with a genome-scale metabolic model (GEM). The vast majority of software that has been developed surrounding GEMs has been done in MATLAB, likely because this form of modeling has origins in engineering (specifically chemical engineering). Although well-suited for metabolic modeling, MATLAB is not open-source and therefore limits the accessibility of this software. Fortunately, the modeling community has implemented the MATLAB COnstrant-Based Reconstruction and Analysis [(COBRA) Toolbox](https://opencobra.github.io/cobratoolbox/stable/) in Python, as [**COBRApy**](https://opencobra.github.io/cobrapy/).
#
# **COBRApy** is still relatively young and therefore lacks some of the functionality of its MATLAB counterparts, but the core utilities are available and quickly expanding. Here, we will demonstrate some of the basic functions and classes of the **COBRApy** package, which should also familiarize the user with the fundamentals of GEM structure and simulation.
#
# Most of the commands and material covered in this tutorial can be found in the [**COBRApy Documentation**](https://cobrapy.readthedocs.io/en/stable/), so we encourage you to reference the documentation if you encounter errors, warnings, or need further detail about something. You can of course always ask us for help too :)
import cobra
import cobra.test
import os
# ## View the global configuration object
#
# Before jumping right into things, it is always nice to see what sort of default settings are in place. **COBRApy** has organized such defaults into a **global configuration object**, which can be viewed or adjusted as needed.
cobra_config = cobra.Configuration()
# view a brief summary of the object
cobra_config
# view the default reaction flux bounds (min, max)
cobra_config.bounds
# ## Import and inspect a small test model
#
# GEMs, as their name implies ("_genome_"-scale), are often quite large, containing thousands of reactions, metabolites, and genes. It is therefore best to begin working with a simplified model that is quick to load and use, and easy to conceptualize.
#
# For this exercise, we will use the `textbook` model that is provided with the **COBRApy** package. This model encompasses the core pathways of central carbon metabolism in the _E. coli_ bacterium.
# the cobra package ships with several test models in different formats
data_dir = cobra.test.data_dir
os.listdir(data_dir)[:10]
# load the "textbook" model from the SBML (.xml) file
model = cobra.io.read_sbml_model(os.path.join(data_dir, "textbook.xml.gz"))
model
# **Note:** SBML ([Systems Biology Markup Language](http://sbml.org/Main_Page)) is an XML-based format commonly used to store GEMs. The aim of SBML is to serve as an open and standardized format to facilitate sharing of models and software.
# list the first few reactions in the model
for x in model.reactions[:10]:
print("%s : %s" % (x.id, x.reaction))
# inspect a reaction (e.g., AKGDH) in more detail
model.reactions.AKGDH
# list the first few metabolites in the model
for x in model.metabolites[:10]:
print("%s : %s" % (x.id, x.formula))
# inspect the 3pg_c metabolite in more detail
model.metabolites.get_by_id('3pg_c')
# ## Add a new reaction to the model
# For this example, we will add the aspartate aminotransferase reaction to enable the synthesis of aspartate:
#
# `L-glutamate + oxaloacetate <==> 2-oxoglutarate + L-aspartate`
# ### Create and edit the reaction object
# create a template reaction and determine what information we need to provide
reaction = cobra.Reaction('ASPAMTR')
reaction
# add the reaction name
reaction.name = 'aspartate aminotransferase'
# +
# we need to find the IDs of the metabolites in the reaction
met_patterns = ['glutamate', 'oxaloacetate', 'oxoglutarate', 'aspartate']
for met in model.metabolites:
if any([x in met.name.lower() for x in met_patterns]):
print("%s : %s" % (met.id, met.name))
# -
# Two interesting observations:
# 1. There are two instances of `2-Oxoglutarate` and `L-Glutamate`
# 2. Aspartate is not yet in the model
#
# For the first point, note that the `_c` and `_e` suffixes represent the compartment to which the metabolite belongs.
# view model compartments
model.compartments
# We want to add our reaction to the cytosol (`c`) compartment, so we will use the `_c` form of the metabolites.
#
# For the second point, we will need to add aspartate to the model.
# ### Create a new metabolite object
# create the aspartate metabolite
asp_c = cobra.Metabolite('asp_c')
asp_c # view its (missing) properties
# fill in some information about the new aspartate metabolite
asp_c.name = 'L-Aspartate'
asp_c.formula = 'C4H6NO4'
asp_c.compartment='c'
# now we can add the metabolites to the new aspartate aminotransferase reaction
reaction.add_metabolites({
model.metabolites.glu__L_c: -1.0,
model.metabolites.oaa_c: -1.0,
model.metabolites.akg_c: 1.0,
asp_c: 1.0
})
# view the reaction details to verify that it looks correct
reaction
# update the reversibility of the reaction (should be reversible)
reaction.reversibility
reaction.reversibility = True # we cannot directly edit the "reversibility" field
# instead we need to change the lower bound of the reaction
reaction.lower_bound = -1000
reaction.reversibility # verify that the reversibilty has been updated
# note that the equation now shows the double-sided arrow "<=>"
reaction
# #### Add a gene-protein-reaction (GPR) rule to the reaction
# aspartate aminotrasferase is encoded by aspC (b0928) in E. coli
reaction.gene_reaction_rule = 'b0928'
reaction
# gene(s) in the GPR rule are automatically added to the "genes" field of the reaction object
reaction.genes
# ### Add the reaction to the model
# add the reaction (provided as a list) to the model
model.add_reactions([reaction])
# verify that the new reaction, metabolite, and gene are now in the model
model.reactions.ASPAMTR
model.metabolites.asp_c
model.genes.b0928.name = 'aspC' # we can also provide the gene name
model.genes.b0928
# ## Flux balance analysis (FBA)
# ### Inspect the optimization objective
# using cobra.util.solver:
from cobra.util.solver import linear_reaction_coefficients
linear_reaction_coefficients(model)
# alternative: use list comprehension
[x for x in model.reactions if x.objective_coefficient != 0]
# view reaction details
model.reactions.Biomass_Ecoli_core
# print entire reaction stoichiometry
model.reactions.Biomass_Ecoli_core.build_reaction_string()
# view the objective direction (maximize or minimize the reaction flux)
model.objective_direction
# ### Perform the optimization
# run FBA
solution = model.optimize()
solution
# view a summary of the returned optimal flux distribution
model.summary()
# get a summary of the fluxes involving a specific metabolite
model.metabolites.atp_c.summary()
# get a summary of the fluxes involving a specific reaction
model.reactions.GAPD.summary()
# ### Change the optimization objective
# let us now optimize the flux through ATPM ("ATP Maintenance"), which is just the hydrolysis of ATP
model.reactions.ATPM.build_reaction_string()
# change the objective to ATPM
model.objective = 'ATPM'
# run FBA with the new objective
solution = model.optimize()
# summarize the results
model.summary()
# note that there is now no metabolic flux through the biomass reaction
#model.reactions.Biomass_Ecoli_core.summary() # gives an error because zero flux
solution.fluxes.Biomass_Ecoli_core
# ## Perform an _in silico_ knock out
# first optimize biomass production to view the initial maximum flux value
model.objective = 'Biomass_Ecoli_core'
biomass_original = model.optimize().objective_value
# knock out the AKGDH reaction
model.reactions.AKGDH.knock_out()
model.reactions.AKGDH # note that the upper and lower bound are now both zero
# The reaction is still present in the model, but it now cannot carry any flux. If we wanted to completely remove it from the model altogether, we could use the `remove_from_model` function: `model.reactions.AKGDH.remove_from_model()`
# now check the reaction again
model.reactions.AKGDH
# in reality, genes are knocked out, not reactions
# knock out a gene, and see what effect it has
model.genes.b0008.knock_out()
model.genes.b0008
inactive_rxns = [rxn.id for rxn in model.reactions if rxn.upper_bound == rxn.lower_bound]
inactive_rxns
model.reactions.ACALD
# need to knock out two isozymes for reaction (ACALD) to be inactivated
model.genes.b0351.knock_out()
model.genes.b1241.knock_out()
inactive_rxns = [rxn.id for rxn in model.reactions if rxn.upper_bound == rxn.lower_bound]
inactive_rxns
|
scripts/COBRApy_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1KBTLgUrzmS7"
# # For External users
#
# You can open this notebook in [Google Colab](https://colab.research.google.com/github/google/meterstick/blob/master/confidence_interval_display_demo.ipynb).
# + [markdown] id="H9ojnghz0b2N"
# ## Installation
#
# You can install from pip for the stable version
# + id="hbwqlbFm1nBo"
#@test {"skip": true}
# !pip install meterstick
# + [markdown] id="MZXKtCHy0CEo"
# or from GitHub for the latest version.
# + id="uOjzL05A1n3c"
#@test {"skip": true}
# !git clone https://github.com/google/meterstick.git
import sys, os
sys.path.append(os.getcwd())
# + [markdown] id="ioi4Gsmr0zmK"
# # Demo Starts
# + id="G_dbf982G612"
import itertools
import numpy as np
import pandas as pd
from meterstick import confidence_interval_display
# + id="puxiUfKdc4b1"
np.random.seed(42)
metrics = ('Click', 'Latency', 'a very very very looooooooooong metric')
ctrl_id = 42
platform = ('Mobile', 'Desktop', 'Tablet')
ctrl_vals = [137, 28, 999.9, 158, 40, 6.66666, -10, -20.1, 33]
expr_ids = (42, 222, 666, 'Experiment Foo')
n_row = len(metrics) * len(platform) * len(expr_ids)
test_df = pd.DataFrame(
itertools.product(expr_ids, platform, metrics),
columns=[
'Experiment_Id',
'Platform',
'Metric',
])
test_df['Control_Value'] = ctrl_vals * (n_row // len(ctrl_vals))
test_df['Ratio'] = 2 * (np.random.rand(n_row) - 0.5)
test_df['CI_Range'] = 2 * np.random.rand(n_row)
test_df['Value'] = test_df.Control_Value * (test_df.Ratio / 100 + 1)
test_df['Country'] = 'US'
ctrl_id = 42
test_df.loc[test_df.Experiment_Id == ctrl_id,
'Value'] = test_df.loc[test_df.Experiment_Id == ctrl_id,
'Control_Value']
# + colab={"height": 717} executionInfo={"elapsed": 431, "status": "ok", "timestamp": 1603603597605, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="joi2JjprON4Y" outputId="0e02748c-2c26-4215-a75b-85fb9a902332"
confidence_interval_display.render(test_df, dims=['Country', 'Platform'])
# + colab={"height": 717} executionInfo={"elapsed": 410, "status": "ok", "timestamp": 1603603637788, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="TaU7Xqfx7jzK" outputId="fec02af3-3dec-48fc-f9a3-b39dd086d7db"
# You can manually specify the columns.
test_df_copy = test_df.copy()
test_df_copy.rename(
columns={
'Experiment_Id': 'expr',
'Metric': 'metric',
'Control_Value': 'control',
'Ratio': 'ratio',
'CI_Range': 'ci',
'Value': 'val'
},
inplace=True)
confidence_interval_display.render(
test_df_copy,
dims=['Country', 'Platform'],
metric='metric',
ratio='ratio',
value='val',
ci_range='ci',
control_value='control',
expr_id='expr')
# + colab={"height": 717} executionInfo={"elapsed": 439, "status": "ok", "timestamp": 1603603641447, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="wAUoIbKjk4DA" outputId="1a3f24c6-c53a-470c-9edc-a994b0b3be33"
test_df_copy = test_df.copy()
test_df_copy['Control_Id'] = None
confidence_interval_display.render(test_df_copy, dims=['Country', 'Platform'])
# + colab={"height": 717} executionInfo={"elapsed": 412, "status": "ok", "timestamp": 1603603644346, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="0r2GRgrJ9hpv" outputId="c9bb099e-a011-4206-9aa7-1d1bcaada749"
# If you don't want to aggregate your dimensions...
confidence_interval_display.render(
test_df,
dims=['Country', 'Platform'],
aggregate_dimensions=False)
# + colab={"height": 717} executionInfo={"elapsed": 409, "status": "ok", "timestamp": 1603603647863, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="gh_CwEoe9m8X" outputId="1678e8a8-cd84-4ef1-e93c-f1857cc60bc7"
# By default we look for Dim_1, Dim2, ... as dimension columns.
test_df.rename(columns={'Country': 'Dim_1', 'Platform': 'Dim_2'}, inplace=True)
confidence_interval_display.render(test_df)
# + colab={"height": 717} executionInfo={"elapsed": 425, "status": "ok", "timestamp": 1603603651949, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="MCgJZN-ZZDz4" outputId="c501db71-cc74-4213-eccb-d61f24bfbb9f"
# You can set control experiment. We'll only display its Value.
confidence_interval_display.render(test_df, ctrl_id=ctrl_id)
# + colab={"height": 685} executionInfo={"elapsed": 415, "status": "ok", "timestamp": 1603603662265, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="I6c7JIsvfcBR" outputId="1576c377-b495-49ce-8a7e-38a604ffc49e"
# Hide the controls if it doesn't matter to you.
confidence_interval_display.render(test_df, ctrl_id=ctrl_id, show_control=False)
# + colab={"height": 717} executionInfo={"elapsed": 421, "status": "ok", "timestamp": 1603603666270, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="FYhPFCYhe2Pt" outputId="5045d7f2-6f38-47ce-dc37-fd3aa9735c2f"
# You can also have multiple controls.
confidence_interval_display.render(
test_df,
ctrl_id={
42: [222],
666: ['Experiment Foo']
})
# + colab={"height": 768} executionInfo={"elapsed": 448, "status": "ok", "timestamp": 1603603669455, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="xErZEoufOR2_" outputId="6f33fe1c-f797-482e-f8d0-48f3685ccaca"
# Auto determines control values if control rows are missing. Your control rows
# are supposed to have the same values in the Control_Value and Value columns.
# If not, we skip. To demonstrate, here we introduce inconsistency in 'Click'.
df = test_df.loc[test_df.Experiment_Id != 42].copy()
df.loc[(df.Experiment_Id == 222) & (df.Metric == 'Click'), 'Control_Value'] = -1
confidence_interval_display.render(
df,
ctrl_id=42,
auto_decide_control_vals=True)
# + colab={"height": 717} executionInfo={"elapsed": 433, "status": "ok", "timestamp": 1603603672824, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="DcWZkt-fVGBG" outputId="26340259-c193-4808-df7f-8c77dc1753a4"
# You can flip the coloring scheme for some metrics.
confidence_interval_display.render(test_df, flip_color=['Latency'])
# + colab={"height": 717} executionInfo={"elapsed": 596, "status": "ok", "timestamp": 1603603677236, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="eoFBNAVZ5ou7" outputId="47b9c761-c64f-4a5a-ecbd-5ad38852017d"
# You can add descriptions for your experiments.
test_df['Description'] = ''
test_df.loc[test_df['Experiment_Id'] == 42, 'Description'] = 'The Answer'
# You can even use raw html.
test_df.loc[test_df['Experiment_Id'] == 666,
'Description'] = '<b>Bold</b>'
confidence_interval_display.render(test_df)
# + colab={"height": 717} executionInfo={"elapsed": 416, "status": "ok", "timestamp": 1603603702299, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="6xeDUqHveDtB" outputId="01ae7395-7fc0-4377-e9f0-4240fdb51466"
# You can customize the formatting of values. By default there are 'percent',
# 'absolute' and 'pp'. You can also provide your own formating string template.
confidence_interval_display.render(
test_df,
metric_formats={
'Ratio': 'pp',
'Value': 'percent'
})
# + colab={"height": 717} executionInfo={"elapsed": 422, "status": "ok", "timestamp": 1603603703160, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="BV8-gF2fwhKX" outputId="f0f91a26-0755-4c7b-ada9-9c73eca0b60c"
# You don't need Value if you don't want to show_control.
test_df2 = test_df.copy()
del test_df2['Value']
confidence_interval_display.render(test_df2, show_control=False)
# + colab={"height": 700} executionInfo={"elapsed": 427, "status": "ok", "timestamp": 1603603706415, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="pLJ7N_vaXqHW" outputId="4e3e484d-37cc-4128-dc57-aa0684955d02"
# You can hide the null control values.
confidence_interval_display.render(
test_df2, show_control=False, hide_null_ctrl=True)
# + colab={"height": 717} executionInfo={"elapsed": 423, "status": "ok", "timestamp": 1603603709179, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="KMRIaPNvmT-V" outputId="78d6fc5b-f40c-4517-d58b-005e1ba0a85b"
# You can order and slice the metrics.
confidence_interval_display.render(test_df, metric_order=['Latency', 'Click'])
# + colab={"height": 717} executionInfo={"elapsed": 480, "status": "ok", "timestamp": 1603603710351, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="wTcWPmT7mr1q" outputId="ada89cc5-0a27-4742-9c4d-6c3fd8a0c8c8"
# You can customize the order of dimension slices.
confidence_interval_display.render(
test_df,
sort_by=[{
'column': 'Dim_2',
'order': ['Mobile', 'Desktop', 'Tablet']
}])
# + colab={"height": 717} executionInfo={"elapsed": 412, "status": "ok", "timestamp": 1603603712642, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="9rGrLYD5GZeO" outputId="df392a24-8acf-470d-c9ea-bfdd352b1c61"
# You can sort by values of a metric. The supported value columns are 'Value',
# 'Ratio', 'CI_Lower', 'CI_Upper'. See the doc of get_formatted_df() also.
confidence_interval_display.render(
test_df,
sort_by=[{
'column': ('Ratio', 'Click'),
'ascending': False
}])
# + colab={"height": 717} executionInfo={"elapsed": 452, "status": "ok", "timestamp": 1603603721016, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="rEZJ4IDpmqWD" outputId="5107c853-9728-445a-bd1b-5f24f3339d74"
# Of course you can sort both by dimesnion and by values.
confidence_interval_display.render(
test_df,
sort_by=[{
'column': 'Dim_2',
'order': ['Mobile', 'Desktop', 'Tablet']
}, {
'column': ('CI_Lower', 'Click'),
}])
# + colab={"height": 717} executionInfo={"elapsed": 486, "status": "ok", "timestamp": 1603603721562, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="3RdDlmTznJ0n" outputId="69c27033-544c-41aa-83d7-02c483be2c11"
pre_agg_df = confidence_interval_display.render(
test_df, return_pre_agg_df=True)
mask = pre_agg_df.applymap(lambda c: bool(
isinstance(c, list) and c[-1] and c[-2] and c[-1] - c[-2] < 1))
formatted_df = confidence_interval_display.render(
test_df, return_formatted_df=True)
formatted_df.mask(
mask,
formatted_df.applymap('<div class="tight-ci">{}</div>'.format),
inplace=True)
extra_css = '.tight-ci {border-color: purple; border-style: dotted;}'
confidence_interval_display.display_formatted_df(formatted_df, extra_css)
# + colab={"height": 330} executionInfo={"elapsed": 398, "status": "ok", "timestamp": 1603603722000, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="Uv-TjCshF_xa" outputId="4922eece-0aaf-4944-ca68-2e1e11082ece"
# Test for no dimensions.
df_no_dim = test_df.groupby(['Experiment_Id', 'Metric']).mean().reset_index()
confidence_interval_display.render(df_no_dim, ctrl_id=42)
|
confidence_interval_display_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: up42-py-test
# language: python
# name: up42-py-test
# ---
# # Example: Radar processing
#
# The processing of radar data - often referred to as SAR data, which stands for Synthetice Aperture Radar - is a topic of its own and quite different from optical data that uses Red, Green, Blue, NIR and similar bands.
# This is the first of a series of jupyter notebooks where example SAR workflows will be explained that can be run on UP42.
# All of the examples are based on a specific type of analysis which is called polarimetry and processes one image at a time.
# In comparison to polarimetric there also is interferometric analysis which leverages the information encoded in suitable pairs of radar image.
#
# In this notebook we want to have a look at the Sentinel-1 GRD full scene block, followed by SNAP Sentinel-1 Polarimetric Processing. Polarimetric SAR Processing turns a Sentinel-1 GRD image into a multichannel GeoTIFF that is ready for analysis.
#
# - Prepare workflow
# - Define aoi and input parameters
# - Run Job
# - Visualize results
import up42
# Authenticate with UP42
up42.authenticate(cfg_file="config.json")
#up42.authenticate(project_id=12345, project_api_key=12345)
# ## Prepare UP42 workflows
#
# Create a new project on UP42 or use an existing one.
S1_SNAP_project = up42.initialize_project()
# Create workflow and check available blocks and data
workflow = S1_SNAP_project.create_workflow(name="S1-GRD_SNAP",
use_existing=True)
print(up42.get_blocks(basic=True))
# Fill the workflow with tasks
input_tasks= ['sobloo-s1-grd-fullscene', 'snap-polarimetric']
workflow.add_workflow_tasks(input_tasks=input_tasks)
workflow.get_parameters_info()
# ## Define aoi and input parameters
#
# The S1 GRD block always delivers the complete images as they are delivered in SAFE format which cannot be clipped.
# The image can be clipped though by the SNAP polarimetric processing blocks. For this it is necessary to supply the same geometry (in this case a bbox) to that block as well.
#
#
input_parameters = {
"sobloo-s1-grd-fullscene:1": {
"bbox": [13.371037, 52.512799, 13.382624, 52.518747],
"ids": None,
"time": "2018-01-01T00:00:00+00:00/2020-12-31T23:59:59+00:00",
"limit": 1,
"zoom_level": 14,
},
"snap-polarimetric:1": {
"bbox": [13.371037, 52.512799, 13.382624, 52.518747],
"mask": None,
"contains": None,
"intersects": None,
"clip_to_aoi": True,
"tcorrection": True,
"linear_to_db": True,
"polarisations": [
"VV"
],
"speckle_filter": True,
"calibration_band": [
"sigma"
]
}
}
# ## Run test query
# Run a test job to query data availability and check the configuration.
test_job = workflow.test_job(input_parameters=input_parameters, track_status=True)
test_results = test_job.get_results_json()
print(test_results)
# ## Run the job
job = workflow.run_job(input_parameters=input_parameters, track_status=True)
# ## Visualize the results
# +
# Download results:
job.download_results()
# Visualize downloaded results
job.plot_results()
# -
# ## Next processing steps
# The output of the workflow is a GeoTIFF file which can be processed by different algorithms. It is for example possible to apply raster tiling and then do some machine learning based analysis at the end.
#
#
|
examples/radar_processing_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Ubuntu Linux)
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
# !pip install nltk
import nltk
nltk.download('punkt')
from data_loader import get_loader
from torchvision import transforms
transform_train = transforms.Compose([
transforms.Resize(256),
transforms.RandomCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406),
(0.229, 0.224, 0.225))])
vocab_threshold = 6
batch_size = 10
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# -
sample_caption = 'A person doing a trick on a rail while riding a skateboard.'
# +
import nltk
sample_tokens = nltk.tokenize.word_tokenize(str(sample_caption).lower())
print(sample_tokens)
# +
sample_caption = []
start_word = data_loader.dataset.vocab.start_word
print('Special start word:', start_word)
sample_caption.append(data_loader.dataset.vocab(start_word))
print(sample_caption)
# -
sample_caption.extend([data_loader.dataset.vocab(token) for token in sample_tokens])
print(sample_caption)
# +
end_word = data_loader.dataset.vocab.end_word
print('Special end word:', end_word)
sample_caption.append(data_loader.dataset.vocab(end_word))
print(sample_caption)
# +
import torch
sample_caption = torch.Tensor(sample_caption).long()
print(sample_caption)
# -
dict(list(data_loader.dataset.vocab.word2idx.items())[:10])
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
# +
# Modify the minimum word count threshold.
vocab_threshold = 6
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# -
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
# +
unk_word = data_loader.dataset.vocab.unk_word
print('Special unknown word:', unk_word)
print('All unknown words are mapped to this integer:', data_loader.dataset.vocab(unk_word))
# -
print(data_loader.dataset.vocab('jfkafejw'))
print(data_loader.dataset.vocab('ieowoqjf'))
# Obtain the data loader (from file). Note that it runs much faster than before!
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_from_file=True)
# +
from collections import Counter
counter = Counter(data_loader.dataset.caption_lengths)
lengths = sorted(counter.items(), key=lambda pair: pair[1], reverse=True)
for value, count in lengths:
print('value: %2d --- count: %5d' % (value, count))
# +
import numpy as np
import torch.utils.data as data
indices = data_loader.dataset.get_train_indices()
print('sampled indices:', indices)
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
images, captions = next(iter(data_loader))
print('images.shape:', images.shape)
print('captions.shape:', captions.shape)
print('images:', images)
print('captions:', captions)
# +
% load_ext autoreload
% autoreload 2
from model import EncoderCNN, DecoderRNN
# -
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# +
embed_size = 512
encoder = EncoderCNN(embed_size)
encoder.to(device)
images = images.to(device)
features = encoder(images)
print('type(features):', type(features))
print('features.shape:', features.shape)
assert type(features)==torch.Tensor, "Encoder output needs to be a PyTorch Tensor."
assert (features.shape[0]==batch_size) & (features.shape[1]==embed_size), "The shape of the encoder output is incorrect."
# +
hidden_size = 512
vocab_size = len(data_loader.dataset.vocab)
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
decoder.to(device)
captions = captions.to(device)
outputs = decoder(features, captions)
print('type(outputs):', type(outputs))
print('outputs.shape:', outputs.shape)
assert type(outputs)==torch.Tensor, "Decoder output needs to be a PyTorch Tensor."
assert (outputs.shape[0]==batch_size) & (outputs.shape[1]==captions.shape[1]) & (outputs.shape[2]==vocab_size), "The shape of the decoder output is incorrect."
|
1_Preliminaries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.5 64-bit (''opencvenv'': conda)'
# language: python
# name: python37564bitopencvenvcondac6bef57599574e11b26dd8fdcebd1d77
# ---
from demo import np_to_torch, pred, scale_input
from dataloader import read_image, read_trimap
from networks.models import build_model
import torch
import numpy as np
import cv2
import matplotlib.pyplot as plt
class Args:
encoder = 'resnet50_GN_WS'
decoder = 'fba_decoder'
weights = 'FBA.pth'
args=Args()
try:
model = build_model(args)
except:
# !gdown https://drive.google.com/uc?id=1T_oiKDE_biWf2kqexMEN7ObWqtXAzbB1
model = build_model(args)
image = read_image('./examples/images/troll.png')
trimap = read_trimap('./examples/trimaps/troll.png')
# +
plt.title('Input Image')
plt.imshow(image)
plt.show()
# transform two channel trimap back to single channel
trimap_im = trimap[:,:,1] + (1-np.sum(trimap,-1))/2
plt.title('Trimap')
plt.imshow(trimap_im, cmap='gray', vmin=0, vmax=1)
plt.show()
# -
fg, bg, alpha = pred(image, trimap, model)
plt.title('Alpha Matte')
plt.imshow(alpha, cmap='gray', vmin=0, vmax=1)
plt.show()
plt.title('Foreground')
plt.imshow(fg)
plt.show()
plt.title('Background')
plt.imshow(bg)
plt.show()
plt.title('Composite')
plt.imshow(fg*alpha[:,:,None])
plt.show()
|
FBA Matting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import requests
sys.path.insert(0, "C:/Users/julie/PycharmProjects/SoundLandscape/")
# -
# # Rules
response = requests.post('http://127.0.0.1:5000/api/music/get_params')
res = response.json()
# # Speed enery
response = requests.post('http://127.0.0.1:5000/api/music/get_speed_energy', json={"speed": 60})
response.json()
# # Driver pref
response = requests.post('http://127.0.0.1:5000/api/music/get_driver_preferences', json={"driver": "julien"})
response.json()
# # Mood
response = requests.post('http://127.0.0.1:5000/api/music/get_mood', json={"mood": "sad"})
response.json()
# # Get day_time
data={"now_time": '2019-03-15T06:03:33+00:00', 'sunrise_time': '2019-03-15T06:03:33+00:00', 'sunset_time': '2019-03-15T17:56:07+00:00'}
response = requests.post('http://127.0.0.1:5000/api/music/get_day_time', json=data)
response.json()
# # Get reco
data = {'popularity': 90, 'seed_genres':['r-n-b'], 'country':'FR'}
response = requests.post('http://127.0.0.1:5000/api/music/get_recommendations', json=data)
response.json()
|
notebooks/07 src.webservice.music.py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="z0B9tfzp2Wiw" executionInfo={"status": "ok", "timestamp": 1629216163904, "user_tz": -540, "elapsed": 2942, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}}
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="ADyg5aKl2aJD" executionInfo={"status": "error", "timestamp": 1629216164406, "user_tz": -540, "elapsed": 507, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="932c4799-9c88-496c-dede-9432f76082e1"
# 데이터 입력
df = pd.read_csv('sample_data/sonar.csv')
# 데이터 분류
dataset = df.values
X = dataset[:,0:60].astype(float)
Y_obj = dataset[:,60]
# 문자열을 숫자로 변환
e = LabelEncoder()
e.fit(Y_obj)
Y = e.transform(Y_obj)
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="IHwvQMZR2brq" executionInfo={"status": "error", "timestamp": 1629216167762, "user_tz": -540, "elapsed": 296, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="367224a9-4a22-4eef-972d-2f2682f0a6f0"
# 전체 데이터에서 학습 데이터와 테스트 데이터로 구분
X_train1, X_test, Y_train1, Y_test = train_test_split(X, Y, test_size=0.2,shuffle=True) ## shuffle=True로 하면 데이터를 섞어서 나눔
## 학습 셋에서 학습과 검증 데이터로 구분
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.2, shuffle=True) ## shuffle=True로 하면 데이터를 섞어서 나눔
# + colab={"base_uri": "https://localhost:8080/", "height": 134} id="qDId3VmU2dQ7" executionInfo={"status": "error", "timestamp": 1629216173225, "user_tz": -540, "elapsed": 311, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="d5a5390e-2866-4b0d-c690-0b1137274d3f"
model=
# 모델 컴파일
loss=
optimizer =
metrics=
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="tYYSoYOH2emt" executionInfo={"status": "error", "timestamp": 1629216194770, "user_tz": -540, "elapsed": 278, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="1c8e6935-3433-4684-9c70-77946594f0a0"
## model fit은 histoy를 반환한다. 훈련중의 발생하는 모든 정보를 담고 있는 딕셔너리.
result=model.fit(X_train, Y_train, epochs=50, batch_size=50, validation_data=(X_valid,Y_valid)) # validation_data=(X_valid,Y_valid)을 추가하여 학습시 검증을 해줌.
## histoy는 딕셔너리이므로 keys()를 통해 출력의 key(카테고리)를 확인하여 무엇을 받고 있는지 확인.
print(result.history.keys())
### result에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### result에서 binary_accuracy와 val_binary_accuracy key를 가지는 값들만 추출
acc = result.history['binary_accuracy']
val_acc = result.history['val_binary_accuracy']
### binary_accuracy와 val_binary_accuracy key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, acc, 'ro', label='Training acc')
plt.plot(epochs, val_acc, 'r', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
## 그래프 띄우기
plt.show()
# + id="BvBzWiZN2j4R"
# model.evalueate를 통해 테스트 데이터로 정확도 확인하기.
## model.evaluate(X_test, Y_test)의 리턴값은 [loss, binary_acuuracy ] -> 위 model.compile에서 metrics=[ keras.metrics.binary_accuracy]옵션을 주어서 binary acuuracy 출력됨.
print("\n Test Accuracy: %.4f" % (model.evaluate(X_test, Y_test)[1]))
|
tensorflow/day3/exercise/03_05_sonar_retrain.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Rabbits and Recurrence Relations
# ## Problem
#
# A sequence is an ordered collection of objects (usually numbers), which are allowed to repeat. Sequences can be finite or infinite. Two examples are the finite sequence (π,−2‾√,0,π) and the infinite sequence of odd numbers (1,3,5,7,9,…). We use the notation an to represent the n-th term of a sequence.
#
# A recurrence relation is a way of defining the terms of a sequence with respect to the values of previous terms. In the case of Fibonacci's rabbits from the introduction, any given month will contain the rabbits that were alive the previous month, plus any new offspring. A key observation is that the number of offspring in any month is equal to the number of rabbits that were alive two months prior. As a result, if Fn represents the number of rabbit pairs alive after the n-th month, then we obtain the Fibonacci sequence having terms Fn that are defined by the recurrence relation Fn=Fn−1+Fn−2 (with F1=F2=1 to initiate the sequence). Although the sequence bears Fibonacci's name, it was known to Indian mathematicians over two millennia ago.
#
# When finding the n-th term of a sequence defined by a recurrence relation, we can simply use the recurrence relation to generate terms for progressively larger values of n. This problem introduces us to the computational technique of dynamic programming, which successively builds up solutions by using the answers to smaller cases.
#
# Given: Positive integers n≤40 and k≤5.
#
# Return: The total number of rabbit pairs that will be present after n months, if we begin with 1 pair and in each generation, every pair of reproduction-age rabbits produces a litter of k rabbit pairs (instead of only 1 pair).
# +
dp = list(range(41))
n,k = map(int,input().split())
dp[0], dp[1] = 1,1
for i in range(2,n+1):
dp[i] = dp[i-1] + k*dp[i-2]
print(dp[n-1])
# -
|
Bioinformatics Stronghold/LEVEL 1/FIB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/aryu99/IBM-Hackathon/blob/master/wind_power_forecasting_final.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="y_dPfP8c7SnP" colab_type="code" colab={}
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
import os
import torch
mpl.rcParams['figure.figsize'] = (8, 6)
mpl.rcParams['axes.grid'] = False
# + id="_7KO2sVC8iaT" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 72} outputId="a01de6c3-ca9b-4c3d-c57b-81d767ee3693"
from google.colab import files
uploaded = files.upload()
# + id="u5xa8qoK8mwj" colab_type="code" colab={}
dataset = pd.read_csv("dataset 3.csv", encoding= 'unicode_escape')
TRAIN_SPLIT = 52561
# + id="Sb_AEYW28mu8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="99177ff8-8e35-49d1-977f-d0d052438eb0"
print (dataset)
# + id="_OAkwKt4eqbT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="19ed2bb1-9920-43dd-ffe9-6a4c222bde59"
dataset.info
# + id="S0MgGGJN8mqc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 369} outputId="caa7997a-9b8f-4fdb-c25f-f921285ed179"
df = dataset
features_considered = ['Wind Speed (m/s)', 'LV ActivePower (kW)', 'Wind Direction (°)']
features = df[features_considered]
features.index = df['Date/Time']
features.head()
features.plot(subplots=True)
# + id="Yz1Bk8yJm_bh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6cb33687-17be-4233-d107-9d75595d9182"
dataset_mean = df.mean(axis = 0)
dataset_std = df.std(axis = 0)
print(dataset_mean[1])
# + id="Nge18Lxo8mjZ" colab_type="code" colab={}
df3 = df
df2 = df
# + id="wfj_fEk78ma7" colab_type="code" colab={}
features_considered = ['Wind Speed (m/s)', 'Wind Direction (°)']
features = df3[features_considered]
features.index = df3['Date/Time']
features.tail()
dataset = features.values
data_mean = dataset[:TRAIN_SPLIT].mean(axis=0)
data_std = dataset[:TRAIN_SPLIT].std(axis=0)
dataset = (dataset-data_mean)/data_std
# + id="q8ei8d0G8lMQ" colab_type="code" colab={}
features_considered_labels = ['Wind Speed (m/s)', 'LV ActivePower (kW)', 'Wind Direction (°)']
features_mod = df2[features_considered_labels]
features_mod.index = df2['Date/Time']
features_mod.tail()
dataset_mod = features_mod.values
data_mean_mod = dataset_mod[:TRAIN_SPLIT].mean(axis=0)
data_std_mod = dataset_mod[:TRAIN_SPLIT].std(axis=0)
dataset_mod = (dataset_mod-data_mean_mod)/data_std_mod
# + id="2vBvWYHF8lfk" colab_type="code" colab={}
# For extracting one data point per hour(Taking steps of 6, as data recorded every 10 min)
def multivariate_data(dataset, target, start_index, end_index, history_size,
target_size, step):
data = []
labels = []
start_index = start_index + history_size
if end_index is None:
end_index = len(dataset) - target_size
for i in range(start_index, end_index):
indices = range(i-history_size, i, step)
data.append(dataset[indices])
labels.append(target[i:i+target_size])
return np.array(data), np.array(labels)
# + id="_gfX1xpP9LiY" colab_type="code" colab={}
past_history = 720
future_target = 432
STEP = 6
BUFFER_SIZE = 10000
BATCH_SIZE = 256
x_train_multi, y_train_multi = multivariate_data(dataset, dataset_mod[:, 1], 0,
TRAIN_SPLIT, past_history,
future_target, STEP)
x_val_multi, y_val_multi = multivariate_data(dataset, dataset_mod[:, 1],
TRAIN_SPLIT, None, past_history,
future_target, STEP)
# + id="8wg9zFXY9LgD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="29895b90-4658-4112-e7c7-a9ceb9755c57"
print ('Single window of past history : {}'.format(x_train_multi[0].shape))
print ('\n Target power to predict : {}'.format(y_train_multi[0].shape))
# + id="LtUTl2KM9LcC" colab_type="code" colab={}
# Conversion into tensors and splitting of data into mini-batches
train_data_multi = tf.data.Dataset.from_tensor_slices((x_train_multi, y_train_multi))
train_data_multi = train_data_multi.cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE).repeat()
val_data_multi = tf.data.Dataset.from_tensor_slices((x_val_multi, y_val_multi))
val_data_multi = val_data_multi.batch(BATCH_SIZE).repeat()
# + id="Gw7khgyP9LV3" colab_type="code" colab={}
def create_time_steps(length):
return list(range(-length, 0))
# + id="KvGTzSuV9LK2" colab_type="code" colab={}
def multi_step_plot(history, true_future, prediction):
plt.figure(figsize=(12, 6))
num_in = create_time_steps(len(history))
num_out = len(true_future)
plt.plot(num_in, np.array(history[:, 1]), label='History')
plt.plot(np.arange(num_out)/STEP, np.array(true_future), 'bo',
label='True Future')
if prediction.any():
plt.plot(np.arange(num_out)/STEP, np.array(prediction), 'ro',
label='Predicted Future')
plt.legend(loc='upper left')
plt.show()
# + id="61UE8s8Q9X9I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="6ee02f15-6c1e-4985-8c1b-3d639c9bd8a1"
for x, y in train_data_multi.take(1):
multi_step_plot(x[0], y[0], np.array([0]))
# + id="CYmJxE5B9X7e" colab_type="code" colab={}
# Defining the model
multi_step_model = tf.keras.models.Sequential()
multi_step_model.add(tf.keras.layers.LSTM(32,
return_sequences=True,
input_shape=x_train_multi.shape[-2:]))
multi_step_model.add(tf.keras.layers.LSTM(16, activation='relu'))
multi_step_model.add(tf.keras.layers.Dense(432))
multi_step_model.compile(optimizer='adam', loss='mae')
# + id="UbfQZfFNW-f4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="d416a8ee-8cf0-425d-b221-76c1bf9cc85b"
multi_step_model.summary()
# + id="nb9vf22n9X6H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="47494136-81a9-434d-e563-c4207c30a106"
for x, y in val_data_multi.take(1):
print (multi_step_model.predict(x).shape)
# + id="Cypaw22W9X3V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2dc62cc3-189e-4245-ce21-b08913bf5883"
EVALUATION_INTERVAL = 200
EPOCHS = 250
multi_step_history = multi_step_model.fit(train_data_multi, epochs=EPOCHS,
steps_per_epoch=EVALUATION_INTERVAL,
validation_data=val_data_multi,
validation_steps=50)
# + id="APwLFNCj9X0e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="a68d621e-8838-47f5-e87f-7f3f781a8dd6"
def plot_train_history(history, title):
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(loss))
plt.figure()
plt.plot(epochs, loss, 'b', label='Training loss')
plt.plot(epochs, val_loss, 'r', label='Validation loss')
plt.title(title)
plt.legend()
plt.show()
plot_train_history(multi_step_history, 'Multi-Step Training and validation loss')
# + id="O5IlQPGQ9XuC" colab_type="code" colab={}
def multivariate_data_predict(dataset, past_window):
data = []
end_index = len(dataset)
start_index = len(dataset) - past_window
for i in range(start_index, end_index):
data.append(dataset[i])
return np.array(data)
# + id="WuMfO0q18l0R" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 72} outputId="79b7e53a-9341-447a-ca9c-4f5da71909b6"
uploaded_predict = files.upload()
# + id="UWDSqHGI-T0w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="1b053c43-8937-46c9-a089-c0d2050073e3"
# Importing the previous 5 days weather data
predict_dataset = pd.read_csv("dataset_predict.csv", encoding= 'unicode_escape')
print(predict_dataset)
# + id="g7HBdV96-TGy" colab_type="code" colab={}
# Normalising the data w.r.t the training dataset
df4 = predict_dataset
predict_features_considered = ['Wind Speed (m/s)', 'Wind Direction (°)']
predict_features = df4[predict_features_considered]
predict_features.index = df4['Date/Time']
predict_features.tail()
predict_dataset = predict_features.values
predict_dataset_mod = (predict_dataset-data_mean)/data_std
# + id="42aFXDVH-TFA" colab_type="code" colab={}
BATCH_SIZE_predict = 120
past_history_predict = 120
x_predict_multi = multivariate_data_predict(predict_dataset_mod, past_history_predict)
# + id="zkvC691m-TDO" colab_type="code" colab={}
x_predict_multi = tf.data.Dataset.from_tensor_slices((x_predict_multi))
x_predict_multi = x_predict_multi.batch(BATCH_SIZE_predict).repeat()
x_predict_multi = x_predict_multi.batch(BATCH_SIZE_predict).repeat()
# + id="_Zzh06o4-TBN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ff55b355-2130-4d7a-d643-765159272852"
# Output predictions for the next 3 days(6 predictions per hour * 72 hours = 432 data points)
predictions_raw = multi_step_model.predict(x_predict_multi.take(1))
predictions_array = (predictions_raw*dataset_std[1]) + dataset_mean[1]
predictions = predictions_array[0]
for i in range(120):
if predictions[i] < 0:
predictions[i] = 0
print(predictions)
# + id="B99SDsLEwiKP" colab_type="code" colab={}
# Function for plotting the predicted values
def output_plot(output):
plt.plot(output)
plt.xlabel("Predictions made every 10 min for the next 72 hours")
plt.ylabel("Active Power Output (kW)")
plt.show()
# + id="F6X4TL6_yl2J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="89c187b9-998a-496f-e31a-f26a1f435b3b"
output_plot(predictions)
# + id="Z1JZaLzk-S6x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="dcba4d89-fb4d-4731-cc63-3bfec746f48a"
from google.colab import drive
drive.mount('/content/gdrive')
# + id="Rb6eH5cD-S25" colab_type="code" colab={}
#using model.save_weights() to save the weights of the model in HDF5 format
multi_step_model.save_weights("/content/gdrive/My Drive/weights_final.h5")
# + id="dod7GrdY-Ssg" colab_type="code" colab={}
multi_step_model.save("/content/gdrive/My Drive/model_final.h5")
|
wind_power_forecasting_final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3alti
# language: python
# name: machinelearning
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import tree
from sklearn.metrics import accuracy_score
import seaborn as sns
sns.set() # setting seaborn default for plots
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Dependicies added
# <NAME> 20 July 2019
# Read train and test data using panda
data = pd.read_csv("train.csv")
test = pd.read_csv('test.csv')
sns.countplot(x='Survived', data=data)
plt.show()
# -
# So far read data and shown how many died and head of data gives info.
#
# 891 data in Trainset / 12 columns . ID is useless, Survived is not feature rather output so 10 feature so far.
data.shape
# Now i have to create a Feature vector to use it later. Let's take look for each feature and comment on them.
#
#
#
# Data Dictionary
#
# Survived: 0 = No, 1 = Yes = No modification. ## OUTPUT - to train
#
# pclass: Ticket class 1 = 1st, 2 = 2nd, 3 = 3rd = No modification , number form & scale is good enough.
#
#
# sex = male female needs to be 0,1
#
# age = maybe use 0-1-2-4 = baby kid etc.. , age is also missing but not much. fill with means of each sex.
#
#
# sibsp: # of siblings / spouses aboard the Titanic
#
# parch: # of parents / children aboard the Titanic
#
# combine sibsp and parch to Family?
# fare = Needs to be adjusted / scaled.
# name = Mr,Mrs,Mister,Miss can be identified as family thingy
#
# ticket: Ticket number = Seems useless ?
#
# cabin: Cabin number = useless for now , missing lot.
#
# embarked: Port of Embarkation C = Cherbourg, Q = Queenstown, S = Southampton
data.isnull().sum()
def bar_chart(feature):
survived = data[data['Survived']==1][feature].value_counts()
dead = data[data['Survived']==0][feature].value_counts()
df = pd.DataFrame([survived,dead])
df.index = ['Survived','Dead']
df.plot(kind='bar',stacked=True, figsize=(10,5))
bar_chart("Sex")
# shows most man died while females survived mostly.
# +
# For Name let's make it useful. We will get titles of names with Regular expression strings ends with .
data['Name'].value_counts()
train_test_data = [data,test] # combining train and test dataset
for dataset in train_test_data:
dataset['Title'] = dataset['Name'].str.extract(' ([A-Za-z]+)\.', expand=False)
# -
# Mr , Miss, Mrs and Rest are Category.
title_mapping = {"Mr": 0, "Miss": 1, "Mrs": 2,
"Master": 3, "Dr": 3, "Rev": 3, "Col": 3, "Major": 3, "Mlle": 3,"Countess": 3,
"Ms": 3, "Lady": 3, "Jonkheer": 3, "Don": 3, "Dona" : 3, "Mme": 3,"Capt": 3,"Sir": 3 }
for dataset in train_test_data:
dataset['Title'] = dataset['Title'].map(title_mapping)
# +
# inplace means True = modify same object, false means returns object which modified
data.drop('Name', axis=1, inplace=True)
test.drop('Name', axis=1, inplace=True)
data.drop('Cabin', axis=1, inplace=True)
test.drop('Cabin', axis=1, inplace=True)
# -
title_sex_map = {"male" : 0 , "female" : 1}
for dataset in train_test_data:
dataset['Sex'] = dataset['Sex'].map(title_sex_map)
#train["Age"].fillna(
data["Age"].fillna(data.groupby("Title")["Age"].transform("median"), inplace=True)
test["Age"].fillna(test.groupby("Title")["Age"].transform("median"), inplace=True)
data.head(30)
# +
for dataset in train_test_data:
dataset.loc[ dataset['Age'] <= 17, 'Age'] = 0,
dataset.loc[(dataset['Age'] > 17) & (dataset['Age'] <= 26), 'Age'] = 1,
dataset.loc[(dataset['Age'] > 26) & (dataset['Age'] <= 36), 'Age'] = 2,
dataset.loc[(dataset['Age'] > 36) & (dataset['Age'] <= 62), 'Age'] = 3,
dataset.loc[ dataset['Age'] > 62, 'Age'] = 4
# age classing them into different scales which fits our data.
# -
emb_mapping = {"S": 0, "C": 1, "Q": 2}
for dataset in train_test_data:
dataset['Embarked'] = dataset['Embarked'].fillna('S')
for dataset in train_test_data:
dataset['Embarked'] = dataset['Embarked'].map(emb_mapping)
# fill missing Fare with according to Where they sitted ( P CLASS gives that)
data["Fare"].fillna(data.groupby("Pclass")["Fare"].transform("median"), inplace=True)
test["Fare"].fillna(test.groupby("Pclass")["Fare"].transform("median"), inplace=True)
# +
data["Fare"]
# need graphic to see which are limits for each category
facet = sns.FacetGrid(data, hue="Survived",aspect=4)
facet.map(sns.kdeplot,'Fare',shade= True)
facet.set(xlim=(0, data['Fare'].max()))
facet.add_legend()
plt.show()
# +
for dataset in train_test_data:
dataset.loc[ dataset['Fare'] <= 17, 'Fare'] = 0,
dataset.loc[(dataset['Fare'] > 17) & (dataset['Fare'] <= 30), 'Fare'] = 1,
dataset.loc[(dataset['Fare'] > 30) & (dataset['Fare'] <= 100), 'Fare'] = 2,
dataset.loc[ dataset['Fare'] > 100, 'Fare'] = 3
# +
# Family sizes without person itself.
data["FamilySize"] = data["SibSp"] + data["Parch"]
test["FamilySize"] = test["SibSp"] + test["Parch"]
data.drop('SibSp', axis=1, inplace=True)
test.drop('SibSp', axis=1, inplace=True)
data.drop('Ticket', axis=1, inplace=True)
test.drop('Ticket', axis=1, inplace=True)
data.drop('Parch', axis=1, inplace=True)
test.drop('Parch', axis=1, inplace=True)
# y VALUE
data.drop('PassengerId', axis=1, inplace=True)
train_data = data.drop('Survived', axis=1)
target = data['Survived']
# +
# Importing Classifier Modules hence this is Classifier model.
# My Cheat Sheat by Sk learn said Linear SVC would be best fit / model but lets see?
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
import numpy as np
# Cross Validation
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
k_fold = KFold(n_splits=10, shuffle=True, random_state=0)
pd.isnull(train_data).sum() > 0
#train_data.info()
#test.info()
#with pd.option_context('display.max_rows', None, 'display.max_columns', None): # more options can be specified also
# print(train_data)
# +
clf = KNeighborsClassifier(n_neighbors = 13)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# kNN Score
round(np.mean(score)*100, 2)
# -
clf = DecisionTreeClassifier()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# decision tree Score
round(np.mean(score)*100, 2)
# +
clf = RandomForestClassifier(n_estimators=13)
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# Random Forest Score
round(np.mean(score)*100, 2)
# +
clf = GaussianNB()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
# Naive Bayes Score
round(np.mean(score)*100, 2)
# +
clf = SVC()
scoring = 'accuracy'
score = cross_val_score(clf, train_data, target, cv=k_fold, n_jobs=1, scoring=scoring)
print(score)
round(np.mean(score)*100,2)
# +
clf = SVC()
clf.fit(train_data, target)
test_data = test.drop("PassengerId", axis=1).copy()
prediction = clf.predict(test_data)
submission = pd.DataFrame({
"PassengerId": test["PassengerId"],
"Survived": prediction
})
submission.to_csv('submission.csv', index=False)
submission = pd.read_csv('submission.csv')
submission.head()
# -
|
kaggleTitanic/titanicproblem.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # COSMOS master catalogue
# ## Preparation of Hyper Suprime-Cam Subaru Strategic Program Catalogues (HSC-SSP) data
#
# This catalogue comes from `dmu0_HSC`.
#
# In the catalogue, we keep:
#
# - The `object_id` as unique object identifier;
# - The position;
# - The g, r, i, z, y (no N921) aperture magnitude in 2” that we aperture correct;
# - The g, r, i, z, y (no N921) kron fluxes and magnitudes.
# - The extended flag that we convert to a stellariy.
#
# **Note**: On ELAIS-N1 the HSC-SSP catalogue does not contain any N816 magnitudes.
#
# We use 2016 as the epoch.
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
import datetime
print("This notebook was executed on: \n{}".format(datetime.datetime.now()))
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
plt.style.use('ggplot')
from collections import OrderedDict
import os
from astropy import units as u
from astropy import visualization as vis
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table
import numpy as np
from herschelhelp_internal.flagging import gaia_flag_column
from herschelhelp_internal.masterlist import nb_astcor_diag_plot, nb_plot_mag_ap_evol, \
nb_plot_mag_vs_apcor, remove_duplicates
from herschelhelp_internal.utils import astrometric_correction, mag_to_flux, aperture_correction
# +
OUT_DIR = os.environ.get('TMP_DIR', "./data_tmp")
try:
os.makedirs(OUT_DIR)
except FileExistsError:
pass
RA_COL = "hsc-udeep_ra"
DEC_COL = "hsc-udeep_dec"
# -
# Pristine HSC catalogue
orig_hsc = Table.read("../../dmu0/dmu0_HSC/data/HSC-PDR1_uDeep_COSMOS.fits")
# ## I - Aperture correction
#
# To compute aperture correction we need to dertermine two parametres: the target aperture and the range of magnitudes for the stars that will be used to compute the correction.
#
# **Target aperture**: To determine the target aperture, we simulate a curve of growth using the provided apertures and draw two figures:
#
# - The evolution of the magnitudes of the objects by plotting on the same plot aperture number vs the mean magnitude.
# - The mean gain (loss when negative) of magnitude is each aperture compared to the previous (except for the first of course).
#
# As target aperture, we should use the smallest (i.e. less noisy) aperture for which most of the flux is captures.
#
# **Magnitude range**: To know what limits in aperture to use when doing the aperture correction, we plot for each magnitude bin the correction that is computed and its RMS. We should then use the wide limits (to use more stars) where the correction is stable and with few dispersion.
# +
bands = ["g", "r", "i", "z", "y", "n921"]
apertures = ["10", "15", "20", "30", "40", "57", "84", "118", "168", "235"]
magnitudes = {}
magnitude_errors ={}
stellarities = {}
for band in bands:
magnitudes[band] = np.array(
[orig_hsc["{}mag_aperture{}".format(band, aperture)] for aperture in apertures]
)
# Some sources have an infinite magnitude
mask = np.isinf(magnitudes[band])
magnitudes[band][mask] = np.nan
try:
magnitude_errors[band] = np.array(
[orig_hsc["{}mag_aperture{}_err".format(band, aperture)] for aperture in apertures]
)
magnitude_errors[band][mask] = np.nan
except KeyError:
print("No error column for a " + band + " band aperture magnitude.")
stellarities[band] = 1 - np.array(orig_hsc["{}classification_extendedness".format(band)])
mag_corr = {}
# -
# ### I.a - g band
nb_plot_mag_ap_evol(magnitudes['g'], stellarities['g'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['gmag_aperture20'], orig_hsc['gmag_aperture40'], stellarities['g'])
# We will use magnitudes between 18.5 and 20.8
# Aperture correction
mag_corr['g'], num, std = aperture_correction(
orig_hsc['gmag_aperture20'], orig_hsc['gmag_aperture40'],
stellarities['g'],
mag_min=18.5, mag_max=20.8)
print("Aperture correction for g band:")
print("Correction: {}".format(mag_corr['g']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ### I.b - r band
nb_plot_mag_ap_evol(magnitudes['r'], stellarities['r'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['rmag_aperture20'], orig_hsc['rmag_aperture40'], stellarities['r'])
# We use magnitudes between 17.6 and 19.7.
# Aperture correction
mag_corr['r'], num, std = aperture_correction(
orig_hsc['rmag_aperture20'], orig_hsc['rmag_aperture40'],
stellarities['r'],
mag_min=17.6, mag_max=19.7)
print("Aperture correction for r band:")
print("Correction: {}".format(mag_corr['r']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ### I.c - i band
nb_plot_mag_ap_evol(magnitudes['i'], stellarities['i'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['imag_aperture20'], orig_hsc['imag_aperture40'], stellarities['i'])
# We use magnitudes between 18.5 and 19.8.
# Aperture correction
mag_corr['i'], num, std = aperture_correction(
orig_hsc['imag_aperture20'], orig_hsc['imag_aperture40'],
stellarities['i'],
mag_min=18.5, mag_max=19.8)
print("Aperture correction for i band:")
print("Correction: {}".format(mag_corr['i']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ### I.d - z band
nb_plot_mag_ap_evol(magnitudes['z'], stellarities['z'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['zmag_aperture20'], orig_hsc['zmag_aperture40'], stellarities['z'])
# We use magnitudes between 17.5 and 19.8.
# Aperture correction
mag_corr['z'], num, std = aperture_correction(
orig_hsc['zmag_aperture20'], orig_hsc['zmag_aperture40'],
stellarities['z'],
mag_min=17.5, mag_max=19.8)
print("Aperture correction for z band:")
print("Correction: {}".format(mag_corr['z']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ### I.e - y band
nb_plot_mag_ap_evol(magnitudes['y'], stellarities['y'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['ymag_aperture20'], orig_hsc['ymag_aperture40'], stellarities['y'])
# We use magnitudes between 17 and 18.7.
# Aperture correction
mag_corr['y'], num, std = aperture_correction(
orig_hsc['ymag_aperture20'], orig_hsc['ymag_aperture40'],
stellarities['y'],
mag_min=17, mag_max=18.7)
print("Aperture correction for y band:")
print("Correction: {}".format(mag_corr['y']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ### I.f - n921 band
nb_plot_mag_ap_evol(magnitudes['n921'], stellarities['n921'], labels=apertures)
# We will use aperture 40 as target.
nb_plot_mag_vs_apcor(orig_hsc['n921mag_aperture20'], orig_hsc['n921mag_aperture40'], stellarities['n921'])
# We use magnitudes between 17 and 18.7.
# Aperture correction
mag_corr['n921'], num, std = aperture_correction(
orig_hsc['n921mag_aperture20'], orig_hsc['n921mag_aperture40'],
stellarities['n921'],
mag_min=17, mag_max=18.7)
print("Aperture correction for n921 band:")
print("Correction: {}".format(mag_corr['n921']))
print("Number of source used: {}".format(num))
print("RMS: {}".format(std))
# ## II - Stellarity
#
# HSC does not provide a 0 to 1 stellarity value but a 0/1 extended flag in each band. We are using the same method as UKIDSS ([cf this page](http://wsa.roe.ac.uk/www/gloss_p.html#dxssource_pstar)) to compute a stellarity based on the class in each band:
#
# \begin{equation*}
# P(star) = \frac{ \prod_{i} P(star)_i }{ \prod_{i} P(star)_i + \prod_{i} P(galaxy)_i }
# \end{equation*}
#
# where $i$ is the band, and with using the same probabilities as UKDISS:
#
# | HSC flag | UKIDSS flag | Meaning | P(star) | P(galaxy) | P(noise) | P(saturated) |
# |:--------:|:-----------:|:----------------|--------:|----------:|---------:|-------------:|
# | | -9 | Saturated | 0.0 | 0.0 | 5.0 | 95.0 |
# | | -3 | Probable galaxy | 25.0 | 70.0 | 5.0 | 0.0 |
# | | -2 | Probable star | 70.0 | 25.0 | 5.0 | 0.0 |
# | 0 | -1 | Star | 90.0 | 5.0 | 5.0 | 0.0 |
# | | 0 | Noise | 5.0 | 5.0 | 90.0 | 0.0 |
# | 1 | +1 | Galaxy | 5.0 | 90.0 | 5.0 | 0.0 |
# We are creating an array containing the extended flag in all band.
# Some sources have no flag in some band, there will be NaN in the array.
hsc_ext_flag = np.array([
orig_hsc[colname] for colname in
['gclassification_extendedness',
'rclassification_extendedness',
'iclassification_extendedness',
'zclassification_extendedness',
'yclassification_extendedness',
'n921classification_extendedness']
])
# +
hsc_pstar = 0.9 * (hsc_ext_flag == 0) + 0.05 * (hsc_ext_flag == 1)
hsc_pgal = 0.05 * (hsc_ext_flag == 0) + 0.9 * (hsc_ext_flag == 1)
# We put back the NaN values
hsc_pstar[np.isnan(hsc_ext_flag)] = np.nan
hsc_pgal[np.isnan(hsc_ext_flag)] = np.nan
# +
stellarity = np.nanprod(hsc_pstar, axis=0) / np.nansum(
[np.nanprod(hsc_pgal, axis=0), np.nanprod(hsc_pstar, axis=0)], axis=0)
stellarity = np.round(stellarity, 3)
# -
vis.hist(stellarity, bins='scott');
orig_hsc.add_column(Column(data=stellarity, name="stellarity"))
# ## II - Column selection
# +
imported_columns = OrderedDict({
"object_id": "hsc-udeep_id",
"ra": "hsc-udeep_ra",
"dec": "hsc-udeep_dec",
"gmag_aperture20": "m_ap_hsc-udeep_g",
"gmag_aperture20_err": "merr_ap_hsc-udeep_g",
"gmag_kron": "m_hsc-udeep_g",
"gmag_kron_err": "merr_hsc-udeep_g",
"rmag_aperture20": "m_ap_hsc-udeep_r",
"rmag_aperture20_err": "merr_ap_hsc-udeep_r",
"rmag_kron": "m_hsc-udeep_r",
"rmag_kron_err": "merr_hsc-udeep_r",
"imag_aperture20": "m_ap_hsc-udeep_i",
"imag_aperture20_err": "merr_ap_hsc-udeep_i",
"imag_kron": "m_hsc-udeep_i",
"imag_kron_err": "merr_hsc-udeep_i",
"zmag_aperture20": "m_ap_hsc-udeep_z",
"zmag_aperture20_err": "merr_ap_hsc-udeep_z",
"zmag_kron": "m_hsc-udeep_z",
"zmag_kron_err": "merr_hsc-udeep_z",
"ymag_aperture20": "m_ap_hsc-udeep_y",
"ymag_aperture20_err": "merr_ap_hsc-udeep_y",
"ymag_kron": "m_hsc-udeep_y",
"ymag_kron_err": "merr_hsc-udeep_y",
"n921mag_aperture20": "m_ap_hsc-udeep_n921",
"n921mag_aperture20_err": "merr_ap_hsc-udeep_n921",
"n921mag_kron": "m_hsc-udeep_n921",
"n921mag_kron_err": "merr_hsc-udeep_n921",
"stellarity": "hsc-udeep_stellarity"
})
catalogue = orig_hsc[list(imported_columns)]
for column in imported_columns:
catalogue[column].name = imported_columns[column]
epoch = 2017
# Clean table metadata
catalogue.meta = None
# -
# Aperture correction
for band in bands:
catalogue["m_ap_hsc-udeep_{}".format(band)] += mag_corr[band]
# +
# Adding flux and band-flag columns
for col in catalogue.colnames:
if col.startswith('m_'):
errcol = "merr{}".format(col[1:])
#Some inf and 99. mags
mask = (catalogue[col] > 90.) | (catalogue[errcol] > 90.)
catalogue[col][mask] = np.nan
catalogue[errcol][mask] = np.nan
flux, error = mag_to_flux(np.array(catalogue[col]), np.array(catalogue[errcol]))
# Fluxes are added in µJy
catalogue.add_column(Column(flux * 1.e6, name="f{}".format(col[1:])))
catalogue.add_column(Column(error * 1.e6, name="f{}".format(errcol[1:])))
# Band-flag column
if 'ap' not in col:
catalogue.add_column(Column(np.zeros(len(catalogue), dtype=bool), name="flag{}".format(col[1:])))
# TODO: Set to True the flag columns for fluxes that should not be used for SED fitting.
# -
catalogue[:10].show_in_notebook()
# ## III - Removal of duplicated sources
# We remove duplicated objects from the input catalogues.
# +
SORT_COLS = [
'merr_ap_hsc-udeep_i', 'merr_ap_hsc-udeep_r', 'merr_ap_hsc-udeep_z',
'merr_ap_hsc-udeep_y', 'merr_ap_hsc-udeep_g','merr_ap_hsc-udeep_n921']
FLAG_NAME = 'hsc-udeep_flag_cleaned'
nb_orig_sources = len(catalogue)
catalogue = remove_duplicates(
catalogue, RA_COL, DEC_COL,
sort_col= SORT_COLS,
flag_name=FLAG_NAME)
nb_sources = len(catalogue)
print("The initial catalogue had {} sources.".format(nb_orig_sources))
print("The cleaned catalogue has {} sources ({} removed).".format(nb_sources, nb_orig_sources - nb_sources))
print("The cleaned catalogue has {} sources flagged as having been cleaned".format(np.sum(catalogue[FLAG_NAME])))
# -
# ## III - Astrometry correction
#
# We match the astrometry to the Gaia one. We limit the Gaia catalogue to sources with a g band flux between the 30th and the 70th percentile. Some quick tests show that this give the lower dispersion in the results.
gaia = Table.read("../../dmu0/dmu0_GAIA/data/GAIA_COSMOS.fits")
gaia_coords = SkyCoord(gaia['ra'], gaia['dec'])
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# +
delta_ra, delta_dec = astrometric_correction(
SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]),
gaia_coords
)
print("RA correction: {}".format(delta_ra))
print("Dec correction: {}".format(delta_dec))
# -
catalogue[RA_COL] += delta_ra.to(u.deg)
catalogue[DEC_COL] += delta_dec.to(u.deg)
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# ## IV - Flagging Gaia objects
catalogue.add_column(
gaia_flag_column(SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]), epoch, gaia)
)
# +
GAIA_FLAG_NAME = "hsc-udeep_flag_gaia"
catalogue['flag_gaia'].name = GAIA_FLAG_NAME
print("{} sources flagged.".format(np.sum(catalogue[GAIA_FLAG_NAME] > 0)))
# -
# ## V - Flagging objects near bright stars
# # VI - Saving to disk
catalogue.write("{}/HSC-UDEEP.fits".format(OUT_DIR), overwrite=True)
|
dmu1/dmu1_ml_COSMOS/1.4.2_HSC-UDEEP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # Python Crash Course
#
# Please note, this is not meant to be a comprehensive overview of Python or programming in general, if you have no programming experience, you should probably take my other course: [Complete Python Bootcamp](https://www.udemy.com/complete-python-bootcamp/?couponCode=PY20) instead.
#
# **This notebook is just a code reference for the videos, no written explanations here**
#
# This notebook will just go through the basic topics in order:
#
# * Data types
# * Numbers
# * Strings
# * Printing
# * Lists
# * Dictionaries
# * Booleans
# * Tuples
# * Sets
# * Comparison Operators
# * if, elif, else Statements
# * for Loops
# * while Loops
# * range()
# * list comprehension
# * functions
# * lambda expressions
# * map and filter
# * methods
# ____
# ## Data types
#
# ### Numbers
1 + 1
1 * 3
1 / 2
2 ** 4
4 % 2
5 % 2
(2 + 3) * (5 + 5)
# ### Variable Assignment
# Can not start with number or special characters
name_of_var = 2
x = 2
y = 3
z = x + y
z
# ### Strings
'single quotes'
"double quotes"
" wrap lot's of other quotes"
# ### Printing
x = 'hello'
x
print(x)
num = 12
name = 'Sam'
print('My number is: {one},x = 'hello' and my name is: {two}'.format(one=num,two=name))
print('My number is: {}, and my name is: {}'.format(num,name))
# ### Lists
[1,2,3]
['hi',1,[1,2]]
my_list = ['a','b','c']
my_list.append('d')
my_list
my_list[0]
my_list[1]
my_list[1:]
my_list[:1]
my_list[0] = 'NEW'
my_list
nest = [1,2,3,[4,5,['target']]]
nest[3]
nest[3][2]
nest[3][2][0]
# ### Dictionaries
d = {'key1':'item1','key2':'item2'}
d
d['key1']
# ### Booleans
True
False
# ### Tuples
t = (1,2,3)
t[0]
t[0] = 'NEW'
# ### Sets
{1,2,3}
{1,2,3,1,2,1,2,3,3,3,3,2,2,2,1,1,2}
# ## Comparison Operators
1 > 2
1 < 2
1 >= 1
1 <= 4
1 == 1
'hi' == 'bye'
# ## Logic Operators
(1 > 2) and (2 < 3)
(1 > 2) or (2 < 3)
(1 == 2) or (2 == 3) or (4 == 4)
# ## if,elif, else Statements
if 1 < 2:
print('Yep!')
if 1 < 2:
print('yep!')
if 1 < 2:
print('first')
else:
print('last')
if 1 > 2:
print('first')
else:
print('last')
if 1 == 2:
print('first')
elif 3 == 3:
print('middle')
else:
print('Last')
# ## for Loops
seq = [1,2,3,4,5]
for item in seq:
print(item)
for item in seq:
print('Yep')
for jelly in seq:
print(jelly+jelly)
# ## while Loops
i = 1
while i < 5:
print('i is: {}'.format(i))
i = i+1
# ## range()
range(5)
for i in range(5):
print(i)
list(range(5))
# ## list comprehension
x = [1,2,3,4]
out = []
for item in x:
out.append(item**2)
print(out)
[item**2 for item in x]
# ## functions
def my_func(param1='default'):
"""
Docstring goes here.
"""
print(param1)
my_func
my_func()
my_func('new param')
my_func(param1='new param')
def square(x):
return x**2
out = square(2)
print(out)
# ## lambda expressions
def times2(var):
return var*2
times2(2)
lambda var: var*2
# ## map and filter
seq = [1,2,3,4,5]
map(times2,seq)
list(map(times2,seq))
list(map(lambda var: var*2,seq))
filter(lambda item: item%2 == 0,seq)
list(filter(lambda item: item%2 == 0,seq))
# ## methods
st = 'hello my name is Sam'
st.lower()
st.upper()
st.split()
tweet = 'Go Sports! #Sports'
tweet.split('#')
tweet.split('#')[1]
d
d.keys()
d.items()
lst = [1,2,3]
lst.pop()
lst
'x' in [1,2,3]
'x' in ['x','y','z']
# # Great Job!
|
01-Introduction to Python-Basics/01-Python Crash Course.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Big Dam Problem: Understanding the Potential Impacts of Future Dam Development
# Contributors: <NAME> & <NAME>
# ## We should give a dam
#
# Dam construction over the past century has dramatically altered our landscapes, threatening ecosystems and endangered species around the world. Thousands of new dams are proposed globally that would affect protected areas such as National Parks, UNESCO World Heritage Sites, and Ramsar Sites. While data is publicly available, there is scant understanding of how these proposed dams will affect these critical protected areas.
#
# Many scientists believe we may be on the cusp of the sixth Mass Extinction in Earth’s history, due to human impact on the environment (1). Since the early 20th Century the majority of major rivers around the globe have been impounded or diverted to harness the power of water for energy production, transportation, flood control, water storage, and even recreation (2, 3). Although we as a society benefit greatly from the construction of these engineering marvels, dams have a dramatic impact on river ecosystems. A dam drastically slows water flow, trapping sediment, fish, and water upstream as the reservoir fills flooding the river valley. As a result both the river habitats and the riparian habitats are lost. Downstream of the dam water flow is strictly controlled, often eliminating the natural annual cycles of flooding that supplied sediment and nutrients to the floodplain. When water flow exceeds the storage capacity of the dam, the water released often causes catastrophic flooding downstream. Even as the water rushes out the necessary sediment, nutrients, and organisms are for the most part still trapped behind the dam. Finally dam construction alters the aesthetics of beautiful landscapes around the world and threatening freshwater and terrestrial species (4).
#
# This project is being undertaken in collaboration with the World Wildlife Foundation for the purpose of identifying critical habitats that will potentially be impacted by future dam construction.
#
# ## What do we already know
# Global Dam Watch (http://globaldamwatch.org/) is a collaboration between academic institutions and non-governmental organizations to collect, analyze, and curate data about current and future dams around the globe. The collaboration maintains three dam databases:
# * GlObal geOreferenced Database of Dams (GOODD) - http://globaldamwatch.org/goodd/
# * Global Reservoir and Dam Database (GRand) - http://globaldamwatch.org/grand/
# * Future Hydropower Reservoirs and Dams (FHReD) - http://globaldamwatch.org/fhred/
#
# These datasets have already contributed to our understanding of how current dam infrastructure effects the movement and storage of water and sediment(5, 6), the impacts on riparian wetlands(7), as well as the preservation and restoration of free-flowing rivers (8).
# ## Exploring the data
#
# These databases paint a clear picture of the distribution of dams globally which allows us now to assess their impacts in different ways. For the larger project we will identify protected areas that will potentially be impacted from the development of new dams around the globe. For this preliminary work we are using the FHReD datase, which contains geographic information about dams such as main river dammed and country as well as dam capacity, project name, and development stage, and the Ramsar Site database (https://rsis.ramsar.org/), which list wetland areas of critical importance as identified under the Convention on Wetlands (1971).
# ## Import packages, define functions, and acquire data
# +
# Imports
import os
import sys
import numpy as np
import numpy.ma as ma
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
import geopandas as gpd
from geopandas import GeoDataFrame as gdf
from geopandas import GeoSeries as gs
from shapely.geometry import Point, Polygon
import contextily as ctx
import earthpy as et
import earthpy.plot as ep
# -
# Check path and set working directory.
wd_path = os.path.join(et.io.HOME, 'earth-analytics', 'data')
if os.path.exists(wd_path):
os.chdir(wd_path)
else:
print("Path does not exist")
# Custon Function 1
def dam_impact(buff_km, country):
""" This function takes a country name, pulls the proposed
dams & ramsar areas for the country, creates a 5km & 10km
buffer around the dams, and returns the total ramsar area covered
by each buffer.
Parameters
----------
buff_km: int or float
The number of kilometers buffer you want around the dam.
country: str
Full name of the country to be analyzed in quotes.
Returns
-------
ramsar_affected: float
Total ramsar area affected in the country by proposed dams with buffer.
"""
# Pull only the data for the country
proposed_dams_cntry = proposed_dams[proposed_dams['Country'] == country]
ramsar_areas_cntry = ramsar_areas[ramsar_areas['country_en'] == country]
# Buffer the dams to 5km & 10km to create gdfs of only the intersections
proposed_dams_cntry['geometry'] = proposed_dams_cntry.buffer(
buff_km * 1000)
try:
data_intersect_cntry = gpd.overlay(
ramsar_areas_cntry, proposed_dams_cntry, how='intersection')
# divide by 1 million because it sq km
ramsar_affected = ((data_intersect_cntry.area.sum())/1000000).round(1)
return ramsar_affected
except IndexError as err:
Dams = None
# Custom Function 2
def autolabel(rects): # width, dataframe, width, label are other
"""Attach a text label above each bar in *rects*, displaying its value.
Parameters
----------
rect: list
The list of values or count of items for the category.
Returns
-------
label: int
The value or count of items in a category.
"""
for rect in rects:
height = rect.get_height()
ax.annotate('{}'.format(height),
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
# +
# Download Data stored on figshare
# Ramsar Sites
et.data.get_data(url="https://ndownloader.figshare.com/files/22507082")
# Future dams
et.data.get_data(url="https://ndownloader.figshare.com/files/22486157")
# Country boundaries
et.data.get_data(url="https://ndownloader.figshare.com/files/22507058")
# +
# Open the ramsar shapefile with geopandas
ramsar_all = gpd.read_file(os.path.join(
"earthpy-downloads", "ramsar-site-data", "ramsar-boundaries",
"features_publishedPolygon.shp"))
# Check the crs of the ramsar sites
print(ramsar_all.crs)
# Open the dams csv files with pandas
fname = os.path.join("earthpy-downloads", "future_dams_2015.csv")
df = pd.read_csv(fname)
# Covert the pandas dataframe to a shapefile for plotting
# Set output path for shp
dams_path = os.path.join('final-project-data', 'fhred-proposed-dams')
if not os.path.exists(dams_path):
os.mkdir(dams_path)
# Define the geometry for the points
geometry = [Point(xy) for xy in zip(df.Lon_Cleaned, df.LAT_cleaned)]
crs = {'init': 'epsg:4326'}
geo_df = gdf(df, crs=crs, geometry=geometry)
geo_df.to_file(driver='ESRI Shapefile', filename=os.path.join(
dams_path, 'proposed_dams.shp'))
# Open the proposed dams shapefile with geopandas
dams_all = gpd.read_file(os.path.join(dams_path, "proposed_dams.shp"))
# Pull only the columns that we need from each gdf to save processing time
proposed_dams = dams_all[['Country',
'Continent', 'Major Basi', 'Stage', 'geometry']]
ramsar_areas = ramsar_all[['country_en', 'geometry']]
# Open country borders shapefile for adding boundary of study area
country_borders_path = os.path.join("earthpy-downloads", "country-borders",
"99bfd9e7-bb42-4728-87b5-07f8c8ac631c2020328-1-1vef4ev.lu5nk.shp")
country_borders = gpd.read_file(country_borders_path)
# -
# # Global Map of Future Dams and Ramsar Sites
#
# In order to visual the scope of the two databases selected for this project, we initially plotted the entire FHReD database with the Ramsar site polygons. At a global scale we can see the proposed dams are concentrated in Africa, Asia, and South America, but the Ramsar sites just fade into the background.
# +
# Create a Global Map of Dams and Ramsar Sites
red_patch = mpatches.Patch(color='red', label='Ramsar Area')
green_dot = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='lightgreen', label='Proposed Dam Site')
black_line = mlines.Line2D([], [], color='black', label='Country Border')
fig, ax = plt.subplots(figsize=(15, 25))
country_borders.plot(ax=ax, color="lightyellow",
edgecolor="black", linewidth=2)
proposed_dams.plot(ax=ax,
markersize=15,
color='lightgreen', legend=True)
ramsar_areas.plot(ax=ax, facecolor='red', legend=True)
ax.set_title(
'Figure 1: Global Proposed Dams & Ramsar Areas', size=20)
ax.set_axis_off()
ax.text(0.5, -0.2, "Data Sources: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/),\nRamsar Sites "
"Information Service (https://rsis.ramsar.org/)",
size=12, ha="center", transform=ax.transAxes)
ax.legend(handles=[red_patch, green_dot, black_line],
fontsize=15,
frameon=True,
loc=('lower right'),
title="LEGEND")
# -
# # Future Dam Construction by Continent and Selected Countries
#
#
#
# ## Visualizing the data at the continent scale
# Stepping down to the continent scale, we compared the number of dams underconstruction (U) and planned (P).
# +
# Getting Number of Dams by Continent and Development State
# Extact the columns needed for analysis
dams_continent = dams_all[['Continent', 'Country', 'Stage']]
# Group and count data by stage.
dams_stage = dams_continent.groupby(['Continent', 'Country', 'Stage'])[
['Stage']].count()
dams_stage
# +
# PLOT - NUMBER OF DAMS PROPOSED VS UNDER CONSTRUCTION BY CONTINENT
# Create a bar plot of the dams by continent and stage of process.
labels = ['Africa', 'Asia', 'Europe', 'N. America', 'Oceania', 'S. America']
proposed = [179, 937, 611, 143, 7, 1188]
under_const = [21, 424, 41, 34, 1, 114]
x = np.arange(len(labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10, 10))
rects1 = ax.bar(x - width/2, proposed, width, label='Proposed')
rects2 = ax.bar(x + width/2, under_const, width, label='Under Construction')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Count', size=15)
ax.set_title('Figure 2: Future Dams by Continent, 2015', size=20)
ax.set_xticks(x)
ax.set_xticklabels(labels, size=15, rotation=45)
ax.legend()
autolabel(rects1)
autolabel(rects2)
ax.text(0.5, -0.2, "Data Source: Global Dam Watch Future Hydropower "
"Reservoirs and Dams Database (http://globaldamwatch.org/fhred/)",
size=12, ha="center", transform=ax.transAxes)
fig.tight_layout()
plt.show()
# -
# ## Visualizing the data at the country scale
# Again, stepping down to the country scale, we compared future dam construction by stage. For this projet we are focusing on Africa as a test case for future analysis with other protected area datasets.
# +
# Extract data by continent.
africa = dams_continent[dams_continent["Continent"] == "Africa"]
# Group and count country data by stage.
africa_stage = africa.groupby(['Country', 'Stage'])[['Stage']].count()
africa_stage
# +
# Create a bar plot of the dams by countries in Africa comparing stage of process.
africa_labels = ['Benin', 'Burkina Faso', 'Gabon', 'Guinea', 'Malawi', 'Mali', 'Morocco',
'Mozambique', 'Namibia', 'Niger', 'Nigeria', 'Rwanda', 'South Africa', 'Zimbabwe']
africa_proposed = [6, 2, 1, 23, 2, 12, 0, 3, 3, 1, 1, 0, 3, 2]
africa_under_const = [0, 0, 1, 0, 0, 1, 2, 1, 0, 1, 2, 1, 0, 1]
x = np.arange(len(africa_labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10, 10))
rects1 = ax.bar(x - width/2, africa_proposed, width, label='Proposed')
rects2 = ax.bar(x + width/2, africa_under_const,
width, label='Under Construction')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Count', size=15)
ax.set_title(
'Figure 3: Future Dam Construction by Selected Countries in Africa', size=20)
ax.set_xticks(x)
ax.set_xticklabels(africa_labels, size=15, rotation=45)
ax.legend()
def autolabel(rects):
"""Attach a text label above each bar in *rects*, displaying its value."""
for rect in rects:
height = rect.get_height()
ax.annotate('{}'.format(height),
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
autolabel(rects1)
autolabel(rects2)
ax.text(0.5, -0.2, "Data Source: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/)",
size=12, ha="center", transform=ax.transAxes)
fig.tight_layout()
plt.show()
# -
# # Overlaying Future Dams and Ramsar Sites in Africa.
# In order to analyze these datasets for one continent we had to choose a coordinate reference system that would minimize spatial distortion in the dataset. Also, we had to check that the country names matched for both datasets (they did not). Once those issues were addressed, the buffer function was run and the results were saved to Pandas dataframe for plotting. An important caveat to the results generated from these data is we have not accounted for the flow direction of the river. We will incorporate this information in the future to further refine our buffer function.
# Change the datas CRS to projected for Africa (WGS 84 World Mercator)
# To make this data more accurate, for the next course we can create
# list of EPSG for each country in Africa to include in function
proposed_dams = proposed_dams.to_crs('epsg:3765')
ramsar_areas = ramsar_areas.to_crs('epsg:3765')
# +
# Get dam impact by African country
# List of African country names
# Data cleaning issues: Removed 'Côte d'Ivoire' bc in Ramsar dataset it's called Cite D'ivore and don't know how to deal with additional ' in a string; also removed Congo bc in Ramsar it's called Congo & Democratic Republic of Congo and in FhRED it's called Congo, Rep.
africa_cntry = ['Algeria', 'Angola', 'Benin', 'Botswana', 'Burkina Faso',
'Burundi', 'Cabo Verde', 'Cameroon', 'Central African Republic',
'Chad', 'Comoros', 'Djibouti', 'Equatorial Guinea', 'Eritrea',
'Ethiopia', 'Gabon', 'Gambia', 'Ghana', 'Guinea', 'Guinea-Bissau',
'Kenya', 'Lesotho', 'Liberia', 'Libya', 'Madagascar', 'Malawi',
'Mali', 'Mauritania', 'Mauritius', 'Morocco', 'Mozambique', 'Namibia',
'Niger', 'Nigeria', 'Rwanda', 'Sao Tome and Principe', 'Senegal',
'Seychelles', 'Sierra Leone', 'Somalia', 'South Africa', 'South Sudan',
'Sudan', 'Tanzania', 'Togo', 'Tunisia', 'Uganda', 'Zambia', 'Zimbabwe']
# Empty Africa dict
africa_dams = {}
# Append dam_impact function data to africa_dams
for i in africa_cntry:
try:
africa_dams[i] = {"5km Buffer Area": dam_impact(
5, i), "10km Buffer Area": dam_impact(10, i)}
except:
dams = None
# +
# Turn it into a pandas dataframe for plotting
africa_df = pd.DataFrame.from_dict(africa_dams)
# Some values in the dataframe are zero and some are NaN, make it the same
africa_df = africa_df.fillna('None')
africa_df.replace({0: 'None'})
# +
# Plot data to illustrate which countires have potential imapcts from proposed dams.
# Create legend so only countries with potential impacts are listed.
blue_patch = mpatches.Patch(color='dodgerblue', label='Beinin')
green_patch = mpatches.Patch(color='green', label='Gabon')
red_patch = mpatches.Patch(color='red', label='Guinea')
teal_patch = mpatches.Patch(color='c', label='Niger')
# Create the figure
fig, ax = plt.subplots(figsize=(8, 8))
africa_df.plot(ax=ax, kind='barh', stacked=True, legend=True)
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_xlabel('Affected Area (km)', size=15)
ax.set_title(
'Figure 4: Total Area of Ramsar Sites in Africa\nPotentially Impacted by Dam Construction', size=20)
ax.text(0.5, -0.2, "Data Sources: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/),\nRamsar Sites "
"Information Service (https://rsis.ramsar.org/)",
size=12, ha="center", transform=ax.transAxes)
ax.legend(handles=[blue_patch, green_patch, red_patch, teal_patch],
fontsize=15,
frameon=True,
loc=('lower right'),
title="Country")
# -
# # Mapped Buffer Results for Guinea
# +
# Analyze Guinea
# Pull only the data for Guinea
proposed_dams_guin = proposed_dams[proposed_dams['Country'] == "Guinea"]
ramsar_areas_guin = ramsar_areas[ramsar_areas['country_en'] == "Guinea"]
guinea_border = country_borders[country_borders['CNTRY_NAME'] == "Guinea"]
# Get the CRS right for plotting
proposed_dams_guin = proposed_dams_guin.to_crs('epsg:3462')
ramsar_areas_guin = ramsar_areas_guin.to_crs('epsg:3462')
guinea_border = guinea_border.to_crs('epsg:3462')
# Buffer the dams to 5km & 10km for plotting
proposed_dams_guin_5k_buff = proposed_dams_guin.buffer(5000)
proposed_dams_guin_10k_buff = proposed_dams_guin.buffer(10000)
# +
# Create a map of the dams and the ramsar sites for Guinea
black_line = mlines.Line2D([], [], color='black', label='Country Border')
yellow_patch = mpatches.Patch(color='yellow', label='Ramsar Area')
green_circle = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='forestgreen', markersize=18,
label='10km Buffer')
lime_circle = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='lime', markersize=12,
label='5km Buffer')
red_dot = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='red', label='Proposed Dam Site')
fig, ax = plt.subplots(figsize=(15, 15))
ramsar_areas_guin.plot(ax=ax, facecolor='yellow')
proposed_dams_guin_10k_buff.plot(facecolor='forestgreen',
ax=ax)
proposed_dams_guin_5k_buff.plot(facecolor='lime',
ax=ax)
proposed_dams_guin.plot(ax=ax,
markersize=5,
color='red')
guinea_border.plot(ax=ax, color="none", edgecolor="black", linewidth=2)
ax.legend(handles=[black_line, yellow_patch, green_circle, lime_circle, red_dot],
fontsize=15,
frameon=True,
loc=('upper right'),
title="LEGEND")
ctx.add_basemap(ax, url=ctx.providers.Stamen.Terrain, zoom=0)
ax.set_axis_off()
ax.set_title(
'Figure 5: Guinea Ramsar Areas, Proposed Dams, and Dam Buffer Areas', size=20)
ax.text(0.5, -0.1, "Data Sources: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/), \n Ramsar Sites "
"Information Service (https://rsis.ramsar.org/)",
size=12, ha="center", transform=ax.transAxes)
# -
# # Conclusions
# These two data sets provide a window into protected areas that are threatened by new dam construction country by country, providing the basic code we can use to expand our analysis. Although there are high numbers of dams that are under construction or planned globally---particularly in Africa, Asia, Europe, and South America---there are not wetlands sites within a 5 km and 10 km buffer zone of these dams in most countries. To fully understand the potential impacts we need to include other designated protected areas. We also need to create a more versatile buffer that is different size upstream and downstream of the dam site and possibly rectangular to accommodate the natural geometry of a river valley.
#
# ## Next Steps
# It is fortunate that we have an entire additional course to explore the question of how dams affect protected areas both in the past, present, and future. Using the current datasets, with more time, we can expand the study area to other continents. We will also take into account flow direction (upstream vs. downstream), the proposed reservoir capacity. We will also expand the protected areas considered to include National Parks, World Heritage Sites, and similar.
#
# There are other datasets available for analysis as we expand this work. For example, the World Database of Protected Areas (WDPA), published by the UN Environment, has information on all protected areas around the world. We chose not to use the WDPA data for this project because it will require some cleaning prior to analysis.
#
# # References
# 1. <NAME>. 2014. The Sixth Extinction. New York: Henry Holt and Company. 320 p.
# 2. <NAME>. 2001. Damage Control: Restoring the Physical Integrity of America’s Rivers in The Annals of the Association of American Geographers, 91(1), p. 1-27.
# 3. <NAME>., <NAME>., <NAME>., <NAME>.. 2014. Fragmentation and Flow Regulation of the World’s Large River Systems in Science, 308(5720), p. 405-408. DOI: 10.1126/science.1107887
# 4. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>. 2001. Biodiversity Impacts of Large Dams. International Union for Conservation of Nature and Natural Resource and the United Nations Environmental Programme Report. http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.392.9398&rep=rep1&type=pdf
# 5. <NAME>., <NAME>. 2016. The contribution of reservoirs to global land surface water storage variations in Journal of Hydrometeorology. 17(1), p. 309-325. https://journals.ametsoc.org/doi/full/10.1175/JHM-D-15-0002.1
# 6. <NAME>., <NAME>., <NAME>., <NAME>. 2018. A model of water and sediment balance as determinants of relative sea level rise in contemporary and future deltas. in Geomorphology. 305, p. 209-220.
# 7. <NAME>., <NAME>., <NAME>., <NAME>. 2017. Hydrological threats to riparian wetlands of international importance - a global quantitative and qualitative analysis. in Hydrologic Earth Systems Science. 21, p. 2799-2815.
# 8. <NAME> al. 2019. Mapping the world's free-flowing rivers. in Nature.569(7755), p. 215-221. https://www.researchgate.net/publication/332961728_Mapping_the_world's_free-flowing_rivers#fullTextFileContent
|
presentations/ea-python-dam-blog.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
import pandas as pd
from bcb import currency
df = currency.get(['USD', 'EUR', 'GBP', 'CHF', 'CAD'], start='2010-01-01', end='2022-01-01')
df.tail(30)
df.plot(figsize=(15, 5));
|
notebooks/currency get strong currencies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
ch = np.zeros((9,9))
ch[::2, 1::2]=1
ch[1::2,::2]= 1
plt.imshow(ch, cmap= "gray", interpolation = "nearest")
# +
import skimage
from skimage import data
import matplotlin.pyplot as plt
camera = data.camera()
camera.dtype
# -
plt.imshow(camera)
camera.shape
# +
import os
from skimage import io
filename = os.path.join(os.getcwd(),"nm.jpg")
pic = io.imread(filename)
plt.imshow(pic)
# -
pic.shape
pic.size
pic.dtype
pic.min(),pic.max()
camera.mean()
pic[20,20]
pic[:10]=0
plt.imshow(pic)
# +
from skimage import exposure
moon = data.moon()
plt.imshow(moon)
v_min, v_max = np.percentile(moon, (0.2, 99.8))
print(v_min, v_max)
better_contrast = exposure.rescale_intensity(
moon, in_range=(v_min, v_max))
# -
coins = data.coins()
from skimage import filters
threshold_value = filters.threshold_otsu(coins)
print(threshold_value)
plt.imshow(coins)
|
image_processing_1.1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
values = df.Prices * df.Amount
df['Values'] = values.where(df.Action == 'Sell', other=-values)
In [39]: df
Out[39]:
# + active=""
# '''
# Prices Amount Action Values
# 0 3 57 Sell 171
# 1 89 42 Sell 3738
# 2 45 70 Buy -3150
# 3 6 43 Sell 258
# 4 60 47 Sell 2820
# 5 19 16 Buy -304
# 6 56 89 Sell 4984
# 7 3 28 Buy -84
# 8 56 69 Sell 3864
# 9 90 49 Buy -4410'''
# -
order_df['Value'] = order_df.apply(lambda row: (row['Prices']*row['Amount']
if row['Action']=='Sell'
else -row['Prices']*row['Amount']),
axis=1)
|
content/Pandas Operations/Multiply two columns in a pandas DataFrame and add the result into a new column - TODO.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <hr style="height:.9px;border:none;color:#333;background-color:#333;" />
# <hr style="height:.9px;border:none;color:#333;background-color:#333;" />
#
# <br><h2> A1_Regression_Analysis</h2>
# <h4>DAT-5303 | Machine Learning</h4>
# <NAME> <br>
# Hult International Business School<br><br><br>
#
# <hr style="height:.9px;border:none;color:#333;background-color:#333;" />
# <hr style="height:.9px;border:none;color:#333;background-color:#333;" />
#
# <br><br>
# <h2>Purpose of this Script</h2><br>
# Regression Model Development
#
# In this assignment, you are tasked with using the information in our course case to build a predictive model on a continuous response variable (Y-variable). This assignment encompasses feature engineering, model preparation, variable selection, and model development.
#
#
# Note to the professor: I am aware that the data is messy and out of order. I did my best to keep it organised but then I got confused and it got messy. In addition to that I just wanted to remind you that I do have dyslexia so any spelling mistakes I've made are because of that!
#
#
#
# <br><br>
#
# <hr style="height:.9px;border:none;color:#333;background-color:#333;" /><br>
# +
# importing libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
from sklearn.model_selection import train_test_split
import statsmodels.formula.api as smf # regression modeling
# new libraries
from sklearn.neighbors import KNeighborsRegressor # KNN for Regression
from sklearn.preprocessing import StandardScaler # standard scaler
# +
# setting pandas print options
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# specifying file name
file = "./__datasets/birthweight_low.xlsx"
# reading the file into Python
weight = pd.read_excel(io = file,
header = 0,
sheet_name = 0)
# printing the first 15 rows of the dataset
weight.head(n=15)
# -
#finding more info from the data set
weight.info()
# formatting and printing the dimensions of the dataset
print(f"""
Size of Original Dataset
------------------------
Observations: {weight.shape[0]}
Features: {weight.shape[1]}
""")
#looping to print the column names
for column in weight:
print(column)
# looking at output for columns
weight.columns
# +
# developing a histogram using HISTPLOT
sns.histplot(data = weight,
x = 'bwght',
kde = True)
# title and axis labels
plt.title(label = " Distribution Birth Weight")
plt.xlabel(xlabel = "Birth Weight") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# displaying the histogram
plt.show()
# -
#formatting and printing the dimensions of the dataset
print(f"""
Size of Original Dataset
------------------------
Observations: {weight.shape[0]}
Features: {weight.shape[1]}
""")
# +
#the #analysing the skewness for Birth Weight
weight['bwght'].skew()
# +
# developing a histogram using HISTPLOT
sns.histplot(data = weight,
x = 'mage',
kde = True)
# title and axis labels
plt.title(label = "Original Distribution Mother Age")
plt.xlabel(xlabel = "mage") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# displaying the histogram
plt.show()
# -
#analysing the skewness for mother age
weight['mage'].skew()
# +
#data discription
print("""
CONTINUOUS: bwght (Y-variable) mage aka Mothers Age,
fage aka Fathers Age,
INTERVAL/COUNT
--------------
meduc, monpre, npvis, feduc, omaps, fmaps, male, mwhte,mblck,moth,fwhte,fblck, foth,cigs, drink
""")
# -
#looking for Null values
weight.isnull().sum(axis = 0)
# +
feduc_median = weight['feduc'].median()
npvis_median = weight['npvis'].median()
meduc_mean = weight['meduc'].median()
# filling the missing values
weight['feduc'] = weight['feduc'].fillna(feduc_median)
weight ['npvis'] = weight['npvis'].fillna(feduc_median)
weight ['meduc'] = weight['meduc'].fillna(feduc_median)
#printing to check forr missing values
print(weight.isnull().sum(axis = 0))
# +
# descriptive statistics for numeric data
weight.describe(include = 'number').round(2)
# alternative code
# housing.describe(include = [int, float]).round(2
# +
# building a base model
lm_best = smf.ols(formula = """bwght ~ mage +
meduc +
fage +
fmaps +
cigs +
drink +
male +
mwhte +
fwhte +
fblck """,
data = weight)
# telling Python to FIT the data to the blueprint
results = lm_best.fit()
# printing a summary of the results
print(results.summary())
# + code_folding=[]
# Visual EDA (Scatterplots)
########################
# setting figure size
fig, ax = plt.subplots(figsize = (10, 8))
# developing a scatterplot
plt.subplot(2, 2, 1)
sns.scatterplot(x = weight['mage'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'mothers age ')
plt.ylabel(ylabel = 'birth wieght')
########################
# developing a scatterplot
plt.subplot(2, 2, 2)
sns.scatterplot(x = weight['meduc'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Mother Education')
plt.ylabel(ylabel = 'Birth Weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 3)
sns.scatterplot(x = weight['monpre'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'month prenatal care began ')
plt.ylabel(ylabel = 'Birth Weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 4)
sns.scatterplot(x = weight['npvis'],
y = weight['bwght'],
color = 'r')
# adding labels but not adding title
plt.xlabel(xlabel = 'total number of prenatal visits')
plt.ylabel(ylabel = 'Birth Weight')
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 1 of 2.png')
plt.show()
########################
# setting figure size
fig, ax = plt.subplots(figsize = (10, 12))
# developing a scatterplot
plt.subplot(3, 2, 1)
sns.scatterplot(x = weight['fage'],
y = weight['bwght'],
color = 'y')
# adding labels but not adding title
plt.xlabel(xlabel = 'Fathers Age')
plt.ylabel(ylabel = 'Birth weight')
########################
# developing a scatterplot
plt.subplot(3, 2, 2)
sns.scatterplot(x = weight['feduc'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'Father Education')
plt.ylabel(ylabel = 'Birth wieght')
########################
# developing a scatterplot
plt.subplot(3, 2, 3)
sns.scatterplot(x = weight['omaps'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'one minute apgar score')
plt.ylabel(ylabel = 'Birth wieght')
########################
# developing a scatterplot
plt.subplot(3, 2, 4)
sns.scatterplot(x = weight['fmaps'],
y = weight['bwght'],
color = 'r')
# adding labels but not adding title
plt.xlabel(xlabel = ' five minute apgar score')
plt.ylabel(ylabel = 'Birth wieght')
########################
# developing a scatterplot
plt.subplot(3, 2, 5)
sns.scatterplot(x = weight['cigs'],
y = weight['bwght'],
color = 'r')
# adding labels but not adding title
plt.xlabel(xlabel = 'avg cigarettes per day')
plt.ylabel(ylabel = 'Birth wieght')
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 2 of 2.png')
plt.show()
# +
# Visual EDA (Scatterplots)
########################
# No need
# setting figure size
fig, ax = plt.subplots(figsize = (10, 8))
# developing a scatterplot
plt.subplot(2, 2, 1)
sns.scatterplot(x = weight['drink'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Number of Drinks ')
plt.ylabel(ylabel = 'Birth weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 2)
sns.scatterplot(x = weight['mwhte'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Mother white')
plt.ylabel(ylabel = 'Birth Weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 3)
sns.scatterplot(x = weight['male'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'Male ')
plt.ylabel(ylabel = 'Birth Weight')
########################
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 1 of 2.png')
plt.show()
########################
# setting figure size
fig, ax = plt.subplots(figsize = (10, 12))
# developing a scatterplot
plt.subplot(3, 2, 1)
sns.scatterplot(x = weight['moth'],
y = weight['bwght'],
color = 'y')
# adding labels but not adding title
plt.xlabel(xlabel = 'Mother Other')
plt.ylabel(ylabel = 'Birth weight')
########################
# developing a scatterplot
plt.subplot(3, 2, 2)
sns.scatterplot(x = weight['fwhte'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'Father White')
plt.ylabel(ylabel = 'Birth wieght')
########################
# developing a scatterplot
plt.subplot(3, 2, 3)
sns.scatterplot(x = weight['foth'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Father Other')
plt.ylabel(ylabel = 'Birth wieght')
########################
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 2 of 2.png')
plt.show()
# +
# Visual EDA (Scatterplots)
########################
# setting figure size
fig, ax = plt.subplots(figsize = (10, 8))
# developing a scatterplot
plt.subplot(2, 2, 1)
sns.scatterplot(x = weight['drink'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Number of Drinks ')
plt.ylabel(ylabel = 'Birth weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 2)
sns.scatterplot(x = weight['mwhte'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Mother white')
plt.ylabel(ylabel = 'Birth Weight')
########################
# developing a scatterplot
plt.subplot(2, 2, 3)
sns.scatterplot(x = weight['male'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'Male ')
plt.ylabel(ylabel = 'Birth Weight')
########################
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 1 of 2.png')
plt.show()
########################
# setting figure size
fig, ax = plt.subplots(figsize = (10, 12))
# developing a scatterplot
plt.subplot(3, 2, 1)
sns.scatterplot(x = weight['moth'],
y = weight['bwght'],
color = 'y')
# adding labels but not adding title
plt.xlabel(xlabel = 'Mother Other')
plt.ylabel(ylabel = 'Birth weight')
########################
# developing a scatterplot
plt.subplot(3, 2, 2)
sns.scatterplot(x = weight['fwhte'],
y = weight['bwght'],
color = 'orange')
# adding labels but not adding title
plt.xlabel(xlabel = 'Father White')
plt.ylabel(ylabel = 'Birth wieght')
########################
# developing a scatterplot
plt.subplot(3, 2, 3)
sns.scatterplot(x = weight['foth'],
y = weight['bwght'],
color = 'g')
# adding labels but not adding title
plt.xlabel(xlabel = 'Father Other')
plt.ylabel(ylabel = 'Birth wieght')
########################
# cleaning up the layout, saving the figures, and displaying the results
plt.tight_layout()
plt.savefig('./__analysis_images/weight Scatterplots 2 of 2.png')
plt.show()
# -
#Defining copy_weight
copy_weight = pd.DataFrame.copy(weight)
#defing the x varibles
x_variables= ['mage','meduc','monpre', 'npvis','fage','feduc' ,'omaps', 'fmaps','cigs' ,'drink',
'male' , 'mwhte' ,'mblck','moth',
'fwhte' ,
'fblck' ,
'foth']
# +
# list of continuous features (including Birthweight)
correlation_data = ["mage","meduc","monpre","npvis","fage","feduc","omaps","fmaps","cigs","drink",
"male","mwhte","mblck","moth","fwhte","fblck","foth","bwght"]
# correlation matrix based on continuous features
weight_corr = copy_weight[correlation_data].corr(method = 'pearson')
# filtering the results to only show correlations with bwght
weight_corr.loc[ : , correlation_data].round(decimals = 2).sort_values(ascending = False,
by ="bwght")
# +
# Applying model in scikit-learn
# Preparing a DataFrame based the the analysis above
ols_data = weight.loc[ : , x_variables]
# Preparing the target variable
weight_target = weight.loc[ : , 'bwght']
###############################################
## setting up more than one train-test split ##
###############################################
# FULL X-dataset (normal Y)
x_train_FULL, x_test_FULL, y_train_FULL, y_test_FULL = train_test_split(
weight_data, # x-variables
weight_target, # y-variable
test_size = 0.25,
random_state = 219)
# OLS p-value x-dataset (normal Y)
x_train_OLS, x_test_OLS, y_train_OLS, y_test_OLS = train_test_split(
ols_data, # x-variables
weight_target, # y-variable
test_size = 0.25,
random_state = 219)
# +
# developing a histogram using HISTPLOT
sns.histplot(data = weight,
x = 'bwght',
kde = True)
# title and axis labels
plt.title(label = "Original Distribution of Birth Weight and Count")
plt.xlabel(xlabel = "Birth Weight") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# displaying the histogram
plt.show()
# -
# log transforming weight to the dataset
weight['log_bwght'] = np.log(weight['bwght'])
weight['log_mage'] = np.log(weight['mage'])
weight['log_meduc'] = np.log(weight['meduc'] + 0.001)
# +
# developing a histogram using HISTPLOT
sns.histplot(data = weight,
x = 'log_bwght',
kde = True)
# title and axis labels
plt.title(label = "Logarithmic Distribution of Birth Weight")
plt.xlabel(xlabel = "Log Birth Weight") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# displaying the histogram
plt.show()
# +
# histogram for Lot_Area (skewed positive)
sns.histplot(data = weight,
x = 'log_mage',
kde = True)
plt.title(label = "Logarithmic Distribution of Birth Weight")
plt.xlabel(xlabel = "Birth Weight") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# rendering the plot
plt.show()
# histogram for Mas_Vnr_Area (zero inflated and skewed positive)
sns.histplot(data = weight,
x = 'log_meduc',
kde = True)
plt.title(label = "Logarithmic Distribution of Birth Weight")
plt.xlabel(xlabel = "Birth Weight") # avoiding using dataset labels
plt.ylabel(ylabel = "Count")
# rendering the plot
plt.show()
# +
# building a base model with a log transformed response variable
# INSTANTIATING a model type
lm_price_qual = smf.ols(formula = """log_mage ~ log_meduc""",
data = weight)
# telling Python to FIT the data to the blueprint
results = lm_price_qual.fit()
# printing a summary of the results
print(results.summary())
# +
# looping to detect features with missing values
for col in weight:
# creating columns with 1s if missing and 0 if not
if weight[col].isnull().astype(int).sum() > 0:
weight['m_'+col] = weight[col].isnull().astype(int)
# -
# summing the missing value flags to check the results of the loop above
weight[ ['m_meduc', 'm_feduc',
'm_log_meduc', 'm_npvis'] ].sum(axis = 0)
# +
# creating a dropped dataset to visualize 'Mas Vnr Area'
df_dropped = weight.dropna()
# displaying the plot for 'Mas Vnr Area'
sns.histplot(x = 'm_feduc',
data = df_dropped,
kde = True)
# title and labels
plt.title('Distribution of Female Education')
# displaying the plot
plt.show()
# -
#Understanding Skewness
skew_mage = weight['mage'].skew()
skew_fage = weight['fage'].skew()
skew_cigs = weight['cigs'].skew()
skew_drink = weight['drink'].skew()
print(f"""
Skewness mage = {skew_mage}
Skewness fage = {skew_fage}
Skewness cigs = {skew_cigs}
Skewness drink = {skew_drink}""")
# log for mother age, father age
weight['log_mage'] = np.log(weight['mage'])
print(weight['log_mage'].skew())
weight['log_fage'] = np.log(weight['fage'])
print(weight['log_fage'].skew())
weight.isnull().any().any()
# +
# developing a scatterplot
sns.scatterplot(x = 'log_mage',
y = 'bwght',
data = weight)
# titles and axis labels
plt.title(label = 'Scatterplot with Interval Data')
plt.xlabel(xlabel = ' Log Mothers Age')
plt.ylabel(ylabel = 'Birth weight')
# displaying the plot
plt.show()
# +
# developing a boxplot
sns.boxplot(x = 'feduc',
y = 'bwght',
data = weight)
# titles and axis labels
plt.title(label = 'Boxplot with Interval Data')
plt.xlabel(xlabel = 'Father Education')
plt.ylabel(ylabel = 'Birth Weight')
# displaying the plot
plt.show()
# +
## comparing log transformation results ##
# setting figure size
fig, ax = plt.subplots(figsize = (10, 8))
## Plot 1: Original X, Original Y ##
sns.boxplot(x = 'mage',
y = 'bwght',
data = weight)
# titles and labels
plt.title(label = 'Original X, Original Y')
plt.xlabel(xlabel = 'Mother Age')
plt.ylabel(ylabel = 'Birth Weight')
## displaying the visual ##
plt.tight_layout()
plt.show()
# -
# +
# Model Example 1 ##
# building a fit model
# blueprinting a model type
lm_full = smf.ols(formula = """ bwght ~ mage +
meduc +
monpre +
npvis +
fage +
feduc +
omaps +
fmaps +
cigs +
drink +
male +
mwhte +
mblck +
moth +
fwhte +
fblck +
foth
""",
data = weight)
#Vaibles cigs & drinks
#log father father eductaion
#cigs * drinks
# telling Python to run the data through the blueprint
results_fit = lm_fit.fit()
# printing the results
results_fit.summary()
# +
# preparing explanatory variable data
weight_data = weight.drop(['omaps',
'fmaps',
'bwght',
'log_bwght'],
axis = 1)
# preparing response variable data
weight_target = weight.loc[ : , 'bwght']
# preparing training and testing sets (all letters are lowercase)
x_train, x_test, y_train, y_test = train_test_split(
weight_data,
weight_target,
test_size = 0.25,
random_state = 219)
# checking the shapes of the datasets
print(f"""
Training Data
-------------
X-side: {x_train.shape}
y-side: {y_train.shape[0]}
Testing Data
------------
X-side: {x_test.shape}
y-side: {y_test.shape[0]}
""")
# +
# applying model in scikit-learn
# Preparing a DataFrame based the the analysis above
ols_data = weight.loc[ : , x_variables]
# Preparing the target variable
weight_target = weight.loc[ : , 'bwght']
###############################################
## setting up more than one train-test split ##
###############################################
# FULL X-dataset (normal Y)
x_train_FULL, x_test_FULL, y_train_FULL, y_test_FULL = train_test_split(
weight_data, # x-variables
weight_target, # y-variable
test_size = 0.25,
random_state = 219)
# OLS p-value x-dataset (normal Y)
x_train_OLS, x_test_OLS, y_train_OLS, y_test_OLS = train_test_split(
ols_data, # x-variables
weight_target, # y-variable
test_size = 0.25,
random_state = 219)
# +
# applying modelin scikit-learn
# preparing x-variables from the OLS model
ols_data = weight.loc[ : , x_variables]
# preparing response variable
birthweight_target = weight.loc[ : , 'bwght']
###############################################
## setting up more than one train-test split ##
###############################################
# FULL X-dataset (normal Y)
x_train_FULL, x_test_FULL, y_train_FULL, y_test_FULL = train_test_split(
weight_data, # x-variables
weight_target, # y-variable
test_size = 0.25,
random_state = 219)
# OLS p-value x-dataset (normal Y)
# for OLS look at p values built for limited sample sizes...
x_train_OLS, x_test_OLS, y_train_OLS, y_test_OLS = train_test_split(
ols_data, # x-variables
birthweight_target, # y-variable
test_size = 0.25,
random_state = 219)
# +
#Removing explanatory variables for better p values
# blueprinting a model type
lm_full = smf.ols(formula = """ bwght ~ mage +
cigs +
drink +
mwhte +
mblck +
moth +
fwhte +
fblck +
foth""",
data = weight)
# telling Python to run the data through the blueprint
results_full = lm_full.fit()
# printing the results
results_full.summary()
# +
import sklearn.linear_model # linear models
# INSTANTIATING a model object
lasso_model = sklearn.linear_model.Lasso(alpha = 1.0,
normalize = True) # default magitude
# FITTING to the training data
lasso_fit = lasso_model.fit(x_train_FULL, y_train_FULL)
# PREDICTING on new data
lasso_pred = lasso_fit.predict(x_test_FULL)
# SCORING the results
print('Lasso Training Score :', lasso_model.score(x_train_FULL, y_train_FULL).round(4))
print('Lasso Testing Score :', lasso_model.score(x_test_FULL, y_test_FULL).round(4))
## the following code has been provided for you ##
# saving scoring data for future use
lasso_train_score = lasso_model.score(x_train_FULL, y_train_FULL).round(4) # using R-square
lasso_test_score = lasso_model.score(x_test_FULL, y_test_FULL).round(4) # using R-square
# displaying and saving the gap between training and testing
print('Lasso Train-Test Gap :', abs(lasso_train_score - lasso_test_score).round(4))
lasso_test_gap = abs(lasso_train_score - lasso_test_score).round(4)
# +
# INSTANTIATING a model object
ard_model = sklearn.linear_model.ARDRegression()
# FITTING the training data
ard_fit = ard_model.fit(x_train_FULL, y_train_FULL)
# PREDICTING on new data
ard_pred = ard_fit.predict(x_test_FULL)
print('Training Score:', ard_model.score(x_train_FULL, y_train_FULL).round(4))
print('Testing Score :', ard_model.score(x_test_FULL, y_test_FULL).round(4))
# saving scoring data for future use
ard_train_score = ard_model.score(x_train_FULL, y_train_FULL).round(4)
ard_test_score = ard_model.score(x_test_FULL, y_test_FULL).round(4)
# displaying and saving the gap between training and testing
print('ARD Train-Test Gap :', abs(ard_train_score - ard_test_score).round(4))
ard_test_gap = abs(ard_train_score - ard_test_score).round(4)
# -
# +
# declaring set of x-variables
x_variables = ['mage','cigs','drink','mwhte','mblck',
'moth','fwhte','fblck','foth']
# looping to make x-variables suitable for statsmodels
for val in x_variables:
print(f"{val} +")
# +
# merging X_train and y_train so that they can be used in statsmodels
birthweight_train = pd.concat([x_train, y_train], axis = 1)
# Step 1: build a model
lm_best = smf.ols(formula = """bwght ~ fage +
cigs +
drink +
mwhte +
mblck +
moth +
fwhte +
fblck +
foth""",
data = birthweight_train)
# Step 2: fit the model based on the data
results = lm_best.fit()
# Step 3: analyze the summary output
print(results.summary())
# +
# INSTANTIATING a model object
from sklearn.linear_model import LinearRegression # linear regression (scikit-learn)
lr = LinearRegression()
# FITTING to the training data
lr_fit = lr.fit(x_train_OLS, y_train_OLS)
# PREDICTING on new data
lr_pred = lr_fit.predict(x_test_OLS)
# SCORING the results
print('OLS Training Score :', lr.score(x_train_OLS, y_train_OLS).round(4))
print('OLS Testing Score :', lr.score(x_test_OLS, y_test_OLS).round(4))
# saving scoring data for future use
lr_train_score = lr.score(x_train_OLS, y_train_OLS).round(4) # using R-square
lr_test_score = lr.score(x_test_OLS, y_test_OLS).round(4) # using R-square
# displaying and saving the gap between training and testing
print('OLS Train-Test Gap :', abs(lr_train_score - lr_test_score).round(4))
lr_test_gap = abs(lr_train_score - lr_test_score).round(4)
# +
#Instantiating a model object
lr = LinearRegression()
#fitting to the training data
lr_fit = lr.fit(x_train_OLS, y_train_OLS)
#predicting on new data
lr_pred = lr_fit.predict(x_test_OLS)
#Scoring the results
print('OLS Training Score :', lr.score(x_train_OLS, y_train_OLS).round(4)) # using R squared
print('OLS Testing Score :', lr.score(x_test_OLS, y_test_OLS).round(4)) #using R squared
lr_train_score = lr.score(x_train_OLS, y_train_OLS).round(4)
lr_test_score = lr.score(x_test_OLS, y_test_OLS).round(4)
#displaying and saving the gap between training and testing
print('OLS Train-Test Gap :' , abs(lr_train_score - lr_test_score).round(4))
lr_test_gap = abs(lr_train_score - lr_test_score).round(4)
# -
print("""I am very lost and having a panic attack with
10 to mins to spare, I've accepted that I'm going to
fail this assignment.
I will book a one on one with you
""")
|
Lozano_Jessica_A1_Regression_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Actor Critic - Syft Duet - Data Scientist 🥁
#
# Contributed by [@Koukyosyumei](https://github.com/Koukyosyumei)
# ## PART 1: Connect to a Remote Duet Server
#
# As the Data Scientist, you want to perform data science on data that is sitting in the Data Owner's Duet server in their Notebook.
#
# In order to do this, we must run the code that the Data Owner sends us, which importantly includes their Duet Session ID. The code will look like this, importantly with their real Server ID.
#
# ```
# import syft as sy
# duet = sy.duet('xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxx')
# ```
#
# This will create a direct connection from my notebook to the remote Duet server. Once the connection is established all traffic is sent directly between the two nodes.
#
# Paste the code or Server ID that the Data Owner gives you and run it in the cell below. It will return your Client ID which you must send to the Data Owner to enter into Duet so it can pair your notebooks.
# +
from itertools import count
from collections import namedtuple
import numpy as np
import torch
import syft as sy
duet = sy.join_duet(loopback=True)
sy.logger.add(sink="./syft_ds.log")
# -
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 0 : Now STOP and run the Data Owner notebook until Checkpoint 1.
sy.load("gym")
sy.load("numpy")
# +
config = {
"gamma": 0.99,
"seed": 543,
"render": False,
"log_interval": 10,
"no_cuda": False,
"log_interval": 1,
"wait_interval": 1,
"dry_run":True,
}
remote_torch = duet.torch
remote_torch.manual_seed(config["seed"])
# +
has_cuda = False
has_cuda_ptr = remote_torch.cuda.is_available()
# lets ask to see if our Data Owner has CUDA
has_cuda = bool(
has_cuda_ptr.get(
request_block=True,
reason="To run test and inference locally",
timeout_secs=3, # change to something slower
)
)
print("Is cuda available ? : ", has_cuda)
use_cuda = not config["no_cuda"] and has_cuda
# now we can set the seed
remote_torch.manual_seed(config["seed"])
device = remote_torch.device("cuda" if use_cuda else "cpu")
# print(f"Data Owner device is {device.type.get()}")
# -
SavedAction = namedtuple("SavedAction", ["log_prob", "value"])
# +
buffer_saved_actions = []
buffer_rewards =[]
class Policy(sy.Module):
"""
implements both actor and critic in one model
"""
def __init__(self, torch_ref):
super(Policy, self).__init__(torch_ref=torch_ref)
self.affine1 = self.torch_ref.nn.Linear(4, 128)
# actor's layer
self.action_head = self.torch_ref.nn.Linear(128, 2)
# critic's layer
self.value_head = self.torch_ref.nn.Linear(128, 1)
# action & reward buffer
# self.saved_actions = []
# self.rewards = []
def forward(self, x):
"""
forward of both actor and critic
"""
x = remote_torch.relu(self.affine1(x))
# actor: choses action to take from state s_t
# by returning probability of each action
action_prob = remote_torch.softmax(self.action_head(x), dim=-1)
# critic: evaluates being in the state s_t
state_values = self.value_head(x)
# return values for both actor and critic as a tuple of 2 values:
# 1. a list with the probability of each action over the action space
# 2. the value from state s_t
return action_prob, state_values
# +
# send our model to remote
policy = Policy(torch)
remote_policy = policy.send(duet)
optimizer = remote_torch.optim.Adam(remote_policy.parameters(), lr=3e-2)
eps = np.finfo(np.float32).eps.item()
# -
# if we have CUDA lets send our model to the GPU
if has_cuda:
remote_policy.cuda(device)
else:
remote_policy.cpu()
# +
# You cannot see the state
def select_action(state):
global buffer_saved_actions
global buffer_rewards
state = remote_torch.from_numpy(state).float()
probs_ptr, state_value_ptr = remote_policy(state)
# create a categorical distribution over the list of probabilities of actions
m = remote_torch.distributions.Categorical(probs_ptr)
# and sample an action using the distribution
action = m.sample()
# save to action buffer
buffer_saved_actions.append(SavedAction(m.log_prob(action),
state_value_ptr))
# the action to take (left or right)
return action.item()
def finish_episode():
"""
Training code. Calculates actor and critic loss and performs backpropagation.
"""
global buffer_saved_actions
global buffer_rewards
gamma = duet.python.Float(config["gamma"])
R = duet.python.Float(0)
policy_losses = duet.python.List([])
value_losses = duet.python.List([])
returns = duet.python.List([])
for r in buffer_rewards[::-1]:
R = r + gamma * R
returns.insert(0, R)
returns = remote_torch.Tensor(returns)
returns = (returns - returns.mean()) / (returns.std() + eps)
for (log_prob, value), R in zip(buffer_saved_actions, returns):
advantage = R - value.item()
# calculate actor (policy) loss
policy_losses.append(-log_prob * advantage)
# calculate critic (value) loss using L1 smooth loss
value_losses.append(remote_torch.nn.functional.smooth_l1_loss(value,
R.reshape(1)))
# reset gradients
optimizer.zero_grad()
# sum up all the values of policy_losses and value_losses
loss = remote_torch.stack(policy_losses).sum() + remote_torch.stack(value_losses).sum()
# perform backprop
loss.backward()
optimizer.step()
# reset rewards and action buffer
del buffer_saved_actions[:]
del buffer_rewards[:]
# -
reward_threshold_ptr = duet.store["reward_threshold"]
reward_threshold = reward_threshold_ptr.get(request_block=True, delete_obj=False)
print(f"reward_threshold is {reward_threshold}")
remote_gym = duet.gym
remote_env = remote_gym.make("CartPole-v0")
remote_env.seed(config["seed"])
# +
running_reward = 10
# run inifinitely many episodes
for i_episode in count(1):
# reset environment and episode reward
state = remote_env.reset()
ep_reward = duet.python.Float(0)
# for each episode, only run 9999 steps so that we don't
# infinite loop while learning
for t in range(1, 10000):
# select action from policy
action = select_action(state)
# take the action
state, reward, done, _ = remote_env.step(action)
buffer_rewards.append(reward)
ep_reward += reward
if done.get(request_block=True):
break
# update cumulative reward
running_reward = 0.05 * ep_reward.get(request_block=True, delete_obj=False) + (1 - 0.05) * running_reward
# perform backprop
finish_episode()
# log results
if i_episode % config["log_interval"] == 0:
print(
"Episode {}\tLast reward: {:.2f}\tAverage reward: {:.2f}".format(
i_episode,
ep_reward.get(request_block=True, delete_obj=False),
running_reward
)
)
# check if we have "solved" the cart pole problem
if running_reward > reward_threshold:
print(
"Solved! Running reward is now {} and "
"the last episode runs to {} time steps!".format(running_reward, t)
)
break
if config["dry_run"]:
break
# -
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 1 : Now STOP and run the Data Owner notebook until Checkpoint 2.
|
packages/syft/examples/duet/reinforcement_learning/Actor_Critic_Syft_Data_Scientist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # `surfs-up`
# + active=""
# {% capture index %}
# {% for page in site.pages %}
# * [{{page.path}}]({{site.baseurl}}/{{page.path}})
#
# {% endfor %}
# {% endcapture %}
#
# {{index |markdownify}}
|
docs/test-index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## An example notebook
#
# Note that due to the `ipython_config.py` you do not need to use `pip install .` in order to import local files.
from mlmax.preprocessing import read_data
d = read_data("s3://sagemaker-sample-data-us-east-1/processing/census/census-income.csv")
d.head()
|
notebooks/example_notebook.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// # Optimisation of analytical functions
// +
// include header files
#include <iostream>
#include "xtensor/xarray.hpp"
#include "xtensor/xnpy.hpp"
#include "xtensor/xio.hpp"
#include "xevo/ga.hpp"
#include "xevo/analytical_functions.hpp"
// -
// ## Rosenbrock function
// Optimise Rosenbrock function with `xevo::ga`.
//
// Rosenbrock function is expressed as:
//
// $$
// f(x_1, x_2) = 100(x_1^2 - x_2) + (1 - x_1)^2 \quad with \quad \bf{X} \quad \in \left[-3, 3\right]
// $$
{
std::array<std::size_t, 2> shape = {40, 2};
xt::xarray<double> X = xt::zeros<double>(shape);
xevo::Rosenbrock_scaled objective_f;
xevo::ga genetic_algorithm;
genetic_algorithm.initialise(X);
std::size_t num_generations = 300;
std::size_t no_frames = 10;
std::size_t skip = static_cast<std::size_t>(std::floor(num_generations / no_frames));
for (auto i{0}; i<num_generations; ++i)
{
genetic_algorithm.evolve(X, objective_f, std::make_tuple(0.05),
std::make_tuple(), std::make_tuple(0.8),
std::make_tuple(0.5, 60.0));
if (i % skip == 0)
{
std::string pop_file = "./output/rose_pop_" + std::to_string(i) + ".npy";
xt::dump_npy(pop_file, X);
}
}
std::cout << "Last pop: \n" << X << "\n" << std::endl;
}
// ## Branin function
// Optimise Branin function with `xevo::ga`.
//
// Branin function is expressed as:
//
// $$
// f(x) = ( x_2 - \frac{5.1}{4\pi^2}x_2 + \frac{5}{\pi}x_1 - 6 )^2 + 10\left[ (1 - \frac{1}{8\pi})\cos{x_1} + 1 \right] + 5x_1\\
// with \quad x_1 \in \left[-5,10\right], x_2 \in \left[0,15\right]
// $$
{
std::array<std::size_t, 2> shape = {40, 2};
xt::xarray<double> X = xt::zeros<double>(shape);
xevo::Branin objective_f;
xevo::ga genetic_algorithm;
genetic_algorithm.initialise(X);
std::size_t num_generations = 300;
auto y = objective_f(X);
auto sort = xt::argsort(y);
double y_best = y(sort(39));
std::size_t stall{0};
for (auto i{0}; i<num_generations; ++i)
{
genetic_algorithm.evolve(X, objective_f, std::make_tuple(0.05),
std::make_tuple(), std::make_tuple(0.85),
std::make_tuple(0.5, 60.0));
y = objective_f(X);
sort = xt::argsort(y);
double y_best_n = y(sort(39));
if ((fabs(y_best - y_best_n) <= 1e-006))
{
if (stall <= 50)
{
++stall;
}
else
{
std::cout << i << std::endl;
break;
}
}
else
{
stall = 0;
}
y_best = y_best_n;
}
std::cout << "Last pop: \n" << X << "\n" << std::endl;
}
|
junbs/optimisation_analytical_functions_ga.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import necessary modules
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
pd.set_option('max_columns', None)
import nltk
import collections as co
from wordcloud import WordCloud, STOPWORDS
# %matplotlib inline
# -
#read loans.csv as a dataframe
loans_df = pd.read_csv('~/Downloads/tanay/loan.csv',low_memory=False, engine='c')
loans_df.head(2)
loans_df.info()
loans_df.describe()
# +
#visual EDA to understand the distribution of loan amount
sns.set_style("whitegrid")
fig, axs = plt.subplots(2,1,figsize=(20,20))
sns.distplot(loans_df.loan_amnt, ax=axs[0], hist=True, kde=True, bins=15)
axs[0].set(xlabel='Loan Amount',
ylabel='% Distribution',title='Density Plot of Loan Amount')
sns.violinplot(loans_df.loan_amnt, ax=axs[1], color='0.6')
axs[1].set(xlabel='Loan Amount',
ylabel='Distribution',title='Violin Plot of Loan Amount')
plt.show()
# -
loans_df['loan_status'].unique()
#define a function to classify loan status into one of the following bins ('Fully Paid', 'Default', 'Current')
def loan_status_bin(text):
if text in ('Fully Paid', 'Does not meet the credit policy. Status:Fully Paid'):
return 'Fully Paid'
elif text in ('Current', 'Issued'):
return 'Current'
elif text in ('Charged Off', 'Default', 'Does not meet the credit policy. Status:Charged Off'):
return 'Default'
elif text in ('Late (16-30 days)', 'Late (31-120 days)', 'In Grace Period'):
return 'Late'
else:
'UNKNOWN BIN'
#create a new attribute 'loan_status_bin' in the dataframe
loans_df['loan_status_bin']=loans_df['loan_status'].apply(loan_status_bin)
loans_df['loan_status_bin'].unique()
# +
#visual EDA of the loan status bin, and a violin plot which captures the loan amount along with the loan status bin
sns.set_style("whitegrid")
fig, axs = plt.subplots(1,2,figsize=(18,8))
loans_df.groupby('loan_status_bin').size().plot(kind='pie', ax=axs[0]);
axs[0].set(title='Pie Plot of Loan Status bin')
sns.violinplot(x=loans_df['loan_status_bin'], y=loans_df['loan_amnt'], ax=axs[1])
axs[1].set(xlabel='Loan Status bin',
ylabel='Loan Amount',title='Violin Plot of Loan Status bin and Loan Amount')
plt.show()
# +
#word cloud
plt.rcParams['figure.figsize'] = (12,12)
loans_df['title'].unique()
list_wc = list()
loans_df['title'].apply(lambda x: list_wc.append(x))
string_wc=str(list_wc)
wordcloud = WordCloud(stopwords=STOPWORDS, background_color='white', max_words=120, width=800, height=500).generate(string_wc)
plt.imshow(wordcloud)
plt.axis('off')
plt.show()
# -
loans_df.desc.unique()
#(loan status bin, term, title)
loans_df.title.head(10)
np.unique(loans_df.emp_length.astype(str))
loans_df.id[loans_df['emp_length'].isnull()].count()
def employment_len(text):
if text=='10+ years':
return '10+ years'
elif text in ('1 year','2 years', '3 years', '4 years','5 years'):
return '1 to 5 years'
elif text in ('6 years', '7 years', '8 years', '9 years'):
return '6 to 9 years'
elif text== '< 1 year':
return 'Less than 1 year'
else:
return 'n/a'
plt.rcParams['figure.figsize'] = (10,10)
loans_df['employment_length']=loans_df['emp_length'].apply(employment_len)
_=loans_df.groupby('employment_length').size().plot(kind='bar')
loans_df[loans_df['annual_inc'].isnull()==True]['annual_inc']
print('Median annual income is {0} and Mean annual income is {1}'.format(loans_df['annual_inc'].median(), loans_df['annual_inc'].mean()))
loans_df.fillna(loans_df.median()['annual_inc'], inplace=True)
loans_df[loans_df['annual_inc'].isnull()==True]['annual_inc'].count()
loans_df_fp=loans_df[loans_df['loan_status_bin']=='Fully Paid']
loans_df_def=loans_df[loans_df['loan_status_bin']=='Default']
loans_df_fp.describe()
loans_df_def.describe()
_=loans_df_fp[['id', 'addr_state']].groupby(by='addr_state')['id'].count().nlargest(10).plot(kind='bar')
_=loans_df_def[['id', 'addr_state']].groupby(by='addr_state')['id'].count().nlargest(10).plot(kind='bar')
ax = sns.regplot(x="loan_amnt", y="annual_inc", data=loans_df_fp)
ax = sns.regplot(x="loan_amnt", y="annual_inc", data=loans_df_def)
|
springboard_modules/5_3_5_data_wrangling_lending_club.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Parsing and manipulating PDB files
# +
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
# + Two important pieces of metadata for a crystal structure are $R$ and $R_{free}$. These measure how well the crystallographic model fits experimental data used to generate the model. The location of these numbers are indicated in pdb files by the following text.
#
# ```
# REMARK 3 R VALUE (WORKING + TEST SET)
#
# and
#
# REMARK 3 FREE R VALUE
# ```
#
# Write a function called **get_R** that takes a string specifying a pdb file as an argument, extracts $R$ and $R_{free}$ from the file, and then returns them as a tuple of floats `(R,R_free)`.
#
#
# +
def get_R(pdb_file):
f = open(pdb_file,"r")
lines = f.readlines()
f.close()
for l in lines:
if l.startswith("REMARK 3 R VALUE (WORKING + TEST SET)"):
R = float(l[47:55])
if l.startswith("REMARK 3 FREE R VALUE "):
R_free = float(l[47:55])
return (R,R_free)
get_R("1gzx.pdb")
# + Create a histogram of all $C_{\alpha,i} \rightarrow C_{\alpha,i+1}$ distances in the pdb file `1stn.pdb`, where $i$ counts along residues in the sequence. (This means the distance between $C_{\alpha}$ for residue 1 and 2, then for 2 and 3, etc.)
# +
# Load file
f = open("1stn.pdb")
lines = f.readlines()
f.close()
# Loop through lines
out = []
for l in lines:
# Grab ATOM entries
if l[:4] == "ATOM":
# Grab CA atoms
if l[13:15] == "CA":
# Grab coordinates
x = float(l[30:38])
y = float(l[38:46])
z = float(l[46:54])
out.append((x,y,z))
# Convert to an Nx3 numpy array
coord = np.array(out)
# dx, dy, dz
steps = coord[1:,:] - coord[:-1,:]
# rmsd
rmsd = np.sqrt(np.sum((steps)**2,axis=1))
fig, ax = plt.subplots()
ax.hist(rmsd)
# -
# Write a function called **center_of_mass** that calculates the center of mass for a pdb file. It should take a string specifying the pdb file as an argument and return a tuple with the center of mass in (x, y, z). The center of mass is the average of the coordinates of the protein atoms, weighted by their mass. The type of each atom is given on the far right of each line of the pdb. You should use the following masses for the atoms:
#
# |atom | mass |
# |-----|----------|
# | `C` | 12.0107 |
# | `N` | 14.0067 |
# | `O` | 15.9994 |
# | `S` | 32.065 |
# +
def center_of_mass(pdb_file):
"""
Get the center of mass from a pdb file.
center of mass is for coordinate D over all atoms i is:
sum(mass_i*D_i)/sum(mass_i)
pdb_file: string indicating the pdb file to analyze
"""
# Masses of atom times
mass_dict = {"C":12.0107,
"N":14.0067,
"O":15.9994,
"S":32.065}
# Read lines from file
f = open(pdb_file,'r')
lines = f.readlines()
f.close()
# Initialize variables
com = np.zeros(3,dtype=np.float)
total_mass = 0.0
# Go through every line
for l in lines:
# Grab ATOM entries
if l[:4] == "ATOM":
# Get atom type, atom mass
atom_type = l[77]
atom_mass = mass_dict[atom_type]
# Extract x, y, and z and multiply by atom mass
# Append each of these to the center of mass
com[0] += float(l[30:38])*atom_mass
com[1] += float(l[38:46])*atom_mass
com[2] += float(l[46:54])*atom_mass
total_mass += atom_mass
return com/total_mass
center_of_mass("1stn.pdb")
# -
# The `HN` hydrogen atom attached to the `N` is often not in crystal structures because it is invisible in the diffraction experiment used to make the model. Unfortunately, the `HN` atom coordinates are necessary to calculate things like structural biologists care about like [Ramachandran plots](https://en.wikipedia.org/wiki/Ramachandran_plot). The missing atom is indicated with the red arrow in the figure below.
#
# 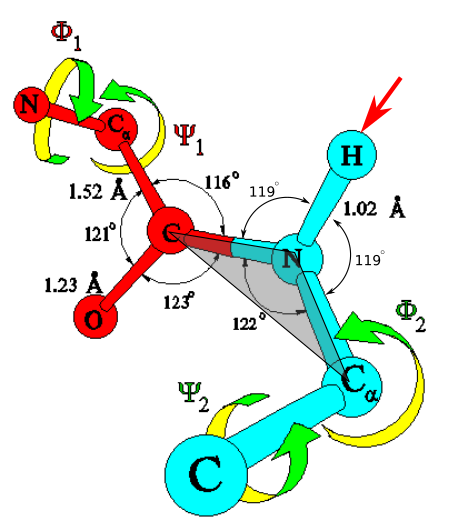
#
# The function below (`calc_hn`) calculates the position of the `HN` atom for the $i^{th}$ residue given the coordinates of the $(i-1)^{th}$ `C` atom (red sphere in picture), the $i^{th}$ `N` atom (cyan sphere in picture) and the $i^{th}$ `CA` atom (cyan sphere in picture). Write a program that takes a pdb file as input, calculates the position of each `HN` atom, and then writes out a new pdb file with the `HN` atoms written out as lines just after the `N` atoms. This means the line encoding the position of `N` for residue 46 would be followed by a new line encoding the position of `HN` for residue 46. You do not have to renumber the atoms in the file (but bonus points if you do).
# +
def calc_hn(CO_i_minus_one,N_i,CA_i):
"""
Calculate the position of the HN proton.
CO_i_minus_one: array of x,y,z coordinates for *previous*
"C" atom
N_i: array of x,y,z coordinates for current "N" atom.
CA_i: array of x,y,z coordinates for current "CA" atom.
Returns: x,y,z array for HN proton.
"""
# Center on N
Ca = CA_i - N_i
Co = CO_i_minus_one - N_i
# Get length of relevant vectors
length_HN = 1.02
length_Ca = np.sqrt(np.sum(Ca**2))
length_Co = np.sqrt(np.sum(Co**2))
# Dot product of H and C
H_dot_C = length_HN*length_Co*np.cos(119.0*np.pi/180.0)
xo = Co[0]
yo = Co[1]
zo = Co[2]
xa = Ca[0]
ya = Ca[1]
za = Ca[2]
Q = length_Ca/length_Co
A = (xo + Q*xa)
B = (yo + Q*ya)
C = (zo + Q*za)
xh = H_dot_C/(xo + yo*B/A + zo*C/A)
yh = xh*B/A
zh = xh*C/A
# Translate HN back to original coordinates
HN_i = np.array((xh,yh,zh)) + N_i
return HN_i
# +
def add_hn(pdb_file,output_pdb_file):
"""
Calculate the positions of the HN atom in a protein pdb file
and write them to a new pdb file.
pdb_file: pdb file without HN atoms
output_pdb_file: pdb file to write out. Warning: if this exists,
it will be overwritten.
"""
# Read lines from pdb file
f = open(pdb_file)
lines = f.readlines()
f.close()
# Dictionary to store information for calculation
df_dict = {"atom":[],
"resid":[],
"x":[],
"y":[],
"z":[]}
# Grab relevant atoms from the lines
for l in lines:
if l[:4] == "ATOM":
# Grab atoms of interest
if l[13:16] in ["N ","CA ","C "]:
# Grab atom and residue number
atom = l[13:16].strip()
resid = l[21:26].strip()
# Record in df_dict
df_dict["atom"].append(atom)
df_dict["resid"].append(resid)
df_dict["x"].append(float(l[30:38]))
df_dict["y"].append(float(l[38:46]))
df_dict["z"].append(float(l[46:54]))
# Convert to data frame
df = pd.DataFrame(df_dict)
# Dict will hold HN positions
HN_dict = {}
# Go over all residues seen
resids = np.unique(df.resid)
for i in range(1,len(resids)):
# Name of i and i-1 residues
r_i = resids[i]
r_i_minus_one = resids[i-1]
# Grab previous CO, N and CA from the data frame
CO_i_minus_one = np.array(df[np.logical_and(df.resid == r_i_minus_one,
df.atom == "C") ].iloc[0,2:])
N_i = np.array(df[np.logical_and(df.resid == r_i,
df.atom == "N") ].iloc[0,2:])
CA_i = np.array(df[np.logical_and(df.resid == r_i,
df.atom == "CA")].iloc[0,2:])
# Calculate position of the HN atom
HN_i = calc_hn(CO_i_minus_one,N_i,CA_i)
# Record it in the dictionary
HN_dict[r_i] = HN_i
out = []
for l in lines:
out.append(l)
# Look for "N" atom and and "HN after that"
if l[:4] == "ATOM" and l[13:16] == "N ":
# See if we calculated HN for this residue
resid = l[21:26].strip()
try:
HN_i = HN_dict[l[21:26].strip()]
except KeyError:
pass
out.append("{}HN {}{:8.3f}{:8.3f}{:8.3f}{}H\n".format(l[:13],l[16:30],
HN_i[0],HN_i[1],HN_i[2],
l[54:77]))
g = open(output_pdb_file,"w")
g.write("".join(out))
g.close()
add_hn("1stn.pdb","junk.pdb")
# -
|
labs/03_molecular-structure/03_structure-files_key.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (SageMath)
# language: python
# name: python3-sage
# resource_dir: /usr/local/share/jupyter/kernels/python3-sage
# ---
writefile("ch12.tex",latex_st) #use after creating a new latex_st1
# +
#PJ: This is a lengthy program that parses LaTeX formulas and extracts substrings from latex_st
from IPython.display import display, Math
from provers import *
import time
def readfile(fn):
fh=open(fn)
st=fh.read()
fh.close()
return st
latex_fn="SPoAchapters.tex"
latex_st=readfile(latex_fn)
print(latex_fn,len(latex_st),'characters')
#latex_fn="SPoAch2-5.tex"
#latex_st+=readfile(latex_fn)
#print("+"+latex_fn,len(latex_st),'characters')
#latex_fn="SPoAch6-9.tex"
#latex_st+=readfile(latex_fn)
#print("+"+latex_fn,len(latex_st),'characters')
def writefile(fn,st):
fh=open(fn,'w')
fh.write(st)
fh.close()
return "Wrote "+str(len(st))+" characters"
# Signature for input and output (the LaTeX symbols can be changed to agree with other conventions)
# Constant operation symbols
Cons="c"; Cond="d"; Iden="1"; Zero="0"; Bot="\\bot"; Top="\\top"
# Prefix unary operation symbols
Fop="f"; Gop="g"; Lneg="\\sim"; Rneg="-"; Dia="\\diamond"; Box="\\box"; Rtri="\\vartriangleright"; Ltri="\\vartriangleleft"
# Postfix unary operation symbols
Star="^*"; Inv="^{-1}"; Conv="\\smallsmile"; Pri="'"
# Infix binary operation symbols
Meet="\\wedge"; Join="\\vee"; Mult="\\cdot"; Lres="/"; Rres="\\backslash"; Omult="\\odot"; Smul="*";
Add="+"; Oadd="\\oplus"; Comp="\\circ"; Rimp="\\to"; Limp="\\leftarrow"; Ominus="\\ominus"; triR="\\triangleright"; triL="\\triangleleft"
# Infix binary relation symbols
Le="\\le"; Ge="\\ge"; Eq="="; Neq="\\ne"
# First-order logic connectives and quantifiers
And="\\text{ and }"; Or="\\text{ or }"; Imp="\\implies"; Not="\\neg"; Iff="\\iff"; All="\\forall"; Exists="\\exists"
VAR = set(["u","v","w","x","y","z"])|set("x_"+str(i) for i in range(10))|set("y_"+str(i) for i in range(10))
CONST = set([Cons,Cond,Iden,Zero,Bot,Top])
PREFIX = [(Fop,11),(Gop,11),(Lneg,11),(Rneg,11),(Dia,11),(Box,11),(Rtri,11),(Ltri,11),(Not,5)] # (symbol, precedence)
POSTFIX = [(Star,12),(Inv,12),(Conv,12),(Pri,12)]
INFIX = [(Mult,10),(Omult,10),(Comp,10),(Rres,9),(Lres,9),(triR,9),(triL,9),(Add,8),(Oadd,8),(Ominus,8),(Meet,7),(Join,7),
(Rimp,6),(Limp,6),(Le,5),(Eq,5),(Neq,5),(And,4),(Or,4),(Imp,3),(Iff,2)]
QUANT = [(All,5),(Exists,5)]
# can add further \newcommand macros in the string below
Macros="" #r"\newcommand{\coimp}{-\!\raisebox{.5pt}{\scriptsize<}\,}"
p9sym={"u":"u","v":"v","w":"w","x":"x","y":"y","z":"z","c":"c","d":"d","1":"1","0":"0","\\bot":"b","\\top":"t","f":"f","g":"g",
"\\sim":"~","-":"-","\\diamond":"dd","\\box":"bx","\\triangleright":" r ","\\triangleleft":" t ","\\vartriangleright":"tr","\\vartriangleleft":"tl",
"^*":"'","^{-1}":"i","\\smallsmile":"'","'":"'","\\wedge":"^","\\vee":" v ","\\cdot":"*","/":"/","\\backslash":"\ ",
"\\odot":"*","*":"*","+":"+","\\oplus":"+","\\ominus":"<-","\\circ":"*","\\to":"->","\\leftarrow":"<-",
"\\le":"<=","\\ge":">=","=":"=","\\ne":"!=","\\text{ and }":" & ","\\text{ or }":" | ",
"\\implies":" -> ","\\iff":" <-> ","\\forall":"all","\\exists":"exists"}
opts=["op(700,infix,\"r\")","op(700,infix,\"t\")"]
################## Parser code (can ignore this) #################
# Terms are read using Vaughn Pratt's top-down parsing algorithm #
symbol_table = {}
def wrap(subt, t): # decide when to add parentheses during printing of terms
return subt.tex() if subt.lbp > t.lbp or len(subt.a)<=1 else "("+subt.tex()+")"
class symbol_base(object):
a = []
def __repr__(self):
return self.tex()
def tex(self):
if len(self.a) == 0: return self.id
if len(self.a) == 1:
if self.id[0]=="^": return wrap(self.a[0],self)+self.id
return self.id+" "+wrap(self.a[0],self)
if len(self.a) == 2:
return wrap(self.a[0],self)+self.id+(" " if self.id[0]=='\\' else "")+wrap(self.a[1],self)
return self.id+" "+self.a[0].id+self.a[1].id+self.a[2].id
def symbol(id, bp=0): # identifier, binding power
if id in symbol_table:
s = symbol_table[id] # look symbol up in table
s.lbp = max(bp, s.lbp) # update left binding power
else:
class s(symbol_base): # create class for this symbol
pass
s.id = id
s.lbp = bp
s.nulld = lambda self: self
symbol_table[id] = s
return s
def advance(id=None):
global token
if id and token.id != id:
raise SyntaxError("Expected "+id+" got "+token.id)
token = next()
def nulld(self): # null denotation
expr = expression()
advance(")")
return expr
def infix(id, bp):
def leftd(self, left): # left denotation
self.a = [left]
self.a.append(expression(bp))
return self
symbol(id, bp).leftd = leftd
def prefix(id, bp):
global token
def nulld(self): # null denotation
global token
if token.id != "(":
self.a = [expression(bp)]
return self
else:
token = next()
self.a = []
if token.id != ")":
while 1:
self.a.append(expression())
if token.id != ",":
break
advance(",")
advance(")")
return self
symbol(id, bp).nulld = nulld
def postfix(id, bp):
def leftd(self,left): # left denotation
self.a = [left]
return self
symbol(id, bp).leftd = leftd
symbol("(").nulld = nulld
symbol(")")
symbol("[").nulld = nulld
symbol("]")
symbol("(end)")
for st in VAR|CONST: symbol(st)
for t in PREFIX: prefix(t[0],t[1])
for t in POSTFIX: postfix(t[0],t[1])
for t in INFIX: infix(t[0],t[1])
for st in VAR:
for t in QUANT: prefix(t[0]+" "+st,t[1])
def tokenize(st):
i = 0
while i<len(st):
tok = st[i] #read single-letter token
j = i+1
if j<len(st) and st[j]=="_": #read subscript
j+=1
if st[j]=="{": j+=1
while j<len(st) and st[j]>='0' and st[j]<='9': j+=1
if j<len(st) and st[j]=="}": j+=1
tok = st[i:j]
elif j<len(st) and st[i]=="^": #read postfix superscript operation
if st[j]=="{": j+=1
if st[j]=="-" or st[j]=="*" or st[j]=="\\": j+=1
if st[j]=="1": j+=1
if st[j-1]=='\\':
while j<len(st) and ((st[j]>='a' and st[j]<='z') or (st[j]>='A' and st[j]<='Z')): j+=1
if j<len(st) and st[j]=="}": j+=1
tok = st[i:j]
elif tok=="{":
tok = st[j]
j+=1
if tok=="\\": #read Latex symbol
while j<len(st) and ((st[j]>='a' and st[j]<='z') or (st[j]>='A' and st[j]<='Z')): j+=1
if st[i]=="{" and st[j]=="}": j+=1
tok = st[i:j]
if tok in ["\\mathbf","\\forall","\\exists"]:
j+=2
if j<len(st) and st[j]=="_": #read subscript
j+=1
if st[j]=="{": j+=1
while j<len(st) and st[j]>='0' and st[j]<='9': j+=1
if j<len(st) and st[j]=="}": j+=1
tok = st[i:j]
elif tok=="\\text":
j+=2
while j<len(st) and st[j]>='a' and st[j]<='z': j+=1
j+=1
if j<len(st) and st[j]=="}": j+=1
tok = st[i:j]
i = j
if tok!=' ':
symb = symbol_table[tok]
if not symb: raise SyntaxError("Unknown operator")
yield symb()
symb = symbol_table["(end)"]
yield symb()
def expression(rbp=0):
global token
t = token
token = next()
left = t.nulld()
while rbp < token.lbp:
t = token
token = next()
left = t.leftd(left)
return left
def parse(str): # e.g., t = parse(r"(p\circ q)\lor \mathbf t")
global token, next
next = tokenize(str.replace("{\\sim}","\\sim ").replace("{-}","-")).__next__
token = next()
return expression()
def show(A, info=True): # display a (list of) formula(s)
st = A if type(A)==str else repr(A)
if info==True: display(Math(Macros+st))
########### end of parser #####################################
# code below assumes 'latex_st' contains the survey (SOASn.tex)
import re
def section(cl,st=latex_st): #find section of the class cl
cl = cl.replace('\\','\\\\').replace('$','\\$').replace('{','\\{')
return re.search(r"(\\hypertarget{"+cl+r"}{.*?)\\hypertarget",st,flags=re.DOTALL).group(1)
def chapter(n,st=latex_st): #find chapter number n
li = re.findall(r"(\\chapter{.*?%%endchapter)",st,flags=re.DOTALL)
return li[n-1]
def allclasses(st=latex_st): #return list of class names in st
li=re.findall(r"\\hypertarget{(.*?)}{",st,flags=re.DOTALL)
print("Number of classes:", len(li))
return li
def classname(cl): #find long class name of the class cl
return re.search(r"\\section{.*?:(.*?)}",section(cl),flags=re.DOTALL).group(1)
def sectiontitle(cl): #find full section title of the class cl
return re.search(r"\\section{(.*?)}",section(cl),flags=re.DOTALL).group(1)
def fulldefinition(cl): #find axioms in full definition of the class cl
ax=re.search(r"\\begin{fulldefinition}(.*?)\\end{fulldefinition}",section(cl),flags=re.DOTALL)
if ax==None: return 'none'
li=re.findall("\$(.*?)\$",ax.group(1),flags=re.DOTALL)
return [s for s in li if (s.find("=")>0 or s.find("\\le")>0) and s.find('\\langle')==-1 and s.find('\\mathbf')==-1]
def finitemembers(cl,info=False): #find fine spectrum in the class cl
fs = re.search(r"\\begin{finitemembers}(.*?)\\end{finitemembers}",section(cl),flags=re.DOTALL)
if info: print(fs)
li = re.findall("\$(.*?)\$",fs.group(1),flags=re.DOTALL)
if info: print(li)
ind = [s.split("_")[1].split("=")[0].strip() for s in li if s.find("_")>=0 and s.find("=")>=0]
ind = [int(s[1:-1]) if s[0]=="{" else int(s) for s in ind if s.find('n')==-1 and s.find('k')==-1]
for i in range(min(8,len(ind))):
if ind[i]!=i+1: raise Exception("index error at: ", i+1)
val = [s.split("_")[1].split("=")[1].strip() for s in li if s.find("_")>=0 and s.find("=")>=0]
return [int(s) for s in val if s!="" and s.find("n")==-1 and s.find("k")==-1]
def subclasses(cl,info=False): #find subclasses of the class cl
fs = re.search(r"\\begin{subclasses}(.*?)\\end{subclasses}",section(cl),flags=re.DOTALL)
if info: print(fs)
return re.findall(r"\\hyperlink{(.*?)}{",fs.group(1),flags=re.DOTALL)
def set_subclasses(cl,sbcli,longname,st=latex_st): #replace subclasses with those in sbcli
mo = section_mo(cl,st)
newsbc = r"\\begin{subclasses}\n"+"\n\n".join(r" \\hyperlink{"+x+"}{"+x+":"+(longname[x] if x in longname.keys() else "")+"}" for x in sbcli)+r"\n\\end{subclasses}"
newst = re.sub(r"\\begin{subclasses}.*?\\end{subclasses}",newsbc,mo.group(1),flags=re.DOTALL)
return st[:mo.start(1)]+newst+st[mo.end(1):]
def subclassposet():
m = re.findall(r"\\hypertarget{(.*?)}{",latex_st,flags=re.DOTALL)
return {s:subclasses(s) for s in m}
def superclasses(cl,info=False): #find subclasses of the class cl
fs = re.search(r"\\begin{superclasses}(.*?)\\end{superclasses}",section(cl),flags=re.DOTALL)
if info: print(fs)
return re.findall(r"\\hyperlink{(.*?)}{",fs.group(1),flags=re.DOTALL)
def set_superclasses(cl,spcli,longname,st=latex_st): #replace superclasses with those in spcli
mo = section_mo(cl,st)
newspc = r"\\begin{superclasses}\n"+"\n\n".join(r" \\hyperlink{"+x+"}{"+x+":"+(longname[x] if x in longname.keys() else "")+"}" for x in spcli)+r"\n\\end{superclasses}"
newst = re.sub(r"\\begin{superclasses}.*?\\end{superclasses}",newspc,mo.group(1),flags=re.DOTALL)
return st[:mo.start(1)]+newst+st[mo.end(1):]
def superclassposet():
m = re.findall(r"\\hypertarget{(.*?)}{",latex_st,flags=re.DOTALL)
return {s:superclasses(s) for s in m}
def finitememberslatex(li): #convert list of numbers to fine spectrum in LaTeX
return ",\n".join("$f_"+(str(i) if i<10 else "{"+str(i)+"}")+" = "+str(li[i-1])+"$" for i in range(1,len(li)+1))+"\n"
def p9out(A): #output formula A in Prover9 format
if A.a==[]: return p9sym[A.id]
if A.id[:7] in ["\\forall","\\exists"]:
return p9sym[A.id[:7]]+" "+A.id[8:]+"("+p9out(A.a[0])+")"
if len(A.a)==1:
#if symbol_table[p9sym[A.id]].lbp!=12:
return p9sym[A.id]+"("+p9out(A.a[0])+")"
#return "("+p9out(A.a[0])+")"+p9sym[A.id]
return "("+p9out(A.a[0])+p9sym[A.id]+p9out(A.a[1])+")"
po=["x<=x","x<=y & y<=x -> x=y","x<=y & y<=z -> x<=z"]
msl=["(x^y)^z=x^(y^z)","x^y=y^x","x^x=x","x^y=x<->x<=y"]
jsl=["(x v y)v z=x v(y v z)","x v y=y v x","x v x=x","x v y=y<->x<=y"]
lat=msl+jsl+["x v(x^y)=x","x^(x v y)=x"]
dlat=lat+["x^(y v z)=(x^y)v(x^z)"]
to=lat+["x^y=x | x^y=y"] #["x<=y | y<=x"]
ba=dlat+["x'v x=t","x'^x=b"]
uo=[]
axioms=[po,jsl,msl,lat,dlat,to,ba,uo]
cli = [allclasses(chapter(i)) for i in range(2,10)]
fam = {x:cx for cx in range(8) for x in cli[cx]}
def fd(cl,info=True,f=fam,new_ax=None):
try:
ax = [p9out(parse(e)) for e in fulldefinition(cl)]
except:
print("############### Error, skipping", cl)
return []
ax = [(x[1:-1] if x[0]=='(' and x[-1]==')' else x) for x in ax]
ch_axioms = axioms[fam[cl]] if new_ax==None else new_ax
if info:
print(ch_axioms+ax)
return ch_axioms+ax
def finespectrum(cl,n,info=True,f=fam,new_ax=None):
# call Prover9 on the translated full definition of cl and find the fine spectrum up to n
if info: print(cl)
ax = fd(cl,new_ax)
if ax==[]: return []
t = time.time()
a = [[1]]+[prover9(ch_axioms+ax,[],10000,10000,k,options=opts) for k in range(2,n+1)]
if info: print("Time: {:.2f}".format(time.time()-t), "sec")
return [len(x) for x in a]
def section_mo(cl,st=latex_st):
cl = cl.replace('\\','\\\\').replace('$','\\$').replace('{','\\{')
return re.search(r"(\\hypertarget{"+cl+r"}{.*?)\\hypertarget",st,flags=re.DOTALL)
def set_finitemembers(cl,li,st=latex_st): #read cl string from latex_st, replace finitemembers entry and return new latex_st
mo = section_mo(cl,st)
newst = re.sub(r"\\begin{finitemembers}.*?\\end{finitemembers}",r"\\begin{finitemembers}\n"+finitememberslatex(li)+r"\\end{finitemembers}",mo.group(1),flags=re.DOTALL)
return st[:mo.start(1)]+newst+st[mo.end(1):]
def set_classname(cl,new_name,st=latex_st): #replace long class name of the class cl
mo = section_mo(cl,st)
newst = re.sub(r"\\section{.*?}",r"\\section{"+new_name+r"}",mo.group(1),flags=re.DOTALL)
return st[:mo.start(1)]+newst+st[mo.end(1):]
ch=jsl
def compareandupdatefs(chax,cl):
li = finitemembers(cl)
sli = [x for x in li if x <= 1000]
n = min(max(len(sli),3),6)
fs = finespectrum(chax,cl,n,True)
print(cl,li)
return li[:n]==fs or fs==[], fs
def sectionnames(st):
return re.findall(r"\\hypertarget{(.*?)}{",st,flags=re.DOTALL)
#axioms=[[],["x<=y->x=y"],po,jsl,msl,lat,dlat,to,ba,uo]
def checkfs(li,info=False):
global latex_st
m=[[] for x in range(10)]
for ch in li:
st1=chapter(ch,latex_st)
m[ch]=sectionnames(st1)
print([len(x) for x in m])
for ch in li:
print(ch,axioms[ch])
for cl in m[ch]:
if fulldefinition(cl)!="none":
fl = compareandupdatefs(axioms[ch-2],cl)
if not fl[0]:
print("***********",fl)
latex_st = set_finitemembers(cl,fl[1],latex_st)
def allclassposets(st=latex_st): #return list of tikz diagrams
return re.findall(r"(\\begin{tikzpicture}\[xscale=1.*?\\end{tikzpicture}\n)",latex_st,flags=re.DOTALL)
def allnodes(st): #return list of node names in st
li=re.findall(r"\\node\((.*?)\)",st,flags=re.DOTALL)
print("Number of nodes:", len(li))
return li
def lowercovers(nd,st): #return list of lowercovers of node name nd in st
edges=re.search(r"\\draw\("+nd+"\)(.*?);",st,flags=re.DOTALL)
return re.findall(r"edge.*?\((.*?)\)",edges.group(1),flags=re.DOTALL) if edges!=None else []
def lc2uc(lc):
uc={x:[] for x in lc}
for x in lc:
for y in lc[x]:
if y in uc: uc[y].append(x)
return uc
def smallmembers(cl): #return list of small algebras of the class cl
sm=re.search(r"\\begin{smallmembers}(.*?)\\end{smallmembers}",section(cl),flags=re.DOTALL)
if sm==None: return 'none'
li=re.findall(r"\\begin{tikzpicture}(.*?)\\end{tikzpicture}",sm.group(1),flags=re.DOTALL)
pl=[list(reversed(re.findall(r"\\node\((.*?);", s,flags=re.DOTALL))) for s in li]
uc=[{int(s[:s.index(")")]):[int(y) for y in re.findall(r"edge.*?\((.*?)\)",s)] for s in x} for x in pl]
xy=[{int(s[:s.index(")")]):(re.search(r"\((.*?),",s).group(1), re.search(r",(.*?)\)",s).group(1)) for s in x} for x in pl]
xy=[{i:tuple((int(z) if z.find(".")==-1 else float(z)) for z in xy[n][i]) for i in xy[n]} for n in range(len(pl))]
c=[[int(s[:s.index(")")]) for s in x if s.find("label=")!=-1] for x in pl]
return [({'uc':uc[n], 'xy':xy[n]} if c[n]==[] else {'uc':uc[n], 'xy':xy[n], 'c':c[n][0]}) for n in range(len(pl))]
def uc2p9(uc):
return [(f"{i}<={j}" if j in uc[i] else f"-({i}<={j})") for i in uc for j in uc]
# -
REqMon=['((x*y)*z)=(x*(y*z))', '(x*1)=x', '(1*x)=x', '((x*y)=z) <-> (y=(x\\ z))']
a=p9(REqMon+["x'=x\ 1"],["x*x'=1"],0,1000)
a
len(latex_st)
fs=finespectrum('BInFL',8)
fs
print(finitememberslatex(fs))
Fld=['((x+y)+z)=(x+(y+z))', '(x+0)=x', '(-(x)+x)=0', '(x+y)=(y+x)', '((x*y)*z)=(x*(y*z))', '(x*1)=x', '(x*y)=(y*x)', '(x*(y+z))=((x*y)+(x*z))', "(x!=0) -> (x*x'=1)","0'=0"]
p9(Fld,[],1000,1000,[6])
fs=finespectrum('InPocrim',8)
InPocrim=['x<=x', 'x<=y & y<=x -> x=y', 'x<=y & y<=z -> x<=z', '-(-(x))=x', '((x*y)<=z) <-> (y<=-((-(z)*x)))', '((x*y)*z)=(x*(y*z))', '(x*y)=(y*x)', '(x*1)=x', 'x<=1']
a=p9(InPocrim,[],1000,1000,[8])
m4diag(a[8])
cpo=allclassposets()
print(len(cpo))
tkz="".join(cpo)
nli=allnodes(tkz)
print(len(nli),len(set(nli)))
appdx=readfile('SPoAappendix.tex')
len(appdx)
#def rep(m): return m.group(1)+"&"+m.group(2)+"\\\\"
#st1=re.sub("\^(\[\[.*?)\|(.*?)\|",rep,st1,flags=re.DOTALL)
def rep(m): return '\\\\'+m.group(1)+'\n\\hyperlink{'+m.group(2)+'}{'+m.group(2)+'}&'
appdx1=re.sub(r'\\\\(.*?)\n(.*?)&',rep,appdx,flags=re.DOTALL)
#print(appdx1[:30])
writefile('SPoAappendix.tex',appdx1)
cls = allclasses()
longname={x:classname(x) for x in cls}
fullname={x:sectiontitle(x) for x in cls}
for x in cls: #x = 'PoUn'
latex_st = set_subclasses(x, sorted(lcp[x]), longname, latex_st)
latex_st = set_superclasses(x, sorted(ucp[x]), longname, latex_st)
latex_st = set_classname(x, fullname[x].replace(':',r'\\index{'+x+'}:'), latex_st)
latex_st = re.sub(r'%\\abbreviation{.*?}\n\n','',latex_st,flags=re.DOTALL)
len(latex_st)
longname['RtHp']
print(section('PoUn',latex_st))
#[longname[x] for x in cls[:20]]
lcp['Pos']
ucp=lc2uc(scp)
lcp=lc2uc(ucp)
[set(lcp[x])-set(scp[x]) for x in lcp if set(lcp[x])-set(scp[x])!=set()]
# +
#[len(set(scp[x])-set(lcp[x])) for x in lcp]
# -
#subclassposet
scp={'Pos': ['pPos', 'PoMag', 'PoImpA', 'PoNUn', 'PoUn', 'Jslat', 'Mslat', 'Set'],
'pPos': ['PoMon', 'pJslat', 'pMslat', 'pSet'],
'PoUn': ['GalPos', 'RPoUn', 'JUn', 'MUn', 'Unar'],
'PoNUn': ['GalPos', 'JNUn', 'MNUn'],
'PoMag': ['LrPoMag', 'PoSgrp', 'JMag', 'MMag'],
'PoSgrp': ['CPoSgrp', 'IdPoSgrp', 'LrPoSgrp', 'PoMon', 'JSgrp', 'MSgrp'],
'PoMon': ['CPoMon', 'IdPoMon', 'IPoMon', 'LrPoMon', 'JMon', 'MMon'],
'IPoMon': ['CIPoMon', 'Polrim', 'IJMon', 'IMMon'],
'IdPoSgrp': ['CIdPoSgrp', 'IdLrPoSgrp', 'IdPoMon', 'IdJSgrp', 'IdMSgrp'],
'IdPoMon': ['CIdPoMon', 'IdLrPoMon', 'IdJMon', 'IdMMon'],
'PoImpA': ['DivPos', 'LrPoMag', 'JImpA', 'MImpA'],
'LrPoMag': ['RPoMag', 'LrPoSgrp', 'LrJMag', 'LrMMag'],
'LrPoSgrp': ['RPoMon', 'LrPoMon', 'IdLrPoSgrp', 'RPoSgrp', 'LrJSgrp', 'LrMSgrp'],
'LrPoMon': ['RPoMon', 'IdLrPoMon', 'Polrim', 'LrJMon', 'LrMMon'],
'Polrim': ['Porim', 'ILrJMon', 'ILrMMon'],
'IdLrPoSgrp': ['IdRPoSgrp', 'IdLrPoMon', 'IdLrJSgrp', 'IdLrMSgrp'],
'IdLrPoMon': ['IdRPoMon', 'IdLrJMon', 'IdLrMMon'],
'RPoUn': ['InPoMon', 'RJUn', 'RMUn'],
'DivPos': ['CDivPos', 'RPoMag', 'DivJslat', 'DivMslat'],
'RPoMag': ['CRPoMag', 'RPoSgrp', 'InPoMag', 'RJMag', 'RMMag'],
'RPoSgrp': ['CRPoSgrp', 'RPoMon', 'IdRPoSgrp', 'InPoSgrp', 'RJSgrp', 'RMSgrp'],
'RPoMon': ['CRPoMon', 'InPoMon', 'IdRPoMon', 'Porim', 'RJMon', 'RMMon'],
'Porim': ['InPorim', 'Pocrim', 'IRJMon', 'IRMMon'],
'IdRPoSgrp': ['CIdRPoSgrp', 'IdRPoMon', 'IdInPoSgrp', 'IdRJSgrp', 'IdRMSgrp'],
'IdRPoMon': ['CIdRPoMon', 'IdInPoMon', 'IdRJMon', 'IdRMMon'],
'GalPos': ['InPos', 'GalJslat', 'GalMslat'],
'InPos': ['InPoMag', 'InLat'],
'InPoMag': ['CyInPoMag', 'InPoSgrp', 'InLMag'],
'InPoSgrp': ['CyInPoSgrp', 'InPoMon', 'IdInPoSgrp', 'InLSgrp'],
'InPoMon': ['CyInPoMon', 'IInPoMon', 'PoGrp', 'IdInPoMon', 'InPorim', 'InLMon'],
'InPorim': ['CyInPorim', 'IInLMon'],
'CyInPoMag': ['CInPoMag', 'CyInPoSgrp', 'CyInLMag'],
'CyInPoSgrp': ['CInPoSgrp', 'CyInPoMon', 'CyIdInPoSgrp', 'CyInLSgrp'],
'CyInPoMon': ['CInPoMon', 'CyInPorim', 'CyIdInPoMon', 'PoGrp', 'CyInLMon'],
'CyInPorim': ['InPocrim', 'CyIInLMon'],
'PoGrp': ['AbPoGrp', 'LGrp'],
'CPoSgrp': ['CIdPoSgrp', 'CRPoSgrp', 'CPoMon', 'CJSgrp', 'CMSgrp'],
'CPoMon': ['CIdPoMon', 'CIPoMon', 'CRPoMon', 'CJMon', 'CMMon'],
'CIPoMon': ['Pocrim', 'CIJMon', 'CIMMon'],
'CIdPoSgrp': ['CIdRPoSgrp', 'CIdPoMon', 'CIdJSgrp', 'CIdMSgrp'],
'CIdPoMon': ['CIdRPoMon', 'CIdJMon', 'CIdMMon'],
'CDivPos': ['CRPoMag', 'BCK', 'CDivJslat', 'CDivMslat'],
'BCK': ['Pocrim', 'HilA', 'BCKJslat', 'BCKMslat'],
'CRPoMag': ['CRPoSgrp', 'CInPoMag', 'CRJMag', 'CRMMag'],
'HilA': ['TarA'],
'TarA': [],
'CRPoSgrp': ['CInPoSgrp', 'CRPoMon', 'CIdRPoSgrp', 'CRJSgrp', 'CRMSgrp'],
'CRPoMon': ['CInPoMon', 'Pocrim', 'CIdRPoMon', 'CRJMon', 'CRMMon'],
'Pocrim': ['InPocrim', 'CIRJMon', 'CIRMMon'],
'CIdRPoSgrp': ['CIdRPoMon', 'CIdInPoSgrp', 'CIdRJSgrp', 'CIdRMSgrp'],
'CIdRPoMon': ['IdRPocrim', 'CIdInPoMon', 'CIdRJMon', 'CIdRMMon'],
'CInPoMag': ['CInPoSgrp', 'CInLMag'],
'CInPoSgrp': ['CInPoMon', 'CIdInPoSgrp', 'CInLSgrp'],
'CInPoMon': ['InPocrim', 'AbPoGrp', 'CIdInPoMon', 'CInLMon'],
'InPocrim': ['CIInLMon'],
'AbPoGrp': ['AbLGrp'],
'Jslat': ['Slat$_1$', 'Slat$_0$', 'pJslat', 'JUn', 'JNUn', 'JMag', 'JImpA', 'Lat'],
'pJslat': ['JMon', 'ubJslat', 'lbJslat', 'pLat'],
'lbJslat': ['IdSrng$_0$', 'bLat'],
'ubJslat': ['IJMon', 'bLat'],
'JUn': ['GalJslat', 'RJUn', 'LUn'],
'JNUn': ['GalJslat', 'LNUn'],
'GalJslat': ['GalLat'],
'JMag': ['JSgrp', 'LrJMag', 'LMag'],
'JSgrp': ['CJSgrp', 'JMon', 'IdJSgrp', 'LrJSgrp', 'LSgrp'],
'JMon': ['CJMon', 'LrJMon', 'IdJMon', 'IJMon', 'IdSrng$_{01}$', 'LMon'],
'IdSrng$_0$': ['IdSrng$_{01}$'],
'IdSrng$_{01}$': ['KA'],
'KA': ['ActA', 'KLat'],
'IJMon': ['CIJMon', 'ILrJMon', 'ILMon'],
'IdJSgrp': ['CIdJSgrp', 'IdJMon', 'IdLrJSgrp', 'IdLSgrp'],
'IdJMon': ['CIdJMon', 'IdLrJMon', 'IdLMon'],
'JImpA': ['LrJMag', 'DivJslat', 'LImpA'],
'LrJMag': ['RJMag', 'LrJSgrp', 'LrLMag'],
'LrJSgrp': ['RJMon', 'LrJMon', 'IdLrJSgrp', 'RJSgrp', 'LrLSgrp'],
'LrJMon': ['RJMon', 'ILrJMon', 'IdLrJMon', 'LrLMon'],
'ILrJMon': ['IRJMon', 'ILrLMon'],
'IdLrJSgrp': ['IdRJSgrp', 'IdLrJMon', 'IdLrLSgrp'],
'IdLrJMon': ['IdRJMon', 'IdLrLMon'],
'RJUn': ['InJMon', 'RLUn'],
'DivJslat': ['CDivJslat', 'RJMag', 'DivLat'],
'RJMag': ['CRJMag', 'RJSgrp', 'RLMag'],
'RJSgrp': ['CRJSgrp', 'RJMon', 'IdRJSgrp', 'RLSgrp'],
'RJMon': ['CRJMon', 'IdRJMon', 'IRJMon', 'RLMon'],
'IRJMon': ['CIRJMon', 'IRLMon'],
'IdRJSgrp': ['CIdRJSgrp', 'IdRJMon', 'IdRLSgrp'],
'IdRJMon': ['CIdRJMon', 'IdRLMon'],
'CJSgrp': ['CJMon', 'CRJSgrp', 'CIdJSgrp', 'CLSgrp'],
'CJMon': ['CIJMon', 'CRJMon', 'CIdJMon', 'CLMon'],
'CIJMon': ['CIRJMon', 'CILMon'],
'CIdJSgrp': ['CIdJMon', 'CIdRJSgrp', 'CIdLSgrp'],
'CIdJMon': ['CIdRJMon', 'CIdLMon'],
'CDivJslat': ['CRJMag', 'BCKJslat', 'CDivLat'],
'BCKJslat': ['BCKLat'],
'CRJMag': ['CRJSgrp', 'CInJMag', 'CRLMag'],
'CRJSgrp': ['CInJSgrp', 'CRJMon', 'CIdRJSgrp', 'CRLSgrp'],
'CRJMon': ['CIRJMon', 'CIdRJMon', 'CRLMon'],
'CIRJMon': ['IdCIRJMon', 'CIRLMon'],
'CIdRJSgrp': ['CIdRJMon', 'CIdRLSgrp'],
'CIdRJMon': ['IdRCIRJMon', 'CIdRLMon'],
'Mslat': ['pMslat', 'MMag', 'MImpA', 'MUn', 'MNUn', 'Lat'],
'pMslat': ['MMon', 'pLat'],
'MUn': ['GalMslat', 'RMUn', 'LUn'],
'MNUn': ['GalMslat', 'LNUn'],
'GalMslat': ['GalLat'],
'MMag': ['MSgrp', 'LrMMag', 'LMag'],
'MSgrp': ['CMSgrp', 'IdMSgrp', 'LrMSgrp', 'LSgrp', 'MMon'],
'MMon': ['CMMon', 'IdMMon', 'IMMon', 'LrMMon', 'LMon'],
'IMMon': ['CIMMon', 'ILrMMon', 'ILMon'],
'IdMSgrp': ['CIdMSgrp', 'IdLrMSgrp', 'IdMMon', 'IdLSgrp'],
'IdMMon': ['CIdMMon', 'IdLrMMon', 'IdLMon'],
'MImpA': ['DivMslat', 'LrMMag', 'LImpA'],
'LrMMag': ['RMMag', 'LrMSgrp', 'LrLMag'],
'LrMSgrp': ['IdLrMSgrp', 'LrMMon', 'RMSgrp', 'LrLSgrp'],
'LrMMon': ['IdLrMMon', 'ILrMMon', 'RMMon', 'LrLMon'],
'ILrMMon': ['IRMMon', 'ILrLMon', 'RtHp'],
'RtHp': ['Hp'],
'IdLrMSgrp': ['IdRMSgrp', 'IdLrMMon', 'IdLrLSgrp'],
'IdLrMMon': ['IdRMMon', 'IdLrLMon'],
'RMUn': ['InMMon', 'RLUn'],
'DivMslat': ['CDivMslat', 'RMMag', 'DivLat'],
'RMMag': ['CRMMag', 'RMSgrp', 'RLMag'],
'RMSgrp': ['CRMSgrp', 'IdRMSgrp', 'RMMon', 'RLSgrp'],
'RMMon': ['IdRMMon', 'IRMMon', 'CRMMon', 'RLMon'],
'IRMMon': ['CIRMMon', 'IRLMon'],
'IdRMSgrp': ['CIdRMSgrp', 'IdRMMon', 'IdRLSgrp'],
'IdRMMon': ['CIdRMMon', 'IdRLMon'],
'CMSgrp': ['CMMon', 'CIdMSgrp', 'CRMSgrp', 'CRLSgrp', 'CLSgrp'],
'CMMon': ['CIMMon', 'CRMMon', 'CIdMMon', 'CLMon'],
'CIMMon': ['CIRMMon', 'CILMon'],
'CIdMSgrp': ['CIdMMon', 'CIdRMSgrp', 'CIdLSgrp'],
'CIdMMon': ['CIdRMMon', 'CIdLMon'],
'CDivMslat': ['CRMMag', 'BCKMslat', 'CDivLat'],
'BCKMslat': ['BCKLat'],
'CRMMag': ['CRMSgrp', 'CInMMag', 'CRLMag'],
'CRMSgrp': ['CIdRMSgrp', 'CRMMon', 'CRLSgrp'],
'CRMMon': ['CIdRMMon', 'CIRMMon', 'CRLMon'],
'CIRMMon': ['CIRL', 'CIRLMon'],
'Hp': ['WaHp', 'BrSlat', 'CGBL'],
'CIdRMSgrp': ['CIdRMMon', 'CIdRLSgrp'],
'CIdRMMon': ['IdRCIRMMon', 'CIdRLMon'],
'BrSlat': ['BrA'],
'Lat': ['pLat', 'LMag', 'LImpA', 'LNUn', 'LUn', 'ModLat', 'JsdLat','MsdLat', 'OLat'],
'pLat': ['LMon', 'pDLat'],
'bLat': ['bModLat', 'CplmLat', 'bDLat'],
'LUn': ['RLUn', 'DLUn'],
'LNUn': ['GalLat', 'DLNUn'],
'LMag': ['LrLMag', 'LSgrp', 'DLMag', 'MultLat'],
'LSgrp': ['CLSgrp', 'LrLSgrp', 'LMon', 'IdLSgrp', 'DLSgrp'],
'LMon': ['CLMon', 'ILMon', 'LrLMon', 'IdLMon', 'DLMon'],
'KLat': ['ActLat'],
'ActLat': ['TrivA'],
'ModLat': ['DLat', 'ComModLat'],
'MultLat': ['LSgrp'],
'ILMon': ['CILMon', 'ILrLMon', 'DILMon'],
'IdLSgrp': ['CIdLSgrp', 'IdLrLSgrp', 'IdLMon', 'DIdLSgrp'],
'IdLMon': ['CIdLMon', 'IdLrLMon', 'DIdLMon'],
'LImpA': ['DivLat', 'LrLMag', 'DLImpA'],
'LrLMag': ['RLMag', 'LrLSgrp', 'DLrLMag'],
'LrLSgrp': ['R$ell$Sgrp', 'LrLMon', 'IdLrLSgrp', 'RLSgrp', 'DLrLSgrp'],
'LrLMon': ['ILrLMon', 'RL', 'IdLrLMon', 'DLrLMon'],
'ILrLMon': ['IRL', 'DILrLMon'],
'IdLrLSgrp': ['IdRLSgrp', 'IdLrLMon', 'DIdLrLSgrp'],
'IdLrLMon': ['IdRL', 'DIdLrLMon'],
'RLUn': ['InLMon', 'DRLUn'],
'DivLat': ['CDivLat', 'RLMag', 'DDivLat'],
'RLMag': ['CRLMag', 'InLMag', 'RLSgrp', 'DRLMag'],
'RLSgrp': ['CRLSgrp', 'InLSgrp', 'RL', 'IdRLSgrp', 'DRLSgrp'],
'RL': ['bRL', 'CRL', 'IRL', 'IdRL', 'DRL', 'ActLat', 'CanRL', 'FL'],
'bRL': ['ILLA', 'MALLA'],
'IRL': ['CIRL', 'IInFL', 'DIRL'],
'IdRLSgrp': ['CIdRLSgrp', 'IdInLSgrp', 'IdRL', 'DIdRLSgrp'],
'IdRL': ['CIdRL', 'IdInFL', 'DIdRL'],
'FL': ['bRL', 'FL$_e$', 'FL$_w$', 'FL$_c$', 'DFL', 'InFL'],
'FL$_c$': ['FL$_{ec}$', 'DFL$_c$'],
'FL$_e$': ['FL$_{ec}$', 'FL$_{ew}$', 'DFL', 'DFL$_e$', 'ILLA'],
'FL$_w$': ['DFL$_w$'],
'FL$_{ec}$': ['DFL$_{ec}$'],
'FL$_{ew}$': ['DFL$_{ew}$'],
'GalLat': ['InLat', 'DGalLat'],
'InLat': ['InLMag', 'DInLat', 'Bilat'],
'InLMag': ['CyInLMag', 'InLSgrp', 'DInLMag'],
'InLSgrp': ['CyInLSgrp', 'InFL', 'IdInLSgrp', 'DInLSgrp'],
'InFL': ['CyInFL', 'IInFL', 'IdInFL', 'DInFL'],
'IInFL': ['CyIInFL', 'DIInFL'],
'CyInLMag': ['CInLMag', 'CyInLSgrp', 'CyDInLMag'],
'CyInLSgrp': ['CInLSgrp', 'CyInFL', 'CyIdInLSgrp', 'CyDInLSgrp'],
'CyInFL': ['CInFL', 'CyIInFL', 'CyIdInFL', 'CyDInFL'],
'CyIInFL': ['CIInFL', 'CyDIInFL'],
'CLSgrp': ['CRLSgrp', 'CLMon', 'CIdLSgrp', 'CDLSgrp'],
'CLMon': ['CILMon', 'CRL', 'CIdLMon', 'CDLMon'],
'CILMon': ['CIRL', 'CDILMon'],
'CIdLSgrp': ['CIdRLSgrp', 'CIdLMon', 'CDIdLSgrp'],
'CIdLMon': ['CIdRL', 'CDIdLMon'],
'CDivLat': ['CRLMag', 'CDDivLat'],
'BCKLat': ['HA'],
'CRLMag': ['CInLMag', 'CRLSgrp', 'CDRLMag'],
'CRLSgrp': ['CInLSgrp', 'CRL', 'CIdRLSgrp', 'CDRLSgrp'],
'CRL': ['CIRL', 'CInFL', 'CIdRL', 'CDRL'],
'CIRL': ['CIInFL', 'CDIRL'],
'CIdRLSgrp': ['CIdInLSgrp', 'CIdRL', 'CDIdRLSgrp'],
'CIdRL': ['CIdInFL', 'CDIdRL'],
'CIdInFL': ['CDIdInFL'],
'CInLMag': ['CInLSgrp', 'CDInLMag'],
'CInLSgrp': ['CInFL', 'CIdInLSgrp', 'CDInLSgrp'],
'CInFL': ['CIInFL', 'CIdInFL', 'CDInFL', 'MALLA'],
'CIInFL': ['CDIInFL'],
'JsdLat': ['SdLat'],
'MsdLat': ['SdLat'],
'SdLat': ['NdLat'],
'NdLat': ['ADLat'],
'ADLat': ['DLat'],
'CplmLat': ['CplmModLat'],
'OLat': ['OModLat'],
'OModLat': ['ModOLat'],
'ModOLat': ['BA'],
'SkLat': ['Lat', 'RecBnd'],
'Bilat': ['TrivA'], # 'DBilat'],
'CanRL': ['TrivA'],
'CplmModLat': ['BA'],
'FRng': ['TrivA'],
'ILLA': ['LLA'],
'LLA': ['TrivA'],
'MALLA': ['LLA'],
'DLat': ['bDLat', 'pDLat', 'DLUn', 'DLNUn', 'DLMag', 'DLImpA', 'ToLat', 'BA', 'pcDLat'],
'pDLat': ['DLMon', 'bDLat', 'pToLat', 'pBA'],
'bDLat': ['BA', 'bToLat', 'DpAlg', 'DdpAlg', 'BoolLat', 'OckA'],
'DLUn': ['DGalLat', 'DRLUn', 'ToUn', 'BUn'],
'DLNUn': ['DGalLat', 'ToNUn', 'BNUn', 'OckA'],
'pcDLat': ['DpAlg'],
'OckA': ['DmA'],
'DmA': ['KLA', 'LA$_n$'],
'DmMon': [],
'DpAlg': ['DDbpAlg', 'StAlg'],
'DdpAlg': ['DDblpAlg'],
'DDblpAlg': ['DblStAlg'],
'StAlg': ['DblStAlg'],
'DblStAlg': ['BA'],
'DLMag': ['DLSgrp', 'DLrLMag', 'ToMag', 'BMag'],
'DLSgrp': ['CDLSgrp', 'DIdLSgrp', 'DLMon', 'DLrLSgrp', 'ToSgrp', 'BSgrp'],
'DLMon': ['CDLMon', 'DIdLMon', 'DILMon', 'DLrLMon', 'ToMon', 'BMon'],
'DILMon': ['ALat', 'CDILMon', 'DILrLMon', 'IToMon', 'BIMon'],
'DIdLSgrp': ['CDIdLSgrp', 'DIdLMon', 'DIdLrLSgrp', 'IdToSgrp', 'BIdSgrp'],
'DIdLMon': ['CDIdLMon', 'DIdLrLMon', 'IdToMon', 'BIdMon'],
'DLImpA': ['CDLSgrp', 'LSgrp', 'DLrLMag', 'DDivLat', 'ToImpA', 'BImpA', 'ImpLat'],
'DLrLMag': ['DRLMag', 'DLrLSgrp', 'LrToMag', 'BLrMag'],
'DLrLSgrp': ['DR$ell$Sgrp', 'DLrLMon', 'DIdLrLSgrp', 'DRLSgrp', 'LrToSgrp', 'BLrSgrp'],
'DLrLMon': ['DRL', 'DILrLMon', 'DIdLrLMon', 'LrToMon', 'BILrMon'],
'DILrLMon': ['DIRL', 'ILrToMon', 'BIdLrSgrp'],
'DIdLrLSgrp': ['DIdRLSgrp', 'DIdLrLMon', 'IdLrToSgrp', 'BIdLrMon'],
'DIdLrLMon': ['DIdRL', 'IdLrToMon', 'BRUn'],
'DRLUn': ['DInLMon', 'RToUn', 'BDivLat'],
'DDivLat': ['DRLMag', 'CDDivLat', 'ToDivLat', 'BRMag'],
'DRLMag': ['CDRLMag', 'DRLSgrp', 'DInLMag', 'RToMag', 'BRSgrp'],
'DRLSgrp': ['CDRLSgrp', 'DRL', 'DIdRLSgrp', 'DInLSgrp', 'RToSgrp', 'BRL'],
'DRL': ['CDRL', 'DFL$_e$', 'DIdRL', 'DIRL', 'DInFL', 'RToMon', 'BIRL', 'GBL'],
'DIRL': ['CDIRL', 'DIInFL', 'IRToMon', 'BIdRSgrp'],
'DIdRLSgrp': ['CDIdRLSgrp', 'DIdRL', 'DIdInLSgrp', 'IdRToSgrp', 'BIdRL'],
'DIdRL': ['CDIdRL', 'DIdInFL', 'IdRToMon', 'BGalA'],
'DGalLat': ['DInLat', 'GalToLat', 'BGalLat'],
'DInLat': ['DInLMag', 'InToLat', 'BInMag'],
'DInLMag': ['CyDInLMag', 'DInLSgrp', 'InToMag', 'BInSgrp'],
'DInLSgrp': ['CyDInLSgrp', 'DInFL', 'DIdInLSgrp', 'InToSgrp', 'BInFL'],
'DInFL': ['CyDInFL', 'DIInFL', 'DLGrp', 'DIdInFL', 'BIInFL'],
'DIInFL': ['CyDIInFL', 'BCyInMag', 'psMV'],
'CyDInLMag': ['CDInLMag', 'CyDInLSgrp', 'CyInToMag', 'BCyInSgrp'],
'CyDInLSgrp': ['CDInLSgrp', 'CyDInFL', 'CyDIdInLSgrp',
'CyInToSgrp', 'BCyInFL'],
'CyDInFL': ['CDInFL', 'CyDIInFL', 'CyDIdInFL', 'LGrp', 'BCyIInFL'],
'CyDIInFL': ['CDIInFL'],
'LGrp': ['NVLGrp'],
'RepLGrp': ['AbLGrp', 'ToGrp'],
'CDLSgrp': ['CDIdLSgrp', 'CDLMon', 'CDRLSgrp', 'CToSgrp', 'BCMon'],
'CDLMon': ['ALat', 'CDIdLMon', 'CDILMon', 'CDRL', 'CToMon', 'BCIMon'],
'CDILMon': ['ALat', 'CDIRL', 'CIToMon', 'BCIdSgrp'],
'CDIdLSgrp': ['CDIdLMon', 'CDIdRLSgrp', 'CIdToSgrp', 'BCIdMon'],
'CDIdLMon': ['CDIdRL', 'CIdToMon', 'BCDivLat'],
'CDDivLat': ['CDRLMag', 'CivToLat', 'BCRMag'],
'CDRLMag': ['CDRLSgrp', 'CDInLMag', 'CRToMag'],
'CDRLSgrp': ['CDInLSgrp', 'CDRL', 'CDIdRLSgrp', 'CRSlSgrp'],
'CDRL': ['CDIRL', 'CDIdRL', 'CDInFL', 'CRSlMon', 'DunnMon'],
'CDIRL': ['DICDIRL', 'CDIInFL', 'CIRSlMon'],
'CDIdRLSgrp': ['CDIdRL', 'CDIdInLSgrp', 'CIdRSlSgrp'],
'CDIdRL': ['DIdRCIRL', 'CDIdInFL', 'CIdRSlMon', 'BCInMag'],
'CDInLMag': ['CDInLSgrp', 'CInToMag'],
'CDInLSgrp': ['CDInFL', 'CDIdInLSgrp', 'CInSlSgrp'],
'CDInFL': ['CDIInFL', 'AbLGrp', 'CDIdInFL', 'CInSlMon', 'DmMon'],
'CDIInFL': ['CIInSlMon'],
'AbLGrp': ['AbToGrp', 'LRng'],
'GBL': ['BLA', 'GBLChn', 'GMV'],
'GMV': ['MV', 'GMVChn'],
'psMV': ['MV'],
'WaHp': ['MV'],
'BrA': ['HA', 'GBA'],
'GBA': ['BA'],
'BoolLat': ['BA'],
'CRSlSgrp': ['CInSlSgrp', 'CIdRSlSgrp', 'CRSlMon', 'CRToSgrp', 'BCRMon'],
'CRSlMon': ['CIdRSlMon', 'CIRSlMon', 'CInSlMon', 'CRToMon', 'BCIRMon'],
'CIRSlMon': ['IMTL', 'CIRToMon', 'CIInSlMon', 'MTLA'],
'MTLA': ['BLA', 'MTLChn'],
'BLA': ['MV', 'HA', 'BLChn'],
'MV': ['BA', 'MVChn'],
'HA': ['GödA'],
'GödA': ['BA', 'GödChn', 'BCIdRSgrp'],
'CIdRSlSgrp': ['CIdInSlSgrp', 'CIdRSlMon', 'CIdRToSgrp', 'BCIdRMon'],
'CIdRSlMon': ['CIdInSlMon', 'CIdRToMon', 'BCInSgrp'],
'CInSlSgrp': ['CIdInSlSgrp', 'CInSlMon', 'CInToSgrp', 'BCInMon'],
'CInSlMon': ['CIdInSlMon', 'IMTL', 'AbLGrp', 'CInToMon', 'CIInSlMon'],
'DunnMon': ['CDIdRL', 'BCIInRMon', 'DmMon'],
'IMTL': ['IMTLChn', 'CIInRToMon'],
'ImpLat': ['GodA', 'MV', 'LGrp'],
'KLA': ['BA'],
'NVLGrp': ['RepLGrp'],
'LA$_n$': ['BA'],
'LRng': ['CLRng', 'ToRng', 'FRng'],
'CLRng': ['CToRng'],
'ToLat': ['pDLat', 'ToMag', 'ToImpA', 'ToNUn', 'ToUn', 'pToLat'],
'pToLat': ['ToMon'],
'ToMag': ['ToSgrp', 'LrToMag'],
'ToSgrp': ['CToSgrp', 'IdToSgrp', 'LrToSgrp', 'ToMon'],
'ToMon': ['CToMon', 'IdToMon', 'IToMon', 'LrToMon'],
'IToMon': ['CIToMon', 'ILrToMon'],
'IdToSgrp': ['CIdToSgrp', 'IdLrToSgrp', 'IdToMon'],
'IdToMon': ['CIdToMon', 'IdLrToMon'],
'ToImpA': ['ToDivLat', 'LrToMag'],
'LrToMag': ['RToMag', 'LrToSgrp'],
'LrToSgrp': ['IdLrToSgrp', 'RToSgrp', 'LrToMon'],
'LrToMon': ['IdLrToMon', 'ILrToMon', 'RToMon'],
'ILrToMon': ['IRToMon'],
'IdLrToSgrp': ['IdRToSgrp', 'IdLrToMon'],
'IdLrToMon': ['IdRToMon'],
'RToUn': ['InToMon'],
'ToDivLat': ['CToDivLat', 'RToMag'],
'RToMag': ['CRToMag', 'InToMag', 'RToSgrp'],
'RToSgrp': ['CRToSgrp', 'IdRToSgrp', 'InToSgrp', 'RToMon'],
'RToMon': ['CRToMon', 'IdRToMon', 'IRToMon', 'InToMon'],
'IRToMon': ['CIRToMon', 'IInRToMon'],
'IdRToSgrp': ['CIdRToSgrp', 'IdInToSgrp', 'IdRToMon'],
'IdRToMon': ['CIdRToMon', 'IdInToMon'],
'ToUn': ['RToUn'],
'ToNUn': ['GalToLat'],
'GalToLat': ['InToLat'],
'InToLat': ['InToMag'],
'InToMag': ['CyInToMag', 'InToSgrp'],
'InToSgrp': ['CyInToSgrp', 'IdInToSgrp', 'InToMon'],
'InToMon': ['CyInToMon', 'IdInToMon', 'IInRToMon'],
'IInRToMon': ['CyIInRToMon'],
'CyInToMag': ['CInToMag', 'CyInToSgrp'],
'CyInToSgrp': ['CInToSgrp', 'CyIdInToSgrp', 'CyInToMon'],
'CyInToMon': ['CInToMon', 'CyIdInToMon', 'CyIInRToMon', 'ToGrp'],
'CyIInRToMon': ['IMTLChn'],
'ToGrp': ['AbToGrp'],
'CToSgrp': ['CIdToSgrp', 'CRToSgrp', 'CToMon'],
'CToMon': ['CIdToMon', 'CIToMon', 'CRToMon'],
'CIToMon': ['CIRToMon'],
'CIdToSgrp': ['CIdRToSgrp', 'CIdToMon'],
'CIdToMon': ['CIdRToMon'],
'CToDivLat': ['CRToMag'],
'CRToMag': ['CInToMag', 'CRToSgrp'],
'CRToSgrp': ['CInToSgrp', 'CIdRToSgrp', 'CRToMon'],
'CRToMon': ['CIdRToMon', 'CIRToMon', 'CInToMon'],
'CIRToMon': ['IMTLChn'],
'CIdRToSgrp': ['CIdInToSgrp', 'CIdRToMon'],
'CIdRToMon': ['CIdInToMon'],
'CInToMag': ['CInToSgrp'],
'CInToSgrp': ['CIdInToSgrp', 'CInToMon'],
'CInToMon': ['CIdInToMon', 'IMTLChn', 'AbLGrp'],
'IMTLChn': ['TrivA'],
'AbToGrp': ['TrivA'],
'ToRng': ['ToFld'],
'CToRng': ['ToFld'],
'ToFld': [],
'BA': ['pBA', 'BMag', 'BImpA', 'BUn', 'BNUn'],
'pBA': ['BMon'],
'BUn': ['BRUn', 'MA', 'CA$_2$', 'BRMod'],
'BNUn': ['BGalLat'],
'MA': ['TA', 'MonA'],
'TA': ['TrivA'],
'MonA': ['TrivA'],
'BMag': ['BSgrp', 'BLrMag'],
'BSgrp': ['BCSgrp', 'BLrSgrp', 'BMon', 'BIdSgrp'],
'BMon': ['BCMon', 'BIMon', 'BLrMon', 'BIdMon'],
'BIMon': ['BCIMon', 'BILrMon'],
'BIdSgrp': ['BCIdSgrp', 'BIdLrSgrp', 'BIdMon'],
'BIdMon': ['BCIdMon', 'BIdLrMon'],
'BImpA': ['BLrMag', 'BDivLat'],
'BLrMag': ['BRMag', 'BLrSgrp'],
'BLrSgrp': ['BRSgrp', 'BLrMon', 'BIdLrSgrp'],
'BLrMon': ['BRL', 'BILrMon', 'BIdLrMon'],
'BILrMon': ['BIRL'],
'BIdLrSgrp': ['BIdRSgrp', 'BIdLrMon'],
'BIdLrMon': ['BIdRL'],
'BRUn': ['BInFL'],
'BDivLat': ['BCDivLat', 'BRMag'],
'BRMag': ['BCRMag', 'BInMag', 'BRSgrp', 'NA'],
'BRSgrp': ['BCRSgrp', 'BInSgrp', 'BRL', 'BIdRSgrp'],
'BRL': ['BCRL', 'BIRL', 'BInFL', 'BIdRL'],
'BIRL': ['BCIRL', 'BIInFL', 'SeqA'],
'BIdRSgrp': ['BCIdRSgrp', 'BIdInSgrp', 'BIdRL'],
'BIdRL': ['BCIdRL', 'BIdInFL'],
'BGalLat': ['BInLat'],
'BInLat': ['BInMag'],
'BInMag': ['BCyInMag', 'BInSgrp'],
'BInSgrp': ['BCyInSgrp', 'BInFL', 'BIdInSgrp'],
'BInFL': ['BCyInFL', 'BIInFL', 'BIdInFL'],
'BIInFL': ['BCyIInFL'],
'BCyInMag': ['BCInMag', 'BCyInSgrp'],
'BCyInSgrp': ['BCInSgrp', 'BCyInFL', 'BCyIdInSgrp'],
'BCyInFL': ['BCInFL', 'BCyIInFL', 'BCyIdInFL'],
'BCyIInFL': ['BCIInFL'],
'BCSgrp': ['BCRSgrp', 'BCMon', 'BCIdSgrp', 'BSlat'],
'BCMon': ['BCIMon', 'BCRL', 'BCIdMon'],
'BCIMon': ['BCIRL'],
'BSlat': ['TrivA'],
'BCIdSgrp': ['BCIdRSgrp', 'BCIdMon'],
'BCIdMon': ['BCIdRL'],
'BCDivLat': ['BCRMag'],
'BCRMag': ['BCInMag', 'BCRSgrp'],
'BCRSgrp': ['BCInSgrp', 'BCRL', 'BCIdRSgrp'],
'BCRL': ['BCIRL', 'BCInFL', 'BCIdRL'],
'BCIRL': ['BCIInFL'],
'BCIdRSgrp': ['BCIdInSgrp', 'BCIdRL'],
'BCIdRL': ['BCIdInFL'],
'BCInMag': ['BCInSgrp'],
'BCInSgrp': ['BCInFL', 'BCIdInSgrp'],
'BCInFL': ['BCIInFL', 'BCIdInFL'],
'BCIInFL': ['TrivA'],
'CA$_2$': ['TrivA'],
'SeqA': ['RA'],
'NA': ['RA'],
'RA': ['IRA'],
'IRA': ['TrivA'],
'BRMod': ['TrivA'],
'Set': ['pSet', 'Unar', 'Mag'],
'pSet': ['Mon'],
'Unar': ['RUnar', 'Mset'],
'AAlg': [],
'BCI': [],
'RtQgrp': ['RtLp', 'Qgrp'],
'Qgrp': ['Lp', 'CQgrp', 'InMag', 'MouQgrp'],
'RtLp': ['Lp'],
'MouQgrp': ['MouLp'],
'Lp': ['MouLp', 'LNeofld'],
'MouLp': ['Grp'],
'Shell': ['Srng$_{01}$'],
'Mag': ['CnjMag', 'Sgrp', 'LtQgrp', 'DivA', 'OrdA', 'QtMag', 'MedMag', 'Qnd', 'BCI', 'RtQgrp', 'Shell', 'Dtoid'],
'Bnd': ['IdMon', 'NBnd', 'RzBnd'],
'NBnd': ['RecBnd'],
'RecBnd': ['Slat'],
'Sgrp': ['Bnd', 'LtCanSgrp', 'CSgrp', 'Mon', 'LrSgrp', 'Sgrp$_0$', 'RegSgrp'],
'Sgrp$_0$': ['Srng$_0$'],
'RegSgrp': ['Bnd', 'InvSgrp'],
'InvSgrp': ['CliffSgrp', 'CInvSgrp'],
'Mon': ['LtCanMon', 'CMon', 'IdMon'],
'CanSgrp': ['CanMon', 'CanCSgrp'],
'CanMon': ['CanCMon', 'Grp'],
'Grp': ['PGrp', 'NlGrp', 'AbGrp', 'pGrp', 'NRng'],
'AbpGrp': ['BGrp'],
'CMag': ['CSgrp'],
'CSgrp': ['Slat', 'CMon', 'CanCSgrp', 'qMV'],
'LtCanSgrp': ['LtCanMon', 'CanSgrp'],
'LtCanMon': ['CanMon'],
'CanCSgrp': ['CanCMon'],
'CInvSgrp': ['AbGrp', 'Slat'],
'CMon': ['CanCMon', 'Slat$_1$', 'Srng'],
'CanCMon': ['AbGrp'],
'AbGrp': ['BGrp', 'pAbGrp', 'Rng'],
'BGrp': ['TrivA'],
'NFld': ['Fld'],
'NRng': ['Rng', 'NRng$_1$'],
'NRng$_1$': ['Rng$_1$'],
'Neofld': ['DivRng'],
'Srng': ['IdSrng', 'Srng$_1$', 'Srng$_0$', 'CSrng'],
'Srng$_1$': ['Srng$_{01}$', 'Sfld'],
'Srng$_0$': ['IdSrng$_0$'],
'Srng$_{01}$': ['IdSrng$_{01}$', 'Rng$_1$'],
'Rng': ['CRng', 'Rng$_1$'],
'Rng$_1$': ['CRng$_1$', 'OreDom', 'NFld', 'RegRng'],
'RegRng': ['DivRng', 'CRegRng'],
'CRegRng': ['Fld'],
'CSrng': ['CSrng$_1$', 'CSrng$_0$'],
'CSrng$_1$': ['CSrng$_{01}$'],
'CSrng$_0$': ['CSrng$_{01}$'],
'CSrng$_{01}$': [],
'CRng': ['CRng$_1$', 'Fld'],
'CRng$_1$': ['BA', 'IntDom'],
'IntDom': ['UFDom'],
'DivRng': ['Fld'],
'Sfld': ['Fld'],
'Fld': [],
'CnjMag': ['CMag'],
'Dtoid': ['Slat'],
'UFDom': ['PIDom'],
'OreDom': ['DivRng'],
'PIDom': ['EucDom'],
'EucDom': ['Fld'],
'Mset': ['Gset'],
'Gset': ['RMod'],
'RMod': ['FVec'],
'FVec': ['NaA'],
'JorA': [],
'LNeofld': ['Neofld'],
'BilinA': ['LieA', 'AAlg'],
'CliffSgrp': ['Grp'],
'LieA': [],
'MedMag': [],
'NlGrp': ['AbGrp'],
'NaA': ['BilinA', 'JorA'],
'OrdA': ['Bnd'],
'pGrp': ['AbpGrp', 'CypGrp'],
'Qnd': [],
'qMV': ['MV','sqMV'],
'sqMV': [],
'QtMag': [],
'TrivA': []}
p=allclasses(chapter(2))
j=allclasses(chapter(3))
m=allclasses(chapter(4))
l=allclasses(chapter(5))
d=allclasses(chapter(6))
t=allclasses(chapter(7))
b=allclasses(chapter(8))
u=allclasses(chapter(9))
# +
jl=['Lat',
'pLat',
'lbLat',
'ubLat',
'LUn',
'LNUn',
'GalLat',
'LMag',
'LSgrp',
'LMon',
'IdSrng$_0$',
'IdSrng$_{01}$',
'KA',
'ILMon',
'IdLSgrp',
'IdLMon',
'LImpA',
'LrLMag',
'LrLSgrp',
'LrLMon',
'ILrLMon',
'IdLrLSgrp',
'IdLrLMon',
'RLUn',
'DivLat',
'RLMag',
'RLSgrp',
'RLMon',
'IRLMon',
'IdRLSgrp',
'IdRLMon',
'CLSgrp',
'CLMon',
'CILMon',
'CIdLSgrp',
'CIdLMon',
'CDivLat',
'BCKL',
'CRLMag',
'CRLSgrp',
'CRLMon',
'CIRLMon',
'CIdRLSgrp',
'CIdRLMon']
ml=['Lat',
'pLat',
'LUn',
'LNUn',
'GalLat',
'LMag',
'LSgrp',
'LMon',
'ILMon',
'IdLSgrp',
'IdLMon',
'LImpA',
'LrLMag',
'LrLSgrp',
'LrLMon',
'ILrLMon',
'RtHp',
'IdLrLSgrp',
'IdLrLMon',
'RLUn',
'DivLat',
'RLMag',
'RLSgrp',
'RLMon',
'IRLMon',
'IdRLSgrp',
'IdRLMon',
'CLSgrp',
'CLMon',
'CILMon',
'CIdLSgrp',
'CIdLMon',
'CDivLat',
'BCKLat',
'CRLMag',
'CRLSgrp',
'CRLMon',
'CIRLMon',
'Hp',
'CIdRLSgrp',
'CIdRLMon',
'BrSlat']
ld=['DLat',
'pDLat',
'bDLat',
'DLUn',
'DLNUn',
'DLMag',
'DLSgrp',
'DLMon',
'',
'',
'',
'',
'DILMon',
'DIdLSgrp',
'DIdLMon',
'DLImpA',
'DLrLMag',
'DLrLSgrp',
'DLrLMon',
'DILrLMon',
'DIdLrLSgrp',
'DIdLrLMon',
'DRLUn',
'DDivLat',
'DRLMag',
'DRLSgrp',
'DRL',
'bDRLat',
'DIRL',
'DIdRLSgrp',
'DIdRL',
'DFL',
'DFL$_c$',
'DFL$_e$',
'DFL$_w$',
'DFL$_{ec}$',
'DFL$_{ew}$',
'DGalLat',
'DInLat',
'DInLMag',
'DInLSgrp',
'DInFL',
'DIInFL',
'CyDInLMag',
'CyDInLSgrp',
'CyDInFL',
'CyDIInFL',
'CDLSgrp',
'CDLMon',
'CDILMon',
'CDIdLSgrp',
'CDIdLMon',
'CDDivLat',
'',
'CDRLMag',
'CDRLSgrp',
'CDRL',
'CDIRL',
'CDIdRLSgrp',
'CDIdRL',
'CDIdInFL',
'CDInLMag',
'CDInLSgrp',
'CDInFL',
'CDIInFL',
'',
'',
'',
'',
'',
'',
'CDdLat',
'',
'',
'',
'DBilat',
'CanDRL',
'',
'',
'',
'',
'',
'']
dt=['ToLat',
'pToLat',
'bToLat',
'ToUn',
'ToNUn',
'',
'',
'',
'',
'',
'',
'',
'',
'',
'ToMag',
'ToSgrp',
'ToMon',
'IToMon',
'IdToSgrp',
'IdToMon',
'ToImpA',
'LrToMag',
'LrToSgrp',
'LrToMon',
'ILrToMon',
'IdLrToSgrp',
'IdLrToMon',
'RToUn',
'ToDivLat',
'RToMag',
'RToSgrp',
'RToMon',
'IRToMon',
'IdRToSgrp',
'IdRToMon',
'GalToLat',
'InToLat',
'InToMag',
'InToSgrp',
'',
'',
'CyInToMag',
'CyInToSgrp',
'',
'',
'ToGrp',
'',
'CToSgrp',
'CToMon',
'CIToMon',
'CIdToSgrp',
'CIdToMon',
'CivToLat',
'CRToMag',
'',
'',
'',
'',
'',
'CInToMag',
'',
'',
'',
'AbToGrp',
'GBLChn',
'GMVChn',
'ToWHp',
'BrA',
'',
'',
'CRToSgrp',
'CRToMon',
'CIRToMon',
'MTLChn',
'BLChn',
'MVChn',
'',
'GödChn',
'CIdRToSgrp',
'CIdRToMon',
'CInToSgrp',
'CInToMon',
'DunnMon',
'CIInRToMon',
'',
'',
'',
'',
'',
'ToRng',
'CToRng',
'','']
db=['BLat', 'pBLat', 'BBLat', 'BUn', 'BNUn', '', '', '', '',
'', '', '', '', '', 'BMag', 'BSgrp', 'BMon', 'BIMon',
'BIdSgrp', 'BIdMon', 'BImpA', 'BLrMag', 'BLrSgrp', 'BILrMon',
'BIdLrSgrp', 'BIdLrMon', 'BRUn', 'BDivLat', 'BRMag', 'BRSgrp',
'BRL', 'BIRL', 'BIdRSgrp', 'BIdRL', 'BGalA', 'BInLat',
'BInMag', 'BInSgrp', 'BInFL', 'BIInFL', 'BCyInMag',
'BCyInSgrp',
'BCyInFL',
'BCyIInFL',
'',
'',
'BCSgrp',
'BCMon',
'BCIMon',
'BCIdSgrp',
'BCIdMon',
'BCDivLat',
'BCRMag',
'',
'',
'',
'',
'',
'BCInMag',
'',
'',
'',
'',
'',
'',
'',
'',
'',
'',
'BCRSgrp',
'BCRMon',
'BCIRMon',
'',
'',
'',
'',
'',
'BCIdRSgrp',
'BCIdRMon',
'BCInSgrp',
'BCInMon',
'',
'BCIInRMon',
'',
'',
'',
'',
'',
'',
'','','']
# -
pj=['Jslat',
'pJslat',
'JUn',
'JNUn',
'JMag',
'JSgrp',
'JMon',
'IJMon',
'IdJSgrp',
'IdJMon',
'JImpA',
'LrJMag',
'LrJSgrp',
'LrJMon',
'ILrJMon',
'IdLrJSgrp',
'IdLrJMon',
'RJUn',
'DivJslat',
'RJMag',
'RJSgrp',
'RJMon',
'IRJMon',
'IdRJSgrp',
'IdRJMon',
'GalJslat',
'InLat',
'InLMag',
'InLSgrp',
'InLMon',
'IInLMon',
'CyInLMag',
'CyInLSgrp',
'CyInLMon',
'CyIInLMon',
'LGrp',
'CJSgrp',
'CJMon',
'CIJMon',
'CIdJSgrp',
'CIdJMon',
'CDivJslat',
'BCKJ',
'CRJMag',
'HilAJ',
'TarAJ',
'CRJSgrp',
'CRJMon',
'CIRJMon',
'CIdRJSgrp',
'CIdRJMon',
'CInLMag',
'CInLSgrp',
'CInLMon',
'CIInLMon',
'AbLGrp']
pm=['Mslat',
'pMslat',
'MUn',
'MNUn',
'MMag',
'MSgrp',
'MMon',
'IMMon',
'IdMSgrp',
'IdMMon',
'MImpA',
'LrMMag',
'LrMSgrp',
'LrMMon',
'ILrMMon',
'IdLrMSgrp',
'IdLrMMon',
'RMUn',
'DivMslat',
'RMMag',
'RMSgrp',
'RMMon',
'IRMMon',
'IdRMSgrp',
'IdRMMon',
'GalMslat',
'',
'',
'',
'',
'',
'',
'',
'',
'',
'',
'CMSgrp',
'CMMon',
'CIMMon',
'CIdMSgrp',
'CIdMMon',
'CDivMslat',
'BCKM',
'CRMMag',
'HilAM',
'TarAM',
'CRMSgrp',
'CRMMon',
'CIRMMon',
'CIdRMSgrp',
'CIdRMMon',
'',
'',
'',
'',
'']
pp={p[i]:([pj[i],pm[i]] if pm[i]!="" else [pj[i]]) for i in range(len(p))}
jj={j[i]:[jl[i]] for i in range(len(j))}
mm={m[i]:[ml[i]] for i in range(len(m))}
ll={l[i]:[ld[i]] for i in range(len(l))}
dd={d[i]:([dt[i],db[i]] if db[i]!="" else [dt[i]]) for i in range(len(d))}
len(pp),len(jj),len(mm),len(ll),len(dd),
cpo=allclassposets()
print(len(cpo))
tkz="".join(cpo)
nli=allnodes(tkz)
print(len(nli),len(set(nli)))
nds = {x:lowercovers(x,tkz) for x in nli}
for x in pp.keys():
nds[x]=nds[x]+pp[x] if x in nds.keys() else pp[x]
for x in jj.keys():
nds[x]=nds[x]+jj[x] if x in nds.keys() else jj[x]
for x in mm.keys():
nds[x]=nds[x]+mm[x] if x in nds.keys() else mm[x]
for x in ll.keys():
nds[x]=nds[x]+ll[x] if x in nds.keys() else ll[x]
for x in dd.keys():
nds[x]=nds[x]+dd[x] if x in nds.keys() else dd[x]
len(nds)
lowercovers('JMon',tkz)
nds['JMon']
scp=subclassposet()
len(scp)
for x in scp.keys():
scp[x]+=([y for y in nds[x] if y!='' and y not in scp[x]] if x in nds.keys() else [])
d=set(s.replace('$\\ell$','L') for s in cli)-set(nli)
len(d)
sc={"a":["b"],"b":[]}
from graphviz import Graph
from IPython.display import display_html
P = Graph()
P.attr('node', shape='circle', width='.15', height='.15', fixedsize='true', fontsize='10')
#P.attr('node', shape='oval', fontsize='10')
for x in sc: P.node(str(x), color='red')
for x in sc: P.edges([(str(x),str(y)) for y in sc[x]])
display_html(P._repr_svg_(),raw=True)
di={0:"a",1:"b"}
for x in di: print(x)
d=set(nli)-set(s.replace('$\\ell$','L') for s in cli)
print(len(d))
d
# %%time
checkfs([1])#,5,6,7,8])
writefile("ch12.tex",latex_st)
print(finitememberslatex(fs))
# %%time
SkLat=['(((x^y)^z)=(x^(y^z)))', '((x^x)=x)', '(((x v y) v z)=(x v (y v z)))', '((x v x)=x)', '((x^(x v y))=x)', '((x v (x^y))=x)', '(((x v y)^y)=y)', '(((x^y) v y)=y)',
'x<=y<->x^y=x&y^x=x']
#a=p9(SkLat,[],1000,1000,[4])
a=p9(SkLat,["x<=y->x^z<=y^z"],100,1000)#,[4])
a
# %%time
fs=finespectrum(lat,"CIRL",5,True)
# %%time
#RtHp=msl+['((x<=y) <-> ((y/x)=1))', '(((x*y)*z)=(x*(y*z)))', '((x*1)=x)', '((1*x)=x)', '((x/(y*z))=((x/z)/y))', '((x/x)=1)', '(((x/y)*y)=((y/x)*x))']+['x<=y -> z*x<=z*y']
RtHp=msl+['((x<=y) <-> ((y/x)=1))', '(((x*y)*z)=(x*(y*z)))', '((x*1)=x)', '((1*x)=x)', '((x/(y*z))=((x/z)/y))', '((x/x)=1)', '(((x/y)*y)=((y/x)*x))',
'((y*z)\ x)=(z\ (y\ x))', '((x\ x)=1)', '((x*(x\ y))=(y*(y\ x)))','((x<=y) <-> ((x\ y)=1))']
#]+['x<=y -> z/y<=z/x']
#a=p9(RtHp,['x<=y -> z/y<=z/x'],0,1000) #provable
a=p9(RtHp,[],1000,1000,[5])#[1, 1, 2, 6, 17] GHp:[1, 1, 2, 5, 10]
#a=p9(RtHp,['x<=y -> z*x<=z*y'],0,1000)
a
# %%time
IdInPoMon=po+['x*x=x','(~(-(x))=x)', '(-(~(x))=x)', '(((x*y)<=z) <-> (y<=~((-(z)*x))))', '(((x*y)<=z) <-> (x<=-((y*~(z)))))', '(((x*y)*z)=(x*(y*z)))', '((x*1)=x)', '((1*x)=x)']
a=p9(IdInPoMon,[],1000,1000,[9])
# %%time
CIdInPoMon=po+['x*y=y*x','x*x=x','(~(-(x))=x)', '(-(~(x))=x)', '(((x*y)<=z) <-> (y<=~((-(z)*x))))', '(((x*y)<=z) <-> (x<=-((y*~(z)))))', '(((x*y)*z)=(x*(y*z)))', '((x*1)=x)', '((1*x)=x)']
a=p9(CIdInPoMon,[],1000,1000,[9])
# %%time
fs=finespectrum(axioms[1],"CInPoMon",9,True) # A328705!!!!!
finespectrum(["c=c"],"AbGrp",8)
finespectrum(["c=c"],"Grp",8)
m=re.findall(r"\\hypertarget{(.*?)}{",latex_st,flags=re.DOTALL)
len(m)
len(Po)
def rep(m): return "\\hypertarget{"+m.group(1)+"}{\\section{"+m.group(1)+": "+m.group(2)+"}}"
st1=re.sub(r"\\hypertarget{(.*?)}{\\section{(.*?)}}",rep,latex_st,flags=re.DOTALL)
len(st1)
st1=st1.replace(r"\abbreviation{",r"%\abbreviation{")
ch=["" for n in range(4)]
n=1
#st1=chapter(n)
st1=latex_st
len(st1)
m=re.findall(r"\\hypertarget{(.*?)}{",st1,flags=re.DOTALL)
len(m)
m
se = {cl:section_mo(cl,st1) for cl in m}
len(se)
[cl for cl in m if se[cl]==None]
len(se)
len(set(m)) #-set(se.keys())
set(Po)-set(m)
st2 = "".join(se[cl].group(1) for cl in Po)
len(st2)
ch = [st1[:se[m[0]].start(1)]+st2+st1[se[m[-1]].end(1):]]
len(ch[0])
len(Po)
set(Po)-set(m),set(m)-set(Po)
print(st1[:2000])
for cl in m:
latex_st = set_classname(cl,m+": "+classname(m),latex_st)
lc=subclassposet()
uc=superclassposet()
for c in lc.keys():
for s in lc[c]:
if s in uc.keys() and c not in uc[s]: print(c,s)
for c in uc.keys():
for s in uc[c]:
if s in lc.keys() and c not in lc[s]: print(c,s)
lc
uc
mm=re.findall(r"\\abbreviation{(.*?)}",latex_st,flags=re.DOTALL)
len(mm)
m=re.findall(r"\\hypertarget{(.*?)}{",latex_st,flags=re.DOTALL)
len(m)
m[54:97]
def finespectralatex(cli):
li = list(reversed(sorted([(finitemembers(s)[:10],s) for s in cli])))
st = r"\begin{tabular}{|l|l|l|}\hline\nName& Fine spectrum& OEIS\\\hline"
return st+"&\\\\\n".join(x[1]+"& "+str(x[0])[1:-1] for x in li)+"\hline\n\end{tabular}\n"
st1=finespectralatex(m)
print(st1[:1000])
writefile("ch3.tex",st1)
[cl for cl in m if fulldefinition(cl)!="none"]
for s in m: print(s.ljust(11, ' '),finitemembers(s))#,info=True))
# +
# %%time
ch=jsl
def compareandupdatefs(chax,cl):
li = finitemembers(cl)
sli = [x for x in li if x <= 1000]
n = min(max(len(sli),3),6)
fs = finespectrum(chax,cl,n,True)
print(cl,li)
return li[:n]==fs, fs
for cl in m[54:97]:
if fulldefinition(cl)!="none":
fl = compareandupdatefs(ch,cl)
if not fl[0]:
print("***********",fl)
latex_st = set_finitemembers(cl,fl[1],latex_st)
# -
fulldefinition("Bnd")
len(latex_st)
writefile("SOASuo.tex",latex_st)
print(finitememberslatex(fs))
# %%time
fs=finespectrum([],'Qnd',5,True)
a=p9(jsl+['(((x->y)->((y->z)->(x->z)))=1)', '((1->x)=x)', '((x->1)=1)', '((x->(x v y))=1)', '(x<=((x->y)->y))'],[],1000,1000,[6])
Po=[[],[
'Pos', #po-algs 10
'pPos',
'PoUn',
'PoNUn',
'PoMag',
'PoSgrp',
'PoMon',
'IPoMon',
'IdPoSgrp',
'IdPoMon',
'PoImpA', #implicative, left-residuated 7
'LrPoMag',
'LrPoSgrp',
'LrPoMon',
'Polrim',
'IdLrPoSgrp',
'IdLrPoMon',
'RPoUn', #residuated 8
'DivPos',
'RPoMag',
'RPoSgrp',
'RPoMon',
'Porim',
'IdRPoSgrp',
'IdRPoMon',
'GalPos', #Galois, involutive 6
'InPos',
'InPoMag',
'InPoSgrp',
'InPoMon',
'InPorim',
#'IdInPoSgrp',
#'IdInPoMon',
'CyInPoMag', #cyclic involutive 5
'CyInPoSgrp',
'CyInPoMon',
'CyInPorim',
#'CyIdInPoSgrp',
#'CyIdInPoMon',
'PoGrp',
'CPoSgrp', #commutative po-algs 5
'CPoMon',
'CIPoMon',
'CIdPoSgrp',
'CIdPoMon',
'CDivPos', #commutative residuated 8
'BCK',
'CRPoMag',
'CRPoSgrp',
'CRPoMon',
'Pocrim',
'CIdRPoSgrp',
'CIdRPoMon',
'CInPoMag', # commutative involutive 5
'CInPoSgrp',
'CInPoMon',
'InPocrim',
'APoGrp',
], #total 54 categories of po-algebras
['Jslat',
'pJslat',
'lbJslat',
'ubJslat',
'JUn',
'JNUn',
'GalJslat',
'JMag',
'JSgrp',
'JMon',
'IdSRng$_0$',
'IdSRng$_{01}$',
'KA',
'IJMon',
'IdJSgrp',
'IdJMon',
'JImpA',
'LrJMag',
'LrJSgrp',
'LrJMon',
'ILrJMon',
'IdLrJSgrp',
'IdLrJMon',
'DivJslat',
'RJMag',
'RJSgrp',
'RJMon',
'IRJMon',
'IdRJSgrp',
'IdRJMon',
'CJSgrp',
'CJMon',
'CIJMon',
'CIdJSgrp',
'CIdJMon',
'CDivJslat',
'BCKJslat',
'CRJMag',
'CRJSgrp',
'CRJMon',
'CIRJMon',
'CIdRJSgrp',
'CIdRJMon',
],
['Mslat',
'pMslat',
'MUn',
'MNUn',
'GalMslat',
'MMag',
'MSgrp',
'MMon',
'IMMon',
'IdMSgrp',
'IdMMon',
'MImpA',
'LrMMag',
'LrMSgrp',
'LrMMon',
'ILrMMon',
'RtHp',
'IdLrMSgrp',
'IdLrMMon',
'DivMslat',
'RMMag',
'RMSgrp',
'RMMon',
'IRMMon',
'IdRMSgrp',
'IdRMMon',
'CMSgrp',
'CMMon',
'CIMMon',
'CIdMSgrp',
'CIdMMon',
'CDivMslat',
'BCKMslat',
'CRMMag',
'CRMSgrp',
'CRMMon',
'CIRMMon',
'Hp',
'CIdRMSgrp',
'CIdRMMon',
'HilA',
'BrSlat',
],
['Lat',
'pLat',
'bLat',
'$\\ell$Un',
'$\\ell$NUn',
'$\\ell$Mag',
'$\\ell$Sgrp',
'$\\ell$Mon',
'KLat',
'MLat',
'MultLat',
'I$\\ell$Mon',
'Id$\\ell$Sgrp',
'Id$\\ell$Mon',
'$\\ell$ImpA',
'Lr$\\ell$Mag',
'Lr$\\ell$Sgrp',
'Lr$\\ell$Mon',
'ILr$\\ell$Mon',
'IdLr$\\ell$Sgrp',
'IdLr$\\ell$Mon',
'DivLat',
'R$\\ell$Mag',
'R$\\ell$Sgrp',
'RL',
'bRLat',
'IRL',
'IdR$\\ell$Sgrp',
'IdRL',
'FL',
'FL$_c$',
'FL$_e$',
'FL$_w$',
'FL$_{ec}$',
'FL$_{ew}$',
'GalLat',
'InLat',
'In$\\ell$Mag',
'In$\\ell$Sgrp',
'InFL',
'IInFL',
'CyIn$\\ell$Mag',
'CyIn$\\ell$Sgrp',
'CyInFL',
'CyIInFL',
'C$\\ell$Sgrp',
'C$\\ell$Mon',
'CI$\\ell$Mon',
'CId$\\ell$Sgrp',
'CId$\\ell$Mon',
'CDivLat',
'BCKlat',
'CR$\\ell$Mag',
'CR$\\ell$Sgrp',
'CRL',
'CIRL',
'CIdR$\\ell$Sgrp',
'CIdRL',
'CIdInFL',
'CIn$\\ell$Mag',
'CIn$\\ell$Sgrp',
'CInFL',
'CIInFL',
'SkLat',
'JsdLat',
'MsdLat',
'SdLat',
'NdLat',
'ADLat',
'CdLat',
'OLat',
'OMLat',
'MOLat',
'Bilat',
'CanRL',
'CdMLat',
'FRng',
'ILLA',
'LLA',
'MALLA',
'psMV',
]
]
Po=['DLat',
'pDLat',
'BDLat',
'D$\\ell$Un',
'D$\\ell$NUn',
'pcDLat',
'OckA',
'DeMA',
'DMMon',
'DpAlg',
'DdpAlg',
'DDblpAlg',
'StAlg',
'DblStAlg',
'D$\\ell$Mag',
'D$\\ell$Sgrp',
'D$\\ell$Mon',
'DI$\\ell$Mon',
'DId$\\ell$Sgrp',
'DId$\\ell$Mon',
'D$\\ell$ImpA',
'DLr$\\ell$Mag',
'DLr$\\ell$Sgrp',
'DLr$\\ell$Mon',
'DILr$\\ell$Mon',
'DIdLr$\\ell$Sgrp',
'DIdLr$\\ell$Mon',
'DDivLat',
'DR$\\ell$Mag',
'DR$\\ell$Sgrp',
'DRL',
'DIRL',
'DIdR$\\ell$Sgrp',
'DIdRL',
'DGalLat',
'DInLat',
'DIn$\\ell$Mag',
'DIn$\\ell$Sgrp',
'DInFL',
'DIInFL',
'CyDIn$\\ell$Mag',
'CyDIn$\\ell$Sgrp',
'CyDInFL',
'CyDIInFL',
'$\\ell$Grp',
'Rep$\\ell$Grp',
'CD$\\ell$Sgrp',
'CD$\\ell$Mon',
'CDI$\\ell$Mon',
'CDId$\\ell$Sgrp',
'CDId$\\ell$Mon',
'CDDivLat',
'CDR$\\ell$Mag',
'CDR$\\ell$Sgrp',
'CDRL',
'CDIRL',
'CDIdR$\\ell$Sgrp',
'CDIdRL',
'CDIn$\\ell$Mag',
'CDIn$\\ell$Sgrp',
'CDInFL',
'CDIInFL',
'A$\\ell$Grp',
'GBL',
'GMV',
'WHp',
'BrA',
'GBA',
'BoolLat',
'CRSlSgrp',
'CRSlMon',
'CIRSlMon',
'MTLA',
'BLA',
'MV',
'HA',
'GödA',
'CIdRSlSgrp',
'CIdRSlMon',
'CInSlSgrp',
'CInSlMon',
'DunnMon',
'CIInRSlMon',
'ImpLat',
'KLA',
'NVLGrp',
'LA$_n$',
'MZrd',
'$\\ell$Rng',
'ORng',
'C$\\ell$Rng',
'CORng',
'OFld',
#'DLE',
#'DLOS',
#'SlMag',
#'SlSgrp',
#'CSlSgrp',
#'SlMon',
#'CSlMon',
#'ISlMon',
#'CISlMon',
#'IdSlSgrp',
#'CIdSlSgrp',
#'IdSlMon',
#'CIdSlMon',
#'SlImpA',
#'LrSlMag',
#'LrSlSgrp',
#'LrSlMon',
#'ILrSlMon',
#'IdLrSlSgrp',
#'IdLrSlMon',
#'SlDivLat',
#'CSlDivLat',
#'RSlMag',
#'CRSlMag',
#'RSlSgrp',
#'RSlMon',
#'IRSlMon',
#'IdRSlSgrp',
#'IdRSlMon',
#'SlUn',
#'SlNUn',
#'SlGalLat',
#'SlInLat',
#'InSlMag',
#'CyInSlMag',
#'CInSlMag',
#'InSlSgrp',
#'CyInSlSgrp',
#'InSlMon',
#'CyInSlMon',
#'IInRSlMon',
#'CyIInRSlMon',
#'Chain',
#'OSgrp',
#'COSgrp',
#'OMon',
#'OMonZ',
#'COMon',
#'OGrp',
#'OAbGrp',
#'OSlat',
#'RLGrp',
'ToLat', #Chn
'pToLat',
'ToMag',
'ToSgrp',
'ToMon',
'IToMon',
'IdToSgrp',
'IdToMon',
'ToImpA',
'LrToMag',
'LrToSgrp',
'LrToMon',
'ILrToMon',
'IdLrToSgrp',
'IdLrToMon',
'DivChn',
'RToMag',
'RToSgrp',
'RToMon',
'IRToMon',
'IdRToSgrp',
'IdRToMon',
'ToUn',
'ToNUn',
'GalChn',
'InChn',
'InToMag',
'InToSgrp',
'InToMon',
'IInRToMon',
'CyInToMag',
'CyInToSgrp',
'CyInToMon',
'CyIInRToMon',
'ToGrp',
'CToSgrp',
'CToMon',
'CIToMon',
'CIdToSgrp',
'CIdToMon',
'CDivChn',
'CRToMag',
'CRToSgrp',
'CRToMon',
'CIRToMon',
'CIdRToSgrp',
'CIdRToMon',
'CInToMag',
'CInToSgrp',
'CInToMon',
'CIInRToMon',
'AbToGrp',
'BA',
'pBA',
'BUn',
'BNUn',
'MA',
'TA',
'MonA',
'BMag',
'BSgrp',
'BMon',
'BIMon',
'BIdSgrp',
'BIdMon',
'BImpA',
'BLrMag',
'BLrSgrp',
'BLrMon',
'BILrMon',
'BIdLrSgrp',
'BIdLrMon',
'BDivLat',
'BRMag',
'BRSgrp',
'BRL',
'BIRL',
'BIdRSgrp',
'BIdRL',
'BGalLat',
'BInLat',
'BInMag',
'BInSgrp',
'BInFL',
'BIInFL',
'BCyInMag',
'BCyInSgrp',
'BCyInFL',
'BCyIInFL',
'BCSgrp',
'BCMon',
'BCIMon',
'BSlat',
'BCIdSgrp',
'BCIdMon',
'BCDivLat',
'BCRMag',
'BCRSgrp',
'BCRL',
'BCIRL',
'BCIdRSgrp',
'BCIdRL',
'BCInMag',
'BCInSgrp',
'BCInFL',
'BCIInFL',
'CA$_2$',
'SeA',
'NA',
'RA',
'IRA',
'DGrp',
'BRMod',
#'BAO',
#'BSp',
#'CanPMon',
'MonoUn',
'Unar',
'AAlg',
'BCI',
'RtQgrp',
'Qgrp',
'MouQgrp',
'Lp',
'MouLp',
'Shells',
'Mag',
'Bnd',
'NBnd',
'RBnd',
'Sgrp',
'Sgrp$_0$',
'RSgrp',
'InvSgrp',
'Mon',
'CanSgrp',
'CanMon',
'Grp',
'ApGrp',
'CMag',
'CSgrp',
'LCanSgrp',
'CanCSgrp',
'CInvSgrp',
'CMon',
'CanCMon',
'AbGrp',
'NFld',
'NRng',
'NRng$_1$',
'Nfld',
'SRng',
'SRng$_1$',
'SRng$_0$',
'SRng$_{01}$',
'Rng',
'Rng$_1$',
'RRng',
'CRRng',
'CRng',
'CRng$_1$',
'IntDom',
'DRng',
'Sfld',
'Fld',
'ConMag',
'Dtoid', #move to Po?
'UFDom',
'OreDom',
'PIDom',
'EucDom',
'MSet',
'Gset',
'RMod',
'FVec',
'JorA',
'LNfld',
'BilinA',
'CliffSgrp',
'LieA',
'MedMag',
'NlGrp',
'NaA',
'OrdA',
'p-Grp',
'Qnd',
'qMV',
'sqMV',
'QtMag',
'Set',
]
[ 'MonoUn',
'Unar',
'BCI',
'RtQgrp',
'Qgrp',
'MouQgrp',
'Lp',
'MouLp',
'Shells',
'Mag',
'Bnd',
'NBnd',
'RBnd',
'Sgrp',
'Sgrp$_0$',
'RSgrp',
'InvSgrp',
'Mon',
'CanSgrp',
'CanMon',
'Grp',
'ApGrp',
'CMag',
'CSgrp',
'LCanSgrp',
'CanCSgrp',
'CInvSgrp',
'CMon',
'CanCMon',
'AbGrp',
'NFld',
'NRng',
'NRng$_1$',
'Nfld',
'SRng',
'SRng$_1$',
'SRng$_0$',
'SRng$_{01}$',
'Rng',
'Rng$_1$',
'RRng',
'CRRng',
'CRng',
'CRng$_1$',
'IntDom',
'DRng',
'Sfld',
'Fld',
'ConMag',
'Dtoid', #move to Po?
'UFDom',
'OreDom',
'PIDom',
'EucDom',
'MSet',
'Gset',
'RMod',
'FVec',
'NaA',
'BilinA',
'LieA',
'JorA',
'AAlg',
'LNfld',
'CliffSgrp',
'MedMag',
'NlGrp',
'OrdA',
'p-Grp',
'Qnd',
'qMV',
'sqMV',
'QtMag',
'Set',
]
m
cl=re.findall(r"\\subsection{(.*?)}",st,flags=re.DOTALL)
cl
len(cl)
de=re.findall(r"\\begin{definition}(.*?)\\end{definition}",st,flags=re.DOTALL)
print(de[6])
n=0
n=st1.find("\\begin{definition}",n+1)
st2=st1[:n]+st1[n:].replace("==","\\end{defintion}\n==",1)
import re
m=re.search("begin{definition}.*?==",st1,flags=re.DOTALL)
m
def rep(m): return "\\begin{tabular}{|ll|}\\\\hline\n^[[Classtype"
st1=re.sub("\^(\[\[Classtype)",rep,st,flags=re.DOTALL)
m=re.search("\^(\[\[Classtype)",st,flags=re.DOTALL)
m
print(st[2300:3100])
def rep(m): return "\\begin{tabular}{|ll|}\\\\hline\n^[[Classtype"
st1=re.sub("\^(\[\[Classtype)",rep,st,flags=re.DOTALL)
def rep(m): return "hline\n\\end{tabular}\n\n\n\\begin{finite"
st1=re.sub("begin{finite",rep,st1,flags=re.DOTALL)
def rep(m): return m.group(1)+"&"+m.group(2)+"\\\\"
st1=re.sub("\^(\[\[.*?)\|(.*?)\|",rep,st1,flags=re.DOTALL)
st1=st1.replace("[[","((").replace("]]","))")
len(st1)
print(st1[2000:3500])
m=re.search(r"=====(.*?)=====.*?Abbreviation: \*\*(.*?)\*\*",st1,flags=re.DOTALL)
m.group(1), m.group(2), m
def rep(m): return "\n\\hrulefill\n%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%\n\\hypertarget{"+m.group(2)+"}{\\subsection{"+m.group(1)+"}}\\ \n\n\\abbreviation{"+m.group(2)+"}"
st2=re.sub(r"=====(.*?)=====.*?Abbreviation: \*\*(.*?)\*\*",rep,st1,flags=re.DOTALL)
len(st2)
print(st2[3000:5200])
def extract(st, env):
# find all environments in st named env and print them
li=re.findall(r"(\\begin{"+env+r"}.*?\\end{"+env+r"})",st,flags=re.DOTALL)
return li
a = extract(st,"examples")
len(a)
a[:20]
|
Python scripts/Editscripts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Installation
# # !pip install pandas
# # !pip install numpy
# # !pip install scipy
# # !pip install implicit
# -
# Imports
import pandas as pd
import numpy as np
import scipy.sparse as sparse
import implicit
import os
import random
# Environment and global variables
pd.set_option('display.max_columns',10)
os.environ['MKL_NUM_THREADS'] = '1'
os.environ['OPENBLAS_NUM_THREADS'] = '1'
# +
# Utility functions
# map the user and item names to contiguous integers and also return the maps
def maptrans(trans):
uniqueusers = np.sort(trans['user'].unique())
uniqueitems = np.sort(trans['item'].unique())
umap = dict(zip(uniqueusers,[i for i in range(len(uniqueusers))])) # this maps username -> index
imap = dict(zip(uniqueitems,[i for i in range(len(uniqueitems))])) # this maps itemname -> index
trans['user'] = trans.apply(lambda row: umap[row['user']], axis = 1)
trans['item'] = trans.apply(lambda row: imap[row['item']], axis = 1)
return (trans,umap,imap)
#return list of similar items, use the item-properties matrix (Q) to do nearest neighbour using cosine similarity
def findsimilaritems(item, item_vecs, n_similar=10):
#Calculate the item vector norms (the vector lengths)
item_norms = np.sqrt((item_vecs * item_vecs).sum(axis=1))
#Calculate the (cosine) similarity score: do dot product of selected content with every other content
#Note: cosine sim = A.B/(norm(A)*norm(B)), since B (item 450) is the same for every item A, we can ignore its norm in this calc
simscores = item_vecs.dot(item_vecs[item]) / item_norms
#Get the top 10 contents (do a sort)
top_idx = np.argpartition(simscores, -n_similar)[-n_similar:]
#Create a descending list of content-score tuples of most similar articles with this article.
similar = sorted(zip(top_idx, simscores[top_idx]/item_norms[item]), key=lambda x: -x[1])
return (similar)
#return the top 10 recommendations chosen based on the person / content vectors
#for contents never interacted with for any given person.
def recommend(user, sparse_user_item, userprefs, itemprops, num_items=10):
# create a template vector, where unrated items = 1, rated items =0
existing_ratings = sparse_user_item[user,:].toarray() # Get existing ratings for target
existing_ratings = existing_ratings.reshape(-1) + 1 # Add 1 to everything, so items with no rating = 1
existing_ratings[existing_ratings > 1] = 0 # make items already rated = 0
# Get dot product of the target user preferences and all item properties ~ P[user]*transpose(Q)
predrats = userprefs[user,:].dot(itemprops.T)
# Items already rated have their predictions multiplied by zero (ie eliminated)
predrats = predrats * existing_ratings
# Sort into descending order of predicted rating and select the topN item indexes
itemids = np.argsort(predrats)[::-1][:num_items]
# Start empty list to store items and scores
recs = []
for item in itemids: recs.append((item, predrats[item]))
return recs
def implicit_testusers(testset, userprefs, itemprops, debug=False):
errs = list([])
#tic = time.perf_counter()
for (indx,(uname,iname,rating)) in testset.iterrows():
if (debug): print('.', end = '')
err = abs(userprefs[uname,:].dot(itemprops[iname,:]) - rating)
errs.append(err)
#print(f"\ntime {time.perf_counter() - tic:0.4f} seconds")
return(errs)
def ahead(arr,r=7,c=7):
with np.printoptions(threshold=np.inf):
print(arr[0:r,0:c])
def sparsity(arr):
return np.isnan(arr).sum()/np.prod(arr.shape)
#1.0 - ( count_nonzero(arr) / float(arr.size) )
# -
path = '/home/mobasshir/recommendation_engine_lab/NUS-Artificial-Intelligence-Training/recommender/Datasets'
os.chdir(path)
trans = pd.read_csv('BookCrossings/BX-Book-Ratings.csv', sep=';', error_bad_lines=False, encoding="latin-1")
print(trans.head())
trans.columns = ['user','isbn','rating']
trans.rating.value_counts()
# +
# trans['rating'] = trans['rating'].apply(lambda x: 5 if x == 0 else x)
# -
trans['item'] = trans.groupby('isbn').grouper.group_info[0]
print(trans['item'])
trans = trans.drop_duplicates()
trans = trans.groupby(['user', 'item', 'isbn']).sum().reset_index()
print(trans)
trans,umap,imap = maptrans(trans)
print(trans)
sparse_item_user = sparse.csr_matrix((trans['rating'].astype(float), (trans['item'],trans['user'])))
sparse_user_item = sparse.csr_matrix((trans['rating'].astype(float), (trans['user'],trans['item'])))
model = implicit.als.AlternatingLeastSquares(factors=20, regularization=0.1, iterations=50)
alpha = 15
data = (sparse_item_user * alpha).astype('double')
model.fit(data)
item_id = 231
similar = model.similar_items(item_id)
for item, score in similar:
print(score,'\t',trans.isbn.loc[trans.item == item].iloc[0], "\n")
item_id = 32070
similar = model.similar_items(item_id)
for item, score in similar:
print(score,'\t',trans.isbn.loc[trans.item == item].iloc[0], "\n")
user_id = 8
recommendations = model.recommend(user_id, sparse_user_item, filter_already_liked_items=True)
for item, score in recommendations:
print(f'{score:0.5f}','\t', trans.isbn.loc[trans.item == item].iloc[0], "\n")
user_id = 26
recommendations = model.recommend(user_id, sparse_user_item, filter_already_liked_items=True)
for item, score in recommendations:
print(f'{score:0.5f}','\t', trans.isbn.loc[trans.item == item].iloc[0], "\n")
item = 26
recommendations = model.recommend(item, sparse_item_user, filter_already_liked_items=True)
for user, score in recommendations:
print(f'{score:0.5f}','\t', user, "\n")
# +
# https://towardsdatascience.com/building-a-collaborative-filtering-recommender-system-with-clickstream-data-dffc86c8c65
# -
|
notebooks/rec_implicit_bookcrossings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from prep.ann_approximation import *
from prep.derivatives import Preprocess_derivatives
import matplotlib.pyplot as plt
# We upload the solution of ordinary differential equation $u \sin x + u' \cos x = 1$ as f and create x - grid. The data is located in https://drive.google.com/drive/folders/1dEUbedeBekephqD61HeYH346ueZCX7gJ?usp=sharing
temp = np.load('Preprocessing/Fill366/fill366.npy')
f = np.stack([temp, temp], axis = 1)
x = np.linspace(0, 4*np.pi, f.shape[0])
# In this step, with the np.stack operation we have added an artificial dimension to the data, to avoid difficulties with ranks of tensorflow tensors, which can occur with the 1D - array (due to differences between tf.Tensor(..., shape=(n, 1), ...), and tf.Tensor(..., shape=(n,), ...)).
print(x.shape, f.shape)
plt.plot(x, f[:, 0], color = 'k')
# The entire process of artificial neural network fitting and further derivatives calculation is encapsulated in the *prep.ann_approximation.Preprocess_field* method, which returns the input function approximation and tuple of calculated derivatives.
approx, derivs_ann = Preprocess_field(f, order=2, steps = (x[1] - x[0], 1))
# To evaluate the quality of the derivatives from the automatic differentiation technique, we use previously implemented method *prep.derivatives.Preprocess_derivatives*, which uses analytical differentiation of polynomials, fit over specified window to the data:
derivs = Preprocess_derivatives(f[:, 0], steps = (x[1] - x[0], 1), smooth=False, mp_poolsize=1, max_order = 2)
# First of all, let's compare the quality of approximation of initial function with ANN:
plt.plot(x, approx[:, 0], color = 'k')
plt.plot(x, f[:, 0], color = 'r')
# To grasp the better understanding of the Preprocess_field output, let's examine it: we obtain a list, where each element is a list (*derivs*), which represents a derivative order. Next, in a "derivative order" list (here, represented by *derivs[0]*), the first element (*derivs[0][0]*) is the list of values for the derivatives, and the second element is the list of coordinates, by which the corrsponding elements of the first element are differentiated.
print(derivs[0], '\n')
print(derivs[0][0], '\n')
print(derivs[0][0][0], '\n')
# Next, we shall plot the function, approximation and derivatives to check the results
plt.plot(x[20:-20], approx[20:-20, 0], color = 'k', linestyle = '-')
plt.plot(x[20:-20], f[20:-20, 0], color = 'k', linestyle = '--')
plt.plot(x[20:-20], derivs[20:-20, 0], color = 'r', linestyle = '-')
plt.plot(x[20:-20], derivs_ann[0][0][0][0].reshape(f.shape)[20:-20, 0], color = 'r', linestyle = '--')
plt.plot(x[20:-20], derivs[20:-20, 1], color = 'b', linestyle = '-')
plt.plot(x[20:-20], derivs_ann[1][0][0][0].reshape(f.shape)[20:-20, 0], color = 'b', linestyle = '--')
# Now, let's create a bit more complex case, where we would try to approximate the solution of wave equation $u_{tt} = c^2 u_{tt}$, with $c = sqrt(0.5)$:
# +
def ic_1(x):
x_max = 5.
return np.sin(x/x_max*np.pi)*np.sin((5-x)/x_max*np.pi)
def ic_2(x):
x_max = 5.; coeff = -1
return coeff * np.sin(x/x_max*np.pi)*np.sin((5-x)/x_max*np.pi)
x_shape = 301; t_shape = 301
x_max = 5; t_max = 1
x_vals = np.linspace(0, x_max, x_shape)
t_vals = np.linspace(0, t_max, t_shape)
delta_x = x_vals[1] - x_vals[0]; delta_t = t_vals[1] - t_vals[0]
k = 0.5
wave_solution = np.empty((t_shape, x_shape))
wave_solution[:, 0] = wave_solution[:, -1] = 0
wave_solution[0, :] = ic_1(x_vals)
wave_solution[1, :] = wave_solution[0, :] + delta_t * ic_2(x_vals)
for t_idx in np.arange(2, t_shape):
for x_idx in np.arange(1, x_shape - 1):
wave_solution[t_idx, x_idx] = k*delta_t**2/delta_x**2 * (wave_solution[t_idx-1, x_idx+1] - 2*wave_solution[t_idx-1, x_idx] + wave_solution[t_idx-1, x_idx-1]) + 2*wave_solution[t_idx-1, x_idx] - wave_solution[t_idx-2, x_idx]
# -
Heatmap(wave_solution)
# We'll use previously introduced methods *Preprocess_field* and *Preprocess_derivatives* to get the tensors of derivatives, obtained with automatic differentiation and analytic differentiation correspondingly:
wave_solution_approximation, wave_derivs_ann = Preprocess_field(wave_solution, order=1, steps = (delta_t, delta_x), training_epochs = 300)
wave_derivs = Preprocess_derivatives(wave_solution, steps = (delta_t, delta_x), smooth=False, mp_poolsize=1, max_order = 1)
Heatmap(wave_solution_approximation)
Heatmap(np.abs(wave_solution_approximation - wave_solution))
# Here, we can notice, that in the centre of the domain the quality of the solution approximation is decent, while near the boundaries errors tend to be higher. The same pattern extends on the derivatives, but the areas of high errors tend to be larger.
plt.plot(wave_solution_approximation[25, :], color = 'r') # red line for the approximation
plt.plot(wave_solution[25, :], color = 'k') # black line for the initial function
Heatmap(wave_derivs_ann[0][0][1][0].reshape(wave_solution.shape))
Heatmap(wave_derivs[:, 1].reshape(wave_solution.shape))
plt.plot(wave_derivs[:, 1].reshape(wave_solution.shape)[25, :], color = 'k') # black line for the derivatives from baseline method
plt.plot(wave_derivs_ann[0][0][1][0].reshape(wave_solution.shape)[25, :], color = 'r') # red line for the derivatives from ann
Heatmap(np.abs(wave_derivs_ann[0][0][1][0].reshape(wave_solution.shape) - wave_derivs[:, 1].reshape(wave_solution.shape)))
|
estar/examples/ann_approximation_experiments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
# Initial imports
import numpy as np
import torch
from captum.attr import IntegratedGradients
from captum.attr import LayerConductance
from captum.attr import NeuronConductance
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import stats
import pandas as pd
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
ig = IntegratedGradients(model)
attributions, delta = ig.attribute(input, baseline, target=0, return_convergence_delta=True)
print('IG Attributions:', attributions)
print('Convergence Delta:', delta)
|
src/visualization/cnn_visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1-minute introduction to Jupyter ##
#
# A Jupyter notebook consists of cells. Each cell contains either text or code.
#
# A text cell will not have any text to the left of the cell. A code cell has `In [ ]:` to the left of the cell.
#
# If the cell contains code, you can edit it. Press <kbd>Enter</kbd> to edit the selected cell. While editing the code, press <kbd>Enter</kbd> to create a new line, or <kbd>Shift</kbd>+<kbd>Enter</kbd> to run the code. If you are not editing the code, select a cell and press <kbd>Ctrl</kbd>+<kbd>Enter</kbd> to run the code.
# Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
#
# Make sure you fill in any place that says `YOUR CODE HERE` or "YOUR ANSWER HERE", as well as your name and collaborators below:
NAME = ""
COLLABORATORS = ""
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "e092d6c6ae4be087700eabdd4622bb8f", "grade": false, "grade_id": "cell-a965c0c70630d0c7", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Assignment 7: Python data types: `dict`, `tuple`, `set`
#
# In this assignment, you should write your code in a **readable** way, and **modularise** chunks of code that would otherwise be repeated.
#
# Modularising a section of program code means to reorganise it in a way that allows it to be reused in another part of the code easily, usually in the form of a function.
#
# Your function definitions should have appropriate **docstrings**.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "cdfdc5e2d3f92eff65d2a80f3d49e8b0", "grade": false, "grade_id": "cell-041351b87d333512", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 1: Numerals to text
#
# In the formal style guides for the English language, it is recommended to spell out singular numbers in full. Applications that follow this guideline have to be able to convert numbers to text in a consistent fashion.
#
# Write a function, `text_numeral(num)` that takes in `num` (`int` < 1000) and returns a `str` of `num` in text format.
#
# ### Expected output
#
# >>> text_numeral(15)
# 'fifteen'
# >>> text_numeral(29)
# 'twenty-nine'
# >>> text_numeral(132)
# 'one hundred and thirty-two'
# >>> text_numeral(250)
# 'two hundred and fifty'
#
# **Hint:** Draw up a list of all the words you will need. Use a `dict` to store mappings of single-word numbers to strings (e.g. one, two, tree, ... , ten, eleven, twelve, ... twenty, thirty, forty, ...).
# Numbers not in this list have to be broken down to combinations of two single-word numbers.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "09c689160f23f9f7447a6dae2ce12807", "grade": false, "grade_id": "cell-ec172748d0a763bc", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Write a function to convert numbers to text numerals
def text_numeral(num:int):
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "91141909f770089da1b2e82a81ed37cb", "grade": true, "grade_id": "cell-1c54094ea29e1cf6", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# ## Cell for manual grading; ignore this cell
#
# YOUR ANSWER HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "5079ebe17b54a2d1a63e1a60881f0c22", "grade": true, "grade_id": "cell-0c7909ca3e617a23", "locked": true, "points": 3, "schema_version": 3, "solution": false, "task": false}
# Run this code cell to validate your function
test_values = {15: 'fifteen',
29: 'twenty-nine',
132: 'one hundred and thirty-two',
250: 'two hundred and fifty',
735: 'seven hundred and thirty-five',
}
for test,ans in test_values.items():
result = text_numeral(test)
print(f'{test}: {result}')
assert result == ans, f'input {test} gave {result}, should be {ans} instead'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "219a7b97350602a2493b779f6295789e", "grade": false, "grade_id": "cell-914b80a8defda316", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 2: Score and grade calculation
#
# The file `testscores.csv` contains test scores for a cohort of students. The first row contains header names.
#
# ### Task 1: Score to grade mapping
#
# **Generate** a `dict`, `grade_for`, that has score 0-100 (`int`) as keys, and the appropriate grade as values.
# You may use a `for` or `while` loop.
#
# The letter grade for each score range is as follows:
#
# A: score >= 70
# B: 70 > score >= 60
# C: 60 > score >= 55
# D: 55 > score >= 50
# E: 50 > score >= 45
# S: 45 > score >= 40
# U: 40 > score
#
# ### Expected output
#
# >>> grade_for[30]
# 'U'
# >>> grade_for[55]
# 'C'
# >>> grade_for[69]
# 'B'
# + deletable=false nbgrader={"cell_type": "code", "checksum": "60ab805a67c10826521688d9a8218cc1", "grade": false, "grade_id": "cell-08b4caba74070014", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Generate grade_for, a dict containing score as key and grade as value
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "88b44e9bc819809a7aab357454d58689", "grade": true, "grade_id": "cell-206387614b7b43e3", "locked": true, "points": 3, "schema_version": 3, "solution": false, "task": false}
# Run this cell to validate your dict
assert grade_for[30] == 'U', f'Grade for 30 should be U'
assert grade_for[55] == 'C', f'Grade for 55 should be C'
assert grade_for[69] == 'B', f'Grade for 69 should be B'
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "85cd5833a616cb49d423d9d6a8eb2d6f", "grade": true, "grade_id": "cell-7d1f75943a2cd036", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# Hidden tests; ignore this cell
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "cc4a4f87b757a23f41ca6b7bb53bc581", "grade": true, "grade_id": "cell-e12a9743d19ddc85", "locked": false, "points": 2, "schema_version": 3, "solution": true, "task": false}
# ## Cell for manual grading; ignore this cell
#
# YOUR ANSWER HERE
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "21f09cad318717530591fdb2b6cfaed3", "grade": false, "grade_id": "cell-acc7e9570795403d", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### Task 2: File reading and score calculation
#
# Write a function, `read_testscores(filename)` that opens `filename` and:
#
# 1. Reads in the student data and returns it as a `list` of `dict`s, with each `dict` representing a student,
# 2. Stores the score data of each student in a `dict` with appropriate keys,
# 3. Calculates the overall score of each student and stores it under an `'overall'` key,
# 4. Determines the grade of each student and stores it under the `'grade'` key.
# 5. Return the `list` of student data
#
# ### Overall score calculation formula
#
# The overall score is calculated using the following formula:
#
# overall = p1/30*15 + p2/40*30 + p3/80*35 + p4/30*20
#
# Where `p1`, `p2`, `p3`, and `p4` are the scores for P1, P2, P3, and P4 respectively.
# The overall score is to be **rounded up** to the nearest integer. You may use the `ceil()` function from the `math` module to do this.
#
# **Hint:** You are recommended to store the scores for P1 to P4 as a quadruple (4-`tuple`) under a `'score'` key, instead of under separate `'p1'` to `'p4'` keys, for easier retrieval.
# You may use the `zip()` function to handle multiple iterable collections in a single loop.
#
# ### Expected output
#
# >>> studentdata = read_testscores('testscores.csv')
# >>> studentdata[0]['class']
# 'Class1'
# >>> studentdata[0]['name']
# 'Student1'
# >>> studentdata[0]['overall']
# 51
# >>> studentdata[0]['grade']
# 'D'
# + deletable=false nbgrader={"cell_type": "code", "checksum": "af40acfc4184723c72f4ed0f04296c2b", "grade": false, "grade_id": "cell-4f04c23647096d3e", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Write a function that opens a file and returns student data
def read_testscores(filename):
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "3420156b8ff446c7a510e37e4513c9c0", "grade": true, "grade_id": "cell-3b1d7481c7ed87c9", "locked": false, "points": 5, "schema_version": 3, "solution": true, "task": false}
# ## Cell for manual grading; ignore this cell
#
# YOUR ANSWER HERE
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "68e9162ba7441ccea08d7a3607075f7a", "grade": true, "grade_id": "cell-532cf5963a408736", "locked": true, "points": 2, "schema_version": 3, "solution": false, "task": false}
# Run this cell to validate your dict
studentdata = read_testscores('testscores.csv')
for k,v in {'class':'Class1',
'name': 'Student1',
'overall': 51,
'grade': 'D'}.items():
assert studentdata[0][k] == v, f'Student 0: key {k} should give value {v} instead of {studentdata[0][k]}'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "d7aa42c5e525073de0fb38873ce3a13e", "grade": false, "grade_id": "cell-0308f0a8e2c9e4b0", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Part 3: Grade analysis
#
# Using `set`s, determine:
#
# 1. Which class(es) have no distinctions (grade 'A')
# 2. Which class(es) have 100% pass (grades A-E)
#
# Print the result.
#
# **Hint:** You should construct a new collection to hold the class grades, then use `set` operators to answer the above questions.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2ca0c42bf21b700041fb00ffad5922ac", "grade": true, "grade_id": "cell-129b553507ccc54c", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# Use sets to determine which class(es) have no distinctions
# + deletable=false nbgrader={"cell_type": "code", "checksum": "796a272670bd313decd972d629e05ebe", "grade": true, "grade_id": "cell-493ec62e26090c99", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# Use sets to determine which class(es) have 100% pass
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "6261f5013b0bc976c1bc84c3ac2e5879", "grade": false, "grade_id": "cell-ad1672e66728167f", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # Feedback and suggestions
#
# Any feedback or suggestions for this assignment?
# + [markdown] deletable=false nbgrader={"cell_type": "markdown", "checksum": "8b2c5aa73b1a793329d6b92780994acd", "grade": true, "grade_id": "cell-63f8f1cb7ff65bb1", "locked": false, "points": 0, "schema_version": 3, "solution": true, "task": false}
# YOUR ANSWER HERE
|
Assignment 7/assignment_07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Imports" data-toc-modified-id="Imports-0.0.1"><span class="toc-item-num">0.0.1 </span>Imports</a></span></li></ul></li></ul></li><li><span><a href="#plot-heatmap-of-ratings" data-toc-modified-id="plot-heatmap-of-ratings-1"><span class="toc-item-num">1 </span>plot heatmap of ratings</a></span></li><li><span><a href="#heatmap-of-rental-prices" data-toc-modified-id="heatmap-of-rental-prices-2"><span class="toc-item-num">2 </span>heatmap of rental prices</a></span></li></ul></div>
# -
# ### Imports
import pandas as pd
from matplotlib import pyplot as plt
import numpy as np
import pandas_explode
pandas_explode.patch()
# Load the data (will take a short while)
calendar = pd.read_csv("airbnb_data/calendar.csv")
demographics = pd.read_csv("airbnb_data/demographics.csv")
econ_state = pd.read_csv("airbnb_data/econ_state.csv")
listings = pd.read_csv("airbnb_data/listings.csv")
real_estate = pd.read_csv("airbnb_data/real_estate.csv")
venues = pd.read_csv("airbnb_data/venues.csv")
display(venues)
# +
# venues["latitude"] = venues["latitude"].multiply(1000).round(0)
# venues["longitude"]= venues["longitude"].multiply(1000).round(0)
# display(venues)
# -
venues_ny = venues.loc[venues['city'] == "new york city"]
# display(venues_ny)
venues_ny_hasRating = venues_ny#.dropna()
# display(venues_ny_hasRating)
df3 = venues_ny_hasRating.types.apply(pd.Series).add_prefix('code_')
# display(df3)
# # plot heatmap of ratings
# df3.reset_index(drop = True)
label = 'point_of_interest'#"university"#"lodging"#"museum"#'restaurant'"church"
notable = np.empty(0)
for idx in df3.index:
if label in df3["code_0"][idx]:
notable = np.append(notable,idx)
# print(notable)
# +
# for idx in df3.index:
# display(df3["code_0"][idx])
# break
# notable.astype(int)
# -
feature = venues_ny_hasRating.loc[notable]
feature
# +
import gmaps
# import gmaps.datasets
gmaps.configure(api_key='AIzaSyDI19Dy-f2Hfl7o-L-AqHRP94lwN_1poXU') # Fill in with your API key
# listing_lat = listings.loc[listings['state'] == "NY"]["latitude"].to_numpy()
# listing_lon = listings.loc[listings['state'] == "NY"]["longitude"].to_numpy()
lat = feature["latitude"].to_numpy()
lon = feature["longitude"].to_numpy()
rat = feature["rating"].to_numpy()
# locations = listings.loc[listings['state'] == "NY"][['latitude', 'longitude']]
locations = [lat, lon]
fig = gmaps.figure()
'''
heatmap_layer = gmaps.heatmap_layer(
feature[['latitude', 'longitude'], weights=feature['rating'],
point_radius = 5
)
'''
heatmap_layer = gmaps.heatmap_layer(
feature[['latitude', 'longitude']], point_radius = 3
)
fig.add_layer(heatmap_layer)
fig
# )
# fig = gmaps.figure()
# fig.add_layer(layer)
# fig
# print("done")
# -
# # heatmap of rental prices
listny = listings.loc[listings["metropolitan"] == 'NYC']
# cleanListing.isnull().sum()
listny.head()
listny.dropna(subset=['price'])
# +
fig2 = gmaps.figure()
heatmap_layer = gmaps.heatmap_layer(
listny[['latitude', 'longitude']], weights=listny['price'], point_radius = 3)
fig2.add_layer(heatmap_layer)
fig2
# -
|
RobertStuff/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import yaml
from run_experiment import choose_dataset_class,choose_task_classes
import torch
import numpy as np
from collections import Counter
# +
# 0. read the embeddings and labesl
yaml_args= yaml.load(open('../SA-config/a-sample-pos-get_cor.yaml'))
dataset_class = choose_dataset_class(yaml_args)
task_class, reporter_class, loss_class = choose_task_classes(yaml_args)
task = task_class(yaml_args)
expt_dataset = dataset_class(yaml_args, task)
# +
# 1. prepare the label matrix.
# train_data = expt_dataset.train_dataset
train_data = expt_dataset.dev_dataset
labels=[]
for obser in train_data.observations:
labels.append(task.labels(obser))
all_labels = torch.cat(labels, 0).numpy().astype(int)
all_labels.shape
all_labels_mat= np.zeros((all_labels.size, all_labels.max()+1))
all_labels_mat[np.arange(all_labels.size),all_labels] = 1
all_labels_mat.shape
# +
# 2. prepare the embedding matrix.
embeddings=[]
for obser in train_data.observations:
embeddings.append(obser.embeddings)
all_embeddings = torch.cat(embeddings, 0).numpy()
all_embeddings.shape
# -
# 3. compute the corrcoef matrix.
# num=204609
# corr_mat_whole = np.corrcoef(all_labels_mat[0:num].T, all_embeddings[0:num].T)
corr_mat_whole = np.corrcoef(all_labels_mat.T, all_embeddings.T)
corr_mat=np.absolute(corr_mat_whole[0:50,50:])
corr_mat.shape
# 4. output the corr_mat data. dump as a pkl.
# +
# 4. output the corr_sum_dim
# we compute the corr score of each dim for all the 50 pos labels.
dim_sum=[]
for i in range(768):
dim_sum.append(np.sum(corr_mat[:,i]))
# corr_sum_dim=np.argsort(dim_sum)[::-1]
# with open('../SA-dim-files/average_corr_dim.tsv','w') as fout:
# for dim in corr_sum_dim:
# fout.write('{} '.format(dim))
# +
# 5. output the weighted_average_corr_dim
# we compute the weighted average corr socre of each dim for all the pos labels
# 5.1 get the distribution/weights
all_labels
result = Counter(all_labels)
weights = [result[i]/len(all_labels) for i in range(50)]
# 5.2 get the weighted score
dim_weighted_average=[]
for i in range(768):
dim_weighted_average.append(np.average(corr_mat[:,i],weights=weights))
# # 5.3 output the dims
# dim_weighted_average_dim=np.argsort(dim_weighted_average)[::-1]
# with open('../SA-dim-files/weighted_average_corr_dim.tsv','w') as fout:
# for dim in dim_weighted_average_dim:
# fout.write('{} '.format(dim))
# -
np.argsort(weights)
weights[8],weights[9],
# +
# 6. output the max_corr
max_val=[]
for i in range(768):
max_val.append(np.max(corr_mat[:,i]))
dim_max_val=np.argsort(max_val)[::-1]
with open('../SA-dim-files/max_corr_dim.tsv','w') as fout:
for dim in dim_weighted_average_dim:
fout.write('{} '.format(dim))
# -
np.max(dim_sum),np.max(dim_weighted_average),np.max(corr_mat),np.max(max_val)
# +
def make_corr_file(layer=1):
# 0. read the embeddings and labesl
yaml_args= yaml.load(open('../SA-config/a-sample-pos-get_cor.yaml'))
yaml_args['model']['model_layer']=layer-1
dataset_class = choose_dataset_class(yaml_args)
task_class, reporter_class, loss_class = choose_task_classes(yaml_args)
task = task_class(yaml_args)
expt_dataset = dataset_class(yaml_args, task)
# 1. prepare the label matrix.
train_data = expt_dataset.train_dataset
labels=[]
for obser in train_data.observations:
labels.append(task.labels(obser))
all_labels = torch.cat(labels, 0).numpy().astype(int)
all_labels.shape
all_labels_mat= np.zeros((all_labels.size, all_labels.max()+1))
all_labels_mat[np.arange(all_labels.size),all_labels] = 1
all_labels_mat.shape
# 2. prepare the embedding matrix.
embeddings=[]
for obser in train_data.observations:
embeddings.append(obser.embeddings)
all_embeddings = torch.cat(embeddings, 0).numpy()
all_embeddings.shape
# 3. compute the corrcoef matrix.
# num=204609
# corr_mat_whole = np.corrcoef(all_labels_mat[0:num].T, all_embeddings[0:num].T)
corr_mat_whole = np.corrcoef(all_labels_mat.T, all_embeddings.T)
corr_mat=np.absolute(corr_mat_whole[0:50,50:])
corr_mat.shape
# 4. output the corr_mat data. dump as a pkl.
# 4. output the corr_sum_dim
# we compute the corr score of each dim for all the 50 pos labels.
dim_sum=[]
for i in range(768):
dim_sum.append(np.sum(corr_mat[:,i]))
corr_sum_dim=np.argsort(dim_sum)[::-1]
with open('../SA-dim-files/average_corr_dim_layer_{}.tsv'.format(layer),'w') as fout:
for dim in corr_sum_dim:
fout.write('{} '.format(dim))
# 5. output the weighted_average_corr_dim
# we compute the weighted average corr socre of each dim for all the pos labels
# 5.1 get the distribution/weights
all_labels
result = Counter(all_labels)
weights = [result[i]/len(all_labels) for i in range(50)]
# 5.2 get the weighted score
dim_weighted_average=[]
for i in range(768):
dim_weighted_average.append(np.average(corr_mat[:,i],weights=weights))
# 5.3 output the dims
dim_weighted_average_dim=np.argsort(dim_weighted_average)[::-1]
with open('../SA-dim-files/weighted_average_corr_dim_layer_{}.tsv'.format(layer),'w') as fout:
for dim in dim_weighted_average_dim:
fout.write('{} '.format(dim))
# 6. output the max_corr
max_val=[]
for i in range(768):
max_val.append(np.max(corr_mat[:,i]))
dim_max_val=np.argsort(max_val)[::-1]
with open('../SA-dim-files/max_corr_dim_layer_{}.tsv'.format(layer),'w') as fout:
for dim in dim_weighted_average_dim:
fout.write('{} '.format(dim))
# -
make_corr_file(layer=1)
make_corr_file(layer=3)
make_corr_file(layer=6)
make_corr_file(layer=9)
make_corr_file(layer=12)
for layer in [1,2,4,5,7,8,10,11]:
make_corr_file(layer=layer)
# ## study the distribution of the embeddings on different layers
# +
def get_corr_map(layer=1):
# 0. read the embeddings and labesl
yaml_args= yaml.load(open('../SA-config/a-sample-pos-get_cor.yaml'))
yaml_args['model']['model_layer']=layer-1
dataset_class = choose_dataset_class(yaml_args)
task_class, reporter_class, loss_class = choose_task_classes(yaml_args)
task = task_class(yaml_args)
expt_dataset = dataset_class(yaml_args, task)
# 1. prepare the label matrix.
train_data = expt_dataset.train_dataset
labels=[]
for obser in train_data.observations:
labels.append(task.labels(obser))
all_labels = torch.cat(labels, 0).numpy().astype(int)
all_labels.shape
all_labels_mat= np.zeros((all_labels.size, all_labels.max()+1))
all_labels_mat[np.arange(all_labels.size),all_labels] = 1
all_labels_mat.shape
# 2. prepare the embedding matrix.
embeddings=[]
for obser in train_data.observations:
embeddings.append(obser.embeddings)
all_embeddings = torch.cat(embeddings, 0).numpy()
all_embeddings.shape
# 3. compute the corrcoef matrix.
# num=204609
# corr_mat_whole = np.corrcoef(all_labels_mat[0:num].T, all_embeddings[0:num].T)
corr_mat_whole = np.corrcoef(all_labels_mat.T, all_embeddings.T)
return corr_mat_whole,all_embeddings,all_labels_mat
# -
corr_map_layer_dict=dict.fromkeys([i+1 for i in range(12)])
for layer in [1,3,6,9,12]:
corr_mat_whole,all_embeddings,all_labels_mat=get_corr_map(layer=layer)
corr_map_layer_dict[layer]={
'corr_mat_whole':corr_mat_whole,
'all_embeddings':all_embeddings,
'all_labels_mat':all_labels_mat
}
for layer in [2,4,5,7,8,10,11]:
corr_mat_whole,all_embeddings,all_labels_mat=get_corr_map(layer=layer)
corr_map_layer_dict[layer]={
'corr_mat_whole':corr_mat_whole,
'all_embeddings':all_embeddings,
'all_labels_mat':all_labels_mat
}
corr_map_layer_dict[1]
for layer in [1,3,6,9,12]:
# emb_mat = np.absolute(corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int))
emb_mat = corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int)
print('layer: ',layer,np.max(emb_mat),np.min(emb_mat),np.mean(np.absolute(emb_mat)),np.median(np.absolute(emb_mat)))
def get_top_dim_from_file(file):
with open(file,'r') as fin:
lines = fin.readlines()
line=lines[0].strip().split()
dims=[int(line[i]) for i in range(len(line))]
return dims
# +
for layer in [1,2,3,4,5,6,7,8,9,10,11,12]:
k=4
# emb_mat = np.absolute(corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int))
emb_mat = np.array(corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int))
dim_file = '../SA-dim-files/average_corr_dim_layer_{}.tsv'.format(layer)
print('layer\t{}\tmax-corr\t{}\tmean-corr\t{} '.format(layer,np.max(np.absolute(emb_mat)),np.mean(np.absolute(emb_mat)),))
# print('layer: ',layer,np.max(emb_mat),np.min(emb_mat),np.max(np.absolute(emb_mat)),np.mean(np.absolute(emb_mat)),np.median(np.absolute(emb_mat)))
dims=get_top_dim_from_file(file=dim_file)
top_emb_mat = emb_mat[dims[0:k]]
top_emb_mat = top_emb_mat[:,dims[0:k]]
# print('layer: ',layer,np.max(top_emb_mat),np.min(top_emb_mat),np.max(np.absolute(top_emb_mat)),np.mean(np.absolute(top_emb_mat)),np.median(np.absolute(top_emb_mat)))
# print(top_emb_mat)
# +
for layer in [1,2,3,4,5,6,7,8,9,10,11,12]:
k=5
# emb_mat = np.absolute(corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int))
emb_mat = np.array(corr_map_layer_dict[layer]['corr_mat_whole'][50:,50:]-np.eye(768, dtype=int))
dim_file = '../SA-dim-files/average_corr_dim_layer_{}.tsv'.format(layer)
# print('layer: ',layer,np.max(emb_mat),np.min(emb_mat),np.max(np.absolute(emb_mat)),np.mean(np.absolute(emb_mat)),np.median(np.absolute(emb_mat)))
dims=get_top_dim_from_file(file=dim_file)
top_emb_mat = emb_mat[dims[0:k]]
top_emb_mat = top_emb_mat[:,dims[0:k]]
print('layer\t{}\tmax-corr\t{}\tmean-corr\t{} '.format(layer,np.max(np.absolute(top_emb_mat)),np.mean(np.absolute(top_emb_mat)),))
# print('layer: ',layer,np.max(top_emb_mat),np.min(top_emb_mat),np.max(np.absolute(top_emb_mat)),np.mean(np.absolute(top_emb_mat)),np.median(np.absolute(top_emb_mat)))
# print(top_emb_mat)
# -
# # 0406
#
# we want to follow 'Identifying and controlling important neurons in neural machine translation'. find the relationship of different dim from layers.
corr_map_layer_dict[1]['all_embeddings'].shape
corr_mat_layer = np.corrcoef(corr_map_layer_dict[1]['all_embeddings'].T,corr_map_layer_dict[2]['all_embeddings'].T)
corr_mat_layer=corr_mat_layer[0:768,768:]
corr_mat_layer
corr_mat_layer.shape
corr_mat_layer=np.absolute(corr_mat_layer)
np.max(corr_mat_layer,axis=1)
np.min(corr_mat_layer),np.max(corr_mat_layer)
# ## build a MaxCorr for each layer
# +
def get_MaxCorr_score_for_one_layer(corr_map_layer_dict,cur_layer):
# 1. get the corr_mat for cur_layer and 11 other layers
layers = [1,2,3,4,5,6,7,8,9,10,11,12]
layers.remove(cur_layer)
corr_mat_layers={}
for layer in layers:
corr_mat_layer = np.corrcoef(corr_map_layer_dict[cur_layer]['all_embeddings'].T,corr_map_layer_dict[layer]['all_embeddings'].T)
corr_mat_layer=np.absolute(corr_mat_layer[0:768,768:])
corr_mat_layers[layer] = corr_mat_layer
# 2. get the MaxCorr score
all_corr_mat=[np.max(corr_mat_layers[layer],axis=1) for layer in layers]
all_corr_mat = np.array(all_corr_mat)
# we first get the max for one layer, and find the max over layers
MaxCorr_sort=np.argsort(np.max(all_corr_mat,axis=0))[::-1]
# we first get the max for one layer, and find the min over layers
MinCorr_sort=np.argsort(np.min(all_corr_mat,axis=0))[::-1]
mean_corr_mat=corr_mat_layers[layers[0]]
for layer in layers[1:]:
mean_corr_mat+=corr_mat_layers[layer]
# we first get the mean all layer, and find the max over layers
MeanCorr_sort=np.argsort(np.max(mean_corr_mat,axis=1))[::-1]
return corr_mat_layers,MaxCorr_sort,MinCorr_sort,MeanCorr_sort
def write_file(cur_layer,MaxCorr_sort,MinCorr_sort,MeanCorr_sort):
with open('../SA-dim-files/Bau_MaxCorr_dim_layer_{}.tsv'.format(cur_layer),'w') as fout:
for dim in MaxCorr_sort:
fout.write('{} '.format(dim))
with open('../SA-dim-files/Bau_MinCorr_dim_layer_{}.tsv'.format(cur_layer),'w') as fout:
for dim in MinCorr_sort:
fout.write('{} '.format(dim))
with open('../SA-dim-files/Bau_MeanCorr_dim_layer_{}.tsv'.format(cur_layer),'w') as fout:
for dim in MeanCorr_sort:
fout.write('{} '.format(dim))
# -
for cur_layer in [1,2,3,4,5,6,7,8,9,10,11,12]:
corr_mat_layers,MaxCorr_sort,MinCorr_sort,MeanCorr_sort = get_MaxCorr_score_for_one_layer(corr_map_layer_dict,cur_layer)
write_file(cur_layer,MaxCorr_sort,MinCorr_sort,MeanCorr_sort)
print('finish',cur_layer)
|
control-tasks/get_cor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Samir4p/Estudos_Ciencia_Dados/blob/main/Enviar_Email.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9iuK2xbqJaT4"
import json
from email.mime.multipart import MIMEMultipart
from email.mime.text import MIMEText
import smtplib
# + id="LBfXFp8XNsJs"
#Corpo da mensagem do email
msg = MIMEMultipart()
message = "Você recebeu um email da Start4p :)"
password = "<PASSWORD>" ####Senha do email que vai enviar a msg
msg['From'] = "xxxxxxxxp@xxxx" ###Endereço email que vai enviar
msg['To'] = "xxxxxxxxxxxxxxx" ###Email que ira receber
msg['Subject'] = "Enviando gmail com Python" ###Assunto do email
# + id="Uzf3k6m4J2Hu"
#Monta conexao envia email
msg.attach(MIMEText(message, 'plain'))
server = smtplib.SMTP('smtp.gmail.com', port=587)
server.starttls()
server.login(msg['From'], password)
server.sendmail(msg['From'], msg['To'], msg.as_string())
server.quit()
|
Enviar_Email.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data tests
#
# The four dimensions are x, y, z, and time
# + cell_style="center"
import numpy as np
import nibabel as nib
import matplotlib.pyplot as plt
# -
brain = nib.load('./pet_data/sub-OAS30100/ses-d0158/pet/sub-OAS30100_ses-d0158_acq-PIB_pet.nii.gz')
brain_data = brain.get_fdata()
brain_data.shape
# + cell_style="center"
print(brain.header)
# +
def show_slices(slices):
""" Function to display row of image slices """
fig, axes = plt.subplots(1, 1, figsize=(10, 10))
axes.imshow(slices[0], cmap="jet", origin="lower")
slices = []
slices.append(brain_data[:, :, 8, 20])
show_slices(slices)
np.amax(slices)
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
fig = plt.figure()
# ims is a list of lists, each row is a list of artists to draw in the
# current frame; here we are just animating one artist, the image, in
# each frame
ims = []
for i in range(brain_data.shape[2]):
im = plt.imshow(brain_data[:, :, i, 20], cmap="jet", origin="lower", animated=True)
ims.append([im])
ani = animation.ArtistAnimation(fig, ims, interval=50, blit=True,
repeat_delay=1000)
ani.save('pet_demo.mp4')
# -
|
get_data_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GDP Per Capita Analysis
# GDP per capita data was taken from world bank
# https://data.worldbank.org/indicator/NY.GDP.PCAP.CD
import pandas as pd
# read "gdp_per_capita.csv" file into a dataframe
data=pd.read_csv("gdp_per_capita_2020July.csv",skiprows=4)
# check the top-five rows the data frame
data.head()
#set index of the dataframe as Country name
data.set_index("Country Name",inplace=True)
#check again top-five rows of the dataframe
data.head()
# ### How to delete a column?
# delete Country Code, Indicator Name and Indicator Code columns
data.drop(labels=["Country Code","Indicator Name","Indicator Code","Unnamed: 64"],axis="columns",inplace=True)
# check the dataframe
data.head()
# ### get data for Italy
italy=data.loc["Italy"]
italy
type(italy)
# get italy's gdp per capita for 2019
italy["2019"]
# ### Find the year that italy has the highest increase of gdb per capita
# - Calculate the yearly percentage increase compared to previous year
# - find the year that has highest increase in terms of percentage.
italy
# calculate the yearly change in percentage
italy_pct_change=italy.pct_change()
# multiply italy_pct_change by 100
italy_pct_change=italy_pct_change*100
italy_pct_change
# find the year that has highest change
italy_pct_change.idxmax()
# ### Find the years that GDP per capita decreased compared to the previous year.
italy_negative=italy_pct_change[italy_pct_change>0]
# get only years
italy_negative.index
# ## Find the year that GDP per capita of the world increased the most compared to previous year
data.head()
# first get the data for world
# there is an index named World
world=data.loc["World"]
# now find the year that has highest increase in percentange
world.pct_change().idxmax()
# ## find top 10 years that have highest standart deviation
# first calculate standart deviation for each year
data_std=data.std()
data_std
# now top-10 years having the hightest standart deviation
data_std.nlargest(10)
# ### Which country has the highest gdp_per_capita in 2019?
data.head()
# get the data for the year 2019
data_2019=data["2019"]
data_2019
# now get the country
data_2019.idxmax()
data_2019.nlargest(1)
# # Rank of Luxembourg
# find the rank of Luxembourg each year
# calculate the rank of each country for each year
data_rank=data.rank(ascending=False)
data_rank
# now get the rank of Luxembourg
lux_rank=data_rank.loc["Luxembourg"]
lux_rank
# plot the rank of luxembourg
lux_rank.plot()
# print the years where Luxembourg ranks top
lux_rank[lux_rank==1]
|
Data Analysis with Python/pandas_workshop-master/gdp_per_capita/gdp_per_capita_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Sample 6.2 Multi-parameter Bayesian model: Globular cluster membership
# +
# %matplotlib inline
# #%pylab
#posteior distribution of normal data
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
import astropy.io.fits as fits
import matplotlib
matplotlib.rc('xtick', labelsize=12)
matplotlib.rc('ytick', labelsize=12)
from astropy.table import Table
import emcee
import corner
gc = Table.read("N6205.vot",format="votable")
# -
print(gc[0])
ind_gc = (gc['parallax_over_error']>3.) &\
( gc['parallax']>-0.03) & ( gc['parallax']<0.2)
print(np.sum(ind_gc))
ind_fld = (gc['parallax_over_error']>3.) &\
( gc['parallax']>-0.03) & ( gc['parallax']>=0.2)
print(np.sum(ind_fld))
# +
fig = plt.figure(figsize=(12,5))
ax = fig.add_subplot(121)
ax.plot(gc['ra'][ind_gc],gc['dec'][ind_gc],'k.',markersize=2,alpha=1)
ax.plot(gc['ra'][ind_fld],gc['dec'][ind_fld],'r.',markersize=1,alpha=0.3)
ax.set_xlabel('RA')
ax.set_ylabel('DEC')
ax = fig.add_subplot(122)
ax.plot(gc['pmra'][ind_gc],gc['pmdec'][ind_gc],'k.',markersize=2,alpha=1)
ax.plot(gc['pmra'][ind_fld],gc['pmdec'][ind_fld],'r.',markersize=1,alpha=1)
ax.set_xlim(-14,10)
ax.set_ylim(-14,10)
ax.set_xlabel('pm_RA')
ax.set_ylabel('pm_DE')
# -
ind_gc2 = ind_gc & (gc['pmra']>-4) & (gc['pmra']<-2.5) & (gc['pmdec']>-3) & (gc['pmdec']<-1.7)
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
h,xedge = np.histogram(gc['parallax'][ind_gc2 | ind_fld],np.arange(0,5,0.01))
ax.step(xedge[1:],h,where='post')
ax.set_xlim(0,1)
ax.set_xlabel('parallax')
# +
fig = plt.figure(figsize=(10,15))
ax = fig.add_subplot(211)
ax.plot(gc['ra'][ind_gc2],gc['dec'][ind_gc2],'k.',markersize=1)
#ax.plot(gc['ra'][ind_fld],gc['dec'][ind_fld],'r.',markersize=1,alpha=0.3)
ax.set_xlabel('RA')
ax.set_ylabel('DEC')
ax = fig.add_subplot(212)
ax.plot(gc['pmra'][ind_gc2],gc['pmdec'][ind_gc2],'k.',markersize=1)
#ax.plot(gc['pmra'][ind_fld],gc['pmdec'][ind_fld],'r.',markersize=1,alpha=0.3)
# ax.set_xlim(-4,-1)
# ax.set_ylim(-4,-1)
ax.set_xlabel(r'$\mu_{RA}$')
ax.set_ylabel(r'$\mu_{DEC}$')
# -
# ## estimate the mean proper motions of N6205
# ## $p(\mu,\Sigma|y)\propto p(y|\mu,\Sigma)p(\mu)p(\sigma_\alpha)p(\sigma_\delta)=\frac{\exp(-{\mu}^T\Sigma^{-1}{\mu})}{2\pi\sigma_\alpha^3\sigma_\delta^3\sqrt{1-\rho^2}}$
y = np.concatenate((gc['pmra'][ind_gc2].data.reshape(np.sum(ind_gc2),1),\
gc['pmdec'][ind_gc2].data.reshape(np.sum(ind_gc2),1)),axis=1)
print(y.shape)
print('mean:',np.mean(y,axis=0))
print('std.:',np.std(y,axis=0))
plt.plot(y[:,0],y[:,1],'k.')
# +
def posterior1(theta,y):
'''
without 1/sigma**2 prior
'''
x = np.zeros_like(y)
x[:,0] = y[:,0]-theta[0]
x[:,1] = y[:,1]-theta[1]
#print(np.mean(y,axis=0),np.mean(x,axis=0))
s1 = theta[2]
s2 = theta[3]
rho = theta[4]
if np.abs(rho)>1 or s1<=0 or s2<=0 :
lnp = -1e50
else:
lnp = np.sum(-1./(2.*(1-rho**2))*(x[:,0]**2/s1**2-2*rho*x[:,0]*x[:,1]/(s1*s2)+x[:,1]**2/s2**2))-\
(y.shape[0])*np.log(s1)-(y.shape[0])*np.log(s2)-0.5*y.shape[0]*np.log(1-rho**2)
return lnp
def posterior2(theta,y):
'''
with 1/sigma**2 prior
'''
mu = np.zeros_like(y)
mu[:,0] = y[:,0]-theta[0]
mu[:,1] = y[:,1]-theta[1]
s1 = theta[2]
s2 = theta[3]
rho = theta[4]
if np.abs(rho)>1 or s1<=0 or s2<=0:
lnp = -1e50
else:
lnp = np.sum(-1./(2.*(1-rho**2))*(mu[:,0]**2/s1**2-2*rho*mu[:,0]*mu[:,1]/(s1*s2)+mu[:,1]**2/s2**2))-\
(2.+y.shape[0])*np.log(s1)-(2.+y.shape[0])*np.log(s2)-0.5*y.shape[0]*np.log(1-rho**2)
return lnp
def mcmc_run(y, p0, lnlikeli = posterior1, ndim = 2, nwalkers = 50):
sampler = emcee.EnsembleSampler(nwalkers, \
ndim, lnlikeli, \
args=[y])
pos, prob, state = sampler.run_mcmc(p0, 100)
sampler.reset()
sampler.run_mcmc(pos, 1000)
samples = sampler.chain[:, :, :].reshape((-1, ndim))
return samples
# -
## always test your ln posterior before MCMC
theta1= [-2.5,-2.5,0.2,0.3,0.]
print(posterior1(theta1,y),posterior2(theta1,y))
theta2= [-0.7,-0.1,0.2,0.3,0.]
print(posterior1(theta2,y),posterior2(theta2,y))
# +
## using posterior1 without 1/sigma**2 prior
nwalkers = 50
ndim = 5
p0=np.zeros((nwalkers,ndim))
p0[:,0] = np.random.rand(nwalkers)-3.
p0[:,1] = np.random.rand(nwalkers)-3.
p0[:,2] = np.random.rand(nwalkers)*0.3
p0[:,3] = np.random.rand(nwalkers)*0.3
p0[:,4] = np.random.rand(nwalkers)*2.-1.
samples1 = mcmc_run(y, p0, lnlikeli = posterior1, ndim = ndim, nwalkers = nwalkers)
fig = corner.corner(samples1,\
labels=[r"$\mu_\alpha$",r"$\mu_\delta$", r"$\sigma_\alpha$", r"$\sigma_\delta$",r"$\rho$"],\
quantiles=[0.16, 0.5, 0.84],\
show_titles=True, \
title_kwargs={"fontsize": 12})
# +
## using posterior2 with 1/sigma**2 prior
samples2 = mcmc_run(y, p0, lnlikeli = posterior2, ndim = ndim, nwalkers = nwalkers)
fig = corner.corner(samples2,\
labels=[r"$\mu_\alpha$",r"$\mu_\delta$", r"$\sigma_\alpha$", r"$\sigma_\delta$",r"$\rho$"],\
quantiles=[0.16, 0.5, 0.84],\
show_titles=True, \
title_kwargs={"fontsize": 12})
# -
|
sample6.2_MultiParamModel_GCpropermotion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TASK 1
m = []
for i in range(2000,3201):
if(i%7==0 and i%5!=0):
m.append(i)
print(m)
# # TASK 2
first_name = input()
last_name = input()
name = list([first_name , " " , last_name])
reverse_name = name[::-1]
reverse_name = reverse_name[0] + reverse_name[1] + reverse_name[2]
print(reverse_name)
# # TASK 3
d = 12
r = d/2
v = (4/3)*3.1416*r**3
print(v)
|
python assignment 1 ineuron.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistical Measures of Performance
#
# This notebook is intended to provide a brief introduction to statistical measures of performance (accuracy, precision, recall (sensitivity) and specificity).
#
# These four statistical Measures are key to get a good summary of your results as they give insights based on four indicators:<br>
# - True Positive (TP): Your result says "True" and your reference says "True".
# - True Negative (TN): Your result says "False" and your reference says "False".
# - False Positive (FP): Your result says "True" and your reference says "False".
# - False Negative (FN): Your result says "False" and your reference says "True".
#
# It's important to note these statistical measures are for binary classification (as the example above - True/False). If you want to use them in a multiclass problem it is possible but you have to take one class and leave the rest of the classes as if they were in one group (further explanation and example shown later in the text).
# ## Sensitivity / recall
#
# ### Definition
# It is the ratio of how much was classified as Positive and how much must have been Positive.<br>
# Sensitivity let you know the ratio of positives, it is specially useful in applications in which a True is crucial and prefered over a False result. As an example, in safety functions for autonomous driving if you need to alert in certain circumstances it is prefered to get an incorrect alert that produces safety instead of not having it and needing it. High sensitivity is crucial for safety functions.
#
#
# ### Formula / Procedure to find it
#
# The equation to compute the sensitivity is as follows: $Sensitivity = \frac{TP}{TP + FN}$ <br>
# ### Visualization
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.graph_objects as go
from plotly.subplots import make_subplots
# ### Creating the data set
data_set = [5, 2, 6, 10, 9, 3, 1, 9, 6, 2, 1, 4, 6, 5, 0, 4, 6,
5, 4, 6, 3, 0, 2, 8, 6, 7, 8, 0, 4, 7, 0, 1, 0, 8,
2, 0, 10, 2, 6, 6, 0, 5, 2, 0, 10, 3, 9, 8, 4, 7, 6,
1, 10, 7, 10, 3, 0, 6, 5, 8, 4, 3, 7, 3, 1, 5, 3, 0,
3, 3, 1, 2, 1, 5, 0, 5, 8, 1, 10, 7, 8, 6, 9, 3, 3,
7, 3, 4, 10, 8, 2, 0, 0, 2, 9, 0, 5, 5, 5, 6]
# Imagine that we what an alert every time we get a datapoint equal to 5 or a greater value.<br>
# For sensitivity comparisson we create two filters (one incorrect as it doesn't take into account the value 5 only greater numbers and one correct).
# +
def sensitivity(actual, predicted, verbosity=True):
true_positives = np.sum([a is p for a,p in zip(actual,predicted) if a])
false_negatives = np.sum([a is not p for a,p in zip(actual,predicted) if a])
sens = true_positives/(true_positives+false_negatives)
if verbosity:
print(f"True positives: {true_positives}\nFalse negatives: {false_negatives}\nSensitivity: {sens}\n")
return sens
def alert_trigger_non_inclusive(data):
return np.array([d>5 for d in data])
def alert_trigger_inclusive(data):
return np.array([d>=5 for d in data])
# -
actual = alert_trigger_inclusive(data_set)
pred = alert_trigger_non_inclusive(data_set)
sensitivity(actual, pred)
# We got a sensitivity of 78%
# It was really bad only by taking out one limit testing condition.
# +
def alert_trigger_inclusive_threshold(data, thresh):
return np.array([d>=thresh for d in data])
def threshold(min_t, max_t, data, func):
df = pd.DataFrame()
measure = list()
current_thresh = list()
actual = alert_trigger_inclusive(data)
for t in range(min_t,max_t):
pred = alert_trigger_inclusive_threshold(data, t)
measure.append(func(actual, pred, verbosity=False))
current_thresh.append(t)
df['func'] = measure
df['threshold'] = current_thresh
return df
# +
df_sens = threshold(0, 10, data_set, sensitivity)
fig = go.Figure(data=go.Scatter(x=df_sens['threshold'],
y=df_sens["func"],
text=[f"Threshold: {_}" for _ in df_sens["threshold"]],
mode='markers+lines'))
fig.update_layout(title_text="Behaviour of Sensitivity with different thresholds",
xaxis=dict(title="Threshold"),
yaxis=dict(title="Sensitivity")
)
fig.show()
# -
# You have to take sensitivity into account if your application is critical and needs True results (prefered).
# ## Specificity
#
# ### Definition
# Specificity is the opposite of sensitivity. It is the ratio of correct classification as negative and all the samples that are indeed negative.
#
# ### Formula / Procedure to find it
#
# The equation to compute the specificity is as follows: $Specificity = \frac{TN}{FP + TN}$ <br>
# ### Creating the data set
# We will be using the same dataset (previously created).
def specificity(actual, predicted, verbosity=True):
true_negatives = np.sum([a is p for a,p in zip(actual,predicted) if not a])
false_positives = np.sum([a is not p for a,p in zip(actual,predicted) if not a])
spec = true_negatives/(true_negatives+false_positives)
if verbosity:
print(f"True negatives: {true_negatives}\nFalse positives: {false_positives}\nSpecificity: {spec}\n")
return spec
actual = alert_trigger_inclusive(data_set)
pred = alert_trigger_non_inclusive(data_set)
specificity(actual, pred)
# +
df_spec = threshold(0, 10, data_set, specificity)
fig = go.Figure(data=go.Scatter(x=df_spec['threshold'],
y=df_spec["func"],
text=[f"Threshold: {_}" for _ in df_sens["threshold"]],
mode='markers+lines'))
fig.update_layout(title_text="Behaviour of Specificity with different thresholds",
xaxis=dict(title="Threshold"),
yaxis=dict(title="Specificity")
)
fig.show()
# -
# As we can see, this classification gets a better score classifing negatives correctly that positives. Getting both is desired and we need a balance between sensitiviy and specificity to get a good classifier but depending on the application you can select the metric needed.<br>
# We can get a really useful visualization having these two metrics (Sensitivity and Specificity): The ROC curve (Receiver Operating Characteristic)<br>
# But we have to modify our classifier to receive thresholds in the classification.
# +
def alert_trigger_inclusive_threshold(data, thresh):
return np.array([d>=thresh for d in data])
def threshold_roc(min_t, max_t, data):
df = pd.DataFrame()
sen_t = list()
spe_t = list()
current_thresh = list()
actual = alert_trigger_inclusive(data)
for t in range(min_t,max_t):
pred = alert_trigger_inclusive_threshold(data, t)
sen_t.append(sensitivity(actual, pred, verbosity=False))
spe_t.append(specificity(actual, pred, verbosity=False))
current_thresh.append(t)
df['sensitivity'] = sen_t
df['specificity'] = spe_t
df['threshold'] = current_thresh
return df
# +
df = threshold_roc(0, 11, data_set)
fig = go.Figure(data=go.Scatter(x=1-df['specificity'],
y=df["sensitivity"],
text=[f"Threshold: {_}" for _ in df["threshold"]],
mode='markers+lines'))
fig.update_layout(title_text="ROC for integers threshold classification [0,10]",
xaxis=dict(title="1 - Specificity"),
yaxis=dict(title="Sensitivity")
)
fig.show()
# -
# As you can see en the plot, while we get closer to the threshold of 5 (the one we know is the right one) we tend to get better results for sensitivity and specificity, reaching the maximum score at threshold 5 with a perfect score of both measures.<br>
# In this case we got a perfect score, in real applications we might determine how good our classifier is by determining the best combination of sensitivity and specificity if we don't get a perfect score.
# You have to take specificity into account if your application is critical and needs False results (prefered).
# ## Precision
#
# ### Definition
# The precision can be seen as the percentage of real True scenarios given the total amounts the classifier considers as True. In other words we can determine in what percentage we can trust the classifier when it gives a True statement.<br>
# It is the ratio of True Positives and the sum of True Positives and False Positives.
#
# ### Formula / Procedure to find it
#
# The equation to compute the precision is as follows: $Precision = \frac{TP}{TP + FP}$ <br>
def precision(actual, predicted, verbosity=True):
true_positives = np.sum([a is p for a,p in zip(actual,predicted) if a])
false_positives = np.sum([a is not p for a,p in zip(actual,predicted) if not a])
prec = true_positives/(true_positives+false_positives)
if verbosity:
print(f"True positives: {true_positives}\nFalse positives: {false_positives}\nPrecision: {prec}\n")
return prec
actual = alert_trigger_inclusive(data_set)
pred = alert_trigger_non_inclusive(data_set)
precision(actual, pred)
# +
df_prec = threshold(0, 10, data_set, precision)
fig = go.Figure(data=go.Scatter(x=df_prec['threshold'],
y=df_prec["func"],
text=[f"Threshold: {_}" for _ in df_sens["threshold"]],
mode='markers+lines'))
fig.update_layout(title_text="Behaviour of Precision with different thresholds",
xaxis=dict(title="Threshold"),
yaxis=dict(title="Precision")
)
fig.show()
# -
# You can get precision to know how much you can trust in a True result from your classifier.
# ## Accuracy
#
# ### Definition
# The accuracy gives an overall hint of how good the classification is performed, you have to take all the correct classifications (Trues or Falses) and divide by the number of samples.
#
# ### Formula / Procedure to find it
#
# The equation to compute the accuracy is as follows: $Accuracy = \frac{TP + TN}{TP + TN + FP + FN}$ <br>
def accuracy(actual, predicted, verbosity=True):
true_positives = np.sum([a is p for a,p in zip(actual,predicted) if a])
true_negatives = np.sum([a is p for a,p in zip(actual,predicted) if not a])
false_positives = np.sum([a is not p for a,p in zip(actual,predicted) if not a])
false_negatives = np.sum([a is not p for a,p in zip(actual,predicted) if a])
acc = (true_positives + true_negatives)/(true_positives+true_negatives+false_positives+false_negatives)
if verbosity:
print(f"True positives: {true_positives}\nTrue negatives: {true_negatives}\n",
f"False positives: {false_positives}\nFalse negatives: {false_negatives}\nAccuracy: {acc}\n")
return acc
actual = alert_trigger_inclusive(data_set)
pred = alert_trigger_non_inclusive(data_set)
accuracy(actual, pred)
# +
df_acc = threshold(0, 10, data_set, accuracy)
fig = go.Figure(data=go.Scatter(x=df_acc['threshold'],
y=df_acc["func"],
text=[f"Threshold: {_}" for _ in df_sens["threshold"]],
mode='markers+lines'))
fig.update_layout(title_text="Behaviour of Accuracy with different thresholds",
xaxis=dict(title="Threshold"),
yaxis=dict(title="Accuracy")
)
fig.show()
# -
# You can observe that Accuracy is not like the other metrics, you get a peak in the right threshold and the it goes down again.<br>
# In conclusion, with accuracy you can get a better idea of the result. If want more details you can use sensitivity and specificity to get the ROC curve and also determine the best threshold that suits your needs.
# ## Excercise
#
# You can test your learning of the introduction of central tendency measures next:
class test:
def __init__(self):
self.questions = list()
self.answers = list()
self.correct_answers = 0
self.score = 0
def add_element(self, q, a):
self.questions.append(q)
self.answers.append(a)
def remove_element(self, index):
self.questions.pop(index)
self.answers.pop(index)
def show_answer(self, index):
print(f"Q{index}: {self.questions[index-1]} - Ans_{index}: {self.answers[index-1]}")
def show_answers(self):
for index, (q, a) in enumerate(zip(self.questions, self.answers)):
print(f"Q{index+1}: {q} - Ans_{index+1}: {a}")
def build_from_csv(self, filename):
df = pd.read_csv(filename)
for index in range(df.shape[0]):
self.add_element(df['Questions'][index], df['Answers'][index])
def visualize_score(self):
fig = go.Figure(data=[go.Pie(labels=["Correct", "Incorrect"],
values=[self.score, 100-self.score],
marker_colors=['rgb(10,100,10)', 'rgb(230,70,70)'],
hole=.3)])
fig.show()
def test(self):
self.correct_answers = 0
for index, (q, a) in enumerate(zip(self.questions, self.answers)):
current_answer = ''
while len(str(current_answer))==0:
current_answer = input(f"Q{index+1}: " + q)
if len(current_answer)>0:
current_answer = np.round(float(current_answer),2)
self.correct_answers += int(current_answer == a)
if a==current_answer:
print("Correct")
else:
print("Incorrect")
self.score = 100*np.sum(self.correct_answers)/len(self.questions)
print(f"Your score: {self.score}")
self.visualize_score()
exam = test()
exam.build_from_csv("https://raw.githubusercontent.com/Ricardo-DG/data_analytics_training/main/measures_test.csv")
# +
# If you would like to see the answers uncomment and run the following line
# exam.show_answers()
# +
# If you would like to see a specific answer uncomment and run the following line
# (make sure to replace "index" with the number of the question you want to know the answer).
# exam.show_answer(index)
# -
score = exam.test()
|
measures_testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 간단한 모델 훈련하기
#
# 데이터셋을 수집, 정제, 포맷팅하고 훈련 세트와 테스트 세트로 분할했습니다. 이전 [노트북](https://github.com/rickiepark/ml-powered-applications/blob/master/notebooks/exploring_data_to_generate_features.ipynb)에서 벡터 특성을 만들었습니다. 이제 첫 번째 간단한 모델을 훈련해 보죠.
#
# 먼저 데이터를 로드하고 포맷팅합니다.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pathlib import Path
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, classification_report
import joblib
import sys
sys.path.append("..")
np.random.seed(35)
import warnings
warnings.filterwarnings('ignore')
from ml_editor.data_processing import (
format_raw_df,
add_text_features_to_df,
get_feature_vector_and_label,
get_split_by_author,
get_vectorized_inputs_and_label,
get_vectorized_series,
train_vectorizer,
)
data_path = Path('../data/writers.csv')
df = pd.read_csv(data_path)
df = format_raw_df(df.copy())
df = df.loc[df["is_question"]].copy()
# -
# 그다음 특성을 추가하고 벡터로 변환하고 훈련/테스트 세트로 나눕니다.
# +
df = add_text_features_to_df(df.copy())
train_df, test_df = get_split_by_author(df, test_size=0.2, random_state=40)
vectorizer = train_vectorizer(train_df)
train_df["vectors"] = get_vectorized_series(train_df["full_text"].copy(), vectorizer)
test_df["vectors"] = get_vectorized_series(test_df["full_text"].copy(), vectorizer)
# -
features = [
"action_verb_full",
"question_mark_full",
"text_len",
"language_question",
]
X_train, y_train = get_feature_vector_and_label(train_df, features)
X_test, y_test = get_feature_vector_and_label(test_df, features)
# 특성과 레이블이 준비되면 `sklearn`을 사용해 몇 줄의 코드로 모델을 훈련할 수 있습니다.
# +
clf = RandomForestClassifier(n_estimators=100, class_weight='balanced', oob_score=True)
clf.fit(X_train, y_train)
y_predicted = clf.predict(X_test)
y_predicted_proba = clf.predict_proba(X_test)
# -
y_train.value_counts()
# ## 측정 지표
#
# 모델을 훈련하고 나면 결과를 평가할 차례입니다. 단순 지표부터 시작해 보죠.
def get_metrics(y_test, y_predicted):
# 진짜 양성 / (진짜 양성 + 가짜 양성)
precision = precision_score(y_test, y_predicted, pos_label=None,
average='weighted')
# 진짜 양성 / (진짜 양성 + 가짜 음성)
recall = recall_score(y_test, y_predicted, pos_label=None,
average='weighted')
# 정밀도와 재현율의 조화 평균
f1 = f1_score(y_test, y_predicted, pos_label=None, average='weighted')
# 진짜 양성 + 진짜 음성 / 전체
accuracy = accuracy_score(y_test, y_predicted)
return accuracy, precision, recall, f1
# +
# 훈련 정확도
# https://datascience.stackexchange.com/questions/13151/randomforestclassifier-oob-scoring-method 참조
y_train_pred = np.argmax(clf.oob_decision_function_,axis=1)
accuracy, precision, recall, f1 = get_metrics(y_train, y_train_pred)
print("훈련 정확도 = %.3f, 정밀도 = %.3f, 재현율 = %.3f, f1 = %.3f" % (accuracy, precision, recall, f1))
# -
accuracy, precision, recall, f1 = get_metrics(y_test, y_predicted)
print("검증 정확도 = %.3f, 정밀도 = %.3f, 재현율 = %.3f, f1 = %.3f" % (accuracy, precision, recall, f1))
# 첫 번째 모델이 잘 동작하는 것 같습니다. 적어도 랜덤 예측보다는 성능이 낫기 때문에 첫 번째 시도로는 고무적입니다. 추후 분석과 사용을 위해 훈련된 모델과 벡터화 객체를 디스크에 저장합니다.
model_path = Path("../models/model_1.pkl")
vectorizer_path = Path("../models/vectorizer_1.pkl")
joblib.dump(clf, model_path)
joblib.dump(vectorizer, vectorizer_path)
# ## 추론 함수
#
#
# 훈련된 모델을 본 적 없는 데이터에서 사용하기 위해 추론 함수를 정의하고 사용합니다. 아래 함수는 임의의 질문을 받고 높은 점수를 받을 추정 확률을 출력합니다.
# +
from ml_editor.model_v1 import get_model_probabilities_for_input_texts
# The inference function expects an array of questions, so we created an array of length 1 to pass a single question
test_q = ["bad question"]
probs = get_model_probabilities_for_input_texts(test_q)
# Index 1 corresponds to the positive class here
print("이 질문이 양성 샘플일 확률: %s" % (probs[0][1]))
|
notebooks/train_simple_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from recohut.datasets import *
from recohut.layers import *
from recohut.models.layerss import *
from recohut.models import *
from recohut.utils import *
from recohut.rl import *
from recohut.visualization import *
# # recohut
# <div id="top"></div>
#
# [![Contributors][contributors-shield]][contributors-url]
# [![Forks][forks-shield]][forks-url]
# [![Stargazers][stars-shield]][stars-url]
# [![Issues][issues-shield]][issues-url]
# [![MIT License][license-shield]][license-url]
#
#
#
# <!-- PROJECT LOGO -->
# <br />
# <div align="center">
# <a href="https://github.com/RecoHut-Projects/recohut">
# <img src="https://github.com/recohut/reco-static/raw/master/media/diagrams/recohut_logo.svg" alt="Logo" width="80" height="80">
# </a>
#
# <!-- <h3 align="center">recohut</h3> -->
#
# <p align="center">
# a python library for building recommender systems.
# <br />
# <a href="https://recohut-projects.github.io/recohut"><strong>Explore the docs »</strong></a>
# <br />
# <br />
# <a href="https://github.com/RecoHut-Projects/recohut/tree/master/tutorials">View Demo</a>
# ·
# <a href="https://github.com/RecoHut-Projects/recohut/issues">Report Bug</a>
# ·
# <a href="https://github.com/RecoHut-Projects/recohut/issues">Request Feature</a>
# </p>
# </div>
#
#
#
# ## About The Project
#
# <img src="https://github.com/recohut/reco-static/raw/master/media/diagrams/recohut_lib_main.svg">
#
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# ### Built With
#
# * [Python](https://www.python.org/)
# * [PyTorch](https://pytorch.org/)
# * [Lightning](https://www.pytorchlightning.ai/)
# * [nbdev](https://github.com/fastai/nbdev)
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- GETTING STARTED -->
# ## Getting Started
#
# To get a local copy up and running follow these simple example steps.
#
# ### Prerequisites
#
# * pytorch
# ```sh
# pip install torch
# ```
# * lightning
# ```sh
# pip install pytorch-lightning
# ```
#
# ### Installation
#
# ```
# pip install recohut
# ```
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- USAGE EXAMPLES -->
# ## Usage
#
# ```python
# # import the required modules
# from recohut.datasets.movielens import ML1mDataModule
# from recohut.models.nmf import NMF
# from recohut.trainers.pl_trainer import pl_trainer
#
# # build the dataset
# class Args:
# def __init__(self):
# self.data_dir = '/content/data'
# self.min_rating = 4
# self.num_negative_samples = 99
# self.min_uc = 5
# self.min_sc = 5
# self.val_p = 0.2
# self.test_p = 0.2
# self.num_workers = 2
# self.normalize = False
# self.batch_size = 32
# self.seed = 42
# self.shuffle = True
# self.pin_memory = True
# self.drop_last = False
# self.split_type = 'stratified'
#
# args = Args()
#
# ds = ML1mDataModule(**args.__dict__)
# ds.prepare_data()
#
# # build the model
# model = NMF(n_items=ds.data.num_items, n_users=ds.data.num_users, embedding_dim=20)
#
# # train and evaluate the matrix factorization model
# pl_trainer(model, ds, max_epochs=5)
# ```
# Check [this](https://github.com/recohut/notebooks/blob/main/nbs/recohut_quick_tutorial.ipynb) quick tutorial.
#
# _For more examples, please refer to the [Documentation](https://recohut-projects.github.io/recohut) and [Tutorials](https://github.com/RecoHut-Projects/recohut/tree/master/tutorials)._
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- ROADMAP -->
# ## Roadmap
#
# - [] RecSys Model Deployment and MLOps features
# - [] RL agents and environment specific to recommender systems
# - [] Visualization utilities and EDA
#
# See the [open issues](https://github.com/RecoHut-Projects/recohut/issues) for a full list of proposed features (and known issues).
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- CONTRIBUTING -->
# ## Contributing
#
# Contributions are what make the open source community such an amazing place to learn, inspire, and create. Any contributions you make are **greatly appreciated**.
#
# If you have a suggestion that would make this better, please fork the repo and create a pull request. You can also simply open an issue with the tag "enhancement".
# Don't forget to give the project a star! Thanks again!
#
# 1. Fork the Project
# 2. Create your Feature Branch (`git checkout -b feature/AmazingFeature`)
# 3. Commit your Changes (`git commit -m 'Add some AmazingFeature'`)
# 4. Push to the Branch (`git push origin feature/AmazingFeature`)
# 5. Open a Pull Request
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- LICENSE -->
# ## License
#
# Distributed under the MIT License. See `LICENSE.txt` for more information.
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- CONTACT -->
# ## Contact
#
# <NAME>.
#
# [@sparsh-ai](https://github.com/RecoHut-Projects/recohut)
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- ACKNOWLEDGMENTS -->
# ## Acknowledgments
#
# * [nbdev team](https://nbdev.fast.ai/tutorial.html) for providing supporting tools to build this library.
# * [colab team](https://colab.research.google.com/) for providing running VMs instances for development and testing.
#
# <p align="right">(<a href="#top">back to top</a>)</p>
#
#
#
# <!-- MARKDOWN LINKS & IMAGES -->
# [contributors-shield]: https://img.shields.io/github/contributors/RecoHut-Projects/recohut.svg?style=for-the-badge
# [contributors-url]: https://github.com/RecoHut-Projects/recohut/graphs/contributors
# [forks-shield]: https://img.shields.io/github/forks/RecoHut-Projects/recohut.svg?style=for-the-badge
# [forks-url]: https://github.com/RecoHut-Projects/recohut/network/members
# [stars-shield]: https://img.shields.io/github/stars/RecoHut-Projects/recohut.svg?style=for-the-badge
# [stars-url]: https://github.com/RecoHut-Projects/recohut/stargazers
# [issues-shield]: https://img.shields.io/github/issues/RecoHut-Projects/recohut.svg?style=for-the-badge
# [issues-url]: https://github.com/RecoHut-Projects/recohut/issues
# [license-shield]: https://img.shields.io/github/license/RecoHut-Projects/recohut.svg?style=for-the-badge
# [license-url]: https://github.com/RecoHut-Projects/recohut/blob/master/LICENSE.txt
# [product-screenshot]: https://github.com/recohut/reco-static/raw/master/media/diagrams/recohut_lib_main.svg
|
nbs/index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from solana.rpc.api import Client
solana_client = Client("https://api.devnet.solana.com")
solana_client.is_connected()
# -
from solana.publickey import PublicKey
solana_client.get_balance(PublicKey(1))
solana_client.get_block_commitment(5)
solana_client.get_block_time(5)
solana_client.get_cluster_nodes()
solana_client.get_confirmed_block(1)
solana_client.get_confirmed_block(1, encoding="base64")
solana_client.get_confirmed_blocks(5, 10)
solana_client.get_signatures_for_address("Vote111111111111111111111111111111111111111", limit=1)
solana_client.get_epoch_info()
solana_client.get_epoch_schedule()
solana_client.get_fee_calculator_for_blockhash("BaQSR194dC4dZaRxATtxYyEwDkk7VgqUY8NVNkub8HFZ")
solana_client.get_recent_blockhash()
solana_client.get_fee_rate_governor()
solana_client.get_slot()
solana_client.get_fees()
solana_client.get_first_available_block()
solana_client.get_genesis_hash()
solana_client.get_identity()
solana_client.get_inflation_governor()
solana_client.get_inflation_rate()
solana_client.get_largest_accounts()
solana_client.get_leader_schedule()
solana_client.get_minimum_balance_for_rent_exemption(50)
filter_opts = {"memcmp": {"offset": 4, "bytes": "3Mc6vR"}}
solana_client.get_program_accounts("<KEY>", data_size=17, filter_opts=filter_opts)
solana_client.get_recent_blockhash()
signatures = ["5VERv8NMvzbJMEkV8xnrLkEaWRtSz9CosKDYjCJjBRnbJLgp8uirBgmQpjKhoR4tjF3ZpRzrFmBV6UjKdiSZkQUW", "5j7s6NiJS3JAkvgkoc18WVAsiSaci2pxB2A6ueCJP4tprA2TFg9wSyTLeYouxPBJEMzJinENTkpA52YStRW5Dia7"]
solana_client.get_signature_statuses(signatures)
solana_client.get_slot_leader()
solana_client.get_stake_activation("CYRJWqiSjLitBAcRxPvWpgX3s5TvmN2SuRY3eEYypFvT", epoch=1)
solana_client.get_supply()
solana_client.get_token_account_balance("7fUAJdStEuGbc3sM84cKRL6yYaaSstyLSU4ve5oovLS7")
solana_client.get_transaction_count()
solana_client.get_minimum_ledger_slot()
solana_client.get_version()
solana_client.request_airdrop(PublicKey(1), 10000)
solana_client.set_log_filter("solana_core=debug")
solana_client.get_vote_accounts()
solana_client.get_account_info(PublicKey(1))
solana_client.get_account_info(PublicKey(1), encoding="jsonParsed")
from solana.rpc.api import DataSlice
solana_client.get_account_info(PublicKey(1), data_slice=DataSlice(1,1))
|
notebooks/JSON RPC API.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
import random
import requests
import functools
limit = lambda x, a, b: a if (x < a) else b if (x > b) else x
num_marks = 25
marks = [limit(round(random.normalvariate(50, 15)), 0, 100) for _ in range(num_marks)]
marks = [43, 32, 38, 64, 62, 41, 20, 40, 52, 45, 57, 50, 60, 70, 68, 50, 38, 44, 55, 57, 45, 50, 44, 44, 66]
print(marks)
# +
num_games = 10
our_scores = [random.randint(0, 4) for _ in range(num_games)]
their_scores = [random.randint(0, 4) for _ in range(num_games)]
our_scores = [4, 4, 0, 4, 4, 0, 0, 4, 2, 4]
their_scores = [2, 3, 3, 2, 4, 0, 3, 0, 0, 1]
print('our scores:', our_scores)
print('their scores:', their_scores)
# -
python_sequence = """TTTACCAAAAACATAACCTTTAGCTAAAACCAGTATTAAAGGCAATGCCTGCCCAGTGAG
ACCTTCTTCAACGGCCGCGGTACCCTAACCGTGCAAAGGTAGCGTAATCACTTGTCTATT
AATTGTAGACCCGTATGAAAGGCCACATGAAAGTCAGACTGTCTCTTGTAATTAATCAAT
TAAACTGATCTTCCAGTACAAAAGCTGAAATGAACATATAAGACCAGAAGACCCTGTGAA
GCTTAAATTAACCTACTAAAACCCATAGTAGCTACTTTCAGTTGGGGCGACTTTGGAACA
AAACAAAACTTCCAAGCACCATGAGCTATCCCTCATACACCAGGCCAACAAGCCACCACA
AGACCCAGTAACACTGATAACCGAACCAAGTTACTCCAGGGATAACAGCGCCATCTTCTT
TAAGAGCCCATATCAAAAAGAAGGTTTACGACCTCGATGTTGGATCAGGACACCCAGGTG
GTGCAACCGCTACCAAAGGTTCGTTTGTTCAACGATTAACAGTCCCACGTGATCTGAGTT
CAGACCGGAGCAATCCAGGTCAGTTTCTATCTATAAAAAGCCCTTTCTAGTACGAAAGGA
CCGAAAGAGCAAAGCCAATACCAAAAGCACGCTTTAACAAAAAATATAAATAAACTCAAT"""
url_base = 'http://192.168.2.101:5000/'
def test_answer(func_name, *parameters):
correct = False
if func_name in globals():
try:
func = globals()[func_name]
answer = func(*parameters)
except:
pass
else:
try:
r = requests.post('http://localhost:5000/test_' + func_name, json=dict(parameters=parameters, answer=answer))
except requests.ConnectionError:
r = requests.post(url_base + 'test_' + func_name, json=dict(parameters=parameters, answer=answer))
if r.status_code == 200:
correct = r.json().get('correct', False)
return(correct)
test_top_marks = functools.partial(test_answer, 'top_marks')
test_gc_content = functools.partial(test_answer, 'gc_content')
test_team_points = functools.partial(test_answer, 'team_points')
test_top_marks(marks)
test_gc_content(python_sequence)
test_team_points(our_scores, their_scores)
# **Problem 1**: Write a function `top_marks(marks)` that takes as its single parameter a list of integers. This list will contain at least 5 marks. It should return the top 5 marks from this list. You can use the `marks` list in this notebook as sample input, the correct answer is `[73, 69, 68, 68, 68]`.
# **Problem 2**: The [GC content](https://en.wikipedia.org/wiki/GC-content) of a DNA molecule is the percentage of bases in the molecule that are guanine or cytosine bases. Write a function `gc_content(sequence)` that returns the integer GC content of the string `sequence`. You can test using the `python_sequence` string in this notebook for which the correct answer is `42`.
# **Problem 3:** You manage a soccer team. The scores from recent matches are in two lists, `our_scores` and `their_scores`. The lists have the same number of elements and the elements of the lists are matched, so e.g. `our_scores[0]` is the score your team achieved in the first game of the season, and `their scores[0]` is the score your opponents achieved in that same match. If your score is less than their score, you lose, and get zero points. If your score is greater than their score, you win, and get 2 points. If the scores are equal, you drew and get 1 point.
#
# Write a function `team_points(our_scores, their_scores)` that takes two list parameters and returns the number of points your team was awarded based on the two lists of scores. You can use the `our_scores` and `their_scores` lists as inputs to test with, in which case the number of points returned should be `14`.
# You can use the test functions above to test if your code is correct.
|
Day9ThreeProblems.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import htmltag as HT
import random
from htmltag import table, td, tr
# Constants
maxImgs = 20
def genImageID(begin,stop):
count = 0
listNum = []
start= begin
end = stop
while count < maxImgs:
num = random.randrange(start,end)
if num in listNum: continue
listNum.append(num)
count += 1
return listNum
imageID = genImageID(1,100)
links = ["http://pachy.cs.uic.edu:5000/api/image/src/"+str(i)+"/?resize_pix_w=500" for i in imageID[0:maxImgs]]
imgTags = []
radioShare = HT.input
for url in links:
imgTags.append(HT.img(src = url,alt = "Unavailable"))
# +
# logic to create the radio buttons
hiddenField = []
shareRadio = []
notShareRadio = []
for i in range(maxImgs):
hiddenField.append(HT.input(type='hidden',name=imageID[i],value=imageID[i]))
shareRadio.append(HT.input(type='radio',value='share',name=imageID[i]) + "Share")
notShareRadio.append(HT.input(type='radio',value='noShare',name=imageID[i]) + "Do Not Share")
# +
tdTags = []
for i in range(maxImgs):
tdTags.append(HT.td(HT.center(HT.HTML(hiddenField[i]),HT.HTML(shareRadio[i]),HT.HTML(notShareRadio[i])),HT.HTML(imgTags[i])))
trTags = []
for i in range(0,maxImgs,2):
trTags.append(HT.tr(HT.HTML(tdTags[i]),HT.HTML(tdTags[i+1])))
# -
bodyTxt = HT.table(HT.HTML(' \n'.join(trTags)),border="1")
print(bodyTxt)
# +
headFile = open("files/header.txt","r")
tailFile = open("files/tail.txt","r")
outputFile = open("files/sampleMTurk.question","w")
for line in headFile:
outputFile.write(line)
outputFile.write(bodyTxt)
for line in tailFile:
outputFile.write(line)
headFile.close()
tailFile.close()
outputFile.close()
# -
|
script/.ipynb_checkpoints/GenerateMTurkFileAPI-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational Autoencoder
#
# Variational Autoencoders (VAEs) are dimensionality reduction devices that code inputs into the latent parameters of a statistical process that supposedly gave rise to them. In practice, this is typically a normal distribution. The decoding involves sampling from the statistical distribution with a given latent parameter and chasing the result through a decoder network.
#
# The difference to classical autoencoders is that variational autoencoders map an input to a function (the distribution) and classical autoencoders map an input to a vector.
#
# c.f. this exellent post https://github.com/yaniv256/VAEs-in-Economics/blob/master/Notebooks/One_Dimensional_VAE_Workshop.ipynb
#
# which uses Chapter 8 in http://faculty.neu.edu.cn/yury/AAI/Textbook/Deep%20Learning%20with%20Python.pdf
#
# #### Despite all my rage, I am still just a rat in a cage, and this implementation continues to throw an out of memory error. I am done blaming myself. I blame tensorflow for being so unbelievably unstable.
# +
import tensorflow as tf
physical_devices = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
def gpu_test():
print("GPU test")
with tf.device('GPU:0'):
print("Executing basic TF graph on GPU:0")
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print("Success.")
print(c)
print("GPUs found:")
physical_devices = tf.config.experimental.list_physical_devices('GPU')
print('\n'.join('[%i]\t'%i +str(dev) for i,dev in enumerate(physical_devices)))
gpu_test()
# -
# from src.resourcelims import memory
#
# """
# Check GPU
# """
# from tensorflow.python.client import device_lib
# import tensorflow as tf
# from keras import backend
#
# #tf.debugging.set_log_device_placement(True)
# #tf.config.experimental_run_functions_eagerly(True)
# #physical_devices = tf.config.experimental.list_physical_devices('GPU')
# #tf.config.experimental.set_memory_growth(physical_devices[0], True)
#
#
#
# @memory(0.8)
# def gpu_test():
#
# print("TENSORFLOW")
# with tf.device('GPU:0'):
#
# a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
# b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
# c = tf.matmul(a, b)
#
# print(c)
# #print("LOCAL DEVICES",device_lib.list_local_devices())
# print("PHYSICAL DEVICES")
# print('\n'.join([str(x) for x in tf.config.list_physical_devices()]))
#
# #print(backend.tensorflow_backend._get_available_gpus())
#
# gpu_test()
# +
import keras
from keras import layers
from keras import backend as K
from keras.models import Model
from keras.datasets import mnist
import numpy as np
import tensorflow as tf
img_shape = (28, 28, 1)
batch_size = 16
latent_dim = 2
#@memory(0.8)
def make_model():
with tf.device('GPU:0'):
"""
Encoder
"""
input_img = keras.Input(shape=img_shape)
#x = layers.Conv2D(16, 3, padding='same', activation='relu')(input_img)
#x = layers.Conv2D(32, 3, padding='same', activation='relu', strides=(2, 2))(x)
#x = layers.Conv2D(32, 3, padding='same', activation='relu')(x)
#x = layers.Conv2D(32, 3, padding='same', activation='relu')(x)
x = layers.Conv2D(32, 3, padding='same', activation='relu')(input_img)
x = layers.Conv2D(64, 3, padding='same', activation='relu', strides=(2, 2))(x)
x = layers.Conv2D(64, 3, padding='same', activation='relu')(x)
x = layers.Conv2D(64, 3, padding='same', activation='relu')(x)
shape_before_flattening = K.int_shape(x)
x = layers.Flatten()(x)
x = layers.Dense(32, activation='relu')(x)
#x = layers.Dense(16, activation='relu')(x)
# encoder output (latent parameters)
z_mean = layers.Dense(latent_dim)(x)
z_log_var = layers.Dense(latent_dim)(x)
"""
Sampling
"""
def sampling(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(K.shape(z_mean)[0], latent_dim),
mean=0., stddev=1.)
return z_mean + K.exp(z_log_var) * epsilon
# sample from latent distribution
z = layers.Lambda(sampling)([z_mean, z_log_var])
"""
Decoder
"""
decoder_input = layers.Input(K.int_shape(z)[1:])
x = layers.Dense(np.prod(shape_before_flattening[1:]), activation='relu')(decoder_input)
x = layers.Reshape(shape_before_flattening[1:])(x)
x = layers.Conv2DTranspose(32, 3, padding='same', activation='relu', strides=(2, 2))(x)
x = layers.Conv2D(1, 3, padding='same', activation='sigmoid')(x)
decoder = Model(decoder_input, x)
# decoder output
z_decoded = decoder(z)
"""
Custom Loss
"""
class CustomVariationalLayer(keras.layers.Layer):
def vae_loss(self, x, z_decoded):
x = K.flatten(x)
z_decoded = K.flatten(z_decoded)
# reconstruction loss
xent_loss = keras.metrics.binary_crossentropy(x, z_decoded)
# regularization loss
kl_loss = -5e-4 * K.mean(1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return K.mean(xent_loss + kl_loss)
def call(self, inputs):
x = inputs[0]
z_decoded = inputs[1]
loss = self.vae_loss(x, z_decoded)
self.add_loss(loss, inputs=inputs)
return x
# loss
y = CustomVariationalLayer()([input_img, z_decoded])
"""
Training (on MNIST)
"""
# compile model (input; loss)
vae = Model(input_img, y)
vae.compile(optimizer='rmsprop', loss=None)
return vae
vae = make_model()
vae.summary()
# +
#from src.resourcelims import memory
#tf.config.run_functions_eagerly(True)
#@memory(0.90)
def train():
with tf.device('GPU:0'):
(x_train, _), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_train = x_train.reshape(x_train.shape + (1,))
x_test = x_test.astype('float32') / 255.
x_test = x_test.reshape(x_test.shape + (1,))
vae.fit(x=x_train, y=None, shuffle=True,
epochs=10,
batch_size=batch_size,
validation_data=(x_test, None))
return vae
vae = train()
# -
# This error is due to incompatibility in cuda, cudnn and Nvidia drivers or memory growth issue. The memory growth issue should be addressed so it's the former issues, most likely. TO DO.
|
Deep Learning - Variational Autoencoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part 1: Import, Load Data.
# * ### Import libraries
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.050276, "end_time": "2021-05-12T06:37:35.575327", "exception": false, "start_time": "2021-05-12T06:37:35.525051", "status": "completed"} tags=[]
# import standard libraries
import numpy as np
import pandas as pd
import seaborn as sns
sns.set()
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.model_selection import train_test_split, GridSearchCV
import sklearn.metrics as metrics
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, roc_auc_score
from sklearn.linear_model import LogisticRegression
import warnings
warnings.filterwarnings('ignore')
# + [markdown] papermill={"duration": 0.020256, "end_time": "2021-05-12T06:37:35.617997", "exception": false, "start_time": "2021-05-12T06:37:35.597741", "status": "completed"} tags=[]
# * ### Read data from ‘.csv’ file
# + papermill={"duration": 0.073936, "end_time": "2021-05-12T06:37:35.712323", "exception": false, "start_time": "2021-05-12T06:37:35.638387", "status": "completed"} tags=[]
# read data from '.csv' file
df = pd.read_csv('gender.csv')
# -
# ## Part 2: Exploratory Data Analysis.
# + [markdown] papermill={"duration": 0.021453, "end_time": "2021-05-12T06:37:35.942783", "exception": false, "start_time": "2021-05-12T06:37:35.921330", "status": "completed"} tags=[]
# * ### Info
# + papermill={"duration": 0.034193, "end_time": "2021-05-12T06:37:35.999559", "exception": false, "start_time": "2021-05-12T06:37:35.965366", "status": "completed"} tags=[]
# print the full summary of the dataset
df.info()
# -
# Dataset consists of 66 rows and 5 columns;
#
# has 1 datatype: object(5);
#
# has no missing values.
# * ### Head
# preview of the first 5 lines of the loaded data
df.head()
# * ### Rename Columns
# columns rename
df.columns = ['color', 'music', 'beverage', 'softdrink', 'gender']
df.columns
# * ### Columns visualisation
# columns visualisation
fig, axes = plt.subplots(nrows=5, ncols=1, figsize=(10,18))
for i in range(len(df.columns)):
sns.countplot(data=df, x=df.iloc[:,i],ax=axes[i])
# * ### 'gender' attribute value counts
# 'gender' value counts
df['gender'].value_counts()
# There are 33 of 'Female' and 33 of 'Male' in our dataset. This means that our dataset is balanced.
# + [markdown] papermill={"duration": 0.02239, "end_time": "2021-05-12T06:37:36.285297", "exception": false, "start_time": "2021-05-12T06:37:36.262907", "status": "completed"} tags=[]
# * ### Encode the Data
# + papermill={"duration": 1.286573, "end_time": "2021-05-12T06:37:37.594309", "exception": false, "start_time": "2021-05-12T06:37:36.307736", "status": "completed"} tags=[]
# label encoding
le_color = LabelEncoder()
data = df
data['color'] = le_color.fit_transform(df.color.values)
le_music = LabelEncoder()
data['music'] = le_music.fit_transform(df.music.values)
le_beverage = LabelEncoder()
data['beverage'] = le_beverage.fit_transform(df.beverage.values)
le_softdrink = LabelEncoder()
data['softdrink'] = le_softdrink.fit_transform(df.softdrink.values)
le_gender = LabelEncoder()
data['gender'] = le_gender.fit_transform(df.gender.values)
data.head()
# -
|
ML-101 Modules/Module 03/Lesson 02/Practice 1/Gender - Practice Code Part 1&2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Корректность проверена на Python 3.6:**
# + pandas 0.23.4
# + numpy 1.15.4
# + sklearn 0.20.2
import warnings
warnings.filterwarnings('ignore')
# # Sklearn
# ## sklearn.grid_search
# документация: http://scikit-learn.org/stable/modules/grid_search.html
# +
from sklearn import model_selection, datasets, linear_model, metrics
import numpy as np
import pandas as pd
# -
# ### Генерация датасета
iris = datasets.load_iris()
train_data, test_data, train_labels, test_labels = model_selection.train_test_split(iris.data, iris.target,
test_size = 0.3,random_state = 0)
# ### Задание модели
classifier = linear_model.SGDClassifier(random_state = 0, tol=1e-3)
# ### Генерация сетки
classifier.get_params().keys()
parameters_grid = {
'loss' : ['hinge', 'log', 'squared_hinge', 'squared_loss'],
'penalty' : ['l1', 'l2'],
'max_iter' : np.arange(5,10),
'alpha' : np.linspace(0.0001, 0.001, num = 5),
}
cv = model_selection.StratifiedShuffleSplit(n_splits=10, test_size = 0.2, random_state = 0)
# ### Подбор параметров и оценка качества
# #### Grid search
grid_cv = model_selection.GridSearchCV(classifier, parameters_grid, scoring = 'accuracy', cv = cv)
# %%time
grid_cv.fit(train_data, train_labels)
grid_cv.best_estimator_
print(grid_cv.best_score_)
print(grid_cv.best_params_)
grid_cv.cv_results_
# #### Randomized grid search
randomized_grid_cv = model_selection.RandomizedSearchCV(classifier, parameters_grid, scoring = 'accuracy', cv = cv, n_iter = 20,
random_state = 0)
# %%time
randomized_grid_cv.fit(train_data, train_labels)
print(randomized_grid_cv.best_score_)
print(randomized_grid_cv.best_params_)
# +
# #!/usr/bin/python
from datetime import datetime
def control_sum(str):
return sum(map(ord, list(str)))
def check(reply):
def _is_number(str):
try:
int(str)
return True
except ValueError:
return False
if "Control sum:" not in reply:
return False
parts = reply.split("Control sum:")
received_current_time = parts[0].strip()
received_control_sum = parts[1].strip()
if not _is_number(received_control_sum):
return False
else:
received_control_sum = int(received_control_sum)
expected_control_sum = control_sum(received_current_time)
return expected_control_sum == received_control_sum
current_time = datetime.now().strftime('%Y-%m-%d %H:%M:%S')
print(current_time)
print("Control sum: " + str(control_sum(current_time)))
#print check(current_time + '\n' + "Control sum: " + str(control_sum(current_time)))
|
SupervisedLearning/materials/notebooks/3-1.ScikitGrigSearch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data_folder = "C:\\Users\\spars\\Documents\\Master\\JHU\TML\\HomePriceBeastNew\\"
merged_home_data_time_series = pd.read_csv(f"{data_folder}merged_home_data_time_series.csv", low_memory=True, thousands=',')
merged_home_data_time_series.columns
def transform_types_and_chunk(data, non_float_cols, date_col, dummy_cols):
for x in data.columns:
if x not in non_float_cols:
data[x] = pd.to_numeric(data[x])
data = pd.get_dummies(data, columns = dummy_cols)
data[date_col] = pd.to_datetime(data[date_col])
post_covid_frame = data[data[date_col] >= "2020-03-01"]
train_frame = data[data[date_col] < "2019-12-01"]
test_frame = data[((data[date_col] >= "2019-12-01") & (data[date_col] < "2020-03-01"))]
return train_frame, test_frame, post_covid_frame
# +
non_float_cols = ["state_code", "county_name", "period_begin", "state_code_dummy"]
#convert these columns to encoding.
dummy_cols = ["state_code_dummy"]
#Keep original state code data after dummifyto
merged_home_data_time_series['state_code_dummy'] = merged_home_data_time_series['state_code']
train_frame, test_frame, post_covid_frame = transform_types_and_chunk(merged_home_data_time_series,
non_float_cols,
"period_begin",
dummy_cols)
# -
train_frame.columns
post_covid_frame[['state_code', 'county_name', 'period_begin', 'inventory']].to_csv(f"{data_folder}post_covid_subset_frame.csv")
test_frame[['state_code', 'county_name', 'period_begin', 'inventory']].to_csv(f"{data_folder}test_subset_frame.csv")
# +
def convert_frame_to_numpy(df, remove_cols, target_prefix, related_prefixes, J,H):
#assemble lag variables.
y_lag_cols = [f'{target_prefix}_lag_{j}' for j in range(J,0,-1)]
y_lead_cols = [target_prefix] + [f'{target_prefix}_lead_{h}' for h in range(1,H+1,1)]
flat_drop = []
x_rel_cols = []
for related_prefix in related_prefixes:
curr_prefix = [f'{related_prefix}_lag_{j}' for j in range(J,0,-1)]
x_rel_cols.append(curr_prefix)
flat_drop = flat_drop + curr_prefix
other_cols = [x for x in df.columns if x not in y_lag_cols\
+ y_lead_cols + flat_drop + remove_cols + related_prefixes]
print(f"Length of other columns = {len(other_cols)}")
print(other_cols)
def get_label_row(row):
label = np.array([row[remove_cols].values])
return label
def get_xvec_row(row):
x = np.array([row[y_lag_cols].values])
#Removing sale price in inventory models
for x_rel in x_rel_cols:
x = np.append(x,[row[x_rel].values],axis=0)
stat_val = row[other_cols].values
stat_val = np.tile(stat_val,[J,1])
stat_val = np.transpose(stat_val)
x = np.append(x,stat_val,axis=0)
return x
def get_yvec_row(row):
y = np.array([row[y_lead_cols].values])
return y
X = np.array(df.apply(get_xvec_row, axis = 1))
y = np.array(df.apply(get_yvec_row, axis = 1))
label = np.array(df.apply(get_label_row, axis = 1))
return X,y, label
remove_cols = ["county_name", "period_begin", "state_code"]
target_prefix = 'inventory'
related_prefix = ['week_num','month','week_offset']
J=5
H=3
# -
X_train, y_train, train_label = convert_frame_to_numpy(train_frame,
remove_cols,
target_prefix,
related_prefix,
J,H)
X_test, y_test, test_label = convert_frame_to_numpy(test_frame,
remove_cols,
target_prefix,
related_prefix,
J,H)
X_post_covid, y_post_covid, post_covid_label = convert_frame_to_numpy(post_covid_frame,
remove_cols,
target_prefix,
related_prefix,
J,H)
stack_range = np.array(X_post_covid[0]).shape[0]
print(f"Stack Range : {stack_range}")
def chunks(a, size):
arr = iter(a)
for v in arr:
tmp = [ v ]
for i,v in zip( range( size - 1 ), arr ):
tmp.append( v )
yield tmp
X_train_stack = list(chunks(np.vstack(X_train), stack_range))
X_test_stack = list(chunks(np.vstack(X_test), stack_range))
label_train_stack = np.expand_dims(np.vstack(train_label),axis=2)
label_test_stack = np.expand_dims(np.vstack(test_label),axis=2)
y_train_stack = np.expand_dims(np.vstack(y_train),axis=2)
y_test_stack = np.expand_dims(np.vstack(y_test),axis=2)
X_train_swap = np.array(X_train_stack).swapaxes(0,1).swapaxes(0,2)
X_test_swap = np.array(X_test_stack).swapaxes(0,1).swapaxes(0,2)
label_train_swap = np.array(label_train_stack).swapaxes(0,1)
label_test_swap = np.array(label_test_stack).swapaxes(0,1)
y_train_swap = np.array(y_train_stack).swapaxes(0,1)
y_test_swap = np.array(y_test_stack).swapaxes(0,1)
# +
X_post_covid_stack = list(chunks(np.vstack(X_post_covid), stack_range))
label_post_covid_stack = np.expand_dims(np.vstack(post_covid_label),axis=2)
y_post_covid_stack = np.expand_dims(np.vstack(y_post_covid),axis=2)
X_post_covid_swap = np.array(X_post_covid_stack).swapaxes(0,1).swapaxes(0,2)
label_post_covid_swap = np.array(label_post_covid_stack).swapaxes(0,1)
y_post_covid_swap = np.array(y_post_covid_stack).swapaxes(0,1)
# -
with open(f"{data_folder}all_model_data.npy", 'wb') as f:
np.save(f, X_train_swap.astype(float))
np.save(f, y_train_swap.astype(float))
np.save(f, X_test_swap.astype(float))
np.save(f, y_test_swap.astype(float))
with open(f"{data_folder}all_model_labels_mapping.npy", 'wb') as f:
np.save(f, label_train_swap)
np.save(f, label_test_swap)
with open(f"{data_folder}post_covid_inv_data.npy", 'wb') as f:
np.save(f, X_post_covid_swap.astype(float))
np.save(f, y_post_covid_swap.astype(float))
with open(f"{data_folder}post_covid_inv_labels_mapping.npy", 'wb') as f:
np.save(f, label_post_covid_swap)
X_post_covid_swap.shape
|
Inventory_model_data_prep.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Linear Regression
#
# In this tutorial, we are going to demonstrate how to use the `abess` package to carry out best subset selection
# in linear regression with both simulated data and real data.
#
# ## Linear Regression
#
# Our package `abess` implement a polynomial algorithm in the for best-subset selection problem:
#
# $$
# \min_{\beta\in \mathbb{R}^p} \frac{1}{2n} ||y-X\beta||^2_2,\quad \text{s.t.}\ ||\beta||_0\leq s,
# $$
#
# where $\| \cdot \|_2$ is the $\ell_2$ norm, $\|\beta\|_0=\sum_{i=1}^pI( \beta_i\neq 0)$ is the $\ell_0$ norm of $\beta$, and the sparsity level $s$ is usually an unknown non-negative integer.
# Next, we present an example to show how to use the `abess` package to solve a simple problem.
#
# ## Simulated Data Example
#
# ### Fixed Support Size Best Subset Selection
#
# We generate a design matrix $X$ containing 300 observation and each observation has 1000 predictors. The response variable $y$ is linearly related to the first, second, and fifth predictors in $X$:
#
# $$y = 3X_1 + 1.5X_2 + 2X_5 + \epsilon,$$
#
# where $\epsilon$ is a standard normal random variable.
# +
import numpy as np
from abess.datasets import make_glm_data
np.random.seed(0)
n = 300
p = 1000
real_coef = np.zeros(p)
real_coef[[0, 1, 4]] = 3, 1.5, 2
data1 = make_glm_data(n, p, "gaussian", k = 3, coef_ = real_coef)
# -
print(data1.x.shape)
print(data1.y.shape)
# Use `abessLm` to fit the data, with a fixed support size:
from abess import abessLm
model = abessLm(support_size = [3])
model.fit(data1.x, data1.y)
# After fitting, the predicted coefficients are stored in `model.coef_`:
# +
print("shape:", model.coef_.shape)
ind = np.nonzero(model.coef_)
print("predicted non-zero: ", ind)
print("predicted coef: ", model.coef_[ind])
# -
# From the result, we know that `abess` found which 3 predictors are useful among all 1000 variables. Besides, the predicted coefficients of them are quite close to the real ones.
#
# ### Adaptive Best Subset Selection
#
# However, we may not know the true sparsity level in real world data, and thus we need to determine the most proper one from a large range. Suppose that we believe the real sparsity level is between 0 to 30 (so that `range(0, 31)`):
# +
model = abessLm(support_size = range(31))
model.fit(data1.x, data1.y)
ind = np.nonzero(model.coef_)
print("predicted non-zero: ", ind)
print("predicted coef: ", model.coef_[ind])
# -
# The program can adaptively choose the sparsity level that best fit the data. It is not surprising that it choose 3 variables, the same as the last section.
# ## Real data example
#
# ### Hitters Dataset
#
# Now we focus on real data on the `Hitters` dataset: [https://www.kaggle.com/floser/hitters](https://www.kaggle.com/floser/hitters).
# We hope to use sevral predictors related to the performance of the baseball atheltes last year to predict their salary.
#
# First, let's have a look at this dataset. There are 19 variables except `Salary` and 322 observations.
# +
import pandas as pd
data2 = pd.read_csv('./Hitters.csv')
print(data2.shape)
# -
print(data2.head(5))
# Since the dataset contains some missing values, we simply drop those rows with missing values. Then we have 263 observations remains:
data2 = data2.dropna()
print(data2.shape)
# What is more, before fitting, we need to transfer the character variables to dummy variables:
data2 = pd.get_dummies(data2)
data2 = data2.drop(['League_A', 'Division_E', 'NewLeague_A'], axis = 1)
print(data2.shape)
print(data2.head(5))
# ### Model Fitting
#
# As what we do in simulated data, an adaptive best subset can be formed easily:
# +
x = np.array(data2.drop('Salary', axis = 1))
y = np.array(data2['Salary'])
model = abessLm(support_size = range(20))
model.fit(x, y)
# -
# The result can be showed:
ind = np.nonzero(model.coef_)
print("non-zero:\n", data2.columns[ind])
print("coef:\n", model.coef_)
# Automatically, variables $Hits$, $CRBI$, $PutOuts$, $League\_N$ are chosen in the model (the chosen sparsity level is 4).
# ### More on the results
#
# We can also plot the path of abess process:
# +
import matplotlib.pyplot as plt
pt = np.zeros((20, 19))
ic = np.zeros(20)
for sz in range(20):
model = abessLm(support_size = [sz])
model.fit(x, y)
pt[sz, :] = model.coef_
ic[sz] = model.ic_
for i in range(19):
plt.plot(pt[:, i], label = i)
plt.xlabel('support_size')
plt.ylabel('coefficients')
# plt.legend() # too long to plot
plt.show()
# -
# Besides, we can also generate a graph about the tuning value. Remember that we used the default EBIC to tune the support size.
plt.plot(ic, 'o-')
plt.xlabel('support_size')
plt.ylabel('EBIC')
plt.show()
# In EBIC criterion, `support_size = 4` has the lowest value, so the process adaptively choose 4 variables. Note that under other information criterion, the result may be different.
#
# ## R tutorial
#
# For R tutorial, please view [https://abess-team.github.io/abess/articles/v01-abess-guide.html](https://abess-team.github.io/abess/articles/v01-abess-guide.html).
|
docs/Tutorial/LinearRegression.ipynb
|
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # set_covering3
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/set_covering3.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/set_covering3.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Set covering in Google CP Solver.
Problem from
<NAME>: 'Optimization Models for Decision Making', page 302f
http://ioe.engin.umich.edu/people/fac/books/murty/opti_model/junior-7.pdf
10 senators making a committee, where there must at least be one
representative from each group:
group: senators:
southern 1 2 3 4 5
northern 6 7 8 9 10
liberals 2 3 8 9 10
conservative 1 5 6 7
democrats 3 4 5 6 7 9
republicans 1 2 8 10
The objective is to minimize the number of senators.
Compare with the following models:
* MiniZinc: http://www.hakank.org/minizinc/set_covering3_model.mzn (model)
http://www.hakank.org/minizinc/set_covering3.mzn (data)
* Comet : http://www.hakank.org/comet/set_covering3.co
* ECLiPSe : http://www.hakank.org/eclipse/set_covering3.ecl
* SICStus : http://hakank.org/sicstus/set_covering3.pl
* Gecode : http://hakank.org/gecode/set_covering3.cpp
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver("Set covering")
#
# data
#
num_groups = 6
num_senators = 10
# which group does a senator belong to?
belongs = [
[1, 1, 1, 1, 1, 0, 0, 0, 0, 0], # 1 southern
[0, 0, 0, 0, 0, 1, 1, 1, 1, 1], # 2 northern
[0, 1, 1, 0, 0, 0, 0, 1, 1, 1], # 3 liberals
[1, 0, 0, 0, 1, 1, 1, 0, 0, 0], # 4 conservative
[0, 0, 1, 1, 1, 1, 1, 0, 1, 0], # 5 democrats
[1, 1, 0, 0, 0, 0, 0, 1, 0, 1] # 6 republicans
]
#
# declare variables
#
x = [solver.IntVar(0, 1, "x[%i]" % i) for i in range(num_senators)]
#
# constraints
#
# number of assigned senators (to minimize)
z = solver.Sum(x)
# ensure that each group is covered by at least
# one senator
for i in range(num_groups):
solver.Add(
solver.SumGreaterOrEqual(
[x[j] * belongs[i][j] for j in range(num_senators)], 1))
objective = solver.Minimize(z, 1)
#
# solution and search
#
solution = solver.Assignment()
solution.Add(x)
solution.AddObjective(z)
collector = solver.LastSolutionCollector(solution)
solver.Solve(
solver.Phase(x, solver.INT_VAR_DEFAULT, solver.INT_VALUE_DEFAULT),
[collector, objective])
print("z:", collector.ObjectiveValue(0))
print("x:", [collector.Value(0, x[i]) for i in range(num_senators)])
for j in range(num_senators):
if collector.Value(0, x[j]) == 1:
print("Senator", j + 1, "belongs to these groups:", end=" ")
for i in range(num_groups):
if belongs[i][j] == 1:
print(i + 1, end=" ")
print()
print()
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
|
examples/notebook/contrib/set_covering3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
while True:
try:
money_spent = int(input('How much money was spent?: '))
num_transactions = int(input('How many transactions?: '))
average_value = money_spent / num_transactions
print("The average value per day is " + str(average_value) + '$')
break
except:
ValueError
print('Type in a number')
# -
def mulai(update, context):
update.message.reply_text('Absen Start!')
while True:
if pycron.is_now('*/1 * * * *'):
absen()
time.sleep(5)
# +
import math
while True:
try:
number_of_kids = int(input('How much money was spent?: '))
num_transactions = int(input('How many transactions?: '))
average_value = money_spent / num_transactions
print("The average value per day is " + str(average_value) + '$')
break
except:
ValueError
print('Type in a number')
# -
# +
from time import sleep
def counting_to_ten(self):
#Counting up to 10
for i in range(20):
print(i+1)
sleep(1)
#stop counting at 5
if i == 10:
break
def close_function(self):
counting_to_ten()
# +
from collections import Counter
z = ['blue', 'red', 'blue', 'yellow', 'blue', 'red']
l = [(1, 2), (1, 3), (1, 4), (1, 10)]
Counter(l)
# -
|
StackOverflow2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NFmOh482SyEF"
# ## Assignment 3: Dealing with overfitting
# + [markdown] colab_type="text" id="AjzAuO3oSvsI"
# Today we work with [Fashion-MNIST dataset](https://github.com/zalandoresearch/fashion-mnist) (*hint: it is available in `torchvision`*).
#
# Your goal for today:
# 1. Train a FC (fully-connected) network that achieves >= 0.885 test accuracy.
# 2. Cause considerable overfitting by modifying the network (e.g. increasing the number of network parameters and/or layers) and demonstrate in in the appropriate way (e.g. plot loss and accurasy on train and validation set w.r.t. network complexity).
# 3. Try to deal with overfitting (at least partially) by using regularization techniques (Dropout/Batchnorm/...) and demonstrate the results.
#
# __Please, write a small report describing your ideas, tries and achieved results in the end of this file.__
#
# *Note*: Tasks 2 and 3 are interrelated, in task 3 your goal is to make the network from task 2 less prone to overfitting. Task 1 is independent from 2 and 3.
#
# *Note 2*: We recomment to use Google Colab or other machine with GPU acceleration.
# + colab={} colab_type="code" id="_KBld6VOSwhW"
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
import torchsummary
from IPython.display import clear_output
from matplotlib import pyplot as plt
from matplotlib.pyplot import figure
import numpy as np
import os
device = 'cuda:0' if torch.cuda.is_available() else 'cpu'
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="EdLOG0XqS_g5" outputId="1a58887c-24fc-4315-bb85-bdc88f4f485e"
# Technical function
def mkdir(path):
if not os.path.exists(root_path):
os.mkdir(root_path)
print('Directory', path, 'is created!')
else:
print('Directory', path, 'already exists!')
root_path = 'fmnist'
mkdir(root_path)
# + colab={"base_uri": "https://localhost:8080/", "height": 397, "referenced_widgets": ["a00cbbf2385c426bb848399f3c13b70f", "4c7a7ac1286649c4804fefd359a1be74", "9faad8d0c45746ab8e06e120bb3ec0b6", "590134f878a74adb98fd129816fde03c", "92685fe1840a4555a6962c006ea90c23", "aa0dc032d1644bad8e741cf1696d9a70", "4296a7e9ab704f019451001dd12c4f46", "29b0a8c468e6410fb390b903559d6ef5", "fb66cde27d0849bba4947c024e198f7f", "bebcff464bfc44ffa82e311a39cae7d7", "b9d32ac7b88c4f4e900b160f05c016a3", "<KEY>", "<KEY>", "<KEY>", "9761d073bcef4c15ad64b01d7bef3561", "<KEY>", "<KEY>", "583e6f0878e04a0bbebf5d80d9d712dc", "0078023356f142bdbe4e13a42df460fb", "01658e0aebd84db9912ea456bd1ad030", "54a1be4ee00c44a0b72192135f62fa16", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "029f1768c5734beab39a592b44f75a9d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>"]} colab_type="code" id="qt6LE7XaTDT9" outputId="ab967f1e-8bf2-4199-cbd7-75806359ee1b"
download = True
train_transform = transforms.ToTensor()
test_transform = transforms.ToTensor()
transforms.Compose((transforms.ToTensor()))
fmnist_dataset_train = torchvision.datasets.FashionMNIST(root_path,
train=True,
transform=train_transform,
target_transform=None,
download=download)
fmnist_dataset_test = torchvision.datasets.FashionMNIST(root_path,
train=False,
transform=test_transform,
target_transform=None,
download=download)
# + colab={} colab_type="code" id="71YP0SPwTIxD"
train_loader = torch.utils.data.DataLoader(fmnist_dataset_train,
batch_size=128,
shuffle=True,
num_workers=2)
test_loader = torch.utils.data.DataLoader(fmnist_dataset_test,
batch_size=256,
shuffle=False,
num_workers=2)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="v_YFmF7NTWrQ" outputId="6b517f52-5fc5-482e-cc8e-cd6b3f1b72f1"
len(fmnist_dataset_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="aHca15bOTY4B" outputId="7eb477ef-816d-418c-f5c3-ade63d4cf915"
for img, label in train_loader:
print(img.shape)
# print(img)
print(label.shape)
print(label.size(0))
break
# + [markdown] colab_type="text" id="b6OOOffHTfX5"
# ### Task 1
# Train a network that achieves $\geq 0.885$ test accuracy. It's fine to use only Linear (`nn.Linear`) layers and activations/dropout/batchnorm. Convolutional layers might be a great use, but we will meet them a bit later.
# + colab={} colab_type="code" id="ftpkTjxlTcFx"
class TinyNeuralNetwork(nn.Module):
def __init__(self, input_shape=28*28, num_classes=10, input_channels=1):
super(self.__class__, self).__init__()
self.model = nn.Sequential(
nn.Flatten(), # This layer converts image into a vector to use Linear layers afterwards
# Your network structure comes here
nn.Linear(input_shape, num_classes)
)
def forward(self, inp):
out = self.model(inp)
# -
torchsummary.summary(TinyNeuralNetwork().to(device), (28*28,))
# + [markdown] colab_type="text" id="544PGKEnjPr5"
# Your experiments come here:
# + colab={"base_uri": "https://localhost:8080/", "height": 607} colab_type="code" id="i3POFj90Ti-6" outputId="82e7e921-541b-4657-f78d-563de48b07c7"
model = TinyNeuralNetwork().to(device)
opt = # YOUR CODE HERE
loss_func = # YOUR CODE HERE
# Your experiments, training and validation loops here
# + [markdown] colab_type="text" id="L7ISqkjmCPB1"
# ### Task 2: Overfit it.
# Build a network that will overfit to this dataset. Demonstrate the overfitting in the appropriate way (e.g. plot loss and accurasy on train and test set w.r.t. network complexity).
#
# *Note:* you also might decrease the size of `train` dataset to enforce the overfitting and speed up the computations.
# + colab={} colab_type="code" id="H12uAWiGBwJx"
class OverfittingNeuralNetwork(nn.Module):
def __init__(self, input_shape=28*28, num_classes=10, input_channels=1):
super(self.__class__, self).__init__()
self.model = nn.Sequential(
nn.Flatten(), # This layer converts image into a vector to use Linear layers afterwards
# Your network structure comes here
nn.Linear(input_shape, num_classes)
)
def forward(self, inp):
out = self.model(inp)
# + colab={"base_uri": "https://localhost:8080/", "height": 449} colab_type="code" id="JgXAKCpvCwqH" outputId="8d29ad18-3f0c-4161-8bcd-004d24ba771c"
torchsummary.summary(OverfittingNeuralNetwork().to(device), (28*28,))
# +
model = OverfittingNeuralNetwork().to(device)
opt = # YOUR CODE HERE
loss_func = # YOUR CODE HERE
# Your experiments, come here
# -
# ### Task 3: Fix it.
# Fix the overfitted network from the previous step (at least partially) by using regularization techniques (Dropout/Batchnorm/...) and demonstrate the results.
class FixedNeuralNetwork(nn.Module):
def __init__(self, input_shape=28*28, num_classes=10, input_channels=1):
super(self.__class__, self).__init__()
self.model = nn.Sequential(
nn.Flatten(), # This layer converts image into a vector to use Linear layers afterwards
# Your network structure comes here
nn.Linear(input_shape, num_classes)
)
def forward(self, inp):
out = self.model(inp)
torchsummary.summary(FixedNeuralNetwork().to(device), (28*28,))
# +
model = FixedNeuralNetwork().to(device)
opt = # YOUR CODE HERE
loss_func = # YOUR CODE HERE
# Your experiments, come here
# + [markdown] colab_type="text" id="dMui_uLJ7G0d"
# ### Conclusions:
# _Write down small report with your conclusions and your ideas._
|
assignments/assignment0_03_DL/assignment0_03_Dealing_with_overfitting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:REL560] *
# language: python
# name: conda-env-REL560-py
# ---
sources_dir = "/Users/jacobbarrett/Documents/GitHub/REL560-SDA-Periodicals/Periodicals/LibM"
sources_dir
import os
os.listdir(sources_dir)
periodical_list = os.listdir(sources_dir)
periodical_list
# What metadata is here?
# - Publication date (YYYYMMDD)
# - Volume Number
# - Issue Number
# - Periodical Title Key
#
# What can we use to split out the bits of info?
#
# - Dashes
# split keydate | volume | issue
#
# How do we separate the date and title key?
# - regex to grab only letters
#
# How to separate YYYYMMDD?
# first 4 (0-3) are year
# 4th and 5th positions are month
# 6th and 7th are day
import re
import csv
# +
# [Periodical key, year, month, day, volume, issue]
with open('../Data/LibM-metadata.csv', 'w', newline='') as csvfile:
csvwriter = csv.writer(csvfile, delimiter=',')
csvwriter.writerow(["file_name", "periodical_key", "year", "month", "day", "volume", "issue"])
for each in periodical_list:
each = os.path.splitext(each)[0]
first_split = each.split('-')
print(first_split)
title = re.match('[A-Za-z]*', first_split[0]).group(0)
print(title)
date = re.findall(r'\d+', first_split[0])[0]
print(date)
csvwriter.writerow([each, title, date[0:4], date[4:6], date[6:8], first_split[1], first_split[2]])
# -
|
code/2022-02-Extract-Metadata-LibM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="NoNMujanjNuu" colab_type="code" colab={}
#Given a map consisting of known poses and a start and end pose, find the optimal path between using A*
#Generate the relative motion in se2 between poses.
#This is straight line motion.
#Also implements cubic interpolation for a smooth trajectory across all points in path.
# + id="VqXKrUKwt05L" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy as np
import random
import scipy.interpolate
import heapq #https://docs.python.org/3/library/heapq.html
# + id="VMJWsOvIvGeJ" colab_type="code" colab={}
#Loading poses from the ground truth file
def load_poses(pose_gt_file) :
pose_gt = np.loadtxt(pose_gt_file, delimiter = ",")
return pose_gt[1:, 1:3]
poses = load_poses('../dataset/data/ground_truth/groundtruth_2012-01-08.csv')
# + id="jQQn9T3U4yA2" colab_type="code" colab={}
#Astar and path functions
class PriorityQueue:
def __init__(self):
self.elements = []
def empty(self):
return len(self.elements) == 0
def put(self, item, priority):
heapq.heappush(self.elements, (priority, item))
def get(self):
return heapq.heappop(self.elements)[1]
class Astar :
# This class implements A* search along a network defined by several points
# Poses is an array of coordinates
# k defines how many nearest neighbors to look at during A* search
# The primary usage of this class is the find_path function:
# Required parameters:
# start_idx:
# goal_idx
def __init__(self, poses) :
self.poses = poses
self.full_tree = scipy.spatial.KDTree(self.poses)
def _extract_path(self, cur_node, parent_idx, start_idx, sparse_poses):
next_idx = cur_node
path = [self.full_tree.query(sparse_poses[next_idx])[1]]
while next_idx != start_idx:
next_idx = parent_idx[next_idx]
path.append(self.full_tree.query(sparse_poses[next_idx])[1])
return path[::-1]
def find_path(self, full_start_idx, full_goal_idx, sparseness=1, k=5):
sparse_poses = poses[0::sparseness, :]
visit_queue = PriorityQueue()
visited_flag, queueed_flag = np.zeros(sparse_poses.shape[0]), np.zeros(sparse_poses.shape[0])
g_score, h_score = np.full(sparse_poses.shape[0], np.inf), np.full(sparse_poses.shape[0], np.inf)
parent_idx = np.zeros(sparse_poses.shape[0], dtype='int')
sparse_tree = scipy.spatial.KDTree(sparse_poses)
start_idx = sparse_tree.query(poses[full_start_idx])[1]
goal_idx = sparse_tree.query(poses[full_goal_idx])[1]
# initialize
goal = sparse_poses[goal_idx]
g_score[start_idx] = 0
visit_queue.put(start_idx, np.inf)
queueed_flag[start_idx] = 1
optimal = False
while not visit_queue.empty():
cur_node = visit_queue.get()
visited_flag[cur_node] = 1
if cur_node == goal_idx:
optimal = True
break
# find neighbours
neighbors = sparse_tree.query(sparse_poses[cur_node], k=k)
for nb_cur_dist, nb_idx in zip(neighbors[0][1:], neighbors[1][1:]):
if visited_flag[nb_idx] == 1:
continue
temp_dist = g_score[cur_node] + np.linalg.norm(sparse_poses[cur_node] - sparse_poses[nb_idx])
# temp_dist = g_score[cur_node] + nb_cur_dist ## this not work
if g_score[nb_idx] > temp_dist:
g_score[nb_idx] = temp_dist
parent_idx[nb_idx] = cur_node
f_score = g_score[nb_idx] + np.linalg.norm(sparse_poses[nb_idx] - goal)
# put into queen
if queueed_flag[nb_idx] == 0:
visit_queue.put(nb_idx, f_score)
queueed_flag[nb_idx] = 1
path = self._extract_path(cur_node, parent_idx, start_idx, sparse_poses)
path[0] = full_start_idx
path[-1] = full_goal_idx
return path, optimal
def find_local_path(self, start_pose, path, steps=5) :
set_trace()
path_tree = scipy.spatial.KDTree(self.poses[path])
path_idx = path_tree.query(start_pose)[1]
start_idx = self.full_tree.query(self.poses[path[path_idx]])[1]
if path_idx + 5 < len(path) :
goal_idx =self.full_tree.query(self.poses[path[path_idx + steps]])[1]
else :
goal_idx =self.full_tree.query(self.poses[path[-1]])[1]
local_path, _ = self.find_path(start_idx, goal_idx)
return local_path
def total_dist_fun(poses) :
total_dist = 0
curr_point = poses[0]
for idx in range(1, poses.shape[0]) :
total_dist += np.linalg.norm(curr_point - poses[idx])
curr_point = poses[idx]
return total_dist
# + id="cvOUf76S5zFZ" colab_type="code" colab={}
#construct A* instance
astar = Astar(poses)
# + id="Mh_A8aKdvwEt" colab_type="code" outputId="5e84feb2-7cc6-4ab2-a4de-c873f3d7b96f" colab={"base_uri": "https://localhost:8080/", "height": 51}
#Test A*
start_idx = np.random.randint(poses.shape[0])
goal_idx = np.random.randint(poses.shape[0])
path, optimal = astar.find_path(start_idx, goal_idx, sparseness=10, k=50)
# + id="CkqvtjJ65x4Y" colab_type="code" outputId="f538c889-e5dd-43a1-b615-213b9161cf75" colab={"base_uri": "https://localhost:8080/", "height": 635}
#Plot computed path
plt.figure(figsize=(16,9))
plt.scatter(poses[:,1], poses[:,0], s=1)
plt.scatter(poses[path,1], poses[path,0], c='y', s=20)
plt.scatter(poses[start_idx,1], poses[start_idx,0], marker='o', c='g', s=500, label='start')
plt.scatter(poses[goal_idx,1], poses[goal_idx,0], marker='*', c='r', s=750, label='goal')
plt.legend()
plt.title('Ground Truth Position of Nodes with Overlaid A* Path')
plt.xlabel('East (m)')
plt.ylabel('North (m)')
plt.axis('equal')
# + id="CRtQebcnjeSM" colab_type="code" colab={}
#SE(2) functions
def matrix_log_SO2(SO2_mat) :
#ln(R) in SO(3) = theta
return np.arctan2(SO2_mat[1,0], SO2_mat[0, 0])
def matrix_log_SE2(SE2_mat) :
theta = matrix_log_SO2(SE2_mat[0:2, 0:2])
if (theta < 1e-6) :
A = 1
B = 0
else :
A = np.sin(theta)/theta
B = (1-np.cos(theta))/theta
v_inv = 1/(A**2 + B**2) * np.array([[A, B], [-B, A]])
mat_log = np.array(np.matmul(v_inv, SE2_mat[0:2, 2]))
mat_log = np.append(mat_log, theta)
return mat_log
def matrix_exp_so2(theta) :
#reconstruct R.
return np.array([[np.cos(theta), -np.sin(theta)], [np.sin(theta), np.cos(theta)]])
def matrix_exp_se2(twist) :
theta = twist[-1]
R = matrix_exp_so2(theta)
#V converges to I2
if (theta < 1e-6) :
V = np.eye(2)
else:
V = 1/theta * np.array([[np.sin(theta), -(1 - np.cos(theta))], [(1-np.cos(theta)), np.sin(theta)]])
mat_exp = np.zeros((3,3))
mat_exp[0:2, 0:2] = R
mat_exp[0:2, 2] = np.matmul(V, twist[0:2])
mat_exp[2, 2] = 1
return mat_exp
def get_twist_SE2(Xstart, pos_end, pos_future=None) :
Xend = np.zeros((3,3))
Xend[-1,-1] = 1
Xend[0:2, 2] = pos_end
#compute end direction (face in direction of future step i.e. end+1)
if not pos_future is None:
next_displacement = pos_future - pos_end
next_theta = np.arctan2(next_displacement[1], next_displacement[0])
Xend[0:2, 0:2] = np.array([[np.cos(next_theta), -np.sin(next_theta)], [np.sin(next_theta), np.cos(next_theta)]])
else :
Xend[0:2, 0:2] = Xstart[0:2, 0:2]
# set_trace()
twist_SE2 = matrix_log_SE2(np.matmul(np.linalg.inv(Xstart), Xend))
return twist_SE2, Xend
def twist_motion(Xstart, twist, s=1) :
return np.matmul(Xstart, s * matrix_exp_se2(twist))
# + id="XkTTjfR_EaGT" colab_type="code" outputId="5aeff481-c2f9-461b-e6e9-85e0f6b4ea12" colab={"base_uri": "https://localhost:8080/", "height": 136}
print('testing exponential map for SE2')
thetas = [0, 1e-4, np.pi/2, np.pi, 15*np.pi/8, 4.5 * np.pi]
for theta in thetas :
test_SE2 = np.array([[np.cos(theta), -np.sin(theta), 1.5], [np.sin(theta), np.cos(theta), 2], [0, 0, 1]])
twist = matrix_log_SE2(test_SE2)
SE2_res = matrix_exp_se2(twist)
assert(np.sum(test_SE2 - SE2_res) < 1e-6)
print('passed theta = ', theta)
# + id="LsDb3F7fKBol" colab_type="code" outputId="fdeaf8f7-a326-4ea7-b495-7fb751b784e0" colab={"base_uri": "https://localhost:8080/", "height": 51}
#Generation of twists and executing path
print('testing motion twist generation')
pos_end = np.array([x1, y1])
pos_future = np.array([x2, y2])
twists = []
Xstart = np.eye(3)
Xstart[0:2, 2] = poses[path[0]]
poses = np.array(poses)
for pose_idx, path_idx in enumerate(path[1:-1]) :
twist, Xstart = get_twist_SE2(Xstart, poses[pose_idx], poses[path_idx + 1])
twists.append(twist)
#print(twist)
twist, Xend = get_twist_SE2(Xstart, poses[-1])
twists.append(twist)
Xk = np.eye(3)
Xk[0:2, 2] = poses[path[0]]
for twist in twists :
Xk = twist_motion(Xk, twist)
assert(np.sum(Xk - Xend) < 1e-6)
print('passed')
# + id="PaYwXvSfeO4k" colab_type="code" outputId="e53f3ada-f8bd-4ec8-ffd2-124154658d74" colab={"base_uri": "https://localhost:8080/", "height": 985}
#Cubic interpolation of poses.
poss = np.array(poses[path])
velocities = np.zeros(poss.shape)
T = np.zeros(poss.shape[0])
total_time = 100
total_dist = total_dist_fun(poss)
cum_dist = 0
velocities[0] = 0
for i in range(1, poss.shape[0] - 1) :
seg_dist = np.linalg.norm(poss[i+1] - poss[i])
velocities[i] = (((poss[i+1] - poss[i]) / seg_dist) + velocities[i-1])/2
T[i] = total_time * cum_dist/total_dist
cum_dist += seg_dist
T[-1] = total_time
velocities[-1] = 0
print(velocities)
a = np.zeros((poss.shape[0], 4, poss.shape[1]))
for j in range(0, poss.shape[0]-1) :
del_Tj = T[j+1] - T[j]
a[j, 0] = poss[j]
a[j, 1] = velocities[j]
a[j, 2] = (3 * poss[j+1] - 3 * poss[j] - 2 * velocities[j] * del_Tj - velocities[j+1] * del_Tj)/ (del_Tj**2)
a[j, 3] = (2 * poss[j] + (velocities[j] + velocities[j+1]) * del_Tj - 2 * poss[j + 1]) / (del_Tj**3)
del_t = 0.005
pos_x = [a[0,0][0]]
pos_y = [a[0,0][1]]
vel_x = [0]
vel_y = [0]
total_trial = 100
for t in np.arange(del_t, total_trial, del_t) :
j = np.argmax(T > t)-1
delta_t = t - T[j]
pos_t = a[j, 0] + a[j, 1]* delta_t + a[j, 2] * (delta_t**2) + a[j, 3] * (delta_t**3)
pos_x.append(pos_t[0])
pos_y.append(pos_t[1])
vel_x.append((pos_x[-1] - pos_x[-2])/del_t)
vel_y.append((pos_y[-1] - pos_y[-2])/del_t)
t = np.arange(0, total_trial, del_t)
plt.figure(figsize=(16,9))
plt.plot(t[1:405], pos_x[1:405], linestyle='-', c='r', label='interpolated x position')
# plt.scatter(t[0:400], pos_y[0:400], label='y position')
plt.scatter(T[1:14], poss[1:14,0], c='b')
plt.plot(T[1:14], poss[1:14,0], linestyle='-', c='g', label='no interpolation')
# plt.scatter(T[0:10], poss[0:10,1], label='y no interp')
plt.legend()
plt.title('position with cubic interpolation of via points')
plt.xlabel('Time (s)')
plt.ylabel('Position (m)')
plt.figure()
plt.scatter(t[2:500], vel_x[2:500])
plt.scatter(t[2:500], vel_y[2:500])
|
src/planning/Motion_Planning_Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="_b0A-ElAnHj2" colab_type="code" colab={}
import pandas as pd
import joblib
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# + id="xuyNn2VlnHm9" colab_type="code" colab={}
file_url = 'https://raw.githubusercontent.com/PacktWorkshops/The-Data-Science-Workshop/master/Chapter11/dataset/breast-cancer-wisconsin.data'
# + id="MSH55O2Qn3nn" colab_type="code" colab={}
col_names = ['Sample code number','Clump Thickness','Uniformity of Cell Size','Uniformity of Cell Shape','Marginal Adhesion','Single Epithelial Cell Size',
'Bare Nuclei','Bland Chromatin','Normal Nucleoli','Mitoses','Class']
# + id="qHM7W8jTnHye" colab_type="code" colab={}
df = pd.read_csv(file_url, header=None, names=col_names, na_values='?')
# + id="HZ5iAGhvntZU" colab_type="code" outputId="e1ab5f63-dd7e-4b9d-d6fa-c185fca8e268" executionInfo={"status": "ok", "timestamp": 1574652400671, "user_tz": -660, "elapsed": 2805, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
df.head()
# + id="pHG3ADzWpIXU" colab_type="code" colab={}
df.fillna(0, inplace=True)
# + id="bjMOEqmuntdD" colab_type="code" colab={}
y = df.pop('Class')
# + id="HHs8bS9qntgL" colab_type="code" colab={}
X = df.drop('Sample code number', axis=1)
# + id="8fB7TTDuoZw1" colab_type="code" outputId="f0219dd4-a5f8-4e4c-d909-8f74d173b63a" executionInfo={"status": "ok", "timestamp": 1574652400674, "user_tz": -660, "elapsed": 2792, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
X.head()
# + id="TnmZuUKbnti9" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=888)
# + id="rM2D75s0otzY" colab_type="code" colab={}
rf_model = RandomForestClassifier(random_state=1)
# + id="VAOTJvwCot_C" colab_type="code" outputId="1e6f7e74-ae56-4269-f7fd-dfc531fedbae" executionInfo={"status": "ok", "timestamp": 1574652401023, "user_tz": -660, "elapsed": 3128, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 190}
rf_model.fit(X_train, y_train)
# + id="rMI1mOihouBq" colab_type="code" outputId="dd35bd04-d606-4bcc-8e9c-8e54e5f185a1" executionInfo={"status": "ok", "timestamp": 1574652401024, "user_tz": -660, "elapsed": 3122, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
rf_model.predict([X_test.iloc[0,]])
# + id="E3wJTFm2qgUV" colab_type="code" outputId="3e59acd8-28e2-4fa0-8d77-fa768064eeb1" executionInfo={"status": "ok", "timestamp": 1574652401027, "user_tz": -660, "elapsed": 3117, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
joblib.dump(rf_model, "model.pkl")
# + id="kl29WUo3W4D5" colab_type="code" colab={}
import socket
import threading
import requests
import json
from flask import Flask, jsonify, request
import numpy as np
# + id="FHV797Dlqac_" colab_type="code" outputId="2ff13d41-9be1-4217-ac32-3759c232900a" executionInfo={"status": "ok", "timestamp": 1574652401029, "user_tz": -660, "elapsed": 3110, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
ip_address = socket.gethostbyname(socket.gethostname())
ip_address
# + id="rBMrb4UNqb3U" colab_type="code" colab={}
app = Flask(__name__)
# + id="YoseBWhgqcd7" colab_type="code" colab={}
trained_model = joblib.load("model.pkl")
# + id="ZwPHtIQKqn5c" colab_type="code" colab={}
@app.route('/api', methods=['POST'])
def predict():
data = request.get_json()
prediction = trained_model.predict(data)
str_pred = np.array2string(prediction)
return jsonify(str_pred)
# + id="6iLQtxEEz6vK" colab_type="code" outputId="ca7b029e-e96d-4b19-d1e0-ec301e80083a" executionInfo={"status": "ok", "timestamp": 1574652401032, "user_tz": -660, "elapsed": 3098, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
flask_thread = threading.Thread(target=app.run, kwargs={'host':'0.0.0.0','port':80})
flask_thread.start()
# + id="_KmZqH-mqn72" colab_type="code" outputId="e6c3a62a-bdd7-400d-9f5d-a8fe5f8d755f" executionInfo={"status": "ok", "timestamp": 1574652401032, "user_tz": -660, "elapsed": 3092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
record = X_test.iloc[0,].to_list()
record
# + id="iL7ezCzKqn-Y" colab_type="code" colab={}
j_data = json.dumps([record])
# + id="uRU36nLVq1Vh" colab_type="code" colab={}
headers = {'content-type': 'application/json', 'Accept-Charset': 'UTF-8'}
# + id="ORrZn6Y3q1cl" colab_type="code" outputId="ab0c1a64-66f9-4d3e-b4e3-7b4f5ff82c92" executionInfo={"status": "ok", "timestamp": 1574652402130, "user_tz": -660, "elapsed": 4177, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCYY-iGjUIqBSnlLoszfZTN7rU7FRNg05Rdt9Ii3A=s64", "userId": "11809607246124237079"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
r = requests.post(f"http://{ip_address}/api", data=j_data, headers=headers)
r.text
|
Chapter18/Exercise18.02/Exercise18_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
from sklearn.cluster import KMeans
import warnings
warnings.filterwarnings('ignore') # Turn off the warnings.
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import os
from sklearn.datasets import make_blobs, make_moons
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.cluster import DBSCAN
from scipy.cluster.hierarchy import linkage, dendrogram, fcluster
# %matplotlib inline
# -
df = sns.load_dataset('iris')
X= df.drop(columns=['species'])
Y= df['species']
head_X = X.columns
df.head()
X
Y
kmeans = KMeans(n_clusters= 3, random_state= 123)
kmeans.fit(X)
rs= pd.Series(kmeans.labels_)
rs
case0 = Y[rs==0]
case1 = Y[rs==1]
case2 = Y[rs==2]
print(case0.value_counts())
print(case2.value_counts())
print(case1.value_counts())
pd.DataFrame(kmeans.cluster_centers_,columns = head_X, index= ['cluster 0','Cluster 1','Cluster 3'])
pd.DataFrame(kmeans.cluster_centers_,columns = head_X)
np.round(pd.DataFrame(kmeans.cluster_centers_,columns = head_X, index= ['cluster 0','Cluster 1','Cluster 3']))
learnedLabels = ['Virginica','Setosa','Versicolor']
learnedLabels
sns.countplot(case0).set_title("Cluster 0")
plt.show()
sns.countplot(case1).set_title("Cluster 1")
plt.show()
sns.countplot(case2).set_title("Cluster 2")
plt.show()
# +
# For a given observation of X, predict the species from what we have learned.
# Case #1.
X_test = {'sepal_length': [4.5] ,'sepal_width': [3.0] , 'petal_length': [5.0] ,'petal_width': [1.5] } # Only X is given.
X_test = pd.DataFrame(X_test)
predCluster = kmeans.predict(X_test)[0]
print("Predicted cluster {} with the most probable label '{}'".format(predCluster,learnedLabels[predCluster]))
# -
X_test = print(pd.DataFrame(X_test))
print(X_test)
# Dataset #1.
X1, label1 = make_blobs(n_samples=200, n_features=2, centers=2, cluster_std = 5, random_state=123)
plt.scatter(X1[:,0],X1[:,1], c= label1, alpha=0.7 )
plt.title('Dataset #1 : Original')
plt.show()
agglo = AgglomerativeClustering(n_clusters=2)
agglo.fit(X1)
myColors = {0:'red',1:'green'} # Define a color palette: 0~1.
plt.scatter(X1[:,0],X1[:,1], c= pd.Series(agglo.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #1 : Agglomerative')
plt.show()
myLinkage = linkage(X1,method='single') # Cluster hierarchically using single linkage.
plt.figure(figsize=(20,5))
dendrogram(myLinkage)
plt.show()
# +
# Dataset #1 and clusters by cutting the dendrogram.
labels = fcluster(myLinkage, 5, criterion='distance') # Cut at the height (distance) = 5 <= change this value at will.
pd.Series(labels).value_counts()
# +
# Dataset #2.
X2, label2 = make_moons(n_samples=200, noise=0.08, random_state=123)
plt.scatter(X2[:,0],X2[:,1], c= label2, alpha=0.7 )
plt.title('Dataset #2 : Original')
plt.show()
# -
agglo = AgglomerativeClustering(n_clusters=2)
agglo.fit(X2)
myColors = {0:'red',1:'green'} # Define a color palette: 0~1.
plt.scatter(X2[:,0],X2[:,1], c= pd.Series(agglo.labels_).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #2 : Agglomerative')
plt.show()
# Dataset #2 and show dendrogram.
myLinkage = linkage(X2,method='single') # Cluster hierarchically using single linkage.
plt.figure(figsize=(20,5))
dendrogram(myLinkage)
plt.show()
# +
# Dataset #2 and clusters by cutting the dendrogram.
labels = fcluster(myLinkage, 0.23, criterion='distance') # Cut at the height (distance) = 0.23 <= change this value at will.
pd.Series(labels).value_counts()
# -
myColors = {1:'red',2:'green'} # Define a color palette: 1~2.
plt.scatter(X2[:,0],X2[:,1], c= pd.Series(labels).apply(lambda x: myColors[x]), alpha=0.7 )
plt.title('Dataset #2 : Hierarchical')
plt.show()
# # First Diagram and the clustering
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
# %matplotlib inline
import scipy.cluster.hierarchy as sch
from sklearn.cluster import AgglomerativeClustering
dataset= make_blobs(n_samples= 200, centers= 4, n_features= 2, cluster_std= 1.6,random_state= 50)
point = dataset[0]
dendrogram= sch.dendrogram(sch.linkage(point,method= 'ward'))
dendrogram= sch.dendrogram(sch.linkage(point,method= 'single'))
hc = AgglomerativeClustering(n_clusters=4, affinity ='euclidean', linkage='single')
hc
y_hc= hc.fit_predict(point)
print(y_hc)
# +
# this will plot the graphic the data set
plt.scatter(point[y_hc ==0,0], point[y_hc == 0,1], color = 'red', s=80)
plt.scatter(point[y_hc == 1,0], point[y_hc == 1,1], color = 'blue', s =80)
plt.scatter(point[y_hc == 2,0], point[y_hc == 2,1], color = 'green', s =80)
plt.scatter(point[y_hc == 3,0], point[y_hc == 3,1], color = 'yellow', s =80)
plt.show()
# -
# # First Clustering and then diagram
kmean = KMeans(n_clusters= 4)
clusters = kmean.cluster_centers_
y_km=kmean.fit_predict(point)
# Ploting the graph
plt.scatter(point[y_km == 0,0], point[y_km ==0,1], s = 80, color= 'red')
plt.scatter(point[y_km == 1,0], point[y_km ==1,1], s = 80, color= 'blue')
plt.scatter(point[y_km == 2,0], point[y_km ==2,1], s = 80, color= 'green')
plt.scatter(point[y_km == 3,0], point[y_km ==3,1], s = 80, color= 'black')
|
Rafay notes/Samsung Course/Chapter 5/Class Work/Lecture-2-22-oct-2021/Pratice of class work.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Classification de documents
# ## Imports
# + tags=[]
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
import seaborn as sn
from pprint import pprint
import numpy as np
import nltk
nltk.download('stopwords')
from sklearn.datasets import fetch_20newsgroups
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import confusion_matrix, classification_report
from scikitplot.metrics import plot_confusion_matrix
import pandas as pd
import re
import operator
# -
# ## Charger le dataset 20 newsgroups
# Pour plus d'information : https://scikit-learn.org/0.19/datasets/twenty_newsgroups.html
news = fetch_20newsgroups(subset='all')
print("Number of articles: " + str(len(news.data)))
print("Number of categories: " + str(len(news.target_names)))
labels = news.target_names
print(labels)
# + jupyter={"outputs_hidden": true} tags=[]
# Exemples d'articles et de labels
for i, article in enumerate(news.data[:10]):
print(f'===== {labels[news.target[i]]} =====')
print(article.replace('\n', ' '), '\n')
# + [markdown] tags=[]
# ## Création d'un modèle de machine learning avec Scikit-Learn
# Pour plus d'information :
# - Pipeline : https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html
# - TfidfVectorizer : https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfVectorizer.html
# - MultinomialNB : https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.MultinomialNB.html
#
# Un article de blog qui explique le TFIDF:
# - https://medium.com/analytics-vidhya/tf-idf-term-frequency-technique-easiest-explanation-for-text-classification-in-nlp-with-code-8ca3912e58c3
#
# Un article de blog qui explique les naive bayes:
# - https://towardsdatascience.com/naive-bayes-classifier-explained-54593abe6e18
# -
# ### Séparer le dataset en features et target (X, y) et en train et test
# Plus d'information : https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html
# Nettoyage des textes
texts = [re.sub('[^a-z]+', ' ', t.lower()).strip() for t in news.data]
# Mapping des targets
targets = np.array([labels[t] for t in news.target])
# +
X_train, X_test, y_train, y_test = train_test_split(texts, targets, test_size=0.2, random_state=11)
print("Training set size:", len(X_train))
print("Test set size:", len(X_test))
# -
# ### Entrainer un modèle de machine learning sur les données d'entrainement
# Définition du type de modèle
classifier = Pipeline([
('vectorizer', TfidfVectorizer(stop_words=stopwords.words('english'), min_df=50, max_df=0.5)),
('classifier', MultinomialNB()),
])
# Entrainement du modèle
classifier.fit(X_train, y_train)
# ### Qu'est ce qu'il s'est passé ?
# #### Le TFIDF calcule le score IDF de chaque mot du corpus
#
# + tags=[]
feature_names = classifier.named_steps['vectorizer'].get_feature_names_out()
idf_scores = classifier.named_steps['vectorizer'].idf_
# -
# Taille du vocabulaire (nombre total de mots différents)
len(feature_names)
# Score IDF de chaque terme du vocabulaire (score haut, mot raire)
for i in range(0, 10):
print(feature_names[i], ':', round(idf_scores[i], 2))
# + tags=[]
# Les 10 mots avec le score IDF le plus haut
for word, score in sorted(zip(feature_names, idf_scores), key=operator.itemgetter(1), reverse=True)[:20]:
print(word, round(score, 2))
# -
# #### Le TF-IDF transforme chaque document en vecteur de la taille du vocabulaire et donc le score est le TFIDF (fréquence du terme dans le document * idf)
tmp = classifier.named_steps['vectorizer'].transform(X_train[:10])
pd.DataFrame(tmp.toarray(), columns=classifier.named_steps['vectorizer'].get_feature_names_out())
# #### Le modèle naïf bayésien apprend la corrélation entre chaque mot et chaque catégorie
pd.DataFrame(classifier.named_steps['classifier'].feature_log_prob_, index=labels, columns=feature_names).T
# #### On peut ainsi découvrir les termes les plus contributifs pour un label donné
pd.DataFrame(classifier.named_steps['classifier'].feature_log_prob_, index=labels, columns=feature_names).T.sort_values(by='alt.atheism', ascending=False).head(20)
# ### Prédire les targets des données de test à l'aide du modèle entrainé
y_pred = classifier.predict(X_test)
# Aperçu des targets prédites
y_pred[:20]
# Aperçu des targets réelles
y_test[:20]
# ### Evaluer le modèle
# + [markdown] tags=[]
# #### Générer un rapport de classification
# Pour plus d'information sur la précision, le recall et le f1-score : https://fr.wikipedia.org/wiki/Pr%C3%A9cision_et_rappel
# -
print(classification_report(y_test, y_pred))
# ### Générer une matrice de confusion
plot_confusion_matrix(y_test, y_pred, figsize=(10, 10), labels=labels, x_tick_rotation=90)
|
module4/s1_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import re
import string
import pymorphy2
from scipy.sparse import *
import enchant
import stop_words
from nltk.stem.snowball import RussianStemmer
# %matplotlib inline
def log_progress(sequence, every=None, size=None, name='Items'):
from ipywidgets import IntProgress, HTML, VBox
from IPython.display import display
is_iterator = False
if size is None:
try:
size = len(sequence)
except TypeError:
is_iterator = True
if size is not None:
if every is None:
if size <= 200:
every = 1
else:
every = int(size / 200) # every 0.5%
else:
assert every is not None, 'sequence is iterator, set every'
if is_iterator:
progress = IntProgress(min=0, max=1, value=1)
progress.bar_style = 'info'
else:
progress = IntProgress(min=0, max=size, value=0)
label = HTML()
box = VBox(children=[label, progress])
display(box)
index = 0
try:
for index, record in enumerate(sequence, 1):
if index == 1 or index % every == 0:
if is_iterator:
label.value = '{name}: {index} / ?'.format(
name=name,
index=index
)
else:
progress.value = index
label.value = u'{name}: {index} / {size}'.format(
name=name,
index=index,
size=size
)
yield record
except:
progress.bar_style = 'danger'
raise
else:
progress.bar_style = 'success'
progress.value = index
label.value = "{name}: {index}".format(
name=name,
index=str(index or '?')
)
EXCLUDE_SYMBOLS_STR = u''.join(['№', '«', 'ђ', '°', '±', '‚', 'ћ', '‰', '…', '»', 'ѓ', 'µ', '·', 'ґ', 'њ', 'ї', 'џ', 'є', '‹',
'‡', '†', '¶', 'ќ', '€', '“', 'ў', '§', '„', '”', '\ufeff', '’', 'љ', '›', '•', '—', '‘',
'\x7f', '\xad', '¤', '\xa0'])
GRAMMS = ['NOUN', 'ADJF', 'ADJS', 'PRTF', 'PRTS', 'GRND', 'ADVB']
# [**Data**](http://study.mokoron.com/)
# ---
DATA_PATH = 'positive.csv'
df = pd.read_csv(DATA_PATH, header=None, sep=';',
names=['id', 'tdate', 'tmane', 'ttext', 'ttype', 'trep', 'trtw', 'tfav', 'tstcount',
'tfol', 'tfrien', 'listcount'])
print(df.shape)
df.head()
df.dtypes
# ---
# +
corpus, texts = [], []
for message in log_progress(df.ttext.dropna().as_matrix()):
sp = message.split('\n')
corpus += sp
texts += sp
# break
print(len(corpus), len(texts))
# -
corpus[1040]
# **Уберем ссылки:**
corpus = [re.sub(r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', u'', doc) for doc in corpus]
corpus = [re.sub(' +' , ' ', doc) for doc in corpus]
print(len(corpus))
corpus[1040]
# **Уберем лишние пробелы:**
corpus = np.asarray([doc.strip().strip('\t').replace('\n', u'') for doc in corpus])
print(len(corpus))
corpus[1040]
# **Уберем другие лишние символы:**
string.punctuation
regex_punct = re.compile('[%s]' % re.escape(string.punctuation))
regex_dig = re.compile('[%s]' % re.escape(string.digits))
regex_symb = re.compile('[%s]' % re.escape(EXCLUDE_SYMBOLS_STR))
regex_struct = re.compile('[%s]' % string.printable + string.whitespace)
emoji_pattern = re.compile("["
"\U0001F600-\U0001F64F" # emoticons
"\U0001F300-\U0001F5FF" # symbols & pictographs
"\U0001F680-\U0001F6FF" # transport & map symbols
"\U0001F1E0-\U0001F1FF" # flags (iOS)
"]+")
corpus = [regex_punct.sub('', doc) for doc in corpus]
print(len(corpus))
corpus = [regex_dig.sub('', doc) for doc in corpus]
print(len(corpus))
corpus = [regex_symb.sub(' ', doc) for doc in corpus]
print(len(corpus))
corpus = [regex_struct.sub('', doc) for doc in corpus]
print(len(corpus))
corpus = [re.sub(' +' , ' ', doc.strip()) for doc in corpus]
print(len(corpus))
corpus[1040]
texts = np.asarray([txt for doc, txt in zip(corpus, texts) if len(doc) > 0])
print(len(texts))
corpus = np.asarray([doc for doc in corpus if len(doc) > 0])
print(len(corpus))
corpus = [doc.lower() for doc in corpus]
print(len(corpus))
corpus[1040]
corpus = [emoji_pattern.sub('', doc) for doc in corpus]
print(len(corpus))
corpus[100]
morph = pymorphy2.MorphAnalyzer()
# +
corpus_tokens = []
inds_to_drop = []
for i, sentence in enumerate(log_progress(corpus[:])):
tmp_tokens = []
sp = sentence.split()
for word in sp:
if word not in stop_words.get_stop_words('ru'):
if morph.word_is_known(word):
tmp_tokens.append(word)
if len(tmp_tokens) > 0:
corpus_tokens.append(tmp_tokens)
else:
inds_to_drop.append(i)
# break
print(len(corpus_tokens), len(texts))
# -
print(len(texts))
texts = [t for i, t in enumerate(texts) if i not in inds_to_drop]
print(len(texts))
# + active=""
# dict_ru = enchant.Dict("ru_RU")
# + active=""
# corpus_tokens_ench = []
# inds_to_drop = []
#
# for i, tokens in enumerate(log_progress(corpus_tokens)):
# tmp_tokens = []
# for word in tokens:
# if dict_ru.check(word):
# tmp_tokens.append(word)
# if len(tmp_tokens) > 0:
# corpus_tokens_ench.append(tmp_tokens)
# else:
# inds_to_drop.append(i)
#
# print(len(corpus_tokens_ench), len(texts))
# + active=""
# print(len(texts))
# texts = [t for i, t in enumerate(texts) if i not in inds_to_drop]
# print(len(texts))
# -
stemmer = RussianStemmer()
# +
corpus_tokens_stem = []
for i, tokens in enumerate(log_progress(corpus_tokens[:])):
tmp = [stemmer.stem(word) for word in tokens]
corpus_tokens_stem.append(tmp)
# break
print(len(corpus_tokens_stem))
# -
corpus_tokens_stem[100]
# ---
# +
corpus_tokens_stem = np.asarray(corpus_tokens_stem)
texts = np.asarray(texts)
print(len(corpus_tokens_stem), len(texts))
# -
diffs = np.asarray([len(tokens) for tokens in corpus_tokens_stem])
print(len(diffs))
plt.figure(figsize=(15, 5))
plt.hist(diffs, bins=100)
plt.grid(True)
plt.xlim(0, 30)
corpus_tokens_stem[diffs == 2]
np.unique(corpus_tokens_stem).shape[0] / float(corpus_tokens_stem.shape[0])
# +
stat = {}
tmp_corp, tmp_texts = [], []
for i, tokens in enumerate(log_progress(corpus_tokens_stem)):
s = ' '.join(tokens)
if stat.get(s, None) is None:
stat[s] = True
tmp_corp.append(tokens)
tmp_texts.append(texts[i])
corpus_tokens_stem = tmp_corp
texts = tmp_texts
print(len(corpus_tokens_stem), len(texts))
# +
corpus_tokens_stem = np.asarray(corpus_tokens_stem)
texts = np.asarray(texts)
print(len(corpus_tokens_stem), len(texts))
# -
diffs = np.asarray([len(tokens) for tokens in corpus_tokens_stem])
print(len(diffs))
plt.figure(figsize=(15, 5))
plt.hist(diffs, bins=100)
plt.grid(True)
plt.xlim(0, 10)
corpus_tokens_stem[diffs == 7]
corpus_tokens_stem[diffs >= 8].shape
corp4learning = corpus_tokens_stem[diffs >= 8]
texts4learning = texts[diffs >= 8]
# +
tmp_corp = []
for tokens in corp4learning:
tmp_corp.append([t for t in tokens if len(t) > 2])
corp4learning = tmp_corp
# -
vocab = np.unique(np.concatenate(corp4learning).flatten()).tolist()
print(len(vocab))
|
class 2/p3. preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
# load data
booze = pd.read_csv("drinks.csv")
# pretty printing
booze
# series vs. dataframe
print(type(booze.country))
# Selecting
booze[booze.country == 'Canada']
# Sorting
booze.sort_values(by='beer_servings')
# Automatic statistics w/ describe()
booze.describe()
|
Section 1/Video 1.5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from gensim.models import word2vec
import seaborn as sns
import matplotlib.pyplot as plt
import re
import nltk
nltk.download('stopwords')
df_path = '../Data/movies_df.csv'
df = pd.read_csv(df_path)
df['overview'][:5]
# ## Clean data
def clean(text): # use regular expression to remove specific characters
text = re.sub("\'", " ", text)
text = re.sub("[^a-zA-Z]"," ",text)
text = ' '.join(text.split())
text = text.lower()
return text
df['clean_overview'] = df['overview'].astype(str).apply(lambda x: clean(x))
df.head()
def word_counts(text, terms):
all_words = ' '.join([t for t in text])
all_words = all_words.split()
fdist = nltk.FreqDist(all_words)
words_df = pd.DataFrame({'word':list(fdist.keys()), 'count':list(fdist.values())})
d = words_df.nlargest(columns="count", n = terms)
plt.figure(figsize=(20,30))
ax = sns.barplot(data=d, x= "count", y = "word")
ax.set(ylabel = 'Word')
plt.show()
word_counts(df['clean_overview'], 200)
stpwrds = set(nltk.corpus.stopwords.words('english'))
def remove_stops(text):
cleaned = [w for w in text.split() if not w in stpwrds]
return ' '.join(cleaned)
df['clean_overview'] = df['clean_overview'].astype(str).apply(lambda x: remove_stops(x))
word_counts(df['clean_overview'], 200)
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
tfidf_vectorizer = TfidfVectorizer(max_df=0.8, max_features=10000)
df.columns
X = df['clean_overview']
y = df[['Family', 'Animation',
'History', 'Documentary', 'Adventure', 'Western', 'Crime', 'Drama',
'Horror', 'Science Fiction', 'Romance', 'War', 'Mystery', 'Fantasy',
'Action', 'TV Movie', 'Thriller', 'Comedy', 'Music']]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
tfidf_xtrain = tfidf_vectorizer.fit_transform(X_train)
tfidf_xtest = tfidf_vectorizer.transform(X_test)
# +
from sklearn.svm import SVC
from sklearn.multiclass import OneVsRestClassifier
from sklearn.metrics import classification_report
# -
model = OneVsRestClassifier(SVC(gamma = 'auto',kernel = 'linear'))
model.fit(tfidf_xtrain, y_train)
predictions = model.predict(tfidf_xtest)
print(classification_report(predictions,y_test))
# ### Trying something different for classifiers
cm = df.loc[0]
ls = df.loc[1]
def make_genre_list(s):
genre_list = ['Family', 'Animation',
'History', 'Documentary', 'Adventure', 'Western', 'Crime', 'Drama',
'Horror', 'Science Fiction', 'Romance', 'War', 'Mystery', 'Fantasy',
'Action', 'TV Movie', 'Thriller', 'Comedy', 'Music']
gen = []
for g in genre_list:
if s.loc[g] == 1:
gen.append(g)
return gen
make_genre_list(cm)
make_genre_list(ls)
df['genre_list'] = df.apply(lambda row : make_genre_list(row), axis = 1)
from sklearn.preprocessing import MultiLabelBinarizer
# +
multilabel_binarizer = MultiLabelBinarizer()
multilabel_binarizer.fit(df['genre_list'])
y = multilabel_binarizer.transform(df['genre_list'])
# -
X = df['clean_overview']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
tfidf_xtrain = tfidf_vectorizer.fit_transform(X_train)
tfidf_xtest = tfidf_vectorizer.transform(X_test)
model = OneVsRestClassifier(SVC(gamma = 'auto', kernel = 'linear'))
model.fit(tfidf_xtrain, y_train)
predictions = model.predict(tfidf_xtest)
print(classification_report(predictions,y_test))
len(y_test)
len(predictions)
# +
true = 0
false = 0
pred = []
test = []
for index, row in enumerate(y_test):
if np.any(row & predictions[index]):
pred.append(True)
test.append(True)
else:
pred.append(False)
test.append(True)
# -
from sklearn.linear_model import LogisticRegression
logovr = OneVsRestClassifier(LogisticRegression())
logovr.fit(tfidf_xtrain, y_train)
predicted = logovr.predict(tfidf_xtest)
print(classification_report(predicted, y_test))
mbi = multilabel_binarizer.inverse_transform(predicted)
y_test_df = pd.DataFrame()
y_test_df['genres'] = y_test.apply(lambda row : make_genre_list(row), axis = 1)
ytgenres = list(y_test_df['genres'])
for i in range(len(mbi)):
print(f'{mbi[i]}\t\t\t\t{ytgenres[i]}')
df.head()
genre_df = df[['Family', 'Animation',
'History', 'Documentary', 'Adventure', 'Western', 'Crime', 'Drama',
'Horror', 'Science Fiction', 'Romance', 'War', 'Mystery', 'Fantasy',
'Action', 'TV Movie', 'Thriller', 'Comedy', 'Music']]
genre_counts = genre_df.apply(pd.Series.value_counts)
genre_counts = genre_counts.loc[1]
gct = genre_counts.transpose()
gct = pd.DataFrame(gct)
gct.columns
gct = gct.reset_index()
gct.columns
gct.columns = ['genres', 'count']
plt.figure(figsize=(15,20))
sns.barplot( x = 'count',y = 'genres', data=gct)
|
Scripts/DataAnalysis_aaron_TFIDF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ctyzNbUYHqWx" colab_type="text"
# to do if there is time:
#
# - least similar song search for funsies
#
# - create visualizations
# + [markdown] id="TMDW0v1XROm_" colab_type="text"
# ## Imports
# + id="8lgM2EHbRQ-_" colab_type="code" outputId="98545e6e-9b45-4eb2-b6a2-e3293895f647" colab={"base_uri": "https://localhost:8080/", "height": 214}
# !pip install spotipy
# https://spotipy.readthedocs.io/en/2.12.0/
# + id="T4i1KpT2RTrV" colab_type="code" outputId="4a483980-868e-4260-f637-ddad307cba11" colab={"base_uri": "https://localhost:8080/", "height": 552}
# !pip install spotify
# + id="QD1XyjiAQ9j5" colab_type="code" colab={}
import pandas as pd
import numpy as np
import spotipy
from spotipy import oauth2, Spotify
from sklearn.neighbors import NearestNeighbors
from sklearn.preprocessing import StandardScaler
# + [markdown] id="QKT360cWRL78" colab_type="text"
# ## Spotify Authentication
# + id="LwLcujQSRY3X" colab_type="code" colab={}
# Spotify API authentication
cid ='555816fae63a45f29f50f99bbc81393d'
secret ='008dc4e8c3e3470c97392be068578540'
credentials = oauth2.SpotifyClientCredentials(client_id=cid, client_secret=secret)
sp = spotipy.Spotify(client_credentials_manager=credentials)
# + [markdown] id="zS_Y96qkRdi-" colab_type="text"
# ## Track Info Functions
# + id="o05CapOeRfne" colab_type="code" colab={}
def fetch_info(artist_name, track_name):
"""
function to retrieve relevant info on user's input from the Spotify API
user inputs artist name and track name
fxn gets info including the unique track id, album name, id, and cover image,
a sample of the track if exists, and audio feature data for the track
"""
data = sp.search(q=f'artist:{artist_name} track:{track_name}')
artist_name = data['tracks']['items'][0]['artists'][0]['name']
track_name = data['tracks']['items'][0]['name']
track_id = data['tracks']['items'][0]['id']
album_name = data['tracks']['items'][0]['album']['name']
album_id = data['tracks']['items'][0]['album']['id']
album_cover_link = data['tracks']['items'][0]['album']['images'][0]['url']
track_sample = data['tracks']['items'][0]['preview_url']
audio_features = sp.audio_features(track_id)
audio_features = audio_features[0] # changes the provided list from spotify into a dictionary
irrelevant = ["id", "uri", "analysis_url", "type", "track_href"]
for key in irrelevant:
del audio_features[key]
#audio_features = jsonify(audio_features) in FLASK app
return track_name, artist_name, track_id, album_name, album_id, album_cover_link, track_sample, audio_features
# + id="aNKwTE-ARqHu" colab_type="code" outputId="e8ea3cf1-8ac0-4520-8407-2789c082f760" colab={"base_uri": "https://localhost:8080/", "height": 372}
fetch_info("Tame Impala", "Borderline")
# + id="isZc7w-XRtY6" colab_type="code" colab={}
def track_feat(track_id):
"""
function to retrieve relevant info from track ID and store it in a df
"""
track_features = sp.audio_features(track_id)
track_features = pd.DataFrame(track_features[0], index=[0])
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
return track_features
# + id="D81nrWunR9vh" colab_type="code" outputId="8cfcadfc-2a76-4b6a-cfd2-d022d94c04e6" colab={"base_uri": "https://localhost:8080/", "height": 77}
track_feat('5hM5arv9KDbCHS0k9uqwjr')
# + [markdown] id="NSjjoYHCSRmI" colab_type="text"
# ## Database
#
# + id="S7GAgg5LSS0S" colab_type="code" outputId="6427a2a0-2d41-4e5c-c040-771b9672322e" colab={"base_uri": "https://localhost:8080/", "height": 385}
# 5/27 created new dataset with top tracks from the past two years
tracks = pd.read_csv("https://raw.githubusercontent.com/Build-Week-Spotify-Song-Suggester-2/datascience/master/SpotifyTrackFeatures.csv", error_bad_lines=False)
print(tracks.shape)
tracks.head()
# + id="AkhQPvNeSV0-" colab_type="code" outputId="ee92b801-f367-4ea8-eca6-cdbf7ba5a740" colab={"base_uri": "https://localhost:8080/", "height": 336}
tracks = pd.DataFrame(tracks)
tracks.dtypes
# + id="0HQ6X9kOPIbj" colab_type="code" outputId="ea92c365-eb09-4229-e719-f6657e14238b" colab={"base_uri": "https://localhost:8080/", "height": 34}
tracks = tracks.drop_duplicates(['track_id'])
tracks.shape
# + [markdown] id="N63HqMS0Sslp" colab_type="text"
# ## KNN
#
# section for creating and tweaking KNN model
# + id="vn6T5f5MStzJ" colab_type="code" colab={}
# scale data and create model
mdf = tracks # dataset
# track_id = '5hM5arv9KDbCHS0k9uqwjr'
target = ['track_id']
features = ['danceability',
'energy',
'key',
'loudness',
'mode',
'speechiness',
'acousticness',
'instrumentalness',
'liveness',
'valence',
'tempo'] # duration, popularity, and time are not good predictors of similarity
# features = features.danceability.str.strip(" ","")
X = mdf[features] # .astype('float64')
y = mdf[target]
# Scale the features to a mean of 0 and a standard deviation of 1.
#scaler = StandardScaler()
# X = scaler.fit_transform(X)
nn = NearestNeighbors(n_neighbors=11, algorithm='kd_tree').fit(X, y) # 11 for top 10 similarities
# + id="VapqmazwTEVi" colab_type="code" colab={}
def testmodel(track_id, ouput = mdf):
"""
function for testing KNN
"""
mdf = tracks
# target = ['track_id']
features = ['danceability',
'energy',
'key',
'loudness',
'mode',
'speechiness',
'acousticness',
'instrumentalness',
'liveness',
'valence',
'tempo']
# get track audio features from track id and put into a df
track_features = sp.audio_features(track_id)
track_features = pd.DataFrame(track_features[0], index=[0])
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
# apply model to get prediciton
pred = nn.kneighbors(track_features)
return pred
# + id="fYy2Kgzm74LX" colab_type="code" colab={}
track_features = sp.audio_features('0SUClY63fA1awioMFtMYeE')
track_features = pd.DataFrame(track_features[0], index=[0])
# + id="Twa4y63J8Dhu" colab_type="code" outputId="4dc32788-0cd2-48bf-efae-c201e4ee5a38" colab={"base_uri": "https://localhost:8080/"}
track_features
# + id="2kY3BexHTYCI" colab_type="code" outputId="a032734e-0266-4ae4-9a67-3c84788bd71d" colab={"base_uri": "https://localhost:8080/"}
testmodel('0SUClY63fA1awioMFtMYeE', mdf)
# run model on two tracks to make sure it is actually working/returning different data
# + id="1bqNS4NjTVTx" colab_type="code" outputId="529bd8fc-9508-4a10-94c3-1fd13fe80f8b" colab={"base_uri": "https://localhost:8080/"}
testmodel('5hM5arv9KDbCHS0k9uqwjr', mdf)
# cool it works, lets try and change the output into something a human will understand
# + id="HVXJ5ssKTkaZ" colab_type="code" colab={}
def predict(track_id, ouput = mdf):
"""
track recommender function
takes track_id input and ouputs a list of 10 similar tracks with artist name
"""
mdf = tracks
features = ['danceability',
'energy',
'key',
'loudness',
'mode',
'speechiness',
'acousticness',
'instrumentalness',
'liveness',
'valence',
'tempo']
# get track audio features via track id and put into a df
track_features = sp.audio_features(track_id)
track_features = pd.DataFrame(track_features[0], index=[0])
#track_features = track_features.astype(({'track_id': 'int32'}))
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
pred = nn.kneighbors(track_features)
return_rec = {}
for index,i in enumerate(pred[1][0]):
return_rec[str(index)] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
# probably have to jsonify this in FLASK
return return_rec
# + id="i3rQJRE98tpg" colab_type="code" colab={}
track_features = sp.audio_features('5hM5arv9KDbCHS0k9uqwjr')
track_features = pd.DataFrame(track_features[0], index=[0])
#track_features = track_features.astype(({'track_id': 'int32'}))
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
pred = nn.kneighbors(track_features)
# + id="88XFmBKa9JJw" colab_type="code" outputId="6593e53d-dbcd-43b9-941b-0cf8d0deee84" colab={"base_uri": "https://localhost:8080/"}
pred
# + id="imfVzvST8zSY" colab_type="code" colab={}
return_rec = {}
for index,i in enumerate(pred[1][0]):
return_rec[str(index)] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
# probabrely have to jsonify this in FLASK
# + id="zmDy4l5JAfFR" colab_type="code" outputId="2de05632-2b1b-4064-d3ec-bd9f6723f2ac" colab={"base_uri": "https://localhost:8080/"}
return_rec
# + id="0VaFFP-w9O9n" colab_type="code" outputId="6573bc76-d525-4943-8b98-36107e05ef24" colab={"base_uri": "https://localhost:8080/"}
return_rec = {}
arr = pred[1][0]
for index,i in enumerate(arr):
return_rec[str(index)] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
return_rec
# + id="qgTO3mNs9ZOC" colab_type="code" outputId="ca7a4011-f424-47c6-d66e-f86d9325d5df" colab={"base_uri": "https://localhost:8080/"}
return_rec = {}
for i in arr:
return_rec[i] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
return_rec
# + id="hB73ZVbh9xsM" colab_type="code" outputId="268a6cbf-4596-49f5-f56c-ec26bf2d647d" colab={"base_uri": "https://localhost:8080/"}
return_rec
# + id="VLkrViq3TuSN" colab_type="code" outputId="d4f1aac5-6d1c-4031-84dc-726ef3e9547a" colab={"base_uri": "https://localhost:8080/"}
predict('0SUClY63fA1awioMFtMYeE', mdf)
# so model works but readable output function isnt working as intended
# + id="77IsfjSAT6hx" colab_type="code" outputId="ddca48dd-d99c-4458-dd07-c240c18420f5" colab={"base_uri": "https://localhost:8080/"}
predict('5hM5arv9KDbCHS0k9uqwjr', mdf)
# seems to be returning just a list of tracks from the dataset instead of the actual model output(?)
# + id="gWFhiZAWGvYJ" colab_type="code" outputId="c7f4cf8f-1747-41c0-b2f8-cbd65de6de22" colab={"base_uri": "https://localhost:8080/"}
list_of_predictions = pred[1][0].tolist()
list_of_predictions
# + id="6M9GhKSnF32V" colab_type="code" colab={}
def predict(track_id, ouput = mdf):
"""
track recommender function
takes track_id input and ouputs a list of 10 similar tracks with artist name
"""
mdf = tracks
features = ['danceability',
'energy',
'key',
'loudness',
'mode',
'speechiness',
'acousticness',
'instrumentalness',
'liveness',
'valence',
'tempo']
# get track audio features via track id and put into a df
track_features = sp.audio_features(track_id)
track_features = pd.DataFrame(track_features[0], index=[0])
#track_features = track_features.astype(({'track_id': 'int32'}))
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
pred = nn.kneighbors(track_features)
return_rec = {}
arr = pred[1][0]
for i in arr:
return_rec[i] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
# probably have to jsonify this in FLASK
return return_rec
# + [markdown] id="R1YGPZw5G-38" colab_type="text"
# ### add scaling IGNORE
#
# + id="kGRQJz10HAVU" colab_type="code" colab={}
def Spredict(track_id, ouput = mdf):
"""
track recommender function
takes track_id input and ouputs a list of 10 similar tracks with artist name
"""
scaler = StandardScaler()
mdf = tracks
features = ['danceability',
'energy',
'key',
'loudness',
'mode',
'speechiness',
'acousticness',
'instrumentalness',
'liveness',
'valence',
'tempo']
# get track audio features via track id and put into a df
track_features = sp.audio_features(track_id)
track_features = pd.DataFrame(track_features[0], index=[0])
#track_features = track_features.astype(({'track_id': 'int32'}))
track_features = track_features.drop(columns=["id", "uri", "analysis_url", "type", "track_href", "duration_ms", "time_signature"])
scaled_feat = scaler.fit_transform(track_features)
pred = nn.kneighbors(scaled_feat)
return_rec = {}
for index,i in enumerate(pred[1][0]):
return_rec[str(index)] = {"track_name" : tracks['track_name'].iloc[i],
"artist" : tracks['artist_name'].iloc[i]}
# probably have to jsonify this in FLASK
return return_rec
# + id="XAyxvs2WHp2N" colab_type="code" outputId="e7b5ac31-64a3-481f-e886-3a070ae555d9" colab={"base_uri": "https://localhost:8080/"}
# test scaling
# input song not scaled
# Tame Impala Borderline
predict('5hM5arv9KDbCHS0k9uqwjr', mdf)
# + id="CalRyedaHZhu" colab_type="code" outputId="e3a8da8b-7cdc-4824-ae8f-d87fd370e04a" colab={"base_uri": "https://localhost:8080/"}
# input song scaled
Spredict('5hM5arv9KDbCHS0k9uqwjr', mdf)
# + id="92YlZo76Hjqg" colab_type="code" outputId="981f91a0-8c2e-4eea-99b5-7e35d3d161b2" colab={"base_uri": "https://localhost:8080/"}
# britney spears toxic
# no scaling
predict('4fbaKWFRghusXd4bSBvvfN', mdf)
# + id="20jArDGsH8sv" colab_type="code" outputId="75c16b69-c325-424f-9ea9-2441e2c86dbf" colab={"base_uri": "https://localhost:8080/"}
# input song scaled
Spredict('4fbaKWFRghusXd4bSBvvfN', mdf)
# + id="tUc1bwIzMlhk" colab_type="code" colab={}
# + [markdown] id="CxUigVc62VaS" colab_type="text"
# ## KNN v2
# created to help compare our KNN vs NN models
# scaled , using slightly different input system
# got incredibly similar results to first KNN
# + id="CFPTaIpW2rvK" colab_type="code" colab={}
# test scaling and remake model
# + id="FamDKslC2XO2" colab_type="code" outputId="0d5935c5-2914-423d-d5c3-337864d12898" colab={"base_uri": "https://localhost:8080/", "height": 367}
modeldf = tracks.copy()
modeldf["track_index"] = modeldf.index +1
dictionary = modeldf[["artist_name", "track_name", "track_id", "track_index"]]
modeldf.head()
# + id="cyk6JhWu5hE-" colab_type="code" outputId="6cf7e88b-cb78-4b85-bd17-b0468ab12e11" colab={"base_uri": "https://localhost:8080/", "height": 217}
modeldf = modeldf.drop(columns=['artist_name','track_id', 'track_name','duration_ms', 'mode', 'time_signature'])
modeldf.head()
# + id="7oyYdNtl2rZa" colab_type="code" colab={}
def predictv2(track_index, ouput = mdf):
"""
track recommender function
takes track_id input and ouputs a list of 10 similar tracks with artist name
"""
#scale
scaler = StandardScaler()
df_scaled = scaler.fit_transform(modeldf)
# input track index to actual track index
input_dict = dictionary[dictionary['track_index']==track_index]
given_index = input_dict.index
# KNN model, 11 for top 10
nn = NearestNeighbors(n_neighbors=11, algorithm='kd_tree')
nn.fit(df_scaled)
pred = nn.kneighbors(df_scaled[given_index])
# create list of recommended track indexes
pred_list = pred[1][0].tolist()
# output top 10 recommended tracks
top10 = []
for item in pred_list:
rec_track = dictionary['track_id'].iloc[item]
top10.append(rec_track)
return return_rec
# + id="K8thqAUt7yFY" colab_type="code" outputId="fc7d16d2-e863-469e-b27d-089f6905481f" colab={"base_uri": "https://localhost:8080/", "height": 52}
print(dictionary[dictionary['track_id']=='5hM5arv9KDbCHS0k9uqwjr'])
# + id="OOQKPgoU78w7" colab_type="code" outputId="11142cc3-b20a-4b31-a020-bf5b9055289e" colab={"base_uri": "https://localhost:8080/", "height": 212}
predictv2(118604)
# + id="kuryoy1z8GwB" colab_type="code" outputId="f1cbb02f-8ead-4431-b75c-7768ecb976cd" colab={"base_uri": "https://localhost:8080/", "height": 212}
predict('5hM5arv9KDbCHS0k9uqwjr', mdf)
# returns essentially the same resuls as the 1st KNN
# + [markdown] id="Fw8rxyyZMmKR" colab_type="text"
# ## K Means
#
# did not turn out as planned, not using
# + id="8XwEUYdpPzmH" colab_type="code" colab={}
from sklearn.preprocessing import scale
from sklearn.preprocessing import MinMaxScaler
from sklearn.cluster import KMeans
# + id="4AVNrBQwMxNa" colab_type="code" outputId="50a24f84-2cd2-447f-f855-584b5b7c316d" colab={"base_uri": "https://localhost:8080/", "height": 367}
tracks.head()
# + id="7aMYj_-HPfm5" colab_type="code" colab={}
numeric_features = ['acousticness','danceability','energy','instrumentalness',
'key', 'liveness', 'loudness','mode','speechiness', 'tempo', 'valence']
tracks[numeric_features] = tracks[numeric_features].astype(float)
# + id="qCQDffMrPk5d" colab_type="code" colab={}
scaler = MinMaxScaler()
numeric_features_scaled = scaler.fit_transform(tracks[numeric_features])
# + id="TLP_HMfsMnsw" colab_type="code" colab={}
Sum_of_squared_distances = []
Ks = range(1,15)
for n in Ks:
km = KMeans(n_clusters=n)
km = km.fit(numeric_features_scaled)
Sum_of_squared_distances.append(km.inertia_)
# + id="Nc6_vcbNMnu1" colab_type="code" colab={}
# score with elbow plot
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(15, 5))
plt.plot(Ks, Sum_of_squared_distances, 'gx-')
plt.xlabel('K')
plt.ylabel('Sum of Squared Distances')
plt.ylim(1000, 100000)
plt.title('Elbow Curve for Optimal K')
plt.show()
# + id="vEnccRrURGit" colab_type="code" outputId="2f3765d1-e194-4574-8dcc-590f0d11ee77" colab={"base_uri": "https://localhost:8080/", "height": 247}
# score with silhouette
# https://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html
from sklearn.metrics import silhouette_score
for n_clusters in range(2,15):
clusterer = KMeans(n_clusters=n_clusters)
predictions = clusterer.fit_predict(numeric_features_scaled)
centers = clusterer.cluster_centers_
score = silhouette_score (numeric_features_scaled, predictions, metric='euclidean')
print (f'n_clusters = {n_clusters}, silhouette score = {score})')
# + id="d_5Q0ll0bgFn" colab_type="code" outputId="3a935835-fcf6-447b-d951-b5668e9e4405" colab={"base_uri": "https://localhost:8080/", "height": 70}
np.random.seed(42)
kmeans = KMeans(n_clusters=5)
kmeans.fit(numeric_features_scaled)
# + id="eKJxmg2dbyXT" colab_type="code" outputId="3c31d480-00b1-4452-c92e-792b8caea74f" colab={"base_uri": "https://localhost:8080/", "height": 424}
# visualize
# PCA
import seaborn as sns
from sklearn.decomposition import PCA
labels_kmeans = kmeans.predict(numeric_features_scaled)
pca = PCA(n_components=2)
principal_components = pca.fit_transform(numeric_features_scaled)
# plotting clusters
pc = pd.DataFrame(principal_components)
pc['label'] = labels_kmeans
pc.columns = ['x', 'y','label']
cluster = sns.lmplot(data=pc, x='x', y='y', hue='label',
fit_reg=False, legend=True, legend_out=True)
# + id="OCD1CJMtcGnB" colab_type="code" colab={}
|
Notebooks/Spotify_Rec_KNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of the Curse Log
# ## Introduction
# With this worksheet, you will learn the first steps with Jupyter, Python, pandas and matplotlib using a practical example: We execute an analysis of the futuristic data set "curse log" – a log of time-tracked swearwords events gathered with the microchip implant "FUTURE 2000" from several humans.
#
# We managed to get the log of uttered curses of some users. We also have some user profile data that we can combine with the curse log to find out, who the cursers are.
# ## Tasks
# * We want to find out
# * which curses are the most popular
# * at what hour of the day most curses are made.
# * which people are in the TOP 10 cursers' list
# * if men curse more often than woman
# * the favorite curse for each job
#
# As a starting point, we have a log from several users recorded in a file that lists the time stamp, the curse word and the id of the user's profile each curse:
#
# ```
# timestamp,curse,profile_id
# 2132-12-31 14:47:43,The A-word,0
# 2132-12-31 13:13:56,The F-word,0
# 2132-12-31 13:03:05,The S-word,0
# 2132-12-31 12:30:34,The F-word,0
# 2132-12-31 12:29:02,The S-word,0
# ```
#
# Hint: This data is a data set from the future. But since time machines aren't invented yet, we are working with a generated / synthetic dataset based on a real data set from an other domain.
#
# Let's get to know the tools we use!
# ## Jupyter
# Jupyter offers us code and documentation in executable **cells**.
#
# ##### Code execution
#
# 1. select the next cell (mouse click or arrow keys).
# 1. execute the cell with a `Ctrl`+`Enter`.
# 1. move the cell again with `Shift`+`Enter`. What's the difference between the output of results?
"Hello World"
# ##### Create new cell
# 1. if it hasn't happened yet, select this cell.
# 1. enter **command mode**, selectable with `ESC` key.
# 1. create a **new cell** after this text with the `b` key.
# 1. change the **cell type** to "Markdown" with key `m`.
# 1. switch to **edit mode** with `Enter` *(note the color to the left of the cell, which turns green instead of blue)*.
# 1. write a text, which you then "execute" with `Ctrl` + `Enter`.
# This is a text
# ## Python
# We look at very basic functions:
#
# - variable assignments
# - value range accesses
# - method calls
# #### Assign text to a variable
# 1. **assign** the text **value** "Hello World" to the **variable** `text` by using the syntax `<variable> = <value>`.
# 1. type the variable `text` in the next line and execute the cell.
text = "Hello World"
text
# ##### Access values
# 1. access the first letter in `text` with `[0]`.
text[0]
# ##### Select last value
# 1. access the last letter in `text` with `[-1]`.
text[-1]
# ##### Select value ranges
# 1. access a range of `text` with the **slice** `[2:5]`.
text[2:4]
# #### Auto completion
# 1. append a `.` to `text` and look at the functions with the `Tab` key.
# 1. execute the **method** `upper()` (Tip: Type a `u` in the function overview).
text.upper()
# #### Interactive documentation
# 1. select the `split` function of `text`.
# 1. press `Shift`+`Tab`.
# 1. press `Shift`+`Tab` twice in quick succession.
# 1. press `Shift`+`Tab` four times in quick succession (and then `ESC` to hide)
# 1. split the text in `text` with `split` exactly once (parameter `maxsplit`) apart by using the `l` ("L") as separator (parameter `sep`).
text.split("l",maxsplit=1)
# ## Pandas
# ### Import data
#
# #### Load a module
# 1. import the module `pandas` with `import <module> as <abbreviation>` as abbreviated `pd`
# 1. in the next line, attached a `?` to `pd` and execute the cell to get some information about `pd`.
# +
import pandas as pd
# pd?
# -
# #### Import file
# 1. use the `read_csv` method to read the data from the file `curse_log.csv`.
# 1. write the result into the variable `log`.
# 1. display the first five entries (= curses) in `log` with the `head()` method.
log = pd.read_csv("dataset/curse_log.gz")
log.head()
# #### Getting to know the data
# 1. call `info()` on `log`.
log.info()
# We see that `log` is
# * a **DataFrame** that consists of
# * three columns (so-called **Series**) `timestamp`, `curse` and `profile_id`
# ### Aggregate data
# #### Top curses
# 1. sum up the number of curses in the column `curse` in `log` with the method `value_counts()`.
# 1. save the result in the variable `top_curses`.
# 1. list the result in `top_curses`.
#
# *Note: In this tutorial, we access Series directly with the `.<Series>` notation (e. g. `log.curse`). This works only if the names of the Series are different from the provided functions of a Series. E. g. it doesn't work, when you try to access a Series named `count`, because `count()` is a function of a Series. Here, you have to use the `['<Series name>']` notation (e.g. `log['count']`. When in doubt, always use the `['<Series name>']` notation (but which disables the auto-completion feature)*
top_curses = log.curse.value_counts()
top_curses
# ## matplotlib
# ### Visualization
#
# #### Plot diagram
# 1. tell Jupyter with `%matplotlib inline` to display generated graphics directly in the notebook.
# 1. create a diagram of the Series `top_curses` with `plot()`.
# %matplotlib inline
top_curses.plot()
# #### Create a bar chart
# 1. call the `bar()` sub-method of `plot` for the data in `log`.
top_curses.plot.bar()
# #### Improve the output
# 1. add a `;` to the call above and re-execute it. * What has changed? *
top_curses.plot.bar();
# #### Create a pie chart
# 1. call the `pie()` sub-method of `plot` for the data in `top_curses`.
top_curses.plot.pie();
# #### Create a beautiful pie chart
# 1. create another pie chart, this time directly with `plot()` of the Series `top_cursess` and with the following parameters:
# * `kind="pie"`
# * `figsize=[7,7]`
# * `title="Top curses"`
# * `label=""`
#
# Tip: Use auto completion.
top_curses.plot(
kind='pie',
title="Top curses",
label="",
figsize=[5,5]);
# ## Time series analysis
# ##### View timestamp column
# 1. display the first five entries of the series `timestamp`.
log.timestamp.head()
# #### Data type conversion
# 1. use the pandas function `pd.to_datetime` to convert the column `timestamp` into a real date data type.
# 1. write the result into the new variable `ts` (abbreviation for "timestamp").
# 1. output the first five entries.
ts = pd.to_datetime(log.timestamp)
ts.head()
# #### Assigning data to a Series
# 1. override the values of the `timestamp` Series with the data in `ts`.
# 1. print the first entries.
log['timestamp'] = ts
log.head()
# #### Working with hourly data
# 1. access the date object `dt` of the Series `timestamp`.
# 1. inspect the hours of the `hour` property of the `dt` object.
# 1. store the hours into the new Series `hour` of the `log` DataFrame.
# 1. print out the first five entries.
log['hour'] = log.timestamp.dt.hour
log.head()
# ##### Find out favorite cursing times
# 1. sum up the number of curses per each hour.
# 1. here, switch off the sorting with the parameter `sort=False`.
# 1. save the result in `curses_per_hour`.
curses_per_hour = log.hour.value_counts(sort=False)
curses_per_hour.head()
# #### Visualize the hourly cursing result
# 1. plot a bar chart of the hourly cursing counts.
curses_per_hour.plot.bar();
# ## Merging data sets
# Now it's time to find out, which users are cursing the most. We have another data set in a CSV file `profiles.csv.gz` with the following content:
#
# 
#
# The columns contain this information for all FUTURE 2000 users. It includes the unique identification number of a user's profile (matches `profile_id` in the curse log) as well as the name, birth date, sex and current job.
#
# We combine this data with our `log` DataFrame to check off the remaining items on our to-do list:
#
# Find out
# * which people are in the TOP 10 cursers' list
# * if men curse more often than woman
# * the favorite curse for each job
# So let's do it!
# #### Read in CSV file
# 1. use the `read_csv` method of Pandas to read in the file `profiles.csv.gz` into the DataFrame / variable `profiles`.
# 1. display the first five rows of the DataFrame.
profiles = pd.read_csv("dataset/profiles.csv.gz", sep=";", encoding='latin-1', index_col=0)
profiles.head()
# #### Join datasets
# 1. use the method `join()` on the `log` DataFrame
# 1. as first argument, put in the DataFrame `profiles`
# 1. as second argument, add the parameter `on='profile_id'` to join `log`'s `profile_id` column with the id (=index) column of the `profiles` data set.
# 1. store the result into the variable `curse_profiles`.
# 1. display the first entries of `curse_profiles`.
curse_profiles = log.join(profiles, on='profile_id')
curse_profiles.head()
# ### TOP 10 cursers
# #### Find the users with the most uttered curses
# 1. Count the top 10 curses
curse_profiles.name.value_counts().head(10)
# ### Cursing genders
# #### Get the ratio between male to all curses in percent
# 1. select only men (value `'M'`) with the selector notation `<DataFrame>[<DataFrame>.<Series> == <value>]`
# 1. count the number of returned rows of the Series `sex`.
# 1. divide that number by all entries of the Series `sex` of the DataFrame `curse_profiles`.
curse_profiles[curse_profiles.sex == 'M'].sex.count() / curse_profiles.sex.count()
# ### Favorite curse per job (advanced level)
# #### Grouping data
# 1. group together the `curse_profiles`' data along `job` and `curse` by using `groupby` and the list `['job', 'curse']` as argument.
# 1. count the values for the Series `sex` (or any other left Series).
# 1. store the returned Series into the variable `job_curses`.
# 1. display the first 10 entries of the Series.
job_curses = curse_profiles.groupby(['job', 'curse']).sex.count()
job_curses.head(10)
# #### Find the maximum per group
# 1. group `job_curses` again along `job`.
# 1. use the `transform` method with the argument `'max'`.
# 1. store the result in `max_per_group`.
# 1. print the first rows of the result.
max_per_job = job_curses.groupby('job').transform('max')
max_per_job.head()
max_per_job.sort_values(ascending=False)
# #### Filter maximum group values
# 1. filter with a selector the max values per group.
# 1. store the result in `favorite_curses_per_job`.
# 1. print the first rows.
favorite_curses_per_job = job_curses[job_curses == max_per_job]
favorite_curses_per_job.head()
# #### Count favorite curses
# 1. use `reset_index()` on `favorite_curses_per_job` to get rid of the grouping index.
# 1. count the occuring values for the `curse` Series
favorite_curses_per_job.reset_index().curse.value_counts()
# #### Unstack Series
# 1. use `unstack()` on `favorite_curses_per_job`
# 1. store the result in `favorite_curse_words`
favorite_curse_words = favorite_curses_per_job.unstack()
favorite_curse_words.head()
# #### Identify outliers
# 1. Show the jobs in `favorite_curse_words` that have the curse word `'The S-word'` as favorite.
favorite_curse_words[~favorite_curse_words['The S-word'].isnull()]
# #### The end
# If you read this: Well done! You did it!
#
# ## What's missing
# You have now learned some basics about pandas. This will get us a long way in our daily work. The other important topics that are still missing are:
# * reading in complicated, semi-structured data structures
# * cleansing of poor data
# * merging different data sources with `merge`
# * transforming of DataFrames with `pivot_table`.
#
# ## Summary
# I hope that this mini-tutorial will show you the potential of data analysis using Jupyter, Python, pandas and matplotlib!
#
# I am looking forward to your comments and feedback!
#
#
# ## Contact
#
#
# **<NAME>**
# Blog: https://www.feststelltaste.de
# Mail: <a href="mailto:<EMAIL>"><EMAIL></a>
# Twitter: [@feststelltaste](https://twitter.com/feststelltaste)
# Consulting and training: http://markusharrer.de
|
Curse Log Analysis Solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## **NASA Analysis**
#
# This is a challenge sent by Semantix. The objective of this challenge, is analyze and extract information from NASA request dataset. There is two datasets on this link [NASA request dataset](http://ita.ee.lbl.gov/html/contrib/NASA-HTTP.html) and i opted to handle one by execution. So, if you wish to execute the script to check results from both files available on dataset, stay tuned on 'results' folder, because all sub-folders and files there can be overwritten.
#
# I need to extract this informations:
# - `bytes_per_day` -> (Quantity of Bytes per day.)
# - `frequency_status` -> (Frequency of each http status)
# - qty_http_404_per_day -> (Quantity of http 404 per Day)
# - sum_bytes -> (Sum of all request bytes)
# - `top_20_request` -> (Top twenty requests)
# - top_5_hosts_http_404 -> (Top five hosts with response http 404)
# - total_http_404 -> (Total of http 404)
# - unique_hosts -> (List of unique hosts)
#
# The highlighted items, are informations that did not belongs to the challenge, but its a plus.
#
#
# First, we need to import some packages as bellow
# +
import findspark
findspark.init()
from pyspark.sql import SQLContext
from pyspark import SparkContext
from pyspark.sql.functions import split, regexp_extract, col, desc
from pyspark.sql import Column
import matplotlib.pyplot as plt
from matplotlib.dates import (YEARLY, DateFormatter,
rrulewrapper, RRuleLocator, drange)
import datetime
from matplotlib.ticker import FuncFormatter
import numpy as np
# -
# Once we have imported those packages, lets define some useful functions.
# The first method we are defining is a simple way to setup a spark context.
def setup_spark_context():
return SparkContext("local", "NASA Data Set Analysis")
# Once we have a dataframe with necessary data, we will export a dataframe to a .CSV file on `results` folder. Note that there is a call for `coalesce()` method, used because we want DataFrame to write a single .CSV file. Also, is important to remember the `overwrite` option, which means, every execution the .csv files will be overwritten on `results` folder.
def export_query_to_csv(data_frame, file_name):
data_frame.coalesce(1).write.mode('overwrite').csv('results/' + file_name)
# On the next method, we just iterate over a dictionary of dataframes, and we use the previous method to export every query to a .csv file.
def export_all_queries_to_csv(data_frames):
for key, value in data_frames.items():
export_query_to_csv(value, key)
# So now, lets define a sparkContext, and after, we will create a sqlContext.
sc = setup_spark_context()
sqlContext = SQLContext(sc)
# Now, we need to load the NASA log file to our sqlContext. To do this, we will give the log file place ('data/NASA_access_log_Aug95'). In the python version of this script, you can give the place of the files from anywhere. Here, you must create a folder called `data` and put the NASA log files inside.
# +
sql_log_data = sqlContext.read.text('data/NASA_access_log_Aug95')
# -
# The dataset have informations like:
#
#
# 1. host making the request. A hostname when possible, otherwise the Internet address if the name could not be looked up.
# 2. timestamp in the format "DAY MON DD HH:MM:SS YYYY", where DAY is the day of the week, MON is the name of the month, DD is the day of the month, HH:MM:SS is the time of day using a 24-hour clock, and YYYY is the year. The timezone is -0400.
# 3. request given in quotes.
# 4. HTTP reply code.
# 5. bytes in the reply.
#
# Example:
#
# `in24.inetnebr.com - - [01/Aug/1995:00:00:01 -0400] "GET /shuttle/missions/sts-68/news/sts-68-mcc-05.txt HTTP/1.0" 200 1839`
#
#
# To extract this informations we use Regular Expressions(regexp_extract) from pysparksql and we give a alias for each information.
splited_data_frame = sql_log_data.select(regexp_extract('value', r'^([^\s]+\s)', 1).alias('host'),
regexp_extract('value', r'^.*\[(\d\d/\w{3}/\d{4}:\d{2}:\d{2}:\d{2} -\d{4})]', 1).alias('timestamp'),
regexp_extract('value', r'^.*"\w+\s+([^\s]+)\s+HTTP.*"', 1).alias('request'),
regexp_extract('value', r'^.*"\s+([^\s]+)', 1).cast('integer').alias('http_status'),
regexp_extract('value', r'^.*\s+(\d+)$', 1).cast('integer').alias('content_size_in_bytes'))
# Now, with the purpose of optimizing the queries, we need to persist the data do memory using cache().
splited_data_frame.cache()
# Once the data is cached, lets began to extract the informations we need. First, lets get the unique hots on the dataset.
# To future use, we are saving all the query results to a dictionary called `data_frames`.
# +
data_frames = {}
data_frames['unique_hosts'] = splited_data_frame.groupBy('host').count().filter('count = 1').select('host')
# -
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.groupBy('host').count().filter('count = 1').select('host').show()
# Now, lets list the top 20 requests.
#
data_frames['top_20_request'] = splited_data_frame.groupBy('request').count().sort(desc("count")).limit(20)
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.groupBy('request').count().sort(desc("count")).limit(20).show()
# We need list now, the total of http 404 results.
data_frames['total_http_404'] = splited_data_frame.groupBy('http_status').count().filter('http_status = "404"')
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.groupBy('http_status').count().filter('http_status = "404"').show()
# Now, we will list the frequency of all http status.
data_frames['frequency_status'] = splited_data_frame.groupBy('http_status').count().sort(desc('count'))
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.groupBy('http_status').count().sort(desc('count')).show()
# To a better visualization, lets plot this in a pie chart.
# +
labels = []
qty = []
for item in data_frames['frequency_status'].collect():
labels.append(item[0])
qty.append(item[1])
fig1, ax1 = plt.subplots()
ax1.pie(qty, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Equal aspect ratio ensures that pie is drawn as a circle.
plt.show()
# -
# We can also plot this in anorther form. Like the example below.
# +
fig, ax1 = plt.subplots()
ind = np.arange(1, 9)
plt.bar(ind, qty)
ax1.set_xticks(ind)
ax1.set_xticklabels(labels)
ax1.set_ylim([0, 1750000])
ax1.set_ylabel('Quantity')
ax1.set_xlabel('Http Status')
ax1.set_title('Http Status per day')
plt.show()
# -
# Now, lets list the top hosts answered with http 404
data_frames['top_5_hosts_http_404'] = splited_data_frame.filter('http_status = "404"').groupBy('request').count().sort(col("count").desc()).limit(5)
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.filter('http_status = "404"').groupBy('request').count().sort(col("count").desc()).limit(5).show()
# Now, lest extract the quantity of http 404 per day.
data_frames['qty_http_404_per_day'] = splited_data_frame.filter('http_status = "404"').groupBy(splited_data_frame.timestamp.substr(1, 11).alias('day')).count().sort('day')
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.filter('http_status = "404"').groupBy(splited_data_frame.timestamp.substr(1, 11).alias('day')).count().sort('day').show()
# In a Chart:
# +
dates = []
quantity = []
for item in data_frames['qty_http_404_per_day'].collect():
date_time_obj = datetime.datetime.strptime(item[0], '%d/%b/%Y')
dates.append(date_time_obj)
quantity.append(item[1])
formatter = DateFormatter('%d/%m/%Y')
date1 = datetime.date(1952, 1, 1)
date2 = datetime.date(2004, 4, 12)
delta = datetime.timedelta(days=100)
fig, ax2 = plt.subplots()
plt.plot(dates, quantity, 'b-')
ax2.set_xticks(dates)
ax2.set_xticklabels(dates)
ax2.xaxis.set_major_formatter(formatter)
ax2.xaxis.set_tick_params(rotation=90, labelsize=8)
ax2.set_ylabel('Quantity')
ax2.set_xlabel('Days')
plt.grid()
plt.show()
# -
# Now, lets show the sum of bytes of all requests in the dataset.
data_frames['sum_bytes'] = splited_data_frame.select('content_size_in_bytes').groupBy().sum()
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.select('content_size_in_bytes').groupBy().sum().show()
# In Gigabytes:
kbytes = data_frames['sum_bytes'].collect()[0][0] / 1000
mbytes = kbytes / 1000
mbytes / 1000
# Now, lets extract how many bytes per day.
data_frames['bytes_per_day'] = splited_data_frame.select('content_size_in_bytes', 'timestamp').groupBy(splited_data_frame.timestamp.substr(1, 11).alias('day')).sum().sort('day')
# To see what is the result of this query, lets add a `.show()` method.
splited_data_frame.select('content_size_in_bytes', 'timestamp').groupBy(splited_data_frame.timestamp.substr(1, 11).alias('day')).sum().sort('day').show()
# For a better visualization:
# +
dates = []
quantity = []
for item in data_frames['bytes_per_day'].collect():
date_time_obj = datetime.datetime.strptime(item[0], '%d/%b/%Y')
dates.append(date_time_obj)
quantity.append(item[1])
formatter = DateFormatter('%d/%m/%Y')
date1 = datetime.date(1952, 1, 1)
date2 = datetime.date(2004, 4, 12)
delta = datetime.timedelta(days=100)
fig, ax2 = plt.subplots()
plt.plot(dates, quantity, 'b-')
ax2.set_xticks(dates)
ax2.set_xticklabels(dates)
ax2.xaxis.set_major_formatter(formatter)
ax2.xaxis.set_tick_params(rotation=90, labelsize=8)
ax2.set_ylabel('Quantity')
ax2.set_xlabel('Days')
plt.grid()
plt.show()
# -
# Once we extract all informations, lets export the results calling the following function.
export_all_queries_to_csv(data_frames)
# I have inspirantions on the following links to solve this amazing challenge.
#
# http://www.awesomestats.in/spark-log-analysis/
#
# https://datascienceplus.com/spark-rdds-vs-dataframes-vs-sparksql-part-3-web-server-log-analysis/
#
#
# **Regards**!
|
nasa_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Change this to the number of cores you have available
# %env NUMBA_NUM_THREADS=4
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from chebysolve import largest_eigenpair, iterate_vector
# +
long_axis_size = 10_000
short_axis_size = 25
# Initialise a random generator
random_seed = 12345
rng = np.random.default_rng(random_seed)
# Sample matrix whose product AA^T we want to eigensolve
# Replace this with your data
matrix = rng.integers(10, size=(long_axis_size, short_axis_size)).astype('float64')
# matrix multiplication is imprecise with single precision and doesn't lead to much improvement in
# runtime, so best to use float64.
# Normalised random initial vector
initial_eigenvector_estimate = rng.random(long_axis_size, dtype='float64')
# +
eigenvalue_estimates_with_errors, eigenvector_estimate, converged = largest_eigenpair(
matrix, initial_eigenvector_estimate
)
eigenvalue_estimate = eigenvalue_estimates_with_errors[-1][0]
eigenvalue_error_estimate = eigenvalue_estimates_with_errors[-1][1]
if converged:
print("Converged at largest eigenvalue", eigenvalue_estimate,
"±", eigenvalue_error_estimate, "after",
len(eigenvalue_estimates_with_errors), "iterations.")
print("Eigenvector available in `eigenvector_estimate`.")
else:
print("Failed to converge after", len(eigenvalue_estimates_with_errors), "iterations.")
print("Last estimate was", eigenvalue_estimate, "±", eigenvalue_error_estimate, ".")
print("Eigenvector estimate available in `eigenvector_estimate`.")
# +
# Plot the history of the estimate
fig, axes = plt.subplots(nrows=2, sharex=True, figsize=(8, 6))
eigenvalue_estimates, eigenvalue_error_estimates = list(
zip(*eigenvalue_estimates_with_errors)
)
# Skip first iteration because the error bar is huge
axes[0].plot(
*zip(*enumerate(eigenvalue_estimates)),
marker='o',
linestyle='None'
)
axes[0].set_ylabel('Largest eigenvalue')
axes[1].plot(
*zip(*enumerate(eigenvalue_error_estimates)),
marker='x',
linestyle='None'
)
axes[1].set_ylabel('Error on largest eigenvalue')
axes[1].set_yscale('log')
axes[1].set_xlabel('Iteration')
fig.tight_layout()
plt.show()
# -
# %timeit -n 2 -r 15 iterate_vector(matrix, initial_eigenvector_estimate)
# +
import sys
print(sys.getsizeof(matrix))
print(sys.getsizeof(initial_eigenvector_estimate))
# -
|
chebysolve_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from piquery.piq_hash64 import download_to_hash
dhash = download_to_hash('https://pic.to8to.com/case/1907/25/20190725_69178e36f26803ba58e12f2dkovoe7fd.jpg')
from PIL import Image
from io import BytesIO
import requests
res = requests.get('http://pic.to8to.com/case/1907/25/20190725_217cfd8882c44fd1cdfawnfkgs238rse.jpg')
im = Image.open(BytesIO(res.content))
im.thumbnail((256, 256), Image.ANTIALIAS)
im = im.convert('L' )
dhash
dhash
from piquery.piquery import PIQBuilder
piq = PIQBuilder.buildHashQuery()
piq.queryRepeat('http://pic.to8to.com/case/{}'.format('day_081123/20090218_e465c7bed0567dbd321dNUKB7GBFCQfL.jpg'))
piq.queryRepeat('http://pic.to8to.com/case/{}'.format('day_081212/20090218_fa4d6149338441e6efdeSMhd0uZXWzbs.jpg'))
'http://pic.to8to.com/case/{}'.format('day_081123/20090218_e465c7bed0567dbd321dNUKB7GBFCQfL.jpg')
'http://pic.to8to.com/case/{}'.format('1904/27/20190427_13902c75a62031a238aaq0kpz295rota.jpg')
int(16059513831837940282).bit_length()
len('16059513831837940282')
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="gxzvVBdrarg9" colab_type="text"
# **Initialization**
#
# The next few steps(or code snippets) initialize the colab environment for running the fast.ai course. Each line of code that added are in place to avoid any error, please do not change the position of any snippet if you do not know what they do.
#
# Note:- You will get an error saying that the code is closing in the memory limit of 12Gb. You can click 'terminate other runtimes'.
# This error is popping up because the dataset is taking up most of the space, so don't worry
#
# + id="AJU6OF0maw8j" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 4}], "base_uri": "https://localhost:8080/", "height": 106} outputId="b9c793ec-8977-4139-8d2e-d5fdb4b1c021" executionInfo={"status": "ok", "timestamp": 1521147562289, "user_tz": -330, "elapsed": 32037, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
# !apt-get install -y -qq software-properties-common python-software-properties module-init-tools
# !add-apt-repository -y ppa:alessandro-strada/ppa 2>&1 > /dev/null
# !apt-get update -qq 2>&1 > /dev/null
# !apt-get -y install -qq google-drive-ocamlfuse fuse
from google.colab import auth
auth.authenticate_user()
from oauth2client.client import GoogleCredentials
creds = GoogleCredentials.get_application_default()
import getpass
# !google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret} < /dev/null 2>&1 | grep URL
vcode = getpass.getpass()
# !echo {vcode} | google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret}
# + id="cBKhUk5Va0GD" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# http://pytorch.org/
# pre installation
from os import path
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
accelerator = 'cu80' if path.exists('/opt/bin/nvidia-smi') else 'cpu'
# !pip install -q http://download.pytorch.org/whl/{accelerator}/torch-0.3.0.post4-{platform}-linux_x86_64.whl torchvision
import torch
# + id="rrEX2Hgpa3QG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 18}, {"item_id": 30}, {"item_id": 49}, {"item_id": 106}], "base_uri": "https://localhost:8080/", "height": 3406} outputId="177a3339-d483-43cd-c6f5-2c61ec5c9a54" executionInfo={"status": "ok", "timestamp": 1521147845450, "user_tz": -330, "elapsed": 233425, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
# !pip install fastai
# + id="hPC6hPgGa_Y-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 2}], "base_uri": "https://localhost:8080/", "height": 330} outputId="0022f436-c257-4652-8d1a-588479658c96" executionInfo={"status": "ok", "timestamp": 1521147846964, "user_tz": -330, "elapsed": 1442, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
# !mkdir data && wget http://files.grouplens.org/datasets/movielens/ml-latest-small.zip && unzip ml-latest-small.zip -d data/
# + [markdown] id="xZauz4Y7aoId" colab_type="text"
# ## Movielens
# + id="edB27G_3aoIh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# #%reload_ext autoreload
# #%autoreload 2
# %matplotlib inline
from fastai.learner import *
from fastai.column_data import *
# + [markdown] id="YhpyX_SRaoIt" colab_type="text"
# Data available from http://files.grouplens.org/datasets/movielens/ml-latest-small.zip
# + id="Ds80g2yHaoIv" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
path='data/ml-latest-small/'
# + [markdown] id="yh8aAzzaaoI3" colab_type="text"
# We're working with the movielens data, which contains one rating per row, like this:
# + id="avQh5JVuaoI6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 206} outputId="b7d275a6-c6af-426c-d94c-c13731eb0a2d" executionInfo={"status": "ok", "timestamp": 1521147850682, "user_tz": -330, "elapsed": 834, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
ratings = pd.read_csv(path+'ratings.csv')
ratings.head()
# + [markdown] id="CI_p8Z_baoJG" colab_type="text"
# Just for display purposes, let's read in the movie names too.
# + id="HaEUyVEKaoJI" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 206} outputId="75c207a0-2d70-4638-b572-3366e02933e4" executionInfo={"status": "ok", "timestamp": 1521147851578, "user_tz": -330, "elapsed": 736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
movies = pd.read_csv(path+'movies.csv')
movies.head()
# + [markdown] id="0BrNE-q5aoJQ" colab_type="text"
# ## Create subset for Excel
# + [markdown] id="aYVKuUDWaoJS" colab_type="text"
# We create a crosstab of the most popular movies and most movie-addicted users which we'll copy into Excel for creating a simple example. This isn't necessary for any of the modeling below however.
# + id="-fJkSQhNaoJV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 551} outputId="ea9b4b18-80e1-448c-ee0f-61dfd88fcd2f" executionInfo={"status": "ok", "timestamp": 1521147852447, "user_tz": -330, "elapsed": 731, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
g=ratings.groupby('userId')['rating'].count()
topUsers=g.sort_values(ascending=False)[:15]
g=ratings.groupby('movieId')['rating'].count()
topMovies=g.sort_values(ascending=False)[:15]
top_r = ratings.join(topUsers, rsuffix='_r', how='inner', on='userId')
top_r = top_r.join(topMovies, rsuffix='_r', how='inner', on='movieId')
pd.crosstab(top_r.userId, top_r.movieId, top_r.rating, aggfunc=np.sum)
# + [markdown] id="Kfx2VkaCaoJf" colab_type="text"
# ## Collaborative filtering
# + id="DwfdnPx3aoJh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
val_idxs = get_cv_idxs(len(ratings))
wd=2e-4
n_factors = 50
# + id="vA_phG-OaoJp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
cf = CollabFilterDataset.from_csv(path, 'ratings.csv', 'userId', 'movieId', 'rating')
learn = cf.get_learner(n_factors, val_idxs, 64, opt_fn=optim.Adam)
# + id="Uw_1sRIgaoJ2" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 43}, {"item_id": 68}], "base_uri": "https://localhost:8080/", "height": 180} outputId="22cd107c-3cf8-42b8-b322-658665856b71" executionInfo={"status": "ok", "timestamp": 1521147886380, "user_tz": -330, "elapsed": 29868, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
learn.fit(1e-2, 2, wds=wd, cycle_len=1, cycle_mult=2)
# + [markdown] id="CIkH0YEMaoKD" colab_type="text"
# Let's compare to some benchmarks. Here's [some benchmarks](https://www.librec.net/release/v1.3/example.html) on the same dataset for the popular Librec system for collaborative filtering. They show best results based on [RMSE](http://www.statisticshowto.com/rmse/) of 0.91. We'll need to take the square root of our loss, since we use plain MSE.
# + id="DQ86B35xaoKF" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 34} outputId="b59fad89-9801-41a8-bfeb-0cfa24518f93" executionInfo={"status": "ok", "timestamp": 1521147887287, "user_tz": -330, "elapsed": 832, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
math.sqrt(0.776)
# + [markdown] id="WYFD98xbaoKQ" colab_type="text"
# Looking good - we've found a solution better than any of those benchmarks! Let's take a look at how the predictions compare to actuals for this model.
# + id="tzCTw9s8aoKV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
preds = learn.predict()
# + id="BNXnU8gYaoKb" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}], "base_uri": "https://localhost:8080/", "height": 784} outputId="e5a33cbf-a363-4d97-cc29-fa1ec9a1c4f8" executionInfo={"status": "error", "timestamp": 1521147897028, "user_tz": -330, "elapsed": 8684, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
y=learn.data.val_y
sns.jointplot(preds, y, kind='hex', stat_func=None);
# + [markdown] id="zd-rTE8xaoKl" colab_type="text"
# ## Analyze results
# + [markdown] id="QppvoDQsaoKm" colab_type="text"
# ### Movie bias
# + id="enSxQrVWaoKn" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
movie_names = movies.set_index('movieId')['title'].to_dict()
g=ratings.groupby('movieId')['rating'].count()
topMovies=g.sort_values(ascending=False).index.values[:3000]
topMovieIdx = np.array([cf.item2idx[o] for o in topMovies])
# + id="fEEWZXbIaoKr" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 121} outputId="79e04a28-553d-44c8-f1fa-d8696b6181ea" executionInfo={"status": "ok", "timestamp": 1521147934103, "user_tz": -330, "elapsed": 1329, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
m=learn.model; m.cuda()
# + [markdown] id="BYYeQCTpaoK4" colab_type="text"
# First, we'll look at the movie bias term. Here, our input is the movie id (a single id), and the output is the movie bias (a single float).
# + id="gTxGsWo3aoK6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 531} outputId="372564b4-c442-475c-8f6d-52db52e87524" executionInfo={"status": "error", "timestamp": 1521147947869, "user_tz": -330, "elapsed": 964, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
movie_bias = to_np(m.ib(V(topMovieIdx)))
# + id="1bxr0OpKaoLM" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 166} outputId="32a494a2-237e-4981-aafa-c92b4fb9f18e" executionInfo={"status": "error", "timestamp": 1521147969912, "user_tz": -330, "elapsed": 4479, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
movie_bias
# + id="DdAFVLLkaoLV" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 166} outputId="2f91a320-a39b-46f7-f8d2-eb2d4a3cc3d2" executionInfo={"status": "error", "timestamp": 1521147978049, "user_tz": -330, "elapsed": 853, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
movie_ratings = [(b[0], movie_names[i]) for i,b in zip(topMovies,movie_bias)]
# + [markdown] id="E275orCkaoLf" colab_type="text"
# Now we can look at the top and bottom rated movies. These ratings are corrected for different levels of reviewer sentiment, as well as different types of movies that different reviewers watch.
# + id="sSdOWsFpaoLg" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="8285dcdc-5295-42cc-a3a4-4fd2eb23254a"
sorted(movie_ratings, key=lambda o: o[0])[:15]
# + id="_-7c4VCOaoLt" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="e03a85af-aaa3-4b56-96e4-6cb6e6d08794"
sorted(movie_ratings, key=itemgetter(0))[:15]
# + id="Q6fpxHZoaoL5" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="b7b2a443-6195-4aec-f696-cc91a32f535d"
sorted(movie_ratings, key=lambda o: o[0], reverse=True)[:15]
# + [markdown] id="ipNxDLYaaoMC" colab_type="text"
# ### Embedding interpretation
# + [markdown] id="Rh460mL8aoMC" colab_type="text"
# We can now do the same thing for the embeddings.
# + id="Bm8IoigqaoMG" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}], "base_uri": "https://localhost:8080/", "height": 548} outputId="729aea0c-6ac0-4f64-ea0a-9da711b9b187" executionInfo={"status": "error", "timestamp": 1521148007794, "user_tz": -330, "elapsed": 848, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "106326660524926425973"}}
movie_emb = to_np(m.i(V(topMovieIdx)))
movie_emb.shape
# + [markdown] id="5pglg7pNaoMM" colab_type="text"
# Because it's hard to interpret 50 embeddings, we use [PCA](https://plot.ly/ipython-notebooks/principal-component-analysis/) to simplify them down to just 3 vectors.
# + id="HSsiwtKyaoMN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from sklearn.decomposition import PCA
pca = PCA(n_components=3)
movie_pca = pca.fit(movie_emb.T).components_
# + id="kvWR1kmLaoMT" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="fbf5d245-b32c-4665-adf4-e1a56ac5a1c2"
movie_pca.shape
# + id="ro3nsvQOaoMY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
fac0 = movie_pca[0]
movie_comp = [(f, movie_names[i]) for f,i in zip(fac0, topMovies)]
# + [markdown] id="bighQuJFaoMb" colab_type="text"
# Here's the 1st component. It seems to be 'easy watching' vs 'serious'.
# + id="D_PlpPlNaoMe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="082f4a7e-bbe1-42c3-a414-d4732d7b3cbb"
sorted(movie_comp, key=itemgetter(0), reverse=True)[:10]
# + id="J3InLi6OaoMl" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="68b0b310-eaa3-4a50-a12a-212c1480c47f"
sorted(movie_comp, key=itemgetter(0))[:10]
# + id="PAReed_JaoMp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
fac1 = movie_pca[1]
movie_comp = [(f, movie_names[i]) for f,i in zip(fac1, topMovies)]
# + [markdown] id="2T855Ob8aoMt" colab_type="text"
# Here's the 2nd component. It seems to be 'CGI' vs 'dialog driven'.
# + id="HhFG0fodaoMu" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="b1939532-93b4-486d-812d-006650640ec0"
sorted(movie_comp, key=itemgetter(0), reverse=True)[:10]
# + id="mCq86eSVaoM0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="1e5dc772-46a6-46df-c33d-347fcae0d7ed"
sorted(movie_comp, key=itemgetter(0))[:10]
# + [markdown] id="4DiEWiI1aoM7" colab_type="text"
# We can draw a picture to see how various movies appear on the map of these components. This picture shows the first two components.
# + id="t7Q8_qTMaoM8" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="843b0462-5a24-443c-905a-20e37afa35d2"
idxs = np.random.choice(len(topMovies), 50, replace=False)
X = fac0[idxs]
Y = fac1[idxs]
plt.figure(figsize=(15,15))
plt.scatter(X, Y)
for i, x, y in zip(topMovies[idxs], X, Y):
plt.text(x,y,movie_names[i], color=np.random.rand(3)*0.7, fontsize=11)
plt.show()
# + [markdown] id="1201U-CkaoNK" colab_type="text"
# ## Collab filtering from scratch
# + [markdown] id="FXWsGVweaoNL" colab_type="text"
# ### Dot product example
# + id="IF8jf6NnaoNN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="0caff8c1-169e-4721-f9a9-2a521c5c4d12"
a = T([[1.,2],[3,4]])
b = T([[2.,2],[10,10]])
a,b
# + id="C_tffm2NaoNS" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="b48cad64-45d3-4a01-895a-2f0a4468e5f0"
a*b
# + id="Vu4EAqlgaoNY" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="ba66334f-59d9-40ff-f1aa-24c74c49334a"
(a*b).sum(1)
# + id="KKpPObVAaoNc" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class DotProduct(nn.Module):
def forward(self, u, m): return (u*m).sum(1)
# + id="6l0zAogkaoNf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
model=DotProduct()
# + id="GU9b0CRnaoNj" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="aa0b932a-2448-48b9-d844-6fddb169e104"
model(a,b)
# + [markdown] id="wC0oZrxuaoNm" colab_type="text"
# ### Dot product model
# + id="MSbo4cylaoNn" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
u_uniq = ratings.userId.unique()
user2idx = {o:i for i,o in enumerate(u_uniq)}
ratings.userId = ratings.userId.apply(lambda x: user2idx[x])
m_uniq = ratings.movieId.unique()
movie2idx = {o:i for i,o in enumerate(m_uniq)}
ratings.movieId = ratings.movieId.apply(lambda x: movie2idx[x])
n_users=int(ratings.userId.nunique())
n_movies=int(ratings.movieId.nunique())
# + id="37blaBCeaoNp" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class EmbeddingDot(nn.Module):
def __init__(self, n_users, n_movies):
super().__init__()
self.u = nn.Embedding(n_users, n_factors)
self.m = nn.Embedding(n_movies, n_factors)
self.u.weight.data.uniform_(0,0.05)
self.m.weight.data.uniform_(0,0.05)
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
u,m = self.u(users),self.m(movies)
return (u*m).sum(1)
# + id="d9bRLT4caoNs" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
x = ratings.drop(['rating', 'timestamp'],axis=1)
y = ratings['rating'].astype(np.float32)
# + id="AblAwxcwaoNv" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
data = ColumnarModelData.from_data_frame(path, val_idxs, x, y, ['userId', 'movieId'], 64)
# + id="50xKZlP2aoNz" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
wd=1e-5
model = EmbeddingDot(n_users, n_movies).cuda()
opt = optim.SGD(model.parameters(), 1e-1, weight_decay=wd, momentum=0.9)
# + id="k8vALYNHaoN2" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="142d70ff-4390-43b1-c0bb-82e20906f722"
fit(model, data, 3, opt, F.mse_loss)
# + id="QY6F_-DyaoN7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
set_lrs(opt, 0.01)
# + id="OIPqIIB1aoOE" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="a5ace65d-ddcd-4abb-f592-3dbb0396ace8"
fit(model, data, 3, opt, F.mse_loss)
# + [markdown] id="K4hHVoGBaoOI" colab_type="text"
# ### Bias
# + id="VWXz3cl3aoOJ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}]} outputId="0d601e7c-dda3-4ef0-c296-2b1afba766d2"
min_rating,max_rating = ratings.rating.min(),ratings.rating.max()
min_rating,max_rating
# + id="3eyW9mo8aoOL" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def get_emb(ni,nf):
e = nn.Embedding(ni, nf)
e.weight.data.uniform_(-0.01,0.01)
return e
class EmbeddingDotBias(nn.Module):
def __init__(self, n_users, n_movies):
super().__init__()
(self.u, self.m, self.ub, self.mb) = [get_emb(*o) for o in [
(n_users, n_factors), (n_movies, n_factors), (n_users,1), (n_movies,1)
]]
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
um = (self.u(users)* self.m(movies)).sum(1)
res = um + self.ub(users).squeeze() + self.mb(movies).squeeze()
res = F.sigmoid(res) * (max_rating-min_rating) + min_rating
return res
# + id="xE4m-_z4aoOQ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
wd=2e-4
model = EmbeddingDotBias(cf.n_users, cf.n_items).cuda()
opt = optim.SGD(model.parameters(), 1e-1, weight_decay=wd, momentum=0.9)
# + id="5xHfcK7DaoOU" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="c40ce878-1d9f-45b5-bbd5-810265a0029b"
fit(model, data, 3, opt, F.mse_loss)
# + id="ayPUNtnwaoOX" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
set_lrs(opt, 1e-2)
# + id="wald34VJaoOa" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="c6543c49-e111-4cc5-9e2e-05b857d959c5"
fit(model, data, 3, opt, F.mse_loss)
# + [markdown] id="P4sWJuGkaoOe" colab_type="text"
# ### Mini net
# + id="BMGg0RqzaoOf" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
class EmbeddingNet(nn.Module):
def __init__(self, n_users, n_movies, nh=10, p1=0.05, p2=0.5):
super().__init__()
(self.u, self.m) = [get_emb(*o) for o in [
(n_users, n_factors), (n_movies, n_factors)]]
self.lin1 = nn.Linear(n_factors*2, nh)
self.lin2 = nn.Linear(nh, 1)
self.drop1 = nn.Dropout(p1)
self.drop2 = nn.Dropout(p2)
def forward(self, cats, conts):
users,movies = cats[:,0],cats[:,1]
x = self.drop1(torch.cat([self.u(users),self.m(movies)], dim=1))
x = self.drop2(F.relu(self.lin1(x)))
return F.sigmoid(self.lin2(x)) * (max_rating-min_rating+1) + min_rating-0.5
# + id="LBPXIZQzaoOh" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
wd=1e-5
model = EmbeddingNet(n_users, n_movies).cuda()
opt = optim.Adam(model.parameters(), 1e-3, weight_decay=wd)
# + id="EUMt7SIHaoOj" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="20d387de-ca26-418d-a09d-b6cea664f73d"
fit(model, data, 3, opt, F.mse_loss)
# + id="gnBKWwhSaoOm" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
set_lrs(opt, 1e-3)
# + id="VBj2sIaDaoOo" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}, {}]} outputId="f509eb4a-a171-41c8-deec-edae5bcf46a1"
fit(model, data, 3, opt, F.mse_loss)
# + id="BUsLBXU-aoO7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
|
dl1/lesson5-movielens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### BreastCancerClassifier
# This notebook is a showcase of common classification methods use breast cancer data as an example. The Wisconsin Breast Cancer Database was studied by <NAME> in 1991. There are total 699 instances and 10 attributes plus class attribute. Attributes include
# 1. Sample code number (id number)
# 2. Clump Thickness (1 - 10)
# 3. Uniformity of Cell Size (1 - 10)
# 4. Uniformity of Cell Shape (1 - 10)
# 5. Marginal Adhesion (1 - 10)
# 6. Single Epithelial Cell Size (1 - 10)
# 7. Bare Nuclei (1 - 10)
# 8. Bland Chromatin (1 - 10)
# 9. Normal Nucleoli (1 - 10)
# 10. Mitoses (1 - 10)
# 11. Class: (2 for benign, 4 for malignant)
#
# #### Model
# ###### 1. Scikit-Learn:
# SVC (with linear and rgb kernal), Decission Tree, Random Forest Classifier, Logistic Regression, Extra Trees Classifier, Gradient Boosting Classifier, Neuron Network, KNN Classifier, Gaussian Process Classifier, Logistic Regression, Linear Discriminant Analysis and Ada Boost Classifier.
# As well as ensemble methods such as voting classifier, ensemble tree (logistic regression+ random forest, logistic regression + gradient boosting)
# ###### 2. XGBoost Classifier
# ###### 3. Light GBM Classifier
# ###### 4. Deep Learning via Keras and Tensor flow:
# Three fully connected hidden layers with (32,16,8) mixes with linear, relu and sigmoid activation function.
#
#
# #### Table of Content
# [1.1 General Classifier](#general)
#
# [1.1.1 Cross Validation](#CV_general)
#
# [1.1.2 Prediction](#Pred_general)
#
# [1.1.3 ROC Curve](#ROC_general)
#
# [1.1.4 Learning Curve](#LR_general)
#
# [1.2.1 Ensemble:Voting](#voting)
#
# [1.3.1 Ensemble:Tree](#ensemble_tree)
#
# [2.1 Xgboost](#xgboost)
#
# [3.1 LightGBM](#lightgbm)
#
# [4.1 DeepLearning:TensorFlow and Keras](#tf)
# +
# %matplotlib inline
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import lightgbm as gbm
from lightgbm import LGBMClassifier
import xgboost as xgb
from xgboost import XGBClassifier
from keras.models import Sequential
from keras.layers import Dense
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import train_test_split,cross_val_score, GridSearchCV, StratifiedKFold, learning_curve, StratifiedKFold
from sklearn.preprocessing import LabelEncoder, StandardScaler, OneHotEncoder
from sklearn.pipeline import Pipeline, make_pipeline
from sklearn.metrics import roc_curve, auc, accuracy_score, f1_score, classification_report
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier,GradientBoostingClassifier, ExtraTreesClassifier, VotingClassifier, RandomTreesEmbedding
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.svm import SVC
# -
data = pd.read_csv('data/breastCancer.csv')
# #### Overview
# 699 entries, 10 columns
# #### columns summary
# - id : unique id
# - feature : numerical rating from 1-10 ['clump_thickness', 'size_uniformity', 'shape_uniformity','marginal_adhesion', 'epithelial_size', 'bare_nucleoli','bland_chromatin', 'normal_nucleoli', 'mitoses']
# - class : {2:benign, 4:malignant}
##fillna
bn_mean = np.floor(np.mean(np.int64(data.bare_nucleoli[data.bare_nucleoli!="?"])))
data.replace('?', bn_mean, inplace=True)
data = data.applymap(np.float64)
feature_cols = ['clump_thickness', 'size_uniformity', 'shape_uniformity','marginal_adhesion', 'epithelial_size', 'bare_nucleoli','bland_chromatin', 'normal_nucleoli', 'mitoses']
classes = {2:0,4:1}
data['binary_class'] = data['class'].map(classes)
X = data[feature_cols].values
Y = data['binary_class'].values
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state = 0)
# #### General Scikit-Learn Methods
# <a id='general'></a>
scaler = StandardScaler()
kfold = StratifiedKFold(n_splits=10)
random_state = 2
MLP_params = {'hidden_layer_sizes':(20,),'solver': 'adam', 'learning_rate_init': 0.01,'max_iter':1000}
classifiers = []
classifiers.append(SVC(kernel="linear", C=0.025,probability=True,random_state=random_state)) #predict_proba available when probability=True
classifiers.append(SVC(gamma=2, C=1,probability=True,random_state=random_state))#predict_proba
classifiers.append(DecisionTreeClassifier(random_state=random_state)) #predict_proba
classifiers.append(AdaBoostClassifier(DecisionTreeClassifier(random_state=random_state),random_state=random_state,learning_rate=0.1))
classifiers.append(RandomForestClassifier(random_state=random_state)) #predict_proba
classifiers.append(ExtraTreesClassifier(random_state=random_state)) #predict_proba
classifiers.append(GradientBoostingClassifier(random_state=random_state)) #predict_proba
classifiers.append(MLPClassifier(random_state=random_state,**MLP_params)) #predict_proba
classifiers.append(KNeighborsClassifier()) #predict_proba
classifiers.append(LogisticRegression(random_state = random_state)) #predict_proba
classifiers.append(LinearDiscriminantAnalysis()) #predict_proba
classifiers.append(GaussianProcessClassifier(1.0 * RBF(1.0))) #predict_proba
start = time.time()
cv_results = []
for classifier in classifiers :
c_start = time.time()
print (classifier)
cv_results.append(cross_val_score(classifier, scaler.fit_transform(X_train), y = Y_train, scoring = 'f1', cv = kfold, n_jobs=-1))
c_time = time.time()-c_start
print ('time: {:f}'.format(c_time))
tot_time = time.time()-start
print ('total_time: {:f}'.format(tot_time))
cv_means = []
cv_std = []
for cv_result in cv_results:
cv_means.append(cv_result.mean())
cv_std.append(cv_result.std())
# #### Cross-Validation
# <a id='CV_general'></a>
# Use F1 score to evalute score, the top 5 methods
# 1. SVC with linear kernal
# 2. KNN classifier
# 3. Logistic Regression
# 4. Gaussian Process
# 5. Extra Trees
cv_res = pd.DataFrame({"CrossValF1Means":cv_means,"CrossValerrors": cv_std,\
"Algorithm":["SVC_lm","SVC_rbf","DecisionTreeClassifier","AdaBoostClassifier","RandomForestClassifier",
"ExtraTreesClassifier","GradientBoostingClassifier","MLPClassifier",'KNeighborsClassifier',
'LogisticRegression','LinearDiscriminantAnalysis','GaussianProcessClassifier']})
cv_res.sort_values('CrossValF1Means',ascending=False).head()
g = sns.barplot("CrossValF1Means","Algorithm",data = cv_res.sort_values('CrossValF1Means',ascending=False), palette="Set3",orient = "h",**{'xerr':cv_std})
g.set_xlabel("Accuracy")
g = g.set_title("Cross validation scores")
# #### Prediction
# <a id='Pred_general'></a>
# Compare the accuracy of predictions by different method. Top 5 methods are
# 1. SVC with linear kernal (97.8571%)
# 2. Random Forest
# 3. Extra Trees
# 4. Gradient Boosting
# 5. Nuron Network
plt.figure(figsize=(12,12))
algrithm = ["SVC_lm","SVC_rgb","DecisionTreeClassifier","AdaBoostClassifier","RandomForestClassifier",
"ExtraTreesClassifier","GradientBoostingClassifier","MLPClassifier",'KNeighborsClassifier',
'LogisticRegression','LinearDiscriminantAnalysis','GaussianProcessClassifier']
ROCurve = dict()
cv_res["Accuracy"]=0
for n, classifier in enumerate(classifiers):
name = algrithm[n]
classifier.fit(scaler.transform(X_train),Y_train)
probs = classifier.predict_proba(scaler.transform(X_test))
preds = probs[:,1]
fpr, tpr, threshold = roc_curve(Y_test, preds)
roc_auc = auc(fpr, tpr)
ROCurve[name]=(fpr,tpr,roc_auc)
acc = accuracy_score(Y_test,classifier.predict(scaler.transform(X_test)))
cv_res.loc[n,"Accuracy"] = acc
cv_res.sort_values('Accuracy',ascending=False).head()
g = sns.barplot("Accuracy","Algorithm",data = cv_res.sort_values('Accuracy',ascending=False), palette="Set3",orient = "h")
g.set_xlabel("Accuracy")
g = g.set_title("Test Prediction scores")
# #### ROC Curve
# <a id='ROC_general'></a>
# We want to maximize both precission and recall, not just precission. The ROC area under curve (auc) demonstrated comparision of the true positive rate and false positive between each methods. Top 5 are
# 1. Linear Discriminant Analysis
# 2. Logistic Regression
# 3. Gaussian Process
# 4. Gradien Boosting
# 5. SVC linear kernal
plt.figure(figsize=(16,8))
for C in ROCurve:
#ROCurve[name]=(fpr,tpr,roc_auc)
plt.plot(ROCurve[C][0], ROCurve[C][1], label = '{:s} AUC = {:.3f}'.format(str(C),ROCurve[C][2]))
plt.title('Receiver Operating Characteristic')
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 0.2])
plt.ylim([0.8, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
# #### Learning Curve
# <a id='LR_general'></a>
# Some methods tend to overfitting the training sample.
# For example, SVC rgb, Decission Tree, Ada Boost, Random Forest, Extra Tree, MPL and Gradient Boosting method. For these methods, we should fine tune hyperparameters to reduce the model complexity to avoid overfit.
# In the other hand, the learning curves of SVC linear, KNN, Logistic, LDA and GPC perform better without overfiting or underfiting
#
###Learning Curve
def plot_learning_curve(estimator, title, X, y, ylim=None, cv=None,
n_jobs=-1, train_sizes=np.linspace(.1, 1.0, 5)):
"""Generate a simple plot of the test and training learning curve"""
scaler = StandardScaler()
kfold = StratifiedKFold(n_splits=10)
X = scaler.fit_transform(X)
plt.figure()
plt.title(title)
if ylim is not None:
plt.ylim(*ylim)
plt.xlabel("Training examples")
plt.ylabel("Score")
train_sizes, train_scores, test_scores = learning_curve(
estimator, X, y, cv=kfold, n_jobs=n_jobs, train_sizes=train_sizes, scoring='accuracy', verbose = 0)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.grid()
plt.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
plt.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="g")
plt.plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
plt.plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
plt.legend(loc="best")
plt.show()
#return plt
for n, c in enumerate(algrithm):
plot_learning_curve(classifiers[n],c,X_train,Y_train)
# #### Voting Classifiers
# <a id='voting'></a>
### Voting
voting_estimators=["SVC_lm","RandomForestClassifier",'GaussianProcessClassifier','KNeighborsClassifier','LogisticRegression']
estimators = list()
for n,c in enumerate(algrithm):
if c in voting_estimators:
estimators.append((c,classifiers[n]))
votingC = VotingClassifier(estimators=estimators, voting='soft', n_jobs=-1)
votingC = votingC.fit(scaler.transform(X_train), Y_train)
y_pred = votingC.predict(scaler.transform(X_test))
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(Y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# #### Ensemble of Trees
# <a id='ensemble_tree'></a>
# +
### Ensemble of Trees
from sklearn.preprocessing import OneHotEncoder
from sklearn.ensemble import RandomTreesEmbedding
from sklearn.pipeline import make_pipeline
scaler = StandardScaler()
n_estimator = 10
X = data[feature_cols].values
Y = data['binary_class'].values
X_train, X_test, y_train, y_test = train_test_split(scaler.fit_transform(X), Y, test_size=0.5)
# It is important to train the ensemble of trees on a different subset
# of the training data than the linear regression model to avoid
# overfitting, in particular if the total number of leaves is
# similar to the number of training samples
X_train, X_train_lr, y_train, y_train_lr = train_test_split(X_train,
y_train,
test_size=0.5)
# Unsupervised transformation based on totally random trees
rt = RandomTreesEmbedding(max_depth=3, n_estimators=n_estimator,
random_state=0)
rt_lm = LogisticRegression()
pipeline = make_pipeline(rt, rt_lm)
pipeline.fit(X_train, y_train)
y_pred_rt = pipeline.predict_proba(X_test)[:, 1]
fpr_rt_lm, tpr_rt_lm, _ = roc_curve(y_test, y_pred_rt)
auc_rt_lm = auc(fpr_rt_lm, tpr_rt_lm)
# Supervised transformation based on random forests
rf = RandomForestClassifier(max_depth=3, n_estimators=n_estimator)
rf_enc = OneHotEncoder()
rf_lm = LogisticRegression()
rf.fit(X_train, y_train)
rf_enc.fit(rf.apply(X_train))
rf_lm.fit(rf_enc.transform(rf.apply(X_train_lr)), y_train_lr)
y_pred_rf_lm = rf_lm.predict_proba(rf_enc.transform(rf.apply(X_test)))[:, 1]
fpr_rf_lm, tpr_rf_lm, _ = roc_curve(y_test, y_pred_rf_lm)
auc_rf_lm = auc(fpr_rf_lm, tpr_rf_lm)
grd = GradientBoostingClassifier(n_estimators=n_estimator)
grd_enc = OneHotEncoder()
grd_lm = LogisticRegression()
grd.fit(X_train, y_train)
grd_enc.fit(grd.apply(X_train)[:, :, 0])
grd_lm.fit(grd_enc.transform(grd.apply(X_train_lr)[:, :, 0]), y_train_lr)
y_pred_grd_lm = grd_lm.predict_proba(
grd_enc.transform(grd.apply(X_test)[:, :, 0]))[:, 1]
fpr_grd_lm, tpr_grd_lm, _ = roc_curve(y_test, y_pred_grd_lm)
auc_grd_lm = auc(fpr_grd_lm, tpr_grd_lm)
# The gradient boosted model by itself
y_pred_grd = grd.predict_proba(X_test)[:, 1]
fpr_grd, tpr_grd, _ = roc_curve(y_test, y_pred_grd)
auc_grd = auc(fpr_grd, tpr_grd)
# The random forest model by itself
y_pred_rf = rf.predict_proba(X_test)[:, 1]
fpr_rf, tpr_rf, _ = roc_curve(y_test, y_pred_rf)
auc_rf = auc(fpr_rf, tpr_rf)
'{:s} AUC = {:.3f}'.format(str(C),ROCurve[C][2])
plt.figure(1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_rt_lm, tpr_rt_lm, label='RT + LR AUC = {:.3f}'.format(auc_rt_lm))
plt.plot(fpr_rf, tpr_rf, label='RF AUC = {:.3f}'.format(auc_rf))
plt.plot(fpr_rf_lm, tpr_rf_lm, label='RF + LR AUC = {:.3f}'.format(auc_rf_lm))
plt.plot(fpr_grd, tpr_grd, label='GBT AUC = {:.3f}'.format(auc_grd))
plt.plot(fpr_grd_lm, tpr_grd_lm, label='GBT + LR AUC = {:.3f}'.format(auc_grd_lm))
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve')
plt.legend(loc='best')
plt.show()
plt.figure(2)
plt.xlim(0, 0.2)
plt.ylim(0.8, 1)
plt.plot([0, 1], [0, 1], 'k--')
plt.plot(fpr_rt_lm, tpr_rt_lm, label='RT + LR')
plt.plot(fpr_rf, tpr_rf, label='RF')
plt.plot(fpr_rf_lm, tpr_rf_lm, label='RF + LR')
plt.plot(fpr_grd, tpr_grd, label='GBT')
plt.plot(fpr_grd_lm, tpr_grd_lm, label='GBT + LR')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.title('ROC curve (zoomed in at top left)')
plt.legend(loc='best')
plt.show()
# -
# #### XGBoost
# <a id='xgboost'></a>
###xgboost
from xgboost import XGBClassifier
X = data[feature_cols].values
Y = data['binary_class'].values
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.20, random_state = 0)
xgb = XGBClassifier()
xgb.fit(scaler.transform(X_train), Y_train)
y_pred = xgb.predict(scaler.transform(X_test))
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(Y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# #### LightGBM
# <a id='lightgbm'></a>
###lightGBM
from lightgbm import LGBMClassifier
gbm = LGBMClassifier()
gbm.fit(scaler.transform(X_train), Y_train)
y_pred = gbm.predict(scaler.transform(X_test))
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(Y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# #### Deep Learning : TensorFlow
# <a id='tf'></a>
###TensorFlow
DNN = Sequential()
DNN.add(Dense(32,activation='linear',input_dim=9))
DNN.add(Dense(16,activation='relu'))
DNN.add(Dense(8,activation='relu'))
DNN.add(Dense(1,activation='sigmoid'))
#sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
DNN.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
DNN.fit(scaler.transform(X_train), Y_train, epochs=10, batch_size=16)
y_pred = DNN.predict(scaler.transform(X_test))
predictions = [round(value) for value in y_pred[:,0]]
accuracy = accuracy_score(Y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
|
BreastCancerClassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python e API com Login
#
# ### O 1º Passo de toda API com Login é criar uma conta e pegar suas credenciais
#
# ### No seu código, o 1º passo é sempre estabelecer a conexão com a API, usando seu login e suas credenciais
#
# - Como cada API é uma ferramenta diferente, cada uma delas pode exigir que você faça algum tipo de configuração, que vai estar explicada na API. No nosso caso, teremos que validar um número e criar um número de envio
#
# - Depois, usamos os métodos da API normalmente para fazer o que queremos. No nosso caso, enviar um SMS
# #### 1. Vamos criar um login no Twilio
#
# https://www.twilio.com/docs/libraries/python
# #### 2. Depois do Login, vamos pegar 3 informações:
#
# - ID da Conta
# - Token
# - Número de Envio
# #### 3. Agora vamos validar um número porque no Twilio, enviar SMS para um número válido é de graça
# #### 4. Agora podemos fazer o nosso código de acordo com as orientações do Twilio
# +
from twilio.rest import Client
account_sid = '' # coloque o seu número de account_sid
token = '' # coloque o seu número de token
client = Client(account_sid, token)
remetente = '' # coloque o seu número de remetente
destino = '' # coloque o seu número de destino
message = client.messages.create(
to=destino,
from_=remetente,
body="Aqui fica a mensagem") # mensagem a ser enviada
print(message.sid)
# -
|
sms-login.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model improvement
#
# In this iteration dive deeper into **feature engineering** generating more suitable features for our classification models.
# Moreover, we only focus on selected data mining methods which we find most suiable.
# In particular, we will focus on:
#
# - KNN
# - Random Forest
# - XGBoost
#
# import required packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#from imblearn.over_sampling import RandomOverSampler
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.model_selection import GridSearchCV,train_test_split
from sklearn.metrics import accuracy_score, f1_score, precision_score, recall_score, confusion_matrix
from src.data.make_dataset import merge_data
from src.features.build_features import features,split_data,feature_engineering, drop_unnecessary_ft, get_unnecessary_ft
from xgboost import XGBClassifier
pd.options.display.max_columns =70
# # I. Import whole dataset
# +
# merge the dataset at a whole (train, test & target data all together)
train_values = pd.read_csv('../data/external/train_values.csv', index_col='building_id')
train_target = pd.read_csv('../data/external/train_labels.csv', index_col='building_id')
test_values = pd.read_csv('../data/external/test_values.csv', index_col='building_id')
df=merge_data(train_values, train_target, test_values)
df=df.reset_index()
display(df)
# -
# # II. Preprocess dataset
# apply features function where encoding, scaling and adding means of categorical features from above happens
df=features(df)
display(df)
train_target
# # III. Feature Engineering: Generate new Features
# +
grouped = df
grouped = df.loc[~df.damage_grade.isna()].drop(columns='damage_grade')
grouped = grouped.set_index('building_id')
grouped = grouped.join(train_target['damage_grade'])
dmg = grouped.groupby('damage_grade').agg({'damage_grade':'count'})
grouped = grouped.groupby('damage_grade').sum()
grouped = grouped.join(dmg)
grouped = grouped.iloc[:,:-1].div(grouped.damage_grade, axis=0)
grouped = grouped.transpose()
grouped = grouped.iloc[8:]
labels = ['1', '2', '3']
plt.figure(figsize=(20,13))
for i in range (1,4):
plt.plot(grouped.index, grouped[i], label=labels[i-1])
plt.grid()
plt.xlabel('')
plt.yticks( np.arange(0,1, step=0.1))
plt.xticks(rotation=90 )
plt.ylabel('')
plt.title('')
plt.savefig('feature_importance_per_damage_grade.png')
plt.legend()
# +
data_ges = train_values.join(train_target)
data_ges = data_ges.reset_index()
age_dmggrade = data_ges.groupby(['age','damage_grade']).agg({'building_id':'count'})
age_dmggrade = age_dmggrade.reset_index()
age_dmggrade #= age_dmggrade[age_dmggrade['age']<=150]
geo = data_ges.groupby(['geo_level_1_id','damage_grade']).agg({'building_id':'count'}).reset_index()
found = data_ges.groupby(['foundation_type','damage_grade']).agg({'building_id':'count'}).reset_index()
buildings_1=age_dmggrade[age_dmggrade['damage_grade']==1].building_id.sum()
buildings_2=age_dmggrade[age_dmggrade['damage_grade']==2].building_id.sum()
buildings_3=age_dmggrade[age_dmggrade['damage_grade']==3].building_id.sum()
age_dmggrade['relative'] = age_dmggrade[age_dmggrade['damage_grade']==1].building_id.div(buildings_1).mul(100000)
age_dmggrade.loc[ (age_dmggrade['damage_grade']==2), 'relative'] = age_dmggrade[age_dmggrade['damage_grade']==2].building_id.div(buildings_2).mul(100000)
age_dmggrade.loc[ (age_dmggrade['damage_grade']==3), 'relative'] = age_dmggrade[age_dmggrade['damage_grade']==3].building_id.div(buildings_3).mul(100000)
#age_dmggrade['relative'] = age_dmggrade[age_dmggrade['damage_grade']==3].building_id.div(buildings_3)
age_dmggrade=age_dmggrade[age_dmggrade['age']<100]
plt.figure(figsize=(12, 9), dpi=80)
plt.scatter(age_dmggrade['age'], age_dmggrade['damage_grade'], s=age_dmggrade['relative'], c=np.log10(age_dmggrade['relative']), alpha=0.5)
plt.colorbar()
plt.xlabel('Age of buildings')
plt.ylabel('Damage grade')
plt.savefig('ageofbuildings_clusteredin_damagegrade')
plt.grid()
plt.show()
# -
plt.figure(figsize=(16, 7), dpi=80)
plt.scatter(geo['geo_level_1_id'], geo['damage_grade'], s=geo['building_id'], c=np.log10(geo['building_id']), alpha=0.5)
plt.colorbar()
plt.xticks(np.arange(0, 31, step=1))
plt.xlabel('District Number')
plt.ylabel('Damage grade')
plt.savefig('districtnumber_clusteredin_damagegrade')
plt.grid()
plt.show()
plt.figure(figsize=(12, 9), dpi=80)
plt.scatter(found['foundation_type'], found['damage_grade'], s=found['building_id'], c=np.log10(found['building_id']), alpha=0.5)
plt.colorbar()
plt.xlabel('Age of buildings')
plt.ylabel('Damage grade')
plt.grid()
plt.show()
# In this step we use our ``feature_engineering`` function which creates additional features in order to improve our classification model.
#
# Overall, we increase our features from **70** to **88** feautres in total.
#
#
# +
pd.set_option('display.max_columns', None)
display(df)
df = feature_engineering(df)
display(df)
# -
# ## Dropping unnecessary features due to infrequency
#
# - here we use our functions ``get_unnecessary_ft()`` and ``drop_unnecessary_ft()`` to remove all feautures which fall below a threshold of relative occurrence per damage grade
display(get_unnecessary_ft(df))
df = drop_unnecessary_ft(df)
# ## Dropping correlated features
#
# +
#Correlation
correlated_features = set()
correlation_matrix = df.corr()
for i in range(len(correlation_matrix.columns)):
for j in range(i):
if abs(correlation_matrix.iloc[i, j]) > 0.8:
print(f"The following features are correlated: {correlation_matrix.columns[i]} and {correlation_matrix.columns[j]}. Correlation = {round(abs(correlation_matrix.iloc[i, j]),2)}")
colname = correlation_matrix.columns[j]
correlated_features.add(colname)
print(f"Drop the following features: {correlated_features}")
#drop correlated features
df=df.drop(columns=correlated_features)
# -
# ## Recreate our train and test values
#
# - dropping correlated and low frequent features results into a total number of **62** features for remaining analysis
train_data, train_target, test_data = split_data(df)
train_data
# ### Train and Test Split
# train-test split with stratify
data_train, data_test, target_train, target_test = train_test_split(train_data, train_target, test_size=0.2, random_state=42, stratify=train_target)
display(data_train)
# # IV. Engineered Models without Hyperparameter Tuning
#
# In this step we focus on our most promising candidate models:
# `
# - ``RandomForest``
# - ``KNN``
# - ``XGBoost``
# ## Random Forest (no Rebalance)
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score, StratifiedKFold
# cross-validation using random forest
skfold = StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
rf = RandomForestClassifier()
kf_cv_scores = cross_val_score(rf, data_train, target_train, cv=skfold, scoring='f1_micro')
print("Random Forest CV-average F1-score: %.2f" % kf_cv_scores.mean())
# Holdout validation random forest
rf.fit(data_train,target_train)
prediction = rf.predict(data_test)
f1_mic= f1_score(target_test,prediction,average='micro')
f1_mac= f1_score(target_test,prediction,average='macro')
print("The F1-Score micro on test set: {:.4f}".format(f1_mic))
print("The F1-Score macro on test set: {:.4f}".format(f1_mac))
print('Confusion Matrix : \n' + str(confusion_matrix(target_test,prediction)))
# ## KNN (no Rebalance)
# cross-validation Knn
knn = KNeighborsClassifier()
kf_cv_scores = cross_val_score(knn, data_train, target_train, cv=skfold, scoring='f1_micro')
print("KNN CV-average F1-score: %.2f" % kf_cv_scores.mean())
# Holdout Knn
knn.fit(data_train,target_train)
prediction = knn.predict(data_test)
f1_mic= f1_score(target_test,prediction,average='micro')
f1_mac= f1_score(target_test,prediction,average='macro')
print("The F1-Score micro on test set: {:.4f}".format(f1_mic))
print("The F1-Score macro on test set: {:.4f}".format(f1_mac))
print('Confusion Matrix : \n' + str(confusion_matrix(target_test,prediction)))
# ## xgboost (no Rebalance)
# cross-validation xgboost
xgb = XGBClassifier()
kf_cv_scores = cross_val_score(xgb, data_train, target_train, cv=skfold, scoring='f1_micro')
print("XGBoost CV-average F1-score: %.2f" % kf_cv_scores.mean())
# Holdout xgboost
xgb.fit(data_train,target_train)
prediction = xgb.predict(data_test)
f1_mic= f1_score(target_test,prediction,average='micro')
f1_mac= f1_score(target_test,prediction,average='macro')
print("The F1-Score micro on test set: {:.4f}".format(f1_mic))
print("The F1-Score macro on test set: {:.4f}".format(f1_mac))
print('Confusion Matrix : \n' + str(confusion_matrix(target_test,prediction)))
|
notebooks/Model_Improvement.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # What is an optimization problem?
#
# A general mathematical formulation for **the optimization problems studied on this course** is
# $$
# \begin{align} \
# \min \quad &f(x)\\
# \text{s.t.} \quad & g_j(x) \geq 0\text{ for all }j=1,\ldots,J\\
# & h_k(x) = 0\text{ for all }k=1,\ldots,K\\
# &x\in \mathbb R^n.
# \end{align}
# $$
#
# The above problem can be expressed as
# >Find an $x\in \mathbb R^n$ such that $g_j(x)\geq 0$ for all $j=1,\ldots,J$ and $h_k(x)=0$ for all $k=1,\ldots,K$, and there does not exist $x'\in \mathbb R^n$ such that $f(x')<f(x)$ and $g_j(x')\geq 0$ for all $j=1,\ldots,J$, $h_k(x')=0$ for all $k=1,\ldots,K$.
# + [markdown] slideshow={"slide_type": "subslide"}
# There are three main components to an optimization problem:
# * the variables $x$ are called the **decision variables**,
# * the equalities and inequalities $g_j(x)\geq 0$ and $h_k(x)=0$ are called the **constraints**,
# * the funtion $f(x)$ is called the **objective function**.
#
# Values of decision variables $x^*$ are called **solutions** and a solution is called
# * **feasible** if $g_j(x^*)\geq 0$ for all $j=1,\ldots,J$, $h_k(x^*)=0$ for all $k=1,\ldots,K$,
# * **locally optimal** if $x^*$ is feasible and there exists $r>0$ such that there does not exist a feasible solution $x'\in \operatorname{B}(x^*,r)$ such that $f(x')<f(x^*)$, and
# * **optimal** if $x^*$ is feasible and there does not exist a feasible solution $x'$ such that $f(x')<f(x^*)$.
#
# The problem is called
# * **linear/nonlinear** if the objective function and the constraints of the problem are/are not affinely linear,
# * **multi/unimodal** if the problem has/does not have more than one local optimum,
# * **convex/nonconvex** if the objective and the constraints are that,
# * **continuous/differentiable/twice-differentiable, etc** if the objective and the constraints are that.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why do we study optimization problems?
#
# Because optimization problems arise in various fields engineering, finance, medicine etc.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Mixing problem
# Refinery produces 3 types of gasoline by mixing 3 different grude oil. Each grude oil can be purchased maximum of 5000 barrels per day. Let us assume that octane values and lead concentrations behave linearly in mixing. Refining costs are 4\$ per barrel and the capacity of the refinery is 14000 barrels per day. Demand of gasoline can be increased by (demand grows 10 barrels per day for each \$ used for advertizing). The details of the gasolines and Grude oils can be found in the following tables:
#
# | |Gasoline 1|Gasoline 2|Gasoline 3|
# |---|---|---|---|
# |Sale price|70|60|50|
# |Lower limit for octane|10|8|6|
# |Upper limit for lead|0.01|0.02|0.01|
# |Demand|3000|2000|1000|
# |Refining cost|4|4|4|
#
# | |Grude oil 1|Grude oil 2|Grude oil 3|
# |---|---|---|---|
# |Purchase price|45|35|25|
# |Octane value|12|6|8|
# |Lead concentration|0.005|0.02|0.03|
# |Availability|5000|5000|5000|
#
#
# Determine the production quantities of each type of gasoline, mixing ratios of different grude oil and advertising budget so that the profit is maximized and the demand is met exactly.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Modelling mixing problem as an optimization problem:
# * decision variables
# * $x_{ij}$ = amount of grude oil $i$ used for producing gasoline $j$
# * $y_j$ = the amount of money used for advertizing gasoline $j$
# * constraints
# * gasoline 1 demand: $x_{1,1}+x_{2,1}+x_{3,1}=3000+10y_1$
# * gasoline 2 demand: $x_{1,2}+x_{2,2}+x_{3,2}=2000+10y_2$
# * gasoline 3 demand: $x_{1,3}+x_{2,3}+x_{3,3}=1000+10y_3$
# * grude oil 1 availability: $x_{1,1}+x_{1,2}+x_{1,3}\leq 5000$
# * grude oil 2 availability: $x_{2,1}+x_{2,2}+x_{2,3}\leq 5000$
# * grude oil 3 availability: $x_{3,1}+x_{3,2}+x_{3,3}\leq 5000$
# * gasoline 1 octave value: $\frac{12x_{1,1}+6x_{2,1}+8x_{3,1}}{x_{1,1}+x_{2,1}+x_{3,1}}\geq 10$
# * gasoline 2 octave value: $\frac{12x_{1,2}+6x_{2,2}+8x_{3,2}}{x_{1,2}+x_{2,2}+x_{3,2}}\geq 8$
# * gasoline 3 octave value: $\frac{12x_{1,3}+6x_{2,3}+8x_{3,3}}{x_{1,3}+x_{2,3}+x_{3,3}}\geq 6$
# * gasoline 1 lead value: $\frac{0.005x_{1,1}+0.02x_{2,1}+0.03x_{3,1}}{x_{1,1}+x_{2,1}+x_{3,1}}\leq 0.01$
# * gasoline 2 lead value: $\frac{0.005x_{1,2}+0.02x_{2,2}+0.03x_{3,2}}{x_{1,2}+x_{2,2}+x_{3,2}}\leq 0.02$
# * gasoline 3 lead value: $\frac{0.005x_{1,3}+0.02x_{2,3}+0.03x_{3,3}}{x_{1,3}+x_{2,3}+x_{3,3}}\leq 0.01$
# * objective function:
# $$
# 70(x_{1,1}+x_{2,1}+x_{3,1})+60(x_{1,2}+x_{2,2}+x_{3,2})+50(x_{1,3}+x_{2,3}+x_{3,3})-45(x_{1,1}+x_{1,2}+x_{1,3})-35(x_{2,1}+x_{2,2}+x_{2,3})-25(x_{3,1}+x_{3,2}+x_{3,3})
# $$
# + [markdown] slideshow={"slide_type": "slide"}
# # How to solve optimization problems?
#
# ## Iterative vs. non-iterative methods
#
# Optimal solutions to some optimization problems can be found by defining an explicit formula for it. For example, if the objective function is twice continuously differentiable and there are no constraints, the optimal solution (if exists) can be found by calculating all the zero-points of the gradient and finding the best one of those. In this kind of cases, the optimization problem **can be solved using non-iterative methods.**
#
# **In this course we concentrate on the iterative methods.** Iterative methods are needed, if the problem has constraints, or the problem is in some other way not-well behaved (to be defined later, depending on the context). In iterative methods, the solving the optimizaiton problem starts from a so-called starting solution and then tries to improve the solution iteratively. The optimization algorithm chooses how the solution is changed at each iteration.
#
#
#
# ## What kind of methods will you learn in this course?
# Different optimization problems require different methods. In this course, we study optimization problems, which are
# * non-linear
# * not hugely multimodal
#
# Often the methods cannot guarantee a (global) optimum, but instead **we need to satisfy ourselves with a local optimum**. In addition, it is usually not possible to find the actual optimal solution, but instead **an approximation of the optimal solution**. A feasible solution $x^*$ is called an approximation of a local optimum $x^{**}$ with quality $L>0$, when $\|x^*-x^{**}\|\leq L$.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Line search
#
# Let us study optimization problem $\min_{x\in[a,b]} f(x)$, where $a,b\in\mathbb R$. Let us try to find an approximation of a local optimum to this problem.
# -
#Example objective function
def f(x):
return 2+(1-x)**2
print "The value of the objective function at 0 is " + str(f(0))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Line search with fixed steps
# **input:** the quality $L>0$ of the approximation of the local optimum.
# **output:** an approximation of the local optimum with quality $L$.
# ```
# start with x as the start point of the interval
# loop until stops:
# if the value of the objective is increasing for x+L from x
# stop, because the approximation of the locally optimal solution is x
# increase x by L
# ```
# -
def fixed_steps_line_search(a,b,f,L):
x = a
while f(x)>f(x+L) and x+L<b:
x=x+L
return x
# %timeit fixed_steps_line_search(0.0,3.0,f,1e-3)
#x = fixed_steps_line_search(0,3,f,1e-3); print "optimum is "+str(x)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The method of bisection
# **input:** the quality $L>0$ of the approximation of the local optimum.
# **output:** an approximation of the local optimum with quality $L$.
# ```
# Set x as the start point of interval and y as the end point
# while y-x<2*L:
# if the function is increasing at the mid point between x and y:
# set y as the midpoint between y and x, because a local optimum is before the midpoint
# otherwise:
# set x as the midpoint, because a local optimum is after the midpoint
# return midpoint between x and y
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# The following function is to be completed in class as an exercise
# -
def bisection_line_search(a,b,f,L,epsilon):
x = a
y = b
while y-x>2*L:
if f((x+y)/2+epsilon)>f((x+y)/2-epsilon):
y=(x+y)/2+epsilon
else:
x = (x+y)/2-epsilon
return (x+y)/2
# + [markdown] slideshow={"slide_type": "notes"}
# This is what we should end up. The following function is not shown on the slides.
# + slideshow={"slide_type": "skip"}
def bisection_line_search(a,b,f,L,epsilon):
x = a
y = b
while y-x<2*L:
if f((x+y)/2-epsilon)<f((x+y)/2+epsilon):
y = (x+y)/2+epsilon
else:
x = (x+y)/2-epsilon
return (x+y)/2
# -
# %timeit bisection_line_search(0.0,3.0,f,1e-3,1e-4)
#x = bisection_line_search(0.0,3.0,f,0.01,1e-4); print "x="+str(x)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Golden section search
#
# ### Golden section
#
# Let $a<c<b$ be such that $\frac{b-a}{c-a}=\frac{c-a}{b-c}$. Then it is said that the point $c$ devides interval $[a,b]$ in the ratio of golden section (from the left, mirror from the right). Note that $c=a+\frac{\sqrt{5}-1}2(b-a)\approx a+0.618(b-a)$.
#
# 
#
# There is a theorem that if $a<c<d<b$ and both points divide the interval $[a,b]$ in the ratio of golden section (from right and left), then the point $c$ divides the interval $[a,d]$ in the ratio of golder ration from the left.
#
# 
#
# ### Golden section search algorithm
#
#
# **input:** the quality $L>0$ of the approximation of the local optimum.
# **output:** an approximation of the local optimum with quality $L$.
# ```
# Set x as the start point of interval and y as the end point
# while y-x>2*L:
# Divide the interval [x,y] in the golden section from the letf and right and attain two division points
# If the greater of the division points has a greater function value
# set y as the rightmost division point, because a local optimum is before that
# otherwise:
# set x as the leftmost division point, because a local optimum is after that
# return midpoint between x and y
# ```
# + [markdown] slideshow={"slide_type": "notes"}
# The following function is to be completed in class as an exercise
# + slideshow={"slide_type": "-"}
import math
def golden_section_line_search(a,b,f,L):
x = a
y = b
while y-x>2*L:
if f(x+(math.sqrt(5.0)-1)/2.0*(y-x))<f(y-(math.sqrt(5.0)-1)/2.0*(y-x)):
x = y-(math.sqrt(5.0)-1)/2.0*(y-x)
else:
y = x+(math.sqrt(5.0)-1)/2.0*(y-x)
return (x+y)/2
# -
# %timeit golden_section_line_search(0.0,3.0,f,1e-3)
#x = golden_section_line_search(0.0,3.0,f,0.01); print "x="+str(x)
|
Lecture 2, What is an optimization problem and how to solve them and line search.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# THis is pytorch tutorial.
from __future__ import print_function
import torch
# Construct a 5x3 matrix, uninitialized:
x = torch.empty(5, 3)
print(x)
# +
# Construct a matrix filled zeros and of dtype long:
x = torch.zeros(5, 3, dtype=torch.long)
print(x)
# -
# Construct a tensor directly from data:
x = torch.tensor([5.5, 3])
print(x)
# +
# or create a tensor based on an existing tensor.
# hese methods will reuse properties of the input tensor, e.g. dtype, unless new values are provided by user
x = x.new_ones(5, 3, dtype=torch.double) # new_* methods take in sizes
print(x)
x = torch.randn_like(x, dtype=torch.float) # override dtype!
print(x) # result has the same size
# -
# # H1
# ## H2
# ### H3
print(x.size())
y = torch.rand(5, 3)
print(x)
print(y)
print(x + y)
print(torch.add(x, y))
# +
# Addition: providing an output tensor as argument
result = torch.empty(5, 3)
torch.add(x, y, out=result)
print(result)
# +
# Inplace
y.add_(x)
print(y)
# -
|
content/learning_curves/machine_learning/.ipynb_checkpoints/pytorch_starting_tutorial-checkpoint.ipynb
|