code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # calculate confusion matrix - precision - recall -
# +
import pandas as pd
from sklearn.linear_model import LogisticRegression
x1 = [-1.0, -0.75, -0.5, -0.25, 0.0, 0.25, 0.5, 0.75, 1.0]
x2 = [abs(x) for x in x1]
y = [1, 1, 1, 0, 0, 0, 1, 1, 1]
df = pd.DataFrame({'x1':x1, 'x2':x2, 'y': y})
X = df[['x1']]
y = df['y']
m = LogisticRegression(C=1e10)
m.fit(X, y)
print('score with x1 only : {:4.2f}'.format(m.score(X, y)))
X = df[['x1', 'x2']]
m.fit(X, y)
print('score with x2=abs(x1) : {:4.2f}'.format(m.score(X, y)))
# -
X = df[['x1']]
m = LogisticRegression(C=1e10)
m.fit(X, y)
m.score(X, y)
from sklearn.metrics import precision_score
# +
ypred = m.predict(X)
y
# -
ypred
precision_score(y, ypred)
from sklearn.metrics import recall_score
recall_score(y, ypred)
from sklearn.metrics import confusion_matrix
confusion_matrix(y, ypred)
# +
# top left: TN
# bottom right: TP
# bottom left: FN
# Top right: FP
# TP : surviving passenger correctly predcited
# TN : drowned passenger correctly predicted
# FP : drowned passenger predicted as surviving
# FN : surviving passenger predicted as dead
# -
precision_score(y, ypred)
precision_score(ypred, y)
# +
# probability
m.predict_proba(X)
# -
# part 3: probabilites and threshold values
#
p = m.predict_proba(X)
posp = p[:, 1] # select all rows, 2nd column
threshold = 0.5
for value in posp:
if value > threshold:
print('positive')
else:
print('negative')
# try threshold values 0.0000000001 and 0.9999999999 as well
p = m.predict_proba(X)
posp = p[:, 1] # select all rows, 2nd column
threshold = 0.9
for value in posp:
if value > threshold:
print('positive')
else:
print('negative')
p = m.predict_proba(X)
posp = p[:, 1] # select all rows, 2nd column
threshold = 0.1
for value in posp:
if value > threshold:
print('positive')
else:
print('negative')
|
Week_02/Titanic/5.2.4. Evaluating Classifiers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-datascience
# language: python
# name: venv-datascience
# ---
# # Principal Component Analysis
#
# Let's discuss PCA! Since this isn't exactly a full machine learning algorithm, but instead an unsupervised learning algorithm, we will just have a lecture on this topic, but no full machine learning project (although we will walk through the cancer set with PCA).
#
# ## PCA Review
#
# Remember that **PCA is just a transformation of your data and attempts to find out what features explain the most variance in your data**. For example:
# <img src='PCA.png' />
# # Import Libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# # Data
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
type(cancer)
cancer
cancer.keys()
cancer['data'][:2]
print(cancer['DESCR'])
df = pd.DataFrame(cancer['data'], columns=cancer['feature_names'])
df.head()
# ---------
# ## PCA Visualization
#
# As we've noticed before it is **difficult to visualize high dimensional data, we can use PCA to find the first two principal components**, and visualize the data in this new, two-dimensional space, with a single scatter-plot.
#
# Before we do this though, we'll need to scale our data so that each feature has a single unit variance.
df.shape
# Now we are dealing with 30 features (30 dimensions).
# ### Scaling
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_data = scaler.fit_transform(df)
# # PCA (Dimensionality Reduction)
# PCA with Scikit Learn uses a very similar process to other preprocessing functions that come with SciKit Learn. We instantiate a PCA object, find the principal components using the fit method, then apply the rotation and dimensionality reduction by calling transform().
#
# We can also specify how many components we want to keep when creating the PCA object.
from sklearn.decomposition import PCA
# the module name decomposition makes sense because PCA ensentially decompse the high dimensions data into selected dimensions
pca = PCA(n_components=2)
# now we want to use use 2 components
x_pca = pca.fit_transform(scaled_data)
# check the shape of original data, there are 30 columns / feataures
scaled_data.shape
# now we can see there are only 2 features after PCA
x_pca.shape
# We've reduced 30 dimensions to just 3! Let's plot these two dimensions out!
#
#
plt.figure(figsize=(8,6))
plt.scatter(x_pca[:, 0], x_pca[:, 1], c=cancer['target']); # color by target columns
plt.xlabel('First Principal Component')
plt.ylabel('Second Principal Component')
# Clearly by using these two components we can easily separate these two classes.
#
# ## Interpreting the components
#
# Unfortunately, with this great power of dimensionality reduction, comes the cost of being able to easily understand what these components represent.
#
# * The components correspond to combinations of the original features (They **don't relate ONE TO ONE relationship** with the original features).
# * the components themselves are stored as an attribute of the fitted PCA object:
pca.components_
# In above numpy array,
#
# - each rows represent Principal Component.
# - each columns actually related back original features.
#
# we can visualize this relationship with a heatmap:
# convert into dataframe
df_comp = pd.DataFrame(pca.components_, columns=cancer['feature_names'])
df_comp.head()
plt.figure(figsize=(12,6))
sns.heatmap(df_comp, cmap='plasma');
# This heatmap and the color bar basically represent the correlation between the various feature and the principal component itself.
#
# Referring from heatmap above, we can see that
# * Component 1 is strongly negatively correlated with `mean radius`, `mean perimeter`, `mean_area`, `worst radius`, `worst perimeter`, `worst area`. (blue color)
#
# * Component 1 is strongly postively correlated with `mean fractal dimention`. (yellow color)
# # Explained Variance Ratio of PCA
#
# - Explained Variance Ratio tells us how much information is compressed into first few components.
# - When you are deciding how much components to keep, look at the percent of cumulative variance. Make sure to **retain at least 70%** of dataset's original information
pca.explained_variance_ratio_
# In the above ratio, we can see
# * 44.27% of information is kept in first component and
# * 18.97% in second component.
#
# but in our case total is only 63% of original data. So we might want to use different number of components to caputure at least 70% of original data.
# ### Cumulative Variance
pca.explained_variance_ratio_.sum() # 63% of information is captured in the two components that were returned.
# ---------
# # New PCA model (to cover 70% of original information)
pca = PCA(n_components=4)
x_pca = pca.fit_transform(scaled_data)
pca.components_[:2]
pca.explained_variance_ratio_
# In our new PCA, our 4 components cover nearly 80% of our original information.
#
# **We can even use only first 3 components as the total cover 70% already.**
#
# 0.44272026 + 0.18971182 + 0.09393163 = 0.72636371
pca.explained_variance_ratio_.sum()
df_comp = pd.DataFrame(pca.components_, columns=cancer['feature_names'])
df_comp.head()
plt.figure(figsize=(10,5))
sns.heatmap(df_comp, cmap='plasma');
# # Summary
#
# ### By using those information,
# ### 1) we can select how many first componets we want to use (to cover at least 70%)
# ### 2) which features to use by referring those chosen components
# ### to proceed with Classification modelling
|
Data Science and Machine Learning Bootcamp - JP/19. Principal-Component-Analysis/01-Principal Component Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="JkUviKhSYLMF" slideshow={"slide_type": "slide"}
# ## Climatology and Anomaly in Geoscience: Part 2
# An example of how Python is used to explore global sea surface temperature (SST).
# * <NAME> (<EMAIL>)
# * Department of Geosciences, Princeton University
# * Junior Colloquium, Nov. 15, 2021
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## A short review of the last session
# 1. `xarray` basic usage: `open_dataset`, `sel`, `isel`, `mean`, `plot`, `plot.contourf`, `groupby`.
# 2. Sea surface temperature dataset: ERSST5.
# 3. SST annual climatology and change, monthly climatology.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Today's plan
#
# 1. SST anomaly (from the monthly climatology).
# 2. El Nino index (Nino3.4) time series (`mean` over lon/lat region box).
# 3. What's next after JC.
# 3. Interactive Q&A.
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Start analysis
# + colab={} colab_type="code" id="2nyvaFZnYYtm" slideshow={"slide_type": "fragment"}
# import and configuration
# xarray is the core package we are going to use
import xarray as xr
import matplotlib.pyplot as plt # we also use pyplot directly in some cases
import os
# some configurations on the default figure output
# %config InlineBackend.figure_format ='retina'
plt.rcParams['figure.dpi'] = 120
# + colab={"base_uri": "https://localhost:8080/", "height": 244} colab_type="code" executionInfo={"elapsed": 835, "status": "ok", "timestamp": 1568235456809, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="bik3mA4tYefp" outputId="c95cc7cf-0866-43b3-ca86-44fa0d0320f8" slideshow={"slide_type": "subslide"}
# analysis of last session
# sst data file
# Am I running the notebook on Adroit or not?
if os.uname().nodename.startswith('adroit'):
ifile = '/home/wenchang/JC2021/ersst5_1979-2018.nc'
else:
ifile = 'ersst5_1979-2018.nc'
print('file to be opened:',ifile)
# open the sst dataset
ds = xr.open_dataset(ifile)
# get the sst DataArray
sst = ds.sst
# 40-year annual mean climatology
sst_clim = sst.mean('time')
# difference between the first and last 10-year climatologies
sst_early = sst.sel(time=slice('1979-01', '1988-12')).mean('time')
sst_late = sst.sel(time=slice('2009-01', '2018-12')).mean('time')
dsst = sst_late - sst_early
dsst.attrs['long_name'] = 'SST change from 1979-1988 to 2009-2018'
dsst.attrs['units'] = '$^\circ$C'
# monthly climatology: the time dimension is now replaced by month
sst_mclim = sst.groupby('time.month').mean('time')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 4631, "status": "ok", "timestamp": 1567700569017, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="CDFFxfZBicEF" outputId="3e13395e-1d61-4301-d096-048cab906141" slideshow={"slide_type": "slide"}
# Make 12 subplots using a single command.
sst_mclim.plot.contourf(col='month', col_wrap=6, levels=20,
cmap='Spectral_r', center=False)
# + [markdown] colab_type="text" id="qDimD2IIAp22" slideshow={"slide_type": "slide"}
# ## Calculate monthly anomaly
# Subtract the monthly climatology from the raw SST data.
# + colab={"base_uri": "https://localhost:8080/", "height": 561} colab_type="code" executionInfo={"elapsed": 800, "status": "ok", "timestamp": 1567700593776, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="TQBWe3yxjptK" outputId="4dc7bdc7-1132-451b-8a2d-c61d3214683d" slideshow={"slide_type": "subslide"}
ssta = sst.groupby('time.month') - sst_mclim
# monthly climatology is now removed
# + slideshow={"slide_type": "subslide"}
ssta
# + colab={"base_uri": "https://localhost:8080/", "height": 507} colab_type="code" executionInfo={"elapsed": 1136, "status": "ok", "timestamp": 1567700660707, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="OKZ-I_UJkTMk" outputId="675e0cdd-cd18-4e17-fc3a-3c871b0f5227" slideshow={"slide_type": "subslide"}
# The 1997 winter is a big El Nino season
ssta.sel(time=slice('1997-12', '1998-02')).mean('time').plot.contourf(levels=19)
# + colab={"base_uri": "https://localhost:8080/", "height": 507} colab_type="code" executionInfo={"elapsed": 1136, "status": "ok", "timestamp": 1567700660707, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="OKZ-I_UJkTMk" outputId="675e0cdd-cd18-4e17-fc3a-3c871b0f5227" slideshow={"slide_type": "subslide"}
# The 1998 winter is a big La Nina season
ssta.sel(time=slice('1998-12', '1999-02')).mean('time').plot.contourf(levels=19)
# + colab={"base_uri": "https://localhost:8080/", "height": 507} colab_type="code" executionInfo={"elapsed": 1136, "status": "ok", "timestamp": 1567700660707, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="OKZ-I_UJkTMk" outputId="675e0cdd-cd18-4e17-fc3a-3c871b0f5227" slideshow={"slide_type": "subslide"}
# The 2015 winter is also a big El Nino season
ssta.sel(time=slice('2015-12', '2016-02')).mean('time').plot.contourf(levels=19)
# + [markdown] colab_type="text" id="ROmgatpmA1og" slideshow={"slide_type": "slide"}
# ## Calculate the Nino3.4 index
# * SST anomaly averaged over the Nino3.4 region: 170W-120W, 5S-5N
# 
# http://www.bom.gov.au/climate/enso/indices/oceanic-indices-map.gif
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" executionInfo={"elapsed": 453, "status": "ok", "timestamp": 1567700672119, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="fusZ4HXhuDXt" outputId="d962bf54-3d16-4684-9ca7-a5bf66739104" slideshow={"slide_type": "subslide"}
nino34 = ssta.sel(lon=slice(360-170,360-120),lat=slice(-5,5)).mean(['lon','lat'])
nino34.attrs['long_name'] = 'Nino3.4 index'
nino34.attrs['units'] = '$^\circ$C'
# + slideshow={"slide_type": "subslide"}
nino34
# + colab={"base_uri": "https://localhost:8080/", "height": 492} colab_type="code" executionInfo={"elapsed": 1663, "status": "ok", "timestamp": 1567700675793, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="h7W9LsfhuP2i" outputId="1d8aa14d-32ba-423c-da48-b06c8e816c3a" slideshow={"slide_type": "subslide"}
nino34.plot()
# plt.axvline('2015-12', color='gray', ls='--')
# plt.text('2015-12', 2, '2015-12', rotation=-45, color='gray', )
# plt.axvline('1997-12', color='gray', ls='--')
# plt.text('1997-12', 2, '1997-12', rotation=-45, color='gray', )
# plt.axvline('1982-12', color='gray', ls='--')
# plt.text('1982-12', 2, '1982-12', rotation=-45, color='gray', )
# + [markdown] slideshow={"slide_type": "slide"}
# ## Seasonality of El Nino/La Nina
# + colab={"base_uri": "https://localhost:8080/", "height": 473} colab_type="code" executionInfo={"elapsed": 1155, "status": "ok", "timestamp": 1567700813854, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="Z60v5lhZ1vG2" outputId="74dff819-35de-4ecf-f666-c9eaa2dad22b" slideshow={"slide_type": "slide"}
# compare Nino3.4 index variability for each month
da = nino34.groupby('time.month').std('time') # standard deviation for each month
da.plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 473} colab_type="code" executionInfo={"elapsed": 1155, "status": "ok", "timestamp": 1567700813854, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzexWcchBOvHGw9u_Nm2D-vWc4ApTqQ4uLX1i-=s64", "userId": "02317458745209383076"}, "user_tz": 240} id="Z60v5lhZ1vG2" outputId="74dff819-35de-4ecf-f666-c9eaa2dad22b" slideshow={"slide_type": "slide"}
# compare Nino3.4 index variability for each month
da = nino34.groupby('time.month').std('time') # standard deviation for each month
da = da.roll(month=-5).assign_coords(month=range(6, 18)) # roll the time series to start from Jun
da.plot()
#add more descriptive label to y axis
plt.ylabel('Nino3.4 standard deviation [$^\circ$C]')
#use month names as the x tick labels
plt.xticks(range(6,18), ['Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec', 'Jan', 'Feb', 'Mar', 'Apr', 'May'])
#add grid lines to axes
plt.grid(True)
# plt.bar(da.month, da.values, color='lightgray')
# + [markdown] slideshow={"slide_type": "subslide"}
# December shows the largest variability!
# + [markdown] slideshow={"slide_type": "slide"}
# ## What's next
#
# * Python resource: https://wy2136.github.io/python.html
# * Popular packages: `xarray`, `pandas`, `numpy`, `matplotlib`.
# * Machine learning: `sklearn`, `keras`, `pytorch`.
# * Try to apply these tools to your project.
# * Google usually helps if you have a question.
# * Ask around if google doesn't work.
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Q&A
|
JC2021/07_clim_anom_2021-11-15_practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # High-level CNTK Example
import numpy as np
import os
import sys
import cntk
from cntk.layers import Convolution2D, MaxPooling, Dense, Dropout
from common.params import *
from common.utils import *
# Force one-gpu
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
print("OS: ", sys.platform)
print("Python: ", sys.version)
print("Numpy: ", np.__version__)
print("CNTK: ", cntk.__version__)
print("GPU: ", get_gpu_name())
print(get_cuda_version())
print("CuDNN Version ", get_cudnn_version())
def create_symbol(n_classes=N_CLASSES):
# Weight initialiser from uniform distribution
# Activation (unless states) is None
with cntk.layers.default_options(init = cntk.glorot_uniform(), activation = cntk.relu):
x = Convolution2D(filter_shape=(3, 3), num_filters=50, pad=True)(features)
x = Convolution2D(filter_shape=(3, 3), num_filters=50, pad=True)(x)
x = MaxPooling((2, 2), strides=(2, 2), pad=False)(x)
x = Dropout(0.25)(x)
x = Convolution2D(filter_shape=(3, 3), num_filters=100, pad=True)(x)
x = Convolution2D(filter_shape=(3, 3), num_filters=100, pad=True)(x)
x = MaxPooling((2, 2), strides=(2, 2), pad=False)(x)
x = Dropout(0.25)(x)
x = Dense(512)(x)
x = Dropout(0.5)(x)
x = Dense(n_classes, activation=None)(x)
return x
def init_model(m, labels, lr=LR, momentum=MOMENTUM):
# Loss (dense labels); check if support for sparse labels
loss = cntk.cross_entropy_with_softmax(m, labels)
# Momentum SGD
# https://github.com/Microsoft/CNTK/blob/master/Manual/Manual_How_to_use_learners.ipynb
# unit_gain=False: momentum_direction = momentum*old_momentum_direction + gradient
# if unit_gain=True then ...(1-momentum)*gradient
learner = cntk.momentum_sgd(m.parameters,
lr=cntk.learning_rate_schedule(lr, cntk.UnitType.minibatch) ,
momentum=cntk.momentum_schedule(momentum),
unit_gain=False)
return loss, learner
# %%time
# Data into format for library
x_train, x_test, y_train, y_test = cifar_for_library(channel_first=True, one_hot=True)
# CNTK format
y_train = y_train.astype(np.float32)
y_test = y_test.astype(np.float32)
print(x_train.shape, x_test.shape, y_train.shape, y_test.shape)
print(x_train.dtype, x_test.dtype, y_train.dtype, y_test.dtype)
# %%time
# Placeholders
features = cntk.input_variable((3, 32, 32), np.float32)
labels = cntk.input_variable(N_CLASSES, np.float32)
# Load symbol
sym = create_symbol()
# %%time
loss, learner = init_model(sym, labels)
# %%time
# Main training loop: 49s
loss.train((x_train, y_train),
minibatch_size=BATCHSIZE,
max_epochs=EPOCHS,
parameter_learners=[learner])
# %%time
# Main evaluation loop: 409ms
n_samples = (y_test.shape[0]//BATCHSIZE)*BATCHSIZE
y_guess = np.zeros(n_samples, dtype=np.int)
y_truth = np.argmax(y_test[:n_samples], axis=-1)
c = 0
for data, label in yield_mb(x_test, y_test, BATCHSIZE):
predicted_label_probs = sym.eval({features : data})
y_guess[c*BATCHSIZE:(c+1)*BATCHSIZE] = np.argmax(predicted_label_probs, axis=-1)
c += 1
print("Accuracy: ", 1.*sum(y_guess == y_truth)/len(y_guess))
|
deep-learning/multi-frameworks/notebooks/CNTK_CNN_highAPI.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Generating `publications.json` partitions
# This is a template notebook for generating metadata on publications - most importantly, the linkage between the publication and dataset (datasets are enumerated in `datasets.json`)
#
# Process goes as follows:
# 1. Import CSV with publication-dataset linkages. Your csv should have at the minimum, fields (spelled like the below):
# * `dataset` to hold the dataset_ids, and
# * `title` for the publication title.
#
# Update the csv with these field names to ensure this code will run. We read in, dedupe and format the title
# 2. Match to `datasets.json` -- alert if given dataset doesn't exist yet
# 3. Generate list of dicts with publication metadata
# 4. Write to a publications.json file
# #### Import CSV containing publication-dataset linkages
# Set `linkages_path` to the location of the csv containg dataset-publication linkages and read in csv
import pandas as pd
import os
import datetime
file_name = 'bundesbank_pub_dataset_linkages_sbr_final.csv'
rcm_subfolder = '20190731_bundesbank'
parent_folder = '/Users/sophierand/RichContextMetadata/metadata'
linkages_path = os.path.join(parent_folder,rcm_subfolder,file_name)
# linkages_path = os.path.join(os.getcwd(),'SNAP_DATA_DIMENSIONS_SEARCH_DEMO.csv')
linkages_csv = pd.read_csv(linkages_path)
# Format/clean linkage data - apply `scrub_unicode` to `title` field.
import unicodedata
def scrub_unicode (text):
"""
try to handle the unicode edge cases encountered in source text,
as best as possible
"""
x = " ".join(map(lambda s: s.strip(), text.split("\n"))).strip()
x = x.replace('’', "'").replace('–',"--")
x = x.replace('“', '"').replace('”', '"')
x = x.replace("‘", "'").replace("’", "'").replace("`", "'")
x = x.replace("`` ", '"').replace("''", '"')
x = x.replace('…', '...').replace("\\u2026", "...")
x = x.replace("\\u00ae", "").replace("\\u2122", "")
x = x.replace("\\u00a0", " ").replace("\\u2022", "*").replace("\\u00b7", "*")
x = x.replace("\\u2018", "'").replace("\\u2019", "'").replace("\\u201a", "'")
x = x.replace("\\u201c", '"').replace("\\u201d", '"')
x = x.replace("\\u20ac", "€")
x = x.replace("\\u2212", " - ") # minus sign
x = x.replace("\\u00e9", "é")
x = x.replace("\\u017c", "ż").replace("\\u015b", "ś").replace("\\u0142", "ł")
x = x.replace("\\u0105", "ą").replace("\\u0119", "ę").replace("\\u017a", "ź").replace("\\u00f3", "ó")
x = x.replace("\\u2014", " - ").replace('–', '-').replace('—', ' - ')
x = x.replace("\\u2013", " - ").replace("\\u00ad", " - ")
x = str(unicodedata.normalize("NFKD", x).encode("ascii", "ignore").decode("utf-8"))
# some content returns text in bytes rather than as a str ?
try:
assert type(x).__name__ == "str"
except AssertionError:
print("not a string?", type(x), x)
return x
# Scrub titles of problematic characters, drop nulls and dedupe
linkages_csv = linkages_csv.loc[pd.notnull(linkages_csv.dataset)].drop_duplicates()
linkages_csv = linkages_csv.loc[pd.notnull(linkages_csv.title)].drop_duplicates()
linkages_csv['title'] = linkages_csv['title'].apply(scrub_unicode)
pub_metadata_fields = ['title']
original_metadata_cols = list(set(linkages_csv.columns.values.tolist()) - set(pub_metadata_fields)-set(['dataset']))
# #### Generate list of dicts of metadata
# Read in `datasets.json`. Update `datasets_path` to your local.
import json
# +
datasets_path = '/Users/sophierand/RCDatasets/datasets.json'
with open(datasets_path) as json_file:
datasets = json.load(json_file)
# -
# Create list of dictionaries of publication metadata. `format_metadata` iterrates through `linkages_csv` dataframe, splits the `dataset` field (for when multiple datasets are listed); throws an error if the dataset doesn't exist and needs to be added to `datasets.json`.
def create_pub_dict(linkages_dataframe,datasets):
pub_dict_list = []
for i, r in linkages_dataframe.iterrows():
r['title'] = scrub_unicode(r['title'])
ds_id_list = [f for f in [d.strip() for d in r['dataset'].split(",")] if f not in [""," "]]
for ds in ds_id_list:
check_ds = [b for b in datasets if b['id'] == ds]
if len(check_ds) == 0:
print('dataset {} isnt listed in datasets.json. Please add to file'.format(ds))
required_metadata = r[pub_metadata_fields].to_dict()
required_metadata.update({'datasets':ds_id_list})
pub_dict = required_metadata
if len(original_metadata_cols) > 0:
original_metadata = r[original_metadata_cols].to_dict()
original_metadata.update({'date_added':datetime.datetime.now().strftime('%Y-%m-%d %H:%M:%S')})
pub_dict.update({'original':original_metadata})
pub_dict_list.append(pub_dict)
return pub_dict_list
# Generate publication metadata and export to json
linkage_list = create_pub_dict(linkages_csv,datasets)
# Update `pub_path` to be:
# `<name_of_subfolder>_publications.json`
json_pub_path = os.path.join('/Users/sophierand/RCPublications/partitions/',rcm_subfolder+'_publications.json')
with open(json_pub_path, 'w') as outfile:
json.dump(linkage_list, outfile, indent=2)
json_pub_path
|
metadata/20190731_bundesbank/publications_export_template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# from sklearn
import pandas as pd
import seaborn as sns; sns.set()
import pickle
import matplotlib as plt
# %matplotlib inline
token_data=pd.read_csv("../data/wowcointotal.csv")
token_data.head()
token_data.shape
# +
token_data.rename(columns=lambda x: x.replace(" ","_"), inplace=True)
token_data.rename(columns=lambda x: x.lower(), inplace=True)
## Previous trial
# splitted_date= [str(token_data["Date"][i]).split(" ") for i in range(len(token_data["Date"]))]
# days=[splitted_date[i][0] for i in range(len(splitted_date))]
# hour=[splitted_date[i][1] for i in range(len(splitted_date))]
# hour=[hour[i][:-2]+ "00" for i in range(len(hour))]
# token_data["day"] = days
# token_data["hour"] = hour
# This one line does what I've tried to achieve above
token_data['date_time'] = pd.to_datetime(token_data['date'])
token_data['date_time'] = token_data["date_time"].dt.strftime('%Y-%m-%d')
token_data.drop("date",axis=1,errors="ignore",inplace=True)
token_data.head()
# -
token_data.dtypes
token_data.dtypes
print(token_data.region.value_counts())
print("\n")
print(token_data.time_left_on_auction.value_counts())
sub_US = token_data[token_data["region"]=="US"]
sub_US.shape
sub_US.drop(labels="region",axis=1,inplace=True)
sub_US.head()
sub_US_long = sub_US[sub_US["time_left_on_auction"]=="Long"]
sub_US_short = sub_US[sub_US["time_left_on_auction"]=="Short"]
sub_US_meedium = sub_US[sub_US["time_left_on_auction"]=="Medium"]
sub_US_very_long = sub_US[sub_US["time_left_on_auction"]=="Very Long"]
sub_US_long.shape
sub_US_long_model = sub_US_long.copy()
sub_US_long_model.drop("time_left_on_auction",axis=1,inplace=True,errors="ignore")
sub_US_long_model.head()
sub_US_long_model = sub_US_long_model.groupby("date_time").mean()
sub_US_long_model.head()
sns.set(rc={'figure.figsize':(14,8)})
sub_US_long.plot(x="date_time",y="price")
sub_US_long_model.plot(y="price")
sns.lineplot(x="date_time", y="price", hue="time_left_on_auction", data=sub_US)
sub_US_long_model.dtypes
from statsmodels.tsa.arima_model import ARIMA
model = ARIMA(sub_US_long_model,order=(5,1,0))
model_fit= model.fit(disp=0)
print(model_fit.summary())
from sklearn.metrics import mean_squared_error
from matplotlib import pyplot
# +
X=sub_US_long_model.values
size = int(len(X) * 0.66)
train, test = X[0:size], X[size:len(X)]
history = [x for x in train]
predictions = list()
for t in range(len(test)):
model = ARIMA(history, order=(5,1,0))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
predictions.append(yhat)
obs = test[t]
history.append(obs)
print('predicted=%f, expected=%f' % (yhat, obs))
error = mean_squared_error(test, predictions)
print('Test MSE: %.3f' % error)
# -
# plot
pyplot.plot(test)
pyplot.plot(predictions, color='red')
pyplot.show()
sub_US_long_model.tail()
# +
# # start_index = pd.datetime(2017, 7,15 ).date()
# # end_index = pd.datetime(2017,7,19).date()
# start_index = pd.to_datetime("2017-07-15").strftime('%Y-%m-%d')
# end_index = pd.to_datetime("2017-07-20").strftime('%Y-%m-%d')
# end_index
forecast = model_fit.forecast(steps=8)
# -
pyplot.plot(forecast[0])
filename = 'token_predictor_model.pkl'
pickle.dump(model_fit, open(filename, 'wb'))
|
app/deploy_ml/notebook/warcraft_token_prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quantum Process Tomography with Q# and Python #
# ## Abstract ##
# In this sample, we will demonstrate interoperability between Q# and Python by using the QInfer and QuTiP libraries for Python to characterize and verify quantum processes implemented in Q#.
# In particular, this sample will use *quantum process tomography* to learn about the behavior of a "noisy" Hadamard operation from the results of random Pauli measurements.
# ## Preamble ##
import warnings
warnings.simplefilter('ignore')
# We can enable Q# support in Python by importing the `qsharp` package.
import qsharp
# Once we do so, any Q# source files in the current working directory are compiled, and their namespaces are made available as Python modules.
# For instance, the `Quantum.qs` source file provided with this sample implements a `HelloWorld` operation in the `Microsoft.Quantum.Samples.Python` Q# namespace:
with open('Quantum.qs') as f:
print(f.read())
# We can import this `HelloWorld` operation as though it was an ordinary Python function by using the Q# namespace as a Python module:
from Microsoft.Quantum.Samples.Python import HelloWorld
HelloWorld
# Once we've imported the new names, we can then ask our simulator to run each function and operation using the `simulate` method.
HelloWorld.simulate(pauli=qsharp.Pauli.Z)
# ## Tomography ##
# The `qsharp` interoperability package also comes with a `single_qubit_process_tomography` function which uses the QInfer library for Python to learn the channels corresponding to single-qubit Q# operations.
from qsharp.tomography import single_qubit_process_tomography
# Next, we import plotting support and the QuTiP library, since these will be helpful to us in manipulating the quantum objects returned by the quantum process tomography functionality that we call later.
# %matplotlib inline
import matplotlib.pyplot as plt
import qutip as qt
qt.settings.colorblind_safe = True
# To use this, we define a new operation that takes a preparation and a measurement, then returns the result of performing that tomographic measurement on the noisy Hadamard operation that we defined in `Quantum.qs`.
experiment = qsharp.compile("""
open Microsoft.Quantum.Samples.Python;
open Microsoft.Quantum.Characterization;
operation Experiment(prep : Pauli, meas : Pauli) : Result {
return SingleQubitProcessTomographyMeasurement(prep, meas, NoisyHadamardChannel(0.1));
}
""")
# Here, we ask for 10,000 measurements from the noisy Hadamard operation that we defined above.
estimation_results = single_qubit_process_tomography(experiment, n_measurements=10000)
# To visualize the results, it's helpful to compare to the actual channel, which we can find exactly in QuTiP.
depolarizing_channel = sum(map(qt.to_super, [qt.qeye(2), qt.sigmax(), qt.sigmay(), qt.sigmaz()])) / 4.0
actual_noisy_h = 0.1 * qt.to_choi(depolarizing_channel) + 0.9 * qt.to_choi(qt.hadamard_transform())
# We then plot the estimated and actual channels as Hinton diagrams, showing how each acts on the Pauli operators $X$, $Y$ and $Z$.
fig, (left, right) = plt.subplots(ncols=2, figsize=(12, 4))
plt.sca(left)
plt.xlabel('Estimated', fontsize='x-large')
qt.visualization.hinton(estimation_results['est_channel'], ax=left)
plt.sca(right)
plt.xlabel('Actual', fontsize='x-large')
qt.visualization.hinton(actual_noisy_h, ax=right)
# We also obtain a wealth of other information as well, such as the covariance matrix over each parameter of the resulting channel.
# This shows us which parameters we are least certain about, as well as how those parameters are correlated with each other.
plt.figure(figsize=(10, 10))
estimation_results['posterior'].plot_covariance()
plt.xticks(rotation=90)
# ## Diagnostics ##
for component, version in sorted(qsharp.component_versions().items(), key=lambda x: x[0]):
print(f"{component:20}{version}")
import sys
print(sys.version)
|
samples/interoperability/python/tomography-sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 1 - TEST
#
# ## Overview
# As explained in the [Before week 1: How to take this class](https://nbviewer.jupyter.org/github/suneman/socialdata2021/blob/main/lectures/How_To_Take_This_Class.ipynb) notebook, each week of this class is an IPython notebook like this one. In order to follow the class, you simply start reading from the top, following the instructions.
#
# **Hint**: And (as explained [here](https://nbviewer.jupyter.org/github/suneman/socialdata2021/blob/main/lectures/COVID-update-1.ipynb)) you can ask me - or any of the friendly Teaching Assistants - for help at any point if you get stuck!
#
# ## Intro video
# Below is today's informal intro video, tying everything together and making each and everyone of you feel welcome and loved!
# short video explaining the plans
from IPython.display import YouTubeVideo
YouTubeVideo("U5hVaBtncNc",width=600, height=333)
# ## Today
#
# This first lecture will go over a few different topics to get you started
#
# * First, I'll explain a little bit about what we'll be doing this year (hint, you may want to watch _Minority Report_ if you want to prepare deeply for the class).
# * Second, we'll start by loading some real-world data into your very own computers and getting started with some data analysis.
# ## Part 1: Predictive policing. A case to learn from
#
# For a number of years I've been a little bit obsessed with [predictive policing](https://www.sciencemag.org/news/2016/09/can-predictive-policing-prevent-crime-it-happens). I guess there are various reasons. For example:
#
# * I think it's an interesting application of data science.
# * It connects to popular culture in a big way. Both through TV shows, such as [NUMB3RS](https://en.wikipedia.org/wiki/Numbers_(TV_series)) (it also features in Bones ... or any of the CSI), and also any number of movies, my favorite of which has to be [Minority report](https://www.imdb.com/title/tt0181689/).
# * Predictive policing is also big business. Companies like [PredPol](https://www.predpol.com), [Palantir](https://www.theverge.com/2018/2/27/17054740/palantir-predictive-policing-tool-new-orleans-nopd), and many other companies offer their services law enforcement by analyzing crime data.
# * It hints at the dark sides of Data Science. In these algorithms, concepts like [bias, fairness, and accountability](https://www.smithsonianmag.com/innovation/artificial-intelligence-is-now-used-predict-crime-is-it-biased-180968337/) become incredibly important when the potential outcome of an algorithm is real people going to prison.
# * And, finally there's lots of data available!! Chicago, NYC, and San Francisco all have crime data available freely online.
#
# Below is a little video to pique your interest.
YouTubeVideo("YxvyeaL7NEM",width=800, height=450)
# All this is to say that in the coming weeks, we'll be working to understand crime in San Francisco. We'll be using the SF crime data as a basis for our work on data analysis and data visualization.
#
# We will draw on data from the project [SF OpenData](https://data.sfgov.org), looking into SFPD incidents which have been recorded back since January 2003.
#
# *Reading*
#
# Read [this article](https://www.sciencemag.org/news/2016/09/can-predictive-policing-prevent-crime-it-happens) from science magazine to get a bit deeper sense of the topic.
#
#
# > *Exercise*
# >
# > Answer the following questions in your own words
# >
# > * According to the article, is predictive policing better than best practice techniques for law enforcement? The article is from 2016. Take a look around the web, does this still seem to be the case in 2020? (hint, when you evaluate the evidence consider the source)
# > * List and explain some of the possible issues with predictive policing according to the article.
# # Part 2: Load some crime-data into `pandas`
#
# Go and check out the Python Bootcamp lecture if you don't know what "loading data into Pandas" means. If you're used to using Pandas, then it's finally time to get your hands on some data!!
# > *Exercise*
# >
# > * Go to https://datasf.org/opendata/
# > * Click on "Public Safety"
# > * Download all police incidence reports, historical 2003 to may 2018. You can get everything as a big CSV file if you press the *Export* button (it's a snappy little 456MB file).
# > * Load the data into `pandas` using thie tips and tricks described [here](https://www.shanelynn.ie/python-pandas-read_csv-load-data-from-csv-files/).
# > * Use pandas to generate the following simple statistics
# > - Report the total number of crimes in the dataset
# > - List the various categories of crime
# > - List the number of crimes in each category
# In order to do awesome *predictive policing* later on in the class, we're going to dissect the SF crime-data quite thoroughly to figure out what has been going on over the last years on the San Francisco crime scene.
#
# ---
# > *Exercise*: The types of crime and their popularity over time. The first field we'll dig into is the column "Category".
# > * Use `pandas` to list and then count all the different categories of crime in the dataset. How many are there?
# > * Now count the number of occurrences of each category in the dataset. What is the most commonly occurring category of crime? What is the least frequently occurring?
# > * Create a histogram over crime occurrences. Mine looks like this
# > 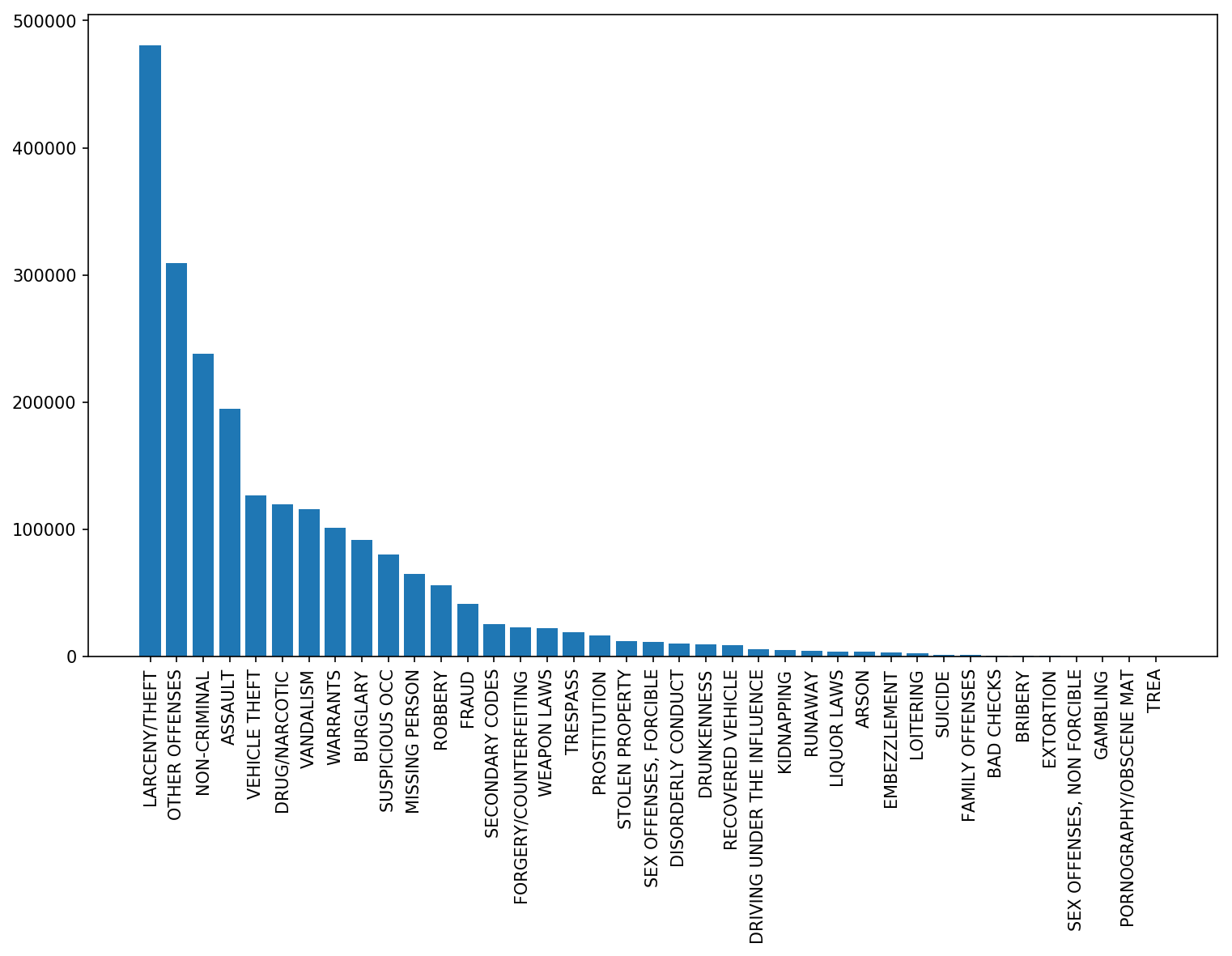
# > * Now it's time to explore how the crime statistics change over time. To start off easily, let's count the number of crimes per year for the years 2003-2017 (since we don't have full data for 2018). What's the average number of crimes per year?
# > * Police chief Suneman is interested in the temporal development of only a subset of categories, the so-called focus crimes. Those categories are listed below (for convenient copy-paste action). Now create bar-charts displaying the year-by-year development of each of these categories across the years 2003-2017.
# >
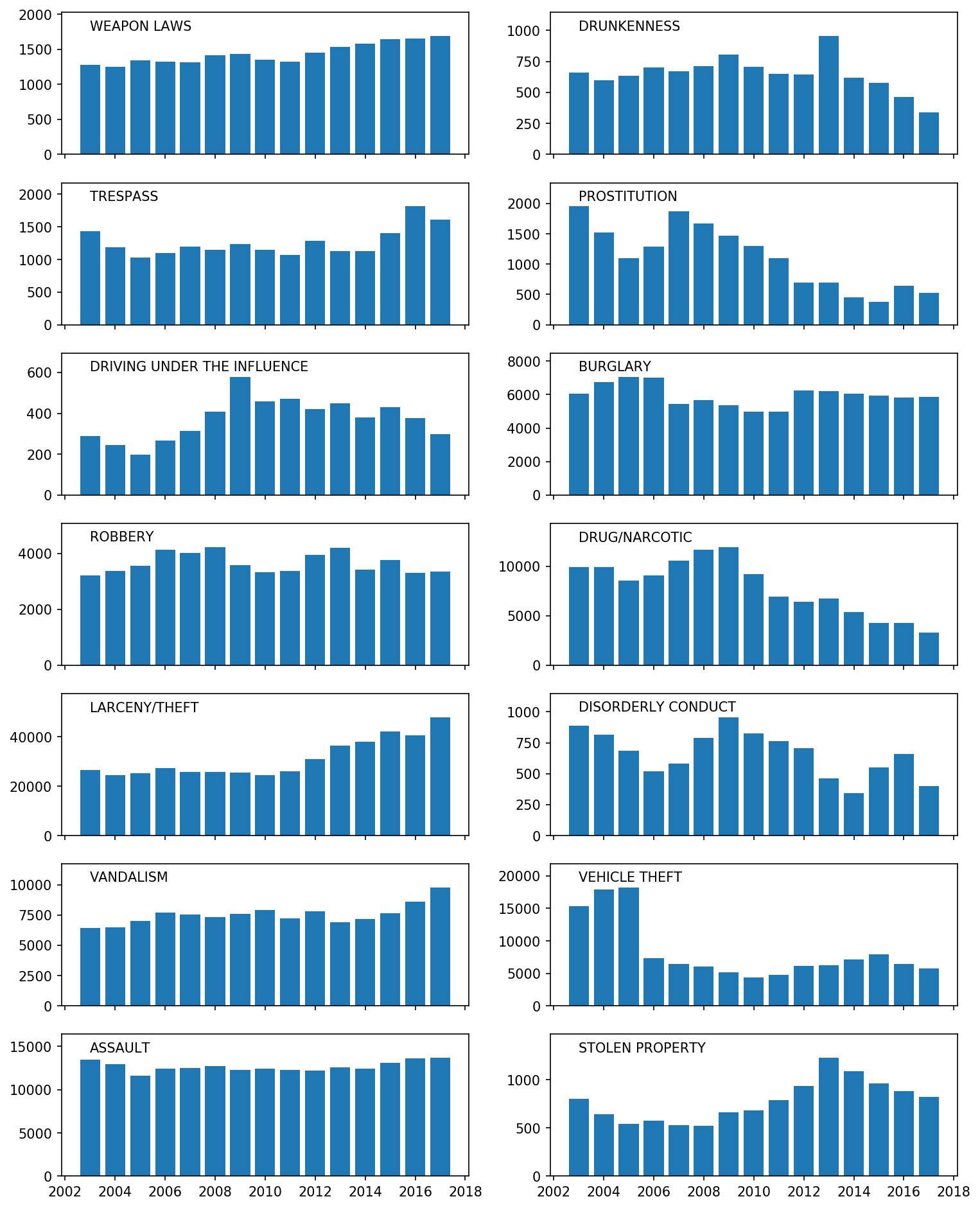
focuscrimes = set(['WEAPON LAWS', 'PROSTITUTION', 'DRIVING UNDER THE INFLUENCE', 'ROBBERY', 'BURGLARY', 'ASSAULT', 'DRUNKENNESS', 'DRUG/NARCOTIC', 'TRESPASS', 'LARCENY/THEFT', 'VANDALISM', 'VEHICLE THEFT', 'STOLEN PROPERTY', 'DISORDERLY CONDUCT'])
# > * My plot looks like this for the 14 focus crimes:
# 
# >
# > (Note that titles are OVER the plots and the axes on the bottom are common for all plots.)
# > * Comment on at least three interesting trends in your plot.
# >
# > Also, here's a fun fact: The drop in car thefts is due to new technology called 'engine immobilizer systems' - get the full story [here](https://www.nytimes.com/2014/08/12/upshot/heres-why-stealing-cars-went-out-of-fashion.html).
|
lectures/Week1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
import datetime
import matplotlib.pyplot as plt
import seaborn as sns
# # 1. EDA
# ## Understanding the data and taking notes for desicions
data = pd.read_csv('regression_data.csv',index_col=None, names= ['id', 'date', 'bedrooms','bathroom','sqft_living', 'sqft_lot','floors', 'waterfront','view', 'condition','grade','sqft_above','sqft_basement', 'yr_built','yr_renovated','zipcode','lat','long','sqft_living15','sqft_lot15','price'])
data.head()
# * drop the zipcode, long & lat columns, as we do not have further information about the location/suburbs
data['yr_renovated'].unique()
data['yr_renovated'].value_counts()
# * Note: lots of unique values, also 0 for not renovated houses --> new column renovated after 1990 True or False
data.dtypes
# * change the date in datetime format
# * make waterfront boolean
data.info()
# * we do not have null values
data.describe()
sns.displot(data['yr_renovated'], bins=150)
data.hist(bins=25,figsize=(15, 15), layout=(5, 4));
plt.show()
|
.ipynb_checkpoints/realestate_Sam-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Prepare fake-factory library for the docker pyspark environment used
# see: https://github.com/jupyter/docker-stacks/tree/master/pyspark-notebook
import os
os.environ['PYSPARK_PYTHON'] = 'python2'
os.environ['PATH'] = os.environ['PATH'].replace('/opt/conda/bin','/opt/conda/envs/python2/bin')
os.environ['CONDA_ENV_PATH'] = '/opt/conda/envs/python2'
# !pip -q install fake-factory
# +
# Prepare fake data to process
from datetime import datetime, timedelta
from faker import Faker
fake = Faker()
import pandas as pd
start = datetime.today()
end = start + timedelta(0.05)
d = [(str(fake.date_time_between_dates(start, end)), fake.ipv4(), fake.free_email_domain()) for x in range(500)]
# +
# Import pyspark
from pyspark import SparkContext
from pyspark.sql import SQLContext
from pyspark.sql import functions as F
# Start a SparkContext, SQLContext
sc = SparkContext('local[*]')
sqlContext = SQLContext(sc)
# Import plotting libs
import seaborn as sns
# %matplotlib inline
# +
# Define the resample function, to perform resampling on timeseries columns
def resample(column, agg_interval=900, time_format='yyyy-MM-dd HH:mm:ss'):
"""
Resamples specified time column of a Spark DataFrame to a specified interval. Interval is given in seconds.lllll
Parameters
----------.
column : Spark Column or string
agg_interval : integer
time_format : string
>>> df = sqlContext.createDataFrame([("2016-01-20 13:32:05",),("2016-01-20 13:50:15",)], ['dt'])
>>> df.select(resample('dt').alias('dt_resampled')).show()
>>> +-------------------+
>>> | dt_resampled|
>>> +-------------------+
>>> |2016-01-20 13:30:00|
>>> |2016-01-20 13:45:00|
>>> +-------------------+
Returns
-------
Spark Column
"""
if type(column)==str:
column = F.col(column)
col_ut = F.unix_timestamp(column, format=time_format) # convert to unix timestamp
col_ut_agg = F.floor(col_ut / agg_interval) * agg_interval # divide the unix timestamp into intervals
col_ts = F.from_unixtime(col_ut_agg) # convert unix timestamp into timestamp
return col_ts
# -
df = sqlContext.createDataFrame(d, ['dt','ip','email_provider'])
df.show(5)
df = df.withColumn('dt_resampled', resample(df.dt, 900))
df.show(5)
df_resampled = df.groupBy('dt_resampled', 'email_provider').count()
df_resampled.show(5)
df_resampled.toPandas() \
.pivot(index='dt_resampled', columns='email_provider', values='count') \
.plot(figsize=[14,5], title='Count emails per 15 minute interval')
|
notebooks/2016-01-20-Spark-resampling.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Key Distribution Kata
#
# The **Quantum Key Distribution** kata is a series of exercises designed to teach you about a neat quantum technology where you can use qubits to exchange secure cryptographic keys. In particular, you will work through implementing and testing a quantum key distribution protocol called [BB84](https://en.wikipedia.org/wiki/BB84).
#
# ### Background
#
# What does a key distribution protocol look like in general? Normally there are two parties, commonly referred to as Alice and Bob, who want to share a random, secret string of bits called a _key_. This key can then be used for a variety of different [cryptographic protocols](https://en.wikipedia.org/wiki/Cryptographic_protocol) like encryption or authentication. Quantum versions of key exchange protocols look very similar, and utilize qubits as a way to securely transmit the bit string.
#
# <img src="./img/qkd-concept.png" alt="General schematic for QKD protocol" width="60%"/>
#
# You can see in the figure above that Alice and Bob have two connections, one quantum channel and one bidirectional classical channel. In this kata you will simulate what happens on the quantum channel by preparing and measuring a sequence of qubits and then perform classical operations to transform the measurement results to a usable, binary key.
#
# There are a variety of different quantum key distribution protocols, however the most common is called [BB84](https://en.wikipedia.org/wiki/BB84) after the initials of the authors and the year it was published. It is used in many existing commercial quantum key distribution devices that implement BB84 with single photons as the qubits.
#
# #### For more information:
# * [Introduction to quantum cryptography and BB84](https://www.youtube.com/watch?v=UiJiXNEm-Go).
# * [QKD summer school lecture on quantum key distribution](https://www.youtube.com/watch?v=oEJOtu0joXk).
# * [Key Distribution Wikipedia article](https://en.wikipedia.org/wiki/Quantum_key_distribution).
# * [BB84 protocol Wikipedia article](https://en.wikipedia.org/wiki/BB84).
# * [Updated version of the BB84 paper](https://www.sciencedirect.com/science/article/pii/S0304397514004241?via%3Dihub).
#
# ---
# ### Instructions
#
# Each task is wrapped in one operation preceded by the description of the task.
#
# Your goal is to fill in the blank (marked with the `// ...` comments)
# with some Q# code that solves the task. To verify your answer, run the cell using Ctrl+Enter (⌘+Enter on macOS).
#
# ## Part I. Preparation
# BB84 protocol loops through the two main steps until Alice and Bob have as much key as they want.
# The first step is to use the quantum channel, where Alice prepares individual qubits and then sends them to Bob to be measured.
# The second step is entirely classical post-processing and communication that takes the measurement results from the quantum step and extracts a classical, random bit string Alice and Bob can use.
# Let's start by looking at how Alice will prepare her qubits for sending to Bob as a part of the quantum phase of the BB84 protocol.
#
# Alice has two choices for each qubit, which basis to prepare it in, and what bit value she wants to encode.
# This leads to four possible states each qubit can be in that Alice sends out.
# The bases she has to choose from are selected such that if an eavesdropper tries to measure a qubit in transit and chooses the wrong basis, then they just get a 0 or 1 with equal probability.
#
# In the first basis, called the computational basis, Alice prepares the states $|0\rangle$ and $|1\rangle$ where $|0\rangle$ represents the key bit value `0` and $|1\rangle$ represents the key bit value `1`.
# The second basis (sometimes called the diagonal or Hadamard basis) uses the states $|+\rangle = \frac{1}{\sqrt2}(|0\rangle + |1\rangle)$ to represent the key bit value `0`, and $|-\rangle = \frac{1}{\sqrt2}(|0\rangle - |1\rangle)$ to represent the key bit value `1`.
#
# ### Task 1.1. Diagonal Basis
#
# Try your hand at converting qubits from the computational basis to the diagonal basis.
#
# **Input:** $N$ qubits (stored in an array of length $N$). Each qubit is either in $|0\rangle$ or in $|1\rangle$ state.
#
# **Goal:** Convert the qubits to the diagonal basis:
# * if `qs[i]` was in state $|0\rangle$, it should be transformed to $|+\rangle = \frac{1}{\sqrt2}(|0\rangle + |1\rangle)$,
# * if `qs[i]` was in state $|1\rangle$, it should be transformed to $|-\rangle = \frac{1}{\sqrt2}(|0\rangle - |1\rangle)$.
# +
%kata T11_DiagonalBasis
operation DiagonalBasis (qs : Qubit[]) : Unit {
// ...
}
# -
# ### Task 1.2. Equal superposition
#
#
# **Input**: A qubit in the $|0\rangle$ state.
#
# **Goal**: Change the qubit state to a superposition state that has equal probabilities of measuring 0 and 1.
#
# > Note that this is not the same as keeping the qubit in the $|0\rangle$ state with 50% probability and converting it to the $|1\rangle$ state with 50% probability!
# +
%kata T12_EqualSuperposition
operation EqualSuperposition (q : Qubit) : Unit {
// ...
}
# -
# ## Part II. BB84 Protocol
# Now that you have seen some of the steps that Alice will use to prepare here qubits, it's time to add in the rest of the steps of the BB84 protocol.
# ### Task 2.1. Generate random array
#
# You saw in part I that Alice has to make two random choices per qubit she prepares, one for which basis to prepare in, and the other for what bit value she wants to send.
# Bob will also need one random bit value to decide what basis he will be measuring each qubit in.
# To make this easier for later steps, you will need a way of generating random boolean values for both Alice and Bob to use.
#
# **Input:** An integer $N$.
#
# **Output** : A `Bool` array of length N, where each element is chosen at random.
#
# > This will be used by both Alice and Bob to choose either the sequence of bits to send or the sequence of bases (`false` indicates $|0\rangle$ / $|1\rangle$ basis, and `true` indicates $|+\rangle$ / $|-\rangle$ basis) to use when encoding/measuring the bits.
# +
%kata T21_RandomArray
operation RandomArray (N : Int) : Bool[] {
// ...
return new Bool[N];
}
# -
# ### Task 2.2. Prepare Alice's qubits
#
# Now that you have a way of generating the random inputs needed for Alice and Bob, it's time for Alice to use the random bits to prepare her sequence of qubits to send to Bob.
#
# **Inputs:**
#
# 1. `qs`: an array of $N$ qubits in the $|0\rangle$ states,
# 2. `bases`: a `Bool` array of length $N$;
# `bases[i]` indicates the basis to prepare the i-th qubit in:
# * `false`: use $|0\rangle$ / $|1\rangle$ (computational) basis,
# * `true`: use $|+\rangle$ / $|-\rangle$ (Hadamard/diagonal) basis.
# 3. `bits`: a `Bool` array of length $N$;
# `bits[i]` indicates the bit to encode in the i-th qubit: `false` = 0, `true` = 1.
#
# **Goal:** Prepare the qubits in the described state.
# +
%kata T22_PrepareAlicesQubits
operation PrepareAlicesQubits (qs : Qubit[], bases : Bool[], bits : Bool[]) : Unit {
// ...
}
# -
# ### Task 2.3. Measure Bob's qubits
#
# Bob now has an incoming stream of qubits that he needs to measure by randomly choosing a basis to measure in for each qubit.
#
# **Inputs:**
#
# 1. `qs`: an array of $N$ qubits;
# each qubit is in one of the following states: $|0\rangle$, $|1\rangle$, $|+\rangle$, $|-\rangle$.
# 2. `bases`: a `Bool` array of length $N$;
# `bases[i]` indicates the basis used to prepare the i-th qubit:
# * `false`: $|0\rangle$ / $|1\rangle$ (computational) basis,
# * `true`: $|+\rangle$ / $|-\rangle$ (Hadamard/diagonal) basis.
#
# **Output:** Measure each qubit in the corresponding basis and return an array of results as boolean values, encoding measurement result `Zero` as `false` and `One` as `true`.
# The state of the qubits at the end of the operation does not matter.
# +
%kata T23_MeasureBobsQubits
operation MeasureBobsQubits (qs : Qubit[], bases : Bool[]) : Bool[] {
// ...
return new Bool[0];
}
# -
# ### Task 2.4. Generate the shared key!
#
# Now, Alice has a list of the bit values she sent as well as what basis she prepared each qubit in, and Bob has a list of bases he used to measure each qubit. To figure out the shared key, they need to figure out when they both used the same basis, and toss the data from qubits where they used different bases. If Alice and Bob did not use the same basis to prepare and measure the qubits in, the measurement results Bob got will be just random with 50% probability for both the `Zero` and `One` outcomes.
#
# **Inputs:**
#
# 1. `basesAlice` and `basesBob`: `Bool` arrays of length $N$
# describing Alice's and Bobs's choice of bases, respectively;
# 2. `measurementsBob`: a `Bool` array of length $N$ describing Bob's measurement results.
#
# **Output:** a `Bool` array representing the shared key generated by the protocol.
#
# > Note that you don't need to know both Alice's and Bob's bits to figure out the shared key!
# +
%kata T24_GenerateSharedKey
function GenerateSharedKey (basesAlice : Bool[], basesBob : Bool[], measurementsBob : Bool[]) : Bool[] {
// ...
return new Bool[0];
}
# -
# ### Task 2.5. Check if error rate was low enough
#
# The main trace eavesdroppers can leave on a key exchange is to introduce more errors into the transmission. Alice and Bob should have characterized the error rate of their channel before launching the protocol, and need to make sure when exchanging the key that there were not more errors than they expected. The "errorRate" parameter represents their earlier characterization of their channel.
#
# **Inputs:**
#
# 1. `keyAlice` and `keyBob`: `Bool` arrays of equal length $N$ describing
# the versions of the shared key obtained by Alice and Bob, respectively.
# 2. `errorRate`: an integer between 0 and 50 - the percentage of the bits that did not match in Alice's and Bob's channel characterization.
#
# **Output:** `true` if the percentage of errors is less than or equal to the error rate, and `false` otherwise.
# +
%kata T25_CheckKeysMatch
function CheckKeysMatch (keyAlice : Bool[], keyBob : Bool[], errorRate : Int) : Bool {
// ...
return false;
}
# -
# ### Task 2.6. Putting it all together
#
# **Goal:** Implement the entire BB84 protocol using tasks 2.1 - 2.5
# and following the comments in the operation template.
#
# > This is an open-ended task, and is not covered by a unit test. To run the code, execute the cell with the definition of the `Run_BB84Protocol` operation first; if it compiled successfully without any errors, you can run the operation by executing the next cell (`%simulate Run_BB84Protocol`).
operation Run_BB84Protocol () : Unit {
// 1. Alice chooses a random set of bits to encode in her qubits
// and a random set of bases to prepare her qubits in.
// ...
// 2. Alice allocates qubits, encodes them using her choices and sends them to Bob.
// (Note that you can not reflect "sending the qubits to Bob" in Q#)
// ...
// 3. Bob chooses a random set of bases to measure Alice's qubits in.
// ...
// 4. Bob measures Alice's qubits in his chosen bases.
// ...
// 5. Alice and Bob compare their chosen bases and use the bits in the matching positions to create a shared key.
// ...
// 6. Alice and Bob check to make sure nobody eavesdropped by comparing a subset of their keys
// and verifying that more than a certain percentage of the bits match.
// For this step, you can check the percentage of matching bits using the entire key
// (in practice only a subset of indices is chosen to minimize the number of discarded bits).
// ...
// If you've done everything correctly, the generated keys will always match, since there is no eavesdropping going on.
// In the next section you will explore the effects introduced by eavesdropping.
}
%simulate Run_BB84Protocol
# ## Part III. Eavesdropping
# ### Task 3.1. Eavesdrop!
#
# In this task you will try to implement an eavesdropper, Eve.
#
# Eve will intercept a qubit from the quantum channel that Alice and Bob are using.
# She will measure it in either the $|0\rangle$ / $|1\rangle$ basis or the $|+\rangle$ / $|-\rangle$ basis, and prepare a new qubit in the state she measured. Then she will send the new qubit back to the channel.
# Eve hopes that if she got lucky with her measurement, that when Bob measures the qubit he doesn't get an error so she won't be caught!
#
# **Inputs:**
#
# 1. `q`: a qubit in one of the following states: $|0\rangle$, $|1\rangle$, $|+\rangle$, $|-\rangle$.
# 2. `basis`: Eve's guess of the basis she should use for measuring.
# Recall that `false` indicates $|0\rangle$ / $|1\rangle$ basis and `true` indicates $|+\rangle$ / $|-\rangle$ basis.
#
# **Output:** the bit encoded in the qubit (`false` for $|0\rangle$ / $|+\rangle$ states, `true` for $|1\rangle$ / $|-\rangle$ states).
#
# > In this task you are guaranteed that the basis you're given matches the one in which the qubit is encoded, that is, if you are given a qubit in state $|0\rangle$ or $|1\rangle$, you will be given `basis = false`, and if you are given a qubit in state $|+\rangle$ or $|-\rangle$, you will be given `basis = true`. This is different from a real eavesdropping scenario, in which you have to guess the basis yourself.
# +
%kata T31_Eavesdrop
operation Eavesdrop (q : Qubit, basis : Bool) : Bool {
// ...
return false;
}
# -
# ### Task 3.2. Catch the eavesdropper
#
# Add an eavesdropper into the BB84 protocol from task 2.6.
#
# Note that now we should be able to detect Eve and therefore we have to discard some of our key bits!
#
# > Similar to task 2.6, this is an open-ended task, and is not covered by a unit test. To run the code, execute the cell with the definition of the `Run_BB84ProtocolWithEavesdropper` operation first; if it compiled successfully without any errors, you can run the operation by executing the next cell (`%simulate Run_BB84ProtocolWithEavesdropper`).
operation Run_BB84ProtocolWithEavesdropper () : Unit {
// ...
}
%simulate Run_BB84ProtocolWithEavesdropper
|
KeyDistribution_BB84/KeyDistribution_BB84.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math as m
import numpy as np
from numpy import genfromtxt
import os
from tqdm import tqdm
num_kp = 22
num_axes = 3
# List all the files in dir
kp_path = "Movements/Kinect/Positions/"
ka_path = "Movements/Kinect/Angles/"
kinect_positions = sorted([kp_path+f for f in os.listdir(kp_path) if f.endswith('.txt')])
kinect_angles = sorted([ka_path+f for f in os.listdir(ka_path) if f.endswith('.txt')])
# +
def Rotx(theta):
return np.matrix([[ 1, 0 , 0 ],
[ 0, m.cos(theta),-m.sin(theta)],
[ 0, m.sin(theta), m.cos(theta)]])
def Roty(theta):
return np.matrix([[ m.cos(theta), 0, m.sin(theta)],
[ 0 , 1, 0 ],
[-m.sin(theta), 0, m.cos(theta)]])
def Rotz(theta):
return np.matrix([[ m.cos(theta), -m.sin(theta), 0 ],
[ m.sin(theta), m.cos(theta) , 0 ],
[ 0 , 0 , 1 ]])
def eulers_2_rot_matrix(x):
gamma_x=x[0];beta_y=x[1];alpha_z=x[2];
return Rotz(alpha_z)*Roty(beta_y)*Rotx(gamma_x)
# -
# convert the data from relative coordinates to absolute coordinates
def rel2abs(p, a, num_frames):
skel = np.zeros((num_kp, num_axes, num_frames))
for i in range(num_frames):
"""
1 Waist (absolute)
2 Spine
3 Chest
4 Neck
5 Head
6 Head tip
7 Left collar
8 Left upper arm
9 Left forearm
10 Left hand
11 Right collar
12 Right upper arm
13 Right forearm
14 Right hand
15 Left upper leg
16 Left lower leg
17 Left foot
18 Left leg toes
19 Right upper leg
20 Right lower leg
21 Right foot
22 Right leg toes
"""
joint = p[:,:,i]
joint_ang = a[:,:,i]
# chest, neck, head
rot_1 = eulers_2_rot_matrix(joint_ang[0,:]*np.pi/180);
joint[1,:] = rot_1@joint[1,:] + joint[0,:]
rot_2 = rot_1*eulers_2_rot_matrix(joint_ang[1,:]*np.pi/180)
joint[2,:] = rot_2@joint[2,:] + joint[1,:]
rot_3 = rot_2*eulers_2_rot_matrix(joint_ang[2,:]*np.pi/180)
joint[3,:] = rot_3@joint[3,:] + joint[2,:]
rot_4 = rot_3*eulers_2_rot_matrix(joint_ang[3,:]*np.pi/180)
joint[4,:] = rot_4@joint[4,:] + joint[3,:]
rot_5 = rot_4*eulers_2_rot_matrix(joint_ang[4,:]*np.pi/180)
joint[5,:] = rot_5@joint[5,:] + joint[4,:]
# left-arm
rot_6 = eulers_2_rot_matrix(joint_ang[2,:]*np.pi/180)
joint[6,:] = rot_6@joint[6,:] + joint[2,:]
rot_7 = rot_6*eulers_2_rot_matrix(joint_ang[6,:]*np.pi/180)
joint[7,:] = rot_7@joint[7,:] + joint[6,:]
rot_8 = rot_7*eulers_2_rot_matrix(joint_ang[7,:]*np.pi/180)
joint[8,:] = rot_8@joint[8,:] + joint[7,:]
rot_9 = rot_8*eulers_2_rot_matrix(joint_ang[8,:]*np.pi/180)
joint[9,:] = rot_9@joint[9,:] + joint[8,:]
# right-arm
rot_10 = eulers_2_rot_matrix(joint_ang[2,:]*np.pi/180)
joint[10,:] = rot_10@joint[10,:] + joint[2,:]
rot_11 = rot_10*eulers_2_rot_matrix(joint_ang[10,:]*np.pi/180)
joint[11,:] = rot_11@joint[11,:] + joint[10,:]
rot_12 = rot_11*eulers_2_rot_matrix(joint_ang[11,:]*np.pi/180)
joint[12,:] = rot_12@joint[12,:] + joint[11,:]
rot_13 = rot_12*eulers_2_rot_matrix(joint_ang[12,:]*np.pi/180)
joint[13,:] = rot_13@joint[13,:] + joint[12,:]
# left-leg
rot_14 = eulers_2_rot_matrix(joint_ang[0,:]*np.pi/180)
joint[14,:] = rot_14@joint[14,:] + joint[0,:]
rot_15 = rot_14*eulers_2_rot_matrix(joint_ang[14,:]*np.pi/180)
joint[15,:] = rot_15@joint[15,:] + joint[14,:]
rot_16 = rot_15*eulers_2_rot_matrix(joint_ang[15,:]*np.pi/180)
joint[16,:] = rot_16@joint[16,:] + joint[15,:]
rot_17 = rot_16*eulers_2_rot_matrix(joint_ang[16,:]*np.pi/180)
joint[17,:] = rot_17@joint[17,:] + joint[16,:]
# right-leg
rot_18 = eulers_2_rot_matrix(joint_ang[0,:]*np.pi/180)
joint[18,:] = rot_18@joint[18,:] + joint[0,:]
rot_19 = rot_18*eulers_2_rot_matrix(joint_ang[18,:]*np.pi/180)
joint[19,:] = rot_19@joint[19,:] + joint[18,:]
rot_20 = rot_19*eulers_2_rot_matrix(joint_ang[19,:]*np.pi/180)
joint[20,:] = rot_20@joint[20,:] + joint[19,:]
rot_21 = rot_20*eulers_2_rot_matrix(joint_ang[20,:]*np.pi/180)
joint[21,:] = rot_21@joint[21,:] + joint[20,:]
skel[:,:,i] = joint
return skel.transpose(2,0,1)
def get_output_name(path):
pstr = "_".join(path.split("/")[-1].split(".")[0].split("_")[:-1])
return f"{pstr}_keypoints.csv"
def main():
# List all the files in dir
kp_path = "Movements/Kinect/Positions/"
ka_path = "Movements/Kinect/Angles/"
kinect_positions = sorted([kp_path+f for f in os.listdir(kp_path) if f.endswith('.txt')])
kinect_angles = sorted([ka_path+f for f in os.listdir(ka_path) if f.endswith('.txt')])
assert len(kinect_positions) == len(kinect_angles)
N = len(kinect_positions)
for i in tqdm(range(N)):
# get a position/Angle file path
pos_path = kinect_positions[i]
ang_path = kinect_angles[i]
# get the output name
op_name = get_output_name(pos_path)
# create the directory you want to save to
op_dir = "Movements/dataset_kp_kinect/"
if not os.path.isdir(op_dir):
os.makedirs(op_dir)
# read the text files
pos_data = genfromtxt(pos_path)
ang_data = genfromtxt(ang_path)
assert pos_data.shape[0] == ang_data.shape[0]
num_frames = pos_data.shape[0]
p_data = pos_data.T.reshape(num_kp, num_axes, -1)
a_data = ang_data.T.reshape(num_kp, num_axes, -1)
# tranform relative coordinates to absolute
skel = rel2abs(p_data, a_data, num_frames)
x = skel.reshape(num_frames, -1)
# save array as ".csv"
np.savetxt(os.path.join(op_dir, op_name), x, delimiter=",")
main()
|
UI-PRMD-Transformed.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.9 64-bit
# name: python3
# ---
# # Authoring Hub Predictors
# NatML provides infrastructure that allows businesses to very quickly deploy machine learning in their products. NatML handles all infrastructure needs, including server provisioning, data transfers, and auto-scaling. As a result, all developers need to do is simply bring their model, along with lightweight code for making predictions server- and client-side. There are two components involved:
#
# 1. **Executor**: The `MLExecutor` is code that is responsible for running the user's ML models on the NatML Hub cloud infrastructure. The executor is responsible for instantiating an inference runtime (like ONNX Runtime or PyTorch) and running inference on features provided by the NatML framework from end-users.
#
# 2. **Predictor**: The predictor is a lightweight component that end-users use to request predictions. The predictor submits prediction workloads to the NatML Hub platform, and post-processes the results client-side into a form easily usable by the end-user.
# ## Writing the Executor
# NatML provides the `MLExecutor` class which all executors must derive from. We will write an executor for the MobileNet v2 model:
# +
from natml import MLExecutor, MLModelData, MLFeature
from onnxruntime import InferenceSession
from typing import List
class MobileNetv2Executor (MLExecutor):
def initialize (self, model_data: MLModelData, graph_path: str):
"""
Initialize the executor.
This method should instantiate an ML inference session using an
inference framework like ONNX Runtime, PyTorch, or TensorFlow.
Parameters:
model_data (MLModelData): Model data.
graph_path (str): Path to ML graph on the file system.
"""
self.__session = InferenceSession(graph_path)
self.__model_data = model_data
def predict (self, *inputs: List[MLFeature]) -> List[MLFeature]:
"""
Make a prediction on one or more input features.
Parameters:
inputs (list): Input features.
Returns:
list: Output features.
"""
pass
# -
# ## Writing the Predictor
# +
from natml import MLModelData, MLModel, MLFeature
from typing import List
class MobileNetv2HubPredictor:
def __init__ (self, model: MLModel):
self.__model = model
def predict (self, *inputs: List[MLFeature]):
pass
# -
# Fetch the model data from Hub
access_key = "<HUB ACCESS KEY>"
model_data = MLModelData.from_hub("@natsuite/mobilenet-v2", access_key)
# Deserialize the model
model = model_data.deserialize()
# Create the predictor
predictor = MobileNetv2HubPredictor(model)
|
examples/authoring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # softmax回归的从零开始实现
# :label:`sec_softmax_scratch`
#
# (**就像我们从零开始实现线性回归一样,**)
# 我们认为softmax回归也是重要的基础,因此(**你应该知道实现softmax回归的细节**)。
# 本节我们将使用刚刚在 :numref:`sec_fashion_mnist`中引入的Fashion-MNIST数据集,
# 并设置数据迭代器的批量大小为256。
#
# + origin_pos=3 tab=["tensorflow"]
import tensorflow as tf
from IPython import display
from d2l import tensorflow as d2l
# + origin_pos=4 tab=["tensorflow"]
batch_size = 256
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
# + [markdown] origin_pos=5
# ## 初始化模型参数
#
# 和之前线性回归的例子一样,这里的每个样本都将用固定长度的向量表示。
# 原始数据集中的每个样本都是$28 \times 28$的图像。
# 在本节中,我们[**将展平每个图像,把它们看作长度为784的向量。**]
# 在后面的章节中,我们将讨论能够利用图像空间结构的特征,
# 但现在我们暂时只把每个像素位置看作一个特征。
#
# 回想一下,在softmax回归中,我们的输出与类别一样多。
# (**因为我们的数据集有10个类别,所以网络输出维度为10**)。
# 因此,权重将构成一个$784 \times 10$的矩阵,
# 偏置将构成一个$1 \times 10$的行向量。
# 与线性回归一样,我们将使用正态分布初始化我们的权重`W`,偏置初始化为0。
#
# + origin_pos=8 tab=["tensorflow"]
num_inputs = 784
num_outputs = 10
W = tf.Variable(tf.random.normal(shape=(num_inputs, num_outputs),
mean=0, stddev=0.01))
b = tf.Variable(tf.zeros(num_outputs))
# + [markdown] origin_pos=9
# ## 定义softmax操作
#
# 在实现softmax回归模型之前,我们简要回顾一下`sum`运算符如何沿着张量中的特定维度工作。
# 如 :numref:`subseq_lin-alg-reduction`和
# :numref:`subseq_lin-alg-non-reduction`所述,
# [**给定一个矩阵`X`,我们可以对所有元素求和**](默认情况下)。
# 也可以只求同一个轴上的元素,即同一列(轴0)或同一行(轴1)。
# 如果`X`是一个形状为`(2, 3)`的张量,我们对列进行求和,
# 则结果将是一个具有形状`(3,)`的向量。
# 当调用`sum`运算符时,我们可以指定保持在原始张量的轴数,而不折叠求和的维度。
# 这将产生一个具有形状`(1, 3)`的二维张量。
#
# + origin_pos=11 tab=["tensorflow"]
X = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
tf.reduce_sum(X, 0, keepdims=True), tf.reduce_sum(X, 1, keepdims=True)
# + [markdown] origin_pos=12
# 回想一下,[**实现softmax**]由三个步骤组成:
#
# 1. 对每个项求幂(使用`exp`);
# 1. 对每一行求和(小批量中每个样本是一行),得到每个样本的规范化常数;
# 1. 将每一行除以其规范化常数,确保结果的和为1。
#
# 在查看代码之前,我们回顾一下这个表达式:
#
# (**
# $$
# \mathrm{softmax}(\mathbf{X})_{ij} = \frac{\exp(\mathbf{X}_{ij})}{\sum_k \exp(\mathbf{X}_{ik})}.
# $$
# **)
#
# 分母或规范化常数,有时也称为*配分函数*(其对数称为对数-配分函数)。
# 该名称来自[统计物理学](https://en.wikipedia.org/wiki/Partition_function_(statistical_mechanics))中一个模拟粒子群分布的方程。
#
# + origin_pos=13 tab=["tensorflow"]
def softmax(X):
X_exp = tf.exp(X)
partition = tf.reduce_sum(X_exp, 1, keepdims=True)
return X_exp / partition # 这里应用了广播机制
# + [markdown] origin_pos=15
# 正如你所看到的,对于任何随机输入,[**我们将每个元素变成一个非负数。
# 此外,依据概率原理,每行总和为1**]。
#
# + origin_pos=17 tab=["tensorflow"]
X = tf.random.normal((2, 5), 0, 1)
X_prob = softmax(X)
X_prob, tf.reduce_sum(X_prob, 1)
# + [markdown] origin_pos=18
# 注意,虽然这在数学上看起来是正确的,但我们在代码实现中有点草率。
# 矩阵中的非常大或非常小的元素可能造成数值上溢或下溢,但我们没有采取措施来防止这点。
#
# ## 定义模型
#
# 定义softmax操作后,我们可以[**实现softmax回归模型**]。
# 下面的代码定义了输入如何通过网络映射到输出。
# 注意,将数据传递到模型之前,我们使用`reshape`函数将每张原始图像展平为向量。
#
# + origin_pos=19 tab=["tensorflow"]
def net(X):
return softmax(tf.matmul(tf.reshape(X, (-1, W.shape[0])), W) + b)
# + [markdown] origin_pos=20
# ## 定义损失函数
#
# 接下来,我们实现 :numref:`sec_softmax`中引入的交叉熵损失函数。
# 这可能是深度学习中最常见的损失函数,因为目前分类问题的数量远远超过回归问题的数量。
#
# 回顾一下,交叉熵采用真实标签的预测概率的负对数似然。
# 这里我们不使用Python的for循环迭代预测(这往往是低效的),
# 而是通过一个运算符选择所有元素。
# 下面,我们[**创建一个数据样本`y_hat`,其中包含2个样本在3个类别的预测概率,
# 以及它们对应的标签`y`。**]
# 有了`y`,我们知道在第一个样本中,第一类是正确的预测;
# 而在第二个样本中,第三类是正确的预测。
# 然后(**使用`y`作为`y_hat`中概率的索引**),
# 我们选择第一个样本中第一个类的概率和第二个样本中第三个类的概率。
#
# + origin_pos=22 tab=["tensorflow"]
y_hat = tf.constant([[0.1, 0.3, 0.6], [0.3, 0.2, 0.5]])
y = tf.constant([0, 2])
tf.boolean_mask(y_hat, tf.one_hot(y, depth=y_hat.shape[-1]))
# + [markdown] origin_pos=23
# 现在我们只需一行代码就可以[**实现交叉熵损失函数**]。
#
# + origin_pos=25 tab=["tensorflow"]
def cross_entropy(y_hat, y):
return -tf.math.log(tf.boolean_mask(
y_hat, tf.one_hot(y, depth=y_hat.shape[-1])))
cross_entropy(y_hat, y)
# + [markdown] origin_pos=26
# ## 分类精度
#
# 给定预测概率分布`y_hat`,当我们必须输出硬预测(hard prediction)时,
# 我们通常选择预测概率最高的类。
# 许多应用都要求我们做出选择。如Gmail必须将电子邮件分类为“Primary(主要邮件)”、
# “Social(社交邮件)”、“Updates(更新邮件)”或“Forums(论坛邮件)”。
# Gmail做分类时可能在内部估计概率,但最终它必须在类中选择一个。
#
# 当预测与标签分类`y`一致时,即是正确的。
# 分类精度即正确预测数量与总预测数量之比。
# 虽然直接优化精度可能很困难(因为精度的计算不可导),
# 但精度通常是我们最关心的性能衡量标准,我们在训练分类器时几乎总会关注它。
#
# 为了计算精度,我们执行以下操作。
# 首先,如果`y_hat`是矩阵,那么假定第二个维度存储每个类的预测分数。
# 我们使用`argmax`获得每行中最大元素的索引来获得预测类别。
# 然后我们[**将预测类别与真实`y`元素进行比较**]。
# 由于等式运算符“`==`”对数据类型很敏感,
# 因此我们将`y_hat`的数据类型转换为与`y`的数据类型一致。
# 结果是一个包含0(错)和1(对)的张量。
# 最后,我们求和会得到正确预测的数量。
#
# + origin_pos=27 tab=["tensorflow"]
def accuracy(y_hat, y): #@save
"""计算预测正确的数量"""
if len(y_hat.shape) > 1 and y_hat.shape[1] > 1:
y_hat = tf.argmax(y_hat, axis=1)
cmp = tf.cast(y_hat, y.dtype) == y
return float(tf.reduce_sum(tf.cast(cmp, y.dtype)))
# + [markdown] origin_pos=28
# 我们将继续使用之前定义的变量`y_hat`和`y`分别作为预测的概率分布和标签。
# 可以看到,第一个样本的预测类别是2(该行的最大元素为0.6,索引为2),这与实际标签0不一致。
# 第二个样本的预测类别是2(该行的最大元素为0.5,索引为2),这与实际标签2一致。
# 因此,这两个样本的分类精度率为0.5。
#
# + origin_pos=29 tab=["tensorflow"]
accuracy(y_hat, y) / len(y)
# + [markdown] origin_pos=30
# 同样,对于任意数据迭代器`data_iter`可访问的数据集,
# [**我们可以评估在任意模型`net`的精度**]。
#
# + origin_pos=31 tab=["tensorflow"]
def evaluate_accuracy(net, data_iter): #@save
"""计算在指定数据集上模型的精度"""
metric = Accumulator(2) # 正确预测数、预测总数
for X, y in data_iter:
metric.add(accuracy(net(X), y), d2l.size(y))
return metric[0] / metric[1]
# + [markdown] origin_pos=33
# 这里定义一个实用程序类`Accumulator`,用于对多个变量进行累加。
# 在上面的`evaluate_accuracy`函数中,
# 我们在(**`Accumulator`实例中创建了2个变量,
# 分别用于存储正确预测的数量和预测的总数量**)。
# 当我们遍历数据集时,两者都将随着时间的推移而累加。
#
# + origin_pos=34 tab=["tensorflow"]
class Accumulator: #@save
"""在n个变量上累加"""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
# + [markdown] origin_pos=35
# 由于我们使用随机权重初始化`net`模型,
# 因此该模型的精度应接近于随机猜测。
# 例如在有10个类别情况下的精度为0.1。
#
# + origin_pos=36 tab=["tensorflow"]
evaluate_accuracy(net, test_iter)
# + [markdown] origin_pos=37
# ## 训练
#
# 如果你看过 :numref:`sec_linear_scratch`中的线性回归实现,
# [**softmax回归的训练**]过程代码应该看起来非常眼熟。
# 在这里,我们重构训练过程的实现以使其可重复使用。
# 首先,我们定义一个函数来训练一个迭代周期。
# 请注意,`updater`是更新模型参数的常用函数,它接受批量大小作为参数。
# 它可以是`d2l.sgd`函数,也可以是框架的内置优化函数。
#
# + origin_pos=40 tab=["tensorflow"]
def train_epoch_ch3(net, train_iter, loss, updater): #@save
"""训练模型一个迭代周期(定义见第3章)"""
# 训练损失总和、训练准确度总和、样本数
metric = Accumulator(3)
for X, y in train_iter:
# 计算梯度并更新参数
with tf.GradientTape() as tape:
y_hat = net(X)
# Keras内置的损失接受的是(标签,预测),这不同于用户在本书中的实现。
# 本书的实现接受(预测,标签),例如我们上面实现的“交叉熵”
if isinstance(loss, tf.keras.losses.Loss):
l = loss(y, y_hat)
else:
l = loss(y_hat, y)
if isinstance(updater, tf.keras.optimizers.Optimizer):
params = net.trainable_variables
grads = tape.gradient(l, params)
updater.apply_gradients(zip(grads, params))
else:
updater(X.shape[0], tape.gradient(l, updater.params))
# Keras的loss默认返回一个批量的平均损失
l_sum = l * float(tf.size(y)) if isinstance(
loss, tf.keras.losses.Loss) else tf.reduce_sum(l)
metric.add(l_sum, accuracy(y_hat, y), tf.size(y))
# 返回训练损失和训练精度
return metric[0] / metric[2], metric[1] / metric[2]
# + [markdown] origin_pos=41
# 在展示训练函数的实现之前,我们[**定义一个在动画中绘制数据的实用程序类**]`Animator`,
# 它能够简化本书其余部分的代码。
#
# + origin_pos=42 tab=["tensorflow"]
class Animator: #@save
"""在动画中绘制数据"""
def __init__(self, xlabel=None, ylabel=None, legend=None, xlim=None,
ylim=None, xscale='linear', yscale='linear',
fmts=('-', 'm--', 'g-.', 'r:'), nrows=1, ncols=1,
figsize=(3.5, 2.5)):
# 增量地绘制多条线
if legend is None:
legend = []
d2l.use_svg_display()
self.fig, self.axes = d2l.plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [self.axes, ]
# 使用lambda函数捕获参数
self.config_axes = lambda: d2l.set_axes(
self.axes[0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# 向图表中添加多个数据点
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
# + [markdown] origin_pos=43
# 接下来我们实现一个[**训练函数**],
# 它会在`train_iter`访问到的训练数据集上训练一个模型`net`。
# 该训练函数将会运行多个迭代周期(由`num_epochs`指定)。
# 在每个迭代周期结束时,利用`test_iter`访问到的测试数据集对模型进行评估。
# 我们将利用`Animator`类来可视化训练进度。
#
# + origin_pos=44 tab=["tensorflow"]
def train_ch3(net, train_iter, test_iter, loss, num_epochs, updater): #@save
"""训练模型(定义见第3章)"""
animator = Animator(xlabel='epoch', xlim=[1, num_epochs], ylim=[0.3, 0.9],
legend=['train loss', 'train acc', 'test acc'])
for epoch in range(num_epochs):
train_metrics = train_epoch_ch3(net, train_iter, loss, updater)
test_acc = evaluate_accuracy(net, test_iter)
animator.add(epoch + 1, train_metrics + (test_acc,))
train_loss, train_acc = train_metrics
assert train_loss < 0.5, train_loss
assert train_acc <= 1 and train_acc > 0.7, train_acc
assert test_acc <= 1 and test_acc > 0.7, test_acc
# + [markdown] origin_pos=45
# 作为一个从零开始的实现,我们使用 :numref:`sec_linear_scratch`中定义的
# [**小批量随机梯度下降来优化模型的损失函数**],设置学习率为0.1。
#
# + origin_pos=47 tab=["tensorflow"]
class Updater(): #@save
"""用小批量随机梯度下降法更新参数"""
def __init__(self, params, lr):
self.params = params
self.lr = lr
def __call__(self, batch_size, grads):
d2l.sgd(self.params, grads, self.lr, batch_size)
updater = Updater([W, b], lr=0.1)
# + [markdown] origin_pos=48
# 现在,我们[**训练模型10个迭代周期**]。
# 请注意,迭代周期(`num_epochs`)和学习率(`lr`)都是可调节的超参数。
# 通过更改它们的值,我们可以提高模型的分类精度。
#
# + origin_pos=49 tab=["tensorflow"]
num_epochs = 10
train_ch3(net, train_iter, test_iter, cross_entropy, num_epochs, updater)
# + [markdown] origin_pos=50
# ## 预测
#
# 现在训练已经完成,我们的模型已经准备好[**对图像进行分类预测**]。
# 给定一系列图像,我们将比较它们的实际标签(文本输出的第一行)和模型预测(文本输出的第二行)。
#
# + origin_pos=51 tab=["tensorflow"]
def predict_ch3(net, test_iter, n=6): #@save
"""预测标签(定义见第3章)"""
for X, y in test_iter:
break
trues = d2l.get_fashion_mnist_labels(y)
preds = d2l.get_fashion_mnist_labels(tf.argmax(net(X), axis=1))
titles = [true +'\n' + pred for true, pred in zip(trues, preds)]
d2l.show_images(
tf.reshape(X[0:n], (n, 28, 28)), 1, n, titles=titles[0:n])
predict_ch3(net, test_iter)
# + [markdown] origin_pos=52
# ## 小结
#
# * 借助softmax回归,我们可以训练多分类的模型。
# * 训练softmax回归循环模型与训练线性回归模型非常相似:先读取数据,再定义模型和损失函数,然后使用优化算法训练模型。大多数常见的深度学习模型都有类似的训练过程。
#
# ## 练习
#
# 1. 在本节中,我们直接实现了基于数学定义softmax运算的`softmax`函数。这可能会导致什么问题?提示:尝试计算$\exp(50)$的大小。
# 1. 本节中的函数`cross_entropy`是根据交叉熵损失函数的定义实现的。它可能有什么问题?提示:考虑对数的定义域。
# 1. 你可以想到什么解决方案来解决上述两个问题?
# 1. 返回概率最大的分类标签总是最优解吗?例如,医疗诊断场景下你会这样做吗?
# 1. 假设我们使用softmax回归来预测下一个单词,可选取的单词数目过多可能会带来哪些问题?
#
# + [markdown] origin_pos=55 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1790)
#
|
submodules/resource/d2l-zh/tensorflow/chapter_linear-networks/softmax-regression-scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (qiime2-2018.4)
# language: python
# name: qiime2-2018.4
# ---
# # Modify genbank parser to add functionality, convert to Python 3
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append('/Users/hughcross/Analysis/repos/genbanking/gb_parsers/')
from genbank_parsers import parse_genbank_features, parse_gb_feature_dict
example1 = parse_genbank_features('/Users/hughcross/Analysis/repos/genbanking/genbank_example.gb')
annot1 = example1[0]
annot1['Blitum_asiaticum:GQ245137']
feats1 = example1[1]
feats1['Blitum_asiaticum:GQ245137']
feats1a = parse_gb_feature_dict(feats1)
feats1a[0]['Blitum_asiaticum:GQ245137']
feats1['Blitum_asiaticum:GQ245137']
feats1#['Blitum_asiaticum:GQ245137'].source
example2 = parse_genbank_features('/Users/hughcross/Documents/Rattus_exulans_isolate_RCIPuk007.gb')
feats2 = parse_gb_feature_dict(example2[1])
feats2
example2[0]
|
notebooks/adding_functionality_to_parser.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# plotting libraries
import matplotlib
import matplotlib.pyplot as plt
# numpy (math) libary
import numpy as np
# Constants
n0 = 3.48 # standard refractive index
n2 = 5e-14 # [cm²/W] intensity-dependent refractive index
e0 = 8.85418782e-12 # vacuum permittivity epsilon_0
c0 = 299792458 # speed of light in vacuum c_0
# Parameters
t = 0.1 # transmission factor, ideally 0.0
B = 0.95 # half-ring loss factor, ideally 1.0. B=0.95 is obtained for a R=10e-6 with losses of 10dB/µm
R = 10e-6 # radius of curvature
s = (1-t**2)**0.5
# +
# intensity of beam, given field amplitude
def Iw(Ew):
return 0.5*e0*n0*c0*Ew**2
# total refractive index
# ---> CHANGE this with propagation index of waveguide,
# aka the effective refractive index of the waveguide/ring n_eff
def n(freq, Ew):
return n0+n2*Iw(Ew)+0*freq
# define transmission factor and half-ring loss factor as
# dependent of frequency and field amplitude
def tf(f, Ew):
return t
def Bf(f, Ew):
return B
# +
# argument of the complex exponential
def psi(freq, Ew):
return ( 2*np.pi*R ) * ( 2*np.pi/c0*freq )* n(freq, Ew)
# transmittivity
def tau(f, Ew):
return -(1-t**2) * B * np.exp( 1J * psi(f, Ew) / 2 ) / (1 - t**2 * B * np.exp( 1J * psi(f, Ew) ) )
def normE(f, Ew):
print("asd")
def phase(f, Ew):
return np.angle(tau(f, Ew))
# +
# frequency study
x = np.linspace((1-3*0.006451613)*200e12,(1+3*0.006451613)*200e12,200)
# magnitude
y = np.abs(tau(x,0))
plt.plot(x, y, 'rx')
plt.show()
# phase
y2 = np.angle(tau(x,0))
plt.plot(x, y2, 'b+')
plt.show()
# +
IFA = 1.0e7 # input field amplitude
IFI = np.square(IFA) # input field intensity
A = [np.abs(tau(200e12,0.0)), ] # normalized output intensity
P = [np.angle(tau(200e12,0.0)), ] # output phase
D = [0.0, ] # un-normalized intensity inside the µ-ring resonator
rounds = 14
for ii in range(0, rounds):
A.append( np.abs(tau(200e12,D[ii])))
P.append( np.angle(tau(200e12,D[ii])) )
D.append( IFA * np.abs(tau(200e12,D[ii])/(1J*s)) )
dA = np.diff(A)
dP = np.diff(P)
dD = np.diff(D)
print(D[-1])
# -
# plot value
if False:
#if True:
plt.plot(range(1,rounds+1),A[1:],'rx')
plt.show()
plt.plot(range(1,rounds+1),P[1:],'rx')
plt.show()
plt.plot(range(1,rounds+1),D[1:],'rx')
plt.show()
# plot variation
if False:
#if True:
plt.plot(range(1,rounds),dA[1:],'rx')
plt.show()
plt.plot(range(1,rounds),dP[1:],'rx')
plt.show()
plt.plot(range(1,rounds),dD[1:],'rx')
plt.show()
# transmittivity 2
def tau2(f, Ew, m):
numerator = -(1-tf(f, Ew)**2) * Bf(f, Ew) * np.exp( 1J * psi(f, m*Ew) / 2 )
denumerator = (1 - tf(f, Ew)**2 * Bf(f, Ew) * np.exp( 1J * psi(f, m*Ew) ) )
return numerator / denumerator
# +
m=1+1
A2 = [np.abs(tau2(200e12, 0.0, m)), ] # normalized output intensity
P2 = [np.angle(tau2(200e12, 0.0, m)), ] # output phase
D2 = [0.0, ] # un-normalized intensity inside the µ-ring resonator
rounds = 14
for ii in range(0, rounds):
A2.append( np.abs(tau2(200e12,D2[ii], m)))
P2.append( np.angle(tau2(200e12,D2[ii], m)) )
D2.append( IFA*np.abs(tau2(200e12,D2[ii],2)/(1J*s)) )
dA2 = np.diff(A2)
dP2 = np.diff(P2)
dD2 = np.diff(D2)
# +
# plot value
plt.plot(range(1,rounds+1),A2[1:],'rx')
plt.show()
plt.plot(range(1,rounds+1),P2[1:],'rx')
plt.show()
plt.plot(range(1,rounds+1),D2[1:],'rx')
plt.show()
# -
plt.plot([0,1],[P[-1],P2[-1]])
plt.show()
def tau_m(f, m, error=1e-16):
Am = [np.abs(tau2(f, 0.0, m)), ] # normalized output intensity
Pm = [np.angle(tau2(f, 0.0, m)), ] # output phase
Dm = [0.0, ] # un-normalized intensity inside the µ-ring resonator
rounds = 4
for ii in range(0, rounds):
Am.append( np.abs(tau2(f, Dm[ii], m)))
Pm.append( np.angle(tau2(f, Dm[ii], m)) )
Dm.append( input_field_amplitude*np.abs(tau2(200e12,Dm[ii],m)/(1J*s)) )
while (Am[-1]-Am[-2]<=error && Pm[-1]-Pm[-2]<=error && Dm[-1]-Dm[-2]<=error)
Am.append( np.abs(tau2(f, Dm[ii], m)))
Pm.append( np.angle(tau2(f, Dm[ii], m)) )
Dm.append( input_field_amplitude*np.abs(tau2(f, Dm[ii], m)/(1J*s)) )
dAm = np.diff(Am)
dPm = np.diff(Pm)
dDm = np.diff(Dm)
#print(Am[-1], Pm[-1], Dm[-1])
return Am[-1], Pm[-1], Dm[-1]
# +
AA = []
PP = []
DD = []
for dd in range(0,10):
a , p, d = cycle(200e12, 1+dd/10)
AA.append(a)
PP.append(p)
DD.append(d)
# +
x = 1+ np.linspace(1,10,10)*0.1
plt.plot(x,PP,'rx')
plt.show()
print(max(PP)-min(PP))
# -
# rensponse of the MZI
x = 1+ np.linspace(1,10,10)*0.1
plt.plot(x, (IFI+np.square(AA)+np.sqrt(AA*IFA)*np.cos(PP-PP[0]) )/IFI )
plt.show()
|
code/Jupyter/all-optical_MZ_modulator/test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import keras
modelI=keras.models.load_model('modelnew.keras')
# +
import cv2
def get_parts_from_image(path, display_img = False ):
img=cv2.imread(path,cv2.IMREAD_GRAYSCALE)
print(path)
if display_img:
plt.axis('off')
plt.imshow(img)
plt.show()
cv2.waitKey(0)
cv2.destroyAllWindows()
train_data=[]
if img is not None:
img=~img
ret,thresh=cv2.threshold(img,127,255,cv2.THRESH_BINARY)
ctrs,ret=cv2.findContours(thresh,cv2.RETR_TREE,cv2.CHAIN_APPROX_SIMPLE)
cnt=sorted(ctrs,key=lambda ctr:cv2.boundingRect(ctr)[0])
w=int(28)
h=int(28)
rects=[]
for c in cnt:
x,y,w,h=cv2.boundingRect(c)
rect=[x,y,w,h]
rects.append(rect)
# print("rects", rects)
bool_rect=[]
for r in rects:
l=[]
for rec in rects:
flag=0
if rec!=r:
if r[0]<(rec[0]+rec[2]+10) and rec[0]<(r[0]+r[2]+10) and r[1]<(rec[1]+rec[3]+10) and rec[1]<(r[1]+r[3]+10):
flag=1
l.append(flag)
if rec==r:
l.append(0)
bool_rect.append(l)
# print("bools",bool_rect)
dump_rect=[]
for i in range(0,len(cnt)):
for j in range(0,len(cnt)):
if bool_rect[i][j]==1:
area1=rects[i][2]*rects[i][3]
area2=rects[j][2]*rects[j][3]
if(area1==min(area1,area2)):
dump_rect.append(rects[i])
# print("dump_rects",dump_rect)
final_rect=[i for i in rects if i not in dump_rect]
# print("Final_rects",final_rect)
for r in final_rect:
x=r[0]
y=r[1]
w=r[2]
h=r[3]
im_crop=thresh[y:y+h+10,x:x+w+10]
im_resize=cv2.resize(im_crop,(28,28))
# cv2.imshow("work",im_resize)
train_data.append(im_resize)
return train_data
# +
import numpy as np
labels_symbols = {
10:'-', 11:'+', 12:'*', 13:'=', 14:'a', 15:'b', 16:'<', 17:'>', 18:'/',19:'√', 20:'(',21:')'
}
for i in range(0,10):
labels_symbols[i] = chr(48+i)
def recognise_parts(train_data):
s=""
for i in range(len(train_data)):
train_data[i]=np.array(train_data[i])
train_data[i]=train_data[i].reshape(1,28,28,1)
# result=modelI.predictclasses(train_data[i])
result=np.argmax(modelI.predict(train_data[i]), axis=1)
s+= labels_symbols[result[0]]
return s
# +
a="NewTest/untitled.jpg"
b="NewTest/untitled1.jpg"
c="NewTest/untitled2.jpg"
d="NewTest/untitled3.jpg"
e="NewTest/untitled4.jpg"
f="NewTest/untitled5.jpg"
g="NewTest/untitled6.jpg"
h="NewTest/untitled7.jpg"
i="Test/expression_new.jpg"
# +
train_data = get_parts_from_image(i, display_img = True)
print(f"number of parts = {len(train_data)}")
plt.rcParams["figure.figsize"] = (20,2)
n_per_line = 10
for i,part in enumerate(train_data):
if i%n_per_line == 0 :
fig, ax = plt.subplots(ncols = n_per_line)
for x in ax: x.axis('off')
ax[i%n_per_line].imshow(part, cmap='gray')
#ax.plot()
plt.rcParams["figure.figsize"] = plt.rcParamsDefault["figure.figsize"]
plt.show()
detected_exp = recognise_parts(train_data)
print(f"Detected Expression : {detected_exp}")
# -
|
.ipynb_checkpoints/Predict-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Duel of sorcerers
# You are witnessing an epic battle between two powerful sorcerers: Gandalf and Saruman. Each sorcerer has 10 spells of variable power in their mind and they are going to throw them one after the other. The winner of the duel will be the one who wins more of those clashes between spells. Spells are represented as a list of 10 integers whose value equals the power of the spell.
# ```
# gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
# saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# ```
# For example:
# 1. The first clash is won by Saruman: 10 against 23, wins 23
# 2. The second clash wins Saruman: 11 against 66, wins 66
# 3. etc.
#
#
# You will create two variables, one for each sorcerer, where the sum of clashes won will be stored. Depending on which variable is greater at the end of the duel, you will show one of the following three results on the screen:
# * Gandalf wins
# * Saruman wins
# * Tie
#
# <img src="images/content_lightning_bolt_big.jpg" width="400">
# ## Solution
# +
# Assign spell power lists to variables
gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# -
# Assign 0 to each variable that stores the victories
ganwins=0
saruwins=0
# +
# Execution of spell clashes
for x in range(len(gandalf)):
if gandalf[x] < saruman[x]:
saruwins+=1
elif gandalf[x]>saruman[x]:
ganwins+=1
# +
# We check who has won, do not forget the possibility of a draw.
if saruwins>ganwins:
print("Winner is Saruman! He won %d out of %d"%(saruwins,len(saruman)))
elif saruwins<ganwins:
print("Winner is Gandalf! He won %d out of %d"%(ganwins,len(saruman)))
elif saruwins==ganwins:
print("It's a tie! Both won the same number of clashes, that's it: %d out of %d"%(ganwins,len(saruman)))
# Print the result based on the winner.
# -
# ## Goals
#
# 1. Treatment of lists
# 2. Use of **for loop**
# 3. Use of conditional **if-elif-else**
# 4. Use of the functions **range(), len()**
# 5. Print
# ## Bonus
#
# 1. Spells now have a name and there is a dictionary that relates that name to a power.
# 2. A sorcerer wins if he succeeds in winning 3 spell clashes in a row.
# 3. Average of each of the spell lists.
# 4. Standard deviation of each of the spell lists.
#
# ```
# POWER = {
# 'Fireball': 50,
# 'Lightning bolt': 40,
# 'Magic arrow': 10,
# 'Black Tentacles': 25,
# 'Contagion': 45
# }
#
# gandalf = ['Fireball', 'Lightning bolt', 'Lightning bolt', 'Magic arrow', 'Fireball',
# 'Magic arrow', 'Lightning bolt', 'Fireball', 'Fireball', 'Fireball']
# saruman = ['Contagion', 'Contagion', 'Black Tentacles', 'Fireball', 'Black Tentacles',
# 'Lightning bolt', 'Magic arrow', 'Contagion', 'Magic arrow', 'Magic arrow']
# ```
#
# Good luck!
# +
# 1. Spells now have a name and there is a dictionary that relates that name to a power.
# variables
POWER = {
'Fireball': 50,
'Lightning bolt': 40,
'Magic arrow': 10,
'Black Tentacles': 25,
'Contagion': 45
}
gandalf = ['Fireball', 'Lightning bolt', 'Lightning bolt', 'Magic arrow', 'Fireball',
'Magic arrow', 'Lightning bolt', 'Fireball', 'Magic arrow', 'Fireball']
saruman = ['Contagion', 'Contagion', 'Black Tentacles', 'Fireball', 'Black Tentacles',
'Lightning bolt', 'Magic arrow', 'Contagion', 'Magic arrow', 'Magic arrow']
gandalfnum=[]
sarumannum=[]
name=""
value=0
# +
# Assign spell power lists to variables
for x in range(len(gandalf)):
spell=gandalf[x]
while spell!=name:
for a,b in POWER.items():
name=a
value=b
if spell==name:
break
gandalfnum.append(b)
for x in range(len(saruman)):
spell=saruman[x]
while spell!=name:
for a,b in POWER.items():
name=a
value=b
if spell==name:
break
sarumannum.append(b)
print("Gandalf: ",gandalfnum)
print("Saruman: ",sarumannum)
# +
# 2. A sorcerer wins if he succeeds in winning 3 spell clashes in a row.
saruwins2=0
ganwins2=0
# Execution of spell clashes
for x in range(len(gandalfnum)-2):
if (gandalfnum[x] < sarumannum[x]) & (gandalfnum[x+1] < sarumannum[x+1]) & (gandalfnum[x+2] < sarumannum[x+2]):
saruwins2+=1
elif (gandalfnum[x] > sarumannum[x]) & (gandalfnum[x+1] > sarumannum[x+1]) & (gandalfnum[x+2] > sarumannum[x+2]):
ganwins2+=1
# check for 3 wins in a row
if saruwins2>ganwins2:
print("Winner is Saruman! He won %d out of %d"%(saruwins2,len(sarumannum)))
elif saruwins2<ganwins2:
print("Winner is Gandalf! He won %d out of %d"%(ganwins2,len(sarumannum)))
elif saruwins2==ganwins2:
print("It's a tie! Both won the same number of clashes, that's it: %d out of %d"%(ganwins2,len(sarumannum)))
# check the winner
# -
# 3. Average of each of the spell lists.
avggan=(sum(gandalfnum)/len(gandalfnum))
print("Average spell list of Gandalf: ",avggan)
avgsar=(sum(sarumannum)/len(sarumannum))
print("Average spell list of Saruman: ",avgsar)
# +
# 4. Standard deviation of each of the spell lists.
summationgan=0
for g in gandalfnum:
summationgan+=((g-avggan)**2)
stdevgan=((summationgan/len(gandalfnum)))**0.5
print("standard deviation for Gandalf is: %.2f" %stdevgan)
summationsar=0
for s in sarumannum:
summationsar+=((s-avgsar)**2)
stdevsar=((summationsar/len(sarumannum)))**0.5
print("standard deviation for Saruman is: %.2f" %stdevsar)
# -
|
duel/duel-Solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 6: Functions and Modules
# February 26, 2021
# Last time we talked about the implementation of repetitive behavior (loops). The lecture covered (pre-test) loops with the ```while``` and ```for``` statements, and the special behavior of the ```break``` and ```continue``` statements. We furthermore discussed the issue of post-test loops in Python.
#
# Today we will discuss functions, modules and packages, have a quick look at the Python’s standard library and the Python Package Index, and introduce the principle of recursion and recursive function definitions. When Python programs become complex, it is a good idea to structure the code using functions, modules, and packages. Proper modularization will keep the code base easier to understand and maintain, and is in fact one of the principles of good software development. Another important principle is the reuse of already existing functionality (rather than implementing it again), so familiarity with the standard libraries is important.
#
# Next time we will cover special data structures in Python, which will allow us to work with more powerful data items than just the individual numbers, strings and Booleans that we have used so far.
# ## Functions
#
# We have seen and used functions before (`print`, `input`, and `bin`, for example), now it is time to have a more systematic look at them and to define functions ourselves. Functions are useful whenever a piece of code forms a self-contained component and is (potentially) used repeatedly in the program. It simplifies coding, testing, debugging and maintenance if this code is only defined once (as a function) and just called when it is needed.
#
# Here is an example of code with unnecessary repetitions:
player1 = input("Please enter name of player: ")
print(f"Hello {player1}, welcome to the game!")
player2 = input("Please enter name of player: ")
print(f"Hello {player2}, welcome to the game!")
player3 = input("Please enter name of player: ")
print(f"Hello {player3}, welcome to the game!")
# Essentially, this code does the same thing three times. This is impractical, for example, if we want to add more players (we have to duplicate a lot of code), or if we want to change the text of the outputs to the players, for instance translate it to Dutch (we would have to do that at all occurrences of the text). So, it might be useful to define functions that ask for a player's name and for displaying a personalized welcome message.
#
# Function definitions in Python have the following basic form:
#
# def <function name>(<function parameter(s)>):
# <do something>
# return <value(s)>
#
# That is, they define the name of the function, which parameters they take as input arguments at runtime (can be none), what they actually do and the value(s) that they return to the calling function (can be none). Functions have to be defined before they can be used. For example, we can define a function that takes a name as a parameter and welcomes a player accordingly:
# +
def greet_player(name):
print(f"Hello {name}, welcome to the game!")
player1 = input("Please enter name of player: ")
greet_player(player1)
player2 = input("Please enter name of player: ")
greet_player(player2)
player3 = input("Please enter name of player: ")
greet_player(player3)
# -
# This already looks better, but we still have redundant code for asking the players to enter their names. We can define another function for that, that has no input parameters, but returns a value to the caller:
# +
def ask_name():
name = input("Please enter name of player: ")
return name
def greet_player(name):
print(f"Hello {name}, welcome to the game!")
player1 = ask_name()
greet_player(player1)
player2 = ask_name()
greet_player(player2)
player3 = ask_name()
greet_player(player3)
# -
# Note that one of our functions contains a return statement (```return name```), and the other does not. When a function does not explicitly define a return value, the ```None``` value (representing “nothingness”) is returned by default. Thus, the return statement can be omitted, as implicitly an (empty) return to the caller will still happen at the end of the function. A return can however also be used to return to the caller at any another point of the function if desired.
#
# Alternatively, if we think that we will never need the actions separately, we could simply define one function that carries out the input and the printout:
# +
def ask_name_and_greet_player():
name = input("Please enter name of player: ")
print(f"Hello {name}, welcome to the game!")
return name
player1 = ask_name_and_greet_player()
player2 = ask_name_and_greet_player()
player3 = ask_name_and_greet_player()
# -
# The functions in the example above are quite short, only one or two lines, and you might wonder if that is actually worth the effort. Well, they were only simple examples to illustrate how functions work. You will soon see in the homework exercises and also in your projects that functions are usually longer and more complex. Below is another example (still small), where the function comprises five lines of code and even a loop:
# +
# function for computing the factorial of a number n
def fac(n):
fac = 1
while n > 0:
fac = fac * n
n = n - 1
return fac
# reading a number from input
n = int(input("Please enter an integer number: "))
# loop computing the factorials for all number from n to 1
while n > 0:
print(f"{n}! is {fac(n)}")
n = n - 1
# -
# This example illustrates another feature of functions: variable names are local to the function. That is, argument names and variables declared inside a function are not visible outside of the function. The area in a program where variables are visible is also called their scope. As a general rule, a variable’s scope is the block that it has been declared in, starting from the point where its name was defined. In the example above, this means that the ```n``` in the ```fac``` function is another ```n``` than the one in the program below the function. The function does not change the value of the n in the calling (part of the) program. This behavior makes a lot of sense sense: If you use a function from a library, where you cannot or do not want to look into its implementation, it is good to know that it will not interfere with your own variables. Generally, data should only be passed to and retrieved from functions through their parameters and return values, respectively, as this is the safest way to avoid undesired side effects from e.g. variables that overwrite each other unintentionally.
#
# Note here that the `global` statement can be used to make variables known across scopes. Functions would have access to its value also without it being passed as a parameter, and they could write results to it without returning it explicitly. The use of the `global` statement is however strongly discouraged for the reasons given above, so we will not discuss it further.
#
# The functions in the examples above have no or only one parameter, but if several inputs are needed, they can simply be defined as comma-separated lists of parameters. For example, a function `add(a,b)` for adding to numbers. When calling the function, the arguments have to passed in the order of the parameters.
#
# Furthermore, functions can have named parameters. They are identified by their name, so their order does not matter when calling the function. They are also given a default value when they are declared, so that when the calling function does not specify them, there is a standard value to work with. Here is an example of a function with one “normal” and two named parameters with default values:
# +
def rectangle_print(letter, columns=3, rows=2):
for i in range(1,rows+1):
print(letter*columns)
rectangle_print("a")
rectangle_print("b", 5, 2)
rectangle_print("c", rows=3)
# -
# The above example illustrates nicely how named parameters can or can not be defined by the caller, depending on what is needed. Note that also the `print` function, which we have used a lot, has named parameters, for example to specify the separator between the string arguments that are passed to print:
print("a", "b", "c", sep="\n")
# In fact, the arbitrary number of strings that can be passed to the print function to be printed to the screen is an example of yet another kind of parameters, the VarArgs (for variable number of arguments) parameters. Parameters become VarArgs by defining them with a single (\*) or double (\*\*) star in front of them. For example, we can use a starred parameter to define a sum function that adds all of its arguments:
# +
def sum_up(*numbers):
s = 0
for n in numbers:
s += n
return s
print(sum_up(1,2,3,4,5))
print(sum_up(1,2,3,4,5,6,7))
# -
# Technically, the arguments for a single-starred parameter form a tuple, and the arguments of a double-starred parameter form a dictionary, hence the latter have to be defined as key-value pairs. What this means will become clear in the next lecture, where we will talk about those data structures in detail.
# ## Modules
#
# Functions make pieces of code in our program file reusable. Modules make it possible to reuse code across different program files. To use the content of another module (file) in a Python program, it has to be imported. We have already seen a module import in the number-guessing game example in the lecture on loops:
# ```
# import random
# number = random.randint(1,10)
# [...]
# ```
#
# The `import` statement in the first line imports the `random` module from Python’s standard library. Afterwards we can use the functions that it provides, for example the `random()` function that generates a random number within a specified range.
#
# Note that import statements can in principle be placed at any point in the program, as long as they happen before using their functions. This is not considered good coding style, however, as imports scattered all over the program make it difficult to see what has already been imported, and might even lead to redundant and conflicting imports. To avoid such problems, all imports that are used in a .py file should be placed at its beginning.
#
# We can also create modules ourselves. There are different forms in which modules can exist. The simplest is a .py file that contains different functions. (In fact, any .py file we create is a module that can potentially be used by other programs.) Generally, it makes sense to create modules for sets of functions that somehow belong together and where it would make sense to distribute them (to other programmers) as a unit.
#
# For example, we might provide the `ask_name()` and `greet_player()` functions from above in a `player_management.py` module, along with other related functions such as, e.g., `show_score()` or `show_game_over()`:
#
# def ask_name():
# name = input("Please enter name of player: ")
# return name
#
# def greet_player(name):
# print(f"Hello {name}, welcome to the game!")
#
# def show_score(name, score):
# print(f"Hello {name}, your current score is {score}.")
#
# def show_game_over(name):
# print(f"GAME OVER! Sorry, {name}...")
#
# The module and its functions might then be used by a game as follows:
#
# +
import lib.player_management
import random
player = player_management.ask_name()
player_management.greet_player(player)
score = random.randint(0,10) # (replace by actual code for playing game)
if score > 0:
player_management.show_score(player,score)
else:
player_management.show_game_over(player)
# -
# Also when loading modules the Python interpreter acts as usual and will step through the module when importing it. The function definitions in the module are read, and thus the functions are subsequently available for use. If the module contains other code, outside of function definitions, it will be executed, too.
#
# Note that when creating own modules, it is advisable to not use names that are already taken by modules from the standard library or other popular collections. Name clashes can lead to errors and unpredictable problem behaviour. So, if you observe weird behavior in relation to your modules and imports, check if maybe there is a name clash. There are ways to influence Python’s importing mechanism in more detail to circumvent problems, but that gets quite technical and furthermore tends to be unstable, so we will not discuss it in this course.
# There are two variations of the `import` statement that you will often see in Python code that is around: The `from … import …` statement can be used to import individual functions from a module, for example:
from random import randint
print(randint(1,10))
# This way, the function is available without using the module name as a prefix. It is not recommended to use this kind of import, however, as it comes with a risk of name clashes and unreliable behavior. The other variation is the ```import``` … ```as``` … statement, which defines an alias for the module name. For example:
import random as rd
print(rd.randint(1,10))
# This is handy in particular for defining shorter aliases for long module names, but it should be used with care, too, to avoid name clashes.
#
# It is important to realize that all Python .py files are modules. However, not all of them are (just) made for being imported into other modules. Some also have code that should be executed when the module is executed as a script or with ```python -m```. Interestingly, a module can discover if it is being executed directly (in Python speech: running in the main scope) or as an import. A pattern that is often used in Python modules is to include such a check, and call a dedicated main function if the method is indeed running in the main scope:
#
# if __name__ == "__main__":
#
# # execute only if running in main scope
# main()
#
# This allows to have a module providing functions that are useful to import by other modules, and at the same time, for example, offering a command line interface that is useful when starting the module as a script, but would only be a nuisance would it be run during a standard import.
# ## Packages
#
# Packages are used to organize sets of modules hierarchically. Technically, they are folders that contains modules and a special `__init__.py` file (to indicate that the modules in this folder form a package). We don’t go into further detail here, but note that when using modules from collections of functionality such as the Python Standard Library, they are distributed as packages.
# ## The Standard Library and other Sources of Functionality
#
# The Python Standard Library, which is distributed with the Python installation, contains ```random``` and a large number of other useful packages, such as the ```calendar```, ```statistics``` or ```io``` libraries. We will see and use some of them in the following lectures. The website https://docs.python.org/3/library/index.html lists what is contained in the standard library of Python and is a good reference during development.
#
# In addition to the packages in the standard library, there are also many packages available from other sources. The Python Package index at https://pypi.python.org/pypi is the central repository for Python packages, currently ca. 290,000 of them. With the Anaconda platform a large number of popular packages has already been installed on your system, but sometimes packages are not yet installed on your machine, so you have to do that yourself before you can use them. How to do this within Anaconda is described at https://docs.anaconda.com/anaconda/navigator/tutorials/manage-packages/#installing-a-package. Alternatively, the basic way to do install new packages is via the command line, using the ```pip``` tool that comes with the Python installation: ```pip install <module>```
# ## Recursion
#
# See Recursion.
#
# Just a joke, but it points to the basic idea of recursion: self-reference. You may recall recursive function definitions from mathematics, for instance:
#
# x$^n$ = x * x$^{n-1}$ if n > 0, and 1 if n = 0
#
# That is, the problem is solved by referring to a smaller instance of the same problem, until it is so small that it is trivial. For example, with this definition 35 is calculated as follows:
#
# 3$^5$ = 3 * 3$^4$ = 3 * 3 * 3$^3$ = 3 * 3 * 3 * 3$^2$ = 3 * 3 * 3 * 3 * 3$^1$ = 3 * 3 * 3 * 3 * 3 * 3$^0$ = 3 * 3 * 3 * 3 * 3 * 1 = 3 * 3 * 3 * 3 * 3 = 3 * 3 * 3 * 9 = 3 * 3 * 27 = 3 * 81 = 243
#
# Recursions in Python (and other programming languages as well) follow the same principle, but the definition in code looks a bit different:
# +
# recursive function to compute x**n
def pow(x,n):
if n > 0:
return x * pow(x,n-1)
else:
return 1
# main program calling the function
x = int(input("Please enter x: "))
n = int(input("Please enter n: "))
x_to_the_power_of_n = pow(x,n)
print(f"{x} ** {n} = {x_to_the_power_of_n}")
# -
# Internally, the following calls and returns of pow(x,n) happen at runtime:
#
# ```
# pow(3,5)
# pow(3,4)
# pow(3,3)
# pow(3,2)
# pow(3,1)
# pow(3,0)
# return 1
# return 3
# return 9
# return 27
# return 81
# return 243
# ```
# Now let us look at an example for which there is not already an operator in Python. For example, the Fibonacci number for an integer n is defined as:
#
# fib(n) = fib(n-1) + fib(n-2) if n > 1, 1 if n = 1 and 0 if n = 0
#
# In Python:
# +
def fib(n):
if n > 1:
return fib(n-1) + fib(n-2)
elif n == 1:
return 1
else:
return 0
n = int(input("Please enter an integer number: "))
print(f"fib({n}) = {fib(n)}")
# -
# The call stack here is a bit more complex than above (only shown for fib(4) to keep it short):
# ```
# fib(4)
# fib(3)
# fib(2)
# fib(1)
# return 1
# fib(0)
# return 0
# return 1
# fib(1)
# return 1
# return 2
# fib(2)
# fib(1)
# return 1
# fib(0)
# return 0
# return 1
# return 3
# ```
# Here is another, non-mathematical example:
# +
def walk_up_and_down(floors):
if floors > 0:
print("Walk one floor up.")
walk_up_and_down(floors-1)
print("Walk one floor down.")
else:
print("Reached the top floor!")
walk_up_and_down(3)
# -
# The call stack here is straightforward again. Note that also without an explicit "return value" statement, the function returns control to its caller when it is finished, but then without returning a value:
# ```
# walk_up_and_down(3)
# walk_up_and_down(2)
# walk_up_and_down(1)
# walk_up_and_down(0)
# (return)
# (return)
# (return)
# (return)
# ```
#
# Each recursion can be implemented by an iterative loop, and vice versa. For the walking-up-and-down example, a corresponding iterative version is:
# +
def walk_up_and_down_iteratively(floors):
i = 0
while i < floors:
print("Walk one floor up.")
i = i + 1
print("Reached the top floor!")
while i > 0:
print("Walk one floor down.")
i = i - 1
walk_up_and_down_iteratively(3)
# -
# Generally, iterative implementations tend to be faster (because they do not have to create several instances of the same method), but sometimes a recursive solution is more elegant (though maybe more difficult to understand).
# ## Exercises
#
# Please use Quarterfall to submit and check your answers.
# ### 1. Leap Years (★★★★☆)
# In our lifetimes (unless we happen to get veeery old) a leap year occurs every four years. But actually, the rule is a bit more involved: A year is a leap year if it is a multiple of 4, but not a multiple of 100, unless it is also a multiple of 400. For example, 1984 and 2000 were leap years, but 1900 and 1985 were not.
# Write a function is_leap_year(year) that tests if the year is a leap year. If so, the function should return True, and False otherwise. Implement the function using only one Boolean expression.
# You can use the code below to test your function (the first line defines a list of years to iterate over in the loop – lists will be explained in detail in the next lecture):
# ```
# tests = [1900, 1984, 1985, 2000, 2018]
#
# for test in tests:
# if is_leap_year(test):
# print(f”{test} is a leap year")
# else:
# print(f”{test} is not a leap year")
# ```
#
# The output should be:
# ```
# 1900 is not a leap year
# 1984 is a leap year
# 1985 is not a leap year
# 2000 is a leap year
# 2018 is not a leap year
# ```
# ### 2. Calculator (★★★☆☆)
# Write a program that acts a simple calculator, asking the user if they want to add, subtract, multiply or divide two arbitrary numbers. Define functions add(x,y), subtract(x,y), multiply(x,y) and divide(x,y) for this. (Normally one would not define functions for these basic operators, but this is just an exercise...) After the user has selected an operation, they are asked to enter the numbers x and y. The program calculates and prints the result. The output should be something like:
# ```
# You have four options:
# 1 - Add
# 2 – Subtract
# 3 - Multiply
# 4 - Divide
# Enter choice (1/2/3/4): 2
# Enter first number: 34
# Enter second number: 53
# 34 - 53 = -19
# ```
# ### 3. Password Generator (★★★★☆)
# People often use passwords that are too short or too simple and can easily be guessed. (“<PASSWORD>”, “Password” and “<PASSWORD>” were the most frequently used passwords in 2017!) Moreover, people tend to use the same password for different services, which makes it easy for criminals to take over other accounts once they have obtained one of the passwords. Thus, it is wise to use passwords that are reasonably long (8 characters minimum), consist of seemingly random sequences of letters (use of special characters is by the way not so important), and have a separate password for each account.
#
# Write a program that helps you to create reasonably good passwords. Therefore define and implement a function create_password(length) that takes the desired length of the password as parameter. If a password shorter than 8 characters is requested, the function should refuse to create it (as it would not be secure). If the requested length is longer, then the function should fill the password with random letters (upper and lower case) and numbers.
# You can use the following code to test your function:
# ```
# print(create_password(4))
# print(create_password(8))
# print(create_password(12))
# print(create_password(16))
# ```
# The output for the test code above should be something like:
# ```
# Too short, please create longer password.
# None
# pk4lU4Cr
# UPFzFg6Pn14r
# ALdVi3yT0khuxzTr
# ```
# Hint: Consider using the `random.choice()` function from the `random` library, which can be used to select a random character from a predefined string.
# ### 4. Basic Statistics (★★★★☆)
# Use the statistics package from the Python standard library to define and implement a function `print_basic_statistics()` with the following characteristics:
# * the function takes arbitrarily many numbers as input
# * the default case is that the function prints the arithmetic mean, median, standard variation and variance of the input data to the screen
# * via a named parameter the calling code should also have the option to select only one of the four to be printed
#
# You can use the following code to test your function:
# ```
# print_basic_statistics(91,82,19,13,44,)
# print_basic_statistics(91,82,19,13,44,73,18,95,17,65,output="median")
# ```
# The output for the test code above should be something like:
# ```
# The mean of (91, 82, 19, 13, 44) is 49.8.
# The median of (91, 82, 19, 13, 44) is 44.0.
# The standard deviation of (91, 82, 19, 13, 44) is 35.6.
# The variance of (91, 82, 19, 13, 44) is 1267.7.
# ```
# ```
# The median of (91, 82, 19, 13, 44, 73, 18, 95, 17, 65) is 54.5.
# ```
# ### 5. Ackermann Function (★★★★☆)
# The Ackermann function (named after the German mathematician <NAME>) grows rapidly already for small inputs. It exists in different variants, one of the common definitions is the following (for two nonnegative integers m and n):
#
# 
#
# Define and implement a (recursive) function ackermann(m,n) that computes the Ackermann function value for two nonnegative integers m and n. Then, write a test program that computes the results for calling the function with growing m and n of the same value, starting with m = n = 0 and incrementing by 1 in each iteration. The output should be something like:
# ```
# ackermann(0,0) = 1
# ackermann(1,1) = 3
# ackermann(2,2) = 7
# [...]
# ```
# What is the last value that your program computes before you get a `RecursionError`? (Hint: It might be that the outputs in the IPython console in Spyder are too verbose to see anything. You can alternatively run your program from the command line to see more.) What does this error mean?
# ## Extras for the Weekend
# CheckiO (https://checkio.org/) is a game where you need to code in Python (or JavaScript) to get further. By now you should know enough Python to try it out and solve the challenges there.
#
# If you want to get your brain twisted with something really geeky, have a look at quines. Quines are programs that print themselves (i.e., their own code) during execution, but it is not allowed that they read in their code from the .py file. Here is an example of a quine in Python:
# ```
# q="\nprint('q='+repr(q)+q)"
# print('q='+repr(q)+q)
# ```
# Can you write another one?
|
Lecturenotes/M06_Functions_and_Modules.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# ## Lagrange Polynomials
#
# This tutorial uses the same example as the [problem formulation](./problem_formulation.ipynb).
#
# [Lagrange polynomials](https://en.wikipedia.org/wiki/Lagrange_polynomial) are not a method for creating orthogonal polynomials.
# Instead it is an interpolation method for creating an polynomial expansion that has the property that each polynomial interpolates exactly one point in space with the value 1 and has the value 0 for all other interpolation values.
#
# To summarize, we need to do the following:
#
# * Generate $Q_1, ..., Q_N = (\alpha_1, \beta_1), ..., (\alpha_N, \beta_N)$, using (pseudo-)random samples or otherwise.
# * Construct a Lagrange polynomial expansion $\Phi_1, ..., \Phi_M$ from the samples $Q_1, ..., Q_N$.
# * Evaluate model predictor $U_1, ..., U_N$ for each sample.
# * Analyze model approximation $u(t; \alpha, \beta) = \sum_m U_m(t) \Phi_n(\alpha, \beta)$ instead of actual model solver.
# %% [markdown]
# ### Evaluation samples
#
# In chaospy, creating low-discrepancy sequences can be done using the `distribution.sample` method.
# Creating quadrature points can be done using `chaospy.generate_quadrature`.
# For example:
# %%
from problem_formulation import joint
samples = joint.sample(5, rule="halton")
# %%
from matplotlib import pyplot
pyplot.scatter(*samples)
pyplot.rc("figure", figsize=[6, 4])
pyplot.xlabel("Parameter $I$ (normal)")
pyplot.ylabel("Parameter $a$ (uniform)")
pyplot.axis([1, 2, 0.1, 0.2])
pyplot.show()
# %% [markdown]
# ### Lagrange basis
#
# Creating an expansion of Lagrange polynomial terms can be done using the following constructor:
# %%
import chaospy
polynomial_expansion = chaospy.expansion.lagrange(samples)
polynomial_expansion[0].round(2)
# %% [markdown]
# On can verify that the returned polynomials follows the property of evaluating 0 for all but one of the samples used in the construction as follows:
# %%
polynomial_expansion(*samples).round(8)
# %% [markdown]
# ### Model approximation
#
# Fitting the polynomials to the evaluations does not need to involve regression.
# Instead it is enough to just multiply the Lagrange polynomial expansion with the evaluations, and sum it up:
# %%
from problem_formulation import model_solver
model_evaluations = numpy.array([
model_solver(sample) for sample in samples.T])
model_approximation = chaospy.sum(
model_evaluations.T*polynomial_expansion, axis=-1).T
model_approximation.shape
# %% [markdown]
# ### Assess statistics
#
# The results is close to what we are used to from the other methods:
# %%
expected = chaospy.E(model_approximation, joint)
variance = chaospy.Var(model_approximation, joint)
# %%
from problem_formulation import coordinates
pyplot.fill_between(coordinates, expected-variance**0.5,
expected+variance**0.5, alpha=0.3)
pyplot.plot(coordinates, expected)
pyplot.axis([0, 10, 0, 2])
pyplot.xlabel("coordinates $t$")
pyplot.ylabel("Model evaluations $u$")
pyplot.show()
# %% [markdown]
# ### Avoiding matrix inversion issues
#
# It is worth noting that the construction of Lagrange polynomials are not always numerically stable.
# For example when using grids along , most often the expansion construction fails:
# %%
nodes, _ = chaospy.generate_quadrature(1, joint)
try:
chaospy.expansion.lagrange(nodes)
except numpy.linalg.LinAlgError as err:
error = err.args[0]
error
# %% [markdown]
# It is impossible to avoid the issue entirely, but in the case of structured grid, it is often possible to get around the problem.
# To do so, the following steps can be taken:
#
# * Create multiple Lagrange polynomials, one for each marginal distribution.
# * Rotate dimensions so each expansion get a dimension corresponding to the marginal it was created for.
# * Use an outer product of the expansions to combine them into a single expansion.
#
# So we begin with making marginal Lagrange polynomials:
# %%
nodes, _ = list(zip(*[chaospy.generate_quadrature(1, marginal)
for marginal in joint]))
expansions = [chaospy.expansion.lagrange(nodes_)
for nodes_ in nodes]
[expansion_.round(4) for expansion_ in expansions]
# %% [markdown]
# Then we rotate the dimensions:
# %%
vars_ = chaospy.variable(len(joint))
expansions = [
expans(q0=var)
for expans, var in zip(expansions, vars_)]
[expans.round(4) for expans in expansions]
# %% [markdown]
# And finally construct the final expansion using an outer product:
# %%
expansion = chaospy.outer(*expansions).flatten()
expansion.round(2)
# %% [markdown]
# The end results can be verified to have the properties we are looking for:
# %%
nodes, _ = chaospy.generate_quadrature(1, joint)
expansion(*nodes).round(8)
|
docs/user_guide/main_usage/lagrange_polynomials.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# Copyright (c) 2020. <NAME> (<EMAIL>)
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
import os
import re
from operator import itemgetter
import pandas as pd
import pickle
import numpy as np
from sklearn.feature_selection import SelectKBest, f_classif, mutual_info_classif
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.utils import compute_class_weight
from tqdm.auto import tqdm
from logger import Logger
from secrets import api_key
from utils import *
class DataGenerator:
def __init__(self, company_code, data_path='./stock_history', output_path='./outputs', strategy_type='original',
update=False, logger: Logger = None):
self.company_code = company_code
self.strategy_type = strategy_type
self.data_path = data_path
self.logger = logger
self.BASE_URL = "https://www.alphavantage.co/query?function=TIME_SERIES_DAILY_ADJUSTED" \
"&outputsize=full&apikey=" + api_key + "&datatype=csv&symbol=" # api key from alpha vantage service
self.output_path = output_path
self.start_col = 'open'
self.end_col = 'eom_26'
self.update = update
self.download_stock_data()
self.df = self.create_features()
self.feat_idx = self.feature_selection()
self.one_hot_enc = OneHotEncoder(sparse=False, categories='auto')
self.one_hot_enc.fit(self.df['labels'].values.reshape(-1, 1))
self.batch_start_date = self.df.head(1).iloc[0]["timestamp"]
self.test_duration_years = 1
self.logger.append_log("{} has data for {} to {}".format(data_path, self.batch_start_date,
self.df.tail(1).iloc[0]['timestamp']))
def log(self, text):
if self.logger:
self.logger.append_log(text)
else:
print(text)
def download_stock_data(self):
path_to_company_data = self.data_path
print("path to company data:", path_to_company_data)
parent_path = os.sep.join(path_to_company_data.split(os.sep)[:-1])
if not os.path.exists(parent_path):
os.makedirs(parent_path)
print("Company Directory created", parent_path)
if not os.path.exists(path_to_company_data):
self.log("Downloading " + self.company_code + " data")
download_save(self.BASE_URL + self.company_code, path_to_company_data, self.logger)
else:
self.log("Data for " + self.company_code + " ready to use")
def calculate_technical_indicators(self, df, col_name, intervals):
# get_RSI(df, col_name, intervals) # faster but non-smoothed RSI
get_RSI_smooth(df, col_name, intervals) # momentum
get_williamR(df, col_name, intervals) # momentum
get_mfi(df, intervals) # momentum
# get_MACD(df, col_name, intervals) # momentum, ready to use +3
# get_PPO(df, col_name, intervals) # momentum, ready to use +1
get_ROC(df, col_name, intervals) # momentum
get_CMF(df, col_name, intervals) # momentum, volume EMA
get_CMO(df, col_name, intervals) # momentum
get_SMA(df, col_name, intervals)
get_SMA(df, 'open', intervals)
get_EMA(df, col_name, intervals)
get_WMA(df, col_name, intervals)
get_HMA(df, col_name, intervals)
get_TRIX(df, col_name, intervals) # trend
get_CCI(df, col_name, intervals) # trend
get_DPO(df, col_name, intervals) # Trend oscillator
get_kst(df, col_name, intervals) # Trend
get_DMI(df, col_name, intervals) # trend
get_BB_MAV(df, col_name, intervals) # volatility
# get_PSI(df, col_name, intervals) # can't find formula
get_force_index(df, intervals) # volume
get_kdjk_rsv(df, intervals) # ready to use, +2*len(intervals), 2 rows
get_EOM(df, col_name, intervals) # volume momentum
get_volume_delta(df) # volume +1
get_IBR(df) # ready to use +1
def create_labels(self, df, col_name, window_size=11):
"""
Data is labeled as per the logic in research paper
Label code : BUY => 1, SELL => 0, HOLD => 2
params :
df => Dataframe with data
col_name => name of column which should be used to determine strategy
returns : numpy array with integer codes for labels with
size = total-(window_size)+1
"""
self.log("creating label with original paper strategy")
row_counter = 0
total_rows = len(df)
labels = np.zeros(total_rows)
labels[:] = np.nan
print("Calculating labels")
pbar = tqdm(total=total_rows)
while row_counter < total_rows:
if row_counter >= window_size - 1:
window_begin = row_counter - (window_size - 1)
window_end = row_counter
window_middle = (window_begin + window_end) / 2
min_ = np.inf
min_index = -1
max_ = -np.inf
max_index = -1
for i in range(window_begin, window_end + 1):
price = df.iloc[i][col_name]
if price < min_:
min_ = price
min_index = i
if price > max_:
max_ = price
max_index = i
if max_index == window_middle:
labels[window_middle] = 0
elif min_index == window_middle:
labels[window_middle] = 1
else:
labels[window_middle] = 2
row_counter = row_counter + 1
pbar.update(1)
pbar.close()
return labels
def create_labels_price_rise(self, df, col_name):
"""
labels data based on price rise on next day
next_day - prev_day
((s - s.shift()) > 0).astype(np.int)
"""
df["labels"] = ((df[col_name] - df[col_name].shift()) > 0).astype(np.int)
df = df[1:]
df.reset_index(drop=True, inplace=True)
def create_label_mean_reversion(self, df, col_name):
"""
strategy as described at "https://decodingmarkets.com/mean-reversion-trading-strategy"
Label code : BUY => 1, SELL => 0, HOLD => 2
params :
df => Dataframe with data
col_name => name of column which should be used to determine strategy
returns : numpy array with integer codes for labels
"""
self.log("creating labels with mean mean-reversion-trading-strategy")
get_RSI_smooth(df, col_name, [3]) # new column 'rsi_3' added to df
rsi_3_series = df['rsi_3']
ibr = get_IBR(df)
total_rows = len(df)
labels = np.zeros(total_rows)
labels[:] = np.nan
count = 0
for i, rsi_3 in enumerate(rsi_3_series):
if rsi_3 < 15: # buy
count = count + 1
if 3 <= count < 8 and ibr.iloc[i] < 0.2: # TODO implement upto 5 BUYS
labels[i] = 1
if count >= 8:
count == 0
elif ibr.iloc[i] > 0.7: # sell
labels[i] = 0
else:
labels[i] = 2
return labels
def create_label_short_long_ma_crossover(self, df, col_name, short, long):
"""
if short = 30 and long = 90,
Buy when 30 day MA < 90 day MA
Sell when 30 day MA > 90 day MA
Label code : BUY => 1, SELL => 0, HOLD => 2
params :
df => Dataframe with data
col_name => name of column which should be used to determine strategy
returns : numpy array with integer codes for labels
"""
self.log("creating label with {}_{}_ma".format(short, long))
def detect_crossover(diff_prev, diff):
if diff_prev >= 0 > diff:
# buy
return 1
elif diff_prev <= 0 < diff:
return 0
else:
return 2
get_SMA(df, 'close', [short, long])
labels = np.zeros((len(df)))
labels[:] = np.nan
diff = df['close_sma_' + str(short)] - df['close_sma_' + str(long)]
diff_prev = diff.shift()
df['diff_prev'] = diff_prev
df['diff'] = diff
res = df.apply(lambda row: detect_crossover(row['diff_prev'], row['diff']), axis=1)
print("labels count", np.unique(res, return_counts=True))
df.drop(columns=['diff_prev', 'diff'], inplace=True)
return res
def create_features(self):
if not os.path.exists(os.path.join(self.output_path, "df_" + self.company_code+".csv")) or self.update:
df = pd.read_csv(self.data_path, engine='python')
df['timestamp'] = pd.to_datetime(df['timestamp'])
df.sort_values('timestamp', inplace=True)
df.reset_index(drop=True, inplace=True)
intervals = range(6, 27) # 21
self.calculate_technical_indicators(df, 'close', intervals)
self.log("Saving dataframe...")
df.to_csv(os.path.join(self.output_path, "df_" + self.company_code+".csv"), index=False)
else:
self.log("Technical indicators already calculated. Loading...")
df = pd.read_csv(os.path.join(self.output_path, "df_" + self.company_code+".csv"))
df['timestamp'] = pd.to_datetime(df['timestamp'])
df.sort_values('timestamp', inplace=True)
df.reset_index(drop=True, inplace=True)
# pickle.load(open(os.path.join(self.output_path, "df_" + self.company_code), "rb"))
prev_len = len(df)
df.dropna(inplace=True)
df.reset_index(drop=True, inplace=True)
self.logger.append_log("Dropped {0} nan rows before label calculation".format(prev_len - len(df)))
if 'labels' not in df.columns or self.update:
if re.match(r"\d+_\d+_ma", self.strategy_type):
short = self.strategy_type.split('_')[0]
long = self.strategy_type.split('_')[1]
df['labels'] = self.create_label_short_long_ma_crossover(df, 'close', short, long)
else:
df['labels'] = self.create_labels(df, 'close')
prev_len = len(df)
df.dropna(inplace=True)
df.reset_index(drop=True, inplace=True)
self.logger.append_log("Dropped {0} nan rows after label calculation".format(prev_len - len(df)))
df.drop(columns=['dividend_amount', 'split_coefficient'], inplace=True)
df.to_csv(os.path.join(self.output_path, "df_" + self.company_code + ".csv"), index=False)
else:
print("labels already calculated")
# pickle.dump(df, open(os.path.join(self.output_path, "df_" + self.company_code), 'wb'))
# console_pretty_print_df(df.head())
self.log("Number of Technical indicator columns for train/test are {}".format(len(list(df.columns)[7:])))
return df
def feature_selection(self):
df_batch = self.df_by_date(None, 10)
list_features = list(df_batch.loc[:, self.start_col:self.end_col].columns)
mm_scaler = MinMaxScaler(feature_range=(0, 1)) # or StandardScaler?
x_train = mm_scaler.fit_transform(df_batch.loc[:, self.start_col:self.end_col].values)
y_train = df_batch['labels'].values
num_features = 225 # should be a perfect square
topk = 350
select_k_best = SelectKBest(f_classif, k=topk)
select_k_best.fit(x_train, y_train)
selected_features_anova = itemgetter(*select_k_best.get_support(indices=True))(list_features)
select_k_best = SelectKBest(mutual_info_classif, k=topk)
select_k_best.fit(x_train, y_train)
selected_features_mic = itemgetter(*select_k_best.get_support(indices=True))(list_features)
common = list(set(selected_features_anova).intersection(selected_features_mic))
self.log("common selected featues:" + str(len(common)) + ", " + str(common))
if len(common) < num_features:
raise Exception(
'number of common features found {} < {} required features. Increase "topK"'.format(len(common),
num_features))
feat_idx = []
for c in common:
feat_idx.append(list_features.index(c))
feat_idx = sorted(feat_idx[0:225])
self.log(str(feat_idx))
return feat_idx
def df_by_date(self, start_date=None, years=5):
if not start_date:
start_date = self.df.head(1).iloc[0]["timestamp"]
end_date = start_date + pd.offsets.DateOffset(years=years)
df_batch = self.df[(self.df["timestamp"] >= start_date) & (self.df["timestamp"] <= end_date)]
return df_batch
def get_data(self, start_date=None, years=5):
df_batch = self.df_by_date(start_date, years)
x = df_batch.loc[:, self.start_col:self.end_col].values
x = x[:, self.feat_idx]
mm_scaler = MinMaxScaler(feature_range=(0, 1)) # or StandardScaler?
x = mm_scaler.fit_transform(x)
dim = int(np.sqrt(x.shape[1]))
x = reshape_as_image(x, dim, dim)
x = np.stack((x,) * 3, axis=-1)
y = df_batch['labels'].values
sample_weights = self.get_sample_weights(y)
y = self.one_hot_enc.transform(y.reshape(-1, 1))
return x, y, df_batch, sample_weights
def get_sample_weights(self, y):
"""
calculate the sample weights based on class weights. Used for models with
imbalanced data and one hot encoding prediction.
params:
y: class labels as integers
"""
y = y.astype(int) # compute_class_weight needs int labels
class_weights = compute_class_weight('balanced', np.unique(y), y)
print("real class weights are {}".format(class_weights), np.unique(y))
print("value_counts", np.unique(y, return_counts=True))
sample_weights = y.copy().astype(float)
for i in np.unique(y):
sample_weights[sample_weights == i] = class_weights[i] # if i == 2 else 0.8 * class_weights[i]
# sample_weights = np.where(sample_weights == i, class_weights[int(i)], y_)
return sample_weights
def get_rolling_data_next(self, start_date=None, window_size_yrs=6, cross_val_split=0.2):
if not start_date:
start_date = self.batch_start_date
x_train, y_train, df_batch_train, sample_weights = self.get_data(start_date, window_size_yrs)
train_end_date = df_batch_train.tail(1).iloc[0]["timestamp"]
test_start_date = train_end_date + pd.offsets.DateOffset(days=1)
test_end_date = test_start_date + pd.offsets.DateOffset(years=self.test_duration_years)
x_test, y_test, df_batch_test, _ = self.get_data(test_start_date, self.test_duration_years)
x_train, x_cv, y_train, y_cv, sample_weights, _ = train_test_split(x_train, y_train, sample_weights,
train_size=1 - cross_val_split,
test_size=cross_val_split,
random_state=2, shuffle=True,
stratify=y_train)
self.logger.append_log("data generated: train duration={}-{}, test_duration={}-{}, size={}, {}, {}".format(
self.batch_start_date, train_end_date, test_start_date, test_end_date, x_train.shape, x_cv.shape,
x_test.shape))
self.batch_start_date = self.batch_start_date + pd.offsets.DateOffset(years=1)
is_last_batch = False
if (self.df.tail(1).iloc[0]["timestamp"] - test_end_date).days < 180: # 6 months
is_last_batch = True
return x_train, y_train, x_cv, y_cv, x_test, y_test, df_batch_train, df_batch_test, \
sample_weights, is_last_batch
# -
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error
# +
diabetes = datasets.load_diabetes()
print(diabetes.keys())
diabetes_x = diabetes.data[:, np.newaxis, 2]
diabetes_x_train = diabetes_x[:-30]
diabetes_x_test = diabetes_x[-30:]
diabetes_y_train = diabetes.target[:-30]
diabetes_y_test = diabetes.target[-30:]
model = linear_model.LinearRegression()
model.fit(diabetes_x_train,diabetes_y_train)
diabetes_y_predicted = model.predict(diabetes_x_test)
print("Mean Squared Error is: ", mean_squared_error(diabetes_y_test,diabetes_y_predicted))
print("Weights: ", model.coef_)
print("Intercept: ", model.intercept_)
#plotting graph with scatter on testing data
plt.scatter(diabetes_x_test,diabetes_y_test)
plt.plot(diabetes_x_test,diabetes_y_predicted)
plt.show()
# +
#dict_keys(['data', 'target', 'DESCR', 'feature_names', 'data_filename', 'target_filename'])
# -
|
linear regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Make sure this code run at the parent level of raw data file
import os
import pandas as pd
os.getcwd()
## setting current directory to the file dir
pwd = 'raw_data'
cwd = os.path.join(os.getcwd(), pwd)
os.chdir(cwd)
## getting all the csv file names
files = [p for p in os.listdir() if p.endswith('.csv')]
## concatenate all the sheets
outf = pd.read_csv(files[0])
print(outf.shape)
for f in files[1:]:
try:
outf = pd.concat([outf , pd.read_csv(f)])
except:
print('Error: {}'.format(f))
else:
print(outf.shape)
err_files = ['export_export_reviews (17).csv', 'export_export_reviews (18).csv']
## using different encoding and separator to read error files
for f in err_files:
data = pd.read_csv(f, sep='\t', encoding='ISO-8859-1')
outf = pd.concat([outf , data])
print(outf.shape)
## setting current directory back the project dir
cwd = os.path.dirname(os.getcwd())
os.chdir(cwd)
## write out the file
outf.to_csv('raw_data.csv', index=False)
|
gathering_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Natural language inference: models
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2021"
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [Sparse feature representations](#Sparse-feature-representations)
# 1. [Feature representations](#Feature-representations)
# 1. [Model wrapper for hyperparameter search](#Model-wrapper-for-hyperparameter-search)
# 1. [Assessment](#Assessment)
# 1. [Hypothesis-only baselines](#Hypothesis-only-baselines)
# 1. [Sentence-encoding models](#Sentence-encoding-models)
# 1. [Dense representations](#Dense-representations)
# 1. [Sentence-encoding RNNs](#Sentence-encoding-RNNs)
# 1. [Other sentence-encoding model ideas](#Other-sentence-encoding-model-ideas)
# 1. [Chained models](#Chained-models)
# 1. [Simple RNN](#Simple-RNN)
# 1. [Separate premise and hypothesis RNNs](#Separate-premise-and-hypothesis-RNNs)
# 1. [Attention mechanisms](#Attention-mechanisms)
# 1. [Error analysis with the MultiNLI annotations](#Error-analysis-with-the-MultiNLI-annotations)
# ## Overview
#
# This notebook defines and explores a number of models for NLI. The general plot is familiar from [our work with the Stanford Sentiment Treebank](sst_01_overview.ipynb):
#
# 1. Models based on sparse feature representations
# 1. Linear classifiers and feed-forward neural classifiers using dense feature representations
# 1. Recurrent neural networks (and, briefly, tree-structured neural networks)
#
# The twist here is that, while NLI is another classification problem, the inputs have important high-level structure: __a premise__ and __a hypothesis__. This invites exploration of a host of neural designs:
#
# * In __sentence-encoding__ models, the premise and hypothesis are analyzed separately, and combined only for the final classification step.
#
# * In __chained__ models, the premise is processed first, then the hypotheses, giving a unified representation of the pair.
#
# NLI resembles sequence-to-sequence problems like __machine translation__ and __language modeling__. The central modeling difference is that NLI doesn't produce an output sequence, but rather consumes two sequences to produce a label. Still, there are enough affinities that many ideas have been shared among these areas.
# ## Set-up
#
# See [the previous notebook](nli_01_task_and_data.ipynb#Set-up) for set-up instructions for this unit.
from collections import Counter
from itertools import product
import nli
import numpy as np
import os
import pandas as pd
from sklearn.exceptions import ConvergenceWarning
from sklearn.linear_model import LogisticRegression
import torch
import torch.nn as nn
import torch.utils.data
from torch_model_base import TorchModelBase
from torch_rnn_classifier import TorchRNNClassifier, TorchRNNModel
from torch_shallow_neural_classifier import TorchShallowNeuralClassifier
import utils
import warnings
utils.fix_random_seeds()
# +
GLOVE_HOME = os.path.join('data', 'glove.6B')
DATA_HOME = os.path.join("data", "nlidata")
SNLI_HOME = os.path.join(DATA_HOME, "snli_1.0")
MULTINLI_HOME = os.path.join(DATA_HOME, "multinli_1.0")
ANNOTATIONS_HOME = os.path.join(DATA_HOME, "multinli_1.0_annotations")
# -
# ## Sparse feature representations
#
# We begin by looking at models based in sparse, hand-built feature representations. As in earlier units of the course, we will see that __these models are competitive__: easy to design, fast to optimize, and highly effective.
# ### Feature representations
#
# The guiding idea for NLI sparse features is that one wants to knit together the premise and hypothesis, so that the model can learn about their relationships rather than just about each part separately.
# With `word_overlap_phi`, we just get the set of words that occur in both the premise and hypothesis.
def word_overlap_phi(t1, t2):
"""
Basis for features for the words in both the premise and hypothesis.
Downcases all words.
Parameters
----------
t1, t2 : `nltk.tree.Tree`
As given by `str2tree`.
Returns
-------
defaultdict
Maps each word in both `t1` and `t2` to 1.
"""
words1 = {w.lower() for w in t1.leaves()}
words2 = {w.lower() for w in t2.leaves()}
return Counter(words1 & words2)
# With `word_cross_product_phi`, we count all the pairs $(w_{1}, w_{2})$ where $w_{1}$ is a word from the premise and $w_{2}$ is a word from the hypothesis. This creates a very large feature space. These models are very strong right out of the box, and they can be supplemented with more fine-grained features.
def word_cross_product_phi(t1, t2):
"""
Basis for cross-product features. Downcases all words.
Parameters
----------
t1, t2 : `nltk.tree.Tree`
As given by `str2tree`.
Returns
-------
defaultdict
Maps each (w1, w2) in the cross-product of `t1.leaves()` and
`t2.leaves()` (both downcased) to its count. This is a
multi-set cross-product (repetitions matter).
"""
words1 = [w.lower() for w in t1.leaves()]
words2 = [w.lower() for w in t2.leaves()]
return Counter([(w1, w2) for w1, w2 in product(words1, words2)])
# ### Model wrapper for hyperparameter search
#
# Our experiment framework is basically the same as the one we used for the Stanford Sentiment Treebank.
#
# For a full evaluation, we would like to search for the best hyperparameters. However, SNLI is very large, so each evaluation is very expensive. To try to keep this under control, we can set the optimizer to do just a few epochs of training during the search phase. The assumption here is that the best parameters actually emerge as best early in the process. This is by no means guaranteed, but it seems like a good way to balance doing serious hyperparameter search with the costs of doing dozens or even thousands of experiments. (See also [the discussion of hyperparameter search in the evaluation methods notebook](evaluation_methods.ipynb#Hyperparameter-optimization).)
def fit_softmax_with_hyperparameter_search(X, y):
"""
A MaxEnt model of dataset with hyperparameter cross-validation.
Parameters
----------
X : 2d np.array
The matrix of features, one example per row.
y : list
The list of labels for rows in `X`.
Returns
-------
sklearn.linear_model.LogisticRegression
A trained model instance, the best model found.
"""
mod = LogisticRegression(
fit_intercept=True,
max_iter=3, ## A small number of iterations.
solver='liblinear',
multi_class='ovr')
param_grid = {
'C': [0.4, 0.6, 0.8, 1.0],
'penalty': ['l1','l2']}
with warnings.catch_warnings():
warnings.simplefilter("ignore")
bestmod = utils.fit_classifier_with_hyperparameter_search(
X, y, mod, param_grid=param_grid, cv=3)
return bestmod
# ### Assessment
# %%time
word_cross_product_experiment_xval = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=word_cross_product_phi,
train_func=fit_softmax_with_hyperparameter_search,
assess_reader=None,
verbose=False)
optimized_word_cross_product_model = word_cross_product_experiment_xval['model']
# `word_cross_product_experiment_xval` consumes a lot of memory, and we
# won't make use of it outside of the model, so we can remove it now.
del word_cross_product_experiment_xval
def fit_optimized_word_cross_product(X, y):
optimized_word_cross_product_model.max_iter = 1000 # To convergence in this phase!
optimized_word_cross_product_model.fit(X, y)
return optimized_word_cross_product_model
# %%time
_ = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=word_cross_product_phi,
train_func=fit_optimized_word_cross_product,
assess_reader=nli.SNLIDevReader(SNLI_HOME))
# As expected `word_cross_product_phi` is reasonably strong. This model is similar to (a simplified version of) the baseline "Lexicalized Classifier" in [the original SNLI paper by Bowman et al.](https://www.aclweb.org/anthology/D15-1075/).
# ## Hypothesis-only baselines
# In an outstanding project for this course in 2016, [<NAME>](https://leonidk.com) observed that [one can do much better than chance on SNLI by processing only the hypothesis](https://leonidk.com/stanford/cs224u.html). This relates to [observations we made in the word-level homework/bake-off](hw_wordentail.ipynb) about how certain terms will tend to appear more on the right in entailment pairs than on the left. In 2018, a number of groups independently (re-)discovered this fact and published analyses: [Poliak et al. 2018](https://www.aclweb.org/anthology/S18-2023/), [Tsuchiya 2018](https://www.aclweb.org/anthology/L18-1239/), [Gururangan et al. 2018](https://www.aclweb.org/anthology/N18-2017/). Let's build on this insight by fitting a hypothesis-only model that seems comparable to the cross-product-based model we just looked at:
def hypothesis_only_unigrams_phi(t1, t2):
return Counter(t2.leaves())
def fit_softmax(X, y):
mod = LogisticRegression(
fit_intercept=True,
solver='liblinear',
multi_class='ovr')
mod.fit(X, y)
return mod
# %%time
_ = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=hypothesis_only_unigrams_phi,
train_func=fit_softmax,
assess_reader=nli.SNLIDevReader(SNLI_HOME))
# Chance performance on SNLI is 0.33 accuracy/F1. The above makes it clear that using chance as a baseline will overstate how much traction a model has actually gotten on the SNLI problem. The hypothesis-only baseline is better for this kind of calibration.
#
# Ideally, for each model one explores, one would fit a minimally different hypothesis-only model as a baseline. To avoid undue complexity, I won't do that here, but we will use the above results to provide informal context, and I will sketch reasonable hypothesis-only baselines for each model we consider.
# ## Sentence-encoding models
#
# We turn now to sentence-encoding models. The hallmark of these is that the premise and hypothesis get their own representation in some sense, and then those representations are combined to predict the label. [Bowman et al. 2015](http://aclweb.org/anthology/D/D15/D15-1075.pdf) explore models of this form as part of introducing SNLI.
# ### Dense representations
#
# Perhaps the simplest sentence-encoding model sums (or averages, etc.) the word representations for the premise, does the same for the hypothesis, and concatenates those two representations for use as the input to a linear classifier.
#
# Here's a diagram that is meant to suggest the full space of models of this form:
#
# <img src="fig/nli-softmax.png" width=800 />
# Here's an implementation of this model where
#
# * The embedding is GloVe.
# * The word representations are summed.
# * The premise and hypothesis vectors are concatenated.
# * A softmax classifier is used at the top.
glove_lookup = utils.glove2dict(
os.path.join(GLOVE_HOME, 'glove.6B.300d.txt'))
# +
def glove_leaves_phi(t1, t2, np_func=np.mean):
"""
Represent `t1` and `t2 as a combination of the vector of their words,
and concatenate these two combinator vectors.
Parameters
----------
t1 : nltk.Tree
t2 : nltk.Tree
np_func : function
A numpy matrix operation that can be applied columnwise,
like `np.mean`, `np.sum`, or `np.prod`. The requirement is that
the function take `axis=0` as one of its arguments (to ensure
columnwise combination) and that it return a vector of a
fixed length, no matter what the size of the tree is.
Returns
-------
np.array
"""
prem_vecs = _get_tree_vecs(t1, glove_lookup, np_func)
hyp_vecs = _get_tree_vecs(t2, glove_lookup, np_func)
return np.concatenate((prem_vecs, hyp_vecs))
def _get_tree_vecs(tree, lookup, np_func):
allvecs = np.array([lookup[w] for w in tree.leaves() if w in lookup])
if len(allvecs) == 0:
dim = len(next(iter(lookup.values())))
feats = np.zeros(dim)
else:
feats = np_func(allvecs, axis=0)
return feats
# -
# %%time
_ = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=glove_leaves_phi,
train_func=fit_softmax_with_hyperparameter_search,
assess_reader=nli.SNLIDevReader(SNLI_HOME),
vectorize=False) # Ask `experiment` not to featurize; we did it already.
# The hypothesis-only counterpart of this model is very clear: we would just encode `t2` with GloVe, leaving `t1` out entirely.
# As an elaboration of this approach, it is worth considering the `VecAvg` model we studied in [sst_03_neural_networks.ipynb](#sst_03_neural_networks.ipynb#The-VecAvg-baseline-from-Socher-et-al.-2013), which updates the initial vector representations during learning.
# ### Sentence-encoding RNNs
#
# A more sophisticated sentence-encoding model processes the premise and hypothesis with separate RNNs and uses the concatenation of their final states as the basis for the classification decision at the top:
#
# <img src="fig/nli-rnn-sentencerep.png" width=800 />
# It is relatively straightforward to extend `torch_rnn_classifier` so that it can handle this architecture:
# #### A sentence-encoding dataset
#
# Whereas `torch_rnn_classifier.TorchRNNDataset` creates batches that consist of `(sequence, sequence_length, label)` triples, the sentence encoding model requires us to double the first two components. The most important features of this is `collate_fn`, which determines what the batches look like:
class TorchRNNSentenceEncoderDataset(torch.utils.data.Dataset):
def __init__(self, prem_seqs, hyp_seqs, prem_lengths, hyp_lengths, y=None):
self.prem_seqs = prem_seqs
self.hyp_seqs = hyp_seqs
self.prem_lengths = prem_lengths
self.hyp_lengths = hyp_lengths
self.y = y
assert len(self.prem_seqs) == len(self.hyp_seqs)
assert len(self.hyp_seqs) == len(self.prem_lengths)
assert len(self.prem_lengths) == len(self.hyp_lengths)
if self.y is not None:
assert len(self.hyp_lengths) == len(self.y)
@staticmethod
def collate_fn(batch):
batch = list(zip(*batch))
X_prem = torch.nn.utils.rnn.pad_sequence(batch[0], batch_first=True)
X_hyp = torch.nn.utils.rnn.pad_sequence(batch[1], batch_first=True)
prem_lengths = torch.tensor(batch[2])
hyp_lengths = torch.tensor(batch[3])
if len(batch) == 5:
y = torch.tensor(batch[4])
return X_prem, X_hyp, prem_lengths, hyp_lengths, y
else:
return X_prem, X_hyp, prem_lengths, hyp_lengths
def __len__(self):
return len(self.prem_seqs)
def __getitem__(self, idx):
if self.y is None:
return (self.prem_seqs[idx], self.hyp_seqs[idx],
self.prem_lengths[idx], self.hyp_lengths[idx])
else:
return (self.prem_seqs[idx], self.hyp_seqs[idx],
self.prem_lengths[idx], self.hyp_lengths[idx],
self.y[idx])
# #### A sentence-encoding model
#
# With `TorchRNNSentenceEncoderClassifierModel`, we create a new `nn.Module` that functions just like the existing `torch_rnn_classifier.TorchRNNClassifierModel`, except that it takes two RNN instances as arguments and combines their final output states to create the classifier input:
class TorchRNNSentenceEncoderClassifierModel(nn.Module):
def __init__(self, prem_rnn, hyp_rnn, output_dim):
super().__init__()
self.prem_rnn = prem_rnn
self.hyp_rnn = hyp_rnn
self.output_dim = output_dim
self.bidirectional = self.prem_rnn.bidirectional
# Doubled because we concatenate the final states of
# the premise and hypothesis RNNs:
self.classifier_dim = self.prem_rnn.hidden_dim * 2
# Bidirectionality doubles it again:
if self.bidirectional:
self.classifier_dim *= 2
self.classifier_layer = nn.Linear(
self.classifier_dim, self.output_dim)
def forward(self, X_prem, X_hyp, prem_lengths, hyp_lengths):
# Premise:
_, prem_state = self.prem_rnn(X_prem, prem_lengths)
prem_state = self.get_batch_final_states(prem_state)
# Hypothesis:
_, hyp_state = self.hyp_rnn(X_hyp, hyp_lengths)
hyp_state = self.get_batch_final_states(hyp_state)
# Final combination:
state = torch.cat((prem_state, hyp_state), dim=1)
# Classifier layer:
logits = self.classifier_layer(state)
return logits
def get_batch_final_states(self, state):
if self.prem_rnn.rnn.__class__.__name__ == 'LSTM':
state = state[0].squeeze(0)
else:
state = state.squeeze(0)
if self.bidirectional:
state = torch.cat((state[0], state[1]), dim=1)
return state
# #### A sentence-encoding model interface
#
# Finally, we subclass `TorchRNNClassifier`. Here, just need to redefine three methods: `build_dataset` and `build_graph` to make use of the new components above:
class TorchRNNSentenceEncoderClassifier(TorchRNNClassifier):
def build_dataset(self, X, y=None):
X_prem, X_hyp = zip(*X)
X_prem, prem_lengths = self._prepare_sequences(X_prem)
X_hyp, hyp_lengths = self._prepare_sequences(X_hyp)
if y is None:
return TorchRNNSentenceEncoderDataset(
X_prem, X_hyp, prem_lengths, hyp_lengths)
else:
self.classes_ = sorted(set(y))
self.n_classes_ = len(self.classes_)
class2index = dict(zip(self.classes_, range(self.n_classes_)))
y = [class2index[label] for label in y]
return TorchRNNSentenceEncoderDataset(
X_prem, X_hyp, prem_lengths, hyp_lengths, y)
def build_graph(self):
prem_rnn = TorchRNNModel(
vocab_size=len(self.vocab),
embedding=self.embedding,
use_embedding=self.use_embedding,
embed_dim=self.embed_dim,
rnn_cell_class=self.rnn_cell_class,
hidden_dim=self.hidden_dim,
bidirectional=self.bidirectional,
freeze_embedding=self.freeze_embedding)
hyp_rnn = TorchRNNModel(
vocab_size=len(self.vocab),
embedding=prem_rnn.embedding, # Same embedding for both RNNs.
use_embedding=self.use_embedding,
embed_dim=self.embed_dim,
rnn_cell_class=self.rnn_cell_class,
hidden_dim=self.hidden_dim,
bidirectional=self.bidirectional,
freeze_embedding=self.freeze_embedding)
model = TorchRNNSentenceEncoderClassifierModel(
prem_rnn, hyp_rnn, output_dim=self.n_classes_)
self.embed_dim = prem_rnn.embed_dim
return model
# #### Simple example
#
# This toy problem illustrates how this works in detail:
def simple_example():
vocab = ['a', 'b', '$UNK']
# Reversals are good, and other pairs are bad:
train = [
[(list('ab'), list('ba')), 'good'],
[(list('aab'), list('baa')), 'good'],
[(list('abb'), list('bba')), 'good'],
[(list('aabb'), list('bbaa')), 'good'],
[(list('ba'), list('ba')), 'bad'],
[(list('baa'), list('baa')), 'bad'],
[(list('bba'), list('bab')), 'bad'],
[(list('bbaa'), list('bbab')), 'bad'],
[(list('aba'), list('bab')), 'bad']]
test = [
[(list('baaa'), list('aabb')), 'bad'],
[(list('abaa'), list('baaa')), 'bad'],
[(list('bbaa'), list('bbaa')), 'bad'],
[(list('aaab'), list('baaa')), 'good'],
[(list('aaabb'), list('bbaaa')), 'good']]
mod = TorchRNNSentenceEncoderClassifier(
vocab,
max_iter=1000,
embed_dim=10,
bidirectional=True,
hidden_dim=10)
X, y = zip(*train)
mod.fit(X, y)
X_test, y_test = zip(*test)
preds = mod.predict(X_test)
print("\nPredictions:")
for ex, pred, gold in zip(X_test, preds, y_test):
score = "correct" if pred == gold else "incorrect"
print("{0:>6} {1:>6} - predicted: {2:>4}; actual: {3:>4} - {4}".format(
"".join(ex[0]), "".join(ex[1]), pred, gold, score))
simple_example()
# #### Example SNLI run
def sentence_encoding_rnn_phi(t1, t2):
"""Map `t1` and `t2` to a pair of lists of leaf nodes."""
return (t1.leaves(), t2.leaves())
def get_sentence_encoding_vocab(X, n_words=None, mincount=1):
wc = Counter([w for pair in X for ex in pair for w in ex])
wc = wc.most_common(n_words) if n_words else wc.items()
if mincount > 1:
wc = {(w, c) for w, c in wc if c >= mincount}
vocab = {w for w, c in wc}
vocab.add("$UNK")
return sorted(vocab)
def fit_simple_sentence_encoding_rnn_with_hyperparameter_search(X, y):
vocab = get_sentence_encoding_vocab(X, mincount=2)
mod = TorchRNNSentenceEncoderClassifier(
vocab,
hidden_dim=300,
embed_dim=300,
bidirectional=True,
early_stopping=True,
max_iter=1)
param_grid = {
'batch_size': [32, 64, 128, 256],
'eta': [0.0001, 0.001, 0.01]}
bestmod = utils.fit_classifier_with_hyperparameter_search(
X, y, mod, cv=3, param_grid=param_grid)
return bestmod
# %%time
sentence_encoder_rnn_experiment_xval = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=sentence_encoding_rnn_phi,
train_func=fit_simple_sentence_encoding_rnn_with_hyperparameter_search,
assess_reader=None,
vectorize=False)
optimized_sentence_encoding_rnn = sentence_encoder_rnn_experiment_xval['model']
# Remove unneeded experimental data:
del sentence_encoder_rnn_experiment_xval
def fit_optimized_sentence_encoding_rnn(X, y):
optimized_sentence_encoding_rnn.max_iter = 1000 # Give early_stopping time!
optimized_sentence_encoding_rnn.fit(X, y)
return optimized_sentence_encoding_rnn
# %%time
_ = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=sentence_encoding_rnn_phi,
train_func=fit_optimized_sentence_encoding_rnn,
assess_reader=nli.SNLIDevReader(SNLI_HOME),
vectorize=False)
# This is above our general hypothesis-only baseline ($\approx$0.65), but it is below the simpler word cross-product model ($\approx$0.75).
#
# A natural hypothesis-only baseline for this model be a simple `TorchRNNClassifier` that processed only the hypothesis.
# ### Other sentence-encoding model ideas
#
# Given that [we already explored tree-structured neural networks (TreeNNs)](sst_03_neural_networks.ipynb#Tree-structured-neural-networks), it's natural to consider these as the basis for sentence-encoding NLI models:
#
# <img src="fig/nli-treenn.png" width=800 />
#
# And this is just the begnning: any model used to represent sentences is presumably a candidate for use in sentence-encoding NLI!
# ## Chained models
#
# The final major class of NLI designs we look at are those in which the premise and hypothesis are processed sequentially, as a pair. These don't deliver representations of the premise or hypothesis separately. They bear the strongest resemblance to classic sequence-to-sequence models.
# ### Simple RNN
#
# In the simplest version of this model, we just concatenate the premise and hypothesis. The model itself is identical to the one we used for the Stanford Sentiment Treebank:
#
# <img src="fig/nli-rnn-chained.png" width=800 />
# To implement this, we can use `TorchRNNClassifier` out of the box. We just need to concatenate the leaves of the premise and hypothesis trees:
def simple_chained_rep_rnn_phi(t1, t2):
"""Map `t1` and `t2` to a single list of leaf nodes.
A slight variant might insert a designated boundary symbol between
the premise leaves and the hypothesis leaves. Be sure to add it to
the vocab in that case, else it will be $UNK.
"""
return t1.leaves() + t2.leaves()
def fit_simple_chained_rnn_with_hyperparameter_search(X, y):
vocab = utils.get_vocab(X, mincount=2)
mod = TorchRNNClassifier(
vocab,
hidden_dim=300,
embed_dim=300,
bidirectional=True,
early_stopping=True,
max_iter=1)
param_grid = {
'batch_size': [32, 64, 128, 256],
'eta': [0.0001, 0.001, 0.01]}
bestmod = utils.fit_classifier_with_hyperparameter_search(
X, y, mod, cv=3, param_grid=param_grid)
return bestmod
# %%time
chained_rnn_experiment_xval = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=simple_chained_rep_rnn_phi,
train_func=fit_simple_chained_rnn_with_hyperparameter_search,
assess_reader=None,
vectorize=False)
optimized_chained_rnn = chained_rnn_experiment_xval['model']
del chained_rnn_experiment_xval
def fit_optimized_simple_chained_rnn(X, y):
optimized_chained_rnn.max_iter = 1000
optimized_chained_rnn.fit(X, y)
return optimized_chained_rnn
# %%time
_ = nli.experiment(
train_reader=nli.SNLITrainReader(SNLI_HOME),
phi=simple_chained_rep_rnn_phi,
train_func=fit_optimized_simple_chained_rnn,
assess_reader=nli.SNLIDevReader(SNLI_HOME),
vectorize=False)
# This model is close to the word cross-product baseline ($\approx$0.75), but it's not better. Perhaps using a GloVe embedding would suffice to push it into the lead.
#
# The hypothesis-only baseline for this model is very simple: we just use the same model, but we process only the hypothesis.
# ### Separate premise and hypothesis RNNs
#
# A natural variation on the above is to give the premise and hypothesis each their own RNN:
#
# <img src="fig/nli-rnn-chained-separate.png" width=800 />
#
# This greatly increases the number of parameters, but it gives the model more chances to learn that appearing in the premise is different from appearing in the hypothesis. One could even push this idea further by giving the premise and hypothesis their own embeddings as well. This could take the form of a simple modification to [the sentence-encoder version defined above](#Sentence-encoding-RNNs).
# ## Attention mechanisms
#
# Many of the best-performing systems in [the SNLI leaderboard](https://nlp.stanford.edu/projects/snli/) use __attention mechanisms__ to help the model learn important associations between words in the premise and words in the hypothesis. I believe [Rocktäschel et al. (2015)](https://arxiv.org/pdf/1509.06664v1.pdf) were the first to explore such models for NLI.
#
# For instance, if _puppy_ appears in the premise and _dog_ in the conclusion, then that might be a high-precision indicator that the correct relationship is entailment.
#
# This diagram is a high-level schematic for adding attention mechanisms to a chained RNN model for NLI:
#
# <img src="fig/nli-rnn-attention.png" width=800 />
#
# Since PyTorch will handle the details of backpropagation, implementing these models is largely reduced to figuring out how to wrangle the states of the model in the desired way.
# ## Error analysis with the MultiNLI annotations
#
# The annotations included with the MultiNLI corpus create some powerful yet easy opportunities for error analysis right out of the box. This section illustrates how to make use of them with models you've trained.
#
# First, we train a chained RNN model on a sample of the MultiNLI data, just for illustrative purposes. To save time, we'll carry over the optimal model we used above for SNLI. (For a real experiment, of course, we would want to conduct the hyperparameter search again, since MultiNLI is very different from SNLI.)
rnn_multinli_experiment = nli.experiment(
train_reader=nli.MultiNLITrainReader(MULTINLI_HOME),
phi=simple_chained_rep_rnn_phi,
train_func=fit_optimized_simple_chained_rnn,
assess_reader=None,
random_state=42,
vectorize=False)
# The return value of `nli.experiment` contains the information we need to make predictions on new examples.
#
# Next, we load in the 'matched' condition annotations ('mismatched' would work as well):
matched_ann_filename = os.path.join(
ANNOTATIONS_HOME,
"multinli_1.0_matched_annotations.txt")
matched_ann = nli.read_annotated_subset(
matched_ann_filename, MULTINLI_HOME)
# The following function uses `rnn_multinli_experiment` to make predictions on annotated examples, and harvests some other information that is useful for error analysis:
def predict_annotated_example(ann, experiment_results):
model = experiment_results['model']
phi = experiment_results['phi']
ex = ann['example']
prem = ex.sentence1_parse
hyp = ex.sentence2_parse
feats = phi(prem, hyp)
pred = model.predict([feats])[0]
gold = ex.gold_label
data = {cat: True for cat in ann['annotations']}
data.update({'gold': gold, 'prediction': pred, 'correct': gold == pred})
return data
# Finally, this function applies `predict_annotated_example` to a collection of annotated examples and puts the results in a `pd.DataFrame` for flexible analysis:
def get_predictions_for_annotated_data(anns, experiment_results):
data = []
for ex_id, ann in anns.items():
results = predict_annotated_example(ann, experiment_results)
data.append(results)
return pd.DataFrame(data)
ann_analysis_df = get_predictions_for_annotated_data(
matched_ann, rnn_multinli_experiment)
# With `ann_analysis_df`, we can see how the model does on individual annotation categories:
pd.crosstab(ann_analysis_df['correct'], ann_analysis_df['#MODAL'])
|
nli_02_models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''3.9.5'': pyenv)'
# name: python3
# ---
# # Quickstart Example with Off-Policy Learners
# ---
# This notebook provides an example of implementing several off-policy learning methods with synthetic logged bandit data.
#
# The example consists of the follwoing four major steps:
# - (1) Generating Synthetic Data
# - (2) Off-Policy Learning
# - (3) Evaluation of Off-Policy Learners
#
# Please see [../examples/opl](../opl) for a more sophisticated example of the evaluation of off-policy learners with synthetic bandit data.
# +
# needed when using Google Colab
# # !pip install obp
# +
from sklearn.ensemble import RandomForestClassifier as RandomForest
from sklearn.linear_model import LogisticRegression
# import open bandit pipeline (obp)
import obp
from obp.dataset import (
SyntheticBanditDataset,
logistic_reward_function,
linear_reward_function,
linear_behavior_policy
)
from obp.policy import (
IPWLearner,
NNPolicyLearner,
Random
)
# -
# obp version
print(obp.__version__)
# ## (1) Generating Synthetic Data
# `obp.dataset.SyntheticBanditDataset` is an easy-to-use synthetic data generator.
#
# It takes
# - number of actions (`n_actions`, $|\mathcal{A}|$)
# - dimension of context vectors (`dim_context`, $d$)
# - reward function (`reward_function`, $q(x,a)=\mathbb{E}[r \mid x,a]$)
# - behavior policy (`behavior_policy_function`, $\pi_b(a|x)$)
#
# as inputs and generates a synthetic logged bandit data that can be used to evaluate the performance of decision making policies (obtained by `off-policy learning`).
# + tags=[]
# generate a synthetic bandit dataset with 10 actions
# we use `logistic function` as the reward function and `linear_behavior_policy` as the behavior policy.
# one can define their own reward function and behavior policy such as nonlinear ones.
dataset = SyntheticBanditDataset(
n_actions=10,
dim_context=5,
tau=0.2, # temperature hyperparameter to control the entropy of the behavior policy
reward_type="binary", # "binary" or "continuous"
reward_function=logistic_reward_function,
behavior_policy_function=linear_behavior_policy,
random_state=12345,
)
# -
# obtain training and test sets of synthetic logged bandit data
n_rounds_train, n_rounds_test = 10000, 10000
bandit_feedback_train = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_train)
bandit_feedback_test = dataset.obtain_batch_bandit_feedback(n_rounds=n_rounds_test)
# the logged bandit data is collected by the behavior policy as follows.
#
# $ \mathcal{D}_b := \{(x_i,a_i,r_i)\}_{i=1}^n$ where $(x,a,r) \sim p(x)\pi_b(a \mid x)p(r \mid x,a) $
# `bandit_feedback` is a dictionary storing synthetic logged bandit feedback
bandit_feedback_train
# ## (2) Off-Policy Learning
# After generating synthetic data, we now train some decision making policies.
#
# To train policies, we use
#
# - `obp.policy.NNPolicyLearner` (Neural Network Policy Learner)
# - `obp.policy.IPWLearner`
#
# For NN Learner, we use
# - Direct Method ("dm")
# - InverseProbabilityWeighting ("ipw")
# - DoublyRobust ("dr")
#
# as its objective functions (`off_policy_objective`).
#
# For IPW Learner, we use *RandomForestClassifier* and *LogisticRegression* implemented in scikit-learn for base machine learning methods.
# A policy is trained by maximizing an OPE estimator as an objective function as follows.
#
# $$ \hat{\pi} \in \arg \max_{\pi \in \Pi} \hat{V} (\pi; \mathcal{D}_{tr}) - \lambda \cdot \Omega (\pi) $$
#
# where $\hat{V}(\cdot; \mathcal{D})$ is an off-policy objective and $\mathcal{D}_{tr}$ is a training bandit dataset. $\Omega (\cdot)$ is a regularization term.
# +
# define NNPolicyLearner with DM as its objective function
nn_dm = NNPolicyLearner(
n_actions=dataset.n_actions,
dim_context=dataset.dim_context,
off_policy_objective="dm",
batch_size=64,
random_state=12345,
)
# train NNPolicyLearner on the training set of logged bandit data
nn_dm.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
)
# obtains action choice probabilities for the test set
action_dist_nn_dm = nn_dm.predict_proba(
context=bandit_feedback_test["context"]
)
# +
# define NNPolicyLearner with IPW as its objective function
nn_ipw = NNPolicyLearner(
n_actions=dataset.n_actions,
dim_context=dataset.dim_context,
off_policy_objective="ipw",
batch_size=64,
random_state=12345,
)
# train NNPolicyLearner on the training set of logged bandit data
nn_ipw.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"],
)
# obtains action choice probabilities for the test set
action_dist_nn_ipw = nn_ipw.predict_proba(
context=bandit_feedback_test["context"]
)
# +
# define NNPolicyLearner with DR as its objective function
nn_dr = NNPolicyLearner(
n_actions=dataset.n_actions,
dim_context=dataset.dim_context,
off_policy_objective="dr",
batch_size=64,
random_state=12345,
)
# train NNPolicyLearner on the training set of logged bandit data
nn_dr.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"],
)
# obtains action choice probabilities for the test set
action_dist_nn_dr = nn_dr.predict_proba(
context=bandit_feedback_test["context"]
)
# + tags=[]
# define IPWLearner with Logistic Regression as its base ML model
ipw_lr = IPWLearner(
n_actions=dataset.n_actions,
base_classifier=LogisticRegression(C=100, random_state=12345)
)
# train IPWLearner on the training set of logged bandit data
ipw_lr.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set
action_dist_ipw_lr = ipw_lr.predict(
context=bandit_feedback_test["context"]
)
# + tags=[]
# define IPWLearner with Random Forest as its base ML model
ipw_rf = IPWLearner(
n_actions=dataset.n_actions,
base_classifier=RandomForest(
n_estimators=30, min_samples_leaf=10, random_state=12345
)
)
# train IPWLearner on the training set of logged bandit data
ipw_rf.fit(
context=bandit_feedback_train["context"],
action=bandit_feedback_train["action"],
reward=bandit_feedback_train["reward"],
pscore=bandit_feedback_train["pscore"]
)
# obtains action choice probabilities for the test set
action_dist_ipw_rf = ipw_rf.predict(
context=bandit_feedback_test["context"]
)
# +
# define Uniform Random Policy as a baseline evaluation policy
random = Random(n_actions=dataset.n_actions,)
# compute the action choice probabilities for the test set
action_dist_random = random.compute_batch_action_dist(
n_rounds=bandit_feedback_test["n_rounds"]
)
# -
# action_dist is a probability distribution over actions (can be deterministic)
action_dist_ipw_lr[:, :, 0]
# ## (3) Evaluation of Off-Policy Learners
# Our final step is the evaluation and comparison of the off-policy learnres.
#
# With synthetic data, we can calculate the policy value of the off-policy learners as follows.
#
# $$V(\pi_e) \approx \frac{1}{|\mathcal{D}_{te}|} \sum_{i=1}^{|\mathcal{D}_{te}|} \mathbb{E}_{a \sim \pi_e(a|x_i)} [q(x_i, a)], \; \, where \; \, q(x,a) := \mathbb{E}_{r \sim p(r|x,a)} [r]$$
#
# where $\mathcal{D}_{te}$ is the test set of logged bandit data.
# + tags=[]
# we calculate the policy values of the trained policies based on the expected rewards of the test data
policy_names = [
"NN Policy Learner with DM",
"NN Policy Learner with IPW",
"NN Policy Learner with DR",
"IPW Learner with Logistic Regression",
"IPW Learner with Random Forest",
"Unifrom Random"
]
action_dist_list = [
action_dist_nn_dm,
action_dist_nn_ipw,
action_dist_nn_dr,
action_dist_ipw_lr,
action_dist_ipw_rf,
action_dist_random
]
for name, action_dist in zip(policy_names, action_dist_list):
true_policy_value = dataset.calc_ground_truth_policy_value(
expected_reward=bandit_feedback_test["expected_reward"],
action_dist=action_dist,
)
print(f'policy value of {name}: {true_policy_value}')
# -
# In fact, NNPolicyLearner maximizing the DM estimator seems the best in this simple setting.
# We can iterate the above process several times to get more relibale results.
#
# Please see [../examples/opl](../opl) for a more sophisticated example of the evaluation of off-policy learners with synthetic bandit data.
|
examples/quickstart/opl.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ekanshchopda/KalpanaLabs/blob/master/Day%208/Copy_of_KalpanaLabs_Day8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="jhXx96TcYBd4"
# # if-else conditions
#
# ---
#
# :
# statements
# -
# -
# elif (condition):
# statements
# -
# -
# elif (condition):
# statements
# -
# -
# else
# statements
# -
# -
# ```
#
# - What does if mean?
# - What does elif mean?
# - What does else mean?
# + [markdown] id="jkkkAsa9nNt8"
# ## Example
# ----
#
# You play games everyday downstairs. You need your parent's permission, right?
#
# Mostly they allow.
#
# But recently it has begun to rain. Now, will they allow?
# + id="4aNaNkG_G145" outputId="85086f16-e68a-4de9-9e11-62baeb3fcb02" colab={"base_uri": "https://localhost:8080/", "height": 34}
itIsRaining = True
if itIsRaining:
print("You cannot go downstairs. Study at home!")
else:
print("Yes, go downstairs. Be back by around 7 PM")
# + [markdown] id="zu2sVtUspL7o"
# ## Example 2
# ---
#
# Let us consider the marking scheme in schools.
#
# | Marks | Grade |
# | --- | ----------- |
# | 0 - 40 | F |
# | 41 -50 | C |
# | 51 -60 | C+ |
# | 61 -70 | B |
# | 71 -80 | B+ |
# | 81 -90 | A |
# | 91 -100 | A+ |
#
# Now, let us creating a grading system!
# + id="p_BTr4qEpBRZ" outputId="8e6466ec-82c3-4cb1-b0a4-ad84e14537b2" colab={"base_uri": "https://localhost:8080/", "height": 34}
marks =
if marks < 41:
grade = "F"
elif marks < 51:
grade = "C"
elif marks < 61:
grade = "C+"
elif marks < 71:
grade = "B"
elif marks < 81:
grade = "B+"
elif marks < 91:
grade = "A"
else:
grade = "A+"
print("Your grade is " + grade)
# + [markdown] id="6j-gS-mdrHIp"
# Another way of writing the same code
# + id="liTlFwarqiSv" outputId="b558d590-1ce2-4027-96d4-8c7f74e1bcb7" colab={"base_uri": "https://localhost:8080/", "height": 34}
marks = 9
if marks < 101 and marks > 89:
grade = "A+"
elif marks < 90 and marks > 79:
grade = "A"
elif marks < 80 and marks > 69:
grade = "B+"
elif marks < 70 and marks > 59:
grade = "B"
elif marks < 60 and marks > 49:
grade = "C+"
elif marks < 50 and marks > 39:
grade = "C"
else:
grade = "F"
print("Your grade is " + grade)
# + id="7HJ7pQJUsAKv"
|
Day 8/Copy_of_KalpanaLabs_Day8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Getting started with fast.ai"
# > "How to make the most of the free deep learning course from fast.ai"
#
# - toc: true
# - branch: master
# - badges: false
# - comments: false
# - categories: [fastai, getting started, online courses]
# - image: images/articles/2012-12-09-getting-started/getting-started-fastai.png
# - hide: false
# ## Welcome!
#
# Summary: I cover some questions you might have regarding the fast.ai course. Then, I offer some advice that would have helped me in the beginning.
#
#
# ### Who is this article for?
# If you're reading this article, I assume you have heard about the terms deep learning, machine learning, or artificial intelligence. Maybe you are not too sure what they mean specifically, but you want to know more. Maybe you've been searching for a while to finally learn about these concepts, but are overwhelmed by the magnitude of courses, books and software available.
#
# Whether you are a software developer planning to add a new skillset, a student wanting to broaden your horizon, or an expert in a completely different field and you want to apply AI in your company or project: The fast.ai online course can be great for you to get started, and it's completely free!
# 
#
# ### About fast.ai
# [fast.ai](https://www.fast.ai) is an organization committed to making deep learning more accessible. It was founded by <NAME> and <NAME>, both distinguished data scientists.
#
# They offer four major elements that can help you get into deep learning.
# - [fastai](https://docs.fast.ai/) (notice the missing dot), a software library that allows creating powerful deep learning models with few lines of code, while nonetheless being very customizable.
# - A [free online course](https://course.fast.ai/) based on the library.
# - A [book](https://www.amazon.com/Deep-Learning-Coders-fastai-PyTorch/dp/1492045527) that is designed to go hand in hand with the course. It's available as a paper book, as an e-book or even for free on [Github](https://github.com/fastai/fastbook).
# - A [forum](https://forums.fast.ai/) with a very helpful community.
#
# For me, what really sets fast.ai apart from other online courses, is their practical approach:
# > We \[are\] always teaching through examples. We ensure that there is a context and a purpose that you can understand intuitively, rather than starting with algebraic symbol manipulation.
#
# This does not mean that the foundations are not covered (they are!), but the order is different than in other books or courses, where you start with basic tools and only get to usable applications at the end of the course - if ever.
# ### What is covered in the course?
#
# 
#
# The course covers major applications of deep learning. There is a certain focus on computer vision, but the other topics like tabular data, natural language processing (NLP), and recommender systems are explained as well. You will be able to create very well performing models in all of these areas, and as early as lecture 2 you can create a working web app that can recognize grizzly bears and brown bears (or anything else you choose).
#
# Later in the course, you will learn the foundations of deep learning. You'll write code for *stochastic gradient descent* and *activation functions* from scratch. Don't worry if you never have heard about these, it's explained very well. You will dig deeper into PyTorch, the underlying software of fastai.
#
# Also, there is a lecture (and a chapter) on Data Ethics. It is taught by <NAME> and will give you a lot of food for thought. I think it's great that fastai encourages you to think about the possible implications of your work early on.
# ### Should I get the book?
#
# Since the book is freely available online in the form of notebooks, you might be wondering if you should get the printed book anyway. My opinion: If you can afford it - yes, you absolutely should get it! Its layout is beautiful, which makes it easier to read than the online version. Also, I personally just like to read a physical copy: my attention span is longer and I tend to take it more seriously.
#
# That being said, you certainly can work with the free online version. It has the added benefit that you will always have the latest version, which is up to date and where bugs are fixed (and there are a few in the book).
#
# 
# ### Do I need to know Python?
#
# Yes - or any other language. The book is called "Deep Learning for Coders", so it will not explain every line of code in detail or teach Python from scratch. In fact, the [course website](https://course.fast.ai/) states:
# > The only prerequisite is that you know how to code (a year of experience is enough), preferably in Python, and that you have at least followed a high school math course.
#
# However, you don't have to be an expert. When I first started with the course, I only had a few weeks of experience with Python. I did however have some (very basic) experience with web languages like JavaScript and PHP, that helped me pick up Python pretty quickly. If you know any programming language, you won't have a problem, since Python is rather accessible and has an easy syntax. If you don't have any programming experience, I recommend investing a few weeks to learn the basics of Python before you tackle fastai. There are wonderful tutorials available for free.
#
# 
# ## I'm in! How do I begin?
# With lesson 1! 😊 You really can get started right away, the interactive content runs on ready-to-use and free platforms like Google Colab, so you don't have to spend time setting up your own machine.
#
# The [course website](https://course.fast.ai/) gives you an overview of the course (you can read it in addition to this article), then you can start with lesson 1.
#
# I enjoyed combining the videos and the book. After every lesson, I ran the associated notebooks on Colab and then read the chapters in the book. The chapters are very similar to the video lessons, but I think it helps consuming the content in a different medium and being able to go back a few pages if you want to read something again.
# ## My suggestions for efficient learning
# ### Watch the videos twice
# I recommend watching the videos twice. On the first view, don't focus on the details too much, just get an overview of the topics covered and take some notes while doing so. This is especially relevant for the later lessons that contain much more code. On the second view, you can take more detailed notes and try to get all the details.
# ### Take notes!
# Taking notes was a gamechanger for me. I think it's the best way to stay active during the videos, and over time the notes will serve as a central knowledge repository. Also, it's great to have a place to jot down your questions, so you can try and answer them later!
#
# I keep notes in Microsoft OneNote, you can of course use any other application or paper. I created a template based on this awesome [forum post](https://forums.fast.ai/t/how-to-do-fastai-study-plans-learning-strategies/39473?u=johannesstutz). The template contains following points:
#
# - **Key Points** from the lecture
# - **Advice** from Jeremy
# - **To-Do** challenges from the further research section of the chapter, ideas for projects
# - **Reading & Exploring** papers that are mentioned, stuff you find on the web but don't have time for at the moment
# - **Questions** that arise during the lecture
# - as the last point, I copy & paste the **questionnaire**.
#
# Please read the above-mentioned forum post for details on each section, it's a really good system.
#
# These notes are meant to be used not just once, but you should refine them and work with them continuously. They can serve you as your go-to resource every time you study. All your open questions, your project ideas, and of course lots of knowledge can live there.
#
# 
# ### Don't get stuck on one thing that you don't understand
# This is an important point and I think Jeremy mentions it as well. You don't have to understand every detail right away. It's often better to move on and revisit the part you didn't quite get later.
# ### Run the code
# Jeremy will ask you to do this, and you really should run the notebooks for yourself. I recommend the clean versions, where there is no text, just code. Predict what the output of a cell will be, and if you were wrong, go back and understand why. Just reading the code in the book is not enough, it can give you the illusion that you understood it, when in reality you could not reproduce it. I fell into this trap more than once...
# ### Do the questionnaire
# Take the time and answer every question in the questionnaire. This might take a while, but it makes sure that you really understood all important concepts. You can find answers to most questions in the [forum](https://forums.fast.ai/t/fastbook-questionnaire-solutions-megathread/76955?u=johannesstutz).
#
# Additionally, I can recommend [aiquizzes.com](https://aiquizzes.com/), where many questions and their answers are made available together with the relevant part of the lecture. You can use this for [spaced repetition learning](https://en.wikipedia.org/wiki/Spaced_repetition), the website will remind you when older questions need reviewing, it's great.
# ### Use other sources
# Fastai, while being excellent, is not the only source of wisdom. I often google for concepts I don't quite get. More often than not I find a blog post, a stack overflow answer or a video explaining it very well or with a different approach than Jeremy.
# ## What to do after the course
# The course covers only around half the book. There will be a part 2 that covers the more advanced chapters, but as of now (December 2020) there is no release date announced. If you want to dig deeper into the material, you should not wait for the course but work through the book on your own.
#
# In addition, I'd suggest picking a project and make it as good as you can. Go on the forums and see if you can help others, or ask for advice. Write about your journey in your own blog - like the one whose first post you are reading here 😊
# ## Thank you for reading
# Since you made it this far, I hope you found this article interesting and I could get you excited for fast.ai! I really can recommend taking the course and reading the book, and if you put in the time you will see amazing results soon.
# Let me know if you have any questions or found this article helpful. I'm on [Twitter](https://twitter.com/daflowjoe), I'd be happy to hear from you there or via mail at <<EMAIL>>
|
_notebooks/2020-12-09-Getting-started-with-fastai.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Business Understanding
#
# I am interested in using a data analysis approach to know more about the situation of Women in Computer Programming. I hope to use the analysis results to provide some useful information to everyone who need this kind of research. The key questions I would like to answer are:
#
# - what is the situation of women in Italy in the programming world compared to man?
# - what is the identikit of women who work in computer programming in Italy? What are their qualifications?
# - Instead in USA, what is the situation of super qualified women in tech ?
#
# ## Data Understanding
#
# The data used in this analysi was Stack Overflow’s developer survey data from 2017 to 2019. Respondents from about 200 countries gave their answers to about 150 survey questions. This notebook attempted to use the survey questions to answer the three questions listed in the Business Understanding section.
#
# ## Gather Data
#
# Data has been gathered by Stack Overflow survey. The following cells import necessary Python libraries, and read them into Pandas Dataframe.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pandasql import sqldf
# ## Italy situation in 2018
# ## Prepare Data
#
# The following cell help me to access Data, select the columns and the values that I need for my analysis
# ## Results for Italy in 2018
# +
stack_2018 = pd.read_csv('2018_survey_results_public.csv')
stack_2018 = stack_2018[['Country','Gender','FormalEducation','DevType','Salary']]
Italy_2018 = stack_2018.loc[stack_2018['Country']=='Italy']
#In this part I used dropna in order to drop all null value in the column Gender. For my analysis I need only to know if the gender is Male of Female#
Ita_2018 = Italy_2018.dropna(subset = ['Gender'])
Ita_18 = Ita_2018[Ita_2018['Gender'].isin(['Male','Female'])]
Ita_Dev18 = Ita_18.dropna(subset = ['DevType'])
Ita_F18 = Ita_Dev18.loc[Ita_Dev18['Gender']=='Female']
Ita_M18 = Ita_Dev18.loc[Ita_Dev18['Gender']=='Male']
# -
# ## Results for Italy in 2018
# A first overview of the total number of people in tech world based on the data of the survey#
print(Ita_Dev18.shape[0])
# A first overview of the number of female in tech world#
print(Ita_F18.shape[0])
# A first overview of the number of male in tech world#
print(Ita_M18.shape[0])
perc_Ita_F18 = Ita_F18.shape[0] / Ita_Dev18.shape[0] *100
#Percentage of women in tech world in Italy #
"{:.2f} %".format(perc_Ita_F18)
# +
func = lambda q : sqldf(q , globals())
q = """
select *
from Ita_Dev18
where FormalEducation like '%Master%' or
FormalEducation like '%Bachelor%' or
FormalEducation like '%Professional%' or
FormalEducation like '%doctoral%';
"""
Dev_OverQual18_Ita = func(q)
# +
### percentage of woman developer overqualified on number of developer women##
Dev_OverQual_F18_Ita = len(Dev_OverQual18_Ita.loc[Dev_OverQual18_Ita['Gender']=='Female'])/Ita_F18.shape[0] * 100
### percentage of man developer overqualified on number of developer man##
Dev_OverQual_M18_Ita = len(Dev_OverQual18_Ita.loc[Dev_OverQual18_Ita['Gender']=='Male'])/Ita_M18.shape[0] * 100
# -
print("{:.2f} %".format(Dev_OverQual_F18_Ita))
print("{:.2f} %".format(Dev_OverQual_M18_Ita))
####### export Ita18 outputs #######
Italy_2018.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\Italy_2018.xlsx', index=False, header=True)
Ita_2018.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaDropnaByGender_2018.xlsx', index=False, header=True)
Ita_18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaMF_18.xlsx', index=False, header=True)
Ita_Dev18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaDev_18.xlsx', index=False, header=True)
Ita_F18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaDev_F18.xlsx', index=False, header=True)
Ita_M18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaDev_M18.xlsx', index=False, header=True)
Dev_OverQual18_Ita.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Ita_18\ItaDev_OverQ18.xlsx', index=False, header=True)
# ## USA situation in 2018
# +
Usa_2018 = stack_2018.loc[stack_2018['Country']=='United States']
Usa_18 = Usa_2018.dropna(subset = ['Gender'])
USA_MF18 = Usa_18[Usa_2018['Gender'].isin(['Male','Female'])]
#In this part I used dropna in order to drop all null value in the column Gender. For my analysis I need only to know if the gender is Male of Female#
Usa_Dev18 = USA_MF18.dropna(subset = ['DevType'])
Usa_F18 = Usa_Dev18.loc[Usa_Dev18['Gender']=='Female']
Usa_M18 = Usa_Dev18.loc[Usa_Dev18['Gender']=='Male']
# -
# ## Results for USA in 2018
# A first overview of the total number of people in tech world in Usa based on the data of the survey#
print(Usa_Dev18.shape[0])
# A first overview of the number of women in tech world in Usa based on the data of the survey#
print(Usa_F18.shape[0])
# A first overview of the total number of men in tech world in Usa based on the data of the survey#
print(Usa_M18.shape[0])
perc_Usa_F18 = Usa_F18.shape[0] / Usa_Dev18.shape[0] *100
"{:.2f} %".format(perc_Usa_F18)
# +
func_1 = lambda d : sqldf(d , globals())
d = """
select *
from Usa_Dev18
where FormalEducation like '%Master%' or
FormalEducation like '%Bachelor%' or
FormalEducation like '%Professional%' or
FormalEducation like '%doctoral%';
"""
Dev_OverQual18_Usa = func_1(d)
# -
Dev_OverQual18_Usa.tail()
### percentage of women developer overqualified on number of women developer##
Dev_OverQual_F18_Usa = len(Dev_OverQual18_Usa.loc[Dev_OverQual18_Usa['Gender']=='Female'])/Usa_F18.shape[0] * 100
### percentage of men developer overqualified on number men developer##
Dev_OverQual_M18_Usa = len(Dev_OverQual18_Usa.loc[Dev_OverQual18_Usa['Gender']=='Male'])/Usa_M18.shape[0] * 100
### percentage of woman developer overqualified on number of developer women##
print("{:.2f} %".format(Dev_OverQual_F18_Usa))
### percentage of man developer overqualified on number of developer man##
print("{:.2f} %".format(Dev_OverQual_M18_Usa))
# +
########### export output Usa 2018 #########################
Usa_2018.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\Usa_2018.xlsx', index=False, header=True)
Usa_18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaDropnaByGender_2018.xlsx', index=False, header=True)
USA_MF18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaMF_18.xlsx', index=False, header=True)
Usa_Dev18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaDev_18.xlsx', index=False, header=True)
Usa_F18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaDev_F18.xlsx', index=False, header=True)
Usa_M18.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaDev_M18.xlsx', index=False, header=True)
Dev_OverQual18_Usa.to_excel(r'C:\Users\moryb\OneDrive\Desktop\Project1\output\output_18\Usa_18\UsaDev_OverQ18.xlsx', index=False, header=True)
|
2018.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started with Thinc Framework
#
# This example shows how to get started with Thinc, using the "hello world" of neural network models: recognizing handwritten digits from the [MNIST dataset](http://yann.lecun.com/exdb/mnist/).
#
#
# This notebook covers:
# - training the model
# - using config files
# - registering custom functions, and
# - wrapping models defined TensorFlow
#
# + pycharm={"is_executing": false} tags=[]
# !pip install ml_datasets "tqdm>=4.41"
# -
# `prefer_gpu` function make sure we're performing operations on GPU if it is available.
#
# The function should be called right after importing Thinc, and it returns a boolean indicating whether the GPU has been activated.
#
# If GPU is not available or not properly setup, the code will still work, just slower :)
#
# + pycharm={"is_executing": false}
from thinc.api import prefer_gpu
prefer_gpu()
# -
import matplotlib.pyplot as plt
import ml_datasets
from tqdm.notebook import tqdm
from thinc.api import Adam, fix_random_seed
# ## Some utility functions
#
# +
def train_model(data, model, optimizer, n_iter, batch_size):
(train_X, train_Y), (test_X, test_Y) = data
for i in range(n_iter):
batches = model.ops.multibatch(batch_size, train_X, train_Y, shuffle=True)
for X, Y in tqdm(batches, leave=False):
Yh, backprop = model.begin_update(X)
backprop(Yh - Y)
model.finish_update(optimizer)
# Evaluate and print progress
correct = 0
total = 0
for X, Y in model.ops.multibatch(batch_size, test_X, test_Y):
Yh = model.predict(X)
correct += (Yh.argmax(axis=1) == Y.argmax(axis=1)).sum()
total += Yh.shape[0]
score = correct / total
print(f" {i} {float(score):.3f}")
def plot_image(data, img_index):
# y_train contains the lables, ranging from 0 to 9
img = data[img_index].reshape(28, 28)
plt.imshow(img, cmap = 'gray')
# -
# # Prepare the dataset
#
# The `ml_datasets` package already packs MNIST dataset properly formated and splitted in training and test tuples.
# Thus, we only have to retrieve it and store in variables to be used.
#
# Each tuple is comprised of **features** and **target** elements, both are numpy arrays.
#
# + pycharm={"is_executing": false} tags=[]
(train_X, train_Y), (test_X, test_Y) = ml_datasets.mnist()
# + tags=[]
print(f"Training size={len(train_X)}, dev size={len(test_X)}")
print("train_X shape:", train_X.shape, "train_Y shape:", train_Y.shape)
# Print the number of training, validation, and test datasets
print(train_X.shape[0], 'train set')
print(test_X.shape[0], 'test set')
# -
# Image index, you can pick any number between 0 and 59,999
plot_image(train_X, img_index=101)
# # Model
#
# Let’s use the following network architecture:
#
# - Two Relu-activated hidden layers,
# - Add dropout after the two hidden layers (to help the model generalize better),
# - A softmax-activated output layer.
#
# The chain combinator is like `Sequential` in PyTorch or Keras: it combines a list of layers together with a feed-forward relationship.
#
# Observe that when we **defined the model**, we don’t infor the input or the output sizes. This is inferred by the framework.
#
# + pycharm={"is_executing": false}
from thinc.api import chain, Relu, Softmax
n_hidden = 32
dropout = 0.2
model = chain(
Relu(nO=n_hidden, dropout=dropout),
Relu(nO=n_hidden, dropout=dropout),
Softmax()
)
# -
# ### Visualization
#
# If you want visualize the model, run the cells in this session.
#
# Make sure you have GraphViz installed and properly configured in your system if you want to visualize the model.
#
# on MacOS:
#
# ```
# brew install graphviz
# ```
#
# + tags=[]
# !pip install pydot graphviz svgwrite
import pydot
from IPython.display import SVG, display
def get_label(layer):
layer_name = layer.name
nO = layer.get_dim("nO") if layer.has_dim("nO") else "?"
nI = layer.get_dim("nI") if layer.has_dim("nI") else "?"
return f"{layer.name}|({nI}, {nO})".replace(">", ">")
def visualize_model(model):
dot = pydot.Dot()
dot.set("rankdir", "LR")
dot.set_node_defaults(shape="record", fontname="arial", fontsize="10")
dot.set_edge_defaults(arrowsize="0.7")
nodes = {}
for i, layer in enumerate(model.layers):
label = get_label(layer)
node = pydot.Node(layer.id, label=label)
dot.add_node(node)
nodes[layer.id] = node
if i == 0:
continue
from_node = nodes[model.layers[i - 1].id]
to_node = nodes[layer.id]
if not dot.get_edge(from_node, to_node):
dot.add_edge(pydot.Edge(from_node, to_node))
display(SVG(dot.create_svg()))
# -
# Before initialization. Check the layers parameters
visualize_model(model)
# After creating the model, we can call the `Model.initialize` method, passing in a small batch of input data `X` and a small batch of output data `Y`. This allows Thinc to infer the missing dimensions.
#
# NOTE!
# We need, however, make sure the data is on the **right device** when passing in the data. This is done by calling `model.ops.asarray` which will e.g. transform the arrays to cupy when running on GPU.
# + pycharm={"is_executing": false} tags=[]
# making sure the data is on the right device,
# converting them to cupy (if we're using that)
# model.ops.xp is an instance of either numpy or cupy,
# depending on whether you run the code on CPU or GPU.
train_X = model.ops.asarray(train_X)
train_Y = model.ops.asarray(train_Y)
test_X = model.ops.asarray(test_X)
test_Y = model.ops.asarray(test_Y)
model.initialize(X=train_X[:5], Y=train_Y[:5])
nI = model.get_dim("nI")
nO = model.get_dim("nO")
print(f"Initialized model with input dimension nI={nI} and output dimension nO={nO}")
# -
# After initialization. Check the layers parameters.
# Dimention inference from data is playing here.
visualize_model(model)
# ### Training the model
#
#
# + jupyter={"outputs_hidden": true} pycharm={"is_executing": false} tags=[]
fix_random_seed(0)
optimizer = Adam(0.001)
batch_size = 128
print("Measuring performance across iterations:")
data = (train_X, train_Y), (test_X, test_Y)
train_model(data, model, optimizer, 20, batch_size)
# -
# # Define the model based on configuration
#
# Here's a config describing the model we defined above. The values in the `hyper_params` section can be referenced in other sections to keep them consistent. The `*` is used for **positional arguments** – in this case, the arguments to the `chain` function, two Relu layers and one softmax layer.
# + pycharm={"is_executing": false}
from thinc.api import Config, registry
CONFIG = """
[hyper_params]
n_hidden = 32
dropout = 0.2
learn_rate = 0.001
[model]
@layers = "chain.v1"
[model.*.relu1]
@layers = "Relu.v1"
nO = ${hyper_params:n_hidden}
dropout = ${hyper_params:dropout}
[model.*.relu2]
@layers = "Relu.v1"
nO = ${hyper_params:n_hidden}
dropout = ${hyper_params:dropout}
[model.*.softmax]
@layers = "Softmax.v1"
[optimizer]
@optimizers = "Adam.v1"
learn_rate = ${hyper_params:learn_rate}
[training]
n_iter = 10
batch_size = 128
"""
config = Config().from_str(CONFIG)
config
# -
# When you call `registry.make_from_config`, Thinc will first create the three layers using the specified arguments populated by the hyperparameters. It will then pass the return values (the layer objects) to `chain`. It will also create an optimizer. All other values, like the training config, will be passed through as a regular dict.
#
# If you want to change a hyperparameter or experiment with a different optimizer, all you need to change is the config. For each experiment you run, you can save a config and you'll be able to reproduce it later.
#
# Your training code can now look like this:
# + pycharm={"is_executing": false, "name": "#%%\n"} tags=[]
loaded_config = registry.make_from_config(config)
model = loaded_config["model"]
model.initialize(X=train_X[:5], Y=train_Y[:5])
visualize_model(model)
# + tags=[]
optimizer = loaded_config["optimizer"]
n_iter = loaded_config["training"]["n_iter"]
batch_size = loaded_config["training"]["batch_size"]
train_model(data, model, optimizer, n_iter, batch_size)
# -
|
notebooks/examples/mnist/thinc_mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# code for loading the format for the notebook
import os
# path : store the current path to convert back to it later
path = os.getcwd()
os.chdir( os.path.join('..', 'notebook_format') )
from formats import load_style
load_style(css_style = 'custom2.css')
# +
os.chdir(path)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# %matplotlib inline
# %load_ext watermark
# %load_ext autoreload
# %autoreload 2
# pip install python-dotenv
# for the dotenv package, it is
# for loading personal API key, as we'll
# see, but you don't have to use it
import json
import requests
from subprocess import call
from dotenv import load_dotenv
from IPython.display import HTML
from IPython.display import Image
from IPython.display import display
from sklearn.metrics import mean_squared_error
from sklearn.metrics import pairwise_distances
# %watermark -a 'Ethen' -d -t -v -p numpy,pandas,matplotlib,sklearn,requests
# -
# # Alternating Least Squares with Weighted Regularization
#
# Recommedantion system is a popular topic in recent years, what is does (or its goal) is to seek to predict the "rating" or "preference" that a user would give to an item. Given the rating, the business will then recommend preferable new items to the user for further purchases. The type of recommendation system that we will be discussing is known as **collaborative filtering**, where the features of the user (e.g. age, gender) or the item (e.g. perishable or not) itself do not play any role in the algorithm. Instead, we rely soley on the ratings that a user gave to the item.
#
# We start by loading some sample data to make this a bit more concrete. For this introduction, we'll be using the [MovieLens dataset](http://grouplens.org/datasets/movielens/). The website has datasets of various sizes, but we just start with the smallest one [MovieLens 100K Dataset](http://files.grouplens.org/datasets/movielens/ml-100k.zip). This dataset consists of 100,000 movie ratings by users (on a 1-5 scale). The main data set `u.data` file consists of four columns (tab-delimited), including user-id (starting at 1), item-id (starting at 1), rating, and timestamp (we won't be using this field).
# +
# download the dataset if it isn't in the same folder
file_dir = 'ml-100k'
file_path = os.path.join(file_dir, 'u.data')
if not os.path.isdir(file_dir):
call(['curl', '-O', 'http://files.grouplens.org/datasets/movielens/ml-100k.zip'])
call(['unzip', 'ml-100k.zip'])
names = ['user_id', 'item_id', 'rating', 'timestamp']
df = pd.read_csv(file_path, sep = '\t', names = names)
print(df.shape)
df.head()
# -
# As we can see, the only data that we have is how each user interacted or rated each item. Given this information, collaborative filtering will start by constructing a user-item matrix with each distinct user being the row, item being the column and the value for each cell will simply be the rating that the user gave to the item. Apart from building the matrix, we will also print out some other information to help us understand our data a bit better.
# +
# create the rating matrix r_{ui}, remember to
# subract the user and item id by 1 since
# the indices starts from 0
n_users = df['user_id'].unique().shape[0]
n_items = df['item_id'].unique().shape[0]
ratings = np.zeros((n_users, n_items))
for row in df.itertuples(index = False):
ratings[row.user_id - 1, row.item_id - 1] = row.rating
# compute the non-zero elements in the rating matrix
matrix_size = np.prod(ratings.shape)
interaction = np.flatnonzero(ratings).shape[0]
sparsity = 100 * (interaction / matrix_size)
print('dimension: ', ratings.shape)
print( 'sparsity: {:.1f}%'.format(sparsity) )
ratings
# +
# change default figure and font size
plt.rcParams['figure.figsize'] = 8, 6
plt.rcParams['font.size'] = 12
plt.hist( np.sum(ratings != 0, axis = 1), histtype = 'stepfilled', bins = 30,
alpha = 0.85, label = '# of ratings', color = '#7A68A6', normed = True )
plt.axvline(x = 10, color = 'black', linestyle = '--')
plt.legend(loc = "upper right")
plt.show()
# -
# From the information above, we know there are 943 unique users, 1682 unique items. Within the rating matrix, only 6.3% of the cells have a value, although we filled in empty ratings as 0, we should not assume these values to truly be zero. More appropriately, they are just missing entries. This kind of sparsity is extremely common in recommendation system, where people seldom give ratings to things that they have purchased. One thing to note is that if the sparsity of the matrix is below 1% (rule of thumb), then the dataset might be too sparse to perform any sort of modeling.
#
# Next the histogram tells us that every user has given at least more than 10 ratings, we will use this to perform the train/test split of the data for testing the algorithm that we'll build later.
#
# One tricky thing about splitting the data into training and testing is that: In supervise machine learning we normally build the trainining and testing holdout set by randomly splitting the rows. In those cases, this idea works, because we have a model with features/target that we are trying to fit a function to. For recommender systems with collaborative filtering (no features), this just won't work anymore, because all of the items/users need to be available when the model is first built. So what we do instead is mask a random sample of the user/item ratings to validate and compare how well the recommender system did in predicting the ratings of those masked values. In our case, given we already know each user has given more than 10 ratings, what we'll do is for every user, we remove 10 of the item ratings and and assign them to the test set. The testing matrix is printed below, as hopefully, you can see that some of the values are indeed different from the original rating matrix.
# +
def create_train_test(ratings):
"""
split into training and test sets,
remove 10 ratings from each user
and assign them to the test set
"""
test = np.zeros(ratings.shape)
train = ratings.copy()
for user in range(ratings.shape[0]):
test_index = np.random.choice( np.flatnonzero(ratings[user]),
size = 10, replace = False )
train[user, test_index] = 0.0
test[user, test_index] = ratings[user, test_index]
# assert that training and testing set are truly disjoint
assert np.all(train * test == 0)
return train, test
train, test = create_train_test(ratings)
del ratings
train
# -
# ## Matrix Factorization
#
# Now that the data preprocessing part has been taken care of, let's get to the more exciting part, the algorithm. The algorithm that we'll introduce today is Matrix Factorization.
#
# Recall that we had a user-item matrix, $R$ where nonzero elements of the matrix are ratings that a user has given an item. What Matrix Factorization does is it decomposes a large matrix into products of matrices, namely, $R = U \times V$. See the picture below taken from a quick [google search](https://www.google.com/search?q=matrix+factorization&source=lnms&tbm=isch&sa=X&ved=0ahUKEwiD8Ku02_vQAhXGsVQKHdpSAW4Q_AUICCgB&biw=1433&bih=732#imgrc=aFtUeC0wtJCkgM%3A) for a better understanding:
#
# 
#
# Matrix factorization assumes that:
#
# - Each user can be described by $d$ features. For example, feature 1 might be a referring to how much each user likes disney movies.
# - Each item, movie in this case, can be described by an analagous set of $d$ features. To correspond to the above example, feature 1 for the movie might be a number that says how close the movie is to a disney movie.
#
# After learning the two smaller matrices, all we have to do is to perform a matrix multiplication of the two matrices and the end result will be a our approximation for the rating the user would give that item (movie).
#
# The cool thing about this is that, we do not know what these features are nor do we have to determine them beforehand, which is why these features are often refer to as latent features. Next, we also don't know how many latent features are optimal for the task at hand. Thus, we simply have to use random search or grid search or other fancy techniques to perform hyperparameter tuning and determine the best number for $d$.
#
# Given all those information, the next question is: how do we learn the user matrix, $U$, and item matrix, $V$? Well, like a lot of machine learning algorithm, by minimizing a loss function.
#
# We start by denoting our $d$ feature user into math by letting a user $u$ take the form of a $1 \times d$-dimensional vector $\textbf{x}_{u}$. These for often times referred to as latent vectors or low-dimensional embeddings. Similarly, an item *i* can be represented by a $1 \times d$-dimensional vector $\textbf{y}_{i}$. And the rating that we predict user $u$ will give for item $i$ is just the dot product of the two vectors
#
# \begin{align}
# \hat r_{ui} &= \textbf{x}_{u} \textbf{y}_{i}^{T} = \sum\limits_{d} x_{ud}y_{di}
# \end{align}
#
# Where $\hat r_{ui}$ represents our prediction for the true rating $r_{ui}$. Next, we will choose a objective function to minimize the square of the difference between all ratings in our dataset ($S$) and our predictions. This produces a objective function of the form:
#
# \begin{align}
# L &= \sum\limits_{u,i \in S}( r_{ui} - \textbf{x}_{u} \textbf{y}_{i}^{T} )^{2} + \lambda \big( \sum\limits_{u} \left\Vert \textbf{x}_{u} \right\Vert^{2} + \sum\limits_{i} \left\Vert \textbf{y}_{i} \right\Vert^{2} \big)
# \end{align}
#
# Note that we've added on two $L_{2}$ regularization terms, with $\lambda$ controlling the strength at the end to prevent overfitting of the user and item vectors. $\lambda$, is another hyperparameter that we'll have to search for to determine the best value. Regularization can a can topic of itself, and if you're confused by this, consider checking out [this documentation](http://nbviewer.jupyter.org/github/ethen8181/machine-learning/blob/master/regularization/regularization.ipynb) that covers it a bit more.
# Now that we formalize our objective function, we'll introduce the **Alternating Least Squares with Weighted Regularization (ALS-WR)** method for optimizing it. The way it works is we start by treating one set of latent vectors as constant. For this example, we'll pick the item vectors, $\textbf{y}_{i}$. We then take the derivative of the loss function with respect to the other set of vectors, the user vectors, $\textbf{x}_{u}$ and solve for the non-constant vectors (the user vectors).
#
# \begin{align}
# \frac{\partial L}{\partial \textbf{x}_{u}}
# &\implies - 2 \sum\limits_{i}(r_{ui} - \textbf{x}_{u} \textbf{y}_{i}^{T} ) \textbf{y}_{i} + 2 \lambda \textbf{x}_{u} = 0 \\
# &\implies -(\textbf{r}_{u} - \textbf{x}_{u} Y^{T} )Y + \lambda \textbf{x}_{u} = 0 \\
# &\implies \textbf{x}_{u} (Y^{T}Y + \lambda I) = \textbf{r}_{u}Y \\
# &\implies \textbf{x}_{u} = \textbf{r}_{u}Y (Y^{T}Y + \lambda I)^{-1}
# \end{align}
#
# To clarify it a bit, let us assume that we have $m$ users and $n$ items, so our ratings matrix is $m \times n$.
#
# - The row vector $\textbf{r}_{u}$ just represents users *u*'s row from the ratings matrix with all the ratings for all the items (so it has dimension $1 \times n$)
# - We introduce the symbol $Y$, with dimensions $n \times d$, to represent all item row vectors vertically stacked on each other
# - Lastly, $I$ is just the identity matrix which has dimension $d \times d$ to ensure the matrix multiplication's dimensionality will be correct when we add the $\lambda$
#
# Now comes the alternating part: With these newly updated user vectors in hand, in the next round, we hold them as constant, and take the derivative of the loss function with respect to the previously constant vectors (the item vectors). As the derivation for the item vectors is quite similar, we will simply list out the end formula:
#
# \begin{align}
# \frac{\partial L}{\partial \textbf{y}_{i}}
# &\implies \textbf{y}_{i} = \textbf{r}_{i}X (X^{T}X + \lambda I)^{-1}
# \end{align}
#
# Then we alternate back and forth and carry out this two-step process until convergence. The reason we alternate is, optimizing user latent vectors, $U$, and item latent vectors $V$ simultaneously is hard to solve. If we fix $U$ or $V$ and tackle one problem at a time, we potentially turn it into a easier sub-problem. Just to summarize it, ALS works by:
#
# 1. Initialize the user latent vectors, $U$, and item latent vectors $V$ with randomly
# 2. Fix $U$ and solve for $V$
# 3. Fix $V$ and solve for $U$
# 4. Repeat step 2 and 3 until convergence
#
# Now that we have our equations, let's program this thing up! We'll set the model to use 20 factors and a regularization value of 0.01 (chosen at random) and train it for 100 iterations and compute the mean square error on the train and test set to assess algorithm convergence.
class ExplicitMF:
"""
Train a matrix factorization model using Alternating Least Squares
to predict empty entries in a matrix
Parameters
----------
n_iters : int
number of iterations to train the algorithm
n_factors : int
number of latent factors to use in matrix
factorization model, some machine-learning libraries
denote this as rank
reg : float
regularization term for item/user latent factors,
since lambda is a keyword in python we use reg instead
"""
def __init__(self, n_iters, n_factors, reg):
self.reg = reg
self.n_iters = n_iters
self.n_factors = n_factors
def fit(self, train, test):
"""
pass in training and testing at the same time to record
model convergence, assuming both dataset is in the form
of User x Item matrix with cells as ratings
"""
self.n_user, self.n_item = train.shape
self.user_factors = np.random.random((self.n_user, self.n_factors))
self.item_factors = np.random.random((self.n_item, self.n_factors))
# record the training and testing mse for every iteration
# to show convergence later (usually, not worth it for production)
self.test_mse_record = []
self.train_mse_record = []
for _ in range(self.n_iters):
self.user_factors = self._als_step(train, self.user_factors, self.item_factors)
self.item_factors = self._als_step(train.T, self.item_factors, self.user_factors)
predictions = self.predict()
test_mse = self.compute_mse(test, predictions)
train_mse = self.compute_mse(train, predictions)
self.test_mse_record.append(test_mse)
self.train_mse_record.append(train_mse)
return self
def _als_step(self, ratings, solve_vecs, fixed_vecs):
"""
when updating the user matrix,
the item matrix is the fixed vector and vice versa
"""
A = fixed_vecs.T.dot(fixed_vecs) + np.eye(self.n_factors) * self.reg
b = ratings.dot(fixed_vecs)
A_inv = np.linalg.inv(A)
solve_vecs = b.dot(A_inv)
return solve_vecs
def predict(self):
"""predict ratings for every user and item"""
pred = self.user_factors.dot(self.item_factors.T)
return pred
@staticmethod
def compute_mse(y_true, y_pred):
"""ignore zero terms prior to comparing the mse"""
mask = np.nonzero(y_true)
mse = mean_squared_error(y_true[mask], y_pred[mask])
return mse
def plot_learning_curve(model):
"""visualize the training/testing loss"""
linewidth = 3
plt.plot(model.test_mse_record, label = 'Test', linewidth = linewidth)
plt.plot(model.train_mse_record, label = 'Train', linewidth = linewidth)
plt.xlabel('iterations')
plt.ylabel('MSE')
plt.legend(loc = 'best')
als = ExplicitMF(n_iters = 100, n_factors = 40, reg = 0.01)
als.fit(train, test)
plot_learning_curve(als)
# From the result, we can see ALS converges after a few sweeps, which is one of the main reason for its popularity. Fast, thus scalable to bigger dataset.
# ## Validating the Recommendation
#
# After training our model, we'll actually look at the recommended result to see if our recommendation algorithm is any good or not. The way we do this is: The MovieLens dataset contains a file called `u.item` with information about each movie. To put the cherry on top, it turns out that there is a website called [themoviedb.org](http://www.themoviedb.org) which has a free API. If we have the IMDB "movie id" for a movie, then we can use this API to return the its poster. Note that you will have to sign up and request for an API key if you wish to reproduce the example below.
# +
# Load in movies' information, stored in the u.item (pipe delimited)
# file, the first element is the id, and the fourth element is the
# movie's IMDB url
idx_to_movie = {}
item_path = os.path.join(file_dir, 'u.item')
with open(item_path, encoding = 'latin1') as f:
for line in f.readlines():
info = line.split('|')
idx = int(info[0]) - 1
movie = info[4]
idx_to_movie[idx] = movie
idx_to_movie[0]
# -
# For example, if we paste the sample url above into our browser, then our url will get redirected. The resulting url contains the IMDB movie ID as the last information in the url starting with "tt". For example, the redirected url for Toy Story is http://www.imdb.com/title/tt0114709/, and the corresponding IMDB movie ID is therefore tt0114709.
# stripping the movie id with requests
response = requests.get(idx_to_movie[0])
movie_id = response.url.split('/')[-2]
print(movie_id)
# From the API documentation:
#
# To build an image URL, we will need 3 pieces of data. The base_url, size and file_path. Simply combine them all and you will have a fully qualified URL. Here’s an example URL:
# https://image.tmdb.org/t/p/w500/8uO0gUM8aNqYLs1OsTBQiXu0fEv.jpg
#
# So to get the base_url, we send a request to the https://api.themoviedb.org/3/configuration using our API key as the parameter, after that we load the information into a dictionary and access the `base_url` key under the `images` key (the base_url is nested under images), and we'll also append the poster size at the end of the base_url.
# +
# here I'm simply loading my API key
# from a .env file, and using dotenv package
# to obtain the value of the variable, you may
# simply change the api_key variable to your key
load_dotenv('.env')
api_key = os.environ.get('API_KEY')
# request the base_url
params = {'api_key': api_key}
response = requests.get('http://api.themoviedb.org/3/configuration', params = params)
# for the poster there are different sizes
# 'w92', 'w154', 'w185', 'w342', 'w500', 'w780', 'original',
# simply change it to the one you prefer
base_url = json.loads(response.text)['images']['base_url'] + 'w185'
base_url
# -
# Next to get the file_path to the poster image, the have to send another request to http://api.themoviedb.org/3/movie/{}/images, where we have to fill in the movie_id for {}. Then the actual path will be stored under the `file_path` key under the `posters` key (accessing the posters key will give you a list of path, we'll simply use the first one).
image_url = 'http://api.themoviedb.org/3/movie/{}/images'.format(movie_id)
response = requests.get(image_url, params = params)
file_path = json.loads(response.text)['posters'][0]['file_path']
file_path
# After that we can simply concatenate the base_url and the file path and display the poster image.
full_path = base_url + file_path
Image(full_path)
# Unfortunately, there are also other edge-cases, where we have to search for the title and grab the poster image there. e.g.
# +
movie_url = 'http://us.imdb.com/M/title-exact?Fox%20and%20the%20Hound,%20The%20(1981)'
response = requests.get(movie_url)
# see how the url is different this time,
# we have to strip out the title between the = sign
# and pass it to the movie search API using the title
# as a query parameter
print(response.url)
query = response.url.split('=')[-2]
params['query'] = query
response = requests.get('http://api.themoviedb.org/3/search/movie', params = params)
json.loads(response.text)
file_path = json.loads(response.text)['results'][0]['poster_path']
full_path = base_url + file_path
Image(full_path)
# -
# In the following section, we'll train the model again, use [themoviedb.org](http://www.themoviedb.org)'s API to grab the movie posters and visualize the top 5 most similar movies (measured by cosine distance) for some input movie. These similar movies will serve as our recommendation for a given input movie.
#
# Side note, the images are displayed using HTML so multiple images can be displayed in one line, idea taken from this [Stackoverflow question](http://stackoverflow.com/questions/19471814/display-multiple-images-in-one-ipython-notebook-cell/27795087#27795087).
class Recommendation:
"""
Train the ALS model to be able to
recommend the top-k similar movies for a given query
Parameters
----------
k: int
number of top similar movies to recommend for
a given query
model: ExplicitMF
the recommendation algorithm
idx_to_movie: dictionary
a dictionary mapping movie id to their imdb url
"""
def __init__(self, model, k, idx_to_movie):
self.k = k
self.model = model
self.idx_to_movie = idx_to_movie
def fit(self, train, test):
# train the model and compute the pairwise cosine distance
self.model.fit(train, test)
self.sim_mat = pairwise_distances(self.model.item_factors, metric = 'cosine')
# also get the base url for the API
# assuming API key is stored in .env file
load_dotenv('.env')
api_key = os.environ.get('API_KEY')
self.params = {'api_key': api_key}
response = requests.get('http://api.themoviedb.org/3/configuration',
params = self.params)
self.base_url = json.loads(response.text)['images']['base_url'] + 'w185'
return self
def recommend(self, query):
movie_url = idx_to_movie[query]
input_poster = self._get_poster(movie_url)
input_image = '<img src={} />'.format(input_poster)
recommended_images = self._get_top_k_recommendation(query)
# display query movie and for that movie top-k recommendations
display( HTML('<font size=5> Input </font>') )
input_image = '<img src={} />'.format(input_poster)
display( HTML(input_image) )
display( HTML('<font size=5> Recommendation </font>') )
display( HTML(recommended_images) )
def _get_poster(self, movie_url):
"""get the poster image for a given movie url"""
try:
response = requests.get(movie_url)
movie_id = response.url.split('/')[-2]
image_url = 'http://api.themoviedb.org/3/movie/{}/images'.format(movie_id)
response = requests.get(image_url, params = self.params)
file_path = json.loads(response.text)['posters'][0]['file_path']
except KeyError:
try:
response = requests.get(movie_url)
query = response.url.split('=')[-2]
self.params['query'] = query
response = requests.get('http://api.themoviedb.org/3/search/movie',
params = self.params)
json.loads(response.text)
file_path = json.loads(response.text)['results'][0]['poster_path']
self.params.pop('query')
except KeyError:
# sometimes the url just doesn't work
# return an empty string to not mess up the display
file_path = ''
full_path = self.base_url + file_path
return full_path
def _get_top_k_recommendation(self, query):
"""
sort the similarity vector to get the most similar,
remember the smaller the cosine, the more similar they are
but the query item itself will always be the most similar,
so exclude that from the output
"""
sim = self.sim_mat[query]
top_k_item_idx = np.argsort(sim)[1:self.k + 1]
recommended_images = ''
for item_idx in top_k_item_idx:
movie_url = self.idx_to_movie[item_idx]
recommended_poster = self._get_poster(movie_url)
recommended_images += """
<img style='width: 150px; height: 250px; margin: 0px;
float: left; border: 1px solid black;'
src={} />
""".format(recommended_poster)
return recommended_images
als = ExplicitMF(n_iters = 100, n_factors = 40, reg = 0.01)
recsys = Recommendation(model = als, k = 5, idx_to_movie = idx_to_movie)
recsys = recsys.fit(train, test)
query = 1
recsys.recommend(query)
query = 500
recsys.recommend(query)
# I'll leave it up to you to determine whether these recommendations are good or not (results will slightly differ across different runs). In practice A/B testing is most likely a better way to judge whether these recommendations are useful, but that's a topic for some other day.
# # Reference
#
# - [Blog: Intro to Recommender Systems: Collaborative Filtering](http://blog.ethanrosenthal.com/2015/11/02/intro-to-collaborative-filtering/)
# - [Blog: Explicit Matrix Factorization: ALS, SGD, and All That Jazz](http://blog.ethanrosenthal.com/2016/01/09/explicit-matrix-factorization-sgd-als/)
# - [Quora: What is the Alternating Least Squares method in recommendation systems?](https://www.quora.com/What-is-the-Alternating-Least-Squares-method-in-recommendation-systems)
|
recsys/1_ALSWR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Genetic algo for lighthouse sensor distribution on arbitrary mesh
import numpy as np
from stl import mesh as meshstl
from mpl_toolkits import mplot3d
import matplotlib.pyplot as plt
from data.plotmesh import plot_mesh
import math
import random
from pyquaternion import Quaternion
from scipy.linalg import qr
import roslib
import rospy
import math
import tf
rospy.init_node('fixed_tf_broadcaster')
# +
# how many sensors would you like to distribute?
sensorsToDistribute = 11
# which mesh would you like to use?
stl_file = 'roboy_models/TestCube/stls/monkey.stl'
#stl_file = 'roboy_models/TestCube/stls/IcoSphere_360.stl'
# +
#Move Lighthouses to
translationLH1 = [-2.,0,2.]
quat1 = Quaternion(axis=[0,0,1],angle=0*np.pi)
global LH1
LH1 = (translationLH1, quat1)
translationLH2 = [2,0.,2.]
quat2 = Quaternion(axis=[1,0,0], angle= -np.pi)
global LH2
LH2 = (translationLH2, quat2)
print(LH1); print(LH2)
# +
from data.rvizMeshVis import meshVisualization
scale = 0.01
position = [0,0,0]
global orientationMesh
orientationMesh = Quaternion(axis=(1,0,0),angle = np.pi*0)
# -
for i in range(5):
meshVisualization(orientationMesh, stl_file, color=(1.,1.,1.,0.9))
# # Preprocess data
# +
from data.trisByDistance import *
#Get mesh vertices and normals
mesh1 = meshstl.Mesh.from_file(('../'+ stl_file))
#mesh1 = meshstl.Mesh.from_file('../src/roboy_models/roboy_2_0_simplified/meshes/CAD/torso.stl')
global triangles
triangles = scale * np.matrix(mesh1.points)
global trianglesBackup
trianglesBackup = triangles
global sortedTriangles
lighthouses = [LH1, LH2]
sortedTriangles = []
normalsNotNorm = mesh1.normals
global normals
normals = []
for normal in normalsNotNorm:
normals.append(1/np.linalg.norm(normal,2)*normal)
normals = np.matrix(normals)
normals = scale * normals
for l in lighthouses:
tris = trisByMinDistanceSortedMap(triangles, l[0])
sortedTriangles.append(tris)
#vertices = np.reshape(triangles,(len(triangles)*3,3))
#Initialize sensors in centers of triangle
sensors = (triangles[:,0:3]+triangles[:,3:6]+triangles[:,6:9])/3
print('%d triangles' %len(triangles))
print('')
#print('%d vertices' %len(vertices))
#print('')
print('%d sensors in centers of triangles' %len(sensors))
print('')
print('%d normals' %len(normals))
#print(normals)
# +
from data.rvizNormalsVis import NormalsVisual
#NormalsVisual(sensors,normals)
# -
# # GA
from deap import algorithms, base, creator, tools
# +
#sensors to dict
global sensor_dict
sensor_dict = list(zip(range(len(sensors)), sensors.tolist()))
global sensorDictBackup
sensorDictBackup = sensor_dict
# +
from data.rvizSensorVis import sensorVisualization
#color = (r,g,b,a)
sensorVisualization(sensor_dict, rate=500, sphereSize=0.03, color=(0,0,1,1))
# -
creator.create("FitnessMax", base.Fitness, weights=(1,)) # 1 -> maximum probblem
creator.create("Individual", list, fitness=creator.FitnessMax)
toolbox = base.Toolbox()
# +
from data.randomSensor import randomSensor
toolbox = base.Toolbox()
# Attribute generator
toolbox.register("attr_bool", randomSensor, sensor_dict)
# Structure initializers
toolbox.register("individual", tools.initRepeat, creator.Individual,
toolbox.attr_bool, sensorsToDistribute)
toolbox.register("population", tools.initRepeat, list, toolbox.individual)
# -
toolbox.attr_bool()
toolbox.individual()
# # Evaluation (Fitness) Function
# +
from data.RayIntersectsTriangle import rayIntersectsTriangle, rayIntersectsTriangleVisual
def FitnessFunction(sensors):
br = tf.TransformBroadcaster()
br.sendTransform((LH1[0][0], LH1[0][1], LH1[0][2]),
(quat1[0], quat1[1], quat1[2], quat1[3]),
rospy.Time.now(),
"lighthouse1",
"world")
br.sendTransform((LH2[0][0], LH2[0][1], LH2[0][2]),
(quat2[0], quat2[1], quat2[2], quat2[3]),
rospy.Time.now(),
"lighthouse2",
"world")
#1. COMPUTE VISIBLE SENSORS AT THE MOMENT
LH1_array = np.asarray(LH1[0])
LH2_array = np.asarray(LH2[0])
#testTriangle = np.squeeze(np.asarray(triangles[0]))
visibleLH1 = 0.0
visibleLH2 = 0.0
angleLH1 = []
for i, nmbr_sensor in enumerate(sensors):
sensor = sensor_dict[nmbr_sensor][1]
#get distance of current sensor and check if intersection
interDistLH1 = rayIntersectsTriangle(LH1_array, sensor,
np.squeeze(np.asarray(triangles[nmbr_sensor])), 'lighthouse1')
distLH1 = np.linalg.norm(np.asarray(sensor) - LH1_array)
interDistLH2 = rayIntersectsTriangle(LH2_array, sensor,
np.squeeze(np.asarray(triangles[nmbr_sensor])), 'lighthouse2')
distLH2 = np.linalg.norm(np.asarray(sensor) - LH2_array)
# get angle between lighthouse vector and normal
normal = np.squeeze(np.asarray(normals[nmbr_sensor]))
#LH1
sensorToLH1 = sensor + (LH1_array - sensor)
angleLH1 = np.dot(sensorToLH1,normal)/(np.linalg.norm(sensorToLH1)*np.linalg.norm(normal))
#angleLH1 = np.arccos(angleLH1)
#LH2
sensorToLH2 = sensor + (LH2_array - sensor)
angleLH2 = np.dot(sensorToLH2,normal)/(np.linalg.norm(sensorToLH2)*np.linalg.norm(normal))
#angleLH2 = np.arccos(angleLH2)
# Might be changed to something different
# Calculate visibility factor depending on angle between normal and lighthouse
visFactorLH1 = angleLH1#np.cos(angleLH1)
visFactorLH2 = angleLH2#np.cos(angleLH2)
#print("Sensor %d has VisFactor %f mit LH1"%(nmbr_sensor, visFactorLH1))
#print("Sensor %d has VisFactor %f mit LH2"%(nmbr_sensor, visFactorLH2))
#print('interDist');print(interDistLH1);print(interDistLH2);print('endinterDist')
isVisible1 = True
isVisible2 = True
# 1st lighthouse
if(visFactorLH1 > 0):
for (j, dist) in sortedTriangles[0]:
if(nmbr_sensor != j):
#print("Testing sensor %i vs tris %i: distance of Sensor %f vs triangle %f" % (i
#, j, distLH1, dist))
tris = triangles[j]
newInterDistLH1 = rayIntersectsTriangle(LH1_array, sensor,
np.squeeze(np.asarray(tris)), 'lighthouse1')#,j)
if(newInterDistLH1 < interDistLH1 and newInterDistLH1 != False):
isVisible1 = False
if(not isVisible1 or dist > distLH1):
# Break if not visible or already checked all tris
# that are located closer to the lighthouse that the sensor
break
if(isVisible1):
visibleLH1 += visFactorLH1
# 2nd lighthouse
if(visFactorLH2 > 0):
for (j, dist) in sortedTriangles[1]:
if(nmbr_sensor != j):
tris = triangles[j]
newInterDistLH2 = rayIntersectsTriangle(LH2_array, sensor,
np.squeeze(np.asarray(tris)), 'lighthouse2')#,j)
if(newInterDistLH2 < interDistLH2 and newInterDistLH2 != False):
isVisible2 = False
if(not isVisible2 or dist > distLH2):
# Break if not visible or already checked all tris
# that are located closer to the lighthouse that the sensor
break
if(isVisible2):
visibleLH2 += visFactorLH2
#print(newInterDistLH1); print(newInterDistLH2)
fractionVisibleLH1 = float(visibleLH1) / sensorsToDistribute
fractionVisibleLH2 = float(visibleLH2) / sensorsToDistribute
#2. COMPUTE EUCLIDEAN DISTANCE OF SENSORS
individual = sensors
dist = 0.0
distTemp = 0.0
distMax = 0.0
for i,ind in enumerate(individual):
ind = np.asarray(sensor_dict[ind][1])
for j in range(i,len(individual)):
if(i != j):
indivi = np.asarray(sensor_dict[individual[j]][1])
distTemp = np.linalg.norm(ind-indivi)
dist += distTemp
if(distTemp > distMax):
distMax = distTemp
#print(dist);print(distMax)
distNorm = dist/(distMax * sensorsToDistribute)
#print(visibleLH1);print(visibleLH2);print('')
return (fractionVisibleLH1 + fractionVisibleLH2 + distNorm),
# +
toolbox.register("evaluate", FitnessFunction)
toolbox.register("mate", tools.cxTwoPoint)
# Independent probability : for each attribute to be mutated.# low~up rondom int
toolbox.register("mutate", tools.mutUniformInt, low=0, up=len(sensors.tolist())-1, indpb=0.2)
toolbox.register("select", tools.selTournament, tournsize=3)
# +
# Creating population
population = toolbox.population(n=100)
# -
hof = tools.HallOfFame(10)
stats = tools.Statistics(lambda ind: ind.fitness.values)
stats.register("avg", np.mean)
stats.register("std", np.std)
stats.register("min", np.min)
stats.register("max", np.max)
# +
from data.algorithmsMod import varAnd
from deap import tools
from data.trafomatrix import getRandomRotationmatrix
from data.bestSensorVis import bestSensorVis
#MODDED VERSION of eaSimple from DEAP
def eaSimpleMod(population, toolbox, cxpb, mutpb, ngen, stats=None,
halloffame=None, verbose=__debug__):
"""This algorithm reproduce the simplest evolutionary algorithm as
presented in chapter 7 of [Back2000]_.
Modded version of DEAP Evolutionary Algorithm Framework
https://github.com/DEAP/deap
"""
global sensor_dict
global triangles
global orientationMesh
logbook = tools.Logbook()
logbook.header = ['gen', 'nevals'] + (stats.fields if stats else [])
# Evaluate the individuals with an invalid fitness
invalid_ind = [ind for ind in population if not ind.fitness.valid]
fitnesses = toolbox.map(toolbox.evaluate, invalid_ind)
for ind, fit in zip(invalid_ind, fitnesses):
ind.fitness.values = fit
if halloffame is not None:
halloffame.update(population)
record = stats.compile(population) if stats else {}
logbook.record(gen=0, nevals=len(invalid_ind), **record)
if verbose:
print logbook.stream
# Begin the generational process
for gen in range(1, ngen + 1):
# Select the next generation individuals
offspring = toolbox.select(population, len(population))
# Vary the pool of individuals
offspring = varAnd(offspring, toolbox, cxpb, mutpb)
# Evaluate the individuals with an invalid fitness
invalid_ind = [ind for ind in offspring if not ind.fitness.valid]
fitnesses = toolbox.map(toolbox.evaluate, invalid_ind)
for ind, fit in zip(invalid_ind, fitnesses):
ind.fitness.values = fit
# Update the hall of fame with the generated individuals
if halloffame is not None:
halloffame.update(offspring)
# Replace the current population by the offspring
population[:] = offspring
# Append the current generation statistics to the logbook
record = stats.compile(population) if stats else {}
logbook.record(gen=gen, nevals=len(invalid_ind), **record)
if verbose:
print logbook.stream
#sensorMovement = tools.selBest(population, k=1)[0]
#bestSensorVis(sensor_dict, sensorMovement, rate=1000, color=(0,1,0,0.8), sphereSize=0.2)
if(gen%1==0):
global sensorDictBackup
global trianglesBackup
sensor_dict = sensorDictBackup
R = getRandomRotationmatrix()
sensorDictNew = []
for sensor in sensor_dict:
sensorDictNew.append(np.squeeze(np.asarray(R.dot(np.array(sensor[1])))).tolist())
sensor_dict = list(zip(range(len(sensorDictNew)), sensorDictNew))
tri1 = R.dot(np.transpose(trianglesBackup[:,0:3]))
tri2 = R.dot(np.transpose(trianglesBackup[:,3:6]))
tri3 = R.dot(np.transpose(trianglesBackup[:,6:9]))
triangles = np.concatenate((tri1.T,tri2.T,tri3.T),axis=1)
# resort the triangles by distance from lighthouses for speedup
global sortedTriangles
lighthouses = [LH1, LH2]
sortedTriangles = []
for l in lighthouses:
tris = trisByMinDistanceSortedMap(triangles, l[0])
sortedTriangles.append(tris)
orientationMesh = Quaternion(matrix=R)
meshVisualization(orientationMesh, stl_file, color=(1.,1.,1.,0.9))
#sensorVisualization(sensor_dict, rate=500, sphereSize=0.03, color=(0,0,1,1))
sensorMovement = tools.selBest(population, k=1)[0]
bestSensorVis(sensor_dict, sensorMovement, rate=1000, color=(0,1,0,0.8), sphereSize=0.2)
return population, logbook
# -
population, log = eaSimpleMod(population,
toolbox,
cxpb=0.5,
mutpb=0.5,
ngen=150,
stats=stats,
halloffame=hof,
verbose=True)
bestSensors = tools.selBest(population, k=1)
print(bestSensors[0])
# +
from data.bestSensorVis import bestSensorVis
#Sensor visualization in RVIZ
orientationMesh = (0,0,0,0)
meshVisualization(orientationMesh, stl_file, color=(1.,1.,1.,0.9))
sensorVisualization(sensorDictBackup, rate=500, sphereSize=0.03, color=(0,0,1,1))
bestSensorVis(sensorDictBackup, bestSensors[0], rate=500, color=(1,0,0,0.8), sphereSize=0.1)
# -
|
GA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: '''Python Interactive'''
# name: 98b5f46f-c131-4b88-9f13-102a8737ee41
# ---
# ## Úkol 1 - Svařované krabičky
#
# Máme firmu na výrobu svařovaných krabiček. Výrobce plechů nám dodává plech o šířce *a* a výšce *b*. Naším úkolem je zjistit, jak velké kusy v rozích odstřihnout, abychom maximalizovali objem krabičky. Krabička nemá víko, má jenom dno a čtyři strany (viz následující obrázek).
#
# 
# Objem naší krabičky tedy bude:
# $$V(x) = x (a-2x) (b-2x) = abx - 2ax^2 - 2bx^2 + 4x^3$$
# A naše úloha tedy bude (předpokládejme, že $a \leq b$):
# $$\underset{x}{\arg\max} \ V(x), \ subject \ to: \ x\in(0,\frac{1}{2}a)$$
# To je jenom složite napsané, že hledáme takový parametr $x$, aby funkce $V(x)$ byla co možná největší. Přitom však $x$ musí být z intervalu $(0,\frac{1}{2}a)$.
#
# Možná vám bude úloha ze začátku připadat složitá, ale ve skutečnosti je to středoškolská matematika. Vykreslíme si funkci objemu v závislosti na *x* pro zvolené *a* a *b*.
import matplotlib.pyplot as plt
import numpy as np
a, b = 1., 2.
x = np.arange(0., a/2, 0.01)
v = x * (a - 2*x) * (b - 2*x)
plt.plot(x, v)
# Optimální řešení se nacházi v bodě, kde je objem největší. Z matematického hlediska musíme vyřešit pouze 2 typy bodů:
#
# 1. Začátek a konec intervalu intervalu ($x=0, x=\frac{1}{2}a$)
# 2. Body, prok teré platí $\frac{dV(x)}{dx} = 0$ a zároveň jsou v našem v intervalu
#
# Derivace naší funkce objemu potom je:
# $$\frac{dV(x)}{dx} = ab-4ax-4bx+12x^2$$
#
# Zkusme si ji vykreslit a podívat se, kde je její hodnota přibližně $0$.
dv = a*b - 4*a*x - 4*b*x + 12*x*x
plt.plot(x, dv)
plt.axhline(0.0, color="r")
# Tento příklad je samozřejmě tak jednoduchý, že ho můžeme vyřešit analyticky, je to polynom 2. řádu.
# koeficienty kvadratického polynomu
poly_a = 12.
poly_b = -4*a - 4*b
poly_c = a*b
diskriminant = poly_b**2 - 4*poly_a*poly_c
x1 = (-poly_b + np.sqrt(diskriminant))/2./poly_a
x2 = (-poly_b - np.sqrt(diskriminant))/2./poly_a
print(f"roots are {x1:2f} and {x2:2f}")
# Musíme ještě ověřit, který z kořenů patří a který nepatří do našeho intervalu $(0, \frac{1}{2}a)$:
solution = x1 if x1 >= 0 and x1 <= a/2 else x2
solution
# Zkusme zakreslit do grafu (funkci objemu jsem vynásobil 20 krát, aby byla v grafu pěkně vidět, protože nám jde hlavně o tvar):
plt.figure()
plt.plot(x, 20*v, label="20*V(x)")
plt.plot(x, dv, label="dV(x)")
plt.axvline(solution, color="g", label="solution")
plt.axhline(0, color="r", label="f(x)=0")
plt.legend()
plt.xlabel("x")
plt.ylabel("20*V(x), dV(x)")
plt.show()
# Zkuste se zamyslet, jestli můžou nastat případy, kdy diskriminant bude záporný? Má tento postup nějaká úskalí?
|
courses/2375004/cvxpy/boxes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df=pd.read_csv('heart.csv')
df.head()
# ##### About the dataset
# age - Age of the patient
#
# sex - Sex of the patient
#
# # cp - Chest pain type ~ 0 = Typical Angina, 1 = Atypical Angina, 2 = Non-anginal Pain, 3 = Asymptomatic
#
# trtbps - Resting blood pressure (in mm Hg)
#
# chol - Cholestoral in mg/dl fetched via BMI sensor
#
# fbs - (fasting blood sugar > 120 mg/dl) ~ 1 = True, 0 = False
#
# restecg - Resting electrocardiographic results ~ 0 = Normal, 1 = ST-T wave normality, 2 = Left ventricular hypertrophy
#
# thalachh - Maximum heart rate achieved
#
# oldpeak - Previous peak
#
# slp - Slope
#
# caa - Number of major vessels
#
# thall - Thalium Stress Test result ~ (0,3)
#
# exng - Exercise induced angina ~ 1 = Yes, 0 = No
#
# output - Target variable
df.isnull().sum()
import seaborn as sns
sns.countplot(df['output'],data=df)
sns.countplot(df['sex'],data=df)
sns.countplot(df['fbs'],data=df)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(20,20))
sns.heatmap(df.corr(),annot=True,cmap="YlGnBu")
df.corr()
sns.barplot(x=df['sex'],y=df['output'],data=df)
sns.countplot(df['cp'],data=df)
sns.countplot(df['thall'],data=df)
sns.countplot(df['caa'],data=df)
sns.countplot(df['restecg'],data=df)
sns.countplot(df['slp'],data=df)
sns.countplot(df['exng'],data=df)
sns.distplot(df['thalachh'],color='green')
sns.distplot(df['trtbps'],color='green')
# ###### violin plot tells us both in kernel density estimator and boxplot
# +
sns.violinplot(df['fbs'],df['output'],data=df,palette='rainbow')
# -
print('no.of zeroes in age {} '.format((df['age']==0).sum()))
print('no.of zeroes in thalachh {}'.format((df['thalachh']==0).sum()))
print('no.of zeros in trtbps {}'.format((df['trtbps']==0).sum()))
print('no.of zeros in chol {}'.format((df['chol']==0).sum()))
print('no.of zeros in trtbps {}'.format((df['trtbps']==0).sum()))
# ### Data Preprocessing
df=pd.get_dummies(df,columns=['sex','cp','fbs','restecg','exng','slp','caa','thall'])
df.head()
from sklearn.preprocessing import StandardScaler
sc=StandardScaler()
from sklearn.model_selection import train_test_split
x=df.drop(['output'],axis=1)
y=df['output']
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.3,random_state=42)
from sklearn.ensemble import RandomForestClassifier
rfc=RandomForestClassifier()
'''n_estimators=100,
*,
criterion='gini',
max_depth=None,
min_samples_split=2,
min_samples_leaf=1,
min_weight_fraction_leaf=0.0,
max_features='auto',
max_leaf_nodes=None,
min_impurity_decrease=0.0,
min_impurity_split=None,
n_jobs=None,
random_state=None,
verbose=0,
max_samples=None,'''
# +
rand={'n_estimators':[int(x) for x in np.linspace(start = 100, stop = 1000, num = 10)],
'criterion' : ["gini", "entropy"],
'max_depth':[int(x) for x in np.linspace(100,1000,10)],
'min_samples_split':[2,4,6,8,10,12,14],
'max_features':["auto", "sqrt", "log2"]}
# -
# +
#from sklearn.model_selection import GridSearchCV
# +
#gc=GridSearchCV(estimator=rfc,param_grid=rand,scoring='accuracy',n_jobs=-1,cv=5)
# +
#gc.fit(x_train,y_train)
# -
from sklearn.model_selection import RandomizedSearchCV
ra=RandomizedSearchCV(estimator=rfc,param_distributions=rand,scoring='accuracy',n_jobs=-1,n_iter=100,cv=5,random_state=45)
ra.fit(x_train,y_train)
ra.best_estimator_
ra.best_score_
from sklearn.ensemble import RandomForestClassifier
rf=RandomForestClassifier(max_depth=200, max_features='sqrt',min_samples_leaf=6,min_samples_split=15, n_estimators=700,random_state=36)
rf.fit(x_train,y_train)
ypred=rf.predict(x_test)
from sklearn.metrics import confusion_matrix,accuracy_score
cm=confusion_matrix(y_test,ypred)
cm
score=accuracy_score(y_test,ypred)
score
from sklearn.naive_bayes import GaussianNB
x_train=sc.fit_transform(x_train)
x_test=sc.transform(x_test)
nb=GaussianNB()
nb.fit(x_train,y_train)
y_pred=nb.predict(x_test)
cm=confusion_matrix(y_test,y_pred)
cm
print('score:{}'.format(accuracy_score(y_test,y_pred)))
import pickle
file=open('Heart.pkl','wb')
pickle.dump(rfc,file)
x.columns
|
heart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize Otter
import otter
grader = otter.Notebook()
# -
# # A 2016 Election Analysis
# +
import numpy as np
from datascience import *
# These lines set up graphing capabilities.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
# -
# In this assignment, you will be analyzing a dataset of polling data from the 2016 Presidential Election. If you follow politics, you may know that many polls strongly underestimated <NAME>'s performance in several key states, causing him to win the election despite many models showing it would be a Clinton victory. We will try to investigate and see if these errors are simple polling errors, or whether there were more serious miscalculations.
#
# This data was taken from FiveThirtyEight, a respected organization that focuses on statistical and social analysis to predict election outcomes. You can find the raw source of the data, as well as more information about the dataset, [here](https://www.kaggle.com/fivethirtyeight/2016-election-polls).
#
# _Acknowledgement: This assignment was originally developed and contributed by <NAME>, the UCSB Data Science Fellow (2020-21 cohort)._
# + [markdown] deletable=false editable=false
# Let's import the required libraries and load our dataset from 'presidential_polls.csv' into a variable called original_data.
#
# <!--
# BEGIN QUESTION
# name: q0
# manual: false
# -->
# -
original_data = ...
# Looking at the columns, there are many different features of this data. Luckily, we are only interested in a few of them:
# * forecastdate: The date the forecast was uploaded to FiveThirtyEight
# * adjpoll_clinton / adjpoll_trump: the adjusted/calculated percentage of people voting for each candidate. Since each poll was random, it likely oversurveyed people of different demographics - for example, 80% of people asked could have been members of Party A, when it is known that the true population proporion is 50%. Adjusting the average allows the bias to be removed and gives a more accurate prediction.
# * state: the state where the entry was polled
# * grade: The grade that the pollster has on FiveThirtyEight, ranging from A to F with +/-
original_data.labels
# ## Question 1 - A General Focus on Wisconsin
#
# + [markdown] deletable=false editable=false
#
# ### 1.1 - Load the data
# Let's spend some time looking at data from Wisconsin - one of the more surprising results of the election. Let's load all polls that took place in Wisconsin into a variable called wisconsin.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# manual: false
# -->
#
# +
wisconsin = ...
wisconsin
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# ### 1.2 - Plotting polling averages
#
# There are clearly many polls that take place in Wisconsin - luckily, they are in order chronologically, so we don't have to worry about sorting them. However, we should probably visualize the results in a more meaningful way. Plot the `adjpoll_clinton` and `adjpoll_trump` columns both in one plot.
#
# Comment on what this visualization represents: e.g., what is the x-axis and the y-axis.
#
# <!--
# BEGIN QUESTION
# name: q1_2a
# manual: true
# -->
#
# -
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# Comment on the pattern that you notice.
#
#
# <!--
# BEGIN QUESTION
# name: q1_2b
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# ### 1.3 - Array of difference between Clinton, Trump
#
# To further understand this relationship, create an array that contains the difference between the percent that voted for Clinton and those that voted for Trump. Then, run the cell below to plot this relationship.
#
# <!--
# BEGIN QUESTION
# name: q1_3
# manual: false
# -->
#
# -
adj_diff_wisconsin = ...
plt.title('Difference between Clinton Vote Percentage and Trump Vote Percentage')
plt.plot(adj_diff_wisconsin)
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# ### 1.4 - Examining the differences
#
# Based on the above visualization, who do you expect to win the election? How confident are you in your analysis?
#
# <!--
# BEGIN QUESTION
# name: q1_4
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# ### Question 2.1 - A Look at high-ranking pollsters
#
# Trump won Wisconsin by just under 1% in 2016. This may be surprising! FiveThirtyEight also ranks pollsters, based on factors such as historical reliability and inherent bias.
#
# Let's select only the A+, A, and A- rated pollsters, to see if they had a better take on the result. Note: you will need to use the [datascience.are.contained_in](http://data8.org/datascience/predicates.html) module. Then, plot the adjusted averages again.
#
# <!--
# BEGIN QUESTION
# name: q2_1
# manual: true
# -->
#
# -
high_rankings = ...
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
# -
...
# + [markdown] deletable=false editable=false
# <!-- BEGIN QUESTION -->
#
# ### Question 2.2
#
# Do you notice a significant difference depending on the rankings? Did they do a better job at predicting the true outcome of the election?
#
# <!--
# BEGIN QUESTION
# name: q2_2
# manual: true
# -->
#
# -
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# # Question 3: Open Ended Analysis
#
# 1. Pick one of the following states: Pennsylvania, Ohio, Florida, Michigan
# 2. Perform an analysis, similar to what we did for Wisconsin, on the state. Then, look up and find the actual result of the state.
# 3. Answer the following questions in some depth (feel free to use the resources below as a starting point for your research):
# - What are some possible explanations for the discrepancies between the polls and the results of the election?
# - To what extent should we be trusting the polls' predictions?
#
# <!--
# BEGIN QUESTION
# name: q3
# manual: true
# -->
#
# +
# Code here - feel free to add as many cells as you need
# SOLTUION NO PROMPT
# -
# <!-- END QUESTION -->
#
#
#
# # Interested in learning more?
#
# The subject of elections and polling has only gotten more relevant since 2016. Here are some resources to help you dve deeper into the topic!
#
# [Why 2016 Election Polls Missed Their Mark](https://www.pewresearch.org/fact-tank/2016/11/09/why-2016-election-polls-missed-their-mark/): a brief but informative article from Pew Research
#
# FiveThirtyEight issued a [rebuttal](https://fivethirtyeight.com/features/the-polls-are-all-right/) defnding their record, including the 2016 election. In fact, their predictions in 2018 would go on to be extremely accurate. Feel free to follow the hyperlinks throughout the article, as many lead to interesting articles and papers.
#
#
# Finally, if you're interested in a more academic approach, see [this paper](https://eprints.soton.ac.uk/413658/1/JenningsWlezienPollingErrors.pdf), which conducted an analysis of thousands of polls in the last 60+ years and came to some interesting conclusions.
#
# # Your Exploration / Future Work
#
# It's your turn. You now have a chance to experiment with everything that you have learned in this class so far.
#
# Take a look at additional attributes that are given in this dataset and write down any other questions that you want to answer. Then, give it your best shot at answering them.
#
# What other datasets could you bring in for additional exploration? Can the dataset we used earlier in this course be helpful with any of the exploration?
#
# If you found/will use them, remember to include the link to the source and the data file with your submission.
#
# Feel free to use the `Insert` menu above to add as many extra cells as you need.
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
# + [markdown] deletable=false editable=false
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
# + deletable=false editable=false
# Save your notebook first, then run this cell to export your submission.
grader.export("election-predictions.ipynb")
# -
#
|
project2/election-predictions/election-predictions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
posts = json.load(open('posts.json'))
len(posts), posts[0]
text = '\n'.join(posts).lower()
len(text)
characters = sorted(list(set(text)))
char_lookup = characters.index
len(characters)
# +
from tqdm import tqdm
maxlen = 100
X, Y = [], []
for i in tqdm(range(len(text) - maxlen)):
x = text[i:i+maxlen]
y = text[i+maxlen]
X.append([char_lookup(c) for c in x])
Y.append(char_lookup(y))
len(X), len(Y)
# +
import numpy as np
import keras.utils
X = np.reshape(X, (len(X), maxlen)) / len(characters) # normalized seq input
Y = keras.utils.to_categorical(Y)
X.shape, Y.shape
# -
from keras.models import Sequential
from keras.layers import Embedding, LSTM, Dense, Flatten, Dropout
model = Sequential()
model.add(Embedding(len(characters), 40, input_length=maxlen, name="Embedding"))
model.add(LSTM(256, name="LSTM"))
model.add(Dropout(0.2, name="Dropout"))
model.add(Dense(len(characters), activation='softmax', name="FullyConnected"))
print(model.summary())
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(X, Y, epochs=1, batch_size=256, shuffle=True)
char_reverse_lookup = characters.__getitem__
char_reverse_lookup(0)
# +
def __make_sentence__(length=420):
seq = X[np.random.randint(0, len(X))].copy().tolist()
ret = ''.join([char_reverse_lookup(int(i)) for i in seq])
for i in tqdm(range(length)):
xhat = np.reshape(seq, (-1, 100)) / len(seq)
yhat = model.predict(xhat)
result = char_reverse_lookup(np.argmax(yhat))
ret += result
seq = seq[1:]
seq.append(np.argmax(yhat))
return ret
model.make_sentence = __make_sentence__
model.make_sentence()
# -
model.fit(X, Y, epochs=50, batch_size=256, shuffle=True)
|
SAD NN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0
# language: julia
# name: julia-1.7
# ---
# # Quantile Regressions
#
# This notebook illustrates quantile regressions.
#
# As an alternative, consider the [QuantileRegressions.jl](https://github.com/pkofod/QuantileRegressions.jl) package.
# ## Load Packages and Extra Functions
# +
using LinearAlgebra, Printf, DelimitedFiles, Statistics
include("jlFiles/printmat.jl")
# +
using Plots
#pyplot(size=(600,400)) #pyplot() or gr()
gr(size=(480,320))
default(fmt = :png)
# -
# ## Loading Data
# +
xx = readdlm("Data/FFdFactors.csv",',',skipstart=1)
xx = xx[:,2] #equity market excess returns
y = xx[2:end] #dependent variable
x = xx[1:end-1] #regressor in an AR(1)
T = size(y,1)
println("Sample size: $T")
# -
# ## A Function for Quantile Regressions
#
# The next cell defines a function `QuanRegrIRLSPs` which estimates a quantile regression. It uses a simple iterative (re-)weighted least squares (IRLS) approach. It can be shown that a linear programming approach gives almost identical results for this data set. However, it seems that the iterative approach is quicker in really large data sets.
#
# The subsequent cells calculate and show the results. The calculations take some time.
# +
"""
QuanRegrIRLSPs(y,x,q=0.5)
Estimate a quantile regression for quantile `q`. The outputs are the point estimates
and three different variance-covariance matrices of the estimates.
# Input
- `y::Vector`: T vector, dependent variable
- `x::Matrix`: TXK, regressors (including any constant)
- `q::Number`: quantile to estimate at, 0<q<1
- `prec::Float64`: convergence criterion, 1e-8
- `epsu::Float64`: lower bound on 1/weight, 1e-6
- `maxiter::Int`: maximum number of iterations, 1000
# Output
- `theta::Vector`: K vector, estimated coefficients
- `vcv::Matrix`: KxK, traditional covariance matrix
- `vcv2::Matrix`: KxK, Powell (1991) covariance matrix
- `vcv3::Matrix`: KxK, Powell (1991) covariance matrix, uniform
"""
function QuantRegrIRLSPs(y,x::Matrix,q=0.5;prec=1e-8,epsu=1e-6,maxiter=1000)
(T,K) = size(x)
xw = copy(x)
(b_old,b,u,iter) = (zeros(K),fill(1e+6,K) .+ prec,zeros(T),0)
while maximum(abs,b - b_old) > prec
copyto!(b_old, b)
b .= (xw'*x)\(xw'*y)
u .= y - x*b
#u .= ifelse.(u.>0,1-q,q).*abs.(u) #as in Python code, divide loss fn by q(1-q) to get it
u .= ifelse.(u.>0,1/q,1/(1-q)).*abs.(u) #abs(u)/q if u>0, abs(u)/(1-q) if u<0
u .= max.(u,epsu) #not smaller than epsu
xw .= x./u
iter = iter + 1
if iter > maxiter
@warn("$iter > maxiter")
b = NaN
break
end
end
res = y - x*b
D = x'x/T
h = 1.06*std(res)/T^0.2 #Silverman (1986) recommendation
fx = exp.(-0.5*((res/h).^2))./(h*sqrt(2*pi)) #Green 7th ed, using N() kernel
f0 = mean(fx)
C = f0*x'x/T
C_1 = inv(C)
vcv = q*(1-q)*C_1*D*C_1/T #variance-covariance matrix
C = (fx.*x)'x/T #Wooldrige 2dn ed, Powell 1991
C_1 = inv(C) #but with Gaussian kernel
vcv2 = q*(1-q)*C_1*D*C_1/T #caputures (x,res) dependence
fx = (abs.(res) .<= h/2)/h #Wooldrige 2nd ed, Powell 1991
C = (fx.*x)'x/T #uniform kernel
C_1 = inv(C)
vcv3 = q*(1-q)*C_1*D*C_1/T
return b, vcv, vcv2, vcv3
end
# +
xGrid = quantile(x,0.01:0.01:0.99) #quantiles of the regressor, for plots
xGrid = [ones(size(xGrid)) xGrid]
qM = [0.01,0.05,0.25,0.5,0.75,0.95,0.99] #quantiles
cx = [ones(T) x] #[constant regressors]
bM = fill(NaN,length(qM),2) #to store regression coeffs
qPred = fill(NaN,length(qM),size(xGrid,1)) #to store predicted values
for i = 1:length(qM)
#local b_q #local/global is needed in script
b_q = QuantRegrIRLSPs(y,cx,qM[i])[1]
bM[i,:] = b_q
qPred[i,:] = xGrid*b_q
end
printblue("quantile regression coefs:\n")
printmat(bM,colNames=["c","slope"],rowNames=string.(qM),cell00="quantile")
printred("\nThe function QuantRegrIRLSPs also outputs different variance-covariance matrices. Compare them.")
# +
lab = permutedims(["quantile $(qM[i])" for i=1:length(qM)])
p1 = plot( xGrid[:,2],qPred',
label = lab,
xlabel = "lagged return",
ylabel = "fitted return",
title = "Fitted values for various quantiles" )
display(p1)
# -
|
Ch24_QuantileRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inference example for trained 2D U-Net model on BraTS.
# Takes a trained model and performs inference on a few validation examples.
# +
import sys
import platform
import os
print("Python version: {}".format(sys.version))
print("{}".format(platform.platform()))
# +
def test_intel_tensorflow():
"""
Check if Intel version of TensorFlow is installed
"""
import tensorflow as tf
print("We are using Tensorflow version {}".format(tf.__version__))
major_version = int(tf.__version__.split(".")[0])
if major_version >= 2:
from tensorflow.python import _pywrap_util_port
print("Intel-optimizations (DNNL) enabled:", _pywrap_util_port.IsMklEnabled())
else:
print("Intel-optimizations (DNNL) enabled:", tf.pywrap_tensorflow.IsMklEnabled())
test_intel_tensorflow()
# -
saved_model_dir = "./output/2d_unet_decathlon"
# +
# Create output directory for images
png_directory = "inference_examples"
if not os.path.exists(png_directory):
os.makedirs(png_directory)
model_filename = os.path.join(saved_model_dir)
# -
# #### Define the DICE coefficient and loss function
# The Sørensen–Dice coefficient is a statistic used for comparing the similarity of two samples. Given two sets, X and Y, it is defined as
#
# \begin{equation}
# dice = \frac{2|X\cap Y|}{|X|+|Y|}
# \end{equation}
# +
import numpy as np
def calc_dice(target, prediction, smooth=0.0001):
"""
Sorensen Dice coefficient
"""
prediction = np.round(prediction)
numerator = 2.0 * np.sum(target * prediction) + smooth
denominator = np.sum(target) + np.sum(prediction) + smooth
coef = numerator / denominator
return coef
def calc_soft_dice(target, prediction, smooth=0.0001):
"""
"""
numerator = 2.0 * np.sum(target * prediction) + smooth
denominator = np.sum(target) + np.sum(prediction) + smooth
coef = numerator / denominator
return coef
# -
# ## Inference Time!
# Inferencing in this example can be done in 3 simple steps:
# 1. Load the data
# 1. Load the Keras model
# 1. Perform a `model.predict` on an input image (or set of images)
# #### Step 1 : Load data
# +
data_path = "../data/decathlon/Task01_BrainTumour/2D_model/"
saved_model_dir = "./output/2d_unet_decathlon"
crop_dim=128 # -1 = Original resolution (240)
batch_size = 128
seed=816 # Change this to see different examples in the test dataset
# -
# # Challenge A: Complete the code
#
# Fill in the dataloader parameters for the testing dataset
# +
from dataloader import DatasetGenerator
# TODO: Fill in the parameters for the dataset generator to return the `testing` data
ds_testing = DatasetGenerator(...)
# -
# #### Step 2 : Load the model
# +
from model import unet
from tensorflow import keras as K
model = K.models.load_model(saved_model_dir, compile=False, custom_objects=unet().custom_objects)
# -
# #### Step 3: Perform prediction on some images.
# The prediction results will be saved in the output directory for images, which is defined by the `png_directory` variable.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_results(ds, idx):
dt = ds.get_dataset().take(1).as_numpy_iterator() # Get some examples (use different seed for different samples)
plt.figure(figsize=(10,10))
for img, msk in dt:
plt.subplot(1, 3, 1)
plt.imshow(img[idx, :, :, 0], cmap="bone", origin="lower")
plt.title("MRI {}".format(idx), fontsize=20)
plt.subplot(1, 3, 2)
plt.imshow(msk[idx, :, :], cmap="bone", origin="lower")
plt.title("Ground truth", fontsize=20)
plt.subplot(1, 3, 3)
# Predict using the TensorFlow model
prediction = model.predict(img[[idx]])
plt.imshow(prediction[0,:,:,0], cmap="bone", origin="lower")
plt.title("Prediction\nDice = {:.4f}".format(calc_dice(msk[idx, :, :], prediction)), fontsize=20)
plt.savefig(os.path.join(png_directory, "prediction_{}.png".format(idx)))
# -
plot_results(ds_testing, 11)
plot_results(ds_testing, 17)
plot_results(ds_testing, 23)
plot_results(ds_testing, 56)
plot_results(ds_testing, 89)
plot_results(ds_testing, 101)
plot_results(ds_testing, 119)
# # Can we perform inference even faster? Hmm..
# Let's find out. Move on the the next tutorial section.
# *Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License. SPDX-License-Identifier: EPL-2.0*
# *Copyright (c) 2019-2020 Intel Corporation*
|
2D/02_Inference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spooky Author Identification
# +
import pandas as pd
pd.set_option('max_columns', None)
train = pd.read_csv("data/train.zip", index_col=['id'])
test = pd.read_csv("data/test.zip", index_col=['id'])
sample_submission = pd.read_csv("data/sample_submission.zip", index_col=['id'])
print(train.shape, test.shape, sample_submission.shape)
print(set(train.columns) - set(test.columns))
# -
train.head()
test.head()
sample_submission.head()
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import cross_val_score
# +
pipeline = Pipeline([
('features', CountVectorizer()),
('clf', LinearSVC())
])
cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3)
# +
from sklearn.ensemble import RandomForestClassifier
pipeline = Pipeline([
('features', CountVectorizer()),
('clf', RandomForestClassifier())
])
cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3)
# -
cross_val_score(pipeline, train.text, train.author,
cv=3, n_jobs=3, scoring='neg_log_loss')
# +
from sklearn.linear_model import LogisticRegression
pipeline = Pipeline([
('features', CountVectorizer()),
('clf', LogisticRegression())
])
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# -
import nltk
# nltk.download('stopwords')
stopwords = nltk.corpus.stopwords.words('english')
print(len(stopwords))
print(stopwords)
# +
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer, HashingVectorizer
params_count_word = {"features__ngram_range": [(1,1), (1,2), (1,3)],
"features__analyzer": ['word'],
"features__max_df":[1.0, 0.9, 0.8, 0.7, 0.6, 0.5],
"features__min_df":[2, 3, 5, 10],
"features__lowercase": [False, True],
"features__stop_words": [None, stopwords]}
params_count_char = {"features__ngram_range": [(1,4), (1,5), (1,6)],
"features__analyzer": ['char'],
"features__max_df":[1.0, 0.9, 0.8, 0.7, 0.6, 0.5],
"features__min_df":[2, 3, 5, 10],
"features__lowercase": [False, True],
"features__stop_words": [None, stopwords]}
# +
import numpy as np
def report(results, n_top=5):
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
# +
from sklearn.model_selection import RandomizedSearchCV
from sklearn.metrics import log_loss
def random_search():
params = {
"clf__C": [0.01, 0.1, 0.3, 1, 3, 10],
"clf__class_weight": [None, 'balanced']
}
params.update(params_count_word)
pipeline = Pipeline([
('features', CountVectorizer()),
('clf', LogisticRegression())
])
random_search = RandomizedSearchCV(pipeline, param_distributions=params,
scoring='neg_log_loss',
n_iter=20, cv=3, n_jobs=4)
random_search.fit(train.text, train.author)
report(random_search.cv_results_)
# random_search()
# +
from sklearn.naive_bayes import MultinomialNB
def random_search():
params = {
"clf__alpha": [0.01, 0.1, 0.5, 1, 2]
}
params.update(params_count_word)
pipeline = Pipeline([
('features', TfidfVectorizer()),
('clf', MultinomialNB())
])
random_search = RandomizedSearchCV(pipeline, param_distributions=params,
scoring='neg_log_loss',
n_iter=20, cv=3, n_jobs=4)
random_search.fit(train.text, train.author)
report(random_search.cv_results_)
# random_search() # Предишния най-добър резултат: -0.469
# -
explore = train.copy()
# +
from nltk.stem import WordNetLemmatizer
from nltk.stem import PorterStemmer
stem = PorterStemmer()
explore['stemmed'] = explore.text.apply(lambda t: " ".join([stem.stem(w) for w in t.split()]))
# +
def random_search():
params = {
"clf__alpha": [0.001, 0.005, 0.01, 0.05, 0.1, 0.3]
}
params.update(params_count_word)
pipeline = Pipeline([
('features', TfidfVectorizer()),
('clf', MultinomialNB())
])
random_search = RandomizedSearchCV(pipeline, param_distributions=params,
scoring='neg_log_loss',
n_iter=20, cv=3, n_jobs=4)
random_search.fit(explore.stemmed, train.author)
report(random_search.cv_results_)
random_search() # -0.423
# +
pipeline = Pipeline([
('features', TfidfVectorizer(ngram_range=(1, 2), min_df=2,
max_df=0.8, lowercase=False)),
('clf', MultinomialNB(alpha=0.01))
])
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# -
# Да пробваме с най-добрите параметри, които сегашното пускане на Random Search намери:
# +
pipeline = Pipeline([
('features', TfidfVectorizer(ngram_range=(1, 3), min_df=2,
max_df=0.5, lowercase=False, analyzer='word')),
('clf', MultinomialNB(alpha=0.1))
])
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# -
import time
from sklearn.model_selection import GridSearchCV
def grid_search(text, params):
pipeline = Pipeline([
('features', TfidfVectorizer()),
('clf', MultinomialNB())
])
search = GridSearchCV(pipeline, param_grid=params,
scoring='neg_log_loss', cv=3, n_jobs=6)
search.fit(text, train.author)
report(search.cv_results_)
results = pd.DataFrame(search.cv_results_)
save_filename = 'grid_search_results_' + str(time.time())
print('Saving grid search results to', save_filename)
results.to_csv(save_filename)
# +
start_time = time.time()
params = {
"features__ngram_range": [(1,1)],#, (1,2), (1,3)],
"features__analyzer": ['word'],
"features__max_df":[1.0, 0.9, 0.8, 0.7, 0.6, 0.5],
"features__min_df":[1, 2, 3, 5, 10],
"features__lowercase": [False, True],
"features__stop_words": [None],
"features__norm": ['l1', 'l2', None],
"clf__alpha": [0.001, 0.005, 0.01, 0.05, 0.1, 0.3]
}
# grid_search(explore.stemmed, params)
print("--- %s seconds ---" % (time.time() - start_time))
# -
# Much hope, no results.. За съжаление след много мъка не успях да подкарам Grid Search да се изпълни с толкова много параматри без да ми свърши RAM-та. Ще пробвам отново randomized search просто с повече итерации.
import time
from sklearn.model_selection import GridSearchCV
def random_search_with_save(text, params, n_iter):
pipeline = Pipeline([
('features', TfidfVectorizer()),
('clf', MultinomialNB())
])
search = RandomizedSearchCV(pipeline, param_distributions=params,
scoring='neg_log_loss', cv=3, n_jobs=6, n_iter=n_iter)
search.fit(text, train.author)
report(search.cv_results_)
results = pd.DataFrame(search.cv_results_)
save_filename = 'random_search_results_' + str(time.time())
print('Saving random search results to', save_filename)
results.to_csv(save_filename)
# +
start_time = time.time()
params = {
"features__ngram_range": [(1,1), (1,2), (1,3), (1,4), (1,5)],
"features__analyzer": ['word'],
"features__max_df":[1.0, 0.9, 0.8, 0.7, 0.6, 0.5],
"features__min_df":[1, 2, 3, 5, 10],
"features__lowercase": [False, True],
"features__stop_words": [None],
"features__norm": ['l1', 'l2', None],
"clf__alpha": [0.001, 0.005, 0.01, 0.05, 0.1, 0.3]
}
for iteration in range(50):
print('\n------------- ITERATION', iteration, '-------------')
random_search_with_save(explore.stemmed, params, n_iter=20)
print("--- %s seconds ---" % (time.time() - start_time))
# -
saved_results = pd.read_csv('random_search_results_1512419845.3117762')
report(saved_results)
# Да пробваме с новонамерените най-добри параметри:
# +
pipeline = Pipeline([
('features', TfidfVectorizer(ngram_range=(1, 2), min_df=1,
max_df=0.9, lowercase=True, analyzer='word', norm='l2')),
('clf', MultinomialNB(alpha=0.05))
])
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# -
# Приятно подобрение. Имам нова идея за Grid Search, нека пробваме и нея и да събмитнем.
import time
from sklearn.model_selection import GridSearchCV
def grid_search(text, params):
pipeline = Pipeline([
('features', TfidfVectorizer()),
('clf', MultinomialNB())
])
search = GridSearchCV(pipeline, param_grid=params,
scoring='neg_log_loss', cv=3, n_jobs=6)
search.fit(text, train.author)
# report(search.cv_results_)
results = pd.DataFrame(search.cv_results_)
save_filename = 'grid_search_results_' + str(time.time())
print('Saving grid search results to', save_filename)
results.to_csv(save_filename)
return results
# +
import numpy as np
def report_best_of_all(results, n_top=5):
results = results.sort_values(by='mean_test_score', ascending=False)
candidates = results.head(5)
for pos_best, (_, candidate) in enumerate(candidates.iterrows()):
print("Model with rank: {0}".format(pos_best+1))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
candidate['mean_test_score'],
candidate['std_test_score']))
print("Parameters: {0}".format(candidate['params']))
print("")
# +
#with stemmed text
def grid_search_ngram_range(ngram_range):
start_time = time.time()
max_dfs = [1.0, 0.9, 0.8, 0.7, 0.6, 0.5]
min_dfs = [1, 2, 3, 5, 10]
params = {
"features__analyzer": ['word'],
"features__lowercase": [False, True],
"features__stop_words": [None],
"features__norm": ['l1', 'l2', None],
"clf__alpha": [0.001, 0.005, 0.01, 0.05, 0.1, 0.3, 1, 2]
}
results = []
for max_df in max_dfs:
for min_df in min_dfs:
print(ngram_range, max_df, min_df)
params["features__ngram_range"] = [ngram_range]
params["features__max_df"] = [max_df]
params["features__min_df"] = [min_df]
result = grid_search(explore.stemmed, params)
results.append(result)
print("--- %s seconds ---" % (time.time() - start_time))
results_concatenated = pd.concat(results, ignore_index=True)
report_best_of_all(results_concatenated)
print("--- %s seconds ---" % (time.time() - start_time))
grid_search_ngram_range((1,1))
# -
grid_search_ngram_range((1,2))
grid_search_ngram_range((1,3))
grid_search_ngram_range((1,4))
grid_search_ngram_range((1,5))
# +
pipeline = Pipeline([
('features', TfidfVectorizer(ngram_range=(1, 2), min_df=1,
max_df=0.5, lowercase=False, analyzer='word', norm='l2')),
('clf', MultinomialNB(alpha=0.05))
])
print(cross_val_score(pipeline, explore.stemmed, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, explore.stemmed, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# +
def grid_search_ngram_range_original_text(ngram_range):
start_time = time.time()
max_dfs = [1.0, 0.9, 0.8, 0.7, 0.6, 0.5]
min_dfs = [1, 2, 3]
params = {
"features__analyzer": ['word'],
"features__lowercase": [False, True],
"features__stop_words": [None],
"features__norm": ['l2'],
"clf__alpha": [0.001, 0.005, 0.01, 0.05, 0.1, 0.3, 1]
}
results = []
for max_df in max_dfs:
for min_df in min_dfs:
print(ngram_range, max_df, min_df)
params["features__ngram_range"] = [ngram_range]
params["features__max_df"] = [max_df]
params["features__min_df"] = [min_df]
result = grid_search(train.text, params)
results.append(result)
print("--- %s seconds ---" % (time.time() - start_time))
results_concatenated = pd.concat(results, ignore_index=True)
report_best_of_all(results_concatenated)
print("--- %s seconds ---" % (time.time() - start_time))
grid_search_ngram_range_original_text((1,2))
# +
pipeline = Pipeline([
('features', TfidfVectorizer(ngram_range=(1, 2), min_df=1,
max_df=0.5, lowercase=False, analyzer='word', norm='l2')),
('clf', MultinomialNB(alpha=0.05))
])
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3))
print(cross_val_score(pipeline, train.text, train.author, cv=3, n_jobs=3,
scoring='neg_log_loss'))
# -
# Получихме същите параметри и дори по-добър резултат.
pipeline = pipeline.fit(train.text, train.author)
test_predictions = pipeline.predict_proba(test.text)
pipeline.classes_
submit_file = pd.DataFrame(test_predictions, columns=['EAP', 'HPL', 'MWS'], index=test.index)
submit_file.head(10)
submit_file.to_csv("predictions.csv")
|
spooky/exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from dotenv import load_dotenv, find_dotenv
from os.path import join, dirname, basename, exists, isdir
### Load environmental variables from the project root directory ###
# find .env automagically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
# now you can get the variables using their names
# Check whether a network drive has been specified
DATABASE = os.environ.get("NETWORK_URL")
if DATABASE == 'None':
pass
else:
pass
#mount network drive here
# set up directory pathsa
CURRENT_DIR = os.getcwd()
PROJ = dirname(dotenv_path) # project root directory
DATA = join(PROJ, 'data') #data directory
RAW_EXTERNAL = join(DATA, 'raw_external') # external data raw directory
RAW_INTERNAL = join(DATA, 'raw_internal') # internal data raw directory
INTERMEDIATE = join(DATA, 'intermediate') # intermediate data directory
FINAL = join(DATA, 'final') # final data directory
RESULTS = join(PROJ, 'results') # output directory
FIGURES = join(RESULTS, 'figures') # figure output directory
PICTURES = join(RESULTS, 'pictures') # picture output directory
# make folders specific for certain data
folder_name = ''
if folder_name != '':
#make folders if they don't exist
if not exists(join(RAW_EXTERNAL, folder_name)):
os.makedirs(join(RAW_EXTERNAL, folder_name))
if not exists(join(INTERMEDIATE, folder_name)):
os.makedirs(join(INTERMEDIATE, folder_name))
if not exists(join(FINAL, folder_name)):
os.makedirs(join(FINAL, folder_name))
print('Standard variables loaded, you are good to go!')
# +
import cobra
import os
import pandas as pd
import cameo
import wget
import ssl
from scipy.stats import pearsonr
#E. coli model:
ssl._create_default_https_context = ssl._create_unverified_context
wget.download("https://raw.githubusercontent.com/BenjaSanchez/notebooks/master/e_coli_simulations/eciML1515.xml")
eColi_Model = cobra.io.read_sbml_model("eciML1515.xml")
os.remove("eciML1515.xml")
# proteomics data:
proteomics_dataset = f"{INTERMEDIATE}/proteomics/proteomics_concentrations.csv"
weights_location = f"{INTERMEDIATE}/proteomics/proteomics_masses.csv"
# -
from collections import namedtuple
from cobra.medium.boundary_types import find_external_compartment
from cobra.io.dict import reaction_to_dict
import pandas as pd
import numpy as np
from simulations.modeling.driven import (
adjust_fluxes2model,
flexibilize_proteomics,
minimize_distance,
)
def reset_real_proteomics(proteomics_dataset):
'''loads set of proteomics data from the provided dataset file into dict of lists'''
data = pd.read_csv(proteomics_dataset, index_col="UP") # yeast
data_dict = {}
for i in range(0,data.shape[1], 3):
uncertainty = data.iloc[:,i:i+3].std(axis=1)
uncertainty_name = data.columns[i]+ "_uncertainty"
data[uncertainty_name] = uncertainty
data_dict[data.columns[i]] = [{'identifier':data.index[j], 'measurement':data.iloc[j,i], 'uncertainty':data[uncertainty_name][j] }\
for j in range(0, len(data.iloc[:,i]))]
data_dict[data.columns[i+1]] = [{'identifier':data.index[j], 'measurement':data.iloc[j,i+1], 'uncertainty':data[uncertainty_name][j] }\
for j in range(0, len(data.iloc[:,i+1]))]
data_dict[data.columns[i+2]] = [{'identifier':data.index[j], 'measurement':data.iloc[j,i+2], 'uncertainty':data[uncertainty_name][j] }\
for j in range(0, len(data.iloc[:,i+2]))]
return data_dict
# +
proteomics_data = reset_real_proteomics(proteomics_dataset)
growth_rates = pd.read_csv(f"{RAW_INTERNAL}/proteomics/growth_conditions.csv")
growth_rates = growth_rates.drop(growth_rates.columns.difference(['Growth condition','Growth rate (h-1)', 'Stdev']), 1)
growth_rates = growth_rates.drop([0,1], axis=0)
# -
# setup
exchange_reaction = "Pyruvate"
exchange_reaction_lowercase = "pyruvate"
# +
def find_exchange_rxn(compound, model):
exchange_reactions = [i for i in model.reactions if "EX" in i.id]
compound_ex_rxn = [i for i in exchange_reactions if compound in i.name]
compound_ex_rxn = [i for i in compound_ex_rxn if len(list(i._metabolites.keys())) == 1 \
& (list(i._metabolites.values())[0] == 1.0) \
& (list(i._metabolites.keys())[0].name == compound + " [extracellular space]")]
return compound_ex_rxn
# minimal medium with pyruvate
pyruvate_growth_rate = list(growth_rates['Growth rate (h-1)'].loc[growth_rates['Growth condition'] == "Pyruvate"])[0]
pyr_model = eColi_Model.copy()
pyr_medium = pyr_model.medium
pyr_medium.pop("EX_glc__D_e_REV", None)
# find Pyruvate
pyr_ex = find_exchange_rxn("Pyruvate", eColi_Model)
print(pyr_ex)
pyr_medium[f'{pyr_ex[0].id}'] = 10
pyr_model.medium = pyr_medium
# pyr_model.medium = minimal_medium(pyr_model).to_dict()
print(pyr_model.optimize())
model = pyr_model
# +
# Flexibilize proteomics
ec_model_1 = model
ec_model_2 = model
ec_model_3 = model
# first
print("Number of proteins originally: ", len(proteomics_data[exchange_reaction_lowercase]))
growth_rate = {"measurement":float(list(growth_rates['Growth rate (h-1)'].loc[growth_rates['Growth condition'] == exchange_reaction])[0]),\
"uncertainty":float(list(growth_rates['Stdev'].loc[growth_rates['Growth condition'] == exchange_reaction])[0])}
new_growth_rate, new_proteomics, warnings = flexibilize_proteomics(ec_model_1, "BIOMASS_Ec_iML1515_core_75p37M", growth_rate, proteomics_data[exchange_reaction_lowercase], [])
print("Number of proteins incorporated: ", len(new_proteomics))
# first
print("Number of proteins originally: ", len(proteomics_data[exchange_reaction_lowercase + "2"]))
growth_rate = {"measurement":float(list(growth_rates['Growth rate (h-1)'].loc[growth_rates['Growth condition'] == exchange_reaction])[0]),\
"uncertainty":float(list(growth_rates['Stdev'].loc[growth_rates['Growth condition'] == exchange_reaction])[0])}
new_growth_rate, new_proteomics, warnings = flexibilize_proteomics(ec_model_2, "BIOMASS_Ec_iML1515_core_75p37M", growth_rate, proteomics_data[exchange_reaction_lowercase + "1"], [])
print("Number of proteins incorporated: ", len(new_proteomics))
# first
print("Number of proteins originally: ", len(proteomics_data[exchange_reaction_lowercase + "2"]))
growth_rate = {"measurement":float(list(growth_rates['Growth rate (h-1)'].loc[growth_rates['Growth condition'] == exchange_reaction])[0]),\
"uncertainty":float(list(growth_rates['Stdev'].loc[growth_rates['Growth condition'] == exchange_reaction])[0])}
new_growth_rate, new_proteomics, warnings = flexibilize_proteomics(ec_model_3, "BIOMASS_Ec_iML1515_core_75p37M", growth_rate, proteomics_data[exchange_reaction_lowercase + "2"], [])
print("Number of proteins incorporated: ", len(new_proteomics))
# +
weights = pd.read_csv(weights_location, index_col = "UP")
# usages of ac proteins
#solution = pyr_model.optimize()
# pyr model uages
def get_usages(prot_int_model, weights):
# get the usages of a model integrated with proteomics
try:
solution = cobra.flux_analysis.pfba(prot_int_model)
except:
print("used normal fba")
solution = prot_int_model.optimize()
abs_usages = pd.Series()
perc_usages = pd.Series()
mass_usages = 0
non_mass_proteins = []
for reaction in prot_int_model.reactions:
if reaction.id.startswith("prot_"):
prot_id = reaction.id.replace("prot_","")
prot_id = prot_id.replace("_exchange","")
abs_usage = solution.fluxes[reaction.id]
abs_usages = abs_usages.append(pd.Series({prot_id:abs_usage}))
perc_usage = solution.fluxes[reaction.id]/reaction.upper_bound
perc_usages = perc_usages.append(pd.Series({prot_id:perc_usage}))
try:
if perc_usage <= 100:
mass_usages += perc_usage/100 * weights[prot_id]
except:
non_mass_proteins.append(prot_id)
return abs_usages, perc_usages, mass_usages, non_mass_proteins
#
abs_usages_1, perc_usages_1, mass_usage_1, non_mass_proteins_1 = get_usages(ec_model_1, weights[f"{exchange_reaction_lowercase}"])
abs_usages_2, perc_usages_2, mass_usage_2, non_mass_proteins_2 = get_usages(ec_model_2, weights[f"{exchange_reaction_lowercase}.1"])
abs_usages_3, perc_usages_3, mass_usage_3, non_mass_proteins_3 = get_usages(ec_model_3, weights[f"{exchange_reaction_lowercase}.2"])
# -
len(non_mass_proteins_1)
print("Mass of Proteins total: ", sum(weights[f"{exchange_reaction}"]))
print("Mass actually used: ", sum(weights[f"{exchange_reaction}"])*(mass_usage_1/sum(weights[f"{exchange_reaction}"])))
abs_usages_df = pd.DataFrame({f"{exchange_reaction_lowercase}": perc_usages_1, f"{exchange_reaction_lowercase}.1": perc_usages_2, f"{exchange_reaction_lowercase}.2": perc_usages_3})
abs_usages_df.to_csv(f"{FINAL}/abs_usages_gecko/{exchange_reaction_lowercase}")
# # Masses
#
# Masses that are actually used seem very low, at 0,9 %
#
# What should I actually do here?
#
# Total protein mass: 117633655349 Dalton
# +
import numpy as np; np.random.seed(42)
import matplotlib.pyplot as plt
import seaborn as sns
df = perc_usages_1.to_frame()
df["perc_usages_2"] = perc_usages_2
df["perc_usages_3"] = perc_usages_3
df.columns = ["Measurement 1", "Measurement 2", "Measurement 3"]
sns.boxplot(x="variable", y="value", data=pd.melt(df[(df > 0) & (df < 100)]))
plt.xlabel('Measurements')
plt.ylabel('Usage of measurement in %')
plt.title('% usage of proteins per ec simulation ')
plt.savefig(f'{FIGURES}/ec_incorporation_perc_usage_box_pyr')
plt.show()
# +
#df['pct'] = df['Location'].div(df.groupby('Hour')['Location'].transform('sum'))
#g = sns.FacetGrid(df, row="pct", hue="pct", aspect=15, height=.5, palette=pal)
perc_incorporation_pyr = pd.melt(df[(df > 0) & (df < 100)])
# Method 1: on the same Axis
sns.distplot( df[(df > 0) & (df < 100)].iloc[:,0] , color="skyblue", label="1", kde=False)
sns.distplot( df[(df > 0) & (df < 100)].iloc[:,1], color="red", label="2", kde=False)
sns.distplot( df[(df > 0) & (df < 100)].iloc[:,2], color="green", label="3", kde=False)
# +
from sklearn.preprocessing import StandardScaler
# standardize data for pca
# #features = ['sepal length', 'sepal width', 'petal length', 'petal width']# Separating out the features
pca_df_all_proteomics_and_pyr = pd.read_csv(proteomics_dataset, index_col="UP").loc[df.index,:]
pca_df_all_proteomics_and_pyr['pyr_1'] = abs_usages_1
pca_df_all_proteomics_and_pyr = pca_df_all_proteomics_and_pyr.T.dropna(axis='columns')
x = pca_df_all_proteomics_and_pyr.values
x = StandardScaler().fit_transform(x)
# run pca
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(x)
principalDf = pd.DataFrame(data = principalComponents, columns = ['principal component 1', 'principal component 2'])
principalDf.index = pca_df_all_proteomics_and_pyr.index
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Principal Component 1', fontsize = 15)
ax.set_ylabel('Principal Component 2', fontsize = 15)
ax.set_title('2 component PCA with zero values', fontsize = 20)
amount = len(principalDf.index)
for i in range(amount):
c = [float(i)/float(amount), 0.0, float(amount-i)/float(amount)] #R,G,B
ax.scatter(principalDf.loc[principalDf.index[i], 'principal component 1']
, principalDf.loc[principalDf.index[i], 'principal component 2']
, color = c
, s = 50)
ax.scatter(principalDf.loc["pyr_1", 'principal component 1']
, principalDf.loc[principalDf.index[i], 'principal component 2']
, color = "green"
, s = 50)
#ax.legend(pca_df_all_proteomics_and_pyr.index)
ax.grid()
plt.savefig(f'{FIGURES}/')
# +
# standardize data for pca
# #features = ['sepal length', 'sepal width', 'petal length', 'petal width']# Separating out the features
pca_df_all_proteomics_and_pyr = pd.read_csv(proteomics_dataset, index_col="UP").loc[df.index,:]
pca_df_all_proteomics_and_pyr['pyr_1'] = abs_usages_1
pca_df_all_proteomics_and_pyr = pca_df_all_proteomics_and_pyr[pca_df_all_proteomics_and_pyr['pyr_1'] > 0]
pca_df_all_proteomics_and_pyr = pca_df_all_proteomics_and_pyr.T.dropna(axis='columns')
x = pca_df_all_proteomics_and_pyr.values
x = StandardScaler().fit_transform(x)
# run pca
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
principalComponents = pca.fit_transform(x)
principalDf = pd.DataFrame(data = principalComponents, columns = ['principal component 1', 'principal component 2'])
principalDf.index = pca_df_all_proteomics_and_pyr.index
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Principal Component 1', fontsize = 15)
ax.set_ylabel('Principal Component 2', fontsize = 15)
ax.set_title('2 component PCA without zero values', fontsize = 20)
amount = len(principalDf.index)
for i in range(amount):
c = [float(i)/float(amount), 0.0, float(amount-i)/float(amount)] #R,G,B
ax.scatter(principalDf.loc[principalDf.index[i], 'principal component 1']
, principalDf.loc[principalDf.index[i], 'principal component 2']
, color = c
, s = 50)
ax.scatter(principalDf.loc["pyr_1", 'principal component 1']
, principalDf.loc[principalDf.index[i], 'principal component 2']
, color = "green"
, s = 50)
ax.grid()
# -
pd.DataFrame({'pyr_1':abs_usages_1, 'pyr_2':abs_usages_2, 'pyr_3':abs_usages_3}).to_csv(f'{INTERMEDIATE}/proteomics/pyruvate_usages.csv')
|
data_science/code/modeling/pyruvate/ec_incorporation_pyruvate.ipynb
|
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題2.3
;; 平⾯上の⻑⽅形の表現を実装せよ
;; (ヒント:練習問題 2.2を利⽤するといいかもしれない)。
;; そのコンストラクタとセレクタを使って、
;; ある⻑⽅形の外周の⻑さと⾯積を計算する⼿続きを書け。
;; 次に、⻑⽅形の異なる表現を実装せよ。
;; 適切な抽象化の壁を使ってシステムを設計し、
;; 外周と⾯積を求める同じ⼿続きがどちらの表現によっても動くようにできるだろうか。
;; <img src="2.03.png" width="50%">
;;
;; <div style="text-align: center;">回答の条件</div>
;; +
; 点のコンストラクタ
(define (make-point x y)(cons x y))
; セレクタ
(define (x-point p)(car p))
(define (y-point p)(cdr p))
(define (print-point p)
(newline)
(display "(")
(display (x-point p))
(display ",")
(display (y-point p))
(display ")"))
; 線分コンストラクタ
(define (make-segment p1 p2)(cons p1 p2))
; セレクタ
(define (start-segment s)(car s))
(define (end-segment s)(cdr s))
; 中点を返す
(define (midpoint-segment s)
(let ((p1 (start-segment s))
(p2 (end-segment s)))
(let ((x1 (x-point p1))
(y1 (y-point p1))
(x2 (x-point p2))
(y2 (y-point p2)))
(make-point (+ (/ (- x2 x1) 2) x1) (+ (/ (- y2 y1) 2) y1))
)
)
)
;; +
; ⻑⽅形コンストラクタ
; 与えられた線分を対角線とした長方形とする。
(define (make-rect p1 p2)(make-segment p1 p2))
; 補助手続き
(define (point-rect pos r)
(let ((p1 (start-segment r))
(p2 (end-segment r)))
(let ((x1 (x-point p1))
(y1 (y-point p1))
(x2 (x-point p2))
(y2 (y-point p2)))
(
cond ((= pos 0) (make-point (min x1 x2) (max y1 y2))) ; top-left
((= pos 1) (make-point (max x1 x2) (max y1 y2))) ; top-right
((= pos 2) (make-point (max x1 x2) (min y1 y2))) ; bottom-right
((= pos 3) (make-point (min x1 x2) (min y1 y2))) ; bottom-left
(else (error "pos error." ()))
)
)
)
)
; セレクタ(長方形の幅を返す)
(define (width-rect r)
(let ((p1 (point-rect 0 r))
(p2 (point-rect 1 r)))
(- (x-point p2) (x-point p1))
)
)
; セレクタ(長方形の高さを返す)
(define (height-rect r)
(let ((p1 (point-rect 3 r))
(p2 (point-rect 0 r)))
(- (y-point p2) (y-point p1))
)
)
; 面積
(define (area-rect r)
(let ((width (width-rect r))
(height (height-rect r)))
(* width height)
)
)
; 外周
(define (perimeter-rect r)
(let ((width (width-rect r))
(height (height-rect r)))
(+ (* width 2) (* height 2))
)
)
(define p1 (make-point 3 3))
(print-point p1)
(define p2 (make-point 11 5))
(print-point p2)
(define p3 (midpoint-segment (make-segment p1 p2)))
(print-point p3)
(newline)
(define r (make-rect p1 p2))
(display (width-rect r))
(newline)
(display (height-rect r))
(newline)
(display (area-rect r))
(newline)
(display (perimeter-rect r))
(newline)
;; +
; 補助手続き:与えられた線分の、x軸に射影したときの幅を返す。
(define (segment-width s)
(let ((p1 (start-segment s))
(p2 (end-segment s)))
(let ((x1 (x-point p1))
(x2 (x-point p2)))
(abs (- x1 x2))
)
)
)
; 補助手続き:与えられた線分の、y軸に射影したときの幅(高さ)を返す。
(define (segment-height s)
(let ((p1 (start-segment s))
(p2 (end-segment s)))
(let ((y1 (y-point p1))
(y2 (y-point p2)))
(abs (- y1 y2))
)
)
)
; ⻑⽅形コンストラクタ
; 与えられた線分を対角線とした長方形
; 内部では中心点、幅、高さを記憶する。
(define (make-rect p1 p2)
(let ((s (make-segment p1 p2)))
(let ((midpoint (midpoint-segment s))
(width (segment-width s))
(height (segment-height s)))
(cons midpoint (cons width height))
)
)
)
; セレクタ(長方形の幅を返す)
(define (width-rect r)
(car (cdr r))
)
; セレクタ(長方形の高さを返す)
(define (height-rect r)
(cdr (cdr r))
)
(define r (make-rect p1 p2))
(display (width-rect r))
(newline)
(display (height-rect r))
(newline)
(display (area-rect r))
(newline)
(display (perimeter-rect r))
(newline)
;; +
; 別回答(コンストラクタのインターフェースが変わってしまっているので解答に適さない)
; コンストラクタ
; 内部は対角線として記憶する。
(define (make-rect center width height)
(let ((half_width (/ width 2))
(half_height (/ height 2)))
(let ((x1 (- (x-point center) half_width))
(y1 (- (y-point center) half_height))
(x2 (+ (x-point center) half_width))
(y2 (+ (y-point center) half_height)))
(let ((p1 (make-point x1 y1))
(p2 (make-point x2 y2)))
(make-segment p1 p2)
)
)
)
)
; 長方形の幅を返す(元のセレクタと同じ定義)
(define (width-rect r)
(let ((p1 (point-rect 0 r))
(p2 (point-rect 1 r)))
(- (x-point p2) (x-point p1))
)
)
; 長方形の高さを返す(元のセレクタと同じ定義)
(define (height-rect r)
(let ((p1 (point-rect 3 r))
(p2 (point-rect 0 r)))
(- (y-point p2) (y-point p1))
)
)
(define p1 (make-point 3 3))
(print-point p1)
(define p2 (make-point 11 5))
(print-point p2)
(newline)
(define r (make-rect (make-point 7 4) 8 2))
(display (width-rect r))
(newline)
(display (height-rect r))
(newline)
(display (area-rect r))
(newline)
(display (perimeter-rect r))
(newline)
|
exercises/2.03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# Merge multi <NAME>
hans = Contributor.objects.search("Hval").exclude(display_name__contains='Olav')
from utils.merge_model_objects import merge_instances
merge_instances(*list(hans))
hans
# +
import re
def re_find(expression, haystack):
try:
return re.search(expression, haystack).groups()[0]
except AttributeError:
return ''
def get_initials_from_file_name(filename):
return re_find(r'-(?P<initials>[A-Z]+)((-NR)?-?\d+)?(-*\d+-of-\d+)?(_\w{7})?\.\w+$', filename)
# -
def active_photographers(when=None):
photographer = Position.objects.get(title='fotograf')
stints = Stint.objects.filter(position=photographer)
if when:
stints = stints.active(when)
contributors = Contributor.objects.filter(pk__in=stints.values_list('contributor', flat=True))
return contributors
import logging
logger = logging.getLogger()
logger.setLevel(logging.ERROR)
from utils.disconnect_signals import disconnect_signals
# Byline images
ImageFile.objects.filter(source_file__startswith='byline-photo').update(category=ImageFile.PROFILE)
# +
# Illustrations
kvammen = Contributor.objects.search('<NAME>')[0]
kvammen.initials = 'ANK'
kvammen.save()
kvammen_illus= ImageFile.objects.filter(source_file__contains='-ANK.jpg')
kvammen_illus.update(category=ImageFile.ILLUSTRATION, contributor=kvammen)
hovland = Contributor.objects.search('<NAME>')[0]
hovland_illus = ImageFile.objects.filter(description__contains='vind Hovland')
hovland_illus.update(category=ImageFile.ILLUSTRATION, contributor=hovland)
# -
# fix some contributor's initials
sjur = Contributor.objects.search('<NAME>').first()
sjur.initials='SGS'
sjur.save()
ingrid = Contributor.objects.search('<NAME>').first()
ingrid.initials = 'IDR'
ingrid.save()
vilde = Contributor.objects.search('<NAME>').first()
vilde.initials = 'VIB'
vilde.save()
aaah = Contributor.objects.search('<NAME>')[0]
ImageFile.objects.filter(contributor=None, source_file__contains='AAAH').update(contributor=aaah)
# Initials lookup dictionary
Contributor.objects.filter(initials=None).update(initials='')
lookup = {cn.initials: cn for cn in Contributor.objects.exclude(initials='') if 1 < len(cn.initials) < 4}
lookup['HHH'] = lookup['HDH']
lookup['TWM'] = lookup['TM']
lookup['AAGS'] = lookup['AS']
lookup['AHA'] = lookup['AH']
lookup['PP'] = lookup['PPB']
lookup['LLIF'] = lookup['LIF']
lookup
# +
def guess_photo_contributor(image_file):
"""Use various methods to determine probable contributor of image"""
name, contributor = None, None
if image_file.contributor:
contributor = image_file.contributor
logger.debug('already assigned')
return contributor
if image_file.exif:
# exif data lookup
exif = image_file.exif
names = [exif.artist, re_find(r'[fF]oto:(.*)^', exif.description)]
name = sorted(names, key=len)[-1]
if name and not contributor:
try:
contributor = Contributor.objects.get(display_name=name)
logger.debug(f'found name {name}')
except Contributor.DoesNotExist:
matches = Contributor.objects.search(name)
if matches.count() == 1:
contributor = matches[0]
logger.debug(f'fuzzy search for name {name} -> {contributor}')
initials = get_initials_from_file_name(image_file.original.name)
if initials and not contributor:
try:
contributor = active_photographers(when=image_file.created).get(initials=initials)
logger.debug(f'found initials {initials}')
except Contributor.DoesNotExist:
# try from lookup list
contributor = lookup.get(initials)
except Contributor.MultipleObjectsReturned:
pass
if not contributor:
# look at bylines for stories
sim = image_file.storyimage_set.first()
if sim:
photo_byline = sim.parent_story.byline_set.filter(credit='photo')
if photo_byline.count() == 1:
contributor = photo_byline.first().contributor
logger.debug(f'found photo byline {photo_byline.first()}')
if not contributor and (name or initials):
logger.warn(f'{image_file} did not find {name} / {initials}')
return contributor
from tqdm import tqdm
def assign_contributor(qs):
disconnect_signals()
success = []
fail = []
proc = tqdm(qs)
for im in proc:
proc.set_description_str(f'{str(im)[:40]:<40}')
cn = guess_photo_contributor(im)
if cn:
im.contributor = cn
im.category = ImageFile.PHOTO
im.save()
success.append(f'{im.pk:<5} {str(cn):<40} {im}')
else:
fail.append(str(im))
print('OK:')
print('\n'.join(success))
print('\nFAIL:')
print(len(fail))
qs = ImageFile.objects.filter(contributor=None)
print('unassigned', qs.count())
assign_contributor(ImageFile.objects.filter(source_file__contains='OV'))
qs = ImageFile.objects.filter(contributor=None)
print('unassigned', qs.count())
# -
pressefoto = ImageFile.objects.filter(category=ImageFile.UNKNOWN).filter(source_file__icontains='pressebilde')
pressefoto.update(category=ImageFile.EXTERNAL)
# +
# Update photographer initials
def get_initials(name):
name = name.translate(str.maketrans('ÆØÅ', 'AOA'))
return ''.join(re.findall(r'[A-Z]', name))[:5]
def is_photographer(cn):
if cn.stint_set.filter(position__title='fotograf').count():
return True
if cn.byline_set.filter(title='foto').count():
return True
if cn.imagefile_set.count():
return True
else:
return False
for cn in Contributor.objects.all().order_by('display_name'):
if is_photographer(cn):
initials = get_initials(str(cn))
if cn.initials != initials:
cn.initials = initials
cn.save()
print(f'{str(cn.initials):<6} {name:<30} {cn.user or "-"}')
elif cn.initials:
cn.initials = ''
cn.save()
name = str(cn)
# +
fix = {
" Prof. <NAME>": "In<NAME>",
". Translated By <NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "Are <NAME>",
"<NAME>": "<NAME>",
"Aud V. Tønnessen": "Aud V. Tønnessen",
"Aud <NAME>ønnesen": "Aud <NAME>ønnesen",
"August Tekrø": "August Tekrø",
"Aurora Sæverud": "Aurora Sæverud",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>-<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Eva–K<NAME>": "Eva-K<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Gun<NAME>id": "Gun<NAME>",
"<NAME>": "<NAME>",
"Hallvard Østtve<NAME>bogen": "Hallvard Østt<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "Hå<NAME>",
"<NAME>": "Id<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Ine Eriksen Søreide": "Ine Eriksen Søreide",
"Ingeborg <NAME>": "Ingeborg <NAME>",
"Ingeborg <NAME> (Leder": "Ingeborg Mar<NAME>",
"Ingeborg Skjelkvåle Ligaarden": "Ingeborg Skjelkvåle Ligaarden",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Ingrid Aune": "Ingrid Aune",
"Ingrid Eidsheim Daae": "Ingrid E. Daae",
"Ingrid Elise Gipling": "Ingrid Gipling",
"Ingrid <NAME>": "Ingrid F<NAME>aker",
"Ingrid K. Fredriksen": "Ingrid Kvamme Fredriksen",
"Ingrid N. Ekeberg": "Ingrid N. Ekeberg",
"Ingrid Stranger–Thorsen": "Ingrid Stranger-Thorsen",
"<NAME>": "<NAME>",
"Iselin Rud–Goksøyr": "Iselin Rud-Goksøyr",
"Iselin Shaw of Tordarroch": "Iselin Shaw of Tordarroch",
"<NAME>": "<NAME>",
"<NAME> Masterstudent": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Katrine Elida Aaland": "Katrine Elida Aaland",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Lars-Christian U. Talseth": "Lars-Christian U. Talseth",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "L<NAME>ensen",
"<NAME>": "Sus<NAME>ensland",
"Line Ørnes Søndergaard": "Line Ør<NAME>øndergaard",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Liv Rønnaug Lilleåsen": "Liv Rønnaug Bjerke Lilleåsen",
"Liv-Kristin <NAME>": "Liv-Kristin <NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME> Nedrelid": "<NAME>",
"Siri Øverland Eriksen": "Siri Øverland Eriksen",
"<NAME>": "<NAME>",
"Solveig Figenschou": "Solveig Figenschou",
"Solve<NAME>": "Solve<NAME>",
"Solveig Schanke Eikum": "Solveig Schanke Eikum",
"<NAME>": "S<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "St<NAME>",
"<NAME>": "<NAME>",
"Studentparlamenett Uio": "Studentparlamentet Uio",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"The<NAME>": "<NAME>",
"Thea Urdal": "Thea Hå<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Tone C. S. Thorgrimsen": "Tone Celine Sundse<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Translation By <NAME>": "<NAME>",
"Universitas´ Juleølpanel": "Universitas´ Juleølpanel",
"Universitas’ Sommerøltestpanel": "Universitas’ Sommerøltestpanel",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"Åshild Støylen": "Åshild Støylen",
"Åsm<NAME>vsthus": "Å<NAME>",
"<NAME>": "Øistein Ø. Svelle",
"Øyvind Østerås": "Øyvind Østerås",
}
# -
cnn = Contributor.objects.extra(where=["CHAR_LENGTH(aliases) > 20"])
for cn in cnn:
name = cn.display_name
if name in fix:
cn.display_name = fix[name]
cn.aliases = ''
print(f'{name:<30} -> {cn.display_name}')
cn.save()
# Disse to personenen er forskjellige
{"<NAME>": "<NAME>"}
helena = Contributor.objects.get(display_name='<NAME>')
lena = Contributor.objects.get_or_create(display_name='<NAME>')[0]
helena.byline_set.filter(credit='photo').update(contributor=lena)
# remove useless contributors
from django.db.models import Count
allcn = Contributor.objects.order_by('display_name').annotate(bl_count=Count('byline', unique=True), stint_count=Count('stint', unique=True))
for cn in allcn.filter(bl_count=0):
if not cn.email and cn.stint_count == 0:
print(f'{str(cn.pk):<6} {str(cn):<40} {cn.bl_count} {cn.stint_count} {cn.user} {cn.email}')
cn.delete()
# find possible misspelled names
from collections import Counter
cnn = Contributor.objects.extra(where=["CHAR_LENGTH(aliases) > 20"])
for cn in cnn:
aliases = Counter(cn.aliases.strip().split('\n'))
aliases = (f'"{k}" ({v}),' for k, v in aliases.items() if v > 1)
print(f'"{cn}": {"|".join(aliases)}')
|
django/notebooks/commit/Categorize photos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EX 10. News Summarization
#
#
# 텍스트 요약
# 상대적으로 긴 문장으로 짧은 문장으로 변환하는 기법을 텍스트 요약이라고 합니다. Ex 10에서는 뉴스 요약을 해보겠습니다.
#
# 텍스트 요약에는 두 가지 방법, Extractive Summarization, Abstractive Summarization이 있습니다. Extractive의 경우 가장 의미있는 문장을 원문 안에서 추출해 내는 방식이고, Abstractive는 원문을 요약하는 새로운 문장을 만들어내는 방식입니다.
#
# 이 프로젝트에서는 원문과 그에 해당하는 요약문을 주고(지도학습) seq2seq와 Attention을 사용하여 원문 -> 요약문으로의 Abstractive Summarization을 실시하겠습니다. 이는 일반적인 번역과 매우 유사한 과정을 거치는 것입니다.
#
#
#
# 평가문항
# 1. Abstractive 모델 구성을 위한 분석단계, 정제단계, 정규화와 불용어 제거, 데이터셋 분리, 인코딩 과정이 빠짐없이 체계적으로 진행되었다.
#
#
# 2. 모델학습이 안정적으로 수렴되었음을 그래프를 통해 확인하였으며, 실제 요약문과 유사한 요약문장을 얻을 수 있었다.
#
#
# 3. Abstractive 요약 결과과 함께 비교해 보았다. 두 요약 결과를 문법완성도 측면과 핵심단어 포함 측면으로 나누어 비교분석 결과를 제시하였다.
#
#
# ## 1. 데이터 수집하기
#
# 이 데이터는 기사의 본문에 해당되는 text와 headlines 두 가지 열로 구성되어져 있습니다.
# | abstractive 요약
#
# Abstractive Extractive 요약
# text 사용 O 사용 O
# headlines 사용 O 사용 X
# +
import nltk
nltk.download('stopwords')
import numpy as np
import pandas as pd
import os
import re
import matplotlib.pyplot as plt
from nltk.corpus import stopwords
from bs4 import BeautifulSoup
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
import urllib.request
print('Done')
# -
import urllib.request
urllib.request.urlretrieve("https://raw.githubusercontent.com/sunnysai12345/News_Summary/master/news_summary_more.csv", filename="news_summary_more.csv")
data = pd.read_csv('news_summary_more.csv', encoding='iso-8859-1')
# +
data.head()
# -
# 모델이 text를 입력받아 context vector를 만들어내고, 출력단이 context vector를 통해 headlines와 가까운 문장을 만들어내는 것이 목표입니다.
data = data[['text', 'headlines']]
data.sample(10)
print('전체 샘플 수:', len(data))
# ### 2.1 중복 샘플과 NULL 값이 존재하는 샘플 제거
#
print('headlines 열에서 중복을 배제한 유일한 샘플의 수 :', data['headlines'].nunique())
print('text 열에서 중복을 배제한 유일한 샘플의 수 :', data['text'].nunique())
# text 중복 샘플 제거
# inplace=True 를 설정하면 DataFrame 타입 값을 return 하지 않고 data 내부를 직접적으로 바꿉니다
data.drop_duplicates(subset = ['text'], inplace=True)
print('전체 샘플수 :', (len(data)))
# Null Check
print(data.isnull().sum())
# 결측치를 제거합니다.
data.dropna(axis=0, inplace=True)
print('전체 샘플수 :', (len(data)))
# ## 2. Preprocessing data
#
#
# 자연어 데이터의 문제를 완화하는 전처리를 시도합니다. "you're"과 "you are "은 같은 말이고, "너는"과 "넌" 역시 같은 말입니다. 또한, '.........'과 '...............' 을 서로 다른 단어으로 생각할 것인지, '.'라는 단어가 여러 개 나열 된 것은 동일하지만 그 개수에서 차이를 보인다고 생각할 것인지에 대해 명확한 규정이 존재하지 않으므로, 어떤 판단기준으로 단어를 선정할 것인지는 엔지니어에 달려 있습니다. 모델이 학습할 때 혼란을 덜어줄 것 같은 전처리를 직관적이지만, 가장 효율적인지 알 수 없는(hueristic) 방법으로 진행하겠습니다.
#
# ### 2.2 정규화 사전
# +
# 정규화 사전 - 축약어를 분해할 때 사용합니다
contractions = {"ain't": "is not", "aren't": "are not","can't": "cannot", "'cause": "because", "could've": "could have", "couldn't": "could not",
"didn't": "did not", "doesn't": "does not", "don't": "do not", "hadn't": "had not", "hasn't": "has not", "haven't": "have not",
"he'd": "he would","he'll": "he will", "he's": "he is", "how'd": "how did", "how'd'y": "how do you", "how'll": "how will", "how's": "how is",
"I'd": "I would", "I'd've": "I would have", "I'll": "I will", "I'll've": "I will have","I'm": "I am", "I've": "I have", "i'd": "i would",
"i'd've": "i would have", "i'll": "i will", "i'll've": "i will have","i'm": "i am", "i've": "i have", "isn't": "is not", "it'd": "it would",
"it'd've": "it would have", "it'll": "it will", "it'll've": "it will have","it's": "it is", "let's": "let us", "ma'am": "madam",
"mayn't": "may not", "might've": "might have","mightn't": "might not","mightn't've": "might not have", "must've": "must have",
"mustn't": "must not", "mustn't've": "must not have", "needn't": "need not", "needn't've": "need not have","o'clock": "of the clock",
"oughtn't": "ought not", "oughtn't've": "ought not have", "shan't": "shall not", "sha'n't": "shall not", "shan't've": "shall not have",
"she'd": "she would", "she'd've": "she would have", "she'll": "she will", "she'll've": "she will have", "she's": "she is",
"should've": "should have", "shouldn't": "should not", "shouldn't've": "should not have", "so've": "so have","so's": "so as",
"this's": "this is","that'd": "that would", "that'd've": "that would have", "that's": "that is", "there'd": "there would",
"there'd've": "there would have", "there's": "there is", "here's": "here is","they'd": "they would", "they'd've": "they would have",
"they'll": "they will", "they'll've": "they will have", "they're": "they are", "they've": "they have", "to've": "to have",
"wasn't": "was not", "we'd": "we would", "we'd've": "we would have", "we'll": "we will", "we'll've": "we will have", "we're": "we are",
"we've": "we have", "weren't": "were not", "what'll": "what will", "what'll've": "what will have", "what're": "what are",
"what's": "what is", "what've": "what have", "when's": "when is", "when've": "when have", "where'd": "where did", "where's": "where is",
"where've": "where have", "who'll": "who will", "who'll've": "who will have", "who's": "who is", "who've": "who have",
"why's": "why is", "why've": "why have", "will've": "will have", "won't": "will not", "won't've": "will not have",
"would've": "would have", "wouldn't": "would not", "wouldn't've": "would not have", "y'all": "you all",
"y'all'd": "you all would","y'all'd've": "you all would have","y'all're": "you all are","y'all've": "you all have",
"you'd": "you would", "you'd've": "you would have", "you'll": "you will", "you'll've": "you will have",
"you're": "you are", "you've": "you have"}
print("정규화 사전의 수: ", len(contractions))
# -
# ### 2.3 불용어 사전
# +
# 불용어 사전 - 문맥과 무관하게 자주 나와 context vector에 담겨있는 정보를 희석할
# 염려가 있는 단어는 사용하지 않도록 합니다.
print('불용어 개수 :', len(stopwords.words('english') ))
print(stopwords.words('english'))
# -
# 불용어 사전에서 의문이 드는 점은, 부정어(no, not, nor, but)등도 포함되어 있다는 점인데, 이들은 사용하도록 하겠습니다. 또한, 뉴스에서 의미를 파악하는데 중요할 것 같은 단어도 사용하도록 하겠습니다
# 불용어 제거는 text 전처리 시에만 호출, headlines는 상대적으로 문장 길이가 짧음
# Abstractive한 문장 요약 결과문이 자연스러운 문장이 되려면 이 불용어들이 Headlines에 남아 있는 게 더 좋을 것 같습니다.
# 데이터 전처리 함수
def preprocess_sentence(sentence, remove_stopwords=True):
sentence = sentence.lower() # 텍스트 소문자화
sentence = BeautifulSoup(sentence, "lxml").text # <br />, <a href = ...> 등의 html 태그 제거
sentence = re.sub(r'\([^)]*\)', '', sentence) # 괄호로 닫힌 문자열 (...) 제거 Ex) my husband (and myself!) for => my husband for
sentence = re.sub('"','', sentence) # 쌍따옴표 " 제거
sentence = ' '.join([contractions[t] if t in contractions else t for t in sentence.split(" ")]) # 약어 정규화
sentence = re.sub(r"'s\b","", sentence) # 소유격 제거. Ex) roland's -> roland
sentence = re.sub("[^a-zA-Z]", " ", sentence) # 영어 외 문자(숫자, 특수문자 등) 공백으로 변환
sentence = re.sub('[m]{2,}', 'mm', sentence) # m이 3개 이상이면 2개로 변경. Ex) ummmmmmm yeah -> umm yeah
# 불용어 제거 (Text)
if remove_stopwords:
tokens = ' '.join(word for word in sentence.split() if not word in stopwords.words('english') if len(word) > 1)
# 불용어 미제거 (headlines)
else:
tokens = ' '.join(word for word in sentence.split() if len(word) > 1)
return tokens
preprocess_sentence("Helloooooo, ....!........what's your name?????????????? not so good?")
# +
temp_text = 'Everything I bought was great, infact I ordered twice and the third ordered was<br />for my mother and father.'
temp_headlines = 'Great way to start (or finish) the day!!!'
print(preprocess_sentence(temp_text))
print(preprocess_sentence(temp_headlines, False)) # 불용어를 제거하지 않습니다.
# -
# ### 2.5 멀티프로세싱
#
# multiprocessing을 이용하여 전체 문장을 전처리합니다
# +
import multiprocessing as mp # 멀티 프로세싱으로 전처리 속도를 획기적으로 줄여봅시다
from multiprocessing import Pool
import numpy as np
import time
from functools import partial # map을 할 때 함수에 여러 인자를 넣어줄 수 있도록 합니다
start = time.time()
# num_cores 만큼 쪼개진 데이터를 전처리하여 반환합니다
def appendTexts(sentences, remove_stopwords):
texts = []
for s in sentences:
texts += preprocess_sentence(s, remove_stopwords),
return texts
def preprocess_data(df, col, remove_stopwords=True):
start_time = time.time()
num_cores = mp.cpu_count() # 컴퓨터의 코어 수를 구합니다
text_data_split = np.array_split(data[col], num_cores) # 코어 수만큼 데이터를 배분하여 병렬적으로 처리할 수 있게 합니다
pool = Pool(num_cores)
processed_data = np.concatenate(pool.map(partial(appendTexts, remove_stopwords=remove_stopwords), text_data_split)) # 각자 작업한 데이터를 하나로 합쳐줍니다
pool.close()
pool.join()
print("{} : {} seconds".format(col, time.time() - start_time))
return processed_data
clean_text = preprocess_data(data,'text') # 클라우드 기준으로 3~4분 정도 소요 됩니다
clean_headlines = preprocess_data(data, 'headlines', remove_stopwords=False) # 클라우드 기준 1분정도 소요됩니다.
# +
#text : 735.3365228176117 seconds
#headlines : 149.51734685897827 seconds
#clean_text = process_data(data, 'text')
#clean_headlines = process_data(data, 'headlines')
# -
print(clean_headlines)
# 텍스트 정제의 과정을 거친 후에는 다시 한번 빈(empty) 샘플이 생겼는지 확인해보는 것이 좋아요. 정제 전에는 데이터가 존재했지만, 정제 과정에서 문장의 모든 단어가 사라지는 경우가 있을 수 있어요. 이렇게 되 면 샘플 자체가 빈 값을 가지게 되겠죠.
# +
#DataFrame으로 저장합니다
data['Text'] = clean_text
data['headlines'] = clean_headlines
# 빈 값을 Null 값으로 변환
data.replace('', np.nan, inplace=True)
# -
# 결측치 확인 74개의 빈 summary 확인되었습니다
data.isnull().sum()
# 결측치 제거
data.dropna(axis=0, inplace=True)
print('전체 샘플수 :', (len(data)))
# ### 2.6 샘플의 최대 길이 정하기
#
# 필요 없는 단어를 모두 솎아낸 데이터를 가지게 되었으니, 이제 훈련에 사용할 샘플의 최대 길이를 정해줄 차례에요.
# +
# 길이 분포 출력
import matplotlib.pyplot as plt
text_len = [len(s.split()) for s in data['text']]
headline_len = [len(s.split()) for s in data['headlines']]
print('텍스트의 최소 길이 : {}'.format(np.min(text_len)))
print('텍스트의 최대 길이 : {}'.format(np.max(text_len)))
print('텍스트의 평균 길이 : {}'.format(np.mean(text_len)))
print('헤드라인의 최소 길이 : {}'.format(np.min(headline_len)))
print('헤드라인의 최대 길이 : {}'.format(np.max(headline_len)))
print('헤드라인의 평균 길이 : {}'.format(np.mean(headline_len)))
plt.subplot(1,2,1)
plt.boxplot(headline_len)
plt.title('Headline')
plt.subplot(1,2,2)
plt.boxplot(text_len)
plt.title('Text')
plt.tight_layout()
plt.show()
plt.title('headline')
plt.hist(headline_len, bins = 40)
plt.xlabel('length of samples')
plt.ylabel('number of samples')
plt.show()
plt.title('Text')
plt.hist(text_len, bins = 40)
plt.xlabel('length of samples')
plt.ylabel('number of samples')
plt.show()
# -
# 데이터의 분포가 넓게 퍼져있음을 알 수 있습니다. 이 프로젝트의 목표는 긍정/부정 분석이 아니고, 뉴스를 요약하는 것이며, 짧은 Text의 경우, padding을 pre로 주게 된다면, 해당 단어가 연이은 padding 뒤에 나와 그에 연관된 context vector가 희석 없이 decoder로 넘어가게 되어 오히려 너무 강한 영향력을 행사할 것 같습니다. 따라서 짧은 텍스트에서 얻을 수 있는 정보는 모두 긴 Text에서 충분히 얻을 수 있다고 가정하고 너무 짧거나 긴 Text를 갖는 데이터포인트를 제거하겠습니다
#
# headline은 6-15 사이에 많은 데이터들이 있고, text는 45-65사이에 대부분의 데이터가 있음을 확인할 수 있습니다!
# 최소, 최대 사이에 있는 샘플을 골라 냅니다.
def below_threshold_len(min_len, max_len, nested_list):
cnt = 0
for s in nested_list:
if(len(s.split()) <= max_len) and (len(s.split()) >= min_len):
cnt = cnt + 1
print(f'전체 샘플 중 길이가 {min_len} 이상,{max_len} 이하인 샘플의 비율:{cnt / len(nested_list)}')
print('text')
below_threshold_len(50, 60, data['text'])
print('--------------------------------------------------------------')
print('headline')
below_threshold_len(5, 12, data['headlines'])
# +
text_max_len = 60; text_min_len = 50
headline_max_len = 11; headline_min_len = 6
# +
data = data[data['text'].apply(lambda x: len(x.split()) <= text_max_len \
and len(x.split()) >= text_min_len)]
data = data[data['headlines'].apply(lambda x: len(x.split()) <= headline_max_len and len(x.split()) >= headline_min_len)]
print('전체 샘플수 :', (len(data)))
# -
data
# ### 2.7 시작 토큰과 종료 토큰 추가하기
#
# 
#
#
# Seq2Seq | decoder_input(입력) | decoder_target (출력, 레이블)
# -----|-----|-----
# 시작토큰 | sostoken +~ |
# 종료토큰 | | ~+ eostoken
#
# 헤드라인 데이터에는 시작 토큰과 종료 토큰을 추가한다.
data['decoder_input'] = data['headlines'].apply(lambda x : 'sostoken '+ x) # 문장 앞에 sostoken을 추가해준다.
data['decoder_target'] = data['headlines'].apply(lambda x : x + ' eostoken') # 문장 뒤에 eostoken을 추가해준다.
data.head()
# 인코더의 입력, 디코더의 입력과 레이블을 각각 다시 Numpy 타입으로 저장 (np.array)
encoder_input = np.array(data['text']) # 인코더의 입력
decoder_input = np.array(data['decoder_input']) # 디코더의 입력
decoder_target = np.array(data['decoder_target']) # 디코더의 레이블
# ### 2.8 훈련 데이터와 테스트 데이터 분리 (# train test split)
#
# np.random.shuffle을 사용하여 섞인 데이터를 8:2 비율로 훈련 데이터와 테스트 데이터로 분리한다.
# +
# Indices 를 설정! (why?? : 순서(index)를 필요로 하기에!)
indices = np.arange(encoder_input.shape[0]) # indices에 인코더를 넣어준다. input shape를 맞춰줘야 한다.
np.random.shuffle(indices) # 랜덤하게 섞어서 과적합을 막는 용도???
print("Indices are :", + indices)
# encoder input과 decoder target을 설정한다.
encoder_input = encoder_input[indices]
decoder_input = decoder_input[indices]
decoder_target = decoder_target[indices]
n_of_val = int(len(encoder_input)*0.2)
print('\n테스트 데이터의 수 :', n_of_val)
# train_set으로 만들어준다! (encoder_input과 decoder_target을 가지고)
# Q: 왜 decoder target이 필요할까??
encoder_input_train = encoder_input[:-n_of_val]
decoder_input_train = decoder_input[:-n_of_val]
decoder_target_train = decoder_target[:-n_of_val]
# test_set 을 만들어준다! (encoder_input과 decoder_target을 가지고)
encoder_input_test = encoder_input[-n_of_val:]
decoder_input_test = decoder_input[-n_of_val:]
decoder_target_test = decoder_target[-n_of_val:]
print('훈련 데이터의 개수 :', len(encoder_input_train))
print('훈련 레이블의 개수 :', len(decoder_input_train))
print('테스트 데이터의 개수 :', len(encoder_input_test))
print('테스트 레이블의 개수 :', len(decoder_input_test))
# -
# ### 2.9 정수 인코딩
#
# 1. Tokenizer() = 텍스트를 토큰화
#
#
# 2. fit_on_texts() = 입력 데이터로부터 단어 집합 생성, 각 단어에 고유한 정수가 부여
#
#
# 3. word_index = 생성된 단어 집합
#
#
# 4. word_counts = 각 단어와 그 단어의 등장 빈도수가 저장되어있는 딕셔너리
#
#
# 5. texts_to_sequences() = 생성된 단어 집합에 기반하여 입력으로 주어진 텍스트 데이터의 단어들을 모두 정수로 변환하는 정수 인코딩을 수행한다.
src_tokenizer = Tokenizer() # 토크나이저 정의 (src를 사용)
src_tokenizer.fit_on_texts(encoder_input_train) # encoder에서 입력된 데이터로부터 단어 집합 생성
# +
# 빈도수가 낮은 단어들은 훈련 데이터에서 제외하고 진행
threshold = 9
total_cnt = len(src_tokenizer.word_index) # 단어의 수
rare_cnt = 0 # 등장 빈도수가 threshold보다 작은 단어의 개수를 카운트
total_freq = 0 # 훈련 데이터의 전체 단어 빈도수 총 합
rare_freq = 0 # 등장 빈도수가 threshold보다 작은 단어의 등장 빈도수의 총 합
# 단어와 빈도수의 쌍(pair)을 key와 value로 받는다.
for key, value in src_tokenizer.word_counts.items():
total_freq = total_freq + value
# 단어의 등장 빈도수가 threshold보다 작으면
if(value < threshold):
rare_cnt = rare_cnt + 1
rare_freq = rare_freq + value
print('단어 집합(vocabulary)의 크기 :', total_cnt)
print('등장 빈도가 %s번 이하인 희귀 단어의 수: %s'%(threshold - 1, rare_cnt))
print('단어 집합에서 희귀 단어를 제외시킬 경우의 단어 집합의 크기 %s'%(total_cnt - rare_cnt))
print("단어 집합에서 희귀 단어의 비율:", (rare_cnt / total_cnt)*100)
print("전체 등장 빈도에서 희귀 단어 등장 빈도 비율:", (rare_freq / total_freq)*100)
# -
# 단어 집합에서 희귀 단어를 제외시킬 경우의 단어 집합의 크기는 20838이다.
src_vocab = 20838
src_tokenizer = Tokenizer(num_words=src_vocab) # 단어 집합의 크기를 20838으로 제한
src_tokenizer.fit_on_texts(encoder_input_train) # 단어 집합 재생성. (encoder를 사용)
# +
# 텍스트 시퀀스를 정수 시퀀스로 변환
encoder_input_train = src_tokenizer.texts_to_sequences(encoder_input_train)
encoder_input_test = src_tokenizer.texts_to_sequences(encoder_input_test)
# 잘 진행되었는지 샘플 출력
print(encoder_input_train[:3])
# +
# 헤드라인 데이터에도 동일한 작업 수행
tar_tokenizer = Tokenizer() # target tokenizer
tar_tokenizer.fit_on_texts(decoder_input_train) # decoder train
threshold = 9
total_cnt = len(tar_tokenizer.word_index) # 단어의 수
rare_cnt = 0 # 등장 빈도수가 threshold보다 작은 단어의 개수를 카운트
total_freq = 0 # 훈련 데이터의 전체 단어 빈도수 총 합
rare_freq = 0 # 등장 빈도수가 threshold보다 작은 단어의 등장 빈도수의 총 합
# 단어와 빈도수의 쌍(pair)을 key와 value로 받는다.
for key, value in tar_tokenizer.word_counts.items():
total_freq = total_freq + value
# 단어의 등장 빈도수가 threshold보다 작으면
if(value < threshold):
rare_cnt = rare_cnt + 1
rare_freq = rare_freq + value
print('단어 집합(vocabulary)의 크기 :', total_cnt)
print('등장 빈도가 %s번 이하인 희귀 단어의 수: %s'%(threshold - 1, rare_cnt))
print('단어 집합에서 희귀 단어를 제외시킬 경우의 단어 집합의 크기 %s'%(total_cnt - rare_cnt))
print("단어 집합에서 희귀 단어의 비율:", (rare_cnt / total_cnt)*100)
print("전체 등장 빈도에서 희귀 단어 등장 빈도 비율:", (rare_freq / total_freq)*100)
# +
# 단어 집합에서 희귀 단어를 제외시킬 경우의 단어 집합의 크기 7919
tar_vocab = 7919
tar_tokenizer = Tokenizer(num_words=tar_vocab)
tar_tokenizer.fit_on_texts(decoder_input_train)
tar_tokenizer.fit_on_texts(decoder_target_train)
# 텍스트 시퀀스를 정수 시퀀스로 변환
decoder_input_train = tar_tokenizer.texts_to_sequences(decoder_input_train)
decoder_target_train = tar_tokenizer.texts_to_sequences(decoder_target_train)
decoder_input_test = tar_tokenizer.texts_to_sequences(decoder_input_test)
decoder_target_test = tar_tokenizer.texts_to_sequences(decoder_target_test)
# 잘 변환되었는지 확인
print('input ',decoder_input_train[:5])
print('decoder ',decoder_target_train[:5])
# +
# 결측치 제거 - eostoken만 남은 경우를 제거한다. 아마 없을 것이다
drop_train = [index for index, sentence in enumerate(decoder_input_train) if len(sentence) == 1]
drop_test = [index for index, sentence in enumerate(decoder_input_test) if len(sentence) == 1]
print('삭제할 훈련 데이터의 개수 :', len(drop_train))
print('삭제할 테스트 데이터의 개수 :', len(drop_test))
encoder_input_train = np.delete(encoder_input_train, drop_train, axis=0)
decoder_input_train = np.delete(decoder_input_train, drop_train, axis=0)
decoder_target_train = np.delete(decoder_target_train, drop_train, axis=0)
encoder_input_test = np.delete(encoder_input_test, drop_test, axis=0)
decoder_input_test = np.delete(decoder_input_test, drop_test, axis=0)
decoder_target_test = np.delete(decoder_target_test, drop_test, axis=0)
print('훈련 데이터의 개수 :', len(encoder_input_train))
print('훈련 레이블의 개수 :', len(decoder_input_train))
print('테스트 데이터의 개수 :', len(encoder_input_test))
print('테스트 레이블의 개수 :', len(decoder_input_test))
# -
# ### Padding
#
# encoder의 경우, padding을 뒤에 넣으면 context vector가 희석되므로 앞에 padding을 넣습니다
encoder_input_train = pad_sequences(encoder_input_train, maxlen=text_max_len, padding='pre')
encoder_input_test = pad_sequences(encoder_input_test, maxlen=text_max_len, padding='pre')
decoder_input_train = pad_sequences(decoder_input_train, maxlen=headline_max_len, padding='post')
decoder_target_train = pad_sequences(decoder_target_train, maxlen=headline_max_len, padding='post')
decoder_input_test = pad_sequences(decoder_input_test, maxlen=headline_max_len, padding='post')
decoder_target_test = pad_sequences(decoder_target_test, maxlen=headline_max_len, padding='post')
# ## 3. 어텐션 메커니즘 사용하기 (추상적 요약)
#
#
#
# 1. Encoder 설계
# 2. Decoder 설계
# 3. Attention 적용
# 4. 인퍼런스 모델
#
#
#
# ### 3.1 Encoder 설계
#
# #### INPUT
#
# text_max_len = 50
#
# #### Embedding
#
# src_vocab = 10838, embedding_dim = 128
#
#
#
#
# ### LSTM
#
# ##### hidden_state : LSTM에서 얼만큼의 수용력(Capacity)를 가질지를 정하는 파라미터
# - 이 파라미터는 LSTM의 용량의 크기나, LSTM에서의 뉴런의 개수 다른 신경망과 마찬가지로 무조건 용량을 많이 준다고 해서 성능이 반드시 올라가지는 않는다.
#
#
# ##### return_sequences : 시퀀스 출력 여부
# return_sequences = True로 설정할 경우 각 time step별 hidden state를 모두 출력하게 된다. Attention을 사용할 때 주로 설정한다.
#
#
# ##### return_state : 시퀀스 출력 여부
# return_state = True 를 한 경우에는 마지막 time step에서의 output(hidden state), hidden state와 cell state가 출력된다
#
#
# ##### dropout
# 0과 1사이 부동소수점. 인풋의 선형적 변형을 실행하는데 드롭시킬(고려하지 않을) 유닛의 비율.
#
#
# ##### recurrent_dropout
#
# - 0과 1사이 부동소수점. 순환 상태의 선형적 변형을 실행하는데 드롭시킬(고려하지 않을) 유닛의 비율.
#
# 1. 'encoder_lstm1 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)'
#
# 2. 'encoder_lstm2 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)'
#
# 3. 'encoder_lstm3 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)'
# +
from tensorflow.keras.layers import Input, LSTM, Embedding, Dense, Concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
# 인코더 설계 시작
embedding_dim = 128
hidden_size = 256
# 인코더 (40,)
encoder_inputs = Input(shape=(text_max_len,))
# 인코더의 임베딩 층 (20000, 128)
enc_emb = Embedding(src_vocab, embedding_dim)(encoder_inputs)
# 인코더의 LSTM 1
encoder_lstm1 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)
encoder_output1, state_h1, state_c1 = encoder_lstm1(enc_emb)
# 인코더의 LSTM 2
encoder_lstm2 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)
encoder_output2, state_h2, state_c2 = encoder_lstm2(encoder_output1)
# 인코더의 LSTM 3
encoder_lstm3 = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.4)
encoder_outputs, state_h, state_c = encoder_lstm3(encoder_output2)
# -
# ### 3.2 Decoder 설계
# 디코더의 임베딩 층과 LSTM을 설계하는 것은 인코더와 거의 동일하다.
#
# 하지만 LSTM의 입력을 정의할 때, initial_state의 인자값으로 인코더의 hidden state와 cell state의 값을 넣어줘야한다.
#
#
#
# ##### INPUT
#
# shape=(None,)
#
# ##### Embedding
#
# tar_vocab = 9700, embedding_dim = 128
#
# #### LSTM
#
# 1. decoder_lstm = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.2)
#
#
#
# #### 출력층
# 디코더의 출력층에서는 다중 클래스 분류 문제를 풀어하므로 Dense의 인자로 tar_vocab을 주고, 활성화 함수로 소프트맥스 함수를 사용한다.
#
# 지금까지 설계한 것은 인코더의 hidden state와 cell state를 디코더의 초기 state로 사용하는 가장 기본적인 seq2seq 구조다.
#
# 그런데 디코더의 출력층을 설계를 살짝 바꿔서 성능을 높일 수 있는 방법이 있는데 바로
# **어텐션** 메커니즘이다
# +
# 디코더 설계
decoder_inputs = Input(shape=(None,))
# 디코더의 임베딩 층
dec_emb_layer = Embedding(tar_vocab, embedding_dim)
dec_emb = dec_emb_layer(decoder_inputs)
# 디코더의 LSTM
decoder_lstm = LSTM(hidden_size, return_sequences=True, return_state=True, dropout=0.4, recurrent_dropout=0.2)
decoder_outputs, _, _ = decoder_lstm(dec_emb, initial_state=[state_h, state_c])
# +
# 디코더의 출력층
decoder_softmax_layer = Dense(tar_vocab, activation='softmax')
decoder_softmax_outputs = decoder_softmax_layer(decoder_outputs)
# 모델 정의
model = Model([encoder_inputs, decoder_inputs], decoder_softmax_outputs)
model.summary()
# -
# ### 3.3 Attention 적용
#
# 일반적인 seq2seq보다는 어텐션 메커니즘을 사용한 seq2seq를 사용하는 것이 더 나은 성능을 얻을 수 있다.
#
# 
#
urllib.request.urlretrieve("https://raw.githubusercontent.com/thushv89/attention_keras/master/src/layers/attention.py", filename="attention.py")
from attention import AttentionLayer
# +
# 어텐션 층(어텐션 함수)
attn_layer = AttentionLayer(name='attention_layer')
# 인코더와 디코더의 모든 time step의 hidden state를 어텐션 층에 전달하고 결과를 리턴
attn_out, attn_states = attn_layer([encoder_outputs, decoder_outputs])
# 어텐션의 결과와 디코더의 hidden state들을 연결
decoder_concat_input = Concatenate(axis=-1, name='concat_layer')([decoder_outputs, attn_out])
# 디코더의 출력층
decoder_softmax_layer = Dense(tar_vocab, activation='softmax')
decoder_softmax_outputs = decoder_softmax_layer(decoder_concat_input)
# 모델 정의
model = Model([encoder_inputs, decoder_inputs], decoder_softmax_outputs)
model.summary()
# -
# 모델 훈련
model.compile(optimizer='rmsprop', loss='sparse_categorical_crossentropy')
es = EarlyStopping(monitor='val_loss', patience=2, verbose=1)
history = model.fit(x=[encoder_input_train, decoder_input_train],
y=decoder_target_train,
validation_data=(
[encoder_input_test, decoder_input_test],decoder_target_test),
batch_size=256,
callbacks=[es],
epochs=50)
# +
# 6. 모델 저장하기
from keras.models import load_model
model.save('attention_seq2seq.h5')
model.save_weights('news_summarizer_weights_1')
# -
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
# #### 4. 인퍼런스 모델 구현
#
# 테스트 단계에서는 정수 인덱스 행렬로 존재하던 텍스트 데이터를 실제 데이터로 복원해야하므로, 필요한 3개의 사전을 아래와 같이 미리 준비해 둡니다.
src_index_to_word = src_tokenizer.index_word # 원문 단어 집합에서 정수 -> 단어를 얻음
tar_word_to_index = tar_tokenizer.word_index # 요약 단어 A집합에서 단어 -> 정수를 얻음
tar_index_to_word = tar_tokenizer.index_word # 요약 단어 집합에서 정수 -> 단어를 얻음
# **seq2seq는 훈련할 때와 실제 동작할 때(인퍼런스 단계)의 방식이 다르므로 그에 맞게 모델 설계를 별개로 진행해야 한다**
#
#
# 훈련 단계
#
# - 디코더의 입력부에 정답이 되는 문장 전체를 한꺼번에 넣고 디코더의 출력과 한 번에 비교할 수 있으므로, 인코더와 디코더를 엮은 통짜 모델 하나만 준비했다.
#
#
# 인퍼런스 단계
#
# - 정답 문장이 없는 인퍼런스 단계에서는 만들어야 할 문장의 길이만큼 디코더가 반복 구조로 동작해야 하기 때문에 부득이하게 인퍼런스를 위한 모델 설계를 별도로 해주어야 합니다. 이때는 인코더 모델과 디코더 모델을 분리해서 설계합니다.
# +
# 인코더 설계
encoder_model = Model(inputs=encoder_inputs, outputs=[encoder_outputs, state_h, state_c])
# 이전 시점의 상태들을 저장하는 텐서
decoder_state_input_h = Input(shape=(hidden_size,))
decoder_state_input_c = Input(shape=(hidden_size,))
dec_emb2 = dec_emb_layer(decoder_inputs)
# 문장의 다음 단어를 예측하기 위해서 초기 상태(initial_state)를 이전 시점의 상태로 사용. 이는 뒤의 함수 decode_sequence()에 구현
# 훈련 과정에서와 달리 LSTM의 리턴하는 은닉 상태와 셀 상태인 state_h와 state_c를 버리지 않음.
decoder_outputs2, state_h2, state_c2 = decoder_lstm(dec_emb2, initial_state=[decoder_state_input_h, decoder_state_input_c])
# +
# 어텐션 메커니즘을 사용하는 출력층을 설계
# 어텐션 함수
decoder_hidden_state_input = Input(shape=(text_max_len, hidden_size))
attn_out_inf, attn_states_inf = attn_layer([decoder_hidden_state_input, decoder_outputs2])
decoder_inf_concat = Concatenate(axis=-1, name='concat')([decoder_outputs2, attn_out_inf])
# 디코더의 출력층
decoder_outputs2 = decoder_softmax_layer(decoder_inf_concat)
# 최종 디코더 모델
decoder_model = Model(
[decoder_inputs] + [decoder_hidden_state_input,decoder_state_input_h, decoder_state_input_c],
[decoder_outputs2] + [state_h2, state_c2])
# -
def decode_sequence(input_seq):
# 입력으로부터 인코더의 상태를 얻음
e_out, e_h, e_c = encoder_model.predict(input_seq)
# <SOS>에 해당하는 토큰 생성
target_seq = np.zeros((1,1))
target_seq[0, 0] = tar_word_to_index['sostoken']
stop_condition = False
decoded_sentence = ''
while not stop_condition: # stop_condition이 True가 될 때까지 루프 반복
output_tokens, h, c = decoder_model.predict([target_seq] + [e_out, e_h, e_c])
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_token = tar_index_to_word[sampled_token_index]
if (sampled_token!='eostoken'):
decoded_sentence += ' '+sampled_token
# <eos>에 도달하거나 최대 길이를 넘으면 중단.
if (sampled_token == 'eostoken' or len(decoded_sentence.split()) >= (headline_max_len-1)):
stop_condition = True
# 길이가 1인 타겟 시퀀스를 업데이트
target_seq = np.zeros((1,1))
target_seq[0, 0] = sampled_token_index
# 상태를 업데이트 합니다.
e_h, e_c = h, c
return decoded_sentence
# #### 모델 테스트
#
#
# 테스트 단계에서는 정수 시퀀스를 텍스트 시퀀스로 변환하여 결과를 확인하는 것이 편하겠죠. 주어진 정수 시퀀스를 텍스트 시퀀스로 변환하는 함수를 만들어볼게요. 함수를 만들 때, Text의 정수 시퀀스에서는 패딩을 위해 사용되는 숫자 0을 제외하고 Summary의 정수 시퀀스에서는 숫자 0, 시작 토큰의 인덱스, 종료 토큰의 인덱스를 출력에서 제외하도록 만들 거예요.
# +
# 원문의 정수 시퀀스를 텍스트 시퀀스로 변환
def seq2text(input_seq):
temp=''
for i in input_seq:
if (i!=0):
temp = temp + src_index_to_word[i]+' '
return temp
# 요약문의 정수 시퀀스를 텍스트 시퀀스로 변환
def seq2summary(input_seq):
temp=''
for i in input_seq:
if ((i!=0 and i!=tar_word_to_index['sostoken']) and i!=tar_word_to_index['eostoken']):
temp = temp + tar_index_to_word[i] + ' '
return temp
# -
# ## 4. 실제 결과와 요약문 비교하기 (추상적 요약)
#
# 원래의 요약문(headlines 열)과 학습을 통해 얻은 추상적 요약의 결과를 비교한다.
for i in range(10):
print("원문 :", seq2text(encoder_input_test[i]))
print("실제 Headline :", seq2summary(decoder_input_test[i]))
print("예측 Headline :", decode_sequence(encoder_input_test[i].reshape(1, text_max_len)))
print("\n")
# ## 5. Summa을 이용해서 추출적 요약해보기
#
# 추상적 요약은 추출적 요약과는 달리 문장의 표현력을 다양하게 가져갈 수 있지만, 추출적 요약에 비해서 난이도가 높다. 반대로 말하면 추출적 요약은 추상적 요약에 비해 난이도가 낮고 기존 문장에서 문장을 꺼내오는 것이므로 잘못된 요약이 나올 가능성이 낮다.
#
# Summa의 summarize()의 인자로 사용되는 값들에 대해서 알아본다.
#
# 1. text (str) : 요약할 테스트.
#
#
# 2. ratio (float, optional) – 요약문에서 원본에서 선택되는 문장 비율. 0~1 사이값
#
#
# 3. words (int or None, optional) – 출력에 포함할 단어 수. 만약, ratio와 함께 두 파라미터가 모두 제공되는 경우 ratio는 무시한다.
#
#
# 4. split (bool, optional) – True면 문장 list / False는 조인(join)된 문자열을 반환
#
#
#
# Summa의 summarize는 문장 토큰화를 별도로 하지 않더라도 내부적으로 문장 토큰화를 수행한다.. 그렇기 때문에 문장 구분이 되어있지 않은 원문을 바로 입력으로 넣을 수 있다. 비율을 적게 주어서 요약문으로 선택되는 문장의 개수를 줄여본다. 원문의 0.005%만을 출력하도록 설정했다.
# +
import requests
from summa.summarizer import summarize
urllib.request.urlretrieve("https://raw.githubusercontent.com/sunnysai12345/News_Summary/master/news_summary_more.csv", filename="news_summary_more.csv")
data = pd.read_csv('news_summary_more.csv', encoding='iso-8859-1')
# -
for i in range(10):
print("실제 HEADLINE : ", data['headlines'][i])
print("추출적 요약 HEADLINE : ", summarize(data['text'][i], ratio=0.5))
print("\n")
|
EXP_10_news_sum.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # LTR Model to Salient Concepts
# In this approach, we use LTR to select candidate phrases for generating summaris.
# # Data Construction
# This step is reading tokens from BookNLP to construct training set.
import numpy as np
from sklearn import linear_model
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.model_selection import KFold
from sklearn.metrics import mean_squared_error, mean_absolute_error
from sklearn.model_selection import cross_val_score
#####################################################################
# DataSet
#####################################################################
train_file = open('../../0.part.tokens.LTR_FT', 'rb')
character = {}
for line in train_file:
terms = line.split('\t')
key = terms[0] + ' ' + terms[1]
if not key in character:
character[key] = []
character[key].append(terms[2])
train_file.close()
samples = character.values()
def sample2Xy(samples, indexes):
X = []
y = []
for index in indexes:
for des in samples[index]:
terms = des.split(' ')
X.append([float(x) for x in terms[1:]])
y.append(float(terms[0]))
X = np.array(X, dtype=np.float32)
y = np.array(y, dtype=np.float32)
return X, y
kf = KFold(n_splits = 5, shuffle = True)
mse_linear_model = []
mae_linear_model = []
mse_tree_model = []
mae_tree_model = []
for train, test in kf.split(samples):
X_train, y_train = sample2Xy(samples, train)
X_test, y_test = sample2Xy(samples, test)
# Linear Regression
clf = linear_model.LinearRegression()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
mse_linear_model.append(mean_squared_error(y_test, y_pred))
mae_linear_model.append(mean_absolute_error(y_test, y_pred))
# Gradient Boosting Regressor
clf = GradientBoostingRegressor(learning_rate=0.05, random_state=1)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
mse_tree_model.append(mean_squared_error(y_test, y_pred))
mae_tree_model.append(mean_absolute_error(y_test, y_pred))
print 'Linear Regression MSE ' + str(abs(np.mean(mse_linear_model)))
print 'Linear Regression MAE ' + str(abs(np.mean(mae_linear_model)))
print 'Gradient Boosting Regressor MSE ' + str(abs(np.mean(mse_tree_model)))
print 'Gradient Boosting Regressor MAE ' + str(abs(np.mean(mae_tree_model)))
i = 495
print y_test[i: i + 10]
print y_pred[i: i + 10]
# # next step
# ## nDCG
# ## MAP
#
|
ranking/LTR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.010968, "end_time": "2021-11-09T00:09:26.192214", "exception": false, "start_time": "2021-11-09T00:09:26.181246", "status": "completed"} tags=[]
# # Word Embeddings
#
# You know at this point that machine learning on text requires that you first represent the text numerically. So far, you've done this with bag of words representations. But you can usually do better with word embeddings.
#
# **Word embeddings** (also called word vectors) represent each word numerically in such a way that the vector corresponds to how that word is used or what it means. Vector encodings are learned by considering the context in which the words appear. Words that appear in similar contexts will have similar vectors. For example, vectors for "leopard", "lion", and "tiger" will be close together, while they'll be far away from "planet" and "castle".
#
# Even cooler, relations between words can be examined with mathematical operations. Subtracting the vectors for "man" and "woman" will return another vector. If you add that to the vector for "king" the result is close to the vector for "queen."
#
# 
#
# These vectors can be used as features for machine learning models. Word vectors will typically improve the performance of your models above bag of words encoding. spaCy provides embeddings learned from a model called Word2Vec. You can access them by loading a large language model like `en_core_web_lg`. Then they will be available on tokens from the `.vector` attribute.
# + papermill={"duration": 15.217082, "end_time": "2021-11-09T00:09:41.419190", "exception": false, "start_time": "2021-11-09T00:09:26.202108", "status": "completed"} tags=[]
import numpy as np
import spacy
# Need to load the large model to get the vectors
nlp = spacy.load('en_core_web_lg')
# + papermill={"duration": 0.04549, "end_time": "2021-11-09T00:09:41.474926", "exception": false, "start_time": "2021-11-09T00:09:41.429436", "status": "completed"} tags=[]
# Disabling other pipes because we don't need them and it'll speed up this part a bit
text = "These vectors can be used as features for machine learning models."
with nlp.disable_pipes():
vectors = np.array([token.vector for token in nlp(text)])
# + papermill={"duration": 0.020553, "end_time": "2021-11-09T00:09:41.505573", "exception": false, "start_time": "2021-11-09T00:09:41.485020", "status": "completed"} tags=[]
vectors.shape
# + [markdown] papermill={"duration": 0.010226, "end_time": "2021-11-09T00:09:41.526424", "exception": false, "start_time": "2021-11-09T00:09:41.516198", "status": "completed"} tags=[]
# These are 300-dimensional vectors, with one vector for each word. However, we only have document-level labels and our models won't be able to use the word-level embeddings. So, you need a vector representation for the entire document.
#
# There are many ways to combine all the word vectors into a single document vector we can use for model training. A simple and surprisingly effective approach is simply averaging the vectors for each word in the document. Then, you can use these document vectors for modeling.
#
# spaCy calculates the average document vector which you can get with `doc.vector`. Here is an example loading the spam data and converting it to document vectors.
# + papermill={"duration": 49.629752, "end_time": "2021-11-09T00:10:31.166578", "exception": false, "start_time": "2021-11-09T00:09:41.536826", "status": "completed"} tags=[]
import pandas as pd
# Loading the spam data
# ham is the label for non-spam messages
spam = pd.read_csv('../input/nlp-course/spam.csv')
with nlp.disable_pipes():
doc_vectors = np.array([nlp(text).vector for text in spam.text])
doc_vectors.shape
# + [markdown] papermill={"duration": 0.010562, "end_time": "2021-11-09T00:10:31.188077", "exception": false, "start_time": "2021-11-09T00:10:31.177515", "status": "completed"} tags=[]
# ## Classification Models
#
# With the document vectors, you can train scikit-learn models, xgboost models, or any other standard approach to modeling.
# + papermill={"duration": 0.317713, "end_time": "2021-11-09T00:10:31.516779", "exception": false, "start_time": "2021-11-09T00:10:31.199066", "status": "completed"} tags=[]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(doc_vectors, spam.label,
test_size=0.1, random_state=1)
# + [markdown] papermill={"duration": 0.01156, "end_time": "2021-11-09T00:10:31.539434", "exception": false, "start_time": "2021-11-09T00:10:31.527874", "status": "completed"} tags=[]
# Here is an example using [support vector machines (SVMs)](https://scikit-learn.org/stable/modules/svm.html#svm). Scikit-learn provides an SVM classifier `LinearSVC`. This works similar to other scikit-learn models.
# + papermill={"duration": 0.307918, "end_time": "2021-11-09T00:10:31.858511", "exception": false, "start_time": "2021-11-09T00:10:31.550593", "status": "completed"} tags=[]
from sklearn.svm import LinearSVC
# Set dual=False to speed up training, and it's not needed
svc = LinearSVC(random_state=1, dual=False, max_iter=10000)
svc.fit(X_train, y_train)
print(f"Accuracy: {svc.score(X_test, y_test) * 100:.3f}%", )
# + [markdown] papermill={"duration": 0.048047, "end_time": "2021-11-09T00:10:31.927503", "exception": false, "start_time": "2021-11-09T00:10:31.879456", "status": "completed"} tags=[]
# ## Document Similarity
#
# Documents with similar content generally have similar vectors. So you can find similar documents by measuring the similarity between the vectors. A common metric for this is the **cosine similarity** which measures the angle between two vectors, $\mathbf{a}$ and $\mathbf{b}$.
#
# $$
# \cos \theta = \frac{\mathbf{a}\cdot\mathbf{b}}{\| \mathbf{a} \| \, \| \mathbf{b} \|}
# $$
#
# This is the dot product of $\mathbf{a}$ and $\mathbf{b}$, divided by the magnitudes of each vector. The cosine similarity can vary between -1 and 1, corresponding complete opposite to perfect similarity, respectively. To calculate it, you can use [the metric from scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.pairwise.cosine_similarity.html) or write your own function.
# + papermill={"duration": 0.036938, "end_time": "2021-11-09T00:10:31.990521", "exception": false, "start_time": "2021-11-09T00:10:31.953583", "status": "completed"} tags=[]
def cosine_similarity(a, b):
return a.dot(b)/np.sqrt(a.dot(a) * b.dot(b))
# + papermill={"duration": 0.049973, "end_time": "2021-11-09T00:10:32.053571", "exception": false, "start_time": "2021-11-09T00:10:32.003598", "status": "completed"} tags=[]
a = nlp("REPLY NOW FOR FREE TEA").vector
b = nlp("According to legend, Emperor <NAME> discovered tea when leaves from a wild tree blew into his pot of boiling water.").vector
cosine_similarity(a, b)
# + [markdown] papermill={"duration": 0.012353, "end_time": "2021-11-09T00:10:32.078961", "exception": false, "start_time": "2021-11-09T00:10:32.066608", "status": "completed"} tags=[]
# # Your Turn
# Word embeddings are incredibly powerful. You know know enough to apply embeddings to **[improve your models and find similar documents](https://www.kaggle.com/kernels/fork/6061026)**.
# + [markdown] papermill={"duration": 0.012403, "end_time": "2021-11-09T00:10:32.103919", "exception": false, "start_time": "2021-11-09T00:10:32.091516", "status": "completed"} tags=[]
# ---
#
#
#
#
# *Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/natural-language-processing/discussion) to chat with other learners.*
|
course/Natural Language Processing/word-vectors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # dask作为算力池
#
# 可以理解为dask的数据结构只是计算框架的一个语法糖,它的本质还是任务调度系统,事实上基本所有分布式计算框架的本质都是任务调度系统.因为分布式计算说白了就是把大任务拆分为小任务然后交给很多机器一起执行,然后再回收结果.
#
# 那我们就可以利用dask的底层api将其作为算力池来使用.
#
# 需要注意的是分布式计算通常都要求函数无[副作用](https://baike.baidu.com/item/%E5%87%BD%E6%95%B0%E5%89%AF%E4%BD%9C%E7%94%A8/22723425?fr=aladdin).也就是说我们定义的函数不应该使用全局变量,不应该改变对象属性.
# ## 使用delayed接口构造任务链
#
# ***注意:delayed接口中不能使用标准dask的数结构***
#
# [delayed]()接口被设计成用于计算标准dask数据结构无法解决的问题.当然通常我们需要使用delayed的场景也都用不到这个接口.这个接口的作用相当于是向集群发布一条由python函数组成的命令,因此使用delayed接口就分解为了两个问题:
#
# + 怎样构造由python函数组成的命令(计算图)
#
# + 集群如何执行由python函数组成的命令
from dask.distributed import Client
client = Client('localhost:8786')
# ### 构造由python函数组成的命令
#
#
# dask通过[delayed](https://docs.dask.org/en/latest/delayed-api.html#)接口注册函数构造计算图,使用计算图来控制分发运算.在构造好计算图后,执行`.compute()`接口发布命令计算结果.下面的例子是官方给出的典型计算---每个元素加一加上每个元素加2之后再所有结果合起来相加
# +
import dask
def inc(x):
return x + 1
def double(x):
return x + 2
def add(x, y):
return x + y
data = [1, 2, 3, 4, 5]
output = []
for x in data:
a = dask.delayed(inc)(x)
b = dask.delayed(double)(x)
c = dask.delayed(add)(a, b)
output.append(c)
total = dask.delayed(sum)(output)
# -
total.compute()
# #### 使用装饰器`delayed`构造计算图
#
# `delayed`也可以作为装饰器,这样在定义函数时就可以同时构造计算图了.这种方式其实并不推荐,因为这样反而降低了灵活性,不利于扩展
# +
@dask.delayed
def inc1(x):
return x + 1
@dask.delayed
def double1(x):
return x + 2
@dask.delayed
def add1(x, y):
return x + y
data = [1, 2, 3, 4, 5]
output = []
for x in data:
a = inc1(x)
b = double1(x)
c = add1(a, b)
output.append(c)
total1 = dask.delayed(sum)(output)
# -
total1.compute()
# #### 自定义计算图
#
# 实际上dask可以用户自定义计算图,我们通过将计算过程编码到一个dict对象构造计算图,然后使用`dask.threaded.get`接口将计算图送入集群来计算,`get`的第一个参数是计算图,第二个是计算图中定义的key.注意计算图中的key可以是字符串或者tuple.更多关于自定义计算图的优化方法需要看下[相关文档](https://docs.dask.org/en/latest/optimize.html)
from dask.threaded import get
# +
def inc(i):
return i + 1
def add(a, b):
return a + b
# -
d = {'x': 1,
'y': (inc, 'x'),
'z': (add, 'y', 10)}
get(d, 'x')
get(d, 'z')
# #### 计算图的高级接口
#
# [计算图的高级接口](https://docs.dask.org/en/latest/high-level-graphs.html)允许我们为计算分层
# +
from dask.highlevelgraph import HighLevelGraph
import pandas as pd
import operator
layers = {
'read-csv': {('read-csv', 0): (pd.read_csv, 'myfile.0.csv'),
('read-csv', 1): (pd.read_csv, 'myfile.1.csv'),
('read-csv', 2): (pd.read_csv, 'myfile.2.csv'),
('read-csv', 3): (pd.read_csv, 'myfile.3.csv')},
'add': {('add', 0): (operator.add, ('read-csv', 0), 100),
('add', 1): (operator.add, ('read-csv', 1), 100),
('add', 2): (operator.add, ('read-csv', 2), 100),
('add', 3): (operator.add, ('read-csv', 3), 100)},
'filter':{('filter', 0): (lambda part: part[part.name == 'Alice'], ('add', 0)),
('filter', 1): (lambda part: part[part.name == 'Alice'], ('add', 1)),
('filter', 2): (lambda part: part[part.name == 'Alice'], ('add', 2)),
('filter', 3): (lambda part: part[part.name == 'Alice'], ('add', 3))}
}
dependencies = {'read-csv': set(),
'add': {'read-csv'},
'filter': {'add'}}
graph = HighLevelGraph(layers, dependencies)
# -
# #### 计算图可视化
#
# `visualize()`接口可以将计算图可视化,这需要安装可选依赖[python-graphviz](https://www.graphviz.org/)
total1.visualize()
# ### 集群执行由python函数组成的命令
#
# 要让集群可以执行命令,需要整个集群的python环境有对应需要的全部依赖,如果我们使用的swarm模式或者k8s模式部署的集群,那我们就需要为镜像单独安装依赖.一个比较好的方式是继承镜像重新打包:
#
# ```dockerfile
# FROM daskdev/dask:latest
# RUN conda install pytorch
#
# ENTRYPOINT ["tini", "-g", "--", "/usr/bin/prepare.sh"]
# ```
|
src/数据清洗篇/工具介绍/dask/dask作为算力池.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Ubuntu Linux)
# language: python
# name: python3
# ---
# #### CW-09
# Atabak, Ehsan, Krista
# - Makefiles provides directions for compiling and linking a program. Makefiles provide efficiency when a program links with multiple header files and header files have been edited. The program does not run the entire program all over, instead recompiles portions of program that are affected by the edited header files.
#
# Describe how the code is organized here. Why is this a reasonable structure?
#
# - Header file is that declares a library with predefined methods.
# - Source file is the code written by the user.
# - Object files are the output files from compilation.
# - Compiling is when source code is converted to machine code.
# - Linking occurs during compilation. It is when the various libraries are combined to be executed.
|
C_Compilation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Reference: [this colab pytorch implementation](https://colab.research.google.com/drive/18EyozusBSgxa5oUBmlzXrp9fEbPyOUoC)
# !git clone https://github.com/tkipf/pygcn.git
import sys
sys.path.insert(0,'pygcn')
# +
import numpy as np
import matplotlib.pyplot as plt
from pygcn.utils import accuracy
import torch
from torch import nn, optim
from torch.utils import data
# -
from pygcn.utils import load_data
adj, features, labels, idx_train, idx_val, idx_test = load_data(path='pygcn/data/cora/')
print(features.shape, adj.shape, labels.shape, torch.max(labels))
# # FF NN with 1 hidden layer
net = nn.Sequential(nn.Linear(1433, 100), nn.ReLU(), nn.Linear(100, 7))
from pygcn.utils import accuracy
def test(model):
# Testa il modello sulla porzione del dataset di test
y_pred = model(features[idx_test])
acc_test = accuracy(y_pred, labels[idx_test])
print("Accuracy:",
"accuracy= {:.4f}".format(acc_test.item()))
test(net)
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters())
# +
import tqdm
loss_history = np.zeros(1000)
for epoch in tqdm.trange(1000):
optimizer.zero_grad()
outputs = net(features[idx_train])
loss = criterion(outputs, labels[idx_train])
loss.backward()
optimizer.step()
loss_history[epoch] = loss.detach().numpy()
# -
plt.plot(loss_history)
plt.show()
test(net)
# # Graph Convolutional Network
# +
import math
import torch
from torch.nn.parameter import Parameter
from torch.nn.modules.module import Module
class GraphConvolution(Module):
"""
Simple GCN layer, similar to https://arxiv.org/abs/1609.02907
"""
def __init__(self, in_features, out_features):
super(GraphConvolution, self).__init__()
self.weight = Parameter(torch.FloatTensor(in_features, out_features))
self.bias = Parameter(torch.FloatTensor(out_features))
self.reset_parameters()
def reset_parameters(self):
stdv = 1. / math.sqrt(self.weight.size(1))
self.weight.data.uniform_(-stdv, stdv)
self.bias.data.uniform_(-stdv, stdv)
def forward(self, input, adj):
support = torch.mm(input, self.weight)
output = torch.spmm(adj, support)
return output + self.bias
# -
# ## Two layer GCN forward model
# $Z = f(X,A)=softmax\biggr(\hat A\text{ ReLU}\big(\hat AXW^{(0)}\big)W^{(1)}\biggr)$
# +
import torch.nn.functional as F
class GCN(nn.Module):
def __init__(self, nfeat, nhid, nclass):
super(GCN, self).__init__()
self.gc1 = GraphConvolution(nfeat, nhid)
self.gc2 = GraphConvolution(nhid, nclass)
def forward(self, x, adj):
x = F.relu(self.gc1(x, adj))
x = self.gc2(x, adj)
return F.log_softmax(x, dim=1)
# -
gcn = GCN(1433, 50, 7)
optimizer_gcn = optim.Adam(gcn.parameters())
criterion = nn.CrossEntropyLoss()
def test(model):
y_pred = model(features, adj) # Usiamo tutto il dataset
acc_test = accuracy(y_pred[idx_test], labels[idx_test]) # Mascheriamo sulla parte di test
print("Accuracy:",
"accuracy= {:.4f}".format(acc_test.item()))
test(gcn)
# +
import tqdm
loss_history = np.zeros(2500)
for epoch in tqdm.trange(2500):
optimizer_gcn.zero_grad()
outputs = gcn(features, adj) # Usiamo tutto il dataset
loss = criterion(outputs[idx_train], labels[idx_train]) # Mascheriamo sulla parte di training
loss.backward()
optimizer_gcn.step()
loss_history[epoch] = loss.detach().numpy()
# -
plt.plot(loss_history)
plt.show()
test(gcn)
loss
outputs.max(1)[1].type_as(labels)
labels
# # (Optional) CUDA-enabled GCN
# +
import math
import torch
from torch.nn.parameter import Parameter
from torch.nn.modules.module import Module
class GraphConvolution(Module):
"""
Simple GCN layer, similar to https://arxiv.org/abs/1609.02907
"""
def __init__(self, in_features, out_features):
super(GraphConvolution, self).__init__()
self.weight = Parameter(torch.FloatTensor(in_features, out_features))
self.bias = Parameter(torch.FloatTensor(out_features))
self.reset_parameters()
def reset_parameters(self):
stdv = 1. / math.sqrt(self.weight.size(1))
self.weight.data.uniform_(-stdv, stdv)
self.bias.data.uniform_(-stdv, stdv)
def forward(self, input, adj):
support = torch.mm(input, self.weight)
output = torch.spmm(adj, support)
return output + self.bias
# +
import torch.nn.functional as F
class GCN(nn.Module):
def __init__(self, nfeat, nhid, nclass, dropout):
super(GCN, self).__init__()
self.gc1 = GraphConvolution(nfeat, nhid)
self.gc2 = GraphConvolution(nhid, nclass)
self.dropout = dropout
def forward(self, x, adj):
x = F.relu(self.gc1(x, adj))
x = F.dropout(x, self.dropout, training=self.training)
x = self.gc2(x, adj)
return F.log_softmax(x, dim=1)
# -
gcn = GCN(1433, 50, 7, 0.2)
optimizer_gcn = optim.Adam(gcn.parameters())
criterion = nn.CrossEntropyLoss()
gcn.to('cuda')
# +
import tqdm
loss_history = np.zeros(2500)
for epoch in tqdm.trange(2500):
optimizer_gcn.zero_grad()
outputs = gcn(features.to('cuda'), adj.to('cuda')) # Usiamo tutto il dataset
loss = criterion(outputs[idx_train].to('cuda'), labels[idx_train].to('cuda')) # Mascheriamo sulla parte di training
loss.backward()
optimizer_gcn.step()
loss_history[epoch] = loss.detach().cpu().numpy()
# -
import matplotlib.pyplot as plt
plt.plot(loss_history)
plt.ylim(0, 0.2)
plt.show()
from pygcn.utils import accuracy
def test(model):
y_pred = model(features.to('cuda'), adj.to('cuda')) # Usiamo tutto il dataset
acc_test = accuracy(y_pred[idx_test], labels[idx_test]) # Mascheriamo sulla parte di test
print("Accuracy:",
"accuracy= {:.4f}".format(acc_test.item()))
test(gcn)
|
Notebooks/GCN/GCN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pLoT
# language: python
# name: plot
# ---
# # Bayes (Part II)
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] tags=[]
# ## The Metropolis-Hastings algorithm
# -
# In the lecture this week, we have seen the _Metropolis-Hastings_ algorithm, which allows us to get samples from an _unnormalized_ distribution function, i.e., a function that returns a probability multiplied by some constant factor. Note that this can be any distribution (although in practice we will need it to get samples from an unnormalized posterior).
#
# First of all, have a look at the description of the algorithm and try to implement it as described here.
# > __**NOTE**__ You can find the implemented algo at the bottom of this notebook, but please try by yourself before looking at it!
def mcmc(nsamples, unnorm_prob_f, proposalf, burnin=100, initial=0.):
"""
Parameters
----------
nsamples: int
Number of samples to draw (includes burnin)
unnorm_prob_f: func
Function taking a point in the support and
returning its unnormalized probability
proposalf: func
Function that takes current position
and returns a proposal for where to move next
burnin: int
Number of initial samples to exclude
initial: float or func
If float, starts from that point.
If func, starts from the output of initial()
Returns
-------
list
A list of samples
"""
# set initial point in variable 'current' and
# calculate current probability ('curr_prob')
# ADD CODE HERE
# we put the samples in list 'states'
states = []
for i in range(nsamples):
# append current position to states
# ADD CODE HERE
# proposes a new point
# ADD CODE HERE
# calculate unnormalized probability
# at proposed point
# ADD CODE HERE
# calculates probability of acceptance
# in variable 'acceptance'
# ADD CODE HERE
if np.random.random_sample() < acceptance:
# if acceptance is higher than random btw 0 and 1,
# accept move, else stay where you are (i.e. do nothing)
# ADD CODE HERE
# return list of samples
return states[burnin:]
# This algorithm has a lot of pros:
# - Suprisingly simple to implement
# - If we get enough samples it is guaranteed to converge to the true posterior!
# - Works for both continuous and discrete parameter spaces
# But it also has some cons:
# - Can be very slow to converge
# - Does not work well with highly dimensional spaces
# - There is no way to tell if it converged to the true posterior
#
# In general, we should get as many samples as possible and run multiple chains with different initial points, to see if they converged to the same distribution.
# + [markdown] tags=[]
# ## Getting samples from an unnormalized normal distribution
# -
# The simplest application is to get samples from a distribution that we can calculate exactly, and see if the algorithm converges to it. Let's take the standard normal distribution, which has density function:
#
# $$
# \varphi (z)= \frac {1}{\sqrt {2\pi }} e^{-{\frac {z^{2}}{2}}}
# $$
#
# let's write a function to calculate this probability density:
def normal(x,mu=0,sigma=1):
"""
Probability density of normal distribution at x
"""
numerator = np.exp((-(x-mu)**2)/(2*sigma**2))
denominator = sigma * np.sqrt(2*np.pi)
return numerator/denominator
# And let's test this function to make sure it does what we expect:
xs = np.linspace(-3,3,1000)
plt.plot(xs, normal(xs))
plt.xlabel('x')
plt.ylabel('density')
plt.show()
# Now suppose that we didn't have the explicit formula for the distribution, but rather some black-box function where $\varphi$ is multiplied by an unknown constant $K$:
#
# $$
# \varphi' (z)= K \frac {1}{\sqrt {2\pi }} e^{-{\frac {z^{2}}{2}}}
# $$
#
# > __**NOTE**__: If $K$ is equal to the normalization constant $\sqrt {2\pi }$, $\varphi' (z)$ becomes just $e^{-{\frac {z^{2}}{2}}}$. This is essentially the trick we will use to sample from the posterior, where we don't have the posterior as such, but we can calculate the posterior multiplied by $P(D)$.
#
# Now we can use our mcmc function above to take samples from the distribution:
samples = mcmc(
# Number of samples
100000,
# Unnormalized density function
lambda x: np.exp(-x**2 / 2),
# Proposal function from point x
lambda x: np.random.normal(loc=x, scale=0.1),
)
# We can plot a histogram of the samples and compare it to the true distribution, and if you wrote the MCMC function correctly, they should look very close:
# plot a histogram of the samples
plt.hist(samples, bins=100, density=True)
# also plot the true distribution, just to be sure!
xs = np.linspace(-3,3,1000)
plt.plot(xs, normal(xs))
plt.show()
# + [markdown] tags=[]
# ## Using MHMC for Bayesian inference: categorization in continuous space
# -
# Now recall the categorization case that we saw in class: we observe some samples from an unknown category, and we have to form a posterior over the position of the unseen category's boundaries. The case we have seen in class has discrete boundaries, and therefore we could in theory calculate the posterior directly: consider all the possible hypotheses (categories) $H$, calculate $P(H) P(D \mid H)$, and sum it all to find $P(D)$.
#
# However, now imagine the case where we have a _continuous_ space, and so infinitely many categories. The probability density of sampling a certain observation from a category is still 0 if the observation lies outside the category and $1/|H|$ otherwise, but now the category $H$ can have a float size (e.g., category [-1, 0.5] has size 1.5). Now it becomes difficult to calculate the $P(D)$, because we can't easily integrate over all categories!
#
# However, if we define some prior over hypotheses (i.e. the categories), we can still easily calculate the numerator of the posterior, namely $P(H) P(D \mid H)$, for specific categories. A simple prior can be defined as follows:
#
# $$
# p(H) = \phi(x=l_H, \mu=0, \sigma=10) \; \phi(x=u_H, \mu=0, \sigma=10)
# $$
#
# where $\phi$ is the probability density function of a normal distribution, $l_H$ is the lower bound, and $u_H$ is the upper bound.
def prior_f(boundaries):
lb, ub = boundaries
return normal(lb, sigma=10) * normal(ub, sigma=10)
# Likelihood can be defined as discussed above:
#
# $$
# p(D \mid H) = \prod_{d \in D} \left( |H|^{-1} \text{ if } d \in H \text{ else } 0 \right)
# $$
#
# where $D$ is a set of observations (points from the unobserved category).
def likelihood_f(data, boundaries):
lb, ub = boundaries
compatible = all([lb < i < ub for i in data])
if compatible:
return 1/(ub-lb)
else:
return 0
# With the prior and the likelihood as defined above, we can write a function that given some data returns the unnormalized posterior function (i.e. a function form a category to the unnormalized posterior probability of the category given the data):
def define_unnorm_posterior(data):
def unnorm_posterior(boundaries):
return (
likelihood_f(data, boundaries) *
prior_f(boundaries)
)
return unnorm_posterior
# Finally, we just need a proposal function that takes a current category and proposes a new one. The proposal function here does this by moving the category up or down and stretching it by some amount:
def proposal_f(current_boundaries):
# transform boundaries linearly
# (i.e., move up and down and stretch)
shift = np.random.normal()
# we want the stretch to be always positive
# to preserve order of upper and lower bound
# (otherwise all samples with wrong order would
# have prob 0 in the likelihood function)
# Moreover, a stretching by some value x
# should be as likely as a contraction by x
# To preserve symmetry!
stretch = np.exp(np.random.normal())
return shift + stretch*current_boundaries
# Let's check the distribution of the upper and lower bounds of the proposals starting from category [-1,1]:
proposed = [proposal_f(np.array([-1,1])) for _ in range(10000)]
lbs, ubs = np.array(proposed).T
plt.hist(lbs, bins=100, alpha=0.2)
plt.hist(ubs, bins=100, alpha=0.2)
plt.xlim(-15, 15)
plt.show()
# Now we are ready to run approximate Bayesian inference on some data! Let's define some data first:
observations = [0.1, 2., 1.1]
# Take samples from the posterior distribution:
samples = mcmc(
# Number of samples
500000,
# Unnormalized density function
define_unnorm_posterior(observations),
# Proposal function from point x
proposal_f,
initial=np.array([-0.5, 0.5])
)
# Let's plot the posterior distributions of upper and lower bound:
lbs, ubs = np.array(samples).T
plt.hist(lbs, bins=100, density=True, label='Lower bound')
plt.hist(ubs, bins=100, density=True, label='Upper bound')
plt.xlim(-12, 12)
plt.scatter(
observations,
[1.]*len(observations),
color='red',
label='observations'
)
plt.legend()
plt.show()
# > __QUESTION__ What happens if we add more observations? Do you see the size effect in continuous space?
#
# > __EXERCISE__ (if there is time left at the end) Write a version of this inference model for 2-d space, where categories are squares and observations are points in 2d space.
# ## Using MHMC in discrete spaces
# We can use Metropolis-Hastings for discrete hypotheses spaces too! The basic idea is the same: we move around the space of hypotheses, calculating the unnormalized posterior for each point, and accepting new moves according to the rule described in the algo. The only complication is that we can't use the normal distribution centered at the current position as a proposal distribution, but we must come up with something different.
# As an example which will be useful in the future, consider the space of sentences generated by a PCFG as the space of hypotheses. For instance, the following PCFG:
#
# $$
# S \rightarrow a | b | Sa | Sb
# $$
#
# with uniform substitution probabilities (each $1/3$). The space of hypotheses then would be: 'a', 'b', 'aa', 'ab', 'ba', 'bb', and so on.
def complete(incomplete_sentence):
result = incomplete_sentence
options = ['a', 'b', 'Sa', 'Sb']
while 'S' in result:
result = np.random.choice(options) + result[1:]
return result
# test function
complete('Saa')
# The probability of a specific sentence of length $n$ then is $4^{-n}$ (can you see why?). Let's verify by sampling a bunch of sentences and finding the proportion of times we sample each one:
n = 10000
samples = [complete('S') for _ in range(n)]
sentences, counts = np.unique(samples, return_counts=True)
argsort = np.argsort(counts)[::-1]
sentences = sentences[argsort]
counts = counts[argsort] / n
plt.bar(x=sentences[:20], height=counts[:20])
plt.xticks(rotation=80)
plt.show()
# Based on this, it is easy to calculate the probability of each sentence:
def prior_f(sentence):
return 4**(-len(sentence))
# example of prior calculation
prior_f('a')
# We want to learn the sentence that produces some data. If we can see the sentence directly, this is easy: just look at the sentence and we're done. However, suppose that we can't see the sentence directly, but we see a 'noisy' version of it, where possibly some characters have been added at the end, where:
# - nothing is added with probability 1/3.
# - 'a' and 'b' are added with probability 1/3 each.
#
# We can calculate the resulting observation probability:
def likelihood_f(observations, sentence):
likelihood = 1
for observation in observations:
# Calculates probability of observation given true sentence
if observation == sentence:
likelihood *= 1/3
elif observation.startswith(sentence):
likelihood *= 1/3**(len(observation)-len(sentence))
else:
return 0
return likelihood
# test the function
print(likelihood_f(['abaa', 'abaaa', 'aba'], 'ab'))
print(likelihood_f(['abaa', 'abaaa', 'abab'], 'ab'))
print(likelihood_f(['abaa', 'abaaa', 'abab'], 'abb'))
# Like in the example above, we can now calculate the unnormalized posterior:
def define_unnorm_posterior(data):
def unnorm_posterior(sentence):
return (
likelihood_f(data, sentence) *
prior_f(sentence)
)
return unnorm_posterior
# Finally, we need a proposal distribution, which brings us from a current sentence to a possible next step. For MHMC to work as written above, it has to be the case that the probability of going from $x$ to $x'$ is the same as the probability of going from $x'$ to $x$. If this is not the case, we need to multiply the value of `acceptance` in `mcmc` by $\frac{P(x' \rightarrow x) }{ P(x \rightarrow x')}$ to balance things out.
#
# In practice, we will use the _subtree-regeneration_ transition distribution proposed at p.153 in Goodman, <NAME>., <NAME>, <NAME>, and <NAME>. “A Rational Analysis of Rule-Based Concept Learning.” Cognitive Science 32, no. 1 (2008): 108–54. https://doi.org/10.1080/03640210701802071. It works as follows, starting from current sentence $x$:
# - Sample a node $n$ at random from the parse tree of $x$
# - Remove everything below $n$ and replace $n$ with appropriate non-terminal, generating a tree $y$ containing non-terminals.
# - Complete $y$ with the PCFG, obtaining new sentence $x'$.
# - The acceptance probability from `mcmc` then gets additionally multiplied by:
#
# \begin{align}
# \frac{p(x' \rightarrow x)}{p(x \rightarrow x')}
# &=
# \frac{ P(\text{sampling node $n$ from $x'$}) }{ P(\text{sampling node $n$ from $x'$}) }
# \frac{ P(x \mid \text{PCFG}) }{ P(x' \mid \text{PCFG}) } \\
# &= \frac{|x'|^{-1}}{|x|^{-1}} \frac{4^{-|x|}}{4^{-|x'|}} \\
# &= \frac{|x|}{|x'|} \frac{4^{-|x|}}{4^{-|x'|}}
# \end{align}
#
def transition_f(current):
n = np.random.randint(len(current))
truncated = 'S' + current[:n]
return complete(truncated)
# test the function
transition_f('aba')
def acceptance_f(current, proposed, unnorm_prob_f):
move_when_lower = (
(unnorm_prob_f(proposed) * len(current) * 0.25**len(current))
/
(unnorm_prob_f(current) * len(proposed) * 0.25**len(proposed))
)
return min(
move_when_lower,
1
)
def mcmc(nsamples, unnorm_prob_f, proposalf, burnin=100, initial=0.):
"""
Same as mcmc above but with different acceptance_f
"""
current = initial() if callable(initial) else initial
curr_prob = unnorm_prob_f(current)
states = []
for i in range(nsamples):
states.append(current)
movement = proposalf(current)
move_prob = unnorm_prob_f(movement)
acceptance = acceptance_f(
current,
movement,
unnorm_prob_f
)
if np.random.random_sample() < acceptance:
current = movement
curr_prob = move_prob
return states[burnin:]
# Define some observations:
observations = ['aabaab', 'aaba', 'aabaaaa']
# Take samples with our algorithm:
samples = mcmc(
100000,
define_unnorm_posterior(observations),
transition_f,
initial='a'
)
# Plot the posterior distribution:
# +
sentences, counts = np.unique(samples, return_counts=True)
argsort = np.argsort(counts)[::-1]
sentences = sentences[argsort]
counts = counts[argsort] / len(samples)
plt.bar(x=sentences, height=counts)
plt.xticks(rotation=80)
plt.show()
# -
# Various things to be noticed:
# - The most likely sentence is the longest sentence that all the observations share, namely 'aaba'
# - The second most likely is 'aab', because it would be a 'suspicious' coincidence if all observations independently produced an 'a' right after 'aab', if the true sentence is indeed 'aab'.
# - Same, but even more so, applies to 'aa' and 'a'.
# - So even though 'aaba' is a priori less likely than 'a', it get a higher posterior probability because of the likelihood!
# ## Homework
# Third (and last) homeworks set! Consider the following CFG (interpret it as a PCFG with uniform probabilities), where each sentence from this grammar is an integer:
# $$
# S \rightarrow 1 | S+1 | S-1
# $$
# Note that there are multiple way to arrive at the same integer. E.g. $I(1+1-1)=I(1)$.
#
# 1. Write a function `prior_f` that takes a sentence produced by the PCFG and returns its probability in the PCFG.
#
# Assume that there is a true (unknown) sentence $H$ that we are trying to infer based on some observations ${o_1, \dots, o_n}$. The observations aren't just the integer defined by the sentence, but rather each observation is equal to the integer defined by the sentence plus some normally distributed noise. The likelihood function therefore is as follows:
# $$
# p(o \mid H) = P(H \mid \text{PCFG}) \; \phi(x= o-I(H), \mu=0, \sigma=2.)
# $$
# as usual, $I$ is the interpretation function, $\phi$ is the density function of the normal distribution. For instance if the true sentence was `1+1`, we might observe 1.94.
#
# 2. Write a function `likelihood_f` that calculates the probability of a list of observations given a sentence.
# > __HINT__ First find the integer $i$ defined by the sentence, then for each observation find the difference between $i$ and the observation, then find the probability of those differences (given that they are samples from a normal with $\mu=0$ and $\sigma=2$), and multiply those probabilities together.
# 3. Write a function to calculate the unnormalized posterior. This is essentially what the `define_unnorm_posterior` function above does.
#
# 4. Write a transition function: a function that takes a current sentence $x$ and returns a randomly generated proposal $x'$. You can modify the subtree-regeneration above to adapt it to this case.
# 5. Using mcmc, draw 100000 samples from $P(H \mid D)$.
# ## Implemented MHMC
def mcmc(nsamples, unnorm_prob_f, proposalf, burnin=100, initial=0.):
"""
Parameters
----------
nsamples: int
Number of samples to draw (includes burnin)
unnorm_prob_f: func
Function taking a point in the support and
returning its unnormalized probability
proposalf: func
Function that takes current position
and returns a proposal for where to move next
burnin: int
Number of initial samples to exclude
initial: float or func
If float, starts from that point.
If func, starts from the output of initial()
Returns
-------
list
A list of samples
"""
current = initial() if callable(initial) else initial
curr_prob = unnorm_prob_f(current)
# we put the samples in list 'states'
states = []
for i in range(nsamples):
# append current position to states
states.append(current)
# proposes a new point
movement = proposalf(current)
# calculate unnormalized probability
# at proposed point
move_prob = unnorm_prob_f(movement)
# calculates probability of acceptance
acceptance = min(move_prob/curr_prob,1)
# if acceptance is higher than random btw 0 and 1,
# accept move, else stay where you are (i.e. do nothing)
if np.random.random_sample() < acceptance:
current = movement
curr_prob = move_prob
# return list of samples
return states[burnin:]
|
book/7_lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing Jupyter with gitpod
print(2 ** 8)
import pandas as pd
import numpy as np
df = pd.DataFrame({ 'A' : 1.,
'B' : pd.Timestamp('20190102'),
'C' : pd.Series(1,index=list(range(4)),dtype='float32'),
'D' : np.array([3] * 4, dtype='int32'),
'E' : pd.Categorical(["car","cat","cam","cap"]),
'F' : 'foobar' })
# ## Display pd version
print(pd.__version__)
df
# +
# %matplotlib inline
import matplotlib.pyplot as plt
price = [100, 250, 380, 500, 700]
number = [1, 2, 3, 4, 5]
plt.plot(price, number)
plt.show()
# -
|
JNTesting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.9 64-bit
# language: python
# name: python3
# ---
# # Importing FB Prophet for Time Series Forecasting
# check prophet version
import prophet
# print version number
print('Prophet %s' % prophet.__version__)
# Loading the Data
# load the car sales dataset
from pandas import read_csv
# load data
# path = 'https://raw.githubusercontent.com/jbrownlee/Datasets/master/monthly-car-sales.csv'
# df = read_csv(path, header=0)
df = read_csv("Electric_Production.csv",header=0)
# summarize shape
print(df.shape)
# show first few rows
print(df.head())
# Visualizing the dataset
# +
# plot the car sales dataset
from matplotlib import pyplot
# plot the time series
df.plot()
pyplot.show()
# -
# # Fitting the FB Prophet model
# +
# fit prophet model on the car sales dataset
from pandas import to_datetime
from prophet import Prophet
# prepare expected column names
df.columns = ['ds', 'y']
df['ds']= to_datetime(df['ds'])
print(df)
# -
# define the model
model = Prophet()
# fit the model
model.fit(df)
from pandas import DataFrame
# define the period for which we want a prediction
future = list()
for i in range(1, 13):
date = '2017-%02d' % i
future.append([date])
future = DataFrame(future)
future.columns = ['ds']
future['ds']= to_datetime(future['ds'])
print(future)
# +
# use the model to make a forecast
forecast = model.predict(future)
# summarize the forecast
print(forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].head())
# -
# Plotting forecast on graph
# plot forecast
model.plot(forecast)
pyplot.show()
# # Out of sample forecast
# define the period for which we want a prediction
future = list()
for i in range(1, 13):
date = '2018-%02d' % i
future.append([date])
future = DataFrame(future)
future.columns = ['ds']
future['ds']= to_datetime(future['ds'])
# use the model to make a forecast
forecast = model.predict(future)
# summarize the forecast
print(forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].head())
# plot forecast
model.plot(forecast)
pyplot.show()
# # Model Evaluation
# create test dataset, remove last 13 months
train = df.drop(df.index[-13:])
print(train.tail())
# +
#from prophet import Prophet
from sklearn.metrics import mean_absolute_error
# define the model
model = Prophet()
# fit the model
model.fit(train)
# define the period for which we want a prediction
future = list()
for i in range(1, 13):
date = '2017-%02d' % i
future.append([date])
future = DataFrame(future)
future.columns = ['ds']
future['ds'] = to_datetime(future['ds'])
# use the model to make a forecast
forecast = model.predict(future)
# calculate MAE between expected and predicted values for december
y_true = df['y'][-12:].values
y_pred = forecast['yhat'].values
mae = mean_absolute_error(y_true, y_pred)
print('MAE: %.3f' % mae)
# -
# Plot of expected vs actual value
# plot expected vs actual
pyplot.plot(y_true, label='Actual')
pyplot.plot(y_pred, label='Predicted')
pyplot.legend()
pyplot.savefig("actual_vs_expected.png")
pyplot.show()
|
main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#importing libraries
import PIL
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
import keras
from keras.models import Sequential
from keras.layers import Convolution2D ,MaxPooling2D, Flatten, Dense,Dropout
from keras.utils import to_categorical
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
#from keras.layers import BatchNormalization
import shutil
import random
import os
import numpy as np
# +
#initialise the CNN model with different layers(used sequential): 3 layers
model = Sequential()
# Note the input shape is the desired size of the image 300x300 with 3 bytes color
# Step 1
model.add(Convolution2D(16,3,3, input_shape=(500, 500, 3),activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
#step 2 Convolution- learn features: put convolution filter on input(5X5) to get tensor output
model.add(Convolution2D(32,3,3, activation="relu"))
#step 3 Normalize the activations of the previous layer at each batch to increase stability and performance
#model.add(BatchNormalization())
#step 4 reduce spatial dimension
model.add(MaxPooling2D(pool_size =(2,2)))
#step 5 learn more features with activation function 'relu'to introduce non-lineraity (& avoid overfitting )
model.add(Convolution2D(64, 3, 3, activation = 'relu'))
#step 6 reduce spatial dimension
model.add(MaxPooling2D(pool_size=(2,2)))
#step 7 multidimensional to linear output
model.add(Flatten())
#step 8 connect every input to every output by weights (dot product)
model.add(Dense(128, activation = 'relu'))
model.add(Dropout(0.2))
model.add(Dense(3, activation = 'softmax'))
# -
#loss function is default for multi-classification problem
opt = keras.optimizers.SGD(learning_rate=0.01)
model.compile(optimizer=opt,loss='categorical_crossentropy', metrics = ['accuracy'])
# +
#augmentation configuration for training and testing
train_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = ImageDataGenerator(rescale = 1./255)
# -
# used this code when I manually put images in folder train under CHEST_xrxay/train.
#now has to be done ramdomly with code because manually picking images to train etc is not good
# Can comment out if need be
#32
training_set = train_datagen.flow_from_directory('CHEST_xray/train',
target_size = (500, 500),
batch_size = 32,
class_mode = 'categorical')
#used this code when I manually put images in folder train under CHEST_xrxay/validate
#now has to be done ramdomly with code because manually picking images is not good
# Can comment out if need be
validation_set = test_datagen.flow_from_directory('CHEST_xray/validate',
target_size = (500, 500),
batch_size = 32,
class_mode = 'categorical')
history=model.fit_generator(training_set,epochs =30,steps_per_epoch=62 ,validation_data = validation_set)
#validation set :Its purpose is to track progress through validation loss and accuracy.
# plot accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# plot loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='upper left')
plt.show()
# +
test_set = test_datagen.flow_from_directory('CHEST_xray/test',
target_size = (500, 500),
batch_size = 32,
class_mode = 'categorical', shuffle= False)
# +
#224
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
#from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score
#from sklearn.metrics import roc_auc
from sklearn.metrics import auc
subfolders=["covid", "normal", "viral_pneumonia"]
# loop over images/files in the subfolders of the test folder
#actual_list=[]
prediction_list=[]
count_covid=0
count_normal=0
count_viral_pneumonia=0
for actual in subfolders:
src_dir='CHEST_xray/test/'+actual
files=os.listdir(src_dir)
print(len(files))
for file in files:
if file !="Thumbs.db":
src_file_path=os.path.join(src_dir, file)
#print(src_file_path)
test_image = image.load_img(src_file_path, target_size = (500, 500))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = model.predict(test_image)
#print(result)
training_set.class_indices
#capturing prediction in an array
if int(result[0][0])==1:
prediction = 'Covid'
prediction_list.append(2)
elif int(result[0][1])==1:
prediction = 'Normal'
prediction_list.append(0)
elif int(result[0][2])==1:
prediction = 'Viral Pneumonia'
prediction_list.append(1)
else:
continue
# capturing actual in an array
if actual=="covid":
count_covid +=1
elif actual=="normal":
count_normal +=1
elif actual=="viral_pneumonia":
count_viral_pneumonia +=1
print(prediction)
actual_list=[2]*count_covid+[0]*count_normal+[1]*count_viral_pneumonia
print(len(actual_list))
print(len(prediction_list))
print(count_covid, count_normal, count_viral_pneumonia)
# calculate metrics
print(accuracy_score(actual_list, prediction_list))
print(precision_score (actual_list, prediction_list, average= 'macro',zero_division="warn"))
print(recall_score (actual_list, prediction_list, average='macro'))
#print(roc_auc_score (actual_list, prediction_list, average='None'))
print(confusion_matrix(actual_list, prediction_list))
#print(confusion_matrix (actual_list, prediction_list))
# +
#224
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
#from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score
#from sklearn.metrics import roc_auc
from sklearn.metrics import auc
subfolders=["covid", "normal", "viral_pneumonia"]
# loop over images/files in the subfolders of the test folder
#actual_list=[]
prediction_list=[]
count_covid=0
count_normal=0
count_viral_pneumonia=0
for actual in subfolders:
src_dir='CHEST_xray/test/'+actual
files=os.listdir(src_dir)
print(len(files))
for file in files:
src_file_path=os.path.join(src_dir, file)
#print(src_file_path)
test_image = image.load_img(src_file_path, target_size = (500, 500))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = model.predict_classes(test_image)
#print(result)
training_set.class_indices
#capturing prediction in an array
if result[0]==0:
#prediction = 'Covid'
prediction_list.append(2)
elif result[0]==1:
#prediction = 'Normal'
prediction_list.append(0)
elif result[0]==2:
#prediction = 'Viral Pneumonia'
prediction_list.append(1)
else:
pass
# capturing actual in an array
if actual=="covid":
count_covid +=1
elif actual=="normal":
count_normal +=1
elif actual=="viral_pneumonia":
count_viral_pneumonia +=1
else:
pass
#print(prediction)
actual_list=[2]*count_covid+[0]*count_normal+[1]*count_viral_pneumonia
print(len(actual_list))
print(len(prediction_list))
print(count_covid, count_normal, count_viral_pneumonia)
# calculate metrics
print(accuracy_score(actual_list, prediction_list))
print(precision_score (actual_list, prediction_list, average= 'macro'))
print(recall_score (actual_list, prediction_list, average='macro'))
#print(roc_auc_score (actual_list, prediction_list, average='None'))
print(confusion_matrix(actual_list, prediction_list))
# -
|
SGD0.01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="biGrpEVhCfa0" executionInfo={"status": "ok", "timestamp": 1612979361103, "user_tz": -180, "elapsed": 2682, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}}
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import scipy
# + [markdown] id="EJHPcclGC4PD"
# # The model
#
# # + Restriction operator is fixed.
# # + Petrov-Galerkin coarse-grid correction.
# # + Both smoothing operators are adjastable $\text{conv}D(A)^{-1}\left(b_k - A_k x_k\right)$.
# # + Serialization of three layers.
# + id="Y_sd-pxGCkOS" executionInfo={"status": "ok", "timestamp": 1612979363601, "user_tz": -180, "elapsed": 1519, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}}
# computes A_k*x_k
def apply_A(x, A, projection_filters, shapes, strides, k):
if k == 0:
return A(x)
else:
for i in range(k-1, -1, -1):
x = tf.nn.conv_transpose(x, projection_filters[i], shapes[i], strides[i], padding="VALID")
x = A(x)
for i in range(k):
x = tf.nn.convolution(x, projection_filters[i], strides[i])
return x
# find diagonal of an emplicitly defined matrix -- works for constant diagonals only
def extract_diagonal(A, projection_filters, shapes, strides, k):
y = np.zeros(shapes[k].numpy())
m = shapes[k].numpy()[1]//2 + 1
y[:, m, m, :] = 1
y = tf.Variable(y, dtype=projection_filters[0].dtype)
D = apply_A(y, A, projection_filters, shapes, strides, k)[0, m, m, 0]
return D
# slightly generalized Jacobi smoother
def smooth_r(x, b, A, D, projection_filters, shapes, strides, smoothing_filter, k):
r = b - apply_A(x, A, projection_filters, shapes, strides, k)
x = x + tf.nn.convolution(r/D, smoothing_filter, padding="SAME")
return x
# for some reason tf.nn.conv_transpose requires shape of an output -- this function compute required shapes
def compute_shapes(J, projection_filters, strides, batch_size=10):
shapes = []
x = tf.Variable(np.random.randn(batch_size, 2**J-1, 2**J-1, 1), dtype=projection_filters[0].dtype)
shapes.append(tf.shape(x))
for filter, stride in zip(projection_filters, strides):
x = tf.nn.convolution(x, filter, stride)
shapes.append(tf.shape(x))
return shapes
# multigrid
def multigrid_r(x, b, A, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post, k=0):
if k == len(projection_filters):
return b/apply_A(tf.ones_like(b), A, projection_filters, shapes, strides, k)
else:
D = extract_diagonal(A, projection_filters, shapes, strides, k)
x = smooth_r(x, b, A, D, projection_filters, shapes, strides, smoothing_filters_pre[k], k)
r = b - apply_A(x, A, projection_filters, shapes, strides, k)
r = tf.nn.convolution(r, projection_filters[k], strides=strides[k])
e = multigrid_r(tf.zeros_like(r), r, A, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post, k=k+1)
e = tf.nn.conv_transpose(e, projection_filters[k], shapes[k], strides=strides[k], padding="VALID")
x = x + e
x = smooth_r(x, b, A, D, projection_filters, shapes, strides, smoothing_filters_post[k], k)
return x
# approximate spectral radius
def stochastic_trace_r(A, J, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post, N_sweeps=10, batch_size=10):
x = tf.Variable(np.random.choice([-1, 1], (batch_size, 2**J-1, 2**J-1, 1)), dtype=projection_filters[0].dtype)
for _ in range(N_sweeps):
x = multigrid_r(x, x*0, A, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post)
return ((tf.norm(x)**2)/batch_size)**(1/(2*N_sweeps))
# + [markdown] id="OWQkMqPSAf03"
# # Equations
# + id="1try7DlJAfXZ" executionInfo={"status": "ok", "timestamp": 1612979370120, "user_tz": -180, "elapsed": 6747, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}}
# equations
poisson_filter = tf.Variable(np.array([[0, -1/4, 0], [-1/4, 1, -1/4], [0, -1/4, 0]]).reshape(3, 3, 1, 1), dtype=tf.float32, trainable=False, name="Poisson")
A_poisson = lambda x: tf.nn.convolution(x, poisson_filter, strides=[1, 1], padding="SAME")
poisson_large_filter = tf.Variable(np.array([[0, 0, 1/60, 0, 0], [0, 0, -16/60, 0, 0], [1/60, -16/60, 1, -16/60, 1/60], [0, 0, -16/60, 0, 0], [0, 0, 1/60, 0, 0]]).reshape(5, 5, 1, 1), dtype=tf.float32, trainable=False, name="Poisson large")
A_poisson_large = lambda x: tf.nn.convolution(x, poisson_large_filter, strides=[1, 1], padding="SAME")
mehrstellen_filter = tf.Variable(np.array([[-1/20, -1/5, -1/20], [-1/5, 1, -1/5], [-1/20, -1/5, -1/20]]).reshape(3, 3, 1, 1), dtype=tf.float32, trainable=False, name="Mehrstellen")
A_mehrstellen = lambda x: tf.nn.convolution(x, mehrstellen_filter, strides=[1, 1], padding="SAME")
epsilon = 10
poisson_anisotropic_filter = tf.Variable(np.array([[0, -1/(2+2*epsilon), 0], [-epsilon/(2+2*epsilon), 1, -epsilon/(2+2*epsilon)], [0, -1/(2+2*epsilon), 0]]).reshape(3, 3, 1, 1), dtype=tf.float32, trainable=False, name="Anisotropic Poisson")
A_anisotropic = lambda x: tf.nn.convolution(x, poisson_anisotropic_filter, strides=[1, 1], padding="SAME")
tau = 3/4
mixed_filter = tf.Variable(np.array([[-tau/8, -1/4, tau/8], [-1/4, 1, -1/4], [tau/8, -1/4, -tau/8]]).reshape(3, 3, 1, 1), dtype=tf.float32, trainable=False, name="Mixed")
A_mixed = lambda x: tf.nn.convolution(x, mixed_filter, strides=[1, 1], padding="SAME")
# + [markdown] id="pgSMtUfnDhOD"
# # Training
# + id="vWSyjZqWDkqZ" executionInfo={"status": "ok", "timestamp": 1612979389388, "user_tz": -180, "elapsed": 7098, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}}
A = A_poisson # change this line to try different linear equation
N_batch = 10
J = 5
N_levels = J - 1
# we are training three layers -- three variables for presmoothing, three variables for postsmoothing, restriction is fixed
proj_var = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1))/2, dtype=tf.float32, trainable=False, name="projection_var")
smoothing_pre_var_1 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="presmoothing_var 1")
smoothing_post_var_1 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="postsmoothing_var 1")
smoothing_pre_var_2 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="presmoothing_var 2")
smoothing_post_var_2 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="postsmoothing_var 2")
smoothing_pre_var_3 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="presmoothing_var 3")
smoothing_post_var_3 = tf.Variable(np.outer([1/2, 1, 1/2], [[1/2], [1], [1/2]]).reshape((3, 3, 1, 1)), dtype=tf.float32, trainable=True, name="postsmoothing_var 3")
smoothing_pre_var = [smoothing_pre_var_1, smoothing_pre_var_2, smoothing_pre_var_3]
smoothing_post_var = [smoothing_post_var_1, smoothing_post_var_2, smoothing_post_var_3]
projection_filters = [proj_var for i in range(N_levels)]
smoothing_filters_pre = [smoothing_pre_var[i%3] for i in range(N_levels)]
smoothing_filters_post = [smoothing_post_var[i%3] for i in range(N_levels)]
variables = smoothing_pre_var + smoothing_post_var
strides = [[2, 2]]*N_levels
shapes = compute_shapes(J, projection_filters, strides, batch_size=N_batch)
# + id="opSx8L2WDylZ" executionInfo={"status": "ok", "timestamp": 1612979472244, "user_tz": -180, "elapsed": 81098, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}}
opt = tf.keras.optimizers.Adam(learning_rate=0.01)
N_epoch = 200
history = np.zeros(N_epoch)
for i in range(N_epoch):
with tf.GradientTape() as tape:
loss = stochastic_trace_r(A, J, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post)
opt.apply_gradients(zip(tape.gradient(loss, variables), variables))
history[i] = loss.numpy()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="GwLJvaI5D1gJ" executionInfo={"status": "ok", "timestamp": 1612979472710, "user_tz": -180, "elapsed": 80370, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}} outputId="ebe6e403-1dd6-4c46-956f-c40b2c197f18"
# training history
final_loss = stochastic_trace_r(A, J, projection_filters, shapes, strides, smoothing_filters_pre, smoothing_filters_post, N_sweeps=10, batch_size=10).numpy()
fig, ax = plt.subplots()
ax.set_yscale("log")
ax.plot(history)
ax.set_xlabel("iteration")
ax.set_ylabel("loss")
print("final loss = ", final_loss);
# + [markdown] id="YkybvW-zDkUc"
# # Model evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="UhxE23eLDma2" executionInfo={"status": "ok", "timestamp": 1612979504725, "user_tz": -180, "elapsed": 27226, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14485856366916106708"}} outputId="3384bd0f-2b51-4542-f581-9d7f74532a4e"
N_batch = 10
for J in [3, 4, 5, 6, 7, 8, 9, 10, 11]:
N_levels = J - 1
# serialization
projection_filters_ = [proj_var for i in range(N_levels)]
smoothing_filters_pre_ = [smoothing_pre_var[i%3] for i in range(N_levels)]
smoothing_filters_post_ = [smoothing_post_var[i%3] for i in range(N_levels)]
strides_ = [[2, 2]]*N_levels
shapes_ = compute_shapes(J, projection_filters_, strides_, batch_size=N_batch)
spectral_radius = stochastic_trace_r(A, J, projection_filters_, shapes_, strides_, smoothing_filters_pre_, smoothing_filters_post_, N_sweeps=10, batch_size=10).numpy()
print("J = ", J, " spectral radius = ", spectral_radius)
|
s3MG(s).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Operadores lógicos
#
# Os operadores lógicos em Python combinam declarações combinam valores lógicos booleanos (True e False)
# ## And
# Retorna True se ambos os valores forem True
x = 1
print (x < 5 and x < 10)
x = 7
print (x < 5 and x < 10)
# ## Or
# Retorna True se algum dos valores forem True
x = 1
print (x < 5 or x < 10)
x = 7
print (x < 5 or x < 10)
# ## Not
# Reverte o resultado, de True para False e de False para True
x = 7
print(not(x < 5))
# ## Operadores Bitwise
# Os operadores Bitwise se assemelham aos operadores lógicos, mas comparam números binários.
# 0 = Falso
# 1 = Verdadeiro
# ## &
# (AND)
#
# Se todos os valores forem 1, retorna 1
# +
x = 1
y = 1
z = x & y
print(z)
# -
# ## |
# (OR)
# (Ou inclusivo)
#
# Se algum dos valores for 1, retorna 1
# +
x = 0
y = 1
z = x | y
print(z)
# -
# ## ^
# (OR)
# (Ou exclusivo)
#
# Se apenas um dos valores for 1, retorna 1
# +
a = 0
b = 1
c = 1
z = a ^ b ^ c
print(z)
# +
a = 0
b = 1
c = 0
z = a ^ b ^ c
print(z)
# -
print(type(1))
|
assunto-6_operadores_logicos/Operadores_logicos_e_bitwise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Bike Share in Washington D.C.
# +
import numpy as np
import pandas as pd
import datetime as dt
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression, Ridge, LinearRegression
from sklearn.metrics import accuracy_score
from sklearn import metrics
from sklearn.metrics import mean_squared_log_error
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import KBinsDiscretizer
from sklearn.preprocessing import MinMaxScaler
import warnings
# %matplotlib inline
warnings.filterwarnings('ignore')
# To see all the columns after doing Feature Engineering, otherwise normally it displays only few columns.
pd.options.display.max_columns = 999
# -
# ## Get the Data
bike = pd.read_csv('./data/train.csv', parse_dates=True, index_col=0)
bike.sample(5)
# +
# Displays number of rows and columns of our DataFrame
bike.shape
# +
# Parsing the datetime
bike['hour'] = bike.index.hour
bike['month'] = bike.index.month
bike['DayOfWeek'] = bike.index.day_name()
bike.sample(5)
# -
bike.shape
# +
# Displays Statistical Information
bike.describe()
# +
# Displays "DataType" information
bike.info()
# +
# Displays unique values
bike.apply(lambda x: len(x.unique()))
# -
# ## Preprocessing Data
# +
# Check for NaN/null values in the DataFrame
bike.isna().sum()
# bike.isnull().sum()
# -
# ## Visualizing the Data
# ### Correlation Matrix
bike.corr()
corr = bike.corr()
plt.figure(figsize=(15, 10))
sns.heatmap(corr,
vmin=-1,
vmax=1,
cmap="PiYG",
annot=True,
annot_kws={'size':12},
)
fig, ax = plt.subplots(figsize=(20, 10))
sns.pointplot(data=bike, x='hour', y='count', hue='DayOfWeek', ax=ax)
plt.title('Count of bikes during weekdays and weekends')
fig, ax = plt.subplots(figsize=(20, 10))
sns.barplot(data=bike, x='hour', y='count')
ax.set(title='Count of bikes during weekdays and weekends')
# +
bike_workday = bike.groupby('workingday').agg({'casual':'sum','registered': 'sum'})
bike_workday.rename(index={0: 'Not WorkDay',1: 'WorkDay'}, inplace=True)
bike_workday
# +
f, axes = plt.subplots(1,2, figsize=(15,10))
for ax, col in zip(axes, bike_workday.columns):
bike_workday[col].plot(kind='pie', autopct='%.2f', ax=ax, title=col, fontsize=15)
ax.legend(loc=3)
# -
fig = plt.subplots(figsize=(10, 5))
sns.barplot(data=bike, x='month', y='count')
plt.title("Count of Bikes during different months.")
fig = plt.subplots(figsize=(12, 7))
sns.barplot(data=bike, x='DayOfWeek', y='count', hue='season', palette=['steelblue'])
sns.despine()
plt.title("Count of Bikes during different Days.")
sns.barplot(x='season', y='count', data=bike)
plt.xticks(ticks=[0, 1, 2, 3], labels=['winter', 'spring', 'summer', 'fall'])
plt.title("Count of Bikes during different Seasons of the year.")
# +
# Realtion between Temperature and Humidity for Usage
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(20, 6)) # ax1, ax2 - Temperature and Humidity
# Temperature and Users, when Temperature increases, no.of Users also increases
sns.scatterplot(x=bike['temp'], y=bike['count'], ax=ax1)
plt.title('Relation between Temperature and Users')
# Humidity and Users , when Humidity increases no.of Users decreases.
sns.scatterplot(x=bike['humidity'], y=bike['count'], ax=ax2)
sns.despine()
plt.title('Relation between Humidity and Users')
# -
# ### Select columns for Feature Matrix (X) and target(y)
X = bike[['holiday', 'workingday', 'temp', 'windspeed', 'season', 'weather', 'DayOfWeek', 'month', 'hour']]
y = bike['count']
X.head()
# ### Splitting data into train and test - not Required while working with Kaggle Dataset
# +
# X_train, X_test, y_train, y_test = train_test_split(X, y)
# +
# X_train.shape, X_test.shape
# -
# ### Feature Engineering
# from sklearn.preprocessing import PolynomialFeatures
from sklearn.ensemble import RandomForestRegressor
fe_eng = ColumnTransformer([
('Scaling', MinMaxScaler(), ['temp', 'windspeed']),
('do-nothing', 'passthrough', ['holiday', 'workingday']),
('One-Hot-Encoding', OneHotEncoder(sparse=False, handle_unknown = 'ignore'), ['season', 'weather', 'DayOfWeek', 'month', 'hour'])
])
# +
# fit the column transformer on the training data
# fe_eng.fit(X_train)
# -
# ## Model selection
model = make_pipeline(
fe_eng,
# PolynomialFeatures(degree=2, include_bias=False), # not Required for Random Forest (bcz increases calculation time)
RandomForestRegressor(n_estimators=30, max_depth=32) # 32
)
# +
# from sklearn.model_selection import cross_validate
# -
# ## Cross Validation
from sklearn.metrics import make_scorer
# +
def rmsle(y, yhat):
yhat[yhat<0]=0
return mean_squared_log_error(y, yhat)**0.5
rmsle_loss = make_scorer(rmsle, greater_is_better=False)
# +
# cv_results = cross_validate(
# estimator=model, # the model you want to evaluate
# X=X, # the training input data
# y=y, # the training output data
# cv=5, # number of cross validation datasets
# scoring=rmsle_loss, # evaluation metric
# return_train_score=True, # return both the score on the training and the cross validated data
# n_jobs=-1 # n_jobs = -1 for using all your processores
# )
# +
# show results in dataframe
# df_cv_results = pd.DataFrame(cv_results)
# df_cv_results
# +
# take a look at the mean of all the results
# df_cv_results.mean()
# +
# fit and transform the Training data
# model.fit(X, y)
# -
# #### Not Required while using Cross_Validation
# +
# Transforming and Predicting Test data
# y_pred = model.predict(X_test)
# +
# y_pred.max(), y_pred.min()
# +
# plt.scatter(y_test, y_pred)
# +
# metrics.mean_absolute_error(y_test, y_pred)
# +
# root mean squared log error(RMSLE)
# y_pred[y_pred<0] = 0
# metrics.mean_squared_log_error(y_test, y_pred)**0.5
# -
# ## Hyperparameter - GridSearchCV
# +
# to get a list of the hyperparamters
model.get_params()
# +
# pick the paramaters you want to search through and the corresponding values you would like to try
param_grid = {
'randomforestregressor__max_depth': [16, 32], # No.of levels in tree
# 'randomforestregressor__max_features': ['auto', 'sqrt'], # no. of features to consider at every split
'randomforestregressor__n_estimators': [30, 50, 100, 150]
}
# -
from sklearn.model_selection import GridSearchCV
# +
# define your gridsearch
grid_cv = GridSearchCV(
estimator=model,
param_grid= param_grid,
cv=5,
return_train_score=True,
scoring=rmsle_loss,
n_jobs=-1
)
# +
# fit all models with all the different hyperparamters
grid_cv.fit(X, y)
# +
# look at different reults that you can check to evaluate the different models
grid_cv.cv_results_.keys()
# -
col=[
'param_randomforestregressor__n_estimators',
'param_randomforestregressor__max_depth',
'mean_train_score',
'mean_test_score',
'rank_test_score'
]
cv_results=pd.DataFrame(grid_cv.cv_results_)
cv_results[col]
cv_results[col].abs()
grid_cv.best_score_
grid_cv.best_params_
# ## Kaggle
bike_test = pd.read_csv('./data/test.csv', parse_dates=True, index_col=0)
bike_test.head()
bike_test.shape
# +
# Parsing the datetime
bike_test['hour'] = bike_test.index.hour
bike_test['month'] = bike_test.index.month
bike_test['DayOfWeek'] = bike_test.index.day_name()
bike_test.sample(5)
# -
X_kaggle = bike_test[['holiday', 'workingday', 'temp', 'windspeed', 'season', 'weather', 'DayOfWeek', 'month', 'hour']]
X_kaggle.head()
bike_test.reset_index(inplace=True)
# +
model.fit(X,y)
# +
# Prediction
y_pred = model.predict(X_kaggle)
y_pred
# -
y_pred[y_pred<0] = 0
y_pred
output = pd.DataFrame({'datetime': bike_test['datetime'], 'count': y_pred})
output.to_csv('./bike_kaggle_submission/bike_kaggle_submission.csv', index=False)
# ## This approach gave me a Kaggle Score - 0.51
|
bike_share_RandomForest_GridSearch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import print_function
# Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import psycopg2
import sys
import datetime as dt
import mp_utils as mp
USE_SQL=0
USE_CSV=1
# +
# below config used on pc70
sqluser = 'alistairewj'
dbname = 'mimic'
schema_name = 'mimiciii'
query_schema = 'SET search_path to public,' + schema_name + ';'
if USE_SQL:
# Connect to local postgres version of mimic
con = psycopg2.connect(dbname=dbname, user=sqluser)
# exclusion criteria:
# - less than 16 years old
# - stayed in the ICU less than 4 hours
# - never have any chartevents data (i.e. likely administrative error)
query = query_schema + \
"""
select
subject_id, hadm_id, icustay_id
from mp_cohort
where excluded = 0
"""
co = pd.read_sql_query(query,con)
# extract static vars into a separate dataframe
df_static = pd.read_sql_query(query_schema + 'select * from mp_static_data', con)
#for dtvar in ['intime','outtime','deathtime']:
# df_static[dtvar] = pd.to_datetime(df_static[dtvar])
vars_static = [u'is_male', u'emergency_admission', u'age',
# services
u'service_any_noncard_surg',
u'service_any_card_surg',
u'service_cmed',
u'service_traum',
u'service_nmed',
# ethnicities
u'race_black',u'race_hispanic',u'race_asian',u'race_other',
# phatness
u'height', u'weight', u'bmi']
# get ~5 million rows containing data from errbody
# this takes a little bit of time to load into memory (~2 minutes)
# # %%time results
# CPU times: user 42.8 s, sys: 1min 3s, total: 1min 46s
# Wall time: 2min 7s
df = pd.read_sql_query(query_schema + 'select * from mp_data', con)
df.drop('subject_id',axis=1,inplace=True)
df.drop('hadm_id',axis=1,inplace=True)
df.sort_values(['icustay_id','hr'],axis=0,ascending=True,inplace=True)
print(df.shape)
# get death information
df_death = pd.read_sql_query(query_schema + """
select
co.subject_id, co.hadm_id, co.icustay_id
, ceil(extract(epoch from (co.outtime - co.intime))/60.0/60.0) as dischtime_hours
, ceil(extract(epoch from (adm.deathtime - co.intime))/60.0/60.0) as deathtime_hours
, case when adm.deathtime is null then 0 else 1 end as death
from mp_cohort co
inner join admissions adm
on co.hadm_id = adm.hadm_id
where co.excluded = 0
""", con)
# get severity scores
df_soi = pd.read_sql_query(query_schema + """
select
co.icustay_id
, case when adm.deathtime is null then 0 else 1 end as death
, sa.saps
, sa2.sapsii
, aps.apsiii
, so.sofa
, lo.lods
, oa.oasis
from mp_cohort co
inner join admissions adm
on co.hadm_id = adm.hadm_id
left join saps sa
on co.icustay_id = sa.icustay_id
left join sapsii sa2
on co.icustay_id = sa2.icustay_id
left join apsiii aps
on co.icustay_id = aps.icustay_id
left join sofa so
on co.icustay_id = so.icustay_id
left join lods lo
on co.icustay_id = lo.icustay_id
left join oasis oa
on co.icustay_id = oa.icustay_id
where co.excluded = 0
""", con)
# get censoring information
df_censor = pd.read_sql_query(query_schema + """
select co.icustay_id, min(cs.charttime) as censortime
, ceil(extract(epoch from min(cs.charttime-co.intime) )/60.0/60.0) as censortime_hours
from mp_cohort co
inner join mp_code_status cs
on co.icustay_id = cs.icustay_id
where cmo+dnr+dni+dncpr+cmo_notes>0
and co.excluded = 0
group by co.icustay_id
""", con)
# -
if USE_CSV:
co = pd.read_csv('df_cohort.csv')
# convert the inclusion flags to boolean
for c in co.columns:
if c[0:10]=='inclusion_':
co[c] = co[c].astype(bool)
df = pd.read_csv('df_data.csv')
df_static = pd.read_csv('df_static_data.csv')
df_censor = pd.read_csv('df_censor.csv')
df_soi = pd.read_csv('df_soi.csv')
df_static.head()
co.columns
# +
idxRem = np.zeros(co.shape[0],dtype=bool)
for c in co.columns:
if 'exclusion_' in c:
print('{:5g} - {:2.2f}% - {}'.format(co[c].sum(),
co[c].mean()*100.0,
c))
idxRem[co[c].values==1] = True
print('{:5g} - {:2.2f}% - {}'.format(np.sum(idxRem),
np.mean(idxRem)*100.0,
'total removed'))
print('{:5g} - {:2.2f}% - {}'.format(np.sum(~idxRem),
np.mean(~idxRem)*100.0,
'final cohort'))
# -
print(df_static.shape)
print(df['icustay_id'].nunique())
# +
# generate k-fold indices
np.random.seed(111)
K = 5 # number of folds
# get unique subject_id
sid = np.sort(np.unique(df_static['subject_id'].values))
# assign k-fold
idxK_sid = np.random.permutation(sid.shape[0])
idxK_sid = np.mod(idxK_sid,K)
# +
var_min, var_max, var_first, var_last, var_sum, var_first_early, var_last_early, var_static = mp.vars_of_interest()
# create window time for each patient
df_tmp=df_death.copy().merge(df_censor, how='left', left_on='icustay_id', right_on='icustay_id')
time_dict = mp.generate_times(df_tmp, T=2, seed=111, censor=True)
# generate windows
df_data = mp.get_design_matrix(df, time_dict, W=8, W_extra=24)
# remove icustay_ids if they were censored (made DNR) before icu admission, or close enough to that
idx = df_censor.loc[df_censor['censortime_hours']<=0, 'icustay_id']
print('Removed {} icustay_id as they were censored on/before ICU admission.'.format((idx.shape[0])))
df_data.drop(idx, axis=0, inplace=True)
# first, the data from static vars from df_static
X = df_data.merge(df_static.set_index('icustay_id')[var_static], how='left', left_index=True, right_index=True)
# next, add in the outcome: death in hospital
X = X.merge(df_death.set_index('icustay_id')[['death']], left_index=True, right_index=True)
# generate K-fold indices
X = X.merge(df_death.set_index('icustay_id')[['subject_id']], left_index=True, right_index=True)
# get indices which map subject_ids in sid to the X dataframe
idxMap = np.searchsorted(sid, X['subject_id'].values)
# use these indices to map the k-fold integers
idxK = idxK_sid[idxMap]
# add idxK to design matrix
X['idxK'] = idxK
# write to file
X.to_csv('X_design_matrix.csv')
|
notebooks/mp-prep-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pathlib import Path
import sys
sys.path.append(str(Path('.').absolute().parent)) #
import hiddenlayer as hl
import matplotlib.pyplot as plt
import pandas as pd
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data
import torchvision
import torchvision.models as models
from PIL import Image, ImageFile
from torchvision import models, transforms
from tqdm.notebook import tqdm
import math
import seaborn as sns
import re
from plasma.training import find_lr, train
from plasma.utils import show_model, plot_history
sns.set(style="whitegrid")
# %matplotlib inline
# -
# ## Visualization of Model Architecture
# +
resnet_50 = torch.hub.load("pytorch/vision", "resnet50")
show_model(re)
# -
# ## Training Neural Networks
# +
from sklearn.datasets import fetch_openml
X, y = fetch_openml('mnist_784', return_X_y=True, as_frame=False)
# +
# https://www.kaggle.com/pinocookie/pytorch-simple-mlp
class MLP(nn.Module):
def __init__(self):
super(MLP, self).__init__()
self.layers = nn.Sequential(
nn.Linear(784, 100),
nn.ReLU(),
nn.Linear(100, 10)
)
def forward(self, x):
# convert tensor (128, 1, 28, 28) --> (128, 1*28*28)
x = x.view(x.size(0), -1)
x = self.layers(x)
return x
simple_mlp = MLP()
# -
train()
|
examples/getLearningRate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Boundary Conditions
#
# In this workbook we are going to modify our fast checkerboard *SOR* solver so that it can manage boundary conditions.
#
# Remember we are solving
# $$u_{xx}+u_{yy}=0$$
# The boundary conditons can be
# * **Dirichlet** boundary conditions where a value of the unknown, _u_, is specified, e.g. $$u(0,y)=273$$
# * **Neumann** boundary conditions where a value of the derivative, _u<sub>x</sub>_, is specified. e.g. $$u_x(0,y)=0.$$
# * **Robin** boundary conditions where the value of a function of the unknown and the derivative is specified, e.g. $$u_x(0,y)-2u(0,y)=1.$$
#
# ## Neumann boundary conditions
# We have already implemented a **Dirichlet** boundary condition in our solver, and **Robin** boundary conditions are very rare, so let us consider **Neumann** conditions.
#
# Supose we have
# $$u_x(0)=0$$
# Since we are already using a 2nd order central finite difference approximation, it seems best to do the same for this partial derivative too,
# $$\frac{\partial u}{\partial x} \approx \frac{u_W-u_E}{2\Delta x}.$$
# At the boundary
# $$\frac{\partial u}{\partial x} =0$$ so
# $$0 = \frac{u_W-u_E}{2\Delta x} \implies u_W=u_E.$$
# This is the so-called _symmetry_ boundary condition.
#
# ### updating the Grid Class
# We are going to need some flags that tell us if a boundary is a Neumann BC or a Dirichlet BC, so the `Grid` class needs extending.
#
# We can define some flags DIRICHLET_BC, NEUMANN_BC and BC_NAME within the class, give every grid left, right, top and bottom flags which are set to *Dirichlet* by default and write a little function to specify boundaries and annother to report them.
# +
import numpy as np
import matplotlib.pyplot as plt
import time
class Grid:
'''Class defining a 2D computational grid. The grid object
contains is a regular cartesian grid with a single variable, u.
It stores information about the number of grid points in the i
and j directions, the ordinates of these points and the bottom
left corner of the gird (the origin) and the top right corner
(the extent).
Written by Prof <NAME>, School of Engineering
(c) 2021 The University of Edinburgh
Licensed under CC-BY-NC.'''
DIRICHLET_BC = 0
NEUMANN_BC = 1
BC_NAME = ['left', 'right', 'top', 'bottom']
def __init__(self,ni,nj):
# set up information about the grid
self.origin = (0.0, 0.0) # bottom left
self.extent = (1.0, 1.0) # top right
self.Ni = ni # grid points in i direction
self.Nj = nj # grid points in j direction
# initialse x,y and u arrays
self.u = np.zeros((nj, ni))
self.x = np.zeros((nj, ni))
self.y = np.zeros((nj, ni))
# boundary conditions (left right top and bottom)
self.BC = [self.DIRICHLET_BC, self.DIRICHLET_BC,
self.DIRICHLET_BC, self.DIRICHLET_BC]
def set_origin(self,x0,y0):
self.origin = (x0, y0)
def set_extent(self,x1,y1):
self.extent = (x1, y1)
def generate(self,Quiet=True):
'''generate a uniformly spaced grid covering the domain from the
origin to the extent. We are going to do this using linspace from
numpy to create lists of x and y ordinates and then the meshgrid
function to turn these into 2D arrays of grid point ordinates.'''
x_ord = np.linspace(self.origin[0], self.extent[0], self.Ni)
y_ord = np.linspace(self.origin[1], self.extent[1], self.Nj)
self.x, self.y = np.meshgrid(x_ord,y_ord)
if not Quiet:
print(self)
def Delta_x(self):
# calculate delta x
return self.x[0,1]-self.x[0,0]
def Delta_y(self):
# calculate delta y
return self.y[1,0]-self.y[0,0]
def find(self,point):
'''find the i and j ordinates of the grid cell which contains
the point (x,y). To do this we calculate the distance from
the point to the origin in the x and y directions and then
divide this by delta x and delta y. The resulting real ordinates
are converted to indices using the int() function.'''
grid_x = (point[0] - self.origin[0])/self.Delta_x()
grid_y = (point[1] - self.origin[1])/self.Delta_y()
return int(grid_x), int(grid_y)
def set_Neumann_bc(self,side):
try:
self.BC[self.BC_NAME.index(side)] = self.NEUMANN_BC
except:
print('error {} must be one of {}'.format(side,self.BC_NAME))
def set_Dirichlet_bc(self,side):
try:
self.BC[self.BC_NAME.index(side)] = self.DIRICHLET_BC
except:
print('error {} must be one of {}'.format(side,self.BC_NAME))
def report_BC(self):
'''compile a string listing the boundary conditions on each side.
We build up a string of four {side name}: {BC type} pairs and
return it'''
# initialise the string
string = ''
# loop over the sides
for side in range(4):
# add the side name
string = string + self.BC_NAME[side]
# and the bounday condition type
if self.BC[side] == self.DIRICHLET_BC:
string = string + ': Dirichlet, '
elif self.BC[side] == self.NEUMANN_BC:
string = string + ': Neumann, '
return string[:-2] +'.' # lose the last comma and space.
def __str__(self):
# describe the object when asked to print it
describe = 'Uniform {}x{} grid from {} to {}.'.format(self.Ni, self.Nj, self.origin, self.extent)
boundaries = self.report_BC()
return describe + '\nBoundaries conditions are - ' + boundaries
# -
test = Grid(11,11)
test.set_Neumann_bc('right')
print(test)
# ## Indirect adressing.
# The SOR solver loops over all the interior cells in `u_new` using whole array opperations.
#
# u_new.flat[centre[::2]] = (1-omega) * u_new.flat[centre[::2]] + \
# omega * C_beta*(u_new.flat[north[::2]]+ \
# u_new.flat[south[::2]]+ \
# beta_sq*(u_new.flat[east[::2]]+ \
# u_new.flat[west[::2]]))
#
# We call this **indirect** adressing because the locations in `u_new` are given by the entries in the `centre[]`, `north[]`, `south[]`, `east[]` and `west[]` arrays. The good news is that this means only need to change the cells refered to in the `east[]` array on the left hand boundary if we want to implement the Neumann boundary condition _u<sub>x</sub>=0_.
#
# The bit of the SOR code that needs modifying is the loop over the interior points that builds the lists.
#
# for j in range(1,mesh.Nj-1):
# for i in range(1,mesh.Ni-1):
# k=i+offset*j # calculate the k value
# # save the south, west, centre, east and north points
# centre.append(k)
# north.append(k+offset)
# east.append(k+1)
# south.append(k-offset)
# west.append(k-1)
#
# If we are next to a boundary **and** it's a Neumann boundary condition we need to adjust the pointer to point to the interior cell. So the west cell would be at k+1 and the east cell at k-1.
#
#
#
# +
def SOR(mesh,tol=0.5e-7,maxit=10000):
'''Sucessive over Relaxation method for the solution of
the Laplace equation with checkerboarding. The solver
assumes a uniform Cartesian mesh and uses the optimal
value of the acceleration parameter, omega.
The method works on the grid stored in the mesh object.
It will continue itterating until the relative difference
between u^{n+1} and u^n is less than tol. It will also stop
if we have done more than maxit itterations.
The solution stored in the mesh.u variable is updated
Written by Prof <NAME>, School of Engineering
(c) 2021 The University of Edinburgh
Licensed under CC-BY-NC.'''
# calculate the optimal value of omega
lamda = (np.cos(np.pi/mesh.Ni)+np.cos(np.pi/mesh.Nj))**2/4
omega = 2/(1+np.sqrt(1-lamda))
# calculate the coefficients
beta = mesh.Delta_x()/mesh.Delta_y()
beta_sq = beta**2
C_beta = 1/(2*(1+beta_sq))
# initialise u_new
u_new = mesh.u.copy()
# build the list of k values
offset = mesh.u.shape[1] # how far away are j+1 and j-1
# create some empty lists
centre = []; north = []; east = []; south=[]; west = []
# loop over all the interior points
for j in range(1,mesh.Nj-1):
for i in range(1,mesh.Ni-1):
k=i+offset*j # calculate the k value
# save the centre point
centre.append(k)
# do the north boundary checking for a Neumann bounary condition
if (j == mesh.Nj-2) and (mesh.BC[2]== mesh.NEUMANN_BC):
north.append(k-offset)
else:
north.append(k+offset)
# do the east boundary checking for a Neumann boundary condition
if (i == mesh.Ni-2) and (mesh.BC[1]== mesh.NEUMANN_BC):
east.append(k-1)
else:
east.append(k+1)
# do the sound boundary checking for a Neumann boundary condition
if (j==1) and (mesh.BC[3]== mesh.NEUMANN_BC):
south.append(k+offset)
else:
south.append(k-offset)
# do the west boundary checking for a Neumann boundary condition
if (i==1) and (mesh.BC[0]== mesh.NEUMANN_BC):
west.append(k+1)
else:
west.append(k-1)
# itteration
for it in range(maxit):
# red squares [::2] means from 0 to n in steps of 2, remember
# we are taking alternate values from the lists centre, east,
# north, west and south and using them as indicies into the
# u_new array.
u_new.flat[centre[::2]] = (1-omega) * u_new.flat[centre[::2]] + \
omega * C_beta*(u_new.flat[north[::2]]+ \
u_new.flat[south[::2]]+ \
beta_sq*(u_new.flat[east[::2]]+ \
u_new.flat[west[::2]]))
# black squares [1::2] means from 1 to n in steps of 2
u_new.flat[centre[1::2]] = (1-omega) * u_new.flat[centre[1::2]] + \
omega * C_beta*(u_new.flat[north[1::2]]+ \
u_new.flat[south[1::2]]+ \
beta_sq*(u_new.flat[east[1::2]]+ \
u_new.flat[west[1::2]]))
# compute the difference between the new and old solutions
err = np.max(np.abs(mesh.u-u_new))/np.max(np.abs(mesh.u))
# update the solution
mesh.u = np.copy(u_new)
# # converged?
if err < tol:
break
if it+1 == maxit:
print('Checkerboard Gauss-Seidel itteration failed to converge, error = {}'.format(err))
return it+1,err
# -
# ## The original test problem
# Were going to use the same test problem as before to make sure it still works!
#
# $$0=u_{xx}+u_{yy}$$
# subject to the boundary conditions
# $$\begin{align*}
# u(x,0)&=0&0\le x\le 2 \\
# u(x,1)&=0&0\le x\le 2 \\
# u(0,y)&=0&0\le y \le 1\\
# u(2,y)&=\sin 2\pi y&0\le y \le 1.
# \end{align*}$$
#
def Example929(ni,nj):
# set up a mesh
mesh = Grid(ni,nj)
mesh.set_extent(2.0,1.0)
mesh.generate()
# now the RHS boundary condition
mesh.u[:,-1]=np.sin(2*np.pi*mesh.y[:,-1])
return mesh
# +
# Test problem on a fine grid
test = Example929(161,81)
print(test)
# run the solver
start = time.process_time()
itt, err = SOR(test,tol=0.5e-14)
stop = time.process_time()
print("Solver took {:.3g} seconds.".format(stop-start))
print('Converged after {} itterations, final residual is {}'.format(itt,err))
# plot the solution
fig, ax1 = plt.subplots()
cmap = plt.get_cmap('PiYG')
cf = ax1.contourf(test.x,test.y,test.u,cmap=cmap)
fig.colorbar(cf, ax=ax1)
ax1.set_title(f'Example 9.29 ({test.Ni} x {test.Nj} grid)')
plt.show()
# -
# ## A new test problem
#
# Now we know we haven't changed the solution to our original test problem we need one that includes Neumann and Dirichlet boundary conditions.
#
# ### Porous Flow
#
# Water seeping through a porous medium can be modelled using the Laplace equation. If we have a "box" _z<sub>0</sub>_ meters tall and _s_ meters wide with a linear pressure gradient _h = z<sub>0</sub> + cx_ applied at the upper boundary, an impermeable base and constant flow boundaries on the left and right. The pressure in the medium can be modlled as
#
# $$h_{zz}+h_{xx}=0$$ with boundary conditions
#
# $$h_z(x,0)=0,\,h_x(0,z)=0\text{ and }h_x(s,z)=0$$
# and
# $$h(x,z_0)=z_0+c x.$$
#
# This test problem has an anlytical solution developed by Toth (1962):
# $$h(x,z) = z_0+\frac{cs}{2}-\frac{4cs}{\pi^2}\sum_{m=0}^\infty
# \frac{\cos[(2m+1)\pi x/s]\cosh[(2m+1)\pi z/s]}
# {(2m+1)^2\cosh[(2m+1)\pi z_0/s]}.$$
#
# Let us consider a 200 wide by 100 m deep medium with _c_ = 0.176.
# +
def porous(ni,nj):
# set up a mesh
mesh = Grid(ni,nj)
mesh.set_extent(200.0,100.0)
mesh.generate()
# set the boundary conditions
mesh.set_Neumann_bc('left')
mesh.set_Neumann_bc('right')
mesh.set_Neumann_bc('bottom')
# now the top boundary condition
mesh.u[-1,:]=100.0 + 0.176 * mesh.x[-1,:]
return mesh
ptest = porous(101,51)
print(ptest)
# run the solver
start = time.process_time()
itt, err = SOR(ptest,tol=0.5e-10,maxit=100000)
stop = time.process_time()
print("Solver took {:.3g} seconds.".format(stop-start))
print('Converged after {} itterations, final residual is {}'.format(itt,err))
# plot the solution Theleft, right and bottom cells contain nothing useful.
fig, ax1 = plt.subplots()
cmap = plt.get_cmap('jet')
cf = ax1.contourf(ptest.x[1:,1:-1],ptest.y[1:,1:-1],ptest.u[1:,1:-1],levels=21, cmap=cmap)
fig.colorbar(cf, ax=ax1)
ax1.set_title(f'Porous media head ({ptest.Ni} x {ptest.Nj} grid)')
plt.show()
# -
# # Further tests
# We can compare this with the analytical solution, and conduct a mesh refinement study. We're going to do the 2nd and need the integrate function again. I'm interested in the integral of head at z=20.0 m from 25.0 to 175.0 m
# +
import scipy.integrate as integrate
from refinement_analysis import refinement_analysis
def integrate_u_dx(mesh,x0,x1,y):
'''Calculate U=\int_{1.0}^{1.95)u(x,0.25) dx using the
u value stored on the grid and simpsons rule'''
# find the left and right grid points
i0,j = mesh.find((x0,y))
i1,j = mesh.find((x1,y))
# add 1 to i1 as we are going to use it as an upper
# bound forarray slicing
i1 = i1 +1
# integrate
return integrate.simps(mesh.u[j,i0:i1],mesh.x[j,i0:i1])
# -
# We then need to set up a sequence of grids based on grid doubling. The number of grid points in each direction should be given by $$N =2^n m + 1$$ where n is the grid index and m is an integer. For our problem we are going to use
# $$N_i=2^n 20 + 1\text{ and }N_j=2^n 10 + 1$$
#
# The following script creates a sequence of grids and solves them all. The upper limit on the `range(5,-1,-1)` is the grid index of the finest grid. Grid 5 takes about 20 minutes to run on my macbook pro. Grid 6, takes a much more reasonable three and a half minutes.
#
# +
import datetime # just seconds may not be enough
# we need some lists u and dx values
U_val = []
dx_val = []
run_time = []
n_pts =[]
for grid_index in range(5,1,-1):
ni = 20*2**grid_index + 1
nj = 10*2**grid_index + 1
n_pts.append(ni*nj)
# set up the problem
test = porous(ni,nj)
print(test)
# solve it with 12 d.p. precision and a lot of itterations
start = time.process_time()
itt, err = SOR(test,tol=0.5e-10,maxit=1000000)
stop = time.process_time()
print("The solver converged after {} itterations, it took {}, final residual is {}" \
.format(itt,datetime.timedelta(seconds=int(stop-start)),err))
# save dx and the integral
dx_val.append(test.Delta_x())
U_val.append(integrate_u_dx(test,75.0,125.0,25.0)/50)
run_time.append(stop-start)
print('Integrated value is ',U_val[-1],'\n')
# -
# lets to the refinement analysis
analysis = refinement_analysis(dx_val,U_val)
analysis.report(r'\int_{75}^{125}h(x,25) dx')
analysis.plot(True,r'$\int_{75}^{125}h(x,25) dx$')
# ### Analysis
#
# The refinement analysis shows
# * _p_ = 1.0, so our 2nd order method has an apparent order of accuracy of 1. Which is very disapointing.
# * The GCI Ratios show that the solution on none of the girds is safely in the asymptotic region where the error is proprtional to Δx<sup>2</sup>. This means need either
# * Our choice of monitor quanity is poor,
# * The SOR solution is not sufficiently converged (i.e. `tol` is too high), or
# * The grids are too coarse.
#
# The We can try running the simulation on a finer grid, but this will need significantly longer to run. Doubling the mesh size will increase the runtime by at least eight (2<sup>4</sup>) times. So we can expect it to run for almost three hours!
#
# We may need a faster solver
|
BoundaryConditions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook compares the numerical evolution with the semianalytical treatment of https://arxiv.org/pdf/1711.09706.pdf
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import UnivariateSpline, interp1d
from scipy.integrate import quad, odeint, solve_ivp
from collections.abc import Sequence
from imripy import halo
from imripy import merger_system as ms
from imripy import inspiral
from imripy import waveform
inspiral.Classic.ln_Lambda=3.
inspiral.Classic.dmPhaseSpaceFraction=1.
# +
def Meff(sp, r=None):
"""
Returns Meff as given by eq (4), default is for r > r_min=r_isco
"""
if r is None:
r = 2.*sp.r_isco()
return np.where(r > sp.r_isco(), sp.m1 - 4.*np.pi*sp.halo.rho_spike*sp.halo.r_spike**3 *sp.r_isco()**(3.-sp.halo.alpha) /(3.-sp.halo.alpha), sp.m1)
def F(sp, r=None):
"""
Returns F as given by eq (5), default is for r > r_min=r_isco
"""
if r is None:
r = 2.*sp.r_isco()
return np.where(r > sp.r_isco(), 4.*np.pi * sp.halo.rho_spike*sp.halo.r_spike**sp.halo.alpha /(3.-sp.halo.alpha), 0.)
def coeffs(sp):
r"""
Calculates the coefficients c_gw, c_df, and \tilde{c} as given by eq (18),(19), and (21)
"""
alpha = sp.halo.alpha
eps = F(sp,2.*sp.r_isco())/Meff(sp)
m2 = sp.m2
if isinstance(m2, (np.ndarray, Sequence)):
m2 = m2[-1]
c_gw = 256./5.* m2 * Meff(sp)**2 * eps**(4./(3.-alpha))
c_df = 8.*np.pi*m2 *sp.halo.rho_spike *sp.halo.r_spike**alpha * 3. \
* Meff(sp)**(-3./2.)* eps**((2.*alpha-3.)/(6.-2.*alpha))
ctild = c_df/c_gw
return c_gw, c_df, ctild
# +
def b_A(sp, x, alpha):
"""
Calculates b_A as given by equation (14), but as a function of x as in eq (15)
"""
eps = F(sp)/Meff(sp)
r = x/eps**(1./(3.-alpha))
omega_s = np.sqrt(Meff(sp, r)/r**3 + F(sp, r)/r**(sp.halo.alpha))
return 4. * r**2 * omega_s**2 / inspiral.Classic.ln_Lambda * (1. + r**2 * omega_s**2)
def f_gw(x, alpha):
"""
Calculates f_gw as given by eq (17) and (20)
"""
return (1.+x**(3.-alpha))**3 / ( 4.*x**3 * ( 1.+ (4.-alpha) *x**(3.-alpha) ) )
def f_df(x, alpha):
"""
Calculates f_dx as given by eq (17) and (20)
"""
return 1. / ( (1.+x**(3.-alpha))**(1./2.) * ( 1.+ (4.-alpha) *x**(3.-alpha) )* x**(-5./2.+alpha) )
def plotDiffEq(sp, r0, r1):
"""
This function plots the differential equation parts of eq(20) and plots them vs the counterpart in the numerical code in between r0 and r1
"""
r = np.geomspace(r0, r1, num=100)
alpha = sp.halo.alpha
eps = F(sp)/Meff(sp)
x = eps**(1./(3.-alpha))*r
c_gw, c_df, ctild = coeffs(sp)
print("c_gw=", c_gw*ms.year_to_pc, "c_df=", c_df*ms.year_to_pc)
l, = plt.loglog(r/sp.r_isco(), np.abs(inspiral.Classic.dE_gw_dt(sp, r))/inspiral.Classic.dE_orbit_da(sp, r), label=r'$dE_{gw}/dt / dE_{orbit}/dR$', alpha=0.5)
plt.loglog(r/sp.r_isco(), c_gw*f_gw(x, alpha)/eps**(1./(3.-alpha)) , label='$c_{gw}f_{gw}$', color=l.get_c(), linestyle='--')
l, = plt.loglog(r/sp.r_isco(), np.abs(inspiral.Classic.dE_df_dt(sp, r))/inspiral.Classic.dE_orbit_da(sp, r), label=r'$dE_{df}/dt / dE_{orbit}/dR$', alpha=0.5)
plt.loglog(r/sp.r_isco(), c_df* f_df(x, alpha)/eps**(1./(3.-alpha)), label='$c_{df}f_{df}$' , color=l.get_c(), linestyle='--')
l, = plt.loglog(r/sp.r_isco(), np.abs(inspiral.Classic.dE_acc_dt(sp, r))/inspiral.Classic.dE_orbit_da(sp, r), label=r'$dE_{acc}/dt / dE_{orbit}/dR$', alpha=0.5)
plt.loglog(r/sp.r_isco(), c_df* f_df(x, alpha)*b_A(sp, x, alpha)/eps**(1./(3.-alpha)), label='$c_{df}f_{df}b_A$' , color=l.get_c(), linestyle='--')
plt.xlabel('$r/r_{ISCO}$')
# +
def J(x, alpha):
"""
Calculates J as in eq (22)
"""
return 4. * x**(11./2. - alpha) / (1. + x**(3.-alpha))**(7./2.)
def K(x, alpha):
"""
Calculates K as given by eq (29), but should coincide with (26f) from https://arxiv.org/pdf/1408.3534.pdf
"""
return (1.+x**(3.-alpha))**(5./2.) * (1. + alpha/3.*x**(3.-alpha)) / (1. + (4.-alpha)*x**(3-alpha) )
def plotPhiprimeprime(sp, r0, r1):
"""
Plots eq (35) and compares it with the counterpart from the numerical simulation
"""
r = np.geomspace(r0, r1, num=100)
alpha = sp.halo.alpha
eps = F(sp)/Meff(sp)
x = eps**(1./(3.-alpha))*r
c_gw, c_df, ctild = coeffs(sp)
Phipp_ana = Meff(sp)**(1./2.) * eps**(3./2./(3.-alpha)) * c_gw*(1.+ctild*J(x, alpha)*(1.+b_A(sp, x, alpha))) *3./4.* K(x,alpha) * x**(-11./2.)
plt.loglog(r/sp.r_isco(), Phipp_ana, label=r'$\ddot{\Phi}^{paper}$' )
#plt.loglog(r/sp.r_isco(), Meff(sp)**(1./2.) * eps**(3./2./(3.-alpha)) \
# * (c_gw*f_gw(x, alpha) + c_df*f_df(x, alpha)) * (3. + alpha*x**(3.-alpha))/(x**(5./2.) * (1.+ x**(3.-alpha))**(1./2.) ), label=r'$\ddot{\Phi}^{paper,ref}$' )
Phipp = (sp.mass(r)/r**3 )**(-1./2.) * (-3.*sp.mass(r)/r**4 + 4.*np.pi *sp.halo.density(r)/r )* inspiral.Classic.da_dt(sp, r)
plt.loglog(r/sp.r_isco(), Phipp, label=r'$\ddot{\Phi}^{code}$', linestyle='--')
plt.loglog(r/sp.r_isco(), np.abs(Phipp - Phipp_ana), label=r'$\Delta \ddot{\Phi}$')
plt.xlabel(r'$r/r_{ISCO}$')
# +
def L(sp, f, accretion=True):
"""
Calculates L as given by eq (48)
If accretion=False, then L' as given by eq (58)
"""
alpha = sp.halo.alpha
#eps = F(sp)/Meff(sp)
#c_gw, c_df, ctild = coeffs(sp)
#c_eps = Meff(sp, 2.*sp.r_isco())**(11./6.-1./3.*alpha) * ctild * eps**((11.-2.*alpha)/(6.-2.*alpha))
c_eps = 5.*np.pi/32. * Meff(sp)**(-(alpha+5.)/3.) * sp.halo.rho_spike * sp.halo.r_spike**(alpha) * inspiral.Classic.ln_Lambda
if accretion:
# TODO: Check prefactor, it's in (36) but not (51)
b_eps = 4.*(np.pi*f * Meff(sp))**(2./3.) / inspiral.Classic.ln_Lambda * (1. + (np.pi*f * Meff(sp))**(2./3.))
else:
b_eps = 0.
deltatild = (1./np.pi**2 / f**2)**(1.-alpha/3.)
return 1. + 4.*c_eps*deltatild**((11.-2.*alpha)/(6.-2.*alpha)) * (1. + b_eps)
def mu(sp, f, f_ini):
"""
Calculates mu as in eq (47) with lower bound f=f_ini
"""
alpha = sp.halo.alpha
prefactor = 16.*np.pi * sp.halo.rho_spike* sp.halo.r_spike**alpha * 5./3./np.pi * (8.*np.pi * Meff(sp)**(2./5))**(-5./3.)
def integrand(y, f):
return (1. + Meff(sp)**(2./3) * (np.pi**2 *f**2)**(1./3.)) / (Meff(sp)/(np.pi**2 * f**2))**((alpha+1)/3.) / np.pi / f * f**(-11./3.) / L(sp, f)
sol = prefactor * odeint(integrand, 0., np.append(f_ini, f), rtol=1e-13, atol=1e-13).flatten()[1:]
return sp.m2 * np.exp(sol)
def getPhaseParameters(sp, ev, f_c = None):
"""
Calculates the terms involved in the derivation of Delta Phi, as the Phi (second part of eq (28b)), 2pi tf (first part of eq(28b), tilde{Phi} (eq(28b)), Delta tilde{Phi} as in eq (56)
for the solution of the numerical evolution
"""
omega_s = sp.omega_s(ev.R)
f_gw = omega_s/np.pi
if f_c is None:
t = ev.t
else:
f_gw = np.unique(np.clip(f_gw, None, f_c))
omega_s = omega_s[:len(f_gw)]
t = ev.t[:len(f_gw)]
if isinstance(sp.m_chirp(), (np.ndarray, Sequence)):
m_chirp = sp.m_chirp()[0]
else:
m_chirp = sp.m_chirp()
omega_gw = interp1d(t, 2*omega_s, kind='cubic', bounds_error=False, fill_value=(0.,0.) )
Phit = np.cumsum([quad(lambda t: omega_gw(t), t[i-1], t[i], limit=200, epsrel=1e-13, epsabs=1e-13)[0] if not i == 0 else 0. for i in range(len(t)) ])
#Phit = odeint(lambda y,t: omega_gw(t), 0., t, atol=1e-13, rtol=1e-13).flatten()
Phi = Phit - Phit[-1]
# Use t_c = t[-1] as reference point
tpt = 2.*np.pi*f_gw * (t - t[-1])
PhiTild = tpt - Phi
return f_gw, Phi, tpt, PhiTild
def plotPhase(sp, ev_acc, ev_nacc, f_c=None, plot_intermediates=False):
"""
Plots the different terms of the derivation of Delta tilde{Phi} semianalytically and compares them to the numerical evolution. Additionally, calculates tilde{Phi}_1 as in eq (57) for both paper and code and compares the delta tilde{Phi} as given by eq (59)
"""
# Code accretion
sp.m2 = ev_acc.m2
f_gw, Phi, tpt, PhiTild = getPhaseParameters(sp, ev_acc, f_c=f_c)
sp.m2 = ev_nacc.m2
f_c = f_gw[-1]
PhiTild0 = (8.*np.pi*sp.m_chirp())**(-5./3.) * (-3./4. * f_gw**(-5./3.) - 5./4. * f_gw * f_c**(-8./3.) + 2.*f_c**(-5./3.))
#PhiTild0 = - 3./4.*(8.*np.pi*sp.m_chirp()*f_gw)**(-5./3.) + 3./4.*(8.*np.pi*sp.m_chirp()*f_c)**(-5./3.)
DeltaPhi = PhiTild - PhiTild0
if plot_intermediates:
plt.plot(f_gw*ms.year_to_pc*3.17e-8, Phi, label=r'$\Phi^{code}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, tpt, label=r'$2\pi t^{code}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, PhiTild, label=r'$\tilde{\Phi}^{code}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, np.abs(DeltaPhi), label=r'$\Delta\tilde{\Phi}^{code}$')
mu_interp = interp1d(f_gw, mu(sp, f_gw, f_gw[0]), kind='cubic', bounds_error=False, fill_value='extrapolate')
#Phi_ana = np.cumsum([quad(lambda f: f**(-8./3.)/L(sp, f)/mu_interp(f), f_gw[i-1], f_gw[i], limit=200, epsrel=1e-13, epsabs=1e-13)[0] if not i == 0 else 0. for i in range(len(f_gw)) ])
Phi_ana = solve_ivp(lambda f,y: f**(-8./3.)/L(sp, f)/mu_interp(f), [f_gw[0], f_gw[-1]], [0.], t_eval=f_gw, atol=1e-13, rtol=1e-13).y[0]
Phi_ana = 10./3. * (8.*np.pi*Meff(sp)**(2./5.))**(-5./3.) * (Phi_ana - Phi_ana[-1])
#tpt_ana = np.cumsum([quad(lambda f: f**(-11./3.)/L(sp, f)/mu_interp(f), f_gw[i-1], f_gw[i], limit=200, epsrel=1e-13, epsabs=1e-13)[0] if not i==0 else 0. for i in range(len(f_gw)) ])
tpt_ana = solve_ivp(lambda f,y: f**(-11./3.)/L(sp, f)/mu_interp(f), [f_gw[0], f_gw[-1]], [0.], t_eval=f_gw, atol=1e-13, rtol=1e-13).y[0]
tpt_ana = 10./3. * (8.*np.pi*Meff(sp)**(2./5.))**(-5./3.) * f_gw * ( tpt_ana - tpt_ana[-1])
PhiTild_ana = tpt_ana - Phi_ana
DeltaPhi_ana = PhiTild_ana - PhiTild0
if plot_intermediates:
plt.plot(f_gw*ms.year_to_pc*3.17e-8, Phi_ana, label=r'$\Phi^{paper}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, tpt_ana, label=r'$2\pi t^{paper}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, PhiTild_ana, label=r'$\tilde{\Phi}^{paper}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, np.abs(DeltaPhi_ana), label=r'$\Delta\tilde{\Phi}^{paper}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, np.abs(DeltaPhi - DeltaPhi_ana), label=r'$\Delta \Delta\tilde{\Phi}$')
# Code no accretion
f_gw_nacc, _, __, PhiTild_nacc = getPhaseParameters(sp, ev_nacc, f_c=f_c)
f_c = f_gw_nacc[-1]
PhiTild0 = (8.*np.pi*sp.m_chirp())**(-5./3.) * (-3./4. * f_gw_nacc**(-5./3.) - 5./4. * f_gw_nacc * f_c**(-8./3.) + 2.*f_c**(-5./3.))
#PhiTild0 = - 3./4.*(8.*np.pi*sp.m_chirp()*f_gw_nacc)**(-5./3.) + 3./4.*(8.*np.pi*sp.m_chirp()*f_c)**(-5./3.)
DeltaPhi_nacc = PhiTild_nacc - PhiTild0
DeltaPhi_naccinterp = interp1d(f_gw_nacc, DeltaPhi_nacc, kind='cubic', bounds_error=False, fill_value=(0.,0.))
deltaPhi = np.abs(DeltaPhi_naccinterp(f_gw) - DeltaPhi)
plt.plot(f_gw*ms.year_to_pc*3.17e-8, deltaPhi, label=r'$\delta\tilde{\Phi}^{code}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, np.abs(deltaPhi/DeltaPhi), label=r'$\delta\tilde{\Phi}^{code}/\Delta\tilde{\Phi}$')
# Paper no accretion
Phi_ana = np.cumsum([quad(lambda f: f**(-8./3.)/L(sp, f, accretion=False), f_gw_nacc[i-1], f_gw_nacc[i], limit=200, epsrel=1e-13, epsabs=1e-13)[0] if not i == 0 else 0. for i in range(len(f_gw_nacc)) ])
#Phi_ana = solve_ivp(lambda f,y: f**(-8./3.)/L(sp, f, accretion=False), [f_gw_nacc[0], f_gw_nacc[-1]], [0.], t_eval=f_gw_nacc, atol=1e-13, rtol=1e-13, method='LSODA').y[0]
Phi_ana = 10./3. * (8.*np.pi*sp.m_chirp())**(-5./3.) * (Phi_ana - Phi_ana[-1])
tpt_ana = np.cumsum([quad(lambda f: f**(-11./3.)/L(sp, f, accretion=False), f_gw_nacc[i-1], f_gw_nacc[i], limit=200, epsrel=1e-13, epsabs=1e-13)[0] if not i==0 else 0. for i in range(len(f_gw_nacc)) ])
#tpt_ana = solve_ivp(lambda f,y: f**(-11./3.)/L(sp, f, accretion=False), [f_gw_nacc[0], f_gw_nacc[-1]], [0.], t_eval=f_gw_nacc, atol=1e-13, rtol=1e-13, method='LSODA').y[0]
tpt_ana = 10./3. * (8.*np.pi*sp.m_chirp())**(-5./3.) * f_gw_nacc * ( tpt_ana - tpt_ana[-1])
PhiTild_nacc_ana = tpt_ana - Phi_ana
PhiTild_nacc_anaInterp = interp1d(f_gw_nacc, PhiTild_nacc_ana, kind='cubic', bounds_error=False, fill_value=(0.,0.))
deltaPhi_ana = np.abs(PhiTild_nacc_anaInterp(f_gw) - PhiTild_ana)
plt.plot(f_gw*ms.year_to_pc*3.17e-8, deltaPhi_ana, label=r'$\delta\tilde{\Phi}^{paper}$')
plt.plot(f_gw*ms.year_to_pc*3.17e-8, np.abs(deltaPhi_ana/DeltaPhi_ana), label=r'$\delta\tilde{\Phi}^{paper}/\Delta\tilde{\Phi}$')
# -
def plotWaveform(sp, ev):
"""
Plots the gravitational waveform of h as given by eq (40) and compares them to the code
"""
f_gw, h, _, Psi = waveform.h_2( sp, ev)
plt.loglog(f_gw*ms.year_to_pc*3.17e-8, h, label=r'$\tilde{h}^{code}$')
alpha = sp.halo.alpha
eps = F(sp,2.*sp.r_isco())/Meff(sp)
A = (5./24.)**(1./2.) * np.pi**(-2./3.) /sp.D * sp.m_chirp()**(5./6.)
plt.loglog(f_gw*ms.year_to_pc*3.17e-8, A*f_gw**(-7./6.) * (L(sp,f_gw))**(-1./2.), label=r'$\tilde{h}^{paper,approx}$')
delta = (Meff(sp)/np.pi**2 / f_gw**2)**(1.-alpha/3.)
chi = 1. + delta*eps/3. + (2.-alpha)/9. *delta**2 * eps**2
x = (delta*eps)**(1./(3.-alpha)) *chi
c_gw, c_df, ctild = coeffs(sp)
plt.loglog(f_gw*ms.year_to_pc*3.17e-8, A*f_gw**(-7./6.) * chi**(19./4.) * (K(x, alpha)* (1. + ctild*J(x, alpha)*(1.+b_A(sp, x, alpha)) ))**(-1./2.), label=r'$\tilde{h}^{paper}$' )
plt.ylabel('h'); plt.xlabel('f')
# ### Define system parameters
m1 = 1e3 *ms.solar_mass_to_pc
m2 = 10. *ms.solar_mass_to_pc
D = 1e3
rho_spike = 226*ms.solar_mass_to_pc
r_spike = 0.54
alpha = 7./3.
sp_1 = ms.SystemProp(m1, m2, halo.Spike( rho_spike, r_spike, alpha), D)
plt.figure()
plotDiffEq(sp_1, sp_1.r_isco(), 1e7*sp_1.r_isco())
plt.legend(); plt.grid()
plt.figure()
plotPhiprimeprime(sp_1, sp_1.r_isco(), 1e5*sp_1.r_isco())
plt.legend(); plt.grid()
R0 = 100.*sp_1.r_isco()
ev_nacc = inspiral.Classic.evolve_circular_binary(sp_1, R0, sp_1.r_isco(), acc=1e-13, accretion=False)
ev_acc = inspiral.Classic.evolve_circular_binary(sp_1, R0, sp_1.r_isco(), acc=1e-13, accretion=True)
plt.figure(figsize=(20,20))
plotPhase(sp_1, ev_acc, ev_nacc, f_c = 0.1*ms.hz_to_invpc, plot_intermediates=False)
plt.legend(); plt.grid()
plt.xlabel('f')
#plt.xscale('log')
#plt.yscale('symlog')
plt.yscale('log')
plt.figure()
plotWaveform(sp_1, ev_acc)
plt.legend(); plt.grid()
# +
plt.figure()
mu_ana = mu(sp_1, sp_1.omega_s(ev_acc.R)/np.pi, sp_1.omega_s(ev_acc.R[0])/np.pi)
plt.loglog(ev_acc.t, mu_ana/m2 -1., label='$\Delta m_2^{paper}/m_2$')
plt.loglog(ev_acc.t, ev_acc.m2/m2 - 1., label="$\Delta m_2^{code}/m_2$", linestyle='--')
plt.loglog(ev_acc.t, np.abs(mu_ana - ev_acc.m2)/m2, label="$\Delta m_2$", linestyle='--')
plt.legend(); plt.grid()
print("mass increase:", ev_acc.m2[-1]/ev_acc.m2[0] -1.)
# -
|
tests/crosschecks1711.09706.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generators Homework
#
# ### Problem 1
#
# Create a generator that generates the squares of numbers up to some number N.
def gensquares(N):
pass
for x in gensquares(10):
print(x)
# ### Problem 2
#
# Create a generator that yields "n" random numbers between a low and high number (that are inputs). <br>Note: Use the random library. For example:
# +
import random
random.randint(1,10)
# -
def rand_num(low,high,n):
pass
for num in rand_num(1,10,12):
print(num)
# ### Problem 3
# Explain a use case for a generator using a yield statement where you would not want to use a normal function with a return statement.<br><br><br><br><br><br>
#
#
# # Great Job!
|
01-Iterators and Generators Homework.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../../pyutils')
import torch
from torch import nn
import torch.nn.functional as F
import numpy as np
import metrics
# -
# # Mean Squared Error (MSE)
#
# The predictions of the model are denoted $\hat{y}$, the true values are denoted $y$.
# This is a 1D version, without average. We can reshape the input and multiply by a constant term to handle average and multi-dimensional data.
#
# # Forward pass
#
# $$l = \sum_{i=1}^n(\hat{y}_i - y_i)^2, \space \hat{y}, y \in \mathbb{R}^n, l \in \mathbb{R}$$
#
# # Backward pass
#
# $$\frac{\partial E}{\partial \hat{y}} = 2 * \frac{\partial E}{\partial l} * (\hat{y} - y)$$
# $$\frac{\partial E}{\partial y} = 2 * \frac{\partial E}{\partial l} * (y - \hat{y})$$
# +
np.random.seed(12)
def mse(pred, target):
return np.sum((pred - target)**2)
def mse_dpred(pred, target, dout):
return 2 * dout * (pred - target)
def mse_dtarget(pred, target, dout):
return 2 * dout * (target - pred)
pred = np.random.randn(26, 4, 8).astype(np.float32)
target = np.random.randn(26, 4, 8).astype(np.float32)
l = mse(pred, target)
loss = np.sum(l**2)
dl = 2*l
dpred = mse_dpred(pred, target, dl)
dtarget = mse_dtarget(pred, target, dl)
tpred = torch.from_numpy(pred).requires_grad_(True)
ttarget = torch.from_numpy(target).requires_grad_(True)
tl = F.mse_loss(tpred, ttarget, reduction='sum')
tloss = torch.sum(tl**2)
tloss.backward()
tdpred = tpred.grad
tdtarget = ttarget.grad
print(metrics.tdist(l, tl.data.numpy()))
print(metrics.tdist(dpred, tdpred.data.numpy()))
print(metrics.tdist(dtarget, tdtarget.data.numpy()))
# -
# # Mean Absolute Error (MAE)
#
# The predictions of the model are denoted $\hat{y}$, the true values are denoted $y$.
# This is a 1D version, without average. We can reshape the input and multiply by a constant term to handle average and multi-dimensional data.
#
# # Forward pass
#
# $$l = \sum_{i=1}^n |\hat{y}_i - y_i|, \space \hat{y}, y \in \mathbb{R}^n, l \in \mathbb{R}$$
#
# # Backward pass
#
# $$\frac{\partial E}{\partial \hat{y}} = \frac{\partial E}{\partial l} * \text{sign}(\hat{y} - y)$$
# $$\frac{\partial E}{\partial y} = \frac{\partial E}{\partial l} * \text{sign}(y - \hat{y})$$
# +
np.random.seed(12)
def mae(pred, target):
return np.sum(np.abs(pred - target))
def mae_dpred(pred, target, dout):
return dout * np.sign(pred - target)
def mae_dtarget(pred, target, dout):
return dout * np.sign(target - pred)
pred = np.random.randn(26, 4, 8).astype(np.float32)
target = np.random.randn(26, 4, 8).astype(np.float32)
l = mae(pred, target)
loss = np.sum(l**2)
dl = 2*l
dpred = mae_dpred(pred, target, dl)
dtarget = mae_dtarget(pred, target, dl)
tpred = torch.from_numpy(pred).requires_grad_(True)
ttarget = torch.from_numpy(target).requires_grad_(True)
tl = F.l1_loss(tpred, ttarget, reduction='sum')
tloss = torch.sum(tl**2)
tloss.backward()
tdpred = tpred.grad
tdtarget = ttarget.grad
print(metrics.tdist(l, tl.data.numpy()))
print(metrics.tdist(dpred, tdpred.data.numpy()))
print(metrics.tdist(dtarget, tdtarget.data.numpy()))
# -
# # Binary Cross Entropy (BCE)
#
# The predictions of the model are denoted $\hat{y}$, they are the probabilities of belonging to class $1$.
# The true values are denoted $y$, they are $1$ if belong to class $1$, or $0$.
#
#
# # Forward pass
#
# $$l = - \sum_{i=1}^N (y_i log(\hat{y_i}) + (1 - y_i) log(1 - \hat{y_i})), \space \hat{y}, y \in \mathbb{R}^N, l \in \mathbb{R}$$
#
# # Backward pass
#
# $$\frac{\partial E}{\partial \hat{y}} = \frac{\partial E}{\partial l} * \frac{\hat{y} - y}{\hat{y}(1 - \hat{y})}$$
# +
np.random.seed(12)
def bce(pred, target):
return - np.sum(target*np.log(pred)+(1-target) * np.log(1-pred))
def bce_dpred(pred, target, dout):
return dout * (pred - target) / (pred * (1 - pred))
pred = np.random.rand(26).astype(np.float32)
target = np.random.randint(0, 2, size=26).astype(np.float32)
l = bce(pred, target)
loss = np.sum(l**2)
dl = 2*l
dpred = bce_dpred(pred, target, dl)
tpred = torch.from_numpy(pred).requires_grad_(True)
ttarget = torch.from_numpy(target)
tl = F.binary_cross_entropy(tpred, ttarget, reduction='sum')
tloss = torch.sum(tl**2)
tloss.backward()
tdpred = tpred.grad
print(metrics.tdist(l, tl.data.numpy()))
print(metrics.tdist(dpred, tdpred.data.numpy()))
# -
# # Cross Entropy (CE)
#
# The predictions of the model are denoted $\hat{y}$, they are the probabilities of belonging to each of the K classes.
# The true values are denoted $y$, a one-hot representation with $1$ for the true class.
#
#
# # Forward pass
#
# $$l = - \sum_{i=1}^N \sum_{j=1}^K [y_{ij} log(\hat{y_{ij}})], \space \hat{y}, y \in \mathbb{R}^{N*K}, l \in \mathbb{R}$$
#
# # Backward pass
#
# $$\frac{\partial E}{\partial \hat{y}} = - \frac{\partial E}{\partial l} * \frac{y}{\hat{y}}$$
# +
np.random.seed(12)
def target2onehot(x, nclasses):
oh = np.zeros((len(x), nclasses)).astype(np.float32)
oh[np.arange(len(x)), x] = 1
return oh
def cross_entropy(pred, target):
return - np.sum(target * np.log(pred))
def cross_entropy_dpred(pred, target, dout):
return - dout * target / pred
pred = np.random.rand(26, 4).astype(np.float32)
pred = pred / np.sum(pred, axis=1, keepdims=True)
target = np.random.randint(0, 4, size=26).astype(np.int)
target = target2onehot(target, 4)
l = cross_entropy(pred, target)
loss = np.sum(l**2)
dl = 2*l
dpred = cross_entropy_dpred(pred, target, dl)
tpred = torch.from_numpy(pred).requires_grad_(True)
ttarget = torch.from_numpy(target)
tl = - torch.sum(ttarget * torch.log(tpred))
tloss = torch.sum(tl**2)
tloss.backward()
tdpred = tpred.grad
print(metrics.tdist(l, tl.data.numpy()))
print(metrics.tdist(dpred, tdpred.data.numpy()))
|
courses/deep_learning/op_loss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import psi4
import numpy as np
import sys
sys.path.append('../lib')
from P4toC4_aux import *
# +
# C4: STO-3G -74.963 319 056 386 7
psi4.set_memory('500 MB')
psi4.core.set_global_option("BASIS", "6-31G*")
psi4.core.set_global_option("SCF_TYPE", "pk")
psi4.core.set_global_option("PUREAM", "True")
psi4.core.set_output_file('output.dat', False)
# O
# H 1 0.96
# H 1 0.96 2 104.5
h2o = psi4.geometry("""
0 1
O 0.000000000000 0.000000000000 -0.065775570547
H 0.000000000000 -0.759061990794 0.521953018286
H 0.000000000000 0.759061990794 0.521953018286
symmetry c1
""")
E, wf = psi4.energy('scf', return_wfn=True)
E
# -
basisset=wf.basisset()
mol=wf.molecule()
for atom in range(mol.natom()):
print(mol.symbol(atom))
basisset.nbf()
# +
# stolen from molden writer v1.5 and modified
mol_string = '\n[GTO]\n'
for atom in range(mol.natom()):
mol_string += f" {atom+1:d} 0\n"
#i = 0
for rel_shell_idx in range(basisset.nshell_on_center(atom)):
#abs_shell_idx = basisset.shell_on_center(atom, rel_shell_idx)
#shell = basisset.shell(abs_shell_idx)
shell = basisset.shell(atom, rel_shell_idx)
mol_string += f" {shell.amchar:s}{shell.nprimitive:5d} 1.00\n"
for prim in range(shell.nprimitive):
mol_string += f"{shell.exp(prim):20.10f} {shell.original_coef(prim):20.10f}\n"
mol_string += '\n'
print(mol_string)
# -
ao=0
for atom in range(mol.natom()):
print(f"Atom {atom}")
for rel_shell_idx in range(basisset.nshell_on_center(atom)):
#abs_shell_idx = basisset.shell_on_center(atom, rel_shell_idx)
#shell = basisset.shell(abs_shell_idx)
shell = basisset.shell(atom, rel_shell_idx)
print(f" {rel_shell_idx} {shell.amchar}")
for ishell in range(basisset.nshell()):
shell = basisset.shell(ishell)
print(f"Shell {ishell} {shell.amchar} {shell.am}")
offset = ao_offset(basisset, verbose=0)
offset
# +
# from write_nbo:
pure_order = [
[1], # s
[103, 101, 102], # p
[255, 252, 253, 254, 251], # d: z2 xz yz x2-y2 xy
[351, 352, 353, 354, 355, 356, 357], # f
[451, 452, 453, 454, 455, 456, 457, 458, 459], #g
[551, 552, 553, 554, 555, 556, 557, 558, 559, 560, 561] #h
]
# from write molden
'''
Reordering expected by Molden
P: x, y, z
5D: D 0, D+1, D-1, D+2, D-2
6D: xx, yy, zz, xy, xz, yz
7F: F 0, F+1, F-1, F+2, F-2, F+3, F-3
10F: xxx, yyy, zzz, xyy, xxy, xxz, xzz, yzz, yyz, xyz
'''
molden_cartesian_order = [
[2,0,1,0,0,0,0,0,0,0,0,0,0,0,0], # p
[0,3,4,1,5,2,0,0,0,0,0,0,0,0,0], # d
[0,4,5,3,9,6,1,8,7,2,0,0,0,0,0], # f
[0,3,4,9,12,10,5,13,14,7,1,6,11,8,2] # g
]
# -
|
dev/Bas_print.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install Augmentor
import Augmentor
p = Augmentor.Pipeline("./red-pandas-curated")
p.flip_left_right(probability=0.5)
p.rotate(probability=0.7, max_left_rotation=15, max_right_rotation=15)
p.zoom(probability=0.5, min_factor=1.1, max_factor=1.6)
p.skew_tilt(probability=0.4,magnitude =0.3)
# p.crop_random(probability=1, percentage_area=0.5)
p.sample(10000)
# p.sample(2500, multi_threaded=False)
# p.process()
|
augment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
##Loading the necessary packages
import numpy as np
import cv2
from imutils.object_detection import non_max_suppression
import pytesseract
from matplotlib import pyplot as plt
#Creating argument dictionary for the default arguments needed in the code.
args = {"image":r"C:\Users\jagadesh\Jupyter Projects\OCR/test1.jpeg", "east":r"C:\Users\jagadesh\Jupyter Projects\OCR\kaggle\east_text\east_text_detection.pb","min_confidence":0.5, "width":320, "height":320}
# +
#Give location of the image to be read.
#"Example-images/ex24.jpg" image is being loaded here.
args['image']=r"C:\Users\jagadesh\Jupyter Projects\OCR/test1.jpeg"
image = cv2.imread(args['image'])
#Saving a original image and shape
orig = image.copy()
(origH, origW) = image.shape[:2]
# set the new height and width to default 320 by using args #dictionary.
(newW, newH) = (args["width"], args["height"])
#Calculate the ratio between original and new image for both height and weight.
#This ratio will be used to translate bounding box location on the original image.
rW = origW / float(newW)
rH = origH / float(newH)
# resize the original image to new dimensions
image = cv2.resize(image, (newW, newH))
(H, W) = image.shape[:2]
# construct a blob from the image to forward pass it to EAST model
blob = cv2.dnn.blobFromImage(image, 1.0, (W, H),
(123.68, 116.78, 103.94), swapRB=True, crop=False)
# +
# load the pre-trained EAST model for text detection
net = cv2.dnn.readNet(args["east"])
# We would like to get two outputs from the EAST model.
#1. Probabilty scores for the region whether that contains text or not.
#2. Geometry of the text -- Coordinates of the bounding box detecting a text
# The following two layer need to pulled from EAST model for achieving this.
layerNames = [
"feature_fusion/Conv_7/Sigmoid",
"feature_fusion/concat_3"]
# -
#Forward pass the blob from the image to get the desired output layers
net.setInput(blob)
(scores, geometry) = net.forward(layerNames)
# +
# Returns a bounding box and probability score if it is more than minimum confidence
def predictions(prob_score, geo):
(numR, numC) = prob_score.shape[2:4]
boxes = []
confidence_val = []
# loop over rows
for y in range(0, numR):
scoresData = prob_score[0, 0, y]
x0 = geo[0, 0, y]
x1 = geo[0, 1, y]
x2 = geo[0, 2, y]
x3 = geo[0, 3, y]
anglesData = geo[0, 4, y]
# loop over the number of columns
for i in range(0, numC):
if scoresData[i] < args["min_confidence"]:
continue
(offX, offY) = (i * 4.0, y * 4.0)
# extracting the rotation angle for the prediction and computing the sine and cosine
angle = anglesData[i]
cos = np.cos(angle)
sin = np.sin(angle)
# using the geo volume to get the dimensions of the bounding box
h = x0[i] + x2[i]
w = x1[i] + x3[i]
# compute start and end for the text pred bbox
endX = int(offX + (cos * x1[i]) + (sin * x2[i]))
endY = int(offY - (sin * x1[i]) + (cos * x2[i]))
startX = int(endX - w)
startY = int(endY - h)
boxes.append((startX, startY, endX, endY))
confidence_val.append(scoresData[i])
# return bounding boxes and associated confidence_val
return (boxes, confidence_val)
# -
# Find predictions and apply non-maxima suppression
(boxes, confidence_val) = predictions(scores, geometry)
boxes = non_max_suppression(np.array(boxes), probs=confidence_val)
# +
##Text Detection and Recognition
# initialize the list of results
results = []
# loop over the bounding boxes to find the coordinate of bounding boxes
for (startX, startY, endX, endY) in boxes:
# scale the coordinates based on the respective ratios in order to reflect bounding box on the original image
startX = int(startX * rW)
startY = int(startY * rH)
endX = int(endX * rW)
endY = int(endY * rH)
#extract the region of interest
r = orig[startY:endY, startX:endX]
#configuration setting to convert image to string.
configuration = ("-l eng --oem 1 --psm 8")
##This will recognize the text from the image of bounding box
text = pytesseract.image_to_string(r, config=configuration)
# append bbox coordinate and associated text to the list of results
results.append(((startX, startY, endX, endY), text))
# +
#Display the image with bounding box and recognized text
orig_image = orig.copy()
# Moving over the results and display on the image
for ((start_X, start_Y, end_X, end_Y), text) in results:
# display the text detected by Tesseract
print("{}\n".format(text))
# Displaying text
text = "".join([x if ord(x) < 128 else "" for x in text]).strip()
cv2.rectangle(orig_image, (start_X, start_Y), (end_X, end_Y),
(0, 0, 255), 2)
cv2.putText(orig_image, text, (start_X, start_Y - 30),
cv2.FONT_HERSHEY_SIMPLEX, 0.7,(0,0, 255), 2)
plt.imshow(orig_image)
plt.title('Output')
plt.show()
# -
|
kaggle/ocr try1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learn Scientific Computing with Python
#
# The textbook used is '**Numerical Python**'.
#
# This notebook is the first one. Just for fun. Here we go.
# +
def fib(n):
"""
Return a list of the first n Fibonacci numbers.
"""
f0, f1 = 0, 1
f = [1]*n
for i in range(1,n):
f[i] = f0 + f1
f0, f1 = f1, f[i]
return f
print(fib(10))
# -
|
Lec000_NumericalPython.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
data = pd.read_csv("lyft.csv")
data.head()
data['date_time'] = data['date'] + ' ' + data['time']
data['date_time'] = pd.to_datetime(data['date_time'])
data = data.set_index(pd.DatetimeIndex(data['date_time']))
data.index
data=data['2018-03-01 00:00:00':'2018-03-31 23:59:59']
data.index
data.columns
# Lyft Line is same type as Pool of Uber
data_lyft_line = data[data.display_name == 'Lyft Line']
data_lyft_line.drop(['can_request_ride','currency', 'display_name',
'end_latitude','end_longitude','is_valid_estimate','primetime_percentage','ride_type',
'start_latitude','start_longitude'], axis=1, inplace=True)
data_lyft_line.columns
# Since Uber uses dollar to estimate and Lyft uses cents, we need to change cents to dollars in Lyft
data_lyft_line['lyft_max_estimate'] = (data_lyft_line['estimated_cost_cents_max']).divide(100)
data_lyft_line['lyft_min_estimate'] = (data_lyft_line['estimated_cost_cents_min']).divide(100)
data_lyft_line.drop(['estimated_cost_cents_max'], axis=1, inplace=True)
data_lyft_line.drop(['estimated_cost_cents_min'], axis=1, inplace=True)
data_lyft_line['estimate'] = (data_lyft_line['lyft_max_estimate'] + data_lyft_line['lyft_min_estimate'])/2
# Also, make the column names shorter
data_lyft_line.rename(columns={"estimate": "lyft_estimate"},inplace=True)
data_lyft_line.rename(columns={"estimated_distance_miles": "lyft_distance"},inplace=True)
data_lyft_line.rename(columns={"estimated_duration_seconds": "lyft_duration"},inplace=True)
data_lyft_line['lyft_price_per_second'] = data_lyft_line['lyft_estimate']/data_lyft_line['lyft_duration']
data_lyft_line.head()
plt.rcParams["figure.figsize"] = (20, 10)
plt.plot(data_lyft_line.index, data_lyft_line['lyft_price_per_second'])
plt.title('Lyft Price Estimate')
plt.ylabel('Price ($)')
plt.show()
data_lyft_line.drop(['date_time'], axis=1, inplace=True)
data_lyft_line.to_csv('lyft_line_March.csv', sep=',')
|
Data Clean Part/DataClean_OneMonth_Lyft_Line.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''especializacao'': conda)'
# language: python
# name: python37964bitespecializacaocondabf03fad2cbc44dc4af89bd01e774ce56
# ---
# + tags=[]
# ****************************************************************
# AULA: Visão Computacional
# Prof: <NAME>, DSc.
# ****************************************************************
import sys
sys.path.append('../')
import argparse
import logging
import util
from tqdm import tqdm
import sys
import datetime
# + tags=[]
# Variaveis
diretorio_base = "../../dataset/dogs_cats/"
tipo = "png"
arquivos = util.obtemTodosOsArquivos(diretorio_base,tipo, True)
dataset = "./dataset_sem_classes.csv"
# Inicia log
logging.basicConfig(level=logging.DEBUG)
print(len(arquivos))
# + tags=[]
if len(arquivos) > 0:
# Obtem informacoes para avaliar o tempo de processamento
inicio = datetime.datetime.now()
logging.info("Iniciando processo em: {}".format(inicio))
try:
with open (dataset, "w") as arquivo_de_dados:
for index, arquivo in tqdm(enumerate(arquivos), total=len(arquivos)): # Cria barra de progressao
arquivo_de_dados.write("{}\n".format(arquivo))
except:
logging.error("Erro ao processar criacao de arquivo de dados.")
sys.exit()
# Fim de processo
fim = datetime.datetime.now()
tempo_de_processamento = (fim-inicio).total_seconds()
logging.info("Tempo de processamento: {}".format(str(tempo_de_processamento)))
|
modulo1/1-scripts-estruturais/notebooks/1-geracao-de-lista-de-base-de-dados-sem-classes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="vSHg24UVDI3I" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 4*
#
# ---
#
# # Logistic Regression
# - do train/validate/test split
# - begin with baselines for classification
# - express and explain the intuition and interpretation of Logistic Regression
# - use sklearn.linear_model.LogisticRegression to fit and interpret Logistic Regression models
#
# Logistic regression is the baseline for classification models, as well as a handy way to predict probabilities (since those too live in the unit interval). While relatively simple, it is also the foundation for more sophisticated classification techniques such as neural networks (many of which can effectively be thought of as networks of logistic models).
# + [markdown] id="jrM_FGFtDI3O" colab_type="text"
# ### Setup
#
# Run the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.
#
# Libraries:
# - category_encoders
# - numpy
# - pandas
# - scikit-learn
# + id="AyU1dzRrDI3Q" colab_type="code" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + [markdown] id="807DRaEtDI3a" colab_type="text"
# # Do train/validate/test split
# + [markdown] id="j1pbQqT7DI3d" colab_type="text"
# ## Overview
# + [markdown] id="QLVkebRcDI3f" colab_type="text"
# ### Predict Titanic survival 🚢
#
# Kaggle is a platform for machine learning competitions. [Kaggle has used the Titanic dataset](https://www.kaggle.com/c/titanic/data) for their most popular "getting started" competition.
#
# Kaggle splits the data into train and test sets for participants. Let's load both:
# + id="J1Rl2bMcDI3h" colab_type="code" colab={}
import pandas as pd
train = pd.read_csv(DATA_PATH+'titanic/train.csv')
test = pd.read_csv(DATA_PATH+'titanic/test.csv')
# + [markdown] id="GWTuT8T4DI3l" colab_type="text"
# Notice that the train set has one more column than the test set:
# + id="E_I-kd66DI3m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="a194b605-e2ab-433f-9bd1-c193e00e6707"
train.shape, test.shape
# + [markdown] id="6N-z28xYDI3q" colab_type="text"
# Which column is in train but not test? The target!
# + id="cIxXLcFvDI3r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="f9c91c74-fdfa-470e-e67a-5bed60e1bd23"
set(train.columns) - set(test.columns)
# + [markdown] id="CwxRnMDvDI3u" colab_type="text"
# ### Why doesn't Kaggle give you the target for the test set?
#
# #### <NAME>, [How (and why) to create a good validation set](https://www.fast.ai/2017/11/13/validation-sets/)
#
# > One great thing about Kaggle competitions is that they force you to think about validation sets more rigorously (in order to do well). For those who are new to Kaggle, it is a platform that hosts machine learning competitions. Kaggle typically breaks the data into two sets you can download:
# >
# > 1. a **training set**, which includes the _independent variables,_ as well as the _dependent variable_ (what you are trying to predict).
# >
# > 2. a **test set**, which just has the _independent variables._ You will make predictions for the test set, which you can submit to Kaggle and get back a score of how well you did.
# >
# > This is the basic idea needed to get started with machine learning, but to do well, there is a bit more complexity to understand. **You will want to create your own training and validation sets (by splitting the Kaggle “training” data). You will just use your smaller training set (a subset of Kaggle’s training data) for building your model, and you can evaluate it on your validation set (also a subset of Kaggle’s training data) before you submit to Kaggle.**
# >
# > The most important reason for this is that Kaggle has split the test data into two sets: for the public and private leaderboards. The score you see on the public leaderboard is just for a subset of your predictions (and you don’t know which subset!). How your predictions fare on the private leaderboard won’t be revealed until the end of the competition. The reason this is important is that you could end up overfitting to the public leaderboard and you wouldn’t realize it until the very end when you did poorly on the private leaderboard. Using a good validation set can prevent this. You can check if your validation set is any good by seeing if your model has similar scores on it to compared with on the Kaggle test set. ...
# >
# > Understanding these distinctions is not just useful for Kaggle. In any predictive machine learning project, you want your model to be able to perform well on new data.
# + [markdown] id="BV38V-ZRDI3v" colab_type="text"
# ### 2-way train/test split is not enough
#
# #### Hastie, Tibshirani, and Friedman, [The Elements of Statistical Learning](http://statweb.stanford.edu/~tibs/ElemStatLearn/), Chapter 7: Model Assessment and Selection
#
# > If we are in a data-rich situation, the best approach is to randomly divide the dataset into three parts: a training set, a validation set, and a test set. The training set is used to fit the models; the validation set is used to estimate prediction error for model selection; the test set is used for assessment of the generalization error of the final chosen model. Ideally, the test set should be kept in a "vault," and be brought out only at the end of the data analysis. Suppose instead that we use the test-set repeatedly, choosing the model with the smallest test-set error. Then the test set error of the final chosen model will underestimate the true test error, sometimes substantially.
#
# #### <NAME> and <NAME>, [Introduction to Machine Learning with Python](https://books.google.com/books?id=1-4lDQAAQBAJ&pg=PA270)
#
# > The distinction between the training set, validation set, and test set is fundamentally important to applying machine learning methods in practice. Any choices made based on the test set accuracy "leak" information from the test set into the model. Therefore, it is important to keep a separate test set, which is only used for the final evaluation. It is good practice to do all exploratory analysis and model selection using the combination of a training and a validation set, and reserve the test set for a final evaluation - this is even true for exploratory visualization. Strictly speaking, evaluating more than one model on the test set and choosing the better of the two will result in an overly optimistic estimate of how accurate the model is.
#
# #### <NAME>, [R for Data Science](https://r4ds.had.co.nz/model-intro.html#hypothesis-generation-vs.hypothesis-confirmation)
#
# > There is a pair of ideas that you must understand in order to do inference correctly:
# >
# > 1. Each observation can either be used for exploration or confirmation, not both.
# >
# > 2. You can use an observation as many times as you like for exploration, but you can only use it once for confirmation. As soon as you use an observation twice, you’ve switched from confirmation to exploration.
# >
# > This is necessary because to confirm a hypothesis you must use data independent of the data that you used to generate the hypothesis. Otherwise you will be over optimistic. There is absolutely nothing wrong with exploration, but you should never sell an exploratory analysis as a confirmatory analysis because it is fundamentally misleading.
# >
# > If you are serious about doing an confirmatory analysis, one approach is to split your data into three pieces before you begin the analysis.
#
#
# #### <NAME>, [Model Evaluation](https://sebastianraschka.com/blog/2018/model-evaluation-selection-part4.html)
#
# > Since “a picture is worth a thousand words,” I want to conclude with a figure (shown below) that summarizes my personal recommendations ...
#
# <img src="https://sebastianraschka.com/images/blog/2018/model-evaluation-selection-part4/model-eval-conclusions.jpg" width="600">
#
# Usually, we want to do **"Model selection (hyperparameter optimization) _and_ performance estimation."** (The green box in the diagram.)
#
# Therefore, we usually do **"3-way holdout method (train/validation/test split)"** or **"cross-validation with independent test set."**
# + [markdown] id="i-F_bY2_DI3v" colab_type="text"
# ### What's the difference between Training, Validation, and Testing sets?
#
# #### <NAME>, [Training, Validation, and Testing Data Sets](https://end-to-end-machine-learning.teachable.com/blog/146320/training-validation-testing-data-sets)
#
# > The validation set is for adjusting a model's hyperparameters. The testing data set is the ultimate judge of model performance.
# >
# > Testing data is what you hold out until very last. You only run your model on it once. You don’t make any changes or adjustments to your model after that. ...
# + [markdown] id="ctBObZLuDI3w" colab_type="text"
# ## Follow Along
#
# > You will want to create your own training and validation sets (by splitting the Kaggle “training” data).
#
# Do this, using the [sklearn.model_selection.train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function:
# + id="SAmBUicBXk4k" colab_type="code" colab={}
random_state=42
# + id="T_vCuyjsDI3x" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, random_state=random_state)
# + id="CXl6sB8dYplN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="53a31fc5-6bdd-4e8e-801e-2ad0fe943f9e"
train.shape, val.shape, test.shape
# + id="ARb-RMTzYSj2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 695} outputId="9dfbab45-0db5-4a88-f6d5-8a28b7e4b73b"
train
# + id="ZFviaGmBYTR8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 827} outputId="4c127011-a9ae-482f-e6a7-9a1026e0e117"
val
# + id="YTgsthp_YXUs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 794} outputId="d7d5a6b1-89a7-4df0-aebd-8493a20a4621"
test
# + [markdown] id="k6rDmwwGDI30" colab_type="text"
# ## Challenge
# + [markdown] id="qr7KOeRDDI30" colab_type="text"
# For your assignment, you'll do a 3-way train/validate/test split.
#
# Then next sprint, you'll begin to participate in a private Kaggle challenge, just for your cohort!
#
# You will be provided with data split into 2 sets: training and test. You will create your own training and validation sets, by splitting the Kaggle "training" data, so you'll end up with 3 sets total.
# + [markdown] id="78MsUJFgDI31" colab_type="text"
# # Begin with baselines for classification
# + [markdown] id="9t7lMnGtDI32" colab_type="text"
# ## Overview
# + [markdown] id="eWANg1w6DI32" colab_type="text"
# We'll begin with the **majority class baseline.**
#
# [<NAME>](https://twitter.com/koehrsen_will/status/1088863527778111488)
#
# > A baseline for classification can be the most common class in the training dataset.
#
# [*Data Science for Business*](https://books.google.com/books?id=4ZctAAAAQBAJ&pg=PT276), Chapter 7.3: Evaluation, Baseline Performance, and Implications for Investments in Data
#
# > For classification tasks, one good baseline is the _majority classifier,_ a naive classifier that always chooses the majority class of the training dataset (see Note: Base rate in Holdout Data and Fitting Graphs). This may seem like advice so obvious it can be passed over quickly, but it is worth spending an extra moment here. There are many cases where smart, analytical people have been tripped up in skipping over this basic comparison. For example, an analyst may see a classification accuracy of 94% from her classifier and conclude that it is doing fairly well—when in fact only 6% of the instances are positive. So, the simple majority prediction classifier also would have an accuracy of 94%.
# + [markdown] id="ErEnqbcGDI33" colab_type="text"
# ## Follow Along
# + [markdown] id="hz31RqyeDI34" colab_type="text"
# Determine majority class
# + id="LOqM5EIZDI34" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 65} outputId="bb0f49bc-74f9-4048-e299-abf9e6577135"
target = 'Survived'
y_train = train[target]
y_train.value_counts(normalize=True)
# + [markdown] id="w3iw1c90DI37" colab_type="text"
# What if we guessed the majority class for every prediction?
# + id="iWqmYjjHDI38" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="f18d5b87-2172-4160-d14e-ac4b7dee702c"
y_train.mode()[0]
# + id="bJqblmGgZwbU" colab_type="code" colab={}
majority_class = y_train.mode()[0]
y_pred = [majority_class] * len(y_train)
# + [markdown] id="pbYCn200DI3-" colab_type="text"
# #### Use a classification metric: accuracy
#
# [Classification metrics are different from regression metrics!](https://scikit-learn.org/stable/modules/model_evaluation.html)
# - Don't use _regression_ metrics to evaluate _classification_ tasks.
# - Don't use _classification_ metrics to evaluate _regression_ tasks.
#
# [Accuracy](https://scikit-learn.org/stable/modules/model_evaluation.html#accuracy-score) is a common metric for classification. Accuracy is the ["proportion of correct classifications"](https://en.wikipedia.org/wiki/Confusion_matrix): the number of correct predictions divided by the total number of predictions.
# + [markdown] id="4qmc5IL0DI3-" colab_type="text"
# What is the baseline accuracy if we guessed the majority class for every prediction?
# + id="c5abyInfDI3_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="709d8b13-4c1d-4085-87bb-3736edb562c5"
from sklearn.metrics import accuracy_score
# Training accuracy of majority class baseline -
# frequency of majority class (aka base rate)
accuracy_score(y_train, y_pred)
# + id="dM9sdEYUDI4B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="3193cf66-56ea-4af2-ad75-d5ff7228819f"
# Validation accuracy of majority class baseline =
# usually similar to Train accuracy, wont be equal
y_val = val[target]
y_pred = [majority_class] * len(y_val)
accuracy_score(y_val, y_pred)
# + [markdown] id="mBbAXCnDDI4D" colab_type="text"
# ## Challenge
# + [markdown] id="F3Q4BDifDI4E" colab_type="text"
# In your assignment, your Sprint Challenge, and your upcoming Kaggle challenge, you'll begin with the majority class baseline. How quickly can you beat this baseline?
# + [markdown] id="mwofokJLDI4E" colab_type="text"
# # Express and explain the intuition and interpretation of Logistic Regression
#
# + [markdown] id="O_ozeChuDI4F" colab_type="text"
# ## Overview
#
# To help us get an intuition for *Logistic* Regression, let's start by trying *Linear* Regression instead, and see what happens...
# + [markdown] id="h5-J2y5aDI4F" colab_type="text"
# ## Follow Along
# + [markdown] id="sYGM00cJDI4H" colab_type="text"
# ### Linear Regression?
# + id="xF8oKrupDI4H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="b42330ab-2ac9-4704-95a3-81ce3653f1ca"
train.describe()
# + id="Lc3DYhlfDI4K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 777} outputId="5f6fc66c-3ae0-4b82-b108-0d51ad0777d4"
# 1. Import estimator class
from sklearn.linear_model import LinearRegression
# 2. Instantiate this class
linear_reg = LinearRegression()
# 3. Arrange X feature matrices (already did y target vectors)
features = ['Pclass', 'Age', 'Fare']
X_train = train[features]
X_val = val[features]
# Impute missing values
from sklearn.impute import SimpleImputer
imputer = SimpleImputer()
X_train_imputed = imputer.fit_transform(X_train)
X_val_imputed = imputer.transform(X_val)
# 4. Fit the model
linear_reg.fit(X_train_imputed, y_train)
# 5. Apply the model to new data.
# The predictions look like this ...
linear_reg.predict(X_val_imputed)
# + id="fY0VVuk6DI4M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 82} outputId="58212bab-22fd-4eb1-9b08-d4ebf96805e2"
# Get coefficients
pd.Series(linear_reg.coef_, features)
# + id="cFulFojADI4O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="93e13e3a-8593-4db8-fd53-86b6490c1888"
test_case = [[1, 5, 500]] # 1st class, 5-year old, Rich
linear_reg.predict(test_case)
# + [markdown] id="43dDu52-DI4Q" colab_type="text"
# ### Logistic Regression!
# + id="3X5NjhVaDI4R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="ad66b9f7-ac11-4d04-8c0b-5046acf1d02d"
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression(solver='lbfgs')
log_reg.fit(X_train_imputed, y_train)
print('Validation Accuracy', log_reg.score(X_val_imputed, y_val))
# + id="lxmXDB-iDI4T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="56929bc3-76a8-4a1b-a504-1bd186bf9e02"
# The predictions look like this
log_reg.predict(X_val_imputed)
# + id="bexaZV1nDI4V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="e4549422-e3a2-4171-ce97-f3fd1d2343ff"
log_reg.predict(test_case)
# + id="ImzmMOamDI4X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="a5eefb64-06e9-4c3b-e7c3-5b28394f3c74"
log_reg.predict_proba(test_case)
# + id="wK8lGZAvDI4Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="66cc8de3-e997-4e79-f8ea-a2cb0f756e3d"
# What's the math?
log_reg.coef_
# + id="pC0TrfR4gcVW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 82} outputId="d43fd7f6-d276-43a0-e1c9-973c06cd1ea3"
pd.Series(log_reg.coef_[0], features)
# + id="1qdcn-MVDI4a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="1e8aaebc-4076-49df-fcab-e069560e4ebc"
log_reg.intercept_
# + id="G2rY1l1gDI4c" colab_type="code" colab={}
# The logistic sigmoid "squishing" function, implemented to accept numpy arrays
import numpy as np
def sigmoid(x):
return 1 / (1 + np.e**(-x))
# + id="Cmui8JxUDI4e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="5e94c85d-e6fa-41c1-a308-fa45cff70a86"
sigmoid(log_reg.intercept_ + np.dot(log_reg.coef_, np.transpose(test_case)))
# + id="HIlFFwSGipZw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="1838e61e-7448-4f0f-fc2a-003f9b0c28a8"
x=0
np.e ** (-x)
# + id="pKt5fjUbi0CW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="b6a736f8-084d-462f-baea-e6510da0eeed"
np.e ** -np.inf
# + [markdown] id="zjQ-3NTeDI4f" colab_type="text"
# So, clearly a more appropriate model in this situation! For more on the math, [see this Wikipedia example](https://en.wikipedia.org/wiki/Logistic_regression#Probability_of_passing_an_exam_versus_hours_of_study).
# + [markdown] id="gvJPe73QDI4g" colab_type="text"
# # Use sklearn.linear_model.LogisticRegression to fit and interpret Logistic Regression models
# + [markdown] id="2PtQAk9vDI4g" colab_type="text"
# ## Overview
#
# Now that we have more intuition and interpretation of Logistic Regression, let's use it within a realistic, complete scikit-learn workflow, with more features and transformations.
# + [markdown] id="8mRz9nMMDI4h" colab_type="text"
# ## Follow Along
#
# Select these features: `['Pclass', 'Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked']`
#
# (Why shouldn't we include the `Name` or `Ticket` features? What would happen here?)
#
# Fit this sequence of transformers & estimator:
#
# - [category_encoders.one_hot.OneHotEncoder](https://contrib.scikit-learn.org/categorical-encoding/onehot.html)
# - [sklearn.impute.SimpleImputer](https://scikit-learn.org/stable/modules/generated/sklearn.impute.SimpleImputer.html)
# - [sklearn.preprocessing.StandardScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
# - [sklearn.linear_model.LogisticRegressionCV](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegressionCV.html)
#
# Get validation accuracy.
# + id="2rlhFbMOnzPr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 98} outputId="5953aa4b-8e1b-4adc-cd31-a71fc48e40d8"
train['Name'].describe()
# if a categorical feature always varies, it is useless for prediction
# + id="mnUXNX1Wn3Mz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 98} outputId="a0d62851-cf85-4f31-a51b-e539940910ff"
train['Ticket'].describe()
# mostly unique, not very usefull for predictions
# + id="favgeHbNnddL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="4b2b7009-b7b7-43b2-b661-293d9909d983"
features = ['Pclass', 'Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked']
target = 'Survived'
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_train.shape, y_train.shape, X_val.shape, y_val.shape
# + id="jHXrXDlroNLb" colab_type="code" colab={}
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegressionCV
# + id="r4hlExr2phHY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="092bdb9f-131b-405a-c818-c74a32860b0f"
X_train.head()
# + id="UVILPvCopi-z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="d75a6e78-41fb-446f-a1d2-bb9227dc3908"
X_val.head()
# + id="0xIWvL4epPVE" colab_type="code" colab={}
encoder = ce.OneHotEncoder(use_cat_names=True)
X_train_encoded = encoder.fit_transform(X_train)
X_val_encoded = encoder.transform(X_val)
# + id="pEe7SIBdppgc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="f332523e-c0c6-4cf6-f4b9-316e15a67d2c"
X_train_encoded.head()
# + id="0DEXEdGXprpb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="6442a1d7-10f1-4202-eb48-ce620ea4734a"
X_val_encoded.head()
# + id="T1E3IdU5qXct" colab_type="code" colab={}
imputer = SimpleImputer(strategy='mean')
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_val_imputed = imputer.transform(X_val_encoded)
# + id="S43_17QUq8O7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="89b296fa-5cbf-401a-c8b7-d3f37ccc6c81"
pd.DataFrame(X_train_imputed, columns=X_train_encoded.columns).head()
# + id="Tm1U9ajFrsBP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="c5a4af94-f291-4666-dbf9-e8de95fbc1ca"
pd.DataFrame(X_val_imputed, columns=X_val_encoded.columns).head()
# + id="ckR_FQ1cr3zW" colab_type="code" colab={}
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train_imputed)
X_val_scaled = scaler.transform(X_val_imputed)
# + id="sjSE3fI2sFgs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="08d3aa47-6f39-4f40-e2c2-f95c2d7496d5"
pd.DataFrame(X_train_scaled, columns=X_train_encoded.columns).head()
# + id="DgvfV1gysKsU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="e6e4b7e8-fc2d-450f-e756-864ea790f375"
pd.DataFrame(X_val_scaled, columns=X_train_encoded.columns).head()
# + id="LARyUzLssUQ8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="49bdbe71-b5d7-4886-e8c9-bf3afbd448a1"
model = LogisticRegressionCV(random_state=random_state, n_jobs=-1)
model.fit(X_train_scaled, y_train)
score_val = model.score(X_val_scaled, y_val)
print(f'Validation Accuracy: {score_val * 100:.2f}%')
# + [markdown] id="rrC87abXDI4i" colab_type="text"
# Plot coefficients:
# + id="pA-soYAGDI4j" colab_type="code" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# + id="rLNSm8TBuC5s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="b93adc65-7e08-4490-eae9-c035250ffbc4"
model.coef_
# + id="_lUOX9SwuNzT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="fc4ecaac-5705-400b-84c0-88b3e9942c8f"
model.coef_[0]
#coefficients for the first class
# + id="glYmrI3buGR8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="05add171-0686-4482-b83e-8a4e37308e59"
X_train_encoded.columns
# + id="zAdVCD4euLAG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 214} outputId="4807c407-bb7b-4111-db71-e4b3207b72ff"
pd.Series(model.coef_[0], X_train_encoded.columns)
# + id="rUKfoWzouY-E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 263} outputId="f56f75b9-2179-4be7-8db1-ffbb897b2349"
coefficients = pd.Series(model.coef_[0], X_train_encoded.columns)
coefficients.plot.barh();
# + id="9JtFYTKSukf3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 263} outputId="51aa993f-d0e8-4598-b065-762bc1344f4a"
coefficients.sort_values().plot.barh();
# + [markdown] id="VcBfIM02DI4k" colab_type="text"
# Generate [Kaggle](https://www.kaggle.com/c/titanic) submission:
# + id="T0-8g6rXDI4l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 349} outputId="ee4e06f8-8399-4e62-c36c-3c1d3a74ac83"
X_test = test[features]
X_test_encoded = encoder.transform(X_test)
X_test_imputed = imputer.transform(X_test_encoded)
X_test_scaled = scaler.transform(X_test_imputed)
y_pred = model.predict(X_test_scaled)
y_pred
# + id="DHu6FSZMvhrt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 32} outputId="131621ce-8e7b-4596-eb91-0d79eab61d2f"
y_pred.shape
# + id="X9wI77Q8vkXy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 411} outputId="a5486844-dcf4-4c7b-b689-2d36d680b654"
submission = test[['PassengerId']].copy()
submission['Survived'] = y_pred
submission
# + id="sbX--U8Rvxgj" colab_type="code" colab={}
submission.to_csv('titanic-submission-01.csv', index=False)
# + [markdown] id="40qcGvJvDI4m" colab_type="text"
# ## Challenge
#
# You'll use Logistic Regression for your assignment, your Sprint Challenge, and optionally for your first model in our Kaggle challenge!
# + [markdown] id="g9N_uHDWDI4n" colab_type="text"
# # Review
#
# For your assignment, you'll use a [**dataset of 400+ burrito reviews**](https://srcole.github.io/100burritos/). How accurately can you predict whether a burrito is rated 'Great'?
#
# > We have developed a 10-dimensional system for rating the burritos in San Diego. ... Generate models for what makes a burrito great and investigate correlations in its dimensions.
#
# - Do train/validate/test split. Train on reviews from 2016 & earlier. Validate on 2017. Test on 2018 & later.
# - Begin with baselines for classification.
# - Use scikit-learn for logistic regression.
# - Get your model's validation accuracy. (Multiple times if you try multiple iterations.)
# - Get your model's test accuracy. (One time, at the end.)
# - Commit your notebook to your fork of the GitHub repo.
# - Watch Aaron's [video #1](https://www.youtube.com/watch?v=pREaWFli-5I) (12 minutes) & [video #2](https://www.youtube.com/watch?v=bDQgVt4hFgY) (9 minutes) to learn about the mathematics of Logistic Regression.
# + [markdown] id="pI0s-Yu1DI4n" colab_type="text"
# # Sources
# - <NAME>, [Training, Validation, and Testing Data Sets](https://end-to-end-machine-learning.teachable.com/blog/146320/training-validation-testing-data-sets)
# - <NAME>, [R for Data Science](https://r4ds.had.co.nz/model-intro.html#hypothesis-generation-vs.hypothesis-confirmation), Hypothesis generation vs. hypothesis confirmation
# - Hastie, Tibshirani, and Friedman, [The Elements of Statistical Learning](http://statweb.stanford.edu/~tibs/ElemStatLearn/), Chapter 7: Model Assessment and Selection
# - <NAME> Guido, [Introduction to Machine Learning with Python](https://books.google.com/books?id=1-4lDQAAQBAJ&pg=PA270), Chapter 5.2.2: The Danger of Overfitting the Parameters and the Validation Set
# - <NAME> Fawcett, [Data Science for Business](https://books.google.com/books?id=4ZctAAAAQBAJ&pg=PT276), Chapter 7.3: Evaluation, Baseline Performance, and Implications for Investments in Data
# - <NAME>, [How (and why) to create a good validation set](https://www.fast.ai/2017/11/13/validation-sets/)
# - <NAME>, [Model Evaluation](https://sebastianraschka.com/blog/2018/model-evaluation-selection-part4.html)
# - <NAME>, ["A baseline for classification can be the most common class in the training dataset."](https://twitter.com/koehrsen_will/status/1088863527778111488)
|
module4-logistic-regression/LS_DS_214.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from local.torch_basics import *
from local.test import *
from local.core import *
from local.layers import *
from local.data.all import *
from local.optimizer import *
from local.learner import *
from local.metrics import *
from local.callback.all import *
from local.vision.all import *
path = untar_data(URLs.CAMVID)
camvid = DataBlock(types=(PILImage, PILMask),
get_items=get_image_files,
splitter=RandomSplitter(),
get_y=lambda o: path/'labels'/f'{o.stem}_P{o.suffix}')
dbunch = camvid.databunch(path/"images", bs=16, item_tfms=Resize((360,480)),
batch_tfms=[*aug_transforms(), Normalize(*imagenet_stats)])
dbunch.show_batch(max_n=9, vmin=1, vmax=30)
codes = np.loadtxt(path/'codes.txt', dtype=str)
dbunch.vocab = codes
# +
name2id = {v:k for k,v in enumerate(codes)}
void_code = name2id['Void']
def acc_camvid(input, target):
input = input.argmax(dim=1)
target = target.squeeze(1)
mask = target != void_code
mask.__class__ = Tensor
return (input[mask]==target[mask]).float().mean()
# -
opt_func = partial(Adam, lr=3e-3, wd=0.01)
learn = unet_learner(dbunch, resnet34, loss_func=CrossEntropyLossFlat(axis=1), opt_func=opt_func, path=path, metrics=acc_camvid)
learn.lr_find()
lr= 1e-2#3e-3
learn.fit_one_cycle(10, slice(lr))
learn.save('stage-1')
learn.show_results(max_n=4, vmin=1, vmax=30)
learn.load('stage-1')
learn.unfreeze()
learn.opt.clear_state()
learn.model
learn.opt.state_dict()
lr = 1e-3
lrs = slice(lr/100,lr)
learn.fit_one_cycle(12, lrs)
learn.show_results(max_n=4, vmin=1, vmax=30, figsize=(15,6))
|
dev/examples/camvid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
data = '''
{
"name":"Chuck",
"phone":{
"type":"int1",
"number":"+1 734 303 4456"
},
"email":{
"hide":"yes"
}
}
'''
info = json.loads(data)
print(type(info))
print('Name:', info['name'])
print('Phone:', info['phone']['number'])
print('Hide:', info['email']['hide'])
# +
import json
datas = '''
[
{
"id":"001",
"name":"Chuck",
"marks":"99"
},
{
"id":"002",
"name":"Donkey",
"marks":"94"
},
{
"id":"003",
"name":"Brunch",
"marks":"90"
}
]
'''
infos = json.loads(datas)
print('User count:', len(infos))
for item in infos:
print('ID:', item['id'])
print('Name:', item['name'])
print('Marks:', item['marks'])
# -
|
Coursera/Using Python to Access Web Data/Week-6/Excercise/Worked-Example-JSON.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/joshdsolis/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments/blob/master/module1-statistics-probability-and-inference/LS_DS_141_Statistics_Probability_and_Inference.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eJGtmni-DezY" colab_type="text"
# # Lambda School Data Science Module 141
# ## Statistics, Probability, and Inference
# + [markdown] id="FMhDKOFND0qY" colab_type="text"
# ## Prepare - examine what's available in SciPy
#
# As we delve into statistics, we'll be using more libraries - in particular the [stats package from SciPy](https://docs.scipy.org/doc/scipy/reference/tutorial/stats.html).
# + id="fQ9rkLJmEbsk" colab_type="code" outputId="937d6c40-d775-4016-9b69-70a82cc8b4c0" colab={"base_uri": "https://localhost:8080/", "height": 4427}
from scipy import stats
dir(stats)
# + id="bxW4SG_gJGlZ" colab_type="code" outputId="e715ad1a-883f-41e2-b070-a1106316f4e7" colab={"base_uri": "https://localhost:8080/", "height": 70}
# As usual, lots of stuff here! There's our friend, the normal distribution
norm = stats.norm()
print(norm.mean())
print(norm.std())
print(norm.var())
# + id="RyNKPt_tJk86" colab_type="code" outputId="db64f558-1945-4fef-f7d7-3184212d8237" colab={"base_uri": "https://localhost:8080/", "height": 70}
# And a new friend - t
t1 = stats.t(5) # 5 is df "shape" parameter
print(t1.mean())
print(t1.std())
print(t1.var())
# + [markdown] id="SRn1zMuaKgxX" colab_type="text"
# 
#
# *(Picture from [Wikipedia](https://en.wikipedia.org/wiki/Student's_t-distribution#/media/File:Student_t_pdf.svg))*
#
# The t-distribution is "normal-ish" - the larger the parameter (which reflects its degrees of freedom - more input data/features will increase it), the closer to true normal.
# + id="seQv5unnJvpM" colab_type="code" outputId="b2f84397-b204-4864-84a1-2b29eb926bbf" colab={"base_uri": "https://localhost:8080/", "height": 70}
t2 = stats.t(30) # Will be closer to normal
print(t2.mean())
print(t2.std())
print(t2.var())
# + [markdown] id="FOvEGMysLaE2" colab_type="text"
# Why is it different from normal? To better reflect the tendencies of small data and situations with unknown population standard deviation. In other words, the normal distribution is still the nice pure ideal in the limit (thanks to the central limit theorem), but the t-distribution is much more useful in many real-world situations.
#
# History sidenote - this is "Student":
#
# 
#
# *(Picture from [Wikipedia](https://en.wikipedia.org/wiki/File:William_Sealy_Gosset.jpg))*
#
# His real name is <NAME>, and he published under the pen name "Student" because he was not an academic. He was a brewer, working at Guinness and using trial and error to determine the best ways to yield barley. He's also proof that, even 100 years ago, you don't need official credentials to do real data science!
# + [markdown] id="1yx_QilAEC6o" colab_type="text"
# ## Live Lecture - let's perform and interpret a t-test
#
# We'll generate our own data, so we can know and alter the "ground truth" that the t-test should find. We will learn about p-values and how to interpret "statistical significance" based on the output of a hypothesis test.
# + id="BuysRPs-Ed0v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="73eb42e6-eecd-4a41-b1de-93458c3956fc"
# TODO - during class, but please help!
survey_data = [0, 1, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 1, 1, 1, 1, 1, 1,
0, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, 1, 1,
1, 0, 1, 0, 1, 1, 0, 1, 0, 1, 1, 0, 0, 0]
import numpy as np
import pandas as pd
df = pd.DataFrame(survey_data)
df.describe()
# + id="ml53p2xVNgbz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 347} outputId="0de629ee-888f-4bb9-dfbb-c97634617c53"
df.plot.hist();
# + id="6z9OuefUNO2z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="feb101cb-a432-4a42-8706-f08564fb09a6"
import scipy
scipy.stats.ttest_1samp(survey_data, 0.5)
# + id="4610IYHRO1iH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3759abdd-b086-48a2-8a6a-5580bc4721bd"
sample_stderr = 0.478518/np.sqrt(len(survey_data))
sample_mean = 0.66
null_hypothesis_mean = 0.5
# We want to calculate: tstat = 2.364321853156195
stderr/(sample_mean-0.5) # Wrong, but conceptually related
t_stat = (sample_mean - null_hypothesis_mean) / sample_stderr
print(t_stat)
# + id="v6LZqGcyUUxp" colab_type="code" colab={}
# Science! Reproducibility...
import random
def make_soda_data(n=50):
# FAIR VERSION
# return pd.DataFrame([random.randint(0,1) for _ in range(n)])
return pd.DataFrame(np.random.binomial(n=1,p=0.51,size=n))
# + id="EkskOHTJU9Ng" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="8e56f969-9504-4a8c-cf90-f30f669468ef"
make_soda_data().describe()
# + id="eTIuwGVHVEQZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="06b6996b-c025-4c63-e02f-c52d2a51d3be"
t_statistics = []
n_experiments = 10
p_values = []
for _ in range (n_experiments):
df = make_soda_data(n=500000)
ttest = scipy.stats.ttest_1samp(df,0.5)
t_statistics.append(ttest.statistic)
p_values.append(ttest.pvalue)
pd.DataFrame(t_statistics).describe()
# + id="lOXqc6OfZV5v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="f90cdfcb-2da4-4c2d-eb4f-deeb0858ceee"
pd.DataFrame(p_values).describe()
# + [markdown] id="egXb7YpqEcZF" colab_type="text"
# ## Assignment - apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="nstrmCG-Ecyk" colab_type="code" colab={}
# TODO - your code here!
import pandas as pd
import scipy
import numpy as np
names = ['party','handicapped infants', 'water project cost sharing',
'adoption of the budget resolution', 'physician fee freeze',
'el salvador aid', 'religious groups in schools',
'anti satellite test ban','aid to nicaraguan contras',
'mx missle', 'immigration','synfuels corp cutback','education spending',
'superfund right to sue','crime','duty free exports',
'export admin act south africa']
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data', header = None)
df=df.rename(columns = {0:'party'})
# + id="t1gXmvMmmYKU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="69c9652a-ae9d-435a-8192-996bb560944e"
df.isna().sum()
# + id="yxjyfXnzmiYk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="bb6c8939-a57c-465a-bf4e-21797db9a5f7"
df.shape
df.head()
# + id="4rxw7sWumoCd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="417f2600-f1f7-465a-df09-750f1245e938"
# Changing '?' to NaN
df = df.replace('?',0.5)
df = df.replace('y', 1)
df = df.replace('n', 0)
df.isna().sum()
# + id="5d0YDmwRqJeF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="874ace52-bcd4-4321-a942-909b63c396e7"
pd.crosstab(df['party'],df[1])
# + id="Ubm3E63wqN0d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="3a5ab79b-f149-4ed5-b942-eb120ed4213a"
df.groupby(["party", 1]).size()
# + id="zsJKSEK4zt8i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="a7b50c83-c897-4950-a51a-d8ad2e4e1195"
df.isna().sum()
# + id="a9sBah7-6UgL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d29a0427-e65c-4c21-cd53-6748e1b8430f"
df.shape
# + id="egYptKVa_1pd" colab_type="code" colab={}
cross = pd.crosstab(df['party'],columns = df[16])
# + id="mV969UgrFAq5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="770ad70c-940f-414e-de71-14c3d4769216"
cross
# + id="L6Ziw-VdAZah" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 397} outputId="16738174-5474-4a72-9153-62da7fb01e16"
cross.plot.bar();
# + id="9vObvOeGE8S9" colab_type="code" colab={}
df_r = df[(df['party']=='republican')]
df_d = df[(df['party']=='democrat')]
# + id="qktqeuOQA393" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="3ed980d3-40bd-432f-9745-1186fe563c68"
scipy.stats.ttest_ind(df_r[1],df_d[1])
# + id="6V26esVhPzCF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="87f49eea-7c65-4fe6-f52d-54b55bac53c0"
scipy.stats.ttest_ind(df_r[2],df_d[2])
# + [markdown] id="4madxysRUi-R" colab_type="text"
# # Democrats support issue adoption of the budget resolution more than Republicans with a p value = 2.872e-76
# + id="9tUCAD33UIwF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="15ae3830-3e87-454a-dbe1-1ca6a516adfe"
scipy.stats.ttest_ind(df_r[3],df_d[3])
# + [markdown] id="wNB3JUFTU_MG" colab_type="text"
# # Republicans support physician fee freeze more than Democrats with a p value = 3.97e-169
# + id="AQfx_Mv-UNnb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="72f1ca19-5b45-4da7-e160-d1d8ab439e49"
scipy.stats.ttest_ind(df_r[4],df_d[4])
# + [markdown] id="160H7zd1VoSM" colab_type="text"
# # There isn't much difference between Republicans and Democrats on water project cost sharing with a p value = 0.93
# + id="Z2QRXGzXU4Wp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="a59d1736-f460-4bff-a437-6d6b0439ffc2"
scipy.stats.ttest_ind(df_r[2],df_d[2])
# + id="v1B-zhMuVgdF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="36af2960-496a-457f-f24c-31cabfcaf61d"
scipy.stats.ttest_ind(df_r[16],df_d[16])
# + id="Z4S34cVVqaPc" colab_type="code" colab={}
|
module1-statistics-probability-and-inference/LS_DS_141_Statistics_Probability_and_Inference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# #### This is forked from https://github.com/ghgr/HFT_Bitcoin, I have added some notes for myself better understanding.
from IPython.display import HTML
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
The raw code for this IPython notebook is by default hidden for easier reading.
To toggle on/off the raw code, click <a href="javascript:code_toggle()">here</a>.''')
import pandas as pd
import json
import random
import datetime
import seaborn as sb
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams['figure.figsize'] = (15.0, 5.0)
import numpy as np
from collections import Counter
import base64
# +
# plt.annotate?
# +
# Basic Function to generate graphs, videos, images???
def annotate(x_point, y_point, x_text, y_text, text):
plt.annotate(text, xy=(x_point, y_point),
xytext=(x_text, y_text),
arrowprops=dict(facecolor='black', shrink=0.05))
def video(fname, mimetype):
from IPython.display import HTML
#video_encoded = open(fname, "rb").read().decode("base64")
with open(fname, 'rb') as image_file:
video_encoded = base64.b64encode(image_file.read())
video_tag = '<video loop autoplay alt="orderbook video" src="data:video/{0};base64,{1}" width=100%>'.format(mimetype, video_encoded)
return HTML(data=video_tag)
def image(fname, width):
from IPython.display import HTML
data = '<img src="%s" width=%d%%>' % (fname, width)
return HTML(data=data)
# -
# # High Frequency Trading in Bitcoin Exchanges
#
# Note: This article is the notebook corresponding to my post [High Frequency Trading in Bitcoin Exchanges](https://www.linkedin.com/pulse/high-frequency-trading-bitcoin-exchanges-eduardo-pena-vina) originally published on Linkedin on July 27, 2017.
#
# ## Introduction
#
# In this post I analyze the **presence and activity of high frequency trading** in a Bitcoin exchange. Since to date this markets are *extremely unregulated*, such behaviour takes places with little to no constraint. I show how **over 99%** of orders placed are **not meant to be filled**, but instead to distort the perception of the market. In addition, I try to spot common HFT strategies, such as *Quote Spoofing*, *Layering* and *Momentum ignition*. Given the anonymous nature of these exchanges these last results are to some extent *subjective*.
#
#
# ## What is High Frequency Trading?
#
# From Wikipedia [1], High-frequency trading (HFT) is a type of algorithmic trading characterized by high speeds, high turnover rates, and high order-to-trade ratios that leverages high-frequency financial data and electronic trading tools.
# ## Methodology
#
# This analysis has been carried out with order data from the Websocket stream from **GDAX**, a US based digital asset exchange [2] owned by Coinbase. It is one of the largest markets (over **42 MM USD/day**) [3] and it exposes a high performance socket where all orders are broadcasted. In addition, it offers some interesting features for data analysis:
#
# 1. Orders are timestamped (as opposed to Bitfinex, for example)
# 2. It has millisecond granularity (again, as opposed to Bitfinex)
# 3. It says whether an order has been matched or cancelled -one could argue that disappearing orders far from the bid/ask spread must have been cancelled (and it's true), but for orders *inside* the spread, this information is necessary.
#
#
# While data has been captured for several days (at the time of this post I'm still capturing data), for the following analysis only data from **July 21, 2017** has been taken. Mind you, there are still **over 2 Million datapoints**.
#
#
# Since the GDAX feed does not explicitly keep information of the current best bid/ask, a little preprocessing is needed. The best bid is the *highest* price for currently open *BUY* orders, while the best ask is the *lowest* price for open *SELL* orders. Although this calculation is not complicated nor particularly slow, it's better to explicitly append the current best bid/ask as additional columns. No further preprocessing has been carried out.
#
# ## Related work
#
# While writing this article, I came across a blog post from <NAME> at *Parasec* [4], who made a **similar analysis in 2014**. While the amount of data differs by orders of magnitude, the findings are the same, especially concerning *flashing orders*. Quoting from his site:
#
# > I collected order book event data from the Bitstamp exchange over a **4 month period**, between July and October (2014), resulting in a dataset of **~33 million individual events**; A minuscule dataset in comparison to the throughput on "conventional" exchanges, see (Nanex: High Frequency Quote Spam) for example.
#
# > While the event dataset consists of ~33 million events, these events can be broken down into individual orders and their types. In total, of the identifiable order types, **there were 14,619,019 individual "flashed orders"** (orders added and later deleted without being hit) **representing 93% of all order book activity**, 707,113 "resting orders" (orders added and not deleted unless hit) and 455,825 "marketable orders" (orders that crossed the book resulting in 1 or more reported trades).
#
# As we'll soon see in this report, I recorded **2,169,450** events in less than one day. That means, the number of events per unit of time is ** 8 times bigger than in 2014**. *Flash orders* are still a majority, **representing over 99% of all order book activity**.
# ## Analysis
#
# ### Data exploration
filename = 'data/2017_07_21_10_52_05.csv.gz'
data = pd.read_csv(filename, parse_dates=['time'], index_col="time")
data['time'] = data.index
data.columns
data.head(10)
# First step, load data in memory (2,169,450 trades) corresponding to 723,880 different orders.
print "Loaded {:,} trades".format(data.shape[0])
print ("Loaded {:,} orders".format(data.order_id.nunique()))
# The provided data feed looks like that:
# There are multiple fields and many NaN (Not-a-Number) values. This happens because all sort of orders are mixed, and not every field is relevant to every order. (For details the reader is kindly pointed to the official documentation [7]).
# We can focus on the most commonly relevant parameters:
keys = ['price','remaining_size','side','reason','type','order_id']
data[keys].head()
# Now the data is clearer. Each order is defined by its **arrival time** (at the server), its **price**, **amount**, **side** (*buy* or *sell*), **status** and **unique id**. We see that for the first 5 orders, 3 of the are cancellations! (more about that soon).
# #### Granularity(n. 间隔尺寸、粒度) of data
# Let's take the time different between arriving orders, and plot as histogram
dt = data.time.diff().iloc[1:].dt
dt = dt.seconds.astype(np.float) + dt.microseconds * 1e-6
dt*=1000
dt.sort_values().diff().hist(bins = np.arange(0,10,0.1));
plt.yscale('log')
plt.xlabel("Delay (ms)")
plt.ylabel("Number of trades")
plt.title("Delay between trades (log scale)")
plt.show()
# We see that a large amount of orders take place simultaneously, and that the smallest *non zero* value of time delay is 1 **ms**. We can directly confirm it with code:
print ("Minimum non zero number of milliseconds: ", dt[dt>0].min())
# #### Analysis of orders by id
data[data.order_id=='9144b925-ecb6-40fa-880b-2e9f83039812'][keys+['order_type']]
# An interesting observation that will be useful later is that each order_id appear at most three times (99.01%). That means, the order arrives, the order waits ("open") and the order is matched or cancelled.
for order, freq in Counter(data.groupby('order_id').size()).most_common():
print (order, freq)
# #### Analysis of orders by outcome
#
# An incoming order has two possible outcomes. Either it is filled (matched with an existing or incoming order) or is is cancelled. During this observation 7,053 orders were filled, while a majority of 714,648 where *Cancelled*. That is a 99.02% of incoming orders were ultimately cancelled!
data.groupby('reason').size()
# #### Grouping orders
# A different exchange, Bitfinex, has been previously analyzed (not shown here for the sake of conciseness). Its data-feed had some major drawbacks that ultimately made me drop it and work with GDAX. Nevertheless, its so called "Raw Book Order" feed, showed interesting information. In that exchange traders don't seem to Send/Cancel many orders, but rather Send/Update/Update/Update/.../Update/Cancel. I don't fully understand why, but I guess it could be to bypass API Limits and to optimize execution of orders (by keeping the place in the order book at a given price).
#
# In any case, I saw that quickly arriving and cancelling orders (flashing orders) at a given price and similar amount are caused by the same individual. Knowing that, let's find orders at a given price and amount. For example, 0.5BTC@$2707.81
w=data[(data.price==2707.81) & (data.remaining_size==0.5)]
w[['price','remaining_size','side','reason','order_id']].tail()
# Let's focus on order 04211005-4140-44c5-98d6-65fcec8eb9e5
data[data.order_id=="04211005-4140-44c5-98d6-65fcec8eb9e5"][['price','remaining_size','side','reason','order_id','type', 'bid','ask']]
# This BUY order was placed **and cancelled 295 ms later** at a level of $2707.81 (when the best ASK price was 2714.51). It seems obvious that this order had no expectation of being filled.
# Let's see if orders are flashed at higher or lower frequencies than 295 ms.
canceled_idx = data.order_id[data.reason=="canceled"]
d = data[data.order_id.isin(canceled_idx)]
g = d.groupby("order_id")
dt = g.time.last() - g.time.first()
duration = dt.dt.seconds.astype(np.float) + dt.dt.microseconds*1e-6
duration*=1000
duration.hist(bins = np.arange(0,300,2));
plt.yscale('log')
plt.title("Number of orders for each lifespan (log scale)")
plt.ylabel("Number of orders (log)")
plt.xlabel("Time from arrival to cancellation (ms)")
plt.show()
# Indeed, there are orders flashing at <5 milliseconds! Let's see some of the fastest flashing orders:
duration[duration>1].sort_values().head()
# For example, let's take order_id 7f043088-f9ad-46f6-bfae-e577db3f7363
data[data.order_id=="7f043088-f9ad-46f6-bfae-e577db3f7363"][keys+['bid','ask']]
# Three milliseconds is an extremely short amount of time considering the GDAX Rate Limits:
#
# > ### Rate Limits
# > 1. We throttle public endpoints by IP: 3 requests per second, up to 6 requests per second in bursts.
# > 2. We throttle private endpoints by user ID: 5 requests per second, up to 10 requests per second in bursts.
# > 3. The FIX API throttles each command type (eg.: NewOrderSingle, OrderCancelRequest) to **30 commands per second**.
#
# What's **Really interesting** is that these patterns seem to exceed the API Limits (the highest, at 30 command per seconds, means 1 order each 33 ms).
# So we can confidently say that **there are bots manipulating the order books**. The next question is, how many? and are they controlled by the same person? Let's find out...
# #### Bots trading
def plotCancelBehaviourForPriceAndSize(price, size, lim=-1):
plt.figure(figsize=(15,5))
#q = data[data.order_id==most_prolific_bots.index[0]]
q = data[((data['size']==size) | (data['remaining_size']==size)) & (data['price']==price)].head(lim)
t=q.time.diff().median()
median_tick_s = t.seconds+t.microseconds*1e-6;
order_type = q.side.iloc[0]
d = data.loc[q.index[0]:q.index[-1]]
d.bid.plot(label="bid price" )
d.ask.plot(label="ask price")
q[q.reason!="canceled"].price.plot(label="%s %.2f BTC @ %.2f" % (order_type, size, price), style='o');
ylim0 = min(d.bid.min(), q.price.min())
ylim1 = max(d.ask.max(), q.price.max())
plt.vlines(q.index[q.reason=="canceled"], ylim0, ylim1, alpha = 0.2);
plt.xlabel("Time")
plt.ylabel("Price (USD)")
plt.title("Cancelled orders along time for %.2f BTC @ $%.2f. Median tick: %.1f milliseconds" % (size, price,median_tick_s*1000))
plt.legend(loc='best', frameon=True)
plt.ylim(d.bid.min(), d.ask.max())
plt.show()
# Let's see the cancel behavior for different price levels and amounts. Red points are SELL orders (not always visible), vertical grey lines are cancellations and the blue and green lines are bid and ask price, respectively.
g = data.groupby(['price','size'])
for (price, size), i in g.size().sort_values(ascending=False).head(5).iteritems():
plotCancelBehaviourForPriceAndSize(price,size, lim=1000)
# There is certainly a cadence of placing and cancelling orders.
# ## HTF Strategies
# The Bocconi Students Investment Club (BSIC) [5] describes some strategies which the HFT traders use to distort the perception of the market. For this post I'll focus on Spoofing, Layering and Momentum Ignition.
#
# ### Spoofing & Layering
#
# Quoting from BSIC [5]:
#
# > Spoofing is a strategy whereby one places limit orders, and removes them before they are executed. By spoofing limit orders, perpetrators hope to distort other trader’s perceptions of market demand and supply. As an example, a large bid limit order could be placed with the intention of being canceled before it is executed. The spoofer would then seek to benefit from prices rising as the result of false optimism others would see in the market structure.
# ### Detection
#
# There is evidence of high frequency spoofing on July 21, 2017 between 09:45:52 and 09:45:56. Let's take a look at the order book. Red points are SELL orders (3 BTC @ $2741.99), vertical grey lines are cancellations and the blue and green lines are bid and ask price, respectively.
# +
size = 3.00
price = 2741.99
plt.figure(figsize=(15,5))
q = data[((data['size']==size) | (data['remaining_size']==size)) & (data['price']==price)].head(1000)
q = q.loc['2017-7-21 9:45:52.000': '2017-7-21 9:45:56.000' ]
t=q.time.diff().median()
median_tick_s = t.seconds+t.microseconds*1e-6;
order_type = q.side.iloc[0]
d = data.loc[q.index[0]:q.index[-1]]
d.bid.plot(label="bid price" ,alpha = 0.5 )
d.ask.plot(label="ask price",alpha = 0.5)
q[q.reason!="canceled"].price.plot(label="%s %.2f BTC @ %.2f" % (order_type, size, price), style='o');
ylim0 = 2739
ylim1 = 2743
plt.vlines(q.index[q.reason=="canceled"], ylim0, ylim1, alpha = 0.2);
plt.xlabel("Time")
plt.ylabel("Price (USD)")
plt.title("Cancelled orders along time for %.2f BTC @ $%.2f. Median tick: %.1f milliseconds" % (size, price,median_tick_s*1000))
plt.legend(loc='best', frameon=True)
plt.ylim(ylim0, ylim1)
plt.show()
# -
# The orders are cancelled around 25 ms after they arrive! They are flashed with a frequency of around 3 Hz. The following animation shows the order book in more or less real time. Notice how a SELL order flashes at level 2741.99 with amount 3 BTC.
# +
def plotOrderBookAtTime(t,xmin, xmax, ymax, fname=False, arrowAtPrice=False):
data_past = data[data.index<=t]
q=data_past.groupby('order_id').type.last()=="open"
q = data_past[data_past.order_id.isin(q[q].index)]
q = q[q.type=="received"]
ask = q[q.side=="sell"].groupby('price')['size'].sum().sort_index(ascending=True)
bid = q[q.side=="buy"].groupby('price')['size'].sum().sort_index(ascending=False)
plt.bar(bid.index, bid, width = 0.01, color='g')
plt.bar(ask.index, ask, width = 0.01, color='r')
#xmin, xmax = np.percentile(bid.index,80), np.percentile(ask.index,20)
#ymin, ymax = 0, max(bid[bid.index>xmin].max(), ask[ask.index<xmax].max())
plt.xlim(xmin, xmax)
plt.ylim(0, ymax)
plt.title("Order book at time: "+str(t))
plt.xlabel("Price (USD)")
plt.ylabel("Depth (BTC)")
if arrowAtPrice:
plt.xticks(list(plt.xticks()[0]) + [arrowAtPrice])
plt.annotate('$%.2f' % (arrowAtPrice), xy=(arrowAtPrice, ymax-4), xytext=(arrowAtPrice, ymax-1),arrowprops=dict(facecolor='black', shrink=0.05))
if fname:
plt.savefig(fname)
plt.close()
else:
plt.show()
if False:
dates = data.loc['2017-7-21 9:45:52.000'].index
dates = dates.append(data.loc['2017-7-21 9:45:53.000'].index)
dates = dates.append(data.loc['2017-7-21 9:45:54.000'].index)
dates = dates.append(data.loc['2017-7-21 9:45:55.000'].index)
dates = dates.append(data.loc['2017-7-21 9:45:56.000'].index)
ts = [t for t in dates]
for i,t in enumerate(ts):
plotOrderBookAtTime(t,2730, 2750, 10, fname = "img/pic_%05d.png" % (i), arrowAtPrice=2741.99)
print ("Done %d/%d" % (i, len(ts)))
video("videos/animation1.mp4","mp4")
# -
# One interesting thing is that neither the bid or ask price moves.
# Also from [5]:
# > More controversial has been the act of layering which carries many similarities to outright spoofing, but differs in that orders are placed evenly across prices with the goal of reserving an early execution priority at each given price level. If the person has no trade to execute at that price point the orders are simply removed. Despite being more benign in nature, the act of layering also distorts market demand and supply perception.
# It seems to be evidence of layering. Let's take a closer look at the minute between July 21, 2017 between 09:41:00 and 9:42:00. Orders seem to push the ASK level downwards, eventually decreasing the BID price. Next, BUY orders are placed at this lowered level, to be sold when the BID price recovers.
# +
q = data.loc['2017-7-21 9:41:00.000': '2017-7-21 9:42:01.000' ]
ymin, ymax = 2739, 2743
q.bid.plot(label="bid price")
q.ask.plot(label="ask price")
plt.xlabel("Time")
plt.ylabel("Price (USD)")
plt.legend(loc='best', frameon=True)
plt.ylim(ymin, ymax)
plt.annotate('Start Layering', xy=('2017-7-21 9:41:08.000', 2742), xytext=('2017-7-21 9:41:12.000', 2742.5),arrowprops=dict(facecolor='black', shrink=0.05))
plt.annotate('Layering Succeeded!', xy=('2017-7-21 9:41:17.600', 2741.25), xytext=('2017-7-21 9:41:05.000', 2740),arrowprops=dict(facecolor='black', shrink=0.05))
plt.annotate('Place BUY Orders at low price', xy=('2017-7-21 9:41:18.400', 2740.15), xytext=('2017-7-21 9:41:19.000', 2739.2),arrowprops=dict(facecolor='black', shrink=0.05))
plt.annotate('Sell at higher price', xy=('2017-7-21 9:41:27.000', 2740.5), xytext=('2017-7-21 9:41:29.000', 2741.15),arrowprops=dict(facecolor='black', shrink=0.05))
plt.show()
# -
# ### Momentum ignition
#
# Still quoting [5]
#
# > Momentum ignition is a strategy in which a trader aims to cause a sharp movement in the price of a stock by using a series of trades, which indicate patterns for high frequency traders, with the motive of attracting other algorithm traders to also trade that stock. The instigator of the whole process knows that after the somewhat “artificially created” rapid price movement, the price reverts to normal and thus the trader profits by taking a position early on and eventually trading out before it fizzles out.
#
# >To detect momentum ignition, it is important to focus on the following three main characteristics as shown in the chart below:
#
# >1. Stable prices and a spike in volume
# >2. A large price movement compared to the intraday volatility
# >3. Reversion to the starting price under a lower volume
# The following picture from zerohedge and Credit Suisse AES Analysis illustrates this behavior.
image("img/momentum.png", 70)
# This behaviour takes place on July 21, 2017 between 12:00:20 and 12:50:01
# +
plt.figure(figsize=(15,10))
plt.subplot(211)
t0 = '2017-7-21 12:00:20.000'
t1 = '2017-7-21 12:50:01.000'
q = data.loc[t0:t1].groupby(pd.TimeGrouper(freq='60S'))
bids = q.bid.last()
asks = q.ask.last()
vol = data[data.type=='match'].loc[t0:t1].groupby(pd.TimeGrouper(freq='60S'))['size'].sum()
bids.plot()
asks.plot()
plt.ylabel("Price (USD)")
annotate('2017-7-21 12:09:20.000', 2772.0, '2017-7-21 12:12:20.000', 2760.6, "1) Short term volume spike\n with no price move")
annotate('2017-7-21 12:24:03.000', 2762.5, '2017-7-21 12:27:20.000', 2765.6, "2) Large price move\n with high volume")
annotate('2017-7-21 12:47:03.000', 2768.0, '2017-7-21 12:42:20.000', 2760.0, "3) Price reversion\n on low volume")
plt.subplot(212)
vol.plot(kind='bar')
plt.ylabel("Volume traded (BTC)")
plt.show()
# -
# Taking a look at the timeframe between July 21, 2007 between 08:53:28 and 08:53:43 there seems to be a mix between momentum ignition and layering (see previous section). Note the spike in volume at 08:53:31.
# +
plt.figure(figsize=(15,10))
t0 = '2017-7-21 8:53:28.000'
t1 = '2017-7-21 8:53:43.000'
q = data.loc[t0:t1]
bids = q.bid
asks = q.ask
#w = data.loc[t0:t1].groupby([pd.TimeGrouper(freq='5S'),'side'])
asks.plot(linestyle='-', marker='X', ms=3, c='r', linewidth=1)
bids.plot(linestyle='-', marker='X', ms=3, c='g', linewidth=1)
plt.yticks(list(plt.yticks()[0]) + [2713.64, 2713.28 ])
annotate('2017-07-21 08:53:30.763',2717.80, '2017-07-21 08:53:31.000',2719,'BUY wall pushed down')
annotate('2017-07-21 08:53:30.763',2713.64, '2017-07-21 08:53:29.173',2710,'SELL 2.8BTC@2713.28')
annotate('2017-07-21 08:53:33.763',2717.00, '2017-07-21 08:53:36.000',2718,'Layering')
annotate('2017-07-21 08:53:37.500',2712.80, '2017-07-21 08:53:34.000',2711,'Layering Succedded!')
annotate('2017-07-21 08:53:40.800',2712.10, '2017-07-21 08:53:41.500',2712.5,'Lucky market maker')
annotate('2017-07-21 08:53:43.000',2709.60, '2017-07-21 08:53:41.000',2708.5,'Low ASK price')
plt.figure(figsize=(15,4))
vol = data[data.type=='match'].loc[t0:t1].groupby(pd.TimeGrouper(freq='1S'))['size'].sum()
vol.plot(kind='bar')
plt.title("MATCHED VOLUME")
plt.tight_layout()
plt.show()
# -
# If we look 5 minutes into the future, until 08:58:00, we see that the price recovers to previous levels.
# +
#plt.figure(figsize=(15,10))
t0 = '2017-7-21 8:53:28.000'
tmp = '2017-7-21 8:53:43.000'
t1 = '2017-7-21 8:58:00.000'
q = data.loc[t0:t1]
bids = q.bid
asks = q.ask
#w = data.loc[t0:t1].groupby([pd.TimeGrouper(freq='5S'),'side'])
asks.plot(linestyle='-', marker='X', ms=3, c='r', linewidth=1)
bids.plot(linestyle='-', marker='X', ms=3, c='g', linewidth=1)
plt.vlines(tmp,2704,2718, linestyles='--')
annotate(tmp,2716.5,'2017-7-21 8:53:55.000', 2717.5, tmp)
annotate('2017-7-21 8:57:7.000',2713.5,'2017-7-21 8:55:55.000', 2716, "Back to previous level")
plt.show()
# -
# ## Conclusion
#
# According to an interview carried out by The Atlantic [6] to <NAME> of the University of Pennsylvania and <NAME> at MIT, this behaviour also happens in traditional trading, and its causes are still matter of dispute. Relevant extract:
# > [...] why would a firm engage in this behavior? Lo and Kearns offered a few theories of their own about what could be happening.
#
# > To be honest, we can't come up with a good reason," Kearns said. What's particularly difficult to explain is how diverse and prevalent the patterns are. If algorithmic traders are simply testing new bots out -- which isn't a bad explanation -- it doesn't seem plausible that they'd do it so often. Alternatively, one could imagine the patterns are generated by some set of systemic information processing mistakes, but then it might be difficult to explain the variety of the patterns.
#
# > "It's possible that the observed patterns are not malicious, in error, or for testing, but for information-gathering," Kearns observed. "One could easily imagine a HFT shop wanting to regularly examine (e.g.) the latency they experienced from the different exchanges under different conditions, including conditions involving high order volume, rapid changes in prices and volumes, etc. And one might want such information not just when getting started, but on a regular basis, since latency and other exchange properties might well be expected to change over time, exhibit seasonality of various kind, etc. The super-HFT groups might even make co-location decisions based on such benchmarks."
#
# ## References
# [1]https://en.wikipedia.org/wiki/Bitfinex
#
# [2] https://www.gdax.com/
#
# [3] Source: https://coinmarketcap.com
#
# [4] http://parasec.net/blog/order-book-visualisation/
#
# [5] http://www.bsic.it/marketmanipulation/
#
# [6] https://www.theatlantic.com/technology/archive/2010/08/explaining-bizarre-robot-stock-trader-behavior/61028/
#
# [7] https://docs.gdax.com/
|
HFT_Bitcoin-master/hftbitcoin.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Node Classification - Introduction
# In this Notebook we are going to examine the process of using Amazon Neptune ML feature to perform RDF Literal string classification in an RDF graph.
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: This notebook take approximately 1 hour to complete</div>
#
# [Neptune ML](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning.html#machine-learning-overview) is a feature of Amazon Neptune that enables users to automate the creation, management, and usage of Graph Neural Network (GNN) machine learning models within Amazon Neptune. Neptune ML is built using [Amazon SageMaker](https://aws.amazon.com/sagemaker/) and [Deep Graph Library](https://www.dgl.ai/) and provides a simple and easy to use mechanism to build/train/maintain these models and then use the predictive capabilities of these models within a Gremlin query to predict elements or property values in the graph.
#
# For this notebook we are going to show how to perform a common machine learning task known as **node classification**. Node classification is a common semi-supervised machine learning task where a model built using existing nodes, which in RDF means existing string Literal values. Where the triple does not exist, we can predict the value of the Literal. Node classification is not unique to GNN based models (look at DeepWalk or node2vec) but the GNN based models in Neptune ML provide additional context to the predictions by combining the connectivity and features of the local neighborhood of a node to create a more predictive model.
#
# Node classification is commonly used to solve many common buisness problems such as:
#
# * [Identifying fradulent transactions](https://aws.amazon.com/blogs/machine-learning/detecting-fraud-in-heterogeneous-networks-using-amazon-sagemaker-and-deep-graph-library/)
# * Predicting group membership in a social or identity network
# * Predicting categories for product recommendation
# * Predicting user churn
#
#
# Neptune ML uses a four step process to automate the process of creating production ready GNN models:
#
# 1. **Load Data** - Data is loaded into a Neptune cluster using any of the normal methods such as SPARQL queries or using the Neptune Bulk Loader.
# 2. **Export Data** - A service call is made specifying the machine learning model type and model configuration parameters. The data and model configuration parameters are then exported from a Neptune cluster to an S3 bucket.
# 3. **Model Training** - A set of service calls are made to pre-process the exported data, train the machine learning model, and then generate an Amazon SageMaker endpoint that exposes the model.
# 4. **Run Queries** - The final step is to use this inference endpoint within our SPARQL queries to infer data using the machine learning model via a SERVICE federated Query call.
#
# 
#
#
# This notebook uses the [MovieLens 100k dataset](https://grouplens.org/datasets/movielens/100k/) provided by [GroupLens Research](https://grouplens.org/datasets/movielens/). This dataset consists of movies, users, and ratings of those movies by users.
#
# 
#
# For this notebook we'll walk through how Neptune ML can predict the group membership of a product in a product knowledge graph. To demonstrate this we'll predict the genre of new movies added to our product knowledge graph. We'll walk through each step of loading and exporting the data, configuring and training the model, and finally we'll show how to use that model to infer the genre of movies using SPARQL queries.
# ## Checking that we are ready to run Neptune ML
# Run the code below to check that this cluster is configured to run Neptune ML.
import neptune_ml_sparql_utils as neptune_ml
neptune_ml.check_ml_enabled()
# If the check above did not say that this cluster is ready to run Neptune ML jobs then please check that the cluster meets all the pre-requisites defined [here](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning.html#machine-learning-overview).
#
# ## Loading the data
#
# The first step in building a Neptune ML model is to load data into the Neptune cluster. Loading data for Neptune ML follows the standard process of ingesting data into Amazon Neptune, for this example we'll be using the Bulk Loader.
#
# We have written a script that automates the process of downloading the data from the MovieLens websites and formatting it to load into Neptune. All you need to provide is an S3 bucket URI that is located in the same region as the cluster.
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: This is the only step that requires any specific input from the user, all remaining cells will automatically propogate the required values.</div>
s3_bucket_uri="s3://<INSERT S3 BUCKET OR PATH>"
# remove trailing slashes
s3_bucket_uri = s3_bucket_uri[:-1] if s3_bucket_uri.endswith('/') else s3_bucket_uri
# Now that you have provided an S3 bucket, run the cell below which will download and format the MovieLens data into a format compatible with Neptune's bulk loader.
response = neptune_ml.prepare_movielens_data_rdf(s3_bucket_uri, clear_staging_area=True)
# This process only takes a few minutes and once it has completed load the data using the `%load` command in the cell below.
# %load -s {response} -f nquads -p OVERSUBSCRIBE --run
# # Export the data and model configuration
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: Before exporting data ensure that Neptune Export has been configured as described here: <a href="https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-service.html#machine-learning-data-export-service-run-export">Neptune Export Service</a></div>
# With our product knowledge graph loaded we are ready to export the data and configuration which will be used to train the ML model.
#
# The export process is triggered by calling to the [Neptune Export service endpoint](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-service.html). This call contains a configuration object which specifies the type of machine learning model to build, in this example node classification, as well as any feature configurations required.
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: The configuration used in this notebook specifies only a minimal set of configuration options meaning that our model's predictions are not as accurate as they could be. The parameters included in this configuration are one of a couple of sets of options available to the end user to tune the model and optimize the accuracy of the resulting predictions.</div>
#
# The configuration options provided to the export service are broken into two main sections, selecting the target and configuring features.
#
# ## Selecting the target
#
# In the first section, selecting the target, we specify what type of machine learning task will be run, and in the case of node classification, what the target node label and property we want to predict. To run a node classification task we are only required to specify the node label and target property to infer.
#
# In this example below we specify the `genre` property on the `movie` node as our target for prediction by including these values in the `targets` sub-parameter of the `additionalParams` object as shown below.
#
# ```
# "additionalParams": {
# "neptune_ml": {
# "targets": [
# {
# "node": "movie",
# "property": "genre"
# }
# ],
# ....
# ```
#
# ## Configuring features
# The second section of the configuration, configuring features, is where we specify details about the types of data stored in our graph and how the machine learning model should interpret that data. In machine learning, each property is known as a feature and these features are used by the model to make predictions.
#
# When data is exported from Neptune all properties of all nodes are included. Each property is treated as a separate feature for the ML model. Neptune ML does its best to infer the correct type of feature for a property, in many cases, the accuracy of the model can be improved by specifying information about the property used for a feature. By default Neptune ML puts features into one of two categories:
#
# * If the feature represents a numerical property (float, double, int) then it is treated as a `numerical` feature type. In this feature type data is represented as a continuous set of numbers. In our example, the `age` of a `user` would best be represented as a numerical feature as the age of a user is best represented as a continuous set of values.
# * All other property types are represented as `category` features. In this feature type, each unique value of data is represented as a unique value in the set of classifications used by the model. In our MovieLens example the `occupation` of a `user` would represent a good example of a `category` feature as we want to group users that all have the same job.
#
# If all of the properties fit into these two feature types then no configuration changes are needed at the time of export. However, in many scenarios these defaults are not always the best choice. In these cases, additional configuration options should be specified to better define how the property should be represented as a feature.
#
# One common feature that needs additional configuration is numerical data, and specifically properties of numerical data that represent chunks or groups of items instead of a continuous stream.
#
# Let's say that instead of wanting `age` to be represented as a set of continuous values we want to represent it as a set of discrete buckets of values (e.g. 18-25, 26-24, 35-44, etc.). In this scenario we want to specify some additional attributes of that feature to bucket this attribute into certain known sets. We achieve this by specifying this feature as a `numerical_bucket`. This feature type takes a range of expected values, as well as a number of buckets, and groups data into buckets during the training process.
#
# Another common feature that needs additional attributes are text features such as names, titles, or descriptions. While Neptune ML will treat these as categorical features by default the reality of these features is that they will likely be unique for each node. For example, since the `title` property of a `movie` node does not fit into a category grouping our model would be better served by representing this type of feature as a `text_word2vec` feature. A `text_word2vec` feature uses techniques from natural language processing to create a vector of data that represents a string of text.
#
# In our export example below we have specified that the `title` property of our `movie` should be exported and trained as a `text_word2vec` feature and that our `age` field should range from 0-100 and that data should be bucketed into 10 distinct groups.
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Important</b>: The example below is an example of a minimal amount of the features of the model configuration parameters and will not create the most accurate model possible. Additional options are available for tuning this configuration to produce an optimal model are described here: <a href="https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-data-export-parameters.html">Neptune Export Process Parameters</a></div>
#
# Running the cell below we set the export configuration and run the export process. Neptune export is capable of automatically creating a clone of the cluster by setting `cloneCluster=True` which takes about 20 minutes to complete and will incur additional costs while the cloned cluster is running. Exporting from the existing cluster takes about 5 minutes but requires that the `neptune_query_timeout` parameter in the [parameter group](https://docs.aws.amazon.com/neptune/latest/userguide/parameters.html) is set to a large enough value (>72000) to prevent timeout errors.
# +
export_params={
"command": "export-rdf",
"params": {
"endpoint": neptune_ml.get_host(),
"useIamAuth": neptune_ml.get_iam(),
"cloneCluster": False,
"profile": "neptune_ml"
},
"outputS3Path": f'{s3_bucket_uri}/neptune-export',
"additionalParams": {
"neptune_ml": {
"version": "v2.0",
"targets": [
{
"node" : "http://aws.amazon.com/neptune/ontology/Movie",
"predicate" : "http://aws.amazon.com/neptune/ontology/hasGenre",
"type": "classification",
"split_rate": [0.7,0.1,0.2]
}
]
}
},
"jobSize": "medium"}
# -
# %%neptune_ml export start --export-url {neptune_ml.get_export_service_host()} --export-iam --wait --store-to export_results
${export_params}
# # ML data processing, model training, and endpoint creation
#
# Once the export job is completed we are now ready to train our machine learning model and create the inference endpoint. Training our Neptune ML model requires three steps.
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: The cells below only configure a minimal set of parameters required to run a model training.</div>
#
# ## Data processing
# The first step (data processing) processes the exported graph dataset using standard feature preprocessing techniques to prepare it for use by DGL. This step performs functions such as feature normalization for numeric data and encoding text features using word2vec. At the conclusion of this step the dataset is formatted for model training.
#
# This step is implemented using a SageMaker Processing Job and data artifacts are stored in a pre-specified S3 location once the job is complete.
#
# Additional options and configuration parameters for the data processing job can be found using the links below:
#
# * [Data Processing](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-processing.html)
# * [dataprocessing command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-dataprocessing.html)
#
# Run the cells below to create the data processing configuration and to begin the processing job.
# +
# The training_job_name can be set to a unique value below, otherwise one will be auto generated
training_job_name=neptune_ml.get_training_job_name('node-classification')
processing_params = f"""
--config-file-name training-data-configuration.json
--job-id {training_job_name}
--s3-input-uri {export_results['outputS3Uri']}
--s3-processed-uri {str(s3_bucket_uri)}/preloading """
# -
# %neptune_ml dataprocessing start --wait --store-to processing_results {processing_params}
# ## Model training
# The second step (model training) trains the ML model that will be used for predictions. The model training is done in two stages. The first stage uses a SageMaker Processing job to generate a model training strategy. A model training strategy is a configuration set that specifies what type of model and model hyperparameter ranges will be used for the model training. Once the first stage is complete, the SageMaker Processing job launches a SageMaker Hyperparameter tuning job. The SageMaker Hyperparameter tuning job runs a pre-specified number of model training job trials on the processed data, and stores the model artifacts generated by the training in the output S3 location. Once all the training jobs are complete, the Hyperparameter tuning job also notes the training job that produced the best performing model.
#
# Additional options and configuration parameters for the data processing job can be found using the links below:
#
# * [Model Training](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-model-training.html)
# * [modeltraining command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-modeltraining.html)
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Information</b>: The model training process takes ~20 minutes</div>
training_params=f"""
--job-id {training_job_name}
--data-processing-id {training_job_name}
--instance-type ml.p3.2xlarge
--s3-output-uri {str(s3_bucket_uri)}/training """
# %neptune_ml training start --wait --store-to training_results {training_params}
# ## Endpoint creation
#
# The final step is to create the inference endpoint which is an Amazon SageMaker endpoint instance that is launched with the model artifacts produced by the best training job. This endpoint will be used by our graph queries to return the model predictions for the inputs in the request. The endpoint once created stays active until it is manually deleted. Each model is tied to a single endpoint.
#
# Additional options and configuration parameters for the data processing job can be found using the links below:
#
# * [Inference Endpoint](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-on-graphs-inference-endpoint.html)
# * [Endpoint command](https://docs.aws.amazon.com/neptune/latest/userguide/machine-learning-api-endpoints.html)
#
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Information</b>: The endpoint creation process takes ~5-10 minutes</div>
# +
endpoint_params=f"""
--id {training_job_name}-2
--model-training-job-id {training_job_name} """
print(endpoint_params)
# -
# %neptune_ml endpoint create --wait --store-to endpoint_results {endpoint_params}
# Once this has completed we get the endpoint name for our newly created inference endpoint. The cell below will set the endpoint name which will be used in the SPARQL queries below.
endpoint=endpoint_results['endpoint']['name']
neptune_role=neptune_ml.get_neptune_ml_role()
# # Predicting values using SPARQL queries
#
# Now that we have our inference endpoint setup let's query our product knowledge graph to show how to predict categorical property values by predicting the the genre for movies, starting with `Empire Strikes Back, The (1980)`.
#
# ## Predicting movie genres
#
# <br>
# <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Object Classification</b> - This model is trained to allow for inferring the `ontology:genre` string literal value of an instance of the `ontology:Movie` rdf:Class.
#
# Example `resource:movie_225` `ontology:genre` `"[to be predicted]"` .
#
# In order to make our prediction, we need to remove some data.
# Run the following query to remove the Genre's for three Movie's:
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
DELETE {
# ?movie ontology:hasGenre ?genre
}
WHERE {
# ?movie ontology:hasGenre ?genre ;
ontology:title ?title .
FILTER (?title IN (
"Empire Strikes Back, The (1980)",
"Mars Attacks! (1996)",
"Independence Day (ID4) (1996)"
))
}
# -
# Now that we have run the above query, three movies do not have any Genre's classified against them.
#
# Run the following query to verify that the Genre's in question were removed:
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
SELECT ?title ?genre WHERE {
# ?movie a ontology:Movie ;
ontology:title ?title.
OPTIONAL {
# ?movie ontology:hasGenre ?genre .
}
FILTER (?title IN (
"Empire Strikes Back, The (1980)",
"Mars Attacks! (1996)",
"Independence Day (ID4) (1996)"
))
}
# -
# As expected there is no Genre in the result set, so let's modify this query to predict the genre for `Empire Strikes Back, The (1980)`. To accomplish this we need to add more statements to our query:
#
# 1. Include a `SERVICE` enclosure to federate the Query to our ML endpoint. To do this you must use a special pre-defined URI which we have designated to use for ML calls:
# `SERVICE <http://aws.amazon.com/neptune/vocab/v01/services/ml#inference> {...} `
#
#
# 2. Add the configuration statements needed to connect to the ML endpoint you have created and specify the type of prediction.
#
# * ` ... ml:config ml:endpoint "{endpoint}" ...` - The ML endpoint that has been created. <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Note</b>: The endpoint values are automatically passed into the queries below</div>
#
# * ` ...ml:config ml:modelType 'OBJECT_CLASSIFICATION' ` - The inference type being used in this prediction. As we are predicting the OBJECT of the `'SUBJECT - PREDICATE - OBJECT'` standard RDF model.
#
#
# 3. Build the model of the RDF statement you are predicting: Add the following statements to the body of the SERVICE enclosure, so that you can build the SUBJECT, PREDICATE and OBJECT of your prediction.
#
# * ` ...ml:config ml:input ?input ` - The input of the prediction is the `'rdf:Subject'` of the predicted full statement using the `'SUBJECT - PREDICATE - OBJECT'` standard RDF model.
#
# * ` ...ml:config ml:predicate ontology:hasGenre ` - The predicate of the prediction is the `'rdf:Predicate'` of the predicted full statement using the `'SUBJECT - PREDICATE - OBJECT'` standard RDF model.
#
# * ` ...ml:config ml:output ?output ` - The output of the prediction is the `'rdf:Object'` of the predicted full statement using the `'SUBJECT - PREDICATE - OBJECT'` standard RDF model.
#
#
# 4. <div style="background-color:#eeeeee; padding:10px; text-align:left; border-radius:10px; margin-top:10px; margin-bottom:10px; "><b>Add optional stataments</b>:</div>
# * ` ...ml:config ml:score ?score ` - The calculated confidence of the prediction from DGL (Deep Graph Library).
#
# * ` ...ml:config ml:limit "3" ` - The number of predictions to return from DGL (Deep Graph Library).
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
SELECT ?title ?genre ?score WHERE {
BIND (resource:movie_172 as ?movie)
# ?movie a ontology:Movie ;
ontology:title ?title.
SERVICE ml:inference {
ml:config ml:endpoint "${endpoint}" ;
# the optional 'score' predicate
ml:outputScore ?score ;
# the optional 'limit' predicate
ml:limit "1" ;
ml:modelType 'OBJECT_CLASSIFICATION' ;
ml:input ?movie ;
ml:predicate ontology:hasGenre ;
ml:output ?genre ;
}
}
# -
# Looking at the results, this prediction seems right for the top result, `Empire Strikes Back, The (1980)` seems to fit well into the `Drama` genre, but the other predictions do not seem correct, so we can set a threshold for the score:
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
SELECT ?title ?genre ?score WHERE {
BIND (resource:movie_172 as ?movie)
# ?movie a ontology:Movie ;
ontology:title ?title.
OPTIONAL {
# ?movie ontology:hasGenre ?genre .
}
SERVICE ml:inference {
ml:config ml:endpoint "${endpoint}" ;
ml:outputScore ?score ;
ml:threshold "0.5D" ;
ml:modelType 'OBJECT_CLASSIFICATION' ;
ml:limit "3" ;
ml:input ?movie ;
ml:predicate ontology:hasGenre ;
ml:output ?genre ;
}
}
# -
# The results that do not meet the threshold are removed.
#
# As we see, depending on the threshold value specified the results where the confidence does not meet that threshold are removed.
#
# So far we have only been working on a single movie, but let's say you wanted to predict the genre for all movies without a `genre`.
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
SELECT ?title ?score ?predictedGenre WHERE {
# ?movie a ontology:Movie ;
ontology:title ?title.
MINUS {
# ?movie ontology:hasGenre ?existingGenre .
}
SERVICE ml:inference {
ml:config ml:endpoint "${endpoint}" ;
ml:outputScore ?score ;
ml:threshold "0.5D" ;
ml:modelType 'OBJECT_CLASSIFICATION' ;
ml:limit "3" ;
ml:input ?movie ;
ml:predicate ontology:hasGenre ;
ml:output ?predictedGenre .
}
}
# -
# We have applied the filters shown above to this query to find the top 3 genres for all our "new" movies, as long as the prediction has >70% confidence.
# Now that we've seen how to use Neptune ML to predict these categories, the next question is, how good are the predictions.
#
# ## Comparing the accuracy of predicted and actual genres
# Run the query below to see the predicted versus actual genres for one of our movies.
# +
# %%sparql
PREFIX ml: <http://aws.amazon.com/neptune/vocab/v01/services/ml#>
PREFIX ontology: <http://aws.amazon.com/neptune/ontology/>
PREFIX resource: <http://aws.amazon.com/neptune/resource#>
SELECT ?title ?existingGenre ?predictedGenre ?score WHERE {
BIND('Boxing Helena (1993)' as ?title)
# ?movie a ontology:Movie ;
ontology:title ?title ;
ontology:hasOriginalGenre ?existingGenre .
SERVICE ml:inference {
ml:config ml:endpoint "${endpoint}" ;
ml:outputScore ?score ;
ml:limit 3 ;
ml:modelType 'OBJECT_CLASSIFICATION' ;
ml:input ?movie ;
ml:predicate ontology:hasGenre ;
ml:output ?predictedGenre .
}
FILTER (?predictedGenre = ?existingGenre)
} limit 10
# -
# Comparing the `existingGenre` versus the `predictedGenre` results we see that our model did a good job of predicting the genres for this Movie.
# # Cleaning up
# Now that you have completed this walkthrough you have created a Sagemaker endpoint which is currently running and will incur the standard charges. If you are done trying out Neptune ML and would like to avoid these recurring costs, run the cell below to delete the inference endpoint.
#
neptune_ml.delete_endpoint(training_job_name)
print(training_job_name)
# In addition to the inference endpoint the CloudFormation script that you used has setup several additional resources. If you are finished then we suggest you delete the CloudFormation stack to avoid any recurring charges. For instructions, see Deleting a Stack on the [Deleting a Stack on the Amazon Web Services CloudFormation Console](https://docs.aws.amazon.com/AWSCloudFormation/latest/UserGuide/cfn-console-delete-stack.html). Be sure to delete the root stack (the stack you created earlier). Deleting the root stack deletes any nested stacks.
|
src/graph_notebook/notebooks/04-Machine-Learning/Neptune-ML-SPARQL/Neptune-ML-01-Introduction-to-Object-Classification-SPARQL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gap Framework - Computer Vision
#
# In this session, we will introduce you to preprocessing image data for computer vision. Preprocessing, storage, retrieval and batch management are all handled by two classes, the <b style='color:saddlebrown'>Image</b> and <b style='color:saddlebrown'>Images</b> class.
#
# Image - represents a single preprocessed image
# Images - represents a collection (or batch) of preprocessed images
# Let's go the directory of the Gap Framework
import os
os.chdir("../")
# %ls
# ### Setup
#
# Let's start by importing the Gap <b style='color:saddlebrown'>vision</b> module.
# import the Gap Vision module
from gapml.vision import Image, Images
# Let's go to a respository of images for sign language. We will use this repository for image preprocessing for computer vision.
# +
from urllib import request
import zipfile
# NOTE: This dataset can be downloaded from www.labs.earth/datasets/sign-lang.zip
os.makedirs("../Training/AITraining/Intermediate/Machine Learning/sign-lang", exist_ok=True)
os.chdir("../Training/AITraining/Intermediate/Machine Learning/sign-lang")
if not os.path.isdir('gestures'):
url = 'http://www.labs.earth/datasets/sign-lang.zip'
request.urlretrieve(url, 'sign-lang.zip')
#unzip file
with zipfile.ZipFile('sign-lang.zip', 'r') as zip_ref:
zip_ref.extractall('./')
# delete file
os.remove('sign-lang.zip')
print('done!')
# -
# The sign language characters (a-z) are labeled 1 .. 26, and 0 is for not a character.
# Each of the training images are under a subdirectory of the corresponding label.
labels = os.listdir("gestures")
print(labels)
# ### Image Class
#
# The <b style='color:saddlebrown'>Image</b> class supports the preprocessing of a single image into machine learning ready data. It can process JPG, PNG, TIF and GIF files. We will start by instantiating an <b style='color:saddlebrown'>Image</b> object for an image in the sign-lang image collection. For parameters, we will give the path to the image, and the label value (1). Labels must be mapped into integer values. For the sign-lang dataset, 1-26 is mapped to the 26 letters of the alphabet, and 0 is for a non-letter.
#
# When we instantiate the image, by default the following will happen:
#
# 1. The image is read in and decompressed, as a numpy array.
# 2. The image is processed according to the configuration parameters or defaults (e.g., resize, normalized, flatten,
# channel conversion).
# 3. The raw image data, processed image data, thumbnail, and metadata are stored to a HDF5 file system.
image = Image('gestures/1/1.jpg', 1)
# ### Image Properties
#
# Let's look at some properties of the <b style='color:saddlebrown'>Image</b> class.
#
# Note how the shape of the ML ready data is (50, 50, 3). We will change that in a bit.
print( image.name ) # The root name of the image (w/o suffix)
print( image.type ) # Type of image (e.g. jpeg)
print( image.dir ) # The directory where the ML ready data will be stored
print( image.size ) # The original size of the image
print( image.shape ) # The shape of the preprocessed image (ML ready data)
print( image.label ) # The label
print( image.time ) # The amount of time (secs) to preprocess the image
# ### Image Data
#
# Let's look at both the raw and ML ready data.
print("Raw Data", image.raw)
print("ML ready data", image.data)
# By default, the number of channels is preserved (e.g., 1 for grayscale, 3 for RGB, and 4 for RGBA), and the data is normalized. On the later, the 0..255 pixel values are rescaled between 0 and 1.
#
# Let's now change the preprocessing of the image to a grayscale image and resize it to 32x32. When we print the shape, you can see the 3rd dimension (channels) is gone - indicating a grayscale image, and the size is now 32 by 32.
# +
image = Image('gestures/1/1.jpg', 1, config=['grayscale', 'resize=(32,32)'])
print( image.shape )
# -
# Let's now say that the image data will be feed into a ANN (not CNN) or to a CNN with a 1D input vector. In this case, we need to feed the ML ready data as a flatten 1D vector. We can do that to. Now when we print the shape you can see its 1024 (32 x 32).
# +
image = Image('gestures/1/1.jpg', 1, config=['grayscale', 'resize=(32,32)', 'flatten', 'raw'])
print( image.shape )
# -
# ### Image Loading
#
# When an image is preprocessed, the ML ready data, raw data and attributes are stored in an HDF5 file. We can subsequently recall (load) the image information from the HDF5 file into an <b style='color:saddlebrown'>Image</b> object.
image = Image() # Create an empty image object
image.load('1.h5')
# Let's see if we get the same properties again.
print( image.name ) # The root name of the image (w/o suffix)
print( image.type ) # Type of image (e.g. jpeg)
print( image.dir ) # The directory where the ML ready data will be stored
print( image.size ) # The original size of the image
print( image.shape ) # The shape of the preprocessed image (ML ready data)
print( image.label ) # The label
# Let's check that we get the raw and ML ready data again.
print("Raw Data", image.raw)
print("ML ready data", image.data)
# ### Thumbnails
#
# We can also generate a store a thumbnail of the original image with the *config* parameter thumb. In the example below, we create a thumbnail with size 16x16
image = Image('gestures/1/1.jpg', 1, config=['grayscale', 'thumb=(16,16)'])
print(image.thumb)
print("Thumb Shape", image.thumb.shape)
# ### Aysnchronous Preprocess
#
# The image data can also be preprocessed asynchronously. In this mode, the parameter *ehandler* is set to an event handler (function) that will be called when the image is done being preprocessed. The image object is passed as a parameter to the event handler.
# +
def func(image):
print("DONE", image.name)
image = Image('gestures/1/1.jpg', 1, ehandler=func)
# -
# Let's cleanup and remove the HDF5 file
os.remove("1.h5")
# ### Remote Image (Url)
#
# The <b style='color:saddlebrown'>Image</b> class (and correspondly the <b style='color:saddlebrown'>Images</b> class), paths to the image file may be specified as an URL; providing the ability to preprocess images stored at remote locations. In this case, an HTTP request is made to retrieve the image data over the network.
# Let's load an image from the CNN news website
image = Image('https://cdn.cnn.com/cnnnext/dam/assets/180727161452-trump-speech-economy-072718-exlarge-tease.jpg', 2)
# Let's look at some properties.
# Let's display some properties of the image that was fetched from a remote location and then preprocessed in ML ready data.
print(image.name)
print(image.size)
print(image.shape)
# ### Raw Image (Pixel)
#
# The <b style='color:saddlebrown'>Image</b> class (and correspondly the <b style='color:saddlebrown'>Images</b> class), paths to the image file may alternatively be the raw pixel input; providing the ability to preprocess images without retreiving from storage, when they are otherwise already in memory.
# +
# import the openCV module
import cv2
# Read the pixel data into memory for an image using openCV
raw = cv2.imread('gestures/1/1.jpg')
# Let's load the image from directly the raw pixel data in memory
image = Image(raw, 1)
# -
# Let's look at some properties.
#
# Note, since this is raw pixel data, the image has no name, and the size is the decompressed (raw) size in memory.
# Let's display some properties of the image that was directly loaded from raw pixel data and then preprocessed in ML ready data.
print(image.name)
print(image.size)
print(image.shape)
# ### Image Augmentation
#
# Image Augmentation is the process of generating (synthesizing) new images from existing images, which can then be used to augment the training process. Augmentation can include, rotation, skew, sharpending and blur of existing images. These new images are then feed into the neural network during training to augment the training set. Rotating and skew aid in recognizing images from different angles, and sharpening and blur help generalize recognition (combat overfitting), as well as recognition under different lightening and time of day conditions.
#
# The <b style='color:saddlebrown'>Image</b> class supports generating new images by rotation. Any degree of rotation can be specified from 0 to 360.
# +
# Let's now rotate it 90 degress
rotated = image.rotate(90)
# Let's now look at the rotated image
cv2.imshow('image',rotated)
cv2.waitKey(0)
# -
# ## Images Class
#
# The <b style='color:saddlebrown'>Images</b> class supports the preprocessing of a collection of images into machine learning ready data. For required parameters, the <b style='color:saddlebrown'>Images</b> class takes a list of images and either a list of corresponding labels, or a single value, where all the images share the same label.
#
# Let's start by creating an <b style='color:saddlebrown'>Images</b> object for all the images under the subfolder 1 (letter A).
# +
# Let's get a list of all the images in the subfolder for the label 1 (letter A)
imgdir = "gestures/1/"
imglst = [imgdir + x for x in os.listdir(imgdir)]
# There should be 1200 images
len(imglst)
# -
# Let's now create an <b style='color:saddlebrown'>Images</b> object and preprocess all the above images.
# 1. Process all 1200 images in the subfolder 1
# 2. Set the label to 1
# 3. Convert them to grayscale.
#
# By default, the image data will be stored in an HDF5 file with the name 'collection.1.h5'.
# +
# Preprocess the set of images
images = Images(imglst, 1, config=['grayscale'])
# Check that the image data is stored in HDF5 file.
os.path.exists("collection.1.h5")
# -
# ### Images Properties
#
# Next, we will show some properties of the <b style='color:saddlebrown'>Images</b> class.
#
# Note, how fast it was to preprocess the set of 1200 images into machine ready data and store them in an HDF5 file.
print( images.name ) # The name of the collection of images
print( images.dir ) # where the ML ready data is stored
print( images.time ) # The length of time to preprocess the collection of images
print( len(images) ) # The len() operator is overridden to return the number of images in the collection
# Let's print the vector of labels
print("LABELS", images.labels)
# The <b style='color:saddlebrown'>Image</b> objects for each corresponding image can be accessed using the [] index operator. Let's get the 33rd one.
# The third Image object
image = images[32]
print(type(image))
print("Name", image.name)
# ### Directories (Subfolders) of Images
#
# The <b style='color:saddlebrown'>Images</b> class can alternately take a list of subfolders (vs. list of images); in which case, all the images under each subfolder are preprocessed into ML ready data. This is useful if your images are separated into subfolders, where each subfolder is a separate class (label) of images. This is a fairly common practice.
#
# In this case, the corresponding label in the same index of the labels parameter will be assigned to each image in the subfolder.
#
# In the example below, we also use the *name* parameter to specify a name (vs. default) for the collection.
# Let's process a list of subfolders of images, and name the collection 'foobar'
images = Images(['gestures/1', 'gestures/2'], [1,2], name='foobar')
# +
# We have two subfolders of 1200 images each, so we should expect 2400 images
print(len(images))
# cleanup
os.remove('foobar.h5')
# -
# ### Aysnchronous Preprocess
#
# A collection of images can also be preprocessed asynchronously. In this mode, the parameter *ehandler* is set to an event handler (function) that will be called when the collection of images is done being preprocessed. The <b style='color:saddlebrown'>Images</b> object is passed as a parameter to the event handler.
# +
def func(images):
print("DONE", images.name, "TIME", images.time)
images = Images(imglst, 1, config=['grayscale'], ehandler=func)
# -
# ### Assemblying a Collection
#
# For performance purposes, one may decide to process subparts of a collection asynchronosly, and then assemble them together into a single collection. The Images class provides support for this assembling subcollections into a single collection using the overridden *+=* operator, to merge other preprocessed collections into a single collection.
#
# In the example below, we first create a collection for the letter 'A' images (label 1) and then separately create a second collection for the letter 'B' images (label 2), and use the *+=* operator to merge the second collection into the first collection.
# +
# Create collection for the letter 'A' images
images = Images(['gestures/1'], 1)
# Create collection for the letter 'B' images and merge it with the letter 'A' image collection
images += Images(['gestures/2'], 2)
# The merged collection should have 2400 images (1200 for 'A', and 1200 for 'B')
print(len(images))
# -
# ### Splitting a Collection into Training and Test Data
#
# The *split* property will split the <b style='color:saddlebrown'>Image</b> objects into training and test data. The list of training image objects is then randomized. When used as a setter, the property takes either 1 or 2 arguments. The first argument is the percentage that is test data, and the optional second argument is the seed for the random shuffle.
# +
# Split the image objects into 80% training and 20% test
images.split = 0.20
# Let's verify that the training set is 80% (960 of 1200) by printing the internal variable _train
print(len(images._train))
# Let's now print the randomized list of image object indices
print("TRAIN INDICES", images._train)
# -
# Let's now add the optional parameter for a random seed.
# +
# Split the image objects into 80% training and 20% test
images.split = 0.20, 42
# Let's now print the randomized list of image object indices
print("TRAIN INDICES", images._train)
# -
# ### Batch Feeding (Batch Gradient Descent)
#
# There are three ways to use the <b style='color:saddlebrown'>Images</b> object to feed a neural network. In batch mode, the entire training set can be ran through the neural network as a single pass, prior to backward probagation and updating the weights using gradient descent. This is known as 'batch gradient descent'.
#
# When the *split* property is used as a getter, it returns the image data and corresponding labels for the training and test set similar to using sci-learn's train_test_split() function.
# +
# Set the percentage and seed, and split the data
images.split = 0.20, 42
# Get the training, test sets and corresponding labels
x_train, x_test, y_train, y_test = images.split
# -
# Let's verify and print the len of the train, test and corresponding labels.
print("x_train", len(x_train))
print("y_train", len(y_train))
print("x_test", len(x_test))
print("y_test", len(y_test))
# Let's verify the contents that the elements are what we expect.
# Each element in x_train list should be a numpy array
print(type(x_train[0]))
# Each element should be in the shape 50 x 50 pixels
print(x_train[0].shape)
# Each elment in y_train should be the label (integer)
print(type(y_train[0]))
# ### Next Iterating (Stochastic Gradient Descent)
#
# Another way of feeding a neural network is to feed one image at a time and do backward probagation, using gradient descent. This is known as stochastic gradient descent.
#
# The *next()* operator supports iterating through the training list one image object at a time. Once all of the entire training set has been iterated through, it is reset and the training set is randomly re-shuffled for the next epoch.
# Let's iterate through the ML ready data and label for each image in the training set
while True:
data, label = next(images)
if data is None: break
print(type(data), label)
# ### Mini-batch generation
#
# Another way of feeding a neural network is through mini-batches. A mini-batch is a subset of the training set, that is greater than one. After each mini-batch is feed, then backward probagation, using gradient descent, is done.
#
# Typically, minibatches are set to sizes like 30, 50, 100, or 200. We will use the *minibatch* property as a setter to set the mini-batch size to 100.
# Set minibatch size to 100 images
images.minibatch = 100
# When we use the *minibatch* property as a getter, it will create a generator.
# +
# Calculate the number of batches
nbatches = len(images) // 100
# process each mini-batch
for _ in range(nbatches):
# Create a generator for the next minibatch
g = images.minibatch
# Get the data, labels for each item in the minibatch
for data, label in g:
pass
# -
# ## Datset of Images
#
# Let's load the entire dataset - that's 27 collections of 1200 images each.
# +
# Prepare each set of labeled data into machine learning ready data
# The images are 50x50, bitdepth=8, 1 channel (grayscale)
total = 0
collections=[]
for label in labels:
# Get a list of all images in the subdirectory for this label (should be 1200 images)
imgdir = "gestures/" + label + "/"
imglst = [imgdir + x for x in os.listdir(imgdir)]
images = Images(imglst, int(label), name='tmp' + label, config=['flatten', 'grayscale', 'raw'])
collections.append(images)
print("Procesed: " + label, "Number of images:", len(images), "Time: ", images.time)
total += images.time
print("average:", total / len(labels))
# -
# Let's verify the preprocessing of our image data
# +
# Let's see how many batches (collections) we have (hint: should be 27)
print(len(collections))
# Let's verify that the items in the collections are an Images object
collection = collections[3]
print(type(collection))
# For a collection, let's see how many image objects we have (hint: should be 1200)
print(len(collection))
# -
# Let's look at the first Image object in this collection.
# Let's get the first Image item and verify it is an Image object
image = collection[0]
print(type(image))
# Let's name view some of the properties and verify that images got processed as expected.
# Let's get some basic information about the image
print(image.name) # the root name of the image
print(image.type) # the image file suffix
print(image.size) # the size of the image on disk
# Let's now check the raw (uncompressed) unprocessed image
print(image.raw.shape)
# Let's look at how the image got processed.
print(image.shape) # Note, that the preprocessed image was flattened into a 1D vector. It was 50x50, and now is 2500.
# Let's now take a look at the image. Remember to hit any key to exit the viewer (i.e., cv2.waitKey(0))
# Let's view the raw image
import cv2
cv2.imshow('image',image.raw)
cv2.waitKey(0)
# # END OF SESSION 3
# some cleanup
os.remove('collection.1.h5')
os.remove('untitled.h5')
os.remove('180727161452-trump-speech-economy-072718-exlarge-tease.h5')
for _ in range(27):
os.remove('tmp' + str(_) + '.h5')
|
train/session 3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: aoc
# language: python
# name: conda-env-aoc-py
# ---
# # Day 15
# +
class Actor:
def __init__(self, grid, x, y):
self.grid = grid
self.x = x
self.y = y
def __str__(self):
return '{0}({1.x}, {1.y})'.format(type(self).__name__, self)
@staticmethod
def fromcode(code, grid, x, y):
if code == 'E':
return Elf(grid, x, y)
elif code == 'G':
return Goblin(grid, x, y)
def go(self):
pass
def move(self):
pass
@property
def reachable(self):
todo = [(self.x + 1, self.y), (self.x - 1, self.y), (self.x, self.y + 1), (self.x, self.y - 1)]
visited = set()
while todo:
pt = todo.pop()
if pt in visited:
continue
visited.add(pt)
x, y = pt
if self.grid[y][x] == '.':
todo.extend([(x + 1, y), (x - 1, y), (x, y + 1), (x, y - 1)])
yield pt
__repr__ = __str__
class Goblin(Actor):
pass
class Elf(Actor):
pass
# -
def parse(f):
grid = [list(line.rstrip()) for line in f]
actors = []
for y, row in enumerate(grid):
for x, block in enumerate(row):
if block in ('E', 'G'):
actors.append(Actor.fromcode(block, grid, x, y))
return actors, grid
from io import StringIO
f = StringIO('''#######
#E..G.#
#...#.#
#.G.#G#
#######''')
actors, grid = parse(f)
actors
set(actors[0].reachable)
|
2018/Advent.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TDA with Python using the Gudhi Library
#
# # Building simplicial complexes from distance matrices
# **Authors :** <NAME> and <NAME>
import numpy as np
import pandas as pd
import pickle as pickle
import gudhi as gd
from pylab import *
# %matplotlib inline
# TDA typically aims at extracting topological signatures from a point cloud in $\mathbb R^d$ or in a general metric space. [Simplicial complexes](https://en.wikipedia.org/wiki/Simplicial_complex) are used in computational geometry to infer topological signatures from data.
#
# This tutorial explains how to build [Vietoris-Rips complexes](https://en.wikipedia.org/wiki/Vietoris%E2%80%93Rips_complex) and [alpha complexes](https://en.wikipedia.org/wiki/Alpha_shape#Alpha_complex) from a matrix of pairwise distances.
#
# ## Vietoris-Rips filtration defined from a matrix of distances
#
# The [$\alpha$-Rips complex](https://en.wikipedia.org/wiki/Vietoris%E2%80%93Rips_complex) of a metric space $\mathbb X$ is an [abstract simplicial complex](https://en.wikipedia.org/wiki/Abstract_simplicial_complex) that can be defined by forming a simplex for every finite subset of $\mathbb X$ that has diameter at most $\alpha$.
#
# 
#
#
# ## Protein binding dataset
#
# The data we will be studying in this notebook represents configurations of protein binding. This example is borrowed from Kovacev-Nikolic et al. [[1]](https://arxiv.org/pdf/1412.1394.pdf).
#
# The paper compares closed and open forms of the maltose-binding protein (MBP), a large biomolecule containing $370$ amino-acid residues. The analysis is not based on geometric distances in $\mathbb R^3$ but on a metric of *dynamical distances* defined by
#
# $$ D_{ij} = 1 - |C_{ij}|, $$
#
# where $C$ is the correlation matrix between residues. Correlation matrices between residues can be found at this [link](https://www.researchgate.net/publication/301543862_corr). We are greatful to the authors for sharing data !
# The next statments load the $14$ correlation matrices with pandas:
path_file = "./datasets/Corr_ProteinBinding/"
files_list = [
'1anf.corr_1.txt',
'1ez9.corr_1.txt',
'1fqa.corr_2.txt',
'1fqb.corr_3.txt',
'1fqc.corr_2.txt',
'1fqd.corr_3.txt',
'1jw4.corr_4.txt',
'1jw5.corr_5.txt',
'1lls.corr_6.txt',
'1mpd.corr_4.txt',
'1omp.corr_7.txt',
'3hpi.corr_5.txt',
'3mbp.corr_6.txt',
'4mbp.corr_7.txt'
]
corr_list = [
pd.read_csv(
path_file + u,
header = None,
delim_whitespace = True
) for u in files_list
]
# The object `corr_list` is a list of $14$ correlation matrices. We can iterate in the list to compute the matrix of distances associated to each configuration:
dist_list = [1 - np.abs(c) for c in corr_list]
# Let's print out the first lines of the first distance matrix:
D = dist_list[0]
D.head()
# ## Vietoris-Rips filtration of Protein binding distance matrix
#
# The `RipsComplex()` function creates a [$1$-skeleton](https://en.wikipedia.org/wiki/N-skeleton) from the point cloud (see the [GUDHI documentation](https://gudhi.inria.fr/python/latest/rips_complex_user.html) for details on the syntax).
skeleton_protein = gd.RipsComplex(
distance_matrix = D.values,
max_edge_length = 0.8
)
# The `max_edge_length` parameter is the maximal diameter: only the edges of length less vers this value are included in the one skeleton graph.
#
# Next, we create the Rips simplicial complex from this one-skeleton graph. This is a filtered Rips complex which filtration function is exacly the diameter of the simplices. We use the `create_simplex_tree()` function:
Rips_simplex_tree_protein = skeleton_protein.create_simplex_tree(max_dimension = 2)
# The `max_dimension` parameter is the maximum dimension of the simplices included in the the filtration. The object returned by the function is a simplex tree, of dimension 2 in this example:
Rips_simplex_tree_protein.dimension()
# We can use the fonctionalites of the simplex tree object to describe the Rips filtration.
# For instance we can check that the 370 points of the first distance matrix are all vertices of the Vietoris-Rips filtration:
Rips_simplex_tree_protein.num_vertices()
# The number of simplices in a Rips complex increases very fast with the number of points and the dimension. There is more than on million of simplexes in the Rips complex:
Rips_simplex_tree_protein.num_simplices()
# Note that this is actually the number of simplices in the "last" Rips complex of the filtration, namely with parameter $\alpha=$ `max_edge_length=`0.8.
# Let's compute the list of simplices in the Rips complex with the `get_filtration()` function:
#
rips_filtration = Rips_simplex_tree_protein.get_filtration()
rips_list = list(rips_filtration)
len(rips_list)
for splx in rips_list[0:400] :
print(splx)
# The integers represent the points in the metric space: the vertex `[2]` corresponds to the point decribed by the second raw (or the second column) in the distance matrix `D`.
#
# The filtration value is the diameter of the simplex, which is zero for the vertices of course. The first edge in the filtration is `[289, 290]`, these two points are the two closest points according to `D`, at distance $0.015$ of each other.
# ### How to define an Alpha complex from a matrix of distance ?
# The [alpha complex filtration](https://en.wikipedia.org/wiki/Alpha_shape#Alpha_complex) of a point cloud in $\mathbb R^p$ is a filtered simplicial complex constructed from the finite cells of a [Delaunay Triangulation](https://en.wikipedia.org/wiki/Delaunay_triangulation).
#
# In our case, the data does not belong to euclideen space $\mathbb R^p$ and we are not in position to directly compute a Delaunay Triangulation in the metric space, using the pairwise distances.
#
# The aim of [Multidimensional Scaling](https://en.wikipedia.org/wiki/Multidimensional_scaling) (MDS) methods is precisely to find a representation of $n$ points in a space $\mathbb R^p$ that preserves as well as possible the pairwise distances between the $n$ points in the original metric space. The are several versions of MDS algorithms, the most popular ones are available in the [sckit-learn library](https://scikit-learn.org/stable/index.html), see this [documention](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.MDS.html).
#
# Let's compute a (classical) MDS representation of the matrix `D` in $\mathbb R^3$:
# +
from sklearn.manifold import MDS
embedding = MDS(n_components = 3, dissimilarity = 'precomputed')
X_transformed = embedding.fit_transform(D)
X_transformed.shape
# -
# Now we can represent this configuration, for instance on the two first axes:
fig = plt.figure()
plt.scatter(X_transformed[:, 0], X_transformed[:, 1], label = 'MDS');
# Of course you should keep in mind that MDS provides an embedding of the data in $\mathbb R^p$ that **approximatively** preserves the distance matrix.
# The main advantage of Alpha complexes is that they contain less simplices than Rips complexes do and so it can be a better option than Rips complexes. As subcomplexes of the Delaunay Triangulation complex, an alpha complex is a geometric simpicial complex.
#
# The `AlphaComplex()` function directly computes the simplex tree representing the Alpha complex:
alpha_complex = gd.AlphaComplex(points = X_transformed)
st_alpha = alpha_complex.create_simplex_tree()
# The point cloud `X_transformed` belongs to $\mathbb R^3$ and so does the Alpha Complex:
st_alpha.dimension()
# As for the Rips complex, the $370$ points are all vertices of the Alpha complex :
print(st_alpha.num_vertices())
# Note that the number of simplexes in the Alpha complex is much smaller then for the Rips complex we computed before:
#
print(st_alpha.num_simplices())
# ### References
# [1] Using persistent homology and dynamical distances to analyze protein binding, <NAME>, <NAME>, <NAME> and <NAME>. Stat Appl Genet Mol Biol 2016 [arxiv link](https://arxiv.org/pdf/1412.1394.pdf).
|
Tuto-GUDHI-simplicial-complexes-from-distance-matrix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="h8mk2niHSn1S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 292} outputId="a5d4c3cf-da68-4663-954b-7af05c3b2869" executionInfo={"status": "ok", "timestamp": 1583270767724, "user_tz": -60, "elapsed": 6856, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
# !pip install --upgrade tables
# + id="u3A-yQfkRcV8" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + id="0N6g75jHRuA7" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="bx7Y7rVDRrnC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4113acf-f438-47ba-8bb7-a23be7101113" executionInfo={"status": "ok", "timestamp": 1583270783645, "user_tz": -60, "elapsed": 639, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
# cd '/content/drive/My Drive/Colab Notebooks/matrix_two/dw_matrix_car'
# + id="0lR1pPBjRvsb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a1655738-2212-4d8b-9ee7-0f7132253fdf" executionInfo={"status": "ok", "timestamp": 1583270786726, "user_tz": -60, "elapsed": 2674, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
# ls
# + id="GomnK0R_SAO_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ae710f6d-e850-4264-e70b-8f70a58a7fdb" executionInfo={"status": "ok", "timestamp": 1583270806873, "user_tz": -60, "elapsed": 3054, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="ptgvzgMESNIj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="8ea1b588-1e2e-4c07-ce11-ba24557b062b" executionInfo={"status": "ok", "timestamp": 1583270816324, "user_tz": -60, "elapsed": 814, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df.columns.values
# + id="8dk7wJB4S0fV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="8ca16136-f33b-4edd-8816-6dd2afaae57a" executionInfo={"status": "ok", "timestamp": 1583270869389, "user_tz": -60, "elapsed": 1057, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df['price_value'].hist(bins=100);
# + id="V1fnLK0_S_0j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="325e43fe-d7dc-43be-861c-d3513927c5b3" executionInfo={"status": "ok", "timestamp": 1583270884903, "user_tz": -60, "elapsed": 868, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df['price_value'].describe()
# + id="UbXEuNHGTHI4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="5c3c7223-6a1a-4e9a-b14c-25d3e5520b60" executionInfo={"status": "ok", "timestamp": 1583270922989, "user_tz": -60, "elapsed": 811, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df['param_marka-pojazdu'].unique()
# + id="PB8f4epPTQdd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="c1f7c9dd-45cb-4ac7-ace1-8574b5b111b3" executionInfo={"status": "ok", "timestamp": 1583270990906, "user_tz": -60, "elapsed": 794, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
df.groupby('param_marka-pojazdu')['price_value'].mean()
# + id="FqAXfDW5ThC4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 401} outputId="94356be8-0889-4c58-f2f7-e297129775c5" executionInfo={"status": "ok", "timestamp": 1583271228458, "user_tz": -60, "elapsed": 1291, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
(
df.groupby('param_marka-pojazdu')['price_value']
.agg(np.mean)
.sort_values(ascending=False)
.head(50)
).plot(kind='bar', figsize=(15,5));
# + id="sCVMMfnRTr5l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 405} outputId="bbc8e34f-68a6-4c6a-eb31-bb54368ec48a" executionInfo={"status": "ok", "timestamp": 1583271312568, "user_tz": -60, "elapsed": 1987, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
(
df.groupby('param_marka-pojazdu')['price_value']
.agg([np.mean, np.median, np.size])
.sort_values(by='mean', ascending=False)
.head(50)
).plot(kind='bar', figsize=(15,5));
# + id="hpZaAKnXUbqM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="e0fc9f66-12d7-4fe2-9c7c-59b2ebf64373" executionInfo={"status": "ok", "timestamp": 1583271326619, "user_tz": -60, "elapsed": 2432, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
(
df.groupby('param_marka-pojazdu')['price_value']
.agg([np.mean, np.median, np.size])
.sort_values(by='mean', ascending=False)
.head(50)
).plot(kind='bar', figsize=(15,5), subplots=True);
# + id="CtNOECZhUym7" colab_type="code" colab={}
def group_and_barplot(feat_groupby, feat_agg='price_value', agg_funcs=[np.mean, np.median, np.size], feat_sort='mean', top=50, subplots=True):
return (
df
.groupby(feat_groupby)[feat_agg]
.agg(agg_funcs)
.sort_values(by=feat_sort, ascending=False)
.head(top)
).plot(kind='bar', figsize=(15,5), subplots=subplots);
# + id="gHK2QbndV4Vf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="d12fa41b-3cc8-4088-ce87-d4f9e8ee7615" executionInfo={"status": "ok", "timestamp": 1583271695246, "user_tz": -60, "elapsed": 2192, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
group_and_barplot('param_marka-pojazdu');
# + id="pHWOAU5ZWGpo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 436} outputId="36254d16-9b57-4763-a878-0551cf9f0a09" executionInfo={"status": "ok", "timestamp": 1583271744919, "user_tz": -60, "elapsed": 1240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
group_and_barplot('param_country-of-origin');
# + id="mZ28tv2NWZA9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 436} outputId="e8c1d54d-61d3-42f9-8c35-68621cc281ed" executionInfo={"status": "ok", "timestamp": 1583271826871, "user_tz": -60, "elapsed": 2140, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
group_and_barplot('param_kraj-pochodzenia', feat_sort='size');
# + id="0ZXClz4fWgs1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="00680fee-6387-41de-fbf8-0e91bf179191" executionInfo={"status": "ok", "timestamp": 1583271897354, "user_tz": -60, "elapsed": 1706, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg7kho4qS_0kFpuVKxuC36fMckRFKoUdbV5tMw2v74=s64", "userId": "15699635066276624141"}}
group_and_barplot('param_kolor', feat_sort='mean');
# + id="eELmTBPtW4k0" colab_type="code" colab={}
|
day2_visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2, os, json, math, time, tqdm
import pandas as pd
import numpy as np
from collections import Counter, OrderedDict, defaultdict
from zipfile import ZipFile
from pycocotools import mask as maskUtils
# ## Deepfashion2 to COCO
def df2coco(path_annos_image:str, num_images:int, output_dir:str):
"""Convert DF2 annotation format to COCO.
Args:
path_annos_image: (str) path of annos and image folders
num_images: (int) number of images == number of annos files
output_dir: (str) path where to save new json
:return: json file with COCO annotations
"""
dataset = {
"info": {},
"licenses": [],
"images": [],
"annotations": [],
"categories": []
}
### Categories
keypoints = [str(i) for i in range(1,295)]
categories = [[1, 'short_sleeved_shirt'],
[2, 'long_sleeved_shirt'],
[3, 'short_sleeved_outwear'],
[4, 'long_sleeved_outwear'],
[5, 'vest'],
[6, 'sling'],
[7, 'shorts'],
[8, 'trousers'],
[9, 'skirt'],
[10, 'short_sleeved_dress'],
[11, 'long_sleeved_dress'],
[12, 'vest_dress'],
[13, 'sling_dress']]
dataset['categories'] = [{'id': categories[i][0],
'name': categories[i][1],
'supercategory': 'clothes',
'keypoints': keypoints,
'skeleton': []} for i in range(len(categories))]
assert len(os.listdir(path_annos_image+'/image')) == len(os.listdir(path_annos_image+'/annos')) == num_images
### Images / Annotations
sub_index = 0 # the index of ground truth instance
for num in tqdm.tqdm(range(1, num_images+1)):
json_name = path_annos_image+'/annos/' + str(num).zfill(6)+'.json'
image_name = path_annos_image+'/image/' + str(num).zfill(6)+'.jpg'
if (num>=0):
shp = cv2.imread(image_name).shape
width, height = shp[1], shp[0]
temp = json.load(open(json_name))
pair_id = temp['pair_id']
source = temp['source']
dataset['images'].append({
'coco_url': '',
'date_captured': '',
'file_name': str(num).zfill(6) + '.jpg',
'flickr_url': '',
'id': num,
'license': 0,
'width': width,
'height': height,
'pair_id': pair_id,
'source': source})
for i in temp.keys(): # for each item
if i == 'source' or i=='pair_id':
continue
else:
points = np.zeros(294 * 3)
sub_index = sub_index + 1
box = temp[i]['bounding_box']
w = box[2]-box[0]
h = box[3]-box[1]
x_1 = box[0]
y_1 = box[1]
bbox = [x_1,y_1,w,h]
cat = temp[i]['category_id']
seg = temp[i]['segmentation']
style = temp[i]['style']
scale = temp[i]['scale']
occlusion = temp[i]['occlusion']
viewpoint = temp[i]['viewpoint']
zoom_in = temp[i]['zoom_in']
landmarks = temp[i]['landmarks']
points_x = landmarks[0::3]
points_y = landmarks[1::3]
points_v = landmarks[2::3]
points_x = np.array(points_x)
points_y = np.array(points_y)
points_v = np.array(points_v)
if cat == 1:
for n in range(0, 25):
points[3 * n] = points_x[n]
points[3 * n + 1] = points_y[n]
points[3 * n + 2] = points_v[n]
elif cat ==2:
for n in range(25, 58):
points[3 * n] = points_x[n - 25]
points[3 * n + 1] = points_y[n - 25]
points[3 * n + 2] = points_v[n - 25]
elif cat ==3:
for n in range(58, 89):
points[3 * n] = points_x[n - 58]
points[3 * n + 1] = points_y[n - 58]
points[3 * n + 2] = points_v[n - 58]
elif cat == 4:
for n in range(89, 128):
points[3 * n] = points_x[n - 89]
points[3 * n + 1] = points_y[n - 89]
points[3 * n + 2] = points_v[n - 89]
elif cat == 5:
for n in range(128, 143):
points[3 * n] = points_x[n - 128]
points[3 * n + 1] = points_y[n - 128]
points[3 * n + 2] = points_v[n - 128]
elif cat == 6:
for n in range(143, 158):
points[3 * n] = points_x[n - 143]
points[3 * n + 1] = points_y[n - 143]
points[3 * n + 2] = points_v[n - 143]
elif cat == 7:
for n in range(158, 168):
points[3 * n] = points_x[n - 158]
points[3 * n + 1] = points_y[n - 158]
points[3 * n + 2] = points_v[n - 158]
elif cat == 8:
for n in range(168, 182):
points[3 * n] = points_x[n - 168]
points[3 * n + 1] = points_y[n - 168]
points[3 * n + 2] = points_v[n - 168]
elif cat == 9:
for n in range(182, 190):
points[3 * n] = points_x[n - 182]
points[3 * n + 1] = points_y[n - 182]
points[3 * n + 2] = points_v[n - 182]
elif cat == 10:
for n in range(190, 219):
points[3 * n] = points_x[n - 190]
points[3 * n + 1] = points_y[n - 190]
points[3 * n + 2] = points_v[n - 190]
elif cat == 11:
for n in range(219, 256):
points[3 * n] = points_x[n - 219]
points[3 * n + 1] = points_y[n - 219]
points[3 * n + 2] = points_v[n - 219]
elif cat == 12:
for n in range(256, 275):
points[3 * n] = points_x[n - 256]
points[3 * n + 1] = points_y[n - 256]
points[3 * n + 2] = points_v[n - 256]
elif cat == 13:
for n in range(275, 294):
points[3 * n] = points_x[n - 275]
points[3 * n + 1] = points_y[n - 275]
points[3 * n + 2] = points_v[n - 275]
num_points = len(np.where(points_v > 0)[0])
dataset['annotations'].append({
'image_id': num,
'category_id': cat,
'id': sub_index,
'bbox': bbox,
'area': w*h,
'segmentation': seg,
'iscrowd': 0,
'num_keypoints':num_points,
'keypoints':points.tolist(),
'style': style,
'scale': scale,
'occlusion': occlusion,
'viewpoint': viewpoint,
'zoom_in': zoom_in
})
with open(output_dir, 'w') as f:
json.dump(dataset, f)
# +
DF2_DATA_DIR = '.../datasets/deepfashion2' #@@@ OVERRIDE: Path of folder with DF2 annotations
# CHECKS
# Check number of images = number of annotation files
images_names_tr = sorted(os.listdir(os.path.join(DF2_DATA_DIR, 'train/image')))
annos_names_tr = sorted(os.listdir(os.path.join(DF2_DATA_DIR, 'train/annos')))
assert len(images_names_tr) == len(annos_names_tr)
images_names_val = sorted(os.listdir(os.path.join(DF2_DATA_DIR, 'validation/image')))
annos_names_val = sorted(os.listdir(os.path.join(DF2_DATA_DIR, 'validation/annos')))
assert len(images_names_val) == len(annos_names_val)
# Check there is 1 and only 1 annotation file for each immage
for i in range(len(images_names_tr)):
assert annos_names_tr[i].split('.')[0] == images_names_tr[i].split('.')[0]
for i in range(len(images_names_val)):
assert annos_names_val[i].split('.')[0] == images_names_val[i].split('.')[0]
# CONVERT!
df2coco(os.path.join(DF2_DATA_DIR, 'train'), len(images_names_tr), 'df2_annos_tr.json')
df2coco(os.path.join(DF2_DATA_DIR, 'validation'), len(images_names_val), 'df2_annos_val.json')
# -
# ## VGG Image Annotator to COCO
def via2coco(via_json_path, ontology_path, img_path, source, output_dir):
"""Convert a json from VGG Image Annotator (VIA) format to COCO.
Args:
via_json_path: (str) path of VIA json file
ontology_path: (str) path of xlsx file with categories
source: (str) source of images (user/shop/IG/user_shop)
img_path: (str) path of folder with images
output_dir: (str) path where to save new json
:return: json file with COCO annotations
"""
via = json.load(open(via_json_path, encoding="utf-8"))
ont_df = pd.read_excel(ontology_path, sheet_name=0, engine="openpyxl")
coco = {
"info": {},
"licenses": [],
"images": [],
"annotations": [],
"categories": []
}
coco['info'] = {
"description": via['_via_settings']['project']['name'],
"version": "1.0",
"VIA_version": via['_via_data_format_version'],
"year": 2022,
"date_created": time.asctime(time.localtime(time.time()))
}
via_cats = via['_via_attributes']['region']['clothing']['options']
for k,v in tqdm.tqdm(via_cats.items()):
rows, cols = np.where(ont_df == v)
if ont_df.columns[cols[0]-1] == 'category':
cat = ont_df.iloc[rows[0], cols[0]-1]
sup = ont_df.iloc[rows[0], cols[0]-2]
else:
cat, sup = '', ont_df.iloc[rows[0], cols[0]-1]
c = {
'id': int(k),
'name': v.strip().replace(' ', '_').lower(),
'category': cat,
'supercategory': sup}
coco['categories'].append(c)
id = 0
for k,v in tqdm.tqdm(via['_via_img_metadata'].items()):
if 'clothing' in v['regions'][0]['region_attributes'].keys():
# Images
k = int(v['filename'][:-4])
assert int(k) == int(v['filename'][:-4])
shp = cv2.imread(os.path.join(img_path, v['filename'])).shape
img = {
"id": int(k),
"file_name": v['filename'],
"width": shp[1],
"height": shp[0],
"source" : source,
"people" : list(v['file_attributes']['people'])}
coco['images'].append(img)
# Annotations
for region in v['regions']:
cat_id = int(region['region_attributes']['clothing'])
assert cat_id in (d['id'] for d in coco['categories'])
points_x = region['shape_attributes']['all_points_x']
points_y = region['shape_attributes']['all_points_y']
seg = [] # There will be just 1 polygon for 1 instance in VIA format
for x, y in zip(points_x, points_y):
seg.append(x)
seg.append(y)
segmentation = [seg]
rles = maskUtils.frPyObjects(segmentation, shp[0], shp[1])
rle = maskUtils.merge(rles) # If a single object consist of multiple parts, merge all parts into 1 RLE
area = maskUtils.area(rle).astype(float)
bbox = list(maskUtils.toBbox(rle))
ann = {
"id": id,
"image_id": int(k),
"category_id": cat_id, # int
"iscrowd": 0,
"area": area, # float
"bbox": bbox, # [x,y,w,h]
"segmentation": segmentation # [x1,y1,x2,y2,...]
}
id += 1
coco['annotations'].append(ann)
with open(output_dir, 'w') as f:
json.dump(coco, f)
return coco
# +
#@@@ OVERRIDE
DIR_VIA_JSON = '.../via.json' # Path of json file with the VIA annotations
DIR_XLSX_ONTOLOGY = './tools/fashion_ontology.xlsx' # Path of excel file with fashion ontolgy
DIR_IMAGES = '.../images' # Path of folder with images (whose annotations have been saved in the VIA json file)
coco = via2coco(DIR_VIA_JSON, DIR_XLSX_ONTOLOGY, DIR_IMAGES, '', './coco.json')
|
converters.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from nb_005 import *
from collections import Counter
# # Wikitext 2
# ## Data
# Download the dataset [here](https://s3.amazonaws.com/research.metamind.io/wikitext/wikitext-2-v1.zip) and unzip it so it's in the folder wikitext.
EOS = '<eos>'
PATH=Path('data/wikitext')
# Small helper function to read the tokens.
def read_file(filename):
tokens = []
with open(PATH/filename, encoding='utf8') as f:
for line in f:
tokens.append(line.split() + [EOS])
return np.array(tokens)
train_tok = read_file('wiki.train.tokens')
valid_tok = read_file('wiki.valid.tokens')
test_tok = read_file('wiki.test.tokens')
len(train_tok), len(valid_tok), len(test_tok)
' '.join(train_tok[4][:20])
cnt = Counter(word for sent in train_tok for word in sent)
cnt.most_common(10)
# Give an id to each token and add the pad token (just in case we need it).
itos = [o for o,c in cnt.most_common()]
itos.insert(0,'<pad>')
vocab_size = len(itos); vocab_size
# Creates the mapping from token to id then numericalizing our datasets.
stoi = collections.defaultdict(lambda : 5, {w:i for i,w in enumerate(itos)})
train_ids = np.array([([stoi[w] for w in s]) for s in train_tok])
valid_ids = np.array([([stoi[w] for w in s]) for s in valid_tok])
test_ids = np.array([([stoi[w] for w in s]) for s in test_tok])
class LanguageModelLoader():
""" Returns a language model iterator that iterates through batches that are of length N(bptt,5)
The first batch returned is always bptt+25; the max possible width. This is done because of they way that pytorch
allocates cuda memory in order to prevent multiple buffers from being created as the batch width grows.
"""
def __init__(self, nums, bs, bptt, backwards=False):
self.bs,self.bptt,self.backwards = bs,bptt,backwards
self.data = self.batchify(nums)
self.i,self.iter = 0,0
self.n = len(self.data)
def __iter__(self):
self.i,self.iter = 0,0
while self.i < self.n-1 and self.iter<len(self):
if self.i == 0:
seq_len = self.bptt + 5 * 5
else:
bptt = self.bptt if np.random.random() < 0.95 else self.bptt / 2.
seq_len = max(5, int(np.random.normal(bptt, 5)))
res = self.get_batch(self.i, seq_len)
self.i += seq_len
self.iter += 1
yield res
def __len__(self): return self.n // self.bptt - 1
def batchify(self, data):
nb = data.shape[0] // self.bs
data = np.array(data[:nb*self.bs])
data = data.reshape(self.bs, -1).T
if self.backwards: data=data[::-1]
return LongTensor(data)
def get_batch(self, i, seq_len):
source = self.data
seq_len = min(seq_len, len(source) - 1 - i)
return source[i:i+seq_len], source[i+1:i+1+seq_len].contiguous().view(-1)
bs,bptt = 20,10
train_dl = LanguageModelLoader(np.concatenate(train_ids), bs, bptt)
valid_dl = LanguageModelLoader(np.concatenate(valid_ids), bs, bptt)
class LMDataBunch():
def __init__(self, train_dl, valid_dl, bs=64, device=None):
self.device = default_device if device is None else device
self.train_dl = DeviceDataLoader(train_dl, self.device, progress_func=tqdm)
self.valid_dl = DeviceDataLoader(valid_dl, self.device, progress_func=tqdm)
@property
def train_ds(self): return self.train_dl.dl.dataset
@property
def valid_ds(self): return self.valid_dl.dl.dataset
data = LMDataBunch(train_dl, valid_dl, bs)
# ## Model
# ### 1. Dropout
# We want to use the AWD-LSTM from [<NAME>](https://arxiv.org/abs/1708.02182). First, we'll need all different kinds of dropouts. Dropout consists into replacing some coefficients by 0 with probability p. To ensure that the averga of the weights remains constant, we apply a correction to the weights that aren't nullified of a factor `1/(1-p)`.
def dropout_mask(x, sz, p):
"Returns a dropout mask of the same type as x, size sz, with probability p to cancel an element."
return x.new(*sz).bernoulli_(1-p)/(1-p)
x = torch.randn(10,10)
dropout_mask(x, (10,10), 0.5)
# Once with have a dropout mask `m`, applying the dropout to `x` is simply done by `x = x * m`. We create our own dropout mask and don't rely on pytorch dropout because we want to nullify the coefficients on the batch dimension but not the token dimension (aka the same coefficients are replaced by zero for each word in the sentence).
#
# Inside a RNN, a tensor x will have three dimensions: seq_len, bs, vocab_size, so we create a dropout mask for the last two dimensions and broadcast it to the first dimension.
class RNNDropout(nn.Module):
def __init__(self, p=0.5):
super().__init__()
self.p=p
def forward(self, x):
if not self.training or not self.p: return x
m = dropout_mask(x.data, (1, x.size(1), x.size(2)), self.p)
return m * x
dp_test = RNNDropout(0.5)
x = torch.randn(2,5,10)
x, dp_test(x)
class WeightDropout(nn.Module):
"A module that warps another layer in which some weights will be replaced by 0 during training."
def __init__(self, module, dropout, layer_names=['weight_hh_l0']):
super().__init__()
self.module,self.dropout,self.layer_names = module,dropout,layer_names
for layer in self.layer_names:
#Makes a copy of the weights of the selected layers.
w = getattr(self.module, layer)
self.register_parameter(f'{layer}_raw', nn.Parameter(w.data))
def _setweights(self):
for layer in self.layer_names:
raw_w = getattr(self, f'{layer}_raw')
self.module._parameters[layer] = F.dropout(raw_w, p=self.dropout, training=self.training)
def forward(self, *args):
self._setweights()
return self.module.forward(*args)
def reset(self):
if hasattr(self.module, 'reset'): self.module.reset()
module = nn.LSTM(20, 20)
dp_module = WeightDropout(module, 0.5)
opt = optim.SGD(dp_module.parameters(), 10)
dp_module.train()
x = torch.randn(2,5,20)
x.requires_grad_(requires_grad=True)
h = (torch.zeros(1,5,20), torch.zeros(1,5,20))
for _ in range(5): x,h = dp_module(x,h)
getattr(dp_module.module, 'weight_hh_l0'),getattr(dp_module,'weight_hh_l0_raw')
target = torch.randint(0,20,(10,)).long()
loss = F.nll_loss(x.view(-1,20), target)
loss.backward()
opt.step()
w, w_raw = getattr(dp_module.module, 'weight_hh_l0'),getattr(dp_module,'weight_hh_l0_raw')
w.grad, w_raw.grad
getattr(dp_module.module, 'weight_hh_l0'),getattr(dp_module,'weight_hh_l0_raw')
class EmbeddingDropout(nn.Module):
"Applies dropout in the embedding layer by zeroing out some elements of the embedding vector."
def __init__(self, emb, dropout):
super().__init__()
self.emb,self.dropout = emb,dropout
self.pad_idx = self.emb.padding_idx
if self.pad_idx is None: self.pad_idx = -1
def forward(self, words, dropout=0.1, scale=None):
if self.training and self.dropout != 0:
size = (self.emb.weight.size(0),1)
mask = dropout_mask(self.emb.weight.data, size, self.dropout)
masked_emb_weight = mask * self.emb.weight
else: masked_emb_weight = self.emb.weight
if scale: masked_emb_weight = scale * masked_emb_weight
return F.embedding(words, masked_emb_weight, self.pad_idx, self.emb.max_norm,
self.emb.norm_type, self.emb.scale_grad_by_freq, self.emb.sparse)
enc = nn.Embedding(100,20, padding_idx=0)
enc_dp = EmbeddingDropout(enc, 0.5)
x = torch.randint(0,100,(25,)).long()
enc_dp(x)
# ### 2. AWD-LSTM
import warnings
def repackage_var(h):
"Detaches h from its history."
return h.detach() if type(h) == torch.Tensor else tuple(repackage_var(v) for v in h)
class RNNCore(nn.Module):
"AWD-LSTM/QRNN inspired by https://arxiv.org/abs/1708.02182"
initrange=0.1
def __init__(self, vocab_sz, emb_sz, n_hid, n_layers, pad_token, bidir=False,
hidden_p=0.2, input_p=0.6, embed_p=0.1, weight_p=0.5, qrnn=False):
super().__init__()
self.bs,self.qrnn,self.ndir = 1, qrnn,(2 if bidir else 1)
self.emb_sz,self.n_hid,self.n_layers = emb_sz,n_hid,n_layers
self.encoder = nn.Embedding(vocab_sz, emb_sz, padding_idx=pad_token)
self.dp_encoder = EmbeddingDropout(self.encoder, embed_p)
if self.qrnn:
#Using QRNN requires cupy: https://github.com/cupy/cupy
from .torchqrnn.qrnn import QRNNLayer
self.rnns = [QRNNLayer(emb_sz if l == 0 else n_hid, (n_hid if l != n_layers - 1 else emb_sz)//self.ndir,
save_prev_x=True, zoneout=0, window=2 if l == 0 else 1, output_gate=True) for l in range(n_layers)]
if weight_p != 0.:
for rnn in self.rnns:
rnn.linear = WeightDropout(rnn.linear, weight_p, layer_names=['weight'])
else:
self.rnns = [nn.LSTM(emb_sz if l == 0 else n_hid, (n_hid if l != n_layers - 1 else emb_sz)//self.ndir,
1, bidirectional=bidir) for l in range(n_layers)]
if weight_p != 0.: self.rnns = [WeightDropout(rnn, weight_p) for rnn in self.rnns]
self.rnns = torch.nn.ModuleList(self.rnns)
self.encoder.weight.data.uniform_(-self.initrange, self.initrange)
self.dropouti = RNNDropout(input_p)
self.dropouths = nn.ModuleList([RNNDropout(hidden_p) for l in range(n_layers)])
def forward(self, input):
sl,bs = input.size()
if bs!=self.bs:
self.bs=bs
self.reset()
raw_output = self.dropouti(self.dp_encoder(input))
new_hidden,raw_outputs,outputs = [],[],[]
for l, (rnn,drop) in enumerate(zip(self.rnns, self.dropouths)):
with warnings.catch_warnings():
#To avoid the warning that comes because the weights aren't flattened.
warnings.simplefilter("ignore")
raw_output, new_h = rnn(raw_output, self.hidden[l])
new_hidden.append(new_h)
raw_outputs.append(raw_output)
if l != self.n_layers - 1: raw_output = drop(raw_output)
outputs.append(raw_output)
self.hidden = repackage_var(new_hidden)
return raw_outputs, outputs
def one_hidden(self, l):
nh = (self.n_hid if l != self.n_layers - 1 else self.emb_sz)//self.ndir
return self.weights.new(self.ndir, self.bs, nh).zero_()
def reset(self):
[r.reset() for r in self.rnns if hasattr(r, 'reset')]
self.weights = next(self.parameters()).data
if self.qrnn: self.hidden = [self.one_hidden(l) for l in range(self.n_layers)]
else: self.hidden = [(self.one_hidden(l), self.one_hidden(l)) for l in range(self.n_layers)]
class LinearDecoder(nn.Module):
"To go on top of a RNN_Core module"
initrange=0.1
def __init__(self, n_out, n_hid, output_p, tie_encoder=None, bias=True):
super().__init__()
self.decoder = nn.Linear(n_hid, n_out, bias=bias)
self.decoder.weight.data.uniform_(-self.initrange, self.initrange)
self.dropout = RNNDropout(output_p)
if bias: self.decoder.bias.data.zero_()
if tie_encoder: self.decoder.weight = tie_encoder.weight
def forward(self, input):
raw_outputs, outputs = input
output = self.dropout(outputs[-1])
decoded = self.decoder(output.view(output.size(0)*output.size(1), output.size(2)))
return decoded, raw_outputs, outputs
class SequentialRNN(nn.Sequential):
def reset(self):
for c in self.children():
if hasattr(c, 'reset'): c.reset()
def get_language_model(vocab_sz, emb_sz, n_hid, n_layers, pad_token, tie_weights=True, qrnn=False, bias=True,
output_p=0.4, hidden_p=0.2, input_p=0.6, embed_p=0.1, weight_p=0.5):
"To create a full AWD-LSTM"
rnn_enc = RNNCore(vocab_sz, emb_sz, n_hid=n_hid, n_layers=n_layers, pad_token=pad_token, qrnn=qrnn,
hidden_p=hidden_p, input_p=input_p, embed_p=embed_p, weight_p=weight_p)
enc = rnn_enc.encoder if tie_weights else None
return SequentialRNN(rnn_enc, LinearDecoder(vocab_sz, emb_sz, output_p, tie_encoder=enc, bias=bias))
tst_model = get_language_model(500, 20, 100, 2, 0)
x = torch.randint(0, 500, (10,5)).long()
z = tst_model(x)
len(z)
# ### 3. Callback to train the model
@dataclass
class RNNTrainer(Callback):
learn:Learner
bptt:int
clip:float=None
alpha:float=0.
beta:float=0.
def on_loss_begin(self, last_output, **kwargs):
#Save the extra outputs for later and only returns the true output.
self.raw_out,self.out = last_output[1],last_output[2]
return last_output[0]
def on_backward_begin(self, last_loss, last_input, last_output, **kwargs):
#Adjusts the lr to the bptt selected
self.learn.opt.lr *= last_input.size(0) / self.bptt
#AR and TAR
if self.alpha != 0.: last_loss += (self.alpha * self.out[-1].pow(2).mean()).sum()
if self.beta != 0.:
h = self.raw_out[-1]
if len(h)>1: last_loss += (self.beta * (h[1:] - h[:-1]).pow(2).mean()).sum()
return last_loss
def on_backward_end(self, **kwargs):
if self.clip: nn.utils.clip_grad_norm_(self.learn.model.parameters(), self.clip)
emb_sz, nh, nl = 400, 1150, 3
model = get_language_model(vocab_size, emb_sz, nh, nl, 0, input_p=0.6, output_p=0.4, weight_p=0.5,
embed_p=0.1, hidden_p=0.2)
learn = Learner(data, model)
learn.opt_fn = partial(optim.Adam, betas=(0.8,0.99))
learn.callbacks.append(RNNTrainer(learn, bptt, clip=0.12, alpha=2, beta=1))
fit_one_cycle(learn, 5e-3, 1, (0.8,0.7), wd=1.2e-6)
|
dev_nb/007_wikitext_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/bereml/iap/blob/master/libretas/1c_reglin.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Regresión lineal
#
# Curso: [Introducción al Aprendizaje Profundo](http://turing.iimas.unam.mx/~ricardoml/course/iap/). Profesores: [Bere](https://turing.iimas.unam.mx/~bereml/) y [Ricardo](https://turing.iimas.unam.mx/~ricardoml/) <NAME>.
#
# ---
# ---
#
# En esta libreta implementaremos el modelo regresión lineal sobre un conjunto de datos de calificaciones, lo que deseamos predecir es que calificación tendrá el alumno en un examen. El conjunto de datos posee dos atributos entrada: la calificación que obtuvo en un examen previo y el número de horas que estudio el alumno para presentar el examen. En este ejemplo consideraremos únicamente un atributo de entrada, el número de horas de estudio.
#
# 
# <div> https://xkcd.com/1725/ </div>
# ## 1 Preparación
#
# ### 1.1 Bibliotecas
# +
# sistema de archivos
import os
# gráficas
import matplotlib.pyplot as plt
# csv
import pandas as pd
# redes neuronales
import torch
# -
# ### 1.2 Auxiliares
URL = 'https://raw.githubusercontent.com/bereml/iap/master/datos/califs.csv'
base_dir = '../datos'
filename = 'califs.csv'
filepath = os.path.join(base_dir, filename)
# ## 2 Datos
# Descargamos los datos.
# ! mkdir {base_dir}
# ! wget -nc {URL} -O {filepath}
# Utilizamos para la lectura del csv e imprimimos los primeros 5 ejemplos.
df = pd.read_csv(filepath)
df.head(5)
# Obtengamos el atributo y la salida:
x = torch.tensor(df.iloc[:, 1].values)
x = x.view(-1, 1).type(torch.float32)
print(x.shape)
x[:5]
y_true = torch.tensor(df.iloc[:, 2].values)
y_true = y_true.view(-1, 1).type(torch.float32)
print(y_true.shape)
y_true[:5]
# número de ejemplos y atributos
m, d = x.shape
m, d
# Grafiquemos para tener una idea de la distribución de los datos:
plt.plot(x.view(-1).numpy(), y_true.view(-1).numpy(),
'.', color='tab:blue', markersize=8)
plt.xlabel('horas de estudio')
plt.ylabel('calificación')
plt.show()
# ## 3 Hipótesis
# Recordemos que dado un conjunto de ejemplos con atributos ($x_1, \dots, x_d$) y salidas $y$, la hipótesis de regresión lineal considerando un plano está dada por:
#
# $$\hat{y} = x_1 w_1 + \dots + x_d w_d + b$$
#
# donde $w_i$ y $b$ son pesos y sesgo (parámetros) del modelo y $\hat{y}$ la salida predicha. Podemos expresar la hipótesis en su forma vectorial como:
#
# $$\hat{y} = x w + b$$
#
# Nuestro trabajo consiste en estimar (aprender) los parámetros $w_i$ y $b$. Por el momento supongamos que proponemos valores para los parámetros y hagamos inferencia:
# +
# parámetros que nos regalo el oráculo
w = torch.tensor([[0.6030]])
b = torch.tensor([0.0496])
# inferencia
y_pred = x @ w + b
# gráfica
plt.plot(x.view(-1).numpy(), y_true.view(-1).numpy(),
'.', color='tab:blue', markersize=8)
plt.plot(x.view(-1).numpy(), y_pred.view(-1).numpy(),
'-', color='tab:green', markersize=8)
plt.xlabel('horas de estudio')
plt.ylabel('calificación')
plt.show()
# -
# ## 4 Función de pérdida
#
# La pérdida para este modelo es el error cuadrático medio y queda expresado de la siguiente manera:
#
# $$ J(w, b) = \frac{1}{2m} \sum_{i=1}^{m}{(\hat{y}^{(i)} - y^{(i)})^2} $$
#
# en su forma vectorial:
#
# $$ J(w, b) = \frac{1}{2m} (\hat{y} - y)^T (\hat{y} - y) $$
#
# Para los parámetros propuestos, la pérdida se puede implementar como:
loss = (y_pred - y_true).T @ (y_pred - y_true) / (2 * m)
loss
# ## 5 Descenso por gradiente
#
# 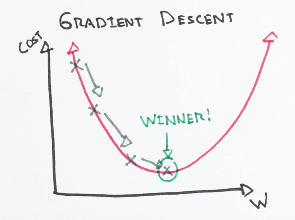
# <div> https://ml-cheatsheet.readthedocs.io/en/latest/gradient_descent.html </div>
#
#
# El algoritmo del gradiente descendente se basa en el gradiente de la pérdida respecto de los parámetros:
#
# $$\frac{\partial J(w_j)}{\partial w_j} = \frac{1}{m} \sum_{i=1}^{m}{x^{(i)}_j(\hat{y}^{(i)} - y^{(i)})}$$
#
# $$\frac{\partial J(b)}{\partial b} = \frac{1}{m} \sum_{i=1}^{m}{(\hat{y}^{(i)} - y^{(i)})}$$
#
# en su forma vectorial:
#
# $$\Delta J(w) = \frac{1}{m} x^T (\hat{y} - y)$$
#
# $$\Delta J(b) = \frac{1}{m} \sum_{i=1}^{m}{(\hat{y} - y)^{(i)}}$$
#
# Para los parámetros propuestos, el computo gradiente se puede implementar como:
grad_w = (x.T @ (y_pred - y_true)) / m
grad_b = (y_pred - y_true).sum() / m
grad_w, grad_b
# ## 6 Entrenamiento
#
# Ahora implementemos todo en una función:
def train(x, y_true, alpha=0.01, steps=10):
"""Fits linear regression."""
# ejemplos, atributos
m, d = x.shape
# inicialización de parámetros
w = torch.zeros(d, 1)
b = torch.zeros(1)
# histórico de pérdidas
loss_hist = []
# ciclo de entrenamiento
for i in range(steps):
# cómputo de la hipótesis
y_pred = x @ w + b
# cómputo de la pérdida
loss = (y_pred - y_true).T @ (y_pred - y_true) / (2 * m)
# cómputo del gradiente
grad_w = (x.T @ (y_pred - y_true)) / m
grad_b = (y_pred - y_true).sum() / m
# actualización de parámetros
w = w - alpha * grad_w
b = b - alpha * grad_b
# histórico de pérdida
loss_hist.append(loss)
return w, b, loss_hist
# Entrenemos un modelo:
w, b, loss_hist = train(x, y_true)
w, b
# Grafiquemos la evolución de la pérdida:
plt.figure()
plt.plot(range(len(loss_hist)), loss_hist, color='tab:red')
plt.show()
# ## 7 Discusión
#
# * ¿De qué otra forma podemos inicilizar los parámetros?
# * ¿Cómo influye la tasa de aprendizaje?
# * ¿Cómo elejimos el número de pasos de entrenamiento?
|
libretas/1c_reglin.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:StockPricePrediction] *
# language: python
# name: conda-env-StockPricePrediction-py
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import pickle
from sklearn.impute import SimpleImputer
import matplotlib.pyplot as plt
sns.set(style="white")
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
import os
os.getcwd()
# +
with open('../../data/interim/features.pickle', 'rb') as file:
features = pickle.load(file)
date = features['Date']
features.drop('Date', inplace=True, axis=1)
# calculate difference between rows
features_diff = features.pct_change().replace([np.inf, -np.inf], np.nan)
# impute NaN values
imp_mean = SimpleImputer(missing_values=np.nan, strategy='mean')
imp_mean.fit(features_diff)
features_diff = imp_mean.transform(features_diff)
# features_diff.dropna()
features_diff = pd.DataFrame(data=features_diff, columns=features.columns)
imp_mean = SimpleImputer(missing_values=np.nan, strategy='mean')
imp_mean.fit(features)
features_ = imp_mean.transform(features)
features_ = pd.DataFrame(data=features_, columns=features.columns)
# +
plt.figure(figsize = (14,10))
ax = sns.heatmap(features_diff.corr(), annot=False, linewidths=.5)
# +
plt.figure(figsize = (14,14))
ax = sns.pairplot(features_.iloc[:, [0,7,8,9,10,11]])
# -
features_.iloc[:, :2]
, hue='Close'
# +
targets_ = pd.DataFrame(features_['Close'].pct_change(), columns=['Close'])
bin_labels = (targets_['Close'] >= 0).astype(np.int32).values[1:]
bin_labels = bin_labels[700:]
features_ = features_.iloc[700:]
# +
import xgboost as xgb
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import balanced_accuracy_score, accuracy_score
x_train, x_test, y_train, y_test = train_test_split(features_.iloc[:-1], bin_labels, test_size=0.2, shuffle=False)
scl = StandardScaler()
scl.fit(x_train)
x_train = pd.DataFrame(scl.transform(x_train), columns=features_.columns.to_list())
x_test = pd.DataFrame(scl.transform(x_test), columns=features_.columns.to_list())
clf = xgb.XGBClassifier(n_estimators=5, max_depth=6, gamma=0.2)
fig, ax = plt.subplots(figsize=(20, 10))
xgb.plot_importance(model, ax=ax)
preds = model.predict(x_test)
accuracy_score(y_test, preds)
# +
from sklearn.linear_model import LogisticRegression
logit = LogisticRegression(solver='lbfgs', max_iter=300, verbose=1)
m = logit.fit(x_train, y_train)
fig, ax = plt.subplots(figsize=(20, 10))
xgb.plot_importance(model, ax=ax)
preds = m.predict(x_test)
balanced_accuracy_score(y_test, preds)
# +
models = ['pseudo-losowy', 'gęsto-połączony', 'GRU', 'LSTM', 'CONV+LSTM']
accuracy = [0.5, 0.744, 0.779, 0.779, 0.779]
precision = [0.497, 0.776, 0.772, 0.793, 0.816]
recall = [0.51, 0.686, 0.791, 0.756, 0.721]
df = pd.DataFrame(data=list(zip(models, accuracy, precision, recall)), columns=['models', 'accuracy', 'precision', 'recall'])
# df = df.sort_values(by='accuracy', ascending=False)
models = df.models.to_list()
df = df.T
# df.set_index('models', inplace=True)
barWidth=0.15
r1 = np.arange(3)
r2 = [x + barWidth for x in r1]
r3 = [x + barWidth for x in r2]
r4 = [x + barWidth for x in r3]
r5 = [x + barWidth for x in r4]
df
# df.iloc[1, :]
# -
def plot():
fig = plt.figure(figsize=(16, 8))
color = ['b', 'orange', 'red', 'green', 'brown']
for i, r in enumerate([r1, r2, r3, r4, r5]):
plt.bar(r, df.iloc[1:, i], color=color[i], width=0.15)
plt.xlabel('Metryka')
plt.ylabel('Wartość')
plt.xticks([r + 0.3 for r in range(3)], ['Dokładność', 'Precyzja', 'Wrażliwość'])
plt.legend(models)
plt.grid(1)
plot()
# df = df.sort_values(by='precision', ascending=False)
# fig = plt.figure(figsize=(10, 6))
# _ = plt.bar(r2, df.iloc[1:, 1], color='orange',width=0.2)
# _ = plt.xlabel('Model')
# _ = plt.ylabel('Precyzja')
df = df.sort_values(by='precision', ascending=False)
# fig = plt.figure(figsize=(10, 6))
_ = plt.bar(df.index, df.precision, color='b')
_ = plt.xlabel('Model')
_ = plt.ylabel('Precyzja')
df = df.sort_values(by='recall', ascending=False)
fig = plt.figure(figsize=(10, 6))
_ = plt.bar(df.index, df.recall, color='b')
_ = plt.xlabel('Model')
_ = plt.ylabel('Wrażliwość')
|
src/visualization/written_project_visualisation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Get From https://github.com/BlaBlaBlazzz/Fake_News_classification/blob/main/Fake_news_identification.ipynb
# Because he said there is a performance issue, so I copy it and test.
import os
from tqdm import tqdm
import glob
import pandas as pd
import numpy as np
glob.glob("*.csv.zip")
# +
#os.system("unzip train.csv.zip")
#os.system("unzip test.csv.zip")
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
submit = pd.read_csv("sample_submission.csv")
train = train.reset_index()
cols = ['title1_zh', 'title2_zh', 'label']
train = train.loc[:,cols]
train.columns = ['text_a', 'text_b', 'label']
train.to_csv("train.tsv", sep="\t", index=False)
train.head()
# -
train.label.value_counts() / len(train)
pip install transformers tqdm boto3 requests regex -q
from torch.utils.data import Dataset
import tensorflow
import torch
torch.cuda.is_available()
import transformers
from transformers import BertTokenizer
#from pytorch_transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained("bert-base-chinese")
# +
from torch.utils.data import Dataset
class FakeNewsIdensification(Dataset):
def __init__(self, mode, tokenizer):
assert mode in ["train", "test"]
self.mode = mode
self.df = pd.read_csv(mode + ".tsv", sep="\t").fillna("")
self.len = len(self.df)
self.label_map = {"agreed":0, "disagreed":1, "unrelated":2}
self.tokenizer = tokenizer
def __getitem__(self, idx):
if self.mode == "test":
text_a, text_b = self.df.iloc[idx,:2].values
label_tensor = None
else:
text_a, text_b, label = self.df.iloc[idx,:].values
label_id = self.label_map[label]
label_tensor = torch.tensor(label_id)
word_pieces = ["[CLS]"]
token_a = self.tokenizer.tokenize(text_a)
word_pieces += token_a
word_pieces += ["[SEP]"]
len_a = len(word_pieces)
token_b = self.tokenizer.tokenize(text_b)
word_pieces += token_b
len_b = len(word_pieces) - len_a
ids = self.tokenizer.convert_tokens_to_ids(word_pieces)
tokens_tensor = torch.tensor(ids)
segments_tensor = torch.tensor([0]*len_a + [1]*len_b, dtype=torch.long)
return (tokens_tensor, segments_tensor, label_tensor)
def __len__(self):
return self.len
trainset = FakeNewsIdensification("train", tokenizer=tokenizer)
# +
from torch.utils.data import DataLoader
from torch.nn.utils.rnn import pad_sequence
def mini_batch(samples):
token_tensors = [s[0] for s in samples]
segment_tensors = [s[1] for s in samples]
if samples[0][2] is not None:
label_ids = torch.stack([s[2] for s in samples])
else:
label_ids = none
token_tensors = pad_sequence(token_tensors, batch_first=True)
segment_tensors = pad_sequence(segment_tensors, batch_first=True)
#Mask_tensor
mask_tensors = torch.zeros(token_tensors.shape, dtype=torch.long)
mask_tensors = mask_tensors.masked_fill(token_tensors != 0, 1)
return token_tensors, segment_tensors, mask_tensors, label_ids
BATCH_SIZE = 64
if __name__ == '__main__':
trainloader = DataLoader(
trainset,
batch_size = BATCH_SIZE,
collate_fn = mini_batch,
num_workers = 0,
pin_memory = True
)
# +
from transformers import BertForSequenceClassification, BertPreTrainedModel
from IPython.display import display, clear_output
PRETRAINED_MODEL_NAME = "bert-base-chinese"
NUM_LABELS = 3
model = BertForSequenceClassification.from_pretrained(
PRETRAINED_MODEL_NAME, num_labels=NUM_LABELS)
clear_output()
model.config
# +
class BertForSequenceClassification(BertPreTrainedModel):
def __init__(self, config, num_labels=3):
super(BertForSequenceClassification, self).__init__(config)
self.num_labels = num_labels
self.bert = BertModel(config) #config -> bert的預訓練參數
#hidden_dropout_prob = 0.1
self.dropout(config.hidden_dropout_prob)
#hidden_size=768
self.linear(config.hidden_size, num_labels)
def forward(self, input_ids, token_type_ids=None, attention_mask=None, label=None):
output = self.bert(input_ids, token_type_ids, attention_mask)
pooled_output = self.dropout(pooled_output) #pooled_output -> 訓練bert輸出
logits = self.classifier(pooled_output)
if labels is not None:
self.loss = CrossEntropyLoss()
loss = loss_fct(logits.view(-1, self.num_labels), labels.view(-1))
return loss
elif self.output_attentions:
return all_attentions, logits
return logits
#if __name__ == '__main__':
# dataloader = DataLoader(trainset, num_workers=0)
# for _ in tqdm(dataloader):
# pass
# +
def get_prediction(model, dataloader, compute_acc=None):
predictions = None
correct = 0
total = 0
with torch.no_grad():
for data in dataloader:
if next(model.parameters()).is_cuda:
data = [t.to("cuda:0") for t in data if t is not None]
token_tensors , segment_tensors, mask_tensors = data[:3]
outputs = model(input_ids = token_tensors, token_type_ids = segment_tensors,
attention_mask = mask_tensors)
print(outputs)
logits = outputs[0]
_, prediction = torch.max(logits.data, 1)
# compute_acc == True
if compute_acc:
labels = data[3]
total += labels.size(0)
correct = (prediction==labels).sum().item()
if predictions is None:
predictions = prediction
else:
predictions = torch.cat(predictions, prediction)
if compute_acc:
accuracy = correct/total
return prediction, accuracy
else:
return prediction
#from tensorflow.python.client import device_lib
#print(device_lib.list_local_devices())
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print("device:", device)
model = model.to(device)
_, acc = get_prediction(model, trainloader, compute_acc=True)
print("classification acc:", acc)
# +
model.eval()
model.train()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-5)
epochs = 5
for epoch in range(epochs):
epoch_loss = 0
if __name__ == '__main__':
for data in trainloader:
token_tensors, segment_tensors, mask_tensors, labels = [t.to(device) for t in data]
optimizer.zero_grad()
output = model(input_ids = token_tensors, token_type_ids = segment_tensors,
attention_mask = mask_tensors, labels = labels)
loss = output[0]
loss = loss.requires_grad_()
loss.backward()
optimizer.step()
epoch_loss += loss.item()
_, acc = get_prediction(model, trainloader, compute_acc=True)
print('[epoch %d] loss: %.3f, acc: %.3f' %(epoch + 1, epoch_loss, acc))
# -
|
Practice/PyTorch/NER_test/Fake_news_identification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Counting sort
# El algoritmo counting sort se basa en determinar para cada elemento x del conjunto, el número de elementos menores a x. Con esa información, es posible poner el elemento x en su posición correcta dentro del conjunto ordenado.
#
# El algoritmo counting sort asume que los elementos que conforman el conjunto son enteros positivos (incluyendo a cero).
# +
# https://www.geeksforgeeks.org/sorting-algorithms/
# %pylab inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import random
# -
def counting_sort_chars(A):
MAX = 256
B = [0 for i in range(len(A))]
C = [0 for i in range(MAX)]
i = 0
while i < len(A):
C[ord(A[i])] += 1
i += 1
i = 1
while i < len(C):
C[i] += C[i-1]
i += 1
i = 0
while i < len(A):
B[C[ord(A[i])]-1] = A[i]
C[ord(A[i])] -= 1
i += 1
return B
# +
TAM = 21
abc = ['a','b','c','d','e','f','g','h','i','j','k','l','m','n','o','p','q','r','s','t','u','v','w','x','y','z']
arr = []
for i in range(1, TAM, 1):
arr.append(abc[random.randint(0, len(abc)-1)])
print(arr)
ans = counting_sort_chars(arr)
print(ans)
# -
class Node:
def __init__(self, number, name, lastname, email, sex):
self.identifier = number
self.name = name
self.last_name = lastname
self.email = email
self.sex = sex[0]
def __str__(self):
return self.name + ":" + self.email
#return self.name
def counting_sort_node(nodeList):
MAX = 128
output = [0 for i in range(len(nodeList))]
count = [0 for i in range(MAX)]
for i in nodeList:
count[ord(i.name[0])] += 1
for i in range(MAX):
count[i] += count[i-1]
for i in range(len(nodeList)):
output[count[ord(nodeList[i].name[0])]-1] = nodeList[i]
count[ord(nodeList[i].name[0])] -= 1
return output
# +
arr = []
file = open("DATA_10.csv", "r")
for line in file:
fields = line.split(",")
arr.append(Node(fields[0], fields[1], fields[2], fields[3], fields[4]))
print("List")
for node in arr:
print(node)
ans = counting_sort_node(arr)
print("\nsort list")
for node in ans:
print(node)
# -
def counting_sort_graph(nodeList):
cont = 0
MAX = 128
output = [0 for i in range(len(nodeList))]
count = [0 for i in range(MAX)]
for i in nodeList:
cont += 1
count[ord(i.name[0])] += 1
for i in range(MAX):
cont += 1
count[i] += count[i-1]
for i in range(len(nodeList)):
cont += 1
output[count[ord(nodeList[i].name[0])]-1] = nodeList[i]
count[ord(nodeList[i].name[0])] -= 1
return cont
TAM = 101
x = list(range(1,TAM,1))
y_omega = []
y_efedeene = []
y_omicron = []
L = []
for num in x:
iter = 1
file = open("DATA_10000.csv", "r")
L = []
for line in file:
fields = line.split(",")
L.append(Node(fields[0], fields[1], fields[2], fields[3], fields[4]))
if iter == num:
break
iter += 1
# average case
y_efedeene.append(counting_sort_graph(L))
# best case
y_omega.append(counting_sort_graph(L))
# worst case
y_omicron.append(counting_sort_graph(L))
# +
fig, ax = plt.subplots(facecolor='w', edgecolor='k')
ax.plot(x, y_omega, marker="o",color="b", linestyle='None')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.grid(True)
ax.legend(["counting sort"])
plt.title('counting sort (best case)')
plt.show()
fig, ax = plt.subplots(facecolor='w', edgecolor='k')
ax.plot(x, y_efedeene, marker="o",color="b", linestyle='None')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.grid(True)
ax.legend(["counting sort"])
plt.title('counting sort (average case)')
plt.show()
fig, ax = plt.subplots(facecolor='w', edgecolor='k')
ax.plot(x, y_omicron, marker="o",color="b", linestyle='None')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.grid(True)
ax.legend(["counting sort"])
plt.title('counting sort (worst case)')
plt.show()
# -
|
EDyA_II/1_sort/3_Counting sort.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: auto-ru
# language: python
# name: auto-ru
# ---
# %load_ext autoreload
# %autoreload 2
# + colab={} colab_type="code" id="k3ZINe3G6kMJ" pycharm={"is_executing": false}
import sys
import json
import pandas as pd
import os
import time
from tqdm.notebook import tqdm
# +
src = os.path.abspath('../src')
if src not in sys.path:
sys.path.append(src)
from spider import scrap
# + [markdown] colab_type="text" id="90YKGNx96kMh"
#
# # VK-APP execute stored procedure
#
# `execute.getInfo` code:
#
# ```
#
# var user = API.users.get({"user_ids": [Args.user], "fields": ["photo_id", "sex", "has_photo", "photo_50", "photo_200_orig", "domain", "status", "last_seen", "followers_count", "common_count", "relation", "verified", "personal", "interests", "can_be_invited_group"]});
#
# var friends = API.firends.get({"user_id": Args.user, "count": 1000});
#
# var followers = API.users.getFollowers({"user_id": Args.user, "count": 1000});
#
# var groups = API.groups.get({"user_id": Args.user, "extended": 1});
#
# var wall = API.wall.get({"owner_id": Args.user, "count": 100, "extended": 1});
#
# return [user, friends, followers, groups, wall];```
# + [markdown] magic_args="7356989"
# %% +79023633381
# %% #Nineteen67tah
# -
scrap()
|
notebooks/scraper_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <span style="color:Maroon">Short Term XgBoost Model
# __Summary:__ <span style="color:Blue">In this code we shall build and test a short term XgBoost Model using Technical Indicators
# Import required libraries
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from xgboost.sklearn import XGBClassifier
from sklearn.model_selection import KFold, GridSearchCV
from itertools import cycle
from sklearn.metrics import roc_curve, auc
from sklearn.preprocessing import label_binarize
from sklearn.metrics import balanced_accuracy_score, make_scorer, classification_report
from sklearn import metrics
import os
import ta
import pickle
from scipy import interp
np.random.seed(0)
# +
# User defined names
index = "BTC-USD"
filename = index+"_hurst_segment_dependent.csv"
date_col = "Date"
# Declare the hyper-parameters for grid search
max_depth = [4, 6]
min_child_weight = [10, 20]
gamma = [0, 0.1]
subsample = [0.8]
colsample_bytree = [0.8]
scale_pos_weight = [1]
learning_rate = [0.05, 0.1]
n_estimators = [200]
reg_alpha = [1e-5, 0.1, 1, 100]
reg_lambda = [0, 0.001, 0.01, 0.1]
# -
# Get current working directory
mycwd = os.getcwd()
print(mycwd)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Data")
# Read the data
df = pd.read_csv(filename, index_col=date_col)
df.index = pd.to_datetime(df.index)
df.head()
#
#
# ## <span style="color:Maroon">Functions
def Split_data_XY(df, dv):
"""
Given a dataset returns two dataframes, X-Dataframe and y-dataframe
"""
X_df = df.drop([dv], axis=1)
y_df = df[dv]
y_labelizer = label_binarize(y_df, classes=[-1, 0, 1])
return X_df, y_df, y_labelizer
def Get_Max_Discretevar(df, var, window=10):
"""
Get maximum value on rolling basis for the variable
"""
df[var+"_max"+str(window)] = df[var].rolling(window=window).max()
return df
def Get_SMA_Continousvar(df, var, window=10):
"""
Get SMA for continous variable
"""
df[var+"_sma"+str(window)] = df[var].rolling(window=window).mean()
return df
def Get_Ratio_Continousvar(df, var, window=10):
"""
Get Ratio for continous variable Min/Max
"""
df[var+"_ratio_minmax"+str(window)] = np.where(np.abs(df[var].rolling(window=window).max()) > 0,
df[var].rolling(window=window).min()/
df[var].rolling(window=window).max(),0)
return df
def Get_std_Continousvar(df, var, window=30):
"""
Get Ratio for continous variable Min/Max
"""
df[var+"_std"+str(window)] = df[var].rolling(window=window).std()
return df
def Generate_Predicted_df(X_train, y_train, X_test, y_test, clf):
"""
Generates Pandas dataframe with predicted values and other columns for P&L analysis
"""
# Train Sample
df_train = pd.DataFrame(y_train)
df_train['Predicted'] = clf.predict(X_train)
df_train['Adj Close'] = X_train['Adj Close']
df_train['Open'] = X_train['Open']
df_train['DVT STD'] = X_train['DVT STD']
df_train["Sample"] = "Train"
# Test Sample
df_test = pd.DataFrame(y_test)
df_test['Predicted'] = clf.predict(X_test)
df_test['Adj Close'] = X_test['Adj Close']
df_test['Open'] = X_test['Open']
df_test['DVT STD'] = X_test['DVT STD']
df_test['Sample'] = "Test"
df = df_train.append(df_test)
return df
#
#
# ## <span style="color:Maroon">Feature Engineering
# Add all technical features
df = ta.add_all_ta_features(df, open="Open", high="High", low="Low", close="Adj Close", volume="Volume")
# Max variable list
max_vars = ['volatility_bbhi', 'volatility_bbli', 'volatility_kchi', 'volatility_kcli', 'trend_psar_up_indicator',
'trend_psar_down_indicator']
for i in range(0, len(max_vars)):
df = Get_Max_Discretevar(df, max_vars[i], 10)
# SMA variable list
sma_vars = ['volume_adi', 'volume_obv', 'volume_cmf', 'volume_fi', 'volume_mfi', 'volume_em', 'volume_sma_em',
'volume_vpt', 'volume_nvi', 'volume_vwap', 'volatility_atr', 'volatility_bbm', 'volatility_bbh',
'volatility_bbl', 'volatility_bbw', 'volatility_bbp', 'volatility_kcc', 'volatility_kch', 'volatility_kcl',
'volatility_kcw', 'volatility_kcp', 'volatility_dcl', 'volatility_dch', 'volatility_dcm', 'volatility_dcw',
'volatility_dcp', 'volatility_ui', 'trend_macd', 'trend_macd_signal', 'trend_macd_diff', 'trend_sma_fast',
'trend_sma_slow', 'trend_ema_fast', 'trend_ema_slow', 'trend_adx', 'trend_adx_pos', 'trend_adx_neg',
'trend_vortex_ind_pos', 'trend_vortex_ind_neg', 'trend_vortex_ind_diff', 'trend_trix', 'trend_mass_index',
'trend_cci', 'trend_dpo', 'trend_kst', 'trend_kst_sig', 'trend_kst_diff', 'trend_ichimoku_conv',
'trend_ichimoku_base', 'trend_ichimoku_a', 'trend_ichimoku_b', 'trend_visual_ichimoku_a',
'trend_visual_ichimoku_b', 'trend_aroon_up', 'trend_aroon_down', 'trend_aroon_ind', 'trend_stc',
'momentum_rsi', 'momentum_stoch_rsi', 'momentum_stoch_rsi_k', 'momentum_stoch_rsi_d', 'momentum_tsi',
'momentum_uo', 'momentum_stoch', 'momentum_stoch_signal', 'momentum_wr', 'momentum_ao', 'momentum_kama',
'momentum_roc', 'momentum_ppo', 'momentum_ppo_signal', 'momentum_ppo_hist', 'others_dr', 'others_dlr',
'others_cr']
for i in range(0, len(sma_vars)):
df = Get_SMA_Continousvar(df, sma_vars[i], window=10)
# Ratio of Min Max variables
for i in range(0, len(sma_vars)):
df = Get_Ratio_Continousvar(df, sma_vars[i], window=10)
# Ratio of std variables
for i in range(0, len(sma_vars)):
df = Get_std_Continousvar(df, sma_vars[i], window=30)
# Drop two features
df = df.drop(['trend_psar_down', 'trend_psar_up'], axis=1)
df = df[df['hurst_150'] > 0]
df.shape
# Drop rows with null values
df.dropna(inplace=True)
df.shape
#
#
# ## <span style="color:Maroon">Divide the data in Segments
df['Segment'].value_counts()
# Break dataset into three segments
df_MeanReverting = df[df['Segment'] == "Mean Reverting"]
df_Trending = df[df['Segment'] == "Trending"]
# Drop Segment variable from all datasets
df.drop("Segment", axis=1, inplace=True)
df_MeanReverting.drop("Segment", axis=1, inplace=True)
df_Trending.drop("Segment", axis=1, inplace=True)
# #### <span style="color:Maroon">Mean Reverting Dataset
# Divide dataset into Train and Test Sample. (5 Fold CV will be used for validation)
df_MeanReverting_Train = df_MeanReverting[df_MeanReverting.index.year <= 2018]
df_MeanReverting_Test = df_MeanReverting[df_MeanReverting.index.year > 2018]
print("Train Sample: ", df_MeanReverting_Train.shape)
print("Test Sample: ", df_MeanReverting_Test.shape)
# #### <span style="color:Maroon">Trending Dataset
df_Trending_Train = df_Trending[df_Trending.index.year <= 2018]
df_Trending_Test = df_Trending[df_Trending.index.year > 2018]
print("Train Sample: ", df_Trending_Train.shape)
print("Test Sample: ", df_Trending_Test.shape)
# #### <span style="color:Maroon">Whole Dataset
df_Train = df[df.index.year <= 2018]
df_Test = df[df.index.year > 2018]
print("Train Sample: ", df_Train.shape)
print("Test Sample: ", df_Test.shape)
#
#
# ## <span style="color:Maroon">XgBoost Model Grid Search
# Grid
grid = {'max_depth': max_depth,
'min_child_weight': min_child_weight,
'gamma': gamma,
'subsample': subsample,
'colsample_bytree': colsample_bytree,
'scale_pos_weight': scale_pos_weight,
'learning_rate': learning_rate,
'n_estimators':n_estimators,
'reg_alpha':reg_alpha,
'reg_lambda':reg_lambda}
# XgBoost Model
scoring = {'Accuracy':make_scorer(balanced_accuracy_score)}
kfold = KFold(n_splits=3)
clf = XGBClassifier( objective= 'multi:softprob', num_classes=3, nthread=4, scale_pos_weight=1, seed=27,
eval_metric='mlogloss')
# Define grid search
grid = GridSearchCV(estimator = clf, param_grid=grid, cv=kfold, scoring=scoring, refit='Accuracy', verbose=1, n_jobs=-1)
#
# #### <span style="color:Maroon">Whole Dataset
# Get X, Y variables
X_train, y_train, y_train_label = Split_data_XY(df_Train, 'Target')
X_test, y_test, y_test_label = Split_data_XY(df_Test, 'Target')
# Fit the grid search model
model = grid.fit(X_train, y_train)
# Get the best xgboost model based on Grid Search
best_xgboost = model.best_estimator_
best_xgboost
# XgBoost model selected using Grid search
clf = best_xgboost
clf.fit(X_train, y_train)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Models")
# +
# Save the model
with open('whole_dataset'+str(index)+'_xgboost_model.pkl', 'wb') as f:
pickle.dump(clf, f)
# load it
with open('whole_dataset'+str(index)+'_xgboost_model.pkl', 'rb') as f:
clf = pickle.load(f)
# -
y_train_out = clf.predict(X_train)
print(classification_report(y_train, y_train_out))
# Confusion Matrix Train Sample
print("Train Sample Confusion Matrix")
pd.crosstab(y_train, y_train_out, rownames=['Actual'], colnames=['Predicted'])
y_test_out = clf.predict(X_test)
print(classification_report(y_test, y_test_out))
# Confusion Matrix Train Sample
print("Test Sample Confusion Matrix")
pd.crosstab(y_test, y_test_out, rownames=['Actual'], colnames=['Predicted'])
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Images")
y_score = clf.predict_proba(X_test)
n_classes = 3
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test_label[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# +
lw = 2
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
classes = [-1,0,1]
plt.figure()
colors = cycle(['aqua', 'darkorange', 'cornflowerblue'])
for i, color in zip(range(n_classes), colors):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(classes[i], roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC xgboost whole df')
plt.legend(loc="lower right")
plt.savefig("xgboost Whole df test" + str(index)+ " ROC curve"+'.png')
plt.show()
# -
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Data")
df_out = Generate_Predicted_df(X_train, y_train, X_test, y_test, clf)
df_out.to_csv('whole_dataset'+str(index)+'_xgboost_model.csv', index=True)
#
# #### <span style="color:Maroon">Trending Dataset
# Get X, Y variables
X_train, y_train, y_train_label = Split_data_XY(df_Trending_Train, 'Target')
X_test, y_test, y_test_label = Split_data_XY(df_Trending_Test, 'Target')
# Fit the grid search model
model = grid.fit(X_train, y_train)
# Get the best xgboost model based on Grid Search
best_xgboost = model.best_estimator_
best_xgboost
# XgBoost model selected using Grid search
clf = best_xgboost
clf.fit(X_train, y_train)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Models")
# +
# Save the model
with open('Trending_dataset'+str(index)+'_xgboost_model.pkl', 'wb') as f:
pickle.dump(clf, f)
# load it
with open('Trending_dataset'+str(index)+'_xgboost_model.pkl', 'rb') as f:
clf = pickle.load(f)
# -
y_train_out = clf.predict(X_train)
print(classification_report(y_train, y_train_out))
# Confusion Matrix Train Sample
print("Train Sample Confusion Matrix")
pd.crosstab(y_train, y_train_out, rownames=['Actual'], colnames=['Predicted'])
y_test_out = clf.predict(X_test)
print(classification_report(y_test, y_test_out))
# Confusion Matrix Train Sample
print("Test Sample Confusion Matrix")
pd.crosstab(y_test, y_test_out, rownames=['Actual'], colnames=['Predicted'])
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Images")
y_score = clf.predict_proba(X_test)
n_classes = 3
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test_label[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# +
lw = 2
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
classes = [-1,0,1]
plt.figure()
colors = cycle(['aqua', 'darkorange', 'cornflowerblue'])
for i, color in zip(range(n_classes), colors):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(classes[i], roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC xgboost Trend df')
plt.legend(loc="lower right")
plt.savefig("xgboost Trending_dataset test" + str(index)+ " ROC curve"+'.png')
plt.show()
# -
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Data")
df_out = Generate_Predicted_df(X_train, y_train, X_test, y_test, clf)
df_out.to_csv('Trending_dataset'+str(index)+'_xgboost_model.csv', index=True)
#
#
# #### <span style="color:Maroon">Mean Reverting Dataset
# Get X, Y variables
X_train, y_train, y_train_label = Split_data_XY(df_MeanReverting_Train, 'Target')
X_test, y_test, y_test_label = Split_data_XY(df_MeanReverting_Test, 'Target')
# Fit the grid search model
model = grid.fit(X_train, y_train)
# Get the best xgboost model based on Grid Search
best_xgboost = model.best_estimator_
best_xgboost
# XgBoost model selected using Grid search
clf = best_xgboost
clf.fit(X_train, y_train)
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Models")
# +
# Save the model
with open('MeanReverting_dataset'+str(index)+'_xgboost_model.pkl', 'wb') as f:
pickle.dump(clf, f)
# load it
with open('MeanReverting_dataset'+str(index)+'_xgboost_model.pkl', 'rb') as f:
clf = pickle.load(f)
# -
y_train_out = clf.predict(X_train)
print(classification_report(y_train, y_train_out))
# Confusion Matrix Train Sample
print("Train Sample Confusion Matrix")
pd.crosstab(y_train, y_train_out, rownames=['Actual'], colnames=['Predicted'])
y_test_out = clf.predict(X_test)
print(classification_report(y_test, y_test_out))
# Confusion Matrix Train Sample
print("Test Sample Confusion Matrix")
pd.crosstab(y_test, y_test_out, rownames=['Actual'], colnames=['Predicted'])
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Images")
y_score = clf.predict_proba(X_test)
n_classes = 3
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test_label[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# +
lw = 2
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
classes = [-1,0,1]
plt.figure()
colors = cycle(['aqua', 'darkorange', 'cornflowerblue'])
for i, color in zip(range(n_classes), colors):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(classes[i], roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC xgboost Mean Reverting')
plt.legend(loc="lower right")
plt.savefig("xgboost MeanReverting_dataset test" + str(index)+ " ROC curve"+'.png')
plt.show()
# -
# Change to data directory
os.chdir("..")
os.chdir(str(os.getcwd()) + "\\Data")
df_out = Generate_Predicted_df(X_train, y_train, X_test, y_test, clf)
df_out.to_csv('MeanReverting_dataset'+str(index)+'_xgboost_model.csv', index=True)
#
#
|
Dev/BTC-USD/Codes/05 Short Term xgboost Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Feature Generation and Dataset Preparation
# +
# Provides a way for us to save cells that we execute & test so that we can
# use them later in our prediction code (avoids having to copy-paste code)
#
# The syntax is as follows:
#
# # %%execute_and_save <filename>
from IPython.core import magic_arguments
from IPython.core.magic import (Magics, magics_class, line_magic,
cell_magic, line_cell_magic)
@magics_class
class SaveScripts(Magics):
@cell_magic
def execute_and_save(self, line, cell):
self.shell.run_cell(cell)
with open(line,'w') as file:
file.write(cell)
ip = get_ipython()
ip.register_magics(SaveScripts)
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg')
font = {'weight' : 'normal',
'size' : 9}
mpl.rc('font', **font)
mpl.rcParams['axes.titlesize'] = 'medium'
mpl.rcParams['axes.labelsize'] = 'medium'
# +
# %%execute_and_save tmp_imports
import numpy as np
import pandas as pd
import math
import os
from datetime import datetime
from datetime import timedelta
# Import our custom that retrieves financial market data from #http://financialmodelingprep.com
import api
import api.tickers as tickers # Information about stock, ETF, etc. tickers
import api.stocks as stocks # Information about stocks and stock indices
from api.stocks import Stock # Information about a particular stock
from api.stocks import Index # Information about a particular index
from api.stocks import TIMEDELTA_QUARTER
from api.stocks import TIMEDELTA_MONTH
from api.stocks import TIMEDELTA_YEAR
from sklearn.preprocessing import MinMaxScaler
# -
# ### Data Retrieval and Caching
sp500_tickers = tickers.get_sp500_tickers()
print("Sample S&P 500 tickers: ",sp500_tickers[:10])
stock_data = { ticker : Stock(ticker) for ticker in sp500_tickers }
# +
from IPython.display import display
def cache_stocks(start,end):
progress = display('',display_id=True)
success_tickers = list()
problem_tickers = list()
stock_list = list(stock_data.keys())
for i, ticker in enumerate(stock_list):
try:
progress.update(f'Caching {ticker} ({i+1}/{len(stock_list)})')
stock_data[ticker].cache_data(start,end)
success_tickers.append(ticker)
except:
problem_tickers.append(ticker)
continue
progress.update('Caching complete!')
print(f'Cached {len(success_tickers)} tickers: {", ".join(success_tickers)}.')
if len(problem_tickers) > 0:
print(f'The following tickers did not have complete data and were not cached: {", ".join(problem_tickers)}.')
# -
# Cache data for the last 15 quarters (90 calendar days / quarter)
end_date = datetime.today().date()
start_date = (end_date - TIMEDELTA_QUARTER*15)
# This is a potentially time-intensive operation
cache_stocks(start_date,end_date)
# ### Feature and Label Preparation
# We will want to smooth out our time-series data so that our algorithms (our neural network and calculations we perform on the time series data) are not susceptible to noise. For instance, we do not want the neural network to learn based upon a single jump in stock price over one or just a few days in the time-series data, as what is of more interest to us is the longer-term performance of a stock (e.g., not on a day-by-day basis, but rather the performance over a quarter).
#
# Further, simple moving averages (SMA) introduce a lag in the data where the smoothed data appears to be delayed relative to the actual trends in the time-series price data. For this reason, we will use an exponentially-weighted moving average. We can experiment with different window sizes to get some smoothing with negligible lag.
# +
# Let's use Apple as an example and see what kind of window size smooths out
# data for the last quarter while generally retaining features.
aapl_data = stock_data['AAPL'].get_historical_prices(start_date,end_date)
aapl_series = aapl_data['close']
aapl_smooth = aapl_series.ewm(span=5).mean()
plt.plot(aapl_series.values,'.',label='Close',color=plt.cm.viridis(.4),markerfacecolor=plt.cm.viridis(.9),markersize=12)
plt.plot(aapl_smooth.values,'-',label='5-day EWMA',color=plt.cm.viridis(0.7))
plt.title('Apple, Inc.')
plt.xlabel('Trading days')
plt.ylabel('Close Price')
plt.legend();
# -
# #### Stock Labels
# +
# %%execute_and_save tmp_benchmark_sp500_gain
# We can use a window size of 5 to smooth out some of the noise
# while retaining fidelity with respect to the general trends
# observed in the data.
# Let's compute the gain of the S&P 500 index over the last
# quarter. We will compare each sualtock's performance over the
# last quarte to this value.
# Note that the S&P 500 index is a capital-weighted index, so
# larger-cap stocks make up a larger portion of the fraction.
# Essentially the question we are asking is whether any given
# stock will outperform the index or "market". Investors can
# choose to invest in index-tracking ETFs instead of a given stock.
def get_sp500_gain_for_interval(interval,offset,output=False):
""" Get the gain for the S&P 500 over the specified interval
Args:
interval: The time interval for gain calculation as a datetime.timedelta
offset: The offset of interval relative to today as a datetime.timedelta
Returns:
The fractional gain or loss over the interval.
"""
end_date = datetime.today().date()
if offset is not None:
end_date -= offset
start_date = end_date - interval
sp500_index = Index('^GSPC')
sp500_time_series = sp500_index.get_historical_data(start_date,end_date)
sp500_close = sp500_time_series['close']
sp500_close_smooth = sp500_close.ewm(span=5).mean()
sp500_end_of_interval = round(sp500_close_smooth.values[-1],2)
sp500_start_of_interval = round(sp500_close_smooth.values[0],2)
sp500_gain_during_interval = round(sp500_end_of_interval / sp500_start_of_interval,4)
if output:
print("Value start of interval: ",sp500_start_of_interval)
print("Value end of interval: ",sp500_end_of_interval)
print("Approximate gain: ",sp500_gain_during_interval)
print("")
plt.plot(sp500_close.values,'.',label='Close',color=plt.cm.viridis(.4),markerfacecolor=plt.cm.viridis(.9),markersize=12)
plt.plot(sp500_close_smooth.values,'-',label='5-day EWMA',color=plt.cm.viridis(0.3))
plt.title('S&P 500 Index')
plt.xlabel('Trading days')
plt.ylabel('Close Price')
plt.legend()
return sp500_gain_during_interval
# -
get_sp500_gain_for_interval(TIMEDELTA_QUARTER,offset=None,output=True);
# We will need to label our data so that we can provide the labels along with training data to our neural network. The labels in this case are generated by looking at the performance of a particular stock against the "market". Since the S&P 500 is a good representation of the US market, we will compare last quarter's performance of each stock that we will use to train the network with that of the S&P 500 index.
# +
# %%execute_and_save tmp_benchmark_label
def get_stock_label_func(p_interval,p_offset):
""" Generates a function that returns a stock label
Args:
p_interval: The prediction interval as a datetime.timedelta
p_offset: The offset of d_interval relative to today as a datetime.timedelta
Returns:
A function that can be called (for a specified stock) to get the stock label
"""
ref_value = get_sp500_gain_for_interval(p_interval,p_offset,output=False)
def get_stock_label(symbol,output=False):
""" Generates a stock label for training and/or validation dataset
Raises:
LookupError: If the stock could not be found
Returns:
An integer value (0 or 1) indicating the stock label
"""
end_date = datetime.today().date()
if p_offset is not None:
end_date -= p_offset
start_date = end_date - p_interval
try:
close_price = stock_data[symbol].get_historical_prices(start_date,end_date)['close']
except:
close_price = Stock(symbol).get_historical_prices(start_date,end_date)['close']
close_price = close_price.ewm(span=3).mean()
stock_gain = close_price.values[-1] / close_price.values[0]
stock_relative_gain = round( (stock_gain) / ref_value,4)
stock_label = 0 if stock_relative_gain < 1 else 1
if output:
print("Gain during interval: ",round(stock_gain,4))
print("Reference value: ",ref_value)
print("Gain relative to reference value: ",stock_relative_gain)
print("Label: ",stock_label)
return stock_label
return get_stock_label
# -
test_get_stock_label = get_stock_label_func(p_interval=TIMEDELTA_QUARTER,p_offset=None)
print('Label for AAPL: ',test_get_stock_label('AAPL'))
print('Label for KSS: ',test_get_stock_label('KSS'))
print('Label for MSFT: ',test_get_stock_label('MSFT'))
print('Label for WELL: ',test_get_stock_label('WELL'))
# #### Stock Features: Categorical
# +
# %%execute_and_save tmp_predictor_categorical
def get_stock_cat_features_func(d_interval,d_offset):
""" Generates a function that returns categorical features for a stock
Args:
d_interval: The data interval as a datetime.timedelta (e.g., 6*TIMEDELTA_QUARTER for 6 quarters of data)
d_offset: The offset of d_interval relative to today as a datetime.timedelta
Returns:
A tuple consisting of array that specifies which categorical feature are to be embedded (as opposed to
stand-alone features) and a function that can be called to get categorical features for a stock. The
array should include the embedding dimension for the feature, or 0 if it is not to be embedded.
"""
# Get list of sectors and map each sector to an index (normalized)
sector_list = np.array(['Energy',
'Consumer Cyclical',
'Real Estate',
'Utilities',
'Industrials',
'Basic Materials',
'Technology',
'Healthcare',
'Financial Services',
'Consumer Defensive'])
industry_list = np.array(['Agriculture',
'Insurance - Life',
'Medical Diagnostics & Research',
'Online Media',
'Oil & Gas - E&P',
'Homebuilding & Construction',
'Oil & Gas - Drilling',
'Oil & Gas - Refining & Marketing',
'Advertising & Marketing Services',
'Utilities - Regulated',
'Consulting & Outsourcing',
'Autos',
'Travel & Leisure',
'Oil & Gas - Integrated',
'Brokers & Exchanges',
'Application Software',
'Manufacturing - Apparel & Furniture',
'Medical Devices',
'Retail - Apparel & Specialty',
'Oil & Gas - Services',
'Consumer Packaged Goods',
'Insurance - Property & Casualty',
'Drug Manufacturers',
'Real Estate Services',
'Airlines',
'Insurance',
'Farm & Construction Machinery',
'Semiconductors',
'Medical Distribution',
'Steel',
'Restaurants',
'Waste Management',
'Entertainment',
'Chemicals',
'REITs',
'Insurance - Specialty',
'Metals & Mining',
'Retail - Defensive',
'Biotechnology',
'Conglomerates',
'Utilities - Independent Power Producers',
'Building Materials',
'Health Care Plans',
'Tobacco Products',
'Oil & Gas - Midstream',
'Transportation & Logistics',
'Business Services',
'Truck Manufacturing',
'Beverages - Non-Alcoholic',
'Personal Services',
'Banks',
'Medical Instruments & Equipment',
'Industrial Distribution',
'Asset Management',
'Forest Products',
'Industrial Products',
'Communication Equipment',
'Packaging & Containers',
'Credit Services',
'Engineering & Construction',
'Computer Hardware',
'Aerospace & Defense',
'Beverages - Alcoholic',
'Health Care Providers',
'Communication Services',
'Employment Services'])
sector_dict = { sector : i for i, sector in enumerate(sector_list)}
industry_dict = { industry : i for i, industry in enumerate(industry_list)}
# SP500 range is on the order of USD 1B to USD 1T, scale accordingly
MIN_MARKET_CAP = 1.0e9
MAX_MARKET_CAP = 1.0e12
# For the specified d_offset we will make a cyclic label corresponding
# to the month of the year (1-12) using sine and cosine functions
end_date = datetime.today().date()
if d_offset is not None:
end_date -= d_offset
# Encoding which month (fractional) the data ends. This is universal
# in that it will work for any intervals and offsets of interest.
month_decimal = end_date.month + end_date.day/30.0;
month_angle = 2*math.pi*month_decimal/12.0
month_x = math.cos(month_angle)
month_y = math.sin(month_angle)
# The feature structure (# of embeddings for each feature or 0 if not to be embedded)
cat_feature_embeddings = [len(sector_list)+1, len(industry_list)+1, 0, 0]
def get_stock_cat_features(symbol):
""" Gets categorical features associated with a paticular stock
Args:
symbol: A stock ticker symbol such as 'AAPL' or 'T'
Raises:
LookupError: If any categorical feature is unavailable of NaN for the stock.
Returns:
Categorical stock features as an array of M x 1 values (for M features). Categorical
features to be embedded are appear first in the returned array
"""
try:
profile = stock_data[symbol].get_company_profile()
except:
profile = Stock(symbol).get_company_profile()
sector = profile.loc[symbol,'sector']
industry = profile.loc[symbol,'industry']
try:
sector_feature = sector_dict[sector]
except:
sector_feature = len(sector_list)
try:
industry_feature = industry_dict[industry]
except:
industry_feature = len(industry_list)
# Get market capitalization corresponding to d_offset
if d_offset is None:
quarter_offset = 0
else:
quarter_offset = int(d_offset / TIMEDELTA_QUARTER)
# Get the "latest" key metrics as of the data interval
try:
key_metrics = stock_data[symbol].get_key_metrics(quarters=1,offset=quarter_offset)
except:
key_metrics = Stock(symbol).get_key_metrics(quarters=1,offset=quarter_offset)
market_cap = key_metrics['Market Cap'][0]
# Scalar value (approx 0-1) corresponding to market capitalization
market_cap_feature = math.log(float(market_cap)/MIN_MARKET_CAP,MAX_MARKET_CAP/MIN_MARKET_CAP)
features = np.array( [sector_feature, industry_feature, market_cap_feature, month_x, month_y],dtype='float32')
if np.isnan(features).any():
raise LookupError
return features
return cat_feature_embeddings, get_stock_cat_features
# -
_, test_get_stock_cat_features = get_stock_cat_features_func(d_interval=4*TIMEDELTA_QUARTER,d_offset=TIMEDELTA_QUARTER)
test_get_stock_cat_features('AMZN')
test_get_stock_cat_features('WELL')
# #### Stock Features: Daily Data
# +
# %%execute_and_save tmp_predictor_daily
def get_stock_daily_features_func(d_interval,d_offset):
""" Generates a function that returns daily features for a stock
Args:
d_interval: The data interval as a datetime.timedelta (e.g., 6*TIMEDELTA_QUARTER for 6 quarters of data)
d_offset: The offset of d_interval relative to today as a datetime.timedelta
Returns:
A function that can be called to get daily features for a stock
"""
end_date = datetime.today().date()
if d_offset is not None:
end_date -= d_offset
start_date = end_date - d_interval
# Th S&P 500 index will have a closing value for every trading day. Each of the stocks
# should also have the same number of values unless they were suspended and didn't trade or
# recently became public.
trading_day_count = len(Index('^GSPC').get_historical_data(start_date,end_date))
def get_stock_daily_features(symbol,output=False):
""" Gets daily features associated with a paticular stock
Args:
symbol: A stock ticker symbol such as 'AAPL' or 'T'
Raises:
LookupError: If any categorical feature is unavailable of NaN for the stock.
Returns:
Daily stock features as an array of M x N values (for M features with N values)
"""
try:
historical_data = stock_data[symbol].get_historical_prices(start_date,end_date)
except:
historical_data = Stock(symbol).get_historical_prices(start_date,end_date)
# Smooth and normalize closing price relative to initial price for data set
close_price = historical_data['close'].ewm(span=5).mean()
close_price = close_price / close_price.iat[0]
close_price = np.log10(close_price)
# Smooth and normalize volume relative to average volume
average_volume = historical_data['volume'].mean()
volume = historical_data['volume'].ewm(span=5).mean()
volume = volume / average_volume
volume = np.log10(volume+1e-6)
# Ensure equal lengths of data (nothing missing)
if len(volume) != len(close_price):
raise LookupError
# Ensure we have the right number of data points for the period
if len(close_price) != trading_day_count:
raise LookupError
features = np.array([close_price, volume],dtype='float32')
if np.isnan(features).any():
raise LookupError
return features
return get_stock_daily_features
# -
test_get_stock_daily_features = get_stock_daily_features_func(1*TIMEDELTA_QUARTER,TIMEDELTA_QUARTER)
test_get_stock_daily_features('AAPL')
# #### Stock Features: Quarterly Data
# +
# %%execute_and_save tmp_predictor_quarterly
def get_stock_quarterly_features_func(d_interval,d_offset):
""" Generates a function that returns quarterly features for a stock
Args:
d_interval: The data interval as a datetime.timedelta (e.g., 6*TIMEDELTA_QUARTER for 6 quarters of data)
d_offset: The offset of d_interval relative to today
Returns:
A function that can be called to get quarterly features for a stock
"""
# Quarterly features can only be used if prediction intervals
if d_interval < TIMEDELTA_QUARTER:
raise ValueError("The specified data interval is less than one quarter")
end_date = datetime.today().date()
if d_offset is not None:
end_date -= d_offset
start_date = end_date - d_interval
quarter_count = int(d_interval / TIMEDELTA_QUARTER)
if d_offset is None:
quarter_offset = 0
else:
quarter_offset = int(d_offset / TIMEDELTA_QUARTER)
price_to_earnings_scaler = MinMaxScaler()
price_to_sales_scaler = MinMaxScaler()
price_to_free_cash_flow_scaler = MinMaxScaler()
dividend_yield_scaler = MinMaxScaler()
price_to_earnings_scaler.fit_transform(np.array([0,200]).reshape(-1, 1))
price_to_sales_scaler.fit_transform(np.array([0,200]).reshape(-1, 1))
price_to_free_cash_flow_scaler.fit_transform(np.array([0,200]).reshape(-1, 1))
dividend_yield_scaler.fit_transform(np.array([0,1]).reshape(-1, 1))
def get_stock_quarterly_features(symbol):
""" Gets quarterly features associated with a paticular stock
Args:
symbol: A stock ticker symbol such as 'AAPL' or 'T'
Raises:
LookupError: If any categorical feature is unavailable of NaN for the stock.
Returns:
Quarterly stock features as an array of M x N values (for M features and N values)
"""
try:
key_metrics = stock_data[symbol].get_key_metrics(quarter_count,quarter_offset)
except:
key_metrics = Stock(symbol).get_key_metrics(quarter_count,quarter_offset)
key_metrics['PE ratio'] = price_to_earnings_scaler.transform(key_metrics['PE ratio'].values.reshape(-1,1))
key_metrics['Price to Sales Ratio'] = price_to_sales_scaler.transform(key_metrics['Price to Sales Ratio'].values.reshape(-1,1))
key_metrics['PFCF ratio'] = price_to_free_cash_flow_scaler.transform(key_metrics['PFCF ratio'].values.reshape(-1,1))
key_metrics['Dividend Yield'] = dividend_yield_scaler.transform(key_metrics['Dividend Yield'].values.reshape(-1,1))
try:
financials = stock_data[symbol].get_income_statement(quarter_count,quarter_offset)
except:
financials = Stock(symbol).get_income_statement(quarter_count,quarter_offset)
# Apply scaling for diluted EPS (we want growth relative to t=0)
financials['EPS Diluted'] = ( financials['EPS Diluted'].astype(dtype='float32') / float(financials['EPS Diluted'].iat[0]) )
features = np.array([
key_metrics['PE ratio'],
key_metrics['Price to Sales Ratio'],
key_metrics['PFCF ratio'],
key_metrics['Dividend Yield'],
financials['EPS Diluted'],
financials['Revenue Growth'],
],dtype='float32')
if np.isnan(features).any():
raise LookupError
return features
return get_stock_quarterly_features
# -
test_get_stock_quarterly_features = get_stock_quarterly_features_func(4*TIMEDELTA_QUARTER,None)
test_get_stock_quarterly_features('AAPL')
test_get_stock_quarterly_features('T')
# ### Generate Training and Testing Data Sets
#
# We have created a custom dataset class that will accept references to the functions we created earlier for extracting categorical, daily and quarterly features and generating a label for a particular stock. To use the dataset, we specify a list of stocks and references to the functions. We are going to create multiple datasets: training and testing datasets for a number of folds.
# !pygmentize model/dataset.py
# #### Generating Datasets: Offsets to Incorporate Seasonality
# We are going to need to specify offsets for our dataset. That is, for a given stock, we will want to generate training data with offsets corresponding to predictions for each of the 4 quarters in order to address seasonality. The total number of training samples will therefore be the number of training stocks multiplied by the number of offsets. Here we pick 12 offsets, one for each month of the year. This will allow for seasonality but will also allow us to generate predictions between fiscal quarter cutoffs.
# +
# An offset of zero corresponds to using the most recent p_interval of data
# for labeling and the preceding d_interval worth of data for features.
# Snap to nearest (last) month (not a strict requirement, but gives us
# some reproducibility for training purposes)
last_offset = timedelta(days=datetime.today().day)
# Produce 12 offsets one month apart from each other
one_month = timedelta(days=30)
offsets = [ last_offset + one_month*i for i in range(12) ]
# -
from model.dataset import StockDataset
# #### Generating Datasets: K-fold Sets for Cross-Validation
# We will split our stock list into K sublists to generate K training and testing datasets, each of which will include offsets as discussed above.
# +
# Prepare list of available stock tickers
stock_list = np.array(list(stock_data.keys()))
stock_count = len(stock_list)
# Split tickers into k-folds (here we use 3 folds) for cross validation
k = 3
k_fold_size = math.ceil(stock_count / k)
k_fold_indices = [ [i, i+k_fold_size] for i in range(0,stock_count,k_fold_size) ]
# List of dataset objects associated with each fold
k_fold_ds_list = list()
for i, fold in enumerate(k_fold_indices):
print(f'Generating dataset for fold {i+1}/{k}')
start, end = k_fold_indices[i][0], k_fold_indices[i][1]
k_fold_ds = StockDataset.from_data(stock_list[start:end],
p_interval=TIMEDELTA_QUARTER,
d_interval=4*TIMEDELTA_QUARTER,
offsets=offsets,
c_features_func_gen=get_stock_cat_features_func,
d_features_func_gen=get_stock_daily_features_func,
q_features_func_gen=get_stock_quarterly_features_func,
label_func_gen=get_stock_label_func,
output=True)
k_fold_ds_list.append(k_fold_ds)
# -
# Concatenate datasets to form new K new train/test sets
for i in range(k):
# Each fold becomes the test dataset
test_ds = k_fold_ds_list[i]
test_ds.to_file(f'data/test-{i}.npz')
# The other folds form the training dataset
train_ds = StockDataset.concat(k_fold_ds_list[:i] + k_fold_ds_list[i+1:])
train_ds.to_file(f'data/train-{i}.npz')
# Generate a seperate full dataset for final model training
train_data_full = StockDataset.concat(k_fold_ds_list)
train_data_full.to_file(f'data/train-full.npz')
# +
# Read our training datasets from disk and check for consistency
datasets = np.append(np.arange(0,k),['full'])
print('Fold Type Samples Features Pos/Neg')
for i in datasets:
for data_type in ['train','test']:
try:
ds = StockDataset.from_file(f'data/{data_type}-{i}.npz')
except:
continue
sample_count = len(ds)
categorical_count = ds.get_categorical_feature_count()
daily_count = ds.get_daily_feature_count()
quarterly_count = ds.get_quarterly_feature_count()
pos_count = ds[:][1].sum()
neg_count = sample_count - pos_count
features = f'{categorical_count}/{daily_count}/{quarterly_count}'
print(f'%4s' % f'{i}' + ' %-5s' % data_type + '%8s' % f'{sample_count}' + '%10s' % f'{features}' + '%11s' % f'{pos_count}/{neg_count}')
# +
# Plot the month encoding to ensure that was correctly
# captured in the aggregate dataset. For our chosen offsets,
# we should see 12 equally spaced points on a unit circle
ds = StockDataset.from_file(f'data/train-full.npz')
x = [train_ds[i][0][0][-2] for i in range(len(train_ds)) ]
y = [train_ds[i][0][0][-1] for i in range(len(train_ds)) ]
plt.figure(figsize=(5,5))
plt.title('Cyclical Offset Encoding')
plt.xlabel('Month X')
plt.ylabel('Month Y')
plt.plot(x,y,marker='.',linestyle='');
# -
# ### Automation
# Concatenate the code associated with the data generator functions (4 functions) that generate labels and categorical, daily and quarterly data for each stock. This will allow us to use the code later for our prediction engine without having to copy-paste from the notebook.
# Generate a file for the categorical, daily and quarterly functions to be used by the prediction code
# !cat tmp_imports tmp_predictor_categorical tmp_predictor_daily tmp_predictor_quarterly > ./model/predictor_helper.py
# !cat tmp_imports tmp_benchmark_sp500_gain tmp_benchmark_label > ./model/benchmark_helper.py
# Generate a file for benchmarking (gets S&P 500 gains)
# !rm tmp_imports tmp_predictor_categorical tmp_predictor_daily tmp_predictor_quarterly
# !rm tmp_benchmark_sp500_gain tmp_benchmark_label
|
data_processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ---
#
# _You are currently looking at **version 1.1** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-social-network-analysis/resources/yPcBs) course resource._
#
# ---
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# # Assignment 3
#
# In this assignment you will explore measures of centrality on two networks, a friendship network in Part 1, and a blog network in Part 2.
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ## Part 1
#
# Answer questions 1-4 using the network `G1`, a network of friendships at a university department. Each node corresponds to a person, and an edge indicates friendship.
#
# *The network has been loaded as networkx graph object `G1`.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
import networkx
friendships = networkx.read_gml('friendships.gml')
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
DEGREE_CENTRALITY = networkx.degree_centrality(friendships)
CLOSENESS_CENTRALITY = networkx.closeness_centrality(friendships)
BETWEENNESS_CENTRALITY = networkx.betweenness_centrality(friendships)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 1
#
# Find the degree centrality, closeness centrality, and normalized betweeness centrality (excluding endpoints) of node 100.
#
# *This function should return a tuple of floats `(degree_centrality, closeness_centrality, betweenness_centrality)`.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_one():
NODE = 100
return (DEGREE_CENTRALITY[NODE],
CLOSENESS_CENTRALITY[NODE], BETWEENNESS_CENTRALITY[NODE])
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# <br>
# #### For Questions 2, 3, and 4, use one of the covered centrality measures to rank the nodes and find the most appropriate candidate.
# <br>
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 2
#
# Suppose you are employed by an online shopping website and are tasked with selecting one user in network G1 to send an online shopping voucher to. We expect that the user who receives the voucher will send it to their friends in the network. You want the voucher to reach as many nodes as possible. The voucher can be forwarded to multiple users at the same time, but the travel distance of the voucher is limited to one step, which means if the voucher travels more than one step in this network, it is no longer valid. Apply your knowledge in network centrality to select the best candidate for the voucher.
#
# *This function should return an integer, the name of the node.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def largest_node(centrality):
"""gets the node with the best (largest score)
Returns:
int: name of the node with the best score
"""
return list(reversed(sorted((value, node)
for (node, value) in centrality.items())))[0][1]
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_two():
return largest_node(DEGREE_CENTRALITY)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 3
#
# Now the limit of the voucher’s travel distance has been removed. Because the network is connected, regardless of who you pick, every node in the network will eventually receive the voucher. However, we now want to ensure that the voucher reaches the nodes in the lowest average number of hops.
#
# How would you change your selection strategy? Write a function to tell us who is the best candidate in the network under this condition.
#
# *This function should return an integer, the name of the node.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_three():
"""Returns the node with the best closeness centrality"""
return largest_node(CLOSENESS_CENTRALITY)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 4
#
# Assume the restriction on the voucher’s travel distance is still removed, but now a competitor has developed a strategy to remove a person from the network in order to disrupt the distribution of your company’s voucher. Identify the single riskiest person to be removed under your competitor’s strategy?
#
# *This function should return an integer, the name of the node.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_four():
"""the node with the highest betweenness centrality"""
return largest_node(BETWEENNESS_CENTRALITY)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ## Part 2
#
# `G2` is a directed network of political blogs, where nodes correspond to a blog and edges correspond to links between blogs. Use your knowledge of PageRank and HITS to answer Questions 5-9.
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
blogs = networkx.read_gml('blogs.gml')
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 5
#
# Apply the Scaled Page Rank Algorithm to this network. Find the Page Rank of node 'realclearpolitics.com' with damping value 0.85.
#
# *This function should return a float.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
PAGE_RANK = networkx.pagerank(blogs)
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_five():
"""Page Rank of realclearpolitics.com"""
return PAGE_RANK['realclearpolitics.com']
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 6
#
# Apply the Scaled Page Rank Algorithm to this network with damping value 0.85. Find the 5 nodes with highest Page Rank.
#
# *This function should return a list of the top 5 blogs in desending order of Page Rank.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def top_five(ranks):
"""gets the top-five blogs by rank"""
top = list(reversed(sorted((rank, node)
for node, rank in ranks.items())))[:5]
return [node for rank, node in top]
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_six():
"""Top 5 nodes by page rank"""
return top_five(PAGE_RANK)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 7
#
# Apply the HITS Algorithm to the network to find the hub and authority scores of node 'realclearpolitics.com'.
#
# *Your result should return a tuple of floats `(hub_score, authority_score)`.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
HUBS, AUTHORITIES = networkx.hits(blogs)
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_seven():
"""HITS score for realclearpolitics.com"""
return HUBS['realclear<EMAIL>'], AUTHORITIES['<EMAIL>clearpolitics.com']
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 8
#
# Apply the HITS Algorithm to this network to find the 5 nodes with highest hub scores.
#
# *This function should return a list of the top 5 blogs in desending order of hub scores.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_eight():
"""Top five blogs by hub scores"""
return top_five(HUBS)
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### Question 9
#
# Apply the HITS Algorithm to this network to find the 5 nodes with highest authority scores.
#
# *This function should return a list of the top 5 blogs in desending order of authority scores.*
# + ein.tags="worksheet-0" slideshow={"slide_type": "-"}
def answer_nine():
"""the top 5 blogs by authorities score"""
return top_five(AUTHORITIES)
|
5_social_networks/3_node_importance/assignment_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="jJQd-lY4CvOx"
import numpy as np
import xgboost as xgb
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.datasets import dump_svmlight_file
### commented out because of warning:
#from sklearn.externals import joblib
### above replaced with this:
import joblib
from sklearn.metrics import precision_score
# + id="I0FvTzvTD7J4"
# First you load the dataset from sklearn, where X will be the data, y – the class labels:
iris = datasets.load_iris()
X = iris.data
y = iris.target
# + id="qUzmx4oEGESP"
# split the data into train and test sets with 80-20% split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# + id="8dHCJ5BdGsbs"
# use DMatrix for xgboost, create the Xgboost specific DMatrix data
# format from the numpy array. Xgboost can work with numpy arrays directly, load data from svmlight files and other formats
dtrain = xgb.DMatrix(X_train, label=y_train)
dtest = xgb.DMatrix(X_test, label=y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="j_9E7D4wHOx6" outputId="3d48d60c-95b1-4f8e-93fa-1eefa6038213"
# use svmlight file for xgboost, If you want to use svmlight for less memory
# consumption, first dump the numpy array into svmlight format and then just pass the filename to DMatrix:
dump_svmlight_file(X_train, y_train, 'dtrain.svm', zero_based=True)
dump_svmlight_file(X_test, y_test, 'dtest.svm', zero_based=True)
dtrain_svm = xgb.DMatrix('dtrain.svm')
dtest_svm = xgb.DMatrix('dtest.svm')
# + id="IDwmjtGdH2ps"
# set xgboost params
param = {
'max_depth': 3, # the maximum depth of each tree
'eta': 0.3, # the training step for each iteration
'silent': 1, # logging mode - quiet
'objective': 'multi:softprob', # error evaluation for multiclass training
'num_class': 3} # the number of classes that exist in this datset
num_round = 20 # the number of training iterations
# + id="NxLPZjEmILZM"
#------------- numpy array ------------------
# training and testing - numpy matrices
bst = xgb.train(param, dtrain, num_round)
preds = bst.predict(dtest)
# + colab={"base_uri": "https://localhost:8080/"} id="S8eD2EboIXi6" outputId="63dd02c3-02d6-43e2-f063-2187ac1826f2"
# extracting most confident predictions
best_preds = np.asarray([np.argmax(line) for line in preds])
print("Numpy array precision:", precision_score(y_test, best_preds, average='macro'))
# + id="f90Ynn1tIvvq"
# ------------- svm file ---------------------
# training and testing - svm file
bst_svm = xgb.train(param, dtrain_svm, num_round)
preds = bst.predict(dtest_svm)
# + colab={"base_uri": "https://localhost:8080/"} id="YYEkLSLhI2oo" outputId="4d07a011-f0b3-4391-dc9a-1098d328dca1"
# extracting most confident predictions
best_preds_svm = [np.argmax(line) for line in preds]
print("Svm file precision:",precision_score(y_test, best_preds_svm, average='macro'))
# --------------------------------------------
# + id="29lFsTJ_JqoW"
# dump the models
bst.dump_model('dump.raw.txt')
bst_svm.dump_model('dump_svm.raw.txt')
# + colab={"base_uri": "https://localhost:8080/"} id="nNowhNq1LnUa" outputId="a7fb0e45-a97b-4493-c4cc-efb9cc95358a"
# save the models for later
joblib.dump(bst, 'bst_model.pkl', compress=True)
joblib.dump(bst_svm, 'bst_svm_model.pkl', compress=True)
|
xgboost/xgboost_jupyter_example_001.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import pickle
from sklearn.preprocessing import StandardScaler, MinMaxScaler
# +
# 之前的测试数据
Test_first=np.load('fft_data14.npy')
Test_first = Test_first[:7,:2251]
Test_first = pd.DataFrame(Test_first)
# print(Test_first.shape)
with open('norm_T_origin.pickle', 'rb') as f:
Test_last = pickle.load(f)
Test_last = pd.DataFrame(Test_last)
Test_last = Test_last.iloc[7:,:]
# print(Test_last.shape)
Test_ori = Test_first.append(Test_last,ignore_index=True)
print(Test_ori.shape)
with open('T_origin-19.pickle', 'rb') as f:
Test = pickle.load(f)
Test = pd.DataFrame(Test)
scaler = MinMaxScaler()
Test = scaler.fit_transform(Test.iloc[:,:-1])
# zif-7
Test = pd.DataFrame(Test)
Test_second = Test.iloc[3:5,:]
print(Test_second.shape)
#zif-9
Test_third = np.load('data/4_zif9.npy')
Test_third = pd.DataFrame(Test_third)
Test_third = Test_third.iloc[:,:2251]
frames = [Test_ori,Test_second,Test_third]
Test = pd.concat(frames)
print(Test.shape)
# -
Test_first = Test.iloc[:2,:]
Test_last = Test.iloc[3:,:]
Test = Test_first.append(Test_last,ignore_index=True)
print(Test.shape)
# +
# Test.to_csv('test_data_19.csv')
# +
# label
with open('T_origin-19.pickle', 'rb') as f:
Test = pickle.load(f)
Test = pd.DataFrame(Test)
labelzif67 = Test.iloc[:3,-1]
labelzif7 = Test.iloc[3:5,-1]
labelzif71and8 = Test.iloc[5:12,-1]
labelzif9 = Test.iloc[12:16,-1]
labelzif90 = Test.iloc[16:,-1]
frames = [labelzif67,labelzif71and8,labelzif90,labelzif7,labelzif9]
label = pd.concat(frames)
# -
print(label)
# +
# label.to_csv('label_19.csv')
# -
|
data_processing/append_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Approximate Training
#
# Recall content of the last section. The core feature of the skip-gram model is the use of softmax operations to compute the conditional probability of generating context word $w_o$ based on the given central target word $w_c$.
#
# $$\mathbb{P}(w_o \mid w_c) = \frac{\text{exp}(\boldsymbol{u}_o^\top \boldsymbol{v}_c)}{ \sum_{i \in \mathcal{V}} \text{exp}(\boldsymbol{u}_i^\top \boldsymbol{v}_c)}.$$
#
# The logarithmic loss corresponding to the conditional probability is given as
#
# $$-\log \mathbb{P}(w_o \mid w_c) =
# -\boldsymbol{u}_o^\top \boldsymbol{v}_c + \log\left(\sum_{i \in \mathcal{V}} \text{exp}(\boldsymbol{u}_i^\top \boldsymbol{v}_c)\right).$$
#
#
# Because the softmax operation has considered that the context word could be any word in the dictionary $\mathcal{V}$, the loss mentioned above actually includes the sum of the number of items in the dictionary size. From the last section, we know that for both the skip-gram model and CBOW model, because they both get the conditional probability using a softmax operation, the gradient computation for each step contains the sum of the number of items in the dictionary size. For larger dictionaries with hundreds of thousands or even millions of words, the overhead for computing each gradient may be too high. In order to reduce such computational complexity, we will introduce two approximate training methods in this section: negative sampling and hierarchical softmax. Since there is no major difference between the skip-gram model and the CBOW model, we will only use the skip-gram model as an example to introduce these two training methods in this section.
#
#
#
# ## Negative Sampling
#
# Negative sampling modifies the original objective function. Given a context window for the central target word $w_c$, we will treat it as an event for context word $w_o$ to appear in the context window and compute the probability of this event from
#
# $$\mathbb{P}(D=1\mid w_c, w_o) = \sigma(\boldsymbol{u}_o^\top \boldsymbol{v}_c),$$
#
# Here, the $\sigma$ function has the same definition as the sigmoid activation function:
#
# $$\sigma(x) = \frac{1}{1+\exp(-x)}.$$
#
# We will first consider training the word vector by maximizing the joint probability of all events in the text sequence. Given a text sequence of length $T$, we assume that the word at time step $t$ is $w^{(t)}$ and the context window size is $m$. Now we consider maximizing the joint probability
#
# $$ \prod_{t=1}^{T} \prod_{-m \leq j \leq m,\ j \neq 0} \mathbb{P}(D=1\mid w^{(t)}, w^{(t+j)}).$$
#
# However, the events included in the model only consider positive examples. In this case, only when all the word vectors are equal and their values approach infinity can the joint probability above be maximized to 1. Obviously, such word vectors are meaningless. Negative sampling makes the objective function more meaningful by sampling with an addition of negative examples. Assume that event $P$ occurs when context word $w_o$ to appear in the context window of central target word $w_c$, and we sample $K$ words that do not appear in the context window according to the distribution $\mathbb{P}(w)$ to act as noise words. We assume the event for noise word $w_k$($k=1, \ldots, K$) to not appear in the context window of central target word $w_c$ is $N_k$. Suppose that events $P and N_1, \ldots, N_K$ for both positive and negative examples are independent of each other. By considering negative sampling, we can rewrite the joint probability above, which only considers the positive examples, as
#
#
# $$ \prod_{t=1}^{T} \prod_{-m \leq j \leq m,\ j \neq 0} \mathbb{P}(w^{(t+j)} \mid w^{(t)}),$$
#
# Here, the conditional probability is approximated to be
# $$ \mathbb{P}(w^{(t+j)} \mid w^{(t)}) =\mathbb{P}(D=1\mid w^{(t)}, w^{(t+j)})\prod_{k=1,\ w_k \sim \mathbb{P}(w)}^K \mathbb{P}(D=0\mid w^{(t)}, w_k).$$
#
#
# Let the text sequence index of word $w^{(t)}$ at time step $t$ be $i_t$ and $h_k$ for noise word $w_k$ in the dictionary. The logarithmic loss for the conditional probability above is
#
# $$
# \begin{aligned}
# -\log\mathbb{P}(w^{(t+j)} \mid w^{(t)})
# =& -\log\mathbb{P}(D=1\mid w^{(t)}, w^{(t+j)}) - \sum_{k=1,\ w_k \sim \mathbb{P}(w)}^K \log\mathbb{P}(D=0\mid w^{(t)}, w_k)\\
# =&- \log\, \sigma\left(\boldsymbol{u}_{i_{t+j}}^\top \boldsymbol{v}_{i_t}\right) - \sum_{k=1,\ w_k \sim \mathbb{P}(w)}^K \log\left(1-\sigma\left(\boldsymbol{u}_{h_k}^\top \boldsymbol{v}_{i_t}\right)\right)\\
# =&- \log\, \sigma\left(\boldsymbol{u}_{i_{t+j}}^\top \boldsymbol{v}_{i_t}\right) - \sum_{k=1,\ w_k \sim \mathbb{P}(w)}^K \log\sigma\left(-\boldsymbol{u}_{h_k}^\top \boldsymbol{v}_{i_t}\right).
# \end{aligned}
# $$
#
# Here, the gradient computation in each step of the training is no longer related to the dictionary size, but linearly related to $K$. When $K$ takes a smaller constant, the negative sampling has a lower computational overhead for each step.
#
#
# ## Hierarchical Softmax
#
# Hierarchical softmax is another type of approximate training method. It uses a binary tree for data structure, with the leaf nodes of the tree representing every word in the dictionary $\mathcal{V}$.
#
# 
#
#
# We assume that $L(w)$ is the number of nodes on the path (including the root and leaf nodes) from the root node of the binary tree to the leaf node of word $w$. Let $n(w,j)$ be the $j$th node on this path, with the context word vector $\mathbf{u}_{n(w,j)}$. We use Figure 10.3 as an example, so $L(w_3) = 4$. Hierarchical softmax will approximate the conditional probability in the skip-gram model as
#
# $$\mathbb{P}(w_o \mid w_c) = \prod_{j=1}^{L(w_o)-1} \sigma\left( [\![ n(w_o, j+1) = \text{leftChild}(n(w_o,j)) ]\!] \cdot \mathbf{u}_{n(w_o,j)}^\top \mathbf{v}_c\right),$$
#
# Here the $\sigma$ function has the same definition as the sigmoid activation function, and $\text{leftChild}(n)$ is the left child node of node $n$. If $x$ is true, $[\![x]\!] = 1$; otherwise $[\![x]\!] = -1$.
# Now, we will compute the conditional probability of generating word $w_3$ based on the given word $w_c$ in Figure 10.3. We need to find the inner product of word vector $\mathbf{v}_c$ (for word $w_c$) and each non-leaf node vector on the path from the root node to $w_3$. Because, in the binary tree, the path from the root node to leaf node $w_3$ needs to be traversed left, right, and left again (the path with the bold line in Figure 10.3), we get
#
# $$\mathbb{P}(w_3 \mid w_c) = \sigma(\mathbf{u}_{n(w_3,1)}^\top \mathbf{v}_c) \cdot \sigma(-\mathbf{u}_{n(w_3,2)}^\top \mathbf{v}_c) \cdot \sigma(\mathbf{u}_{n(w_3,3)}^\top \mathbf{v}_c).$$
#
# Because $\sigma(x)+\sigma(-x) = 1$, the condition that the sum of the conditional probability of any word generated based on the given central target word $w_c$ in dictionary $\mathcal{V}$ be 1 will also suffice:
#
# $$\sum_{w \in \mathcal{V}} \mathbb{P}(w \mid w_c) = 1.$$
#
# In addition, because the order of magnitude for $L(w_o)-1$ is $\mathcal{O}(\text{log}_2|\mathcal{V}|)$, when the size of dictionary $\mathcal{V}$ is large, the computational overhead for each step in the hierarchical softmax training is greatly reduced compared to situations where we do not use approximate training.
#
# ## Summary
#
# * Negative sampling constructs the loss function by considering independent events that contain both positive and negative examples. The gradient computational overhead for each step in the training process is linearly related to the number of noise words we sample.
# * Hierarchical softmax uses a binary tree and constructs the loss function based on the path from the root node to the leaf node. The gradient computational overhead for each step in the training process is related to the logarithm of the dictionary size.
#
# ## Exercises
#
# * Before reading the next section, think about how we should sample noise words in negative sampling.
# * What makes the last formula in this section hold?
# * How can we apply negative sampling and hierarchical softmax in the skip-gram model?
#
# ## Scan the QR Code to [Discuss](https://discuss.mxnet.io/t/2386)
#
# 
|
02_word_embedding/approx-training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # DATA 601 Final Project
# # Severe Thunderstorm Climatology vs Sunspot Number 2019-2020
# ## Python weather analysis application that compares the occurrence of severe thunderstorm activity occurring over the continental Unites States to daily international sunspot number measurements.
# +
#Load modules and invoke WxData API
from __future__ import print_function, division
import numpy as np
import pandas as pd
import sys
#Function that builds a storm report listing for each semi-monthly period between November 2019 and April 2020.
def dataframe_strmrpt(wxdata_df):
maxmag = np.amax(wxdata_df['Magnitude']) #Extract maximum wind gust magnitude for semi-monthly period
meanmag = np.mean(wxdata_df['Magnitude'])
medmag = np.median(wxdata_df['Magnitude'])
strmrpt_nr = len(wxdata_df.Magnitude) #Count number of storm reports for semi-monthly period
return maxmag, meanmag, medmag, strmrpt_nr #Return storm report dataframe and descriptive statistics
# -
# ## Read daily total sunspot number dataset obtained from SILSO.
# +
#Read daily total sunspot number dataset obtained from SILSO
fname1 = 'data\SN_d_tot_V2.0_2019_2020.txt' #Modified Daily total sunspot number dataset obtained from SILSO
#Read SILSO dataset
sunspot_no = np.loadtxt(fname1, dtype='str', usecols=range(0, 4))
#Build Pandas series objects for each column in the original dataset
year,month,day,sunspotnr = np.loadtxt(fname1, usecols=range(0, 4), unpack=True)
year_list = pd.Series(year,dtype='int32')
month_list = pd.Series(month,dtype='int32')
day_list = pd.Series(day,dtype='int32')
sunspot_nr = pd.Series(sunspotnr,dtype='int32')
#Build Pandas dataframe from each series object and print as a table
sunspot_2019_2020 = pd.DataFrame({'Year':year_list,'Month':month_list,'Day':day_list,'Sunspot No.':sunspot_nr})
sunspot_titles = ['Year','Month','Day','Sunspot No.']
sunspot_2019_2020 = sunspot_2019_2020[sunspot_titles]
print(sunspot_2019_2020)
sunspot_2019_2020.to_csv(r'data\sunspot_2019_2020.csv')
# -
# ## EDA: Print storm report and sunspot number lists for November - December 2019
# +
df_wxdata_nov2019e = pd.read_csv("data/wxdata_nov2019e.csv")
print(df_wxdata_nov2019e.head())
maxmag_nov2019e, meanmag_nov2019e, medmag_nov2019e, strmrpt_nr_nov2019e = dataframe_strmrpt(df_wxdata_nov2019e)
print("November 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_nov2019e)
print("Mean wind magnitude: ", meanmag_nov2019e)
print("Median wind magnitude: ", medmag_nov2019e)
print("Number of storm reports: ", strmrpt_nr_nov2019e)
totsunspot_nr_nov2019e = sunspot_2019_2020.iloc[31:46, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_nov2019e = totsunspot_nr_nov2019e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_nov2019e)
print("Mean Sunspot number: ", meansunspot_nr_nov2019e)
print("")
df_wxdata_nov2019l = pd.read_csv("data/wxdata_nov2019l.csv")
print(df_wxdata_nov2019l.head())
maxmag_nov2019l, meanmag_nov2019l, medmag_nov2019l, strmrpt_nr_nov2019l = dataframe_strmrpt(df_wxdata_nov2019l)
print("November 16-30 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_nov2019l)
print("Mean wind magnitude: ", meanmag_nov2019l)
print("Median wind magnitude: ", medmag_nov2019l)
print("Number of storm reports: ", strmrpt_nr_nov2019l)
totsunspot_nr_nov2019l = sunspot_2019_2020.iloc[46:61, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_nov2019l = totsunspot_nr_nov2019l/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_nov2019l)
print("Mean Sunspot number: ", meansunspot_nr_nov2019l)
print("")
df_wxdata_dec2019e = pd.read_csv("data/wxdata_dec2019e.csv")
print(df_wxdata_dec2019e.head())
maxmag_dec2019e, meanmag_dec2019e, medmag_dec2019e, strmrpt_nr_dec2019e = dataframe_strmrpt(df_wxdata_dec2019e)
print("December 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_dec2019e)
print("Mean wind magnitude: ", meanmag_dec2019e)
print("Median wind magnitude: ", medmag_dec2019e)
print("Number of storm reports: ", strmrpt_nr_dec2019e)
totsunspot_nr_dec2019e = sunspot_2019_2020.iloc[61:76, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_dec2019e = totsunspot_nr_dec2019e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_dec2019e)
print("Mean Sunspot number: ", meansunspot_nr_dec2019e)
print("")
df_wxdata_dec2019l = pd.read_csv("data/wxdata_dec2019l.csv")
print(df_wxdata_dec2019l.head())
maxmag_dec2019l, meanmag_dec2019l, medmag_dec2019l, strmrpt_nr_dec2019l = dataframe_strmrpt(df_wxdata_dec2019l)
print("December 16-31 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_dec2019l)
print("Mean wind magnitude: ", meanmag_dec2019l)
print("Median wind magnitude: ", medmag_dec2019l)
print("Number of storm reports: ", strmrpt_nr_dec2019l)
totsunspot_nr_dec2019l = sunspot_2019_2020.iloc[76:92, 3].sum() #Calculate sum of daily sunspot number over 16-day period
meansunspot_nr_dec2019l = totsunspot_nr_dec2019l/16 #Calculate mean daily sunspot number over 16-day period
print("Sunspot number: ", totsunspot_nr_dec2019l)
print("Mean Sunspot number: ", meansunspot_nr_dec2019l)
print("")
# -
# ## EDA: Print storm report and sunspot number lists for January - February 2020
# +
df_wxdata_jan2020e = pd.read_csv("data/wxdata_jan2020e.csv")
maxmag_jan2020e, meanmag_jan2020e, medmag_jan2020e, strmrpt_nr_jan2020e = dataframe_strmrpt(df_wxdata_jan2020e)
print("January 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_jan2020e)
print("Mean wind magnitude: ", meanmag_jan2020e)
print("Median wind magnitude: ", medmag_jan2020e)
print("Number of storm reports: ", strmrpt_nr_jan2020e)
totsunspot_nr_jan2020e = sunspot_2019_2020.iloc[92:107, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_jan2020e = totsunspot_nr_jan2020e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_jan2020e)
print("Mean Sunspot number: ", meansunspot_nr_jan2020e)
print("")
df_wxdata_jan2020l = pd.read_csv("data/wxdata_jan2020l.csv")
maxmag_jan2020l, meanmag_jan2020l, medmag_jan2020l, strmrpt_nr_jan2020l = dataframe_strmrpt(df_wxdata_jan2020l)
strmrpt_nr_jan2020l += 3
print("January 16-31 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_jan2020l)
print("Mean wind magnitude: ", meanmag_jan2020l)
print("Median wind magnitude: ", medmag_jan2020l)
print("Number of storm reports: ", strmrpt_nr_jan2020l)
totsunspot_nr_jan2020l = sunspot_2019_2020.iloc[107:123, 3].sum() #Calculate sum of daily sunspot number over 16-day period
meansunspot_nr_jan2020l = totsunspot_nr_jan2020l/16 #Calculate mean daily sunspot number over 16-day period
print("Sunspot number: ", totsunspot_nr_jan2020l)
print("Mean Sunspot number: ", meansunspot_nr_jan2020l)
print("")
df_wxdata_feb2020e = pd.read_csv("data/wxdata_feb2020e.csv")
maxmag_feb2020e, meanmag_feb2020e, medmag_feb2020e, strmrpt_nr_feb2020e = dataframe_strmrpt(df_wxdata_feb2020e)
print("February 1-14 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_feb2020e)
print("Mean wind magnitude: ", meanmag_feb2020e)
print("Median wind magnitude: ", medmag_feb2020e)
print("Number of storm reports: ", strmrpt_nr_feb2020e)
totsunspot_nr_feb2020e = sunspot_2019_2020.iloc[123:137, 3].sum() #Calculate sum of daily sunspot number over 14-day period
meansunspot_nr_feb2020e = totsunspot_nr_feb2020e/14 #Calculate mean daily sunspot number over 14-day period
print("Sunspot number: ", totsunspot_nr_feb2020e)
print("Mean Sunspot number: ", meansunspot_nr_feb2020e)
print("")
df_wxdata_feb2020l = pd.read_csv("data/wxdata_feb2020l.csv")
maxmag_feb2020l, meanmag_feb2020l, medmag_feb2020l, strmrpt_nr_feb2020l = dataframe_strmrpt(df_wxdata_feb2020l)
print("February 15-29 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_feb2020l)
print("Mean wind magnitude: ", meanmag_feb2020l)
print("Median wind magnitude: ", medmag_feb2020l)
print("Number of storm reports: ", strmrpt_nr_feb2020l)
totsunspot_nr_feb2020l = sunspot_2019_2020.iloc[137:152, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_feb2020l = totsunspot_nr_feb2020l/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_feb2020l)
print("Mean Sunspot number: ", meansunspot_nr_jan2020l)
print("")
# -
# ## EDA: Print storm report and sunspot number lists for March - April 2020
# +
df_wxdata_mar2020e = pd.read_csv("data/wxdata_mar2020e.csv")
maxmag_mar2020e, meanmag_mar2020e, medmag_mar2020e, strmrpt_nr_mar2020e = dataframe_strmrpt(df_wxdata_mar2020e)
print("March 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_mar2020e)
print("Mean wind magnitude: ", meanmag_mar2020e)
print("Median wind magnitude: ", medmag_mar2020e)
print("Number of storm reports: ", strmrpt_nr_mar2020e)
totsunspot_nr_mar2020e = sunspot_2019_2020.iloc[152:167, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_mar2020e = totsunspot_nr_mar2020e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_mar2020e)
print("Mean Sunspot number: ", meansunspot_nr_mar2020e)
print("")
df_wxdata_mar2020l = pd.read_csv("data/wxdata_mar2020l.csv")
maxmag_mar2020l, meanmag_mar2020l, medmag_mar2020l, strmrpt_nr_mar2020l = dataframe_strmrpt(df_wxdata_mar2020l)
print("March 16-31 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_mar2020l)
print("Mean wind magnitude: ", meanmag_mar2020l)
print("Median wind magnitude: ", medmag_mar2020l)
print("Number of storm reports: ", strmrpt_nr_mar2020l)
totsunspot_nr_mar2020l = sunspot_2019_2020.iloc[167:183, 3].sum() #Calculate sum of daily sunspot number over 16-day period
meansunspot_nr_mar2020l = totsunspot_nr_mar2020l/16 #Calculate mean daily sunspot number over 16-day period
print("Sunspot number: ", totsunspot_nr_mar2020l)
print("Mean Sunspot number: ", meansunspot_nr_mar2020l)
print("")
df_wxdata_apr2020e = pd.read_csv("data/wxdata_apr2020e.csv")
maxmag_apr2020e, meanmag_apr2020e, medmag_apr2020e, strmrpt_nr_apr2020e = dataframe_strmrpt(df_wxdata_apr2020e)
print("April 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_apr2020e)
print("Mean wind magnitude: ", meanmag_apr2020e)
print("Median wind magnitude: ", medmag_apr2020e)
print("Number of storm reports: ", strmrpt_nr_apr2020e)
totsunspot_nr_apr2020e = sunspot_2019_2020.iloc[183:198, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_apr2020e = totsunspot_nr_apr2020e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_apr2020e)
print("Mean Sunspot number: ", meansunspot_nr_apr2020e)
print("")
df_wxdata_apr2020l = pd.read_csv("data/wxdata_apr2020l.csv")
maxmag_apr2020l, meanmag_apr2020l, medmag_apr2020l, strmrpt_nr_apr2020l = dataframe_strmrpt(df_wxdata_apr2020l)
print("April 16-30 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_apr2020l)
print("Mean wind magnitude: ", meanmag_apr2020l)
print("Median wind magnitude: ", medmag_apr2020l)
print("Number of storm reports: ", strmrpt_nr_apr2020l)
totsunspot_nr_apr2020l = sunspot_2019_2020.iloc[198:213, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_apr2020l = totsunspot_nr_apr2020l/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_apr2020l)
print("Mean Sunspot number: ", meansunspot_nr_apr2020l)
# +
df_wxdata_nov2020e = pd.read_csv("data/wxdata_nov2020e.csv")
print(df_wxdata_nov2020e.head())
maxmag_nov2020e, meanmag_nov2020e, medmag_nov2020e, strmrpt_nr_nov2020e = dataframe_strmrpt(df_wxdata_nov2020e)
strmrpt_nr_nov2020e += 234
print("November 1-15 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_nov2020e)
print("Mean wind magnitude: ", meanmag_nov2020e)
print("Median wind magnitude: ", medmag_nov2020e)
print("Number of storm reports: ", strmrpt_nr_nov2020e)
totsunspot_nr_nov2020e = sunspot_2019_2020.iloc[397:412, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_nov2020e = totsunspot_nr_nov2020e/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_nov2020e)
print("Mean Sunspot number: ", meansunspot_nr_nov2020e)
print("")
df_wxdata_nov2020l = pd.read_csv("data/wxdata_nov2020l.csv")
print(df_wxdata_nov2020l.head())
maxmag_nov2020l, meanmag_nov2020l, medmag_nov2020l, strmrpt_nr_nov2020l = dataframe_strmrpt(df_wxdata_nov2020l)
strmrpt_nr_nov2020l += 113
print("November 16-30 storm wind summary:")
print("Maximum wind magnitude: ", maxmag_nov2020l)
print("Mean wind magnitude: ", meanmag_nov2020l)
print("Median wind magnitude: ", medmag_nov2020l)
print("Number of storm reports: ", strmrpt_nr_nov2020l)
totsunspot_nr_nov2020l = sunspot_2019_2020.iloc[412:427, 3].sum() #Calculate sum of daily sunspot number over 15-day period
meansunspot_nr_nov2020l = totsunspot_nr_nov2020l/15 #Calculate mean daily sunspot number over 15-day period
print("Sunspot number: ", totsunspot_nr_nov2020l)
print("Mean Sunspot number: ", meansunspot_nr_nov2020l)
print("")
# -
# ## Plot time series of total sunspot number vs storm report number.
import matplotlib.pyplot as plt
import pandas as pd
#Plot sunspot number vs storm report number
tss = pd.Series([totsunspot_nr_nov2019e, totsunspot_nr_nov2019l, totsunspot_nr_dec2019e, totsunspot_nr_dec2019l,
totsunspot_nr_jan2020e, totsunspot_nr_jan2020l, totsunspot_nr_feb2020e, totsunspot_nr_feb2020l,
totsunspot_nr_mar2020e, totsunspot_nr_mar2020l, totsunspot_nr_apr2020e, totsunspot_nr_apr2020l,
totsunspot_nr_nov2020e, totsunspot_nr_nov2020l])
mss = pd.Series([meansunspot_nr_nov2019e, meansunspot_nr_nov2019l, meansunspot_nr_dec2019e, meansunspot_nr_dec2019l,
meansunspot_nr_jan2020e, meansunspot_nr_jan2020l, meansunspot_nr_feb2020e, meansunspot_nr_feb2020l,
meansunspot_nr_mar2020e, meansunspot_nr_mar2020l, meansunspot_nr_apr2020e, meansunspot_nr_apr2020l,
meansunspot_nr_nov2020e, meansunspot_nr_nov2020l])
sr = pd.Series([strmrpt_nr_nov2019e, strmrpt_nr_nov2019l, strmrpt_nr_dec2019e, strmrpt_nr_dec2019l, strmrpt_nr_jan2020e,
strmrpt_nr_jan2020l, strmrpt_nr_feb2020e, strmrpt_nr_feb2020l, strmrpt_nr_mar2020e, strmrpt_nr_mar2020l,
strmrpt_nr_apr2020e, strmrpt_nr_apr2020l,strmrpt_nr_nov2020e, strmrpt_nr_nov2020l])
meanmag = pd.Series([meanmag_nov2019e, meanmag_nov2019l, meanmag_dec2019e, meanmag_dec2019l, meanmag_jan2020e, meanmag_jan2020l,
meanmag_feb2020e, meanmag_feb2020l, meanmag_mar2020e, meanmag_mar2020l, meanmag_apr2020e, meanmag_apr2020l,
meanmag_nov2020e, meanmag_nov2020l])
maxmag = pd.Series([maxmag_nov2019e, maxmag_nov2019l, maxmag_dec2019e, maxmag_dec2019l, maxmag_jan2020e, maxmag_jan2020l,
maxmag_feb2020e, maxmag_feb2020l, maxmag_mar2020e, maxmag_mar2020l, maxmag_apr2020e, maxmag_apr2020l,
maxmag_nov2020e, maxmag_nov2020l])
month = pd.Series(['11/1-15/19','11/16-30/19','12/1-15/19','12/16-31/19','1/1-15/20','1/16-31/20','2/1-14/20','2/15-29/20',
'3/1-15/20','3/16-31/20','4/1-15/20','4/16-30/20','11/1-15/20','11/16-30/20'])
solar_storm_df = pd.DataFrame({'Period':month,'Total Sunspot No.':tss,'Mean Sunspot No.':mss,'Storm Report No.':sr,
'Mean Wind (mph)':meanmag,'Peak Wind (mph)':maxmag})
solar_storm_df = solar_storm_df[['Period','Total Sunspot No.','Storm Report No.','Mean Wind (mph)','Peak Wind (mph)']]
print(solar_storm_df)
solar_storm_df.to_csv(r'data\solar_storm.csv')
|
Notebooks/thunderstorm_sunspot_EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import datetime as dt
cd = dt.date.today()
print(cd)
import pandas as pd
t1 = pd.to_datetime('2001-10-10 00:00:00')
t2 = pd.to_datetime('2001-10-15 00:00:00')
series = pd.date_range(t1,t2,freq='1min')
iso8601 = [t.strftime('%Y%m%dT%H:%M%SZ') for t in series]
iso8601[:5]
|
Y5 Version/Basic rough.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test Your Skills
#
# *Author: <NAME>*
#
# *Copyright © 2017 CATALIT LLC*
# ## Exercise
#
# The [Pima Indians dataset](https://archive.ics.uci.edu/ml/datasets/Pima+Indians+Diabetes) is a very famous dataset distributed by UCI and originally collected from the National Institute of Diabetes and Digestive and Kidney Diseases. It contains data from clinical exams for women age 21 and above of Pima indian origins. The objective is to predict based on diagnostic measurements whether a patient has diabetes.
#
# It has the following features:
#
# - Pregnancies: Number of times pregnant
# - Glucose: Plasma glucose concentration a 2 hours in an oral glucose tolerance test
# - BloodPressure: Diastolic blood pressure (mm Hg)
# - SkinThickness: Triceps skin fold thickness (mm)
# - Insulin: 2-Hour serum insulin (mu U/ml)
# - BMI: Body mass index (weight in kg/(height in m)^2)
# - DiabetesPedigreeFunction: Diabetes pedigree function
# - Age: Age (years)
#
# The last colum is the outcome, and it is a binary variable.
#
# In this exercise we will explore it through the following steps:
#
# 1. Load the `../../../data/diabetes.csv` dataset, use pandas to explore the range of each feature
# - Inspect the features using `sns.pairplot`, can you guess which features are more predictive?
# - Look at the scale of each feature and rescale all of them using the Standard Scaler from Scikit Learn.
# - Prepare your final `X` and `y` variables to be used by a ML model. Make sure you define your target variable well.
# - Split your data in a train/test with a test size of 20% and a `random_state = 22`
# - define a sequential model with at least one inner layer. You will have to make choices for the following things:
# - what is the size of the input?
# - how many nodes will you use in each layer?
# - what is the size of the output?
# - what activation functions will you use in the inner layers?
# - what activation function will you use at output?
# - what loss function will you use?
# - what optimizer will you use?
# - fit your model on the training set, using a validation_split of 0.1
# - test your trained model on the test data from the train/test split
# - check the accuracy score, the confusion matrix and the classification report
# - compare your Neural Network model with another model from scikit-learn
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
df = pd.read_csv('../../../data/diabetes.csv')
df.head()
import seaborn as sns
sns.pairplot(df, hue='Outcome')
from sklearn.preprocessing import StandardScaler
from keras.utils import to_categorical
sc = StandardScaler()
X = sc.fit_transform(df.drop('Outcome', axis=1))
y = df['Outcome'].values
y_cat = to_categorical(y)
X.shape
y_cat.shape
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y_cat,
random_state=22,
test_size=0.2)
from keras.models import Sequential
from keras.layers import Dense
from keras.optimizers import Adam
model = Sequential()
model.add(Dense(32, input_shape=(8,), activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dense(2, activation='softmax'))
model.compile(Adam(lr=0.05),
loss='categorical_crossentropy',
metrics=['accuracy'])
model.summary()
model.fit(X_train, y_train, epochs=20, verbose=2, validation_split=0.1)
y_pred = model.predict(X_test)
y_test_class = np.argmax(y_test, axis=1)
y_pred_class = np.argmax(y_pred, axis=1)
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
pd.Series(y_test_class).value_counts() / len(y_test_class)
accuracy_score(y_test_class, y_pred_class)
print(classification_report(y_test_class, y_pred_class))
confusion_matrix(y_test_class, y_pred_class)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
for mod in [RandomForestClassifier(), SVC(), GaussianNB()]:
mod.fit(X_train, y_train[:, 1])
y_pred = mod.predict(X_test)
print("="*80)
print(mod)
print("-"*80)
print("Accuracy score: {:0.3}".format(accuracy_score(y_test_class,
y_pred)))
print("Confusion Matrix:")
print(confusion_matrix(y_test_class, y_pred))
print()
# -
|
solutions_do_not_open/are_you_really_sure/are_you_really_really_sure/05_Test_your_skills_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.12 64-bit (''soccer'': conda)'
# name: python_defaultSpec_1600863097834
# ---
import xml.etree.ElementTree as ET
import xmltodict
import json
import pandas as pd
# +
paths = [
'data/RBKvFKH20200920/2020-09-20 16_00_00 Rosenborg - Haugesund.xml',
#'data/RBKvFKH20200920/haugesund_player-events_20-09-2020_rosenborg-haugesund.xml',
#'data/RBKvFKH20200920/haugesund_team-events_20-09-2020_rosenborg-haugesund.xml',
#'data/RBKvFKH20200920/rosenborg_player-events_20-09-2020_rosenborg-haugesund.xml',
#'data/RBKvFKH20200920/rosenborg_team-events_20-09-2020_rosenborg-haugesund.xml',
#'data/RBKvFKH20200920/Rosenborg-Haugesund_20-09-2020_effective_time.xml'
]
dicts = []
for path in paths:
tree = ET.parse(path)
xml_data = tree.getroot()
xmlstr = ET.tostring(xml_data, encoding='utf-8', method='xml')
dicts.append(dict(xmltodict.parse(xmlstr)))
# -
paths
def sec_to_min(start, end):
min_start = str(int(start)//60) + ',' + str(int(start)%60) + '\''
min_end = str(int(end)//60) + ',' + str(int(end)%60) + '\''
return min_start + '-' + min_end
# + tags=[]
rosenborg = []
for i in dicts[0]['file']['ROWS']['row']:
if int(i['sort_order']) < 43 and 27 < int(i['sort_order']):
rosenborg.append(i['code'][(i['code'].find(' ')+1):])
haugesund = []
for i in dicts[0]['file']['ROWS']['row']:
if 42 < int(i['sort_order']):
haugesund.append(i['code'][(i['code'].find(' ')+1):])
print(rosenborg)
print(haugesund)
# + tags=[]
from collections import OrderedDict as odict
rbk = odict()
hfk = odict()
for name in rosenborg:
rbk[name] = odict(({'total': 0, 'won':0, 'lost':0}))
for name in haugesund:
hfk[name] = odict(({'total': 0, 'won':0, 'lost':0}))
duels = {} #dict with key = player number and value 3-list with [nr_duels, duels_won, duels_lost]
nr_duels = 0
nr_won = 0
nr_lost = 0
for dictionary in dicts:
for i in dictionary['file']['ALL_INSTANCES']['instance']:
name = i['code'][(i['code'].find(' ')+1):]
if name in rosenborg:
if i['label'][-1]['group'] == 'duels' :
for attr in i['label']:
if attr['text'] == 'Plus':
rbk[name]['total'] += 1
rbk[name]['won'] += 1
if attr['text'] == 'Minus':
rbk[name]['total'] += 1
rbk[name]['lost'] += 1
if attr['text'] == 'Neutral':
rbk[name]['total'] += 1
elif name in haugesund:
if i['label'][-1]['group'] == 'duels' :
for attr in i['label']:
if attr['text'] == 'Plus':
hfk[name]['total'] += 1
hfk[name]['won'] += 1
if attr['text'] == 'Minus':
hfk[name]['total'] += 1
hfk[name]['lost'] += 1
if attr['text'] == 'Neutral':
hfk[name]['total'] += 1
# + tags=[]
for player in list(rbk.keys()):
total = int(rbk[player]['total'])
won = int(rbk[player]['won'])
lost = int(rbk[player]['lost'])
if won+lost > 0 and total > 0:
rbk[player]['percentage'] = won/(won+lost)
rbk[player]['percentage_of_total'] = total * rbk[player]['percentage']
print(player, f'was in {total} duels and won {won}, lost {lost} or won {won/(won+lost):.2f}%')
else:
rbk[player]['percentage'] = 0
rbk[player]['percentage_of_total'] = 0
for player in list(hfk.keys()):
total = int(hfk[player]['total'])
won = int(hfk[player]['won'])
lost = int(hfk[player]['lost'])
if won+lost > 0 and total > 0:
hfk[player]['percentage'] = won/(won+lost)
hfk[player]['percentage_of_total'] = total * hfk[player]['percentage']
print(player, f'was in {total} duels and won {won}, lost {lost} or won {won/(won+lost):.2f}%')
else:
hfk[player]['percentage'] = 0
hfk[player]['percentage_of_total'] = 0
# + tags=[]
nested_rbk = []
nested_hfk = []
for player in list(rbk.keys()):
nested_rbk.append(list(rbk[player].values()))
for player in list(hfk.keys()):
nested_hfk.append(list(hfk[player].values()))
df_rbk = pd.DataFrame(nested_rbk, columns=rbk['<NAME>'].keys(), index = rosenborg)
df_hfk = pd.DataFrame(nested_hfk, columns=hfk['<NAME>'].keys(), index = haugesund)
# + tags=[]
import plotly.graph_objects as go
players = rosenborg
data = []
won = list(df_rbk['percentage_of_total'])
lost= list(df_rbk['total'])
for i in range(len(won)):
lost[i] -= won[i]
fig = go.Figure(data=[
go.Bar(name='won', x=players, y=won, marker=dict(color="white")),
go.Bar(name='lost', x=players, y=lost, marker=dict(color="blue"))
])
# Change the bar mode
fig.update_layout(barmode='stack')
fig.show()
# +
players = haugesund
won = list(df_hfk['percentage_of_total'])
lost= list(df_hfk['total'])
for i in range(len(won)):
lost[i] -= won[i]
fig = go.Figure(data=[
go.Bar(name='won', x=players, y=won, marker=dict(color="blue")),
go.Bar(name='lost', x=players, y=lost, marker=dict(color="white")),
])
# Change the bar mode
fig.update_layout(barmode='stack')
fig.show()
# -
|
soccer_visualization/xml_loader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from scipy import sparse
from sklearn.metrics.pairwise import cosine_similarity
ratings=pd.read_csv("toy_dataset.csv",index_col=0)
ratings.fillna(0, inplace=True)
ratings
# +
def standardize(row):
new_row = (row - row.mean())/(row.max()-row.min())
return new_row
df_std = ratings.apply(standardize).T
print(df_std)
sparse_df = sparse.csr_matrix(df_std.values)
corrMatrix = pd.DataFrame(cosine_similarity(sparse_df),index=ratings.columns,columns=ratings.columns)
corrMatrix
# -
corrMatrix = ratings.corr(method='pearson')
corrMatrix.head(6)
def get_similar(movie_name,rating):
similar_score = corrMatrix[movie_name]*(rating-2.5)
similar_score = similar_score.sort_values(ascending=False)
#print(type(similar_ratings))
return similar_score
# +
action_lover = [("action1",5),("romantic2",1),("romantic3",1)]
similar_scores = pd.DataFrame()
for movie,rating in action_lover:
similar_scores = similar_scores.append(get_similar(movie,rating),ignore_index = True)
similar_scores.head(10)
# -
similar_scores.sum().sort_values(ascending=False)
|
UAS_SISTEM_PENUNJANG_KEPUTUSAN/Collaborative Filtering/Collaborative Filtering Dummy Dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Author: <NAME>
# # Trees
# * A tree is a data structure similar to a linked List but instead of each node pointing simply to the next node in a linear fashion, each node points to a number of nodes. Tree is an example of non-linear data structures. A tree structure is a way of representing the **hierarchical** nature of a structure in a graphical form.
# * The **root** of the tree is the node with no parents. There can be at most one root node in a tree.
# * The _edge_ refers to the link from parent to child.
# * A node with NO children is called _leaf_ node.
# * Children of same parent are called _siblings_.
#
# ### Applications of trees:
# * Manipulate hierarchical data.
# * Make information easy to search (see tree traversal).
# * Manipulate sorted lists of data.
# * As a workflow for compositing digital images for visual effects.
# * Router algorithms
# * Form of a multi-stage decision-making (see business chess).
# ## Binary Tree
# * A tree is called _binary tree_ if each node of the tree has zero, one or two children.
# * Empty tree is also a valid binary tree.
# ### Types of binary trees:
# 1. **Full binary tree**:
# A Binary Tree is full if every node has 0 or 2 children. Following are examples of full binary tree.
# 2. **Complete binary tree**:
# A Binary Tree is complete Binary Tree if all levels are completely filled except possibly the last level and the last level has all keys as left as possible.
# 3. **Perfect Binary Tree**:
# A Binary tree is Perfect Binary Tree in which all internal nodes have two children and all leaves are at same level.
# 4. **Balanced Binary Tree**:
# A binary tree is balanced if height of the tree is O(Log n) where n is number of nodes. For Example, AVL tree maintain O(Log n) height by making sure that the difference between heights of left and right subtrees is 1. Red-Black trees maintain O(Log n) height by making sure that the number of Black nodes on every root to leaf paths are same and there are no adjacent red nodes. Balanced Binary Search trees are performance wise good as they provide O(log n) time for search, insert and delete.
# 5. **A degenerate (or pathological) tree**:
# A Tree where every internal node has one child. Such trees are performance-wise same as linked list.
#
# For visualizations:
# http://www.geeksforgeeks.org/binary-tree-set-3-types-of-binary-tree/
#
# For properties:
# http://www.geeksforgeeks.org/binary-tree-set-2-properties/
# ### Binary tree implementation:
# +
class Node(object):
def __init__(self, data = None):
self.left = None
self.right = None
self.data = data
# for setting left node
def setLeft(self, node):
self.left = node
# for setting right node
def setRight(self, node):
self.right = node
# for getting the left node
def getLeft(self):
return self.left
# for getting right node
def getRight(self):
return self.right
# for setting data of a node
def setData(self, data):
self.data = data
# for getting data of a node
def getData(self):
return self.data
root = Node(1)
root.setLeft(Node(2))
root.setRight(Node(3))
root.left.setLeft(Node(4))
''' This will a create like this
1
/ \
2 3
/
4
'''
# -
# ### Tree traversal:
# Unlike linear data structures (Array, Linked List, Queues, Stacks, etc) which have only one logical way to traverse them, trees can be traversed in different ways. Following are the generally used ways for traversing trees.
#
# * Inorder (left, data, right)
# * Preorder (data, left, right)
# * Postorder (left, right, data)
# ### Implementing tree traversals:
# +
# in this we traverse first to the leftmost node, then print its data and then traverse for rightmost node
def inorder(Tree):
if Tree:
inorder(Tree.getLeft())
print(Tree.getData(), end = ' ')
inorder(Tree.getRight())
return
# in this we first print the root node and then traverse towards leftmost node and then to the rightmost node
def preorder(Tree):
if Tree:
print(Tree.getData(), end = ' ')
preorder(Tree.getLeft())
preorder(Tree.getRight())
return
# in this we first traverse to the leftmost node and then to the rightmost node and then print the data
def postorder(Tree):
if Tree:
postorder(Tree.getLeft())
postorder(Tree.getRight())
print(Tree.getData(), end = ' ')
return
print('Inorder Traversal:')
inorder(root)
print('\nPreorder Traversal:')
preorder(root)
print('\nPostorder Traversal:')
postorder(root)
|
Trees/Tree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [my_projects_env]
# language: python
# name: Python [my_projects_env]
# ---
s='FLuFFy'
print s.lower()
s=['FluFFY','BlaBla']
for i in s:
print i.lower()
|
_posts/Untitled3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # 8 Puzzle solver
# * <NAME> - 97149081
# * In this repository we're going to solve this puzzle using $ A^* $ and $ IDA $
# + [markdown] pycharm={"name": "#%% md\n"}
# #### imports:
# + pycharm={"name": "#%%\n"}
import copy
import pandas as pd
import numpy as np
import collections
import heapq
# + [markdown] pycharm={"name": "#%% md\n"}
# #### Test case 1:
# + pycharm={"name": "#%%\n"}
input_puzzle_1 = [
[1, 2, 3],
[4, 0, 5],
[7, 8, 6]
]
print('Input: ')
print(pd.DataFrame(input_puzzle_1))
print()
desired_output_1 = [
[1, 2, 3],
[4, 5, 0],
[7, 8, 6]
]
print('Desired Output:')
print(pd.DataFrame(desired_output_1))
# -
# #### Test case 2: (Hardest)
# + pycharm={"name": "#%%\n"}
input_puzzle_2 = [
[8, 6, 7],
[2, 5, 4],
[3, 0, 1]
]
print('Input 2: ')
print(pd.DataFrame(input_puzzle_2))
print()
desired_output_2 = [
[6, 4, 7],
[8, 5, 0],
[3, 2, 1]
]
print('Desired Output 2:')
print(pd.DataFrame(desired_output_2))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### code's configs:
# + pycharm={"name": "#%%\n"}
heuristic_method = input("Enter the desired Heuristic method: h1 or h2")
f_function_omega = eval(input("Enter the desired f function omega: 2 is Greedy, "
"0 is Uninformed best-first search, "
"0 < omega <= 1 is A*"))
test_case = eval(input("which test case? 1:easy, 2:hard"))
if test_case == 1:
input_puzzle = input_puzzle_1
desired_output = desired_output_1
elif test_case == 2:
input_puzzle = input_puzzle_2
desired_output = desired_output_2
# -
# ### Matrix to dictionary converter
# + pycharm={"name": "#%%\n"}
class Mat2dict:
def __init__(self, matrix):
self.matrix = matrix
self.dic = {}
def convert(self):
for r in range(len(self.matrix)):
for c in range(len(self.matrix[0])):
key = self.matrix[r][c]
self.dic[key] = [r, c]
return self.dic
# -
# ### the heuristic calculator class:
#
# * H1 heuristic (misplaced tiles)
#
# $ \Sigma_{i=1}^{9} \; if \; currentPuzzleBoard[node_i] \; != \; goalPuzzleBoard[node_i]$
#
# $ then \; h(state_y) = h(state_y) + 1$
#
# * H2 heuristic (manhattan distance)
#
# $ goalPuzzleBoard.find(currentPuzzleBoard[node_i]) $
#
# $ retrieve \; Row \; \& \; Col \; of \; goal $
#
# $ manhattanDistance = |(goal.row - current.row)| + |(goal.col - current.col)| $
#
# $ TotalHeuristic[state_i] = \Sigma_{i=1}^{9} manhattanDistance_i $
# + pycharm={"name": "#%%\n"}
class Heuristic:
def __init__(self, node, current_puzzle, desired_answer, method):
self.method = method
self.node = node
self.current_puzzle = current_puzzle
self.desired_answer = desired_answer
#self.current_puzzle_dict = Mat2dict(self.current_puzzle)
self.desired_answer_dict = Mat2dict(self.desired_answer).convert()
def do(self):
if self.method == 'h1':
return self.h1_misplaced_tiles()
elif self.method == 'h2':
return self.h2_manhattan_distance()
def h1_misplaced_tiles(self):
misplaced_counter = 0
for row in range(len(self.current_puzzle)):
for col in range(len(self.current_puzzle[0])):
if self.current_puzzle[row][col] != self.desired_answer[row][col]:
misplaced_counter += 1
return misplaced_counter
def h2_manhattan_distance(self):
total_distance_counter = 0
for row in range(len(self.current_puzzle)):
for col in range(len(self.current_puzzle[0])):
key = self.current_puzzle[row][col]
correct_row, correct_col = self.desired_answer_dict[key]
total_distance_counter += abs(row - correct_row) + abs(col - correct_col)
return total_distance_counter
# + [markdown] pycharm={"name": "#%% md\n"}
# ### The node class:
#
# * F function is calculated in a such way that you can control how the Heuristic and G-cost
# can perform:
#
# $ FCost(n) = (2-\omega) * GCost(n) + \omega * h(n)$
#
# $ if \; \omega = 2: $
#
# $ then: algorithm \; is \; Greedy \; due \; to \; GCost \; being \; 0:$
#
# $ FCost(n) = 0 + 2 * h(n) $
#
# $ if \; \omega = 0: $
#
# $ then: algorithm \; is \; uninformed \; search \; due \; to \; h(n) \; being \; 0:$
#
# $ FCost(n) = 2 * GCost(n) + 0 $
#
# $ if \; 0 \lt \omega \le 1 : $
#
# $ then: algorithm \; is \; informed \; search(A^*):$
#
# $ FCost(n) = (2-\omega) * GCost(n) + \omega * h(n) $
# + pycharm={"name": "#%%\n"}
class Node:
def __init__(self, current_puzzle, parent=None):
self.current_puzzle = current_puzzle
self.parent = parent
if self.parent:
self.g_cost = self.parent.f_function
self.depth = self.parent.depth + 1
else:
self.g_cost = 0
self.depth = 0
self.h_cost = Heuristic(self, current_puzzle, desired_output, heuristic_method).do()
self.f_function = (2 - f_function_omega) * self.g_cost + f_function_omega * self.h_cost
def __eq__(self, other):
return self.f_function == other.f_function
def __lt__(self, other):
return self.f_function < other.f_function
def get_id(self):
return str(self)
def get_path(self):
node, path = self, []
while node:
path.append(node)
node = node.parent
return list(reversed(path))
def get_position(self, element):
for row in range(len(self.current_puzzle)):
for col in range(len(self.current_puzzle[0])):
if self.current_puzzle[row][col] == element:
return [row, col]
return [0, 0]
# + [markdown] pycharm={"name": "#%% md\n"}
# ### The puzzle solver class:
# + pycharm={"name": "#%%\n"}
class PuzzleSolver:
def __init__(self, start_node):
self.final_state = None
self.start_node = start_node
self.depth = 0
self.visited_nodes = set()
self.expanded_nodes = 0
def solve(self):
queue = [self.start_node]
self.visited_nodes.add(self.start_node.get_id())
while queue:
self.expanded_nodes += 1
node = heapq.heappop(queue)
if node.current_puzzle == desired_output:
self.final_state = node
Result(self.final_state, self.expanded_nodes)
return True
if node.depth + 1 > self.depth:
self.depth = node.depth + 1
for neighbor in NeighborsCalculator(node).get_list_of_neighbors():
if not neighbor.get_id in self.visited_nodes:
self.visited_nodes.add(neighbor.get_id())
heapq.heappush(queue, neighbor)
return False
# + [markdown] pycharm={"name": "#%% md\n"}
# ### result class
# + pycharm={"name": "#%%\n"}
class Result:
def __init__(self, final_state, expanded_nodes):
self.expanded_nodes = expanded_nodes
self.final_state = final_state
self.solved_puzzle = self.final_state.current_puzzle
self.path = self.final_state.get_path()
self.show_puzzles()
self.show_path()
def show_puzzles(self):
print("Inital Puzzle: ")
print(pd.DataFrame(input_puzzle))
print("Result Puzzle: ")
print(pd.DataFrame(self.solved_puzzle))
print("Expected Puzzle: ")
print(pd.DataFrame(desired_output))
print()
print("Number of expanded nodes: {}".format(self.expanded_nodes))
print()
def show_path(self):
counter = 0
while self.path:
counter += 1
step = self.path.pop(0)
print("step {}: ".format(counter))
print(pd.DataFrame(step.current_puzzle))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Neighbors calculator
# + pycharm={"name": "#%%\n"}
class NeighborsCalculator:
def __init__(self, current_state):
self.current_state = current_state
self.puzzle = self.current_state.current_puzzle
self.neighbors = []
def get_list_of_neighbors(self):
row, col = map(int, self.current_state.get_position(0))
#if row or col is None:
# return []
# move right
if col < len(self.puzzle[0]) - 1:
moved_right = copy.deepcopy(self.puzzle)
moved_right[row][col], moved_right[row][col + 1] = moved_right[row][col + 1], moved_right[row][col]
self.neighbors.append(Node(moved_right, self.current_state))
# move left
if col > 0:
moved_left = copy.deepcopy(self.puzzle)
moved_left[row][col], moved_left[row][col - 1] = moved_left[row][col - 1], moved_left[row][col]
self.neighbors.append(Node(moved_left, self.current_state))
# move up
if row > 0:
moved_up = copy.deepcopy(self.puzzle)
moved_up[row][col], moved_up[row - 1][col] = moved_up[row - 1][col], moved_up[row][col]
self.neighbors.append(Node(moved_up, self.current_state))
# move down
if row < len(self.puzzle) - 1:
moved_down = copy.deepcopy(self.puzzle)
moved_down[row][col], moved_down[row + 1][col] = moved_down[row + 1][col], moved_down[row][col]
self.neighbors.append(Node(moved_down, self.current_state))
return self.neighbors
# + pycharm={"name": "#%%\n"}
initial_state = Node(input_puzzle)
PuzzleSolver(initial_state).solve()
# + [markdown] pycharm={"name": "#%% md\n"}
# ### The puzzle solver IDA class:
# + pycharm={"name": "#%%\n"}
class PuzzleSolverIDA:
def __init__(self, start_node, iterate):
self.iterate = iterate
self.final_state = None
self.start_node = start_node
self.depth = 0
self.visited_nodes = set()
self.expanded_nodes = 0
self.f_cutoff = 0
def solve(self):
while True:
self.f_cutoff += self.iterate
queue = [self.start_node]
self.visited_nodes.add(self.start_node.get_id())
while queue:
self.expanded_nodes += 1
node = heapq.heappop(queue)
if node.current_puzzle == desired_output:
self.final_state = node
Result(self.final_state, self.expanded_nodes)
return True
if node.depth + 1 > self.depth:
self.depth = node.depth + 1
for neighbor in NeighborsCalculator(node).get_list_of_neighbors():
if not neighbor.get_id in self.visited_nodes:
if neighbor.f_function <= self.f_cutoff:
self.visited_nodes.add(neighbor.get_id())
heapq.heappush(queue, neighbor)
# + pycharm={"name": "#%%\n"}
initial_state = Node(input_puzzle)
PuzzleSolverIDA(initial_state, 4).solve()
|
8-Puzzle-Solver.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# >>> Work in Progress (Following are the lecture notes of Prof <NAME> - CS229 - Stanford. This is my interpretation of his excellent teaching and I take full responsibility of any misinterpretation/misinformation provided herein.)
# # Lecture Notes
#
# #### Outline:
# - Naive Bayes
# - Laplace smoothing
# - Element models
# - Comments on applying ML
# - SVM
# ## Naive Bayes
# - is a generative type of algorithm
# - To build generative model, P(x|y) and P(y) needs to be modeled. Gaussian discriminative model uses Gaussian and Bernoulli respectively to model this. Naive Bayes uses a different model.
# > - $ x_{j} = \mathbb 1$ {indicator - word j appears in email or not}
# > - $ P(x|y) = \prod\limits_{j=1}^{n}P(x_{j}|y)$ - NB uses the product of conditional probabilities of individual features given in the class label y
# - Parameters of NB model are:
# > - $P(y=1) = \phi_{y} $ - the class prior for y=1
# > - $P(x_{j}=1|y=0) = \phi_{j|y=0}$ - chances of the word appearing in non-spam email.
# > - $P(x_{j}=1|y=1) = \phi_{j|y=1}$ - chances of the word appearing in spam email.
#
# ### Maximum likelihood estimate
# > $\phi_{y} = \frac{\sum\limits_{i=1}^{m}\mathbb 1 \{y^{(i)}=1\}}{m}$
# > $\phi_{j|y=0} = \frac{\sum\limits_{i=1}^{m}\mathbb 1 \{x_{j}^{(i)}=1, y^{(i)}=0\}}{\sum\limits_{i=1}^{m}\mathbb 1 \{y^{(i)}=0\}}$
# ### Laplace smoothing
# - ML conference papers - NIPS - Neural Information Processing Systems
#
# - __NB breaks__ - because the probability of some event that you have not seen is trained as 0, which is statistically wrong
# - the classifier gives wrong result, when it gets such event for the first time
# - instead using __Laplace smoothing__ helps this problem
# - add 1 each to pass and fail scenario
#
# #### Maximum likelihood estimate
# > $\phi_{x=i} = \frac{\sum\limits_{j=1}^{m}\mathbb 1 \{x^{(i)}=j\} + 1} {m + k}$
# - k is the size of dictionary
# > $\phi_{i|y=0} = \frac{\sum\limits_{i=1}^{m}\mathbb 1 \{x_{j}^{(i)}=1, y^{(i)}=0\} + 1}{\sum\limits_{i=1}^{m}\mathbb 1 \{y^{(i)}=0\} + 2}$
# ## Event models for text classification
# - The functionality can be generalized from binary to multinomial features instead
# - Multivariate Bernoulli event
# - Multinomial event
#
# - We can discretize the size of house and transform a Bernoulli event to Multinomial event
# - a rule of thumb is to discretize variables into 10 buckets
#
#
# <img src="images/06_multinomialEventSqFt.png" width=400 height=400>
# $\tiny{\text{YouTube-Stanford-CS229-Andrew Ng}}$
#
# ## NB variation
# - for text classification
# - if the email contains text "Drugs! Buy drugs now",
# - the feature vector will make binary vector of all words appearing in the email
# - it will lose the information that the word "drug" appeared twice
# - each featture only stores information $x_{j} \in \{0,1\}$
# - instead of making a feature vector of 10000 words of dictionary, we can make a feature vector of 4 words(as above) holding word index and $x_{j} \in \{1, 2, ... ,10000\}$
# - algorithm containing feature vector of 10000 is called __"Multivariate Bernoulli algorithm"__
# - algorithm containing feature vector of 4 is called __"Multinomial algorithm"__
# - <NAME> used these 2 names
#
#
#
# ### NB advantage
# - quick to implement
# - computationally efficient
# - no need to implement gradient descent
# - easy to do a quick and dirty type of work
# - SVM or logistic regression does better work in classification problems
# - NB or GDA does not result in very good classification, but is very quick to implement, it is quick to train, it is non-iterative
#
# ## Support Vector Machines (SVM)
# - find classification in the form of non linear decision boundaries
# - SVM does not have many parameters to fiddle with
# - they have very robust packages
# - and does not have parameters like learning rate, etc to fiddle with
#
# <br>
#
# - Let the non-linear feature variables be mapped in vector form as below. This non-linear function can be viewed as a linear function over the variables $\phi(x)$
# - derive an algorithm that can take input features of the $x_{1}, x_{2}$ and map them to much higher dimensional set of feature and then apply linear classifier, similar to logistic regression but different in details. This allows us to learn very non-linear decision boundaries.
#
# <img src="images/06_svmBoundary.png" width=200 height=200>
# $\tiny{\text{YouTube-Stanford-CS229-Andrew Ng}}$
#
#
# ### Outline for SVM
# - Optimal margin classifier (separable case)
#
# - Kernel
# - Kernels will allow us to fit in a large set of features
# - How to take feature vector say $\mathbb R^{2}$ and map it to $\mathbb R^{5}$ or $\mathbb R^{10000}$ or $\mathbb R^{\inf}$, and train the algorithm to this higher feature. Map 2-dimensional feature space to infinite dimensional set of features. What it helps us is relieve us from lot of burden of manually picking up features. ($x^{2}, \space x^{3}, \space \sqrt{x}, \space x_{1}x_{2}^{2/3}, ...$)
#
# - Inseparable case
# ### Optimal margin classifier (separable case)
# - Separable case means data is separable
#
# #### Functional margin of classifier
#
# - How confidently and accurately do you classify an example
# - Using binary classification and logistic regression
# - In logistic classifier:
# > $h_{\theta}(x) = g(\theta^{T}x)$
# - If you turn this into binary classification, if you have this algorithm predict not a probability but predict 0 or 1,
# > - Predict "1", if $\theta^{T}x > 0 \implies h_{\theta}(x) = g(\theta^{T}x) \ge 0.5$)
# > - Predict "0", otherwise
# - If $y^{(i)} = 1$, hope that $\theta^{T}x^{(i)} \gg 0$
# - If $y^{(i)} = 0$, hope that $\theta^{T}x^{(i)} \ll 0$
# - this implies that the prediction is very correct and accurate
#
# #### Geometric margin
# - assuming data is linearly separable,
# - and there are two lines that separates the true and false variables
# - the line that has a bigger separation or geometric margin meaning a physical separation from the training examples is a better choice
#
# - We need to prove the optimal classifier is the algorithm that tries to maximize the geometric margin
# - what SVM and low dimensional classifier do is pose an optimization problem to try and find the line that classify these examples to find bigger separation margin
# #### Notation
# - Labels: $y \in \{-1, +1\} $ to denote class labels
# - SVM will generate hypothesis output value as {-1, +1} instead of probability as in logistic regression
# - g(z) = $\begin{equation}
# \begin{cases}
# +1 & \text{if z $\ge$ 0}\\
# -1 & \text{otherwise}
# \end{cases}
# \end{equation}$
# - instead of smooth transition, here we have a hard transition
#
#
# Previously in logistic regression:
# > $h_{\theta}(x) = g(\theta^{T}x)$
# - where x is $\mathbb R^{n+1} $ and $x_{0} = 1$
#
# In SVM:
# > $h_{W,b}(x) = g(W^{T}x + b)$
# - where x is $\mathbb R^{n} $ and b is $\mathbb R $ and drop $x_{0}=1$
# - The term
#
# Other way to think about is:
# > $
# \begin{bmatrix}
# \theta_{0}\\
# \theta_{1}\\
# \theta_{2}\\
# \theta_{3}
# \end{bmatrix}
# $
# - where $\theta_{0}$ corresponds to b and $\theta_{1}, \theta_{2}, \theta_{3}$ corresponds to W
# #### Functional margin of hyperplane (cont)
# - Functional margin __wrt single training example__
# - Functional margin of hyperplane defined by $(w, b)$ wrt a single training example $(x^{(i)}, y^{(i)})$ is
# > $\hat{\gamma}^{(i)} = y^{(i)} (w^{T}x^{(i)} + b)$
# - If $y^{(i)} = +1$, hope that $(w^{T}x^{(i)} + b) \gg 0$
# - If $y^{(i)} = -1$, hope that $(w^{T}x^{(i)} + b) \ll 0$
# - Combining these two statements:
# - we hope that $\hat{\gamma}^{(i)} \gg 0$
# - If $\hat{\gamma}^{(i)} \gt 0$,
# - implies $h(x^{(i)}) = y^{(i)}$
# - In logistic regression
# - $\gt 0$ means the prediction is atleast a little bit above 0.5 or little bit below 0.5
# - $\gg 0$ means the prediction is either very close to 1 or very close to 0
# - Functional margin __wrt entire training set__ (how well are you doing on the worst example in your training set)
# > $\hat{\gamma} = \min\limits_{i} \hat{\gamma}^{(i)}$
# - where i = 1,..,m - all the training examples
# - here the assumption is training set is linearly separable
# - we can assume this kind of worst-case notion because we are assuming that the boundary is linearly separable
# - we can normalize (w, b), which does not change the classification boundary, it simply rescales the parameters
# > $(w, b) \rightarrow (\frac{w}{\Vert w \Vert}, \frac{b}{\Vert b \Vert})$
# - classification remains the same, with any rescale number
#
# #### Geometric margin of hyperplane (cont)
# - Geometric margin __wrt single training example__
# - Upper half of hyperplane has positive cases and the lower half negative cases
# - Let $(x^{(i)}, y^{(i)})$ be a training example which the classifier is classifying correctly, by predicting it as $h_{w,b}(x) = +1$. It predicts the lower half as $h_{w,b}(x) = -1$.
# - Let the distance from the plane to the training example be the geometric margin
# - Geometric margin of hyperplane (w,b) wrt $(x^{(i)}, y^{(i)})$ be
# > $\hat{\gamma} = \frac{(y^{(i)})(w^{T}x^{(i)} + b)}{\Vert w \Vert}$
# - This is the Euclidean distance between training example and decision boundary
# - For positive examples, $y^{(i)}$ is +1 and the equation reduces to
# > $\hat{\gamma} = \frac{(w^{T}x^{(i)} + b)}{\Vert w \Vert}$
#
#
# <img src="images/06_geometricMargin1.png" width=400 height=400>
# $\tiny{\text{YouTube-Stanford-CS229-Andrew Ng}}$
#
#
# #### Relationship between functional and geometric margin
# > $\gamma^{(i)} = \frac{\hat{\gamma}^{(i)}}{\Vert w \Vert}$
# > i.e., $\text{Geometric Margin} = \frac{\text{Functional Margin}}{\text{Norm of w}}$
#
# #### Geometric margin of hyperplane (cont)
# - Geometric margin __wrt entire training example__
# > $\gamma = \min\limits_{i} \gamma^{(i)}$
#
# #### Optimal margin classifier (cont)
# - what the optimal margin classifier does is to choose w,b to maximize $\gamma$
# > $\max_{\gamma, w, b} \text{s.t.} \frac{(y^{(i)})(w^{T}x^{(i)} + b)}{\Vert w \Vert} \ge \gamma$ for i=1,...,m
# - subject to for every single training example must have the geometric margin greater than or equal to gamma
# - this causes to maximize the worst-case geometric margin
# - this is not a convex optimization problem and cannot be solved using gradient descent or local optima
# - but this can be re-written/reformulated into a equivalent problem which is minimizing norm of w subject to the geometric margin
# > $\min_{w,b} \Vert w \Vert ^{2} \\
# \text{s.t. } y^{(i)}(w^{T}x^{(i)} + b) \ge 1$
# - This is a convex optimization problem and can be solved using optimization packages
# - All this study is for linear separable case only
# - this is a baby SVM
# - Once we learn kernels and apply kernels with optimal margin classifier, we get solution for SVM
|
cs229_ml/lec06-NaiveBayes-SVM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Synopsis:
# -iteratively load data, normalize it, and save it here .../MICCAI_BraTS_2019_Data_Training/MICCAI_BraTS_2019_Data_Training/normalized_hgg
# +
import utils.hgg_utils as hu
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
import nibabel as nib
import pathlib
# -
# # Func: Normalize
def normalize_each_modality_by_its_max(a):
for modality_idx in range( a.shape[-1] ):
a[:, :, :, modality_idx] /= a[:, :, :, modality_idx].max()
# # Func: Save the normalized files
def save_data( multi_mod, affines_list, mod_paths, destination ):
patient_paths = [ x.parent.stem for x in mod_paths ]
mods = [x.name for x in mod_paths]
for modality in range(multi_mod.shape[-1]):
new_file_name = "normalized_" + str(mods[modality])
new_patient_folder = destination.joinpath(patient_paths[modality])
if not new_patient_folder.exists():
new_patient_folder.mkdir()
new_dest = new_patient_folder.joinpath(new_file_name)
a = nib.Nifti1Image( multi_mod[:,:,:,modality], affine=affines_list[modality] )
nib.save( a, new_dest )
# # Normalize and Save the files in folder: .../MICCAI_BraTS_2019_Data_Training/MICCAI_BraTS_2019_Data_Training/normalized_hgg
# +
patient_paths = hu.get_each_hgg_folder()
normalized_hgg = patient_paths[0].parent.parent
normalized_hgg = normalized_hgg.joinpath("normalized_hgg")
#print(normalized_hgg)
if not normalized_hgg.exists():
normalized_hgg.mkdir()
for patient in tqdm(patient_paths):
X, affines, paths = hu.get_a_multimodal_tensor( patient )
normalize_each_modality_by_its_max(X)
save_data(X, affines, paths, normalized_hgg)
# -
|
normalize_and_save_all_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from scipy import misc
from scipy import ndimage
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
# -
# Load image into variable and display it
lion = misc.imread("assets/Test/test.png") # Paste address of image
plt.imshow(lion, cmap = plt.get_cmap('gray'))
plt.show()
# Convert color image to grayscale to help extraction of edges and plot it
lion_gray = np.dot(lion[...,:3], [0.299, 0.587, 0.114])
#lion_gray = lion_gray.astype('int32')
plt.imshow(lion_gray, cmap = plt.get_cmap('gray'))
plt.show()
# Blur the grayscale image so that only important edges are extracted and the noisy ones ignored
lion_gray_blurred = ndimage.gaussian_filter(lion_gray, sigma=1.4) # Note that the value of sigma is image specific so please tune it
plt.imshow(lion_gray_blurred, cmap = plt.get_cmap('gray'))
plt.show()
# Apply Sobel Filter using the convolution operation
# Note that in this case I have used the filter to have a maximum amgnitude of 2, but it can also be changed to other numbers for aggressive edge extraction
# For eg [-1,0,1], [-5,0,5], [-1,0,1]
def SobelFilter(img, direction):
if(direction == 'x'):
Gx = np.array([[-1,0,+1], [-2,0,+2], [-1,0,+1]])
Res = ndimage.convolve(img, Gx)
#Res = ndimage.convolve(img, Gx, mode='constant', cval=0.0)
if(direction == 'y'):
Gy = np.array([[-1,-2,-1], [0,0,0], [+1,+2,+1]])
Res = ndimage.convolve(img, Gy)
#Res = ndimage.convolve(img, Gy, mode='constant', cval=0.0)
return Res
# Normalize the pixel array, so that values are <= 1
def Normalize(img):
#img = np.multiply(img, 255 / np.max(img))
img = img/np.max(img)
return img
# Apply Sobel Filter in X direction
gx = SobelFilter(lion_gray_blurred, 'x')
gx = Normalize(gx)
plt.imshow(gx, cmap = plt.get_cmap('gray'))
plt.show()
# Apply Sobel Filter in Y direction
gy = SobelFilter(lion_gray_blurred, 'y')
gy = Normalize(gy)
plt.imshow(gy, cmap = plt.get_cmap('gray'))
plt.show()
# +
# Apply the Sobel Filter using the inbuilt function of scipy, this was done to verify the values obtained from above
# Also differnet modes can be tried out for example as given below:
#dx = ndimage.sobel(lion_gray_blurred, axis=1, mode='constant', cval=0.0) # horizontal derivative
#dy = ndimage.sobel(lion_gray_blurred, axis=0, mode='constant', cval=0.0) # vertical derivative
dx = ndimage.sobel(lion_gray_blurred, axis=1) # horizontal derivative
dy = ndimage.sobel(lion_gray_blurred, axis=0) # vertical derivative
# -
# Plot the derivative filter values obtained using the inbuilt function
plt.subplot(121)
plt.imshow(dx, cmap = plt.get_cmap('gray'))
plt.subplot(122)
plt.imshow(dy, cmap = plt.get_cmap('gray'))
plt.show()
# Calculate the magnitude of the gradients obtained
Mag = np.hypot(gx,gy)
Mag = Normalize(Mag)
plt.imshow(Mag, cmap = plt.get_cmap('gray'))
plt.show()
# Calculate the magnitude of the gradients obtained using the inbuilt function, again done to verify the correctness of the above value
mag = np.hypot(dx,dy)
mag = Normalize(mag)
plt.imshow(mag, cmap = plt.get_cmap('gray'))
plt.show()
# Calculate direction of the gradients
Gradient = np.degrees(np.arctan2(gy,gx))
# Calculate the direction of the gradients obtained using the inbuilt sobel function
gradient = np.degrees(np.arctan2(dy,dx))
# +
# Do Non Maximum Suppression with interpolation to get a better estimate of the magnitude values of the pixels in the gradient direction
# This is done to get thin edges
def NonMaxSupWithInterpol(Gmag, Grad, Gx, Gy):
NMS = np.zeros(Gmag.shape)
for i in range(1, int(Gmag.shape[0]) - 1):
for j in range(1, int(Gmag.shape[1]) - 1):
if((Grad[i,j] >= 0 and Grad[i,j] <= 45) or (Grad[i,j] < -135 and Grad[i,j] >= -180)):
yBot = np.array([Gmag[i,j+1], Gmag[i+1,j+1]])
yTop = np.array([Gmag[i,j-1], Gmag[i-1,j-1]])
x_est = np.absolute(Gy[i,j]/Gmag[i,j])
if (Gmag[i,j] >= ((yBot[1]-yBot[0])*x_est+yBot[0]) and Gmag[i,j] >= ((yTop[1]-yTop[0])*x_est+yTop[0])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] > 45 and Grad[i,j] <= 90) or (Grad[i,j] < -90 and Grad[i,j] >= -135)):
yBot = np.array([Gmag[i+1,j] ,Gmag[i+1,j+1]])
yTop = np.array([Gmag[i-1,j] ,Gmag[i-1,j-1]])
x_est = np.absolute(Gx[i,j]/Gmag[i,j])
if (Gmag[i,j] >= ((yBot[1]-yBot[0])*x_est+yBot[0]) and Gmag[i,j] >= ((yTop[1]-yTop[0])*x_est+yTop[0])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] > 90 and Grad[i,j] <= 135) or (Grad[i,j] < -45 and Grad[i,j] >= -90)):
yBot = np.array([Gmag[i+1,j] ,Gmag[i+1,j-1]])
yTop = np.array([Gmag[i-1,j] ,Gmag[i-1,j+1]])
x_est = np.absolute(Gx[i,j]/Gmag[i,j])
if (Gmag[i,j] >= ((yBot[1]-yBot[0])*x_est+yBot[0]) and Gmag[i,j] >= ((yTop[1]-yTop[0])*x_est+yTop[0])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] > 135 and Grad[i,j] <= 180) or (Grad[i,j] < 0 and Grad[i,j] >= -45)):
yBot = np.array([Gmag[i,j-1] ,Gmag[i+1,j-1]])
yTop = np.array([Gmag[i,j+1] ,Gmag[i-1,j+1]])
x_est = np.absolute(Gy[i,j]/Gmag[i,j])
if (Gmag[i,j] >= ((yBot[1]-yBot[0])*x_est+yBot[0]) and Gmag[i,j] >= ((yTop[1]-yTop[0])*x_est+yTop[0])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
return NMS
# -
# This is also non-maxima suppression but without interpolation i.e. the pixel closest to the gradient direction is used as the estimate
def NonMaxSupWithoutInterpol(Gmag, Grad):
NMS = np.zeros(Gmag.shape)
for i in range(1, int(Gmag.shape[0]) - 1):
for j in range(1, int(Gmag.shape[1]) - 1):
if((Grad[i,j] >= -22.5 and Grad[i,j] <= 22.5) or (Grad[i,j] <= -157.5 and Grad[i,j] >= 157.5)):
if((Gmag[i,j] > Gmag[i,j+1]) and (Gmag[i,j] > Gmag[i,j-1])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] >= 22.5 and Grad[i,j] <= 67.5) or (Grad[i,j] <= -112.5 and Grad[i,j] >= -157.5)):
if((Gmag[i,j] > Gmag[i+1,j+1]) and (Gmag[i,j] > Gmag[i-1,j-1])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] >= 67.5 and Grad[i,j] <= 112.5) or (Grad[i,j] <= -67.5 and Grad[i,j] >= -112.5)):
if((Gmag[i,j] > Gmag[i+1,j]) and (Gmag[i,j] > Gmag[i-1,j])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
if((Grad[i,j] >= 112.5 and Grad[i,j] <= 157.5) or (Grad[i,j] <= -22.5 and Grad[i,j] >= -67.5)):
if((Gmag[i,j] > Gmag[i+1,j-1]) and (Gmag[i,j] > Gmag[i-1,j+1])):
NMS[i,j] = Gmag[i,j]
else:
NMS[i,j] = 0
return NMS
# Get the Non-Max Suppressed output
NMS = NonMaxSupWithInterpol(Mag, Gradient, gx, gy)
NMS = Normalize(NMS)
plt.imshow(NMS, cmap = plt.get_cmap('gray'))
plt.show()
# Get the Non-max suppressed output on the same image but using the image using the inbuilt sobel operator
nms = NonMaxSupWithInterpol(mag, gradient, dx, dy)
nms = Normalize(nms)
plt.imshow(nms, cmap = plt.get_cmap('gray'))
plt.show()
# Double threshold Hysterisis
# Note that I have used a very slow iterative approach for ease of understanding, a faster implementation using recursion can be done instead
# This recursive approach would recurse through every strong edge and find all connected weak edges
def DoThreshHyst(img):
highThresholdRatio = 0.2
lowThresholdRatio = 0.15
GSup = np.copy(img)
h = int(GSup.shape[0])
w = int(GSup.shape[1])
highThreshold = np.max(GSup) * highThresholdRatio
lowThreshold = highThreshold * lowThresholdRatio
x = 0.1
oldx=0
# The while loop is used so that the loop will keep executing till the number of strong edges do not change, i.e all weak edges connected to strong edges have been found
while(oldx != x):
oldx = x
for i in range(1,h-1):
for j in range(1,w-1):
if(GSup[i,j] > highThreshold):
GSup[i,j] = 1
elif(GSup[i,j] < lowThreshold):
GSup[i,j] = 0
else:
if((GSup[i-1,j-1] > highThreshold) or
(GSup[i-1,j] > highThreshold) or
(GSup[i-1,j+1] > highThreshold) or
(GSup[i,j-1] > highThreshold) or
(GSup[i,j+1] > highThreshold) or
(GSup[i+1,j-1] > highThreshold) or
(GSup[i+1,j] > highThreshold) or
(GSup[i+1,j+1] > highThreshold)):
GSup[i,j] = 1
x = np.sum(GSup == 1)
GSup = (GSup == 1) * GSup # This is done to remove/clean all the weak edges which are not connected to strong edges
return GSup
# The output of canny edge detection
Final_Image = DoThreshHyst(NMS)
plt.imshow(Final_Image, cmap = plt.get_cmap('gray'))
plt.show()
# The output of canny edge detection using the inputs obtaind using the inbuilt sobel operator
# Notice that the output here looks better than the one above, this might be because of the low magnitude of filter value used in our implementation of the Sobel Operator
# Changing the filter to a higher value leads to more aggressive edge extraction and thus a better output.
final_image = DoThreshHyst(nms)
plt.imshow(final_image, cmap = plt.get_cmap('gray'))
plt.show()
|
CaseStudies/Canny_Edge_Detector/Simple_for_Readme.ipynb
|