code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Simulating quantum programs with large number of qubits
#
# The quantum programs with large number of qubits can be efficiently simulated using the [sparse simulator](https://docs.microsoft.com/azure/quantum/machines/sparse-simulator).
#
# The sparse simulator is a simulator that utilizes a sparse representation of quantum state vectors, as opposed to the [full state simulator](https://docs.microsoft.com/azure/quantum/user-guide/machines/full-state-simulator). This feature allows the sparse simulator to minimize the memory footprint used to represent quantum states, thus enabling simulations over a larger number of qubits. The sparse simulator is efficient for representing quantum states that are sparse in the computational basis, that is, quantum states for which most of the amplitude coefficients are zero in the computational basis. As such, sparse simulator enables users to explore larger applications than what can be represented using the full state simulator which will waste both memory and time on an exponentially large number of zero-amplitudes.
#
# For more information about the sparse simulator, please see [<NAME> (arXiv:2105.01533)](https://arxiv.org/abs/2105.01533).
#
# In this sample, we'll look at the advantages of the sparse simulator over the full state simulator.
#
# Below, we'll create a few clusters (groups) of qubits, each prepared in a [GHZ state](https://en.wikipedia.org/wiki/Greenberger%E2%80%93Horne%E2%80%93Zeilinger_state).
# To run the cells below, click the cell and press `<Ctrl+Enter>`. The same shortcut will let you exit from the Edit mode of a Markdown cell like the one you are reading.
# To see the other shortcuts press `<Esc>`, `<h>`. Press `<Ctrl+Shift+p>` to see the command palette. `<Esc>` to exit.
# Below, we'll open a few Q# namespaces to provide access to functions for creating manipulating arrays of data, and for helping to visualize our quantum programs. To learn more, check out the [sample on visualizing quantum programs](https://github.com/microsoft/Quantum/tree/main/samples/diagnostics/visualization) and the [Q# API reference](https://docs.microsoft.com/qsharp/api/qsharp/).
open Microsoft.Quantum.Arrays;
open Microsoft.Quantum.Diagnostics;
/// # Summary
/// Prepares, measures, and resets the GHZ states.
///
/// # Input
/// ## nQubits
/// Number of qubits to allocate (must be a multiple of the nClusters input).
///
/// ## nClusters
/// Number of clusters to break-up qubits into.
operation PrepareAndMeasureGhzState(nQubits : Int, nClusters : Int) : Unit {
Message($"Allocating {nQubits} qubits...");
// Allocate the qubits:
use qs = Qubit[nQubits];
Message("Finished allocating!");
// Number of qubits in every cluster:
let clusterSize = nQubits / nClusters;
Message($"Creating {nClusters} clusters of GHZ states between {clusterSize} qubits...");
// Iterate through the clusters:
for cluster in Chunks(clusterSize, qs) {
// Apply H gate to the first-most qubit in the cluster:
H(Head(cluster));
// Apply CNOT gate to the remaining qubits in the cluster:
ApplyCNOTChain(cluster);
// After this, all qubits in the cluster will have the same state.
}
Message("Finished GHZ clusters!");
// For small states
if nQubits < 15 {
// show the histogram:
DumpMachine();
}
Message("Collapse and show final state:");
// Measure the qubits:
let results = ForEach(M, qs);
Message($"{results}");
Message("Cleaning up...");
ResetAll(qs);
Message("Done");
}
# We'll make the small states displayed conveniently on a histogram.
# For more information about `dump.basisStateLabelingConvention`, please see [Configuration settings](https://docs.microsoft.com/qsharp/api/iqsharp-magic/config#configuration-settings).
%config dump.basisStateLabelingConvention = "bitstring"
# Now we'll run our Q# operation on the full state simulator.
%simulate PrepareAndMeasureGhzState nQubits=6 nClusters=2
# In the cell above increase the `nQubits` input (by multiples of `nClusters`) such that the simulation becomes slow (e.g. `nQubits`: 8, 10, 12, 14, ...; `nClusters`: 2). The simulation is expected to become slow at `nQubits=24` and to take about a minute with `nQubits=26`. It is not recommended to make large steps, otherwise the simulation can run too long.
# After the number of qubits exceeds 15, the state histogram will not be displayed.
# Now we'll run our Q# operation on the sparse simulator, using the [%simulate_sparse](https://docs.microsoft.com/qsharp/api/iqsharp-magic/simulate_sparse) magic command.
%simulate_sparse PrepareAndMeasureGhzState nQubits=6 nClusters=2
# From the histogram above you can see that the sparse simulator only stores non-zero amplitudes, as opposed to the full state simulator (although the visualization functionality allows truncating out amplitudes that are close to zero when printing from the full state simulator as well).
#
# By increasing the `nQubits` parameter you can see that the sparse simulator still provides the result quickly enough, even for qubit registers large enough that the full state simulator slows down.
#
# You can increase the number of qubits up to 1,024. That is the current hardcoded limit in the sparse simulator.
%simulate_sparse PrepareAndMeasureGhzState nQubits=1024 nClusters=8
# Now you see that the sparse simulator enables users to explore larger applications than what can be represented using full state simulator alone.
# ## See Also
# * [Quantum simulators](https://docs.microsoft.com/en-us/azure/quantum/user-guide/machines/)
# * [Q# API reference](https://docs.microsoft.com/qsharp/api/qsharp/)
|
samples/getting-started/simulation/LargeSimulation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
pd.set_option('display.max_columns', 200)
from app.evaluation import complete_columns, drop_text_features, get_accuracy_and_plot_confusion
from xgboost import XGBClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
df = pd.DataFrame.from_csv("data/processed_data.csv")
val_df = pd.DataFrame.from_csv("data/validation_data.csv")
y_train = df.pop("label")
y_val = val_df.pop("label")
df.fillna(0, inplace=True)
val_df.fillna(0, inplace=True)
df, val_df = complete_columns(df, val_df)
df = drop_text_features(df)
val_df = drop_text_features(val_df)
# -
# Encode labels
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder().fit(y_train)
# +
# Train classifier
clf = XGBClassifier()
clf.fit(df.values, y_train)
y_valid_pred = clf.predict(val_df.values)
print get_accuracy_and_plot_confusion(y_val, y_valid_pred, le.classes_)
# -
|
Validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Code to get landmarks from mediapipe and then create a bounding box around dip joints to segment them
# for further analysis
# Use distance transform to further correct the landmarks from mediapipe
# See if you can get tip indices from drawing a convex hull around hand??
# NOTE: Current code works on already segmented masks - this preprocessing step is required!
# +
# import required libraries
import cv2 as cv # opencv
import numpy as np # for numerical calculations
import synapseclient # synapse login etc
import synapseutils # to download files using folder structure
import pandas as pd # data frames
from matplotlib import pyplot as plt # plotting
import mediapipe as mp # for detecting hand landmarks
import seaborn as sns # for violin plot
import os # for listing files in a folder
import timeit # to track program running time
mp_drawing = mp.solutions.drawing_utils
mp_hands = mp.solutions.hands
# +
# login into Synapse
syn = synapseclient.login()
## hand landmark detection using mediapipe
hands = mp_hands.Hands(static_image_mode=True,
max_num_hands=1, # use one hand at a time
min_detection_confidence=0.5)
# +
def getTIPLandmarks(multi_hand_landmarks, img_shape):
# input is results.multi_hand_landmarks
# will focus on all fingers except thumb - i.e index, middle, ring and pinky
for hand_landmarks in multi_hand_landmarks:
index_finger_tip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_TIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_TIP].y* img_shape[0])]
middle_finger_tip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_TIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_TIP].y* img_shape[0])]
ring_finger_tip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_TIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_TIP].y* img_shape[0])]
pinky_tip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_TIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_TIP].y* img_shape[0])]
tips_dict = {'index': index_finger_tip, 'middle': middle_finger_tip, 'ring': ring_finger_tip, 'pinky': pinky_tip}
return(tips_dict)
def getDIPLandmarks(multi_hand_landmarks, img_shape):
# input is results.multi_hand_landmarks
# will focus on all fingers except thumb - i.e index, middle, ring and pinky
for hand_landmarks in multi_hand_landmarks:
index_finger_dip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_DIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_DIP].y* img_shape[0])]
middle_finger_dip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_DIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_DIP].y* img_shape[0])]
ring_finger_dip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_DIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_DIP].y* img_shape[0])]
pinky_dip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_DIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_DIP].y* img_shape[0])]
dips_dict = {'index': index_finger_dip, 'middle': middle_finger_dip, 'ring': ring_finger_dip, 'pinky': pinky_dip}
return(dips_dict)
def getPIPLandmarks(multi_hand_landmarks, img_shape):
# input is results.multi_hand_landmarks
# will focus on all fingers except thumb - i.e index, middle, ring and pinky
for hand_landmarks in multi_hand_landmarks:
index_finger_pip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_PIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.INDEX_FINGER_PIP].y* img_shape[0])]
middle_finger_pip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_PIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.MIDDLE_FINGER_PIP].y* img_shape[0])]
ring_finger_pip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_PIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.RING_FINGER_PIP].y* img_shape[0])]
pinky_pip = [round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_PIP].x* img_shape[1]), round(hand_landmarks.landmark[mp_hands.HandLandmark.PINKY_PIP].y* img_shape[0])]
dips_dict = {'index': index_finger_pip, 'middle': middle_finger_pip, 'ring': ring_finger_pip, 'pinky': pinky_pip}
return(dips_dict)
def getHalfPoints(landmarks_1, landmarks_2):
# input is two landmarks dictionaries, eg., tips_landmarks and dips_landmarks
index_landmarks = np.round(np.add(landmarks_1['index'], landmarks_2['index'])/2)
middle_landmarks = np.round(np.add(landmarks_1['middle'], landmarks_2['middle'])/2)
ring_landmarks = np.round(np.add(landmarks_1['ring'], landmarks_2['ring'])/2)
pinky_landmarks = np.round(np.add(landmarks_1['pinky'], landmarks_2['pinky'])/2)
return_dict = {'index': index_landmarks, 'middle': middle_landmarks, 'ring': ring_landmarks, 'pinky': pinky_landmarks}
return(return_dict)
def getTipsFromHull(tips_landmarks, hull):
# input is TIPLandmarks and hull points
# For each landmark we find the nearest hull point, i.e matching the hull points
# with the tips, so that we can associate which sections of the contour belong
# to which finger
# index finger
index_dist = list(map(lambda x: np.linalg.norm(x-tips_landmarks['index']), hull[0]))
index_hull_point = hull[0][index_dist.index(min(index_dist))][0]
# middle finger
middle_dist = list(map(lambda x: np.linalg.norm(x-tips_landmarks['middle']), hull[0]))
middle_hull_point = hull[0][middle_dist.index(min(middle_dist))][0]
# ring finger
ring_dist = list(map(lambda x: np.linalg.norm(x-tips_landmarks['ring']), hull[0]))
ring_hull_point = hull[0][ring_dist.index(min(ring_dist))][0]
# pinky
pinky_dist = list(map(lambda x: np.linalg.norm(x-tips_landmarks['pinky']), hull[0]))
pinky_hull_point = hull[0][pinky_dist.index(min(pinky_dist))][0]
tips_hulls_dict = {'index': index_hull_point, 'middle': middle_hull_point, 'ring': ring_hull_point, 'pinky': pinky_hull_point}
return(tips_hulls_dict)
def getClosestPixelInHull(pixel_in, contour_section):
# Given a pixel_in and contour_section, output the pixel in the contour_section that is closese to pixel_in
index_dist = list(map(lambda x: np.linalg.norm(x-pixel_in), contour_section[0]))
index_point = contour_section[0][index_dist.index(min(index_dist))][0]
return(index_point)
def locatePixelInList(pixel,input_list):
# Given a contour find the index of the input pixel (mostly one from the convex hull)
temp_list = list([(input_list[0][x][0] == pixel).all() for x in range(len(input_list[0]))])
# gives a list of true/false
pixel_index = temp_list.index(max(temp_list))
# pick the true
return(pixel_index)
# +
def getHand(img, hand= 'left'):
"""
Split the given two-hand image into a single hand image.
:param img: input RGB image
:param hand: 'left' - left half of the picture
'right' - right half of the image
"""
rows, cols, channels = img.shape
new_cols = round(cols/2)
if hand == 'left':
img_cropped = img[:, 0:new_cols-1,:]
elif hand == 'right':
img_cropped = img[:, new_cols:cols-1,:]
else:
print('Returning input image')
img_cropped = img
return img_cropped
def getBinaryImage(im, min_foreground_pixel = 10, max_foreground_pixel = 255):
# Thresholds the given image(segmented mask with black background) to give a black-white binary image
# with the background being white pixels and foreground(hand) being white pixels
### Convert to Gray Scale
imgray = cv.cvtColor(im, cv.COLOR_BGR2GRAY)
### Binary Threshold
ret, thresh = cv.threshold(imgray, min_foreground_pixel, max_foreground_pixel, cv.THRESH_BINARY)
return(thresh)
def getContoursAndHull(black_white_im):
# Given a binary(black and white image with segmented hand mask) give output of contours, hierarchy
# and convex hull points
# returns only the largest contour as this is the one we want for the hand!
### Find contours
contours, hierarchy = cv.findContours(black_white_im, cv.RETR_TREE, cv.CHAIN_APPROX_NONE)
### contour lengths
contour_lengths = [len(cc) for cc in contours]
largest_contour_index = contour_lengths.index(max(contour_lengths))
# subset contours and hierarchy to the largest contour
contours = [contours[largest_contour_index]]
hierarchy = np.array([[hierarchy[0][largest_contour_index]]])
### Convex Hull
# create hull array for convex hull points
hull = []
# calculate points for each contour
for i in range(len(contours)):
# creating convex hull object for each contour
hull.append(cv.convexHull(contours[i], False))
return({'contours':contours, 'hierarchy':hierarchy, 'hull': hull})
def drawConvexHull(black_white_im, contours, hierarchy, hull, draw_Contour = True):
# given a black white image, contours and hull, output a image with contours and hull drawn
# create an empty black image
# drawing = np.zeros((black_white_im.shape[0], black_white_im.shape[1], 3), np.uint8)
drawing = black_white_im
# draw contours and hull points
color_contours = (0, 255, 0) # green - color for contours
color = (255, 0, 0) # blue - color for convex hull
# draw ith contour
if draw_Contour:
cv.drawContours(drawing, contours, -1, color_contours, 1, 8, hierarchy)
# draw ith convex hull object
cv.drawContours(drawing, hull, -1, color, 1, 8)
return(drawing)
# +
def cropImageFromContour(img, cnt):
# Crop the image(img) based on the input of a closed contour(cnt), set of points
# adapted from https://www.life2coding.com/cropping-polygon-or-non-rectangular-region-from-image-using-opencv-python/
mask = np.zeros(img.shape[0:2], dtype=np.uint8)
# draw the contours on the mask
cv.drawContours(mask, [cnt], -1, (255, 255, 255), -1, cv.LINE_AA)
res = cv.bitwise_and(img,img,mask = mask)
rect = cv.boundingRect(cnt) # returns (x,y,w,h) of the rect
cropped = res[rect[1]: rect[1] + rect[3], rect[0]: rect[0] + rect[2]]
# ## To get white background cropped image
# ## create the white background of the same size of original image
# wbg = np.ones_like(img, np.uint8)*255
# cv.bitwise_not(wbg,wbg, mask=mask)
# # overlap the resulted cropped image on the white background
# dst = wbg+res
# dst_cropped = dst[rect[1]: rect[1] + rect[3], rect[0]: rect[0] + rect[2]]
return(cropped)
def cropWarpedRect(img, rect):
#### BEST
# from https://stackoverflow.com/questions/11627362/how-to-straighten-a-rotated-rectangle-area-of-an-image-using-opencv-in-python/48553593#48553593
# Get center, size, and angle from rect
center, size, theta = rect
box = cv.boxPoints(rect)
box = np.int0(box)
width = int(rect[1][0])
height = int(rect[1][1])
src_pts = box.astype("float32")
dst_pts = np.array([[0, height-1],
[0, 0],
[width-1, 0],
[width-1, height-1]], dtype="float32")
M = cv.getPerspectiveTransform(src_pts, dst_pts)
warped = cv.warpPerspective(img, M, (width, height))
if (theta > 45):
warped = cv.rotate(warped, cv.ROTATE_90_CLOCKWISE)
return(warped)
# -
def getContourSubsectionPercentage(contour_in, pixel_in, percent = 5):
# get subsection of contour that is 5%(default) left and right to the input pixel
input_pixel_location = locatePixelInList(pixel_in, contour_in)
nPixel_contour = len(contour_in[0])
section_left = int(max(0, input_pixel_location - round(nPixel_contour*percent/100)))
section_right = int(min(nPixel_contour-1, input_pixel_location + round(nPixel_contour*percent/100)))
print(section_left)
print(section_right)
subContour = [np.array(contour_in[0][section_left:section_right])]
return(subContour)
# +
def getContourSubsection(contour_in, pixel_on_contour, ref_pixel):
# Given a pixel on the contour, split the contour into two sections left of the pixel
# and right of the pixel.
# Then find the closest points in both sections (left and right) to the ref_pixel, and
# then give subset the contour between these two points
# The idea is when we give a contour of the hand, and a point on it corresponding to the
# finger tip, we get two sections of the contour, to the left of the finger enf then right
# We then find the points closest to these
input_pixel_location = locatePixelInList(pixel_on_contour, contour_in)
nPixel_contour = len(contour_in[0])
# roll/shift the array so that input_pixel is the middle of array
contour_rolled = [np.array(np.roll(contour_in[0],2*(round(nPixel_contour/2)-input_pixel_location)))]
section_left = [np.array(contour_rolled[0][0:round(nPixel_contour/2)])]
section_right = [np.array(contour_rolled[0][round(nPixel_contour/2):nPixel_contour])]
closest_pixel_left = getClosestPixelInHull(ref_pixel, section_left)
closest_pixel_right = getClosestPixelInHull(ref_pixel, section_right)
closest_pixel_left_location = locatePixelInList(closest_pixel_left, contour_rolled)
closest_pixel_right_location = locatePixelInList(closest_pixel_right, contour_rolled)
subContour = [np.array(contour_rolled[0][(closest_pixel_left_location-1):closest_pixel_right_location])]
subContour = [np.array(np.roll(subContour[0],-2*(round(nPixel_contour/2)-input_pixel_location)))]
# subContour = [np.array([[pixel_on_contour], [closest_pixel_left], [closest_pixel_right]])]
# return({'left': closest_pixel_left, 'right': closest_pixel_right})
return(subContour)
# +
def getContourSubsectionJoint(contour_in, pixel_on_contour, ref_pixel_1, ref_pixel_2):
# Given a pixel on the contour, split the contour into two sections left of the pixel
# and right of the pixel.
# Then find the closest points in both sections (left and right) to the ref_pixels, and
# then give subset the contour between these two points
# ref_pixel_2 is farther from tip hull than ref_pixel_1
# The idea is when we give a contour of the hand, and a point on it corresponding to the
# finger tip, we get two sections of the contour, to the left of the finger enf then right
# We then find the points closest to these
input_pixel_location = locatePixelInList(pixel_on_contour, contour_in)
nPixel_contour = len(contour_in[0])
# roll/shift the array so that input_pixel is the middle of array
contour_rolled = [np.array(np.roll(contour_in[0],2*(round(nPixel_contour/2)-input_pixel_location)))]
section_left = [np.array(contour_rolled[0][0:round(nPixel_contour/2)])]
section_right = [np.array(contour_rolled[0][round(nPixel_contour/2):nPixel_contour])]
closest_pixel_left_1 = getClosestPixelInHull(ref_pixel_1, section_left)
closest_pixel_right_1 = getClosestPixelInHull(ref_pixel_1, section_right)
closest_pixel_left_1_location = locatePixelInList(closest_pixel_left_1, contour_rolled)
closest_pixel_right_1_location = locatePixelInList(closest_pixel_right_1, contour_rolled)
closest_pixel_left_2 = getClosestPixelInHull(ref_pixel_2, section_left)
closest_pixel_right_2 = getClosestPixelInHull(ref_pixel_2, section_right)
closest_pixel_left_2_location = locatePixelInList(closest_pixel_left_2, contour_rolled)
closest_pixel_right_2_location = locatePixelInList(closest_pixel_right_2, contour_rolled)
subContour_1 = [np.array(contour_rolled[0][(closest_pixel_left_1_location-1):closest_pixel_right_1_location])]
subContour_1 = [np.array(np.roll(subContour_1[0],-2*(round(nPixel_contour/2)-input_pixel_location)))]
subContour_2 = [np.array(contour_rolled[0][(closest_pixel_left_2_location-1):closest_pixel_right_2_location])]
subContour_2 = [np.array(np.roll(subContour_2[0],-2*(round(nPixel_contour/2)-input_pixel_location)))]
subContour = [np.array([np.array([x]) for x in ((set([(tuple(x[0])) for x in subContour_2[0]]))^(set([(tuple(x[0])) for x in subContour_1[0]])))])]
# subContour = {'1': subContour_1, '2': subContour_2}
# subContour = subContour_2
# subContour = [np.array([[pixel_on_contour], [closest_pixel_left], [closest_pixel_right]])]
# return({'left': closest_pixel_left, 'right': closest_pixel_right})
return(subContour)
# +
# In the working folder (..../), have the following folder structure
# ..../hand_segmentation_psorcast Jun 16 2021 08_51 Dan/
# ..../testHullsDIP
# ..../testHullsDIP/left
# ..../testHullsDIP/right
# ..../segmentedDIP
# ..../segmentedDIP/left
# ..../segmentedDIP/left_unrotated
# ..../segmentedDIP/right
# ..../segmentedDIP/right_unrotated
# +
## Getting hands images from Synapse
hand_masks_synapse_id = 'syn25999658' # Folder containing all merged hand images and masks
img_all_directory = 'hand_segmentation_psorcast Jun 16 2021 08_51 Dan/' # Local path for storing images from Synapse
# Hand mask images form slack (check June 16 - Chat with <NAME>, <NAME> and <NAME>)
# Also in https://www.synapse.org/#!Synapse:syn25999658
# Download the curated hand image files from Synapse
# hands_all_files = synapseutils.syncFromSynapse(syn = syn, entity= hand_masks_synapse_id, path= img_all_directory)
all_files = os.listdir(img_all_directory)
fuse_files = list(filter(lambda x: 'fuse' in x, all_files))
orig_files = list(map(lambda x: x[:-11], fuse_files)) # remove ___fuse.png
target_directory = 'testHullsDIP' # directory with mediapipe results
target_dip_directory = 'segmentedDIP' # directory with segmented DIP joints
# +
## Getting full - resolution hand images from synapse
full_res_images_synapse_id = 'syn25837496' # Folder containing full resolution images
img_full_res_directory = 'hand_images_full_res' # local path to download the files into
# Download the curated hand image files from Synapse
# hands_all_files = synapseutils.syncFromSynapse(syn = syn, entity= full_res_images_synapse_id, path= img_full_res_directory)
all_files = os.listdir(img_all_directory)
# +
## Target Synapse directory for segmented dips
dips_syn_target_folder = 'syn26381177'
def customSynapseUpload(file_path, file_name):
# provenance
provenance_set = synapseclient.Activity(
name = file_name,
used = 'syn25999658')
test_entity = synapseclient.File(
file_path,
parent=dips_syn_target_folder)
test_entity = syn.store(test_entity, activity = provenance_set)
# +
#### LEFT HAND
left_fail_index = 0
left_pass_index = 0
left_dip_rects = {} # save image name and rect dictionaries, i.e image vs dip bounding boxes
for index in range(len(fuse_files)):
# update path for current image
current_image_path = img_all_directory + '/' + fuse_files[index]
# current_orig_path = img_all_directory + '/' + orig_files[index]
current_orig_path = img_full_res_directory + '/' + orig_files[index] # take full res image in
print(current_image_path)
# update target path for the contoured image for the current image
current_left_target_path = target_directory + '/' + 'left/' + fuse_files[index]
# update target path for segmented dips from the current image (ROTATED)
current_left_dip_target_path = target_dip_directory + '/' + 'left'
# update target path for segmented dips from the current image (UN-ROTATED)
current_left_dip_target_path_unrotated = target_dip_directory + '/' + 'left_unrotated'
# read images
img = cv.imread(current_image_path)
orig_img = cv.imread(current_orig_path)
# Masks of left and right img (fuse files)
left_img_fuse = getHand(img, 'left')
# Actual photos
left_orig_img = getHand(orig_img, 'left')
# clones of actual images
left_orig_img_clone = left_orig_img.copy()
# resize image [for faster calculation, and mediapipe usually takes in small images of size
# 300 x 300][https://github.com/google/mediapipe/blob/master/mediapipe/calculators/image/scale_image_calculator.cc]
img_shape = left_orig_img.shape
resize_factor = round(300/max(img_shape),3) # resize max(length, width) to 300 pixels
left_orig_img_resize = cv.resize(left_orig_img, None, fx = resize_factor , fy = resize_factor, interpolation = cv.INTER_AREA)
color_contours = (0, 255, 0) # green - color for contours
### left hand
bw_img_fuse = getBinaryImage(left_img_fuse)
# resize the left hand mask to match that of the original image
img_shape = left_orig_img.shape[0:2]
bw_img = cv.resize(bw_img_fuse, (img_shape[1], img_shape[0]), interpolation = cv.INTER_AREA)
###
contours_hull = getContoursAndHull(bw_img)
###
dw_img = drawConvexHull(left_orig_img, contours_hull['contours'], contours_hull['hierarchy'], contours_hull['hull'])
# apply mediapipe to get hand landmarks
results = hands.process(cv.cvtColor(left_orig_img_resize, cv.COLOR_BGR2RGB))
# to draw all landmarks from mediapipe
if not results.multi_hand_landmarks:
cv.imwrite(current_left_target_path, dw_img)
left_dip_rects[orig_files[index]] = {}
left_fail_index = left_fail_index + 1
else:
dw_img = drawConvexHull(left_orig_img, contours_hull['contours'], contours_hull['hierarchy'], contours_hull['hull'], draw_Contour=True)
for hand_landmarks in results.multi_hand_landmarks:
mp_drawing.draw_landmarks(dw_img, hand_landmarks, mp_hands.HAND_CONNECTIONS)
# get tip landmarks from results
tips_landmarks = getTIPLandmarks(results.multi_hand_landmarks, left_orig_img.shape)
# get DIP landmarks from results
dips_landmarks = getDIPLandmarks(results.multi_hand_landmarks, left_orig_img.shape)
# get PIP landmarks from results
pips_landmarks = getPIPLandmarks(results.multi_hand_landmarks, left_orig_img.shape)
## get middle points b/2 dips_landmarks and pips_landmarks
dip_pip_landmarks = getHalfPoints(dips_landmarks, pips_landmarks)
# get points closest to tips from hull
tips_hull = getTipsFromHull(tips_landmarks, contours_hull['hull'])
## get middle points b/w tips_hull and dips_landmarks
tip_dip_landmarks = getHalfPoints(tips_landmarks, dips_landmarks)
# get subcontours for each finger
subContours = {}
# get minimum bounding rectangle(min area) for each subContour
subContourRects = {}
subContourBoxes = {}
for finger_key in tips_hull:
subContours[finger_key] = getContourSubsectionJoint(contours_hull['contours'], tips_hull[finger_key],tip_dip_landmarks[finger_key], dip_pip_landmarks[finger_key])
# subContours[finger_key] = getContourSubsectionPercentage(contours_hull['contours'], tips_hull[finger_key])
rect = cv.minAreaRect(subContours[finger_key][0])
box = cv.boxPoints(rect)
box = np.int0(box)
subContourRects[finger_key] = rect
subContourBoxes[finger_key] = [box]
# draw SubContours finger
for finger_key in subContours:
cv.drawContours(dw_img, subContours[finger_key], -1, (255,0,0), 1, 8, contours_hull['hierarchy'])
cv.drawContours(dw_img, subContourBoxes[finger_key],-1,(0,0,255))
# rotated via rects
cropped_finger = cropWarpedRect(left_orig_img_clone, subContourRects[finger_key])
curr_file_name = 'left_' + finger_key + '_' + fuse_files[index]
curr_path = current_left_dip_target_path + '/' + curr_file_name
if min(cropped_finger.shape) >0 :
cv.imwrite(curr_path, cropped_finger)
# upload this corrected cropped finger to synapse via customSynpaseUpload
customSynapseUpload(curr_path, curr_file_name)
# unrotated fingers
cropped_finger_unrotated = cropImageFromContour(left_orig_img_clone, subContourBoxes[finger_key][0])
curr_file_name = 'left_' + finger_key + '_' + fuse_files[index]
curr_path = current_left_dip_target_path_unrotated + '/' + curr_file_name
if min(cropped_finger_unrotated.shape) >0 :
cv.imwrite(curr_path, cropped_finger_unrotated)
cv.imwrite(current_left_target_path, dw_img)
left_dip_rects[orig_files[index]] = subContourRects
left_pass_index = left_pass_index + 1
print('left fail percent')
print(100*left_fail_index/(left_fail_index+left_pass_index))
# +
#### RIGHT HAND
right_fail_index = 0
right_pass_index = 0
right_dip_rects = {} # save image name and rect dictionaries, i.e image vs nail bounding boxes
for index in range(len(fuse_files)):
# update path for current image
current_image_path = img_all_directory + '/' + fuse_files[index]
# current_orig_path = img_all_directory + '/' + orig_files[index]
current_orig_path = img_full_res_directory + '/' + orig_files[index] # take full res image in
print(current_image_path)
# update target path for the contoured image for the current image
current_right_target_path = target_directory + '/' + 'right/' + fuse_files[index]
# update target path for segmented dips from the current image (ROTATED)
current_right_dip_target_path = target_dip_directory + '/' + 'right'
# update target path for segmented dips from the current image (UN-ROTATED)
current_right_dip_target_path_unrotated = target_dip_directory + '/' + 'right_unrotated'
# read images
img = cv.imread(current_image_path)
orig_img = cv.imread(current_orig_path)
# Masks of left and right img (fuse files)
right_img_fuse = getHand(img, 'right')
# Actual photos
right_orig_img = getHand(orig_img, 'right')
# clones of actual images
right_orig_img_clone = right_orig_img.copy()
# resize image [for faster calculation, and mediapipe usually takes in small images of size
# 300 x 300][https://github.com/google/mediapipe/blob/master/mediapipe/calculators/image/scale_image_calculator.cc]
img_shape = right_orig_img.shape
resize_factor = round(300/max(img_shape),3) # resize max(length, width) to 300 pixels
right_orig_img_resize = cv.resize(right_orig_img, None, fx = resize_factor , fy = resize_factor, interpolation = cv.INTER_AREA)
color_contours = (0, 255, 0) # green - color for contours
### right hand
bw_img_fuse = getBinaryImage(right_img_fuse)
# resize the right hand mask to match that of the original image
img_shape = right_orig_img.shape[0:2]
bw_img = cv.resize(bw_img_fuse, (img_shape[1], img_shape[0]), interpolation = cv.INTER_AREA)
###
contours_hull = getContoursAndHull(bw_img)
###
dw_img = drawConvexHull(right_orig_img, contours_hull['contours'], contours_hull['hierarchy'], contours_hull['hull'])
# apply mediapipe to get hand landmarks
results = hands.process(cv.cvtColor(right_orig_img_resize, cv.COLOR_BGR2RGB))
# to draw all landmarks from mediapipe
if not results.multi_hand_landmarks:
cv.imwrite(current_right_target_path, dw_img)
right_dip_rects[orig_files[index]] = {}
right_fail_index = right_fail_index + 1
else:
dw_img = drawConvexHull(right_orig_img, contours_hull['contours'], contours_hull['hierarchy'], contours_hull['hull'], draw_Contour=True)
for hand_landmarks in results.multi_hand_landmarks:
mp_drawing.draw_landmarks(dw_img, hand_landmarks, mp_hands.HAND_CONNECTIONS)
# get tip landmarks from results
tips_landmarks = getTIPLandmarks(results.multi_hand_landmarks, right_orig_img.shape)
# get DIP landmarks from results
dips_landmarks = getDIPLandmarks(results.multi_hand_landmarks, right_orig_img.shape)
# get PIP landmarks from results
pips_landmarks = getPIPLandmarks(results.multi_hand_landmarks, right_orig_img.shape)
## get middle points b/2 dips_landmarks and pips_landmarks
dip_pip_landmarks = getHalfPoints(dips_landmarks, pips_landmarks)
# get points closest to tips from hull
tips_hull = getTipsFromHull(tips_landmarks, contours_hull['hull'])
## get middle points b/w tips_hull and dips_landmarks
tip_dip_landmarks = getHalfPoints(tips_landmarks, dips_landmarks)
# get subcontours for each finger
subContours = {}
# get minimum bounding rectangle(min area) for each subContour
subContourRects = {}
subContourBoxes = {}
for finger_key in tips_hull:
subContours[finger_key] = getContourSubsectionJoint(contours_hull['contours'], tips_hull[finger_key],tip_dip_landmarks[finger_key], dip_pip_landmarks[finger_key])
# subContours[finger_key] = getContourSubsectionPercentage(contours_hull['contours'], tips_hull[finger_key])
rect = cv.minAreaRect(subContours[finger_key][0])
box = cv.boxPoints(rect)
box = np.int0(box)
subContourRects[finger_key] = rect
subContourBoxes[finger_key] = [box]
# draw SubContours finger
for finger_key in subContours:
cv.drawContours(dw_img, subContours[finger_key], -1, (255,0,0), 1, 8, contours_hull['hierarchy'])
cv.drawContours(dw_img, subContourBoxes[finger_key],-1,(0,0,255))
# rotated via rects
cropped_finger = cropWarpedRect(right_orig_img_clone, subContourRects[finger_key])
curr_file_name = 'right_' + finger_key + '_' + fuse_files[index]
curr_path = current_right_dip_target_path + '/' + curr_file_name
if min(cropped_finger.shape) >0 :
cv.imwrite(curr_path, cropped_finger)
# upload this corrected cropped finger to synapse via customSynpaseUpload
customSynapseUpload(curr_path, curr_file_name)
# unrotated fingers
cropped_finger_unrotated = cropImageFromContour(right_orig_img_clone, subContourBoxes[finger_key][0])
curr_file_name = 'right_' + finger_key + '_' + fuse_files[index]
curr_path = current_right_dip_target_path_unrotated + '/' + curr_file_name
if min(cropped_finger_unrotated.shape) >0 :
cv.imwrite(curr_path, cropped_finger_unrotated)
cv.imwrite(current_right_target_path, dw_img)
right_dip_rects[orig_files[index]] = subContourRects
right_pass_index = right_pass_index + 1
print('right fail percent')
print(100*right_fail_index/(right_fail_index+right_pass_index))
# +
# Minimum bounding rects of dips into a dataframe (left hand)
aa_left = pd.DataFrame.from_dict(left_dip_rects,orient = 'index')
aa_left.columns = 'left_' + aa_left.columns
aa_left['image'] = aa_left.index
aa_left.index = range(len(aa_left.index))
# Minimum bounding rects of dips into a dataframe (right hand)
aa_right = pd.DataFrame.from_dict(right_dip_rects,orient = 'index')
aa_right.columns = 'right_' + aa_right.columns
aa_right['image'] = aa_right.index
aa_right.index = range(len(aa_right.index))
# Merge rects from left and right hands
aa = pd.merge(aa_left, aa_right, on = 'image', how = 'outer')
aa.head()
# +
### Upload nail bounding rects to Synapse
## Write to csv
aa.to_csv('dip_bounding_rects.csv')
# Synapse target folder
syn_target_folder = 'syn22342373'
# Upload results to Synapse
# provenance
provenance_set = synapseclient.Activity(
name='dip_bounding_rects.csv',
description='Minimum bounding rectangle for dips (except thumb). Each rectangle is of the form (center, size, theta)',
used = 'syn25999658',
executed = 'https://github.com/itismeghasyam/psorcast-validation-analysis/blob/main/analysis/handImaging_analysis/dip_segmentation_mediapipe_pipeline.ipynb')
test_entity = synapseclient.File(
'dip_bounding_rects.csv',
description='Minimum bounding rectangle for dips (except thumb). Each rectangle is of the form (center, size, theta)',
parent=syn_target_folder)
test_entity = syn.store(test_entity, activity = provenance_set)
|
analysis/handImaging_analysis/dip_segmentation_mediapipe_pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/davidmuna/Hypothesis-Testing/blob/master/Autolib_Hypothesis_Testing_Notebook_week_4_Core.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="mh5V-mRF29qO" colab_type="text"
# <font color="green">*Working as a data scientist in an electric car-sharing service company.I will work as a Data Scientist to investigate a claim about the blue cars from the provided Autolib dataset.*</font>
# + [markdown] id="m0tgCkvzPLEa" colab_type="text"
# # Autolib Car Sharing Service Hypothesis Testing
#
# * Link to [Report](https://docs.google.com/document/d/1o8l1XDqZUZ0TZUNf6f0WIXxjjizHGrSyBHcgs-plJVM/edit?usp=sharing)
#
# + [markdown] colab_type="text" id="VSnOCZ5-IPCi"
# ## 1.0 Importing our Libraries
# + id="3r-KDLvrO44a" colab_type="code" colab={}
# Importing the pandas and numpy libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="HToJZ2gBPawZ" colab_type="text"
# ## 1.1 Loading the Dataset from our CSV files
#
#
#
# + [markdown] id="LBUr8VUXQX-K" colab_type="text"
#
# * Dataset link for download or access: [http://bit.ly/DSCoreAutolibDataset
#
# * Dataset Description link for download or access:[http://bit.ly/DSCoreAutolibDatasetGlossary
#
#
#
#
# + [markdown] colab_type="text" id="Vd-5Xne-uCui"
#
#
# Creating and reading our DataFrame
#
#
#
#
# + id="1acwmZanRNHs" colab_type="code" colab={}
# reading the dataset and creating a dataframe
autolib = pd.read_csv('http://bit.ly/DSCoreAutolibDataset')
# + [markdown] id="IICLGbvBRYHq" colab_type="text"
# ## 1.2 Previewing our dataframe and accessing information about it
#
# + id="WrwHg-hzR5BY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 179} outputId="5e2ad1db-4d90-4e76-abfa-318ba7225256"
# Previewing our dataframe (3 rows)
autolib.head(3)
# + [markdown] id="RAVzK5O6IR8v" colab_type="text"
# ## 1.3 Accessing Information about our Dataset
# + id="0d2kfemDIYWI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="17766050-e161-4acd-8b94-d4162dd9488c"
#Accessing information about our data
autolib.info()
autolib.shape
#Dataset has no missing data
# + [markdown] id="R9S2o1WXIhq1" colab_type="text"
# ## 1.4 <font color="green">Cleaning and dealing with outliers, anomalies and missing data</font>
# + id="8n6lZYUT0X8O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="6d8fbfa0-b8b7-4aa2-b79c-be2b555eaecf"
# Renaming columns, fixing syntax errors and putting column names them in lower case for standardisation
autolib.columns = autolib.columns.str.strip().str.lower().str.replace(' ', '_').str.replace('1.4', '14').str.replace('_sum', '')
#Changing date column to datetime
autolib['date']= pd.to_datetime(autolib['date'])
#Dropping unwanted columns
autolib1 = autolib.drop(columns=['utilib_taken','utilib_returned','utilib_14_taken','utilib_14_returned','bluecars_returned','slots_taken'])
#Date column is the correct data type, let’s set it as the DataFrame’s index
autolib1 = autolib1.set_index('date')
autolib1.head(3)
# + id="g-qzyoFDBzA3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="ae6d063b-02e2-480e-934d-c05878f8b193"
#Viewing the tail
autolib1.tail(3)
# + id="fGR0p3580fCX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="35733086-efb0-446b-f9fd-6e912d8a31dd"
#Viewing the presense or absense of Outliers using a box plot for bluecars_taken
autolib1.boxplot(column =['bluecars_taken'], grid = False)
# + id="Y9jL-I7vIaJG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="99ac8804-796c-4b09-d1bc-c4ed296148a1"
#Skewness for bluecars_taken
autolib1['bluecars_taken'].skew()
#bluecars_taken is skewed to the right indicating outliers in the upper quantile and that its slightly not normally distributed
# + id="R7d4gU7uIhVY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="f4bdcbca-562f-4936-891d-381de124d818"
#Checking for Anomalies
#Unique values check
print(autolib1.nunique())
# + [markdown] colab_type="text" id="7bCanJLJQLaq"
# * Unique values in the columns looks okay. No anomalies
# * The outliers in n_daily_data_points and bluecars_taken seem to be a large due to the skewness.However, the blucars_taken is slightly skewed so we'll remove the outliers in the upper quantile range and check skewness again for normality.
# * For slots_freed, there are a large number of outliers as well hence we'll retain them
# * There are no missing values
#
#
#
#
#
#
# + [markdown] id="NicC8P9LMjvO" colab_type="text"
# #### 1.5 RESEARCH QUESTION:
#
# TO IDENTIFY THE TOP THREE POSTAL CODE/AREAS THAT HAVE THE HIGHEST NUMBER OF BLUE CARS TAKEN DURING THE WEEKDAY AND DETERMINE IF THEY ARE THE SAME FOR THE 75015 & 75017 POSTAL CODE AREAS.
# + [markdown] id="8izhaW7UNYGJ" colab_type="text"
# ## <font color="green">Univariate Summaries<font/>
# + id="x3l_5T1ewQSw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="74efdb45-0a48-424d-80fc-e7653083145a"
#Identifying number of blucars taken in every area/postal code during the weekend sorted in descending order
#creating a filter variable for weekend
weekend = autolib1['day_type'] == 'weekend'
autolibweekend = autolib1[weekend].groupby('postal_code')['bluecars_taken'].sum().sort_values(ascending = False).reset_index(name = 'bluecars_taken').rename(columns = {'0':''})
autolibweekend = pd.DataFrame(autolibweekend)
#filtering areas 75015 and 75017
area_15 = autolibweekend['postal_code']==75015
area_17 = autolibweekend['postal_code']==75017
autolibweekend[area_15|area_17]
# + colab_type="code" id="XK69rp1M63xj" colab={"base_uri": "https://localhost:8080/", "height": 111} outputId="04aafbd0-9c43-4462-d5fb-33ba04cd5fe8"
#Identifying number of blucars taken in every area/postal code during the weekday sorted in descending order
#Filter variable for weekday
weekday = autolib1['day_type'] == 'weekday'
autolibweekday = autolib1[weekday].groupby('postal_code')['bluecars_taken'].sum().sort_values(ascending = False).reset_index(name = 'bluecars_taken').rename(columns = {'0':''})
autolibweekday = pd.DataFrame(autolibweekday)
area_15 = autolibweekday['postal_code']==75015
area_17 = autolibweekday['postal_code']==75017
#filtering areas 75015 and 75017
area_17and15_weekday = autolibweekday[area_15|area_17]
area_17and15_weekday
# + [markdown] id="xIfJm8TfTy1Q" colab_type="text"
# Our period of choice is weekdays because we identified theres more activity on weekdays compared to weekends.
# + id="2tlewDKR-Ne-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 310} outputId="852c7efc-e020-4fde-fed8-3685e6fc50f9"
#bluecars taken for area/postal codes 75015 and 75017 bar chart on weekdays
sns.set(style="whitegrid")
plt.title("Blue Cars Taken in Area/Postal Code 75015 & 75017 during weekdays")
plt.tight_layout()
ax = sns.barplot(x="postal_code", y="bluecars_taken", data=area_17and15_weekday)
# + id="us9PpwxYQEiQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="ebb80b5a-b5bc-4b43-d66d-6fd62278486d"
#Summary statistics for the dataset
autolib_summarystat = autolib1.drop(columns=['n_daily_data_points','day_type','slots_freed','dayofweek','postal_code'])
autolib_summarystat.describe()
# + id="fZzFii_sRKRF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="a0c5578c-3a12-4042-cece-c80e3459cf38"
print("Population Variance for ", autolib_summarystat.var())
print("Population Median for ", autolib_summarystat.median())
print("Population Mode for ", autolib_summarystat.median())
print("Population Skew for ", autolib_summarystat.skew())
print("Population Kurtosis for ", autolib_summarystat.kurt())
# + [markdown] id="h2aSJyl4Khb7" colab_type="text"
# ## <font color="green">Bivariate Summaries<font/>
# + id="5RHBhwW9SV9e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 453} outputId="5c476cc8-749a-4a3d-b8c7-e19ce757403d"
#Correlation Matrix
autolib_corr = autolib1.drop(columns=['n_daily_data_points','slots_freed'])
autolib_corr= autolib_corr.corr(method='pearson')
#Chosing the size of the heatmap
plt.figure(figsize = (12,7))
#Displaying correlations using the seaborn heatmap
sns.heatmap(autolib_corr,
xticklabels=autolib_corr.columns,
yticklabels=autolib_corr.columns,
cmap='RdBu_r',
annot=True,
linewidth=0.8)
#Theres a strong negative correlation between the postal code/area and the number of blue cars taken with a coefficient of -0.7
# + id="-T1ILSdKGT_Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 524} outputId="4436ece2-b992-43a4-c397-af7c72135038"
#Plotting a time series for bluecars taken during the weekday for postal codes 75015 and 75017
autolib2 = autolib1[weekday]
autolib3 = autolib2.drop(columns=['n_daily_data_points','day_type','slots_freed','dayofweek'])
area_75015 = autolib3['postal_code']==75015
area_75017 = autolib3['postal_code']==75017
autolib75015 = autolib3[area_75015]
autolib75017 = autolib3[area_75017]
axes = autolib75015['bluecars_taken'].plot(marker='.', alpha=0.5, linestyle='-', figsize=(11, 9), subplots=True, color='blue')
for ax in axes:
ax.set_ylabel('Blue Cars Taken')
ax.set_xlabel('Month')
ax.set_title('Time Series of Blue Cars taken in postal code 75015(Blue) and 75017(Red) within the 6 month period')
axes2 = autolib75017['bluecars_taken'].plot(marker='.', alpha=0.5, linestyle='-', figsize=(11, 9), subplots=True, color='red')
for ax in axes2:
ax.set_ylabel('Blue Cars Taken')
ax.set_xlabel('Month')
ax.set_title('Time Series of Blue Cars taken in postal code 75015(Blue) and 75017(Red) within the 6 month period')
# + id="RsaG9JlCV94y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 322} outputId="e764c049-b4ac-4c90-b6e8-e59b355d3c70"
#Checking Time Series Seasonality by day of the weekday
autolib_dayseason = autolib2.drop(columns=['n_daily_data_points','day_type','slots_freed'])
area_season15 = autolib_dayseason['postal_code']==75015
area_season17 = autolib_dayseason['postal_code']==75017
autolib_dayseason15 = autolib_dayseason[area_75015]
autolib_dayseason17 = autolib_dayseason[area_75017]
sns.boxplot(data=autolib_dayseason15, x='dayofweek', y='bluecars_taken');
plt.xlabel("Day of Week (0-Mon, 4-Fri)", fontsize=15)
plt.ylabel("Blue Cars Taken", fontsize=15)
plt.title("Weekly Seasonality for Blue Cars Taken in Area/Postal Code 75015", fontsize=15)
# + [markdown] colab_type="text" id="HgEKsTLtcUs9"
# Weekly Seasonality for Area 75015 indicates a steady increase in number of bluecars taken from Monday to Friday. Friday being the highest.
# + id="fQWkwlMubjbP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 322} outputId="33a895f4-681c-4b77-a92b-36e1974a410f"
sns.boxplot(data=autolib_dayseason17, x='dayofweek', y='bluecars_taken');
plt.xlabel("Day of Week (0-Mon, 4-Fri)", fontsize=15)
plt.ylabel("Blue Cars Taken", fontsize=15)
plt.title("Weekly Seasonality for Blue Cars Taken in Area/Postal Code 75017", fontsize=15)
# + [markdown] id="E5IbfIz4b0x2" colab_type="text"
# Weekly Seasonality for Area 75017 indicates less Bluecars taken on Thursday and Wednesday. So there's no steady increase from Monday to Friday.
# + [markdown] id="l_-ZTLpnJzVz" colab_type="text"
# Overal Analysis indicates that Blue Cars taken in area 75015 is more than Blue Cars taken in area 75017
# + [markdown] colab_type="text" id="UrnkqDAlUmNj"
# ## 1.6 Implementing the solution through Hypothesis Testing
# + [markdown] colab_type="text" id="lvlrvw6hv8k9"
# #### <font color="green">Formulating the Null and Alternative Hypotheses<font/>
# + [markdown] id="rxXFAnwLdq2b" colab_type="text"
#
#
# * Ho : The number of Blue Cars taken in area/postal code 75015 and 75015 are the same
#
#
# * Ha : The number of Blue Cars taken in area/postal code 75015 is different from Blue Cars taken in area/postal code 75017 (CLAIM)
#
#
# + [markdown] id="lJ_KTThjelIq" colab_type="text"
# Writing the Ho and Ha in mathematical form noting that Null Hypotheses are identified by equality
#
# 1. Ho : Bluecarstaken in 75015 = Bluecarstaken in 75017
#
# 2. Ha : Bluecarstaken in 75015 != Bluecarstaken in 75017(CLAIM)
#
#
# + [markdown] colab_type="text" id="OEyzLpWZy7yn"
# #### <font color="green">Choosing Sample Size and Level of Significance as 5%<font/>
# + [markdown] id="PRfECCXl1pJk" colab_type="text"
# Sampling technique used is Stratified Random sampling to break the population into area 75015 and 75017 subgroups and obtaining a simple random sample from each group.
# + id="TNK_R2Xch6gS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="0352a9ce-24d8-448b-8a08-de5244091fa8"
#Merging the two subgroup original population datasets autolib75015 and autolib75017 to population
population = pd.concat([autolib75015,autolib75017])
print("population is ",population.groupby('postal_code').count())
# + id="hJnbZLz-3-p1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="5f10c245-de71-4e34-aa15-2067a829cd10"
# Stratified sample
strat_sample = population.groupby('postal_code', group_keys=False).apply(lambda grouped_subset : grouped_subset.sample(frac=0.3))
#Proportion of the stratified sample
print(strat_sample['postal_code'].value_counts())
# + [markdown] id="K0T_Yfve4M1s" colab_type="text"
# Our sample size is 68 out of 224
# + [markdown] colab_type="text" id="j8YcGQ4i4ryn"
# #### <font color="green">Choosing the Test Statistic and finding the P Value<font/>
# + [markdown] id="9wdymqo35DXt" colab_type="text"
# Test Statistic chosen is the One Sample Z-Test because we have the population standard deviation from univariate summary statistics
# + id="esKsjGwl6-Tq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4ad3453d-2d56-4d36-f8b5-2164c60f47c7"
from scipy.stats import norm
from scipy import stats
import math
pop_mean = 125.93 #From summary statistics in univariate summary
n = 68
sample_mean = strat_sample['bluecars_taken'].mean()
alpha = 0.05
pop_std = 185.42 #From summary statistics in univariate summary
#Calculating Z score
zscore = ( sample_mean-pop_mean ) / 185.42
#Finding the P Value given that the Null Hypothesis is TRUE
pval = stats.norm.cdf(zscore) * 2 #Since its a two tailed test as per the alternative Ha (ie presence of !=)
print(" Z-Score is :", zscore)
print(" P Value is :", pval)
# + id="0ZyaoKmTjf4T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="310be9c7-0684-42c7-bf5b-559ae4976a8f"
#Constructing a confidence interval around the parameter
from scipy.stats import sem, t
from scipy import mean
confidence = 0.95
data = strat_sample['bluecars_taken']
n = len(data)
m = mean(data)
std_err = sem(data)
h = std_err * t.ppf((1 + confidence) / 2, n - 1)
start = m - h
end = m + h
print("The start is :",round(start))
print("The end is :",round(end))
# + [markdown] colab_type="text" id="KTPDblB5BRwK"
# #### <font color="green">Interpreting P Value and drawing a conclusion<font/>
#
# Theres the possibility of having Type Two Errors in this Hypothesis Test in the event that:
#
# * We fail to reject the null hypothesis when in real sense it is false
#
#
#
# + id="LTwNZSEqBa5w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="645fc82f-ce05-4109-d378-b382bd29af3b"
if pval >alpha:
print("We fail to reject null hypothesis")
print("Conclusion:")
print("Theres no sufficient statistical evidence to support the claim that Bluecarstaken in 75015 != Bluecarstaken in 75017 at the 5% level of significance")
else:
print("We reject null the hypothesis")
print("Conclusion:")
print("Theres is sufficient statistical evidence to support the claim that Bluecarstaken in 75015 != Bluecarstaken in 75017 at the 5% level of significance")
# + [markdown] id="IQIvYW_KHm5Z" colab_type="text"
# #### Conclusion:
# Theres no sufficient statistical evidence to support the claim that Bluecarstaken in 75015 != Bluecarstaken in 75017 at the 5% level of significance
#
# * This is a Type II error because the Null Hypothesis is False
#
#
#
|
Autolib_Hypothesis_Testing_Notebook_week_4_Core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CMASS demonstration
#
# A fresh example of CMASS (Compressive MultiAgent System Sensing) algorithm in operation
import numpy as np
import matplotlib.pyplot as plt
from cmass.agent import *
from cmass.controller import *
from cmass.massimulator import *
from cmass.adj_mat_gen import *
from cmass.sensor import *
from cmass.datacenter import *
# Set simulation parameters
t1 = 60. # total simulation time (in seconds)
tc = 0. # time at which cluster control starts
dt = 0.01 # simulation time step
i1 = int(t1 // dt) # total number of iterations
np.random.seed(2) # set constant seed for ease of monitoring
# Set the number of agents and true number of clusters
n_agents = 16
n_clusters = 4
# Generate adjacency matrix
adj_mat = example_squares_16()
# Generate a map {agent : cluster}
cluster_map = {}
for i in range(n_agents):
cluster_map[i] = i // (n_agents // n_clusters)
# +
# Set local control strength
rho = 0.5
# w_i = 4.1, 4.2, 4.3, 4.4, 8.1, 8.2, 8.3, 8.4, 12.1, 12.2, 12.3, 12.4, 16.1, 16.2, 16.3, 16.4
w_disp = 4.
w_per_cluster = np.arange(1, 5) * 0.1
w_nat_matrix = np.array([(i+1) * w_disp + w_per_cluster for i in range(n_clusters)])
nat_freqs = w_nat_matrix.flatten() / rho
# mu_1 = mu_5 = mu_9 = mu_13 = 0.375,
# mu_2 = mu_6 = mu_10 = mu_14 = 0.75,
# mu_3 = mu_7 = mu_11 = mu_15 = 1.125,
# mu_4 = mu_5 = mu_12 = mu_16 = 1.5
mus = np.tile(np.arange(1, n_clusters+1) / n_clusters, n_clusters) * 1.5
# Generate randomized frequencies for sinusoidal cluster control
f0s = 0.5 * np.sqrt(np.random.rand(n_clusters))
# +
# Sensor parameters
n_pixels = 200
exposure_time = .1
fov_top = 20
fov_bot = 0
sensor_resolution = (fov_top - fov_bot) / n_pixels
cs_m = int(1/2.5 * n_pixels) # 2.5x compression
meas_mat = np.random.randn(cs_m, n_pixels) * 1/cs_m
# -
# Initialize the simulator
sim = KuramotoSimulator(n_agents, dt, adj_mat, cluster_map, tc, rho, mus, f0s, nat_freqs)
# Initialize the sensor
sensor = Sensor(n_pixels, exposure_time, fov_top, fov_bot, dt, meas_mat, cs_m)
# Full progress recording at each iteration
thetas = np.zeros((n_agents, i1))
thetas_dot = np.zeros_like(thetas)
# Simulation iterations with recording
current_states = np.asarray(sim.get_all_outputs())
for iter in range(i1):
thetas[:, iter] = current_states
sim.update_all_states()
new_states = np.asarray(sim.get_all_outputs())
theta_dot = np.divide(new_states - current_states, dt)
thetas_dot[:, iter] = theta_dot
sensor.accept_sample(theta_dot)
current_states = new_states
# Visualize
figure_figsize = (6, 3)
figure_dpi = 200
plt.figure(figsize=figure_figsize, dpi=figure_dpi)
colors = ['#BF616A', '#5E81AC' , '#A3BE8C', '#EBCB8B']
for i in range(n_agents):
color = colors[int(i/n_clusters)]
label = f"Cluster #{int(i/n_clusters)}" if i % n_clusters == 0 else ""
plt.plot(np.linspace(0., t1, i1), thetas_dot[i], '-', color=color, label=label)
plt.xlabel(r'$t$', fontsize=16)
plt.ylabel(r'$\dot{\theta}$', fontsize=16)
plt.xlim(0, i1 * dt)
plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left', borderaxespad=0.)
plt.show()
# Get recorded data
cs_rec, full_rec = sensor.get_recording()
# Visualize full recorded data
figure_figsize = (8, 4)
figure_dpi = 200
plt.figure(figsize=figure_figsize, dpi=figure_dpi)
plt.imshow(full_rec, origin='lower', cmap='gray', extent=[0, t1, fov_bot, fov_top])
plt.xlabel(r'$t$', fontsize=16)
plt.ylabel(r'$\dot{\theta}$', fontsize=16)
plt.show()
# Reconstruct each frame
reconst_rec = np.zeros_like(full_rec)
n_frames = full_rec.shape[1]
for i in range(n_frames):
reconst_rec[:, i] = get_reconst_recording(cs_rec[:, i], meas_mat, n_pixels)
# Visualize reconstructed recorded data
figure_figsize = (8, 4)
figure_dpi = 200
plt.figure(figsize=figure_figsize, dpi=figure_dpi)
plt.imshow(reconst_rec, origin='lower', cmap='gray', extent=[0, t1, fov_bot, fov_top])
plt.xlabel(r'$t$', fontsize=16)
plt.ylabel(r'$\dot{\theta}$', fontsize=16)
plt.show()
# Simple cluster visualizer
def show_clusters(rec, delta, min_cluster):
"""
Visualize cluster centroids and get metrics.
Args:
rec: Array of frames
delta: Clusterization threshold
min_clust: Min size of a cluster, anything below would be considered as an outlier
Returns:
accuracy: Calculated accuracy
"""
plt.figure(figsize=figure_figsize, dpi=figure_dpi)
plt.imshow(rec, origin='lower', cmap='gray', extent=[0, t1, fov_bot, fov_top])
cmax = []
frame_ticks = np.arange(0, t1, exposure_time)
for i in range(frame_ticks.size-1):
x = rec[:, i]
c, l = get_cluster_data(x, delta, min_cluster)
if len(c) > 0:
c = c * sensor_resolution
cmax.append(c.size)
t_val = np.full(c.shape, frame_ticks[i]) + exposure_time * 0.5
plt.scatter(t_val, c, color='r', s=0.3)
accuracy = clusterization_accuracy(cmax, n_clusters)
plt.xlabel(r'$t$', fontsize=16)
plt.ylabel(r'$\dot{\theta}$', fontsize=16)
plt.xlim(0, t1)
plt.ylim(fov_bot, fov_top)
plt.show()
return accuracy
# Set delta (hierarchical clusterization threshold) and minimum cluster (below which are outliers)
delta = 8
min_cluster = 2
# Evaluate full original data
original_accuracy = show_clusters(full_rec, delta, min_cluster)
print("Original frames clusterization accuracy:", original_accuracy)
# Evaluate reconstructed data
reconst_accuracy = show_clusters(reconst_rec, delta, min_cluster)
print("Reconstructed frames clusterization accuracy:", reconst_accuracy)
|
example_16nodes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Objective
#
# Combine centroids found from each crop in this proof-of-concept zoom test and investigate the success of this experiment.
#
# # Result
#
# The centroids found from each crop came together very cleanly. This entire process (auto-generating zoomed-in crops, getting the crops annotated, processing annotation data for each image to get consensus annotations, and adding these consensus annotations back to the parent image) has resulted in a **91% detection rate**. Compare with detection rates of 46% and 56% respectively for the non-inverted and inverted full 700 X 450 pixel image.
#
# # Next step
# Clean up code from these zooming + stitching notebooks and add to the pipeline.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ### Load in centroids from the parent image (with crop regions blacked out)
coords_blacked = np.asarray(pd.read_csv('smfish_blacked_coords.csv'))
for coord in coords_blacked:
plt.scatter(coord[0], 450-coord[1], color = 'black')
plt.title('Centroids from blacked parent image only')
plt.show()
# ### Load in centroids from individual crops
#
# In the below plot, all centroids from a given individual crop are of the same color.
# +
total = []
for coord in coords_blacked:
plt.scatter(coord[0], 450-coord[1], color = 'black')
total.append([coord[0], 450-coord[1]])
colors = ['red', 'orange', 'yellow', 'green', 'cyan', 'blue', 'purple', 'violet', 'magenta']
for i in range(9):
coords = np.asarray(pd.read_csv('smfish_' + str(i) + '_coords.csv'))
stats = np.asarray(pd.read_csv('smfish_' + str(i) + '.csv', header = None))
xmin = stats[0][0]
ymin = stats[2][0]
ymax = stats[3][0]
height = ymax - ymin
zoom_factor = stats[4][0]
refit = []
for coord in coords:
x = coord[0]/zoom_factor + xmin
y = height - coord[1]/zoom_factor + ymin
refit.append([x, y])
total.append([x, y])
refit = np.asarray(refit)
for coord in refit:
plt.scatter(coord[0], 450-coord[1], color = colors[i])
plt.title('Centroids from blacked parent image and from all crops')
plt.show()
# -
# ### All found centroids vs. ground truth
#
# Visually, all found centroids are very close to ground truth.
# +
total = np.asarray(total)
np.savetxt('total_coords.csv', total, delimiter=",", comments='', header = "centroid_x,centroid_y")
coords_total = np.asarray(pd.read_csv('total_coords.csv'))
for coord in coords_total:
plt.scatter(coord[0], coord[1], color = '#d2c295')
plt.title('All found centroids saved to total_coords.csv')
plt.show()
# +
ground_truth = np.asarray(pd.read_csv('700.csv'))
for coord in ground_truth:
plt.scatter(coord[0], coord[1], color = '#5e3032')
plt.title('Ground truth')
plt.show()
# -
for coord in coords_total:
plt.scatter(coord[0], coord[1], color = '#d2c295')
for coord in ground_truth:
plt.scatter(coord[0], coord[1], color = '#5e3032')
plt.title('Found centroids superimposed with ground truth')
plt.show()
# ### Metric of success: detection rate
# +
from sklearn.neighbors import KDTree
def get_nnd(coord, kdt):
dist, ind = kdt.query([coord], k=2)
return dist[0][0]
detected = 0
kdt = KDTree(total, leaf_size=2, metric='euclidean')
threshold = 4
for coord in ground_truth:
nnd = get_nnd(coord, kdt)
if nnd <= threshold:
plt.scatter(coord[0], 450-coord[1], facecolors = 'g')
detected += 1
else:
plt.scatter(coord[0], 450-coord[1], facecolors = 'm')
plt.title('Detected (green) vs. undetected (magenta), threshold = 4 px')
plt.show()
print('detected / total = ' + str(detected) + ' / ' + str(len(ground_truth)) + ' = ' + str(round(float(detected)/len(ground_truth), 2)))
# +
from sklearn.neighbors import KDTree
def get_nnd(coord, kdt):
dist, ind = kdt.query([coord], k=2)
return dist[0][0]
correct = 0
kdt = KDTree(ground_truth, leaf_size=2, metric='euclidean')
threshold = 4
for coord in total:
nnd = get_nnd(coord, kdt)
if nnd <= threshold:
plt.scatter(coord[0], 450-coord[1], facecolors = 'g')
correct += 1
else:
plt.scatter(coord[0], 450-coord[1], facecolors = 'm')
plt.title('Correct (green) vs. Incorrect (magenta), threshold = 4 px')
plt.show()
print('correct / total = ' + str(correct) + ' / ' + str(len(total)) + ' = ' + str(round(float(correct)/len(total), 2)))
|
datasets/zoom_test/stitch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import sqlite3
import seaborn as sns
# +
# Set plotting style
plt.style.use('seaborn-white')
# Metadata saved to output plots
METADATA = {
"Contributor": "<NAME>",
"Rights": "CC-BY-SA 4.0"
}
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
blue_line_color = colors[0]
orange_line_color = colors[1]
green_line_color = colors[2]
# -
# %matplotlib inline
# Show all columns
pd.set_option('display.max_columns', None)
# +
START_YEAR = 2019
LAST_YEAR = 2020
DATE_COL = "collision_date"
CRASH_COL = "crashes"
FACTOR_COL = "vehicle_make"
MAKE_1 = "ford"
MAKE_2 = "toyota"
SQL_FILE = "./v2.0.0-converted_with_sts_v4.0.0-switrs.sqlite"
# + tags=[]
# Date conversion function
def date_columns(query):
"""If a date column is included in the query, parse it as a date in the
dataframe."""
dates = []
fields = ["collision_date", "process_date"]
if '*' in query:
dates = fields
else:
for date in fields:
if date in query:
dates.append(date)
if not dates:
dates = None
return dates
# + tags=[]
def run_query(query, sql_file):
""" Read sqlite query results into a pandas DataFrame. """
with sqlite3.connect(sql_file) as con:
# Construct a Dataframe from the results
df = pd.read_sql_query(
query,
con,
parse_dates = date_columns(query),
)
return df
# + tags=[]
# We should be able to just set rcParams, expect Jupyter has a bug:
# https://github.com/jupyter/notebook/issues/3385
#
# So we have to call this function every time we want to plot.
def setup_plot(title=None, xlabel=None, ylabel=None):
"""Set up a simple, single pane plot with custom configuration.
Args:
title (str, optional): The title of the plot.
xlabel (str, optional): The xlabel of the plot.
ylabel (str, optional): The ylabel of the plot.
Returns:
(fig, ax): A Matplotlib figure and axis object.
"""
# Plot Size
plt.rcParams["figure.figsize"] = (12, 7) # (Width, height)
# Text Size
SMALL = 12
MEDIUM = 16
LARGE = 20
HUGE = 28
plt.rcParams["axes.titlesize"] = HUGE
plt.rcParams["figure.titlesize"] = HUGE
plt.rcParams["axes.labelsize"] = LARGE
plt.rcParams["legend.fontsize"] = LARGE
plt.rcParams["xtick.labelsize"] = MEDIUM
plt.rcParams["ytick.labelsize"] = MEDIUM
plt.rcParams["font.size"] = SMALL
# Legend
plt.rcParams["legend.frameon"] = True
plt.rcParams["legend.framealpha"] = 1
plt.rcParams["legend.facecolor"] = "white"
plt.rcParams["legend.edgecolor"] = "black"
# Figure output
plt.rcParams["savefig.dpi"] = 300
# Make the plol
fig, ax = plt.subplots()
ax.set_title(title)
ax.set_xlabel(xlabel)
ax.set_ylabel(ylabel)
# Make the title and label area opaque instead of transparent
fig.patch.set_facecolor(ax.get_facecolor())
return fig, ax
# + tags=[]
def save_plot(fig, filename):
"""Save the plot with metadata and tight layout.
Args:
fig (matplotlib figure): The figure to save.
filename (str): The loction to save the file to.
"""
metadata = METADATA
fig.savefig(
fname=f"{filename}",
bbox_inches="tight",
metadata=metadata,
)
# -
def normalize_dataframe(df, factor_col=FACTOR_COL, end_date='2019-07-01'):
"""Normalize the dataframe by setting the mean to 1 over the first few months."""
ford_mean = df[(df[FACTOR_COL]=='ford') & (df.index < end_date)].mean()
toyota_mean = df[(df[FACTOR_COL]=='toyota') & (df.index < end_date)].mean()
df_normed = df.copy(deep=True)
df_normed.loc[df[factor_col]=='ford', "total"] /= ford_mean[0]
df_normed.loc[df[FACTOR_COL]=='toyota', "total"] /= toyota_mean[0]
return df_normed
def make_ford_vs_toyota_plot(df, fig, ax, toyota_label_xy, ford_label_xy, stay_at_home_y, norm_label_y=None, y_lim=None, factor_col=FACTOR_COL,):
sns.lineplot(data=df, x="collision_date", y="total", hue=factor_col, drawstyle="steps-post", linewidth=2)
# Add labels instead of legend
ax.get_legend().remove()
ax.text(x=pd.Timestamp(toyota_label_xy[0]), y=toyota_label_xy[1], s="Toyota", color=orange_line_color, fontsize=30)
ax.text(x=pd.Timestamp(ford_label_xy[0]), y=ford_label_xy[1], s="Ford", color=blue_line_color, fontsize=30)
# Add Stay at home order
ax.axvline(x=pd.Timestamp("2020-03-19"), color="red", linewidth=2, zorder=1, label="Stay at home order")
ax.text(x=pd.Timestamp("2020-03-27"), y=stay_at_home_y, s="Stay-at-home Order", color="red", fontsize=24)
# Add mean line
if norm_label_y is not None:
ax.hlines(y=1, xmin=pd.Timestamp("2019-01-01"), xmax=pd.Timestamp("2019-06-30"), linewidth=3, color=green_line_color, zorder=2)
ax.text(x=pd.Timestamp("2018-12-14"), y=norm_label_y, s="Mean normalized from\nJanuary through June", color=green_line_color, fontsize=24)
# Remove date label
ax.xaxis.label.set_visible(False)
if y_lim is not None:
ax.set_ylim(*y_lim)
return fig, ax
# # Crashes by Week
#
# Let's start by looking at the number of incidents per month:
# +
query = f"""
SELECT collision_date
, {FACTOR_COL}
, count(1) as total
FROM collisions AS c
LEFT JOIN parties as p
ON p.case_id = c.case_id
WHERE {DATE_COL} IS NOT NULL
AND {DATE_COL} BETWEEN '{START_YEAR}-01-01' AND '{LAST_YEAR}-11-30'
AND p.{FACTOR_COL} in ('{MAKE_1}', '{MAKE_2}')
GROUP BY 1, 2
ORDER BY 1, total DESC
"""
df = run_query(query, SQL_FILE)
# -
df_grouped = df.groupby([pd.Grouper(key=DATE_COL, freq='W-MON'), FACTOR_COL]).sum().reset_index(FACTOR_COL)
# +
fig, ax = setup_plot(
title="Collisions During the COVID-19 Pandemic",
ylabel="Collisions / Week",
xlabel="",
)
fig, ax = make_ford_vs_toyota_plot(
df_grouped, fig, ax,
toyota_label_xy=("2019-03-01", 2400),
ford_label_xy=("2019-03-01", 1400),
stay_at_home_y=2000,
)
for ext in ("svg", "png"):
save_plot(fig, f"/tmp/covid_pandemic_ford_vs_toyota_collisions.{ext}")
# -
# ## Normalized
df_normed = normalize_dataframe(df_grouped)
# +
fig, ax = setup_plot(
title="Normalized Collisions Rates During the COVID-19 Pandemic",
ylabel="Normalized Collision Rate",
xlabel="",
)
fig, ax = make_ford_vs_toyota_plot(
df_normed, fig, ax,
toyota_label_xy=("2020-08-01", 0.27),
ford_label_xy=("2020-08-01", 0.63),
stay_at_home_y=1,
norm_label_y=0.7,
y_lim=(0, 1.25),
)
for ext in ("svg", "png"):
file_name = f"/tmp/covid_pandemic_normalized_ford_vs_toyota_collisions.{ext}"
save_plot(fig, file_name)
# -
# ## Is this a truck thing?
# +
query4 = f"""
SELECT collision_date
, {FACTOR_COL}
, count(1) as total
FROM collisions AS c
LEFT JOIN parties as p
ON p.case_id = c.case_id
WHERE {DATE_COL} IS NOT NULL
AND {DATE_COL} BETWEEN '{START_YEAR}-01-01' AND '{LAST_YEAR}-11-30'
AND p.{FACTOR_COL} in ('{MAKE_1}', '{MAKE_2}')
AND statewide_vehicle_type = 'pickup or panel truck'
GROUP BY 1, 2
ORDER BY 1, total DESC
"""
df4 = run_query(query4, SQL_FILE)
# -
df4_grouped = df4.groupby([pd.Grouper(key=DATE_COL, freq='W-MON'), FACTOR_COL]).sum().reset_index(FACTOR_COL)
df4_normed = normalize_dataframe(df4_grouped)
# +
fig, ax = setup_plot(
title="Normalized Collisions Rates During the COVID-19 Pandemic\nTrucks Only",
ylabel="Normalized Collision Rate",
xlabel="",
)
fig, ax = make_ford_vs_toyota_plot(
df4_normed, fig, ax,
toyota_label_xy=("2020-08-01", 0.35),
ford_label_xy=("2020-08-01", 1.0),
stay_at_home_y=1.3,
norm_label_y=0.5,
y_lim=(0, 1.5),
)
for ext in ("svg", "png"):
file_name = f"/tmp/covid_pandemic_normalized_ford_vs_toyota_collisions_trucks.{ext}"
save_plot(fig, file_name)
# -
# ## Is this a county thing?
# +
query2 = f"""
SELECT collision_date
, {FACTOR_COL}
, count(1) as total
FROM collisions AS c
LEFT JOIN parties as p
ON p.case_id = c.case_id
WHERE {DATE_COL} IS NOT NULL
AND {DATE_COL} BETWEEN '{START_YEAR}-01-01' AND '{LAST_YEAR}-11-30'
AND p.{FACTOR_COL} in ('{MAKE_1}', '{MAKE_2}')
AND county_location = 'contra costa'
GROUP BY 1, 2
ORDER BY 1, total DESC
"""
df2 = run_query(query2, SQL_FILE)
# -
df2_grouped = df2.groupby([pd.Grouper(key=DATE_COL, freq='W-MON'), FACTOR_COL]).sum().reset_index(FACTOR_COL)
df2_normed = normalize_dataframe(df2_grouped)
# + tags=[]
fig, ax = setup_plot(
title="Normalized Collisions Rates During the COVID-19 Pandemic\nContra Costa County",
ylabel="Normalized Collision Rate",
xlabel="",
)
fig, ax = make_ford_vs_toyota_plot(
df2_normed, fig, ax,
toyota_label_xy=("2020-08-01", 0.12),
ford_label_xy=("2020-08-01", 1.0),
stay_at_home_y=1.3,
norm_label_y=0.5,
y_lim=(0, 1.5),
)
for ext in ("svg", "png"):
file_name = f"/tmp/covid_pandemic_normalized_ford_vs_toyota_collisions_contra_costa.{ext}"
save_plot(fig, file_name)
# -
# ## Is this an Age thing?
# +
query3 = f"""
SELECT collision_date
, {FACTOR_COL}
, party_age
FROM collisions AS c
LEFT JOIN parties as p
ON p.case_id = c.case_id
WHERE {DATE_COL} IS NOT NULL
AND {DATE_COL} BETWEEN '{START_YEAR}-01-01' AND '{LAST_YEAR}-11-30'
AND p.{FACTOR_COL} in ('{MAKE_1}', '{MAKE_2}')
"""
df3 = run_query(query3, SQL_FILE)
# +
fig, ax = setup_plot(
title="Age Distribution",
ylabel="Probability",
xlabel="",
)
sns.histplot(data=df3, x="party_age", hue=FACTOR_COL, stat="probability", common_norm=False, multiple="layer", binwidth=5, binrange=(16, 100), linewidth=2, zorder=1)
plt.tick_params(labelleft = False)
ax.text(x=72, y=0.035, s="Toyota", color=orange_line_color, fontsize=30)
ax.text(x=45, y=0.095, s="Ford", color=blue_line_color, fontsize=30)
ax.get_legend().remove()
# Remove date label
ax.xaxis.label.set_visible(False)
for ext in ("svg", "png"):
file_name = f"/tmp/covid_pandemic_ford_vs_toyota_collisions_age_distribution.{ext}"
save_plot(fig, file_name)
# +
query5 = f"""
SELECT collision_date
, {FACTOR_COL}
, count(1) as total
FROM collisions AS c
LEFT JOIN parties as p
ON p.case_id = c.case_id
WHERE {DATE_COL} IS NOT NULL
AND {DATE_COL} BETWEEN '{START_YEAR}-01-01' AND '{LAST_YEAR}-11-30'
AND p.{FACTOR_COL} in ('{MAKE_1}', '{MAKE_2}')
AND p.party_age BETWEEN 30 AND 50
GROUP BY 1, 2
ORDER BY 1, total DESC
"""
df5 = run_query(query5, SQL_FILE)
# -
df5_grouped = df5.groupby([pd.Grouper(key=DATE_COL, freq='W-MON'), FACTOR_COL]).sum().reset_index(FACTOR_COL)
df5_normed = normalize_dataframe(df5_grouped)
# + tags=[]
fig, ax = setup_plot(
title="Normalized Collisions Rates During the COVID-19 Pandemic\nDrivers Aged 30 to 50",
ylabel="Normalized Collision Rate",
xlabel="",
)
fig, ax = make_ford_vs_toyota_plot(
df5_normed, fig, ax,
toyota_label_xy=("2020-08-01", 0.12),
ford_label_xy=("2020-08-01", 1.0),
stay_at_home_y=1.3,
norm_label_y=0.55,
y_lim=(0, 1.5),
)
for ext in ("svg", "png"):
file_name = f"/tmp/covid_pandemic_normalized_ford_vs_toyota_collisions_age_30_to_50.{ext}"
save_plot(fig, file_name)
|
files/switrs-covid/SWITRS Ford vs Toyota During Lockdown.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression, Lasso, LogisticRegression
from sklearn.ensemble import GradientBoostingRegressor, GradientBoostingClassifier
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from sklearn.preprocessing import PolynomialFeatures, StandardScaler
from sklearn.pipeline import Pipeline
import scipy.special
# # NLSYM DATA
# +
# Preprocess data
df = pd.read_csv("data/card.csv")
data_filter = df['educ'].values >= 6
T = df['educ'].values[data_filter]
Z = df['nearc4'].values[data_filter]
y = df['lwage'].values[data_filter]
# Impute missing values with mean, add dummy columns
# I excluded the columns 'weights' as we don't know what it is
X_df = df[['exper', 'expersq']].copy()
X_df['fatheduc'] = df['fatheduc'].fillna(value=df['fatheduc'].mean())
X_df['fatheduc_nan'] = df['fatheduc'].isnull()*1
X_df['motheduc'] = df['motheduc'].fillna(value=df['motheduc'].mean())
X_df['motheduc_nan'] = df['motheduc'].isnull()*1
X_df[['momdad14', 'sinmom14', 'reg661', 'reg662',
'reg663', 'reg664', 'reg665', 'reg666', 'reg667', 'reg668', 'reg669', 'south66']] = df[['momdad14', 'sinmom14',
'reg661', 'reg662','reg663', 'reg664', 'reg665', 'reg666', 'reg667', 'reg668', 'reg669', 'south66']]
X_df[['black', 'smsa', 'south', 'smsa66']] = df[['black', 'smsa', 'south', 'smsa66']]
columns_to_scale = ['fatheduc', 'motheduc', 'exper', 'expersq']
scaler = StandardScaler()
X_df[columns_to_scale] = scaler.fit_transform(X_df[columns_to_scale])
X = X_df.values[data_filter]
true_fn = lambda x: np.zeros(x.shape[0])
# -
X_df.columns
# # ANALYSIS
# ### Defining some hyperparameters
random_seed = 123459 # random seed for each experiment
N_SPLITS = 10 # number of splits for cross-fitting
COV_CLIP = 20/(X.shape[0]) # covariance clipping in driv
print(COV_CLIP)
# ### Defining some generic non-parametric regressors and classifiers
# +
from utilities import RegWrapper
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LassoCV, LogisticRegressionCV
from xgboost import XGBClassifier, XGBRegressor
from xgb_utilities import XGBWrapper
# XGB forest models for Regression and Classification
model = lambda: XGBWrapper(XGBRegressor(gamma=0.001, n_estimators=100, min_child_weight=50, n_jobs=10),
early_stopping_rounds=5, eval_metric='rmse', binary=False)
model_clf = lambda: RegWrapper(XGBWrapper(XGBClassifier(gamma=0.001, n_estimators=100, min_child_weight=50, n_jobs=10),
early_stopping_rounds=5, eval_metric='logloss', binary=True))
# -
# ### Some utility functions
def nuisance_diagnostic(cate, nuisance_model, property_name, property_fn,
index_names=None, statistic=np.std, threshold=None):
std = statistic([property_fn(ns) for ns in cate.fitted_nuisances[nuisance_model]], axis=0)
if hasattr(std, '__len__'):
if threshold is None:
coefs = np.argmax(std).flatten()
else:
coefs = np.argwhere(std >= threshold).flatten()
if index_names is None:
index_names = np.arange(std.shape[0])
for high_var in coefs:
plt.figure(figsize=(4,3))
plt.title("{}: {}[{}] Across Folds".format(nuisance_model, property_name, index_names[high_var]))
plt.plot([property_fn(ns)[high_var] for ns in cate.fitted_nuisances[nuisance_model]])
plt.xlabel('fold')
plt.ylabel('property')
plt.show()
else:
plt.figure(figsize=(4,3))
plt.title("{}: {} Across Folds".format(nuisance_model, property_name))
plt.plot([property_fn(ns) for ns in cate.fitted_nuisances[nuisance_model]])
plt.xlabel('fold')
plt.ylabel('property')
plt.show()
# # ATE via DMLATEIV
# +
from dml_ate_iv import DMLATEIV
np.random.seed(random_seed)
# We need to specify models to be used for each of these residualizations
model_Y_X = lambda: model() # model for E[Y | X]
model_T_X = lambda: model() # model for E[T | X]. We use a regressor since T is continuous
model_Z_X = lambda: model_clf() # model for E[Z | X]. We use a classifier since Z is binary
dmlate = DMLATEIV(model_Y_X(), model_T_X(), model_Z_X(),
n_splits=N_SPLITS, # n_splits determines the number of splits to be used for cross-fitting.
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)
# -
# We fit DMLATEIV with these models
dmlate.fit(y, T, X, Z)
# We call effect() to get the ATE
ta_effect = dmlate.effect()
# Comparison with true ATE
print("ATE Estimate: {:.3f}".format(ta_effect))
print("Standard error: {:.3f}".format(dmlate.std))
# We can call normal_effect_interval to get confidence intervals based
# based on the asympotic normal approximation
lower, upper = dmlate.normal_effect_interval(lower=2.5, upper=97.5)
# Comparison with true ATE
print("ATE Estimate Interval: ({:.3f}, {:.3f})".format(lower, upper))
# # ATE and CATE via DMLIV
# +
from dml_iv import DMLIV
from utilities import SelectiveLasso, SeparateModel
from sklearn.linear_model import LassoCV, LogisticRegressionCV
from econml.utilities import hstack
np.random.seed(random_seed)
# For DMLIV we also need a model for E[T | X, Z]. To allow for heterogeneity in the compliance, i.e.
# T = beta(X)*Z + gamma(X)
# we train a separate model for Z=1 and Z=0. The model for Z=1 learns the
# quantity beta(X) + gamma(X) and the model for Z=0 learns gamma(X).
model_T_XZ = lambda: SeparateModel(model(), model())
# We now specify the features to be used for heterogeneity. We will fit a CATE model of the form
# theta(X) = <theta, phi(X)>
# for some set of features phi(X). The featurizer needs to support fit_transform, that takes
# X and returns phi(X). We need to include a bias if we also want a constant term.
dmliv_featurizer = lambda: PolynomialFeatures(degree=1, include_bias=True)
# Then we need to specify a model to be used for fitting the parameters theta in the linear form.
# This model will minimize the square loss:
# (Y - E[Y|X] - <theta, phi(X)> * (E[T|X,Z] - E[T|X]))**2
dmliv_model_effect = lambda: LinearRegression(fit_intercept=False)
# Potentially with some regularization on theta. Here we use an ell_1 penalty on theta
# If we also have a prior that there is no effect heterogeneity we can use a selective lasso
# that does not penalize the constant term in the CATE model
#dmliv_model_effect = lambda: SelectiveLasso(np.arange(1, X.shape[1]+1), LassoCV(cv=5, fit_intercept=False))
# We initialize DMLIV with all these models and call fit
cate = DMLIV(model_Y_X(), model_T_X(), model_T_XZ(),
dmliv_model_effect(), dmliv_featurizer(),
n_splits=N_SPLITS, # number of splits to use for cross-fitting
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)
# -
cate.fit(y, T, X, Z)
# To get the CATE at every X we call effect(X)
dml_effect = cate.effect(X)
plt.hist(dml_effect, label='est')
plt.legend()
plt.show()
# To get the parameter theta we call coef_. The first entry is the intercept of the CATE model
print(cate.coef_)
try:
print(cate.effect_model.lasso_model.alpha_)
plt.plot(cate.effect_model.lasso_model.alphas_, cate.effect_model.lasso_model.mse_path_)
plt.show()
except:
print("NA")
# We can average the CATE to get an ATE
print("ATE Estimate: {:.3f}".format(np.mean(dml_effect)))
# We can also see how it compares to the true CATE at each target point and calculate MSE
plt.title("DMLIV CATE as Function of {}".format(X_df.columns[np.argmax(np.abs(cate.coef_[1:]))]))
plt.scatter(X[:, np.argmax(np.abs(cate.coef_[1:]))], dml_effect, label='est')
plt.scatter(X[:, np.argmax(np.abs(cate.coef_[1:]))], true_fn(X), label='true', alpha=.2)
plt.legend()
plt.show()
# #### Some Diagnostics of the Fitted Nuisance Models Across Folds
# The property .fitted_nuisances is a dictionary of the form:
# {'name_of_nuisance': [fitted_model_fold_1, fitted_model_fold_2, ...]}
# then we can access all properties of each of the fitted models for each fold.
# If for instance all nuisances have a linear form we can look at the standard deviation
# of the coefficients of each of the nuisance model across folds to check for stability
try:
nuisance_diagnostic(cate, 'model_T_XZ', 'coef', lambda ns: ns.coef_.flatten(),
[c+"_0" for c in X_df.columns] + [c+"_1" for c in X_df.columns])
nuisance_diagnostic(cate, 'model_T_X', 'coef', lambda ns: ns.coef_.flatten(), X_df.columns)
nuisance_diagnostic(cate, 'model_Y_X', 'coef', lambda ns: ns.coef_.flatten(), X_df.columns)
except:
print("Unavailable")
try:
for i in range(N_SPLITS):
plt.plot([c0 - c1 for c0, c1 in zip(cate.fitted_nuisances['model_T_XZ'][i].model0.coef_.flatten(),
cate.fitted_nuisances['model_T_XZ'][i].model1.coef_.flatten())])
plt.title("Difference in coefficients betwen model0 and model1")
plt.show()
except:
print("Unavailable")
# # ATE via DRIV
# +
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from utilities import SubsetWrapper, StatsModelLinearRegression, ConstantModel
from sklearn.dummy import DummyRegressor
np.random.seed(random_seed)
# For DRIV we need a model for predicting E[T*Z | X]. We use a classifier
model_TZ_X = lambda: model()
# We also need a model for the final regression that will fit the function theta(X)
# If we want to fit an ATE, we simply fit a constant functin theta(X) = theta
# We can do this with a pipeline where the preprocessing step only creates a bias column
# and the regression step fits a linear regression with no intercept.
# To get normal confidence intervals easily we can use a statsmodels linear regression
# wrapped in an sklearn interface
const_driv_model_effect = lambda: ConstantModel()
# As in OrthoDMLIV we need a perliminary estimator of the CATE.
# We use a DMLIV estimator with no cross-fitting (n_splits=1)
dmliv_prel_model_effect = DMLIV(model_Y_X(), model_T_X(), model_T_XZ(),
dmliv_model_effect(), dmliv_featurizer(),
n_splits=1, binary_instrument=True, binary_treatment=True)
const_dr_cate = DRIV(model_Y_X(), model_T_X(), model_Z_X(), # same as in DMLATEIV
dmliv_prel_model_effect, # preliminary model for CATE, must support fit(y, T, X, Z) and effect(X)
model_TZ_X(), # model for E[T * Z | X]
const_driv_model_effect(), # model for final stage of fitting theta(X)
cov_clip=COV_CLIP, # covariance clipping to avoid large values in final regression from weak instruments
n_splits=N_SPLITS, # number of splits to use for cross-fitting
binary_instrument=True, # a flag whether to stratify cross-fitting by instrument
binary_treatment=True # a flag whether to stratify cross-fitting by treatment
)
# -
const_dr_cate.fit(y, T, X, Z, store_final=True)
# To get the statsmodel summary we look at the effect_model, which is the pipeline, we then look
# at the reg step of the pipeline which is the statsmodel wrapper and then we look
# at the model attribute of the statsmodel wrapper and print the summary()
const_dr_cate.effect_model.summary()
# #### Some diagnostics of the fitted nuisance models across folds
# The property .fitted_nuisances is a dictionary of the form:
# {'name_of_nuisance': [fitted_model_fold_1, fitted_model_fold_2, ...]}
# then we can access all properties of each of the fitted models for each fold.
# If for instance all nuisances have a linear form we can look at the standard deviation
# of the coefficients of each of the nuisance model across folds to check for stability
try:
[print("Model {} max std of coefs: {}".format(name,
np.max(np.std([ns.coef_ for ns in nuisance_insts], axis=0))))
if hasattr(nuisance_insts[0], 'coef_') else None
for name, nuisance_insts in const_dr_cate.fitted_nuisances.items()]
except:
print("Unavailable")
try:
nuisance_diagnostic(const_dr_cate, 'model_TZ_X', 'coef', lambda ns: ns.coef_.flatten(), X_df.columns)
nuisance_diagnostic(const_dr_cate, 'model_TZ_X', 'predict', lambda ns: ns.predict(X), np.arange(X.shape[0]))
nuisance_diagnostic(const_dr_cate, 'model_T_X', 'coef', lambda ns: ns.coef_.flatten(), X_df.columns)
nuisance_diagnostic(const_dr_cate, 'prel_model_effect', 'effect', lambda ns: ns.effect(X), np.arange(X.shape[0]))
except:
print("Unavailable")
# ## Projecting CATE to a pre-chosen subset of variables in final model
# +
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from utilities import SubsetWrapper
np.random.seed(random_seed)
# We could also fit a projection on a subset of the features by using the
# subset wrapper from our utilities.
# Example: including everything for expository purposes, but any array-like of indices would work
subset_names = set(['motheduc'])
# list of indices of features X to use in the final model
feature_inds = np.argwhere([(x in subset_names) for x in X_df.columns.values]).flatten()
print(feature_inds)
# Because we are projecting to a low dimensional model space, we can
# do valid inference and we can use statsmodel linear regression to get all
# the hypothesis testing capability
proj_driv_model_effect = lambda: SubsetWrapper(StatsModelLinearRegression(),
feature_inds # list of indices of features X to use in the final model
)
# -
proj_dr_cate = const_dr_cate.refit_final(proj_driv_model_effect())
# To get the CATE at every X we call effect(X[:, feature_inds])
proj_dr_effect = proj_dr_cate.effect(X[:, feature_inds])
# To get the statsmodel summary we look at the effect_model, which is
# an instance of SubsetWrapper, we look at the model of the SubsetWrapper which is
# and instance of the pipeline, we then look at the reg step of the pipeline which is the statsmodel wrapper and
# call summary() of the wrapper (most prob there is a better API for this, but we can go with this for now :)
proj_dr_cate.effect_model.summary(alpha=.05, xname=['const']+list(X_df.columns[feature_inds]))
# +
# We can also evaluate coverage and create prediction intervals using statsmodels attributes
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
res = proj_dr_cate.effect_model.model
predictions = res.get_prediction(PolynomialFeatures(degree=1, include_bias=True).fit_transform(X[:, feature_inds]))
frame = predictions.summary_frame(alpha=0.05)
pred = frame['mean']
iv_l = frame['mean_ci_lower']
iv_u = frame['mean_ci_upper']
fig, ax = plt.subplots(figsize=(3,4))
order = np.argsort(X[:, feature_inds[0]])
ax.plot(X[order, feature_inds[0]], iv_u[order], 'C3--', label="Upper 95% CI")
ax.plot(X[order, feature_inds[0]], iv_l[order], 'C3--', label="Lower 95% CI")
ax.plot(X[order, feature_inds[0]], pred[order], 'C0--.', label="Prediction")
ax.legend(loc='best')
plt.xlabel("Mother's Education (scaled)")
plt.ylabel("Treatment Effect")
#plt.savefig("NLSYM_momeduc_linear_projection_2.pdf", dpi=300, bbox_inches='tight')
plt.show()
# -
# ## Random Forest Based CATE and Tree Explainer
# +
from dml_iv import DMLIV
from dr_iv import DRIV, ProjectedDRIV
from utilities import SubsetWrapper
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
np.random.seed(random_seed)
rf_driv_model_effect = lambda: RandomForestRegressor(n_estimators=5000, max_depth=2, min_impurity_decrease=0.01,
min_samples_leaf=100, bootstrap=True)
# -
rf_dr_cate = const_dr_cate.refit_final(rf_driv_model_effect())
# +
import seaborn as sns
rf_dr_effect = rf_dr_cate.effect(X)
plt.figure(figsize=(3,4))
sns.distplot(rf_dr_effect)
plt.xlabel("Treatment Effect")
plt.ylabel("Frequency")
plt.savefig("NLSYM_hte_distribution_2.pdf", dpi=300, bbox_inches='tight')
plt.show()
# +
import shap
import pandas as pd
Xdf = pd.DataFrame(X, columns=X_df.columns)
# explain the model's predictions using SHAP values
explainer = shap.TreeExplainer(rf_dr_cate.effect_model)
shap_values = explainer.shap_values(Xdf)
# visualize the first prediction's explanation (use matplotlib=True to avoid Javascript)
shap.force_plot(explainer.expected_value, shap_values[0,:], Xdf.iloc[0,:], matplotlib=True)
# -
plt.figure(figsize=(4,4))
shap.summary_plot(shap_values, Xdf, plot_type="violin", max_display=10, show=False, auto_size_plot=False)
plt.show()
plt.figure(figsize=(5,4))
shap.summary_plot(shap_values, Xdf, plot_type='bar', max_display=10, show=False, auto_size_plot=False)
plt.xlabel("mean(|SHAP value|)\n average impact on model output magnitude")
plt.savefig("NLSYM_shap_summary_bar.pdf", dpi=300, bbox_inches='tight')
plt.show()
|
prototypes/dml_iv/NLSYM_GBM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computer Vision
#
# ## Manipulating Images
#
#
# ### Load an Image
#
# +
# %matplotlib inline
from matplotlib.pyplot import imshow
from PIL import Image
import numpy as np
import skimage.color as sc
# !curl https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/graeme2.jpg -o img.jpg
i = np.array(Image.open('img.jpg'))
imshow(i)
# -
# ### Examine Numerical Properties of the Image
#
type(i)
i.dtype
i.shape
i_mono = sc.rgb2gray(i)
imshow(i_mono, cmap='gray')
i_mono.shape
# ### View Pixel Value Distributions
# Plot a histogram
# +
def im_hist(img):
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8, 6))
fig.clf()
ax = fig.gca()
ax.hist(img.flatten(), bins = 256)
plt.show()
im_hist(i_mono)
# -
# Plot a cumulative histogram
# +
def im_cdf(img):
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(8, 6))
fig.clf()
ax = fig.gca()
ax.hist(img.flatten(), bins = 256, cumulative=True)
plt.show()
im_cdf(i_mono)
# -
# ### Equalize the Image
# Use skimage library to equalize the image.
# +
from skimage import exposure
i_eq = exposure.equalize_hist(i_mono)
imshow(i_eq, cmap='gray')
# -
# View the histogram and CDF plots:
im_hist(i_eq)
im_cdf(i_eq)
# ## Denoising with Filters
#
# ### Add Noise
import skimage
i_n = skimage.util.random_noise(i_eq)
imshow(i_n, cmap="gray")
# ### Use a Gaussian Filter
def gauss_filter(im, sigma = 10):
from scipy.ndimage.filters import gaussian_filter as gf
import numpy as np
return gf(im, sigma = sigma)
i_g = gauss_filter(i_n)
imshow(i_g, cmap="gray")
# ### Use a Median Filter
#
def med_filter(im, size = 10):
from scipy.ndimage.filters import median_filter as mf
import numpy as np
return mf(im, size = size)
i_m = med_filter(i_n)
imshow(i_m, cmap="gray")
#
# ### Extract Features
#
# #### Sobel Edge Detection
#
#
# +
def edge_sobel(image):
from scipy import ndimage
import skimage.color as sc
import numpy as np
image = sc.rgb2gray(image) # Convert color image to gray scale
dx = ndimage.sobel(image, 1) # horizontal derivative
dy = ndimage.sobel(image, 0) # vertical derivative
mag = np.hypot(dx, dy) # magnitude
mag *= 255.0 / np.amax(mag) # normalize (Q&D)
mag = mag.astype(np.uint8)
return mag
i_edge = edge_sobel(i_m)
imshow(i_edge, cmap="gray")
# -
# #### Harris Corner Detection
#
# +
def corner_harr(im, min_distance = 10):
from skimage.feature import corner_harris, corner_peaks
mag = corner_harris(im)
return corner_peaks(mag, min_distance = min_distance)
harris = corner_harr(i_eq, 10)
def plot_harris(im, harris, markersize = 20, color = 'red'):
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure(figsize=(6, 6))
fig.clf()
ax = fig.gca()
ax.imshow(np.array(im).astype(float), cmap="gray")
ax.plot(harris[:, 1], harris[:, 0], 'r+', color = color, markersize=markersize)
return 'Done'
plot_harris(i_eq, harris)
# -
# The corner detection algorithm has identified the eyes in the image.
# ## Image Classification
#
# ### Custom Vision API
# https://www.customvision.ai/projects
# Install the Custom Vision SDK
# ! pip install azure-cognitiveservices-vision-customvision
PREDICTION_KEY = 'YOUR_PREDICTION_KEY'
ENDPOINT='https://YOUR_REGION.api.cognitive.microsoft.com'
PROJECT_ID = 'YOUR_PROJECT_ID'
# +
from azure.cognitiveservices.vision.customvision.prediction import CustomVisionPredictionClient
import matplotlib.pyplot as plt
from PIL import Image
import requests
from io import BytesIO
# %matplotlib inline
# Use two test images
test_img1_url = 'http://www.pachd.com/free-images/food-images/apple-01.jpg'
test_img2_url = 'http://www.pachd.com/free-images/food-images/carrot-02.jpg'
# Create an instance of the prediction service
predictor = CustomVisionPredictionClient(PREDICTION_KEY, endpoint=ENDPOINT)
# Get a prediction for image 1
result1 = predictor.predict_image_url(PROJECT_ID, url=test_img1_url)
# The results include a prediction for each tag, in descending order of probability - so we'll get the first one
prediction1 = result1.predictions[0].tag_name + ": {0:.2f}%".format(result1.predictions[0].probability * 100)
# Get a prediction for image 2
result2 = predictor.predict_image_url(PROJECT_ID, url=test_img2_url)
prediction2 = result2.predictions[0].tag_name + ": {0:.2f}%".format(result2.predictions[0].probability * 100)
# Download the images so we can show them
response = requests.get(test_img1_url)
img1 = Image.open(BytesIO(response.content))
response = requests.get(test_img2_url)
img2 = Image.open(BytesIO(response.content))
# Create a figure
fig = plt.figure(figsize=(16, 8))
# Subplot for first image and its predicted class
a=fig.add_subplot(1,2,1)
imgplot = plt.imshow(img1)
a.set_title(prediction1)
# Subplot for second image and its predicted class
a=fig.add_subplot(1,2,2)
imgplot = plt.imshow(img2)
a.set_title(prediction2)
plt.show()
# -
# ## Use the Computer Vision API
# https://portal.azure.com.
#
#
visionURI = 'YOUR_REGION.api.cognitive.microsoft.com'
visionKey = 'YOUR_KEY'
# ### Get An Image from a URL
#
# +
# %matplotlib inline
from matplotlib.pyplot import imshow
from PIL import Image
import requests
from io import BytesIO
img_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/graeme2.jpg'
# Get the image and show it
response = requests.get(img_url)
img = Image.open(BytesIO(response.content))
imshow(img)
# -
# ### Use the Computer Vision API to Get Image Features
#
# +
def get_image_features(img_url):
import http.client, urllib.request, urllib.parse, urllib.error, base64, json
headers = {
# Request headers.
'Content-Type': 'application/json',
'Ocp-Apim-Subscription-Key': visionKey,
}
params = urllib.parse.urlencode({
# Request parameters. All of them are optional.
'visualFeatures': 'Categories,Description,Color',
'language': 'en',
})
body = "{'url':'" + img_url + "'}"
try:
# Execute the REST API call and get the response.
conn = http.client.HTTPSConnection(visionURI)
conn.request("POST", "/vision/v1.0/analyze?%s" % params, body, headers)
response = conn.getresponse()
data = response.read().decode("UTF-8")
# 'data' contains the JSON response.
parsed = json.loads(data)
if response is not None:
return parsed
conn.close()
except Exception as e:
print('Error:')
print(e)
jsonData = get_image_features(img_url)
desc = jsonData['description']['captions'][0]['text']
print(desc)
# -
# Get the full response
# View the full details returned
import http.client, urllib.request, urllib.parse, urllib.error, base64, json
print (json.dumps(jsonData, sort_keys=True, indent=2))
# Let's try with a different image:
# +
img_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/soccer.jpg'
# Get the image and show it
response = requests.get(img_url)
img = Image.open(BytesIO(response.content))
imshow(img)
jsonData = get_image_features(img_url)
desc = jsonData['description']['captions'][0]['text']
print(desc)
# -
# ## Using the Face API
#
#
# ### Create a Face API Service
# https://portal.azure.com.
#
faceURI = "https://YOUR_REGION.api.cognitive.microsoft.com/face/v1.0/"
faceKey = "YOUR_KEY"
# Install the Face SDK package. This makes it easier to work with.
#
# !pip install cognitive_face
# !pip install pillow
# Detect a face in an image:
# +
# %matplotlib inline
import requests
from io import BytesIO
from matplotlib.pyplot import imshow
from PIL import Image, ImageDraw
import cognitive_face as CF
# Set URI and Key
CF.Key.set(faceKey)
CF.BaseUrl.set(faceURI)
# Detect faces in an image
img_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/graeme1.jpg'
result = CF.face.detect(img_url)
# Get the ID of the first face detected
face1 = result[0]['faceId']
print ("Face 1:" + face1)
# Get the image
response = requests.get(img_url)
img = Image.open(BytesIO(response.content))
# Add rectangles for each face found
color="blue"
if result is not None:
draw = ImageDraw.Draw(img)
for currFace in result:
faceRectangle = currFace['faceRectangle']
left = faceRectangle['left']
top = faceRectangle['top']
width = faceRectangle['width']
height = faceRectangle['height']
draw.line([(left,top),(left+width,top)],fill=color, width=5)
draw.line([(left+width,top),(left+width,top+height)],fill=color , width=5)
draw.line([(left+width,top+height),(left, top+height)],fill=color , width=5)
draw.line([(left,top+height),(left, top)],fill=color , width=5)
# show the image
imshow(img)
# -
# Compare with another image
# +
# Get the image to compare
img2_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/graeme2.jpg'
response2 = requests.get(img2_url)
img2 = Image.open(BytesIO(response2.content))
# Detect faces in a comparison image
result2 = CF.face.detect(img2_url)
# Assume the first face is the one we want to compare
face2 = result2[0]['faceId']
print ("Face 2:" + face2)
def verify_face(face1, face2):
# By default, assume the match is unverified
verified = "Not Verified"
color="red"
if result2 is not None:
# compare the comparison face to the original one we retrieved previously
verify = CF.face.verify(face1, face2)
# if there's a match, set verified and change color to green
if verify['isIdentical'] == True:
verified = "Verified"
color="lightgreen"
# Display the second face with a red rectange if unverified, or green if verified
draw = ImageDraw.Draw(img2)
for currFace in result2:
faceRectangle = currFace['faceRectangle']
left = faceRectangle['left']
top = faceRectangle['top']
width = faceRectangle['width']
height = faceRectangle['height']
draw.line([(left,top),(left+width,top)] , fill=color, width=5)
draw.line([(left+width,top),(left+width,top+height)] , fill=color, width=5)
draw.line([(left+width,top+height),(left, top+height)] , fill=color, width=5)
draw.line([(left,top+height),(left, top)] , fill=color, width=5)
# show the image
imshow(img2)
# Display verification status and confidence level
print(verified)
print ("Confidence Level: " + str(verify['confidence']))
verify_face(face1, face2)
# -
# And another?
# +
# Get the image to compare
img2_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/graeme3.jpg'
response2 = requests.get(img2_url)
img2 = Image.open(BytesIO(response2.content))
# Detect faces in a comparison image
result2 = CF.face.detect(img2_url)
# Assume the first face is the one we want to compare
face2 = result2[0]['faceId']
print ("Face 2:" + face2)
verify_face(face1, face2)
# -
# And another?
# +
# Get the image to compare
img2_url = 'https://raw.githubusercontent.com/MicrosoftLearning/AI-Introduction/master/files/satya.jpg'
response2 = requests.get(img2_url)
img2 = Image.open(BytesIO(response2.content))
# Detect faces in a comparison image
result2 = CF.face.detect(img2_url)
# Assume the first face is the one we want to compare
face2 = result2[0]['faceId']
print ("Face 2:" + face2)
verify_face(face1, face2)
# -
# No match!
|
Artificial-Intelligence (AI)/Introduction to Artificial Intelligence (AI)/DAT263x-demos/Vision/Images - Demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] heading_collapsed=true
# # Imports
# + code_folding=[] hidden=true
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import (accuracy_score,
make_scorer,
classification_report,
roc_curve,
roc_auc_score)
from sklearn.preprocessing import StandardScaler
from sklearn.dummy import DummyClassifier, DummyRegressor
from sklearn.model_selection import (cross_val_score,
train_test_split,
GridSearchCV,
KFold,
StratifiedKFold,
learning_curve,
validation_curve)
from sklearn.pipeline import make_pipeline
# %matplotlib inline
plt.style.use('fivethirtyeight')
sns.set_context('notebook')
# -
# # Data
data = datasets.load_breast_cancer()
X = data['data']
y = data['target']
df = pd.DataFrame(np.concatenate([X, y[:, np.newaxis]], axis=1), columns=list(
data['feature_names']) + ['label'])
df.head()
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=1, test_size=0.3, stratify=y)
X_train.shape, y_train.shape, X_test.shape, y_test.shape
log_model = make_pipeline(StandardScaler(), LogisticRegression())
print(
f'3-fold CV accuracy : {cross_val_score(log_model, X_train, y_train, cv=3).mean():.2%}')
log_model.fit(X_train, y_train)
print(f'Test accuracy : {accuracy_score(y_test, log_model.predict(X_test)):.2%}')
# # DummyModel - Baseline model
# - DummyClassifier: strategies can be: stratified, uniform, most_frequent, constant
# - DummyRegressor: strategies can be: mean, constant median, quantile
# Create dummy classifier
baseline_clf = DummyClassifier(strategy='stratified')
baseline_clf.fit(X_train, y_train)
accuracy_score(y_test, baseline_clf.predict(X_test))
# # Cross Validation
# We can either use cv=int or specify strategy to generate the folds such as KFold object or generator
kf = KFold(n_splits=5, shuffle=True, random_state=1)
cross_val_score(log_model, X_train, y_train, n_jobs=-1, scoring='accuracy', cv=kf).mean()
cross_val_score(log_model, X_train, y_train, n_jobs=-1, scoring='accuracy', cv=5).mean()
# # Make scorer
# We can use custom scoring function and then wrap it up using `make_scorer()` from sklearn.metrics to provide for GridSearchCV() or cross_val_score
# +
#make_scorer()
# -
# # Classification report
print(classification_report(y_test, log_model.predict(X_test), target_names=['no', 'yes']))
# # Nested CV
# It's better to use to get an unbiased CV metric by having an inner CV such as GridSearchCV to select the best hyperparameters and then use outer CV such as cross_val_score to get the model's performance using the best hyperparameters we got from GridSearchCV
# # Learning Curves
# +
kf = StratifiedKFold(n_splits=10, shuffle=True, random_state=1)
train_sizes, train_scores, test_scores = learning_curve(LogisticRegression(),
X_train, y_train,
scoring='accuracy',
cv=kf,
n_jobs=-1)
train_mean = 1 - train_scores.mean(axis=1)
test_mean = 1 - test_scores.mean(axis=1)
train_std = train_scores.std(axis=1)
test_std = test_scores.std(axis=1)
# -
plt.plot(train_sizes, train_mean, label='Training score')
plt.plot(train_sizes, test_mean, label='Test score')
plt.fill_between(train_sizes, train_mean - train_std, train_mean + train_std, color='#DDDDDD')
plt.fill_between(train_sizes, test_mean - test_std, test_mean + test_std, color='#FDDDDD')
plt.legend()
# # ROC
fpr, tpr, _ = roc_curve(y_test, log_model.predict(X_test))
plt.plot(fpr, tpr)
plt.plot([0,1], ls='--');
print(f'AUC : {roc_auc_score(y_test, log_model.predict_proba(X_test)[:, 1]):.2%}')
# # Validation Curve
# +
train_scores, test_scores = validation_curve(LogisticRegression(),
X_train, y_train,
param_name='C',
param_range=np.logspace(-4, 4, 10),
scoring='accuracy',
cv=kf,
n_jobs=-1)
train_mean = 1 - train_scores.mean(axis=1)
test_mean = 1 - test_scores.mean(axis=1)
train_std = train_scores.std(axis=1)
test_std = test_scores.std(axis=1)
# -
plt.plot(np.logspace(-4, 4, 10), train_mean, label='Training score')
plt.plot(np.logspace(-4, 4, 10), test_mean, label='Test score')
# plt.fill_between(np.logspace(-4, 4, 100), train_mean - train_std, train_mean + train_std)
# plt.fill_between(np.logspace(-4, 4, 100), test_mean - test_std, test_mean + test_std)
plt.legend();
|
notebooks/Model Evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import numpy as np
import pandas as pd
import scipy.stats as stats
# Disable warnings from printing
from warnings import filterwarnings
filterwarnings('ignore')
# -
# Read the sessions data
sessions_data = pd.read_csv("../Data/sessions.csv")
# Group all null user_ids into "NAN" group
sessions_data[sessions_data.user_id.isnull()].user_id = "NAN"
# Get unique user ids to store results
sessions_data_ids = sessions_data.loc[:, ["user_id"]].drop_duplicates().reset_index().drop("index", axis=1)
# Get user ids and actions for action processing
sessions_data_action = sessions_data.loc[:, ["user_id", "action"]]
# +
# Processing for Action attribute
# Replace null values with 'NAN'
sessions_data_action.action = sessions_data_action.action.fillna("NAN")
# Replace infrequent values with 'Other' to avoid overfitting and speed up computation
THRESHOLD = 0.005*sessions_data_action.shape[0]
counts = sessions_data_action.action.value_counts()
sessions_data_action.action = sessions_data_action.action.apply(lambda x: 'Other' if counts[x] < THRESHOLD else x)
# +
# Get counts of each action for each user
unique_actions = sessions_data_action["action"].unique()
for act in unique_actions:
sessions_data_ids = sessions_data_ids.join(
sessions_data_action.loc[sessions_data_action.action == act, :].groupby(['user_id']).size().to_frame(),
on="user_id")
sessions_data_ids.rename(columns={0 : "action_" + act}, inplace=True)
sessions_data_ids = sessions_data_ids.fillna(0)
# -
# Get number of sessions of each user
sessions_data_ids = sessions_data_ids.join(
sessions_data_action.groupby(['user_id'])["user_id"].size().to_frame(),
on="user_id")
sessions_data_ids.rename(columns={0 : "NumSessions"}, inplace=True)
# +
# Get mean and std of distribution of counts of actions for each user
sessions_data_ids["NumActionsMean"] = \
sessions_data_ids.loc[:, "action_lookup":"action_similar_listings_v2"].mean(axis=1)
sessions_data_ids["NumActionsStd"] = \
sessions_data_ids.loc[:, "action_lookup":"action_similar_listings_v2"].std(axis=1)
# -
# Get user ids and action_type for action_type processing
sessions_data_action_type = sessions_data.loc[:, ["user_id", "action_type"]]
# +
# Processing for action_type attribute
# Replace null values with 'NAN'
sessions_data_action_type.action_type = sessions_data_action_type.action_type.fillna("NAN")
sessions_data_action_type.action_type = sessions_data_action_type.action_type.replace(
{
'-unknown-': 'NAN'
}
)
# Replace infrequent values with 'Other' to avoid overfitting and speed up computation
THRESHOLD = 0.005*sessions_data_action_type.shape[0]
counts = sessions_data_action_type.action_type.value_counts()
sessions_data_action_type.action_type = \
sessions_data_action_type.action_type.apply(lambda x: 'Other' if counts[x] < THRESHOLD else x)
# +
# Get counts of each action_type for each user
unique_actions = sessions_data_action_type["action_type"].unique()
for act in unique_actions:
sessions_data_ids = sessions_data_ids.join(
sessions_data_action_type.loc[sessions_data_action_type.action_type == act, :].groupby(
['user_id']).size().to_frame(),
on="user_id")
sessions_data_ids.rename(columns={0 : "action_type_" + act}, inplace=True)
sessions_data_ids = sessions_data_ids.fillna(0)
# +
# Get mean and std of distribution of counts of action_type for each user
sessions_data_ids["NumActionTypeMean"] = \
sessions_data_ids.loc[:, "action_type_NAN":"action_type_Other"].mean(axis=1)
sessions_data_ids["NumActionTypeStd"] = \
sessions_data_ids.loc[:, "action_type_NAN":"action_type_Other"].std(axis=1)
# +
# Repeat procedure for action_detail
# Get user ids and action_detail for action_detail processing
sessions_data_action_detail = sessions_data.loc[:, ["user_id", "action_detail"]]
# Processing for action_detail attribute
# Replace null values with 'NAN'
sessions_data_action_detail.action_detail = sessions_data_action_detail.action_detail.fillna("NAN")
sessions_data_action_detail.action_detail = sessions_data_action_detail.action_detail.replace(
{
'-unknown-': 'NAN'
}
)
# Replace infrequent values with 'Other' to avoid overfitting and speed up computation
THRESHOLD = 0.005*sessions_data_action_detail.shape[0]
counts = sessions_data_action_detail.action_detail.value_counts()
sessions_data_action_detail.action_detail = \
sessions_data_action_detail.action_detail.apply(lambda x: 'Other' if counts[x] < THRESHOLD else x)
# Get counts of each action_type for each user
unique_actions = sessions_data_action_detail["action_detail"].unique()
for act in unique_actions:
sessions_data_ids = sessions_data_ids.join(
sessions_data_action_detail.loc[sessions_data_action_detail.action_detail == act, :].groupby(
['user_id']).size().to_frame(),
on="user_id")
sessions_data_ids.rename(columns={0 : "action_detail_" + act}, inplace=True)
sessions_data_ids = sessions_data_ids.fillna(0)
# +
# Get mean and std of distribution of counts of action_detail for each user
sessions_data_ids["NumActionDetailMean"] = \
sessions_data_ids.loc[:, "action_detail_NAN":"action_detail_listing_reviews"].mean(axis=1)
sessions_data_ids["NumActionDetailStd"] = \
sessions_data_ids.loc[:, "action_detail_NAN":"action_detail_listing_reviews"].std(axis=1)
# +
# Repeat the procedure for device_type
# Get user ids and device_type for device_type processing
sessions_data_device_type = sessions_data.loc[:, ["user_id", "device_type"]]
# Processing for device_type attribute
# Replace null values with 'NAN'
sessions_data_device_type.device_type = sessions_data_device_type.device_type.fillna("NAN")
sessions_data_device_type.device_type = sessions_data_device_type.device_type.replace(
{
'-unknown-': 'NAN'
}
)
# Replace infrequent values with 'Other' to avoid overfitting and speed up computation
THRESHOLD = 0.005*sessions_data_device_type.shape[0]
counts = sessions_data_device_type.device_type.value_counts()
sessions_data_device_type.device_type = \
sessions_data_device_type.device_type.apply(lambda x: 'Other' if counts[x] < THRESHOLD else x)
# Get counts of each action_type for each user
unique_actions = sessions_data_device_type["device_type"].unique()
for act in unique_actions:
sessions_data_ids = sessions_data_ids.join(
sessions_data_device_type.loc[sessions_data_device_type.device_type == act, :].groupby(
['user_id']).size().to_frame(),
on="user_id")
sessions_data_ids.rename(columns={0 : "device_type_" + act}, inplace=True)
sessions_data_ids = sessions_data_ids.fillna(0)
# +
# Get mean and std of distribution of counts of device_type for each user
sessions_data_ids["NumDeviceTypeMean"] = \
sessions_data_ids.loc[:, "device_type_Windows Desktop":"device_type_Tablet"].mean(axis=1)
sessions_data_ids["NumDeviceTypeStd"] = \
sessions_data_ids.loc[:, "device_type_Windows Desktop":"device_type_Tablet"].std(axis=1)
# +
# Get user ids and secs_elapsed for secs_elapsed processing
sessions_data_secs_elapsed = sessions_data.loc[:, ["user_id", "secs_elapsed"]]
sessions_data_secs_elapsed.secs_elapsed = sessions_data_secs_elapsed.secs_elapsed.fillna(0)
# Get simple stats on secs_elapsed
tmp = sessions_data_secs_elapsed.groupby('user_id').aggregate(
[
np.mean, np.std, np.median, stats.skew
]
)
tmp.columns = ["secs_elapsed_mean", "secs_elapsed_std", "secs_elapsed_median", "secs_elapsed_skew"]
sessions_data_ids = sessions_data_ids.join(tmp, on="user_id")
# -
# Save processed Output
sessions_data_ids.to_csv("../Data/sessions_processed.csv", header=True)
|
Source/SessionsDataProcessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
# +
pickle_file = 'SVHN.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
#train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
#print('Training set', train_dataset.shape, train_labels.shape)
#print('Validation set', valid_dataset.shape, valid_labels.shape)
#print('Test set', test_dataset.shape, test_labels.shape)
# +
pickle_file = 'SVHN1.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset1 = save['train_dataset1']
del save # hint to help gc free up memory
pickle_file = 'SVHN2.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset2 = save['train_dataset2']
del save # hint to help gc free up memory
pickle_file = 'SVHN3.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset3 = save['train_dataset3']
del save # hint to help gc free up memory
# -
train_dataset = np.concatenate((train_dataset1, train_dataset2, train_dataset3), axis=0)
del train_dataset1, train_dataset2, train_dataset3
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# +
image_size = 32
num_labels = 10
num_channels = 1 # grayscale
def reformat(dataset, labels):
dataset = dataset.reshape(
(-1, image_size, image_size, num_channels)).astype(np.float32)
labels = labels.astype(np.int32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# +
def LecunLCN(X, image_shape, threshold=1e-4, radius=7, use_divisor=True):
"""Local Contrast Normalization"""
"""[http://yann.lecun.com/exdb/publis/pdf/jarrett-iccv-09.pdf]"""
# Get Gaussian filter
filter_shape = (radius, radius, image_shape[3], 1)
#self.filters = theano.shared(self.gaussian_filter(filter_shape), borrow=True)
filters = gaussian_filter(filter_shape)
X = tf.convert_to_tensor(X, dtype=tf.float32)
# Compute the Guassian weighted average by means of convolution
convout = tf.nn.conv2d(X, filters, [1,1,1,1], 'SAME')
# Subtractive step
mid = int(np.floor(filter_shape[1] / 2.))
# Make filter dimension broadcastable and subtract
centered_X = tf.sub(X, convout)
# Boolean marks whether or not to perform divisive step
if use_divisor:
# Note that the local variances can be computed by using the centered_X
# tensor. If we convolve this with the mean filter, that should give us
# the variance at each point. We simply take the square root to get our
# denominator
# Compute variances
sum_sqr_XX = tf.nn.conv2d(tf.square(centered_X), filters, [1,1,1,1], 'SAME')
# Take square root to get local standard deviation
denom = tf.sqrt(sum_sqr_XX)
per_img_mean = tf.reduce_mean(denom)
divisor = tf.maximum(per_img_mean, denom)
# Divisise step
new_X = tf.truediv(centered_X, tf.maximum(divisor, threshold))
else:
new_X = centered_X
return new_X
def gaussian_filter(kernel_shape):
x = np.zeros(kernel_shape, dtype = float)
mid = np.floor(kernel_shape[0] / 2.)
for kernel_idx in xrange(0, kernel_shape[2]):
for i in xrange(0, kernel_shape[0]):
for j in xrange(0, kernel_shape[1]):
x[i, j, kernel_idx, 0] = gauss(i - mid, j - mid)
return tf.convert_to_tensor(x / np.sum(x), dtype=tf.float32)
def gauss(x, y, sigma=3.0):
Z = 2 * np.pi * sigma ** 2
return 1. / Z * np.exp(-(x ** 2 + y ** 2) / (2. * sigma ** 2))
# -
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == labels)
/ predictions.shape[0])
# +
batch_size = 64
patch_size = 5
depth1 = 16
depth2 = 32
depth3 = 64
num_hidden1 = 64
num_hidden2 = 16
shape=[batch_size, image_size, image_size, num_channels]
# Construct a 7-layer CNN.
# C1: convolutional layer, batch_size x 28 x 28 x 16, convolution size: 5 x 5 x 1 x 16
# S2: sub-sampling layer, batch_size x 14 x 14 x 16
# C3: convolutional layer, batch_size x 10 x 10 x 32, convolution size: 5 x 5 x 16 x 32
# S4: sub-sampling layer, batch_size x 5 x 5 x 32
# C5: convolutional layer, batch_size x 1 x 1 x 64, convolution size: 5 x 5 x 32 x 64
# Dropout
# F6: fully-connected layer, weight size: 64 x 16
# Output layer, weight size: 16 x 10
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.int64, shape=(batch_size))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.get_variable("W1", shape=[patch_size, patch_size, num_channels, depth1],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
layer1_biases = tf.Variable(tf.constant(1.0, shape=[depth1]), name='B1')
layer2_weights = tf.get_variable("W2", shape=[patch_size, patch_size, depth1, depth2],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth2]), name='B2')
layer3_weights = tf.get_variable("W3", shape=[patch_size, patch_size, depth2, num_hidden1],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden1]), name='B3')
layer4_weights = tf.get_variable("W4", shape=[num_hidden1, num_hidden2],\
initializer=tf.contrib.layers.xavier_initializer())
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden2]), name='B4')
layer5_weights = tf.get_variable("W5", shape=[num_hidden2, num_labels],\
initializer=tf.contrib.layers.xavier_initializer())
layer5_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]), name='B5')
# Model.
def model(data, keep_prob, shape):
LCN = LecunLCN(data, shape)
conv = tf.nn.conv2d(LCN, layer1_weights, [1,1,1,1], 'VALID', name='C1')
hidden = tf.nn.relu(conv + layer1_biases)
lrn = tf.nn.local_response_normalization(hidden)
sub = tf.nn.max_pool(lrn, [1,2,2,1], [1,2,2,1], 'SAME', name='S2')
conv = tf.nn.conv2d(sub, layer2_weights, [1,1,1,1], padding='VALID', name='C3')
hidden = tf.nn.relu(conv + layer2_biases)
lrn = tf.nn.local_response_normalization(hidden)
sub = tf.nn.max_pool(lrn, [1,2,2,1], [1,2,2,1], 'SAME', name='S4')
conv = tf.nn.conv2d(sub, layer3_weights, [1,1,1,1], padding='VALID', name='C5')
hidden = tf.nn.relu(conv + layer3_biases)
hidden = tf.nn.dropout(hidden, keep_prob)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer4_weights) + layer4_biases)
return tf.matmul(hidden, layer5_weights) + layer5_biases
# Training computation.
logits = model(tf_train_dataset, 0.9375, shape)
loss = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
#optimizer = tf.train.AdagradOptimizer(0.01).minimize(loss)
global_step = tf.Variable(0)
learning_rate = tf.train.exponential_decay(0.05, global_step, 10000, 0.95)
optimizer = tf.train.AdagradOptimizer(learning_rate).minimize(loss, global_step=global_step)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(model(tf_train_dataset, 1.0, shape))
valid_prediction = tf.nn.softmax(model(tf_valid_dataset, 1.0, shape))
test_prediction = tf.nn.softmax(model(tf_test_dataset, 1.0, shape))
saver = tf.train.Saver()
num_steps = 100001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size)]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
save_path = saver.save(session, "CNN_1.ckpt")
print("Model saved in file: %s" % save_path)
# +
batch_size = 64
patch_size1 = 5
patch_size2 = 3
patch_size3 = 1
depth1 = 16
depth2 = 32
depth3 = 96
num_hidden1 = 64
num_hidden2 = 32
shape=[batch_size, image_size, image_size, num_channels]
# Construct a Inception Module CNN.
# C1: convolutional layer, batch_size x 28 x 28 x 16, convolution size: 5 x 5 x 1 x 16
# P2: max pool layer, batch_size x 14 x 14 x 16
# C3: convolutional layer, batch_size x 10 x 10 x 32, convolution size: 5 x 5 x 16 x 32
# Inception Module:
# Inception & Concat: output batch_size x 5 x 5 x 96
# C6: convolutional layer, batch_size x 5 x 5 x 32, convolution size: 1 x 1 x 96 x 32
# C7: convolutional layer, batch_size x 1 x 1 x 64, convolution size: 5 x 5 x 32 x 64
# Dropout layer
# F8: fully-connected layer, weight size: 64 x 16
# Output layer, weight size: 16 x 10
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.int64, shape=(batch_size))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.get_variable("W1", shape=[patch_size1, patch_size1, num_channels, depth1],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
layer1_biases = tf.Variable(tf.constant(1.0, shape=[depth1]), name='B1')
layer2_weights = tf.get_variable("W2", shape=[patch_size1, patch_size1, depth1, depth2],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth2]), name='B2')
layer6_weights = tf.get_variable("W6", shape=[patch_size3, patch_size3, depth3, depth2],\
initializer=tf.contrib.layers.xavier_initializer())
layer6_biases = tf.Variable(tf.constant(1.0, shape=[depth2]), name='B6')
layer3_weights = tf.get_variable("W3", shape=[patch_size1, patch_size1, depth2, num_hidden1],\
initializer=tf.contrib.layers.xavier_initializer())
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden1]), name='B3')
layer4_weights = tf.get_variable("W4", shape=[num_hidden1, num_hidden2],\
initializer=tf.contrib.layers.xavier_initializer())
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden2]), name='B4')
layer5_weights = tf.get_variable("W5", shape=[num_hidden2, num_labels],\
initializer=tf.contrib.layers.xavier_initializer())
layer5_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]), name='B5')
# Inception Variables.
incep_11_w = tf.get_variable("I11W", shape=[1, 1, depth2, 16],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_11_b = tf.Variable(tf.constant(1.0, shape=[16]), name='I11B')
incep_12_w = tf.get_variable("I12W", shape=[patch_size1, patch_size1, 16, 32],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_12_b = tf.Variable(tf.constant(1.0, shape=[32]), name='I12B')
incep_21_w = tf.get_variable("I21W", shape=[1, 1, depth2, 16],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_21_b = tf.Variable(tf.constant(1.0, shape=[16]), name='I21B')
incep_22_w = tf.get_variable("I22W", shape=[patch_size2, patch_size2, 16, 32],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_22_b = tf.Variable(tf.constant(1.0, shape=[32]), name='I22B')
incep_31_w = tf.get_variable("I31W", shape=[1, 1, depth2, 16],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_31_b = tf.Variable(tf.constant(1.0, shape=[16]), name='I31B')
incep_41_w = tf.get_variable("I41W", shape=[1, 1, depth2, 16],\
initializer=tf.contrib.layers.xavier_initializer_conv2d())
incep_41_b = tf.Variable(tf.constant(1.0, shape=[16]), name='I41B')
# Model.
def model(data, keep_prob, shape):
LCN = LecunLCN(data, shape)
conv = tf.nn.conv2d(LCN, layer1_weights, [1,1,1,1], 'VALID', name='C1')
hidden = tf.nn.relu(conv + layer1_biases)
lrn = tf.nn.local_response_normalization(hidden)
sub = tf.nn.max_pool(lrn, [1,2,2,1], [1,2,2,1], 'SAME', name='P2')
conv = tf.nn.conv2d(sub, layer2_weights, [1,1,1,1], padding='VALID', name='C3')
hidden = tf.nn.relu(conv + layer2_biases)
lrn = tf.nn.local_response_normalization(hidden)
# Inception Module
hidden = Inception(lrn)
conv = tf.nn.conv2d(hidden, layer6_weights, [1,1,1,1], padding='SAME', name='C6')
hidden = tf.nn.relu(conv + layer6_biases)
lrn = tf.nn.local_response_normalization(hidden)
conv = tf.nn.conv2d(lrn, layer3_weights, [1,1,1,1], padding='VALID', name='C7')
hidden = tf.nn.relu(conv + layer3_biases)
lrn = tf.nn.local_response_normalization(hidden)
hidden = tf.nn.dropout(lrn, keep_prob)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer4_weights) + layer4_biases)
return tf.matmul(hidden, layer5_weights) + layer5_biases
def Inception(input_tensor):
# Branch 1
conv1 = tf.nn.conv2d(input_tensor, incep_11_w, [1,1,1,1], padding='SAME', name='I1C1')
hidden1 = tf.nn.relu(conv1 + incep_11_b)
lrn1 = tf.nn.local_response_normalization(hidden1)
conv1 = tf.nn.conv2d(lrn1, incep_12_w, [1,2,2,1], padding='SAME', name='I1C2')
hidden1 = tf.nn.relu(conv1 + incep_12_b)
lrn1 = tf.nn.local_response_normalization(hidden1)
# Branch 2
conv2 = tf.nn.conv2d(input_tensor, incep_21_w, [1,1,1,1], padding='SAME', name='I2C1')
hidden2 = tf.nn.relu(conv2 + incep_21_b)
lrn2 = tf.nn.local_response_normalization(hidden2)
conv2 = tf.nn.conv2d(lrn2, incep_22_w, [1,2,2,1], padding='SAME', name='I2C2')
hidden2 = tf.nn.relu(conv2 + incep_22_b)
lrn2 = tf.nn.local_response_normalization(hidden2)
# Branch 3
sub3 = tf.nn.avg_pool(input_tensor, [1,3,3,1], [1,2,2,1], 'SAME', name='I3P1')
conv3 = tf.nn.conv2d(sub3, incep_31_w, [1,1,1,1], padding='SAME', name='I3C2')
hidden3 = tf.nn.relu(conv3 + incep_31_b)
lrn3 = tf.nn.local_response_normalization(hidden3)
# Branch 4
conv4 = tf.nn.conv2d(input_tensor, incep_41_w, [1,2,2,1], padding='SAME', name='I4C1')
hidden4 = tf.nn.relu(conv4 + incep_41_b)
lrn4 = tf.nn.local_response_normalization(hidden4)
# Concat
output_tensor = tf.concat(3, [lrn1, lrn2, lrn3, lrn4], name='CONCAT')
return output_tensor
# Training computation.
logits = model(tf_train_dataset, 1, shape)
loss = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
#optimizer = tf.train.AdagradOptimizer(0.01).minimize(loss)
global_step = tf.Variable(0)
learning_rate = tf.train.exponential_decay(0.01, global_step, 10000, 0.95)
optimizer = tf.train.AdagradOptimizer(learning_rate).minimize(loss, global_step=global_step)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(model(tf_train_dataset, 1.0, shape))
valid_prediction = tf.nn.softmax(model(tf_valid_dataset, 1.0, shape))
test_prediction = tf.nn.softmax(model(tf_test_dataset, 1.0, shape))
saver = tf.train.Saver()
num_steps = 100001
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
reader = tf.train.NewCheckpointReader("CNN_1.ckpt")
reader.get_variable_to_shape_map()
#saver.restore(session, "CNN_1.ckpt")
print("Model restored.")
#tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size)]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
#print(predictions[:])
if (step % 500 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
save_path = saver.save(session, "CNN_2.ckpt")
print("Model saved in file: %s" % save_path)
|
2_CNN_single.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Philosphy: Lets learn programming language to solve a problem and not to solve coding complexities.
# ### Mall Customer Segmentation Data
# https://www.kaggle.com/vjchoudhary7/customer-segmentation-tutorial-in-python
#
# #### Description of dataset:
# This file contains the basic information (ID, age, gender, income, spending score) about the customer
#
# #### Explore the dataset for follwoing questons:
# - Does age of a customer impact spending nature ?
# - Does Annual Income play a role in spendings by a customer ?
# Pythonic way to read a file
with open('Mall_Customers.csv',"r") as file: # f is a file handler, while "r" is the mode (r for read)
for line in file:
print(line)
# ### Refer below for more details on filehandlers in python
# https://docs.python.org/3.7/tutorial/inputoutput.html
"8,Female,23,18,94".split(",")
# +
# Initiate lists to store individual column values
read_list = []
customerID =[]
gender = []
age =[]
annual_income_dollar = []
spending_score = []
# Use Python I/O file handlers
with open('Mall_Customers.csv') as file:
for f in file:
temp = f.split(',') # splitting each line as values are separated by ','
print(temp)
customerID.append(temp[0]) # appending each value to respective list
gender.append(temp[1])
age.append(temp[2])
annual_income_dollar.append(temp[3])
spending_score.append(temp[4])
# Create dictionary to combine data
mall_dict = {customerID[0]:customerID[1:], gender[0]:gender[1:],age[0]:age[1:],
annual_income_dollar[0]: annual_income_dollar[1:],
spending_score[0]:spending_score[1:]}
# -
mall_dict
# +
temporary_list = []
for x in spending_score[1:]:
temp = x.split('\n')
temp = temp[0]
temp = int(temp)
temporary_list.append(temp)
print(temporary_list[0:5])
listComprehension = [int(x.split('\n')[0]) for x in spending_score[1:]]
print(listComprehension[0:5])
# -
l = [0,2,4,6]
l1 = []
for element in l:
temp = element + 10
temp = temp *2
l1.append(temp) # operation
print(l1)
l2 = [ (element +10)*2 for element in l]
l2
mall_dict['spending_score'] = [int(x.split('\n')[0]) for x in spending_score[1:]]
#print(mall_dict)
# __List Comprehensions__. List comprehensions is a pythonic way to provide a concise way to create lists. It consists of brackets containing an expression followed by a for clause, then zero or more for or if clauses.
mall_dict = {customerID[0]:customerID[1:], gender[0]:gender[1:],age[0]:[int(x) for x in age[1:]],
annual_income_dollar[0]: [int(x) for x in annual_income_dollar[1:]]
, spending_score[0]: [int(x.split('\n')[0]) for x in spending_score[1:]]}
print(mall_dict)
# ### Now thats tedious !!!! To Write so many lines to export a file :(
# ### Libraries
# Often times, we need either internal or external help for complicated computation tasks. In these occasions, we need to _import libraries_.
#
# #### Built in Packages
# Python provides many built-in packages to prevent extra work on some common and useful functions
# We will use __math__ as an example.
#
# #### Most Commonly used Packags for Machine Learning in Python:
# - Pandas http://pandas.pydata.org/pandas-docs/stable/reference/index.html
# - Numpy https://docs.scipy.org/doc/numpy/user/quickstart.html
# - Scipy https://docs.scipy.org/doc/scipy/reference/
# - Matplotlib https://matplotlib.org/contents.html
# - Scikit learn https://scikit-learn.org/stable/user_guide.html
# ### Pandas (Python Data Analysis Library) is a great package for data structures: DataFrame)
# A great library to slic and dice data and visualize it in a columnar format. Its API provides wide range of functionalities that makes data analysis a fun and true strength of Python as prefered choice over other programming languages.
# Export the Mall dataset using Pandas
import pandas as pd # loading pandas library and giving it an alise pd to make code less verbose
import numpy as np
df = pd.read_csv(filepath_or_buffer='Mall_Customers.csv')
type(df)
df.head()
df.tail()
df.columns
print(df.head()) # top n reocrds
print(df.tail(n=10)) # last n records
print(df.shape) # rows x columns
df.describe(include = "all")
df.groupby(['Gender']).mean()
ex = df.groupby(['Gender'])["Age"].max()
ex
ex.columns
df.columns
df.rename(columns = {'Annual Income (k$)':'Income' , 'Spending Score (1-100)': 'Score'},inplace= True)
df.head()
df.columns =['ID','Gender','Age','income', 'score']
df.head()
# +
a = df.head()
for row in a.index:
print(row, end = " ")
# +
df_top = df.head()
list(df_top.index)
# -
df.index.tolist()
# Rename a single or multiple columns
df.rename(columns={'CustomerID':'id','Spending Score (1-100)':'score','Annual Income (k$)':'income'},inplace=True)
df.columns
# ### Slicinig a data frame
# It is one the most common operation done while analyzing the data. Slicing of a datafrmae can happen using either name of columns / rows or location index of the columns.
df_new = df['id'] # to select a specific column by its name
df_new
df_new = df.loc[0:10,['Gender','Age']] # uisng poistional slicing of dataframe along with name of columns
df_new
df_new = df.iloc[0:10,0:2] # using iloc for positional slicing along rows and columns
df_new
# ### Filter & Sort dataframe
# Filtering or sorting dataframe is commonly done to extract specific values basis a rule. Filtering / Sorting can happen basis one more columns
l = [0,1,2,3]
l[::-1]
df.head()
df.iloc[::-1]
df_new = df[df['Gender'] == 'Male']
df_new
# Using & operator, don't forget to wrap the sub-statements with ()
df_new = df[(df['Gender'] == 'Male') | (df['Age'] >= 20)]
df_new
#Sorting of data frame basis a single columns
df_new = df.sort_values(by = ['Score'], ascending=[False])
df_new
#Sorting of data frame basis multiple columns
df_new = df.sort_values(by = ['Gender','Income'], ascending=[True,False])
df_new
# ### Data Visualization
# Visualization is one of the most important things of data analysis. Besides just producing ___readable___ plots, we should make an effort to improve the overall attractiveness of the plots. `matplotlib` is a powerful package for ___Python___ users. Let's start with an example.
from matplotlib import pyplot as plt
# %matplotlib inline
from numpy.polynomial.polynomial import polyfit
# this magic line is important to plot graph within jupyter notebook cell
# # Problems to solve
# - Does age of a customer impact spending nature ?
# - Does Annual Income play a role in spendings by a customer ?
x , y = df['Age'] , df.Score
b0, b1 = polyfit(x, y, 1)
plt.figure(figsize= (10,6))
plt.scatter(x = df['Age'], y = df.Score)
plt.plot(x, b0 + b1 * x)
plt.xlabel('Age')
plt.ylabel('Spending Score')
# set title
plt.title('Relationship between Age & Spending Score')# df.score - is another method to access a specific column of data frame
import seaborn as sns; sns.set(color_codes=True)
ax = sns.regplot(x="Age", y="Score", data=df)
ax.figure.set_figwidth(16)
ax.figure.set_figheight(8)
# The above graph does not have any labels and is inconclusive of any information. Lets add some context to the above scatter plot.
plt.figure(figsize=(16,8)) # Adding size to the graph- width by height
# Use `+` as marker; color set as `g` (green); size proportion to Y values
plt.scatter(x = df['Age'], y = df.Score, c='r')
# set x/y labels
plt.xlabel('Age')
plt.ylabel('Spending Score')
# set title
plt.title('Age vs Spending Score')
# ### Anatomy of Figure
# Before we go deeper, let's take a look at the structure of a figure in `matplotlib`:
#
# <img width=700 src="https://matplotlib.org/_images/anatomy1.png">
#
# As you can see, we have done things for many of the elements. There's no need to memorize them, though. We can always Google and find information on specific parts that we want to update.
fig, ax = plt.subplots()
#The Figure is like a canvas, and the Axes is a part of that canvas on which we will make a particular visualization.
fig, ax = plt.subplots()
fig.set_figwidth(16)
fig.set_figheight(8)
# What we just did, applying to `ax`
ax.scatter(x = df['Age'], y = df.Score, c='g')
#ax.plot(x = df['Age'], y = df.score, linestyle='dashed', color='k')
#ax.axis([0, 10, 0, 35])
ax.set_xlabel('Age')
ax.set_ylabel('Spending Score')
ax.set_title('Age Vs Spending Score')
means = df.groupby('Gender').mean()
print(means)
type(means)
print(means.index)
# +
fig, ax = plt.subplots()
ax.bar(x = means.index, height = means['Score'])
#ax.barh(x = df['Gender'], height = np.mean(df.score))
ax.set_title('Mean of Spending Score')
ax.set_xlabel('Gender')
ax.set_ylabel('Mean of Spending Score')
ax.set_ylim(top = 60)
for i, v in enumerate(means['Score']):
ax.text(i, v+2, s = int(v), color='blue', fontweight='bold')
# -
l = ["a","b","c"]
for pos , value in enumerate(l):
print("Position is: %s"%(pos))
print("Value is: %s"%(value))
|
60-120.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # BNN on Pynq
#
# This notebook covers how to use Binary Neural Networks on Pynq.
# It shows an example of handwritten digit recognition using a binarized neural network composed of 4 fully connected layers with 1024 neurons each, trained on the MNIST dataset of handwritten digits.
# In order to reproduce this notebook, you will need an external USB Camera connected to the PYNQ Board.
#
# ## 1.Instantiate the classifier
# Creating a classifier will automatically download the correct bitstream onto the device and load the weights trained on the specified dataset. By default there is one set of weights for the LFC network, the MNIST
# +
import bnn
print(bnn.available_params(bnn.NETWORK_LFC))
classifier = bnn.PynqBNN(network=bnn.NETWORK_LFC)
# -
# # 2. Download the network parameters
# The parameters of the network are downloaded in the Programmable logic memory, storing the trained weights on the MNIST dataset.
classifier.load_parameters("chars_merged")
# ## 3. Load the image from the camera
# The image is captured from the external USB camera and stored locally. The image is then enhanced in contract and brightness to remove background noise.
# The resulting image should show the digit on a white background:
# +
from PIL import Image as PIL_Image
from PIL import ImageEnhance
from PIL import ImageOps
orig_img_path = '/home/xilinx/image.jpg'
# !fswebcam --no-banner --save {orig_img_path} -d /dev/video0 2> /dev/null # Loading the image from the webcam
img = PIL_Image.open(orig_img_path).convert("L") # convert in black and white
#Image enhancement
contr = ImageEnhance.Contrast(img)
img = contr.enhance(3) # The enhancement values (contrast and brightness)
bright = ImageEnhance.Brightness(img) # depends on backgroud, external lights etc
img = bright.enhance(4.0)
#img = img.rotate(180) # Rotate the image (depending on camera orientation)
#Adding a border for future cropping
img = ImageOps.expand(img,border=80,fill='white')
threshold = 180
img = img.point(lambda p: p > threshold and 255)
img
# -
# ## 4. Crop and scale the image
# +
from PIL import Image as PIL_Image
import numpy as np
import math
from scipy import misc
#Find bounding box
inverted = ImageOps.invert(img)
box = inverted.getbbox()
img_new = img.crop(box)
width, height = img_new.size
ratio = min((28./height), (28./width))
background = PIL_Image.new('RGB', (28,28), (255,255,255))
if(height == width):
img_new = img_new.resize((28,28))
elif(height>width):
img_new = img_new.resize((int(width*ratio),28))
background.paste(img_new, (int((28-img_new.size[0])/2),int((28-img_new.size[1])/2)))
else:
img_new = img_new.resize((28, int(height*ratio)))
background.paste(img_new, (int((28-img_new.size[0])/2),int((28-img_new.size[1])/2)))
background
img_data=np.asarray(background)
img_data = img_data[:,:,0]
misc.imsave('/home/xilinx/img_webcam.png', img_data)
# -
# ## 5. Convert to BNN input format
# The image is resized to comply with the MNIST standard. The image is resized at 28x28 pixels and the colors inverted.
# +
from array import *
from PIL import Image as PIL_Image
img_load = PIL_Image.open('/home/xilinx/img_webcam.png').convert("L")
# Convert to BNN input format
# The image is resized to comply with the MNIST standard. The image is resized at 28x28 pixels and the colors inverted.
#Resize the image and invert it (white on black)
smallimg = ImageOps.invert(img_load)
smallimg = smallimg.rotate(0)
data_image = array('B')
pixel = smallimg.load()
for x in range(0,28):
for y in range(0,28):
if(pixel[y,x] == 255):
data_image.append(255)
else:
data_image.append(1)
# Setting up the header of the MNIST format file - Required as the hardware is designed for MNIST dataset
hexval = "{0:#0{1}x}".format(1,6)
header = array('B')
header.extend([0,0,8,1,0,0])
header.append(int('0x'+hexval[2:][:2],16))
header.append(int('0x'+hexval[2:][2:],16))
header.extend([0,0,0,28,0,0,0,28])
header[3] = 3 # Changing MSB for image data (0x00000803)
data_image = header + data_image
output_file = open('/home/xilinx/image.images-idx3-ubyte', 'wb')
data_image.tofile(output_file)
output_file.close()
smallimg
# -
# # 6. Launching BNN in hardware
# The image is passed in the PL and the inference is performed
class_out= classifier.inference("/home/xilinx/image.images-idx3-ubyte")
print("Class number: {0}".format(class_out))
print("Class name: {0}".format(classifier.class_name(class_out)))
|
notebooks/Char_Recognition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#queries
# -
import numpy as np
import pandas as pd
# +
from sqlalchemy import create_engine
from sqlalchemy import Table, MetaData
#engine = create_engine('sqlite://', echo=False)
engine = create_engine('mysql+pymysql://oasis:oasis@mysql/oasis', echo=False)
metadata = MetaData(bind=engine)
# -
# denormalize events
q = """
SELECT e.*, i.*, a.*, c.*
FROM events e
LEFT JOIN items i ON i.id = e.item_id
LEFT JOIN artifacts a ON a.id = i.artifact_id
LEFT JOIN categories c ON c.id = a.cat_id
LIMIT 3
"""
pd.read_sql_query(q, engine)
# best items for winning fights, by player
q = """
SELECT p.name, a.name, e.action, min(c.name), COUNT(*) as actions
FROM events e
LEFT JOIN players p ON p.id = e.player_id
LEFT JOIN races r ON r.id = p.race_id
LEFT JOIN items i ON i.id = e.item_id
LEFT JOIN artifacts a ON a.id = i.artifact_id
LEFT JOIN categories c ON c.id = a.cat_id
where e.action = 'win' and a.name IS NOT NULL
group by a.name, p.name, e.action
order by actions desc
"""
pd.read_sql_query(q, engine)
# best items for winning fights
q = """
SELECT a.name, e.action, min(c.name), COUNT(*) as actions
FROM events e
LEFT JOIN items i ON i.id = e.item_id
LEFT JOIN artifacts a ON a.id = i.artifact_id
LEFT JOIN categories c ON c.id = a.cat_id
where e.action = 'win' and a.name IS NOT NULL
group by a.name, e.action
order by actions desc
"""
pd.read_sql_query(q, engine)
# money makers
q = """
SELECT p.*, w.*, r.name
FROM wallets w
LEFT JOIN players p ON p.id = w.player_id
LEFT JOIN races r ON r.id = p.race_id
order by amount desc
"""
pd.read_sql_query(q, engine)
# +
# actions each minute
q = """
SELECT MIN(t.ts) AS ts,
t.h,
t.m,
t.action,
count(*) as count
FROM
( SELECT e.ts AS ts,
DATE(e.ts) AS d,
HOUR(e.ts) AS h,
ROUND(MINUTE(e.ts), -1) AS m,
e.action AS action
FROM events e
LEFT JOIN players p ON p.id = e.player_id ) AS t
GROUP BY t.d,
t.h,
t.m,
t.action
ORDER BY ts ASC
LIMIT 10
"""
pd.read_sql_query(q, engine)
# -
# actions by user
q = """
SELECT name,
race,
(win -lose) AS net_win,
(sell-buy) AS net_sell,
trade
FROM
(SELECT p.name,
MIN(r.name) as race,
count(CASE
WHEN e.action = 'win' THEN 1
END) win,
count(CASE
WHEN e.action = 'lose' THEN 1
END) lose,
count(CASE
WHEN e.action = 'sell' THEN 1
END) sell,
count(CASE
WHEN e.action = 'buy' THEN 1
END) buy,
count(CASE
WHEN e.action LIKE "trade%%" THEN 1
END) trade
FROM events e
LEFT JOIN players p ON p.id = e.player_id
LEFT JOIN races r ON r.id = p.race_id
GROUP BY p.name) o
"""
pd.read_sql_query(q, engine)
|
demos/oasis/demo/simulator/eda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jana0601/A_Summer_school_archiv-/blob/main/Copy_of_toy_example_students.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3d979c7f"
# First we download and import all neccessary packages
# + id="5aVhK5fwUFZr" colab={"base_uri": "https://localhost:8080/"} outputId="082c7f86-3123-41fd-f96b-736ee2e988e1"
# !git clone --depth=1 https://github.com/ntapiam/dagstuhl.git
# %cd dagstuhl
# + id="3f09593d"
import isstorch # Feature extraction procedure
from tqdm.notebook import trange # Pretty printing
# Some utilities
from utils import generate_examples, compute_signatures
# Import PyTorch and shorthand some components
import torch
import torch.nn as nn
from torch.optim import Adam
from torch.utils.data import DataLoader, TensorDataset
from sklearn.model_selection import train_test_split
# Utils for plotting
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] id="XUVWT0xoUa4e"
# Set the default `torch` device depending on whether there is a GPU available.
# You can change this by going to `Runtime > Change runtime type` and selecting among the available accelerators (None means CPU).
#
# We also run a random example to measure speed and check that everything is working OK.
# + id="0727924b" colab={"base_uri": "https://localhost:8080/"} outputId="f76ef0e2-ce6d-4bd8-89c1-27992ecc5d2c"
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
x = torch.randn(1000, 100, device=device)
if torch.cuda.is_available():
with torch.no_grad():
# %time isstorch.compute(x); torch.cuda.synchronize('cuda')
x = x.to('cpu')
with torch.no_grad():
# %time isstorch.compute(x);
# + [markdown] id="73a55913"
# Let's look at some samples. The function `generate_samples` outputs a different set of data each time.
#
# The dataset is generated from an ARMA(1,1) model with mean equal to 0.5 and different parameters for each class.
# More precisely, each sample in the class is generated by the recursion
# $$
# x_{t+1} = \phi x_t + 0.5 + \epsilon_t + \theta\epsilon_{t-1}
# $$
# where $\phi$ and $\theta$ are class dependent.
# + id="6637121d" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="31f5226d-991f-48ed-f0e0-24344d44dc09"
X, labels = generate_examples()
plt.plot(range(100), X[0], label="class 0")
plt.plot(range(100), X[500], label="class 1")
plt.xlabel('t')
plt.legend()
plt.show()
# + [markdown] id="27d94aa5"
# Now, we compute the signature features using our `pytorch` implementation.
# + id="f9a2bd78" colab={"base_uri": "https://localhost:8080/"} outputId="832333b7-2fce-4400-afba-1fd2a504da03"
with torch.no_grad():
# %time sigs = compute_signatures(X, level=4) # Hyperparameter chosen here!
# Play with it and see how it
# changes the results
sigs.shape # Check how many features we obtain
# + [markdown] id="5f796922"
# In order to train and test our Neural Network, we split the data into a training set and a test set using the auxiliary function `train_test_split` from the `sklearn` package. The stratification ensures a balanced distribution of both classes in each split.
#
# We use 70% of the data for training. Try modifying this value to see how the results change.
#
# We need to wrap the data in a PyTorch `DataLoader` [(docs)](https://pytorch.org/docs/stable/data.html).
# We also load the data in batches.
# + id="0f11e74f"
train_sigs, test_sigs, train_labels, test_labels = train_test_split(sigs, labels, stratify=labels, train_size=0.7)
train_dataloader = DataLoader(TensorDataset(train_sigs, train_labels), batch_size=100)
test_dataloader = DataLoader(TensorDataset(test_sigs, test_labels), batch_size=100)
# + [markdown] id="3f294f52"
# We build our model using by subclassing `torch.nn.Module`.
#
# To see all possible layers and activations, please see the `torch.nn` [(docs)](https://pytorch.org/docs/stable/nn.html).
# + id="f2b3e163"
class DenseNet(torch.nn.Module):
def __init__(self, hidden_layers=1,width=10,in_features=3):
super().__init__()
self.batch_norm = nn.BatchNorm1d(in_features)
input_layer = nn.Sequential(
nn.Linear(in_features=in_features,out_features=width),
nn.ReLU()
)
layers = [input_layer]
for k in range(hidden_layers):
layers.append(nn.Sequential(
nn.Linear(in_features=width,out_features=width),
nn.ReLU()
)
)
layers.append(nn.Sequential(
nn.Linear(in_features=width,out_features=2),
nn.Tanh()
)
)
self.stack = nn.Sequential(*layers)
def forward(self, x):
x = self.batch_norm(x)
return self.stack(x)
# + id="fWehjQjH4QWN"
class DenseNet(nn.Module):
def __init__(self, hidden_layers=1, width=10, in_features=3):
super().__init__()
self.batch_norm = nn.BatchNorm1d(in_features)
input_layer = nn.Sequential(
nn.Linear(in_features=in_features, out_features=width),
nn.ReLU()
)
layers = [input_layer]
for k in range(hidden_layers):
layers.append(nn.Sequential(
nn.Linear(in_features=width, out_features=width),
nn.ReLU()
)
)
layers.append(nn.Sequential(
nn.Linear(in_features=width, out_features=2),
nn.Tanh()
)
)
self.stack = nn.Sequential(*layers)
def forward(self, x):
x = self.batch_norm(x)
return self.stack(x)
# + [markdown] id="31146f47"
# Next, we set up the training and testing loops
# + id="524e7150"
def train_loop(model, dataloader, loss_fn, optimizer, device):
#pass
### FILL IN
## Some pointers:
## - Load the data in batches by iterating through `dataloader`
## - Make a prediction using `model` and evaluate it using `loss_fn`
## - Backpropagate the error through the network
## - Finally, use `optimizer` to modify the weights (remember to reset them at each batch)
for X,y in dataloader:
X, y = X.to(device),y.to(device)
pred = model(X)
loss = loss_fn(pred, y)
optimizer.zero_grad()
loss.backward()
optimizer.step()
def test_loop(model, dataloader, loss_fn, device):
### FILL IN
## Some pointers:
## - Load the data by iterating through `dataloader`
## - Make a prediction using `model` and evaluate it using `loss_fn`
## - Compute the number of correct guesses by comparing your predictions to the 'ground truth'
## - Return the accuracy and average loss value of your model.
# assume we have X,y from batch
total_samples = len(dataloader.dataset)
test_loss,correct = 0,0
with torch.no_grad():
for X,y in dataloader:
pred = model(X)
test_loss += loss_fn(pred,y).item()
correct += (pred.argmax(dim = 1) == y).type(torch.float).sum().item()
return correct/total_samples, test_loss / total_samples
# + [markdown] id="7f7d5136"
# Setup our model, loss function and optimizer [(docs)](https://pytorch.org/docs/stable/optim.html#algorithms).
#
# Train the network and output the result.
# + id="lDQUVXyt172M"
# + id="eb4fe2ca" colab={"base_uri": "https://localhost:8080/", "height": 116, "referenced_widgets": ["0999da7fc66d4ea3afbaf538c601f979", "fefd938113944be395758bbc451be5e1", "4941fadfe322451b87e2c3f7aae7a089", "<KEY>", "c5199827baee45fb9ec63ab9667ccce5", "f0b69e7da8ba4d78ab4a88dbe6bc48c3", "<KEY>", "<KEY>", "<KEY>", "2c51a57febdd4476a344e3bab0042a82", "ef393024a7d849839e69774ae9580a4c"]} outputId="cea05e40-a3a4-4a88-b86a-aed740fb63d0"
# %%time
model = DenseNet(hidden_layers=1,width=20,in_features=len(sigs[0]))## FILL IN (remember to call .to(device) if training on the GPU!)
optimizer = Adam(model.parameters(), lr=1e-3) # One possible choice of optimizer
loss_fn = nn.CrossEntropyLoss() ## We choose the Negative Log Likelihood loss
epochs = 100 # Hyperparameter, feel free to change it
losses = [] # Record the losses during training for some a posteriori analysis
accuracies = [] # Same with accuracies
# The following is some pretty printing
with trange(epochs) as pbar:
for t in pbar:
train_loop(model, train_dataloader, loss_fn, optimizer, device)
acc, avg_loss = test_loop(model, test_dataloader, loss_fn, device)
accuracies.append(acc)
losses.append(avg_loss)
pbar.set_description(f"Training")
pbar.set_postfix(loss=avg_loss, accuracy=acc)
# Run the test loop once more to verify our accuracy
acc, avg_loss = test_loop(model, test_dataloader, loss_fn, device)
print(f"\nDone! Accuraccy: {acc:.2%}, Loss: {avg_loss:f}")
# + [markdown] id="0NEYm84TuEHp"
# Now, we can make a predicition on our test data (or even new inputs), which will be given as a probability distribution over class labels.
#
# Note that we apply the exponential mapping in order to obtain (true) probabilities, as the network outputs negative log probas.
# + id="EurkCtLfsE-F"
preds = model(test_sigs[:10]).exp()
# + [markdown] id="IYdBTLwjuf8n"
# We check that they sum to 1 for each example.
# + id="crgNwQUrunrK" colab={"base_uri": "https://localhost:8080/"} outputId="6542a5f1-6a6a-4c3c-95fe-f926fe409e8f"
torch.allclose(preds.sum(dim=1), torch.ones(10)) # Need to use `allclose` to account for floating point errors
# + [markdown] id="HGv4o4ocwUWe"
# The predicted label is the one with the highest probability.
# + id="Zda9btrxuxYT" colab={"base_uri": "https://localhost:8080/"} outputId="899f5436-9638-4c59-9389-bafa15bdbe14"
preds.argmax(dim=1), test_labels[:10]
# + [markdown] id="1xFUPRc90ENd"
# The following cell plots the evolution of the total loss and accuracy across training epochs.
# + id="xTL6ueOcWbIW" colab={"base_uri": "https://localhost:8080/", "height": 310} outputId="e188b86f-aa87-4400-bcd6-e52a9f89b237"
sns.set_theme() # Prettify plots
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(range(epochs), losses, color='tab:red')
ax2.plot(range(epochs), accuracies, color='tab:blue')
ax1.set_xlabel('epochs')
ax1.set_ylabel('losses')
ax2.set_ylim(bottom=0, top=1)
ax2.set_ylabel('accuracy')
fig.tight_layout()
plt.plot()
# + [markdown] id="toHZnQ2s0PLw"
# Next, we compare with the same model but trained on raw time-series data.
# + id="2fecveXkdKOB"
#X, labels = generate_examples()
train_data, test_data, train_labels, test_labels = train_test_split(X.flatten(start_dim=1), labels, stratify=labels, train_size=0.7)
#train_sigs, test_sigs, train_labels, test_labels = train_test_split(sigs, labels, stratify=labels, train_size=0.7)
train_dataloader = DataLoader(TensorDataset(train_data, train_labels), batch_size=100)
test_dataloader = DataLoader(TensorDataset(test_data, test_labels), batch_size=100)
# + id="ndZYJ0zfdaPm" colab={"base_uri": "https://localhost:8080/", "height": 116, "referenced_widgets": ["ea606941e4734c7fbde47c3916bc4678", "f2134d778dc74160876d9667457cc114", "bf132b3af14c4b14b1b2ce0b69be69c2", "86ad3e9150254a16a7d2592b58490b8b", "<KEY>", "e244d513f0144d34859a56220960c076", "fcb1880962af4e179337a06185791c1e", "7e777305ba76405ba1ac8a06122a9649", "4f7def823ef0400492aaa22574d21183", "477859779b9e4d8aab22405776fecd2e", "<KEY>"]} outputId="1dc0579b-8438-4a3c-9d4b-935a0d8b5e16"
# %%time
in_features=100
new_model = DenseNet(hidden_layers=1, width=20, in_features=len(X.flatten(start_dim=1)[0]))## Use the same model as before with the appropriate parameteres
new_optimizer = Adam(new_model.parameters(), lr=1e-3) # Initialize optimizer
new_loss_fn = nn.CrossEntropyLoss()
accuracies2 = []
losses2 = []
with trange(epochs) as pbar:
for t in pbar:
train_loop(new_model, train_dataloader, new_loss_fn, new_optimizer, device)
acc, avg_loss = test_loop(new_model, test_dataloader, new_loss_fn, device)
pbar.set_description(f"Training")
pbar.set_postfix(loss=avg_loss, accuracy=acc)
accuracies2.append(acc)
losses2.append(avg_loss)
acc, avg_loss = test_loop(new_model, test_dataloader, new_loss_fn, device)
print(f"\nDone! Accuraccy: {acc:.2%}, Loss: {avg_loss:f}")
# + colab={"base_uri": "https://localhost:8080/", "height": 310} id="_CW2Q3_u7_N3" outputId="c99c8a10-ec45-4b45-f732-47bdf183b128"
sns.set_theme() # Prettify plots
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(range(epochs), losses2, color='tab:red')
ax2.plot(range(epochs), accuracies2, color='tab:blue')
ax1.set_xlabel('epochs')
ax1.set_ylabel('losses')
ax2.set_ylim(bottom=0, top=1)
ax2.set_ylabel('accuracy')
fig.tight_layout()
plt.plot()
|
Copy_of_toy_example_students.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# <center>*Copyright <NAME> 2017*</center>
# <center>*For more information, visit us at www.pieriandata.com*</center>
# # Stock Market Analysis Project
#
# ### Please Note: You are free to treat this as a full exercise, or just view the solutions video as a code along project. This project is meant to be pretty challenging as it will introduce a few new concepts through some hints!
#
# Welcome to your first capstone project! This project is meant to cap off the first half of the course, which mainly dealt with learning the libraries that we use in this course, the second half of the course will deal a lot more with quantitative trading techniques and platforms.
#
# We'll be analyzing stock data related to a few car companies, from Jan 1 2012 to Jan 1 2017. Keep in mind that this project is mainly just to practice your skills with matplotlib, pandas, and numpy. Don't infer financial trading advice from the analysis we do here!
#
# ### Part 0: Import
#
# ** Import the various libraries you will need-you can always just come back up here or import as you go along :) **
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# ____
# ## Part 1: Getting the Data
#
# ### Tesla Stock (Ticker: TSLA on the NASDAQ)
#
# ** *Note! Not everyone will be working on a computer that will give them open access to download the stock information using pandas_datareader (firewalls, admin permissions, etc...). Because of this, the csv file for the Tesla is provided in a data folder inside this folder. It is called Tesla_Stock.csv. Feel free to just use this with read_csv! **
#
# ** Use pandas_datareader to obtain the historical stock information for Tesla from Jan 1, 2012 to Jan 1, 2017.**
#
#
import pandas_datareader.data as web
import datetime
start = datetime.datetime(2012, 1, 1)
end = datetime.datetime(2017, 1, 1)
tesla_stock = web.DataReader('TSLA', 'google', start, end)
tesla_stock.head()
# CSV files is used since Google Finance does not provide full data.
tesla_stock = pd.read_csv('./Tesla_Stock.csv',
index_col= 'Date')
tesla_stock.head()
# ### Other Car Companies
#
# ** Repeat the same steps to grab data for Ford and GM (General Motors), **
ford_stock = pd.read_csv('./Ford_Stock.csv', index_col= 'Date')
ford_stock.head()
gm_stock = pd.read_csv('./GM_Stock.csv', index_col= 'Date')
gm_stock.head()
# ## Part 2: Visualizing the Data
#
# ** Time to visualize the data. **
#
# ** Follow along and recreate the plots below according to the instructions and explanations. **
# ____
#
# ** Recreate this linear plot of all the stocks' Open price ! Hint: For the legend, use label parameter and plt.legend()**
# +
# Code Here
fig = plt.figure(figsize = (12, 6))
plt.title('Open')
tesla_stock['Open'].plot(label = 'Tesla')
ford_stock['Open'].plot(label = 'Ford')
gm_stock['Open'].plot(label = 'GM')
plt.legend()
# -
# ____
# ** Plot the Volume of stock traded each day.**
# +
fig = plt.figure(figsize = (12, 6))
plt.title('Volume')
tesla_stock['Volume'].plot(label = 'Tesla')
ford_stock['Volume'].plot(label = 'Ford')
gm_stock['Volume'].plot(label = 'GM')
plt.legend()
# -
# ** Interesting, looks like Ford had a really big spike somewhere in late 2013. What was the date of this maximum trading volume for Ford?**
#
# **Bonus: What happened that day? **
ford_stock['Volume'].argmax()
# +
# http://money.cnn.com/2013/12/18/news/companies/ford-profit/index.html
# -
# ____
# ** The Open Price Time Series Visualization makes Tesla look like its always been much more valuable as a company than GM and Ford. But to really understand this we would need to look at the total market cap of the company, not just the stock price. Unfortunately our current data doesn't have that information of total units of stock present. But what we can do as a simple calcualtion to try to represent total money traded would be to multply the Volume column by the Open price. Remember that this still isn't the actual Market Cap, its just a visual representation of the total amount of money being traded around using the time series. (e.g. 100 units of stock at \$10 each versus 100000 units of stock at $1 each)**
# ** Create a new column for each dataframe called "Total Traded" which is the Open Price multiplied by the Volume Traded.**
# Code Here
tesla_stock['Total Traded'] = tesla_stock['Open'] * tesla_stock['Volume']
ford_stock['Total Traded'] = ford_stock['Open'] * ford_stock['Volume']
gm_stock['Total Traded'] = gm_stock['Open'] * gm_stock['Volume']
# ** Plot this "Total Traded" against the time index.**
# +
# Code here
fig = plt.figure(figsize = (12, 6))
plt.title('Total Traded')
tesla_stock['Total Traded'].plot(label = 'Tesla')
ford_stock['Total Traded'].plot(label = 'Ford')
gm_stock['Total Traded'].plot(label = 'GM')
plt.legend()
# -
# ** Interesting, looks like there was huge amount of money traded for Tesla somewhere in early 2014. What date was that and what happened? **
tesla_stock['Total Traded'].argmax()
# +
# http://money.cnn.com/2014/02/25/investing/tesla-record-high/
# -
# ____
# ** Let's practice plotting out some MA (Moving Averages). Plot out the MA50 and MA200 for GM. **
# +
# Code here
# -
fig = plt.figure(figsize = (12, 6))
gm_stock.rolling(window = 50).mean()['Open'].plot(label = 'MA50')
gm_stock.rolling(window = 200).mean()['Open'].plot(label = 'MA200')
plt.legend()
# ______
# ** Finally lets see if there is a relationship between these stocks, after all, they are all related to the car industry. We can see this easily through a scatter matrix plot. Import scatter_matrix from pandas.plotting and use it to create a scatter matrix plot of all the stocks'opening price. You may need to rearrange the columns into a new single dataframe. Hints and info can be found here: https://pandas.pydata.org/pandas-docs/stable/visualization.html#scatter-matrix-plot **
from pandas.plotting import scatter_matrix
df = pd.concat([tesla_stock['Open'], ford_stock['Open'], gm_stock['Open']], axis = 1)
df.columns = ['Tesla', 'Ford', 'GM']
df.head()
scatter_matrix(df, figsize = (10, 10), hist_kwds = {'bins' : 100})
# _____
# ### Bonus Visualization Task! (Note: This is hard!)
# **Let's now create a candlestick chart! Watch the video if you get stuck on trying to recreate this visualization, there are quite a few steps involved!Refer to the video to understand how to interpret and read this chart. Hints: https://matplotlib.org/examples/pylab_examples/finance_demo.html**
#
# ** Create a CandleStick chart for Ford in January 2012 (too many dates won't look good for a candlestick chart)**
start = '2012-01'
end = '2012-02'
ford_candle = ford_stock.loc[start:end]
# +
# To be continued
# -
# ____
# # Part 3: Basic Financial Analysis
#
# Now it is time to focus on a few key financial calculations. This will serve as your transition to the second half of the course. All you need to do is follow along with the instructions, this will mainly be an exercise in converting a mathematical equation or concept into code using python and pandas, something we will do often when working with quantiative data! If you feel very lost in this section, don't worry! Just go to the solutions lecture and treat it as a code-along lecture, use whatever style of learning works best for you!
#
# Let's begin!
# ____
# ## Daily Percentage Change
# First we will begin by calculating the daily percentage change. Daily percentage change is defined by the following formula:
# $ r_t = \frac{p_t}{p_{t-1}} -1$
# This defines r_t (return at time t) as equal to the price at time t divided by the price at time t-1 (the previous day) minus 1. Basically this just informs you of your percent gain (or loss) if you bought the stock on day and then sold it the next day. While this isn't necessarily helpful for attempting to predict future values of the stock, its very helpful in analyzing the volatility of the stock. If daily returns have a wide distribution, the stock is more volatile from one day to the next. Let's calculate the percent returns and then plot them with a histogram, and decide which stock is the most stable!
# ** Create a new column for each dataframe called returns. This column will be calculated from the Close price column. There are two ways to do this, either a simple calculation using the .shift() method that follows the formula above, or you can also use pandas' built in pct_change method. **
tesla_stock['returns'] = (tesla_stock['Close'] / tesla_stock['Close'].shift(1)) - 1
tesla_stock.head()
ford_stock['returns'] = (ford_stock['Close'] / ford_stock['Close'].shift(1)) - 1
gm_stock['returns'] = (gm_stock['Close'] / gm_stock['Close'].shift(1)) - 1
# ** Now plot a histogram of each companies returns. Either do them separately, or stack them on top of each other. Which stock is the most "volatile"? (as judged by the variance in the daily returns we will discuss volatility in a lot more detail in future lectures.)**
# +
# Separately
fig = plt.figure(0)
tesla_stock['returns'].plot(kind = 'hist', bins = 50)
plt.title('Tesla')
plt.show()
fig = plt.figure(1)
ford_stock['returns'].plot(kind = 'hist', bins = 50)
plt.title('Ford')
plt.show()
fig = plt.figure(2)
gm_stock['returns'].plot(kind = 'hist', bins = 50)
plt.title('GM')
plt.show()
# -
# On one graph.
fig = plt.figure(figsize = (12, 10))
tesla_stock['returns'].plot(kind = 'hist',
bins = 50,
label = 'Tesla',
alpha = 0.5)
ford_stock['returns'].plot(kind = 'hist',
bins = 50,
label = 'Ford',
alpha = 0.8)
gm_stock['returns'].plot(kind = 'hist',
bins = 50,
label = 'GM',
alpha = 0.4)
plt.legend()
# ** Try also plotting a KDE instead of histograms for another view point. Which stock has the widest plot? **
fig = plt.figure(figsize = (12, 10))
tesla_stock['returns'].plot(kind = 'kde',
label = 'Tesla',
alpha = 0.5)
ford_stock['returns'].plot(kind = 'kde',
label = 'Ford',
alpha = 0.8)
gm_stock['returns'].plot(kind = 'kde',
label = 'GM',
alpha = 0.4)
plt.legend()
# ** Try also creating some box plots comparing the returns. **
box_df = pd.concat([tesla_stock['returns'], ford_stock['returns'], gm_stock['returns']], axis = 1)
box_df.columns = ['Tesla', 'Ford', 'GM']
box_df.plot(kind = 'box', figsize = (12, 10))
plt.legend()
# ## Comparing Daily Returns between Stocks
#
# ** Create a scatter matrix plot to see the correlation between each of the stocks daily returns. This helps answer the questions of how related the car companies are. Is Tesla begin treated more as a technology company rather than a car company by the market?**
# +
from pandas.plotting import scatter_matrix
scatter_matrix(box_df, figsize = (10, 10), hist_kwds={'bins':50})
# -
# ** It looks like Ford and GM do have some sort of possible relationship, let's plot just these two against eachother in scatter plot to view this more closely!**
fig = plt.figure(figsize = (12, 8))
plt.scatter(ford_stock['returns'], gm_stock['returns'])
# ____
# ## Cumulative Daily Returns
#
# Great! Now we can see which stock was the most wide ranging in daily returns (you should have realized it was Tesla, our original stock price plot should have also made that obvious).
#
# With daily cumulative returns, the question we are trying to answer is the following, if I invested $1 in the company at the beginning of the time series, how much would is be worth today? This is different than just the stock price at the current day, because it will take into account the daily returns. Keep in mind, our simple calculation here won't take into account stocks that give back a dividend. Let's look at some simple examples:
# Lets us say there is a stock 'ABC' that is being actively traded on an exchange. ABC has the following prices corresponding to the dates given
# Date Price
# 01/01/2018 10
# 01/02/2018 15
# 01/03/2018 20
# 01/04/2018 25
# **Daily Return** : Daily return is the profit/loss made by the stock compared to the previous day. (This is what ew just calculated above). A value above one indicates profit, similarly a value below one indicates loss. It is also expressed in percentage to convey the information better. (When expressed as percentage, if the value is above 0, the stock had give you profit else loss). So for the above example the daily returns would be
# Date Daily Return %Daily Return
# 01/01/2018 10/10 = 1 -
# 01/02/2018 15/10 = 3/2 50%
# 01/03/2018 20/15 = 4/3 33%
# 01/04/2018 25/20 = 5/4 20%
# **Cumulative Return**: While daily returns are useful, it doesn't give the investor a immediate insight into the gains he had made till date, especially if the stock is very volatile. Cumulative return is computed relative to the day investment is made. If cumulative return is above one, you are making profits else you are in loss. So for the above example cumulative gains are as follows
# Date Cumulative Return %Cumulative Return
# 01/01/2018 10/10 = 1 100 %
# 01/02/2018 15/10 = 3/2 150 %
# 01/03/2018 20/10 = 2 200 %
# 01/04/2018 25/10 = 5/2 250 %
# The formula for a cumulative daily return is:
#
# $ i_i = (1+r_t) * i_{t-1} $
#
# Here we can see we are just multiplying our previous investment at i at t-1 by 1+our percent returns. Pandas makes this very simple to calculate with its cumprod() method. Using something in the following manner:
#
# df[daily_cumulative_return] = ( 1 + df[pct_daily_return] ).cumprod()
#
# ** Create a cumulative daily return column for each car company's dataframe.**
tesla_stock['Cumulative Return'] = (1 + tesla_stock['returns']).cumprod()
tesla_stock.head()
ford_stock['Cumulative Return'] = (1 + ford_stock['returns']).cumprod()
gm_stock['Cumulative Return'] = (1 + gm_stock['returns']).cumprod()
# ** Now plot the Cumulative Return columns against the time series index. Which stock showed the highest return for a $1 invested? Which showed the lowest?**
fig = plt.figure(figsize = (12, 6))
tesla_stock['Cumulative Return'].plot(label = 'Tesla')
ford_stock['Cumulative Return'].plot(label = 'Ford')
gm_stock['Cumulative Return'].plot(label = 'GM')
plt.legend()
# # Great Job!
#
# That is it for this very basic analysis, this concludes this half of the course, which focuses much more on learning the tools of the trade. The second half of the course is where we really dive into functionality designed for time series, quantitative analysis, algorithmic trading, and much more!
|
07-Stock-Market-Analysis-Capstone-Project/Stock Market Analysis Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import sys
sys.path.append('..')
import functools
import math
import numpy as np
import torch
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import sobol_seq
import bvbq_torch
# -
mixmeans_test = torch.tensor([[ 1.8033],
[ -1.4296],
[ -1.9691],
[ 1.2147],
[ -6.9166],
[-11.5672],
[ -1.5860],
[ -4.3302],
[ 8.7410],
[ -1.3793],
[ -4.8774],
[ 2.3949]])
mixvars_test = torch.ones((12,1))
mixweights_test = torch.ones(12)/12
# +
def logprob_mixgaussians(x):
return bvbq_torch.distributions.MixtureDiagonalNormalDistribution.logprob_(
x,mixmeans_test,mixvars_test,mixweights_test).reshape(-1,1)
def logprob_t(x,nu):
normalizer = jax.scipy.special.gammaln((nu+1)/2) \
-jax.scipy.special.gammaln(nu/2) \
-0.5*jnp.log(jnp.pi*nu)
main_term = -(nu+1)/2*jnp.log(1+x**2/nu)
return main_term + normalizer
xdata = torch.linspace(-20,20,51).reshape(-1,1)
ydata = logprob_mixgaussians(xdata)
logprobgp = bvbq_torch.gp.SimpleGP(1,kind='smatern32',noise=0,zeromax=True)
logprobgp.mean = -10.0
logprobgp.fix_mean()
logprobgp.fix_noise()
logprobgp.set_data(xdata,ydata,empirical_params=False)
# logprobgp.optimize_params(tol=1e-1)
# -
ydata.shape
gp = bvbq_torch.gp.SimpleGP(1,kind='smatern32',noise=1e-2)
gp.set_data(xdata,ydata)
gp.mean = -10.0
gp.fix_mean()
gp.fix_noise()
gp.optimize_params(tol=1e-1)
xplot = np.linspace(-20,20,201).reshape(-1,1)
yplot1 = logprob_mixgaussians(xplot).detach().numpy()
# yplot1 = logprob_t(xplot,2.5)
yplot2 = logprobgp.predict(xplot,return_cov=False).detach().numpy()
plt.plot(xplot,np.exp(yplot1),'b')
plt.plot(xplot,np.exp(yplot2),'r')
samples1 = bvbq_torch.distributions.MixtureDiagonalNormalDistribution.sample_(100000,
mixmeans_test,
mixvars_test,
mixweights_test).detach().numpy()
plt.hist(samples1.flatten(),density=True,bins=100);
bvbq_torch.distributions.DiagonalNormalDistribution.sample_
# +
def mcbq_dmvn_relbo(logprobgp,mean,var,mixmeans,mixvars,mixweights,
nsamples=100,logdelta=-20,reg=1.0):
term1 = bvbq_torch.bayesquad.separable_dmvn_bq(logprobgp,mean,var,return_var=False) #Variance
samples = bvbq_torch.distributions.DiagonalNormalDistribution.sample_(nsamples,mean,var)
term2_ = bvbq_torch.distributions.MixtureDiagonalNormalDistribution.logprob_(
samples,mixmeans,mixvars,mixweights)
term2 = -bvbq_torch.utils.logbound(term2_,logdelta).mean()
term3 = 0.5*torch.sum(torch.log(2*math.pi*math.e*var)) #Entropy
return term1 + term2 + reg*term3
def mcbq_mixdmvn_gradboost_delbodw(weight,logprobgp,mean,var,
mixmeans,mixvars,mixweights,
nsamples=1000):
weight = bvbq_torch.utils.tensor_convert(weight)
mixmeans_up = torch.vstack([mixmeans,mean])
mixvars_up = torch.vstack([mixvars,var])
mixweights_up = torch.hstack([(1-weight)*mixweights,weight])
samplesprevious = bvbq_torch.distributions.MixtureDiagonalNormalDistribution.sample_(
nsamples,mixmeans,mixvars,mixweights)
samplesproposal = bvbq_torch.distributions.DiagonalNormalDistribution.sample_(nsamples,mean,var)
term1 = bvbq_torch.bayesquad.separable_dmvn_bq(logprobgp,mean,var,return_var=False)
term2 = -bvbq_torch.bayesquad.separable_mixdmvn_bq(logprobgp,mixmeans,
mixvars,mixweights,
return_var=False)
term3 = -bvbq_torch.distributions.MixtureDiagonalNormalDistribution.logprob_(
samplesproposal,mixmeans_up,mixvars_up,mixweights_up).mean()
term4 = bvbq_torch.distributions.MixtureDiagonalNormalDistribution.logprob_(
samplesprevious,mixmeans_up,mixvars_up,mixweights_up).mean()
return term1 + term2 + term3 + term4
# -
nmixtures = 3
ndim = 1
mixmeans = torch.randn(nmixtures,1)
mixvars = torch.distributions.HalfNormal(1.0).sample((nmixtures,1))
mixweights = torch.distributions.Dirichlet(torch.ones(nmixtures)).sample()
mean = torch.randn((1,))
var = torch.distributions.HalfNormal(1.0).sample((1,))
# +
alpha_relbo = 1e-1
maxiter_relbo = 100
alpha_boost = 1e-1
maxiter_boost = 100
maxiter = 20
mixmeans = torch.zeros(1,ndim)
mixvars = (20.0/3)**2*torch.ones(1,ndim)
mixweights = torch.ones(1)
for k in range(maxiter):
mean0 = bvbq_torch.distributions.MixtureDiagonalNormalDistribution.sample_(1,mixmeans,mixvars,mixweights)[0,:]
var0 = torch.distributions.HalfNormal(1.0).sample((ndim,))
rawvar0 = torch.log(torch.exp(var0)-1)
optimizer = torch.optim.Adam([mean0,rawvar0],lr=alpha_relbo)
mean0.requires_grad = True
rawvar0.requires_grad = True
# reg = torch.rand(1)
for i in range(maxiter_relbo):
optimizer.zero_grad()
var0 = torch.log(torch.exp(rawvar0)+1)
reg = torch.rand(1)
relbo = mcbq_dmvn_relbo(logprobgp,mean0,var0,mixmeans,mixvars,mixweights,reg=reg)
loss = -relbo
loss.backward()
optimizer.step()
mean = mean0.detach()
rawvar = rawvar0.detach()
var = torch.log(torch.exp(rawvar)+1.0)
weight = torch.tensor(1e-4)
for j in range(maxiter_boost):
grad = mcbq_mixdmvn_gradboost_delbodw(weight,
logprobgp,
mean,
var,
mixmeans,
mixvars,
mixweights,
nsamples=1000)
dweight = alpha_boost/(k+1)*grad
weight += dweight
weight = torch.clamp(weight,1e-4,1-1e-4)
mixmeans = torch.vstack([mixmeans,mean])
mixvars = torch.vstack([mixvars,var])
mixweights = torch.hstack([(1-weight)*mixweights,weight])
print(mean,var,weight)
distrib = bvbq_torch.distributions.MixtureDiagonalNormalDistribution(mixmeans,mixvars,mixweights)
# maxiter = kwargs.get("maxiter_nc",300)
# lr = kwargs.get("lr_nc",0.1)
# distrib_constant = kwargs.get("distrib_constant_nc",1.0)
# distrib_type = kwargs.get("distrib_type_nc","HN")
# verbose = kwargs.get("verbose",1)
# nsamples_nc = kwargs.get("n_samples_nc",100)
# if self.num_mixtures == 0:
# raise NotImplementedError
# else:
# mu0 = self._sample_from_current(1,flatten=True)
# if distrib_type == "HN":
# distrib = torch.distributions.HalfNormal
# elif distrib_type == "HC":
# distrib = torch.distributions.HalfCauchy
# cov0 = distrib(torch.tensor([distrib_constant],device=self.device)).\
# sample((self.dim,)).flatten()
# raw_cov0 = utils.invsoftplus(cov0)
# theta0 = torch.cat([mu0,raw_cov0])
# theta0.requires_grad = True
# optimizer = torch.optim.Adam([theta0],lr=lr)
# for i in range(maxiter):
# optimizer.zero_grad()
# theta0.requires_grad = True
# loss = self._evaluate_loss(theta0,nsamples=nsamples_nc,
# verbose=verbose)
# if verbose >= 2:
# print(theta0)
# print(loss)
# loss.backward()
# optimizer.step()
# mu = theta0[:self.dim].detach()
# cov = (utils.softplus(theta0[self.dim:])).detach() #covariance
# if verbose >= 1:
# print(loss)
# return mu,cov
# -
distrib = bvbq_torch.distributions.MixtureDiagonalNormalDistribution(mixmeans,mixvars,mixweights)
xplot = np.linspace(-20,20,201).reshape(-1,1)
yplot1 = logprob_mixgaussians(xplot).detach().numpy()
yplot2 = logprobgp.predict(xplot,return_cov=False).detach().numpy()
yplot3 = distrib.logprob(xplot).detach().numpy().reshape(-1,1)
plt.plot(xplot,np.exp(yplot1),'b')
plt.plot(xplot,np.exp(yplot2),'r')
plt.plot(xplot,np.exp(yplot3),'g')
distrib.logprob(xplot).detach().numpy().reshape(-1,1)
|
toybox/old/bvbq_torch_test_bvbq_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Biblioteca para trabalho com grafos
# +
#criar um grafo
import networkx as nx
G = nx.Graph()
#criar os nodos
#um a um
G.add_node(1)
#lista de nodos
G.add_nodes_from([2, 3])
#cria as arestas
#uma a uma
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e)
#uma lista
G.add_edges_from([(1, 2), (1, 3)])
print(G.number_of_nodes())
print(G.number_of_edges())
print(list(G.nodes))
print(list(G.edges))
G.remove_node(2)
print(list(G.nodes))
print(list(G.edges))
# -
|
.ipynb_checkpoints/pytrigo-9-networkx-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="uoJsVjtCMunI"
# <table align="center">
# <td align="center"><a target="_blank" href="http://introtodeeplearning.com">
# <img src="http://introtodeeplearning.com/images/colab/mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="https://colab.research.google.com/github/aamini/introtodeeplearning/blob/master/lab1/Part2_Music_Generation.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/colab.png?v2.0" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/aamini/introtodeeplearning/blob/master/lab1/Part2_Music_Generation.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/github.png" height="70px" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# </table>
#
# # Copyright Information
# + colab={} colab_type="code" id="bUik05YqMyCH"
# Copyright 2020 MIT 6.S191 Introduction to Deep Learning. All Rights Reserved.
#
# Licensed under the MIT License. You may not use this file except in compliance
# with the License. Use and/or modification of this code outside of 6.S191 must
# reference:
#
# © MIT 6.S191: Introduction to Deep Learning
# http://introtodeeplearning.com
#
# + [markdown] colab_type="text" id="O-97SDET3JG-"
# # Lab 1: Intro to TensorFlow and Music Generation with RNNs
#
# # Part 2: Music Generation with RNNs
#
# In this portion of the lab, we will explore building a Recurrent Neural Network (RNN) for music generation. We will train a model to learn the patterns in raw sheet music in [ABC notation](https://en.wikipedia.org/wiki/ABC_notation) and then use this model to generate new music.
# + [markdown] colab_type="text" id="rsvlBQYCrE4I"
# ## 2.1 Dependencies
# First, let's download the course repository, install dependencies, and import the relevant packages we'll need for this lab.
# + colab={"base_uri": "https://localhost:8080/", "height": 344} colab_type="code" id="riVZCVK65QTH" outputId="68f7b646-3b10-48b1-a508-6fc36b1ac0e4"
# Import Tensorflow 2.0
# #%tensorflow_version 2.x
import tensorflow as tf
# Download and import the MIT 6.S191 package
# #!pip install mitdeeplearning
import mitdeeplearning as mdl
# Import all remaining packages
import numpy as np
import os
import time
import functools
from IPython import display as ipythondisplay
from tqdm import tqdm
# #!apt-get install abcmidi timidity > /dev/null 2>&1
# Check that we are using a GPU, if not switch runtimes
# using Runtime > Change Runtime Type > GPU
assert len(tf.config.list_physical_devices('GPU')) > 0
# + [markdown] colab_type="text" id="_ajvp0No4qDm"
# ## 2.2 Dataset
#
# 
#
# We've gathered a dataset of thousands of Irish folk songs, represented in the ABC notation. Let's download the dataset and inspect it:
#
# + colab={"base_uri": "https://localhost:8080/", "height": 253} colab_type="code" id="P7dFnP5q3Jve" outputId="a89aa492-8039-4620-917c-a16f77e9c73c"
# Download the dataset
songs = mdl.lab1.load_training_data()
# Print one of the songs to inspect it in greater detail!
example_song = songs[0]
print("\nExample song: ")
print(example_song)
# + [markdown] colab_type="text" id="hKF3EHJlCAj2"
# We can easily convert a song in ABC notation to an audio waveform and play it back. Be patient for this conversion to run, it can take some time.
# + colab={"base_uri": "https://localhost:8080/", "height": 60} colab_type="code" id="11toYzhEEKDz" outputId="c254efd1-dae4-4198-cd6d-5420eb318755"
# Convert the ABC notation to audio file and listen to it
mdl.lab1.play_song(example_song)
# + [markdown] colab_type="text" id="7vH24yyquwKQ"
# One important thing to think about is that this notation of music does not simply contain information on the notes being played, but additionally there is meta information such as the song title, key, and tempo. How does the number of different characters that are present in the text file impact the complexity of the learning problem? This will become important soon, when we generate a numerical representation for the text data.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="IlCgQBRVymwR" outputId="0333c209-8da9-44ba-b2bd-b1adb05bb67a"
# Join our list of song strings into a single string containing all songs
songs_joined = "\n\n".join(songs)
# Find all unique characters in the joined string
vocab = sorted(set(songs_joined))
print("There are", len(vocab), "unique characters in the dataset")
# + [markdown] colab_type="text" id="rNnrKn_lL-IJ"
# ## 2.3 Process the dataset for the learning task
#
# Let's take a step back and consider our prediction task. We're trying to train a RNN model to learn patterns in ABC music, and then use this model to generate (i.e., predict) a new piece of music based on this learned information.
#
# Breaking this down, what we're really asking the model is: given a character, or a sequence of characters, what is the most probable next character? We'll train the model to perform this task.
#
# To achieve this, we will input a sequence of characters to the model, and train the model to predict the output, that is, the following character at each time step. RNNs maintain an internal state that depends on previously seen elements, so information about all characters seen up until a given moment will be taken into account in generating the prediction.
# + [markdown] colab_type="text" id="LFjSVAlWzf-N"
# ### Vectorize the text
#
# Before we begin training our RNN model, we'll need to create a numerical representation of our text-based dataset. To do this, we'll generate two lookup tables: one that maps characters to numbers, and a second that maps numbers back to characters. Recall that we just identified the unique characters present in the text.
# + colab={} colab_type="code" id="IalZLbvOzf-F"
### Define numerical representation of text ###
# Create a mapping from character to unique index.
# For example, to get the index of the character "d",
# we can evaluate `char2idx["d"]`.
char2idx = {u:i for i, u in enumerate(vocab)}
# Create a mapping from indices to characters. This is
# the inverse of char2idx and allows us to convert back
# from unique index to the character in our vocabulary.
idx2char = np.array(vocab)
# + [markdown] colab_type="text" id="tZfqhkYCymwX"
# This gives us an integer representation for each character. Observe that the unique characters (i.e., our vocabulary) in the text are mapped as indices from 0 to `len(unique)`. Let's take a peek at this numerical representation of our dataset:
# + colab={"base_uri": "https://localhost:8080/", "height": 435} colab_type="code" id="FYyNlCNXymwY" outputId="5bae686f-02cb-432d-f12f-40a565c17b8f"
print('{')
for char,_ in zip(char2idx, range(20)):
print(' {:4s}: {:3d},'.format(repr(char), char2idx[char]))
print(' ...\n}')
# + colab={} colab_type="code" id="g-LnKyu4dczc"
### Vectorize the songs string ###
'''TODO: Write a function to convert the all songs string to a vectorized
(i.e., numeric) representation. Use the appropriate mapping
above to convert from vocab characters to the corresponding indices.
NOTE: the output of the `vectorize_string` function
should be a np.array with `N` elements, where `N` is
the number of characters in the input string
'''
def vectorize_string(string):
# TODO
return np.array([char2idx[s] for s in string])
vectorized_songs = vectorize_string(songs_joined)
# + [markdown] colab_type="text" id="IqxpSuZ1w-ub"
# We can also look at how the first part of the text is mapped to an integer representation:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="l1VKcQHcymwb" outputId="0a56bacc-f407-44eb-e4d4-a0a34cb5ae15"
print ('{} ---- characters mapped to int ----> {}'.format(repr(songs_joined[:10]), vectorized_songs[:10]))
# check that vectorized_songs is a numpy array
assert isinstance(vectorized_songs, np.ndarray), "returned result should be a numpy array"
# + [markdown] colab_type="text" id="hgsVvVxnymwf"
# ### Create training examples and targets
#
# Our next step is to actually divide the text into example sequences that we'll use during training. Each input sequence that we feed into our RNN will contain `seq_length` characters from the text. We'll also need to define a target sequence for each input sequence, which will be used in training the RNN to predict the next character. For each input, the corresponding target will contain the same length of text, except shifted one character to the right.
#
# To do this, we'll break the text into chunks of `seq_length+1`. Suppose `seq_length` is 4 and our text is "Hello". Then, our input sequence is "Hell" and the target sequence is "ello".
#
# The batch method will then let us convert this stream of character indices to sequences of the desired size.
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" id="LF-N8F7BoDRi" outputId="f949be0d-dee6-454b-cfed-2f1d39b9dd9d"
### Batch definition to create training examples ###
def get_batch(vectorized_songs, seq_length, batch_size):
# the length of the vectorized songs string
n = vectorized_songs.shape[0] - 1
# randomly choose the starting indices for the examples in the training batch
idx = np.random.choice(n-seq_length, batch_size)
'''TODO: construct a list of input sequences for the training batch'''
input_batch = [vectorized_songs[id:id+seq_length] for id in idx] # TODO
'''TODO: construct a list of output sequences for the training batch'''
output_batch = [vectorized_songs[id+1:id+1+seq_length] for id in idx] # TODO
# x_batch, y_batch provide the true inputs and targets for network training
x_batch = np.reshape(input_batch, [batch_size, seq_length])
y_batch = np.reshape(output_batch, [batch_size, seq_length])
return x_batch, y_batch
# Perform some simple tests to make sure your batch function is working properly!
test_args = (vectorized_songs, 10, 2)
if not mdl.lab1.test_batch_func_types(get_batch, test_args) or \
not mdl.lab1.test_batch_func_shapes(get_batch, test_args) or \
not mdl.lab1.test_batch_func_next_step(get_batch, test_args):
print("======\n[FAIL] could not pass tests")
else:
print("======\n[PASS] passed all tests!")
# + [markdown] colab_type="text" id="_33OHL3b84i0"
# For each of these vectors, each index is processed at a single time step. So, for the input at time step 0, the model receives the index for the first character in the sequence, and tries to predict the index of the next character. At the next timestep, it does the same thing, but the RNN considers the information from the previous step, i.e., its updated state, in addition to the current input.
#
# We can make this concrete by taking a look at how this works over the first several characters in our text:
# + colab={"base_uri": "https://localhost:8080/", "height": 290} colab_type="code" id="0eBu9WZG84i0" outputId="2516460c-8900-4ea5-beb8-eba1769a6f5c"
x_batch, y_batch = get_batch(vectorized_songs, seq_length=5, batch_size=1)
for i, (input_idx, target_idx) in enumerate(zip(np.squeeze(x_batch), np.squeeze(y_batch))):
print("Step {:3d}".format(i))
print(" input: {} ({:s})".format(input_idx, repr(idx2char[input_idx])))
print(" expected output: {} ({:s})".format(target_idx, repr(idx2char[target_idx])))
# + [markdown] colab_type="text" id="r6oUuElIMgVx"
# ## 2.4 The Recurrent Neural Network (RNN) model
# + [markdown] colab_type="text" id="m8gPwEjRzf-Z"
# Now we're ready to define and train a RNN model on our ABC music dataset, and then use that trained model to generate a new song. We'll train our RNN using batches of song snippets from our dataset, which we generated in the previous section.
#
# The model is based off the LSTM architecture, where we use a state vector to maintain information about the temporal relationships between consecutive characters. The final output of the LSTM is then fed into a fully connected [`Dense`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense) layer where we'll output a softmax over each character in the vocabulary, and then sample from this distribution to predict the next character.
#
# As we introduced in the first portion of this lab, we'll be using the Keras API, specifically, [`tf.keras.Sequential`](https://www.tensorflow.org/api_docs/python/tf/keras/models/Sequential), to define the model. Three layers are used to define the model:
#
# * [`tf.keras.layers.Embedding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Embedding): This is the input layer, consisting of a trainable lookup table that maps the numbers of each character to a vector with `embedding_dim` dimensions.
# * [`tf.keras.layers.LSTM`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/LSTM): Our LSTM network, with size `units=rnn_units`.
# * [`tf.keras.layers.Dense`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense): The output layer, with `vocab_size` outputs.
#
#
# <img src="https://raw.githubusercontent.com/aamini/introtodeeplearning/2019/lab1/img/lstm_unrolled-01-01.png" alt="Drawing"/>
# + [markdown] colab_type="text" id="rlaOqndqBmJo"
# ### Define the RNN model
#
# Now, we will define a function that we will use to actually build the model.
# + colab={} colab_type="code" id="8DsWzojvkbc7"
def LSTM(rnn_units):
return tf.keras.layers.LSTM(
rnn_units,
return_sequences=True,
recurrent_initializer='glorot_uniform',
recurrent_activation='sigmoid',
stateful=True,
)
# + [markdown] colab_type="text" id="IbWU4dMJmMvq"
# The time has come! Fill in the `TODOs` to define the RNN model within the `build_model` function, and then call the function you just defined to instantiate the model!
# + colab={} colab_type="code" id="MtCrdfzEI2N0"
### Defining the RNN Model ###
'''TODO: Add LSTM and Dense layers to define the RNN model using the Sequential API.'''
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = tf.keras.Sequential([
# Layer 1: Embedding layer to transform indices into dense vectors
# of a fixed embedding size
tf.keras.layers.Embedding(vocab_size, embedding_dim, batch_input_shape=[batch_size, None]),
# Layer 2: LSTM with `rnn_units` number of units.
# TODO: Call the LSTM function defined above to add this layer.
LSTM(rnn_units),
# Layer 3: Dense (fully-connected) layer that transforms the LSTM output
# into the vocabulary size.
# TODO: Add the Dense layer.
# '''TODO: DENSE LAYER HERE'''
tf.keras.layers.Dense(vocab_size)
])
return model
# Build a simple model with default hyperparameters. You will get the
# chance to change these later.
model = build_model(len(vocab), embedding_dim=256, rnn_units=1024, batch_size=32)
# + [markdown] colab_type="text" id="-ubPo0_9Prjb"
# ### Test out the RNN model
#
# It's always a good idea to run a few simple checks on our model to see that it behaves as expected.
#
# First, we can use the `Model.summary` function to print out a summary of our model's internal workings. Here we can check the layers in the model, the shape of the output of each of the layers, the batch size, etc.
# + colab={"base_uri": "https://localhost:8080/", "height": 271} colab_type="code" id="RwG1DD6rDrRM" outputId="76677ce8-a309-43fb-a3bb-f9ff0189fc0a"
model.summary()
# + [markdown] colab_type="text" id="8xeDn5nZD0LX"
# We can also quickly check the dimensionality of our output, using a sequence length of 100. Note that the model can be run on inputs of any length.
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="C-_70kKAPrPU" outputId="47aa155d-968d-4b67-c593-705be647450c"
x, y = get_batch(vectorized_songs, seq_length=100, batch_size=32)
pred = model(x)
print("Input shape: ", x.shape, " # (batch_size, sequence_length)")
print("Prediction shape: ", pred.shape, "# (batch_size, sequence_length, vocab_size)")
# + [markdown] colab_type="text" id="mT1HvFVUGpoE"
# ### Predictions from the untrained model
#
# Let's take a look at what our untrained model is predicting.
#
# To get actual predictions from the model, we sample from the output distribution, which is defined by a `softmax` over our character vocabulary. This will give us actual character indices. This means we are using a [categorical distribution](https://en.wikipedia.org/wiki/Categorical_distribution) to sample over the example prediction. This gives a prediction of the next character (specifically its index) at each timestep.
#
# Note here that we sample from this probability distribution, as opposed to simply taking the `argmax`, which can cause the model to get stuck in a loop.
#
# Let's try this sampling out for the first example in the batch.
# + colab={"base_uri": "https://localhost:8080/", "height": 126} colab_type="code" id="4V4MfFg0RQJg" outputId="d3181507-31ec-45bf-f83d-1d068450ecd0"
sampled_indices = tf.random.categorical(pred[0], num_samples=1)
sampled_indices = tf.squeeze(sampled_indices,axis=-1).numpy()
sampled_indices
# + [markdown] colab_type="text" id="LfLtsP3mUhCG"
# We can now decode these to see the text predicted by the untrained model:
# + colab={"base_uri": "https://localhost:8080/", "height": 108} colab_type="code" id="xWcFwPwLSo05" outputId="1ec8f353-1d8a-49e0-8baf-e543d17fa641"
print("Input: \n", repr("".join(idx2char[x[0]])))
print()
print("Next Char Predictions: \n", repr("".join(idx2char[sampled_indices])))
# + [markdown] colab_type="text" id="HEHHcRasIDm9"
# As you can see, the text predicted by the untrained model is pretty nonsensical! How can we do better? We can train the network!
# + [markdown] colab_type="text" id="LJL0Q0YPY6Ee"
# ## 2.5 Training the model: loss and training operations
#
# Now it's time to train the model!
#
# At this point, we can think of our next character prediction problem as a standard classification problem. Given the previous state of the RNN, as well as the input at a given time step, we want to predict the class of the next character -- that is, to actually predict the next character.
#
# To train our model on this classification task, we can use a form of the `crossentropy` loss (negative log likelihood loss). Specifically, we will use the [`sparse_categorical_crossentropy`](https://www.tensorflow.org/api_docs/python/tf/keras/losses/sparse_categorical_crossentropy) loss, as it utilizes integer targets for categorical classification tasks. We will want to compute the loss using the true targets -- the `labels` -- and the predicted targets -- the `logits`.
#
# Let's first compute the loss using our example predictions from the untrained model:
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="4HrXTACTdzY-" outputId="75b85a48-ea60-4326-e7d5-6b947e6c0756"
### Defining the loss function ###
'''TODO: define the loss function to compute and return the loss between
the true labels and predictions (logits). Set the argument from_logits=True.'''
def compute_loss(labels, logits):
loss = tf.keras.losses.sparse_categorical_crossentropy(labels, logits, from_logits=True) # TODO
return loss
'''TODO: compute the loss using the true next characters from the example batch
and the predictions from the untrained model several cells above'''
example_batch_loss = compute_loss(y, pred) # TODO
print("Prediction shape: ", pred.shape, " # (batch_size, sequence_length, vocab_size)")
print("scalar_loss: ", example_batch_loss.numpy().mean())
# + [markdown] colab_type="text" id="0Seh7e6eRqd7"
# Let's start by defining some hyperparameters for training the model. To start, we have provided some reasonable values for some of the parameters. It is up to you to use what we've learned in class to help optimize the parameter selection here!
# + colab={} colab_type="code" id="JQWUUhKotkAY"
### Hyperparameter setting and optimization ###
# Optimization parameters:
num_training_iterations = 2000 # Increase this to train longer
batch_size = 4 # Experiment between 1 and 64
seq_length = 100 # Experiment between 50 and 500
learning_rate = 5e-3 # Experiment between 1e-5 and 1e-1
# Model parameters:
vocab_size = len(vocab)
embedding_dim = 256
rnn_units = 1024 # Experiment between 1 and 2048
# Checkpoint location:
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "my_ckpt")
# + [markdown] colab_type="text" id="5cu11p1MKYZd"
# Now, we are ready to define our training operation -- the optimizer and duration of training -- and use this function to train the model. You will experiment with the choice of optimizer and the duration for which you train your models, and see how these changes affect the network's output. Some optimizers you may like to try are [`Adam`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adam?version=stable) and [`Adagrad`](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/Adagrad?version=stable).
#
# First, we will instantiate a new model and an optimizer. Then, we will use the [`tf.GradientTape`](https://www.tensorflow.org/api_docs/python/tf/GradientTape) method to perform the backpropagation operations.
#
# We will also generate a print-out of the model's progress through training, which will help us easily visualize whether or not we are minimizing the loss.
# + colab={"base_uri": "https://localhost:8080/", "height": 559} colab_type="code" id="F31vzJ_u66cb" outputId="c4995286-b68a-4738-f806-c7bb8b805c64"
### Define optimizer and training operation ###
'''TODO: instantiate a new model for training using the `build_model`
function and the hyperparameters created above.'''
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size)
'''TODO: instantiate an optimizer with its learning rate.
Checkout the tensorflow website for a list of supported optimizers.
https://www.tensorflow.org/api_docs/python/tf/keras/optimizers/
Try using the Adam optimizer to start.'''
optimizer = tf.keras.optimizers.Adam(
learning_rate=learning_rate, beta_1=0.9, beta_2=0.999, epsilon=1e-07, amsgrad=False,
name='Adam')
# TODO
@tf.function
def train_step(x, y):
# Use tf.GradientTape()
with tf.GradientTape() as tape:
'''TODO: feed the current input into the model and generate predictions'''
y_hat = model(x)
'''TODO: compute the loss!'''
loss = compute_loss(y, y_hat)
# Now, compute the gradients
'''TODO: complete the function call for gradient computation.
Remember that we want the gradient of the loss with respect all
of the model parameters.
HINT: use `model.trainable_variables` to get a list of all model
parameters.'''
grads = tape.gradient(loss, model.trainable_variables)
# Apply the gradients to the optimizer so it can update the model accordingly
optimizer.apply_gradients(zip(grads, model.trainable_variables))
return loss
##################
# Begin training!#
##################
history = []
plotter = mdl.util.PeriodicPlotter(sec=2, xlabel='Iterations', ylabel='Loss')
if hasattr(tqdm, '_instances'): tqdm._instances.clear() # clear if it exists
for iter in tqdm(range(num_training_iterations)):
# Grab a batch and propagate it through the network
x_batch, y_batch = get_batch(vectorized_songs, seq_length, batch_size)
loss = train_step(x_batch, y_batch)
# Update the progress bar
history.append(loss.numpy().mean())
plotter.plot(history)
# Update the model with the changed weights!
if iter % 100 == 0:
model.save_weights(checkpoint_prefix)
# Save the trained model and the weights
model.save_weights(checkpoint_prefix)
# + [markdown] colab_type="text" id="kKkD5M6eoSiN"
# ## 2.6 Generate music using the RNN model
#
# Now, we can use our trained RNN model to generate some music! When generating music, we'll have to feed the model some sort of seed to get it started (because it can't predict anything without something to start with!).
#
# Once we have a generated seed, we can then iteratively predict each successive character (remember, we are using the ABC representation for our music) using our trained RNN. More specifically, recall that our RNN outputs a `softmax` over possible successive characters. For inference, we iteratively sample from these distributions, and then use our samples to encode a generated song in the ABC format.
#
# Then, all we have to do is write it to a file and listen!
# + [markdown] colab_type="text" id="JIPcXllKjkdr"
# ### Restore the latest checkpoint
#
# To keep this inference step simple, we will use a batch size of 1. Because of how the RNN state is passed from timestep to timestep, the model will only be able to accept a fixed batch size once it is built.
#
# To run the model with a different `batch_size`, we'll need to rebuild the model and restore the weights from the latest checkpoint, i.e., the weights after the last checkpoint during training:
# + colab={"base_uri": "https://localhost:8080/", "height": 271} colab_type="code" id="LycQ-ot_jjyu" outputId="31da208e-e114-42ff-c329-1ea86a8d272d"
'''TODO: Rebuild the model using a batch_size=1'''
model = build_model(vocab_size, embedding_dim, rnn_units, batch_size=1)
# Restore the model weights for the last checkpoint after training
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
model.build(tf.TensorShape([1, None]))
model.summary()
# + [markdown] colab_type="text" id="I9b4V2C8N62l"
# Notice that we have fed in a fixed `batch_size` of 1 for inference.
# + [markdown] colab_type="text" id="DjGz1tDkzf-u"
# ### The prediction procedure
#
# Now, we're ready to write the code to generate text in the ABC music format:
#
# * Initialize a "seed" start string and the RNN state, and set the number of characters we want to generate.
#
# * Use the start string and the RNN state to obtain the probability distribution over the next predicted character.
#
# * Sample from multinomial distribution to calculate the index of the predicted character. This predicted character is then used as the next input to the model.
#
# * At each time step, the updated RNN state is fed back into the model, so that it now has more context in making the next prediction. After predicting the next character, the updated RNN states are again fed back into the model, which is how it learns sequence dependencies in the data, as it gets more information from the previous predictions.
#
# 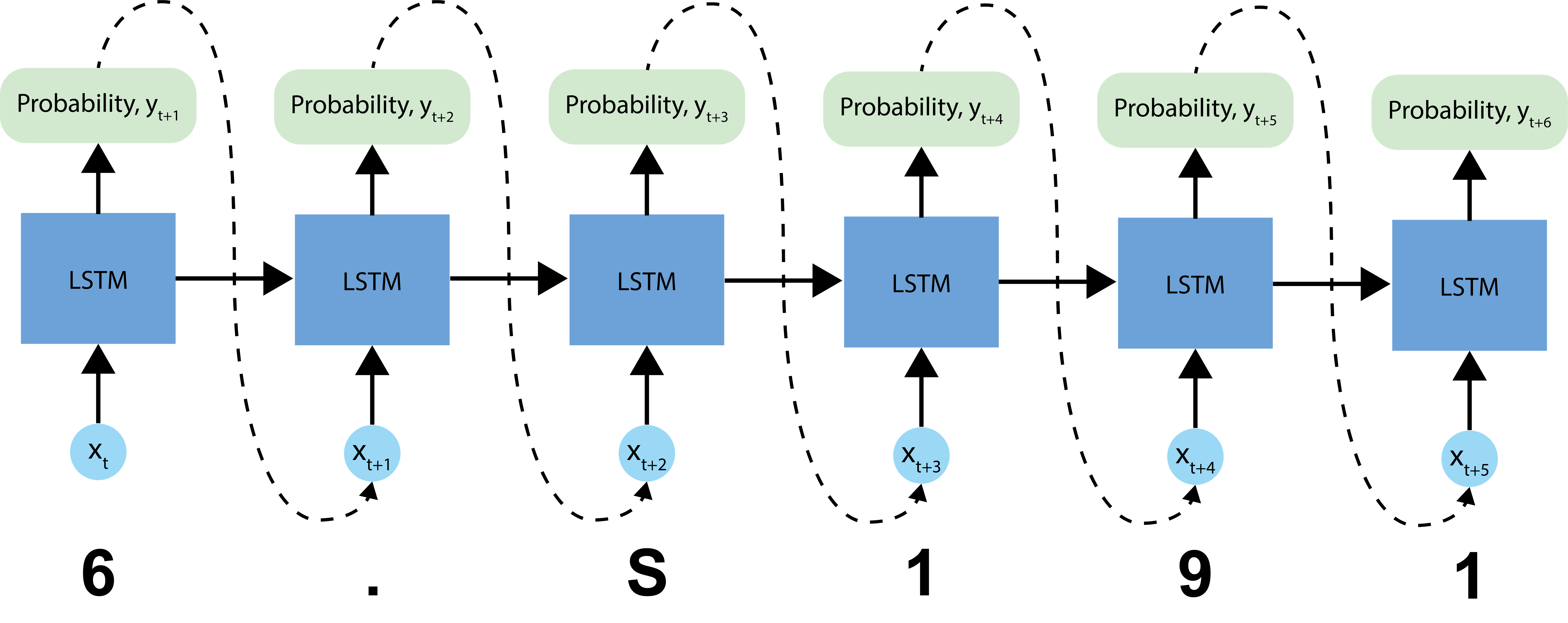
#
# Complete and experiment with this code block (as well as some of the aspects of network definition and training!), and see how the model performs. How do songs generated after training with a small number of epochs compare to those generated after a longer duration of training?
# + colab={} colab_type="code" id="WvuwZBX5Ogfd"
### Prediction of a generated song ###
def generate_text(model, start_string, generation_length=1000):
# Evaluation step (generating ABC text using the learned RNN model)
'''TODO: convert the start string to numbers (vectorize)'''
input_eval = [vectorize_string(start_string)]
input_eval = tf.expand_dims(input_eval, 0)
# Empty string to store our results
text_generated = []
# Here batch size == 1
model.reset_states()
tqdm._instances.clear()
for i in tqdm(range(generation_length)):
'''TODO: evaluate the inputs and generate the next character predictions'''
predictions = model(input_eval)
# Remove the batch dimension
predictions = tf.squeeze(predictions, 0)
'''TODO: use a multinomial distribution to sample'''
predicted_id = tf.random.categorical(predictions, num_samples=1)[-1,0].numpy()
# Pass the prediction along with the previous hidden state
# as the next inputs to the model
input_eval = tf.expand_dims([predicted_id], 0)
'''TODO: add the predicted character to the generated text!'''
# Hint: consider what format the prediction is in vs. the output
text_generated.append(idx2char[predicted_id])
return (start_string + ''.join(text_generated))
# + colab={"base_uri": "https://localhost:8080/", "height": 53} colab_type="code" id="ktovv0RFhrkn" outputId="5a1de778-2039-4c2c-8957-b5b7f4c8d0d1"
'''TODO: Use the model and the function defined above to generate ABC format text of length 1000!
As you may notice, ABC files start with "X" - this may be a good start string.'''
generated_text = generate_text(model, start_string="X", generation_length=1000) # TODO
# generated_text = generate_text('''TODO''', start_string="X", generation_length=1000)
# + [markdown] colab_type="text" id="AM2Uma_-yVIq"
# ### Play back the generated music!
#
# We can now call a function to convert the ABC format text to an audio file, and then play that back to check out our generated music! Try training longer if the resulting song is not long enough, or re-generating the song!
# + colab={} colab_type="code" id="LrOtG64bfLto"
### Play back generated songs ###
generated_songs = mdl.lab1.extract_song_snippet(generated_text)
print(generated_songs)
for i, song in enumerate(generated_songs):
# Synthesize the waveform from a song
waveform = mdl.lab1.play_song(song)
# If its a valid song (correct syntax), lets play it!
if waveform:
print("Generated song", i)
ipythondisplay.display(waveform)
# + [markdown] colab_type="text" id="HgVvcrYmSKGG"
# ## 2.7 Experiment and **get awarded for the best songs**!!
#
# Congrats on making your first sequence model in TensorFlow! It's a pretty big accomplishment, and hopefully you have some sweet tunes to show for it.
#
# If you want to go further, try to optimize your model and submit your best song! Tweet us at [@MITDeepLearning](https://twitter.com/MITDeepLearning) or [email us](mailto:<EMAIL>) a copy of the song (if you don't have Twitter), and we'll give out prizes to our favorites!
#
# Consider how you may improve your model and what seems to be most important in terms of performance. Here are some ideas to get you started:
#
# * How does the number of training epochs affect the performance?
# * What if you alter or augment the dataset?
# * Does the choice of start string significantly affect the result?
#
# Have fun and happy listening!
#
#
# 
#
#
#
# + colab={} colab_type="code" id="jX9zPJkAPI11"
# Example submission by a previous 6.S191 student (credit: <NAME>)
# %%html
<blockquote class="twitter-tweet"><a href="https://twitter.com/AdibChristian/status/1090030964770783238?ref_src=twsrc%5Etfw">January 28, 2019</a></blockquote>
<script async src="https://platform.twitter.com/widgets.js" charset="utf-8"></script>
# + colab={} colab_type="code" id="zFdnDAcV_Ml1"
|
lab1/Part2_Music_Generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# This notebook is part of the `nbsphinx` documentation: https://nbsphinx.readthedocs.io/.
# -
# # Code Cells
#
# ## Code, Output, Streams
#
# An empty code cell:
# Two empty lines:
# Leading/trailing empty lines:
# +
# 2 empty lines before, 1 after
# -
# A simple output:
6 * 7
# The standard output stream:
print('Hello, world!')
# Normal output + standard output
print('Hello, world!')
6 * 7
# The standard error stream is highlighted and displayed just below the code cell.
# The standard output stream comes afterwards (with no special highlighting).
# Finally, the "normal" output is displayed.
# +
import sys
print("I'll appear on the standard error stream", file=sys.stderr)
print("I'll appear on the standard output stream")
"I'm the 'normal' output"
# -
# <div class="alert alert-info">
#
# **Note:**
#
# Using the IPython kernel, the order is actually mixed up,
# see https://github.com/ipython/ipykernel/issues/280.
#
# </div>
# ## Cell Magics
#
# IPython can handle code in other languages by means of [cell magics](https://ipython.readthedocs.io/en/stable/interactive/magics.html#cell-magics):
# + language="bash"
# for i in 1 2 3
# do
# echo $i
# done
# -
# ## Special Display Formats
#
# See [IPython example notebook](https://nbviewer.jupyter.org/github/ipython/ipython/blob/master/examples/IPython Kernel/Rich Output.ipynb).
# ### Local Image Files
#
# See also [SVG support for LaTeX](markdown-cells.ipynb#SVG-support-for-LaTeX).
from IPython.display import Image
i = Image(filename='images/notebook_icon.png')
i
display(i)
# ### Image URLs
Image(url='https://www.python.org/static/img/python-logo-large.png')
Image(url='https://www.python.org/static/img/python-logo-large.png', embed=True)
Image(url='https://jupyter.org/assets/nav_logo.svg')
# ### Math
from IPython.display import Math
eq = Math(r'\int\limits_{-\infty}^\infty f(x) \delta(x - x_0) dx = f(x_0)')
eq
display(eq)
from IPython.display import Latex
Latex(r'This is a \LaTeX{} equation: $a^2 + b^2 = c^2$')
# + language="latex"
# \begin{equation}
# \int\limits_{-\infty}^\infty f(x) \delta(x - x_0) dx = f(x_0)
# \end{equation}
# -
# ### Plots
#
# The output formats for Matplotlib plots can be customized.
# You'll need separate settings for the Jupyter Notebook application and for `nbsphinx`.
#
# If you want to use SVG images for Matplotlib plots,
# add this line to your IPython configuration file:
#
# ```python
# c.InlineBackend.figure_formats = {'svg'}
# ```
#
# If you want SVG images, but also want nice plots when exporting to LaTeX/PDF, you can select:
#
# ```python
# c.InlineBackend.figure_formats = {'svg', 'pdf'}
# ```
#
# If you want to use the default PNG plots or HiDPI plots using `'png2x'` (a.k.a. `'retina'`),
# make sure to set this:
#
# ```python
# c.InlineBackend.rc = {'figure.dpi': 96}
# ```
#
# This is needed because the default `'figure.dpi'` value of 72
# is only valid for the [Qt Console](https://qtconsole.readthedocs.io/).
#
# If you are planning to store your SVG plots as part of your notebooks,
# you should also have a look at the `'svg.hashsalt'` setting.
#
# For more details on these and other settings, have a look at
# [Default Values for Matplotlib's "inline" Backend](https://nbviewer.jupyter.org/github/mgeier/python-audio/blob/master/plotting/matplotlib-inline-defaults.ipynb).
#
# The configuration file `ipython_kernel_config.py` can be either
# in the directory where your notebook is located
# (see the [ipython_kernel_config.py](ipython_kernel_config.py) in this directory),
# or in your profile directory
# (typically `~/.ipython/profile_default/ipython_kernel_config.py`).
# To find out your IPython profile directory, use this command:
#
# python3 -m IPython profile locate
#
# A local `ipython_kernel_config.py` in the notebook directory
# also works on https://mybinder.org/.
# Alternatively, you can create a file with those settings in a file named
# `.ipython/profile_default/ipython_kernel_config.py` in your repository.
#
# To get SVG and PDF plots for `nbsphinx`,
# use something like this in your `conf.py` file:
#
# ```python
# nbsphinx_execute_arguments = [
# "--InlineBackend.figure_formats={'svg', 'pdf'}",
# "--InlineBackend.rc={'figure.dpi': 96}",
# ]
# ```
#
# In the following example, `nbsphinx` should use an SVG image in the HTML output
# and a PDF image for LaTeX/PDF output.
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=[6, 3])
ax.plot([4, 9, 7, 20, 6, 33, 13, 23, 16, 62, 8]);
# Alternatively, the figure format(s) can also be chosen directly in the notebook
# (which overrides the setting in `nbsphinx_execute_arguments` and in the IPython configuration):
# %config InlineBackend.figure_formats = ['png']
fig
# If you want to use PNG images, but with HiDPI resolution,
# use the special `'png2x'` (a.k.a. `'retina'`) format
# (which also looks nice in the LaTeX output):
# %config InlineBackend.figure_formats = ['png2x']
fig
# ### Pandas Dataframes
#
# [Pandas dataframes](https://pandas.pydata.org/pandas-docs/stable/getting_started/dsintro.html#dataframe)
# should be displayed as nicely formatted HTML tables (if you are using HTML output).
import numpy as np
import pandas as pd
df = pd.DataFrame(np.random.randint(0, 100, size=[5, 4]),
columns=['a', 'b', 'c', 'd'])
df
# For LaTeX output, however, the plain text output is used by default.
#
# To get nice LaTeX tables, a few settings have to be changed:
pd.set_option('display.latex.repr', True)
# This is not enabled by default because of
# [Pandas issue #12182](https://github.com/pandas-dev/pandas/issues/12182).
#
# The generated LaTeX tables utilize the `booktabs` package, so you have to make sure that package is [loaded in the preamble](https://www.sphinx-doc.org/en/master/latex.html) with:
#
# \usepackage{booktabs}
#
# In order to allow page breaks within tables, you should use:
pd.set_option('display.latex.longtable', True)
# The `longtable` package is already used by Sphinx,
# so you don't have to manually load it in the preamble.
#
# Finally, if you want to use LaTeX math expressions in your dataframe, you'll have to disable escaping:
pd.set_option('display.latex.escape', False)
# The above settings should have no influence on the HTML output, but the LaTeX output should now look nicer:
df = pd.DataFrame(np.random.randint(0, 100, size=[10, 4]),
columns=[r'$\alpha$', r'$\beta$', r'$\gamma$', r'$\delta$'])
df
# ### YouTube Videos
from IPython.display import YouTubeVideo
YouTubeVideo('WAikxUGbomY')
# ### Arbitrary JavaScript Output (HTML only)
# + language="javascript"
#
# var text = document.createTextNode("Hello, I was generated with JavaScript!");
# // Content appended to "element" will be visible in the output area:
# element.appendChild(text);
# -
# ### Unsupported Output Types
#
# If a code cell produces data with an unsupported MIME type, the Jupyter Notebook doesn't generate any output.
# `nbsphinx`, however, shows a warning message.
display({
'text/x-python': 'print("Hello, world!")',
'text/x-haskell': 'main = putStrLn "Hello, world!"',
}, raw=True)
# ## ANSI Colors
#
# The standard output and standard error streams may contain [ANSI escape sequences](https://en.wikipedia.org/wiki/ANSI_escape_code) to change the text and background colors.
print('BEWARE: \x1b[1;33;41mugly colors\x1b[m!', file=sys.stderr)
print('AB\x1b[43mCD\x1b[35mEF\x1b[1mGH\x1b[4mIJ\x1b[7m'
'KL\x1b[49mMN\x1b[39mOP\x1b[22mQR\x1b[24mST\x1b[27mUV')
# The following code showing the 8 basic ANSI colors is based on http://tldp.org/HOWTO/Bash-Prompt-HOWTO/x329.html.
# Each of the 8 colors has an "intense" variation, which is used for bold text.
# +
text = ' XYZ '
formatstring = '\x1b[{}m' + text + '\x1b[m'
print(' ' * 6 + ' ' * len(text) +
''.join('{:^{}}'.format(bg, len(text)) for bg in range(40, 48)))
for fg in range(30, 38):
for bold in False, True:
fg_code = ('1;' if bold else '') + str(fg)
print(' {:>4} '.format(fg_code) + formatstring.format(fg_code) +
''.join(formatstring.format(fg_code + ';' + str(bg))
for bg in range(40, 48)))
# -
# ANSI also supports a set of 256 indexed colors.
# The following code showing all of them is based on [http://bitmote.com/index.php?post/2012/11/19/Using-ANSI-Color-Codes-to-Colorize-Your-Bash-Prompt-on-Linux](https://web.archive.org/web/20190109005413/http://bitmote.com/index.php?post/2012/11/19/Using-ANSI-Color-Codes-to-Colorize-Your-Bash-Prompt-on-Linux).
# +
formatstring = '\x1b[38;5;{0};48;5;{0}mX\x1b[1mX\x1b[m'
print(' + ' + ''.join('{:2}'.format(i) for i in range(36)))
print(' 0 ' + ''.join(formatstring.format(i) for i in range(16)))
for i in range(7):
i = i * 36 + 16
print('{:3} '.format(i) + ''.join(formatstring.format(i + j)
for j in range(36) if i + j < 256))
# -
# You can even use 24-bit RGB colors:
# +
start = 255, 0, 0
end = 0, 0, 255
length = 79
out = []
for i in range(length):
rgb = [start[c] + int(i * (end[c] - start[c]) / length) for c in range(3)]
out.append('\x1b['
'38;2;{rgb[2]};{rgb[1]};{rgb[0]};'
'48;2;{rgb[0]};{rgb[1]};{rgb[2]}mX\x1b[m'.format(rgb=rgb))
print(''.join(out))
|
doc/code-cells.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
print(__doc__)
# Code source: <NAME>
# Modified for documentation by <NAME>
# License: BSD 3 clause
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from sklearn import datasets
from sklearn.decomposition import PCA
# %matplotlib inline
# import some data to play with
iris = datasets.load_iris()
X = iris.data[:, :2] # we only take the first two features.
Y = iris.target
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
plt.figure(2, figsize=(8, 6))
plt.clf()
# Plot the training points
plt.scatter(X[:, 0], X[:, 1], c=Y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
# To getter a better understanding of interaction of the dimensions
# plot the first three PCA dimensions
fig = plt.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=110)
X_reduced = PCA(n_components=3).fit_transform(iris.data)
ax.scatter(X_reduced[:, 0], X_reduced[:, 1], X_reduced[:, 2], c=Y,
cmap=plt.cm.Paired)
ax.set_title("First three PCA directions")
ax.set_xlabel("1st eigenvector")
ax.w_xaxis.set_ticklabels([])
ax.set_ylabel("2nd eigenvector")
ax.w_yaxis.set_ticklabels([])
ax.set_zlabel("3rd eigenvector")
ax.w_zaxis.set_ticklabels([])
plt.show()
|
Algorithm_Physic/M2_PTSC_17/.ipynb_checkpoints/Iris-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="r0WpWpygRnTv"
# # Instruções
#
# - IMPORTANTE: primeiro clique no círculo preto com a seta branca abaixo.
#
# - Faça as atividades em sequência, escrevendo ou alterando o trecho de código em cada célula para obter o resultado esperado.
#
# - Use **SHIFT + ENTER** para executar o código. O resultado será exibido logo abaixo.
#
# - Então valide o resultado com **SHIFT + ENTER** novamente, na célula que contém o comando **validar()**. Não remova ou altere esta linha.
#
# - Você pode desfazer a alteração em qualquer célula usando **CONTROL + Z**.
# + id="GoSb_6yyujWG"
# Não se preocupe com este código. E nem faça alterações nele, por favor.
import builtins
import io
from IPython.core.display import display, HTML
# practice core v2.2
_INPUT = []
_PRINT = []
def input(*args, **kwargs):
s = builtins.input(*args, **kwargs)
_INPUT.append(s)
return s
def print(*args, **kwargs):
s = _sprint(*args, **kwargs)
_PRINT.append(s)
return builtins.print(s)
def _sprint(*args, end='', **kwargs):
sio = io.StringIO()
builtins.print(*args, **kwargs, end=end, file=sio)
return sio.getvalue()
def _correct(msg='Correto!'):
display(HTML('<p style="background-color: LawnGreen; padding: 8px 8px 8px 8px;"><b>{}</b></p>'.format(msg)))
def _incorrect(msg):
display(HTML('<p style="background-color: Tomato; padding: 8px 8px 8px 8px;">{}</p>'.format(msg)))
def _hint(msg):
display(HTML('<p style="background-color: LightBlue; padding: 8px 8px 8px 8px;">{}</p>'.format(msg)))
def _out(value): # expects a single value to be matched against the last cell result
if repr(_) == repr(value):
return 0
if _ == '':
_incorrect('Não houve resultado, verifique se você executou seu código com SHIFT+ENTER.')
return 1
if type(_) != type(value):
if type(value) == int:
_hint('Dica: o resultado precisa ser do tipo inteiro.')
elif type(value) == float:
_hint('Dica: o resultado precisa ser do tipo real.')
elif type(value) == bool:
_hint('Dica: o resultado precisa ser do tipo lógico.')
elif type(value) == str:
_hint('Dica: o resultado precisa ser do tipo string.')
else:
_hint('Dica: o resultado precisa ser de outro tipo.')
_incorrect('Resultado incorreto, tente novamente.')
return 1
def _src(terms): # expects a list of strings to be matched against the last cell executed
result = 0
for term in terms:
if term + ' ' not in _i:
_hint('Dica: verifique se você usou "{}" no seu código.'.format(term))
result = 1
return result
def _inp(lines): # expects a list of strings to be matched against the read input
if repr(_INPUT) == repr(lines):
return 0
if len(_INPUT) == 0:
_incorrect('Não houve entrada, verifique se você usou o comando "input()" no seu código.')
return 1
if len(_INPUT) != len(lines):
if len(lines) == 1:
_hint('Dica: é esperada 1 entrada do usuário.')
else:
_hint('Dica: são esperadas {} entradas do usuário.'.format(len(lines)))
else:
_incorrect('Entrada com dados diferentes do que foi solicitado, tente novamente.')
return 1
def _pri(lines): # expects a list of strings to be matched against the printed output
if repr(_PRINT) == repr(lines):
return 0
if len(_PRINT) == 0:
_incorrect('Não houve saída, verifique se você usou o comando "print()" em seu código.')
return 1
if len(_PRINT) != len(lines):
if len(lines) == 1:
_hint('Dica: é esperada 1 linha de saída.')
else:
_hint('Dica: são esperadas {} linhas de saída.'.format(len(lines)))
else:
_incorrect('Saída incorreta, tente novamente.')
return 1
def _var(variables): # expects a dictionary of variable names and their respective values
result = 0
for name, value in variables.items():
if name not in globals():
_hint('Dica: a variável "{}" precisa ser declarada.'.format(name))
result = 1
continue
if repr(globals()[name]) == repr(value):
continue # OK by now
if type(globals()[name]) != type(value):
if type(value) == int:
_hint('Dica: a variável "{}" precisa ser do tipo inteiro.'.format(name))
elif type(value) == float:
_hint('Dica: a variável "{}" precisa ser do tipo real.'.format(name))
elif type(value) == bool:
_hint('Dica: a variável "{}" precisa ser do tipo lógico.'.format(name))
elif type(value) == str:
_hint('Dica: a variável "{}" precisa ser do tipo string.'.format(name))
else:
_hint('Dica: a variável "{}" precisa ser de outro tipo.'.format(name))
else:
_incorrect('Resultado incorreto para a variável "{}".'.format(name))
result = 1
return result
def _typ(variables): # expects a dictionary of variable names and their respective types
result = 0
for name, typ in variables.items():
if name not in globals():
_hint('Dica: a variável "{}" precisa ser declarada.'.format(name))
result = 1
continue
if type(globals()[name]) == typ:
continue # OK by now
if typ == int:
_hint('Dica: a variável "{}" precisa ser do tipo inteiro.'.format(name))
elif typ == float:
_hint('Dica: a variável "{}" precisa ser do tipo real.'.format(name))
elif typ == bool:
_hint('Dica: a variável "{}" precisa ser do tipo lógico.'.format(name))
elif typ == str:
_hint('Dica: a variável "{}" precisa ser do tipo string.'.format(name))
else:
_hint('Dica: a variável "{}" precisa ser de outro tipo.'.format(name))
result = 1
return result
def _clear():
global _INPUT, _PRINT
_INPUT = []
_PRINT = []
for name in set(globals()) - _GLOBALS:
if not name.startswith('_'):
#builtins.print('remove:', name)
globals().pop(name)
def validar(n=None):
if n == None:
_incorrect('É preciso indicar o número da atividade. Volte o comando "validar" como estava antes!')
elif _test(n) == 0:
_correct()
_clear()
_GLOBALS = set(globals())
_GLOBALS.add('_GLOBALS')
_GLOBALS.add('_test')
# + id="2mi_jwCBZSVh"
def _test(n):
if n == 1: return _typ({'c': int}) + _pri(['1', '2', '3'])
elif n == 2: return _typ({'c': int}) + _pri(['1', '2', '3'])
elif n == 3: return _typ({'c': int}) + _pri(['0', '1', '2'])
elif n == 4: return _typ({'c': int}) + _pri(['5', '4', '3'])
elif n == 5: return _typ({'c': int}) + _pri(['7', '8', '9'])
elif n == 6: return _typ({'c': int}) + _pri(['c agora é 2', 'c agora é 4', 'c agora é 6', 'c agora é 8'])
elif n == 7: return _typ({'c': int}) + _pri(['1 * 5 = 5', '2 * 5 = 10', '3 * 5 = 15', '4 * 5 = 20', '5 * 5 = 25'])
elif n == 8: return _typ({'c': int}) + _pri(['1', '2', '3', '4', '3', '2', '1'])
elif n == 9: return _typ({'a': int, 'c': int}) + _pri(['5050'])
elif n == 10: return _typ({'soma': int, 'c': int}) + _pri(['1683']) + _src(['if', '%'])
elif n == 11: return _typ({'soma': int, 'c': int}) + _pri(['1683'])
elif n == 12: return _typ({'quantos': int}) + _pri(['71'])
elif n == 13: return _typ({'n': int}) + _pri(['1', '2', '3', '4'])
elif n == 14: return _typ({'n': int}) + _pri(['-5', '-4', '-3', '-2', '-1'])
elif n == 15: return _var({'a': 21, 'b': 18}) + _inp(['21', '18']) + _pri(['21', '20', '19', '18'])
elif n == 16: return _typ({'v': int}) + _inp(['8', '3', '2', '0']) + _pri(['8', '2', '0'])
elif n == 17: return _typ({'v': int, 'produto': int}) + _inp(['8', '-4', '2', '0']) + _pri(['-64'])
elif n == 18: return _typ({'v1': int, 'v2': int}) + _inp(['3', '5', '7', '2', '1', '1']) + _pri(['v1 < v2', 'v1 > v2'])
elif n == 19: return _pri([' 1',' 2',' 4',' 8',' 16',' 32',' 64',' 128',' 256',' 512','1024'])
elif n == 20: return _inp(['240', '15', '27', '0']) + _pri(['36.00', '64.80', 'fim'])
elif n == 21: return _typ({'x': int, 'y': int, 'z': int}) + _inp(['42', '38']) + _pri(['39', '40', '41'])
elif n == 22: return _typ({'maior': int}) + _inp(['3', '5', '4', '7', '1']) + _pri(['7'])
elif n == 23: return _typ({'n': int}) + _inp(['50']) + _pri(['1275', '42925', '25.5'])
elif n == 24: return _typ({'m': int, 'n': int}) + _inp(['14', '21']) + _pri(['294'])
elif n == 25: return _inp(['9.3', '1.7', '4.1', '2.3', '5.6', '0']) + _pri(['1.7'])
# + [markdown] id="2nj3eRk6Q1_n"
# # Repetição contada
# + [markdown] id="oglGzfYvQhV6"
# ## Prática 1
# Busque e corrija o erro de sintaxe no código para obter a seguinte saída:
# ```
# 1
# 2
# 3
# ```
# + id="0n0wf32_QhV9"
c = 1
while c <= 3
print(c)
c = c + 1
# + id="-lpiUZikQhWH"
validar(1)
# + [markdown] id="GiWgucBqQz7O"
# ## Prática 2
# Busque e corrija o erro de sintaxe no código para obter a seguinte saída:
# ```
# 1
# 2
# 3
# ```
# + id="BxAeYwOSQ1Zu"
while c <= 3:
print(c)
c = c + 1
# + id="pVmNnTTAQ1yQ"
validar(2)
# + [markdown] id="9pWnRIliQhWP"
# ## Prática 3
# Conserte o erro de lógica no código para obter a seguinte saída:
# ```
# 0
# 1
# 2
# ```
# + id="_AjYoA3KQhWR"
c = 0
while c >= 2:
print(c)
c = c + 1
# + id="i8JjgD4xQhWZ"
validar(3)
# + [markdown] id="L3Wo_gvCQhWv"
# ## Prática 4
# Conserte o erro de lógica no código para obter a seguinte saída:
# ```
# 5
# 4
# 3
# ```
# + id="tXq-tLemQhWx"
c = 5
while c <= 3:
print(c)
c = c - 1
# + id="UvWTrUuGQhW4"
validar(4)
# + [markdown] id="ahLYEpjZQhW_"
# ## Prática 5
# Conserte o erro de lógica no código para obter a seguinte saída:
# ```
# 7
# 8
# 9
# ```
# + id="p8MO-Y8HQhXA"
c = 7
while c <= 9:
c = c + 1
print(c)
# + id="t46KlnZuQhXG"
validar(5)
# + [markdown] id="KfwoGIw2QhXO"
# ## Prática 6
# Escreva um trecho de código que defina a variável **c** como contador. Use ela para gerar a seguinte saída:
# ```
# c agora é 2
# c agora é 4
# c agora é 6
# c agora é 8
# ```
# + id="Q1zFJrKoQhXP"
# + id="TL4wNK6MQhXV"
validar(6)
# + [markdown] id="KgCBAclYVLjm"
# ## Prática 7
# Escreva um trecho de código que defina a variável **c** como contador. Use ela para mostrar a seguinte saída:
# ```
# 1 * 5 = 5
# 2 * 5 = 10
# 3 * 5 = 15
# 4 * 5 = 20
# 5 * 5 = 25
# ```
# + id="BKysIQddVLjs"
# + id="0YMMPya_VLj9"
validar(7)
# + [markdown] id="QyfeRxAzVMoA"
# ## Prática 8
# Escreva um trecho de código que defina a variável **c** como contador. Gere a seguinte saída, usando dois laços em sequência:
# ```
# 1
# 2
# 3
# 4
# 3
# 2
# 1
# ```
# + id="GYIR7SZ4VMoC"
# + id="342SY256VMoK"
validar(8)
# + [markdown] id="80Pcdw4oV-O6"
# ## Prática 9
# Escreva um trecho de código que defina a variável **a** como acumulador e **c** como contador. Use ambas para calcular a soma dos inteiros de 0 a 100, que deve ser impressa apenas no final.
# + id="ROdXECfQV_FN"
# + id="WiZwS0ztV_FT"
validar(9)
# + [markdown] id="GkSguw6zV-_u"
# ## Prática 10
# Escreva um trecho de código que defina a variável **soma** como acumulador e **c** como contador. Use ambas para calcular e imprimir ao final a soma dos múltiplos de 3 entre 3 e 99. Use um **if** e o operador **%** para testar se **c** é múltiplo de 3.
# + id="chLXVFlYV_AG"
# + id="aizuDbb4V_AM"
validar(10)
# + [markdown] id="MEb6FN-LV-58"
# ## Prática 11
# Escreva um trecho de código que defina a variável **soma** como acumulador e **c** como contador. Use ambas para calcular e imprimir ao final a soma da sequência 3, 6, 9, 12, ... 99. Não use um **if**, mas sim um passo maior para **c**.
# + id="oTZAIW-dV-5-"
# + id="fLJJkYnHV-6D"
validar(11)
# + [markdown] id="RO2lEnx0Y9Mw"
# ## Prática 12
# Escreva um trecho de código que conte na variável **quantos** quantos são os números múltiplos de 7 que também são pares no intervalo de 1 a 1000. Imprima esta quantidade ao final.
# + id="oFUhGuT7V-2F"
# + id="vBhnKBZcV-2L"
validar(12)
# + [markdown] id="B5u2q_QlV-0M"
# ## Prática 13
# Escreva um trecho de código que solicite um número **n**. Apenas se **n** for maior do que zero, imprima os números de 1 a **n** (inclusive). Teste com a entrada 4.
#
# + id="5J1Kqp_iV-0N"
# + id="zoaZoITvV-0W"
validar(13)
# + [markdown] id="5bMIbnh2V-t8"
# ## Prática 14
# Escreva um trecho de código que solicite um número inteiro negativo **n**. Se o usuário digitar 0 ou maior, avise imprimindo uma mensagem de erro. Caso contrário, imprima os números entre **n** e -1 (inclusive). Teste com a entrada -5.
#
# + id="HGBjsBi0V-t9"
# + id="ZVmCWzjmV-uH"
validar(14)
# + [markdown] id="3eojGRIRV-oq"
# ## Prática 15
# Escreva um trecho de código que leia dois número inteiros **a** e **b**. Se **a** for menor ou igual a **b** imprima os valores do primeiro até o segundo (incluindo ambos) em ordem crescente. Caso contrário imprima os valores de **a** até **b** em ordem decrescente. Teste com 21 e 18, nesta ordem.
# + id="HdJRowt8V-os"
# + id="aJd7LMVIV-oy"
validar(15)
# + [markdown] id="gUkGYdtVjQSB"
# # Repetição até terminar
# + [markdown] id="NBwYsRYfb8Ny"
# ## Prática 16
# Faça um programa que repita até encontrar o valor zero, sempre lendo um valor inteiro para a variável **v** dentro do laço de repetição. Se o valor lido for par, imprima também ele na saída. Teste com a sequência 8, 3, 2, 0.
#
# + id="w9iGThEDV-lM"
# + id="2vzqkenbV-lU"
validar(16)
# + [markdown] id="15h8guiO8SjC"
# ## Prática 17
# Faça um programa que repita lendo valores inteiros, colocados na variável **v**. Ele deve parar ao encontrar um valor zero. Use um acumulador **produto** para calcular o produto de todos os valores digitados (menos o último valor zero). Apresente o produto ao final. Teste com a sequência 8, -4, 2, 0.
#
# + id="jDwkjxsW8TIr"
# + id="CIpMJWJs8Tkb"
validar(17)
# + [markdown] id="pXkqWtELcXYS"
# ## Prática 18
# Faça um programa que repita lendo dois valores inteiros **v1** e **v2** de cada vez. Ele deve parar ao encontrar valores iguais em **v1** e **v2**. Dentro do laço, apresente a mensagem **v1 < v2** ou **v1 > v2** conforme o caso. Teste com as entradas 3, 5, 7, 2, 1, 1.
#
# + id="hLJzVcKjV-gf"
# + id="5p_qyf7_V-gp"
validar(18)
# + [markdown] id="RoDTlruV0-x6"
# # Formatação de números
# + [markdown] id="nqHUpJ27dHWW"
# ## Prática 19
# Escreva um programa que imprime as potências inteiras de dois, indo de 1 até 1024. Imprima a saída alinhada à direita e com preenchida com espaços, usando para isso a função **format** dentro do comando **print()**. Dica: use um acumulador que é sempre multiplicado por 2.
# + id="JLHhnxEUV-PC"
# + id="Eu7NE5UnV-PR"
validar(19)
# + [markdown] id="IxXJyb0Td7TR"
# ## Prática 20
# Faça um programa que leia um número real chamado **preço**. Então faça uma repetição, que lê um **desconto** inteiro. O laço termina se desconto for zero, imprimindo **fim**. Caso contrário, deve ser impresso o preço com o desconto aplicado, usando duas casas decimais. Teste com a seguinte entrada: 240, 15, 27, 0.
# + id="QYiTlCVyd7Tb"
# + id="-tzlOH7vd7Tp"
validar(20)
# + [markdown] id="P7Lac0L643_K"
# # Exercícios
# + [markdown] id="AaW1CAqP5Cf2"
# ## Exercício 21
# Faça um programa que leia dois valores inteiros **x** e **y**. Se **x** for maior do que **y**, troque estes dois valores (usando uma variável auxiliar **z**). Então o programa deve imprimir todos os inteiros no intervalo aberto (x,y), ou seja, excluindo tanto **x** como **y**. Teste com 42 e 38.
#
# + id="fBnkW1hD4VL0"
# + id="xeFq_H9F6XfX"
validar(21)
# + [markdown] id="m-pU0-R1--O1"
# ## Exercício 22
# Faça um programa que leia 5 valores inteiros e ao final imprima o maior. Você deve criar uma variável chamada **maior** e colocar o primeiro valor lido nela. Para os demais valores lidos, faça uma repetição, sempre comparando com **maior** e atualizando esta variável se for o caso. Teste com a sequência 3, 5, 4, 7 e 1.
# + id="l42Yhbmt97o2"
# + id="DlTEf7kn-_Fg"
validar(22)
# + [markdown] id="rq-eoBAMASQl"
# ## Exercício 23
# Faça um programa que calcule e imprima o somatório, a soma dos quadrados (use **c * c**) e a média aritmética entre os **n** primeiros números inteiros positivos. Teste com **n=50**.
# + id="6nDjjseb_7Sb"
# + id="6dOaYciEATIb"
validar(23)
# + [markdown] id="20-YqgpZAT07"
# ## Exercício 24
# Faça um programa que multiplica dois números inteiros **m** e **n**, sem o recurso da operação de multiplicação, apresentando o resultado no final. Teste com 14 e 21.
# + id="NLajAZnhAUU1"
# + id="Po4qm9vVAUpl"
validar(24)
# + [markdown] id="d6pkQhwQAVaR"
# ## Exercício 25
# Faça um programa que localiza o menor entre vários valores reais positivos. O conjunto de valores informados termina quando o usuário informar zero. Teste com a sequência 9.3, 1.7, 4.1, 2.3, 5.6 e 0.
# + id="xGjLXj23AVss"
# + id="-hUXkVJAAV7U"
validar(25)
|
04_Python_Praticas_e_exercicios_de_programacao_com_lacos_de_repeticao.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# #### **Title**: Linked pointer cross-section example
#
# **Description**: A linked streams example demonstrating how to use PointerXY stream linked across two plots.
#
# **Dependencies**: Bokeh
#
# **Backends**: [Bokeh](./PointerX.ipynb)
import numpy as np
import holoviews as hv
from holoviews import opts
hv.extension('bokeh')
# +
opts.defaults(opts.Curve(width=100))
# Create two images
x,y = np.meshgrid(np.linspace(-5,5,101), np.linspace(5,-5,101))
img1 = hv.Image(np.sin(x**2+y**2))
img2 = hv.Image(np.sin(x**2+y**3))
# Declare PointerX and dynamic VLine
posx = hv.streams.PointerX(x=0)
vline = hv.DynamicMap(lambda x: hv.VLine(x or -100), streams=[posx])
# Declare cross-sections at PointerX location
# Using apply syntax
crosssection1 = img1.apply.sample(x=posx.param.x)
# By constructing a DynamicMap
crosssection2 = hv.DynamicMap(lambda x: img2.sample(x=x if x else 0), streams=[posx])
# Combine images, vline and cross-sections
((img1 * vline) << crosssection1) + ((img2 * vline) << crosssection2)
# -
# <center><img src="https://assets.holoviews.org/gifs/examples/streams/bokeh/linked_pointer_crosssection.gif" width=600></center>
|
examples/reference/streams/bokeh/PointerX.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Check data for any mouse ID with duplicate time points and remove any data associated with that mouse ID.
# ## Use the cleaned data for the remaining steps.
# Generate a summary statistics table consisting of the mean, median, variance, standard deviation, and SEM of the tumor volume for each drug regimen.
# Generate a bar plot using both Pandas's `DataFrame.plot()` and Matplotlib's `pyplot` that shows the number of total mice for each treatment regimen throughout the course of the study.
b
#**NOTE:** These plots should look identical.
# +
# Generate a pie plot using both Pandas's `DataFrame.plot()` and Matplotlib's `pyplot` that shows the distribution of female or male mice in the study.
# **NOTE:** These plots should look identical.
# -
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens: Capomulin, Ramicane, Infubinol, and Ceftamin. Calculate the quartiles and IQR and quantitatively determine if there are any potential outliers across all four treatment regimens.
# +
# Using Matplotlib, generate a box and whisker plot of the final tumor
# volume for all four treatment regimens and highlight any potential outliers
# in the plot by changing their color and style.
# **Hint**: All four box plots should be within the same figure.
# Use this [Matplotlib documentation page]
# (https://matplotlib.org/gallery/pyplots/boxplot_demo_pyplot.html#sphx-glr-gallery-pyplots-boxplot-demo-pyplot-py)
# for help with changing the style of the outliers.
# -
# Select a mouse that was treated with Capomulin and generate a line plot of time point
# versus tumor volume for that mouse.
# Generate a scatter plot of mouse weight versus average tumor volume
# for the Capomulin treatment regimen.
# Calculate the correlation coefficient and linear regression model between mouse
# weight and average tumor volume for the Capomulin treatment.
#Plot the linear regression model on top of the previous scatter plot.
# Look across all previously generated figures and tables and
# write at least three observations or inferences that can be made from the data.
# Include these observations at the top of notebook.
# +
# Here are some final considerations:
# You must use proper labeling of your plots, to include properties such as:
# plot titles, axis labels, legend labels, _x_-axis and _y_-axis limits, etc.
# See the [starter workbook](Pymaceuticals/pymaceuticals_starter.ipynb)
# for help on what modules to import and expected format of the notebook.
# +
|
.ipynb_checkpoints/MatPlotLib_HW-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ML Gdansk (1 VI 2020)
# +
import os
import sys
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import tensorflow_datasets as tfds
data_path = "data_location3"
sys.path.insert(0, "/home/jupyter/cache_disk2/MLG1VI2020")
from ds_utils_MLG import *
print(f'TensorFlow version: {tf.__version__}')
# -
TARGET_SIZE = (224,224,3)
N_Classes = 196
from tensorflow.keras.applications.mobilenet_v2 import MobileNetV2, preprocess_input
from tensorflow.keras.layers import Input, Dense, Dropout, GlobalAveragePooling2D, Activation, Lambda, Flatten
from tensorflow.keras.models import Model, load_model
# ### 1. Download and prepare dataset (make TFRecords)
(ds_train, ds_valid, ds_test), ds_info = tfds.load(
'cars196',
split=['train', 'test[50%:]', 'test[:50%]'],
as_supervised=True,
with_info=True)
ds_info
# See what we got:
display_images_and_labels_from_dataset(dataset=ds_train)
def get_filtered_resized_dataset(dataset, filter_label_value):
dataset = dataset.filter(lambda image, label: tf.equal(label, filter_label_value))
return dataset.map(lambda image, label: (tf.cast(tf.image.resize(image, TARGET_SIZE[:-1]),tf.uint8), label))
some_filtered_ds = get_filtered_resized_dataset(dataset=ds_train, filter_label_value=194)
display_images_and_labels_from_dataset(some_filtered_ds)
# +
from tqdm.notebook import tqdm as ntqdm
def produce_TFRs_from_filtered_datasets(dataset, subset):
for idx in ntqdm( range(N_Classes) ):
label = str(idx).zfill(3)
some_filtered_dataset = get_filtered_resized_dataset(dataset,filter_label_value=idx)
dataset_to_TFR(some_filtered_dataset, prefix_target_filename=f'{subset}_{label}', features_keys=['img','label'])
print(f'TFR for class {idx} finished!')
# -
produce_TFRs_from_filtered_datasets(dataset=ds_train, subset=f"{data_path}/train_L_")
produce_TFRs_from_filtered_datasets(dataset=ds_valid, subset=f"{data_path}/valid_L_")
produce_TFRs_from_filtered_datasets(dataset=ds_test, subset=f"{data_path}/test_L_")
|
materials/MLGdansk2020VI/data_producer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="F3Eu8-Ndwz45"
# # Data Exploratory
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" id="n0VvAXJowz46"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
#import os
#for dirname, _, filenames in os.walk('/kaggle/input'):
# for filename in filenames:
# print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + id="h-abzjZe0aV5"
import zipfile
path_to_zip_file='/content/drive/My Drive/Data/Vision/Histopathologic_Cancer/train.zip'
directory_to_extract_to='/content/train/'
with zipfile.ZipFile(path_to_zip_file, 'r') as zip_ref:
zip_ref.extractall(directory_to_extract_to)
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" id="eAZN789vwz49" outputId="c35e8e2b-a696-4e93-9bc2-b2e66c838405" colab={"base_uri": "https://localhost:8080/", "height": 204}
import pandas as pd
path2csv="/content/drive/My Drive/Data/Vision/Histopathologic_Cancer/train_labels.csv"
labels_df=pd.read_csv(path2csv)
labels_df.head()
# + id="tJua4yCSwz5B" outputId="72070089-e462-4d48-d03f-fd061dbc7c57" colab={"base_uri": "https://localhost:8080/", "height": 68}
print(labels_df['label'].value_counts())
# + id="hMokHD5fwz5E" outputId="3c1b2f7f-85e0-4a69-d3c8-6463bdcb1870" colab={"base_uri": "https://localhost:8080/", "height": 265}
# %matplotlib inline
labels_df['label'].hist();
# + id="U-WUMVRUwz5H" outputId="398c1eb1-63b7-4eea-a7f2-e230c07c0ac7" colab={"base_uri": "https://localhost:8080/", "height": 574}
import matplotlib.pylab as plt
from PIL import Image, ImageDraw
import numpy as np
import os
# %matplotlib inline
# data is stored here
path2train=directory_to_extract_to
# show images in gray-scale, if you want color change it to True
color=False
# get ids for malignant images
malignantIds = labels_df.loc[labels_df['label']==1]['id'].values
plt.rcParams['figure.figsize'] = (10.0, 10.0)
plt.subplots_adjust(wspace=0, hspace=0)
nrows,ncols=3,3
for i,id_ in enumerate(malignantIds[:nrows*ncols]):
full_filenames = os.path.join(path2train , id_ +'.tif')
# load image
img = Image.open(full_filenames)
# draw a 32*32 rectangle
draw = ImageDraw.Draw(img)
draw.rectangle(((32, 32), (64, 64)),outline="green")
plt.subplot(nrows, ncols, i+1)
if color is True:
plt.imshow(np.array(img))
else:
plt.imshow(np.array(img)[:,:,0],cmap="gray")
plt.axis('off')
# + id="k-hqHsAWwz5K" outputId="467badb7-81d0-45bf-e348-161ac6bb74e7" colab={"base_uri": "https://localhost:8080/", "height": 51}
print("image shape:", np.array(img).shape)
print("pixel values range from %s to %s" %(np.min(img), np.max(img)))
# + [markdown] id="mZbdi8jcwz5M"
# # Create a Custom Dataset
# + [markdown] id="-2fq4AgBwz5N"
# A traditional method for working with a dataset would be to load all images into NumPy
# arrays. Since we are dealing with a relatively large dataset, this would be a waste of our
# computer resources. If you are short on RAM, this would be impossible. Luckily, PyTorch
# has a powerful tool to handle large datasets.
#
# We can create a custom Dataset class by subclassing the PyTorch Dataset class. When
# creating a custom Dataset class, make sure to define two essential functions: __len__ and
# __getitem__. The __len__ function returns the dataset's length. This function is callable
# with the Python len function. The __getitem__ function returns an image at the specified
# index.
# + [markdown] id="Ic3hQyE0wz5N"
# In step 1, we imported the required packages. The PIL package is imported to load images.
# From torch.utils.data, we import Dataset as the base class of our custom dataset. The
# pandas package is imported to load the CSV files. We also used torchvision for data
# transformation.
#
# Then, we defined the Dataset class. In the __init__ function, we received the path to the
# data and the data_type. The data_type can be either train or test. The class, like any
# PyTorch dataset, has the __len__ and __getitem__ functions. The __len__ function
# returns the length of the dataset. The __getitem__ function also returns the transformed
# image at the given index and its corresponding label.
# + id="nA6e4sdO-9qf" outputId="ff481b00-64b9-4b13-d7c1-5c5f3d8cf8e6" colab={"base_uri": "https://localhost:8080/", "height": 419}
labels_df
# + id="FJscQhWtwz5O"
import torch
from PIL import Image
from torch.utils.data import Dataset
import pandas as pd
import torchvision.transforms as transforms
import os
# fix torch random seed
torch.manual_seed(0)
class histoCancerDataset(Dataset):
def __init__(self, data_dir, transform,data_type="train"):
# path to images
path2data=os.path.join(data_dir,data_type)
print(path2data)
# get list of images
filenames = os.listdir(path2data) # all file in path
print(filenames[:5])
# get the full path to images
self.full_filenames = [os.path.join(path2data, f) for f in filenames]
print(self.full_filenames[:5])
# labels are in a csv file named train_labels.csv
#path2csvLabels=os.path.join(data_dir,"train_labels.csv")
path2csvLabels=path2csv
print(path2csvLabels)
labels_df=pd.read_csv(path2csvLabels)
print(labels_df)
# set data frame index to id
labels_df.set_index("id", inplace=True)
print(labels_df)
# obtain labels from data frame
# <NAME> toan bo file name trong training voi label value tu labels_df
for filename in filenames[:3]:
print('---------')
print('filename[:-4]: ',filename[:-4])
print('labels_df.loc[filename[:-4]]: ',labels_df.loc[filename[:-4]])
print('labels_df.loc[filename[:-4]].values[0]: ',labels_df.loc[filename[:-4]].values[0])
print(type(labels_df.loc[filename[:-4]].values[0]))
self.labels = [labels_df.loc[filename[:-4]].values[0] for filename in filenames]
print(self.labels[:5])
self.transform = transform
def __len__(self):
# return size of dataset
return len(self.full_filenames)
def __getitem__(self, idx):
# open image, apply transforms and return with label
image = Image.open(self.full_filenames[idx]) # PIL image
image = self.transform(image)
return image, self.labels[idx]
# + [markdown] id="QhPlVf7owz5R"
# In step 2, we defined the transformation function. For now, we only convert PIL images into
# Pytorch tensors in the transformation function.
# + id="qRZ4yq3-wz5S"
import torchvision.transforms as transforms
data_transformer = transforms.Compose([transforms.ToTensor()])
# + id="IExO4neKwz5U" outputId="1149733a-fb57-4efa-f4f3-f676b60d4e40" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#data_dir = "../input/histopathologic-cancer-detection/" # must contain label and all images
data_dir='/content/'
histo_dataset = histoCancerDataset(data_dir, data_transformer, "train")
print(len(histo_dataset))
# + id="fBCvkd82wz5X" outputId="11b2c9cb-8a1b-4303-e700-610f47a0a329" colab={"base_uri": "https://localhost:8080/", "height": 163}
histo_dataset.labels
# + id="ZqqWO_9Lwz5Z" outputId="70bcd213-b338-433b-b6b7-0ef1a2aa58a2"
histo_dataset.full_filenames
# + [markdown] id="ffdJtanywz5c"
# In step 4, we can see that the dataset returns images in the (Channels, Height, Width) format
# and pixel values are normalized to the range [0.0, 1.0]. This is the result of transforms.
# ToTensor() converts a PIL image into the range [0, 255] to torch.FloatTensor of
# shape (C x H x W) in the range [0.0, 1.0]. It is common to use this formatting when working
# with images in PyTorch.
# + id="N11CyPHPwz5d" outputId="1d237cd8-b722-4b74-d215-c1cec02020ad"
# load an image
img,label=histo_dataset[9]
print(img.shape,torch.min(img),torch.max(img))
# + id="E3QI1GZ1wz5f" outputId="6d3eae2c-4101-4b6e-a47e-2a54c77d7e8d"
# load an image
img,label=histo_dataset[1]
print(img.shape,torch.min(img),torch.max(img))
# + id="KrluEqT-wz5h" outputId="ba8e2024-674f-4f96-c982-77e4ca9753e2"
img
# + [markdown] id="Y581LmJvwz5k"
# # Split Dataset
# + [markdown] id="AJf6-wctwz5l"
# In step 1, we used the random_split function from torch.utils.data to split the
# dataset. This will return two datasets: train_ds and val_ds with lengths of 176020 and
# 44005, respectively
# + id="I9WmV53Uwz5m" outputId="a6f80989-eaa7-48a0-9f73-11aae42450a1"
from torch.utils.data import random_split
len_histo=len(histo_dataset)
len_train=int(0.8*len_histo)
len_val=len_histo-len_train
train_ds,val_ds=random_split(histo_dataset,[len_train,len_val])
print("train dataset length:", len(train_ds))
print("validation dataset length:", len(val_ds))
# + id="Yuf72oM6wz5o" outputId="d6bce8e1-ae31-40af-8b3c-e4796c0387e5"
train_ds
# + id="d5bGTgeWwz5s" outputId="eded2db4-7e14-41f0-c73f-d92f2524f98b"
i=0
for x,y in train_ds:
print(x.shape,y)
i+=1
if i == 5:
break
# + id="X7DQoZoPwz5u" outputId="1e8ce96b-5d4c-44c7-eb19-164c44f4e122"
for x,y in val_ds:
print(x.shape,y)
break
# + id="1kvQC5B1wz5x" outputId="5be1642f-d856-439d-f7e3-456fcbd06d28"
from torchvision import utils
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(0)
def show(img,y,color=False):
# convert tensor to numpy array
npimg = img.numpy()
# Convert to H*W*C shape
npimg_tr=np.transpose(npimg, (1,2,0))
if color==False:
npimg_tr=npimg_tr[:,:,0]
plt.imshow(npimg_tr,interpolation='nearest',cmap="gray")
else:
# display images
plt.imshow(npimg_tr,interpolation='nearest')
plt.title("label: "+str(y))
grid_size=4
rnd_inds=np.random.randint(0,len(train_ds),grid_size)
print("image indices:",rnd_inds)
x_grid_train=[train_ds[i][0] for i in rnd_inds]
y_grid_train=[train_ds[i][1] for i in rnd_inds]
x_grid_train=utils.make_grid(x_grid_train, nrow=4, padding=2)
print(x_grid_train.shape)
plt.rcParams['figure.figsize'] = (10.0, 5)
show(x_grid_train,y_grid_train)
# + id="EZ13dVu3wz5z" outputId="780beb0b-10f1-4043-dc67-19a641bd5dbe"
train_ds[0]
# + id="n6t09qqhwz51" outputId="dcfc18ec-e554-423a-dd41-bb797a449a4c"
grid_size=4
rnd_inds=np.random.randint(0,len(val_ds),grid_size)
print("image indices:",rnd_inds)
x_grid_val=[val_ds[i][0] for i in range(grid_size)]
y_grid_val=[val_ds[i][1] for i in range(grid_size)]
x_grid_val=utils.make_grid(x_grid_val, nrow=4, padding=2)
print(x_grid_val.shape)
show(x_grid_val,y_grid_val)
# + [markdown] id="8kjzg5d4wz54"
# # Transform Data
# + [markdown] id="cZYTEhcbwz54"
# Image transformation and image augmentation are necessary for training deep learning
# models. By using image transformations, we can expand our dataset or resize and
# normalize it to achieve better model performance. Typical transformations include
# horizontal and vertical flipping, rotation, and resizing. The good news is that we can use
# various image transformations for our binary classification model without making label
# changes. For instance, if we rotate or flip a malignant image, it will remain malignant. In
# this recipe, you will learn how to use the torchvision package to perform on-the-fly
# image transformation during training.
# + [markdown] id="ob_uixkZwz54"
# In step 1, RandomHorizontalFlip and RandomVerticalFlip will flip the image
# horizontally and vertically with a probability of 0.5, respectively.
# The RandomRotation function rotates images in the range of [-45,45] degrees.
# Also, RandomSizedCrop crops a square image randomly in the range of [72, 96] and then
# resizes it to the original size of 96*96.
# + id="OGPd7xnqwz55"
#1. First, let's define the following transformations for the training dataset:
train_transformer = transforms.Compose([
transforms.RandomHorizontalFlip(p=0.5),
transforms.RandomVerticalFlip(p=0.5),
transforms.RandomRotation(45),
transforms.RandomResizedCrop(96,scale=(0.8,1.0),ratio=(1.0,1.0)),
transforms.ToTensor()])
# + [markdown] id="ICDvQmycwz57"
# In step 2, we used transforms.ToTensor to normalize the images in the range [0, 1] and
# convert them into tensors. It is common to normalize images to zero-mean and unitvariance by subtracting the mean and diving into the standard deviation of the pixel values.
# However, here, we opt not to do any further normalization.
# The validation dataset was only normalized to the range [0, 1] without any data
# augmentation.
# + id="GpPqs2Lbwz58"
#2. For the validation dataset, we don't need any augmentation. So, we only convert
#the images into tensors in the transforms function:
val_transformer = transforms.Compose([transforms.ToTensor()])
# + [markdown] id="6z1ZrSALwz5_"
# Finally, in step 3, we defined the transformations and overwrote the transform functions of
# train_ds and val_ds.
# + id="rzjYPObqwz5_"
# overwrite the transform functions
train_ds.transform=train_transformer
val_ds.transform=val_transformer
# + id="P2d4fNkhwz6C" outputId="32515bc9-e2d4-4060-bcdd-12b1198f9912"
dir(train_ds)
# + id="CJUME9czwz6E" outputId="c8ee39e9-67a0-42fc-9d44-f591b155613e"
len(train_ds)
# + [markdown] id="oab5Frh8wz6G"
# # Create Dataloader
# + [markdown] id="i2swumVGwz6G"
# We are ready to create a PyTorch dataloader. If we do not use dataloaders, we have to write
# code to loop over datasets and extract a data batch. This process can be made automatically
# using a PyTorch Dataloader.
# + [markdown] id="khyI4IiSwz6H"
# In step 1, when defining a PyTorch Dataloader, we need to define the batch size. The batch
# size determines the number of images to be extracted from the dataset in each iteration. The
# typical values of the batch size for classification tasks are in the range of [8-128]. Also, note
# that we do not need to shuffle the validation data during evaluation.
#
# When iterating over a dataloader, it will automatically extract batches of data from the
# dataset. As seen in step 2, for the train dataloader, we extracted batches of images that were
# 32 in size. This was 64 for the validation dataloader since we set the batch size to 64.
# + id="Lgf3Q9_2wz6H"
from torch.utils.data import DataLoader
train_dl = DataLoader(train_ds, batch_size=32, shuffle=True)
val_dl = DataLoader(val_ds, batch_size=64, shuffle=False)
# + id="-QJ3gHwJwz6J" outputId="9330cc77-6787-4601-e7d4-e651c61eb188"
len(train_dl) # 5501 batchs, each batch have 32 image --> total 176032 images
# + id="5kEDKCQXwz6L" outputId="4be35cc9-b777-41ee-e0a5-5522088e4da8"
# extract a batch from trainin data
for x, y in train_dl:
print(x.shape)
print(y.shape)
break
# + id="3jFZ4ZQUwz6N" outputId="dbcd4816-b66f-46e1-873c-50867c90b47e"
x
# + id="W_N4AoEPwz6P" outputId="1a4deb0f-2e19-4171-8f6c-18d85c7f2003"
y
# + id="G-bjCUDLwz6R" outputId="395a6fdb-e3fc-4462-fafd-1fb011ced56a"
# extract a batch from validation data
for x, y in val_dl:
print(x.shape)
print(y.shape)
break
# + [markdown] id="iGxzcAs4wz6T"
# # Building Classification Model
# + [markdown] id="MeuTXPBWwz6U"
# ## dumb baselines
# + [markdown] id="uwgG7nxtwz6U"
# In step 1, we created dumb baselines. We should always establish a baseline before moving
# forward with complex models. This will help us find out how well we are performing. We
# can create dumb baselines by creating random predictions, all-zero predictions, or all-one
# predictions. Let's create simple baselines using these predictions. It is interesting to see that
# we can get an accuracy of 0.60 with dumb predictions.
# + id="M1AhB79-wz6U" outputId="25425c64-59ef-4b53-9e0b-146bf6ac4e8b"
# get labels for validation dataset
y_val=[y for _,y in val_ds]
def accuracy(labels, out):
return np.sum(out==labels)/float(len(labels))
# accuracy all zero predictions
acc_all_zeros=accuracy(y_val,np.zeros_like(y_val))
# accuracy all ones predictions
acc_all_ones=accuracy(y_val,np.ones_like(y_val))
# accuracy random predictions
acc_random=accuracy(y_val,np.random.randint(2,size=len(y_val)))
print("accuracy random prediction: %.2f" %acc_random)
print("accuracy all zero prediction: %.2f" %acc_all_zeros)
print("accuracy all one prediction: %.2f" %acc_all_ones)
# + [markdown] id="SisKNDeMwz6W"
# ## find Output size
# + [markdown] id="7dblynydwz6X"
# Let's implement a helper function to calculate the output size of a CNN layer
#
# In step 2, we developed findConv2DOutShape to automatically compute the output size of
# - a CNN and pooling layer. The inputs to this function are:
# - H_in: an integer representing the height of input data
# - W_in: an integer representing the width of input data
# - conv: an object of the CNN layer
# - pool: an integer representing the pooling size and default to 2
# + id="Q1_48V12wz6X" outputId="d15cb970-e1c3-42a7-e484-d333a816694a"
import torch.nn as nn
def findConv2dOutShape(H_in,W_in,conv,pool=2):
# get conv arguments
kernel_size=conv.kernel_size
stride=conv.stride
padding=conv.padding
dilation=conv.dilation
# Ref: https://pytorch.org/docs/stable/nn.html
H_out=np.floor((H_in+2*padding[0]-dilation[0]*(kernel_size[0]-1)-1)/stride[0]+1)
W_out=np.floor((W_in+2*padding[1]-dilation[1]*(kernel_size[1]-1)-1)/stride[1]+1)
if pool:
H_out/=pool
W_out/=pool
return int(H_out),int(W_out)
# example
conv1 = nn.Conv2d(3, 8, kernel_size=3) # in channel: 3 (R,P,G), out channel: 8 (filter)
h,w=findConv2dOutShape(96,96,conv1)
print(h,w)
# + [markdown] id="d3_0EMoHwz6Z"
# ## Define Model
# + [markdown] id="cyZyzdHiwz6Z"
# In this section, we will build a model for our binary classification task. Our model is
# comprised of four convolutional neural networks (CNNs) and two fully connected layers,
# as shown in the following diagram:
#
# In step 3, we implemented the CNN model using the nn.Module class. In the __init__
# function, we define the layers of the model. We provide the model parameters as a Python
# dictionary to this function. We recommend using variables for the model parameters and a
# dictionary to define them outside the class. This way, in the case of parameter changes, we
# do not have to make changes inside the Net class. This will also make future
# hyperparameter searches easier.
#
# For most layers, it is important to specify the number of inputs and outputs of the layer. For
# instance, the first CNN layer has C_in=3 channels as input and init_f=8 filters as output,
# as defined in the following code:
#
# self.conv1 = nn.Conv2d(C_in, init_f, kernel_size=3)
#
# Similarly, the second CNN layer receives init_f=8 channels as input and 2*init_f=16
# channels as output. Thus, by providing the number of output channels of the previous layer
# as the number of input channels to the next layer, we can define each layer. However, this
# becomes tricky when it comes to nn.Linear layers. The linear layer accepts a 2D tensor.
# That is why we need the view method in the forward function to reshape the 4D tensor
# into a 2D tensor:
#
# flatten/reshape
#
# x = x.view(-1, self.num_flatten)
#
# How do we get the value of self.num_flatten? Well, PyTorch does not automatically
# calculate this value. It is our duty to find this value and provide it to the linear layer. One
# approach would be to print (x.size()) to get the output shape of each layer.
#
# The other option is using the findConv2DOutShape function. The output size of the fourth
# CNN layer (together with the pooling layer) is h,w and there are 8*init_f output
# channels. Therefore, after flattening, we get self.num_flatten=h*w*8*init_f.
#
# We also added a dropout layer before the output layer to reduce the overfitting problem in
# deep learning models. Notice that we set the training = self.training argument in
# the F.dropout function. The self.training parameter is automatically set to True
# during training and False at evaluation. This will bypass the dropout layer at the
# deployment time.
#
# The dropout layer is only applied during training. At deployment, the
# dropout layer should be deactivated. To do so, make sure to set training
# argument of the dropout layer to False at deployment.
#
# The last layer is F.log_softmax(x, dim=1) (with two outputs), which is equivalent to
# . We will explain why we choose this output in the next section when
# defining the loss function. Keep in mind that if we want to get the output probability value,
# we need to use the exponential operation.
# + id="Dqv_9AkEwz6a"
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self, params):
super(Net, self).__init__()
C_in,H_in,W_in=params["input_shape"] #3x96x96
init_f=params["initial_filters"] #8
num_fc1=params["num_fc1"] #100
num_classes=params["num_classes"] #2
self.dropout_rate=params["dropout_rate"] #0.25
self.conv1 = nn.Conv2d(C_in, init_f, kernel_size=3) #3,8,3
h,w=findConv2dOutShape(H_in,W_in,self.conv1) # 96x96,self.conv1
print(h,w) # 47x47
self.conv2 = nn.Conv2d(init_f, 2*init_f, kernel_size=3)
h,w=findConv2dOutShape(h,w,self.conv2)
print(h,w) # 22x22
self.conv3 = nn.Conv2d(2*init_f, 4*init_f, kernel_size=3)
h,w=findConv2dOutShape(h,w,self.conv3)
print(h,w) # 10x10
self.conv4 = nn.Conv2d(4*init_f, 8*init_f, kernel_size=3)
h,w=findConv2dOutShape(h,w,self.conv4)
print(h,w) # 4x4
# compute the flatten size
self.num_flatten=h*w*8*init_f # 4x4x8x8
print(self.num_flatten)#1024
self.fc1 = nn.Linear(self.num_flatten, num_fc1)
self.fc2 = nn.Linear(num_fc1, num_classes)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv3(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv4(x))
x = F.max_pool2d(x, 2, 2)
#The linear layer accepts a 2D tensor.That is why we need the view method in the
#forward function to reshape the 4D tensor into a 2D tensor:
x = x.view(-1, self.num_flatten)
x = F.relu(self.fc1(x))
x=F.dropout(x, self.dropout_rate)
x = self.fc2(x)
return F.log_softmax(x, dim=1)
# + [markdown] id="vbiyIDW_wz6e"
# In step 4, we constructed an object of the Net class. The input shape is the shape of the
# image tensors (3,96,96). We choose initial_filters=8 filters for the first CNN layer. The
# number of filters is automatically doubled in the next layers. Also, there are num_fc1=100
# units in the fully connected layer.
# + id="1z3QORVkwz6e" outputId="da68e4bb-6bb1-485d-cb09-40f1b6511ee7"
# dict to define model parameters
params_model={
"input_shape": (3,96,96),
"initial_filters": 8,
"num_fc1": 100,
"dropout_rate": 0.25,
"num_classes": 2,
}
# create model
cnn_model = Net(params_model)
# + [markdown] id="iOPyh0LIwz6h"
# In step 5, if you have a CUDA device, always move the model to the CUDA device to take
# advantage of the computation acceleration.
# + id="ND1i0VM_wz6h"
# move model to cuda/gpu device
if torch.cuda.is_available():
device = torch.device("cuda")
cnn_model=cnn_model.to(device)
# + id="5eNQI88cwz6j" outputId="76d83559-94ee-447d-efcb-beff0a3095c2"
print(cnn_model)
# + id="3bgfbiXkwz6l" outputId="de934f82-3230-4df7-d561-2cd50509634a"
print(next(cnn_model.parameters()).device)
# + id="_Ql9Vib-wz6p" outputId="d4b18d11-fa9e-47fd-8242-f7420eea4947"
# !pip install torchsummary
# + id="foYEXoeRwz6u" outputId="aabbe29c-f32f-4ee7-db73-e50bc8990949"
from torchsummary import summary
summary(cnn_model, input_size=(3, 96, 96))
#Conv2d-1 : -1, 8 filter, 94 height , 94 width
#Conv2d-2 : -1, 16 filter, 45 height, 45 width
# + [markdown] id="5INW7Be1wz6w"
# Then, we showed the model summary. The summary shows the output shape and the
# number of parameters per layer. However, this does not include the functional layers. It is
# interesting to see that the fully connected layer has the highest number of parameters
# (102500), which is not efficient. Therefore, state-of-the-art deep learning models eliminate
# fully connected layers in their design.
# + [markdown] id="hgVKrBRUwz6w"
# # Loss function
# + [markdown] id="HnrkK3crwz6x"
# Output Activation | Number of Outputs | Loss Function
#
# None |1 |nn.BCEWithLogitsLoss
#
# Sigmoid| 1 |nn.BCELoss
#
# None |2 |nn.CrossEntropyLoss
#
# log_softmax |2 |nn.NLLLoss
#
# We recommend using the log_softmax function as it is easier to expand to multi-class
# classification. PyTorch combines the log and softmax operations into one function due to
# numerical stability and speed.
#
# We use log_softmax as the output and nn.NLLLoss as the negative log-likelihood
# loss. An important argument in defining the loss function to pay attention to is the
# reduction, which specifies the reduction to apply to the output. There are three options to
# choose from: none, sum, and mean. We choose reduction=sum so that the output loss will
# be summed. Since we will process the data in batches, this will return the sum of loss
# values per batch of data.
# + id="-gboKMA5wz6x"
loss_func = nn.NLLLoss(reduction="sum")
# + [markdown] id="MvBUcuTBwz6z"
# Next, we will use the loss in an example:
#
# In step 2, we calculate the loss using an example with n=8 samples and c=2 classes.
#
# In step 3, we compute the gradients for the example in step 2. Later, we will use the
# backward method to compute the gradients of the loss with respect to the model
# parameters.
# + id="nrjYocv5wz60" outputId="e0895f89-c1fa-4700-94a8-7fb8f71fb4fa"
# fixed random seed
torch.manual_seed(0)
n,c=8,2
y = torch.randn(n, c, requires_grad=True)
print(y)
ls_F = nn.LogSoftmax(dim=1)
y_out=ls_F(y)
print(y_out)
print(y_out.shape)
target = torch.randint(c,size=(n,))
print(target)
print(target.shape)
# + id="Fnx72kR_wz62" outputId="fdef0017-27f7-4996-f942-e79982f7b9ca"
loss = loss_func(y_out, target)
print(loss.item())
# + id="H1uW6yujwz65" outputId="da53f4a3-0987-4a2c-e8c5-67534e08eec7"
loss.backward()
print (y.data)
# + [markdown] id="WQICOjIrwz67"
# # Defining Optimizer
# + [markdown] id="g9WgbzJqwz67"
# The torch.optim package provides the implementation of common optimizers. The
# optimizer will hold the current state and will update the parameters based on the
# computed gradients. For binary classification tasks, SGD and Adam optimizers are used the
# most.
#
# Another helpful tool in the torch.optim package is learning schedules. Learning
# schedules are useful tools for automatically adjusting the learning rate during training to
# improve model performance.
#
# In step 1, we defined the Adam optimizer. The important parameters of the optimizer class
# are the model's parameters and the learning rate. cnn_model.parameters() returns an
# iterator over module parameters that are passed to the optimizer. The learning rate will
# determine the amount to update by. In PyTorch, we can set a different learning rate per
# layer, but in this chapter, we'll choose to set one learning rate for all the layers. In step 2, we
# developed a helper function that returns the current value of the learning rate.
# + id="z-YuHDATwz68"
#1. First, let's define an object of the Adam optimizer with a learning rate of 3e-4:
from torch import optim
opt = optim.Adam(cnn_model.parameters(), lr=3e-4)
# + id="sYfGythZwz69" outputId="b37ddb7c-f103-4f0c-d374-70edd4830dd9"
#2. We can read the current value of the learning rate using the following function:
# get learning rate
def get_lr(opt):
for param_group in opt.param_groups:
return param_group['lr']
current_lr=get_lr(opt)
print('current lr={}'.format(current_lr))
# + [markdown] id="7dGGMwaYwz7C"
# In step 3, we used the ReduceLROnPlateau method from
# the torch.optim.lr_scheduler package. This scheduler reads a metric quantity and if
# no improvement is seen for a patience number of epochs, the learning rate is reduced by a
# factor of 0.5. The mode argument defines whether the metric quantity is increasing or
# decreasing during training. For instance, if we monitor the loss value, we set mode='min'.
# If we monitor the accuracy, we should set mode='max'.
#
# In step 4, we showed you how the learning rate schedule works by passing it a constant
# value during a loop. As we saw, after every 20 iterations, the scheduler reduces the learning
# rate by half.
# + id="zuRQuQpMwz7C"
# 3. Next, we will define a learning scheduler using the ReduceLROnPlateau method:
from torch.optim.lr_scheduler import ReduceLROnPlateau
# define learning rate scheduler
lr_scheduler = ReduceLROnPlateau(opt, mode='min',factor=0.5, patience=20,verbose=1)
# + id="QgkSWWeQwz7E" outputId="87fed30f-9e2c-4d10-c24a-64619fd0ad7e"
# 4. Then, we will learn how the learning rate schedule works using the following example:
for i in range(100):
lr_scheduler.step(1)
# + [markdown] id="yO0rIdb7wz7G"
# # Training and Evaluation
# + [markdown] id="RY2ZzlTdwz7G"
# So far, we've created the datasets, built the model, and defined the loss function and
# optimizer. In this recipe, we'll implement the training and validation scripts. The training
# and validation scripts can be long and repetitive. For better code readability and to avoid
# code repetition, we'll build a few helper functions first.
# + [markdown] id="kxvvl2UYwz7G"
# 1. First, let's develop a helper function to count the number of correct predictions
# per data batch:
#
# In step 1, we counted the number of correct predictions per batch to be used for calculating
# the model accuracy.
# + id="-KNU0cjdwz7H"
def metrics_batch(output, target):
# get output class
pred = output.argmax(dim=1, keepdim=True)
# compare output class with target class
corrects=pred.eq(target.view_as(pred)).sum().item()
return corrects
# + id="-vhf9iPEwz7K"
n,c=8,2
output = torch.randn(n, c, requires_grad=True)
print (output)
print(output.shape)
#target = torch.randint(c,size=(n,))
target = torch.ones(n,dtype=torch.long)
print(target.shape)
metrics_batch(output,target)
# + [markdown] id="2HSE96DKwz7M"
# 2. Then, we will develop a helper function to compute the loss value per batch of
# data:
#
# In step 2, the loss_batch function receives the objects of the loss function and optimizer,
# the model output, and the ground truth label. We only perform the backpropagation step
# during training.
# + id="mDTxewUbwz7M"
def loss_batch(loss_func, output, target, opt=None):
# get loss
loss = loss_func(output, target)
# get performance metric
metric_b = metrics_batch(output,target)
if opt is not None:
opt.zero_grad()
loss.backward()
opt.step()
return loss.item(), metric_b
# + [markdown] id="bCduFrx9wz7O"
# 3. Next, we develop a helper function to compute the loss value and the
# performance metric for the entire dataset, also called an epoch
#
# In step 3, the loss_epoch function receives the objects of the model, the loss function, a
# Dataloader, and the optimizer. We used the Dataloader to fetch batches of data from the
# dataset. Then, we moved the batch to the CUDA device and got the model's output. We
# also declared two variables to keep the loss and number of correct predictions after a batch
# is executed. You can see how the choice of reduction="sum" in the definition of the loss
# function is deployed to accumulate the loss over the entire dataset. The sanity_check flag
# breaks the loop to quickly execute the function in the case of debugging
# + id="hhCeWAxKwz7O"
# define device as a global variable
device = torch.device("cuda")
def loss_epoch(model,loss_func,dataset_dl,sanity_check=False,opt=None):
running_loss=0.0
running_metric=0.0
len_data=len(dataset_dl.dataset)
for xb, yb in dataset_dl:
# move batch to device
xb=xb.to(device)
yb=yb.to(device)
# get model output
output=model(xb)
# get loss per batch
loss_b,metric_b=loss_batch(loss_func, output, yb, opt)
# update running loss
running_loss+=loss_b
# update running metric
if metric_b is not None:
running_metric+=metric_b
# break the loop in case of sanity check
if sanity_check is True:
break
# average loss value
loss=running_loss/float(len_data)
# average metric value
metric=running_metric/float(len_data)
return loss, metric
# + [markdown] id="VS2h0pBqwz7R"
# 4. Let's develop the train_val function in the following code blocks.The function is too long, so we will present it in a few steps. Keep in mind that all
# the following code blocks belong to one function.
#
# In step 4, we passed the parameters as a Python dictionary. This will improve the code's
# readability. We defined two dictionaries to record the loss and metric values during
# training for visualization purposes.
# + id="M-idRNN2wz7R"
def train_val(model, params):
# extract model parameters
num_epochs=params["num_epochs"]
loss_func=params["loss_func"]
opt=params["optimizer"]
train_dl=params["train_dl"]
val_dl=params["val_dl"]
sanity_check=params["sanity_check"]
lr_scheduler=params["lr_scheduler"]
path2weights=params["path2weights"]
# history of loss values in each epoch
loss_history={
"train": [],
"val": [],
}
# histroy of metric values in each epoch
metric_history={
"train": [],
"val": [],
}
'''
An important step is to track the model's performance during training to avoid overfitting.
To this end, we evaluate the model on the validation dataset after each training epoch.
Then, we compared the validation loss with the best validation loss, val_loss <
best_loss. If the validation loss has improved, we store and make copies of the weights as
the best model weights.
'''
# a deep copy of weights for the best performing model
best_model_wts = copy.deepcopy(model.state_dict())
# initialize best loss to a large value
best_loss=float('inf')
# main loop
for epoch in range(num_epochs):
# get current learning rate
current_lr=get_lr(opt)
print('Epoch {}/{}, current lr={}'.format(epoch, num_epochs - 1, current_lr))
# train model on training dataset
model.train()
train_loss, train_metric=loss_epoch(model,loss_func,train_dl,sanity_check,opt)
# collect loss and metric for training dataset
loss_history["train"].append(train_loss)
metric_history["train"].append(train_metric)
# evaluate model on validation dataset
model.eval()
with torch.no_grad():
val_loss, val_metric=loss_epoch(model,loss_func,val_dl,sanity_check)
# store best model
if val_loss < best_loss:
best_loss = val_loss
best_model_wts = copy.deepcopy(model.state_dict())
# store weights into a local file
torch.save(model.state_dict(), path2weights)
print("Copied best model weights!")
# collect loss and metric for validation dataset
loss_history["val"].append(val_loss)
metric_history["val"].append(val_metric)
'''
The learning rate scheduler monitors the validation loss and reduces the learning rate by a
factor of two
Also, every time the learning rate is reduced, we would like to continue training from the
best weights:
'''
# learning rate schedule
lr_scheduler.step(val_loss)
if current_lr != get_lr(opt):
print("Loading best model weights!")
model.load_state_dict(best_model_wts)
print("train loss: %.6f, dev loss: %.6f, accuracy: %.2f" %(train_loss,val_loss,100*val_metric))
print("-"*10)
# load best model weights
model.load_state_dict(best_model_wts)
return model, loss_history, metric_history
# + [markdown] id="EVNhSHPKwz7T"
# 5. Let's set the sanity_check flag to True and run the code.
#
# Developing and training deep learning models is time-consuming. The last thing you want
# is an error popping up somewhere in the middle of training. We recommend performing a
# sanity check before starting the actual training. We created a flag called sanity_check for
# this purpose. If set to True, the training loop breaks after one batch in an epoch. Therefore,
# we get to see the whole loop in a short amount of time.
#
# In step 5, the sanity check should quickly train and evaluate the model for 100 epochs.
# Everything looks normal. Even the learning rate schedule was activated and reduced the
# learning rate from 3e-4 to 7.5e-5. The training loss converges to near zero, which makes it
# clear that we can overfit to one batch. Don't worry about accuracy. This is a sanity check,
# after all.
# + id="2RFViDGRwz7T"
# Define the objects for the optimization, loss, and learning rate schedule:
import copy
loss_func = nn.NLLLoss(reduction="sum")
opt = optim.Adam(cnn_model.parameters(), lr=3e-4)
lr_scheduler = ReduceLROnPlateau(opt, mode='min',factor=0.5, patience=20,verbose=1)
# Define the training parameters and call the train_val helper function:
params_train={
"num_epochs": 100,
"optimizer": opt,
"loss_func": loss_func,
"train_dl": train_dl,
"val_dl": val_dl,
"sanity_check": True,
"lr_scheduler": lr_scheduler,
#"path2weights": "./models/weights.pt",
"path2weights": "weights.pt"
}
# train and validate the model
cnn_model,loss_hist,metric_hist=train_val(cnn_model,params_train)
# + [markdown] id="DbHq76T6wz7X"
# This will print out the following output:
#
# Epoch 0/99, current lr=0.0003
#
# Copied best model weights!
#
# train loss: 0.000129, dev loss: 0.001024, accuracy: 0.05
#
# ....
#
# Epoch 1/99, current lr=0.0003
#
# Copied best model weights!
#
# train loss: 0.000125, dev loss: 0.001021, accuracy: 0.05
# + [markdown] id="K1xfi9q7wz7X"
# 6. Let's plot the training validation's progress using the returned values, that
# is, loss_hist and metric_hist:
#
# In step 6, we saw that, at some point, the learning rate schedule will reduce the learning rate
# after the validation loss does not improve for 20 epochs. When this happens, the model is
# reloaded with the last best weights. That is why we see sudden jumps in the loss and
# accuracy curves at these points.
#
# We stored the best model weights (state_dict) during training by tracking the model's
# performance on the validation dataset. The weights are located in
# the ./models/weights.pt path. This is useful in case the program is stopped for any
# reason – at least you have a copy of the best weights.
# + id="GY0Tmyxbwz7Y"
# Train-Validation Progress
num_epochs=params_train["num_epochs"]
# plot loss progress
plt.title("Train-Val Loss")
plt.plot(range(1,num_epochs+1),loss_hist["train"],label="train")
plt.plot(range(1,num_epochs+1),loss_hist["val"],label="val")
plt.ylabel("Loss")
plt.xlabel("Training Epochs")
plt.legend()
plt.show()
# plot accuracy progress
plt.title("Train-Val Accuracy")
plt.plot(range(1,num_epochs+1),metric_hist["train"],label="train")
plt.plot(range(1,num_epochs+1),metric_hist["val"],label="val")
plt.ylabel("Accuracy")
plt.xlabel("Training Epochs")
plt.legend()
plt.show()
# + [markdown] id="Qfi1yAA6wz7Z"
# 7. By doing this, we've made sure that all the elements are implemented correctly.
# Let's set the flag to sanity_check: False and run the code. The training will
# start and we will see its progress, as shown in the following code block:
# + id="_YX7O0Iwwz7a"
# %%time
import copy
loss_func = nn.NLLLoss(reduction="sum")
opt = optim.Adam(cnn_model.parameters(), lr=3e-4)
lr_scheduler = ReduceLROnPlateau(opt, mode='min',factor=0.5, patience=20,verbose=1)
params_train={
"num_epochs": 2,
"optimizer": opt,
"loss_func": loss_func,
"train_dl": train_dl,
"val_dl": val_dl,
"sanity_check": False,
"lr_scheduler": lr_scheduler,
"path2weights": "./weights.pt",
}
# train and validate the model
cnn_model,loss_hist,metric_hist=train_val(cnn_model,params_train)
# + id="BG25ajLjwz7c"
# Train-Validation Progress
num_epochs=params_train["num_epochs"]
# plot loss progress
plt.title("Train-Val Loss")
plt.plot(range(1,num_epochs+1),loss_hist["train"],label="train")
plt.plot(range(1,num_epochs+1),loss_hist["val"],label="val")
plt.ylabel("Loss")
plt.xlabel("Training Epochs")
plt.legend()
plt.show()
# plot accuracy progress
plt.title("Train-Val Accuracy")
plt.plot(range(1,num_epochs+1),metric_hist["train"],label="train")
plt.plot(range(1,num_epochs+1),metric_hist["val"],label="val")
plt.ylabel("Accuracy")
plt.xlabel("Training Epochs")
plt.legend()
plt.show()
# + [markdown] id="GXEIIWofwz7d"
# There's more...
#
# Once you get a baseline performance using your first model, you can start tuning
# hyperparameters. Hyperparameter tuning is a technique that's used to improve model
# performance. An effective way of doing this is by using a random search. We defined the
# hyperparameters as variables so that you can easily play with them. Try different values for
# the hyperparameters and see how the model's performance is affected. As an example, try
# increasing the number of CNN filters by setting initial_filters = 16 and retraining
# the model.
# + id="HJ2DJb_vwz7d"
# + id="BqDDDHPowz7h"
# + id="XkVqWIJ9wz7i"
# + id="cX9xHzGuwz7k"
|
Chapter02/Kaggle_histopathologic-cancer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# Load data, rename columns and drop missing data - testing
load = (
pd.read_csv("data/raw/hour.csv",header = None)
.rename(columns={0:"instant",1:"dteday",2:"season",3:"yr",4:"mnth",5:"hr",6:"holiday",7:"weekday",
8:"workingday",9:"weathersit",10:"temp",11:"atemp",12:"hum",13:"windspeed",
14:"casual",15:"registered",16:"cnt"})
.dropna()
.rename(index = lambda x: x + 1)
)
# +
def load_and_process(path):
load = (
pd.read_csv(path,header = None)
.rename(columns={0:"instant",1:"dteday",2:"season",3:"yr",4:"mnth",5:"hr",6:"holiday",7:"weekday",
8:"workingday",9:"weathersit",10:"temp",11:"atemp",12:"hum",13:"windspeed",
14:"casual",15:"registered",16:"cnt"})
.dropna()
.rename(index = lambda x: x + 1)
)
return load_and_process
load_and_process("data/raw/hour.csv")
# -
|
analysis/.ipynb_checkpoints/Dante-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# # Plotting methods
# +
import pandas as pd
import matplotlib.pyplot as plt
df=pd.read_csv("heart.csv")
df.head(2)
# -
fig, axes=plt.subplots(ncols=2)
df.plot.scatter(x="age",y="chol",ax=axes[0]);
df.plot.scatter(x="age",y="trestbps",ax=axes[1]);
fig.tight_layout();
fig2=axes[0].get_figure()
fig2=plt.savefig("plots.png",bbox_inches="tight",transparent=True)
fig2
with plt.style.context("default"):
fig,axes=plt.subplots(ncols=3,sharey=True,
gridspec_kw={"width_ratios":[2,1,1],"wspace":0})
y="age"
xs=["chol","trestbps","thalach"]
for x,ax in zip(xs,axes):
ax.scatter(df[x],df[y])
ax.set_xlabel(x)
axes[0].set_ylabel(y);
fig.savefig("output.pdf")
|
Vizuallizationexplore/Basics/PlottingMethords.ipynb
|
(* -*- coding: utf-8 -*-
(* --- *)
(* jupyter: *)
(* jupytext: *)
(* text_representation: *)
(* extension: .ml *)
(* format_name: light *)
(* format_version: '1.5' *)
(* jupytext_version: 1.14.4 *)
(* kernelspec: *)
(* display_name: OCaml 4.07.1 *)
(* language: OCaml *)
(* name: ocaml-jupyter *)
(* --- *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* <center> *)
(* *)
(* <h1 style="text-align:center"> Lambda Calculus : Syntax </h1> *)
(* <h2 style="text-align:center"> CS3100 Fall 2019 </h2> *)
(* </center> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Review *)
(* *)
(* ### Last time *)
(* *)
(* * Higher Order Functions *)
(* *)
(* ### Today *)
(* *)
(* * Lambda Calculus: Basis of FP! *)
(* + Origin, Syntax, substitution, alpha equivalence *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Computability *)
(* *)
(* <h3> In 1930s </h3> *)
(* *)
(* * What does it mean for the function $f : \mathbb{N} \rightarrow \mathbb{N}$ to be *computable*? *)
(* * **Informal definition:** A function is computable if using pencil-and-paper you can compute $f(n)$ for any $n$. *)
(* * Three different researchers attempted to formalise *computability*. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## <NAME> *)
(* *)
(* <img src="images/turing.jpg" style="float:left" width="200"> *)
(* *)
(* <div style="float:left;width:75%"> *)
(* *)
(* * Defined an idealised computer -- **The Turing Machine** (1935) *)
(* * A function is computable if and only if it can be computed by a turning machine *)
(* * A programming language is turing complete if: *)
(* + It can map every turing machine to a program. *)
(* + A program can be written to emulate a turing machine. *)
(* + It is a superset of a known turning complete language. *)
(* *)
(* </div> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## <NAME> *)
(* *)
(* <img src="images/church.jpg" style="float:left" width="200"> *)
(* *)
(* <div style="float:left;width:75%"> *)
(* *)
(* * Developed the **λ-calculus** as a formal system for mathematical logic (1929 - 1932). *)
(* * Postulated that a function is computable (in the intuitive sense) if and only if it can be written as a lambda term (1935). *)
(* * Church was Turing's PhD advisor! *)
(* * Turing showed that the systems defined by Church and his system were equivalent. *)
(* + **Church-Turing Thesis** *)
(* *)
(* </div> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## <NAME> *)
(* *)
(* <img src="images/godel.jpg" style="float:left" width="200"> *)
(* *)
(* <div style="float:left;width:75%"> *)
(* *)
(* * Defined the class of **general recursive functions** as the smallest set of functions containing *)
(* + all the constant functions *)
(* + the successor function and *)
(* + closed under certain operations (such as compositions and recursion). *)
(* * He postulated that a function is computable (in the intuitive sense) if and only if it is general recursive. *)
(* *)
(* </div> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Impact of Church-Turing thesis *)
(* *)
(* * The **“Church-Turing Thesis”** is by itself is one of the most important ideas on computer science *)
(* + The impact of Church and Turing’s models goes far beyond the thesis itself. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Impact of Church-Turing thesis *)
(* *)
(* * Oddly, however, the impact of each has been in almost completely separate communities *)
(* + Turing Machines $\Rightarrow$ Algorithms & Complexity *)
(* + Lambda Calculus $\Rightarrow$ Programming Languages *)
(* * Not accidental *)
(* + Turing machines are quite low level $\Rightarrow$ well suited for measuring resources (**efficiency**). *)
(* + Lambda Calculus is quite high level $\Rightarrow$ well suited for abstraction and composition (**structure**). *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Programming Language Expressiveness *)
(* *)
(* * So what language features are needed to express all computable functions? *)
(* + *What's the minimal language that is Turing Complete?* *)
(* * Observe that many features that we have seen in this class were syntactic sugar *)
(* + **Multi-argument functions** - simulate using partial application *)
(* + **For loop, while loop** - simulate using recursive functions *)
(* + **Mutable heaps** - simulate using functional maps and pass around. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Functional Heap *)
(* +
type ('k,'v) heap = 'k -> 'v option
let empty_heap : ('k,'v) heap = fun k -> None
let set (h : ('k,'v) heap) (x : 'k) (v : 'v) : ('k,'v) heap =
fun k -> if k = x then Some v else h k
let get (h : ('k,'v) heap) (x : 'k) : 'v option = h x
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Functional Heap *)
(* -
let _ =
let h = set empty_heap "a" 0 in
let h = set h "b" 1 in
(get h "a", get h "b", get h "c")
(* + [markdown] slideshow={"slide_type": "fragment"}
(* * You can imagine passing around the heap as an **implicit extra argument** to every function. *)
(* + The issue of storing values of different types, default values, etc. can be orthogonally addressed. *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* <center> *)
(* *)
(* <h1 style="text-align:center"> All you need is <strike> Love</strike> <i> Functions.</i> </h1> *)
(* </center> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lambda Calculus : Syntax *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* e & ::= & x & \text{(Variable)} \\ *)
(* & \mid & \lambda x.e & \text{(Abstraction)} \\ *)
(* & \mid & e~e & \text{(Application)} *)
(* \end{array} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* * This grammar describes ASTs; not for parsing (ambiguous!) *)
(* * Lambda expressions also known as lambda **terms** *)
(* * $\lambda x.e$ is like `fun x -> e` *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* <center> *)
(* *)
(* <h2 style="text-align:center"> That's it! Nothing but higher order functions </h2> *)
(* </center> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Why Study Lambda Calculus? *)
(* *)
(* * It is a "core" language *)
(* + Very small but still Turing complete *)
(* * But with it can explore general ideas *)
(* + Language features, semantics, proof systems, algorithms, ... *)
(* * Plus, higher-order, anonymous functions (aka lambdas) are now very popular! *)
(* + C++ (C++11), PHP (PHP 5.3.0), C# (C# v2.0), Delphi (since 2009), Objective C, Java 8, Swift, Python, Ruby (Procs), ... *)
(* + and functional languages like OCaml, Haskell, F#, ... *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Three Conventions *)
(* *)
(* 1. Scope of $\lambda$ extends as far right as possible *)
(* + Subject to scope delimited by parentheses *)
(* + $\lambda x. \lambda y.x~y~$ is the same as $\lambda x.(\lambda y.(x~y))$ *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* 2. Function Application is left-associative *)
(* + `x y z` is `(x y) z` *)
(* + Same rule as OCaml *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* 3. As a convenience, we use the following syntactic sugar for local declarations *)
(* + `let x = e1 in e2` is short for $(\lambda x.e2) ~e1$. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lambda calculus interpreter in OCaml *)
(* *)
(* * In Assignment 2, you will be implementing a lambda calculus interpreter in OCaml. *)
(* * What is the Abstract Syntax Tree (AST)? *)
(* *)
(* ```ocaml *)
(* type expr = *)
(* | Var of string *)
(* | Lam of string * expr *)
(* | App of expr * expr *)
(* ``` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lambda expressions in OCaml *)
(* *)
(* * $y~$ is `Var "y"` *)
(* * $\lambda x.x~$ is `Lam ("x", Var "x")` *)
(* * $\lambda x. \lambda y.x ~y~$ is `Lam ("x",(Lam("y",App (Var "x", Var "y"))))` *)
(* * $(\lambda x.\lambda y.x ~y) ~\lambda x.x ~x~$ is *)
(* *)
(* ```ocaml *)
(* App *)
(* (Lam ("x", Lam ("y",App (Var "x", Var "y"))), *)
(* Lam ("x", App (Var "x", Var "x"))) *)
(* ``` *)
(* + slideshow={"slide_type": "slide"}
#use "init.ml";;
(* -
parse "y";;
parse "\\x.x";;
parse "\\x.\\y.x y";;
parse "(\\x.\\y.x y) \\x. x x";;
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 1 *)
(* *)
(* $\lambda x.(y ~z)$ and $\lambda x.y ~z$ are equivalent. *)
(* *)
(* 1. True *)
(* 2. False *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 1 *)
(* *)
(* $\lambda x.(y ~z)$ and $\lambda x.y ~z$ are equivalent. *)
(* *)
(* 1. True ✅ *)
(* 2. False *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 2 *)
(* *)
(* What is this term’s AST? $\lambda x.x ~x$ *)
(* *)
(* 1. `App (Lam ("x", Var "x"), Var "x")` *)
(* 2. `Lam (Var "x", Var "x", Var "x")` *)
(* 3. `Lam ("x", App (Var "x", Var "x"))` *)
(* 4. `App (Lam ("x", App ("x", "x")))` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 2 *)
(* *)
(* What is this term’s AST? $\lambda x.x ~x$ *)
(* *)
(* 1. `App (Lam ("x", Var "x"), Var "x")` *)
(* 2. `Lam (Var "x", Var "x", Var "x")` *)
(* 3. `Lam ("x", App (Var "x", Var "x"))` ✅ *)
(* 4. `App (Lam ("x", App ("x", "x")))` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 3 *)
(* *)
(* This term is equivalent to which of the following? *)
(* *)
(* $\lambda x.x ~a ~b$ *)
(* *)
(* 1. $(\lambda x.x) ~(a ~b)$ *)
(* 2. $(((\lambda x.x) ~a) ~b)$ *)
(* 3. $\lambda x.(x ~(a ~b))$ *)
(* 4. $(\lambda x.((x ~a) ~b))$ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 3 *)
(* *)
(* This term is equivalent to which of the following? *)
(* *)
(* $\lambda x.x ~a ~b$ *)
(* *)
(* 1. $(\lambda x.x) ~(a ~b)$ *)
(* 2. $(((\lambda x.x) ~a) ~b)$ *)
(* 3. $\lambda x.(x ~(a ~b))$ *)
(* 4. $(\lambda x.((x ~a) ~b))$ ✅ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Free Variables *)
(* *)
(* In *)
(* *)
(* ```ocaml *)
(* λx. x y *)
(* ``` *)
(* *)
(* * The first `x` is the binder. *)
(* * The second `x` is a **bound** variable. *)
(* * The `y` is a **free** variable. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Free Variables *)
(* *)
(* Let $FV(t)$ denote the free variables in a term $t$. *)
(* *)
(* We can define $FV(t)$ inductively over the definition of terms as follows: *)
(* *)
(* \\[ *)
(* \begin{array}{rcl} *)
(* FV(x) & = & \{x\} \\ *)
(* FV(\lambda x.t_1) & = & FV(t_1) \setminus \{x\} \\ *)
(* FV(t_1 ~t_2) & = & FV(t_1) ~\cup~ FV(t_2) *)
(* \end{array} *)
(* \\] *)
(* *)
(* If $FV(t) = \emptyset$ then we say that $t$ is a **closed** term. *)
(* *)
(* $ *)
(* \newcommand{\cg}[1]{\color{green}{#1}} *)
(* \newcommand{\cr}[1]{\color{red}{#1}} *)
(* \newcommand{\cb}[1]{\color{blue}{#1}} *)
(* $ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 4 *)
(* *)
(* What are the free variables in the following? *)
(* *)
(* 1. $\lambda x.x ~(\lambda y. y)$ *)
(* 2. $x ~y ~z$ *)
(* 3. $\lambda x. (\lambda y. y) ~x ~y$ *)
(* 4. $\lambda x. (\lambda y. x) ~y$ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 4 *)
(* *)
(* What are the free variables in the following? *)
(* *)
(* $ *)
(* \begin{array}{ll} *)
(* 1. ~\lambda x.x ~(\lambda y. y) & \{\} \\ *)
(* 2. ~\cr{x ~y ~z} & \{x,y,z\} \\ *)
(* 3. ~\lambda x. (\lambda y. y) ~x ~\cr{y} & \{y\} \\ *)
(* 4. ~\lambda x. (\lambda y. x) ~\cr{y} & \{y\} *)
(* \end{array} *)
(* $ *)
(* + slideshow={"slide_type": "slide"}
free_variables "\\x.x (\\y. y)";;
free_variables "x y z";;
free_variables "\\x.(\\y. y) x y";;
free_variables "\\x.(\\y.x) y";;
(* + [markdown] slideshow={"slide_type": "slide"}
(* # $\alpha$-equivalence *)
(* *)
(* Lambda calculus uses **static scoping** (just like OCaml) *)
(* *)
(* \\[ *)
(* \lambda \cg{x}. \cg{x} ~(\lambda \cr{x}. \cr{x}) *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* This is equivalent to: *)
(* *)
(* \\[ *)
(* \lambda \cg{x}. \cg{x} ~(\lambda \cr{y}. \cr{y}) *)
(* \\] *)
(* *)
(* * Renaming bound variables consistently preserves meaning *)
(* + This is called as **𝛼-renaming** or **𝛼-conversion**. *)
(* * If a term $t_1$ is obtained by 𝛼-renaming another term $t_2$ then $t_1$ and $t_2$ are said to be **𝛼-equivalent**. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 5 *)
(* *)
(* Which of the following equivalences hold? *)
(* *)
(* 1. $\lambda x. x ~(\lambda y. y) ~y =_{\alpha} \lambda y. y ~(\lambda x. x) ~x$ *)
(* 2. $\lambda x. x ~(\lambda y. y) ~y =_{\alpha} \lambda y. y ~(\lambda x. x) ~y$ *)
(* 3. $(\lambda x. x ~(\lambda y. y) ~y) =_{\alpha} \lambda w. w ~(\lambda w. w) ~y$ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz 5 *)
(* *)
(* Which of the following equivalences hold? *)
(* *)
(* 1. $\lambda x. x ~(\lambda y. y) ~y =_{\alpha} \lambda y. y ~(\lambda x. x) ~x~$ ❌ *)
(* 2. $\lambda x. x ~(\lambda y. y) ~y =_{\alpha} \lambda y. y ~(\lambda x. x) ~y~$ ❌ *)
(* 3. $\lambda x. x ~(\lambda y. y) ~y =_{\alpha} \lambda w. w ~(\lambda w. w) ~y~$ ✅ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution *)
(* *)
(* * In order to formally define $\alpha$-equivalence, we need to define **substitutions**. *)
(* * Substitution replaces **free** occurrences of a variable $x$ with a lambda term $N$ in some other term $M$. *)
(* + We write it as $M[N/x]$. (read "N for x in M"). *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* For example, *)
(* *)
(* \\[ *)
(* (\lambda x.x ~y)[(\lambda z.z)/y] = \lambda x.x ~(\lambda z.z) *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* <h4> Substitution is quite subtle. So we will start with our intuitions and see how things break and finally work up to the correct example. <h4> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution: Take 1 *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* x[s/x] & = & s \\ *)
(* y[s/x] & = & y & \text{if } x \neq y\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda y.t_1[s/x] \\ *)
(* (t_1 ~t_2)[s/x] & = & (t_1[s/x]) ~(t_2[s/x]) *)
(* \end{array} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* This definition works for most examples. For example, *)
(* *)
(* \\[ *)
(* (\lambda y.x)[(\lambda z.z~w)/x] = \lambda y.\lambda z.z ~w *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution: Take 1 *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* x[s/x] & = & s \\ *)
(* y[s/x] & = & y & \text{if } x \neq y\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda y.t_1[s/x] \\ *)
(* (t_1 ~t_2)[s/x] & = & (t_1[s/x]) ~(t_2[s/x]) *)
(* \end{array} *)
(* \\] *)
(* *)
(* However, it fails if the substitution is on the bound variable: *)
(* *)
(* \\[ *)
(* (\lambda x.x)[y/x] = \lambda x.y *)
(* \\] *)
(* *)
(* The **identity** function has become a **constant** function! *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution: Take 2 *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* x[s/x] & = & s \\ *)
(* y[s/x] & = & y & \text{if } x \neq y\\ *)
(* (\lambda x.t_1)[s/x] & = & \lambda x.t_1\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda y.t_1[s/x] & \text{if } x \neq y\\ *)
(* (t_1 ~t_2)[s/x] & = & (t_1[s/x]) ~(t_2[s/x]) *)
(* \end{array} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* However, this is not quite right. For example, *)
(* *)
(* \\[ *)
(* (\lambda x.y)[x/y] = \lambda x.x *)
(* \\] *)
(* *)
(* * The **constant** function has become a **identity** function. *)
(* * The problem here is that the free $x$ gets **captured** by the binder $x$. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution: Take 3 *)
(* *)
(* Capture-avoiding substitution *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* x[s/x] & = & s \\ *)
(* y[s/x] & = & y & \text{if } x \neq y\\ *)
(* (\lambda x.t_1)[s/x] & = & \lambda x.t_1\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda y.t_1[s/x] & \text{if } x \neq y \text{ and } y \notin FV(s)\\ *)
(* (t_1 ~t_2)[s/x] & = & (t_1[s/x]) ~(t_2[s/x]) *)
(* \end{array} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* * Unfortunately, this made substitution a partial function *)
(* + There is no valid rule for $(\lambda x.y)[x/y]$ *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Substitution: Take 4 *)
(* *)
(* Capture-avoiding substitution + totality *)
(* *)
(* \\[ *)
(* \begin{array}{rcll} *)
(* x[s/x] & = & s \\ *)
(* y[s/x] & = & y & \text{if } x \neq y\\ *)
(* (\lambda x.t_1)[s/x] & = & \lambda x.t_1\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda y.t_1[s/x] & \text{if } x \neq y \text{ and } y \notin FV(s)\\ *)
(* (\lambda y.t_1)[s/x] & = & \lambda w.t_1[w/y][s/x] & \text{if } x \neq y \text{ and } y \in FV(s) \text { and } w \text{ is fresh}\\ *)
(* (t_1 ~t_2)[s/x] & = & (t_1[s/x]) ~(t_2[s/x]) *)
(* \end{array} *)
(* \\] *)
(* *)
(* * A **fresh** binder is different from every other binder in use **(generativity)**. *)
(* * In the case above, *)
(* *)
(* \\[ *)
(* w \text{ is fresh } \equiv w \notin FV(t_1) \cup FV(s) \cup \{x\} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* Now our example works out: *)
(* *)
(* \\[ *)
(* (\lambda x.y)[x/y] = \lambda w.x *)
(* \\] *)
(* + slideshow={"slide_type": "slide"}
substitute "\\y.x" "x" "\\z.z w"
(* -
substitute "\\x.x" "x" "y"
substitute "\\x.y" "y" "x"
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## $\alpha$-equivalence formally *)
(* *)
(* $=_{\alpha}$ is an equivalence (reflexive, transitive, symmetric) relation such that: *)
(* *)
(* $ *)
(* \newcommand{\inferrule}[2]{\displaystyle{\frac{#1}{#2}}} *)
(* $ *)
(* *)
(* \\[ *)
(* \begin{array}{cc} *)
(* \inferrule{}{x =_{\alpha} x} \quad & \quad \inferrule{M =_{\alpha} M' \quad N =_{\alpha} N'}{M ~N =_{\alpha} M' ~N'} *)
(* \end{array} *)
(* \\] *)
(* *)
(* <br> *)
(* *)
(* \\[ *)
(* \inferrule{z \notin FV(M) \cup FV(N) \quad M[z/x] =_{\alpha} N[z/y]}{\lambda x.M =_{\alpha} \lambda y.N} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Convention *)
(* *)
(* From now on, *)
(* *)
(* * Unless stated otherwise, we identify lambda terms up to α-equivalence. *)
(* + when we speak of lambda terms being **equal**, we mean that they are α-equivalent *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* <center> *)
(* *)
(* <h1 style="text-align:center"> Fin. </h1> *)
(* </center> *)
|
lectures/lec8/lec8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib as plt
import seaborn as sns
from sklearn.svm import SVC
# ## Question 5) String kernels
# *Note: data taken from https://cseweb.ucsd.edu/classes/wi17/cse151-a/hw5.pdf along with inspiration for this problem.*
#
# As we've seen, kernels have wide applications. We've seen kernels be a time efficient way to lift a feature matrix in many different contexts like PCA, Perceptrons, SVM's and Ridge Regression. One great property of kernels is the ability to supply any valid kernel function to one of our kernelized model it will work within that space.
# For this problem, we will be working with **string kernels**, which apply kernel functions to text data so that models like SVM can work with them. Remember kernel functions are essentially similarity functions. Therefore, string kernel functions generally tell us how similar two strings are.
#
# ### Question 5a)
# First, let's take a look at our data.
train_data = pd.read_csv('string_train.csv')
train_data.head()
# The sequence is a sequence of amino acids, and the classification is whether or not the sequence belongs to a particular protein family or not.
# Visualize the number of positive and negative sequences we have using a barplot.
# +
# START TODO
# END TODO
# -
# Answer the following questions.
# * **Brainstorm some ways to deal with this text data. How can we compute similarity between two strings?**
# * **Does the imbalance in data matter? Why or why not?**
# ### Question 5b)
# For this problem, we are going to be using the **Spectrum Kernel**, one of the most simple string kernels. The basic idea behind the kernel is that two strings with more common substrings will be more similar. What is a substring? A substring is any contiguous sequence of characters within a string. The kernel function the **p-spectrum kernel** expresses this idea. $k(s_1, s_2)$ counts all size $p$ substrings that are present in both the string $s_1$ and the string $s_2$.
#
# Fill out the code below to finish the implementation of the kernel.
#
#
# *Note 1: There are many other valid string kernels, many of which are much better and domain specific than this one. For more string kernels, check here https://people.eecs.berkeley.edu/~jordan/kernels/0521813972c11_p344-396.pdf. In fact, <NAME> has a whole book on kernels for pattern recognition that might be fun to look at.*
# *Note 2: Section 1.4.6.2.2 https://scikit-learn.org/stable/modules/svm.html#kernel-functions might be useful.*
def p_spectrum(s, t, p):
num_in_common = ...
# START TODO
num_in_common = 0
for i in range(len(s) - p + 1):
curr = s[i:i+p]
if curr in t:
num_in_common += 1
# END TODO
return num_in_common
# Now that we have our kernel function defined, let's actually use it to classify our dataset. We will be using an SVM classifier. Unfortunately, scikit learn doesn't support string kernel functions for SVM's, but there are a few work arounds. The first work around is precomputing the **Gram matrix** $K$ for our data, and then running an SVM on it. Remember, all we need is the Gram matrix to do classification.
#
# *Note: it will take a few seconds to a minute to run this part of the code.*
# +
X_train = np.array(train_data["Sequence"])
y_train = np.array(train_data["Classification"])
def compute_gram_matrix(X_one, X_two, y, p=3):
# START COMPUTE GRAM MATRIX
K = np.zeros((X_one.shape[0], X_two.shape[0]))
for s_index, s in enumerate(X_one):
for t_index, t in enumerate(X_two):
K[s_index][t_index] = p_spectrum(s, t, p)
# END COMPUTE GRAM MATRIX
return K
# -
K_train = ...
# START TODO
K_train = compute_gram_matrix(X_train, X_train, y_train)
# END TODO
print (K_train.shape)
K_train
# ### Question 5c)
# Now let's try some classifiers out. Let's start with the SVM classifier. Complete the below code to fit an SVM model with our precomputed Gram matrix and print out the accuracy on the train set.
# +
clf = SVC(kernel='precomputed')
clf.fit(K_train, y_train)
score = clf.score(K_train, y_train)
print ("Accuracy on the training data: " + str(score))
# -
# Now calculate the accuracy on the testing data.
# +
test_data = pd.read_csv('string_test.csv')
X_test = np.array(test_data["Sequence"])
y_test = np.array(test_data["Classification"])
# START TODO
K_test = compute_gram_matrix(X_test, X_train, y_test)
y_pred = clf.predict(K_test)
# END TODO
score = clf.score(K_test, y_test)
print ("Accuracy on the testing data: " + str(score))
# -
# Answer the following questions.
# * **What was your accuracy on the test dataset? Was it different from the training dataset?**
# * **Can you think of any improvements to our model?**
# Congrats! You just used string kernels to classify a real dataset of Amino Acid sequences! Hopefully you can see the power of kernels from this example. Further applications include Graph kernels, Tree kernels, kernels for images... the possibilities are endless!
# If you want to learn more, read some of <NAME>'s stuff on this subject here: https://people.eecs.berkeley.edu/~jordan/kernels/0521813972pre_pi-xiv.pdf
# If you're interested in string kernels, here are some good papers to read that build upon the simple p-spectrum kernel we used in this example!
# * Mismatch kernels - https://papers.nips.cc/paper/2179-mismatch-string-kernels-for-svm-protein-classification.pdf
# * Gappy kernels - https://www.semanticscholar.org/paper/A-fast-%2C-large-scale-learning-method-for-protein-Kuksa-Huang/bd5a49164b7d0a9179ef5cb39148279825877a7f
# * Motif kernels - https://almob.biomedcentral.com/articles/10.1186/1748-7188-1-21
# * More spectrum kernels - https://pubmed.ncbi.nlm.nih.gov/11928508/
|
final_kernels_project_blank/prob5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import scipy as stats
import seaborn as sns
import pandas_profiling
titanic=sns.load_dataset('titanic')
titanic.tail(2)
titanic.tail(2)
titanic.groupby('sex').count()
titanic.groupby('sex')[['survived']].sum()
titanic.groupby('sex')[['survived']].aggregate('sum').unstack()
titanic.groupby(['sex','class'])[['survived']].sum().unstack()
#pivote Table syntax
titanic.pivot_table('survived',columns='class')
titanic.pivot_table('survived',index='sex',columns='class')
#multilevel pivote table
age=pd.cut(titanic['age'],[0,18,30,80])
titanic.pivot_table('survived',['sex',age,'who'],'class')
fare=pd.qcut(titanic['fare'],4)
titanic.pivot_table('survived',['sex',age,'who'],[fare,'class'])
#additional pivote table
titanic.pivot_table(index='sex',columns='class',
aggfunc={'survived':sum,'fare':'min'})
titanic.pivot_table('survived',index='sex',columns='class',margins=True)
|
pd_pioting_table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# The general environment setup
#
# Importing libraries
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import csv #to convert to csv files
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# -
# **In this code an example file 'datahosp.txt' is created in the format as specified with details of 5 patients and is added as test data.
# In the first cell below the file read and converted into list.
# Each line is seperated using split function and the new list contains each line seperated by commas as an element.**
df = open('../input/incubyte/datahosp.txt', "r")
lines=df.read()
lines=list(lines.split('\n'))
d=[]
for i in lines:
line=i.split('|')
d.append(line)
for i in range(len(d)):
del d[i][0]
print(d)
df.close()
# **The above data is converted into a csv file by creating a new csv file in write mode as shown below:**
with open('protagonist.csv', 'w', newline='') as file:
writer = csv.writer(file)
writer.writerows(d)
meta=pd.read_csv('protagonist.csv')
meta.head()
# **To sort according to countries we use below code to get list of countries to use in queries:**
index=meta['Country'].unique()
f=dict(enumerate(index))
print(f)
# **To create seperate tables for each country we can use below conditions.Two tables are created as examples:**
TableINDIA=meta[meta['Country']==index[1]]
TableUSA=meta[meta['Country']==index[0]]
print(TableUSA)
print(TableINDIA)
# **We can also filter data by generating queries.For example below code is example of query used to get data using patient name:**
print(meta.query('Customer_Name=="Alex"'))
|
patientdetails.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
# import some dependencies
import numpy as np
import matplotlib.pyplot as plt
try:
import seaborn as sns
sns.set()
except ImportError:
pass
import torch
from torch.autograd import Variable
import pyro
import pyro.infer
import pyro.optim
import pyro.distributions as dist
torch.manual_seed(101)
# -
# # Inference in Pyro: From Stochastic Functions to Marginal Distributions
#
# Stochastic functions induce a joint probability distribution $p(y, z \; \vert \; x)$ over their latent variables $z$ and return values $y$, and this joint distribution induces a marginal distribution over return values of the function. However, for non-primitive stochastic functions, we can no longer explicitly compute the marginal probability of an output $p(y \; \vert \; x)$ or draw samples from the marginal distribution over return values $y \sim p (y \; \vert \; x)$.
#
# In its most general formulation, *inference* in a universal probabilistic programming language like Pyro is the problem of constructing this marginal distribution given an arbitrary boolean constraint so that we can perform these computations. The constraint can be a deterministic function of the return value, the internal randomness, or both.
#
# *Bayesian inference* or *posterior inference* is an important special case of this more general formulation that admits tractable approximations. In Bayesian inference, the return value is always the values of some subset internal `sample` statements, and the constraint is an equality constraint on the other internal `sample` statements. Much of modern machine learning can be cast as approximate Bayesian inference and expressed succinctly in a language like Pyro.
#
# To motivate the rest of this tutorial, let's first build a generative model for a simple physical problem so that we can use Pyro's inference machinery to solve it.
# ## A Simple Example
#
# Suppose we are trying to figure out how much something weighs, but the scale we're using is unreliable and gives slightly different answers every time we weigh the same object. We could try to compensate for this variability by integrating the noisy measurement information with a guess based on some prior knowledge about the object, like its density or material properties. The following model encodes this process:
def scale(guess):
# The prior over weight encodes our uncertainty about our guess
weight = pyro.sample("weight", dist.normal, guess, Variable(torch.ones(1)))
# This encodes our belief about the noisiness of the scale:
# the measurement fluctuates around the true weight
return pyro.sample("measurement", dist.normal, weight, Variable(torch.Tensor([0.75])))
# ## Representing Marginal Distributions
#
# Before we actually try using our model to estimate an object's weight, let's try analyzing our model's behavior. In particular, we can use importance sampling to simulate the marginal distribution of measurement values we'd expect to see a priori for a given guess.
#
# Marginalization in Pyro with `pyro.infer.Marginal` is split into two steps. First, we collect a number of weighted execution traces of the model. Then, we can collapse those traces into a histogram over possible return values given a particular set of arguments.
#
# Collecting execution traces can be done either through sampling or, for models with only discrete latent variables, exact enumeration. To create a basic importance sampler over execution traces (using the prior as the proposal distribution), we can write:
posterior = pyro.infer.Importance(scale, num_samples=100)
# `posterior` is not a particularly useful object on its own, though advanced users can call it with arguments for `scale` to sample a raw execution trace. Instead, `posterior` is meant to be consumed by `pyro.infer.Marginal`, which creates a primitive stochastic function with the same input and output types as `scale`.
# +
guess = Variable(torch.Tensor([8.5]))
marginal = pyro.infer.Marginal(posterior)
print(marginal(guess))
# -
# When called with an input `guess`, `marginal` first uses `posterior` to generate a sequence of weighted execution traces given `guess`, then builds a histogram over return values from the traces, and finally returns a sample drawn from the histogram. Calling `marginal` with the same arguments more than once will sample from the same histogram.
plt.hist([marginal(guess).data[0] for _ in range(100)], range=(5.0, 12.0))
plt.title("P(measurement | guess)")
plt.xlabel("weight")
plt.ylabel("#")
# `pyro.infer.Marginal` also accepts the optional keyword argument `sites=[name1, name2, ...]` that provides a list of names of latent variables. When `sites` is specified, `marginal` will return a dictionary where the keys are the names in `sites` and the values are values at those sites from a single execution trace. This is useful because we may wish to compute many different marginals from the same posterior object.
# ## Conditioning Models on Data
#
# The real utility of probabilistic programming is in the ability to condition generative models on observed data and infer the latent factors that might have produced that data. In Pyro, we separate the expression of conditioning from its evaluation via inference, making it possible to write a model once and condition it on many different observations. Pyro supports constraining a model's internal `sample` statements to be equal to a given set of observations.
#
# Consider `scale` once again. Suppose we want to sample from the marginal distribution of `weight` given input `guess = 8.5`, but now we have observed that `measurement == 9.5`. Pyro provides the function `pyro.condition` to allow us to constrain the values of sample statements. `pyro.condition` is a higher-order function that takes a model and a dictionary of data and returns a new model that has the same input and output signatures but always uses the given values at observed `sample` statements:
conditioned_scale = pyro.condition(
scale, data={"measurement": Variable(torch.Tensor([9.5]))})
# Because it behaves just like an ordinary Python function, conditioning can be deferred or parametrized with Python's `lambda` or `def`:
def deferred_conditioned_scale(measurement, *args, **kwargs):
return pyro.condition(scale, data={"measurement": measurement})(*args, **kwargs)
# In some cases it might be more convenient to pass observations directly to individual `pyro.sample` statements instead of using `pyro.condition`. The optional `obs` keyword argument is reserved by `pyro.sample` for that purpose; in addition, `pyro.observe` is an alias for `pyro.sample` with `obs`:
# +
# equivalent to pyro.condition(scale, data={"measurement": Variable(torch.ones(1))})
def scale_obs(guess):
weight = pyro.sample("weight", dist.normal, guess, Variable(torch.ones(1)))
# here we attach an observation measurement == 9.5
return pyro.sample("measurement", dist.normal, weight, Variable(torch.ones(1)),
obs=Variable(torch.Tensor([9.5])))
# equivalent to scale_obs:
def scale_obs(guess):
weight = pyro.sample("weight", dist.normal, guess, Variable(torch.ones(1)))
# here we attach an observation measurement == 9.5
return pyro.observe("measurement", dist.normal, Variable(torch.Tensor([9.5])),
weight, Variable(torch.ones(1)))
# -
# However, hardcoding is not usually recommended due to its invasive non-compositional nature. By contrast, using `pyro.condition`, conditioning may be composed freely to form multiple complex queries on probabilistic models without modifying the underlying model. The only restriction is that a single site may only be constrained once.
# +
def scale2(guess):
weight = pyro.sample("weight", dist.normal,
guess, Variable(torch.ones(1)))
tolerance = torch.abs(
pyro.sample("tolerance", dist.normal, Variable(torch.zeros(1)), Variable(torch.ones(1))))
return pyro.sample("measurement", dist.normal, weight, tolerance)
# conditioning composes:
# the following are all equivalent and do not interfere with each other
conditioned_scale2_1 = pyro.condition(
pyro.condition(scale2, data={"weight": Variable(torch.Tensor([9.2]))}),
data={"measurement": Variable(torch.Tensor([9.5]))})
conditioned_scale2_2 = pyro.condition(
pyro.condition(scale2, data={"measurement": Variable(torch.Tensor([9.5]))}),
data={"weight": Variable(torch.Tensor([9.2]))})
conditioned_scale2_3 = pyro.condition(
scale2, data={"weight": Variable(torch.Tensor([9.2])),
"measurement": Variable(torch.Tensor([9.5]))})
# -
# In addition to `pyro.condition` for incorporating observations, Pyro also contains `pyro.do`, an implementation of Pearl's `do`-operator used for causal inference with an identical interface to `pyro.condition`. `condition` and `do` can be mixed and composed freely, making Pyro a powerful tool for model-based causal inference. See the [causal inference tutorial](http://pyro.ai/examples/causal_inference.html) for more details about `pyro.do` and a simple example of causal inference in a model of disease diagnosis.
# ## Flexible Approximate Inference With Guide Functions
#
# Let's return to `deferred_conditioned_scale`. Now that we have constrained `measurement` against some data, we can use Pyro's approximate inference algorithms to estimate the distribution over `weight` given `guess` and `measurement == data`. We saw earlier how to use importance sampling to do this for `scale`; we can use exactly the same constructs with a conditioned model:
# +
guess = Variable(torch.Tensor([8.5]))
measurement = Variable(torch.Tensor([9.5]))
conditioned_scale = pyro.condition(scale, data={"measurement": measurement})
marginal = pyro.infer.Marginal(
pyro.infer.Importance(conditioned_scale, num_samples=100), sites=["weight"])
# The marginal distribution concentrates around the data
print(marginal(guess))
plt.hist([marginal(guess)["weight"].data[0] for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#")
# -
# However, this approach is extremely computationally inefficient because the prior distribution over `weight` may be very far from the true distribution over weights, especially if our initial `guess` is not very good.
#
# Therefore, some inference algorithms in Pyro, like `pyro.infer.Importance` and `pyro.infer.SVI`, allow us to use arbitrary stochastic functions, which we will call *guide functions* or *guides*, as approximate posterior distributions. Guide functions must satisfy these two criteria to be valid approximations for a particular model:
# 1. all unobserved sample statements that appear in the model appear in the guide.
# 2. the guide has the same input signature as the model (i.e. takes the same arguments)
#
# Guide functions can serve as programmable, data-dependent proposal distributions for importance sampling, rejection sampling, sequential Monte Carlo, MCMC, and independent Metropolis-Hastings, and as variational distributions or inference networks for stochastic variational inference. Currently, only importance sampling and stochastic variational inference are implemented in Pyro, but we plan to add other algorithms in the future.
#
# Although the precise meaning of the guide is different across different inference algorithms, the guide function should generally be chosen so that it closely approximates the distribution over all unobserved `sample` statements in the model. The simplest guide for `deferred_conditioned_scale` matches the prior distribution over `weight`:
# +
def scale_prior_guide(guess):
return pyro.sample("weight", dist.normal, guess, Variable(torch.ones(1)))
posterior = pyro.infer.Importance(conditioned_scale,
guide=scale_prior_guide,
num_samples=10)
marginal = pyro.infer.Marginal(posterior, sites=["weight"])
# -
# Can we do better than the prior? In the case of `scale`, it turns out that the true posterior distribution over `weight` given `guess` and `measurement` can be written directly as:
# +
def scale_posterior_guide(measurement, guess):
# note that torch.size(measurement, 0) is the total number of measurements
# that we're conditioning on
a = (guess + torch.sum(measurement)) / (measurement.size(0) + 1.0)
b = Variable(torch.ones(1)) / (measurement.size(0) + 1.0)
return pyro.sample("weight", dist.normal, a, b)
posterior = pyro.infer.Importance(deferred_conditioned_scale,
guide=scale_posterior_guide,
num_samples=20)
marginal = pyro.infer.Marginal(posterior, sites=["weight"])
plt.hist([marginal(measurement, guess)["weight"].data[0] for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#")
# -
# ## Parametrized Stochastic Functions and Variational Inference
#
# Although we could write out the exact posterior distribution for `scale`, in general it is intractable to specify a guide that is a good approximation to the posterior distribution of an arbitrary conditioned stochastic function. What we can do instead is use the top-level function `pyro.param` to specify a *family* of guides indexed by named parameters, and search for the member of that family that is the best approximation. This approach to approximate posterior inference is called *variational inference*.
#
# `pyro.param` is a frontend for Pyro's key-value *parameter store*, which is described in more detail in the documentation. Like `pyro.sample`, `pyro.param` is always called with a name as its first argument. The first time `pyro.param` is called with a particular name, it stores its argument in the parameter store and then returns that value. After that, when it is called with that name, it returns the value from the parameter store regardless of any other arguments. It is similar to `simple_param_store.setdefault` here, but with some additional tracking and management functionality.
#
# ```python
# simple_param_store = {}
# a = simple_param_store.setdefault("a", torch.randn(1))
# ```
#
# For example, we can parametrize `a` and `b` in `scale_posterior_guide` instead of specifying them by hand:
def scale_parametrized_guide(guess):
a = pyro.param("a", Variable(torch.randn(1) + guess.data.clone(), requires_grad=True))
b = pyro.param("b", Variable(torch.randn(1), requires_grad=True))
return pyro.sample("weight", dist.normal, a, torch.abs(b))
# Pyro is built to enable *stochastic variational inference*, a powerful and widely applicable class of variational inference algorithms with three key characteristics:
# 1. Parameters are always real-valued tensors
# 2. We compute Monte Carlo estimates of a loss function from samples of execution histories of the model and guide
# 3. We use stochastic gradient descent to search for the optimal parameters.
#
# Combining stochastic gradient descent with PyTorch's GPU-accelerated tensor math and automatic differentiation allows us to scale variational inference to very high-dimensional parameter spaces and massive datasets.
#
# Pyro's SVI functionality is described in detail in the [SVI tutorial](http://pyro.ai/examples/svi_part_i.html). Here is a very simple example applying it to `scale`:
# +
pyro.clear_param_store()
svi = pyro.infer.SVI(model=conditioned_scale,
guide=scale_parametrized_guide,
optim=pyro.optim.SGD({"lr": 0.001}),
loss="ELBO")
losses = []
for t in range(1000):
losses.append(svi.step(guess))
plt.plot(losses)
plt.title("ELBO")
plt.xlabel("step")
plt.ylabel("loss")
# -
# Note that optimization will update the guide parameters, but does not produce a posterior distribution object itself. Once we find good parameter values, we can just use the guide as a representation of the model's approximate posterior for downstream tasks.
#
# For example, we can use the optimized guide as an importance distribution for estimating the marginal distribution over `weight` with many fewer samples than the prior:
# +
posterior = pyro.infer.Importance(conditioned_scale, scale_parametrized_guide, num_samples=10)
marginal = pyro.infer.Marginal(posterior, sites=["weight"])
plt.hist([marginal(guess)["weight"].data[0] for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#")
# -
# We can also sample from the guide directly as an approximate posterior:
plt.hist([scale_parametrized_guide(guess).data[0] for _ in range(100)], range=(5.0, 12.0))
plt.title("P(weight | measurement, guess)")
plt.xlabel("weight")
plt.ylabel("#")
# ## Next Steps
#
# In the [Variational Autoencoder tutorial](http://pyro.ai/examples/vae.html), we'll see how models like `scale` can be augmented with deep neural networks and use stochastic variational inference to build a generative model of images.
|
tutorial/source/intro_part_ii.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression Project - Ecommerce clothing store
# **<NAME>**
#
# This is a project for [Machine Learning Bootcamp](https://www.udemy.com/course/python-for-data-science-and-machine-learning-bootcamp) course on Udemy. All data being used are generated(fake).
#
# An Ecommerce company based in New York City that sells clothing online is at a fork in the road. Although, it is an online clothing store, they also have in-store style and clothing advice sessions. Customers come in to the store, have sessions/meetings with a personal stylist, then they can go home and order either on a mobile app or website for the clothes they want.
#
#
# The company is trying to decide whether to focus their efforts on their mobile app experience or their website. We'll explore the data and apply linear regression technique to provide the company with the insights to better their decision process
# ## Imports
#
# Let's make the necessary imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ## Read the data
#
cust_data = pd.read_csv('../input/new-york-ecommerce-customers/Ecommerce Customers')
# The csv has Customer info, such as Email, Address, and their color Avatar. It also has numerical value columns:
#
# * **Avg. Session Length**: Average session of in-store style advice sessions.
# * **Time on App**: Average time spent on App in minutes
# * **Time on Website**: Average time spent on Website in minutes
# * **Length of Membership**: How many years the customer has been a member.
#
# Let's explore the data a little bit.
cust_data.head()
cust_data.describe()
cust_data.info()
cust_data.isnull().sum()
#the dataset has no null values.
plt.rcParams["patch.force_edgecolor"] = True
sns.set_style('whitegrid')
sns.jointplot(x='Time on Website',y='Yearly Amount Spent',data=cust_data,kind='hex')
sns.jointplot(x='Time on App',y='Yearly Amount Spent',data=cust_data,kind='hex')
# By looking at both the graphs above, we see that the time spent on the mobile app is far lesser than time spent on the website for the same amount of money spent. We can double check this relationship by plotting a heatmap of the correlations of data columns.
sns.heatmap(cust_data.corr(),cmap = 'GnBu', annot=True)
# Between the App and the website, the correlation with the yearly amount spent is much higher in the case of App at 0.5 as compared to the website which is -0.0026. Another finding is the Length of membership has the highest correlation at 0.81.
sns.lmplot(x='Length of Membership',y='Yearly Amount Spent',data=cust_data)
plt.xlim(0,cust_data['Length of Membership'].max()+1)
cust_data.columns
# ## Splitting the data
#
# We split the dataset into train and test data.
X = cust_data[[ 'Avg. Session Length', 'Time on App', 'Time on Website', 'Length of Membership']]
y = cust_data['Yearly Amount Spent']
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state=101)
# ## Training the model
from sklearn.linear_model import LinearRegression
linear_model = LinearRegression()
linear_model.fit(X_train,y_train)
# ## Model predictions
#
# Let's test the model against the true y values
predictions = linear_model.predict(X_test)
plt.scatter(y_test,predictions,edgecolors='r')
plt.xlabel('Y test')
plt.ylabel('Predicted Y values')
# ## Conclusion
#
# Let's evaluate the co-efficients
cust_coef_df = pd.DataFrame(linear_model.coef_,index=X.columns,columns=['Coefficient'])
cust_coef_df
# Interpreting the coefficients:
#
# * Holding all other features fixed, a 1 unit increase in Avg. Session Length is associated with an increase of **25.98** total dollars spent.
# * Holding all other features fixed, a 1 unit increase in Time on App is associated with an increase of **38.59** total dollars spent.
# * Holding all other features fixed, a 1 unit increase in Time on Website is associated with an increase of **0.19** total dollars spent.
# * Holding all other features fixed, a 1 unit increase in Length of Membership is associated with an increase of **61.27** total dollars spent.
# > Answer : By the data we see that each minute spent on the App would result in more dollars spent by the customer, the company has a choice, it can either focus entirely on the app or improve the website as it is performing poorly. In addition to the above, the company can also introduce measures to lengthen the existing customers' memberships.
# #### **Thanks for reading!**
#
# #### **Cheers!**
#
# <NAME>
|
Ecommerce Clothing store/linear-regression-project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Experiment 1.2 - Xgboost with hyperparameter tuning
#
# Use a method to tune hyperparameters for the xgboost model.
#
# https://www.mikulskibartosz.name/xgboost-hyperparameter-tuning-in-python-using-grid-search/
#
# https://machinelearningmastery.com/tune-number-size-decision-trees-xgboost-python/
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
# import load_data function from helper file
# %load_ext autoreload
# %autoreload 2
# fix system path
import sys
sys.path.append("/home/jovyan/work")
# +
from src.features.helper_functions import load_sets
X_train, y_train, X_val, y_val, X_test = load_sets()
# -
print(X_train)
print(X_val)
# ## Create grid search parameters
# pip install xgboost -- run this if xgboost doesn't load
from xgboost import XGBClassifier
from sklearn.model_selection import GridSearchCV
# instatiate xgboost
model = XGBClassifier(seed=42)
# define parameters to try
parameters = {
'max_depth': range (2, 10, 1),
'n_estimators': range(60, 220, 40),
'learning_rate': [0.1, 0.01, 0.05]
}
grid_search = GridSearchCV(
estimator=model,
param_grid=parameters,
scoring = 'roc_auc', # use roc_auc to score
n_jobs = -1, # use all available processors
cv = 10, # no of folds for cross validation
verbose=3 # display all info as the model trains
)
# train
grid_result = grid_search.fit(X_train, y_train)
# +
# from sklearn.model_selection import StratifiedKFold
# kfold = StratifiedKFold(n_splits=10, shuffle=True, random_state=7)
# summarise results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
# -
# ## Train the model using best parameters
# Train using best parameters
model = XGBClassifier(max_depth = 2, n_estimators = 140, learning_rate = 0.05)
model_best_params = model.fit(X_train, y_train)
from src.features.helper_functions import save_model
save_model(model_best_params, 'rex_xgb_gridsearch')
# ## Make predictions using validation set
# predict class
y_train_preds = model_best_params.predict(X_train)
y_val_preds = model_best_params.predict(X_val)
# predict proabilities
y_train_preds_prob = model_best_params.predict_proba(X_train)
y_val_preds_prob = model_best_params.predict_proba(X_val)
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
# +
print(mse(y_train, y_train_preds, squared=False))
print(mae(y_train, y_train_preds))
print(mse(y_val, y_val_preds, squared=False))
print(mae(y_val, y_val_preds))
# +
from sklearn.metrics import accuracy_score
accuracy = accuracy_score(y_val, y_val_preds)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# -
from sklearn.metrics import roc_auc_score ,recall_score, precision_score
precision=precision_score(y_val, y_val_preds)
recall=recall_score(y_val, y_val_preds)
roc=roc_auc_score(y_val, y_val_preds)
print("Precision: %.2f%% " % (precision *100))
print("Recall: %.2f%% " % (recall * 100))
print("AUC: %.2f%% " % (roc *100))
roc_auc_score(y_train, y_train_preds_prob[:,1])
roc_auc_score(y_val, y_val_preds_prob[:,1])
# +
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(model_best_params, X_val, y_val, cmap=plt.cm.Blues)
plt.show()
# +
from sklearn.metrics import classification_report
class_report = classification_report(y_val, y_val_preds)
print(class_report)
# -
# About 250 are being falsely classed as 1, whereas only 11 are being wrongly classed as 0. This may be due to the imbalance majority classs 1, overpowering the 0's.
# Plot roc curve using code from here https://www.analyticsvidhya.com/blog/2020/06/auc-roc-curve-machine-learning/
# +
from sklearn.metrics import roc_curve
# roc curve for models
fpr, tpr, thresh = roc_curve(y_val, y_val_preds_prob[:,1], pos_label=1)
# roc curve for tpr = fpr
random_probs = [0 for i in range(len(y_val))]
p_fpr, p_tpr, _ = roc_curve(y_val, random_probs, pos_label=1)
# +
# use matplotlib to plot ROC curve
plt.style.use('seaborn')
# plot roc curves
plt.plot(fpr, tpr, linestyle='--',color='orange', label='XGBoost best params')
plt.plot(p_fpr, p_tpr, linestyle='--', color='blue')
# title
plt.title('ROC curve')
# x label
plt.xlabel('False Positive Rate')
# y label
plt.ylabel('True Positive rate')
plt.legend(loc='best')
plt.savefig('../src/visualisation/Rez XGBoost best params',dpi=300)
plt.show()
# -
# Slight improvement over base model, the AUC is higher at 0.702, but that is still under the current best score on the leaderboard.
#
# More paramaters can be tuned, try a few more before moving onto a different model e.g. SVM.
# ## Make predictions with test set for Kaggle submission
y_test_preds = model_best_params.predict(X_test)
y_test_preds_prob = model_best_params.predict_proba(X_test)
print(y_test_preds)
print(y_test_preds_prob)
unique_elements, counts_elements = np.unique(y_test_preds, return_counts=True)
print(np.asarray((unique_elements, counts_elements)))
# convert X_test back to df
df_test = pd.DataFrame(X_test)
print(df_test)
# output predictions
id_col = df_test.iloc[:,[0]]
print(id_col)
id_col.rename(columns = {0:'Id'}, inplace = True)
print(id_col.columns)
print(id_col.info())
# need to change Id to int
id_col = id_col.Id.astype(int)
probabilities = pd.DataFrame(y_test_preds_prob[:,1], columns = ['TARGET_5Yrs'])
print(probabilities)
# +
# concat columns
output = pd.concat([id_col,probabilities], axis=1)
#df_output = pd.DataFrame(output_list, columns = ['Id','TARGET_5Yrs'])
print(output.head())
# -
# save to csv
output.to_csv('../data/processed/output_xgboost_best_params_wk1.csv',index=False)
# # Kaggle AUC = 0.70615
|
notebooks/tith_reasmey-10845345-week1_xgboosthptuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: test
# language: python
# name: test
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import ipywidgets
# %matplotlib inline
# -
# ## 전처리
# ### step 1: Image frame 단위 raw 데이터 로딩 및 중간 단계 dataset file 생성
# * inputs:
# * 태깅파일: ~_interpolation.json
# * 이미지파일: ~KM_0000000000.jpg
# * 감정종류: 8가지 ('happiness', 'afraid', 'neutral', 'surprise', 'sadness', 'contempt', 'anger', 'disgust')
# * 예시: {'happiness': 0, 'afraid': 0, 'neutral': 10, 'surprise': 0, 'sadness': 0, 'contempt': 0, 'anger': 0, 'disgust': 0} 일 경우 'neutral'이 선정
# * 중간단계 CSV dataset file 생성
# * 태깅데이터에서 전안 얼굴 검출 되는 이미지만 추출
# 중단 단계 dataset file 결과 예시
raw_tag_info = pd.read_csv('../datasets/kor_multi_modal/prep_part1_frontal_face.csv')
raw_tag_info.head(5)
# ### step 2: emotion class 조정 및 최종 dataset file 생성
# * emotion class 조정
# * emotion mapping
# * ASIS: 8 Catogory ('happiness', 'afraid', 'neutral', 'surprise', 'sadness', 'contempt', 'anger', 'disgust')
# * TO: 7 Category (0=Angry, 1=Disgust, 2=Fear, 3=Happy, 4=Sad, 5=Surprise, 6=Neutral)
# * mapped Category: 0=anger, 1=disgust&contempt, 2=afraid, 3=happiness, 4=sadness, 5=surprise, 6=neutral
# * 최종 CSV dataset file 생성
# * 태깅데이터에서 얼굴 검출 되는 이미지만 추출
# 최종 dataset file 예시: emotion, pixels 항목만 사용
df = pd.read_csv('../datasets/kor_multi_modal/prep_col_all_mdf_frontal_face.csv')
df.head(5)
# ## Model
# 1. face detection
# 2. emotion detection
# 3. tflite 변환 테스팅
#
# ### 1. face detection
#
# * OpenCV Haar cascade 방식
# * 사용이유: CPU 상 검출 시간이 빠르고, 적용하기 용이함
# * 버전: haarcascade_frontalface_alt2
# * 설명 내용 출처: https://thebook.io/006939/ch13/02/
# 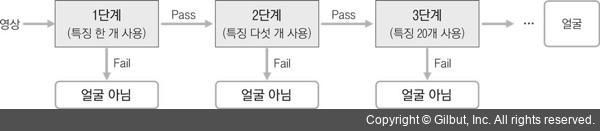
# 캐스케이드 구조 1단계에서는 얼굴 검출에 가장 유용한 유사-하르 필터 하나를 사용하여, 얼굴이 아니라고 판단되면 이후의 유사-하르 필터 계산은 수행하지 않습니다. 1단계를 통과하면 2단계에서 유사-하르 필터 다섯 개를 사용하여 얼굴이 아닌지를 검사하고, 얼굴이 아니라고 판단되면 이후 단계의 검사는 수행하지 않습니다.
# <img src='https://thebook.io/img/006939/p411.jpg' width=300 />
# <center>얼굴 검출에 유용한 유사-하르 필터의 예</center>
#
# * Dlib HOG(Histogram of Oriented Gradients) based 방식
# * 사용이유: CPU 상 OpenCV Haar cascade 방식보다 성능이 좋음, 적용하기 용이
# * 설명 내용 출처: https://medium.com/@jongdae.lim/기계-학습-machine-learning-은-즐겁다-part-4-63ed781eee3c
# * 검출방식 비교: https://seongkyun.github.io/study/2019/03/25/face_detection/
#
# <img src='https://miro.medium.com/max/1022/1*dP0Ixs4vHGUKCScufH9_Vw.jpeg' width=300 />
# <center>1) 이미지를 흑백으로 변환</center>
# <img src='https://miro.medium.com/max/1600/1*lsNRg_1oOELFcug_AjlkqQ.gif' width=300 />
# <center>2) 이미지를 16x16 픽셀의 작은 정사각형들로 분해 한 후 각 정사각형에서, 그래디언트가 주요 방향(윗쪽, 우상쪽, 오른쪽, 등)을 얼마나 가리키고 있는지 카운팅합니다.</center>
# <center>그런 다음 이 사각형을 가장 강한 화살표 방향을 나타내는 벡터로 변환 합니다.</center>
# <img src='https://miro.medium.com/max/1600/1*HtgQZ4guaIo8wflbsR1MLw.png' width=500 />
#
# ### 2. emotion detection
# * mini_XCEPTION 모델
# * 사용이유: 인식시간이 빠르고, 모델이 가볍기에 온디바이스에 적용가능
# * 학습내용:
# * 학습데이터: AIhub 멀티모달 데이터(http://www.aihub.or.kr/content/555)로 학습
# * 클래스 밸런싱된 48 x 48 gray scale image 128,499만여개에서 샘플링하여 사용
# * 학습된 모델을 tflite로 변환
# * 모델출처: https://github.com/oarriaga/face_classification
fig = plt.figure(figsize=(15, 15))
plt.imshow(plt.imread('../images/simple_model_structure.png'))
# ## 전처리
# ### step 1: Image frame 단위 raw 데이터 로딩 및 중간 단계 dataset file 생성
# 한 파일당 약 2~3시간 소요
# python preprocess_jj.py <file_number> <input_folder> <output_folder>
예시: !python pretrain_jj.py 1 /tf/notebooks/datasets/emotion/multi-modal ../datasets/kor_multi_modal
# !cat pretrain_jj.py
# 중단 단계 dataset file 결과 예시
raw_tag_info = pd.read_csv('../datasets/kor_multi_modal/prep_part1_frontal_face.csv')
raw_tag_info.head(5)
# ### step 2: emotion class 조정
# +
import os
import json
import io
import numpy as np
import operator
import collections
import base64
import pandas as pd
from glob import glob
from IPython.display import HTML
import ray
import matplotlib.pyplot as plt
import copy
import logging
import cv2
from utils.inference import load_detection_model
from utils.inference import detect_faces
from utils.inference import apply_offsets
detection_model_path = '../trained_models/detection_models/haarcascade_frontalface_alt2.xml'
logger = logging.getLogger()
logger.setLevel(logging.DEBUG)
# multi-modal 데이터셋 zip파일의 part 번호 (1~5)
part_num = '1'
# input 폴더
base_folder = '/tf/notebooks/datasets/emotion/multi-modal/part'+part_num+'/'
# output 파일경로
prep_trainset_path='../datasets/kor_multi_modal/prep_part'+part_num+'.csv'
prep_trainset_frontal_face_path='../datasets/kor_multi_modal/prep_part'+part_num+'_frontal_face.csv'
prep_trainset_mid_path='../datasets/kor_multi_modal/prep_part'+part_num+'_mid.csv'
# %matplotlib inline
# +
raw_tag_info = pd.read_csv(prep_trainset_frontal_face_path)
raw_tag_info_df = pd.DataFrame(raw_tag_info)
# raw_tag_info_df["emotion"].value_counts().plot.bar()
# raw_tag_info_df["emotion"].value_counts()
## disgust와 contempt는 disgust 하나로 합침
raw_tag_info_df = raw_tag_info_df.replace('contempt', 'disgust')
## balancing sampling
min_emotion_cnt = np.min(raw_tag_info_df["emotion"].value_counts())
sampled_tag_info_df = raw_tag_info_df.groupby("emotion", group_keys=False).apply(lambda df: df.sample(min_emotion_cnt))
sampled_tag_info_df = sampled_tag_info_df.reset_index(drop=True)
# sampled_tag_info_df["emotion"].value_counts()
# sampled_tag_info_df.count()
# sampled_tag_info_df.head()
### Real-time Convolutional Neural Networks for Emotion and Gender Classification 모델을 위한 전처리 부분
import cv2
import re
import numpy as np
import matplotlib.pyplot as plt
from multiprocessing import Pool
import json
import multiprocessing
# num_cores = multiprocessing.cpu_count()
num_cores = 5
def img_path_to_pixel(x):
img_path, face_rect = x[0], x[1]
face_rect = json.loads(face_rect.replace("'", "\""))
img = cv2.imread(img_path, cv2.IMREAD_GRAYSCALE)
img_shape = img.shape
# face_rect 부분 잘라내기
# ex) 'max_x': 1125, 'max_y': 798, 'min_x': 708, 'min_y': 267
min_x = face_rect['min_x']
max_x = face_rect['max_x']
min_y = face_rect['min_y']
max_y = face_rect['max_y']
if not isinstance(min_x, int) or not isinstance(max_x, int) or not isinstance(min_y, int) or not isinstance(max_y, int) or \
min_x <= 0 or max_x <= 0 or min_y <= 0 or max_y <= 0:
return None
# print(min_y,max_y, min_x,max_x)
img = img[min_y:max_y, min_x:max_x]
# 이미지 축소 시에는 INTER_AREA, https://076923.github.io/posts/Python-opencv-8/
img = cv2.resize(img, dsize=(48, 48), interpolation=cv2.INTER_AREA)
# split array to ' ' str
img = ' '.join(img.astype(int).astype(str).reshape(-1))
return img
def multi_process_sub(df_split):
return df_split[['img_path', 'face_rect']].apply(img_path_to_pixel, axis=1)
def multi_process(df, func):
df_split = np.array_split(df, num_cores)
pool = Pool(num_cores)
df = pd.concat(pool.map(func, df_split))
pool.close()
pool.join()
return df
# # %time sampled_tag_info_df['img_path_new'] = sampled_tag_info_df[['img_path','face_rect']].apply(img_path_to_pixel, axis=1)
# %time sampled_tag_info_df['img_path_new'] = multi_process(sampled_tag_info_df, multi_process_sub)
# sampled_tag_info_df.shape
# 이상 데이터 drop
sampled_tag_info_df = sampled_tag_info_df.dropna().reset_index(drop=True)
# sampled_tag_info_df.shape
sampled_tag_info_df['img_path'] = sampled_tag_info_df['img_path_new']
prep_tag_info_df = sampled_tag_info_df.drop(['img_path_new', 'face_rect'], axis=1)
# prep_tag_info_df["emotion"].value_counts().plot.bar()
# * CSV dataset file 생성
# * 포멧: emotion
# * emotion mapping
# * ASIS: 8 Catogory ('happiness', 'afraid', 'neutral', 'surprise', 'sadness', 'contempt', 'anger', 'disgust')
# * TO: 7 Category (0=Angry, 1=Disgust, 2=Fear, 3=Happy, 4=Sad, 5=Surprise, 6=Neutral)
# * mapped Category: 0=anger, 1=disgust&contempt, 2=afraid, 3=happiness, 4=sadness, 5=surprise, 6=neutral
def emotion_mapping(emotion):
if emotion == 'anger':
return 0
elif emotion == 'disgust' or emotion == 'contempt':
return 1
elif emotion == 'afraid':
return 2
elif emotion == 'happiness':
return 3
elif emotion == 'sadness':
return 4
elif emotion == 'surprise':
return 5
elif emotion == 'neutral':
return 6
else:
return 6
prep_tag_info_df['mapped_emotion'] = prep_tag_info_df['emotion'].map(emotion_mapping)
# prep_tag_info_df['mapped_emotion'].value_counts()
prep_tag_info_df['emotion'] = prep_tag_info_df['mapped_emotion']
prep_tag_info_df = prep_tag_info_df.drop(['mapped_emotion'], axis=1)
# prep_tag_info_df.head()
prep_tag_info_df.to_csv(prep_trainset_mid_path, index=False)
tmp_df = pd.read_csv(prep_trainset_mid_path)
tmp_df.head()
# -
# 최종 dataset file 예시: emotion, pixels 항목만 사용
df = pd.read_csv('../datasets/kor_multi_modal/prep_col_all_mdf_frontal_face.csv')
df.head(5)
# ## 모델
# ### 1. face detection
model_path = '../trained_models/detection_models/haarcascade_frontalface_alt2.xml'
detection_model = cv2.CascadeClassifier(model_path)
# ### 2. emotion detection
# +
# 추후 공개
# -
# ### 3. TFlite 모델 파일 테스팅
# !ls models/
demo_df = pd.read_csv("../data/demo.csv")
# +
# %%time
def f(x):
emotion_map = {
0:'anger'
,1:'disgust, contempt'
,2:'afraid'
,3:'happiness'
,4:'sadness'
,5:'surprise'
,6:'neutral'
}
row = demo_df.iloc[x]
pixels = row['pixels']
emotion = row['emotion']
img = np.asarray(pixels.split(" "), dtype=np.int16).reshape((48, 48))
img = img / 255.0
input_data = tf.cast(img, tf.float32)
input_data = input_data[tf.newaxis,:,:,tf.newaxis]
fig = plt.figure(figsize=(10, 10))
fig.add_subplot(2, 1, 1)
print('Label: ', emotion_map[emotion], plt.imshow(img, 'gray'))
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path="../models/converted_model_final.tflite")
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
# Test model on random input data.
input_shape = input_details[0]['shape']
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
# The function `get_tensor()` returns a copy of the tensor data.
# Use `tensor()` in order to get a pointer to the tensor.
output_data = interpreter.get_tensor(output_details[0]['index'])
print('Predicted label: ', emotion_map[np.argmax(output_data)], output_data)
fig.add_subplot(2, 1, 2)
plt.bar(emotion_map.values(), output_data[0]*100)
ipywidgets.interactive(f, x = ipywidgets.IntSlider(0, 0, demo_df.shape[0]-1, 1))
# -
|
src/00_vision_model_total_process.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ------------------------------------------------------------------------------------------------------
# ## Primordial Black Hole Evaporation -- Purely Gravitational Interacting Dark Matter.
# ## Authors: <NAME>, <NAME>, <NAME>, <NAME>
#
# ## Based on: arXiv:2107.xxxxx
# ## ------------------------------------------------------------------------------------------------------
#
# ### Basic notebook to show how to calculate the relic density of a fermionic dark matter particle produced only by Hawking radiation.
import Omega_h2_onlyDM as pbh
import numpy as np
# #### Set the parameters
MPBH = 1e2 # PBH initial mass in grams
ast = 0.0 # BH initial angular momentum parameter, ast = 0. -> Schwarzschild, ast > 0. -> Kerr.
beta = 10**(-17.5) # Parameter describing the initial BH energy density over total energy density
mdm = 1e7 # Dark matter mass in GeV
# The class FrInPBH is where the $\Omega h^2$ is calculated
foh2 = pbh.FrInPBH(np.log10(MPBH), ast, np.log10(beta), np.log10(mdm),2)
# This is how you get the relic density
# +
foh2.Omegah2()
|
PBH-DM/Example_onlyDM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # LinkedIn - Send invitation from gsheet
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/LinkedIn/LinkedIn_Send_invitation_from_gsheet.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #linkedin #invitation #naas_drivers #gsheet #automation #scheduler
# + [markdown] papermill={} tags=[]
# **Author:** [<NAME>](https://www.linkedin.com/in/valentin-goulet-3a3070152/)
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Import libraries
# + papermill={} tags=[]
import naas
from naas_drivers import linkedin, gsheet
# + [markdown] papermill={} tags=[]
# ### Setup your LinkedIn
# <a href='https://www.notion.so/LinkedIn-driver-Get-your-cookies-d20a8e7e508e42af8a5b52e33f3dba75'>How to get your cookies ?</a>
# + papermill={} tags=[]
# LinkedIn cookies
LI_AT = 'YOUR_COOKIE_LI_AT'
JSESSIONID = 'YOUR_COOKIE_JSESSIONID'
# LinkedIn limit invitations up to 100 per week (Becareful !)
add_per_launch = 4
# + [markdown] papermill={} tags=[]
# ### Setup your Google Sheet
# 👉 The spreadsheet needs to contains profil's url on the 1st column of the file.
# + papermill={} tags=[]
spreadsheet_id = "YOUR_SPREADSHEET_ID"
sheet_name = "YOUR_SHEET_NAME"
profile_col_name = "url"
# + [markdown] papermill={} tags=[]
# ### Schedule your notebook
# + papermill={} tags=[]
# Scheduler your invitation everyday at 8:00 AM
naas.scheduler.add(cron="0 8 * * *")
# Uncomment the line below to delete your scheduler
# naas.scheduler.delete()
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Get all list of profiles
# + papermill={} tags=[]
df = gsheet.connect(spreadsheet_id).get(sheet_name=sheet_name)
# + [markdown] papermill={} tags=[]
# ### Get restricted list
# + papermill={} tags=[]
# Alert when last than 20 urls remains in the gsheet
if len(df) < 20:
email_to = "YOUR_EMAIL"
subject = "Invite LinkedIn alert : " + str(len(df)) + " lines left in the Linkedin's url database"
content = "You can add more lines to the gsheet or update the Notebook to set a new spreadsheet !"
naas.notification.send(email_to=email_to, subject=subject, html=content)
df = df.head(add_per_launch)
print("Invits will be send to :")
df
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Send the invitation and delete profil from Gsheet
# + papermill={} tags=[]
for index, row in df.iterrows():
profile = row[profile_col_name]
result = linkedin.connect(LI_AT, JSESSIONID).invitation.send(recipient_url=profile)
print(f"Invitation sent to : {profile}")
# Suppression de la ligne du gsheet
gsheet.connect(spreadsheet_id).delete(sheet_name=sheet_name, rows=[2])
|
LinkedIn/LinkedIn_Send_invitation_from_gsheet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from keplersplinev2 import *
#Make a fake light curve with a transit
t = np.linspace(0, 20, 1000)
tno = np.copy(t)
t[200:] = t[200:] + 4
f = 0.02 * np.sin(2 * np.pi * t / (13.4)) + np.random.normal(1, 0.003, len(t)) -0.05 * np.sin(2 * np.pi * t / (38.4))
f[600:620] = f[600:620] - 0.05
# +
#Calculate a keplerspline with the default bkspacing of 1.5 days between knots:
s = keplersplinev2(t, f)
fig, ax = plt.subplots(figsize=(16, 8))
plt.plot(t, f,'.')
plt.plot(t, s,'o')
# +
#If we want a more aggressive spline, we can use a smaller value for bkspace, but in this case it eats the transit:
s = keplersplinev2(t, f, bkspace = 0.5)
fig, ax = plt.subplots(figsize=(16, 8))
plt.plot(t, f,'.')
plt.plot(t, s,'o')
# +
#So let's create a mask. It's just an array of True/False values, where "True" means "use this point in the fit",
#and "False" means "Ignore this point"
input_mask = np.ones_like(t, dtype=np.bool)
input_mask[595:625] = False
smasked = keplersplinev2(t, f, input_mask = input_mask, bkspace = 0.5)
fig, ax = plt.subplots(figsize=(16, 8))
plt.plot(t, f,'.')
plt.plot(t, smasked,'o')
# +
#We can also use the return_metadata=True tag in keplersplinev2
input_mask = np.ones_like(t, dtype=np.bool)
input_mask[595:625] = False
smasked, metadata = keplersplinev2(t, f, input_mask = input_mask, bkspace = 1.5, return_metadata=True)
fig, ax = plt.subplots(figsize=(16, 8))
plt.plot(t, f,'.')
plt.plot(t, smasked,'o')
# -
#Metadata has the following attributes:
# metadata.light_curve_mask = light_curve_mask
# metadata.input_light_curve_mask = all_input_mask
# metadata.bkspace = bkspace
# metadata.likelihood_term = likelihood_term
# metadata.penalty_term = penalty_term
# metadata.bic = bic
print(metadata.bkspace)
# +
#You can also ask keplerspline to choose the best bkspace value using choosekeplersplinev2
s, metadata = choosekeplersplinev2(t, f, return_metadata=True)
fig, ax = plt.subplots(figsize=(16, 8))
plt.plot(t, f,'.')
plt.plot(t, s,'o')
# -
#To find out which bkspace choosekeplersplinev2 chose, grab the metadata
print(metadata.bkspace)
|
keplersplinev2_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="eDRfJeA9i540" colab={"base_uri": "https://localhost:8080/"} outputId="eb115cb0-2826-4f8c-a203-4bc491b7726a"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import arviz as az
from statsmodels.tsa import stattools
import statsmodels.api as sm
import pymc3 as pm
import pymc
import sys
sys.path.insert(0, '..')
from utils.plot_lib import set_default
set_default(figsize=(6, 4))
# + [markdown] id="WBG3Vsj2S3xV"
# ### Mixture models for density estimation
#
# The galaxies dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="V7VdfuBLShRW" outputId="df4dcdf3-0ad0-4702-a075-2ae02a5d4e04"
# !wget http://www.key2stats.com/Velocities_for_82_Galaxies_1152_58.csv -O galaxy.csv --no-check-certificate
# + colab={"base_uri": "https://localhost:8080/", "height": 223} id="j76qLqfXTOgJ" outputId="4ddc701b-df98-4eb3-b8ca-da13f60812e1"
# Read dataset - galaxies
galaxy = pd.read_csv('galaxy.csv')
galaxy.pop('Unnamed: 0')
galaxy.pop('X')
# Checking dataframe
print("The shape of the dataset is: {}".format(galaxy.shape))
galaxy.head()
# + [markdown] id="P_alHv5ljLpp"
# These are velocities of galaxies measured in six well-separated conic sections of the Corona Borealis.
# So we want to understand whether this distribution is multimodal, how many modes are present, and in general provide
# a density estimator for this different velocities.
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="1EF48VnKjPkY" outputId="eeea79d3-b421-4c7d-bd33-b42bcd7fa994"
from scipy.stats import norm
## Clear the environment and load required libraries
np.random.seed(1)
### Loading data and setting up global variables
KK = 6 # We're going to be using six components in our density estimators using mixture models.
x = galaxy['dat'].to_numpy() # Data
n = len(x)
sns.histplot(x, stat= 'density', bins = 6)
plt.title('Approximate density of the data')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="zE6mgAt8VwfV" outputId="a25b5ff2-bfc3-4070-bd91-832ca9d77181"
### First, compute the "Maximum Likelihood" density estimate associated with a location mixture of 6 Gaussian distributions using the EM algorithm
## Initialize the parameters
## Initialize the parameters
w = np.ones(KK) / KK # Assign equal weight to each component to start with
mu = norm.rvs(loc = np.mean(x), scale = np.std(x), size = KK) # Random cluster centers randomly spread over the support of the data
sigma = np.std(x) / KK # Initial standard deviation
print('The initial guess for mu are: \n{}'.format(mu))
print('The initial guess for sigma are: {}'.format(sigma))
# + [markdown] id="FqZi45dOUNKC"
# ### MLE EM algorithm
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="eH5GGs5RAl4o" outputId="bab4817a-27f8-4f80-87fa-297b909ebb6d"
from scipy.stats import lognorm
s = 0 # Counter of the current iteration
sw = False # Decide when the algorithm will end
KL = np.NINF # Value of Q to monitor convergence
KL_out = []
epsilon = 10e-6 # Stopping criteria
## Checking convergence of the algorithm
while(not sw):
## Expectation (E) step
v = np.zeros((n, KK))
for k in range(KK):
v[:, k] = np.log(w[k]) + norm.logpdf(loc = mu[k], scale = sigma, x = x)
for i in range(n):
v[i, :] = np.exp(v[i, :] - max(v[i, : ])) / np.sum(np.exp(v[i, :] - max(v[i, :]))) # Go from logs to actual weights in a numerically stable manner
## Maximization (M) step
# Weights
w = np.mean(v, axis = 0)
# Mus
mu = np.zeros(KK)
for k in range(KK):
for i in range(n):
# Update Mu
mu[k] = mu[k] + (v[i,k] * x[i])
mu[k] = mu[k] / np.sum(v[:, k])
# Standard deviations
sigma = 0
for i in range(n):
for k in range(KK):
sigma = sigma + (v[i,k] * ((x[i] - mu[k])**2))
sigma = np.sqrt(sigma / np.sum(v))
##Check convergence
KLn = 0 # Value of Q function at current iteration
for i in range(n):
for k in range(k):
KLn = KLn + v[i, k]*(np.log(w[k]) + norm.logpdf(loc = mu[k], scale =sigma, x = x[i]))
# If relative error is less than epsilon stop
if(np.abs(KLn - KL) / np.abs(KLn) < epsilon):
sw = True
KL = KLn
KL_out.append(KL)
s = s + 1
print('The iteration {} has a Q value of {}'.format(s, KLn))
fig, axs = plt.subplots(2)
axs[0].plot(KL_out[0 : s], c = 'red')
axs[0].set_ylabel('Q')
axs[0].axis(xmin = 1, xmax = max(10, s))
axs[1].set_title("Q-value over iterations")
# Estimate of the density
xx = np.linspace(5000, 37000, num = 300)
nxx = len(xx)
density_EM = np.zeros(nxx)
for s_ in range(nxx):
for k in range(KK):
density_EM[s_] = density_EM[s_] + w[k] * norm.pdf(loc = mu[k], scale = sigma, x = xx[s_])
# Plot the initial guess for the density
axs[1].plot(xx, density_EM, 'blue', label = 'Density estimated')
axs[1] = sns.scatterplot(np.array(x), np.zeros(len(x)))
axs[1].set_ylabel('x')
axs[1].set_ylabel('Density')
axs[1].set_title("s = {}, Q = {}".format(s, np.round(KL_out[s - 1], 4)))
fig.subplots_adjust(hspace=0.5)
plt.show()
# + [markdown] id="HTL9WA6WaX8_"
# ### Bayesian EM algorithm
#
# This is the same model used in L5A.
# + id="ZEy-XZ2yaZbG"
### Get a "Bayesian" kernel density estimator based on the same location mixture of 6 normals
## Priors set up using an "empirical Bayes" approach
aa = np.ones(KK) # Uniform prior on w
eta = np.mean(x) # Mean for the prior on mu_k
tau = np.sqrt(np.var(x)) # Standard deviation for the prior for mu_l
dd = 2 # Inverse gamma prior for sigma_2, parameter d
qq = np.var(x) / KK # Inverse gamma prior for sigma_2, parameter q
## Initialize the parameters
## Initialize the parameters
w = np.ones(KK) / KK # Assign equal weight to each component to start with
mu = norm.rvs(loc = np.mean(x), scale = np.std(x), size = KK, random_state = 1) # Random cluster centers randomly spread over the support of the data
sigma = np.std(x) / KK # Initial standard deviation
cc = np.random.choice(np.arange(KK), n, p = w)
# + colab={"base_uri": "https://localhost:8080/", "height": 798} id="LmDKH9ktacHX" outputId="2da5e229-830f-4bcd-f24e-3219270b60e9"
from scipy.stats import dirichlet
from scipy.stats import invgamma
from scipy.stats import gamma
# Number of iterations of the sampler
rrr = 12000 # Number of iterations
burn = 3000 # Burning period
# Storing the samples
cc_out = np.zeros((rrr, n)) # Store indicators
w_out = np.zeros((rrr, KK)) # Sample of the weights
mu_out = np.zeros((rrr, KK)) # Sample of mus
sigma_out = np.zeros(rrr) # Sample of sigmas
logpost = np.zeros(rrr) # Used to monitor convergence
for s in range(rrr):
# Sample the indicators
cc = np.zeros(n)
for i in range(n):
v = np.zeros(KK)
for k in range(KK):
v[k] = np.log(w[k]) + norm.logpdf(loc = mu[k], scale = sigma, x = x[i]) # Compute the log of the weights
v = np.exp(v - max(v)) / np.sum(np.exp(v - max(v))) # Go from logs to actual weights in a numerically stable manner
cc[i] = np.random.choice(np.arange(KK), 1, p = v) # C_i sample
# Sample the weights
counts = np.zeros(KK)
for k in range(KK):
sum_k = np.sum(cc == k)
counts[k] = int(sum_k)
w = dirichlet.rvs(alpha = aa + counts, size = 1)[0]
# Sample the means
for k in range(KK):
nk = np.sum(cc == k)
xsumk = np.sum(x[cc == k])
tau2_hat = 1 / (nk / sigma**2 + 1 / tau**2)
mu_hat = tau2_hat * (xsumk / sigma**2 + eta / tau**2)
mu[k] = norm.rvs(loc = mu_hat, scale = np.sqrt(tau2_hat), size = 1)
# Sample the variances
dd_star = dd + n / 2
mu_temp = [mu[int(c_i)] for c_i in cc] # Create vector of mus
qq_star = qq + np.sum((x - mu_temp)**2) / 2
sigma = np.sqrt(1 / gamma.rvs(a = dd_star, scale = 1 / qq_star, size = 1))
# Store samples
cc_out[s, :] = cc
w_out[s] = w
mu_out[s, :] = mu
sigma_out[s] = sigma
for i in range(n):
# Computing logposterior likelihood term
logpost[s] = logpost[s] + np.log(w[int(cc[i])]) + norm.logpdf(loc = mu[int(cc[i])], scale = sigma, x = x[i])
# W term
logpost[s] = logpost[s] + dirichlet.logpdf(alpha = aa, x = w)
# Mu term
for k in range(KK):
logpost[s] = logpost[s] + norm.logpdf(loc = eta, scale = tau, x = mu[k])
# Sigma term
logpost[s] = logpost[s] + gamma.logpdf(a = dd, scale = 1 / qq, x = 1 / sigma**2) - 4 * np.log(sigma)
if s / 500 == np.floor(s / 500):
print('Current iteration is: {}'.format(s))
## Plot the logposterior distribution for various samples
fig, ax = plt.subplots(1, 1)
ax.plot(np.arange(len(logpost)), logpost, 'r-', lw=1, alpha=0.6, label='Trace plot') # Trace plot of data
ax.legend(loc='best', frameon=False)
# plot density estimate of the posterior
plt.title('Trace plot of Logposterior')
plt.show()
# + id="4A6Qw8hLgzRu"
# Values to sample
xx = np.linspace(5000, 37000, num = 300)
nxx = len(xx)
density_mcmc = np.zeros((rrr-burn, len(xx)))
for s in range(rrr-burn):
for k in range(KK):
density_mcmc[s, :] = density_mcmc[s, :] + \
w_out[s + burn, k] * norm.pdf(loc = mu_out[s + burn, k], scale = sigma_out[s + burn], x = xx)
density_mcmc_m = np.mean(density_mcmc, axis = 0)
density_mcmc_lq = np.quantile(density_mcmc, 0.025, axis = 0)
density_mcmc_uq = np.quantile(density_mcmc, 0.975, axis = 0)
# + colab={"base_uri": "https://localhost:8080/", "height": 381} id="CNMBd9txhfix" outputId="bdbdaaca-f7fb-4a3f-f8be-f5924c16066b"
## Plot the final result distribution for various samples
fig, ax = plt.subplots(1, 1)
# Bayesian plot
ax.plot(xx, density_mcmc_m, lw=2, alpha=0.6, label='MCMC') # Trace plot of data
# EM algorithm
ax.plot(xx, density_EM, 'blue', label = 'EM')
# KDE plot
ax = sns.kdeplot(x, bw_adjust = 0.5, label = 'KDE')
# Plotting original data
ax.scatter(np.array(x), np.zeros((x.shape[0])), label = 'Original data')
# Plotting uncertainty
plt.fill_between(xx, density_mcmc_uq, density_mcmc_lq, alpha=0.2,
label='Uncertainty Interval')
ax.legend(loc='best', frameon=False)
# plot density estimate of the posterior
plt.title('Trace plot of Logposterior')
plt.show()
|
C3/L6_density_estimation.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Sellke construction of the SIR model
# <NAME> (@slwu89), 2021-11-30
#
# ## Introduction
#
# The Sellke construction is an alternative stochastic process to the standard stochastic epidemic
# process that can be used to sample the final size distribution of an SIR epidemic in a finite population,
# originally described in [On the asymptotic distribution of the size of a stochastic epidemic](https://doi.org/10.2307/3213811).
#
# Rather than sampling next events and times under a race condition to update the system as in
# standard stochastic epidemics, the Sellke construction samples at once a "threshold" value
# for each susceptible individual. The "infection pressure" generated by the infectious population
# is tracked, and when it exceeds the threshold for a given susceptible person, they become infected.
#
# This construction is similar to a [random time change](http://bactra.org/notebooks/random-time-changes.html) such
# that the thresholds are independent and identically distributed exponential random variates with a mean of 1,
# and infection pressure is the cumulative hazard $\beta \, I/N$.
#
# ## Libraries
using Distributions
using Random
using BenchmarkTools
using Plots
# ## Sellke model
#
# Code to draw a final epidemic size from a standard continuous-time Markov SIR model
# is below. The Sellke construction can handle non-exponentially distributed infectious
# periods, which simply require the the infectious durations to be sampled from the appropriate
# distribution. Another nice feature of the Sellke epidemic model is that parameters can
# easily be changed without needing to draw new random numbers.
#
# Because the construction samples the final epidemic size, and then creates a corresponding
# temporal trajectory, we do not specify the time domain.
#
# This particular implementation is described in the paper [How big is an outbreak likely to be? Methods for epidemic final-size calculation](https://royalsocietypublishing.org/doi/full/10.1098/rspa.2012.0436).
function sellke(u0, p)
(S, I, R) = u0
N = S + I + R
(β, c, γ) = p
λ = β*c/N
Q = rand(Exponential(), S)
sort!(Q)
T0 = rand(Exponential(1/γ), I)
T = rand(Exponential(1/γ), S)
ST0 = sum(T0)
Y = [ST0; ST0 .+ cumsum(T[1:end-1])]
Z = findfirst(Q .> Y*λ)
if isnothing(Z)
Z = S + I # entire population infected
else
Z = Z + I - 1
end
TT = [T0; T] # all infectious periods
QQ = [T0 * 0; Q] # all thresholds
R = T[1:I] # recovery times of the initial infectives
# max num of events possible
max = I + 2*S
t = zeros(max)
St = zeros(max)
It = zeros(max)
It[1:I] = (1:I)
St[1:I] = N .- (1:I)
tt = 0
La = 0
j = I+1
k = j
while It[k-1] > 0
(minR, i) = findmin(R)
dtprop = minR-tt
Laprop = La + (λ * It[k-1] * dtprop)
if j > length(QQ) # only recoveries remain
R = R[setdiff(1:length(R), i)]
tt = minR
t[k] = minR
It[k] = It[k-1]-1
St[k] = St[k-1]
La = Laprop
k = k+1
else # infections
if QQ[j] > Laprop
R = R[setdiff(1:length(R), i)]
tt = minR
t[k] = minR
It[k] = It[k-1]-1
St[k] = St[k-1]
La = Laprop
else
tt = tt + ((QQ[j]-La)/(Laprop-La))*dtprop
La = QQ[j]
t[k] = tt
It[k] = It[k-1]+1
St[k] = St[k-1]-1
R = [R; tt+TT[j]]
j = j+1
end
k = k+1
end
end
trajectory = hcat(t, St, It, N .- (St + It))
trajectory = trajectory[I:k-1, :]
return trajectory
end
# ## Initial conditions
u0 = [990,10,0]; # S,I,R
# ## Parameter values
p = [0.05,10.0,0.25]; # β,c,γ
# ## Random number seed
#
# We set a random number seed for reproducibility.
Random.seed!(1234);
# ## Running the model
out = sellke(u0, p)
# ## Plotting
#
# We plot a histogram of final epidemic sizes.
plot(out[:, 1], out[:, 2:end],
label=["S" "I" "R"],
xlabel="Time",
ylabel="Number")
# ## Benchmarking
@benchmark sellke(u0, p)
|
notebook/sellke/sellke.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Deep Autoencoders
# #### by <NAME> as a part of a <a href="https://www.google-melange.com/gsoc/project/details/google/gsoc2014/khalednasr92/5657382461898752">GSoC 2014 project</a> mentored by <NAME> and <NAME>
# This notebook illustrates how to train and evaluate a deep autoencoder using Shogun. We'll look at both regular fully-connected autoencoders and convolutional autoencoders.
# ## Introduction
# A (single layer) [autoencoder](http://deeplearning.net/tutorial/dA.html#autoencoders) is a neural network that has three layers: an input layer, a hidden (encoding) layer, and a decoding layer. The network is trained to reconstruct its inputs, which forces the hidden layer to try to learn good representations of the inputs.
#
# In order to encourage the hidden layer to learn good input representations, certain variations on the simple autoencoder exist. Shogun currently supports two of them: Denoising Autoencoders [1] and Contractive Autoencoders [2]. In this notebook we'll focus on denoising autoencoders.
#
# For denoising autoencoders, each time a new training example is introduced to the network, it's randomly corrupted in some mannar, and the target is set to the original example. The autoencoder will try to recover the orignal data from it's noisy version, which is why it's called a denoising autoencoder. This process will force the hidden layer to learn a good representation of the input, one which is not affected by the corruption process.
#
# A deep autoencoder is an autoencoder with multiple hidden layers. Training such autoencoders directly is usually difficult, however, they can be pre-trained as a stack of single layer autoencoders. That is, we train the first hidden layer to reconstruct the input data, and then train the second hidden layer to reconstruct the states of the first hidden layer, and so on. After pre-training, we can train the entire deep autoencoder to fine-tune all the parameters together. We can also use the autoencoder to initialize a regular neural network and train it in a supervised manner.
#
# In this notebook we'll apply deep autoencoders to the USPS dataset for handwritten digits. We'll start by loading the data and dividing it into a training set and a test set:
# +
# %pylab inline
# %matplotlib inline
import os
SHOGUN_DATA_DIR=os.getenv('SHOGUN_DATA_DIR', '../../../data')
from scipy.io import loadmat
from shogun import features, MulticlassLabels, Math
# load the dataset
dataset = loadmat(os.path.join(SHOGUN_DATA_DIR, 'multiclass/usps.mat'))
Xall = dataset['data']
# the usps dataset has the digits labeled from 1 to 10
# we'll subtract 1 to make them in the 0-9 range instead
Yall = np.array(dataset['label'].squeeze(), dtype=np.double)-1
# 4000 examples for training
Xtrain = features(Xall[:,0:4000])
Ytrain = MulticlassLabels(Yall[0:4000])
# the rest for testing
Xtest = features(Xall[:,4000:-1])
Ytest = MulticlassLabels(Yall[4000:-1])
# initialize the random number generator with a fixed seed, for repeatability
Math.init_random(10)
# -
# ## Creating the autoencoder
# Similar to regular neural networks in Shogun, we create a [deep autoencoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html) using an array of [NeuralLayer](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CNeuralLayer.html)-based classes, which can be created using the utility class [NeuralLayers](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CNeuralLayers.html). However, for deep autoencoders there's a restriction that the layer sizes in the network have to be symmetric, that is, the first layer has to have the same size as the last layer, the second layer has to have the same size as the second-to-last layer, and so on. This restriction is necessary for pre-training to work. More details on that can found in the following section.
#
# We'll create a 5-layer deep autoencoder with following layer sizes: 256->512->128->512->256. We'll use [rectified linear neurons](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CNeuralRectifiedLinearLayer.html) for the hidden layers and [linear neurons](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CNeuralLinearLayer.html) for the output layer.
# +
from shogun import NeuralLayers, DeepAutoencoder
layers = NeuralLayers()
layers = layers.input(256).rectified_linear(512).rectified_linear(128).rectified_linear(512).linear(256).done()
ae = DeepAutoencoder(layers)
# -
# ## Pre-training
# Now we can pre-train the network. To illustrate exactly what's going to happen, we'll give the layers some labels: L1 for the input layer, L2 for the first hidden layer, and so on up to L5 for the output layer.
#
# In pre-training, an autoencoder will formed for each encoding layer (layers up to the middle layer in the network). So here we'll have two autoencoders: L1->L2->L5, and L2->L3->L4. The first autoencoder will be trained on the raw data and used to initialize the weights and biases of layers L2 and L5 in the deep autoencoder. After the first autoencoder is trained, we use it to transform the raw data into the states of L2. These states will then be used to train the second autoencoder, which will be used to initialize the weights and biases of layers L3 and L4 in the deep autoencoder.
#
# The operations described above are performed by the the [pre_train()](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#acf6896cb166afbba063fd1257cb8bc97) function. Pre-training parameters for each autoencoder can be controlled using the [pt_* public attributes](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#a6389a6f19b8854c64e1b6be5aa0c1fc4) of [DeepAutoencoder](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html). Each of those attributes is an [SGVector](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1SGVector.html) whose length is the number of autoencoders in the deep autoencoder (2 in our case). It can be used to set the parameters for each autoencoder indiviually. [SGVector's set_const()](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1SGVector.html#a8bce01a1fc41a734d9b5cf1533fd7a2a) method can also be used to assign the same parameter value for all autoencoders.
#
# Different noise types can be used to corrupt the inputs in a denoising autoencoder. Shogun currently supports 2 [noise types](http://www.shogun-toolbox.org/doc/en/latest/namespaceshogun.html#af95cf5d3778127a87c8a67516405d863): dropout noise, where a random portion of the inputs is set to zero at each iteration in training, and gaussian noise, where the inputs are corrupted with random gaussian noise. The noise type and strength can be controlled using [pt_noise_type](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#af6e5d2ade5cb270cc50565d590f929ae) and [pt_noise_parameter](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#adbdff6c07fa7dd70aaf547e192365075). Here, we'll use dropout noise.
# +
from shogun import AENT_DROPOUT, NNOM_GRADIENT_DESCENT
ae.pt_noise_type.set_const(AENT_DROPOUT) # use dropout noise
ae.pt_noise_parameter.set_const(0.5) # each input has a 50% chance of being set to zero
ae.pt_optimization_method.set_const(NNOM_GRADIENT_DESCENT) # train using gradient descent
ae.pt_gd_learning_rate.set_const(0.01)
ae.pt_gd_mini_batch_size.set_const(128)
ae.pt_max_num_epochs.set_const(50)
ae.pt_epsilon.set_const(0.0) # disable automatic convergence testing
# uncomment this line to allow the training progress to be printed on the console
#from shogun import MSG_INFO; ae.io.set_loglevel(MSG_INFO)
# start pre-training. this might take some time
ae.pre_train(Xtrain)
# -
# ## Fine-tuning
# After pre-training, we can train the autoencoder as a whole to fine-tune the parameters. Training the whole autoencoder is performed using the [train()](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CAutoencoder.html#ace3eb6cc545affcbfa31d754ffd087dc) function. Training parameters are controlled through the [public attributes](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#pub-attribs), same as a regular neural network.
# +
ae.put('noise_type', AENT_DROPOUT) # same noise type we used for pre-training
ae.put('noise_parameter', 0.5)
ae.put('max_num_epochs', 50)
ae.put('optimization_method', NNOM_GRADIENT_DESCENT)
ae.put('gd_mini_batch_size', 128)
ae.put('gd_learning_rate', 0.0001)
ae.put('epsilon', 0.0)
# start fine-tuning. this might take some time
_ = ae.train(Xtrain)
# -
# ## Evaluation
# Now we can evaluate the autoencoder that we trained. We'll start by providing it with corrupted inputs and looking at how it will reconstruct them. The function [reconstruct()](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#ae8c2d565cf2ea809103d0557c57689c7) is used to obtain the reconstructions:
# +
# get a 50-example subset of the test set
subset = Xtest[:,0:50].copy()
# corrupt the first 25 examples with multiplicative noise
subset[:,0:25] *= (random.random((256,25))>0.5)
# corrupt the other 25 examples with additive noise
subset[:,25:50] += random.random((256,25))
# obtain the reconstructions
reconstructed_subset = ae.reconstruct(features(subset))
# plot the corrupted data and the reconstructions
figure(figsize=(10,10))
for i in range(50):
ax1=subplot(10,10,i*2+1)
ax1.imshow(subset[:,i].reshape((16,16)), interpolation='nearest', cmap = cm.Greys_r)
ax1.set_xticks([])
ax1.set_yticks([])
ax2=subplot(10,10,i*2+2)
ax2.imshow(reconstructed_subset[:,i].reshape((16,16)), interpolation='nearest', cmap = cm.Greys_r)
ax2.set_xticks([])
ax2.set_yticks([])
# -
# The figure shows the corrupted examples and their reconstructions. The top half of the figure shows the ones corrupted with multiplicative noise, the bottom half shows the ones corrupted with additive noise. We can see that the autoencoders can provide decent reconstructions despite the heavy noise.
#
# Next we'll look at the weights that the first hidden layer has learned. To obtain the weights, we can call the [get_layer_parameters()]() function, which will return a vector containing both the weights and the biases of the layer. The biases are stored first in the array followed by the weights matrix in column-major format.
# +
# obtain the weights matrix of the first hidden layer
# the 512 is the number of biases in the layer (512 neurons)
# the transpose is because numpy stores matrices in row-major format, and Shogun stores
# them in column major format
w1 = ae.get_layer_parameters(1)[512:].reshape(256,512).T
# visualize the weights between the first 100 neurons in the hidden layer
# and the neurons in the input layer
figure(figsize=(10,10))
for i in range(100):
ax1=subplot(10,10,i+1)
ax1.imshow(w1[i,:].reshape((16,16)), interpolation='nearest', cmap = cm.Greys_r)
ax1.set_xticks([])
ax1.set_yticks([])
# -
# Now, we can use the autoencoder to initialize a supervised neural network. The network will have all the layer of the autoencoder up to (and including) the middle layer. We'll also add a softmax output layer. So, the network will look like: L1->L2->L3->Softmax. The network is obtained by calling [convert_to_neural_network()](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CDeepAutoencoder.html#a8c179cd9a503b2fa78b9bfe10ae473e5):
# +
from shogun import NeuralSoftmaxLayer
nn = ae.convert_to_neural_network(NeuralSoftmaxLayer(10))
nn.put('max_num_epochs', 50)
nn.put('labels', Ytrain)
_ = nn.train(Xtrain)
# -
# Next, we'll evaluate the accuracy on the test set:
# +
from shogun import MulticlassAccuracy
predictions = nn.apply_multiclass(Xtest)
accuracy = MulticlassAccuracy().evaluate(predictions, Ytest) * 100
print("Classification accuracy on the test set =", accuracy, "%")
# -
# ## Convolutional Autoencoders
# Convolutional autoencoders [3] are the adaptation of autoencoders to images (or other spacially-structured data). They are built with convolutional layers where each layer consists of a number of feature maps. Each feature map is produced by convolving a small filter with the layer's inputs, adding a bias, and then applying some non-linear activation function. Additionally, a max-pooling operation can be performed on each feature map by dividing it into small non-overlapping regions and taking the maximum over each region. In this section we'll pre-train a [convolutional network](http://deeplearning.net/tutorial/lenet.html) as a stacked autoencoder and use it for classification.
#
# In Shogun, convolutional autoencoders are constructed and trained just like regular autoencoders. Except that we build the autoencoder using [CNeuralConvolutionalLayer](http://www.shogun-toolbox.org/doc/en/latest/classshogun_1_1CNeuralConvolutionalLayer.html) objects:
# +
from shogun import DynamicObjectArray, NeuralInputLayer, NeuralConvolutionalLayer, CMAF_RECTIFIED_LINEAR
conv_layers = DynamicObjectArray()
# 16x16 single channel images
conv_layers.append_element(NeuralInputLayer(16,16,1))
# the first encoding layer: 5 feature maps, filters with radius 2 (5x5 filters)
# and max-pooling in a 2x2 region: its output will be 10 8x8 feature maps
conv_layers.append_element(NeuralConvolutionalLayer(CMAF_RECTIFIED_LINEAR, 5, 2, 2, 2, 2))
# the second encoding layer: 15 feature maps, filters with radius 2 (5x5 filters)
# and max-pooling in a 2x2 region: its output will be 20 4x4 feature maps
conv_layers.append_element(NeuralConvolutionalLayer(CMAF_RECTIFIED_LINEAR, 15, 2, 2, 2, 2))
# the first decoding layer: same structure as the first encoding layer
conv_layers.append_element(NeuralConvolutionalLayer(CMAF_RECTIFIED_LINEAR, 5, 2, 2))
# the second decoding layer: same structure as the input layer
conv_layers.append_element(NeuralConvolutionalLayer(CMAF_RECTIFIED_LINEAR, 1, 2, 2))
conv_ae = DeepAutoencoder(conv_layers)
# -
# Now we'll pre-train the autoencoder:
# +
conv_ae.pt_noise_type.set_const(AENT_DROPOUT) # use dropout noise
conv_ae.pt_noise_parameter.set_const(0.3) # each input has a 30% chance of being set to zero
conv_ae.pt_optimization_method.set_const(NNOM_GRADIENT_DESCENT) # train using gradient descent
conv_ae.pt_gd_learning_rate.set_const(0.002)
conv_ae.pt_gd_mini_batch_size.set_const(100)
conv_ae.pt_max_num_epochs[0] = 30 # max number of epochs for pre-training the first encoding layer
conv_ae.pt_max_num_epochs[1] = 10 # max number of epochs for pre-training the second encoding layer
conv_ae.pt_epsilon.set_const(0.0) # disable automatic convergence testing
# start pre-training. this might take some time
conv_ae.pre_train(Xtrain)
# -
# And then convert the autoencoder to a regular neural network for classification:
# +
conv_nn = ae.convert_to_neural_network(NeuralSoftmaxLayer(10))
# train the network
conv_nn.put('epsilon', 0.0)
conv_nn.put('max_num_epochs', 50)
conv_nn.put('labels', Ytrain)
# start training. this might take some time
_ = conv_nn.train(Xtrain)
# -
# And evaluate it on the test set:
# +
predictions = conv_nn.apply_multiclass(Xtest)
accuracy = MulticlassAccuracy().evaluate(predictions, Ytest) * 100
print("Classification accuracy on the test set =", accuracy, "%")
# -
# ## References
# - [1] [Stacked Denoising Autoencoders: Learning Useful Representations in a Deep Network with a Local Denoising Criterion, Vincent, 2010](http://jmlr.org/papers/volume11/vincent10a/vincent10a.pdf)
# - [2] [Contractive Auto-Encoders: Explicit Invariance During Feature Extraction, Rifai, 2011](http://machinelearning.wustl.edu/mlpapers/paper_files/ICML2011Rifai_455.pdf)
# - [3] [Stacked Convolutional Auto-Encoders for Hierarchical Feature Extraction, <NAME>, 2011](http://www.idsia.ch/~ciresan/data/icann2011.pdf)
|
doc/ipython-notebooks/neuralnets/autoencoders.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp models.MultiInputNet
# -
# # MultiInputNet
#
# > This is an implementation created by <NAME> - <EMAIL>. It can be used to combine different types of deep learning models into a single one that will accept multiple inputs from a MixedDataLoaders.
#export
from tsai.imports import *
from tsai.models.layers import *
from tsai.models.utils import *
#export
class MultiInputNet(Module):
def __init__(self, *models, c_out=None, reshape_fn=None, multi_output=False, joint_head=None, device=None, **kwargs):
r"""
Args:
models : list of models (one model per dataloader in dls). They all must have a head.
c_out : output layer size.
reshape_fn : callable to transform a 3d input into a 2d input (Noop, Reshape(-1), GAP1d())
multi_output : determines if the model creates M+1 output (one per model plus a combined one), or just a single output (combined one).
joint_head : allows you to pass a custom joint head. If None a MLP will be created (you can pass 'layers' to this default head using kwargs)
device : cpu or cuda. If None, default_device() will be chosen.
kwargs : head kwargs
"""
c_out = ifnone(c_out, get_layers(models[0], cond=is_linear)[-1].out_features)
self.M = len(models)
self.m = []
self.backbones = nn.ModuleList()
self.heads = nn.ModuleList()
head_nf = 0
min_nf = np.inf
for i, model in enumerate(models):
self.heads.append(model.head)
model.head = Identity()
self.backbones.append(model)
self.m.append(Sequential(self.backbones[-1], self.heads[-1]))
head_nf += model.head_nf
min_nf = min(min_nf, model.head_nf)
self.head_nf = head_nf
if joint_head is None:
if not 'layers' in kwargs: layers = [head_nf, min_nf, min_nf]
head = create_fc_head(layers=layers, c_out=c_out, **kwargs)
else:
head = joint_head(self.head_nf, c_out, **kwargs)
self.heads.append(head)
self.multi_output = multi_output
self.m.append(self)
self.reshape = ifnone(reshape_fn, GAP1d())
self.concat = Concat(dim=1)
device = ifnone(device, default_device())
self.to(device=device)
def forward(self, xs):
xs = tuple(*xs) if len(xs) == 1 else xs
out = []
for k in range(self.M):
x = xs[k]
# Create separate features
feat = self.backbones[k](*x) if isinstance(x, (list, tuple, L)) else self.backbones[k](x)
# Process features separately
if self.training and self.multi_output: out.append(self.heads[k](feat))
# Concat features
if feat.ndim == 3: feat = self.reshape(feat)
concat_feats = feat if k==0 else self.concat([concat_feats, feat])
# Process joint features
out.append(self.heads[-1](concat_feats))
if self.training and self.multi_output: return out
else: return out[0]
from fastai.data.transforms import *
from tsai.data.all import *
from tsai.models.utils import *
from tsai.models.XCM import *
from tsai.models.TabModel import *
dsid = 'NATOPS'
X, y, splits = get_UCR_data(dsid, split_data=False)
ts_features_df = get_ts_features(X, y)
# +
# raw ts
tfms = [None, [Categorize()]]
batch_tfms = TSStandardize()
ts_dls = get_ts_dls(X, y, splits=splits, tfms=tfms, batch_tfms=batch_tfms)
ts_model = build_ts_model(XCM, dls=ts_dls, window_perc=.5)
# ts features
cat_names = None
cont_names = ts_features_df.columns[:-2]
y_names = 'target'
tab_dls = get_tabular_dls(ts_features_df, cat_names=cat_names, cont_names=cont_names, y_names=y_names, splits=splits)
tab_model = build_tabular_model(TabModel, dls=tab_dls)
# mixed
mixed_dls = get_mixed_dls(ts_dls, tab_dls)
MultiModalNet = MultiInputNet(ts_model, tab_model)
learn = Learner(mixed_dls, MultiModalNet, metrics=[accuracy, RocAuc()])
learn.fit_one_cycle(1, 1e-3)
# -
#hide
out = create_scripts(); beep(out)
|
nbs/130_models.MultiInputNet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import numpy.linalg as nplin
import itertools
import timeit
np.random.seed(0)
def operators(s):
#generate terms in the energy function
n_seq,n_var = s.shape
ops = np.zeros((n_seq,n_var+int(n_var*(n_var-1)/2.0)))
jindex = 0
for index in range(n_var):
ops[:,jindex] = s[:,index]
jindex +=1
for index in range(n_var-1):
for index1 in range(index+1,n_var):
ops[:,jindex] = s[:,index]*s[:,index1]
jindex +=1
return ops
def energy_ops(ops,w):
return np.sum(ops*w[np.newaxis,:],axis=1)
def generate_seqs(n_var,n_seq,n_sample=30,g=1.0):
samples = np.random.choice([1.0,-1.0],size=(n_seq*n_sample,n_var),replace=True)
ops = operators(samples)
n_ops = ops.shape[1]
#w_true = g*(np.random.rand(ops.shape[1])-0.5)/np.sqrt(float(n_var))
w_true = np.random.normal(0.,g/np.sqrt(n_var),size=n_ops)
sample_energy = energy_ops(ops,w_true)
p = np.exp(sample_energy)
p /= np.sum(p)
out_samples = np.random.choice(np.arange(n_seq*n_sample),size=n_seq,replace=True,p=p)
return w_true,samples[out_samples] #,p[out_samples],sample_energy[out_samples]
def eps_machine(s,eps_scale=0.1,max_iter=151,alpha=0.1):
MSE = np.zeros(max_iter)
KL = np.zeros(max_iter)
E_av = np.zeros(max_iter)
n_seq,n_var = s.shape
ops = operators(s)
n_ops = ops.shape[1]
cov_inv = np.eye(ops.shape[1])
np.random.seed(13)
w = np.random.rand(n_ops)-0.5
#w_iter = np.zeros((max_iter,n_ops))
for i in range(max_iter):
eps_scale = np.random.rand()/np.max([1.,np.max(np.abs(w))])
energies_w = energy_ops(ops,w)
probs_w = np.exp(-energies_w*(1-eps_scale))
z_data = np.sum(probs_w)
probs_w /= z_data
ops_expect_w = np.sum(probs_w[:,np.newaxis]*ops,axis=0)
#if i%int(10) == 0:
E_exp = (probs_w*energies_w).sum()
#KL[i] = -E_exp - np.log(z_data) + np.sum(np.log(np.cosh(w*eps_scale))) + n_var*np.log(2.)
#if i > 50 and KL[i] >= KL[i-1]: break
E_av[i] = energies_w.mean()
MSE[i] = ((w-w_true)**2).mean()
sec_order = w*eps_scale
w += alpha*cov_inv.dot((ops_expect_w - sec_order))
#w_iter[i,:] = w
#return MSE,KL,E_av,w_iter
return MSE,-E_av,w
def hopfield_model(s):
ops = operators(s)
w = np.mean(ops,axis=0)
#print('hopfield error ',nplin.norm(w-w_true))
return w
def boltzmann_machine_exact(s,s_all,max_iter=150,alpha=5e-2,cov=False):
n_seq,n_var = s.shape
ops = operators(s)
cov_inv = np.eye(ops.shape[1])
ops_obs = np.mean(ops,axis=0)
ops_model = operators(s_all)
n_ops = ops.shape[1]
np.random.seed(13)
w = np.random.rand(n_ops)-0.5
for iterate in range(max_iter):
energies_w = energy_ops(ops_model,w)
probs_w = np.exp(energies_w)
probs_w /= np.sum(probs_w)
if iterate%10 == 0:
#print(iterate,nplin.norm(w-w_true)) #,nplin.norm(spin_cov_w-spin_cov_obs))
MSE = ((w-w_true)**2).mean()
#print(iterate,MSE)
w += alpha*cov_inv.dot(ops_obs - np.sum(ops_model*probs_w[:,np.newaxis],axis=0))
#print('final',iterate,MSE)
return w
# +
#print('Hopfield:')
#w1 = hopfield_model(seqs)
#print(((w1-w_true)**2).mean())
# -
nn_var = 6
n_var_list = np.linspace(50,100,nn_var).astype(int)
n_var_list
# +
max_iter = 100
n_seq = 10000
#n_var = 10
g = 1.
run_time1 = np.zeros(nn_var)
run_time2 = np.zeros(nn_var)
for i,n_var in enumerate(n_var_list):
n_var = int(n_var)
w_true,seqs = generate_seqs(n_var,n_seq,g=g)
print('esp machine:')
start_time1 = timeit.default_timer()
MSE,E_av,w = eps_machine(seqs,eps_scale=0.1,max_iter=max_iter)
run_time1[i] = timeit.default_timer() - start_time1
print('run_time:',n_var,run_time1[i],MSE[-1])
#print('Boltzmann (exact):')
#start_time2 = timeit.default_timer()
#s_all = np.asarray(list(itertools.product([1.0, -1.0], repeat=n_var)))
#w2 = boltzmann_machine_exact(seqs,s_all,cov=False)
#run_time2[i] = timeit.default_timer() - start_time2
#print('run_time:',run_time2[i])
# -
print(run_time1)
np.savetxt('run_time_eps_random_50_100.txt',(n_var_list,run_time1),fmt='%f')
|
Ref/fig2_compare_all_method/fig3d_time/1main_time_run-eps_random_m50-100_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in **Data Science Interview Questions** series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/DataScience_Interview_Questions)**
# </i></small></small>
# # Predictive Modeling ➞ <span class='label label-default'>19 Questions</span>
# ### 1. (Given a Dataset) Analyze this dataset and give me a model that can predict this response variable.
# <span class='label label-default'>Solution</span>
#
# - Problem Determination ➞ Data Cleaning ➞ Feature Engineering ➞ Modeling
#
# - Benchmark Models
# - Linear Regression (Ridge or Lasso) for regression
# - Logistic Regression for Classification
#
# - Advanced Models
# - Random Forest, Boosting Trees, and so on
# - Scikit-Learn, XGBoost, LightGBM, CatBoost
#
# - Determine if the problem is classification or regression.
#
# - Plot and visualize the data.
#
# - Start by fitting a simple model (multivariate regression, logistic regression), do some feature engineering accordingly, and then try some complicated models. Always split the dataset into train, validation, test dataset and use cross validation to check their performance.
#
# - Favor simple models that run quickly and you can easily explain.
#
# - Mention cross validation as a means to evaluate the model.
# ### 2. What could be some issues if the distribution of the test data is significantly different than the distribution of the training data?
# <span class='label label-default'>Solution</span>
#
# - The model that has high training accuracy might have low test accuracy. Without further knowledge, it is hard to know which dataset represents the population data and thus the generalizability of the algorithm is hard to measure. This should be mitigated by repeated splitting of train vs. test dataset (as in cross validation).
# - When there is a change in data distribution, this is called the dataset shift. If the train and test data has a different distribution, then the classifier would likely overfit to the train data.
# - This issue can be overcome by using a more general learning method.
# - This can occur when:
# - $P(y|x)$ are the same but $P(x)$ are different. (covariate shift)
# - $P(y|x)$ are different. (concept shift)
# - The causes can be:
# - Training samples are obtained in a biased way. (sample selection bias)
# - Train is different from test because of temporal, spatial changes. (non-stationary environments)
# - Solution to covariate shift
# - importance weighted cv
# ### 3. What are some ways I can make my model more robust to outliers?
# <span class='label label-default'>Solution</span>
#
# - We can have regularization such as L1 or L2 to reduce variance (increase bias).
# - Changes to the algorithm:
# - Use tree-based methods instead of regression methods as they are more resistant to outliers. For statistical tests, use non parametric tests instead of parametric ones.
# - Use robust error metrics such as MAE or Huber Loss instead of MSE.
# - Changes to the data:
# - Winsorizing the data
# - Transforming the data (e.g. log)
# - Remove them only if you’re certain they’re anomalies and not worth predicting
# ### What are some differences you would expect in a model that minimizes squared error, versus a model that minimizes absolute error? In which cases would each error metric be appropriate?
# <span class='label label-default'>Solution</span>
#
# - MSE is more strict to having outliers. MAE is more robust in that sense, but is harder to fit the model for because it cannot be numerically optimized. So when there are less variability in the model and the model is computationally easy to fit, we should use MAE, and if that’s not the case, we should use MSE.
# - MSE: easier to compute the gradient, MAE: linear programming needed to compute the gradient
# - MAE more robust to outliers. If the consequences of large errors are great, use MSE
# - MSE corresponds to maximizing likelihood of Gaussian random variables
# ### 5. What error metric would you use to evaluate how good a binary classifier is? What if the classes are imbalanced? What if there are more than 2 groups?
# <span class='label label-default'>Solution</span>
#
# - Accuracy: proportion of instances you predict correctly.
# - Pros: intuitive, easy to explain
# - Cons: works poorly when the class labels are imbalanced and the signal from the data is weak
# - ROC curve and AUC: plot false-positive-rate (fpr) on the x axis and true-positive-rate (tpr) on the y axis for different threshold. Given a random positive instance and a random negative instance, the AUC is the probability that you can identify who's who.
# - Pros: Works well when testing the ability of distinguishing the two classes.
# - Cons: can’t interpret predictions as probabilities (because AUC is determined by rankings), so can’t explain the uncertainty of the model, and it doesn't work for multi-class case.
# - logloss/deviance/cross entropy:
# - Pros: error metric based on probabilities
# - Cons: very sensitive to false positives, negatives
# - When there are more than 2 groups, we can have k binary classifications and add them up for logloss. Some metrics like AUC is only applicable in the binary case.
# ### 6. What are various ways to predict a binary response variable? Can you compare two of them and tell me when one would be more appropriate? What’s the difference between these? (SVM, Logistic Regression, Naive Bayes, Decision Tree, etc.)
# <span class='label label-default'>Solution</span>
#
# - Things to look at: N, P, linearly separable, features independent, likely to overfit, speed, performance, memory usage and so on.
# - Logistic Regression
# - features roughly linear, problem roughly linearly separable
# - robust to noise, use l1,l2 regularization for model selection, avoid overfitting
# - the output come as probabilities
# - efficient and the computation can be distributed
# - can be used as a baseline for other algorithms
# - (-) can hardly handle categorical features
# - SVM
# - with a nonlinear kernel, can deal with problems that are not linearly separable
# - (-) slow to train, for most industry scale applications, not really efficient
# - Naive Bayes
# - computationally efficient when P is large by alleviating the curse of dimensionality
# - works surprisingly well for some cases even if the condition doesn’t hold
# - with word frequencies as features, the independence assumption can be seen reasonable. So the algorithm can be used in text categorization
# - (-) conditional independence of every other feature should be met
# - Tree Ensembles
# - good for large N and large P, can deal with categorical features very well
# - non parametric, so no need to worry about outliers
# - GBT’s work better but the parameters are harder to tune
# - RF works out of the box, but usually performs worse than GBT
# - Deep Learning
# - works well for some classification tasks (e.g. image)
# - used to squeeze something out of the problem
# ### 7. What is regularization and where might it be helpful? What is an example of using regularization in a model?
# <span class='label label-default'>Solution</span>
#
# - Regularization is useful for reducing variance in the model, meaning avoiding overfitting.
# - For example, we can use L1 regularization in Lasso regression to penalize large coefficients and automatically select features, or we can also use L2 regularization for Ridge regression to penalize the feature coefficients.
# ### 8. Why might it be preferable to include fewer predictors over many?
# <span class='label label-default'>Solution</span>
#
# - When we add irrelevant features, it increases model's tendency to overfit because those features introduce more noise. When two variables are correlated, they might be harder to interpret in case of regression, etc.
# - curse of dimensionality
# - adding random noise makes the model more complicated but useless
# - computational cost
# - Ask someone for more details.
# ### 9. Given training data on tweets and their retweets, how would you predict the number of retweets of a given tweet after 7 days after only observing 2 days worth of data?
# <span class='label label-default'>Solution</span>
#
# - Build a time series model with the training data with a seven day cycle and then use that for a new data with only 2 days data.
# - Ask someone for more details.
# - Build a regression function to estimate the number of retweets as a function of time t
# - to determine if one regression function can be built, see if there are clusters in terms of the trends in the number of retweets
# - if not, we have to add features to the regression function
# - features + # of retweets on the first and the second day ➞ predict the seventh day
# - https://en.wikipedia.org/wiki/Dynamic_time_warping
# ### 10. How could you collect and analyze data to use social media to predict the weather?
# <span class='label label-default'>Solution</span>
#
# - We can collect social media data using twitter, Facebook, instagram API’s.
# - Then, for example, for twitter, we can construct features from each tweet, e.g. the tweeted date, number of favorites, retweets, and of course, the features created from the tweeted content itself.
# - Then use a multivariate time series model to predict the weather.
# - Ask someone for more details.
# ### 11. How would you construct a feed to show relevant content for a site that involves user interactions with items?
# <span class='label label-default'>Solution</span>
#
# - We can do so using building a recommendation engine.
# - The easiest we can do is to show contents that are popular other users, which is still a valid strategy if for example the contents are news articles.
# - To be more accurate, we can build a content based filtering or collaborative filtering. If there’s enough user usage data, we can try collaborative filtering and recommend contents other similar users have consumed. If there isn’t, we can recommend similar items based on vectorization of items (content based filtering).
# ### 12. How would you design the people you may know feature on LinkedIn or Facebook?
# <span class='label label-default'>Solution</span>
#
# - Find strong unconnected people in weighted connection graph
# - Define similarity as how strong the two people are connected
# - Given a certain feature, we can calculate the similarity based on
# - friend connections (neighbors)
# - Check-in’s people being at the same location all the time.
# - same college, workplace
# - Have randomly dropped graphs test the performance of the algorithm
# - Ref. News Feed Optimization
# - Affinity score: how close the content creator and the users are
# - Weight: weight for the edge type (comment, like, tag, etc.). Emphasis on features the company wants to promote
# - Time decay: the older the less important
# ### 13. How would you predict who someone may want to send a Snapchat or Gmail to?
# <span class='label label-default'>Solution</span>
#
# - for each user, assign a score of how likely someone would send an email to
# - the rest is feature engineering:
# - number of past emails, how many responses, the last time they exchanged an email, whether the last email ends with a question mark, features about the other users, etc.
# - Ask someone for more details.
# - People who someone sent emails the most in the past, conditioning on time decay.
# ### 14. How would you suggest to a franchise where to open a new store?
# <span class='label label-default'>Solution</span>
#
# - build a master dataset with local demographic information available for each location.
# - local income levels, proximity to traffic, weather, population density, proximity to other businesses
# - a reference dataset on local, regional, and national macroeconomic conditions (e.g. unemployment, inflation, prime interest rate, etc.)
# - any data on the local franchise owner-operators, to the degree the manager
# - identify a set of KPIs acceptable to the management that had requested the analysis concerning the most desirable factors surrounding a franchise
# - quarterly operating profit, ROI, EVA, pay-down rate, etc.
# - run econometric models to understand the relative significance of each variable
# - run machine learning algorithms to predict the performance of each location candidate
# ### 15. In a search engine, given partial data on what the user has typed, how would you predict the user’s eventual search query?
# <span class='label label-default'>Solution</span>
#
# - Based on the past frequencies of words shown up given a sequence of words, we can construct conditional probabilities of the set of next sequences of words that can show up (n-gram). The sequences with highest conditional probabilities can show up as top candidates.
# - To further improve this algorithm,
# - we can put more weight on past sequences which showed up more recently and near your location to account for trends
# - show your recent searches given partial data
# - Personalize and localize the search
# - Use the user's historical search data
# - Use the historical data from the local region
# ### 16. Given a database of all previous alumni donations to your university, how would you predict which recent alumni are most likely to donate?
# <span class='label label-default'>Solution</span>
#
# - Based on frequency and amount of donations, graduation year, major, etc, construct a supervised regression (or binary classification) algorithm.
# ### 17. You’re Uber and you want to design a heatmap to recommend to drivers where to wait for a passenger. How would you approach this?
# <span class='label label-default'>Solution</span>
#
# - Based on the past pickup location of passengers around the same time of the day, day of the week (month, year), construct
# - Ask someone for more details.
# - Based on the number of past pickups
# - account for periodicity (seasonal, monthly, weekly, daily, hourly)
# - special events (concerts, festivals, etc.) from tweets
# ### 18. How would you build a model to predict a March Madness bracket?
# <span class='label label-default'>Solution</span>
#
# - One vector each for team A and B. Take the difference of the two vectors and use that as an input to predict the probability that team A would win by training the model. Train the models using past tournament data and make a prediction for the new tournament by running the trained model for each round of the tournament
# - Some extensions:
# - Experiment with different ways of consolidating the 2 team vectors into one (e.g concantenating, averaging, etc)
# - Consider using a RNN type model that looks at time series data.
# ### 19. You want to run a regression to predict the probability of a flight delay, but there are flights with delays of up to 12 hours that are really messing up your model. How can you address this?
# <span class='label label-default'>Solution</span>
#
# - This is equivalent to making the model more robust to outliers.
# - See **Question 3**.
|
02_Predictive_Modeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to NumPy
#
# NumPy is one of the two most important libraries in Python for data science, along with pandas (which we will cover in Section 4). NumPy is a crucial library for effectively loading, storing, and manipulating in-memory data in Python, all of which will be at the heart of what you do with data science in Python.
#
# Datasets come from a huge range of sources and in a wide range of formats, such as text documents, images, sound clips, numerical measurements, and nearly anything else. Despite this variety, however, the start of data science is to think of all data fundamentally as arrays of numbers.
#
# For example, the words in documents can be represented as the numbers that encode letters in computers or even the frequency of particular words in a collection of documents. Digital images can be thought of as two-dimensional arrays of numbers representing pixel brightness or color. Sound files can be represented as one-dimensional arrays of frequency versus time. However, no matter what form our data takes, in order to analyze it, our first step will be to transform it into arrays of numbers—which is where NumPy comes in (and pandas down the road).
#
# NumPy is short for *Numerical Python*, and it provides an efficient means of storing and operating on dense data buffers in Python. Array-oriented computing in Python goes back to 1995 with the Numeric library. Scientific programming in Python took off over the next 10 years, but the collections of libraries splintered. The NumPy project began in 2005 as a means of bringing the Numeric and NumArray projects together around a single array-based framework.
#
# Some examples in this section are drawn from the *Python Data Science Handbook* by <NAME> (content available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook)) and *Python for Data Analysis* by <NAME>. Text from the *Python Data Science Handbook* is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode); code is released under the [MIT license](https://opensource.org/licenses/MIT).
#
# Let's get started exploring NumPy! Our first step will be to import NumPy using `np` as an alias:
import numpy as np
# Get used to this convention — it's a common convention in Python, and it's the way we will use and refer to NumPy throughout the rest of this course.
# ## Built-In Help
#
# There's a lot to learn about NumPy, and it can be tough to remember it all the first time through. Don't worry! IPython — the underlying program that enables notebooks like this one to interact with Python—has you covered.
#
# First off, IPython gives you the ability to quickly explore the contents of a package like NumPy by using the tab-completion feature. So, if you want to see all of the functions available with NumPy, type this:
#
# ```ipython
# In [2]: np.<TAB>
# ```
# When you do so, a drop-down menu will appear next to the `np.`
#
# > **Exercise**
# Place your cursor after the period and press <TAB>:
np.
# From the drop-down menu, you can select any function to run. Better still, you can select any function and view the built-in help documentation for it. For example, to view the documentation for the NumPy `add()` function, you can run this code:
#
# ```ipython
# In [3]: np.add?
# ```
# Try this with a few different functions. Remember, these functions are just like ones you wrote in Section 2; the documentation will help explain what parameters you can (or should) provide the function, in addition to what output you can expect.
#
# > **Exercise**
# +
# Replace 'add' below with a few different NumPy function names and look over the documentation:
# np.add?
# -
# For more detailed documentation (along with additional tutorials and other resources), visit [www.numpy.org](http://www.numpy.org).
#
# Now that you know how to quickly get help while you are working on your own, let's return to storing data in arrays.
# ## NumPy arrays: a specialized data structure for analysis
#
# > **Learning goal:** By the end of this subsection, you should have a basic understanding of what NumPy arrays are and how they differ from the other Python data structures you have studied thus far.
#
# We started the discussion in this section by noting that data science starts by representing data as arrays of numbers.
#
# "Wait!" you might be thinking. "Can't we just use Python lists for that?"
#
# Depending on the data, yes, you could (and you will use lists as a part of working with data in Python). But to see what we might want to use a specialized data structure for, let's look a little more closely at lists.
# ### Lists in Python
#
# Python lists can hold just one kind of object. Let's use one to create a list of just integers:
myList = list(range(10))
myList
# Remember list comprehension? We can use it to probe the data types of items in a list:
[type(item) for item in myList]
# Of course, a really handy feature of Python lists is that they can hold heterogeneous types of data in a single list object:
myList2 = [True, "2", 3.0, 4]
[type(item) for item in myList2]
# However, this flexibility comes at a price. Each item in a list is really a separate Python object (the list is an object itself, true, but mostly it is an object that serves as a container for the memory pointers to the constituent objects). That means that each item in a list must contain its own type info, reference count, and other information. All of this information can become expensive in terms of memory and performance if we are dealing with hundreds of thousands or millions of items in a list. Moreover, for many uses in data science, our arrays just store a single type of data (such as integers or floats), which means that all of the object-related information for items in such an array would be redundant. It can be much more efficient to store data in a fixed-type array.
#
# <img align="left" style="padding-right:10px;" src="Graphics/Sec3_array_vs_list.png">
#
# Enter the fixed-type, NumPy-style array.
# ### Fixed-type arrays in Python
#
# At the level of implementation by the computer, the `ndarray` that is part of the NumPy package contains a single pointer to one contiguous block of data. This is efficient memory-wise and computationally. Better still, NumPy provides efficient *operations* on data stored in `ndarray` objects.
#
# (Note that we will pretty much use “array,” “NumPy array,” and “ndarray” interchangeably throughout this section to refer to the ndarray object.)
# #### Creating NumPy arrays method 1: using Python lists
#
# There are multiple ways to create arrays in NumPy. Let's start by using our good old familiar Python lists. We will use the `np.array()` function to do this (remember, we imported NumPy as '`np`'):
# Create an integer array:
np.array([1, 4, 2, 5, 3])
# Remember that, unlike Python lists, NumPy constrains arrays to contain a single type. So, if data types fed into a NumPy array do not match, NumPy will attempt to *upcast* them if possible. To see what we mean, here NumPy upcasts integers to floats:
np.array([3.14, 4, 2, 3])
# > **Exercise**
# What happens if you construct an array using a list that contains a combination of integers, floats, and strings?
# If you want to explicitly set the data type of your array when you create it, you can use the `dtype` keyword:
np.array([1, 2, 3, 4], dtype='float32')
# > **Exercise**
# Try this using a different dtype.
# Remember that you can always refer to the documentation with the command np.array.
# Most usefully for a lot of applications in data science, NumPy arrays can explicitly be multidimensional (like matrices or tensors). Here's one way of creating a multidimensional array using a list of lists:
# nested lists result in multi-dimensional arrays
np.array([range(i, i + 3) for i in [2, 4, 6]])
# The inner lists in a list of lists are treated as rows of the two-dimensional array you created.
# #### Creating NumPy arrays method 2: building from scratch
#
# In practice, it is often more efficient to create arrays from scratch using functions built into NumPy, particularly for larger arrays. Here are a few examples; these example will help introduce you to several useful NumPy functions.
# Create an integer array of length 10 filled with zeros
np.zeros(10, dtype=int)
# Create a 3x5 floating-point array filled with ones
np.ones((3, 5), dtype=float)
# Create a 3x5 array filled with 3.14
# The first number in the tuple gives the number of rows
# The second number in the tuple sets the number of columns
np.full((3, 5), 3.14)
# Create an array filled with a linear sequence
# Starting at 0, ending at 20, stepping by 2
# (this is similar to the built-in Python range() function)
np.arange(0, 20, 2)
# Create an array of five values evenly spaced between 0 and 1
np.linspace(0, 1, 5)
# Create a 3x3 array of uniformly distributed
# random values between 0 and 1
np.random.random((3, 3))
# Create a 3x3 array of normally distributed random values
# with mean 0 and standard deviation 1
np.random.normal(0, 1, (3, 3))
# Create a 3x3 array of random integers in the interval [0, 10)
np.random.randint(0, 10, (3, 3))
# Create a 3x3 identity matrix
np.eye(3)
# Create an uninitialized array of three integers
# The values will be whatever happens to already exist at that memory location
np.empty(3)
# Now take a couple of minutes to go back and play with these code snippets, changing the parameters. These functions are the bread-and-butter of creating NumPy arrays and you will want to become comfortable with them.
# Below is a table listing out several of the array-creation functions in NumPy.
#
# | Function | Description |
# |:--------------|:------------|
# | `array` | Converts input data (list, tuple, array, or other sequence type) to an ndarray either |
# | | by inferring a dtype or explicitly specifying a dtype. Copies the input data by default. |
# | `asarray` | Converts input to ndarray, but does not copy if the input is already an ndarray. |
# | `arange` | Similar to the built-in `range()` function but returns an ndarray instead of a list. |
# | `ones`, `ones_like` | Produces an array of all 1s with the given shape and dtype. |
# | | `ones_like` takes another array and produces a ones-array of the same shape and dtype. |
# | `zeros`, `zeros_like` | Similar to `ones` and `ones_like` but producing arrays of 0s instead. |
# | `empty`, `empty_like` | Creates new arrays by allocating new memory, but does not populate with any values
# | | like `ones` and `zeros`. |
# | `full`, `full_like` | Produces an array of the given shape and dtype with all values set to the indicated “fill value.” |
# | | `full_like` takes another array and produces a a filled array of the same shape and dtype. |
# | `eye`, `identity` | Create a square $N \times N$ identity matrix (1s on the diagonal and 0s elsewhere) |
# ### NumPy data types
#
# The standard NumPy data types are listed in the following table. Note that when constructing an array, they can be specified using a string:
#
# ```python
# np.zeros(8, dtype='int16')
# ```
#
# Or they can be specified directly using the NumPy object:
#
# ```python
# np.zeros(8, dtype=np.int16)
# ```
# | Data type | Description |
# |:--------------|:------------|
# | ``bool_`` | Boolean (True or False) stored as a byte |
# | ``int_`` | Default integer type (same as C ``long``; normally either ``int64`` or ``int32``)|
# | ``intc`` | Identical to C ``int`` (normally ``int32`` or ``int64``)|
# | ``intp`` | Integer used for indexing (same as C ``ssize_t``; normally either ``int32`` or ``int64``)|
# | ``int8`` | Byte (-128 to 127)|
# | ``int16`` | Integer (-32768 to 32767)|
# | ``int32`` | Integer (-2147483648 to 2147483647)|
# | ``int64`` | Integer (-9223372036854775808 to 9223372036854775807)|
# | ``uint8`` | Unsigned integer (0 to 255)|
# | ``uint16`` | Unsigned integer (0 to 65535)|
# | ``uint32`` | Unsigned integer (0 to 4294967295)|
# | ``uint64`` | Unsigned integer (0 to 18446744073709551615)|
# | ``float_`` | Shorthand for ``float64``.|
# | ``float16`` | Half-precision float: sign bit, 5 bits exponent, 10 bits mantissa|
# | ``float32`` | Single-precision float: sign bit, 8 bits exponent, 23 bits mantissa|
# | ``float64`` | Double-precision float: sign bit, 11 bits exponent, 52 bits mantissa|
# | ``complex_`` | Shorthand for ``complex128``.|
# | ``complex64`` | Complex number, represented by two 32-bit floats|
# | ``complex128``| Complex number, represented by two 64-bit floats|
# If these data types seem a lot like those in C, that's because NumPy is built in C.
#
# > **Takeaway:** NumPy arrays are a data structure similar to Python lists that provide high performance when storing and working on large amounts of homogeneous data—precisely the kind of data that you will encounter frequently in doing data science. NumPy arrays support many data types beyond those discussed in this course. With all of that said, however, don’t worry about memorizing all of the NumPy dtypes. **It’s often just necessary to care about the general kind of data you’re dealing with: floating point, integer, Boolean, string, or general Python object.**
# ## Working with NumPy arrays: the basics
#
# > **Learning goal:** By the end of this subsection, you should be comfortable working with NumPy arrays in basic ways.
#
# Now that you know how to create arrays in NumPy, you need to get comfortable manipulating them for two reasons. First, you will work with NumPy arrays as part of your exploration of data science. Second, our other important Python data-science tool, pandas, is actually built around NumPy. Getting good at working with NumPy arrays will pay dividends in the next section (Section 4) and beyond: NumPy arrays are the building blocks for the `Series` and `DataFrame` data structures in the Python pandas library and you will use them *a lot* in data science. To get comfortable with array manipulation, we will cover five specifics:
# - **Arrays attributes**: Assessing the size, shape, and data types of arrays
# - **Indexing arrays**: Getting and setting the value of individual array elements
# - **Slicing arrays**: Getting and setting smaller subarrays within a larger array
# - **Reshaping arrays**: Changing the shape of a given array
# - **Joining and splitting arrays**: Combining multiple arrays into one and splitting one array into multiple arrays
# ### Array attributes
# First, let's look at some array attributes. We'll start by defining three arrays filled with random numbers: one one-dimensional, another two-dimensional, and the last three-dimensional. Because we will be using NumPy's random number generator, we will set a *seed* value in order to ensure that you get the same random arrays each time you run this code:
# +
import numpy as np
np.random.seed(0) # seed for reproducibility
a1 = np.random.randint(10, size=6) # One-dimensional array
a2 = np.random.randint(10, size=(3, 4)) # Two-dimensional array
a3 = np.random.randint(10, size=(3, 4, 5)) # Three-dimensional array
# -
# Each array has attributes ``ndim`` (the number of dimensions of an array), ``shape`` (the size of each dimension of an array), and ``size`` (the total number of elements in an array).
#
# > **Exercise:**
# Change the values in this code snippet to look at the attributes for a1, a2, and a3:
print("a3 ndim: ", a3.ndim)
print("a3 shape:", a3.shape)
print("a3 size: ", a3.size)
# Another useful array attribute is the `dtype`, which we already encountered earlier in this section as a means of determining the type of data in an array:
print("dtype:", a3.dtype)
# > **Exercise:**
# Explore the dtype for the other arrays.
# What dtypes do you predict them to have?
print("dtype:", a3.dtype)
# ### Indexing arrays
#
# Indexing in NumPy is pretty similar to indexing lists in standard Python. In fact, indices in one-dimensional arrays work exactly as they do with Python lists:
a1
a1[0]
a1[4]
# As with regular Python lists, in order to index from the end of the array, you can use negative indices:
a1[-1]
a1[-2]
# > **Exercise:**
# Do multidimensional NumPy arrays work like Python lists of lists?
# Try a few combinations like a2[1][1] or a3[0][2][1] and see what comes back
# You might have noticed that we can treat multidimensional arrays like lists of lists. But a more common means of accessing items in multidimensional arrays is to use a comma-separated tuple of indices.
#
# (Yes, we realize that these comma-separated tuples use square brackets rather than the parentheses the name might suggest, but they are nevertheless referred to as tuples.)
a2
a2[0, 0]
a2[2, 0]
a2[2, -1]
# You can also modify values by use of this same comma-separated index notation:
a2[0, 0] = 12
a2
# Remember, once defined, NumPy arrays have a fixed data type. So, if you attempt to insert a float into an integer array, the value will be silently truncated.
a1[0] = 3.14159
a1
# > **Exercise:**
# What happens if you try to insert a string into a1?
# Hint: try both a string like '3' and one like 'three'
# ### Slicing arrays
# Similar to how you can use square brackets to access individual array elements, you can also use them to access subarrays. You do this with the *slice* notation, marked by the colon (`:`) character. NumPy slicing syntax follows that of the standard Python list; so, to access a slice of an array `a`, use this notation:
# ``` python
# a[start:stop:step]
# ```
# If any of these are unspecified, they default to the values ``start=0``, ``stop=``*``size of dimension``*, ``step=1``.
# Let's take a look at accessing sub-arrays in one dimension and in multiple dimensions.
# #### One-dimensional slices
a = np.arange(10)
a
a[:5] # first five elements
a[5:] # elements after index 5
a[4:7] # middle sub-array
a[::2] # every other element
a[1::2] # every other element, starting at index 1
# > **Exercise:**
# How would you access the *last* five elements of array a?
# How about every other element of the last five elements of a?
# Hint: Think back to list indexing in Python
# Be careful when using negative values for ``step``. When ``step`` has a negative value, the defaults for ``start`` and ``stop`` are swapped and you can use this functionality to reverse an array:
a[::-1] # all elements, reversed
a[5::-2] # reversed every other from index 5
# > **Exercise:**
# How can you create a slice that contains every third element of a
# descending from the second-to-last element to the second element of a?
# #### Multidimensional slices
#
# Multidimensional slices use the same slice notation of one-dimensional subarrays mixed with the comma-separated notation of multidimensional arrays. Some examples will help illustrate this.
a2
a2[:2, :3] # two rows, three columns
a2[:3, ::2] # all rows, every other column
# Finally, subarray dimensions can even be reversed together:
a2[::-1, ::-1]
# > **Exercise:**
# Now try to show 2 rows and 4 columns with every other element?
# #### Accessing array rows and columns
# One thing you will often need to do in manipulating data is accessing a single row or column in an array. You can do this through a combination of indexing and slicing; specifically by using an empty slice marked by a single colon (``:``). Again, some examples will help illustrate this.
print(a2[:, 0]) # first column of x2
print(a2[0, :]) # first row of x2
# In the case of row access, the empty slice can be omitted for a more compact syntax:
print(a2[0]) # equivalent to a2[0, :]
# > **Exercise:**
# How would you access the third column of a3?
# How about the third row of a3?
# #### Slices are no-copy views
# It's important to know that slicing produces *views* of array data, not *copies*. This is a **huge** difference between NumPy array slicing and Python list slicing. With Python lists, slices are only shallow copies of lists; if you modify a copy, it doesn't affect the parent list. When you modify a NumPy subarray, you modify the original list. Be careful: this can have ramifications when you are trying to just work with a small part of a large dataset and you don’t want to change the whole thing. Let's look more closely.
print(a2)
# Extract a $2 \times 2$ subarray from `a2`:
a2_sub = a2[:2, :2]
print(a2_sub)
# Now modify this subarray:
a2_sub[0, 0] = 99
print(a2_sub)
# `a2` is now modified as well:
print(a2)
# > **Exercise:**
# Now try reversing the column and row order of a2_sub
# Does a2 look the way you expected it would after that manipulation?
# The fact that slicing produces views rather than copies is useful for data-science work. As you work with large datasets, you will often find that it is easier to access and manipulate pieces of those datasets rather than copying them entirely.
# #### Copying arrays
# Instead of just creating views, sometimes it is necessary to copy the data in one array to another. When you need to do this, use the `copy()` method:
a2_sub_copy = a2[:2, :2].copy()
print(a2_sub_copy)
# If we now modify this subarray, the original array is not touched:
a2_sub_copy[0, 0] = 42
print(a2_sub_copy)
print(a2)
# ### Reshaping arrays
# Another way in which you will need to manipulate arrays is by reshaping them. This involves changing the number and size of dimensions of an array. This kind of manipulation can be important in getting your data to meet the expectations of machine learning programs or APIs.
#
# The most flexible way of doing this kind of manipulation is with the `reshape` method. For example, if you want to put the numbers 1 through 9 in a $3 \times 3$ grid, you can do the following:
grid = np.arange(1, 10).reshape((3, 3))
print(grid)
# Another common manipulation you will do in data science is converting one-dimensional arrays into two-dimensional row or column matrices. This can be a common necessity when doing linear algebra for machine learning. While you can do this by means of the `reshape` method, an easier way is to use the `newaxis` keyword in a slice operation:
# +
a = np.array([1, 2, 3])
# row vector via reshape
a.reshape((1, 3))
# -
# row vector via newaxis
a[np.newaxis, :]
# column vector via reshape
a.reshape((3, 1))
# column vector via newaxis
a[:, np.newaxis]
# You will see this type of transformation a lot in the remainder of this course.
# ### Joining and splitting arrays
#
# Another common data-manipulation need in data science is combining multiple datasets; learning first how to do this with NumPy arrays will help you in the next section (Section 4) when we do this with more complex data structures. You will many times also need to split a single array into multiple arrays.
# #### Joining arrays
# To join arrays in NumPy, you will most often use `np.concatenate`, which is the method we will cover here. If you find yourself in the future needing to specifically join arrays in mixed dimensions (a rarer case), read the documentation on `np.vstack`, `np.hstack`, and `np.dstack`.
# ##### `np.concatenate()`
#
# `np.concatenate` takes a tuple or list of arrays as its first argument:
a = np.array([1, 2, 3])
b = np.array([3, 2, 1])
np.concatenate([a, b])
# You can also concatenate more than two arrays at once:
c = [99, 99, 99]
print(np.concatenate([a, b, c]))
# `np.concatenate` can also be used for two-dimensional arrays:
grid = np.array([[1, 2, 3],
[4, 5, 6]])
# concatenate along the first axis, which is the default
np.concatenate([grid, grid])
# > **Exercise:**
# Recall that axes are zero-indexed in NumPy.
# What do you predict np.concatenate([grid, grid], axis=1) will produce?
# #### Splitting arrays
# In order to split arrays into multiple smaller arrays, you can use the functions ``np.split``, ``np.hsplit``, ``np.vsplit``, and ``np.dsplit``. As above, we will only cover the most commonly used function (`np.split`) in this course.
# ##### `np.split()`
# Let's first examine the case of a one-dimensional array:
a = [1, 2, 3, 99, 99, 3, 2, 1]
a1, a2, a3 = np.split(a, [3, 5])
print(a1, a2, a3)
# Notice that *N* split-points produces to *N + 1* subarrays. In this case it has formed the subarray `a2` with `a[3]` and `a[4]` (the element just before position 5 [remember how Python indexing goes], the second input in the tuple) as elements. `a1` and `a3` pick up the leftover portions from the original array `a`.
# > **Exercise:**
grid = np.arange(16).reshape((4, 4))
grid
# What does np.split(grid, [1, 2]) produce?
# What about np.split(grid, [1, 2], axis=1)?
# > **Takeaway:** Manipulating datasets is a fundamental part of preparing data for analysis. The skills you learned and practiced here will form building blocks for the most sophisticated data-manipulation you will learn in later sections in this course.
# ## Fancy indexing
#
# So far, we have explored how to access and modify portions of arrays using simple indices like `arr[0]`) and slices like `arr[:5]`. Now it is time for fancy indexing, in which we pass an array of indices to an array in order to access or modify multiple array elements at the same time.
#
# Let's try it out:
# +
rand = np.random.RandomState(42)
arr = rand.randint(100, size=10)
print(arr)
# -
# Suppose you need to access three different elements. Using the tools you currently have, your code might look something like this:
[arr[3], arr[7], arr[2]]
# With fancy indexing, you can pass a single list or array of indices to do the same thing:
ind = [3, 7, 4]
arr[ind]
# Another useful aspect of fancy indexing is that the shape of the output array reflects the shape of the *index arrays* you supply, rather than the shape of the array you are accessing. This is handy because there will be many times in a data scientist's life when they want to grab data from an array in a particular manner, such as to pass it to a machine learning API. Let's examine this property with an example:
ind = np.array([[3, 7],
[4, 5]])
arr[ind]
# `arr` is a one-dimensional array, but `ind`, your index array, is a $2 \times 2$ array, and that is the shape the results comes back in.
#
# > **Exercise:**
# What happens when your index array is bigger than the target array?
# Hint: you could use a large one-dimensional array or something fancier like ind = np.arange(0, 12).reshape((6, 2))
# Fancy indexing also works in multiple dimensions:
arr2 = np.arange(12).reshape((3, 4))
arr2
# As with standard indexing, the first index refers to the row and the second to the column:
row = np.array([0, 1, 2])
col = np.array([2, 1, 3])
arr2[row, col]
# What did you actually get as your final result here? The first value in the result array is `arr2[0, 2]`, the second one is `arr2[1, 1]`, and the third one is `arr2[2, 3]`.
#
# The pairing of indices in fancy indexing follows all the same broadcasting rules we covered earlier. Thus, if you combine a column vector and a row vector within the indices, you get a two-dimensional result:
arr2[row[:, np.newaxis], col]
# Here, each row value is matched with each column vector, exactly as we saw in broadcasting of arithmetic operations.
#
# > **Exercise:**
# Now try broadcasting this on your own.
# What do you get with row[:, np.newaxis] * col?
# Or row[:, np.newaxis] * row? col[:, np.newaxis] * row?
# What about col[:, np.newaxis] * row?
# Hint: think back to the broadcast rules
# **The big takeaway:** It is always important to remember that fancy indexing returns values reflected by the *broadcasted shape of the indices*, and not the shape of the array being indexed.
# ### Combined indexing
#
# You can also combine fancy indexing with the other indexing schemes you have learned. Consider `arr2` again:
print(arr2)
# Now combine fancy and simple indices:
arr2[2, [2, 0, 1]]
# What did you get back? The elements at positions 2, 0, and 1 of row 2 (the third row).
#
# You can also combine fancy indexing with slicing:
arr2[1:, [2, 0, 1]]
# Again, consider what you got back as output: the elements at positions 2, 0, and 1 of each row after the first one (so the second and third rows).
#
# Of course, you can also combine fancy indexing with masking:
mask = np.array([1, 0, 1, 0], dtype=bool)
arr2[row[:, np.newaxis], mask]
# ### Modifying values using fancy indexing
#
# Fancy indexing is, of course, not just for accessing parts of an array, but also for modifying parts of an array:
ind = np.arange(10)
arr = np.array([2, 1, 8, 4])
ind[arr] = 99
print(ind)
# You can also use a ufunc here and subtract 10 from each element of the array:
ind[arr] -= 10
print(ind)
# Be cautious when using repeated indices with operations like these. They might not always produce the results you expect. For example:
ind = np.zeros(10)
ind[[0, 0]] = [4, 6]
print(ind)
# Where did the 4 go? The result of this operation is to first assign `ind[0] = 4`, followed by `ind[0] = 6`. So the result is that `ind[0]` contains the value 6.
#
# But not every operation repeats the way you might think it should:
arr = [2, 3, 3, 4, 4, 4]
ind[arr] += 1
ind
# We might have expected that `ind[3]` would contain the value 2 and `ind[4]` would contain the value 3. After all, that is how many times each index is repeated. So what happened?
#
# This happened because `ind[arr] += 1` is really shorthand for `ind[arr] = ind[arr] + 1`. `ind[arr] + 1` is evaluated, and then the result is assigned to the indices in `ind`. So, similar to the previous example, this is not augmentation that happens multiple times, but an assignment, which can lead to potentially counterintuitive results.
#
# But what if you want an operation to repeat? To do this, use the `at()` method of ufuncs:
ind = np.zeros(10)
np.add.at(ind, arr, 1)
print(ind)
# > **Exercise:**
# What does np.subtract.at(ind, arr, 1) give you?
# Play around with some of the other ufuncs we have seen.
# > **Takeaway:** Fancy indexing enables you to select and manipulate several array members at once. This type of programmatic data manipulation is common in data science: often what you want to do with your data you want to do on several data points at once.
# ## Sorting arrays
#
# So far we have just worried about accessing and modifying NumPy arrays. Another huge thing you will need to do as a data scientist is sort array data. Sorting is often an important means of teasing out the structure in data (such as outlying data points).
#
# Although you could use Python's built-in `sort` and `sorted` functions, they will not work nearly as efficiently as NumPy's `np.sort` function.
#
# `np.sort` returns a sorted version of an array without modifying the input:
a = np.array([2, 1, 4, 3, 5])
np.sort(a)
# In order to sort the array in-place, use the `sort` method directly on arrays:
a.sort()
print(a)
# A related function is `argsort`, which returns the *indices* of the sorted elements rather than the elements themselves:
a = np.array([2, 1, 4, 3, 5])
b = np.argsort(a)
print(b)
# The first element of this result gives the index of the smallest element, the second value gives the index of the second smallest, and so on. These indices can then be used (via fancy indexing) to reconstruct the sorted array:
a[b]
# ### Sorting along rows or columns
#
# A useful feature of NumPy's sorting algorithms is the ability to sort along specific rows or columns of a multidimensional array using the `axis` argument. For example:
rand = np.random.RandomState(42)
table = rand.randint(0, 10, (4, 6))
print(table)
# Sort each column of the table
np.sort(table, axis=0)
# Sort each row of the table
np.sort(table, axis=1)
# Bear in mind that this treats each row or column as an independent array; any relationships between the row or column values will be lost doing this kind of sorting.
# ## Partial sorting: partitioning
#
# Sometimes you don't need to sort an entire array, you just need to find the *k* smallest values in the array (often when looking at the distance of data points from one another). NumPy supplies this functionality through the `np.partition` function. `np.partition` takes an array and a number *k*; the result is a new array with the smallest *k* values to the left of the partition, and the remaining values to the right (in arbitrary order):
arr = np.array([7, 2, 3, 1, 6, 5, 4])
np.partition(arr, 3)
# Note that the first three values in the resulting array are the three smallest in the array, and the remaining array positions contain the remaining values. Within the two partitions, the elements have arbitrary order.
#
# Similarly to sorting, we can partition along an arbitrary axis of a multidimensional array:
np.partition(table, 2, axis=1)
# The result is an array where the first two slots in each row contain the smallest values from that row, with the remaining values filling the remaining slots.
#
# Finally, just as there is an `np.argsort` that computes indices of the sort, there is an `np.argpartition` that computes indices of the partition. We'll see this in action in the following section when we discuss pandas.
#
# > **Takeaway:** Sorting your data is a fundamental means of exploring it and answering questions about it. The sorting algorithms in NumPy provide you with a fast, computationally efficient way of doing this on large amounts of data and with fine-grain control.
# ## Efficient computation on NumPy arrays: Universal functions
#
# > **Learning goal:** By the end of this subsection, you should have a basic understanding of what NumPy universal functions are and how (and why) to use them.
#
# Some of the properties that make Python great to work with for data science (its dynamic, interpreted nature, for example) can also make it slow. This is particularly true with looping. These small performance hits can add up to minutes (or longer) when dealing with truly huge datasets.
#
# When we first examined loops in Section 2, you probably didn't notice any delay: the loops were short enough that Python’s relatively slow looping wasn’t an issue. Consider this function, which calculates the reciprocal for an array of numbers:
# +
import numpy as np
np.random.seed(0)
def compute_reciprocals(values):
output = np.empty(len(values))
for i in range(len(values)):
output[i] = 1.0 / values[i]
return output
values = np.random.randint(1, 10, size=5)
compute_reciprocals(values)
# -
# Running this loop, it was probably difficult to even discern that execution wasn't instantaneous.
#
# But let’s try it on a much larger array. To empirically do this, we'll time this with IPython's `%timeit` magic command.
big_array = np.random.randint(1, 100, size=1000000)
# %timeit compute_reciprocals(big_array)
# You certainly noticed that delay. The slowness of this looping becomes noticeable when we repeat many small operations many times.
#
# The performance bottleneck is not the operations themselves, but the type-checking and function dispatches that Python performs on each cycle of the loop. In the case of the `compute_reciprocals` function above, each time Python computes the reciprocal, it first examines the object's type and does a dynamic lookup of the correct function to use for that type. Such is life with interpreted code. However, were we working with compiled code instead (such as in C), the object-type specification would be known before the code executes, and the result could be computed much more efficiently. This is where NumPy universal functions come into play.
# ### Ufuncs
#
# Universal functions in NumPy (often shortened to *ufuncs*) provide a statically typed, compiled function for many of the operations that we will need to run in the course of manipulating and analyzing data.
#
# Let's examine what this means in practice. Let's find the reciprocals of `big_array` again, this time using a built-in NumPy division ufunc on the array:
# %timeit (1.0 / big_array)
# That’s orders of magnitude better.
#
# Ufuncs can be used between a scalar and an array and between arrays of arbitrary dimensions.
#
# Computations vectorized by ufuncs are almost always more efficient than doing the same computation using Python loops. This is especially true on large arrays. When possible, try to use ufuncs when operating on NumPy arrays, rather than using ordinary Python loops.
#
# Ufuncs come in two flavors: *unary ufuncs*, which use a single input, and *binary ufuncs*, which operate on two inputs. The common ufuncs we'll look at here encompass both kinds.
# #### Array arithmetic
#
# Many NumPy ufuncs use Python's native arithmetic operators, so you can use the standard addition, subtraction, multiplication, and division operators that we covered in Section 1:
a = np.arange(4)
print("a =", a)
print("a + 5 =", a + 5)
print("a - 5 =", a - 5)
print("a * 2 =", a * 2)
print("a / 2 =", a / 2)
print("a // 2 =", a // 2) # floor division
# There are also ufuncs for negation, exponentiation, and the modulo operation:
print("-a = ", -a)
print("a ** 2 = ", a ** 2)
print("a % 2 = ", a % 2)
# You can also combine these ufuncs using the standard order of operations:
-(0.5*a + 1) ** 2
# The Python operators are not actually the ufuncs, but are rather wrappers around functions built into NumPy. So the `+` operator is actually a wrapper for the `add` function:
np.add(a, 2)
# Here is a cheat sheet for the equivalencies between Python operators and NumPy ufuncs:
#
# | Operator | Equivalent ufunc | Description |
# |:--------------|:--------------------|:--------------------------------------|
# |``+`` |``np.add`` |Addition (e.g., ``1 + 1 = 2``) |
# |``-`` |``np.subtract`` |Subtraction (e.g., ``3 - 2 = 1``) |
# |``-`` |``np.negative`` |Unary negation (e.g., ``-2``) |
# |``*`` |``np.multiply`` |Multiplication (e.g., ``2 * 3 = 6``) |
# |``/`` |``np.divide`` |Division (e.g., ``3 / 2 = 1.5``) |
# |``//`` |``np.floor_divide`` |Floor division (e.g., ``3 // 2 = 1``) |
# |``**`` |``np.power`` |Exponentiation (e.g., ``2 ** 3 = 8``) |
# |``%`` |``np.mod`` |Modulus/remainder (e.g., ``9 % 4 = 1``)|
#
# Python Boolean operators also work; we will explore those later in this section.
# #### Absolute value
#
# NumPy also understands Python's built-in absolute value function:
a = np.array([-2, -1, 0, 1, 2])
abs(a)
# This corresponds to the NumPy ufunc `np.absolute` (which is also available under the alias `np.abs`):
np.absolute(a)
np.abs(a)
# #### Exponents and logarithms
#
# You will need to use exponents and logarithms a lot in data science; these are some of the most common data transformations for machine learning and statistical work.
a = [1, 2, 3]
print("a =", a)
print("e^a =", np.exp(a))
print("2^a =", np.exp2(a))
print("3^a =", np.power(3, a))
# The basic `np.log` gives the natural logarithm; if you need to compute base-2 or base-10 logarithms, NumPy also provides those:
a = [1, 2, 4, 10]
print("a =", a)
print("ln(a) =", np.log(a))
print("log2(a) =", np.log2(a))
print("log10(a) =", np.log10(a))
# There are also some specialized versions of these ufuncs to help maintain precision when dealing with very small inputs:
a = [0, 0.001, 0.01, 0.1]
print("exp(a) - 1 =", np.expm1(a))
print("log(1 + a) =", np.log1p(a))
# These functions give more precise values than if you were to use the raw `np.log` or `np.exp` on very small values of `a`.
# #### Specialized ufuncs
#
# NumPy has many other ufuncs. Another source for specialized and obscure ufuncs is the submodule `scipy.special`. If you need to compute some specialized mathematical or statistical function on your data, chances are it is implemented in `scipy.special`.
from scipy import special
# Gamma functions (generalized factorials) and related functions
a = [1, 5, 10]
print("gamma(a) =", special.gamma(a))
print("ln|gamma(a)| =", special.gammaln(a))
print("beta(a, 2) =", special.beta(a, 2))
# > **Takeaway:** Universal functions in NumPy provide you with computational functions that are faster than regular Python functions, particularly when working on large datasets that are common in data science. This speed is important because it can make you more efficient as a data scientist and it makes a broader range of inquiries into your data tractable in terms of time and computational resources.
# ## Aggregations
#
# > **Learning goal:** By the end of this subsection, you should be comfortable aggregating data in NumPy.
#
# One of the first things you will find yourself doing with most datasets is computing the summary statistics for the data in order to get a general overview of your data before exploring it further. These summary statistics include the mean and standard deviation, in addition to other aggregates, such as the sum, product, median, minimum and maximum, or quantiles of the data.
#
# NumPy has fast built-in aggregation functions for working on arrays that are the subject of this sub-section.
# ### Summing the values of an array
#
# You can use the built-in Python `sum` function to sum up the values in an array.
import numpy as np
myList = np.random.random(100)
sum(myList)
# If you guessed that there is also a built-in NumPy function for this, you guessed correctly:
np.sum(myList)
# And if you guessed that the NumPy version is faster, you are doubly correct:
large_array = np.random.rand(1000000)
# %timeit sum(large_array)
# %timeit np.sum(large_array)
# For all their similarity, bear in mind that `sum` and `np.sum` are not identical; their optional arguments have different meanings, and `np.sum` is aware of multiple array dimensions.
# ### Minimum and maximum
#
# Just as Python has built-in `min` and `max` functions, NumPy has similar, vectorized versions:
np.min(large_array), np.max(large_array)
# You can also use `min`, `max`, and `sum` (and several other NumPy aggregates) as methods of the array object itself:
print(large_array.min(), large_array.max(), large_array.sum())
# ### Multidimensional aggregates
#
# Because you will often treat the rows and columns of two-dimensional arrays differently (treating columns as variables and rows as observations of those variables, for example), it can often be desirable to aggregate array data along a row or column. Let's consider a two-dimensional array:
md = np.random.random((3, 4))
print(md)
# Unless you specify otherwise, each NumPy aggregation function will compute the aggregate for the entire array. Hence:
md.sum()
# Aggregation functions take an additional argument specifying the *axis* along which to compute the aggregation. For example, we can find the minimum value within each column by specifying `axis=0`:
md.min(axis=0)
# > **Exercise:**
# What do you get when you try md.max(axis=1)?
# Remember that the `axis` keyword specifies the *dimension of the array that is to be collapsed*, not the dimension that will be returned. Thus specifying `axis=0` means that the first axis will be the one collapsed: for two-dimensional arrays, this means that values within each column will be aggregated.
# ### Other aggregation functions
#
# The table below lists other aggregation functions in NumPy. Most NumPy aggregates have a '`NaN`-safe' version, which computes the result while ignoring missing values marked by the `NaN` value.
#
# |Function Name | NaN-safe Version | Description |
# |:------------------|:--------------------|:----------------------------------------------|
# | ``np.sum`` | ``np.nansum`` | Compute sum of elements |
# | ``np.prod`` | ``np.nanprod`` | Compute product of elements |
# | ``np.mean`` | ``np.nanmean`` | Compute mean of elements |
# | ``np.std`` | ``np.nanstd`` | Compute standard deviation |
# | ``np.var`` | ``np.nanvar`` | Compute variance |
# | ``np.min`` | ``np.nanmin`` | Find minimum value |
# | ``np.max`` | ``np.nanmax`` | Find maximum value |
# | ``np.argmin`` | ``np.nanargmin`` | Find index of minimum value |
# | ``np.argmax`` | ``np.nanargmax`` | Find index of maximum value |
# | ``np.median`` | ``np.nanmedian`` | Compute median of elements |
# | ``np.percentile`` | ``np.nanpercentile``| Compute rank-based statistics of elements |
# | ``np.any`` | N/A | Evaluate whether any elements are true |
# | ``np.all`` | N/A | Evaluate whether all elements are true |
#
# We will see these aggregates often throughout the rest of the course.
# > **Takeaway:** Aggregation is the primary means you will use to explore you data, not just when using NumPy, but particularly in conjunction with pandas, the Python library you will learn about in the next section, which builds off of NumPy and thus off of everything you have learned thus far.
# ## Computation on arrays with broadcasting
#
# > **Learning goal:** By the end of this subsection, you should have a basic understanding of how broadcasting works in NumPy (and why NumPy uses it).
#
# Another means of vectorizing operations is to use NumPy's *broadcasting* functionality: creating rules for applying binary ufuncs like addition, subtraction, or multiplication on arrays of different sizes.
#
# Before, when we performed binary operations on arrays of the same size, those operations were performed on an element-by-element basis.
first_array = np.array([3, 6, 8, 1])
second_array = np.array([4, 5, 7, 2])
first_array + second_array
# Broadcasting enables you to perform these types of binary operations on arrays of different sizes. Thus, you could just as easily add a scalar (which is really just a zero-dimensional array) to an array:
first_array + 5
# Similarly, you can add a one-dimensional array to a two-dimensional array:
one_dim_array = np.ones((1))
one_dim_array
two_dim_array = np.ones((2, 2))
two_dim_array
one_dim_array + two_dim_array
# So far, so easy. But you can use broadcasting on arrays in more complicated ways. Consider this example:
# +
horizontal_array = np.arange(3)
vertical_array = np.arange(3)[:, np.newaxis]
print(horizontal_array)
print(vertical_array)
# -
horizontal_array + vertical_array
# ### Rules of broadcasting
# Broadcasting ollows a set of rules to determine the interaction between the two arrays:
# - **Rule 1**: If the two arrays differ in their number of dimensions, the shape of the one with fewer dimensions is *padded* with ones on its leading (left) side.
# - **Rule 2**: If the shape of the two arrays does not match in any dimension, the array with shape equal to 1 in that dimension is stretched to match the other shape.
# - **Rule 3**: If, in any dimension, the sizes disagree and neither is equal to 1, NumPy raises an error.
#
# Let's see these rules in action to better understand them.
# #### Broadcasting example 1
#
# Let's look at adding a two-dimensional array to a one-dimensional array:
two_dim_array = np.ones((2, 3))
one_dim_array = np.arange(3)
# Let's consider an operation on these two arrays. The shape of the arrays are:
#
# - `two_dim_array.shape = (2, 3)`
# - `one_dim_array.shape = (3,)`
#
# We see by rule 1 that the array `one_dim_array` has fewer dimensions, so we pad it on the left with ones:
#
# - `two_dim_array.shape -> (2, 3)`
# - `one_dim_array.shape -> (1, 3)`
#
# By rule 2, we now see that the first dimension disagrees, so we stretch this dimension to match:
#
# - `two_dim_array.shape -> (2, 3)`
# - `one_dim_array.shape -> (2, 3)`
#
# The shapes match, and we see that the final shape will be `(2, 3)`:
two_dim_array + one_dim_array
# > **Exercise:**
# Flip this around. Try adding these with two_dim_array = np.ones((3, 2))
# and one_dim_array = np.arange(3)[:, np.newaxis].
# What do you get?
# #### Broadcasting example 2
#
# Let's examine what happens when both arrays need to be broadcast:
vertical_array = np.arange(3).reshape((3, 1))
horizontal_array = np.arange(3)
# Again, we'll start by writing out the shape of the arrays:
#
# - `vertical_array.shape = (3, 1)`
# - `horizontal_array.shape = (3,)`
#
# Rule 1 says we must pad the shape of `horizontal_array ` with ones:
#
# - `vertical_array.shape -> (3, 1)`
# - `horizontal_array.shape -> (1, 3)`
#
# And rule 2 tells us that we upgrade each of these ones to match the corresponding size of the other array:
#
# - `vertical_array.shape -> (3, 3)`
# - `horizontal_array.shape -> (3, 3)`
#
# Because the result matches, these shapes are compatible. We can see this here:
vertical_array + horizontal_array
# #### Broadcasting example 3
#
# Here's what happens with incompatible arrays:
M = np.ones((3, 2))
i = np.arange(3)
# This is just a slightly different situation than in the first example: the matrix ``M`` is transposed.
# How does this affect the calculation? The shape of the arrays are:
#
# - ``M.shape = (3, 2)``
# - ``i.shape = (3,)``
#
# Again, rule 1 tells us that we must pad the shape of ``i`` with ones:
#
# - ``M.shape -> (3, 2)``
# - ``i.shape -> (1, 3)``
#
# By rule 2, the first dimension of ``i`` is stretched to match that of ``M``:
#
# - ``M.shape -> (3, 2)``
# - ``i.shape -> (3, 3)``
#
# Now we hit Rule 3: the final shapes do not match and the two arrays are incompatible:
M + i
# ### Broadcasting in practice
# Ufuncs enable you to avoid using slow Python loops; broadcasting builds on that.
#
# A common data practice is to *center* an array of data. For example, if we have an array of 10 observations, each of which consists of three values (called features in this context), we might want to center that data so that we have the differences from the mean rather than the raw data itself. Doing this can help us better compare the different values.
#
# We'll store this in a $10 \times 3$ array:
T = np.random.random((10, 3))
T
# Now compute the mean of each feature using the ``mean`` aggregate across the first dimension:
Tmean = T.mean(0)
Tmean
# Finally, center ``T`` by subtracting the mean. (This is a broadcasting operation.)
T_centered = T - Tmean
T_centered
# This is not just faster, but easier than writing a loop to do this.
# > **Takeaway:** The data you will work with in data science invariably comes in different shapes and sizes (at least in terms of the arrays in which you work with that data). The broadcasting functionality in NumPy enables you to use binary functions on irregularly fitting data in a predictable way.
# ## Comparisons, masks, and Boolean logic in NumPy
#
# > **Learning goal:** By the end of this subsection, you should be comfortable with and understand how to use Boolean masking in NumPy in order to answer basic questions about your data.
#
# *Masking* is when you want to manipulate or count or extract values in an array based on a criterion. For example, counting all the values in an array greater than a certain value is an example of masking. Boolean masking is often the most efficient way to accomplish these types of tasks in NumPy and it plays a large part in cleaning and otherwise preparing data for analysis (see Section 5).
# ### Example: Counting Rainy Days
#
# Let's see masking in practice by examining the monthly rainfall statistics for Seattle. The data is in a CSV file from data.gov. To load the data, we will use pandas, which we will formally introduce in Section 4.
# +
import numpy as np
import pandas as pd
# Use pandas to extract rainfall as a NumPy array
rainfall_2003 = pd.read_csv('Data/Observed_Monthly_Rain_Gauge_Accumulations_-_Oct_2002_to_May_2017.csv')['RG01'][ 2:14].values
rainfall_2003
# -
# Let’s break down what we just did in the code cell above. The rainfall data contains monthly rainfall totals from several rain gauges around the city of Seattle; we selected the first one. From that gauge, we then selected the relevant months for the first full calendar year in the dataset, 2003. That range of months started at the third row of the CSV file (remember, Python zero-indexes!) and ran through the thirteenth row, hence `2:14]`.
#
# You now have an array containing 12 values, each of which records the monthly rainfall in inches from January to December 2003.
#
# Commonly in data science, you will want to take a quick first exploratory look at the data. In this case, a bar chart is a good way to do this. To generate this bar chart, we will use Matplotlib, another important data-science tool that we will introduce formally later in the course. (This also brings up another widely used Python convention you should adopt: `import matplotlib.pyplot as plt`.)
# %matplotlib inline
import matplotlib.pyplot as plt
plt.bar(np.arange(1, len(rainfall_2003) + 1), rainfall_2003)
# To briefly interpret the code snippet above, we passed two parameters to the bar function in pyplot: the first defining the index for the x-axis and the second defining the data to use for the bars (the y-axis). To create the index, we use the NumPy function `arange` to create a sequence of numbers (this is the same `arange` we encountered earlier in this section). We know that the length of our array is 12, but it can be a good habit to get into to programmatically pass the length of an array in case it changes or you don’t know it with specificity. We also added 1 to both the start and the end of the `arange` to accommodate for Python zero-indexing (because there is no “month-zero” in the calendar).
#
# Looking at the chart above (and as residents can attest), Seattle can have lovely, sunny summers. However, this is only a first glimpse of the data. There are still several questions we would like to answer, such as in how many months did it rain, or what was the average precipitation in those months? We would use masking to answer those questions. (We will also return to this example dataset to demonstrate concepts throughout the rest of this section.) Before we dive deeper in explaining what masking is, we should briefly touch on comparison operators in NumPy.
# ### Comparison operators as ufuncs
#
# In addition to the computational operators as ufuncs that we have already encountered, NumPy also implements comparison operators such as `<` (less than) and `>` (greater than) as element-wise ufuncs. All of the standard Python comparison operations are available:
simple_array = np.array([1, 2, 3, 4, 5])
simple_array < 2 # less than
simple_array >= 4 # greater than or equal
simple_array == 2 # equal
# It is also possible to do an element-wise comparison of two arrays, and to include compound expressions:
(2 * simple_array) == (simple_array ** 2)
# As with the arithmetic operators, these comparison operators are wrappers for the NumPy ufuncs: when you write ``x < 3``, NumPy actually uses ``np.less(x, 3)``. Here is a summary of the comparison operators and their equivalent ufuncs:
#
# | Operator | Equivalent ufunc || Operator | Equivalent ufunc |
# |:--------------|:--------------------||:--------------|:--------------------|
# |``==`` |``np.equal`` ||``!=`` |``np.not_equal`` |
# |``<`` |``np.less`` ||``<=`` |``np.less_equal`` |
# |``>`` |``np.greater`` ||``>=`` |``np.greater_equal`` |
# Just like the arithmetic ufuncs, the comparison ufuncs work on arrays of any size and shape.
rand = np.random.RandomState(0)
two_dim_array = rand.randint(10, size=(3, 4))
two_dim_array
two_dim_array < 6
# The result is a Boolean array, and NumPy provides a number of straightforward patterns for working with these Boolean results.
# ## Working with Boolean arrays
#
# Given a Boolean array, there are a host of useful operations you can do.
# We'll work with `two_dim_array`, the two-dimensional array we created earlier.
print(two_dim_array)
# ### Counting entries
#
# To count the number of ``True`` entries in a Boolean array, ``np.count_nonzero`` is useful:
# how many values less than 6?
np.count_nonzero(two_dim_array < 6)
# We see that there are eight array entries that are less than 6.
# Another way to get at this information is to use ``np.sum``; in this case, ``False`` is interpreted as ``0``, and ``True`` is interpreted as ``1``:
np.sum(two_dim_array < 5)
# The benefit of `sum()` is that, like with other NumPy aggregation functions, this summation can be done along rows or columns as well:
# how many values less than 5 in each row?
np.sum(two_dim_array < 5, axis=1)
# This counts the number of values less than 5 in each row of the matrix.
#
# If we're interested in quickly checking whether any or all the values are true, we can use (you guessed it) ``np.any`` or ``np.all``:
# Are there any values less than zero?
np.any(two_dim_array < 0)
# > **Exercise:**
# Now check to see if all values less than 10?
# Hint: use np.all()
# ``np.all`` and ``np.any`` can be used along particular axes as well. For example:
# are all values in each row less than 7?
np.all(two_dim_array < 7, axis=1)
# Here, all the elements in the first and third rows are less than 7, while this is not the case for the second row.
#
# **A reminder:** Python has built-in `sum()`, `any()`, and `all()` functions. These have a different syntax than the NumPy versions, and, in particular, will fail or produce unintended results when used on multidimensional arrays. Be sure that you are using `np.sum()`, `np.any()`, and `np.all()` for these examples.
# ### Boolean operators
#
# We've already seen how we might count, say, all months with rain less than four inches, or all months with more than two inches of rain. But what if we want to know about all months with rain less than four inches and greater than one inch? This is accomplished through Python's *bitwise logic operators*, `&`, `|`, `^`, and `~`. Like with the standard arithmetic operators, NumPy overloads these as ufuncs which work element-wise on (usually Boolean) arrays.
#
# For example, we can address this sort of compound question as follows:
np.sum((rainfall_2003 > 0.5) & (rainfall_2003 < 1))
# So we see that there are two months with rainfall between 0.5 and 1.0 inches.
# Note that the parentheses here are important—because of operator-precedence rules, with parentheses removed, this expression would be evaluated as follows, which results in an error:
rainfall_2003 > (0.5 & rainfall_2003) < 1
# Using the equivalence of *A AND B and NOT (NOT A OR NOT B)* (which you might remember if you've taken an introductory logic course), we can compute the same result in a different manner:
np.sum(~((rainfall_2003 <= 0.5) | (rainfall_2003 >= 1)))
# Combining comparison operators and Boolean operators on arrays can lead to a wide range of efficient logical operations.
#
# The following table summarizes the bitwise Boolean operators and their equivalent ufuncs:
# | Operator | Equivalent ufunc || Operator | Equivalent ufunc |
# |:--------------|:--------------------||:--------------|:--------------------|
# |``&`` |``np.bitwise_and`` ||| |``np.bitwise_or`` |
# |``^`` |``np.bitwise_xor`` ||``~`` |``np.bitwise_not`` |
# Using these tools, you can start to answer the types of questions we listed above about the Seattle rainfall data. Here are some examples of results we can compute when combining masking with aggregations:
print("Number of months without rain:", np.sum(rainfall_2003 == 0))
print("Number of months with rain: ", np.sum(rainfall_2003 != 0))
print("Months with more than 1 inch: ", np.sum(rainfall_2003 > 1))
print("Rainy months with < 1 inch: ", np.sum((rainfall_2003 > 0) &
(rainfall_2003 < 1)))
# ## Boolean arrays as masks
#
# In the prior section, we looked at aggregates computed directly on Boolean arrays.
# A more powerful pattern is to use Boolean arrays as masks, to select particular subsets of the data themselves.
# Returning to our `two_dim_array` array from before, suppose we want an array of all values in the array that are less than 5:
two_dim_array
# You can obtain a Boolean array for this condition easily:
two_dim_array < 5
# Now, to *select* these values from the array, you can simply index on this Boolean array. This is the *masking* operation:
two_dim_array[two_dim_array < 5]
# What is returned is a one-dimensional array filled with all the values that meet your condition. Put another way, these are all the values in positions at which the mask array is ``True``.
#
# You can use masking as a way to compute some relevant statistics on the Seattle rain data:
# +
# Construct a mask of all rainy months
rainy = (rainfall_2003 > 0)
# Construct a mask of all summer months (June through September)
months = np.arange(1, 13)
summer = (months > 5) & (months < 10)
print("Median precip in rainy months in 2003 (inches): ",
np.median(rainfall_2003[rainy]))
print("Median precip in summer months in 2003 (inches): ",
np.median(rainfall_2003[summer]))
print("Maximum precip in summer months in 2003 (inches): ",
np.max(rainfall_2003[summer]))
print("Median precip in non-summer rainy months (inches):",
np.median(rainfall_2003[rainy & ~summer]))
# -
# > **Takeaway:** By combining Boolean operations, masking operations, and aggregates, you can quickly answer questions similar to those we posed about the Seattle rainfall data about any dataset. Operations like these will form the basis for the data exploration and preparation for analysis that will by our primary concerns in Sections 4 and 5.
|
Educator Resources/Reactor Workshops/Data Science/Track 1/Student_Notebook-Section_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="l9xBZPgkD9Ts"
TRAIN_PATH = "/content/drive/MyDrive/CovidDataset/Train"
VAL_PATH = "/content/drive/MyDrive/CovidDataset/Val"
# + id="UMKMAuJkFNST"
import numpy as np
import matplotlib.pyplot as plt
import keras
from keras.layers import *
from keras.models import *
from keras.preprocessing import image
# + id="_Nhtr7cYFQkJ"
# VGG16 Model in Keras
model = Sequential()
model.add(Conv2D(64, kernel_size=(3,3), activation='relu', input_shape=(224, 224, 3)))
model.add(Conv2D(64, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(128, (3,3), activation='relu'))
model.add(Conv2D(128, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(256, (3,3), activation='relu'))
model.add(Conv2D(256, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(Conv2D(512, (3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2), strides=(7, 7)))
model.add(Flatten())
model.add(Dense(4096, activation='relu'))
model.add(Dense(4096, activation='relu'))
model.add(Dense(1000, activation='relu'))
model.add(Dense(1, activation='softmax'))
model.compile(loss=keras.losses.binary_crossentropy, optimizer='adam', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="qGt43AyvHtZ8" outputId="642c4732-d2ff-4bec-e26e-64dcf6683389"
model.summary()
# + id="E1hLnd75HxZu"
# Train from scratch
train_datagen = image.ImageDataGenerator(
rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True
)
test_dataset = image.ImageDataGenerator(rescale=1./255)
# + colab={"base_uri": "https://localhost:8080/"} id="-z-3Dp1FH0eu" outputId="31e8f284-3987-41ff-873c-8bed273ba757"
train_generator = train_datagen.flow_from_directory(
'/content/drive/MyDrive/CovidDataset/Train',
target_size = (224, 224),
batch_size = 16,
class_mode = 'binary'
)
# + colab={"base_uri": "https://localhost:8080/"} id="tQLkZ7ORH3M5" outputId="05014c92-e1a6-4c7d-d870-c4f15b356537"
train_generator.class_indices
# + colab={"base_uri": "https://localhost:8080/"} id="vXY--BSkH5qc" outputId="2682b0d5-3460-4719-bdf8-a9c84b340b77"
validation_generator = test_dataset.flow_from_directory(
'/content/drive/MyDrive/CovidDataset/Val',
target_size = (224, 224),
batch_size = 16,
class_mode = 'binary'
)
# + id="VfDvgrqDIwLT"
trainImagesX = '/content/drive/MyDrive/CovidDataset/Train/Covid'
trainImagesY = '/content/drive/MyDrive/CovidDataset/Train/Normal'
testImagesX = '/content/drive/MyDrive/CovidDataset/Val/Covid'
testImagesY = '/content/drive/MyDrive/CovidDataset/Val/Normal'
# + colab={"base_uri": "https://localhost:8080/"} id="vyJgu0GnJYkW" outputId="33d9f212-7724-496b-889a-900c7301a693"
hist = model.fit_generator(
train_generator,
steps_per_epoch=8,
epochs=50,
validation_data = validation_generator,
validation_steps=2
)
# + colab={"base_uri": "https://localhost:8080/"} id="t84k-6tKNBMZ" outputId="9a8d0935-06ed-4f7f-b4fc-5b326198143e"
# Loss is very high
model.evaluate_generator(train_generator)
# + colab={"base_uri": "https://localhost:8080/"} id="XBkYPeXxNUQA" outputId="114aedda-7e75-4ef7-9ca2-3d1ef33e8bb0"
model.evaluate_generator(validation_generator)
# + id="QWd1nZ2tNY5V"
# Test Images
y_actual = []
y_test = []
# + id="hmmysAnUNbZa"
import os
# + id="8Cx3b2S6NypC"
for i in os.listdir("/content/drive/MyDrive/CovidDataset/Val/Normal/"):
img = image.load_img("/content/drive/MyDrive/CovidDataset/Val/Normal/"+i, target_size=(224, 224))
img = image.img_to_array(img)
img = np.expand_dims(img, axis=0)
p = model.predict(img)
classes_x=np.argmax(p, axis=1)
y_test.append(p[0, 0])
y_actual.append(1)
# + id="Zwrw_8ejN2iU"
for i in os.listdir("/content/drive/MyDrive/CovidDataset/Val/Covid/"):
img = image.load_img("/content/drive/MyDrive/CovidDataset/Val/Covid/"+i, target_size=(224, 224))
img = image.img_to_array(img)
img = np.expand_dims(img, axis=0)
p = model.predict(img)
classes_x=np.argmax(p, axis=1)
y_test.append(p[0, 0])
y_actual.append(0)
# + id="24sqjvQcN7Hr"
y_actual = np.array(y_actual)
y_test = np.array(y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="uukOLmNjN8YU" outputId="35e7efaa-160e-4a91-ccbf-b8ff4799b6d7"
y_actual
# + colab={"base_uri": "https://localhost:8080/"} id="JaSHOlJpN_XO" outputId="1de4cde2-dad6-4626-f4bb-32aeae4e2c6c"
y_test
# + id="riW9Viw3OD9Y"
from sklearn.metrics import confusion_matrix
# + id="Sbc-CArnOBON"
cm = confusion_matrix(y_actual, y_test)
# + id="jQO_gXZnOPa_"
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="4zYoKWmWOVz6" outputId="1b078dd3-eb62-42a8-8a98-753d7b06f704"
sns.heatmap(cm, cmap="plasma", annot=True)
# + id="rJWcNB7IOmz1"
def cnn_evaluation(model, history, train_images, train_labels, test_images, test_labels, class_names, model_name=None):
"""
Evaluates the performance of a CNN with loss and accuracy plots, a confusion matrix and a classification report for the training and test sets.
"""
train_acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
train_loss = history.history['loss']
val_loss = history.history['val_loss']
epch = range(1, len(train_acc) + 1)
plt.plot(epch, train_acc, 'g.', label='Training Accuracy')
plt.plot(epch, val_acc, 'g', label='Validation acc')
plt.title('Accuracy')
plt.legend()
plt.figure()
plt.plot(epch, train_loss, 'r.', label='Training loss')
plt.plot(epch, val_loss, 'r', label='Validation loss')
plt.title('Loss')
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="xVlbad9KOqQ_" outputId="4bdb18ac-ff97-48cb-dc5b-80c05048da5f"
cnn_evaluation(model, hist, trainImagesX, trainImagesY, testImagesX, testImagesY, class_names=['covid', 'no findings'])
|
Experiments and Weights/Third Week/VGG16_Major_Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import all dependencies
import pandas as pd
import numpy as np
import tensorflow as tf
from sklearn import preprocessing
#from pandas_profiling import ProfileReport
import matplotlib.pyplot as plt
tf.random.set_seed(123)
np.random.seed(123)
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Restrict TensorFlow to only use the fourth GPU
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
# +
#tf.config.threading.set_intra_op_parallelism_threads(6)
# -
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
# +
# read csv in the dataframea
df = pd.read_csv("../ane_energy/df_merged.csv",parse_dates=['dt_start_utc'],index_col='dt_start_utc')
# +
# show if some nans in our dataframe
print(np.any(np.isnan(df)))
# -
df.head()
# +
# lets have a look about our data, how its distributed
df.describe()
# +
#profile = ProfileReport(df, title="df :: Pandas Profiling Report", explorative=True)
#profile.to_file("Pandas_Report_epex_da_de.html")
# +
# helps us to find and replace inappropriate characters
import re
regex = re.compile(r"\[|\]|<", re.IGNORECASE)
df.columns = [regex.sub("_", col) if any(x in str(col) for x in set(('[', ']', '<'))) else col for col in df.columns.values]
# +
# Feature Importances
from numpy import loadtxt
from xgboost import XGBRegressor
from matplotlib import pyplot
model = XGBRegressor()
model.fit(df,df['rebap_eur_mwh'])
(pd.Series(model.feature_importances_, index=df.columns)
.nlargest(7)
.plot(kind='barh'));
# +
# defined function which preprocesses the data suitable for forecasting
def custom_ts_multi_data_prep(dataset, target, start, end, window, horizon):
X = []
y = []
start = start + window
if end is None:
end = len(dataset) - horizon
for i in range(start, end):
indices = range(i-window, i)
X.append(dataset[indices])
indicey = range(i+1, i+1+horizon)
y.append(target[indicey])
return np.array(X), np.array(y)
# +
# cut the last 24 Timestamps data for forecasting
validate = df.tail(24)
df.drop(df.tail(24).index,inplace=True)
# +
# drop the columns we dont need
df_1 = df.drop("rebap_eur_mwh", axis= 1)
# +
# MinMaxScaler to scale down the values.
# The neural network converges sooner when it exposes the same scaled features and gives better accuracy
x_scaler = preprocessing.MinMaxScaler()
y_scaler = preprocessing.MinMaxScaler()
dataX = x_scaler.fit_transform(df_1)
dataY = y_scaler.fit_transform(df[['rebap_eur_mwh']])
# +
# As we are doing multiple-step forecasting,
# let’s allow the model to see past 96 hours of data and forecast the 24 hrs after data;
# for that, we set the horizon to 24.
hist_window = 96
horizon = 24
TRAIN_SPLIT = 10000
x_train_multi, y_train_multi = custom_ts_multi_data_prep(
dataX, dataY, 0, TRAIN_SPLIT, hist_window, horizon)
x_val_multi, y_val_multi= custom_ts_multi_data_prep(
dataX, dataY, TRAIN_SPLIT, None, hist_window, horizon)
# -
print ('Single window of past history')
print(x_train_multi[0])
print ('\n Target horizon')
print (y_train_multi[0])
df.shape
# +
# Prepare the training data and validation data using the TensorFlow data function,
# which faster and efficient way to feed data for training.
BATCH_SIZE = 16
BUFFER_SIZE = 4
train_data_multi = tf.data.Dataset.from_tensor_slices((x_train_multi, y_train_multi))
train_data_multi = train_data_multi.cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE).repeat()
val_data_multi = tf.data.Dataset.from_tensor_slices((x_val_multi, y_val_multi))
val_data_multi = val_data_multi.batch(BATCH_SIZE).repeat()
# +
# Build and compile the model
lstm_multi = tf.keras.models.Sequential()
lstm_multi.add(tf.keras.layers.LSTM(150,input_shape=x_train_multi.shape[-2:],return_sequences=True))
lstm_multi.add(tf.keras.layers.Dropout(0.2)),
lstm_multi.add(tf.keras.layers.LSTM(units=100,return_sequences=False)),
lstm_multi.add(tf.keras.layers.Dropout(0.2)),
lstm_multi.add(tf.keras.layers.Dense(units=horizon)),
lstm_multi.compile(optimizer='adam', loss='mse')
# +
# save model
model_path = r'\Chapter_7\LSTM_Multivariatex.h5"
# +
# train the model
EVALUATION_INTERVAL = 150
EPOCHS = 150
history = lstm_multi.fit(train_data_multi, epochs=EPOCHS,steps_per_epoch=EVALUATION_INTERVAL,validation_data=val_data_multi, validation_steps=50,verbose =1,
callbacks =[tf.keras.callbacks.EarlyStopping(monitor='val_loss', min_delta=0, patience=10, verbose=1, mode='min'),tf.keras.callbacks.ModelCheckpoint(model_path,monitor='val_loss', save_best_only=True, mode='min', verbose=0)])
# +
# Load the saved model
Trained_model = tf.keras.models.load_model(model_path)
# + tags=[]
# plot our train and validation loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train loss', 'validation loss'], loc='upper left')
plt.rcParams["figure.figsize"] = [16,9]
plt.show()
# -
# Show the model architecture
Trained_model.summary()
# +
# Prepare the testing data for the last 96 hrs and check the prediction against it
# by visualizing the actual and predicted values.
data_val = x_scaler.fit_transform(df_1.tail(96))
# -
val_rescaled = data_val.reshape(1, data_val.shape[0], data_val.shape[1])
Predicted_results = Trained_model.predict(val_rescaled)
Predicted_results
Predicted_results_Inv_trans = y_scaler.inverse_transform(Predicted_results)
Predicted_results_Inv_trans
# +
# Finally, evaluate the result with standard performance metrics.
from sklearn import metrics
def timeseries_evaluation_metrics_func(y_true, y_pred):
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
print('Evaluation metric results:-')
print(f'MSE is : {metrics.mean_squared_error(y_true, y_pred)}')
print(f'MAE is : {metrics.mean_absolute_error(y_true, y_pred)}')
print(f'RMSE is : {np.sqrt(metrics.mean_squared_error(y_true, y_pred))}')
print(f'MAPE is : {mean_absolute_percentage_error(y_true, y_pred)}')
print(f'R2 is : {metrics.r2_score(y_true, y_pred)}',end='\n\n')
# +
# Results of the metrics
timeseries_evaluation_metrics_func(validate['rebap_eur_mwh'],Predicted_results_Inv_trans[0])
# +
# Plot the actual vs predicted data
plt.plot( list(validate['rebap_eur_mwh']))
plt.plot( list(Predicted_results_Inv_trans[0]))
plt.title("Actual vs Predicted")
plt.ylabel("rebap_eur_mwh")
plt.legend(('Actual','predicted'))
plt.show()
|
4_Modelling/Time_Series_Models/Predict_06_Hours/LSTM_Multivariate_Horizon_Style_06H.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon Fine Food Reviews Analysis
#
#
# Data Source: https://www.kaggle.com/snap/amazon-fine-food-reviews <br>
#
# EDA: https://nycdatascience.com/blog/student-works/amazon-fine-foods-visualization/
#
#
# The Amazon Fine Food Reviews dataset consists of reviews of fine foods from Amazon.<br>
#
# Number of reviews: 568,454<br>
# Number of users: 256,059<br>
# Number of products: 74,258<br>
# Timespan: Oct 1999 - Oct 2012<br>
# Number of Attributes/Columns in data: 10
#
# Attribute Information:
#
# 1. Id
# 2. ProductId - unique identifier for the product
# 3. UserId - unqiue identifier for the user
# 4. ProfileName
# 5. HelpfulnessNumerator - number of users who found the review helpful
# 6. HelpfulnessDenominator - number of users who indicated whether they found the review helpful or not
# 7. Score - rating between 1 and 5
# 8. Time - timestamp for the review
# 9. Summary - brief summary of the review
# 10. Text - text of the review
#
#
# #### Objective:
# Given a review, determine whether the review is positive (rating of 4 or 5) or negative (rating of 1 or 2).
#
# <br>
# [Q] How to determine if a review is positive or negative?<br>
# <br>
# [Ans] We could use Score/Rating. A rating of 4 or 5 can be cosnidered as a positive review. A rating of 1 or 2 can be considered as negative one. A review of rating 3 is considered nuetral and such reviews are ignored from our analysis. This is an approximate and proxy way of determining the polarity (positivity/negativity) of a review.
#
#
#
# # [1]. Reading Data
# ## [1.1] Loading the data
#
# The dataset is available in two forms
# 1. .csv file
# 2. SQLite Database
#
# In order to load the data, We have used the SQLITE dataset as it is easier to query the data and visualise the data efficiently.
# <br>
#
# Here as we only want to get the global sentiment of the recommendations (positive or negative), we will purposefully ignore all Scores equal to 3. If the score is above 3, then the recommendation wil be set to "positive". Otherwise, it will be set to "negative".
# +
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import sqlite3
import pandas as pd
import numpy as np
import nltk
import string
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import confusion_matrix
from sklearn import metrics
from sklearn.metrics import roc_curve, auc
from nltk.stem.porter import PorterStemmer
import re
# Tutorial about Python regular expressions: https://pymotw.com/2/re/
import string
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
from nltk.stem.wordnet import WordNetLemmatizer
from gensim.models import Word2Vec
from gensim.models import KeyedVectors
import pickle
from tqdm import tqdm
import os
# +
# using SQLite Table to read data.
con = sqlite3.connect('database.sqlite')
# filtering only positive and negative reviews i.e.
# not taking into consideration those reviews with Score=3
# SELECT * FROM Reviews WHERE Score != 3 LIMIT 500000, will give top 500000 data points
# you can change the number to any other number based on your computing power
# filtered_data = pd.read_sql_query(""" SELECT * FROM Reviews WHERE Score != 3 LIMIT 500000""", con)
# for tsne assignment you can take 5k data points
filtered_data = pd.read_sql_query(""" SELECT * FROM Reviews WHERE Score != 3 LIMIT 100000""", con)
# Give reviews with Score>3 a positive rating(1), and reviews with a score<3 a negative rating(0).
def partition(x):
if x < 3:
return 0
return 1
#changing reviews with score less than 3 to be positive and vice-versa
actualScore = filtered_data['Score']
positiveNegative = actualScore.map(partition)
filtered_data['Score'] = positiveNegative
print("Number of data points in our data", filtered_data.shape)
filtered_data.head(3)
# -
display = pd.read_sql_query("""
SELECT UserId, ProductId, ProfileName, Time, Score, Text, COUNT(*)
FROM Reviews
GROUP BY UserId
HAVING COUNT(*)>1
""", con)
print(display.shape)
display.head()
display[display['UserId']=='AZY10LLTJ71NX']
display['COUNT(*)'].sum()
# # [2] Exploratory Data Analysis
# ## [2.1] Data Cleaning: Deduplication
#
# It is observed (as shown in the table below) that the reviews data had many duplicate entries. Hence it was necessary to remove duplicates in order to get unbiased results for the analysis of the data. Following is an example:
display= pd.read_sql_query("""
SELECT *
FROM Reviews
WHERE Score != 3 AND UserId="AR5J8UI46CURR"
ORDER BY ProductID
""", con)
display.head()
# As it can be seen above that same user has multiple reviews with same values for HelpfulnessNumerator, HelpfulnessDenominator, Score, Time, Summary and Text and on doing analysis it was found that <br>
# <br>
# ProductId=B000HDOPZG was Loacker Quadratini Vanilla Wafer Cookies, 8.82-Ounce Packages (Pack of 8)<br>
# <br>
# ProductId=B000HDL1RQ was Loacker Quadratini Lemon Wafer Cookies, 8.82-Ounce Packages (Pack of 8) and so on<br>
#
# It was inferred after analysis that reviews with same parameters other than ProductId belonged to the same product just having different flavour or quantity. Hence in order to reduce redundancy it was decided to eliminate the rows having same parameters.<br>
#
# The method used for the same was that we first sort the data according to ProductId and then just keep the first similar product review and delelte the others. for eg. in the above just the review for ProductId=B000HDL1RQ remains. This method ensures that there is only one representative for each product and deduplication without sorting would lead to possibility of different representatives still existing for the same product.
#Sorting data according to ProductId in ascending order
sorted_data=filtered_data.sort_values('ProductId', axis=0, ascending=True, inplace=False, kind='quicksort', na_position='last')
#Deduplication of entries
final=sorted_data.drop_duplicates(subset={"UserId","ProfileName","Time","Text"}, keep='first', inplace=False)
final.shape
#Checking to see how much % of data still remains
(final['Id'].size*1.0)/(filtered_data['Id'].size*1.0)*100
# <b>Observation:-</b> It was also seen that in two rows given below the value of HelpfulnessNumerator is greater than HelpfulnessDenominator which is not practically possible hence these two rows too are removed from calcualtions
# +
display= pd.read_sql_query("""
SELECT *
FROM Reviews
WHERE Score != 3 AND Id=44737 OR Id=64422
ORDER BY ProductID
""", con)
display.head()
# -
final=final[final.HelpfulnessNumerator<=final.HelpfulnessDenominator]
# +
#Before starting the next phase of preprocessing lets see the number of entries left
print(final.shape)
#How many positive and negative reviews are present in our dataset?
final['Score'].value_counts()
# -
# # [3] Preprocessing
# ## [3.1]. Preprocessing Review Text
#
# Now that we have finished deduplication our data requires some preprocessing before we go on further with analysis and making the prediction model.
#
# Hence in the Preprocessing phase we do the following in the order below:-
#
# 1. Begin by removing the html tags
# 2. Remove any punctuations or limited set of special characters like , or . or # etc.
# 3. Check if the word is made up of english letters and is not alpha-numeric
# 4. Check to see if the length of the word is greater than 2 (as it was researched that there is no adjective in 2-letters)
# 5. Convert the word to lowercase
# 6. Remove Stopwords
# 7. Finally Snowball Stemming the word (it was obsereved to be better than Porter Stemming)<br>
#
# After which we collect the words used to describe positive and negative reviews
# +
# printing some random reviews
sent_0 = final['Text'].values[0]
print(sent_0)
print("="*50)
sent_1000 = final['Text'].values[1000]
print(sent_1000)
print("="*50)
sent_1500 = final['Text'].values[1500]
print(sent_1500)
print("="*50)
sent_4900 = final['Text'].values[4900]
print(sent_4900)
print("="*50)
# +
# remove urls from text python: https://stackoverflow.com/a/40823105/4084039
sent_0 = re.sub(r"http\S+", "", sent_0)
sent_1000 = re.sub(r"http\S+", "", sent_1000)
sent_150 = re.sub(r"http\S+", "", sent_1500)
sent_4900 = re.sub(r"http\S+", "", sent_4900)
print(sent_0)
# +
# https://stackoverflow.com/questions/16206380/python-beautifulsoup-how-to-remove-all-tags-from-an-element
from bs4 import BeautifulSoup
soup = BeautifulSoup(sent_0, 'lxml')
text = soup.get_text()
print(text)
print("="*50)
soup = BeautifulSoup(sent_1000, 'lxml')
text = soup.get_text()
print(text)
print("="*50)
soup = BeautifulSoup(sent_1500, 'lxml')
text = soup.get_text()
print(text)
print("="*50)
soup = BeautifulSoup(sent_4900, 'lxml')
text = soup.get_text()
print(text)
# +
# https://stackoverflow.com/a/47091490/4084039
import re
def decontracted(phrase):
# specific
phrase = re.sub(r"won't", "will not", phrase)
phrase = re.sub(r"can\'t", "can not", phrase)
# general
phrase = re.sub(r"n\'t", " not", phrase)
phrase = re.sub(r"\'re", " are", phrase)
phrase = re.sub(r"\'s", " is", phrase)
phrase = re.sub(r"\'d", " would", phrase)
phrase = re.sub(r"\'ll", " will", phrase)
phrase = re.sub(r"\'t", " not", phrase)
phrase = re.sub(r"\'ve", " have", phrase)
phrase = re.sub(r"\'m", " am", phrase)
return phrase
# -
sent_1500 = decontracted(sent_1500)
print(sent_1500)
print("="*50)
#remove words with numbers python: https://stackoverflow.com/a/18082370/4084039
sent_0 = re.sub("\S*\d\S*", "", sent_0).strip()
print(sent_0)
#remove spacial character: https://stackoverflow.com/a/5843547/4084039
sent_1500 = re.sub('[^A-Za-z0-9]+', ' ', sent_1500)
print(sent_1500)
# +
# https://gist.github.com/sebleier/554280
# we are removing the words from the stop words list: 'no', 'nor', 'not'
# <br /><br /> ==> after the above steps, we are getting "br br"
# we are including them into stop words list
# instead of <br /> if we have <br/> these tags would have revmoved in the 1st step
stopwords= set(['br', 'the', 'i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've",\
"you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', \
'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their',\
'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', \
'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', \
'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', \
'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after',\
'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further',\
'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more',\
'most', 'other', 'some', 'such', 'only', 'own', 'same', 'so', 'than', 'too', 'very', \
's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', \
've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn',\
"hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'mustn',\
"mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", \
'won', "won't", 'wouldn', "wouldn't"])
# -
# Combining all the above stundents
from tqdm import tqdm
preprocessed_reviews = []
# tqdm is for printing the status bar
for sentance in tqdm(final['Text'].values):
sentance = re.sub(r"http\S+", "", sentance)
sentance = BeautifulSoup(sentance, 'lxml').get_text()
sentance = decontracted(sentance)
sentance = re.sub("\S*\d\S*", "", sentance).strip()
sentance = re.sub('[^A-Za-z]+', ' ', sentance)
# https://gist.github.com/sebleier/554280
sentance = ' '.join(e.lower() for e in sentance.split() if e.lower() not in stopwords)
preprocessed_reviews.append(sentance.strip())
preprocessed_reviews[1500]
final['Cleaned_Text']=preprocessed_reviews
print(final.shape)
print(final.head())
# # [4] Featurization
# ## [4.1] BAG OF WORDS
# +
from sklearn.metrics import accuracy_score,confusion_matrix
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(final, final['Score'], test_size=0.25, random_state=0)
X_train, X_cv, y_train, y_cv = train_test_split(X_train, y_train, test_size=0.25)
print(X_train.shape, y_train.shape)
print(X_cv.shape, y_cv.shape)
print(X_test.shape, y_test.shape)
count_vect = CountVectorizer(min_df = 10) #in scikit-learn
final_X_train = count_vect.fit_transform(X_train['Cleaned_Text'].values)
final_X_test = count_vect.transform(X_test['Cleaned_Text'].values)
final_X_cv = count_vect.transform(X_cv['Cleaned_Text'].values)
# +
print("some feature names ", count_vect.get_feature_names()[:10])
print('='*50)
final_X_train = preprocessing.normalize(final_X_train)
final_X_test = preprocessing.normalize(final_X_test)
final_X_cv = preprocessing.normalize(final_X_cv)
print("the type of count vectorizer ",type(final_X_train))
print("the shape of out text BOW vectorizer ",final_X_train.get_shape())
print("the number of unique words ", final_X_train.get_shape()[1])
# +
from sklearn.metrics import roc_auc_score
from sklearn.naive_bayes import BernoulliNB,MultinomialNB
import matplotlib.pyplot as plt
from sklearn.model_selection import GridSearchCV
train_auc = []
cv_auc = []
alpha_range = [1e-7, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1, 1e1, 1e2, 1e3, 1e4, 1e5]
parameters = {'alpha' : [1e-7, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1, 1e1, 1e2, 1e3, 1e4, 1e5]}
nb = MultinomialNB()
clf = GridSearchCV(nb, parameters, cv=3, scoring='roc_auc')
clf.fit(final_X_train, y_train)
optimal_alpha = alpha_range[clf.best_index_]
print("Best Alpha value:", optimal_alpha)
train_auc= clf.cv_results_['mean_train_score']
train_auc_std= clf.cv_results_['std_train_score']
cv_auc = clf.cv_results_['mean_test_score']
cv_auc_std= clf.cv_results_['std_test_score']
plt.semilogx(alpha_range, train_auc, label='Train AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
plt.gca().fill_between(alpha_range,train_auc - train_auc_std,train_auc + train_auc_std,alpha=0.2,color='darkblue')
plt.semilogx(alpha_range, cv_auc, label='CV AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
plt.gca().fill_between(alpha_range,cv_auc - cv_auc_std,cv_auc + cv_auc_std,alpha=0.2,color='darkorange')
plt.legend()
plt.xlabel("hyperparameter: Alpha")
plt.ylabel("AUC")
plt.title("ERROR PLOTS")
plt.show()
# +
# top 10 features
import operator
nb = MultinomialNB(alpha=optimal_alpha).fit(final_X_train, y_train)
pos_imp_features = nb.feature_log_prob_[1,:]
neg_imp_features = nb.feature_log_prob_[0,:]
imp_features = {}
feature_names= count_vect.get_feature_names()
for i in range(len(feature_names)):
imp_features[feature_names[i]] = pos_imp_features[i]
names_diff_sorted = sorted(imp_features.items(), key = operator.itemgetter(1), reverse = True)
print("Postive top 10 important features are:")
for i in range(10):
print(names_diff_sorted[i])
for i in range(len(feature_names)):
imp_features[feature_names[i]] = neg_imp_features[i]
names_diff_sorted = sorted(imp_features.items(), key = operator.itemgetter(1), reverse = True)
print("\n\nNegative top 10 important features are:")
for i in range(10):
print(names_diff_sorted[i])
# -
def plot_confusion_matrix(test_y, predict_y):
C = confusion_matrix(test_y, predict_y)
print("Percentage of misclassified points ", (len(test_y)-np.trace(C))/len(test_y)*100)
# C = 9,9 matrix, each cell (i,j) represents number of points of class i are predicted class j
A =(((C.T)/(C.sum(axis=1))).T)
#divid each element of the confusion matrix with the sum of elements in that column
# C = [[1, 2],
# [3, 4]]
# C.T = [[1, 3],
# [2, 4]]
# C.sum(axis = 1) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =1) = [[3, 7]]
# ((C.T)/(C.sum(axis=1))) = [[1/3, 3/7]
# [2/3, 4/7]]
# ((C.T)/(C.sum(axis=1))).T = [[1/3, 2/3]
# [3/7, 4/7]]
# sum of row elements = 1
B =(C/C.sum(axis=0))
#divid each element of the confusion matrix with the sum of elements in that row
# C = [[1, 2],
# [3, 4]]
# C.sum(axis = 0) axis=0 corresonds to columns and axis=1 corresponds to rows in two diamensional array
# C.sum(axix =0) = [[4, 6]]
# (C/C.sum(axis=0)) = [[1/4, 2/6],
# [3/4, 4/6]]
labels = [0,1]
cmap=sns.light_palette("green")
# representing A in heatmap format
print("-"*50, "Confusion matrix", "-"*50)
plt.figure(figsize=(10,5))
sns.heatmap(C, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
print("-"*50, "Precision matrix", "-"*50)
plt.figure(figsize=(10,5))
sns.heatmap(B, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
print("Sum of columns in precision matrix",B.sum(axis=0))
# representing B in heatmap format
print("-"*50, "Recall matrix" , "-"*50)
plt.figure(figsize=(10,5))
sns.heatmap(A, annot=True, cmap=cmap, fmt=".3f", xticklabels=labels, yticklabels=labels)
plt.xlabel('Predicted Class')
plt.ylabel('Original Class')
plt.show()
print("Sum of rows in Recall matrix",A.sum(axis=1))
# +
from sklearn.metrics import roc_curve, auc
nb = MultinomialNB(alpha=optimal_alpha)
# fitting the model
nb.fit(final_X_train, y_train)
# predict the response
pred = nb.predict(final_X_test)
train_fpr, train_tpr, thresholds = roc_curve(y_train, nb.predict_proba(final_X_train)[:,1])
test_fpr, test_tpr, thresholds = roc_curve(y_test, nb.predict_proba(final_X_test)[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC = %0.5f" %(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC = %0.5f" %(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("hyperparameter: Alpha")
plt.ylabel("AUC")
plt.title("ERROR PLOTS")
plt.show()
# -
plot_confusion_matrix(y_train, nb.predict(final_X_train))
plot_confusion_matrix(y_test, nb.predict(final_X_test))
# ### Observations:
# 1. Used 100k datapoints in building this model.
# 2. with best alpha of 0.1, the model has an roc_auc score of ~96% and ~94% using train and test data respectively which gives confidence that the model is neither overfitting nor underfitting.
# 3. By looking at the above observations, the model is performing good for positive data points but NOT OK on predicting the negative reviews. May be if we can change the BOW using ngrams, the model may perform better.
# ## [4.2] TF-IDF
# +
tf_idf_vect = TfidfVectorizer(ngram_range=(1,2), min_df=10)
tfidf_X_train = tf_idf_vect.fit_transform(X_train['Cleaned_Text'].values)
tfidf_X_test = tf_idf_vect.transform(X_test['Cleaned_Text'].values)
tfidf_X_cv = tf_idf_vect.transform(X_cv['Cleaned_Text'].values)
print("some sample features(unique words in the corpus)",tf_idf_vect.get_feature_names()[0:10])
print('='*50)
tfidf_X_train = preprocessing.normalize(tfidf_X_train)
tfidf_X_test = preprocessing.normalize(tfidf_X_test)
tfidf_X_cv = preprocessing.normalize(tfidf_X_cv)
print("the type of count vectorizer ",type(tfidf_X_train))
print("the shape of out text TFIDF vectorizer ",tfidf_X_train.get_shape())
print("the number of unique words including both unigrams and bigrams ", tfidf_X_train.get_shape()[1])
# +
train_auc = []
cv_auc = []
alpha_range = [1e-7, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1, 1e1, 1e2, 1e3, 1e4, 1e5]
parameters = {'alpha' : [1e-7, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1, 1e1, 1e2, 1e3, 1e4, 1e5]}
nb = MultinomialNB()
clf = GridSearchCV(nb, parameters, cv=3, scoring='roc_auc')
clf.fit(tfidf_X_train, y_train)
optimal_alpha = alpha_range[clf.best_index_]
print("Best Alpha value:", optimal_alpha)
train_auc= clf.cv_results_['mean_train_score']
train_auc_std= clf.cv_results_['std_train_score']
cv_auc = clf.cv_results_['mean_test_score']
cv_auc_std= clf.cv_results_['std_test_score']
plt.semilogx(alpha_range, train_auc, label='Train AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
plt.gca().fill_between(alpha_range,train_auc - train_auc_std,train_auc + train_auc_std,alpha=0.2,color='darkblue')
plt.semilogx(alpha_range, cv_auc, label='CV AUC')
# this code is copied from here: https://stackoverflow.com/a/48803361/4084039
plt.gca().fill_between(alpha_range,cv_auc - cv_auc_std,cv_auc + cv_auc_std,alpha=0.2,color='darkorange')
plt.legend()
plt.xlabel("hyperparameter: Alpha")
plt.ylabel("AUC")
plt.title("ERROR PLOTS")
plt.show()
# +
nb = MultinomialNB(alpha=optimal_alpha).fit(tfidf_X_train, y_train)
pos_imp_features = nb.feature_log_prob_[1,:]
neg_imp_features = nb.feature_log_prob_[0,:]
imp_features = {}
feature_names= tf_idf_vect.get_feature_names()
for i in range(len(feature_names)):
imp_features[feature_names[i]] = pos_imp_features[i]
names_diff_sorted = sorted(imp_features.items(), key = operator.itemgetter(1), reverse = True)
print("Postive top 10 important features are:")
for i in range(10):
print(names_diff_sorted[i])
for i in range(len(feature_names)):
imp_features[feature_names[i]] = neg_imp_features[i]
names_diff_sorted = sorted(imp_features.items(), key = operator.itemgetter(1), reverse = True)
print("\n\nNegative top 10 important features are:")
for i in range(10):
print(names_diff_sorted[i])
# +
from sklearn.metrics import roc_curve, auc
nb = MultinomialNB(alpha=optimal_alpha)
# fitting the model
nb.fit(tfidf_X_train, y_train)
# predict the response
pred = nb.predict(tfidf_X_test)
train_fpr, train_tpr, thresholds = roc_curve(y_train, nb.predict_proba(tfidf_X_train)[:,1])
test_fpr, test_tpr, thresholds = roc_curve(y_test, nb.predict_proba(tfidf_X_test)[:,1])
plt.plot(train_fpr, train_tpr, label="train AUC = %0.5f" %(auc(train_fpr, train_tpr)))
plt.plot(test_fpr, test_tpr, label="test AUC = %0.5f" %(auc(test_fpr, test_tpr)))
plt.legend()
plt.xlabel("hyperparameter: Alpha")
plt.ylabel("AUC")
plt.title("ERROR PLOTS")
plt.show()
# -
plot_confusion_matrix(y_train, nb.predict(tfidf_X_train))
plot_confusion_matrix(y_test, nb.predict(tfidf_X_test))
# # Observations
# 1. We have used 100k datapoints only to build the model
# 2. with best alpha of 0.1, the model has roc_auc of ~98% and ~95% using train and test data respectively which gives confidence that the model is neither overfitting nor underfitting.
# 3. The AUC for the tf-idf is ~0.96 which is very good compared to BOW (unigram) with AUC of ~0.94. In this case, tf-idf performed better than BOW (unigram) but still the tf-idf also needs to perform better on negative reviews as well.
# 4. If we can train the model with ngrams, the model may perform better.
# +
# Pretty print
from prettytable import PrettyTable
x = PrettyTable()
x.field_names = ["Vectorizer", "Model", "Hyper Parameter", "AUC"]
x.add_row(["BOW", 'MNB', 0.1 , 0.94])
x.add_row(["TF-IDF", 'MNB', 0.1 , 0.96])
print(x)
# -
|
Amazon-fine-food-reviews/04 Amazon Fine Food Reviews Analysis_NaiveBayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 03: NYC 311 Service Request Analysis
#
# You don't need to limit yourself to the number of rows/cells provided. You can add additional rows in each section to add more lines of code.
#
# **Happy coding!**
#import required libraries
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# %matplotlib inline
#get the data
sr_df = pd.read_csv('311_Service_Requests_from_2010_to_Present.csv')
#view first five records
sr_df.head()
#view the info
sr_df.info()
#check the number of observations
sr_df.shape
#describe
sr_df.describe()
#view the columns
sr_df.columns
#plot bar graph for complaint type breakdown
sr_df['Complaint Type'].value_counts().plot(kind='bar',alpha=0.6,figsize=(12,15))
plt.title('Complaint Type Breakdown')
plt.xlabel('Complaint Type')
plt.ylabel('Number of Complaints')
plt.show()
#count the city with highest complaints
sr_df['City'].value_counts()
#plot the bar graph for complaint by city
sr_df['City'].value_counts().plot(kind='bar',alpha=0.6,figsize=(15,15))
plt.title('Complaint By City')
plt.xlabel('City')
plt.ylabel('Number of Complaints')
plt.show()
#display the complaint type, city together
sr_df[['Complaint Type','City']].head(10)
#group data by complaint type
group_ComplaintType=sr_df.groupby('Complaint Type')
TopComplaintType=group_ComplaintType.get_group('Blocked Driveway')
TopComplaintType.shape
#get Nan values from dataset
sr_df.isnull().sum()
#remove blank values in the city column
sr_df['City'].dropna(inplace=True)
sr_df['City'].shape
sr_df['City'].head()
#count null city values in group for top complaint type
TopComplaintType['City'].isnull().sum()
#Fill those null city values with 'unknown city'
TopComplaintType['City'].fillna('Unknown City',inplace=True)
# plot scatter diagram to show all cities raised complaint type ='Blocked Driveway'
plt.figure(figsize=(15,15))
plt.scatter(TopComplaintType['Complaint Type'],TopComplaintType['City'])
plt.title('List of cities raised complaint type Blocked Driveway')
plt.xlabel('Complaint Type')
plt.ylabel('Cities')
plt.show()
#find the top 10 major complaint type and the counts
group_ComplaintType['Complaint Type'].value_counts().nlargest(10)
#fill null Location Type with 'Unknown Location'
sr_df['Location Type'].fillna('Unknown Location',inplace=True)
sr_df['Location Type'].values
#count null values in Top Complaint group
TopComplaintType['Location Type'].isnull().sum()
TopComplaintType['Location Type'].fillna('Unknown Location',inplace=True)
#Heat map for complaint type against location type for any pattern
plt.figure(figsize=(5,5))
plt.scatter(TopComplaintType['Complaint Type'],TopComplaintType['Location Type'])
plt.title('complaint type against location type')
#plt.xlabel('Complaint Type')
#plt.ylabel('Cities')
plt.show()
|
Project 3- NYC 311 Service Request Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# You can put here Jupyter Notebook files and explanatory materials (remember about naming convention divided with '-', for e.g. 'reproducible-neuroimaging.ipynb').
|
notebooks/jupyter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 1 - Python program to create Bankaccount class with deposit, withdraw function.
# +
class Bank_Account:
def __init__(self):
self.balance = 0
print("Hello, Your account is created.")
def deposit(self):
amount = int(input('Enter the amount to deposit: '))
self.balance += amount
print('Your New Balance = %d' %self.balance)
def withdraw(self):
amount = int(input('Enter the amount to withdraw:'))
if (amount > self.balance):
print('Insufficient Balance!!')
else:
self.balance -= amount
print('Your Remaining Balance = %d', self.balance)
def enquiry(self):
print('Your Balance = %d', self.balance)
account = Bank_Account()
# -
account.deposit()
account.withdraw()
# # Assignment 2 - Python program to calculate volume and surface area of cone
# +
import math
pi = math.pi
# Function to calculate Volume of Cone
def volume(r, h):
return (1 / 3) * pi * r * r * h
# Function To Calculate Surface Area of Cone
def surfacearea(r, s):
return pi * r * s + pi * r * r
radius = float(5)
height = float(12)
slat_height = float(13)
print( "Volume Of Cone : ", volume(radius, height) )
print( "Surface Area Of Cone : ", surfacearea(radius, slat_height) )
|
Day 6 Assignments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p38
# language: python
# name: p38
# ---
# # 07.04 - TENSORFLOW
# !wget --no-cache -O init.py -q https://raw.githubusercontent.com/rramosp/ai4eng.v1/main/content/init.py
import init; init.init(force_download=False); init.get_weblink()
# +
import numpy as np
import matplotlib.pyplot as plt
from local.lib import mlutils
from IPython.display import Image
try:
# %tensorflow_version 2.x
print ("Using TF2 in Google Colab")
except:
pass
import tensorflow as tf
# %matplotlib inline
# -
# ## A dataset (again)
## KEEPOUTPUT
from sklearn.datasets import make_moons
X,y = make_moons(300, noise=.15)
plt.scatter(X[:,0][y==0], X[:,1][y==0], color="blue", label="class 0", alpha=.5)
plt.scatter(X[:,0][y==1], X[:,1][y==1], color="red", label="class 1", alpha=.5)
# ## A neural network in tensorflow, 4 neurons in the hidden layer, 1 output
model = tf.keras.Sequential([
tf.keras.layers.Dense(4, activation='tanh'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer=tf.keras.optimizers.SGD(learning_rate=.5),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=False),metrics=['accuracy'])
## KEEPOUTPUT
model.fit(X,y, epochs=10, batch_size=1)
## KEEPOUTPUT
predict = lambda X: (model.predict(X)[:,0]>.5).astype(int)
mlutils.plot_2Ddata_with_boundary(predict, X, y)
# ## A bigger network
#
# - different activation functions
# - different optimizer
model = tf.keras.Sequential([
tf.keras.layers.Dense(20, activation='tanh'),
tf.keras.layers.Dense(50, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=.01),
loss=tf.keras.losses.BinaryCrossentropy(from_logits=False),metrics=['accuracy'])
## KEEPOUTPUT
model.fit(X,y, epochs=10, batch_size=1)
## KEEPOUTPUT
predict = lambda X: (model.predict(X)[:,0]>.5).astype(int)
mlutils.plot_2Ddata_with_boundary(predict, X, y)
# ## Cross entropy - multiclass classification
#
# follow [THIS EXAMPLE](https://www.tensorflow.org/tutorials/keras/classification) in TensorFlow doc site. Observe that:
#
# - labels corresponding to a 10-class classification problem
# - the network contains 10 output neurons, one per output class
# - the loss function is `SparseCategoricalCrossEntropy`
#
# Observe how **cross entropy** works with 4 classes:
#
# - first we convert the output to a one-hot encoding
# - we create a network with two output neurons with sigmoid activation
# - interpret each neuron's output as elements of a probability distribution
# - normalize the probability distribution (must add up to one)
# - we consider network output is better when it yields more probability to the correct class
# **expected classes for five data points**
## KEEPOUTPUT
y = np.random.randint(4, size=5)
y
# **convert it to one hot encoding**
## KEEPOUTPUT
y_ohe = np.eye(4)[y].astype(int)
y_ohe
# **simulate some neural network output with NO ACTIVATION function**
#
# with 10 output neurons, so for each input element (we have five) we have 4 outputs.
#
# this is called **LOGITS** in Tensorflow
## KEEPOUTPUT
y_hat = np.round(np.random.normal(size=y_ohe.shape), 2)
y_hat
# **normalize LOGITS**. This is the **SOFTMAX function**
#
# **LOGITS** obtained from network last layer with no activation
#
# $$\hat{\mathbf{y}}^{(i)} = [\hat{y}^{(i)}_0, \hat{y}^{(i)}_1,...,\hat{y}^{(i)}_9]$$
#
# **SOFTMAX ACTIVATION**
#
# $$\hat{\bar{\mathbf{y}}}^{(i)} = [\hat{\bar{y}}^{(i)}_0, \hat{\bar{y}}^{(i)}_1,...,\hat{\bar{y}}^{(i)}_9]$$
#
# with
#
# $$\hat{\bar{y}}^{(i)}_k = \frac{e^{\hat{y}^{(i)}_k}}{\sum_{j=0}^9e^{\hat{y}^{(i)}_j}}$$
#
#
# this ensures:
#
# - $\sum_{k=0}^9 \hat{\bar{y}}^{(i)}_k=1$
# - $0 \le \hat{\bar{y}}^{(i)}_k \le 1$
#
# this way, for each input we have a nice probability distribution in its outputs.
#
# This is implemented in **Tensorflow**
## KEEPOUTPUT
y_hatb = tf.nn.softmax(y_hat).numpy()
y_hatb
# check sums
## KEEPOUTPUT
y_hatb.sum(axis=1)
# how would you now measure how closely `y_hatb` is to the expected output on `y_ohe`?
#
# **cross entropy**: just take the probability assigned to the correct class (and pass it through a log function)
#
# $$\text{loss}(\bar{\mathbf{y}}^{(i)}, \hat{\bar{\mathbf{y}}}^{(i)}) = -\sum_{k=0}^9 \bar{y}^{(i)}_k\log(\hat{\bar{y}}^{(i)}_k)$$
#
# where $\bar{\mathbf{y}}^{(i)}$ is the one-hot encoding of the expected class (label) for data point $i$.
#
# observe that,
#
# - in the one-hot encoding $\bar{\mathbf{y}}^{(i)}$ only one of the elements will be 1, and the rest will be 0's, so the sumation above is only taking the log of the probability of the correct label.
# - the negative sign accounts for logs of values <1 are negative and we will later want to **minimize** the loss
#
# This is implemented in **Tensorflow**
## KEEPOUTPUT
tf.keras.losses.categorical_crossentropy(y_ohe, y_hatb)
# Observe that TensorFlow also implements the corresponding **sparse** convenience function that works directly with our labels
## KEEPOUTPUT
tf.keras.losses.sparse_categorical_crossentropy(y, y_hatb)
|
content/NOTES 07.04 - NEURAL NETWORKS WITH TENSORFLOW.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Clustering Techniques Writeup
# ### September 26, 2016
# ### K-Means Clustering
#
# K-Means clustering is a regression technique which involves 'fitting' a number $n$ of given values from a dataset around a pre-defined number of $k$ clusters. The K-means clustering process seeks to minimize the Euclidean distance from each point $n$ from its associated $k$ cluster. Note that although the $k$ clusters inhabit the same coordinate space as the $n$ data points, the $k$ cluster locations are not necessarily derived from the set of $n$ points.
#
# The process of K-means involves first defining the number of $k$ clusters. Defining the number of $k$ means is a non-trivial problem that is dependent on the specific experiment at hand. Analysis with an elbow plot showing explained variance as a function of number of clusers allows the designer to designate the $k$ number by choosing the point where adding additional $k$ clusters has diminishing returns for explained variance.
#
#
# The actual algorithm behind K-means clustering is relatively simple. Given $n$ data points and $k$ clusters, we're looking for µ<sub>k</sub> (cluster means that minimize the Euclidean distance between the $n$ data points and said cluster). Effectively, the objective function we're trying to minimize can be written as (courtesy SciKit):
#
# $$
# \begin{align}
# \sum\limits_{i=0}^n \min_{µ_j \subseteq C} ||(x_j - µ_i)||^2
# \end{align}
# $$
#
# For the analysis, we must first choose $k$ starting mean µ's. This is often achieved by randomly sampling $k$ points from the dataset arbitrarily. Below, we have an image of the data set at the start, with arbitrarily placed means. *(Image sequence courtesy <NAME>, from Project Rhea)*
#
# 
#
# After choosing the initial set of starting means, we can apply a three step iterative process to eventually 'converge' on finalized means:
#
# 1) **Fit n points to µ means:** Calculate the Euclidean distances and assign each $n$ to its closest µ. In this case, all points are closest to the blue node, so all $n$ are assigned to the blue µ.
#
# 
#
# 2) **Define new µ's by finding average:** Find the center of each cluster and assign those as new µ's. Only the blue µ moves, as the other nodes have no values assigned to them.
#
# 
#
# 3) **Repeat until convergence:** Repeat until there is no change in the µ positions. I've shown the final converged image below.
#
# 
#
# **PROS:**
# 1. Oft-implemented, existing documentation.
# 2. Has SciKit implementation in Python.
#
# **CONS:**
# 1. Defining number of means is a non-trivial process.
# 2. Does not necessarily converge to global solution, as randomized starting positions will alter final result for different results.
# 3. Does not set the means to positions to some value from the dataset (if we're interested in brain clusters, we want our clusters to be centered around some set of pre-defined nodes, not some weighted average determined by k-means.)
# 4. Assumes similarly sized 'clusters', which is a very large assumption to be making for brain areas.
# ### K-Medoid Clustering
#
# K-Medoids clustering is similar to K-means in that both start with a known, pre-defined number of 'k' clusters. However, K-medoids sets the 'k' clusters to points that occur naturally in the dataset of 'n' points. Thus, while K-means finds new 'means' and fits each 'n' point to these new means by reducing some summed error, K-medoids instead seeks to maximize some measure of similarity (eg: the similarity matrix) between each points and its medoid (which itself is one of the pre-existing points).
#
# The process for K-Medoids is similar to K-Means; again, the problem of defining the number of 'k' means is a difficult process. From there, an additional difficulty lies in how to define the 'similarity' between two points. A way to do so would be to make a similarity matrix that is has each corresponding value as the inverse of the Euclidean distance between the points. Aside from this, a similar process would be applied iteratively to converge to medoids that maximize the similarity between their respective nodes.
#
# The specific name of the algorithmic approach is Partitioning Around Medoids, or PAM. The steps are, specifically:
# 1) Arbitrarily select 'k' of the 'n' nodes as the medoids.
# 2) Calculate the total 'similarity' (eg, by finding the inverse of the total of the distances between all 'n' and their closest 'k', or by using some other measure).
# 3) Swap one 'n' with one 'k', and recalculate the 'similarity' measure. If the 'similarity' increased, keep the new configuration and continue. Otherwise, return to the previous configuration.
#
# **PROS:**
# 1. Simple, like K-means.
#
# **CONS:**
# 1. Has similiar faults as the K-means methodology (defining 'k', not necessarily global, assumes equal sized regions)
# 2. Recalculating similarity at each step relative to all other points is very computationally intensive
# ### Spectral Clustering
# Up until now, we've focused on clustering techniques centered around compactness. Spectral clustering is a different dataset fitting technique that focuses on connectivity instead.
#
#
# The actual algorithm behind K-means clustering is relatively simple. Given 'n' data points and 'k' clusters, we're looking for µk (cluster means that minimize the Euclidean distance between the 'n' data points and said cluster).
#
# **Pros:**
# 1. abc
# 2. abc
#
# **Cons:**
# 1. abc
# 2. abc
# ### Louvain Method of Community Detection
# The Louvain method of community detection is a clustering technique that relies on monitoring the relative connectiveness of communities.
#
# From Blondel, Guillaume, Lambiotte and Lefebvre's seminal paper that defined the Louvain method, they defined first a modularity value. From the paper, "the modularity of a partition is a scalar value between -1 and 1 that measures the density of links inside communities as compared to links between communities." In the case of weighted nodes (where for instance they used the number of calls between between two phone users), the modularity is defined as:
#
# $$
# \begin{align}
# Q = \frac{1}{2m} \sum\limits_{i, \ j} \big[ A_{i, \ j} - \frac{k_i k_j}{2m} \big] \ \delta \big( c_i , c_j \big)
# \end{align}
# $$
#
# In this case, the $A_{i, \ j}$ refers to the weight of the edge between $i$ and $j$. From there, they defined $k_i$ as the $\sum_{j} A_{i, \ j}$ as the "weights of the edges attached to vertex $i$". $c_i$ is the 'community' to which node $i$ is assigned, while the $\delta$-function $\delta \big( u, v \big)$ is 1 if $u$ is $v$, and 0 otherwise. $m$ is defined to be $\frac{1}{2}\sum_{i, \ j} A_{i, \ j}$ In lay speech, then, what this calculates is 'how connected' some node $i$ is relative some node $j$ (since we're subtracting from the edge weight of $ij$ the edge weights of all connections from $i$ and $j$).
#
# From there, the paper progresses to explain how this traditional definition of modularity takes a significant amount of storage power to process. Data analytics on the original data using continuous modularity calls take more processing power than is truly necessary. What Blondel, Guillaume, Lambiotte, and Lefebvre did was define an algorithmically (and computationally) more efficient method to measure the *change* in modularity. It does so by calculating a $\Delta Q$ value that is the *change* in modularity by moving (inserting) some node $i$ from an 'isolated neighborhood' into $C$:
#
# $$
# \begin{align}
# \Delta Q = \bigg[ \frac{\sum_{in} + \ k_{i, \ in}}{2m} - \big( \frac{\sum_{tot} + \ k_i}{2m} \big)^2 \bigg] - \bigg[ \frac{\sum_{in}}{2m} - \big( \frac{\sum_{tot}}{2m} \big)^2 - \big( \frac{k_i}{2m} \big)^2 \bigg]
# \end{align}
# $$
#
# A similar equation is provided for the removal of a node $i$ from the neighborhood $C$.
#
# Application-wise, the process for Louvain's method is similar to the approach for all the other regression techniques (K-means, K-medoids). The approach first assigns and puts each node $N$ into a unique community (so it's in its own community). They then calculate the $\Delta Q$ by adding the node $i$ into community $j$; if after all similar additions into other neighborhoods, the $\Delta Q$ is maximized by adding i to j, that process is fulfilled. Otherwise, the system is reverted to before the addition.
#
# By applying this until a local minimum occurs (when no changes to $i$ increase or decrease $\Delta Q$, the Louvain method can create a very solid mapping.
#
# **Pros:**
# 1. Computationally efficient, their calculations in the paper show a significant reduction in computation time (running on 6.3M nodes, their algorithm only took 197 seconds. The only comparable algorithm that even converged was the Wakita and Tsumuri's CNN implementation method, which took almost three times as long).
# 2. Accuracy seems to be high, according to their own discussion and results.
#
# **Cons:**
# 1. Requires weighted edges (we haven't generated these yet). If our weights are defined by the number of connections within the epsilon ball, what's the point of using weighted edges over K-means or K-medoids? Isn't that just an extra layer of complexity?
# 2. Requires a C-based workaround that allows us to run the methods in Python (so I won't be able to use these lovely Markdown/Python notebooks). Not too big a problem, but it's nice when packets like SciKit already has implementations of the other methods. Link: http://perso.crans.org/aynaud/communities/
#
# Link to Louvain's Method paper: http://arxiv.org/abs/0803.0476
#
# + language="latex"
# \begin{align}
# \nabla \times \vec{\mathbf{B}} -\, \frac1c\, \frac{\partial\vec{\mathbf{E}}}{\partial t} & = \frac{4\pi}{c}\vec{\mathbf{j}} \\
# \nabla \cdot \vec{\mathbf{E}} & = 4 \pi \rho \\
# \nabla \times \vec{\mathbf{E}}\, +\, \frac1c\, \frac{\partial\vec{\mathbf{B}}}{\partial t} & = \vec{\mathbf{0}} \\
# \nabla \cdot \vec{\mathbf{B}} & = 0
# \end{align}
# -
|
examples/Jupyter/.ipynb_checkpoints/Clustering Techniques-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CDC NDI Mortality - Syft Duet - Data Owner 🎸
# This worksheet is intended to illustrate functionality of a shared statistical platform, using a partially synthetic public-use dataset that mirrors the restricted-use dataset. Ultimately, these processes would apply to the restricted-use data.
#
# Sample data compiled from the public-use linked mortality files share at https://www.cdc.gov/nchs/data-linkage/mortality.htm provided by the National Center for Health Statistics (NCHS).
# ## PART 1: Launch a Duet Server and Connect
#
# As a Data Owner, you want to allow someone else to perform data science on data that you own and likely want to protect.
#
# In order to do this, we must load our data into a locally running server within this notebook. We call this server a "Duet".
#
# To begin, you must launch Duet and help your Duet "partner" (a Data Scientist) connect to this server.
#
# You do this by running the code below and sending the code snippet containing your unique Server ID to your partner and following the instructions it gives!
# +
import syft as sy
duet = sy.launch_duet(loopback=True)
# -
sy.load("pandas")
sy.load("statsmodels")
sy.load("numpy")
duet.requests.add_handler(action="accept")
# +
import zipfile
import requests
import io
from urllib.request import urlopen
csv_file = "mort_match_nhis_all_years.csv"
zip_file = f"{csv_file}.zip"
url = f"https://raw.githubusercontent.com/madhavajay/datasets/main/{zip_file}"
res = requests.get(url)
z = zipfile.ZipFile(io.BytesIO(res.content))
with z.open(z.namelist()[0], 'r') as myfile:
binaryCSV = myfile.read()
# +
import pandas as pd
df = pd.read_csv(io.BytesIO(binaryCSV), encoding='utf-8')
df[df.MORTSTAT == 0].head()
# -
len(df)
df_ptr = df.send(duet, tags=["df"])
df
# +
# local stats
# Select the records that died by cancer that were eligible for linkage
# 002-Malignant neoplasms (C00-C97)
cancer = df[(df.UCOD_LEADING == 2) & (df.ELIGSTAT == 1)]
# Select the records that died due to heart disease and were eligible for linkage
# 001-Diseases of heart (I00-I09, I11, I13, I20-I51)
heart = df[(df.UCOD_LEADING == 1) & (df.ELIGSTAT == 1)]
# -
# Compute simple means and for the cancer and heart subgroups that had diabetes
# listed as a multiple cause of death
cancer["DIABETES"].mean()
# Compute simple means and standard deviations for the cancer and heart subgroups
# that had diabetes as a multiple cause of death
heart["DIABETES"].mean()
# +
# Sample means data should account for weights. Write a custom function that uses the weights.
def weighted_mean(dx, key, weight_key="WGT_NEW"):
w = dx[weight_key]
v = dx[key]
return (w * v).sum() / w.sum()
weighted_mean(cancer, "DIABETES"), weighted_mean(heart, "DIABETES")
# +
# Example of a small subgroup (sample size = 6)
# Cancer-deaths from males aged 47 who died in 2015
# We should check for small cell sizes here
subgroup = cancer[(cancer.SEX == 1) & (cancer.AGE_P == 47) & (cancer.DODYEAR == 2015)]
print(subgroup["DIABETES"].mean())
print(weighted_mean(subgroup, "DIABETES"))
print(len(subgroup))
# These stats are problematic, as the subgroup is too small to report (n=6)
subgroup
# +
import statsmodels.api as sm
from statsmodels.genmod.generalized_linear_model import GLM
from statsmodels.genmod.families import Binomial
# Drop any missing values in the dataset (those under 18)
df = df.dropna(subset=["MORTSTAT"])
# Keep only the eligible portion
df = df[df.ELIGSTAT == 1]
# Ignore people > 80
df = df[df.AGE_P <= 80]
# A person is alive if MORTSTAT==0
df["is_alive"] = df.MORTSTAT == 0
# Assign a helpful column for sex (0==male, 1==female)
df["sex"] = "male"
df.loc[df.SEX == 2, "sex"] = "female"
x = df["AGE_P"]
_x = sm.add_constant(x)
_y = df["is_alive"]
results = GLM(_y, _x, family=Binomial()).fit()
print(results.summary())
# -
predict_x = range(x.min(), x.max() + 1, 1)
preds = results.predict(sm.add_constant(predict_x))
# share predictions
preds.send(duet, tags=["preds"])
try:
import pylab as plt
import seaborn as sns
plt.figure(figsize=(12, 5))
plt.plot(predict_x, preds, "k", lw=3, label="Best Fit for all data")
sns.lineplot(data=df, x="AGE_P", y="is_alive", hue="sex", err_style="bars")
sns.despine()
except ImportError:
print("Cant import seaborn try:\n!pip install seaborn")
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 1 : Well done!
|
packages/syft/examples/data-science/pandas/CDC_NDI_Mortality_Syft_Data_Owner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# ### This is taking two versions of Webservices' lists in .csv format and yielding their difference
#
# <u><b>Assumption:</b></u>
#
# The code assumes that the services' list will be in the following format->
# <table border = 1>
# <th>Parent Service</th>
# <th>API Name</th>
# <tr>
# <td>Root Service Category 1</td>
# <td>Service 1</td>
# </tr>
# </table>
import pandas as pd
# Assign the two Services' lists to two dataframes
df2 = pd.read_csv("Version2.csv")
df1 = pd.read_csv("Version1.csv")
df2.head()
df1.head()
d1 = df1.drop(['Parent Service'], axis = 1)
d2 = df2.drop(['Parent Service'], axis = 1)
# +
# Compare the two dataframes by the following method:
# Concat the two dataframes
# Reset the indices of the rows accordingly
# Group the same rows
# Get the indices of unique records
# Filter them
df = pd.concat([d1, d2])
df = df.reset_index(drop=True)
df_gpby = df.groupby(list(df.columns))
idx = [x[0] for x in df_gpby.groups.values() if len(x) == 1]
df_new = df.reindex(idx)
df_new.head()
# -
# Save the dataframe to an Excel sheet
writer = pd.ExcelWriter('C:\\Users\\Sayak\\Desktop\\output.xlsx')
df_new.to_excel(writer,'Sheet1')
writer.save()
|
Comparing+two+dataframes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.017247, "end_time": "2021-11-20T14:47:44.230791", "exception": false, "start_time": "2021-11-20T14:47:44.213544", "status": "completed"} tags=[]
# **This notebook is an exercise in the [Intermediate Machine Learning](https://www.kaggle.com/learn/intermediate-machine-learning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/introduction).**
#
# ---
#
# + [markdown] papermill={"duration": 0.014452, "end_time": "2021-11-20T14:47:44.260674", "exception": false, "start_time": "2021-11-20T14:47:44.246222", "status": "completed"} tags=[]
# As a warm-up, you'll review some machine learning fundamentals and submit your initial results to a Kaggle competition.
#
# # Setup
#
# The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
# + papermill={"duration": 1.270213, "end_time": "2021-11-20T14:47:45.548207", "exception": false, "start_time": "2021-11-20T14:47:44.277994", "status": "completed"} tags=[]
# Set up code checking
import os
if not os.path.exists("../input/train.csv"):
os.symlink("../input/home-data-for-ml-course/train.csv", "../input/train.csv")
os.symlink("../input/home-data-for-ml-course/test.csv", "../input/test.csv")
from learntools.core import binder
binder.bind(globals())
from learntools.ml_intermediate.ex1 import *
print("Setup Complete")
# + [markdown] papermill={"duration": 0.015309, "end_time": "2021-11-20T14:47:45.579201", "exception": false, "start_time": "2021-11-20T14:47:45.563892", "status": "completed"} tags=[]
# You will work with data from the [Housing Prices Competition for Kaggle Learn Users](https://www.kaggle.com/c/home-data-for-ml-course) to predict home prices in Iowa using 79 explanatory variables describing (almost) every aspect of the homes.
#
# 
#
# Run the next code cell without changes to load the training and validation features in `X_train` and `X_valid`, along with the prediction targets in `y_train` and `y_valid`. The test features are loaded in `X_test`. (_If you need to review **features** and **prediction targets**, please check out [this short tutorial](https://www.kaggle.com/dansbecker/your-first-machine-learning-model). To read about model **validation**, look [here](https://www.kaggle.com/dansbecker/model-validation). Alternatively, if you'd prefer to look through a full course to review all of these topics, start [here](https://www.kaggle.com/learn/machine-learning).)_
# + papermill={"duration": 0.283, "end_time": "2021-11-20T14:47:45.877822", "exception": false, "start_time": "2021-11-20T14:47:45.594822", "status": "completed"} tags=[]
import pandas as pd
from sklearn.model_selection import train_test_split
# Read the data
X_full = pd.read_csv('../input/train.csv', index_col='Id')
X_test_full = pd.read_csv('../input/test.csv', index_col='Id')
# Obtain target and predictors
y = X_full.SalePrice
features = ['LotArea', 'YearBuilt', '1stFlrSF', '2ndFlrSF', 'FullBath', 'BedroomAbvGr', 'TotRmsAbvGrd']
X = X_full[features].copy()
X_test = X_test_full[features].copy()
# Break off validation set from training data
X_train, X_valid, y_train, y_valid = train_test_split(X, y, train_size=0.8, test_size=0.2,
random_state=0)
# + [markdown] papermill={"duration": 0.014846, "end_time": "2021-11-20T14:47:45.908219", "exception": false, "start_time": "2021-11-20T14:47:45.893373", "status": "completed"} tags=[]
# Use the next cell to print the first several rows of the data. It's a nice way to get an overview of the data you will use in your price prediction model.
# + papermill={"duration": 0.038154, "end_time": "2021-11-20T14:47:45.961590", "exception": false, "start_time": "2021-11-20T14:47:45.923436", "status": "completed"} tags=[]
X_train.head()
# + [markdown] papermill={"duration": 0.015445, "end_time": "2021-11-20T14:47:45.994978", "exception": false, "start_time": "2021-11-20T14:47:45.979533", "status": "completed"} tags=[]
# The next code cell defines five different random forest models. Run this code cell without changes. (_To review **random forests**, look [here](https://www.kaggle.com/dansbecker/random-forests)._)
# + papermill={"duration": 0.026699, "end_time": "2021-11-20T14:47:46.037517", "exception": false, "start_time": "2021-11-20T14:47:46.010818", "status": "completed"} tags=[]
from sklearn.ensemble import RandomForestRegressor
# Define the models
model_1 = RandomForestRegressor(n_estimators=50, random_state=0)
model_2 = RandomForestRegressor(n_estimators=100, random_state=0)
model_3 = RandomForestRegressor(n_estimators=100, criterion='mae', random_state=0)
model_4 = RandomForestRegressor(n_estimators=200, min_samples_split=20, random_state=0)
model_5 = RandomForestRegressor(n_estimators=100, max_depth=7, random_state=0)
models = [model_1, model_2, model_3, model_4, model_5]
# + [markdown] papermill={"duration": 0.015699, "end_time": "2021-11-20T14:47:46.069327", "exception": false, "start_time": "2021-11-20T14:47:46.053628", "status": "completed"} tags=[]
# To select the best model out of the five, we define a function `score_model()` below. This function returns the mean absolute error (MAE) from the validation set. Recall that the best model will obtain the lowest MAE. (_To review **mean absolute error**, look [here](https://www.kaggle.com/dansbecker/model-validation).)_
#
# Run the code cell without changes.
# + papermill={"duration": 5.334584, "end_time": "2021-11-20T14:47:51.420206", "exception": false, "start_time": "2021-11-20T14:47:46.085622", "status": "completed"} tags=[]
from sklearn.metrics import mean_absolute_error
# Function for comparing different models
def score_model(model, X_t=X_train, X_v=X_valid, y_t=y_train, y_v=y_valid):
model.fit(X_t, y_t)
preds = model.predict(X_v)
return mean_absolute_error(y_v, preds)
for i in range(0, len(models)):
mae = score_model(models[i])
print("Model %d MAE: %d" % (i+1, mae))
# + [markdown] papermill={"duration": 0.017109, "end_time": "2021-11-20T14:47:51.455661", "exception": false, "start_time": "2021-11-20T14:47:51.438552", "status": "completed"} tags=[]
# # Step 1: Evaluate several models
#
# Use the above results to fill in the line below. Which model is the best model? Your answer should be one of `model_1`, `model_2`, `model_3`, `model_4`, or `model_5`.
# + papermill={"duration": 0.02999, "end_time": "2021-11-20T14:47:51.503140", "exception": false, "start_time": "2021-11-20T14:47:51.473150", "status": "completed"} tags=[]
# Fill in the best model
best_model = model_3
# Check your answer
step_1.check()
# + papermill={"duration": 0.025613, "end_time": "2021-11-20T14:47:51.547193", "exception": false, "start_time": "2021-11-20T14:47:51.521580", "status": "completed"} tags=[]
# Lines below will give you a hint or solution code
#step_1.hint()
#step_1.solution()
# + [markdown] papermill={"duration": 0.018144, "end_time": "2021-11-20T14:47:51.583960", "exception": false, "start_time": "2021-11-20T14:47:51.565816", "status": "completed"} tags=[]
# # Step 2: Generate test predictions
#
# Great. You know how to evaluate what makes an accurate model. Now it's time to go through the modeling process and make predictions. In the line below, create a Random Forest model with the variable name `my_model`.
# + papermill={"duration": 0.030659, "end_time": "2021-11-20T14:47:51.632850", "exception": false, "start_time": "2021-11-20T14:47:51.602191", "status": "completed"} tags=[]
# Define a model
my_model = model_3 # Your code here
# Check your answer
step_2.check()
# + papermill={"duration": 0.026953, "end_time": "2021-11-20T14:47:51.679446", "exception": false, "start_time": "2021-11-20T14:47:51.652493", "status": "completed"} tags=[]
# Lines below will give you a hint or solution code
#step_2.hint()
#step_2.solution()
# + [markdown] papermill={"duration": 0.019048, "end_time": "2021-11-20T14:47:51.718090", "exception": false, "start_time": "2021-11-20T14:47:51.699042", "status": "completed"} tags=[]
# Run the next code cell without changes. The code fits the model to the training and validation data, and then generates test predictions that are saved to a CSV file. These test predictions can be submitted directly to the competition!
# + papermill={"duration": 4.611844, "end_time": "2021-11-20T14:47:56.349378", "exception": false, "start_time": "2021-11-20T14:47:51.737534", "status": "completed"} tags=[]
# Fit the model to the training data
my_model.fit(X, y)
# Generate test predictions
preds_test = my_model.predict(X_test)
# Save predictions in format used for competition scoring
output = pd.DataFrame({'Id': X_test.index,
'SalePrice': preds_test})
output.to_csv('submission.csv', index=False)
# + [markdown] papermill={"duration": 0.019441, "end_time": "2021-11-20T14:47:56.390849", "exception": false, "start_time": "2021-11-20T14:47:56.371408", "status": "completed"} tags=[]
# # Submit your results
#
# Once you have successfully completed Step 2, you're ready to submit your results to the leaderboard! First, you'll need to join the competition if you haven't already. So open a new window by clicking on [this link](https://www.kaggle.com/c/home-data-for-ml-course). Then click on the **Join Competition** button. _(If you see a "Submit Predictions" button instead of a "Join Competition" button, you have already joined the competition, and don't need to do so again.)_
#
# Next, follow the instructions below:
# 1. Begin by clicking on the **Save Version** button in the top right corner of the window. This will generate a pop-up window.
# 2. Ensure that the **Save and Run All** option is selected, and then click on the **Save** button.
# 3. This generates a window in the bottom left corner of the notebook. After it has finished running, click on the number to the right of the **Save Version** button. This pulls up a list of versions on the right of the screen. Click on the ellipsis **(...)** to the right of the most recent version, and select **Open in Viewer**. This brings you into view mode of the same page. You will need to scroll down to get back to these instructions.
# 4. Click on the **Output** tab on the right of the screen. Then, click on the file you would like to submit, and click on the **Submit** button to submit your results to the leaderboard.
#
# You have now successfully submitted to the competition!
#
# If you want to keep working to improve your performance, select the **Edit** button in the top right of the screen. Then you can change your code and repeat the process. There's a lot of room to improve, and you will climb up the leaderboard as you work.
#
# + [markdown] papermill={"duration": 0.01933, "end_time": "2021-11-20T14:47:56.430074", "exception": false, "start_time": "2021-11-20T14:47:56.410744", "status": "completed"} tags=[]
# # Keep going
#
# You've made your first model. But how can you quickly make it better?
#
# Learn how to improve your competition results by incorporating columns with **[missing values](https://www.kaggle.com/alexisbcook/missing-values)**.
# + [markdown] papermill={"duration": 0.020436, "end_time": "2021-11-20T14:47:56.470169", "exception": false, "start_time": "2021-11-20T14:47:56.449733", "status": "completed"} tags=[]
# ---
#
#
#
#
# *Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/intermediate-machine-learning/discussion) to chat with other learners.*
|
course/Feature Engineering/exercise-introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mengwangk/trading-playground/blob/master/notebook/pytorch/transfer_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="4DWAnogqX5nh" colab_type="code" colab={}
from __future__ import print_function, division
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import numpy as np
import torchvision
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import time
import os
import copy
plt.ion() # interactive mode
# + [markdown] id="_Xcgx8KZX5nm" colab_type="text"
# ## Load Data
# + id="nn-cw_34X5nn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="916d6654-4115-499a-8283-8175522029b9"
import os
import zipfile
from six.moves import urllib
import zipfile
DATASET_URL = "https://download.pytorch.org/tutorial/hymenoptera_data.zip"
DATASET_PATH = os.path.join(os.getcwd(), "data", "")
DATASET_FILE = os.path.join(DATASET_PATH, "hymenoptera_data.zip")
def fetch_data(url=DATASET_URL, path=DATASET_PATH, zip_path=DATASET_FILE) -> bool:
if not os.path.isdir(path):
os.makedirs(path)
urllib.request.urlretrieve(url, zip_path)
if (os.path.isfile(zip_path)):
print(f'Download path: {zip_path}')
return True
else:
return False
def extract_zip(zip_file=DATASET_FILE, unzip_folder=DATASET_PATH):
with zipfile.ZipFile(zip_file, 'r') as zip_ref:
zip_ref.extractall(unzip_folder)
if fetch_data():
extract_zip()
else:
print('Error downloading')
# + id="O3q5TenTX5ns" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ed1a5710-6e59-4dac-e0a1-159ccada4dfd"
# Data augmentation and normalization for training
# Just normalization for validation
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
}
data_dir = 'data/hymenoptera_data'
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x),
data_transforms[x])
for x in ['train', 'val']}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x], batch_size=4,
shuffle=True, num_workers=4)
for x in ['train', 'val']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val']}
class_names = image_datasets['train'].classes
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(class_names)
# + [markdown] id="jZZELcS4X5nv" colab_type="text"
# ## Visualize a few images
# + id="lLTXdwHHX5nw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 175} outputId="b55d6969-06f2-4985-b65a-bedf57fc2ecc"
def imshow(inp, title=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
# Get a batch of training data
inputs, classes = next(iter(dataloaders['train']))
# Make a grid from batch
out = torchvision.utils.make_grid(inputs)
imshow(out, title=[class_names[x] for x in classes])
# + [markdown] id="6GGr0pS-X5nz" colab_type="text"
# ## Train the model
# + id="-Q6-gjalX5n0" colab_type="code" colab={}
def train_model(model, criterion, optimizer, scheduler, num_epochs=25):
since = time.time()
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
scheduler.step()
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
epoch_loss = running_loss / dataset_sizes[phase]
epoch_acc = running_corrects.double() / dataset_sizes[phase]
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, epoch_loss, epoch_acc))
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(
time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
return model
# + [markdown] id="QLCxrtUZX5n2" colab_type="text"
# ## Visualize the model
# + id="sGTQLFXOauaE" colab_type="code" colab={}
def visualize_model(model, num_images=6):
was_training = model.training
model.eval()
images_so_far = 0
fig = plt.figure()
with torch.no_grad():
for i, (inputs, labels) in enumerate(dataloaders['val']):
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
for j in range(inputs.size()[0]):
images_so_far += 1
ax = plt.subplot(num_images//2, 2, images_so_far)
ax.axis('off')
ax.set_title('predicted: {}'.format(class_names[preds[j]]))
imshow(inputs.cpu().data[j])
if images_so_far == num_images:
model.train(mode=was_training)
return
model.train(mode=was_training)
# + [markdown] id="3hqAtO6yavTK" colab_type="text"
# ## Fine tuning the convnet
# + id="GEHJ5UQcazJt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="1c6d3310-0c27-4959-a4db-b35fcf05acb7"
model_ft = models.resnet18(pretrained=True)
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs, 2)
model_ft = model_ft.to(device)
criterion = nn.CrossEntropyLoss()
# Observe that all parameters are being optimized
optimizer_ft = optim.SGD(model_ft.parameters(), lr=0.001, momentum=0.9)
# Decay LR by a factor of 0.1 every 7 epochs
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_ft, step_size=7, gamma=0.1)
# + [markdown] id="Ds8sAdjRa6o8" colab_type="text"
# ## Train and evaluate
# + id="q4FZp8VMa_EW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2176} outputId="a41f4a9c-ed98-4d64-e9e8-ce831dfa68de"
model_ft = train_model(model_ft, criterion, optimizer_ft, exp_lr_scheduler,
num_epochs=25)
# + id="RG3GTQ9Fa_bu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 821} outputId="515c35f1-49e8-4891-b2ea-a2a389746692"
visualize_model(model_ft)
# + [markdown] id="TKvj5nfQb4xx" colab_type="text"
# ## ConvNet as fixed feature extractor
# + id="cCBPb7uRcAYt" colab_type="code" colab={}
model_conv = torchvision.models.resnet18(pretrained=True)
for param in model_conv.parameters():
param.requires_grad = False
# Parameters of newly constructed modules have requires_grad=True by default
num_ftrs = model_conv.fc.in_features
model_conv.fc = nn.Linear(num_ftrs, 2)
model_conv = model_conv.to(device)
criterion = nn.CrossEntropyLoss()
# Observe that only parameters of final layer are being optimized as
# opposed to before.
optimizer_conv = optim.SGD(model_conv.fc.parameters(), lr=0.001, momentum=0.9)
# Decay LR by a factor of 0.1 every 7 epochs
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_conv, step_size=7, gamma=0.1)
# + [markdown] id="21-FrkyNcIiV" colab_type="text"
# ## Train and evaluate
# + id="NqTx6qcDcMWg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2176} outputId="7b1e6b97-75cf-48c6-d490-83f2848d8fde"
model_conv = train_model(model_conv, criterion, optimizer_conv,
exp_lr_scheduler, num_epochs=25)
# + id="qf6frsKFcMs_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 821} outputId="4e493656-4421-4dc2-de35-3f093fa09b47"
visualize_model(model_conv)
plt.ioff()
plt.show()
# + id="1WlbI2PedT9D" colab_type="code" colab={}
|
notebook/pytorch/transfer_learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Moneyball Project: UEFA Euro 2020 Fantasy Football
# Passion project to leverage data-driven decision making for team selection in [UEFA Euro 2020 Fantasy Football](https://gaming.uefa.com/en/uefaeuro2020fantasyfootball/overview)
#
# ## Data Preparation and Cleansing
# -----------------------------
# ### Purpose
# Initial exploration on available dataset, aggregating and merging to dataframe for further exploration.
#
# ### Author
# [<NAME>](https://github.com/christianwbsn)
#
#
# ## 1. Import Library
# +
import pandas as pd
import numpy as np
import seaborn as sns
import json
import re
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_row',50)
# -
import difflib
from tqdm import tqdm
from nltk import everygrams
DATA_DIR = "../data"
# ## 2. Common Function
def camel_to_snake(name):
name = re.sub(" ", "", name)
name = re.sub('(.)([A-Z][a-z]+)', r'\1_\2', name)
return re.sub('([a-z0-9])([A-Z])', r'\1_\2', name).lower()
def extract_date(date):
return pd.Series([date.year, date.month, date.day])
def euro_fantasy_score(df):
# Not covered by dataset
# Common - Goal from outside the box 2 points
# Common - Winning a penalty 2 points
# Common - Conceding a penalty -1 points
# Common - Own Goal -2 points
# common
score = 1
if df["min"] >= 60:
score += 1
if df["assists"] > 0:
score += (df["assists"] * 3)
if df["penalty_kick_miss"] > 0:
score -= (df["penalty_kick_miss"] * 2)
if df["yellow_cards"] > 0:
score -= 1
if df["red_cards"] > 0:
score -= 3
# position specific
if df["position"] == "F":
score += (df["goals"] * 4)
if df["position"] == "M":
score += (df["goals"] * 5)
if df["min"] >= 60 and df["clean_sheet"] > 0:
score += 1
if df["position"] == "D":
score += (df["goals"] * 6)
if df["min"] >= 60 and df["clean_sheet"] > 0:
score += 4
score -= (df['goals_allowed'] // 2)
if df["position"] == "GK":
score += (df["goals"] * 6)
score += (df["penalty_kick_saved"] * 5)
if df["min"] >= 60 and df["clean_sheet"] > 0:
score += 4
score += (df["saves"] // 3)
score -= (df["goals_allowed"] // 2)
return score
# ## 3. Dataset Exploration
# ### 3.1 Main Dataset
## Using this dataset as SSOT for player name and team name
main_df = pd.read_csv("{}/interim/md_4_df.csv".format(DATA_DIR))
main_df["date"] = pd.to_datetime(main_df["date"])
main_df["match"] = main_df["team_name"] + ',' + main_df['opponent_name']
main_df["match"] = main_df["match"].apply(lambda x: ' '.join(sorted(x.split(","))))
main_df["match"] = main_df["match"] + '_' + main_df["date"].dt.strftime('%Y-%m-%d')
max_points = main_df.groupby("match").max()["points"].to_dict()
main_df["is_star_of_match"] = main_df.apply(lambda x: 1 if x["points"] == max_points[x["match"]] else 0, axis=1)
# ### 3.2 Euro 2020 Dataset
# #### 3.2.1 Players
# #### 3.2.1.1 Appending last matchday data
with open('{}/raw/euro-2020/players_4.json'.format(DATA_DIR))as f:
data = json.load(f)
players = data["data"]["value"]["playerList"]
old_players_df = pd.json_normalize(players)
old_players_df.rename(camel_to_snake, axis=1, inplace=True)
with open('{}/raw/euro-2020/players_5.json'.format(DATA_DIR))as f:
data = json.load(f)
players = data["data"]["value"]["playerList"]
players_df = pd.json_normalize(players)
players_df.rename(camel_to_snake, axis=1, inplace=True)
players_df = players_df[players_df["trained"]!='']
players_df = pd.merge(players_df, old_players_df[["p_f_name", "g_s", "assist", "y_c", "r_c", "p_m"]],
on="p_f_name", suffixes=("", "_last_md"))
players_df["g_s"] = players_df["g_s"] - players_df["g_s_last_md"]
players_df["assist"] = players_df["assist"] - players_df["assist_last_md"]
players_df["y_c"] = players_df["y_c"] - players_df["y_c_last_md"]
players_df["r_c"] = players_df["r_c"] - players_df["r_c_last_md"]
players_df["p_m"] = players_df["p_m"] - players_df["p_m_last_md"]
players_df = players_df.drop(["g_s_last_md", "assist_last_md", "y_c_last_md", "r_c_last_md", "p_m_last_md"], axis=1)
players_df["date"] = players_df["current_matches_list"].apply(lambda x: x[0]["matchDate"])
players_df["opponent_name"] = players_df["current_matches_list"].apply(lambda x: x[0]["vsTSCode"])
all_players_name = main_df["player"].unique()
def get_closest_match(name):
# return closest match for join operation
return ''.join(list(difflib.get_close_matches(name, all_players_name, n=1, cutoff=0.7)))
players_df["closest_match"] = players_df["p_f_name"].apply(get_closest_match)
players_df["player"] = players_df.apply(lambda x: x["closest_match"] if x["closest_match"] != "" else x["p_f_name"], axis=1)
players_df["date"] = pd.to_datetime(players_df["date"])
players_df[["year", "month", "day"]] = players_df["date"].apply(extract_date)
main_df.head()
players_df.head()
with open('{}/raw/euro-2020/fixtures.json'.format(DATA_DIR))as f:
data = json.load(f)
fixtures = data["data"]["value"][2]["match"]
fixtures_df = pd.json_normalize(fixtures)
fixtures_df["atName"] = fixtures_df["atName"].apply(lambda x: x.strip())
fixtures_df["htName"] = fixtures_df["htName"].apply(lambda x: x.strip())
def heuristic_minutes_played(df):
if (df["g_s"] == 0) and (df["assist"] == 0) and (df["y_c"] == 0) and (df["r_c"] == 0) and (df["last_gd_points"] == 2):
return 90
elif df["last_gd_points"] >= 2:
return 90
elif df["last_gd_points"] == 0:
return 0
else:
return 59
players_df["min"] = players_df.apply(heuristic_minutes_played, axis=1)
players_df.rename(columns={"t_name": "team_name", "g_s": "goals", "assist": "assists",
"y_c": "yellow_cards", "r_c" : "red_cards", "last_gd_points": "points",
"p_m": "penalty_kick_miss"}, inplace=True)
players_df["league_name"] = "European Championship 2020"
fixtures_df.head()
fixtures_df[fixtures_df["htName"] == "Belgium"]["htScore"]
def goals_allowed(df):
if df["team_name"] in fixtures_df["htName"].values:
return int(fixtures_df[fixtures_df["htName"] == df["team_name"]].reset_index()["atScore"])
else:
return int(fixtures_df[fixtures_df["atName"] == df["team_name"]].reset_index()["htScore"])
players_df["goals_allowed"] = players_df.apply(goals_allowed, axis=1)
players_df["clean_sheet"] = players_df["goals_allowed"].apply(lambda x: 1 if x == 0 else 0)
players_df["game_started"] = players_df["min"].apply(lambda x: 1 if x >= 60 else 0)
players_df.shape
# if players have multiple position choose the most common position
position = main_df.groupby("player").agg(position=('position', "first")).to_dict()["position"]
players_df["position"] = players_df["player"].apply(lambda x: position[x]
if x in position.keys() else "")
players_df.shape
players_df
def update_data(df):
main_df_columns = ["player", "date", "league_name", "game_started",
"team_name", "opponent_name", "position", "goals_allowed", "clean_sheet",
"year", "month", "day", "min", "goals", "assists",
"penalty_kick_miss","yellow_cards", "red_cards", "saves", "points"]
return df[main_df_columns]
players_df = players_df[~pd.isnull(players_df["date"])]
new_train = update_data(players_df)
new_train["match"] = new_train["team_name"] + ',' + new_train['opponent_name']
new_train["match"] = new_train["match"].apply(lambda x: ' '.join(sorted(x.split(","))))
new_train["match"] = new_train["match"] + '_' + new_train["date"].dt.strftime('%Y-%m-%d')
max_points = new_train.groupby("match").max()["points"].to_dict()
max_points
new_train["is_star_of_match"] = new_train.apply(lambda x: 1 if x["points"] == max_points[x["match"]] else 0, axis=1)
main_df = pd.concat([main_df, new_train])
main_df = main_df.drop(["match"], axis=1)
main_df = main_df.fillna(0)
main_df.to_csv("{}/interim/md_5_df.csv".format(DATA_DIR), index=False)
# #### 3.2.1.2 Generating test data
with open('{}/raw/euro-2020/players_6.json'.format(DATA_DIR))as f:
data = json.load(f)
players = data["data"]["value"]["playerList"]
players_df = pd.json_normalize(players)
players_df.rename(camel_to_snake, axis=1, inplace=True)
players_df = players_df[players_df["trained"]!='']
players_df.head()
players_df["date"] = players_df["upcoming_matches_list"].apply(lambda x: x[0]["matchDate"])
players_df["opponent_name"] = players_df["upcoming_matches_list"].apply(lambda x: x[0]["vsTSCode"])
players_df = players_df[~pd.isnull(players_df["opponent_name"])]
all_players_name = main_df["player"].unique()
def get_closest_match(name):
# return closest match for join operation
return ''.join(list(difflib.get_close_matches(name, all_players_name, n=1, cutoff=0.7)))
players_df["closest_match"] = players_df["p_f_name"].apply(get_closest_match)
players_df["player"] = players_df.apply(lambda x: x["closest_match"] if x["closest_match"] != "" else x["p_f_name"], axis=1)
players_df["date"] = pd.to_datetime(players_df["date"])
players_df[["year", "month", "day"]] = players_df["date"].apply(extract_date)
main_df.head()
players_df.head()
players_df.rename(columns={"t_name": "team_name"}, inplace=True)
players_df["league_name"] = "European Championship 2020"
players_df.shape
# if players have multiple position choose the most common position
position = main_df.groupby("player").agg(position=('position',
lambda x: x.value_counts().sort_index().sort_values(ascending=False).index[0])).to_dict()["position"]
players_df["position"] = players_df["player"].apply(lambda x: position[x]
if x in position.keys() else "")
players_df.shape
def generate_test_data(df):
main_df_columns = ["player", "date", "league_name",
"team_name", "opponent_name", "position",
"year", "month", "day"]
return df[main_df_columns]
test = generate_test_data(players_df)
main_df = pd.concat([main_df, test])
main_df = pd.merge(main_df, players_df[["player", "value", "skill"]], on=["player"], how="left")
def get_agg_before(df):
merged_df = df.copy()
merged_df = pd.merge(merged_df, df, on=["player", "team_name"])
merged_df = merged_df[merged_df['date_y'] < merged_df["date_x"]]
merged_df["is_scoring"] = merged_df["goals_y"].apply(lambda x: 1 if x > 0 else 0)
merged_df["is_assisting"] = merged_df["assists_y"].apply(lambda x: 1 if x > 0 else 0)
merged_df["significant_min_play"] = merged_df["min_y"].apply(lambda x: 1 if x > 45 else 0)
merged_df_1 = merged_df.groupby(["player", "team_name", "date_x"]).agg(
prev_mean_points=("points_y", "mean"),
prev_mean_goals=("goals_y", "median"),
prev_mean_assists=("assists_y", "mean"),
prev_max_points=("points_y", "max"),
prev_std_points=("points_y", "std"),
prev_std_goals=("goals_y", "std"),
prev_std_assists=("assists_y", "std"),
prev_median_min=("min_y", "median"),
prev_ratio_starter=("game_started_y", "mean"),
count_played=("date_y","nunique"),
goal_consistency=("is_scoring", "mean"),
assist_consistency=("is_assisting", "mean"),
clean_sheet_consistency=("clean_sheet_y", "mean"),
star_of_match_consistency=("is_star_of_match_y", "mean"),
star_of_match_count=("is_star_of_match_y", "sum")
)
merged_df_1 = merged_df_1.reset_index()
merged_df_1.rename(columns={"date_x": "date"}, inplace=True)
merged_df_2 = merged_df.groupby(["team_name", "date_x"]).agg(count_team_played=("date_y", "nunique"))
merged_df_2 = merged_df_2.reset_index()
merged_df_2.rename(columns={"date_x": "date"}, inplace=True)
merged_df_3 = merged_df[merged_df["opponent_name_x"] == merged_df["opponent_name_y"]]
merged_df_3 = merged_df_3.groupby(["player", "team_name", "date_x"]).agg(prev_max_goal_to_specific_opp=("goals_y", "max"),
prev_max_points_to_specific_opp=("points_y", "max"),
prev_mean_points_to_specific_opp=("points_y", "mean"))
merged_df_3 = merged_df_3.reset_index()
merged_df_3.rename(columns={"date_x": "date", "opponent_name_y": "opponent_name"}, inplace=True)
merged_df_4 = merged_df.sort_values(by=["date_y"], ascending=False)
merged_df_4 = merged_df_4.groupby(["player", "team_name", "date_x"]).head(3)
merged_df_4 = merged_df_4.groupby(["player", "team_name", "date_x"]).agg(last_three_match_mean_points=("points_y", "mean"),
last_three_match_std_points=("points_y", "std"),
last_three_match_max_points=("points_y", "max"),
last_three_match_significant_min_play=("significant_min_play", "sum"),
last_three_match_goal_rate=("goals_y", "mean"))
merged_df_4 = merged_df_4.reset_index()
merged_df_4.rename(columns={"date_x": "date"}, inplace=True)
merged_df = pd.merge(merged_df_1, merged_df_2, on=["team_name", "date"], how="left")
merged_df = pd.merge(merged_df, merged_df_3, on=["player", "team_name", "date"], how="left")
merged_df = pd.merge(merged_df, merged_df_4, on=["player", "team_name", "date"], how="left")
merged_df["prev_ratio_played"] = merged_df["count_played"] / merged_df["count_team_played"]
return merged_df
agg = get_agg_before(main_df)
main_df.head()
main_df = main_df.sort_values(["player", "date"])
main_df['last_md_points'] = main_df.groupby("player")["points"].shift()
main_df['last_md_goals'] = main_df.groupby("player")["goals"].shift()
main_df['last_md_assists'] = main_df.groupby("player")["assists"].shift()
main_df = main_df.drop(["goals", "assists", "shots", "shots_on_goal", "crosses", "fouls_drawn",
"fouls_committed", "tackles_won", "interceptions", "yellow_cards", "red_cards",
"penalty_kick_miss", "clean_sheet", "goals_allowed", "accurate_passes",
"shots_assisted", "shootout_goals", "shootout_misses", "game_started", "saves", "wins",
"penalty_kick_saved", "shootout_saves"], axis=1)
main_df = pd.merge(main_df, agg, how="left", on=["player", "team_name", "date"])
main_df.columns
players_df.to_csv("{}/interim/fantasy_euro.csv".format(DATA_DIR), index=False)
main_df
# ## 3.3 National Team FIFA Rank Dataset
main_df["date"].describe()
fifa_rank = pd.read_csv("{}/raw/historical-match-and-rank/fifa_ranking-2021-05-27.csv".format(DATA_DIR))
CUTOFF_DATE = "2018-01-01"
fifa_rank = fifa_rank[fifa_rank["rank_date"] > CUTOFF_DATE]
fifa_rank = fifa_rank[["country_full", "rank", "total_points", "rank_date"]]
fifa_rank["rank_date"] = pd.to_datetime(fifa_rank["rank_date"])
fifa_rank = fifa_rank.sort_values(by=["country_full", "rank_date"])
# get fifa rank closest to the match date
df_with_rank = pd.merge(main_df[["team_name", "date"]], fifa_rank, how="left", left_on="team_name", right_on="country_full")
df_with_rank["time_diff"] = df_with_rank.apply(lambda x: (x['date']-x['rank_date']).total_seconds(), axis=1)
df_with_rank = df_with_rank[df_with_rank["time_diff"] > 0] # filter out rank after match
df_with_rank = df_with_rank.sort_values(by=["team_name", "time_diff"], ascending=False)
df_with_rank = df_with_rank.groupby(["team_name", "date"]).agg(prev_team_highest_rank=("rank", "min"),
team_rank=("rank", "last"),
team_total_points=("total_points", "last")).reset_index()
main_df = pd.merge(main_df, df_with_rank, how="left", on=["team_name", "date"])
# get fifa rank closest to the match date
df_with_rank = pd.merge(main_df[["opponent_name", "date"]], fifa_rank, how="left", left_on="opponent_name", right_on="country_full")
df_with_rank["time_diff"] = df_with_rank.apply(lambda x: (x['date']-x['rank_date']).total_seconds(), axis=1)
df_with_rank = df_with_rank[df_with_rank["time_diff"] > 0] # filter out rank after match
df_with_rank = df_with_rank.sort_values(by=["opponent_name", "time_diff"], ascending=False)
df_with_rank = df_with_rank.groupby(["opponent_name", "date"]).agg(prev_opponent_highest_rank=("rank", "min"),
opponent_rank=("rank", "last"),
opponent_total_points=("total_points", "last")).reset_index()
main_df = pd.merge(main_df, df_with_rank, how="left", on=["opponent_name", "date"])
main_df.head()
main_df.to_csv("{}/interim/main.csv".format(DATA_DIR), index=False)
historical_matches = pd.read_csv("{}/raw/historical-match-and-rank/international-footbal-match.csv".format(DATA_DIR))
historical_matches["date"] = pd.to_datetime(historical_matches["date"])
historical_matches = historical_matches[historical_matches["date"] > "2010-01-01"]
historical_matches["match"] = historical_matches["home_team"] + ',' + historical_matches['away_team']
historical_matches["match"] = historical_matches["match"].apply(lambda x: ' '.join(sorted(x.split(","))))
def get_match_result(df):
if df["home_score"] > df["away_score"]:
return df["home_team"]
elif df["away_score"] > df["home_score"]:
return df["away_team"]
else:
return "Draw"
historical_matches["result"] = historical_matches.apply(get_match_result, axis=1)
historical_matches["margin"] = historical_matches.apply(lambda x: abs(x["home_score"] - x["away_score"]), axis=1)
historical_matches.head()
def get_all_historical_matches(df, team, opp, date):
name_tuple = ' '.join(sorted([team, opp]))
hist = df[(df['match'] == name_tuple) & (df["date"] < date)]
hth = hist["result"].value_counts()
team_win, opp_win, draw = 0, 0, 0
if "Draw" in hth.keys():
draw = hth["Draw"]
if team in hth.keys():
team_win = hth[team]
if opp in hth.keys():
opp_win = hth[opp]
max_margin = hist["margin"].max()
team_score = hist[hist['home_team'] == team]["home_score"].sum() + hist[hist['away_team'] == team]["away_score"].sum()
opp_score = hist[hist['home_team'] == opp]["home_score"].sum() + hist[hist['away_team'] == opp]["away_score"].sum()
return pd.Series([team_win, opp_win, draw, team_score, opp_score, max_margin])
main_df[["hth_team_win", "hth_opp_win", "hth_draw", "hth_team_score", "hth_opp_score", "htt_max_margin"]] = main_df.apply(lambda x: get_all_historical_matches(historical_matches, x["team_name"], x["opponent_name"], x["date"]), axis=1)
# ## 3.4 Transfermarkt Dataset
# ### 3.4.1 National Team Level
euro = pd.read_excel("{}/raw/transfermarkt/transfermarkt-market-value.xlsx".format(DATA_DIR), sheet_name=0)
nations_league = pd.read_excel("{}/raw/transfermarkt/transfermarkt-market-value.xlsx".format(DATA_DIR), sheet_name=1)
euro_qual = pd.read_excel("{}/raw/transfermarkt/transfermarkt-market-value.xlsx".format(DATA_DIR), sheet_name=2)
wc_euro_qual = pd.read_excel("{}/raw/transfermarkt/transfermarkt-market-value.xlsx".format(DATA_DIR), sheet_name=3)
nations_league["league_name"] = "UEFA Nations League"
euro_qual["league_name"] = "European Championship Qualifiers"
wc_euro_qual["league_name"] = "European World Cup Qualifiers"
euro["league_name"] = "European Championship 2020"
euro = euro.drop(["EURO participations"], axis=1)
euro.rename(columns={"Average Age": "Age"}, inplace=True)
def preprocess_market_value(text):
match = re.sub("€", "", text)
match = re.search("(\d+(?:\.\d+)?)", text)
val = float(match.group())
num = text[match.end():]
if num == "bn":
val *= 10e9
elif num == "m":
val *= 10e6
elif num == "Th.":
val *= 10e3
return val
mv_df = pd.concat([nations_league, euro_qual, wc_euro_qual, euro])
mv_df["market_value"] = mv_df["Market Value"].apply(preprocess_market_value)
mv_df["mean_market_value"] = mv_df["Average Market Value"].apply(preprocess_market_value)
mv_df = mv_df.drop_duplicates(subset=["Club", "league_name"], keep="first")
mv_df = mv_df[["Club", "league_name", "Age", "market_value", "mean_market_value"]]
mv_df.rename(columns={"Club" : "team_name", "Age": "mean_squad_age"}, inplace=True)
main_df = pd.merge(main_df, mv_df, how="left", on=["team_name", "league_name"])
main_df.rename(columns={"mean_squad_age" : "team_mean_squad_age",
"mean_market_value": "team_mean_market_value",
"market_value" : "team_market_value"
}, inplace=True)
mv_df.rename(columns={"team_name" : "opponent_name"}, inplace=True)
main_df = pd.merge(main_df, mv_df, how="left", on=["opponent_name", "league_name"])
main_df.rename(columns={"mean_squad_age" : "opponent_mean_squad_age",
"mean_market_value": "opponent_mean_market_value",
"market_value" : "opponent_market_value"
}, inplace=True)
# ## 3.6 FIFA Dataset
fifa_21 = pd.read_csv("{}/raw/fifa/fifa-players_21.csv".format(DATA_DIR))
fifa_20 = pd.read_csv("{}/raw/fifa/players_20.csv".format(DATA_DIR))
fifa_19 = pd.read_csv("{}/raw/fifa/players_19.csv".format(DATA_DIR))
fifa_18 = pd.read_csv("{}/raw/fifa/players_18.csv".format(DATA_DIR))
fifa_21["nationality"] = fifa_21["nationality"].apply(lambda x: x.strip())
fifa_20["nationality"] = fifa_20["nationality"].apply(lambda x: x.strip())
fifa_19["nationality"] = fifa_19["nationality"].apply(lambda x: x.strip())
fifa_18["nationality"] = fifa_18["nationality"].apply(lambda x: x.strip())
fifa_21 = fifa_21[fifa_21['nationality'].isin(main_df['team_name'].unique())]
fifa_20 = fifa_20[fifa_20['nationality'].isin(main_df['team_name'].unique())]
fifa_19 = fifa_19[fifa_19['nationality'].isin(main_df['team_name'].unique())]
fifa_18 = fifa_18[fifa_18['nationality'].isin(main_df['team_name'].unique())]
# +
fifa_21['len_name'] = fifa_21["long_name"].apply(lambda x: len(x.split(" ")))
fifa_21['len_short_name'] = fifa_21["short_name"].apply(lambda x: len(x.split(" ")))
fifa_21['min_char_in_name'] = fifa_21['long_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_21['min_char_in_short_name'] = fifa_21['short_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_20['len_name'] = fifa_20["long_name"].apply(lambda x: len(x.split(" ")))
fifa_20['len_short_name'] = fifa_20["short_name"].apply(lambda x: len(x.split(" ")))
fifa_20['min_char_in_name'] = fifa_20['long_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_20['min_char_in_short_name'] = fifa_20['short_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_19['len_name'] = fifa_19["long_name"].apply(lambda x: len(x.split(" ")))
fifa_19['len_short_name'] = fifa_19["short_name"].apply(lambda x: len(x.split(" ")))
fifa_19['min_char_in_name'] = fifa_19['long_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_19['min_char_in_short_name'] = fifa_19['short_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_18['len_name'] = fifa_18["long_name"].apply(lambda x: len(x.split(" ")))
fifa_18['len_short_name'] = fifa_18["short_name"].apply(lambda x: len(x.split(" ")))
fifa_18['min_char_in_name'] = fifa_18['long_name'].apply(lambda x: min(len(y) for y in x.split()))
fifa_18['min_char_in_short_name'] = fifa_18['short_name'].apply(lambda x: min(len(y) for y in x.split()))
# +
def join_tuple_string(strings_tuple):
return ' '.join(strings_tuple)
def create_unigram_bigram_trigram_quadgram(text, x):
token_list = text.split(" ")
tuple_gram = list(everygrams(token_list, 2, x))
result = map(join_tuple_string, tuple_gram)
if x > 2:
return list(result) + [' '.join(token_list[::len(token_list)-1])]
return list(result)
def calculate_closest_token(df):
everygram = create_unigram_bigram_trigram_quadgram(df["long_name"], df['len_name'])
closest = difflib.get_close_matches(df["short_name"], everygram, n=1)
return ''.join(closest)
# -
fifa_21['closest_match'] = fifa_21.apply(calculate_closest_token, axis=1)
fifa_20['closest_match'] = fifa_20.apply(calculate_closest_token, axis=1)
fifa_19['closest_match'] = fifa_19.apply(calculate_closest_token, axis=1)
fifa_18['closest_match'] = fifa_18.apply(calculate_closest_token, axis=1)
name_mapping = {
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'D.Rice': "<NAME>",
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Azpilicueta': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'van de Beek': '<NAME>',
'<NAME>': '<NAME>',
'Duje Ćaleta-Car': 'D<NAME>-Car',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'H<NAME>hanoglu':'H<NAME>',
'<NAME>': '<NAME>',
'İ<NAME>an': 'Il<NAME>ündogan',
'<NAME>ci': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Palhinha': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Oyarzabal': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Ok<NAME>okuşlu': 'Ok<NAME>okuslu',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>ökçü': 'Orkun Kökcü',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Šime Vrsaljko': 'Sime Vrsaljko',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': 'Ugurcan Çakir',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'Yusuf Yazıcı': 'Yusuf Yazici',
'Çağlar Söyüncü': 'Çaglar Söyüncü',
'<NAME>': "<NAME>",
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>änen': '<NAME>änen',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': "<NAME>",
'<NAME>': "<NAME>",
"<NAME>": "<NAME>",
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'João <NAME>': '<NAME>',
'<NAME>':'<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>' : '<NAME>',
'Thiago':'<NAME>',
'<NAME>' : "<NAME>",
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
'<NAME>': '<NAME>',
"Morata":'<NAME>',
"<NAME>" :"<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>": "<NAME>",
"<NAME>":"<NAME>",
"<NAME>": "<NAME>"
}
def heuristic_match(df):
if df["len_short_name"] == 1:
return df["short_name"]
if len(df["closest_match"].split()) == 0:
return df["short_name"]
elif df["min_char_in_short_name"] >= 4:
return df["short_name"]
elif df["len_name"] > 3:
return df["closest_match"]
elif df["min_char_in_name"] >= 3:
return df["closest_match"]
else:
return df["long_name"]
fifa_21["player"] = fifa_21.apply(heuristic_match, axis=1)
fifa_20["player"] = fifa_20.apply(heuristic_match, axis=1)
fifa_19["player"] = fifa_19.apply(heuristic_match, axis=1)
fifa_18["player"] = fifa_18.apply(heuristic_match, axis=1)
def map_name(name):
global name_mapping
if name in name_mapping.keys():
return name_mapping[name]
else:
return name
fifa_21['player'] = fifa_21['player'].apply(map_name)
fifa_20['player'] = fifa_20['player'].apply(map_name)
fifa_19['player'] = fifa_19['player'].apply(map_name)
fifa_18['player'] = fifa_18['player'].apply(map_name)
col = ["player", "nationality", "work_rate", "age", "height_cm", "weight_kg", "league_rank", "overall", "potential", "wage_eur",
"international_reputation", "pace", "shooting", "passing", "dribbling", "defending", 'nation_position', 'nation_jersey_number',
"physic", "attacking_crossing", "attacking_finishing", "attacking_heading_accuracy", "attacking_short_passing",
"attacking_volleys", "skill_dribbling", "skill_curve", "skill_fk_accuracy", "skill_long_passing", "skill_ball_control",
"movement_acceleration", "movement_sprint_speed", "movement_agility", "movement_reactions", "movement_balance", "power_shot_power",
"power_jumping", "power_stamina","power_strength", "power_long_shots", "mentality_aggression", "mentality_interceptions",
"mentality_positioning", "mentality_vision", "mentality_penalties", "mentality_composure", "defending_standing_tackle",
"defending_sliding_tackle", "goalkeeping_diving", "goalkeeping_handling", "goalkeeping_kicking", "goalkeeping_positioning", "goalkeeping_reflexes"]
fifa_21 = fifa_21[col]
fifa_20 = fifa_20[col]
fifa_19 = fifa_19[col]
fifa_18 = fifa_18[col]
fifa_21.rename(columns={"nationality": "team_name"}, inplace=True)
fifa_20.rename(columns={"nationality": "team_name"}, inplace=True)
fifa_19.rename(columns={"nationality": "team_name"}, inplace=True)
fifa_18.rename(columns={"nationality": "team_name"}, inplace=True)
fifa_21 = fifa_21.drop_duplicates(subset=["player", "team_name"], keep="first")
fifa_20 = fifa_20.drop_duplicates(subset=["player", "team_name"], keep="first")
fifa_19 = fifa_19.drop_duplicates(subset=["player", "team_name"], keep="first")
fifa_18 = fifa_18.drop_duplicates(subset=["player", "team_name"], keep="first")
fifa_21["year"] = 2021
fifa_20["year"] = 2020
fifa_19["year"] = 2019
fifa_18["year"] = 2018
fifa = pd.concat([fifa_21, fifa_20, fifa_19, fifa_18])
main_df = pd.merge(main_df, fifa, how="left", on=["player", "team_name", "year"])
# ## Feature Engineering
main_df.head()
# +
main_df["diff_team_points"] = main_df['team_total_points'] - main_df['opponent_total_points']
main_df["diff_team_ranking"]= main_df['team_rank'] - main_df['opponent_rank']
main_df["diff_team_market_value"] = main_df['team_market_value'] - main_df['opponent_market_value']
main_df["diff_team_mean_market_value"] = main_df['team_mean_market_value'] - main_df['opponent_mean_market_value']
main_df["diff_team_mean_squad_age"] = main_df['team_mean_squad_age'] - main_df['opponent_mean_squad_age']
main_df["diff_team_ranking"]= main_df['team_rank'] - main_df['opponent_rank']
main_df["is_senior"] = main_df["age"] > main_df["team_mean_squad_age"]
main_df["is_imbalanced"]= main_df['diff_team_ranking'].apply(lambda x: abs(x) > 7)
main_df["big_match"] = (main_df["team_rank"] + main_df["opponent_rank"]) <= 20
main_df["gap_to_potential"] = main_df["potential"] - main_df["overall"]
main_df["roi"] = main_df["points"] / main_df["value"]
main_df["more_likely_to_win"] = (main_df["hth_team_win"] - main_df["hth_opp_win"]) >= 3
main_df["work_rate"] = main_df['work_rate'].fillna("")
main_df[["attacking_work_rate", "defending_work_rate"]] = main_df["work_rate"].apply(lambda x: pd.Series(x.split("/")))
main_df = main_df.drop(["work_rate"], axis=1)
# -
main_df.drop_duplicates(subset=["player", "date"], inplace=True)
main_df.to_csv("{}/processed/dataset_md6.csv".format(DATA_DIR), index=False)
|
notebooks/md6-preparing-and-cleansing-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sweep a option in QComponent, get Ansys HFSS eigenmode solution matrix.
# # Prerequisite
# You need to have a working local installation of Ansys
# ## 1. Perform the necessary imports and create a QDesign in Metal first.
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
from qiskit_metal.analyses.sweep_options.sweeping import Sweeping
# +
# Create the design in Metal
# Create a design by specifying the chip size and open Metal GUI.
design = designs.DesignPlanar({}, True)
design.chips.main.size['size_x'] = '2mm'
design.chips.main.size['size_y'] = '2mm'
gui = MetalGUI(design)
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
from qiskit_metal.qlibrary.connectors.open_to_ground import OpenToGround
from qiskit_metal.qlibrary.interconnects.meandered import RouteMeander
# -
# ### In this example, the design consists of 1 qubit and 1 CPW connected to OpenToGround.
# +
# Allow running the same cell here multiple times to overwrite changes
design.overwrite_enabled = True
# Remove all qcomponents from GUI.
design.delete_all_components()
# So as to demonstrate the quality factor outputs easily, the
#subtrate material type is being changed to FR4_epoxy from the
#default of silicon
design.chips.main.material = 'FR4_epoxy'
q1 = TransmonPocket(
design,
'Q1',
options=dict(pad_width='425 um',
pocket_height='650um',
connection_pads=dict(
readout=dict(loc_W=+1, loc_H=+1, pad_width='200um'))))
otg = OpenToGround(design,
'open_to_ground',
options=dict(pos_x='1.75mm', pos_y='0um', orientation='0'))
readout = RouteMeander(
design, 'readout',
Dict(
total_length='6 mm',
hfss_wire_bonds = True,
fillet='90 um',
lead=dict(start_straight='100um'),
pin_inputs=Dict(start_pin=Dict(component='Q1', pin='readout'),
end_pin=Dict(component='open_to_ground', pin='open')),
))
gui.rebuild()
gui.autoscale()
# -
# ## 2. Must open Ansys HFSS at this point.
# ### Registration occurs automatically within the method "sweep_one_option_get_eigenmode_solution_data".
#
# ### Must have an open Ansys project and HFSS eigenmode design before executing render_design. Can do it manually or automate it.
#
# Manually: To create a new Ansys project, click "New", the leftmost icon right below the file menu.
#
# Automate: Use the cells below to open the Ansys application. The method sweep_one_option_get_eigenmode_solution_data will connect to ansys, activate a HFSS design, and add a setup.
# This cell will open the Ansys application. Run this cell, if Ansys, is not already open.
# A project will be opened using this command.
hfss = design.renderers.hfss
#hfss.open_ansys()
# +
### for render_design()
# Render every QComponent in QDesign.
render_qcomps = []
# Identify which kind of pins in Ansys.
# Follow details from renderer in
# QHFSSRenderer.render_design.
# No pins are open, so don't need to utilize ender_endcaps.
render_endcaps = []
#List of tuples of jj's that shouldn't be rendered.
#Follow details from renderer in QHFSSRenderer.render_design.
render_ignored_jjs = [('Q1','rect_jj')]
# Either calculate a bounding box based on the location of
# rendered geometries or use chip size from design class.
render_box_plus_buffer = True
sweep = Sweeping(design)
# +
# Pass a dict with arguments to be used add Setup to project.
# If you don't pass all the arguments, the default is determined by
# QHFSSRenderer's default_options.
# The name of setup will be "Sweep_em_setup".
# If a setup named "Sweep_em_setup" exists in the project, it will be deleted,
# and a new setup will be added with the arguments from setup_args.
em_setup_args = Dict(min_freq_ghz=None,
n_modes=None,
max_delta_f=None,
max_passes=None,
min_passes=None,
min_converged=None,
pct_refinement=None,
basis_order=None)
# Example of updating the keys that we are interested in.
em_setup_args = Dict(min_freq_ghz=5,
n_modes=2,
max_passes=9,)
# -
# ## 3 Metal pass information to Ansys, and gets a solution matrix.
# ### - Connect to Ansys HFSS, eigenmode solution.
# ### - Rebuild QComponents in Metal.
# ### - Render QComponents within HFSS and setup.
# ### - Delete/Clear the HFSS between each calculation of solution matrix.
# ### - Calculate solution matrix for each value in option_sweep.
#
# #### Return a dict and return code. If the return code is zero, there were no errors detected.
# #### The dict has: key = each value used to sweep, value = Dict with frequency, kappa_over_2pis, and quality_factor.
#
# #### This could take minutes based size of design.
# +
#The argument "leave_last_design" is true by default.
#Note: The method will connect to Ansys, activate_eigenmode_design(), add_eigenmode_setup().
all_sweeps, return_code = sweep.sweep_one_option_get_eigenmode_solution_data(
readout.name,
'total_length',
['5.8mm', '6mm', '6.2mm'],
render_qcomps,
render_endcaps,
render_ignored_jjs,
box_plus_buffer_render=render_box_plus_buffer,
design_name="GetEigenModeSolution",
setup_args=em_setup_args)
# +
if return_code ==0:
print(f'{all_sweeps.keys()} \n')
for key in all_sweeps.keys():
print(f'\nkey={key}')
option_name = all_sweeps[key]['option_name']
print(f'option_name[\'{key}\'][\'option_name\']={option_name}')
print('frequency:')
print(all_sweeps[key]['frequency'])
print('kappa_over_2pis:')
print(all_sweeps[key]['kappa_over_2pis'])
print('quality_factor:')
print(all_sweeps[key]['quality_factor'])
# -
# For example, just one group of solution data.
all_sweeps['6mm']
# +
# In HFSS, after the last sweep, should the design be cleared? The argument "leave_last_design" is true by default.
# If you want to delete after last sweep, change argument to false.
leave_last_design = False
all_sweeps, return_code = sweep.sweep_one_option_get_eigenmode_solution_data(
q1.name,
'connection_pads.readout.pad_width',
['100um', '110um', '120um'],
render_qcomps,
render_endcaps,
render_ignored_jjs,
box_plus_buffer_render=render_box_plus_buffer,
design_name="GetEigenModeSolution",
leave_last_design=leave_last_design,
setup_args=em_setup_args)
# +
# Uncomment next line if you would like to close the gui
# gui.main_window.close()
|
tutorials/6 Analysis/Sweep_an_option_get_hfss_eigenmode_matrix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from datetime import datetime, timedelta
import math
import os
import matplotlib.pyplot as plt
# -
filename = os.listdir('../dash-2019-coronavirus/cumulative_data/')
sheet_name = [i.replace('.csv', '') for i in filename if i.endswith('.csv')]
dfs = {sheet_name: pd.read_csv('../dash-2019-coronavirus/cumulative_data/{}.csv'.format(sheet_name))
for sheet_name in sheet_name}
DailyData = dfs[sheet_name[0]][['date_day']]
for region in sheet_name:
#DailyData[region] = dfs[region]['New']
DailyData.loc[:, region] = dfs[region].loc[:,'New']
DailyData.set_index('date_day', inplace=True)
DailyData = DailyData.sort_index()
DailyData = DailyData.drop(DailyData.iloc[-1].name)
DailyData.tail()
DailyData.columns
# +
fig = plt.figure(figsize=(24,24), constrained_layout=True)
axs = fig.subplots(nrows=12, ncols=12)
for i in range(len(DailyData.columns)):
axs.flat[i].plot(DailyData.index, DailyData.iloc[:,i], color='tab:blue')
axs.flat[i].get_xaxis().set_ticks([])
axs.flat[i].get_yaxis().set_ticks([])
axs.flat[i].annotate(DailyData.iloc[:,i].name, (0.05, 0.8),xycoords='axes fraction', va='center', ha='left')
# -
DailyData.iloc[:,10].name
def bray_curtis_distance(table, sample1_id, sample2_id):
numerator = 0
denominator = 0
sample1_counts = table[sample1_id]
sample2_counts = table[sample2_id]
for sample1_count, sample2_count in zip(sample1_counts, sample2_counts):
numerator += abs(sample1_count - sample2_count)
denominator += sample1_count + sample2_count
return numerator / denominator
# +
from skbio.stats.distance import DistanceMatrix
from numpy import zeros
def table_to_distances(table, pairwise_distance_fn):
sample_ids = table.columns
num_samples = len(sample_ids)
data = zeros((num_samples, num_samples))
for i, sample1_id in enumerate(sample_ids):
for j, sample2_id in enumerate(sample_ids[:i]):
data[i,j] = data[j,i] = pairwise_distance_fn(table, sample1_id, sample2_id)
return DistanceMatrix(data, sample_ids)
# -
bc_dm = table_to_distances(DailyData, bray_curtis_distance)
print(bc_dm)
bc_dm.data
bc_dm.plot(cmap='Greens')
from skbio.stats.ordination import pcoa
bc_pc = pcoa(bc_dm)
bc_pc
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca.fit(bc_dm.data)
print(pca.explained_variance_)
|
coronavirus_country_PCoA/.ipynb_checkpoints/COVID-19_analysis-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Exploring a new dataset
# - For each new dataset you encounter, you'll need to understand what state it is in
# - Exploration and cleaning will have to occur with all datasets
# - The purpose of this notebook is to function as an exploration checklist
# + [markdown] slideshow={"slide_type": "slide"}
# ## Import libraries
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + [markdown] slideshow={"slide_type": "slide"}
# ## Read in dataset
# - In this dataset, target refers to plant type which have been assigned a number between 0-2
# - There are options in pandas to explicitly specific datatypes here
# + slideshow={"slide_type": "fragment"}
df = pd.read_csv('../data/iris.csv')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Examine the data
# - Need to ensure the datatypes are correct for each column
# - Presence of unexpected datatypes may indicate the presence of errors
# + slideshow={"slide_type": "fragment"}
df.head()
# + slideshow={"slide_type": "fragment"}
df.columns
# + slideshow={"slide_type": "slide"}
df.info()
# + slideshow={"slide_type": "fragment"}
df.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Convert datatypes
# - sepal length was already a float but I'm converting it as an example
# - `errors='coerce'` will replace non-numeric data with nan (not a number)
# + slideshow={"slide_type": "fragment"}
df['sepal length (cm)'] = pd.to_numeric(df['sepal length (cm)'], errors='coerce')
df['target'] = df['target'].astype('category')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Examine categorical variables
# + slideshow={"slide_type": "fragment"}
df['target'].value_counts()
# + slideshow={"slide_type": "fragment"}
df['target'].value_counts().plot(kind='barh', color="blue", alpha=.65)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualise relationships
# - Look at the distribution of the data
# - Look for outliers or minority classes
# + slideshow={"slide_type": "slide"}
sns.pairplot(data=df) #, hue='target'
# + slideshow={"slide_type": "slide"}
df.plot(kind='box', subplots=True, layout=(2,2), sharex=False, sharey=False)
plt.show()
# + slideshow={"slide_type": "slide"}
df.corr()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Missing values
# ### Finding them
# - Check for missing values: they are indicated by a `nan`
# - Be careful, some datasets may not leave a blank space for us to infer missing values
# - Instead, you may have to look for signs of them, such as them being recorded as:
# - 0s
# - Spaces
# - You can either replace them or remove them
# - Consider whether missing values are random or if there is a systemic bias in the dataset
# - Removing data with missing values may result in you throwing too much information away
# + slideshow={"slide_type": "slide"}
# examine every column
pd.notnull(df).all()
# + slideshow={"slide_type": "fragment"}
# Examine rows with missing values
df[df['target'].isnull()]
# + [markdown] slideshow={"slide_type": "slide"}
# ### Replacing missing values
# - Simplest methods involve replacing missing values with:
# - 0s
# - mean/mean
# - min
# - You can also try to use more advanced techinques to try and infer the missing values
# + slideshow={"slide_type": "fragment"}
df = df.fillna(0)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Dropping missing values
# - Can be done at the column or row level
# - Column: axis=1
# - row: axis=0
# - You can choose how strict you want to be when filtering out missing values
# + slideshow={"slide_type": "fragment"}
# Columns: delete if all or any of the values in them are missing
df = df.dropna(axis=1, how='all')
df = df.dropna(axis=1, how='any')
# Rows: Delete if any are missing or if there are less than 2 non-missing balues
df = df.dropna(axis=0, how='any')
df.dropna(thresh=2)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Eliminating duplicate rows
# -
# keep default='first', other options include 'last' and False
df[df.duplicated(keep='first')]
# + slideshow={"slide_type": "fragment"}
# don't do this if there's a reason there may be duplicates!
df = df.drop_duplicates()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visualise relationships and categorical variables
# + slideshow={"slide_type": "fragment"}
sns.lmplot(x="sepal length (cm)", y="sepal width (cm)", hue="target", data=df, fit_reg=True)
sns.lmplot(x="petal length (cm)", y="petal width (cm)", hue="target", data=df, fit_reg=True)
|
notebooks/exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import sklearn
from sklearn import preprocessing
from sklearn.model_selection import KFold,train_test_split
import time
import lightgbm as lgb
import seaborn as sns
import matplotlib.pyplot as plt
import os
# -
train = pd.read_csv("Train_Processed_4.23.csv")
test = pd.read_csv("test_new_feature.csv")
print(train.shape, test.shape)
print(train.describe())
train.head()
#check for missing values in train data
print(train.isnull().sum().sort_values(ascending=False))
#check for missing values in test data
print(test.isnull().sum().sort_values(ascending=False))
#drop the missing values
train = train.drop(train[train.isnull().any(1)].index, axis = 0)
# +
def add_distances_features(df):
df['euclidean'] = (df['abs_diff_longitude'] ** 2 + df['abs_diff_latitude'] ** 2) ** 0.5
df['manhattan'] = df['abs_diff_longitude'] + df['abs_diff_latitude']
def add_time_features(df):
df['pickup_datetime'] = pd.to_datetime(df['pickup_datetime'], format='%Y-%m-%d %H:%M:%S %Z')
df['year'] = df['pickup_datetime'].apply(lambda x: x.year)
df['month'] = df['pickup_datetime'].apply(lambda x: x.month)
df['day'] = df['pickup_datetime'].apply(lambda x: x.day)
df['hour'] = df['pickup_datetime'].apply(lambda x: x.hour)
df['weekday'] = df['pickup_datetime'].apply(lambda x: x.weekday())
# Drop 'pickup_datetime' as we won't need it anymore
df = df.drop('pickup_datetime', axis=1)
add_distances_features(train)
add_time_features(train)
add_distances_features(test)
add_time_features(test)
# -
y_train = train.fare_amount.values
X_train = train.drop(['fare_amount'],axis=1)
# X_train, X_test, y_train, y_test = train_test_split(X_train, y_train, test_size=0.33, random_state=42)
test.head()
# +
dropped_columns = ['fare_amount','Unnamed: 0', 'key', 'pickup_datetime']
dropped_columns1 = ['key', 'pickup_datetime']
X_train = train.drop(dropped_columns, axis=1)
test_key = test['key']
X_test = test.drop(dropped_columns1,axis =1)
# -
X_test.head()
params = {
'objective': "regression",
'metric': "rmse",
'num_boost_round':1000,
'learning_rate': 0.034,
'num_leaves':31 ,
'max_depth':-1,
'subsample':0.8,
'colsample_bytree':0.6,
'min_split_gain':0.5 ,
'min_child_weight':1,
'min_child_samples':10,
'scale_pos_weight':1,
'num_threads':4,
'boosting_type':"gbdt",
'zero_as_missing':True,
'seed':0,
'eval_freq':50
}
train_set = lgb.Dataset(X_train, y_train, silent=True)
model = lgb.train(params, train_set = train_set)
# +
# from sklearn.model_selection import GridSearchCV
# #decision tree Parameters Adjustment:
# model_lgb = lgb.LGBMRegressor(objective='regression',num_leaves=50,
# learning_rate=0.1, n_estimators=43, max_depth=6,
# metric='rmse', bagging_fraction = 0.8,feature_fraction = 0.8)
# params_test1={
# 'max_depth': range(3,8,2),
# 'num_leaves':range(20, 110, 30)
# }
# gsearch1 = GridSearchCV(estimator=model_lgb, param_grid=params_test1, scoring='neg_mean_squared_error', cv=5, verbose=1, n_jobs=4)
# gsearch1.fit(X_train, y_train)
# gsearch1.grid_scores_, gsearch1.best_params_, gsearch1.best_score_
# -
pred_test_y = np.zeros(X_test.shape[0])
pred_test_y = model.predict(X_test, num_iteration = model.best_iteration)
print(pred_test_y)
submission = pd.DataFrame({"key": test_key, "fare_amount": pred_test_y.round(2)})
submission['fare_amount'] = pred_test_y
submission.to_csv('submission_LGB.csv', index=False)
submission.head(20)
|
src/LightGBM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
import os
import sys
sys.path.append('../src')
from skew import *
import xgboost as xgb
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error, mean_squared_log_error, mean_absolute_error
import pickle
data_dir = '../data/house-prices-advanced-regression-techniques'
# -
df = pd.read_csv(os.path.join(data_dir,'train.csv'))
df.head()
params_xgb = {'subsample': 0.6,
'learning_rate': 0.01,
'min_child_weight': 5,
'colsample_bytree': 0.7,
'objective': 'reg:squarederror',
'n_estimators': 1000,
'max_depth': 5}
# # Unskewed, normalized, encoded, feature selection on continuous columns only after encoding
# +
target = 'SalePrice'
mockup_skew = SkewAnalysis()
nan, processed = mockup_skew.handle_nan(df.copy(), target)
unskew, tr = mockup_skew.unskew(processed)
normedEncoded = mockup_skew.normalize_and_encode(unskew)
focus_col = [i for i in df.columns if df.loc[:,i].dtype!='object']
feature = mockup_skew.feature_selection(normedEncoded, focus_col)
feature.head()
# +
data = feature.copy()
train_col = [i for i in data.columns if i!=target]
X_train, X, y_train, y = train_test_split(data[train_col], data[target], test_size = 0.1, random_state=0, shuffle=True)
X_val, X_test, y_val, y_test = train_test_split(X, y, test_size = 0.5, random_state=42, shuffle=True)
# +
reg = xgb.XGBRegressor(**params_xgb).fit(X_train,
y_train,
early_stopping_rounds = 25,
eval_metric = ['rmse'],
eval_set = [(X_train, y_train), (X_val, y_val)],
verbose = False)
# Test data
preds, gt, index = mockup_skew.decode(reg.predict(X_test), y_test)
mae = mean_absolute_error(preds, gt)
rmsle = np.sqrt(mean_squared_log_error(preds, gt))
rmse = np.sqrt(mean_squared_error(preds, gt))
# Training metrics
results = reg.evals_result()
# print(results['validation_0'])
epochs = len(results['validation_0']['rmse'])
x_axis = range(0, epochs)
# Plot log loss
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(20,7))
ax1.plot(x_axis, results['validation_0']['rmse'], label='Train')
ax1.plot(x_axis, results['validation_1']['rmse'], label='Val')
ax1.set_xlabel('epoch', fontsize=15)
ax1.set_ylabel('RMSE', fontsize=15)
ax1.legend(fontsize=15)
ax2.plot(gt,'b-', label='Ground truth')
ax2.plot(preds,'r-', label='Prediction')
ax2.set_ylabel('Price prediction', fontsize=15)
ax2.set_title('MAE = %.2f - RMSE = %.2f - RMSLE = %.6f'%(mae, rmse, rmsle), fontsize=20)
ax2.legend(fontsize=15)
plt.tight_layout()
plt.show()
# -
# # Kurtosis check
train = pd.concat([X_train, y_train],axis=1)
# +
kurt = {}
col = [i for i in train.columns if 'x' not in i]
for i in col:
kurt[i] = stats.kurtosis(train.loc[:,i], fisher=False)
kurt_mean_ref = np.mean(list(kurt.values()))-3
# kurt_mean_ref = kurt['TotalBsmtSF'] - 3
print(kurt_mean_ref)
plt.figure(figsize=(10,7))
plt.bar(kurt.keys(), kurt.values())
plt.xticks(rotation=90)
plt.show()
# -
removed_row = []
for idx in feature.index:
df_kurt = train.loc[[i for i in df.index if i!=idx],:]
kurt = {}
for i in col:
kurt[i] = stats.kurtosis(df_kurt.loc[:,i], fisher=False)
kurt_mean = np.mean(list(kurt.values()))-3
# kurt_mean = kurt['TotalBsmtSF'] - 3
if abs(kurt_mean)<abs(kurt_mean_ref):
kurt_mean_ref = kurt_mean
removed_row.append(idx)
train = df.drop([idx])
print(len(removed_row))
train.head()
# +
kurt = {}
for i in col:
kurt[i] = stats.kurtosis(df.loc[:,i], fisher=False)
kurt_mean_ref = np.mean(list(kurt.values()))-3
print(kurt_mean_ref)
plt.figure(figsize=(10,7))
plt.bar(kurt.keys(), kurt.values())
plt.xticks(rotation=90)
plt.show()
# -
X_train = train[train_col]
y_train = train[target]
# +
reg = xgb.XGBRegressor(**params_xgb).fit(X_train,
y_train,
early_stopping_rounds = 25,
eval_metric = ['rmse'],
eval_set = [(X_train, y_train), (X_val, y_val)],
verbose = False)
# Test data
preds, gt, index = mockup_skew.decode(reg.predict(X_test), y_test)
mae = mean_absolute_error(preds, gt)
rmsle = np.sqrt(mean_squared_log_error(preds, gt))
rmse = np.sqrt(mean_squared_error(preds, gt))
# Training metrics
results = reg.evals_result()
# print(results['validation_0'])
epochs = len(results['validation_0']['rmse'])
x_axis = range(0, epochs)
# Plot log loss
fig, (ax1,ax2) = plt.subplots(1, 2, figsize=(20,7))
ax1.plot(x_axis, results['validation_0']['rmse'], label='Train')
ax1.plot(x_axis, results['validation_1']['rmse'], label='Val')
ax1.set_xlabel('epoch', fontsize=15)
ax1.set_ylabel('RMSE', fontsize=15)
ax1.legend(fontsize=15)
ax2.plot(gt,'b-', label='Ground truth')
ax2.plot(preds,'r-', label='Prediction')
ax2.set_ylabel('Price prediction', fontsize=15)
ax2.set_title('MAE = %.2f - RMSE = %.2f - RMSLE = %.6f'%(mae, rmse, rmsle), fontsize=20)
ax2.legend(fontsize=15)
plt.tight_layout()
plt.show()
# -
|
notebook/kurtosis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook illustrates the entire VQA pipeline using a TbD-Net. It loads the appropriate models necessary for performing the task, then allows a user to input a natural-language question and provide an image, get an answer back, and view intermediary attention masks.
# +
from pathlib import Path
from itertools import groupby
import numpy as np
from scipy.misc import imresize
import matplotlib.pyplot as plt
# %matplotlib inline
from tbd.module_net import load_tbd_net
from utils.clevr import load_vocab
from utils.generate_programs import load_program_generator, generate_single_program
from utils.download_pretrained_models import download
from utils.extract_features import load_feature_extractor, extract_image_feats
# -
# Now let's make sure we have all the models we need, and download them if not. We'll also load all of them here.
#
# *Note*: if you do not have our program generator model, our TbD-Net model, and PyTorch's ResNet-101 model, these will all be downloaded.
# +
vocab = load_vocab(Path('data/vocab.json'))
model_path = Path('./models')
program_generator_checkpoint = 'program_generator.pt'
tbd_net_checkpoint = 'clevr-reg-hres.pt'
if not (model_path / tbd_net_checkpoint).exists():
download(tbd_net_checkpoint)
tbd_net = load_tbd_net(model_path / tbd_net_checkpoint, vocab)
program_generator = load_program_generator(model_path / program_generator_checkpoint)
feature_extractor = load_feature_extractor()
# -
def run_question_and_image(question, img_path):
''' Utility function to forward a question and image through a TbD-Net model
and return the image and the intermediate attention masks.
Parameters
----------
tbd_net : torch.nn.Module
The TbD-Net module to forward the question and image through.
question : str
A natural-language question to produce the answer to.
img_path : Union[pathlib.Path, str]
The path to the image to use to answer `question`.
Returns
-------
Tuple[numpy.ndarray, str, List[Tuple[str, numpy.ndarray]]]
A tuple of (img, answer, [(operation, attention), ...])
Note that some of the intermediates will be `None` rather than a tuple, which
indicates a break in the logic chain. For example, in the question:
"What color is the cube to the left of the sphere and right of the cylinder?"
We have 3 distinct chains of reasoning. We first localize the sphere and look left.
We then localize the cylinder and look right. Thirdly, we look at the intersection
of these two, and find the cube. We'll encounter a None at the beginning of the
question and after the `left` and `right` operations.
'''
prog_var = generate_single_program(question, program_generator, vocab)
img, feat_var = extract_image_feats(img_path, feature_extractor)
answer, intermediaries = tbd_net.forward_and_return_intermediates(prog_var, feat_var)
return (img, answer, intermediaries)
# There's one more function to set up. We want to be able to visualize our intermediate attention masks, so we define a function that takes the intermediaries from our `run_question_and_image` function above and displays the attention masks in a logical structure.
def display_tree(intermediaries, interp='lanczos', img=None):
''' Convenience function for displaying intermediary outputs in as tree-like a
structure as it's worth computing.
Parameters
----------
intermdiaries : List[Tuple[str, numpy.ndarray]]
The [(operation, attention), ...] list to display.
interp : str
The type of interpolation to use. In our paper, we use lanczos interpolation
for the smoothed images and nearest for the rest of our visualizations.
Legal options are: (‘nearest’, ‘lanczos’, ‘bilinear’, ‘bicubic’, ‘cubic’).
img : numpy.ndarray, shape=(r, c, 3), optional (default=None)
The image to display under each attention mask.
Returns
-------
Tuple[matplotlib.figure.Figure, matplotlib.axes.Axes]
The figure and axes objects the visualization is displayed on.
'''
if img is not None:
color_to_gray = [0.2126, 0.7152, 0.0722] # luminance-preserving transform
gray = np.dot(imresize(img, (224, 224), 'bicubic'), color_to_gray)
num_rows = intermediaries.count(None) # None indicates a break in logic
x = intermediaries
# get the maximum run between None entries, which is the max number of columns
num_cols = max(len(tuple(j)) for _, j in groupby(x, key=lambda x: x is None))
# create our plots
fig, ax = plt.subplots(num_rows, num_cols, figsize=(4*num_cols, 4*num_rows),
squeeze=False)
row, col = 0, 0
intermediaries = intermediaries[1:] # the first operation (index 0) is always None
for i, intermediate in enumerate(intermediaries):
if intermediate is None:
while col < num_cols:
# remove all the axes that don't have a plot so we don't
# display annoying white boxes
ax[row, col].remove()
col += 1
row += 1
col = 0
continue
ax[row, col].set_title(intermediate[0])
interp = interp if interp else 'nearest'
if img is not None:
# if the user provides an image to underlay, display it with
# overlaid attention masks
ax[row, col].imshow(gray, cmap='gray')
ax[row, col].imshow(imresize(intermediate[1], (224, 224), interp), alpha=0.5)
else:
# otherwise just display the attention mask
ax[row, col].imshow(imresize(intermediate[1], (224, 224), interp))
ax[row, col].axis('off')
col += 1
while col < num_cols:
ax[row, col].remove()
col += 1
return (fig, ax)
# In the cell below, provide your own question and image!
img_path = Path('data/test_im_000012.png')
question = ' '.join(['What color is the big object that is left of the large metal sphere',
'and right of the green metal thing?'])
img, prediction, intermediaries = run_question_and_image(question, img_path)
# Let's visualize the outputs to see where our model was looking during each stage of the reasoning process.
# +
fig, ax = plt.subplots()
ax.imshow(img)
ax.axis('off')
fig, ax = display_tree(intermediaries, interp='lanczos', img=img)
# -
# Was the model right?
prediction
# Yes!
|
full-vqa-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="MDcEqde2UtVJ"
# # Домашнее задание "Производная функции нескольких аргументов".
# + id="hkGWP9e5TgmG"
import numpy as np
from sympy import *
from scipy.optimize import approx_fprime
# + [markdown] id="Brjig4o4UnHe"
# ### Уровень 0:
# + [markdown] id="xOv4al86VAw1"
# Посчитайте частные производные функций:
# + [markdown] id="fCyKnRYIUw2j"
#
# 1) $f(x,y)=2x^2y^3 + 1/x + y^2x + 7$, а для этой функции также градиент в точке $(1,2)$
#
# 2) $f(x,y)=x^2y - sin(xy) + cos(x^2) + 6y$
# -
x = symbols('x')
y = symbols('y')
# first function
f = 2 * x**2 * y**3 + 1/x + y**2 * x + 7
f1x = diff(f, x)
print(format(f1x))
f1y = diff(f, y)
print(format(f1y))
# second function
f = x**2 * y - sin(x * y) + cos(x**2) + 6 * y
f1x = diff(f, x)
print(format(f1x))
f1y = diff(f, y)
print(format(f1y))
# +
# gradient of the first function
def func(t):
return 2 * t[0] ** 2 * t[1] ** 3 + 1 / t[0] + t[1] ** 2 * t[0] + 7
eps = np.sqrt(np.finfo(float).eps)
grad = approx_fprime([1, 2], func, [eps, eps])
print(grad)
# + [markdown] id="COzir0ZGVLtz"
# ### Уровень 1:
# + [markdown] id="VIW3H6ChVOZQ"
# Градиентный спуск своими руками:
# + id="rFpy298AUwXs"
def f(t):
return (t[0] ** 2) + (t[1] ** 2)
# + id="-QoutSlMcZ7M"
x0 = np.array([100, 200])
learning_rate = 0.1
# + id="kBQCpgyEcZ9j"
eps = np.sqrt(np.finfo(float).eps)
x = x0
for i in range(100):
# Calculate gradient
grad = approx_fprime(x, f, [eps, eps])
# Update x with gradient
x = np.array([
x[0] - learning_rate * grad[0],
x[1] - learning_rate * grad[1]
])
print("Minimum is in: ", x)
print("Minimum value is: ", f(x))
# + [markdown] id="eAw4uXx_UnJk"
#
# Проверка с помощью встроенной функции numpy:
# + id="_O-rTJ1yciMI"
from scipy.optimize import minimize
res = minimize(f, x0, method='nelder-mead', options={'xtol': 1e-8, 'disp': True})
print(res)
# -
|
HW_5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Requirement of the notebook
# <font color='red'>To run this notebook, you will have to create an anaconda environment with python==3.6 and R, install the packages 'mvtnorm' and 'condMVNorm' in R and 'rpy2==2.9.4' in Python. </font>
# # 3.2 Comparisons of the different estimators
# ## Reproduction of figures 4 and 5
# +
import warnings
warnings.filterwarnings('ignore')
from acv_explainers import ACVTree
import shap
import matplotlib.pyplot as plt
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import roc_auc_score
import random
import time
import numpy as np
import pandas as pd
import seaborn as sns
random.seed(2021)
np.random.seed(2021)
from experiments.exp_linear import *
# Fixed the seed and plot env
random.seed(1)
np.random.seed(1)
plt.style.use(['ggplot'])
# -
def bar_plot(values_1, values_2, values_3, values_4, labels):
x = np.arange(len(labels)) # the label locations
width = 0.2 # the width of the bars
fig, ax = plt.subplots(dpi=250)
rects1 = ax.bar(x - width/2, values_1, width, label='exact')
rects2 = ax.bar(x + width/2, values_2, width, label='Monte_Carlo ')
rects3 = ax.bar(x + 1.5*width, values_3, width, label='Plug-in ')
rects4 = ax.bar(x + 2.5*width, values_4, width, label='SHAP')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Shapley values')
# ax.set_title('Scores by group and gender')
ax.set_xticks(x)
ax.set_xticklabels(labels)
ax.legend()
def autolabel(rects):
"""Attach a text label above each bar in *rects*, displaying its height."""
for rect in rects:
height = rect.get_height()
ax.annotate('{}'.format(height),
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom', color='black')
autolabel(rects1)
autolabel(rects2)
autolabel(rects3)
autolabel(rects4)
fig.tight_layout()
plt.show()
# ### Load data and model
# +
p = 0.8
n = 10000
d = 3
cov = p*np.ones(shape=(d, d)) - (p - 1)*np.eye(d)
C = [[]]
b = np.random.randn(d, d)
cov = np.dot(b, b.T)
exp = ExperimentsLinear(mean=[0, 0, 0], cov=cov, coefs=[0.5, 10, -4], n=n, C=C)
# -
seed = np.random.randint(0, 10000)
tree = DecisionTreeRegressor(min_samples_leaf=10, random_state=seed)
tree = exp.fit_model(tree)
seed
# ### Shapley values computation with the different algorithms (exact, shap, plugin, monte_carlo)
# +
ind = exp.data[np.random.randint(0, n)]
t0 = time.perf_counter()
sh_e = exp.tree_shap(tree, ind, algo='exact')
# print(sh_e)
t1 = time.perf_counter()
print('Exact - Elapsed time = {}'.format(t1-t0))
t0 = time.perf_counter()
sh_o = exp.tree_shap(tree, ind, algo='plugin')
# print(sh_o)
t1 = time.perf_counter()
print('Plug-in - Elapsed time = {}'.format(t1-t0))
t0 = time.perf_counter()
sh_mc = exp.tree_shap(tree, ind, algo='monte_carlo', N=50000)
# print(sh_mc)
t1 = time.perf_counter()
print('Monte carlo - Elapsed timetime = {}'.format(t1-t0))
t0 = time.perf_counter()
sh_alg = exp.tree_shap(tree, ind, algo='shap', N=50000)
# print(sh_alg)
t1 = time.perf_counter()
print('Monte carlo - Elapsed timetime = {}'.format(t1-t0))
bar_plot(np.round(sh_e, 2), np.round(sh_mc, 2), np.round(sh_o, 2), np.round(sh_alg, 2),\
['V_{}'.format(i) for i in range(len(sh_e))])
# -
print('x = {}'.format(ind))
# +
S = [1]
nb = 100
x_0 = np.linspace(-5, 5, nb)
sample = []
sample_lundb = []
sample_ours = []
sample_mc = []
sample_exact = []
for x in tqdm(x_0):
a = sampleMVN(1, exp.mean, exp.cov, [1, 2], [0], x)
sample.append(np.concatenate(([x], a[0])))
sample_lundb.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="shap"))
sample_ours.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="plugin"))
sample_mc.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="monte_carlo", N=n))
sample_exact.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="exact"))
plt.figure(dpi=250)
plt.plot(x_0, sample_lundb, label='SHAP')
plt.plot(x_0, sample_ours, label='Plug-In')
plt.plot(x_0, sample_exact, label='exact')
plt.plot(x_0, sample_mc, label='Monte Carlo')
plt.ylabel('E[ f(x) | X_1 ]')
plt.xlabel('X_0 range')
plt.legend()
# +
S = [2]
nb = 100
x_0 = np.linspace(-5, 5, nb)
sample = []
sample_lundb = []
sample_ours = []
sample_mc = []
sample_exact = []
for x in tqdm(x_0):
a = sampleMVN(1, exp.mean, exp.cov, [1, 2], [0], x)
sample.append(np.concatenate(([x], a[0])))
sample_lundb.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="shap"))
sample_ours.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="plugin"))
sample_mc.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="monte_carlo", N=n))
sample_exact.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="exact"))
plt.figure(dpi=250)
plt.plot(x_0, sample_lundb, label='SHAP')
plt.plot(x_0, sample_ours, label='Plug-In')
plt.plot(x_0, sample_exact, label='exact')
plt.plot(x_0, sample_mc, label='Monte Carlo')
plt.ylabel('E[ f(x) | X_2 ]')
plt.xlabel('X_0 range')
plt.legend()
# -
# ### Figure 4: Estimation of expectation $\mathbb{E} \left[ f(\boldsymbol{X}) | X_2 \right]$ with different observation and $X_0 \in \left[-4, 4\right]$.
# +
S = [1, 2]
nb = 100
x_0 = np.linspace(-5, 5, nb)
sample = []
sample_lundb = []
sample_ours = []
sample_mc = []
sample_exact = []
for x in tqdm(x_0):
a = sampleMVN(1, exp.mean, exp.cov, [1, 2], [0], x)
sample.append(np.concatenate(([x], a[0])))
sample_lundb.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="shap"))
sample_ours.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="plugin"))
sample_mc.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="monte_carlo", N=n))
sample_exact.append(exp.cond_exp_tree(x=sample[-1], tree=tree.tree_, S=S, algo="exact"))
plt.figure(dpi=250)
plt.plot(x_0, sample_lundb, label='SHAP')
plt.plot(x_0, sample_ours, label='Plug-In')
plt.plot(x_0, sample_exact, label='exact')
plt.plot(x_0, sample_mc, label='Monte Carlo')
plt.ylabel('E[ f(x) | X_1, X_2 ]')
plt.xlabel('X_0 range')
plt.legend()
# -
# ### Errors distribution of Shapley values between the exact and the different estimators
data_shap = exp.shap_fit_metric(tree, exp.data_test, nb=200, plot=True)
# ### Figure 5: Errors distribution between the true SV with exact expectation and the corresponding algorithm (SHAP, Plug-In, MC) over 200 new observations generated by $P_{\boldsymbol{X}}$.
|
notebooks/experiments_paper/comparisons_of_the_different_estimators.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Bit Classification with Deep Learning
# Deep Learning (DL) is a subset of Machine Learning that uses Neural Network inspired architecture to make predictions. Convolutional Neural Networks (CNN) are a type of DL model that is effective in learning patterns in 2-dimensional data such as images. Images of drill bit types are used to train a classifier to identify common drill bit types. See [Machine Learning for Engineers course website](https://apmonitor.com/pds) for the source code to [Image Classification: Bits and Cracks](https://apmonitor.com/pds/index.php/Main/BitClassification).
#
# <img align=left width=400px src='https://apmonitor.com/pds/uploads/Main/bit_classification.png'>
# ### Setup
# Import the following Python modules. Use ```pip``` to install any missing packages. For example, if there is an error: ```ModuleNotFoundError: No module named 'cv2'```, add a new cell and run the following command in another cell: ```pip install opencv-python```. An online search *how to install python cv2* may be needed if the name of the install package isn't the same as the import name. See additional information on [how to install and manage Python packages](https://apmonitor.com/pds/index.php/Main/InstallPythonPackages).
# + id="yF35Ku0Ms-O6"
import os
import zipfile
import urllib.request
import cv2
import re
import numpy as np
import random
import matplotlib.pyplot as plt
from tensorflow.keras.preprocessing import image
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D
# +
#pip install opencv-python
# -
# ### Labeled Photos
# Download photos from [Drill Bit Images (bit_photos.zip)](http://apmonitor.com/pds/uploads/Main/bit_photos.zip). Download and extract the images by either selecting the [link](http://apmonitor.com/pds/uploads/Main/bit_photos.zip) or running the code cell below to download and unzip the archive.
#
# <img align=left width=400px src='https://apmonitor.com/pds/uploads/Main/bit_types.png'>
# +
# download bit_photos.zip
file = 'bit_photos.zip'
url = 'http://apmonitor.com/pds/uploads/Main/'+file
urllib.request.urlretrieve(url, file)
# extract archive and remove bit_photos.zip
with zipfile.ZipFile(file, 'r') as zip_ref:
zip_ref.extractall('./')
os.remove(file)
# -
# The archive contains two folders, a test folder and train folder with subdirectories corresponding to the possible drill bit types (PDC, Roller Cone, and Spoon). The images are found within each subdirectory. The tree structure of the folders is:
#
# ```
# ├───test
# │ ├───PDC
# │ ├───Roller Cone
# │ └───Spoon
# └───train
# ├───PDC
# ├───Roller Cone
# └───Spoon
# ```
# Import the photos into the Python session. The first step is to process the images into a format that 1) makes the data readable to the model, and 2) provides more training material for the model to learn. For example, the ```train_processor``` variable scales the data so that it can be a feature (input) for the model, but also takes each images and augments it so that the model can learn from multiple variations of the same picture. It flips it horizontally, rotates it, and shifts it, and more to make sure the model learns from the shape of the bit rather than the orientation or size.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 23726, "status": "ok", "timestamp": 1618065873113, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjhknOaA2vSO9txrYBB1U01ff4hmNe3wj5xrjP9Nzg=s64", "userId": "16032387266776692701"}, "user_tz": 360} id="topbCwEWsr3q" outputId="a4116caa-a435-41f2-d2d8-71ca1e307d8c"
# Data processing
train_processor = ImageDataGenerator(rescale = 1./255, \
horizontal_flip = True, zoom_range = 0.2, \
rotation_range = 10, shear_range = 0.2, \
height_shift_range = 0.1, width_shift_range = 0.1)
test_processor = ImageDataGenerator(rescale = 1./255)
# Load data
train = train_processor.flow_from_directory('train', \
target_size = (256, 256), batch_size = 32, \
class_mode = 'categorical', shuffle = True)
test = test_processor.flow_from_directory('test', \
target_size = (256 ,256), batch_size = 32, \
class_mode = 'categorical', shuffle = False)
# -
# ### Model Building
# Next step is to build the CNN model with options for building the model. This includes the number of convolutional layers, fully connected dense layers, the number of nodes in each layer, and the number of training epochs. For more information on these parameters and Convolutional Neural Networks in general, see [Computer Vision with Deep Learning](https://apmonitor.com/pds/index.php/Main/VisionDeepLearning). Change these parameters and analyze the performance of the model.
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3288116, "status": "ok", "timestamp": 1618026147600, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "16032387266776692701"}, "user_tz": 360} id="9-jCm9Y3Ehbs" outputId="c99cfeb8-dd44-4da7-e0b7-e97ba1455875"
# choose model parameters
num_conv_layers = 2
num_dense_layers = 1
layer_size = 64
num_training_epochs = 20
# +
# Initiate model variable
model = Sequential()
# begin adding properties to model variable
# e.g. add a convolutional layer
model.add(Conv2D(layer_size, (3, 3), input_shape=(256,256, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# add additional convolutional layers based on num_conv_layers
for _ in range(num_conv_layers-1):
model.add(Conv2D(layer_size, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# reduce dimensionality
model.add(Flatten())
# add fully connected "dense" layers if specified
for _ in range(num_dense_layers):
model.add(Dense(layer_size))
model.add(Activation('relu'))
# add output layer
model.add(Dense(3))
model.add(Activation('softmax'))
# compile the sequential model with all added properties
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'],
)
# use the data already loaded previously to train/tune the model
model.fit(train,
epochs=num_training_epochs,
validation_data = test)
# save the trained model
model.save(f'bits.h5')
# -
# ### Model Testing
# The model is trained and saved as an `h5` file. The last line of the printed output contains the accuracy for both the training and testing data.
#
# ```
# Epoch 19/20
# 2/2 [====] - 3s 2s/step - loss: 0.8785 - accuracy: 0.6202 - val_loss: 0.9091 - val_accuracy: 0.6000
# Epoch 20/20
# 2/2 [====] - 3s 2s/step - loss: 0.9028 - accuracy: 0.5721 - val_loss: 0.8648 - val_accuracy: 0.6667
# ```
#
# The ```val_accuracy``` is the accuracy on the test images that are not included in the training. Hyperparameter optimization can be used to improve the accuracy by adjusting the CNN architecture, training selections, or other parameters. The function ```make_prediction``` takes the file path to a drill bit photo as an input and produces a classification result.
# + colab={"base_uri": "https://localhost:8080/", "height": 286} executionInfo={"elapsed": 3080, "status": "ok", "timestamp": 1617992171243, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjhknOaA2vSO9txrYBB1U01ff4hmNe3wj5xrjP9Nzg=s64", "userId": "16032387266776692701"}, "user_tz": 360} id="Chpl4ZrSPtHc" outputId="cb9bd334-7945-4a5d-d765-6abd8f0f0eec"
btype = ['PDC', 'Roller Cone', 'Spoon'] # possible output values
def make_prediction(image_fp):
im = cv2.imread(image_fp) # load image
plt.imshow(im)
img = image.load_img(image_fp, target_size = (256,256))
img = image.img_to_array(img)
image_array = img / 255. # scale the image
img_batch = np.expand_dims(image_array, axis = 0)
predicted_value = btype[model.predict(img_batch).argmax()]
true_value = re.search(r'(PDC)|(Roller Cone)|(Spoon)', image_fp)[0]
out = f"""Predicted Bit Type: {predicted_value}
True Bit Type: {true_value}
Correct?: {predicted_value == true_value}"""
return out
# -
# randomly select type (1-3) and image number (1-5)
i = random.randint(0,2); j = random.randint(1,5)
b = btype[i]; im = b.replace(' ','_').lower() + '_' + str(j) + '.jpg'
test_image_filepath = r'./test/'+b+'/'+im
print(make_prediction(test_image_filepath))
# The validation accuracy as well as individual testing shows that there are misclassifications.
#
# Here are a few things that can improve the accuracy for this application:
# - <b>More photos!</b> This is the most important thing, Machine Learning typically requires many photos that are representative of what the classifiers see. At this point, there are not nearly enough photos for the model to learn each bit type.
# - <b>Background Clutter.</b> Most of the images in this set have the background removed. To train a classifier to identify bit types in the field, more photos with realistic backgrounds are needed. Synthetic backgrounds can also be added.
# - <b>Hyperparameter Optimization.</b> To make the best model, the best parameters must be selected to maximize the accuracy (hyperparameter optimization). Packages such as [Hyperopt](http://hyperopt.github.io/hyperopt/) can intelligently try different parameter combinations to increase the accuracy without more data.
# <img align=left width=550px src='https://apmonitor.com/pds/uploads/Main/activity.png'>
# **Activity**: Replace the photos in the bit classification case study to build a classifier to distinguish photos of concrete with a crack (Positive) or no crack (Negative).
#
# ```python
# url=http://apmonitor.com/pds/uploads/Main/concrete_cracks.zip
# ```
#
# <img width=400px align=left src='http://apmonitor.com/pds/uploads/Main/concrete_cracks.png'>
|
Bit_Classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import time
import os
import torch
import torch.nn as nn
from torch.nn.modules.module import Module
from torch.autograd import Variable
from torch.utils.data import Dataset, DataLoader
import argparse
import torch.nn.functional as F
from torch.utils.data.dataloader import _use_shared_memory
from torch.nn import Parameter
import torchvision.models as models
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import scipy
from scipy.sparse import coo_matrix
import pdb
from tensorboardX import SummaryWriter
import os
import csv
import matplotlib.pyplot as plt
import matplotlib
import matplotlib.colors as colors
from matplotlib.backends.backend_agg import FigureCanvasAgg as FigureCanvas
from matplotlib.figure import Figure
import networkx as nx
import sklearn.metrics as metrics
import torch
import torch.nn as nn
from torch.autograd import Variable
from tensorboardX import SummaryWriter
import argparse
import os
import pickle
import random
import shutil
import time
import cross_val
import encoders
import gen.feat as featgen
import gen.data as datagen
from graph_sampler import GraphSampler
import load_data
import util
# -
class Csv_Logger():
def __init__(self, log_path, log_header):
self.log_path = os.path.abspath(log_path)
self.log_header = log_header
self.initialized = False
def _initialize(self):
self.initialized = True
parent_dir = os.path.abspath(os.path.join(self.log_path, '..'))
os.makedirs(parent_dir, exist_ok=True)
with open(self.log_path, "w") as f:
writer = csv.DictWriter(f, self.log_header)
writer.writeheader()
def write_row(self, **kwargs):
if not self.initialized:
self._initialize()
with open(self.log_path, "a") as f:
writer = csv.DictWriter(f, self.log_header)
writer.writerow(kwargs)
def load_data():
dayX=np.load('../preprocessed_data/daysX.npy').reshape(-1,24,9413).astype(int) #(2738, 9413, 24)
ydaysD=np.load('../preprocessed_data/ydaysD.npy').astype(int) #(2738, 1)
x_train, x_test, y_train, y_test = train_test_split(dayX, ydaysD, test_size=0.2)
y_test=y_test.squeeze()
y_train=y_train.squeeze()
adj_load=scipy.io.loadmat('AdjG.mat')['AdjG']
adj=adj_load.astype(float)
return adj,x_train, x_test, y_train, y_test
class Dataset(Dataset):
# Custom dataset
def __init__(self, features, labels):
labels=labels[:,np.newaxis]
self.features = [torch.from_numpy(x) for x in features]
self.labels = [torch.from_numpy(x) for x in labels]
assert len(self.features) == len(self.labels)
def __len__(self):
return len(self.features)
def __getitem__(self, item): #for-loop yield
return self.features[item], self.labels[item]
# +
def get_adjacency(order_num,adj,device):
#Identity
ret =[]
c=scipy.sparse.identity(9413)
for i in range(order_num+1): #indices, values
temp=coo_matrix(c)
values=temp.data
indices = np.vstack((temp.row, temp.col))
i = torch.LongTensor(indices).to(device)
v = torch.FloatTensor(values).to(device)
adj_v=torch.sparse.FloatTensor(i, v, torch.Size(temp.shape))
adj_v.require_grad=False
adj_v.to(device)
ret.append(adj_v)
c=c*adj
return ret
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
# -
class TAGCN_layer(Module):
def __init__(self, in_channels, out_channels,degree,adj_matrices, bias=True,include_identity=False):
super(TAGCN_layer, self).__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.weight = Parameter(torch.FloatTensor(degree+1,in_channels,out_channels))
self.degree=degree
self.bias = Parameter(torch.FloatTensor(out_channels))#parameter and variable
self.reset_parameters_xavier()
self.adj_matrices=adj_matrices
self.include_identity=include_identity
def reset_parameters_xavier(self):
nn.init.xavier_normal_(self.weight.data,gain=0.02) # Implement Xavier Uniform, 0.02
# nn.init.xavier_uniform_(self.weight.data,gain=0.02) # Implement Xavier Uniform, 0.02
nn.init.constant_(self.bias.data, 0.0)
def forward(self, inputs):
# x = inputs # samples by channels by 9413
[number_of_samples,in_channels, number_of_nodes]=inputs.shape
assert (in_channels==self.in_channels)
output=torch.zeros(number_of_nodes,number_of_samples,self.out_channels).to(device)
for out_channel in range(self.out_channels):
for i in range(self.degree):
test=torch.sum(self.weight[i+1,:,out_channel]*inputs.transpose(1,2),dim=2).view(number_of_nodes,-1)
output[:,:,out_channel] += torch.spmm(self.adj_matrices[i+1],test).to(device)
output[:,:,out_channel]=output[:,:,out_channel]+self.bias[out_channel]
return output.view(number_of_samples,self.out_channels,number_of_nodes).squeeze()
# +
class Trainer:
def __init__(self, model, optimizer,loader, max_epochs=100,load_path=None):
self.model = model
self.loader = loader
self.train_losses = []
self.val_losses = []
self.predictions = []
self.predictions_test = []
self.generated_logits = []
self.generated = []
self.generated_logits_test = []
self.generated_test = []
self.epochs = 0
self.max_epochs = max_epochs
self.optimizer = optimizer
self.criterion = nn.CrossEntropyLoss()
self.testaccuracies=[]
self.trainaccuracies=[]
self.name='./logs/'+str(time.time())+'.csv'
self.logger = Csv_Logger(self.name, ['episode', 'test_accuracy', 'train_accuracy', 'train_loss'])
if load_path is not None:
self.model.load_state_dict(
torch.load(
load_path,
map_location=lambda storage,
loc: storage))
def train(self,device):
self.model.train() # set to training mode
epoch_loss = 0
correct = 0
for batch_num, (features,labels) in enumerate(self.loader):
self.optimizer.zero_grad()
bs=features.shape[0]
features=features.to(device).float()
labels=labels.to(device).long().squeeze()
self.model=self.model.to(device)
out=self.model.forward(features)
out=out.view(bs,7)
pred = out.data.max(1, keepdim=True)[1]
predicted = pred.eq(labels.data.view_as(pred))
correct += predicted.sum()
loss = self.criterion(out,labels)
loss.backward()
self.optimizer.step()
epoch_loss += loss.item()
epoch_loss = epoch_loss / (batch_num + 1)
writer.add_scalar('loss',epoch_loss,self.epochs)
accuracy_train = correct.cpu().numpy() / len(self.loader.dataset)
writer.add_scalar('train accuracy',accuracy_train,self.epochs)
self.epochs += 1
print('[TRAIN] Epoch [%d/%d] Loss: %.4f Accuracy: %.4f'
% (self.epochs, self.max_epochs, epoch_loss,accuracy_train))
self.train_losses.append(epoch_loss)
self.trainaccuracies.append(accuracy_train)
def save_model(self, path):
torch.save(self.model.state_dict(), path)
def inference(self,inference_loader,device):
self.model = self.model.eval()
bs=64
y_true=[]
y_pred=[]
with torch.no_grad():
correct=0
for batch_num, (features,labels) in enumerate(inference_loader):
# self.optimizer.zero_grad()
bs=features.shape[0]
features=features.to(device).float()
labels=labels.to(device).long().squeeze()
self.model=self.model.to(device)
out=self.model.forward(features)
out=out.view(bs,7)
pred = out.data.max(1, keepdim=True)[1]
predicted = pred.eq(labels.data.view_as(pred))
correct += predicted.sum()
y_true=y_true+labels.cpu().numpy().tolist()
y_pred=y_pred+pred.cpu().numpy().reshape(-1).tolist()
conf_matrix=confusion_matrix(y_true,y_pred)
accuracy_test = correct.cpu().numpy() / len(inference_loader.dataset)
self.testaccuracies.append(accuracy_test)
print('[Test] Epoch [%d/%d] Accuracy: %.4f'
% (self.epochs, self.max_epochs,accuracy_test))
self.logger.write_row(episode=self.epochs, test_accuracy=accuracy_test,train_accuracy=self.trainaccuracies[-1],train_loss=self.train_losses[-1])
# print('Confusion Matrix: ',conf_matrix)
# +
def evaluate(dataset, model, args, name='Validation', max_num_examples=None):
model.eval()
labels = []
preds = []
for batch_idx, data in enumerate(dataset):
adj = Variable(data['adj'].float(), requires_grad=False).cuda()
h0 = Variable(data['feats'].float()).cuda()
labels.append(data['label'].long().numpy())
batch_num_nodes = data['num_nodes'].int().numpy()
assign_input = Variable(data['assign_feats'].float(), requires_grad=False).cuda()
ypred = model(h0, adj, batch_num_nodes, assign_x=assign_input)
_, indices = torch.max(ypred, 1)
preds.append(indices.cpu().data.numpy())
if max_num_examples is not None:
if (batch_idx+1)*args.batch_size > max_num_examples:
break
labels = np.hstack(labels)
preds = np.hstack(preds)
result = {'prec': metrics.precision_score(labels, preds, average='macro'),
'recall': metrics.recall_score(labels, preds, average='macro'),
'acc': metrics.accuracy_score(labels, preds),
'F1': metrics.f1_score(labels, preds, average="micro")}
print(name, " accuracy:", result['acc'])
return result
def gen_prefix(args):
if args.bmname is not None:
name = args.bmname
else:
name = args.dataset
name += '_' + args.method
if args.method == 'soft-assign':
name += '_l' + str(args.num_gc_layers) + 'x' + str(args.num_pool)
name += '_ar' + str(int(args.assign_ratio*100))
if args.linkpred:
name += '_lp'
else:
name += '_l' + str(args.num_gc_layers)
name += '_h' + str(args.hidden_dim) + '_o' + str(args.output_dim)
if not args.bias:
name += '_nobias'
if len(args.name_suffix) > 0:
name += '_' + args.name_suffix
return name
def gen_train_plt_name(args):
return 'results/' + gen_prefix(args) + '.png'
def log_assignment(assign_tensor, writer, epoch, batch_idx):
plt.switch_backend('agg')
fig = plt.figure(figsize=(8,6), dpi=300)
# has to be smaller than args.batch_size
for i in range(len(batch_idx)):
plt.subplot(2, 2, i+1)
plt.imshow(assign_tensor.cpu().data.numpy()[batch_idx[i]], cmap=plt.get_cmap('BuPu'))
cbar = plt.colorbar()
cbar.solids.set_edgecolor("face")
plt.tight_layout()
fig.canvas.draw()
data = np.fromstring(fig.canvas.tostring_rgb(), dtype=np.uint8, sep='')
data = data.reshape(fig.canvas.get_width_height()[::-1] + (3,))
writer.add_image('assignment', data, epoch)
def log_graph(adj, batch_num_nodes, writer, epoch, batch_idx, assign_tensor=None):
plt.switch_backend('agg')
fig = plt.figure(figsize=(8,6), dpi=300)
for i in range(len(batch_idx)):
ax = plt.subplot(2, 2, i+1)
num_nodes = batch_num_nodes[batch_idx[i]]
adj_matrix = adj[batch_idx[i], :num_nodes, :num_nodes].cpu().data.numpy()
G = nx.from_numpy_matrix(adj_matrix)
nx.draw(G, pos=nx.spring_layout(G), with_labels=True, node_color='#336699',
edge_color='grey', width=0.5, node_size=300,
alpha=0.7)
ax.xaxis.set_visible(False)
plt.tight_layout()
fig.canvas.draw()
data = np.fromstring(fig.canvas.tostring_rgb(), dtype=np.uint8, sep='')
data = data.reshape(fig.canvas.get_width_height()[::-1] + (3,))
writer.add_image('graphs', data, epoch)
# log a label-less version
#fig = plt.figure(figsize=(8,6), dpi=300)
#for i in range(len(batch_idx)):
# ax = plt.subplot(2, 2, i+1)
# num_nodes = batch_num_nodes[batch_idx[i]]
# adj_matrix = adj[batch_idx[i], :num_nodes, :num_nodes].cpu().data.numpy()
# G = nx.from_numpy_matrix(adj_matrix)
# nx.draw(G, pos=nx.spring_layout(G), with_labels=False, node_color='#336699',
# edge_color='grey', width=0.5, node_size=25,
# alpha=0.8)
#plt.tight_layout()
#fig.canvas.draw()
#data = np.fromstring(fig.canvas.tostring_rgb(), dtype=np.uint8, sep='')
#data = data.reshape(fig.canvas.get_width_height()[::-1] + (3,))
#writer.add_image('graphs_no_label', data, epoch)
# colored according to assignment
assignment = assign_tensor.cpu().data.numpy()
fig = plt.figure(figsize=(8,6), dpi=300)
num_clusters = assignment.shape[2]
all_colors = np.array(range(num_clusters))
for i in range(len(batch_idx)):
ax = plt.subplot(2, 2, i+1)
num_nodes = batch_num_nodes[batch_idx[i]]
adj_matrix = adj[batch_idx[i], :num_nodes, :num_nodes].cpu().data.numpy()
label = np.argmax(assignment[batch_idx[i]], axis=1).astype(int)
label = label[: batch_num_nodes[batch_idx[i]]]
node_colors = all_colors[label]
G = nx.from_numpy_matrix(adj_matrix)
nx.draw(G, pos=nx.spring_layout(G), with_labels=False, node_color=node_colors,
edge_color='grey', width=0.4, node_size=50, cmap=plt.get_cmap('Set1'),
vmin=0, vmax=num_clusters-1,
alpha=0.8)
plt.tight_layout()
fig.canvas.draw()
data = np.fromstring(fig.canvas.tostring_rgb(), dtype=np.uint8, sep='')
data = data.reshape(fig.canvas.get_width_height()[::-1] + (3,))
writer.add_image('graphs_colored', data, epoch)
def train(dataset, model, args, same_feat=True, val_dataset=None, test_dataset=None, writer=None,
mask_nodes = True):
writer_batch_idx = [0, 3, 6, 9]
optimizer = torch.optim.Adam(filter(lambda p : p.requires_grad, model.parameters()), lr=0.001)
iter = 0
best_val_result = {
'epoch': 0,
'loss': 0,
'acc': 0}
test_result = {
'epoch': 0,
'loss': 0,
'acc': 0}
train_accs = []
train_epochs = []
best_val_accs = []
best_val_epochs = []
test_accs = []
test_epochs = []
val_accs = []
for epoch in range(args.num_epochs):
begin_time = time.time()
avg_loss = 0.0
model.train()
print('Epoch: ', epoch)
for batch_idx, data in enumerate(dataset):
model.zero_grad()
adj = Variable(data['adj'].float(), requires_grad=False).cuda()
h0 = Variable(data['feats'].float(), requires_grad=False).cuda()
label = Variable(data['label'].long()).cuda()
batch_num_nodes = data['num_nodes'].int().numpy() if mask_nodes else None
assign_input = Variable(data['assign_feats'].float(), requires_grad=False).cuda()
ypred = model(h0, adj, batch_num_nodes, assign_x=assign_input)
if not args.method == 'soft-assign' or not args.linkpred:
loss = model.loss(ypred, label)
else:
loss = model.loss(ypred, label, adj, batch_num_nodes)
loss.backward()
nn.utils.clip_grad_norm(model.parameters(), args.clip)
optimizer.step()
iter += 1
avg_loss += loss
#if iter % 20 == 0:
# print('Iter: ', iter, ', loss: ', loss.data[0])
# log once per XX epochs
if epoch % 10 == 0 and batch_idx == len(dataset) // 2 and args.method == 'soft-assign' and writer is not None:
log_assignment(model.assign_tensor, writer, epoch, writer_batch_idx)
log_graph(adj, batch_num_nodes, writer, epoch, writer_batch_idx, model.assign_tensor)
avg_loss /= batch_idx + 1
elapsed = time.time() - begin_time
if writer is not None:
writer.add_scalar('loss/avg_loss', avg_loss, epoch)
if args.linkpred:
writer.add_scalar('loss/linkpred_loss', model.link_loss, epoch)
print('Avg loss: ', avg_loss, '; epoch time: ', elapsed)
result = evaluate(dataset, model, args, name='Train', max_num_examples=100)
train_accs.append(result['acc'])
train_epochs.append(epoch)
if val_dataset is not None:
val_result = evaluate(val_dataset, model, args, name='Validation')
val_accs.append(val_result['acc'])
if val_result['acc'] > best_val_result['acc'] - 1e-7:
best_val_result['acc'] = val_result['acc']
best_val_result['epoch'] = epoch
best_val_result['loss'] = avg_loss
if test_dataset is not None:
test_result = evaluate(test_dataset, model, args, name='Test')
test_result['epoch'] = epoch
if writer is not None:
writer.add_scalar('acc/train_acc', result['acc'], epoch)
writer.add_scalar('acc/val_acc', val_result['acc'], epoch)
writer.add_scalar('loss/best_val_loss', best_val_result['loss'], epoch)
if test_dataset is not None:
writer.add_scalar('acc/test_acc', test_result['acc'], epoch)
print('Best val result: ', best_val_result)
best_val_epochs.append(best_val_result['epoch'])
best_val_accs.append(best_val_result['acc'])
if test_dataset is not None:
print('Test result: ', test_result)
test_epochs.append(test_result['epoch'])
test_accs.append(test_result['acc'])
matplotlib.style.use('seaborn')
plt.switch_backend('agg')
plt.figure()
plt.plot(train_epochs, util.exp_moving_avg(train_accs, 0.85), '-', lw=1)
if test_dataset is not None:
plt.plot(best_val_epochs, best_val_accs, 'bo', test_epochs, test_accs, 'go')
plt.legend(['train', 'val', 'test'])
else:
plt.plot(best_val_epochs, best_val_accs, 'bo')
plt.legend(['train', 'val'])
plt.savefig(gen_train_plt_name(args), dpi=600)
plt.close()
matplotlib.style.use('default')
return model, val_accs
def prepare_data(graphs, args, test_graphs=None, max_nodes=0):
random.shuffle(graphs)
if test_graphs is None:
train_idx = int(len(graphs) * args.train_ratio)
test_idx = int(len(graphs) * (1-args.test_ratio))
train_graphs = graphs[:train_idx]
val_graphs = graphs[train_idx: test_idx]
test_graphs = graphs[test_idx:]
else:
train_idx = int(len(graphs) * args.train_ratio)
train_graphs = graphs[:train_idx]
val_graphs = graph[train_idx:]
print('Num training graphs: ', len(train_graphs),
'; Num validation graphs: ', len(val_graphs),
'; Num testing graphs: ', len(test_graphs))
print('Number of graphs: ', len(graphs))
print('Number of edges: ', sum([G.number_of_edges() for G in graphs]))
print('Max, avg, std of graph size: ',
max([G.number_of_nodes() for G in graphs]), ', '
"{0:.2f}".format(np.mean([G.number_of_nodes() for G in graphs])), ', '
"{0:.2f}".format(np.std([G.number_of_nodes() for G in graphs])))
# minibatch
dataset_sampler = GraphSampler(train_graphs, normalize=False, max_num_nodes=max_nodes,
features=args.feature_type)
train_dataset_loader = torch.utils.data.DataLoader(
dataset_sampler,
batch_size=args.batch_size,
shuffle=True,
num_workers=args.num_workers)
dataset_sampler = GraphSampler(val_graphs, normalize=False, max_num_nodes=max_nodes,
features=args.feature_type)
val_dataset_loader = torch.utils.data.DataLoader(
dataset_sampler,
batch_size=args.batch_size,
shuffle=False,
num_workers=args.num_workers)
dataset_sampler = GraphSampler(test_graphs, normalize=False, max_num_nodes=max_nodes,
features=args.feature_type)
test_dataset_loader = torch.utils.data.DataLoader(
dataset_sampler,
batch_size=args.batch_size,
shuffle=False,
num_workers=args.num_workers)
return train_dataset_loader, val_dataset_loader, test_dataset_loader, \
dataset_sampler.max_num_nodes, dataset_sampler.feat_dim, dataset_sampler.assign_feat_dim
def syn_community1v2(args, writer=None, export_graphs=False):
# data
graphs1 = datagen.gen_ba(range(40, 60), range(4, 5), 500,
featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float)))
for G in graphs1:
G.graph['label'] = 0
if export_graphs:
util.draw_graph_list(graphs1[:16], 4, 4, 'figs/ba')
graphs2 = datagen.gen_2community_ba(range(20, 30), range(4, 5), 500, 0.3,
[featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float))])
for G in graphs2:
G.graph['label'] = 1
if export_graphs:
util.draw_graph_list(graphs2[:16], 4, 4, 'figs/ba2')
graphs = graphs1 + graphs2
train_dataset, val_dataset, test_dataset, max_num_nodes, input_dim, assign_input_dim = prepare_data(graphs, args)
if args.method == 'soft-assign':
print('Method: soft-assign')
model = encoders.SoftPoolingGcnEncoder(
max_num_nodes,
input_dim, args.hidden_dim, args.output_dim, args.num_classes, args.num_gc_layers,
args.hidden_dim, assign_ratio=args.assign_ratio, num_pooling=args.num_pool,
bn=args.bn, linkpred=args.linkpred, assign_input_dim=assign_input_dim).cuda()
elif args.method == 'base-set2set':
print('Method: base-set2set')
model = encoders.GcnSet2SetEncoder(input_dim, args.hidden_dim, args.output_dim, 2,
args.num_gc_layers, bn=args.bn).cuda()
else:
print('Method: base')
model = encoders.GcnEncoderGraph(input_dim, args.hidden_dim, args.output_dim, 2,
args.num_gc_layers, bn=args.bn).cuda()
train(train_dataset, model, args, val_dataset=val_dataset, test_dataset=test_dataset,
writer=writer)
def syn_community2hier(args, writer=None):
# data
feat_gen = [featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float))]
graphs1 = datagen.gen_2hier(1000, [2,4], 10, range(4,5), 0.1, 0.03, feat_gen)
graphs2 = datagen.gen_2hier(1000, [3,3], 10, range(4,5), 0.1, 0.03, feat_gen)
graphs3 = datagen.gen_2community_ba(range(28, 33), range(4,7), 1000, 0.25, feat_gen)
for G in graphs1:
G.graph['label'] = 0
for G in graphs2:
G.graph['label'] = 1
for G in graphs3:
G.graph['label'] = 2
graphs = graphs1 + graphs2 + graphs3
train_dataset, val_dataset, test_dataset, max_num_nodes, input_dim, assign_input_dim = prepare_data(graphs, args)
if args.method == 'soft-assign':
print('Method: soft-assign')
model = encoders.SoftPoolingGcnEncoder(
max_num_nodes,
input_dim, args.hidden_dim, args.output_dim, args.num_classes, args.num_gc_layers,
args.hidden_dim, assign_ratio=args.assign_ratio, num_pooling=args.num_pool,
bn=args.bn, linkpred=args.linkpred, args=args, assign_input_dim=assign_input_dim).cuda()
elif args.method == 'base-set2set':
print('Method: base-set2set')
model = encoders.GcnSet2SetEncoder(input_dim, args.hidden_dim, args.output_dim, 2,
args.num_gc_layers, bn=args.bn, args=args, assign_input_dim=assign_input_dim).cuda()
else:
print('Method: base')
model = encoders.GcnEncoderGraph(input_dim, args.hidden_dim, args.output_dim, 2,
args.num_gc_layers, bn=args.bn, args=args).cuda()
train(train_dataset, model, args, val_dataset=val_dataset, test_dataset=test_dataset,
writer=writer)
def pkl_task(args, feat=None):
with open(os.path.join(args.datadir, args.pkl_fname), 'rb') as pkl_file:
data = pickle.load(pkl_file)
graphs = data[0]
labels = data[1]
test_graphs = data[2]
test_labels = data[3]
for i in range(len(graphs)):
graphs[i].graph['label'] = labels[i]
for i in range(len(test_graphs)):
test_graphs[i].graph['label'] = test_labels[i]
if feat is None:
featgen_const = featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float))
for G in graphs:
featgen_const.gen_node_features(G)
for G in test_graphs:
featgen_const.gen_node_features(G)
train_dataset, test_dataset, max_num_nodes = prepare_data(graphs, args, test_graphs=test_graphs)
model = encoders.GcnEncoderGraph(
args.input_dim, args.hidden_dim, args.output_dim, args.num_classes,
args.num_gc_layers, bn=args.bn).cuda()
train(train_dataset, model, args, test_dataset=test_dataset)
evaluate(test_dataset, model, args, 'Validation')
def benchmark_task(args, writer=None, feat='node-label'):
graphs = load_data.read_graphfile(args.datadir, args.bmname, max_nodes=args.max_nodes)
if feat == 'node-feat' and 'feat_dim' in graphs[0].graph:
print('Using node features')
input_dim = graphs[0].graph['feat_dim']
elif feat == 'node-label' and 'label' in graphs[0].node[0]:
print('Using node labels')
for G in graphs:
for u in G.nodes():
G.node[u]['feat'] = np.array(G.node[u]['label'])
else:
print('Using constant labels')
featgen_const = featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float))
for G in graphs:
featgen_const.gen_node_features(G)
train_dataset, val_dataset, test_dataset, max_num_nodes, input_dim, assign_input_dim = \
prepare_data(graphs, args, max_nodes=args.max_nodes)
if args.method == 'soft-assign':
print('Method: soft-assign')
model = encoders.SoftPoolingGcnEncoder(
max_num_nodes,
input_dim, args.hidden_dim, args.output_dim, args.num_classes, args.num_gc_layers,
args.hidden_dim, assign_ratio=args.assign_ratio, num_pooling=args.num_pool,
bn=args.bn, dropout=args.dropout, linkpred=args.linkpred, args=args,
assign_input_dim=assign_input_dim).cuda()
elif args.method == 'base-set2set':
print('Method: base-set2set')
model = encoders.GcnSet2SetEncoder(
input_dim, args.hidden_dim, args.output_dim, args.num_classes,
args.num_gc_layers, bn=args.bn, dropout=args.dropout, args=args).cuda()
else:
print('Method: base')
model = encoders.GcnEncoderGraph(
input_dim, args.hidden_dim, args.output_dim, args.num_classes,
args.num_gc_layers, bn=args.bn, dropout=args.dropout, args=args).cuda()
train(train_dataset, model, args, val_dataset=val_dataset, test_dataset=test_dataset,
writer=writer)
evaluate(test_dataset, model, args, 'Validation')
def benchmark_task_val(args, writer=None, feat='node-label'):
all_vals = []
graphs = load_data.read_graphfile(args.datadir, args.bmname, max_nodes=args.max_nodes)
if feat == 'node-feat' and 'feat_dim' in graphs[0].graph:
print('Using node features')
input_dim = graphs[0].graph['feat_dim']
elif feat == 'node-label' and 'label' in graphs[0].node[0]:
print('Using node labels')
for G in graphs:
for u in G.nodes():
G.node[u]['feat'] = np.array(G.node[u]['label'])
else:
print('Using constant labels')
featgen_const = featgen.ConstFeatureGen(np.ones(args.input_dim, dtype=float))
for G in graphs:
featgen_const.gen_node_features(G)
for i in range(10):
train_dataset, val_dataset, max_num_nodes, input_dim, assign_input_dim = \
cross_val.prepare_val_data(graphs, args, i, max_nodes=args.max_nodes)
if args.method == 'soft-assign':
print('Method: soft-assign')
model = encoders.SoftPoolingGcnEncoder(
max_num_nodes,
input_dim, args.hidden_dim, args.output_dim, args.num_classes, args.num_gc_layers,
args.hidden_dim, assign_ratio=args.assign_ratio, num_pooling=args.num_pool,
bn=args.bn, dropout=args.dropout, linkpred=args.linkpred, args=args,
assign_input_dim=assign_input_dim).cuda()
elif args.method == 'base-set2set':
print('Method: base-set2set')
model = encoders.GcnSet2SetEncoder(
input_dim, args.hidden_dim, args.output_dim, args.num_classes,
args.num_gc_layers, bn=args.bn, dropout=args.dropout, args=args).cuda()
else:
print('Method: base')
model = encoders.GcnEncoderGraph(
input_dim, args.hidden_dim, args.output_dim, args.num_classes,
args.num_gc_layers, bn=args.bn, dropout=args.dropout, args=args).cuda()
_, val_accs = train(train_dataset, model, args, val_dataset=val_dataset, test_dataset=None,
writer=writer)
all_vals.append(np.array(val_accs))
all_vals = np.vstack(all_vals)
all_vals = np.mean(all_vals, axis=0)
print(all_vals)
print(np.max(all_vals))
print(np.argmax(all_vals))
def arg_parse():
parser = argparse.ArgumentParser(description='GraphPool arguments.')
io_parser = parser.add_mutually_exclusive_group(required=False)
io_parser.add_argument('--dataset', dest='dataset',
help='Input dataset.')
benchmark_parser = io_parser.add_argument_group()
benchmark_parser.add_argument('--bmname', dest='bmname',
help='Name of the benchmark dataset')
io_parser.add_argument('--pkl', dest='pkl_fname',
help='Name of the pkl data file')
softpool_parser = parser.add_argument_group()
softpool_parser.add_argument('--assign-ratio', dest='assign_ratio', type=float,
help='ratio of number of nodes in consecutive layers')
softpool_parser.add_argument('--num-pool', dest='num_pool', type=int,
help='number of pooling layers')
parser.add_argument('--linkpred', dest='linkpred', action='store_const',
const=True, default=False,
help='Whether link prediction side objective is used')
parser.add_argument('--datadir', dest='datadir',
help='Directory where benchmark is located')
parser.add_argument('--logdir', dest='logdir',
help='Tensorboard log directory')
parser.add_argument('--cuda', dest='cuda',
help='CUDA.')
parser.add_argument('--max-nodes', dest='max_nodes', type=int,
help='Maximum number of nodes (ignore graghs with nodes exceeding the number.')
parser.add_argument('--lr', dest='lr', type=float,
help='Learning rate.')
parser.add_argument('--clip', dest='clip', type=float,
help='Gradient clipping.')
parser.add_argument('--batch-size', dest='batch_size', type=int,
help='Batch size.')
parser.add_argument('--epochs', dest='num_epochs', type=int,
help='Number of epochs to train.')
parser.add_argument('--train-ratio', dest='train_ratio', type=float,
help='Ratio of number of graphs training set to all graphs.')
parser.add_argument('--num_workers', dest='num_workers', type=int,
help='Number of workers to load data.')
parser.add_argument('--feature', dest='feature_type',
help='Feature used for encoder. Can be: id, deg')
parser.add_argument('--input-dim', dest='input_dim', type=int,
help='Input feature dimension')
parser.add_argument('--hidden-dim', dest='hidden_dim', type=int,
help='Hidden dimension')
parser.add_argument('--output-dim', dest='output_dim', type=int,
help='Output dimension')
parser.add_argument('--num-classes', dest='num_classes', type=int,
help='Number of label classes')
parser.add_argument('--num-gc-layers', dest='num_gc_layers', type=int,
help='Number of graph convolution layers before each pooling')
parser.add_argument('--nobn', dest='bn', action='store_const',
const=False, default=True,
help='Whether batch normalization is used')
parser.add_argument('--dropout', dest='dropout', type=float,
help='Dropout rate.')
parser.add_argument('--nobias', dest='bias', action='store_const',
const=False, default=True,
help='Whether to add bias. Default to True.')
parser.add_argument('--method', dest='method',
help='Method. Possible values: base, base-set2set, soft-assign')
parser.add_argument('--name-suffix', dest='name_suffix',
help='suffix added to the output filename')
parser.set_defaults(datadir='data',
logdir='log',
dataset='syn1v2',
max_nodes=1000,
cuda='1',
feature_type='default',
lr=0.001,
clip=2.0,
batch_size=20,
num_epochs=1000,
train_ratio=0.8,
test_ratio=0.1,
num_workers=1,
input_dim=10,
hidden_dim=20,
output_dim=20,
num_classes=2,
num_gc_layers=3,
dropout=0.0,
method='base',
name_suffix='',
assign_ratio=0.1,
num_pool=1
)
return parser.parse_args()
# +
# export scalar data to JSON for external processing
class prog_args_init:
def __init__(self):
self.datadir='data'
self.logdir='log'
self.dataset='syn1v2'
self.max_nodes=1000
self.cuda='1'
self.feature_type='default'
self.lr=0.001
self.clip=2.0
self.batch_size=20
self.num_epochs=1000
self.train_ratio=0.8
self.test_ratio=0.1
self.num_workers=0
self.input_dim=10
self.hidden_dim=20
self.output_dim=20
self.num_classes=2
self.num_gc_layers=3
self.dropout=0.0
self.method='base'
self.name_suffix=''
self.assign_ratio=0.1
self.num_pool=1
self.bmname=None
self.bias=None
self.pkl_fname=None
self.bn=None
self.linkpred=None
prog_args=prog_args_init()
path = os.path.join(prog_args.logdir, gen_prefix(prog_args))
if os.path.isdir(path):
print('Remove existing log dir: ', path)
shutil.rmtree(path)
writer = SummaryWriter(path)
#writer = None
os.environ['CUDA_VISIBLE_DEVICES'] = prog_args.cuda
print('CUDA', prog_args.cuda)
if prog_args.bmname is not None:
benchmark_task_val(prog_args, writer=writer)
elif prog_args.pkl_fname is not None:
pkl_task(prog_args)
elif prog_args.dataset is not None:
if prog_args.dataset == 'syn1v2':
syn_community1v2(prog_args, writer=writer)
if prog_args.dataset == 'syn2hier':
syn_community2hier(prog_args, writer=writer)
writer.close()
# +
# adj,x_train, x_test, y_train, y_test=load_data()
# number_of_nodes=9413
# number_of_classes=7
# writer = SummaryWriter()
# -
# #hyperparameters
# EPOCHS=400
# lr=1e-3
# # wd=1.2e-6
# # wd=1e-6
# # batch_size=2048
# batch_size=256
# include_identity=False
# +
# device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# train_dataset=Dataset(x_train,y_train)
# train_loader = DataLoader(train_dataset,shuffle=True, batch_size=batch_size)
# test_dataset=Dataset(x_test,y_test)
# test_loader = DataLoader(test_dataset,shuffle=False, batch_size=batch_size)
# +
# epoch=0
# for degree in [3]: #,3,4 #1,2,3,4
# # for dropout_rate in [0.2,0.5,0.8]:
# print('Degree: ', degree)
# adj_matrices=get_adjacency(degree,adj,device)
# model=nn.Sequential(TAGCN_layer(24,1,degree,adj_matrices,include_identity),nn.ELU(),nn.Linear(number_of_nodes,number_of_classes))
# # model=nn.Sequential(TAGCN_layer(24,1,degree,adj_matrices,include_identity),nn.ReLU(),nn.BatchNorm1d(number_of_nodes), nn.Linear(number_of_nodes,number_of_classes))
# # model=nn.Sequential(TAGCN_layer(24,1,degree,adj_matrices,include_identity),nn.ReLU(),nn.Linear(number_of_nodes,number_of_classes))
# # model=nn.Sequential(TAGCN_layer(24,24,degree,adj_matrices,include_identity),nn.ELU(),TAGCN_layer(24,8,degree,adj_matrices,include_identity),nn.ELU(),TAGCN_layer(8,1,degree,adj_matrices,include_identity),nn.ELU(),nn.Linear(number_of_nodes,number_of_classes))
# # model=nn.Sequential(TAGCN_layer(24,1,degree,adj_matrices,include_identity),nn.ELU(),nn.Linear(number_of_nodes,number_of_classes))
# print('Number of parameters: ', count_parameters(model))
# AdamOptimizer = torch.optim.Adam(model.parameters())
# trainer=Trainer(model,AdamOptimizer,train_loader, max_epochs=EPOCHS)
# epoch=0
# while epoch<EPOCHS:
# ts=time.time()
# epoch+=1
# trainer.train(device)
# trainer.inference(test_loader,device)
# te=time.time()
# print(te-ts)
# # del adj_matrices
# # print(np.mean(trainer.testaccuracies))
# # print(np.var(trainer.testaccuracies))
# -
plt.figure()
plt.plot(range(1, trainer.epochs + 1), trainer.train_losses, label='Training losses')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
trainer.train_losses
print(adj.shape)
print('Test Accuracies')
print(trainer.testaccuracies)
print('Train Losses')
print(trainer.train_losses)
print('Train Accuracies')
print(trainer.trainaccuracies)
|
TAGCN_main-pooling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process raw WAV files
# This notebook clips WAV files in the specified 'directory' to a specified 'clip_duration' starting from a time calculated from a time specified in the WAV filename and a specified 'watch_offset'.
# #### Import statements
import os.path
from os import listdir
import wave
from audioop import ratecv
import datetime
import time
# #### Variable declarations
# directory – path to directory containing WAV files to be processed
directory = ""
# set clip duration
clip_duration = datetime.timedelta(minutes=5)
# set watch offset
watch_offset = datetime.timedelta(minutes=2, seconds=25)
watch_start_time = ''
# set time change offset
time_shift = datetime.timedelta(hours=0)
# specify specific start time
specific_start_time = {'enabled':True, 'hour':16, 'minute':45, 'second':10}
# set directories
directory = '/Volumes/jPassport/alpine soundscapes/field recordings/2016-03-07/'
os.chdir(directory)
clip_directory = directory + 'clipped/'
if os.path.exists(clip_directory):
os.rmdir(clip_directory)
os.mkdir(clip_directory)
# find all WAV files in directory
#WAV_files = [f for f in listdir(directory) if os.path.isfile(os.path.join(directory, f)) and f.split('.')[1] == 'WAV']
WAV_files = [f for f in listdir(directory) if os.path.isfile(os.path.join(directory, f)) and f.split('.')[1] == 'WAV' and f == '160301-164729.WAV']
print(WAV_files)
# +
# clip all WAV files in the specified directory
cliplog_path = './clipped/cliplog.txt'
log_count = 0
while os.path.exists(cliplog_path):
log_count = log_count + 1
cliplog_path = './clipped/cliplog_{0}.txt'.format(log_count)
cliplog = open(cliplog_path, 'w')
text = 'This operation was performed on %s' % str(datetime.datetime.now())
printlog(cliplog, text)
text = 'The watch offset is set to: %s' % str(watch_offset)
printlog(cliplog, text)
printlog(cliplog, ' ')
text = 'Found the following WAV files:'
printlog(cliplog, text)
printlog(cliplog, ' ')
for WAV_file in WAV_files:
text = WAV_file
printlog(cliplog, text)
printlog(cliplog, ' ')
text = 'Processing...'
printlog(cliplog, text)
printlog(cliplog, ' ')
for WAV_file in WAV_files:
clip_offset, start_time = determine_clip_offset(WAV_file)
clip_WAV_file(WAV_file, clip_offset, start_time, cliplog)
printlog(cliplog, ' ')
printlog(cliplog, ' ')
text = 'Done clipping all WAV files in the specified directory.'
printlog(cliplog, text)
cliplog.close()
# -
# open WAV file
def open_WAV_file(path, mode):
WAV_file = wave.open(path, mode)
return WAV_file
# +
# determine clip_offset from time in filename
def determine_clip_offset(filename):
time_string = filename.split('.')[0]
time_year = int('20' + time_string[0:2])
time_month = int(time_string[2:4])
time_day = int(time_string[4:6])
time_hour = int(time_string[7:9])
time_minute = int(time_string[9:11])
time_second = int(time_string[11:13])
machine_start_time = datetime.datetime(year=time_year, month=time_month, day=time_day, hour=time_hour, minute=time_minute, second=time_second)
global watch_start_time
watch_start_time = (machine_start_time + time_shift) - watch_offset
if specific_start_time['enabled']:
start_time = watch_start_time.replace(hour=specific_start_time['hour'], minute=specific_start_time['minute'], second=specific_start_time['second'])
end_time = start_time + clip_duration
clip_offset = start_time - watch_start_time
else:
a, r = divmod(watch_start_time.minute, 5)
start_minute = a*5 + 5
if start_minute == 60:
start_minute = 0
start_hour = watch_start_time.hour + 1
else:
start_hour = watch_start_time.hour
start_time = watch_start_time.replace(hour=start_hour, minute=start_minute, second=0)
end_time = start_time + clip_duration
clip_offset = start_time - watch_start_time
return clip_offset, start_time
# +
# clip WAV file
def clip_WAV_file(filename, clip_offset, start_time, cliplog):
clip_filename = start_time.strftime('%y%m%d-%H%M%S.WAV')
if os.path.exists(os.path.join(clip_directory, clip_filename)):
os.remove(os.path.join(clip_directory, clip_filename))
waveread = open_WAV_file(os.path.join(directory, filename), 'rb')
wavewrite = open_WAV_file(os.path.join(clip_directory, clip_filename), 'wb')
text = filename
printlog(cliplog, text)
text = 'watch start: %s' % str(watch_start_time)
printlog(cliplog, text)
text = 'start offset: %s' % str(clip_offset)
printlog(cliplog, text)
text = filename + ' --> will be clipped as --> ' + clip_filename
printlog(cliplog, text)
params = waveread.getparams()
framerate = params[2]
nframes_read = int(clip_duration.total_seconds()*framerate)
nreads = 100
buffer_size = nframes_read / nreads
nframes_write = nframes_read
wavewrite.setparams((params[0], params[1], framerate, nframes_write, params[4], params[5]))
# report if time_shift is enabled
if time_shift:
text = 'NOTE: The clip was shifted %s hours forward! (daylight savings)' % str(time_shift)
printlog(cliplog, text)
# check if clip offset should be shifted forward
recording_length = float(params[3])/framerate
clip_offset_shift = 0
while (clip_offset.total_seconds() + clip_duration.total_seconds()) > recording_length - 10:
clip_offset = clip_offset - datetime.timedelta(seconds=1)
clip_offset_shift = clip_offset_shift + 1
if clip_offset_shift:
text = 'NOTE: The clip offset was shifted %i seconds forward!' % clip_offset_shift
printlog(cliplog, text)
startpos = int(clip_offset.total_seconds()*framerate)
waveread.setpos(startpos)
print('Clipping'),
for w in xrange(nreads):
write_buffer = waveread.readframes(buffer_size)
wavewrite.writeframes(write_buffer)
print('.'),
print(' done.')
wavewrite.close()
waveread.close()
# -
# print and log text
def printlog(file, text):
print(text)
file.write(text)
file.write('\n')
# check time offset
os.chdir(directory)
# find all WAV files in directory
WAV_files = [f for f in listdir(directory) if os.path.isfile(os.path.join(directory, f)) and f.split('.')[1] == 'WAV']
for WAV_file in WAV_files:
clip_offset, start_time = determine_clip_offset(WAV_file)
print('%s:' % WAV_file)
print('offset: %s ' % str(clip_offset))
print('start : %s ' % str(start_time))
print('\n')
|
process recordings/Process raw WAV files.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# ### LASSO
df_train = pd.read_csv('../data/train_con_features.csv', index_col='Unnamed: 0')
df_test = pd.read_csv('../data/test_con_features.csv', index_col='Unnamed: 0')
df_train.head()
# +
lst_propiedades = ['antiguedad', 'habitaciones', 'garages', 'banos', 'metroscubiertos', 'metrostotales',
'gimnasio', 'usosmultiples', 'piscina', 'escuelascercanas', 'centroscomercialescercanos',
'cantidad_amenities', 'cantidad_servicios', 'relacion_metros', 'metros_por_habitacion',
'metros_por_bano', 'metros_por_amenities', 'es_avenida']
lst_tipos_propiedades = ['es_Apartamento', 'es_Casa_en_condominio', 'es_Casa', 'es_Terreno',
'es_Terreno_comercial', 'es_Local_Comercial', 'es_Quinta_Vacacional',
'es_Oficina_comercial', 'es_Edificio', 'es_Casa_uso_de_suelo',
'es_Local_en_centro_comercial', 'es_Bodega_comercial', 'es_Otros',
'es_Villa', 'es_Duplex', 'es_Inmuebles_productivos_urbanos',
'es_Departamento_Compartido', 'es_Nave_industrial', 'es_Rancho',
'es_Terreno_industrial', 'es_Huerta', 'es_Lote', 'es_Hospedaje', 'es_Garage']
lst_cantidad_palabras = ['seguridad_descripcion', 'moderno_descripcion', 'hermoso_descripcion', 'estado_descripcion',
'seguridad_titulo', 'moderno_titulo', 'hermoso_titulo', 'estado_titulo',
'relacion_palabras_descripcion', 'cantidad_palabras_descripcion']
lst_ciudades = ['provincia_top5', 'es_ciudad_capital', 'ciudad_turistica_top15']
columnas_numericas = lst_propiedades + lst_tipos_propiedades + lst_cantidad_palabras
columnas_numericas_y_precio = lst_propiedades + lst_tipos_propiedades + lst_cantidad_palabras
columnas_numericas_y_precio.append('precio')
# -
X = df_train[columnas_numericas_y_precio].drop(columns=['precio']).values
y = df_train['precio'].values
# +
# Métrica de evaluación
#def RMSLE(actual, pred):
# return (np.mean((np.log(actual + 1) - np.log(pred + 1)) ** 2)) **.5
#alphas = [0.02, 0.3, 0.5, 0.7, 0.9, 1.5]
#for un_alpha in alphas:
# lasso_fit_alpha, lasso_pred_alpha = tunning_alpha(X_train, y_train, X_test, un_alpha)
# lasso_rmsle = RMSLE(y_test, lasso_pred_alpha)
# lasso_rmsle_train = RMSLE(y_train, lasso_fit_alpha.predict(X_train))
# print(f"RMSLE Lasso (train): {lasso_rmsle_train:.5f}", "con alpha =", un_alpha)
# print(f"RMSLE Lasso: {lasso_rmsle:.5f}", "con alpha =", un_alpha)
# print("------------------------------")
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.2, random_state=0)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# +
from sklearn import linear_model
def tunning_alpha (Datos_train, val_train, Datos_test, un_alpha):
lasso = linear_model.Lasso(alpha=un_alpha, fit_intercept=False, tol=0.00001, max_iter=10000000,positive=True)
lasso_fit = lasso.fit(Datos_train,val_train)
lasso_pred = lasso_fit.predict(Datos_test)
return (lasso_fit, lasso_pred)
# -
alphas = np.random.rand(9)
display(alphas)
# +
from sklearn.metrics import mean_absolute_error
alphas = [0.02, 0.3, 0.5, 0.7, 0.9, 1.5]
for un_alpha in alphas:
lasso_fit_alpha, lasso_pred_alpha = tunning_alpha(X_train, y_train, X_test, un_alpha)
lasso_mae = mean_absolute_error(y_test, lasso_pred_alpha)
lasso_mae_train = mean_absolute_error(y_train, lasso_fit_alpha.predict(X_train))
print(f"MAE Lasso (train): {lasso_mae_train:.5f}", "con alpha =", un_alpha)
print(f"MAE Lasso: {lasso_mae:.5f}", "con alpha =", un_alpha)
print("------------------------------")
# -
# Cambiando el parámetro ***alpha*** practicamente los mismos resultados
df_train_sumbit = df_train[columnas_numericas_y_precio]
df_test_submit = df_test[columnas_numericas]
lasso_pred_sumbit = linear_model.Lasso(alpha=1.5,fit_intercept=False, tol=0.00001, max_iter=10000000, positive=True)\
.fit(df_train_sumbit[columnas_numericas], df_train_sumbit['precio'])\
.predict(df_test_submit[columnas_numericas])
resultado_submit = pd.DataFrame(index=df_test.index)
resultado_submit['id'] = df_test['id']
resultado_submit['target'] = lasso_pred_sumbit
display(resultado_submit.head())
resultado_submit.to_csv('../data/sumbitLasso.csv',index=False)
# #### Score en Kaggle (features => score):
# - *['habitaciones', 'garages', 'gimnasio', 'usosmultiples', 'piscina', 'escuelascercanas', 'centroscomercialescercanos'] =>* ***1433282.27514*** con alpha = 0.1
# - *['antiguedad', 'habitaciones', 'garages', 'gimnasio', 'usosmultiples', 'piscina','escuelascercanas', 'centroscomercialescercanos', 'cantidad_amenities', 'cantidad_servicios', 'relacion_metros', 'metros_por_amenities'] =>* ***1400340.32238*** con alpha = 0.1
#
# **3.** *['antiguedad', 'habitaciones', 'garages', 'banos', 'metroscubiertos', 'metrostotales', 'gimnasio', 'usosmultiples', 'piscina', 'escuelascercanas', 'centroscomercialescercanos', 'cantidad_amenities', 'cantidad_servicios', 'relacion_metros', 'metros_por_habitacion', 'metros_por_bano', 'metros_por_amenities', 'es_avenida', 'es_Apartamento', 'es_Casa_en_condominio', 'es_Casa', 'es_Terreno', 'es_Terreno_comercial', 'es_Local_Comercial', 'es_Quinta_Vacacional', 'es_Oficina_comercial', 'es_Edificio', 'es_Casa_uso_de_suelo', 'es_Local_en_centro_comercial', 'es_Bodega_comercial', 'es_Otros', 'es_Villa', 'es_Duplex', 'es_Inmuebles_productivos_urbanos', 'es_Departamento_Compartido', 'es_Nave_industrial', 'es_Rancho', 'es_Terreno_industrial', 'es_Huerta', 'es_Lote', 'es_Hospedaje', 'es_Garage', 'seguridad_descripcion', 'moderno_descripcion', 'hermoso_descripcion', 'estado_descripcion', 'seguridad_titulo', 'moderno_titulo', 'hermoso_titulo', 'estado_titulo', 'relacion_palabras_descripcion', 'cantidad_palabras_descripcion'] =>* ***1129210.77576*** con alpha=1.5,fit_intercept=False, tol=0.00001, max_iter=10000000, positive=True
#
|
Lasso.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pybullet]
# language: python
# name: conda-env-pybullet-py
# ---
# # Simulating data with PyBullet
# Random ShapeNet objects on a ShapeNet table inside a SUNCG room...
# Below, we detail the schema for the scene description
#
# `scene_description` schema:
# * room_description (for SUNCG house/room)
# * house_id
# * room_id
# * table_description
# * mesh_filename
# * position (3D)
# * orientation (quaternion)
# * scale
# * object_descriptions (list)
# * mesh_filename (using the ShapeNet subdirectory)
# * position (3D)
# * orientation (quaternion)
# * scale of object
# ## Imports
# +
import time
import os, sys
import json
import glob
import numpy as np
import matplotlib.pyplot as plt
import cv2
# my libraries
import simulation_util as sim_util
# pybullet
import pybullet as p
import pybullet_data
# suncg
import pybullet_suncg.simulator as suncg_sim
# for reloading libraries and debugging
from importlib import reload
# -
# ## Load some SUNCG stuff
# +
suncg_dir = '/data/suncg/v1/'
# House lists
training_houses_filename = '/data/tabletop_dataset_v3/training_suncg_houses.json'
test_houses_filename = '/data/tabletop_dataset_v3/test_suncg_houses.json'
train_houses = json.load(open(training_houses_filename))
test_houses = json.load(open(test_houses_filename))
# Room types I'm considering
valid_room_types = set(['Living_Room', 'Kitchen', 'Room', 'Dining Room', 'Office'])
# Room objects to filter out
nyuv2_40_classes_filter_list = ['desk', 'chair', 'table', 'person', 'otherstructure', 'otherfurniture']
coarse_grained_classes_filter_list = ['desk', 'chair', 'table', 'person', 'computer', 'bench_chair',
'ottoman', 'storage_bench', 'pet']
# -
# ## Loading ShapeNet stuff
# +
shapenet_filepath = '/data/ShapeNetCore.v2/'
# Create a dictionary of name -> synset_id
temp = json.load(open(shapenet_filepath + 'taxonomy.json'))
taxonomy_dict = {x['name'] : x['synsetId'] for x in temp}
# weirdly, the synsets in the taxonomy file are not the same as what's in the ShapeNetCore.v2 directory. Filter this out
synsets_in_dir = os.listdir(shapenet_filepath)
synsets_in_dir.remove('taxonomy.json')
synsets_in_dir.remove('README.txt')
taxonomy_dict = {k:v for (k,v) in taxonomy_dict.items() if v in synsets_in_dir}
# useful synsets for simulation
useful_named_synsets = [
'ashcan,trash can,garbage can,wastebin,ash bin,ash-bin,ashbin,dustbin,trash barrel,trash bin',
'bag,traveling bag,travelling bag,grip,suitcase',
'birdhouse',
'bottle',
'bowl',
'camera,photographic camera',
'can,tin,tin can',
'cap',
'clock',
'computer keyboard,keypad',
'dishwasher,dish washer,dishwashing machine',
'display,video display',
'helmet',
'jar',
'knife',
'laptop,laptop computer',
'loudspeaker,speaker,speaker unit,loudspeaker system,speaker system',
'microwave,microwave oven',
'mug',
'pillow',
'printer,printing machine',
'remote control,remote',
'telephone,phone,telephone set',
'cellular telephone,cellular phone,cellphone,cell,mobile phone',
'washer,automatic washer,washing machine'
]
print("Number of synsets: {0}".format(len(useful_named_synsets)))
# List of train/test tables
training_tables_filename = '/data/tabletop_dataset_v3/training_shapenet_tables.json'
test_tables_filename = '/data/tabletop_dataset_v3/test_shapenet_tables.json'
train_tables = json.load(open(training_tables_filename))
test_tables = json.load(open(test_tables_filename))
# List of train/test object instances
training_instances_filename = '/data/tabletop_dataset_v3/training_shapenet_objects.json'
test_instances_filename = '/data/tabletop_dataset_v3/test_shapenet_objects.json'
train_models = json.load(open(training_instances_filename))
test_models = json.load(open(test_instances_filename))
# -
# debugging. only run if I made a change to SUNCG code, or need to disconnect from PyBullet
sim.disconnect()
suncg_sim = reload(suncg_sim)
# # Generate scenes
# +
simulation_params = {
'is_shapenetsem' : False,
# scene stuff
'min_num_objects_per_scene' : 10,
'max_num_objects_per_scene' : 25,
'simulation_steps' : 1000,
# House stuff
'house_ids' : train_houses, # test_houses
# room stuff
'valid_room_types' : valid_room_types,
'min_xlength' : 3.0, # Note: I believe this is in meters
'min_ylength' : 3.0,
# table stuff
'valid_tables' : train_tables, # test_tables
'max_table_height' : 1.0, # measured in meters
'min_table_height' : 0.75,
'table_init_factor' : 0.9, # this multiplicative factor limits how close you can initialize to wall
# object stuff
'object_ids' : train_models, # test_models
'max_xratio' : 1/4,
'max_yratio' : 1/4,
'max_zratio' : 1/3,
'delta' : 1.0,
# stuff
'max_initialization_tries' : 100,
# Camera/Frustum parameters
'img_width' : 640,
'img_height' : 480,
'near' : 0.01,
'far' : 100,
'fov' : 45, # vertical field of view in angles
# other camera stuff
'max_camera_rotation' : np.pi / 15., # Max rotation in radians
# other stuff
'taxonomy_dict' : taxonomy_dict,
'nyuv2_40_classes_filter_list' : nyuv2_40_classes_filter_list,
'coarse_grained_classes_filter_list' : coarse_grained_classes_filter_list,
}
# -
# Create simulator
sim = suncg_sim.Simulator(mode='gui',
suncg_data_dir_base=suncg_dir,
shapenet_data_dir_base=shapenet_filepath,
params=simulation_params,
verbose=False)
# +
# Note: This will continue trying to generate a single scene until the task is finished. If it fails,
# it will spit out why.
def keep_trying_to_generate_single_scene(sim):
try:
scenes = sim.generate_scenes(1)
print("Done")
except TimeoutError as e:
print(str(e))
keep_trying_to_generate_single_scene(sim)
except:
print("Errored out. Not due to timer, but something else...")
keep_trying_to_generate_single_scene(sim)
keep_trying_to_generate_single_scene(sim)
# -
# Debug, just to visualize stuff
scenes = sim.generate_scenes(1)
# ## Actual Data Generation
def save_img_dict(img_dict, view_num, save_dir):
# RGB
rgb_filename = save_dir + f"rgb_{view_num:05d}.jpeg"
cv2.imwrite(rgb_filename, cv2.cvtColor(img_dict['rgb'], cv2.COLOR_RGB2BGR))
# Depth
depth_filename = save_dir + f"depth_{view_num:05d}.png"
cv2.imwrite(depth_filename, sim_util.saveable_depth_image(img_dict['depth']))
# Segmentation
seg_filename = save_dir + f"segmentation_{view_num:05d}.png"
sim_util.imwrite_indexed(seg_filename, img_dict['seg'].astype(np.uint8))
# +
total_num_scenarios = 10
views_per_scene = 7
save_path = '/data/tabletop_dataset_v3/training_set/'
scene_num = 0 # start from here
while scene_num < total_num_scenarios:
# Sample scene
try:
sim = suncg_sim.Simulator(mode='gui',
suncg_data_dir_base=suncg_dir,
shapenet_data_dir_base=shapenet_filepath,
params=simulation_params,
verbose=False)
scene_description = sim.generate_scenes(1)[0]
except TimeoutError as e: # Scene took longer than 45 seconds to generate, or errored out
print(str(e))
sim.disconnect()
continue
except:
print("Errored out. Not due to timer, but something else...")
sim.disconnect()
continue
# Make directory
save_dir = save_path + f"scene_{scene_num:05d}/"
if not os.path.exists(save_dir):
os.makedirs(save_dir)
# Dictionary to save views
scene_description['views'] = {}
# Background-only view
sim.reset()
sim.load_house_room(scene_description)
valid_background_view = False
num_tries = 0
while not valid_background_view:
if num_tries > simulation_params['max_initialization_tries']:
break # this will force the entire scene to start over
num_tries += 1
# Sample the view
img_dict = sim.sample_room_view()
# Make sure it's valid. MUST have at least 2 SUNCG objects (e.g. walls/floor/fridge/couch)
unique_labels = np.unique(img_dict['orig_seg_img'])
valid_background_view = unique_labels.shape[0] >= 2
if not valid_background_view:
print("No valid background views...")
sim.disconnect()
continue
else:
save_img_dict(img_dict, 0, save_dir)
scene_description['views']['background'] = img_dict['view_params']
# Background-table view
sim.load_table(scene_description)
valid_table_view = False
num_tries = 0
while not valid_table_view:
if num_tries > simulation_params['max_initialization_tries']:
break # this will force the entire scene to start over
num_tries += 1
# Sample the view
img_dict = sim.sample_table_view()
# Make sure it's valid
unique_labels = np.unique(img_dict['seg'])
valid_table_view = 1 in unique_labels and np.count_nonzero(img_dict['seg'] == 1) > 75
if not valid_table_view:
print("No valid table views...")
sim.disconnect()
continue
else:
save_img_dict(img_dict, 1, save_dir)
scene_description['views']['background+table'] = img_dict['view_params']
# Sample background-table-object views and save
sim.load_objects(scene_description)
scene_description['views']['background+table+objects'] = []
valid_views = False
view_num = 2; num_tries = 0
while not valid_views: #view_num < views_per_scene:
if num_tries > simulation_params['max_initialization_tries']:
break # this will force the entire scene to start over
num_tries += 1
# Sample the view
img_dict = sim.sample_table_view()
# Make sure it's valid
unique_labels = np.unique(img_dict['seg'])
unique_object_labels = set(unique_labels).difference({0,1})
valid = (0 in unique_labels and # background is in view
1 in unique_labels and # table is in view
len(unique_object_labels) >= 1 # at least 1 objects in view
)
for label in unique_object_labels: # Make sure these labels are large enough
if np.count_nonzero(img_dict['seg'] == label) < 75:
valid = False
if not valid:
continue # sample another scene
### Save stuff ###
save_img_dict(img_dict, view_num, save_dir)
scene_description['views']['background+table+objects'].append(img_dict['view_params'])
# increment
view_num += 1
if view_num >= views_per_scene:
valid_views = True
if not valid_views:
print("Tried to sample view too many times...")
sim.disconnect()
continue
# Scene Description
scene_description_filename = save_dir + 'scene_description.txt'
with open(scene_description_filename, 'w') as save_file:
json.dump(scene_description, save_file)
# increment
scene_num += 1
if scene_num % 10 == 0:
print(f"Generated scene {scene_num}!")
sim.disconnect()
# -
# ## Scene saving/loading
save_filename = '/home/chrisxie/Desktop/scene_description.txt'
# +
scenes = [sim.export_scene_to_dictionary()]
# Before saving, chop of absolute filepaths
for scene in scenes:
scene['table']['mesh_filename'] = scene['table']['mesh_filename'].replace(shapenet_filepath, '')
for object_desc in scene['object_descriptions']:
object_desc['mesh_filename'] = object_desc['mesh_filename'].replace(shapenet_filepath, '')
# Serialize this JSON file
with open(save_filename, 'w') as save_file:
json.dump(scenes, save_file)
# -
# Load the json file
# scenes = json.load(open(save_filename, 'r'))
scene_description = json.load(open(save_filename, 'r'))[0]
# # Load a scene
#
# Given a scene description as I've described, load it in PyBullet
# +
# scene_description = scenes[0]
# -
sim.load_scene(scene_description)
# ## Sample camera positions
img_dict = sim.sample_table_view()
# +
# Plot the sampled camera stuff
# %matplotlib inline
plt.figure(1, figsize=(20,60))
# RGB image
plt.subplot(1,3,1)
plt.imshow(img_dict['rgb'])
plt.title('RGB')
# Depth image
plt.subplot(1,3,2)
plt.imshow(img_dict['depth'], cmap='gray')
plt.title('Linear depth')
# Segmentation image
print(np.unique(img_dict['seg']))
plt.subplot(1,3,3)
plt.imshow(img_dict['seg'])
plt.title('Segmentation')
# plt.savefig(f'/home/chrisxie/Desktop/ex{i}.png', bbox_inches='tight')
# i += 1
# -
# ### Debugging stuff
temp = p.getDebugVisualizerCamera()
camera_yaw, camera_pitch, camera_distance, camera_target = temp[8:]
print("Yaw: {0}".format(camera_yaw))
print("Pitch: {0}".format(camera_pitch))
print("Distance: {0}".format(camera_distance))
print("Target: {0}".format(camera_target))
# Set camera to look at origin
p.resetDebugVisualizerCamera(1, 0, 0, np.array([0,0,0]))
|
simulation/simulating_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test Geometries
# -----------------
# This notebook contains tests for functions to translate on C to test the se2dr code
import numpy as np
import matplotlib.pylab as plt
# +
x = np.arange(-100.0, 100.0, 0.1)
y = np.arange(-100.0, 100.0, 0.1)
X, Y = np.meshgrid(x, y)
Z = lambda theta:((-np.sin(theta)*X+np.cos(theta)*Y))
# +
fig, ax = plt.subplots(figsize=(6,6))
angle=45.0*(np.pi/180.0)
cf = ax.contourf(X,Y,Z(angle))
fig.colorbar(cf, ax=ax)
plt.plot([-100*np.cos(angle),100*np.cos(angle)],[-100*np.sin(angle),100*np.sin(angle)],"k")
plt.plot([0,-50*np.sin(angle)],[0,50*np.cos(angle)],"k")
plt.grid()
plt.show()
# -
np.sin(90)
|
PythonCodes/Utilities/TestGeometries/Test Geometries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Now we use advanced _import_ methods
#
#
# Check out directory structure
#
# Check out the contents of `/option_models/__init__.py`
#
# ```python
# from . import bsm
# from . import normal
# from . import sabr
# ```
#
# +
# when directory is imported, __init__.py is executed
# from __init__.py all sub modules are imported as well
import option_models as opt
# -
# you may call
opt.bsm.price, opt.bsm.Model
from option_models import bsm
# You can even do
#from option_models import bsm as bs
bsm.price, bsm.Model
# The objects imported different ways are all same (as class type)
bsm.Model == opt.bsm.Model
# you can create instances
bsm_model1 = bsm.Model(vol=0.2, texp=1)
bsm_model2 = opt.bsm.Model(vol=0.2, texp=1)
# But class instances are diffferent !!
bsm_model1 == bsm_model2
# ## You cal also import a particular class or functions
from option_models.bsm import Model
Model == bsm.Model
# However this is confusing. which Model?
# +
# Always better to be explicit
from option_models.bsm import Model as BsmModel
from option_models.normal import Model as NormalModel
BsmModel == bsm.Model, NormalModel == opt.normal.Model
# -
# ## And more coming..
|
py/HW3/Demo_Advanced_Import.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import time
import copy
from itertools import product
import numpy as np
import matplotlib.pyplot as plt
from cycler import cycler
color_list = ['#1f77b4','#ff7f0e','#2ca02c','#d62728','#9467bd','#8c564b','#e377c2','#7f7f7f','#bcbd22','#17becf']
marker_list = ['o','x','d','v','^','<','>']
plt.rc('axes', prop_cycle=(cycler('color',color_list)))
from LRT import LRT
from LRT import moments
from LRT import figfuns
from LRT import PT
from LRT import PCTBIN
# -
# **Setup parameter struct:**
par = LRT.ParStruct()
SAMPLE = '_p100'
# ## Load data
# +
# a. load
data = LRT.dataStruct()
data.logY = np.transpose(np.genfromtxt(f'data/logY{SAMPLE}.csv',delimiter=','))
T, par.N = data.logY.shape
# b. rescale
Y = np.exp(data.logY)
Y_lev = Y
Y = Y / np.mean(Y[0,:])
data.logY = np.log(Y)
# c. save simple data moments
mean_Y = np.ones(par.T)
mean_Y_lev = np.ones(par.T)
mean_logY = np.ones(par.T)
var_logY = np.ones(par.T)
for t in range(par.T):
mean_Y[t] = np.mean(Y[t,:])
mean_Y_lev[t] = np.mean(Y_lev[t,:])
mean_logY[t] = np.mean(data.logY[t,:])
var_logY[t] = np.var(data.logY[t,:])
np.savetxt("data/mean_Y.txt",mean_Y,delimiter=',',fmt='%f',newline='\r\n')
np.savetxt("data/mean_Y_lev.txt",mean_Y_lev,delimiter=',',fmt='%f',newline='\r\n')
np.savetxt("data/mean_logY.txt",mean_logY,delimiter=',',fmt='%f',newline='\r\n')
np.savetxt("data/var_logY.txt",var_logY,delimiter=',',fmt='%f',newline='\r\n')
# d. upgrade simulation for large sample
par.simN = max([par.N,par.simN])
# -
# ## Histograms
ages = np.arange(par.agemin, par.agemax+1)
# **Histograms** of income **levels**.
pcts = np.percentile(data.logY.flatten(), q=[0.1, 99.9])
for i,t in enumerate([0,15,29]):
fig,ax = figfuns.new()
ax.hist(data.logY[t,:],bins=300)
ax.set_ylabel('observations')
ax.set_xlabel('log income ($y_{it}$)')
ax.set_xlim(pcts)
figfuns.save(fig,ax,name=f'hist_inc_level_age{ages[t]}')
plt.close(fig)
# **Histograms** of income **growth**
growth = data.logY[1:, :] - data.logY[:-1, :]
pcts = np.percentile(growth.flatten(), q=[0.1, 99.9])
for i,t in enumerate([0,14,28]):
fig,ax = figfuns.new()
ax.hist(growth[t,:], bins=500)
ax.set_ylabel('observations')
ax.set_xlabel('log income differences ($y_{i'+str(t+1)+'} - y_{i'+str(t)+'}$)')
ax.set_xlim(pcts)
figfuns.save(fig, ax, name=f'hist_inc_growth_age{ages[t+1]}')
plt.close(fig)
# ## Load ABB
# +
# a. load abb
data_abbsim = LRT.dataStruct()
data_abbsim.logY = np.transpose(np.genfromtxt('data/abb_sim.csv',delimiter=','))
# b. setup
par_abb = LRT.ParStruct()
par_abb.T, par_abb.N = data_abbsim.logY.shape
# c. normalize
Y = np.exp(data_abbsim.logY)
Y = Y / np.mean(Y[0,:])
data_abbsim.logY = np.log(Y)
# d. add model details
model_abb = LRT.modelStruct()
model_abb.data = data_abbsim
model_abb.type = 'ABB'
model_abb.name = 'ABB'
model_abb.name_short = 'ABB'
model_abb.par = copy.deepcopy(par)
# -
# # Estimate and simulate
# ## LRT
# +
models_LRT = []
for depth,name_short,name in zip([7,10],
['LRT_depth_7','LRT_depth_10'],
['LRT (depth 7)','LRT (depth 10)']):
# a. settings
par.k = 5
par.k_lead = 30
par.depth = depth
# b. estimate
model = LRT.estimate(par,data,name)
model.name_short = name_short
models_LRT.append(model)
prefmodel = models_LRT[-1]
# -
# **Save:**
# +
# a. number of leafs
np.savetxt("data/num_leafs.txt",prefmodel.num_leafs,delimiter=',',fmt='%d',newline='\r\n')
# b. initial grouping
prob_G_ini = np.ones(prefmodel.num_leafs[0])
for i in range(prefmodel.num_leafs[0]):
I = (prefmodel.G[0] == prefmodel.uniqueG[0][i])
prob_G_ini[i] = I.sum()/par.N
np.savetxt("data/prob_G_ini.txt",prob_G_ini,delimiter=',',fmt='%f',newline='\r\n')
# c. prediction and transitions
for t in range(0,par.T):
# i. save ypred
filename = "data/ypred_G_t{:d}.txt".format(t)
np.savetxt(filename, prefmodel.ypred_G[t],delimiter=',',fmt='%f',newline='\r\n')
# ii. distribution over groups
maxG = np.amax(prefmodel.uniqueG[t])
count = np.bincount(prefmodel.G[t], minlength=maxG+1)
minnum = np.min(count[prefmodel.uniqueG[t]])
p50num = np.int(np.median(count[prefmodel.uniqueG[t]]))
maxnum = np.max(count[prefmodel.uniqueG[t]])
if t == 0:
continue
# iii. save trans
trans = np.copy(prefmodel.trans_obs[t])
filename = "data/trans_t{:d}.txt".format(t)
np.savetxt(filename,trans,delimiter=',',fmt='%d',newline='\r\n')
# -
# **Simulate:**
for model in models_LRT:
model.data = LRT.simulate(model.par, model, data)
# ## PT
# +
# a. estimates
model_PT = PT.estimate(par,data)
# b. simulate
model_PT.data = PT.simulate(model_PT.par, model_PT, data)
# c. save
np.savetxt("output/PT_estimates.txt",np.array([model_PT.sigma_psi,model_PT.sigma_xi,model_PT.P_ini_std]),
delimiter=',',fmt='%f',newline='\r\n')
# -
# ## PCTBIN
# +
# a. estimate
model_PCTBIN = PCTBIN.estimate(par,data, num_bins=20)
# b. simulate
model_PCTBIN.data = PCTBIN.simulate(model_PCTBIN,data)
# -
# ## List of models
# +
models = [models_LRT[0],models_LRT[1],model_PT,model_PCTBIN,model_abb]
# colors and markers
for i,model in enumerate(models):
model.color = color_list[i]
model.marker = marker_list[i]
# -
# ## Group means
for model in models_LRT + [model_PCTBIN]:
fig, ax = figfuns.new()
ages = np.arange(par.agemin, par.agemax+1)
for t in range(par.T):
y = model.ypred_G[t]
age = ages[t]*np.random.normal(loc=1.0,scale=0.0,size=y.shape)
ax.plot(age, y, linestyle='', marker='o',markersize=2,color=model.color)
ax.set_ylabel('log income ($\mu_{gt}$)')
ax.set_xlabel('age')
figfuns.save(fig,ax,f'group_means_of_income_{model.name_short}')
plt.close()
# ## Transition rates
# +
def censor_trans_mat(trans_mat, trans_obs):
Y = copy.deepcopy(trans_mat)
I = (trans_obs<=5) & (trans_obs>0)
Y = Y**0.75
return Y
def plot_trans_mat(model,t):
fig,ax = figfuns.new()
ax.grid(False)
Y = censor_trans_mat(model.trans[t], model.trans_obs[t])
ax.imshow(Y, cmap='hot_r', interpolation='nearest')
ax.set_xlabel(f'groups $t={t+1}$')
ax.set_ylabel(f'groups $t={t}$')
ax.set_yticks([])
ax.set_xticks([])
ax.set_yticklabels([])
fig.tight_layout()
figfuns.save(fig,ax,name=f'trans_mat_{model.name_short}')
plt.close()
# -
t = 10
for model in [models_LRT[0],models_LRT[1],model_PCTBIN]:
plot_trans_mat(model,10)
# # Moments
moments.calc_all(par,data,printprogress=True)
for model in models:
print(model.name)
moments.calc_all(model.par,model.data,printprogress=True)
# # Figures
# ## Levels (life-cycle profile)
figfuns.age_profile(par, data, models, 0, varname_str='logY', prefix='compare')
# ## Changes (life-cycle profile)
figfuns.age_profile(par, data, models, 1,
varname_str='dlogY', prefix='compare')
# ## 5-year changes (life-cycle profile)
figfuns.age_profile(par, data, models, 5,
varname_str='dlogY', prefix='compare')
# ## 10-year changes (life-cycle profile)
figfuns.age_profile(par, data, models, 10,
varname_str='dlogY', prefix='compare')
# ## Changes (over recent income and age-groups)
for model in models:
k = 1
for mom_str in ['mean','var','skew','kurt']:
figfuns.REpercs_profile(par, data, model, k, mom_str,
varname_str='dlogY',
age_grp_str='detail',
perc_grp_str='RE', modelname= model.name_short)
# ## 5-year changes (over RE and age-groups)
for model in models:
k = 5
for mom_str in ['mean','var','skew','kurt']:
figfuns.REpercs_profile(par, data, model, k, mom_str,
varname_str='dlogY',
age_grp_str='detail',
perc_grp_str='RE', modelname=model.name_short)
# ## 10-year changes (over RE and age-groups)
for model in models:
k = 10
for mom_str in ['mean','var','skew','kurt']:
figfuns.REpercs_profile(par, data, model, k, mom_str,
varname_str='dlogY',
age_grp_str='detail',
perc_grp_str='RE', modelname=model.name_short)
# ## Covariances over age
for model in models:
prefix = model.name_short
figfuns.cov(par, data, model, prefix=prefix)
# ## Autocorrelations
for model in models:
figfuns.autocorr(par, data, model, k_list=[1,2,5],
varname_str='dlogY',
modelname=model.name_short)
for model in models:
figfuns.autocorr(par, data, model, k_list=[1,2,5],
varname_str='logY',
modelname=model.name_short)
# ## Covariances of levels and future growth
for model in models:
figfuns.cov_YdY(par,data,model,prefix=model.name_short)
# ## Heterogeneity
figfuns.heterogenous(par, data, models,
'dlogY', 0, 'life-time growth (demeaned)',
prefix='compare', bounds=(-1.5,1.5))
figfuns.heterogenous(par, data, models,
'autocorr', 1, 'first-order autocorr.',
prefix='compare', bounds=(-1,1))
figfuns.heterogenous(par, data, models,
'autocorr_level', 1, 'first-order autocorr. (levels)',
prefix='compare', bounds=(-.25,1))
figfuns.heterogenous(par, data, models,
'std_dlogY', 1, 'std. of income growth',
prefix='compare', bounds=(0,0.5))
# ## Compare fit across models for single age group over RE percs
for k,iage in product([1,5],[0,4]):
mom_str = 'mean'
figfuns.REpercs_profile_compare_models(par,data,models,k,mom_str,
varname_str='dlogY',age_grp_str='detail',perc_grp_str='RE',iage=iage,
prefix='compare')
# ## Within vs. between
for mom in [2,3,4]:
figfuns.moments_within_groups(data, models[1], marker_list, color_list,
central_moment=mom, kk=[1,5,10],
prefix=models[1].name_short)
|
2_1_main_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building your Recurrent Neural Network - Step by Step
#
# Welcome to Course 5's first assignment! In this assignment, you will implement key components of a Recurrent Neural Network in numpy.
#
# Recurrent Neural Networks (RNN) are very effective for Natural Language Processing and other sequence tasks because they have "memory". They can read inputs $x^{\langle t \rangle}$ (such as words) one at a time, and remember some information/context through the hidden layer activations that get passed from one time-step to the next. This allows a unidirectional RNN to take information from the past to process later inputs. A bidirectional RNN can take context from both the past and the future.
#
# **Notation**:
# - Superscript $[l]$ denotes an object associated with the $l^{th}$ layer.
#
# - Superscript $(i)$ denotes an object associated with the $i^{th}$ example.
#
# - Superscript $\langle t \rangle$ denotes an object at the $t^{th}$ time-step.
#
# - **Sub**script $i$ denotes the $i^{th}$ entry of a vector.
#
# Example:
# - $a^{(2)[3]<4>}_5$ denotes the activation of the 2nd training example (2), 3rd layer [3], 4th time step <4>, and 5th entry in the vector.
#
# #### Pre-requisites
# * We assume that you are already familiar with `numpy`.
# * To refresh your knowledge of numpy, you can review course 1 of this specialization "Neural Networks and Deep Learning".
# * Specifically, review the week 2 assignment ["Python Basics with numpy (optional)"](https://www.coursera.org/learn/neural-networks-deep-learning/item/Zh0CU).
#
#
# #### Be careful when modifying the starter code
# * When working on graded functions, please remember to only modify the code that is between the
# ```Python
# #### START CODE HERE
# ```
# and
# ```Python
# #### END CODE HERE
# ```
# * In particular, Be careful to not modify the first line of graded routines. These start with:
# ```Python
# # GRADED FUNCTION: routine_name
# ```
# * The automatic grader (autograder) needs these to locate the function.
# * Even a change in spacing will cause issues with the autograder.
# * It will return 'failed' if these are modified or missing."
# ## <font color='darkblue'>Updates for 3a</font>
#
# #### If you were working on the notebook before this update...
# * The current notebook is version "3a".
# * You can find your original work saved in the notebook with the previous version name ("v3")
# * To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#
# #### List of updates
#
# * "Forward propagation for the basic RNN", added sections to clarify variable names and shapes:
# - "Dimensions of $x^{\langle t \rangle}$"
# - "Hidden State $a$",
# - "Dimensions of hidden state $a^{\langle t \rangle}$"
# - "Dimensions of prediction $y^{\langle t \rangle}$"
# * `rnn_cell_forward`:
# * Added additional hints.
# * Updated figure 2.
# * `rnn_forward`
# - Set `xt` in a separate line of code to clarify what code is expected; added additional hints.
# - Clarifies instructions to specify dimensions (2D or 3D), and clarifies variable names.
# - Additional Hints
# - Clarifies when the basic RNN works well.
# - Updated figure 3.
# * "About the gates" replaced with "overview of gates and states":
# - Updated to include conceptual description of each gate's purpose, and an explanation of each equation.
# - Added sections about the cell state, hidden state, and prediction.
# - Lists variable names that are used in the code, and notes when they differ from the variables used in the equations.
# - Lists shapes of the variables.
# - Updated figure 4.
# * `lstm_forward`
# - Added instructions, noting the shapes of the variables.
# - Added hints about `c` and `c_next` to help students avoid copy-by-reference mistakes.
# - Set `xt` in a separate line to make this step explicit.
# * `dimension`
# - clarified use of $x^{(i)\langle t \rangle}$ in dimension description
# * `Backward description`
# - Updated figure 6 and 7
# * `rnn_cell_backward`
# - changed 'dtanh' to 'dz' to avoid confusion with the output of the tanh
# - fixed naming of 'b' in test section, changed to 'ba', updated results
# * `rnn_backward`
# - modified instructions in comments to sum da and gradients from previous timesteps
# * `lstm_cell_backward`
# - updated equations and description to fix errors.
# - Added Figure 8
# - modified equations to use the term dlower_case_gamma vs dGamma - the previous naming confused the location of the gate derivative.
# - removed redundant lines for equations 7-10, changed equation numbers to match new equation numbers
# * `lstm_backward`
# - removed comment listing dc as argument.
# - added da_prevt and dc_prevt to for loop and recalculated results
# - in test, added "Wy" and "by" zero fill as it is required for lstm_forward.
#
# * Renamed global variables so that they do not conflict with local variables within the function.
# * Spelling, grammar and wording corrections.
# * For unit tests, updated print statements and "expected output" for easier comparisons.
# * Many thanks to mentors and students for suggested improvements and fixes in the assignments for course 5!
# Let's first import all the packages that you will need during this assignment.
import numpy as np
from rnn_utils import *
# ## 1 - Forward propagation for the basic Recurrent Neural Network
#
# Later this week, you will generate music using an RNN. The basic RNN that you will implement has the structure below. In this example, $T_x = T_y$.
# <img src="images/RNN.png" style="width:500;height:300px;">
# <caption><center> **Figure 1**: Basic RNN model </center></caption>
# ### Dimensions of input $x$
#
# #### Input with $n_x$ number of units
# * For a single timestep of a single input example, $x^{(i) \langle t \rangle }$ is a one-dimensional input vector.
# * Using language as an example, a language with a 5000 word vocabulary could be one-hot encoded into a vector that has 5000 units. So $x^{(i)\langle t \rangle}$ would have the shape (5000,).
# * We'll use the notation $n_x$ to denote the number of units in a single timestep of a single training example.
# #### Time steps of size $T_{x}$
# * A recurrent neural network has multiple time steps, which we'll index with $t$.
# * In the lessons, we saw a single training example $x^{(i)}$ consist of multiple time steps $T_x$. For example, if there are 10 time steps, $T_{x} = 10$
# #### Batches of size $m$
# * Let's say we have mini-batches, each with 20 training examples.
# * To benefit from vectorization, we'll stack 20 columns of $x^{(i)}$ examples.
# * For example, this tensor has the shape (5000,20,10).
# * We'll use $m$ to denote the number of training examples.
# * So the shape of a mini-batch is $(n_x,m,T_x)$
# #### 3D Tensor of shape $(n_{x},m,T_{x})$
# * The 3-dimensional tensor $x$ of shape $(n_x,m,T_x)$ represents the input $x$ that is fed into the RNN.
#
# #### Taking a 2D slice for each time step: $x^{\langle t \rangle}$
# * At each time step, we'll use a mini-batches of training examples (not just a single example).
# * So, for each time step $t$, we'll use a 2D slice of shape $(n_x,m)$.
# * We're referring to this 2D slice as $x^{\langle t \rangle}$. The variable name in the code is `xt`.
# ### Definition of hidden state $a$
#
# * The activation $a^{\langle t \rangle}$ that is passed to the RNN from one time step to another is called a "hidden state."
#
# ### Dimensions of hidden state $a$
#
# * Similar to the input tensor $x$, the hidden state for a single training example is a vector of length $n_{a}$.
# * If we include a mini-batch of $m$ training examples, the shape of a mini-batch is $(n_{a},m)$.
# * When we include the time step dimension, the shape of the hidden state is $(n_{a}, m, T_x)$
# * We will loop through the time steps with index $t$, and work with a 2D slice of the 3D tensor.
# * We'll refer to this 2D slice as $a^{\langle t \rangle}$.
# * In the code, the variable names we use are either `a_prev` or `a_next`, depending on the function that's being implemented.
# * The shape of this 2D slice is $(n_{a}, m)$
# ### Dimensions of prediction $\hat{y}$
# * Similar to the inputs and hidden states, $\hat{y}$ is a 3D tensor of shape $(n_{y}, m, T_{y})$.
# * $n_{y}$: number of units in the vector representing the prediction.
# * $m$: number of examples in a mini-batch.
# * $T_{y}$: number of time steps in the prediction.
# * For a single time step $t$, a 2D slice $\hat{y}^{\langle t \rangle}$ has shape $(n_{y}, m)$.
# * In the code, the variable names are:
# - `y_pred`: $\hat{y}$
# - `yt_pred`: $\hat{y}^{\langle t \rangle}$
# Here's how you can implement an RNN:
#
# **Steps**:
# 1. Implement the calculations needed for one time-step of the RNN.
# 2. Implement a loop over $T_x$ time-steps in order to process all the inputs, one at a time.
# ## 1.1 - RNN cell
#
# A recurrent neural network can be seen as the repeated use of a single cell. You are first going to implement the computations for a single time-step. The following figure describes the operations for a single time-step of an RNN cell.
#
# <img src="images/rnn_step_forward_figure2_v3a.png" style="width:700px;height:300px;">
# <caption><center> **Figure 2**: Basic RNN cell. Takes as input $x^{\langle t \rangle}$ (current input) and $a^{\langle t - 1\rangle}$ (previous hidden state containing information from the past), and outputs $a^{\langle t \rangle}$ which is given to the next RNN cell and also used to predict $\hat{y}^{\langle t \rangle}$ </center></caption>
#
# #### rnn cell versus rnn_cell_forward
# * Note that an RNN cell outputs the hidden state $a^{\langle t \rangle}$.
# * The rnn cell is shown in the figure as the inner box which has solid lines.
# * The function that we will implement, `rnn_cell_forward`, also calculates the prediction $\hat{y}^{\langle t \rangle}$
# * The rnn_cell_forward is shown in the figure as the outer box that has dashed lines.
# **Exercise**: Implement the RNN-cell described in Figure (2).
#
# **Instructions**:
# 1. Compute the hidden state with tanh activation: $a^{\langle t \rangle} = \tanh(W_{aa} a^{\langle t-1 \rangle} + W_{ax} x^{\langle t \rangle} + b_a)$.
# 2. Using your new hidden state $a^{\langle t \rangle}$, compute the prediction $\hat{y}^{\langle t \rangle} = softmax(W_{ya} a^{\langle t \rangle} + b_y)$. We provided the function `softmax`.
# 3. Store $(a^{\langle t \rangle}, a^{\langle t-1 \rangle}, x^{\langle t \rangle}, parameters)$ in a `cache`.
# 4. Return $a^{\langle t \rangle}$ , $\hat{y}^{\langle t \rangle}$ and `cache`
#
# #### Additional Hints
# * [numpy.tanh](https://www.google.com/search?q=numpy+tanh&rlz=1C5CHFA_enUS854US855&oq=numpy+tanh&aqs=chrome..69i57j0l5.1340j0j7&sourceid=chrome&ie=UTF-8)
# * We've created a `softmax` function that you can use. It is located in the file 'rnn_utils.py' and has been imported.
# * For matrix multiplication, use [numpy.dot](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html)
#
#
# +
# GRADED FUNCTION: rnn_cell_forward
def rnn_cell_forward(xt, a_prev, parameters):
"""
Implements a single forward step of the RNN-cell as described in Figure (2)
Arguments:
xt -- your input data at timestep "t", numpy array of shape (n_x, m).
a_prev -- Hidden state at timestep "t-1", numpy array of shape (n_a, m)
parameters -- python dictionary containing:
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
ba -- Bias, numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
Returns:
a_next -- next hidden state, of shape (n_a, m)
yt_pred -- prediction at timestep "t", numpy array of shape (n_y, m)
cache -- tuple of values needed for the backward pass, contains (a_next, a_prev, xt, parameters)
"""
# Retrieve parameters from "parameters"
Wax = parameters["Wax"]
Waa = parameters["Waa"]
Wya = parameters["Wya"]
ba = parameters["ba"]
by = parameters["by"]
### START CODE HERE ### (≈2 lines)
# compute next activation state using the formula given above
a_next = np.tanh(np.dot(Waa,a_prev)+np.dot(Wax,xt)+ba)
# compute output of the current cell using the formula given above
yt_pred = softmax(np.dot(Wya,a_next)+by)
### END CODE HERE ###
# store values you need for backward propagation in cache
cache = (a_next, a_prev, xt, parameters)
return a_next, yt_pred, cache
# +
np.random.seed(1)
xt_tmp = np.random.randn(3,10)
a_prev_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Waa'] = np.random.randn(5,5)
parameters_tmp['Wax'] = np.random.randn(5,3)
parameters_tmp['Wya'] = np.random.randn(2,5)
parameters_tmp['ba'] = np.random.randn(5,1)
parameters_tmp['by'] = np.random.randn(2,1)
a_next_tmp, yt_pred_tmp, cache_tmp = rnn_cell_forward(xt_tmp, a_prev_tmp, parameters_tmp)
print("a_next[4] = \n", a_next_tmp[4])
print("a_next.shape = \n", a_next_tmp.shape)
print("yt_pred[1] =\n", yt_pred_tmp[1])
print("yt_pred.shape = \n", yt_pred_tmp.shape)
# -
# **Expected Output**:
# ```Python
# a_next[4] =
# [ 0.59584544 0.18141802 0.61311866 0.99808218 0.85016201 0.99980978
# -0.18887155 0.99815551 0.6531151 0.82872037]
# a_next.shape =
# (5, 10)
# yt_pred[1] =
# [ 0.9888161 0.01682021 0.21140899 0.36817467 0.98988387 0.88945212
# 0.36920224 0.9966312 0.9982559 0.17746526]
# yt_pred.shape =
# (2, 10)
#
# ```
# ## 1.2 - RNN forward pass
#
# - A recurrent neural network (RNN) is a repetition of the RNN cell that you've just built.
# - If your input sequence of data is 10 time steps long, then you will re-use the RNN cell 10 times.
# - Each cell takes two inputs at each time step:
# - $a^{\langle t-1 \rangle}$: The hidden state from the previous cell.
# - $x^{\langle t \rangle}$: The current time-step's input data.
# - It has two outputs at each time step:
# - A hidden state ($a^{\langle t \rangle}$)
# - A prediction ($y^{\langle t \rangle}$)
# - The weights and biases $(W_{aa}, b_{a}, W_{ax}, b_{x})$ are re-used each time step.
# - They are maintained between calls to rnn_cell_forward in the 'parameters' dictionary.
#
#
# <img src="images/rnn_forward_sequence_figure3_v3a.png" style="width:800px;height:180px;">
# <caption><center> **Figure 3**: Basic RNN. The input sequence $x = (x^{\langle 1 \rangle}, x^{\langle 2 \rangle}, ..., x^{\langle T_x \rangle})$ is carried over $T_x$ time steps. The network outputs $y = (y^{\langle 1 \rangle}, y^{\langle 2 \rangle}, ..., y^{\langle T_x \rangle})$. </center></caption>
#
# **Exercise**: Code the forward propagation of the RNN described in Figure (3).
#
# **Instructions**:
# * Create a 3D array of zeros, $a$ of shape $(n_{a}, m, T_{x})$ that will store all the hidden states computed by the RNN.
# * Create a 3D array of zeros, $\hat{y}$, of shape $(n_{y}, m, T_{x})$ that will store the predictions.
# - Note that in this case, $T_{y} = T_{x}$ (the prediction and input have the same number of time steps).
# * Initialize the 2D hidden state `a_next` by setting it equal to the initial hidden state, $a_{0}$.
# * At each time step $t$:
# - Get $x^{\langle t \rangle}$, which is a 2D slice of $x$ for a single time step $t$.
# - $x^{\langle t \rangle}$ has shape $(n_{x}, m)$
# - $x$ has shape $(n_{x}, m, T_{x})$
# - Update the 2D hidden state $a^{\langle t \rangle}$ (variable name `a_next`), the prediction $\hat{y}^{\langle t \rangle}$ and the cache by running `rnn_cell_forward`.
# - $a^{\langle t \rangle}$ has shape $(n_{a}, m)$
# - Store the 2D hidden state in the 3D tensor $a$, at the $t^{th}$ position.
# - $a$ has shape $(n_{a}, m, T_{x})$
# - Store the 2D $\hat{y}^{\langle t \rangle}$ prediction (variable name `yt_pred`) in the 3D tensor $\hat{y}_{pred}$ at the $t^{th}$ position.
# - $\hat{y}^{\langle t \rangle}$ has shape $(n_{y}, m)$
# - $\hat{y}$ has shape $(n_{y}, m, T_x)$
# - Append the cache to the list of caches.
# * Return the 3D tensor $a$ and $\hat{y}$, as well as the list of caches.
#
# #### Additional Hints
# - [np.zeros](https://docs.scipy.org/doc/numpy/reference/generated/numpy.zeros.html)
# - If you have a 3 dimensional numpy array and are indexing by its third dimension, you can use array slicing like this: `var_name[:,:,i]`.
# +
# GRADED FUNCTION: rnn_forward
def rnn_forward(x, a0, parameters):
"""
Implement the forward propagation of the recurrent neural network described in Figure (3).
Arguments:
x -- Input data for every time-step, of shape (n_x, m, T_x).
a0 -- Initial hidden state, of shape (n_a, m)
parameters -- python dictionary containing:
Waa -- Weight matrix multiplying the hidden state, numpy array of shape (n_a, n_a)
Wax -- Weight matrix multiplying the input, numpy array of shape (n_a, n_x)
Wya -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
ba -- Bias numpy array of shape (n_a, 1)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
Returns:
a -- Hidden states for every time-step, numpy array of shape (n_a, m, T_x)
y_pred -- Predictions for every time-step, numpy array of shape (n_y, m, T_x)
caches -- tuple of values needed for the backward pass, contains (list of caches, x)
"""
# Initialize "caches" which will contain the list of all caches
caches = []
# Retrieve dimensions from shapes of x and parameters["Wya"]
n_x, m, T_x = x.shape
n_y, n_a = parameters["Wya"].shape
### START CODE HERE ###
# initialize "a" and "y_pred" with zeros (≈2 lines)
a = np.zeros((n_a,m,T_x))
y_pred = np.zeros((n_y,m,T_x))
# Initialize a_next (≈1 line)
a_next = a0
# loop over all time-steps of the input 'x' (1 line)
for t in range(T_x):
# Update next hidden state, compute the prediction, get the cache (≈2 lines)
xt = x[:,:,t]
a_next, yt_pred, cache = rnn_cell_forward(xt, a_next, parameters)
# Save the value of the new "next" hidden state in a (≈1 line)
a[:,:,t] = a_next
# Save the value of the prediction in y (≈1 line)
y_pred[:,:,t] = yt_pred
# Append "cache" to "caches" (≈1 line)
np.append(caches,cache)
### END CODE HERE ###
# store values needed for backward propagation in cache
caches = (caches, x)
return a, y_pred, caches
# +
np.random.seed(1)
x_tmp = np.random.randn(3,10,4)
a0_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Waa'] = np.random.randn(5,5)
parameters_tmp['Wax'] = np.random.randn(5,3)
parameters_tmp['Wya'] = np.random.randn(2,5)
parameters_tmp['ba'] = np.random.randn(5,1)
parameters_tmp['by'] = np.random.randn(2,1)
a_tmp, y_pred_tmp, caches_tmp = rnn_forward(x_tmp, a0_tmp, parameters_tmp)
print("a[4][1] = \n", a_tmp[4][1])
print("a.shape = \n", a_tmp.shape)
print("y_pred[1][3] =\n", y_pred_tmp[1][3])
print("y_pred.shape = \n", y_pred_tmp.shape)
print("caches[1][1][3] =\n", caches_tmp[1][1][3])
print("len(caches) = \n", len(caches_tmp))
# -
# **Expected Output**:
#
# ```Python
# a[4][1] =
# [-0.99999375 0.77911235 -0.99861469 -0.99833267]
# a.shape =
# (5, 10, 4)
# y_pred[1][3] =
# [ 0.79560373 0.86224861 0.11118257 0.81515947]
# y_pred.shape =
# (2, 10, 4)
# caches[1][1][3] =
# [-1.1425182 -0.34934272 -0.20889423 0.58662319]
# len(caches) =
# 2
# ```
# Congratulations! You've successfully built the forward propagation of a recurrent neural network from scratch.
#
# #### Situations when this RNN will perform better:
# - This will work well enough for some applications, but it suffers from the vanishing gradient problems.
# - The RNN works best when each output $\hat{y}^{\langle t \rangle}$ can be estimated using "local" context.
# - "Local" context refers to information that is close to the prediction's time step $t$.
# - More formally, local context refers to inputs $x^{\langle t' \rangle}$ and predictions $\hat{y}^{\langle t \rangle}$ where $t'$ is close to $t$.
#
# In the next part, you will build a more complex LSTM model, which is better at addressing vanishing gradients. The LSTM will be better able to remember a piece of information and keep it saved for many timesteps.
# ## 2 - Long Short-Term Memory (LSTM) network
#
# The following figure shows the operations of an LSTM-cell.
#
# <img src="images/LSTM_figure4_v3a.png" style="width:500;height:400px;">
# <caption><center> **Figure 4**: LSTM-cell. This tracks and updates a "cell state" or memory variable $c^{\langle t \rangle}$ at every time-step, which can be different from $a^{\langle t \rangle}$. </center></caption>
#
# Similar to the RNN example above, you will start by implementing the LSTM cell for a single time-step. Then you can iteratively call it from inside a "for-loop" to have it process an input with $T_x$ time-steps.
# ### Overview of gates and states
#
# #### - Forget gate $\mathbf{\Gamma}_{f}$
#
# * Let's assume we are reading words in a piece of text, and plan to use an LSTM to keep track of grammatical structures, such as whether the subject is singular ("puppy") or plural ("puppies").
# * If the subject changes its state (from a singular word to a plural word), the memory of the previous state becomes outdated, so we "forget" that outdated state.
# * The "forget gate" is a tensor containing values that are between 0 and 1.
# * If a unit in the forget gate has a value close to 0, the LSTM will "forget" the stored state in the corresponding unit of the previous cell state.
# * If a unit in the forget gate has a value close to 1, the LSTM will mostly remember the corresponding value in the stored state.
#
# ##### Equation
#
# $$\mathbf{\Gamma}_f^{\langle t \rangle} = \sigma(\mathbf{W}_f[\mathbf{a}^{\langle t-1 \rangle}, \mathbf{x}^{\langle t \rangle}] + \mathbf{b}_f)\tag{1} $$
#
# ##### Explanation of the equation:
#
# * $\mathbf{W_{f}}$ contains weights that govern the forget gate's behavior.
# * The previous time step's hidden state $[a^{\langle t-1 \rangle}$ and current time step's input $x^{\langle t \rangle}]$ are concatenated together and multiplied by $\mathbf{W_{f}}$.
# * A sigmoid function is used to make each of the gate tensor's values $\mathbf{\Gamma}_f^{\langle t \rangle}$ range from 0 to 1.
# * The forget gate $\mathbf{\Gamma}_f^{\langle t \rangle}$ has the same dimensions as the previous cell state $c^{\langle t-1 \rangle}$.
# * This means that the two can be multiplied together, element-wise.
# * Multiplying the tensors $\mathbf{\Gamma}_f^{\langle t \rangle} * \mathbf{c}^{\langle t-1 \rangle}$ is like applying a mask over the previous cell state.
# * If a single value in $\mathbf{\Gamma}_f^{\langle t \rangle}$ is 0 or close to 0, then the product is close to 0.
# * This keeps the information stored in the corresponding unit in $\mathbf{c}^{\langle t-1 \rangle}$ from being remembered for the next time step.
# * Similarly, if one value is close to 1, the product is close to the original value in the previous cell state.
# * The LSTM will keep the information from the corresponding unit of $\mathbf{c}^{\langle t-1 \rangle}$, to be used in the next time step.
#
# ##### Variable names in the code
# The variable names in the code are similar to the equations, with slight differences.
# * `Wf`: forget gate weight $\mathbf{W}_{f}$
# * `Wb`: forget gate bias $\mathbf{W}_{b}$
# * `ft`: forget gate $\Gamma_f^{\langle t \rangle}$
# #### Candidate value $\tilde{\mathbf{c}}^{\langle t \rangle}$
# * The candidate value is a tensor containing information from the current time step that **may** be stored in the current cell state $\mathbf{c}^{\langle t \rangle}$.
# * Which parts of the candidate value get passed on depends on the update gate.
# * The candidate value is a tensor containing values that range from -1 to 1.
# * The tilde "~" is used to differentiate the candidate $\tilde{\mathbf{c}}^{\langle t \rangle}$ from the cell state $\mathbf{c}^{\langle t \rangle}$.
#
# ##### Equation
# $$\mathbf{\tilde{c}}^{\langle t \rangle} = \tanh\left( \mathbf{W}_{c} [\mathbf{a}^{\langle t - 1 \rangle}, \mathbf{x}^{\langle t \rangle}] + \mathbf{b}_{c} \right) \tag{3}$$
#
# ##### Explanation of the equation
# * The 'tanh' function produces values between -1 and +1.
#
#
# ##### Variable names in the code
# * `cct`: candidate value $\mathbf{\tilde{c}}^{\langle t \rangle}$
# #### - Update gate $\mathbf{\Gamma}_{i}$
#
# * We use the update gate to decide what aspects of the candidate $\tilde{\mathbf{c}}^{\langle t \rangle}$ to add to the cell state $c^{\langle t \rangle}$.
# * The update gate decides what parts of a "candidate" tensor $\tilde{\mathbf{c}}^{\langle t \rangle}$ are passed onto the cell state $\mathbf{c}^{\langle t \rangle}$.
# * The update gate is a tensor containing values between 0 and 1.
# * When a unit in the update gate is close to 1, it allows the value of the candidate $\tilde{\mathbf{c}}^{\langle t \rangle}$ to be passed onto the hidden state $\mathbf{c}^{\langle t \rangle}$
# * When a unit in the update gate is close to 0, it prevents the corresponding value in the candidate from being passed onto the hidden state.
# * Notice that we use the subscript "i" and not "u", to follow the convention used in the literature.
#
# ##### Equation
#
# $$\mathbf{\Gamma}_i^{\langle t \rangle} = \sigma(\mathbf{W}_i[a^{\langle t-1 \rangle}, \mathbf{x}^{\langle t \rangle}] + \mathbf{b}_i)\tag{2} $$
#
# ##### Explanation of the equation
#
# * Similar to the forget gate, here $\mathbf{\Gamma}_i^{\langle t \rangle}$, the sigmoid produces values between 0 and 1.
# * The update gate is multiplied element-wise with the candidate, and this product ($\mathbf{\Gamma}_{i}^{\langle t \rangle} * \tilde{c}^{\langle t \rangle}$) is used in determining the cell state $\mathbf{c}^{\langle t \rangle}$.
#
# ##### Variable names in code (Please note that they're different than the equations)
# In the code, we'll use the variable names found in the academic literature. These variables don't use "u" to denote "update".
# * `Wi` is the update gate weight $\mathbf{W}_i$ (not "Wu")
# * `bi` is the update gate bias $\mathbf{b}_i$ (not "bu")
# * `it` is the forget gate $\mathbf{\Gamma}_i^{\langle t \rangle}$ (not "ut")
# #### - Cell state $\mathbf{c}^{\langle t \rangle}$
#
# * The cell state is the "memory" that gets passed onto future time steps.
# * The new cell state $\mathbf{c}^{\langle t \rangle}$ is a combination of the previous cell state and the candidate value.
#
# ##### Equation
#
# $$ \mathbf{c}^{\langle t \rangle} = \mathbf{\Gamma}_f^{\langle t \rangle}* \mathbf{c}^{\langle t-1 \rangle} + \mathbf{\Gamma}_{i}^{\langle t \rangle} *\mathbf{\tilde{c}}^{\langle t \rangle} \tag{4} $$
#
# ##### Explanation of equation
# * The previous cell state $\mathbf{c}^{\langle t-1 \rangle}$ is adjusted (weighted) by the forget gate $\mathbf{\Gamma}_{f}^{\langle t \rangle}$
# * and the candidate value $\tilde{\mathbf{c}}^{\langle t \rangle}$, adjusted (weighted) by the update gate $\mathbf{\Gamma}_{i}^{\langle t \rangle}$
#
# ##### Variable names and shapes in the code
# * `c`: cell state, including all time steps, $\mathbf{c}$ shape $(n_{a}, m, T)$
# * `c_next`: new (next) cell state, $\mathbf{c}^{\langle t \rangle}$ shape $(n_{a}, m)$
# * `c_prev`: previous cell state, $\mathbf{c}^{\langle t-1 \rangle}$, shape $(n_{a}, m)$
# #### - Output gate $\mathbf{\Gamma}_{o}$
#
# * The output gate decides what gets sent as the prediction (output) of the time step.
# * The output gate is like the other gates. It contains values that range from 0 to 1.
#
# ##### Equation
#
# $$ \mathbf{\Gamma}_o^{\langle t \rangle}= \sigma(\mathbf{W}_o[\mathbf{a}^{\langle t-1 \rangle}, \mathbf{x}^{\langle t \rangle}] + \mathbf{b}_{o})\tag{5}$$
#
# ##### Explanation of the equation
# * The output gate is determined by the previous hidden state $\mathbf{a}^{\langle t-1 \rangle}$ and the current input $\mathbf{x}^{\langle t \rangle}$
# * The sigmoid makes the gate range from 0 to 1.
#
#
# ##### Variable names in the code
# * `Wo`: output gate weight, $\mathbf{W_o}$
# * `bo`: output gate bias, $\mathbf{b_o}$
# * `ot`: output gate, $\mathbf{\Gamma}_{o}^{\langle t \rangle}$
# #### - Hidden state $\mathbf{a}^{\langle t \rangle}$
#
# * The hidden state gets passed to the LSTM cell's next time step.
# * It is used to determine the three gates ($\mathbf{\Gamma}_{f}, \mathbf{\Gamma}_{u}, \mathbf{\Gamma}_{o}$) of the next time step.
# * The hidden state is also used for the prediction $y^{\langle t \rangle}$.
#
# ##### Equation
#
# $$ \mathbf{a}^{\langle t \rangle} = \mathbf{\Gamma}_o^{\langle t \rangle} * \tanh(\mathbf{c}^{\langle t \rangle})\tag{6} $$
#
# ##### Explanation of equation
# * The hidden state $\mathbf{a}^{\langle t \rangle}$ is determined by the cell state $\mathbf{c}^{\langle t \rangle}$ in combination with the output gate $\mathbf{\Gamma}_{o}$.
# * The cell state state is passed through the "tanh" function to rescale values between -1 and +1.
# * The output gate acts like a "mask" that either preserves the values of $\tanh(\mathbf{c}^{\langle t \rangle})$ or keeps those values from being included in the hidden state $\mathbf{a}^{\langle t \rangle}$
#
# ##### Variable names and shapes in the code
# * `a`: hidden state, including time steps. $\mathbf{a}$ has shape $(n_{a}, m, T_{x})$
# * 'a_prev`: hidden state from previous time step. $\mathbf{a}^{\langle t-1 \rangle}$ has shape $(n_{a}, m)$
# * `a_next`: hidden state for next time step. $\mathbf{a}^{\langle t \rangle}$ has shape $(n_{a}, m)$
# #### - Prediction $\mathbf{y}^{\langle t \rangle}_{pred}$
# * The prediction in this use case is a classification, so we'll use a softmax.
#
# The equation is:
# $$\mathbf{y}^{\langle t \rangle}_{pred} = \textrm{softmax}(\mathbf{W}_{y} \mathbf{a}^{\langle t \rangle} + \mathbf{b}_{y})$$
#
# ##### Variable names and shapes in the code
# * `y_pred`: prediction, including all time steps. $\mathbf{y}_{pred}$ has shape $(n_{y}, m, T_{x})$. Note that $(T_{y} = T_{x})$ for this example.
# * `yt_pred`: prediction for the current time step $t$. $\mathbf{y}^{\langle t \rangle}_{pred}$ has shape $(n_{y}, m)$
# ### 2.1 - LSTM cell
#
# **Exercise**: Implement the LSTM cell described in the Figure (4).
#
# **Instructions**:
# 1. Concatenate the hidden state $a^{\langle t-1 \rangle}$ and input $x^{\langle t \rangle}$ into a single matrix:
#
# $$concat = \begin{bmatrix} a^{\langle t-1 \rangle} \\ x^{\langle t \rangle} \end{bmatrix}$$
#
# 2. Compute all the formulas 1 through 6 for the gates, hidden state, and cell state.
# 3. Compute the prediction $y^{\langle t \rangle}$.
#
# #### Additional Hints
# * You can use [numpy.concatenate](https://docs.scipy.org/doc/numpy/reference/generated/numpy.concatenate.html). Check which value to use for the `axis` parameter.
# * The functions `sigmoid()` and `softmax` are imported from `rnn_utils.py`.
# * [numpy.tanh](https://docs.scipy.org/doc/numpy/reference/generated/numpy.tanh.html)
# * Use [np.dot](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html) for matrix multiplication.
# * Notice that the variable names `Wi`, `bi` refer to the weights and biases of the **update** gate. There are no variables named "Wu" or "bu" in this function.
# +
# GRADED FUNCTION: lstm_cell_forward
def lstm_cell_forward(xt, a_prev, c_prev, parameters):
"""
Implement a single forward step of the LSTM-cell as described in Figure (4)
Arguments:
xt -- your input data at timestep "t", numpy array of shape (n_x, m).
a_prev -- Hidden state at timestep "t-1", numpy array of shape (n_a, m)
c_prev -- Memory state at timestep "t-1", numpy array of shape (n_a, m)
parameters -- python dictionary containing:
Wf -- Weight matrix of the forget gate, numpy array of shape (n_a, n_a + n_x)
bf -- Bias of the forget gate, numpy array of shape (n_a, 1)
Wi -- Weight matrix of the update gate, numpy array of shape (n_a, n_a + n_x)
bi -- Bias of the update gate, numpy array of shape (n_a, 1)
Wc -- Weight matrix of the first "tanh", numpy array of shape (n_a, n_a + n_x)
bc -- Bias of the first "tanh", numpy array of shape (n_a, 1)
Wo -- Weight matrix of the output gate, numpy array of shape (n_a, n_a + n_x)
bo -- Bias of the output gate, numpy array of shape (n_a, 1)
Wy -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
Returns:
a_next -- next hidden state, of shape (n_a, m)
c_next -- next memory state, of shape (n_a, m)
yt_pred -- prediction at timestep "t", numpy array of shape (n_y, m)
cache -- tuple of values needed for the backward pass, contains (a_next, c_next, a_prev, c_prev, xt, parameters)
Note: ft/it/ot stand for the forget/update/output gates, cct stands for the candidate value (c tilde),
c stands for the cell state (memory)
"""
# Retrieve parameters from "parameters"
Wf = parameters["Wf"] # forget gate weight
bf = parameters["bf"]
Wi = parameters["Wi"] # update gate weight (notice the variable name)
bi = parameters["bi"] # (notice the variable name)
Wc = parameters["Wc"] # candidate value weight
bc = parameters["bc"]
Wo = parameters["Wo"] # output gate weight
bo = parameters["bo"]
Wy = parameters["Wy"] # prediction weight
by = parameters["by"]
# Retrieve dimensions from shapes of xt and Wy
n_x, m = xt.shape
n_y, n_a = Wy.shape
### START CODE HERE ###
# Concatenate a_prev and xt (≈1 line)
concat = np.concatenate((a_prev,xt), axis=0)
# Compute values for ft (forget gate), it (update gate),
# cct (candidate value), c_next (cell state),
# ot (output gate), a_next (hidden state) (≈6 lines)
ft = sigmoid(np.dot(Wf,concat)+bf) # forget gate
it = sigmoid(np.dot(Wi,concat)+bi) # update gate
cct = np.tanh(np.dot(Wc,concat)+bc) # candidate value
c_next = ft*c_prev+it*cct # cell state
ot = sigmoid(np.dot(Wo,concat)+bo) # output gate
a_next = ot*np.tanh(c_next) # hidden state
# Compute prediction of the LSTM cell (≈1 line)
yt_pred = softmax(np.dot(Wy,a_next)+by)
### END CODE HERE ###
# store values needed for backward propagation in cache
cache = (a_next, c_next, a_prev, c_prev, ft, it, cct, ot, xt, parameters)
return a_next, c_next, yt_pred, cache
# +
np.random.seed(1)
xt_tmp = np.random.randn(3,10)
a_prev_tmp = np.random.randn(5,10)
c_prev_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wf'] = np.random.randn(5, 5+3)
parameters_tmp['bf'] = np.random.randn(5,1)
parameters_tmp['Wi'] = np.random.randn(5, 5+3)
parameters_tmp['bi'] = np.random.randn(5,1)
parameters_tmp['Wo'] = np.random.randn(5, 5+3)
parameters_tmp['bo'] = np.random.randn(5,1)
parameters_tmp['Wc'] = np.random.randn(5, 5+3)
parameters_tmp['bc'] = np.random.randn(5,1)
parameters_tmp['Wy'] = np.random.randn(2,5)
parameters_tmp['by'] = np.random.randn(2,1)
a_next_tmp, c_next_tmp, yt_tmp, cache_tmp = lstm_cell_forward(xt_tmp, a_prev_tmp, c_prev_tmp, parameters_tmp)
print("a_next[4] = \n", a_next_tmp[4])
print("a_next.shape = ", c_next_tmp.shape)
print("c_next[2] = \n", c_next_tmp[2])
print("c_next.shape = ", c_next_tmp.shape)
print("yt[1] =", yt_tmp[1])
print("yt.shape = ", yt_tmp.shape)
print("cache[1][3] =\n", cache_tmp[1][3])
print("len(cache) = ", len(cache_tmp))
# -
# **Expected Output**:
#
# ```Python
# a_next[4] =
# [-0.66408471 0.0036921 0.02088357 0.22834167 -0.85575339 0.00138482
# 0.76566531 0.34631421 -0.00215674 0.43827275]
# a_next.shape = (5, 10)
# c_next[2] =
# [ 0.63267805 1.00570849 0.35504474 0.20690913 -1.64566718 0.11832942
# 0.76449811 -0.0981561 -0.74348425 -0.26810932]
# c_next.shape = (5, 10)
# yt[1] = [ 0.79913913 0.15986619 0.22412122 0.15606108 0.97057211 0.31146381
# 0.00943007 0.12666353 0.39380172 0.07828381]
# yt.shape = (2, 10)
# cache[1][3] =
# [-0.16263996 1.03729328 0.72938082 -0.54101719 0.02752074 -0.30821874
# 0.07651101 -1.03752894 1.41219977 -0.37647422]
# len(cache) = 10
# ```
# ### 2.2 - Forward pass for LSTM
#
# Now that you have implemented one step of an LSTM, you can now iterate this over this using a for-loop to process a sequence of $T_x$ inputs.
#
# <img src="images/LSTM_rnn.png" style="width:500;height:300px;">
# <caption><center> **Figure 5**: LSTM over multiple time-steps. </center></caption>
#
# **Exercise:** Implement `lstm_forward()` to run an LSTM over $T_x$ time-steps.
#
# **Instructions**
# * Get the dimensions $n_x, n_a, n_y, m, T_x$ from the shape of the variables: `x` and `parameters`.
# * Initialize the 3D tensors $a$, $c$ and $y$.
# - $a$: hidden state, shape $(n_{a}, m, T_{x})$
# - $c$: cell state, shape $(n_{a}, m, T_{x})$
# - $y$: prediction, shape $(n_{y}, m, T_{x})$ (Note that $T_{y} = T_{x}$ in this example).
# - **Note** Setting one variable equal to the other is a "copy by reference". In other words, don't do `c = a', otherwise both these variables point to the same underlying variable.
# * Initialize the 2D tensor $a^{\langle t \rangle}$
# - $a^{\langle t \rangle}$ stores the hidden state for time step $t$. The variable name is `a_next`.
# - $a^{\langle 0 \rangle}$, the initial hidden state at time step 0, is passed in when calling the function. The variable name is `a0`.
# - $a^{\langle t \rangle}$ and $a^{\langle 0 \rangle}$ represent a single time step, so they both have the shape $(n_{a}, m)$
# - Initialize $a^{\langle t \rangle}$ by setting it to the initial hidden state ($a^{\langle 0 \rangle}$) that is passed into the function.
# * Initialize $c^{\langle t \rangle}$ with zeros.
# - The variable name is `c_next`.
# - $c^{\langle t \rangle}$ represents a single time step, so its shape is $(n_{a}, m)$
# - **Note**: create `c_next` as its own variable with its own location in memory. Do not initialize it as a slice of the 3D tensor $c$. In other words, **don't** do `c_next = c[:,:,0]`.
# * For each time step, do the following:
# - From the 3D tensor $x$, get a 2D slice $x^{\langle t \rangle}$ at time step $t$.
# - Call the `lstm_cell_forward` function that you defined previously, to get the hidden state, cell state, prediction, and cache.
# - Store the hidden state, cell state and prediction (the 2D tensors) inside the 3D tensors.
# - Also append the cache to the list of caches.
# +
# GRADED FUNCTION: lstm_forward
def lstm_forward(x, a0, parameters):
"""
Implement the forward propagation of the recurrent neural network using an LSTM-cell described in Figure (4).
Arguments:
x -- Input data for every time-step, of shape (n_x, m, T_x).
a0 -- Initial hidden state, of shape (n_a, m)
parameters -- python dictionary containing:
Wf -- Weight matrix of the forget gate, numpy array of shape (n_a, n_a + n_x)
bf -- Bias of the forget gate, numpy array of shape (n_a, 1)
Wi -- Weight matrix of the update gate, numpy array of shape (n_a, n_a + n_x)
bi -- Bias of the update gate, numpy array of shape (n_a, 1)
Wc -- Weight matrix of the first "tanh", numpy array of shape (n_a, n_a + n_x)
bc -- Bias of the first "tanh", numpy array of shape (n_a, 1)
Wo -- Weight matrix of the output gate, numpy array of shape (n_a, n_a + n_x)
bo -- Bias of the output gate, numpy array of shape (n_a, 1)
Wy -- Weight matrix relating the hidden-state to the output, numpy array of shape (n_y, n_a)
by -- Bias relating the hidden-state to the output, numpy array of shape (n_y, 1)
Returns:
a -- Hidden states for every time-step, numpy array of shape (n_a, m, T_x)
y -- Predictions for every time-step, numpy array of shape (n_y, m, T_x)
c -- The value of the cell state, numpy array of shape (n_a, m, T_x)
caches -- tuple of values needed for the backward pass, contains (list of all the caches, x)
"""
# Initialize "caches", which will track the list of all the caches
caches = []
### START CODE HERE ###
Wy = parameters['Wy'] # saving parameters['Wy'] in a local variable in case students use Wy instead of parameters['Wy']
# Retrieve dimensions from shapes of x and parameters['Wy'] (≈2 lines)
n_x, m, T_x = x.shape
n_y, n_a = Wy.shape
# initialize "a", "c" and "y" with zeros (≈3 lines)
a = np.zeros((n_a,m,T_x))
c = np.zeros((n_a,m,T_x))
y = np.zeros((n_y,m,T_x))
# Initialize a_next and c_next (≈2 lines)
a_next = a0
c_next = np.zeros((n_a,m))
# loop over all time-steps
for t in range(T_x):
# Get the 2D slice 'xt' from the 3D input 'x' at time step 't'
xt = x[:,:,t]
# Update next hidden state, next memory state, compute the prediction, get the cache (≈1 line)
a_next, c_next, yt, cache = lstm_cell_forward(xt, a_next, c_next, parameters)
# Save the value of the new "next" hidden state in a (≈1 line)
a[:,:,t] = a_next
# Save the value of the next cell state (≈1 line)
c[:,:,t] = c_next
# Save the value of the prediction in y (≈1 line)
y[:,:,t] = yt
# Append the cache into caches (≈1 line)
np.append(caches,cache)
### END CODE HERE ###
# store values needed for backward propagation in cache
caches = (caches, x)
return a, y, c, caches
# +
np.random.seed(1)
x_tmp = np.random.randn(3,10,7)
a0_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wf'] = np.random.randn(5, 5+3)
parameters_tmp['bf'] = np.random.randn(5,1)
parameters_tmp['Wi'] = np.random.randn(5, 5+3)
parameters_tmp['bi']= np.random.randn(5,1)
parameters_tmp['Wo'] = np.random.randn(5, 5+3)
parameters_tmp['bo'] = np.random.randn(5,1)
parameters_tmp['Wc'] = np.random.randn(5, 5+3)
parameters_tmp['bc'] = np.random.randn(5,1)
parameters_tmp['Wy'] = np.random.randn(2,5)
parameters_tmp['by'] = np.random.randn(2,1)
a_tmp, y_tmp, c_tmp, caches_tmp = lstm_forward(x_tmp, a0_tmp, parameters_tmp)
print("a[4][3][6] = ", a_tmp[4][3][6])
print("a.shape = ", a_tmp.shape)
print("y[1][4][3] =", y_tmp[1][4][3])
print("y.shape = ", y_tmp.shape)
print("caches[1][1][1] =\n", caches_tmp[1][1][1])
print("c[1][2][1]", c_tmp[1][2][1])
print("len(caches) = ", len(caches_tmp))
# -
# **Expected Output**:
#
# ```Python
# a[4][3][6] = 0.172117767533
# a.shape = (5, 10, 7)
# y[1][4][3] = 0.95087346185
# y.shape = (2, 10, 7)
# caches[1][1][1] =
# [ 0.82797464 0.23009474 0.76201118 -0.22232814 -0.20075807 0.18656139
# 0.41005165]
# c[1][2][1] -0.855544916718
# len(caches) = 2
# ```
# Congratulations! You have now implemented the forward passes for the basic RNN and the LSTM. When using a deep learning framework, implementing the forward pass is sufficient to build systems that achieve great performance.
#
# The rest of this notebook is optional, and will not be graded.
# ## 3 - Backpropagation in recurrent neural networks (OPTIONAL / UNGRADED)
#
# In modern deep learning frameworks, you only have to implement the forward pass, and the framework takes care of the backward pass, so most deep learning engineers do not need to bother with the details of the backward pass. If however you are an expert in calculus and want to see the details of backprop in RNNs, you can work through this optional portion of the notebook.
#
# When in an earlier [course](https://www.coursera.org/learn/neural-networks-deep-learning/lecture/0VSHe/derivatives-with-a-computation-graph) you implemented a simple (fully connected) neural network, you used backpropagation to compute the derivatives with respect to the cost to update the parameters. Similarly, in recurrent neural networks you can calculate the derivatives with respect to the cost in order to update the parameters. The backprop equations are quite complicated and we did not derive them in lecture. However, we will briefly present them below.
# Note that this notebook does not implement the backward path from the Loss 'J' backwards to 'a'. This would have included the dense layer and softmax which are a part of the forward path. This is assumed to be calculated elsewhere and the result passed to rnn_backward in 'da'. It is further assumed that loss has been adjusted for batch size (m) and division by the number of examples is not required here.
# This section is optional and ungraded. It is more difficult and has fewer details regarding its implementation. This section only implements key elements of the full path.
# ### 3.1 - Basic RNN backward pass
#
# We will start by computing the backward pass for the basic RNN-cell and then in the following sections, iterate through the cells.
#
# <img src="images/rnn_backward_overview_3a_1.png" style="width:500;height:300px;"> <br>
# <caption><center> **Figure 6**: RNN-cell's backward pass. Just like in a fully-connected neural network, the derivative of the cost function $J$ backpropagates through the time steps of the RNN by following the chain-rule from calculus. Internal to the cell, the chain-rule is also used to calculate $(\frac{\partial J}{\partial W_{ax}},\frac{\partial J}{\partial W_{aa}},\frac{\partial J}{\partial b})$ to update the parameters $(W_{ax}, W_{aa}, b_a)$. The operation can utilize the cached results from the forward path. </center></caption>
# Recall from lecture, the shorthand for the partial derivative of cost relative to a variable is dVariable. For example, $\frac{\partial J}{\partial W_{ax}}$ is $dW_{ax}$. This will be used throughout the remaining sections.
#
# <img src="images/rnn_cell_backward_3a_4.png" style="width:500;height:300px;"> <br>
# <caption><center> **Figure 7**: This implementation of rnn_cell_backward does not include the output dense layer and softmax which are included in rnn_cell_forward.
#
# $da_{next}$ is $\frac{\partial{J}}{\partial a^{\langle t \rangle}}$ and includes loss from previous stages and current stage output logic. This addition will be part of your implementation of rnn_backward. </center></caption>
# ##### Equations
# To compute the rnn_cell_backward you can utilize the following equations. It is a good exercise to derive them by hand. Here, $*$ denotes element-wise multiplication while the absence of a symbol indicates matrix multiplication.
#
# $a^{\langle t \rangle} = \tanh(W_{ax} x^{\langle t \rangle} + W_{aa} a^{\langle t-1 \rangle} + b_{a})\tag{-}$
#
# $\displaystyle \frac{\partial \tanh(x)} {\partial x} = 1 - \tanh^2(x) \tag{-}$
#
# $\displaystyle {dW_{ax}} = da_{next} * ( 1-\tanh^2(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a}) ) x^{\langle t \rangle T}\tag{1}$
#
# $\displaystyle dW_{aa} = da_{next} * (( 1-\tanh^2(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a}) ) a^{\langle t-1 \rangle T}\tag{2}$
#
# $\displaystyle db_a = da_{next} * \sum_{batch}( 1-\tanh^2(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a}) )\tag{3}$
#
# $\displaystyle dx^{\langle t \rangle} = da_{next} * { W_{ax}}^T ( 1-\tanh^2(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a}) )\tag{4}$
#
# $\displaystyle da_{prev} = da_{next} * { W_{aa}}^T ( 1-\tanh^2(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a}) )\tag{5}$
#
# #### Implementing rnn_cell_backward
# The results can be computed directly by implementing the equations above. However, the above can optionally be simplified by computing 'dz' and utlilizing the chain rule.
# This can be further simplified by noting that $\tanh(W_{ax}x^{\langle t \rangle}+W_{aa} a^{\langle t-1 \rangle} + b_{a})$ was computed and saved in the forward pass.
#
# To calculate dba, the 'batch' above is a sum across the horizontal (axis= 1) axis. Note that you should use the keepdims = True option.
#
# It may be worthwhile to review Course 1 [Derivatives with a computational graph](https://www.coursera.org/learn/neural-networks-deep-learning/lecture/0VSHe/derivatives-with-a-computation-graph) through [Backpropagation Intuition](https://www.coursera.org/learn/neural-networks-deep-learning/lecture/6dDj7/backpropagation-intuition-optional), which decompose the calculation into steps using the chain rule.
# Matrix vector derivatives are described [here](http://cs231n.stanford.edu/vecDerivs.pdf), though the equations above incorporate the required transformations.
#
# Note rnn_cell_backward does __not__ include the calculation of loss from $y \langle t \rangle$, this is incorporated into the incoming da_next. This is a slight mismatch with rnn_cell_forward which includes a dense layer and softmax.
#
# Note: in the code:
# $\displaystyle dx^{\langle t \rangle}$ is represented by dxt,
# $\displaystyle d W_{ax}$ is represented by dWax,
# $\displaystyle da_{prev}$ is represented by da_prev,
# $\displaystyle dW_{aa}$ is represented by dWaa,
# $\displaystyle db_{a}$ is represented by dba,
# dz is not derived above but can optionally be derived by students to simplify the repeated calculations.
#
def rnn_cell_backward(da_next, cache):
"""
Implements the backward pass for the RNN-cell (single time-step).
Arguments:
da_next -- Gradient of loss with respect to next hidden state
cache -- python dictionary containing useful values (output of rnn_cell_forward())
Returns:
gradients -- python dictionary containing:
dx -- Gradients of input data, of shape (n_x, m)
da_prev -- Gradients of previous hidden state, of shape (n_a, m)
dWax -- Gradients of input-to-hidden weights, of shape (n_a, n_x)
dWaa -- Gradients of hidden-to-hidden weights, of shape (n_a, n_a)
dba -- Gradients of bias vector, of shape (n_a, 1)
"""
# Retrieve values from cache
(a_next, a_prev, xt, parameters) = cache
# Retrieve values from parameters
Wax = parameters["Wax"]
Waa = parameters["Waa"]
Wya = parameters["Wya"]
ba = parameters["ba"]
by = parameters["by"]
### START CODE HERE ###
# compute the gradient of the loss with respect to z (optional) (≈1 line)
dz = None
# compute the gradient of the loss with respect to Wax (≈2 lines)
dxt = da_next*np.dot(np.transpose(Wax),(1-(np.tanh(np.dot(Wax,xt)+np.dot(Waa,a_prev)+ba)**2)))
dWax = da_next*np.dot((1-(np.tanh(np.dot(Wax,xt)+np.dot(Waa,a_prev)+ba)**2)),np.transpose(xt))
# compute the gradient with respect to Waa (≈2 lines)
da_prev = da_next*np.dot(np.transpose(Waa),(1-(np.tanh(np.dot(Wax,xt)+np.dot(Waa,a_prev)+ba)**2)))
dWaa = da_next*np.dot((1-(np.tanh(np.dot(Wax,xt)+np.dot(Waa,a_prev)+ba)**2)),np.transpose(a_prev))
# compute the gradient with respect to b (≈1 line)
dba = da_next*np.reduce_sum((1-(np.tanh(np.dot(Wax,xt)+np.dot(Waa,a_prev)+ba)**2)))
### END CODE HERE ###
# Store the gradients in a python dictionary
gradients = {"dxt": dxt, "da_prev": da_prev, "dWax": dWax, "dWaa": dWaa, "dba": dba}
return gradients
# +
np.random.seed(1)
xt_tmp = np.random.randn(3,10)
a_prev_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wax'] = np.random.randn(5,3)
parameters_tmp['Waa'] = np.random.randn(5,5)
parameters_tmp['Wya'] = np.random.randn(2,5)
parameters_tmp['ba'] = np.random.randn(5,1)
parameters_tmp['by'] = np.random.randn(2,1)
a_next_tmp, yt_tmp, cache_tmp = rnn_cell_forward(xt_tmp, a_prev_tmp, parameters_tmp)
da_next_tmp = np.random.randn(5,10)
gradients_tmp = rnn_cell_backward(da_next_tmp, cache_tmp)
print("gradients[\"dxt\"][1][2] =", gradients_tmp["dxt"][1][2])
print("gradients[\"dxt\"].shape =", gradients_tmp["dxt"].shape)
print("gradients[\"da_prev\"][2][3] =", gradients_tmp["da_prev"][2][3])
print("gradients[\"da_prev\"].shape =", gradients_tmp["da_prev"].shape)
print("gradients[\"dWax\"][3][1] =", gradients_tmp["dWax"][3][1])
print("gradients[\"dWax\"].shape =", gradients_tmp["dWax"].shape)
print("gradients[\"dWaa\"][1][2] =", gradients_tmp["dWaa"][1][2])
print("gradients[\"dWaa\"].shape =", gradients_tmp["dWaa"].shape)
print("gradients[\"dba\"][4] =", gradients_tmp["dba"][4])
print("gradients[\"dba\"].shape =", gradients_tmp["dba"].shape)
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# **gradients["dxt"][1][2]** =
# </td>
# <td>
# -1.3872130506
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dxt"].shape** =
# </td>
# <td>
# (3, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da_prev"][2][3]** =
# </td>
# <td>
# -0.152399493774
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da_prev"].shape** =
# </td>
# <td>
# (5, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWax"][3][1]** =
# </td>
# <td>
# 0.410772824935
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWax"].shape** =
# </td>
# <td>
# (5, 3)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWaa"][1][2]** =
# </td>
# <td>
# 1.15034506685
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWaa"].shape** =
# </td>
# <td>
# (5, 5)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dba"][4]** =
# </td>
# <td>
# [ 0.20023491]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dba"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# </table>
# #### Backward pass through the RNN
#
# Computing the gradients of the cost with respect to $a^{\langle t \rangle}$ at every time-step $t$ is useful because it is what helps the gradient backpropagate to the previous RNN-cell. To do so, you need to iterate through all the time steps starting at the end, and at each step, you increment the overall $db_a$, $dW_{aa}$, $dW_{ax}$ and you store $dx$.
#
# **Instructions**:
#
# Implement the `rnn_backward` function. Initialize the return variables with zeros first and then loop through all the time steps while calling the `rnn_cell_backward` at each time timestep, update the other variables accordingly.
# * Note that this notebook does not implement the backward path from the Loss 'J' backwards to 'a'.
# * This would have included the dense layer and softmax which are a part of the forward path.
# * This is assumed to be calculated elsewhere and the result passed to rnn_backward in 'da'.
# * You must combine this with the loss from the previous stages when calling rnn_cell_backward (see figure 7 above).
# * It is further assumed that loss has been adjusted for batch size (m).
# * Therefore, division by the number of examples is not required here.
def rnn_backward(da, caches):
"""
Implement the backward pass for a RNN over an entire sequence of input data.
Arguments:
da -- Upstream gradients of all hidden states, of shape (n_a, m, T_x)
caches -- tuple containing information from the forward pass (rnn_forward)
Returns:
gradients -- python dictionary containing:
dx -- Gradient w.r.t. the input data, numpy-array of shape (n_x, m, T_x)
da0 -- Gradient w.r.t the initial hidden state, numpy-array of shape (n_a, m)
dWax -- Gradient w.r.t the input's weight matrix, numpy-array of shape (n_a, n_x)
dWaa -- Gradient w.r.t the hidden state's weight matrix, numpy-arrayof shape (n_a, n_a)
dba -- Gradient w.r.t the bias, of shape (n_a, 1)
"""
### START CODE HERE ###
# Retrieve values from the first cache (t=1) of caches (≈2 lines)
(caches, x) = None
(a1, a0, x1, parameters) = None
# Retrieve dimensions from da's and x1's shapes (≈2 lines)
n_a, m, T_x = None
n_x, m = None
# initialize the gradients with the right sizes (≈6 lines)
dx = None
dWax = None
dWaa = None
dba = None
da0 = None
da_prevt = None
# Loop through all the time steps
for t in reversed(range(None)):
# Compute gradients at time step t.
# Remember to sum gradients from the output path (da) and the previous timesteps (da_prevt) (≈1 line)
gradients = None
# Retrieve derivatives from gradients (≈ 1 line)
dxt, da_prevt, dWaxt, dWaat, dbat = gradients["dxt"], gradients["da_prev"], gradients["dWax"], gradients["dWaa"], gradients["dba"]
# Increment global derivatives w.r.t parameters by adding their derivative at time-step t (≈4 lines)
dx[:, :, t] = None
dWax += None
dWaa += None
dba += None
# Set da0 to the gradient of a which has been backpropagated through all time-steps (≈1 line)
da0 = None
### END CODE HERE ###
# Store the gradients in a python dictionary
gradients = {"dx": dx, "da0": da0, "dWax": dWax, "dWaa": dWaa,"dba": dba}
return gradients
# +
np.random.seed(1)
x_tmp = np.random.randn(3,10,4)
a0_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wax'] = np.random.randn(5,3)
parameters_tmp['Waa'] = np.random.randn(5,5)
parameters_tmp['Wya'] = np.random.randn(2,5)
parameters_tmp['ba'] = np.random.randn(5,1)
parameters_tmp['by'] = np.random.randn(2,1)
a_tmp, y_tmp, caches_tmp = rnn_forward(x_tmp, a0_tmp, parameters_tmp)
da_tmp = np.random.randn(5, 10, 4)
gradients_tmp = rnn_backward(da_tmp, caches_tmp)
print("gradients[\"dx\"][1][2] =", gradients_tmp["dx"][1][2])
print("gradients[\"dx\"].shape =", gradients_tmp["dx"].shape)
print("gradients[\"da0\"][2][3] =", gradients_tmp["da0"][2][3])
print("gradients[\"da0\"].shape =", gradients_tmp["da0"].shape)
print("gradients[\"dWax\"][3][1] =", gradients_tmp["dWax"][3][1])
print("gradients[\"dWax\"].shape =", gradients_tmp["dWax"].shape)
print("gradients[\"dWaa\"][1][2] =", gradients_tmp["dWaa"][1][2])
print("gradients[\"dWaa\"].shape =", gradients_tmp["dWaa"].shape)
print("gradients[\"dba\"][4] =", gradients_tmp["dba"][4])
print("gradients[\"dba\"].shape =", gradients_tmp["dba"].shape)
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# **gradients["dx"][1][2]** =
# </td>
# <td>
# [-2.07101689 -0.59255627 0.02466855 0.01483317]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dx"].shape** =
# </td>
# <td>
# (3, 10, 4)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da0"][2][3]** =
# </td>
# <td>
# -0.314942375127
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da0"].shape** =
# </td>
# <td>
# (5, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWax"][3][1]** =
# </td>
# <td>
# 11.2641044965
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWax"].shape** =
# </td>
# <td>
# (5, 3)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWaa"][1][2]** =
# </td>
# <td>
# 2.30333312658
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWaa"].shape** =
# </td>
# <td>
# (5, 5)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dba"][4]** =
# </td>
# <td>
# [-0.74747722]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dba"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# </table>
# ## 3.2 - LSTM backward pass
# ### 3.2.1 One Step backward
# The LSTM backward pass is slighltly more complicated than the forward one.
#
# <img src="images/LSTM_cell_backward_rev3a_5.png" style="width:500;height:400px;"> <br>
# <caption><center> **Figure 8**: lstm_cell_backward. Note the output functions, while part of the lstm_cell_forward, are not included in lstm_cell_backward </center></caption>
#
# The equations for the LSTM backward pass are provided below. (If you enjoy calculus exercises feel free to try deriving these from scratch yourself.)
# ### 3.2.2 gate derivatives
# Note the location of the gate derivatives ($\gamma$..) between the dense layer and the activation function (see graphic above). This is convenient for computing parameter derivatives in the next step.
#
# $d\gamma_o^{\langle t \rangle} = da_{next}*\tanh(c_{next}) * \Gamma_o^{\langle t \rangle}*\left(1-\Gamma_o^{\langle t \rangle}\right)\tag{7}$
#
# $dp\widetilde{c}^{\langle t \rangle} = \left(dc_{next}*\Gamma_u^{\langle t \rangle}+ \Gamma_o^{\langle t \rangle}* (1-\tanh^2(c_{next})) * \Gamma_u^{\langle t \rangle} * da_{next} \right) * \left(1-\left(\widetilde c^{\langle t \rangle}\right)^2\right) \tag{8}$
#
# $d\gamma_u^{\langle t \rangle} = \left(dc_{next}*\widetilde{c}^{\langle t \rangle} + \Gamma_o^{\langle t \rangle}* (1-\tanh^2(c_{next})) * \widetilde{c}^{\langle t \rangle} * da_{next}\right)*\Gamma_u^{\langle t \rangle}*\left(1-\Gamma_u^{\langle t \rangle}\right)\tag{9}$
#
# $d\gamma_f^{\langle t \rangle} = \left(dc_{next}* c_{prev} + \Gamma_o^{\langle t \rangle} * (1-\tanh^2(c_{next})) * c_{prev} * da_{next}\right)*\Gamma_f^{\langle t \rangle}*\left(1-\Gamma_f^{\langle t \rangle}\right)\tag{10}$
#
# ### 3.2.3 parameter derivatives
#
# $ dW_f = d\gamma_f^{\langle t \rangle} \begin{bmatrix} a_{prev} \\ x_t\end{bmatrix}^T \tag{11} $
# $ dW_u = d\gamma_u^{\langle t \rangle} \begin{bmatrix} a_{prev} \\ x_t\end{bmatrix}^T \tag{12} $
# $ dW_c = dp\widetilde c^{\langle t \rangle} \begin{bmatrix} a_{prev} \\ x_t\end{bmatrix}^T \tag{13} $
# $ dW_o = d\gamma_o^{\langle t \rangle} \begin{bmatrix} a_{prev} \\ x_t\end{bmatrix}^T \tag{14}$
#
# To calculate $db_f, db_u, db_c, db_o$ you just need to sum across the horizontal (axis= 1) axis on $d\gamma_f^{\langle t \rangle}, d\gamma_u^{\langle t \rangle}, dp\widetilde c^{\langle t \rangle}, d\gamma_o^{\langle t \rangle}$ respectively. Note that you should have the `keepdims = True` option.
#
# $\displaystyle db_f = \sum_{batch}d\gamma_f^{\langle t \rangle}\tag{15}$
# $\displaystyle db_u = \sum_{batch}d\gamma_u^{\langle t \rangle}\tag{16}$
# $\displaystyle db_c = \sum_{batch}d\gamma_c^{\langle t \rangle}\tag{17}$
# $\displaystyle db_o = \sum_{batch}d\gamma_o^{\langle t \rangle}\tag{18}$
#
# Finally, you will compute the derivative with respect to the previous hidden state, previous memory state, and input.
#
# $ da_{prev} = W_f^T d\gamma_f^{\langle t \rangle} + W_u^T d\gamma_u^{\langle t \rangle}+ W_c^T dp\widetilde c^{\langle t \rangle} + W_o^T d\gamma_o^{\langle t \rangle} \tag{19}$
#
# Here, to account for concatenation, the weights for equations 19 are the first n_a, (i.e. $W_f = W_f[:,:n_a]$ etc...)
#
# $ dc_{prev} = dc_{next}*\Gamma_f^{\langle t \rangle} + \Gamma_o^{\langle t \rangle} * (1- \tanh^2(c_{next}))*\Gamma_f^{\langle t \rangle}*da_{next} \tag{20}$
#
# $ dx^{\langle t \rangle} = W_f^T d\gamma_f^{\langle t \rangle} + W_u^T d\gamma_u^{\langle t \rangle}+ W_c^T dp\widetilde c^{\langle t \rangle} + W_o^T d\gamma_o^{\langle t \rangle}\tag{21} $
#
# where the weights for equation 21 are from n_a to the end, (i.e. $W_f = W_f[:,n_a:]$ etc...)
#
# **Exercise:** Implement `lstm_cell_backward` by implementing equations $7-21$ below.
#
#
# Note: In the code:
#
# $d\gamma_o^{\langle t \rangle}$ is represented by `dot`,
# $dp\widetilde{c}^{\langle t \rangle}$ is represented by `dcct`,
# $d\gamma_u^{\langle t \rangle}$ is represented by `dit`,
# $d\gamma_f^{\langle t \rangle}$ is represented by `dft`
#
def lstm_cell_backward(da_next, dc_next, cache):
"""
Implement the backward pass for the LSTM-cell (single time-step).
Arguments:
da_next -- Gradients of next hidden state, of shape (n_a, m)
dc_next -- Gradients of next cell state, of shape (n_a, m)
cache -- cache storing information from the forward pass
Returns:
gradients -- python dictionary containing:
dxt -- Gradient of input data at time-step t, of shape (n_x, m)
da_prev -- Gradient w.r.t. the previous hidden state, numpy array of shape (n_a, m)
dc_prev -- Gradient w.r.t. the previous memory state, of shape (n_a, m, T_x)
dWf -- Gradient w.r.t. the weight matrix of the forget gate, numpy array of shape (n_a, n_a + n_x)
dWi -- Gradient w.r.t. the weight matrix of the update gate, numpy array of shape (n_a, n_a + n_x)
dWc -- Gradient w.r.t. the weight matrix of the memory gate, numpy array of shape (n_a, n_a + n_x)
dWo -- Gradient w.r.t. the weight matrix of the output gate, numpy array of shape (n_a, n_a + n_x)
dbf -- Gradient w.r.t. biases of the forget gate, of shape (n_a, 1)
dbi -- Gradient w.r.t. biases of the update gate, of shape (n_a, 1)
dbc -- Gradient w.r.t. biases of the memory gate, of shape (n_a, 1)
dbo -- Gradient w.r.t. biases of the output gate, of shape (n_a, 1)
"""
# Retrieve information from "cache"
(a_next, c_next, a_prev, c_prev, ft, it, cct, ot, xt, parameters) = cache
### START CODE HERE ###
# Retrieve dimensions from xt's and a_next's shape (≈2 lines)
n_x, m = None
n_a, m = None
# Compute gates related derivatives, you can find their values can be found by looking carefully at equations (7) to (10) (≈4 lines)
dot = None
dcct = None
dit = None
dft = None
# Compute parameters related derivatives. Use equations (11)-(18) (≈8 lines)
dWf = None
dWi = None
dWc = None
dWo = None
dbf = None
dbi = None
dbc = None
dbo = None
# Compute derivatives w.r.t previous hidden state, previous memory state and input. Use equations (19)-(21). (≈3 lines)
da_prev = None
dc_prev = None
dxt = None
### END CODE HERE ###
# Save gradients in dictionary
gradients = {"dxt": dxt, "da_prev": da_prev, "dc_prev": dc_prev, "dWf": dWf,"dbf": dbf, "dWi": dWi,"dbi": dbi,
"dWc": dWc,"dbc": dbc, "dWo": dWo,"dbo": dbo}
return gradients
# +
np.random.seed(1)
xt_tmp = np.random.randn(3,10)
a_prev_tmp = np.random.randn(5,10)
c_prev_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wf'] = np.random.randn(5, 5+3)
parameters_tmp['bf'] = np.random.randn(5,1)
parameters_tmp['Wi'] = np.random.randn(5, 5+3)
parameters_tmp['bi'] = np.random.randn(5,1)
parameters_tmp['Wo'] = np.random.randn(5, 5+3)
parameters_tmp['bo'] = np.random.randn(5,1)
parameters_tmp['Wc'] = np.random.randn(5, 5+3)
parameters_tmp['bc'] = np.random.randn(5,1)
parameters_tmp['Wy'] = np.random.randn(2,5)
parameters_tmp['by'] = np.random.randn(2,1)
a_next_tmp, c_next_tmp, yt_tmp, cache_tmp = lstm_cell_forward(xt_tmp, a_prev_tmp, c_prev_tmp, parameters_tmp)
da_next_tmp = np.random.randn(5,10)
dc_next_tmp = np.random.randn(5,10)
gradients_tmp = lstm_cell_backward(da_next_tmp, dc_next_tmp, cache_tmp)
print("gradients[\"dxt\"][1][2] =", gradients_tmp["dxt"][1][2])
print("gradients[\"dxt\"].shape =", gradients_tmp["dxt"].shape)
print("gradients[\"da_prev\"][2][3] =", gradients_tmp["da_prev"][2][3])
print("gradients[\"da_prev\"].shape =", gradients_tmp["da_prev"].shape)
print("gradients[\"dc_prev\"][2][3] =", gradients_tmp["dc_prev"][2][3])
print("gradients[\"dc_prev\"].shape =", gradients_tmp["dc_prev"].shape)
print("gradients[\"dWf\"][3][1] =", gradients_tmp["dWf"][3][1])
print("gradients[\"dWf\"].shape =", gradients_tmp["dWf"].shape)
print("gradients[\"dWi\"][1][2] =", gradients_tmp["dWi"][1][2])
print("gradients[\"dWi\"].shape =", gradients_tmp["dWi"].shape)
print("gradients[\"dWc\"][3][1] =", gradients_tmp["dWc"][3][1])
print("gradients[\"dWc\"].shape =", gradients_tmp["dWc"].shape)
print("gradients[\"dWo\"][1][2] =", gradients_tmp["dWo"][1][2])
print("gradients[\"dWo\"].shape =", gradients_tmp["dWo"].shape)
print("gradients[\"dbf\"][4] =", gradients_tmp["dbf"][4])
print("gradients[\"dbf\"].shape =", gradients_tmp["dbf"].shape)
print("gradients[\"dbi\"][4] =", gradients_tmp["dbi"][4])
print("gradients[\"dbi\"].shape =", gradients_tmp["dbi"].shape)
print("gradients[\"dbc\"][4] =", gradients_tmp["dbc"][4])
print("gradients[\"dbc\"].shape =", gradients_tmp["dbc"].shape)
print("gradients[\"dbo\"][4] =", gradients_tmp["dbo"][4])
print("gradients[\"dbo\"].shape =", gradients_tmp["dbo"].shape)
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# **gradients["dxt"][1][2]** =
# </td>
# <td>
# 3.23055911511
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dxt"].shape** =
# </td>
# <td>
# (3, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da_prev"][2][3]** =
# </td>
# <td>
# -0.0639621419711
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da_prev"].shape** =
# </td>
# <td>
# (5, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dc_prev"][2][3]** =
# </td>
# <td>
# 0.797522038797
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dc_prev"].shape** =
# </td>
# <td>
# (5, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWf"][3][1]** =
# </td>
# <td>
# -0.147954838164
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWf"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWi"][1][2]** =
# </td>
# <td>
# 1.05749805523
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWi"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWc"][3][1]** =
# </td>
# <td>
# 2.30456216369
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWc"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWo"][1][2]** =
# </td>
# <td>
# 0.331311595289
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWo"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbf"][4]** =
# </td>
# <td>
# [ 0.18864637]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbf"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbi"][4]** =
# </td>
# <td>
# [-0.40142491]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbi"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbc"][4]** =
# </td>
# <td>
# [ 0.25587763]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbc"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbo"][4]** =
# </td>
# <td>
# [ 0.13893342]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbo"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# </table>
# ### 3.3 Backward pass through the LSTM RNN
#
# This part is very similar to the `rnn_backward` function you implemented above. You will first create variables of the same dimension as your return variables. You will then iterate over all the time steps starting from the end and call the one step function you implemented for LSTM at each iteration. You will then update the parameters by summing them individually. Finally return a dictionary with the new gradients.
#
# **Instructions**: Implement the `lstm_backward` function. Create a for loop starting from $T_x$ and going backward. For each step call `lstm_cell_backward` and update the your old gradients by adding the new gradients to them. Note that `dxt` is not updated but is stored.
def lstm_backward(da, caches):
"""
Implement the backward pass for the RNN with LSTM-cell (over a whole sequence).
Arguments:
da -- Gradients w.r.t the hidden states, numpy-array of shape (n_a, m, T_x)
caches -- cache storing information from the forward pass (lstm_forward)
Returns:
gradients -- python dictionary containing:
dx -- Gradient of inputs, of shape (n_x, m, T_x)
da0 -- Gradient w.r.t. the previous hidden state, numpy array of shape (n_a, m)
dWf -- Gradient w.r.t. the weight matrix of the forget gate, numpy array of shape (n_a, n_a + n_x)
dWi -- Gradient w.r.t. the weight matrix of the update gate, numpy array of shape (n_a, n_a + n_x)
dWc -- Gradient w.r.t. the weight matrix of the memory gate, numpy array of shape (n_a, n_a + n_x)
dWo -- Gradient w.r.t. the weight matrix of the save gate, numpy array of shape (n_a, n_a + n_x)
dbf -- Gradient w.r.t. biases of the forget gate, of shape (n_a, 1)
dbi -- Gradient w.r.t. biases of the update gate, of shape (n_a, 1)
dbc -- Gradient w.r.t. biases of the memory gate, of shape (n_a, 1)
dbo -- Gradient w.r.t. biases of the save gate, of shape (n_a, 1)
"""
# Retrieve values from the first cache (t=1) of caches.
(caches, x) = caches
(a1, c1, a0, c0, f1, i1, cc1, o1, x1, parameters) = caches[0]
### START CODE HERE ###
# Retrieve dimensions from da's and x1's shapes (≈2 lines)
n_a, m, T_x = None
n_x, m = None
# initialize the gradients with the right sizes (≈12 lines)
dx = None
da0 = None
da_prevt = None
dc_prevt = None
dWf = None
dWi = None
dWc = None
dWo = None
dbf = None
dbi = None
dbc = None
dbo = None
# loop back over the whole sequence
for t in reversed(range(None)):
# Compute all gradients using lstm_cell_backward
gradients = None
# Store or add the gradient to the parameters' previous step's gradient
da_prevt = None
dc_prevt = None
dx[:,:,t] = None
dWf = None
dWi = None
dWc = None
dWo = None
dbf = None
dbi = None
dbc = None
dbo = None
# Set the first activation's gradient to the backpropagated gradient da_prev.
da0 = None
### END CODE HERE ###
# Store the gradients in a python dictionary
gradients = {"dx": dx, "da0": da0, "dWf": dWf,"dbf": dbf, "dWi": dWi,"dbi": dbi,
"dWc": dWc,"dbc": dbc, "dWo": dWo,"dbo": dbo}
return gradients
# +
np.random.seed(1)
x_tmp = np.random.randn(3,10,7)
a0_tmp = np.random.randn(5,10)
parameters_tmp = {}
parameters_tmp['Wf'] = np.random.randn(5, 5+3)
parameters_tmp['bf'] = np.random.randn(5,1)
parameters_tmp['Wi'] = np.random.randn(5, 5+3)
parameters_tmp['bi'] = np.random.randn(5,1)
parameters_tmp['Wo'] = np.random.randn(5, 5+3)
parameters_tmp['bo'] = np.random.randn(5,1)
parameters_tmp['Wc'] = np.random.randn(5, 5+3)
parameters_tmp['bc'] = np.random.randn(5,1)
parameters_tmp['Wy'] = np.zeros((2,5)) # unused, but needed for lstm_forward
parameters_tmp['by'] = np.zeros((2,1)) # unused, but needed for lstm_forward
a_tmp, y_tmp, c_tmp, caches_tmp = lstm_forward(x_tmp, a0_tmp, parameters_tmp)
da_tmp = np.random.randn(5, 10, 4)
gradients_tmp = lstm_backward(da_tmp, caches_tmp)
print("gradients[\"dx\"][1][2] =", gradients_tmp["dx"][1][2])
print("gradients[\"dx\"].shape =", gradients_tmp["dx"].shape)
print("gradients[\"da0\"][2][3] =", gradients_tmp["da0"][2][3])
print("gradients[\"da0\"].shape =", gradients_tmp["da0"].shape)
print("gradients[\"dWf\"][3][1] =", gradients_tmp["dWf"][3][1])
print("gradients[\"dWf\"].shape =", gradients_tmp["dWf"].shape)
print("gradients[\"dWi\"][1][2] =", gradients_tmp["dWi"][1][2])
print("gradients[\"dWi\"].shape =", gradients_tmp["dWi"].shape)
print("gradients[\"dWc\"][3][1] =", gradients_tmp["dWc"][3][1])
print("gradients[\"dWc\"].shape =", gradients_tmp["dWc"].shape)
print("gradients[\"dWo\"][1][2] =", gradients_tmp["dWo"][1][2])
print("gradients[\"dWo\"].shape =", gradients_tmp["dWo"].shape)
print("gradients[\"dbf\"][4] =", gradients_tmp["dbf"][4])
print("gradients[\"dbf\"].shape =", gradients_tmp["dbf"].shape)
print("gradients[\"dbi\"][4] =", gradients_tmp["dbi"][4])
print("gradients[\"dbi\"].shape =", gradients_tmp["dbi"].shape)
print("gradients[\"dbc\"][4] =", gradients_tmp["dbc"][4])
print("gradients[\"dbc\"].shape =", gradients_tmp["dbc"].shape)
print("gradients[\"dbo\"][4] =", gradients_tmp["dbo"][4])
print("gradients[\"dbo\"].shape =", gradients_tmp["dbo"].shape)
# -
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# **gradients["dx"][1][2]** =
# </td>
# <td>
# [0.00218254 0.28205375 -0.48292508 -0.43281115]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dx"].shape** =
# </td>
# <td>
# (3, 10, 4)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da0"][2][3]** =
# </td>
# <td>
# 0.312770310257
# </td>
# </tr>
# <tr>
# <td>
# **gradients["da0"].shape** =
# </td>
# <td>
# (5, 10)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWf"][3][1]** =
# </td>
# <td>
# -0.0809802310938
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWf"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWi"][1][2]** =
# </td>
# <td>
# 0.40512433093
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWi"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWc"][3][1]** =
# </td>
# <td>
# -0.0793746735512
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWc"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWo"][1][2]** =
# </td>
# <td>
# 0.038948775763
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dWo"].shape** =
# </td>
# <td>
# (5, 8)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbf"][4]** =
# </td>
# <td>
# [-0.15745657]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbf"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbi"][4]** =
# </td>
# <td>
# [-0.50848333]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbi"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbc"][4]** =
# </td>
# <td>
# [-0.42510818]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbc"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbo"][4]** =
# </td>
# <td>
# [ -0.17958196]
# </td>
# </tr>
# <tr>
# <td>
# **gradients["dbo"].shape** =
# </td>
# <td>
# (5, 1)
# </td>
# </tr>
# </table>
# ### Congratulations !
#
# Congratulations on completing this assignment. You now understand how recurrent neural networks work!
#
# Let's go on to the next exercise, where you'll use an RNN to build a character-level language model.
#
|
Building_a_Recurrent_Neural_Network_Step_by_Step.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="5o3CCzV_NZ2s" colab_type="text"
# # What is a Gaussian Mixture Model?
#
# A Gaussian mixture model (GMM) is a latent variable model of continuous data. It assumes that each data point comes from one of several different Gaussian distributions. The modeler assumes she knows the total number of Gaussians in the mixture.
#
#
# * List item
# * List item
#
#
# 
#
# The figure on the left is a directed acyclic graph (DAG). The figure on the right is the same model represented using [plate notation](https://en.wikipedia.org/wiki/Plate_notation). Plate notation takes a set of nodes in the DAG that repeat and collapses them over one dimension into a single node. The "plates" represent a single dimension.
#
# There are two plates in our GMM. One plate has size N=3 for the number of data points and one of size K = 2 for the number of components. Next to each plate is a sampling statement that shows how to sample the vector of variables for that plate from conditional probability distributions.
#
# Plate notation is ideal for a specific model class like a GMM because the number of nodes in the DAG can vary from problem to problem, while the plate notation stays the same.
#
# ## A GMM as a Causal Model
#
# You have probabbly never heard of a GMM being described as a causal model. Indeed in most cases it would likely perform poorly as a causal model. Later, we'll discuss how well this would fair as a causal model.
#
# However, for now, let's just realize the fact that we have a probabilistic generative model on a directed acyclic graph, so we can assume this is a causal model just by assuming the DAG represents causality.
#
# In this figure $X_1$, $X_2$ and $X_3$ are observed continuous random variables. The fact that they are observed is indicated by grey.
#
# $Z_1$, $Z_2$, and $Z_3$ are latent (unobserved) discrete random variables. The fact that they are latent is indicated by the white color of the node.
#
# Each observed node $X_i$ is sampled from either a Normal distribution with mean $\mu_1$ or a Normal distribution with mean $\mu_2$.
#
# ### So what is the causal generative story?
# The _causal generative story_ is simply this; $Z_i$ causes $X_i$. $Z_i$ is a switch mechanism that causes $X_i$ to have a value of either $\mu_1$ plus noise or $\mu_2$ plus noise or $\mu_3$ plus noise.
#
# ## Greeks vs. Romans
#
# We see two kinds of variable names in this representation. Those named with letters from the Greek letters, and those named with letters from the Roman alphabet.
#
# So what should we think about these Greek letters? They don't show up in the causal generative story. Why are they in the graph?
#
# Here is how to understand the differences between the Greeks and the Romans.
#
# 1. The Roman letters X and Z are the causally-related components of our data generating process.
# 2. The Greek letters $\alpha$, $\theta$ $\sigma$, $\sigma_0$ are parameters or weights. These are merely parameters of the **causal Markov kernels**.
#
# A **causal Markov kernel** is just another name for the probability distribution of a variable conditional on its parents in the causal DAG. The actual causal mechanism between the parents and the child determines (the word in the literature is "entails") this probability distribution. If the causal model is correct, the causal Markov kernels should be invariant across data and domain.
#
# The parameters of the causal Markov kernel are explicitly in the graph because **we are thinking like Bayesians**. In a previous lecture, we said that we should use probability to represent any uncertainty there is in elements of our "data creation myth." Generally, in probabilistic graphical models, random variables get their own nodes. So a Bayesian using graphical modeling will represent parameters as random variables and thus nodes. Explicitly modeling parameters in the graph structure allows them to use techniques from [Bayesian hierarchical modeling](https://en.wikipedia.org/wiki/Bayesian_hierarchical_modeling) to model uncertainty in these parameters.
#
# However, from our causal perspective, explicit representations of these parameters distract us from the causal relationships we are assuming in our model. We can get a view of those relationships by ignoring the Greek letters.
#
# 
#
#
# + [markdown] id="4N-4lJUHNZ2x" colab_type="text"
# ### Simple example
#
# The following shows how to implement our GMM in Pyro.
#
# Fist lets import a bunch of things, not all of which will be needed.
# + id="GqnKa7krf-SM" colab_type="code" colab={}
# To install Pyro
# #!pip3 install torch torchvision
# #!pip3 install pyro-ppl
import os
from collections import defaultdict
import torch
import numpy as np
import scipy.stats
from torch.distributions import constraints
from matplotlib import pyplot
# %matplotlib inline
import pyro
import pyro.distributions as dist
from pyro import poutine
from pyro.infer.autoguide import AutoDelta
from pyro.optim import Adam
from pyro.infer import SVI, TraceEnum_ELBO, config_enumerate, infer_discrete
smoke_test = ('CI' in os.environ)
assert pyro.__version__.startswith('1.2.0')
pyro.enable_validation(True)
# + [markdown] id="4rHqDNLwMgsO" colab_type="text"
# Next, let's specify the model. The`config_enumerate` decorator is used in inference. We don't need to worry about it for our learning purposes.
# + id="c_482mpuP9x1" colab_type="code" colab={}
K = 2
@config_enumerate
def model(N):
# Global variables.
α = 0.5
θ = pyro.sample('θ', dist.Dirichlet(α * torch.ones(K)))
σ = 1.0
σ_O = 10.
with pyro.plate('components', K):
μ = pyro.sample('μ', dist.Normal(0., σ_O))
with pyro.plate('data', N):
# Local variables.
Z = pyro.sample('Z', dist.Categorical(θ))
X = pyro.sample('X', dist.Normal(μ[Z], σ))
return {'X': X, 'Z': Z}
# + [markdown] id="2HVpjtqQK2Sw" colab_type="text"
# Notice how Pyro has a `pyro.plate` context manager that captures the "plate" abstraction in plate notation. Also notice how the tensor representation provided by a deep generative modeling framework makes it convenient to capture variables within plates as vectors.
# + [markdown] id="lFiAYBn9NbBE" colab_type="text"
# Now let's generate from the model.
# + id="zSICDB2gNeox" colab_type="code" outputId="c6afd9aa-21f6-41c8-b25c-3173defd2504" colab={"base_uri": "https://localhost:8080/", "height": 34}
model(4)
# + [markdown] id="x0o4uLSMLgMz" colab_type="text"
# # Interventions
#
# Since this is a causal model, we can apply interventions.
#
# Pyro has a `pyro.do` function that will take in a model, and return a modified model that reflects the intervention. It does this by replacing whatever sampling statement was used to generate the intervention target in the model with a statment that fixes that value to the intervention value.
#
# In the following code, I set 10 values of Z to [0, 1, 1, 0, 1, 1, 1, 1, 1, 1]. Then I generate from the model.
# + id="CZP_iy96IqPE" colab_type="code" outputId="63e11590-acbe-48df-a19b-311cc98c31ef" colab={"base_uri": "https://localhost:8080/", "height": 52}
intervention = torch.tensor([0, 1, 1, 0, 1, 1, 1, 1, 1, 1])
intervention_model = pyro.do(model, data={'Z': intervention})
intervention_model(10)
# + [markdown] id="qVZ-Y6g1N_D_" colab_type="text"
# Note the Z values are exactly what the intervention set them to. The X values are forward generated from the Z values.
# + [markdown] id="Hy2q1JAu7zAj" colab_type="text"
# # Training the Greeks
#
#
# + [markdown] id="IJtSsFXtOPRb" colab_type="text"
# In latent variable modeling, the modeler generally doesn't know the values of the Greek variables. In our case, we used probability distributions to capture that uncertainty.
#
# In practice, modelers try to infer their values from training data (i.e., values of Z's and X's). In other words, we treat the Greeks as weights in a training step.
#
# There are several ways to learn these parameters from data. Getting maximum likelihood estimates using expectation maximization is a common way. Here, since we are thinking as Bayesians, we use Bayesian inference.
#
# A Bayesian inference algorithm will treat the probability distributions we gave to the unknown Greek letters as a prior distribution. Given data, an inference algorithm will update these distributions.
#
# The following uses an [approximate Bayesian algorithm](https://en.wikipedia.org/wiki/Approximate_Bayesian_computation) called [stochastic variational inference](http://pyro.ai/examples/svi_part_i.html) (SVI). SVI makes good use of the gradient-descent based optimization infrastructure of a deep learning framework like Pyro. The following inference implementation will find [MAP estimates](https://en.wikipedia.org/wiki/Maximum_a_posteriori_estimation) of the Greek letters -- these are Bayesian analogs to maximum likelihood estimates.
#
# Do not be intimidated by the following code. This is not unlike most deep learning code you see with deep learning libraries. `TraceEnum_ELBO` and `SVI` are abstractions for stochastic variational inference. I encourage you to learn more about Bayesian inference algorithms. After all, knowledge of these algorithms tends to correlate with salary. However, in these AltDeep causal modeling courses we only need a high-level understanding of inference.
#
# + id="cg390HbW8P2g" colab_type="code" outputId="37239aed-2bbd-4a0e-d47d-4029f5878936" colab={"base_uri": "https://localhost:8080/", "height": 141}
data = torch.tensor([0., 1., 10., 11., 12.])
N = len(data)
K = 2 # Fixed number of components.
evidence_model = pyro.condition(model, data={'X': data})
optim = pyro.optim.Adam({'lr': 0.1, 'betas': [0.8, 0.99]})
elbo = TraceEnum_ELBO(max_plate_nesting=1)
def init_loc_fn(site):
if site["name"] == "θ":
# Initialize weights to uniform.
return torch.ones(K) / K
if site["name"] == "μ":
return data[torch.multinomial(torch.ones(N) / N, K)]
raise ValueError(site["name"])
def initialize(seed):
global global_guide, svi
pyro.set_rng_seed(seed)
pyro.clear_param_store()
global_guide = AutoDelta(poutine.block(evidence_model, expose=['θ', 'μ']),
init_loc_fn=init_loc_fn)
svi = SVI(evidence_model, global_guide, optim, loss=elbo)
return svi.loss(evidence_model, global_guide, N)
# Choose the best among 100 random initializations.
loss, seed = min((initialize(seed), seed) for seed in range(100))
initialize(seed)
print('seed = {}, initial_loss = {}'.format(seed, loss))
# Register hooks to monitor gradient norms.
gradient_norms = defaultdict(list)
for name, value in pyro.get_param_store().named_parameters():
value.register_hook(lambda g, name=name: gradient_norms[name].append(g.norm().item()))
losses = []
for i in range(200 if not smoke_test else 2):
loss = svi.step(N)
losses.append(loss)
print('.' if i % 100 else '\n', end='')
map_estimates = global_guide(N)
θ = map_estimates['θ']
μ = map_estimates['μ']
print('/n')
print('θ = {}'.format(θ.data.numpy()))
print('μ = {}'.format(μ.data.numpy()))
# + [markdown] id="-UYD_5ibRtAX" colab_type="text"
# Now that we have estimates for the value of our Greeks, we can replace their distributions in the model with these estimates values. An even more ideal approach would be to sample them from distributions in `pyro.distributions` that were close in shape to the posteriors of these Greeks.
#
# ## A word of caution on inference
#
# There is much to say about Bayesian inference. This is not a course on inference so I don't say much and leave it to you to experiment with various inference abstractions in Pyro.
#
# However, there are some points worth mentioning when it comes to inferring the values of "Greeks" in causal models. Firstly, getting these Greek letters right is of supreme importance in the common causal inference task of *inferring causal effects*, meaning quantifying the degree to which a cause influences an effect.
#
# The above inference algorithm assumes latent Z's, which is the usual case for GMM's. Even if our causal model were a good one, trying to train model parameters when causes like Z are latent can lead to problems when trying to estimate these causal effects accurately. We address this in the "Identification and Estimation" part of the causal modeling curriculum.
#
# Also, as a general rule, if you want an accurate estimation of the Greek variables, you should avoid approximate Bayesian algorithms in favor of exact ones (like MCMC approaches). Approximate algorithms often ignore important nonlinearities in the causal mechanisms in exchange for speed and scalability.
#
# That said, if all we care about is getting reasonably good predictions of interventions, we might be okay if we had a good causal model. Further, we could start with a basic GMM, then apply the **iterative refutation algorithm** (see lecture notes in Model-based Inference om Machine Learning) to iterate on this model. Each iteration we could retrain the model using new data from actual intervention experiments from the previous intervention, gradually overcoming estimation problems.
|
tutorials/1-introduction/3_GMM_lecture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# <NAME>
# BTECH/60036/18
# Visualize the difference between Linear,Ridge Regression,Logistic Regression with the help of a plot.Assume appropriate values for α
# -
# NOTE:
# Ridge and Lasso are two hyper-parameters of Logistic Regression. Ridge and Lasso both aim to regularize the terms. The main difference, however, is that Lasso finds the absolute value of the Beta coefficients, and Ridge squares Beta. The Lasso method punishes high values of Beta by making them 0 if they are not relevant, therefore ending up with fewer features in the model that we started with. Lasso tends to have a more significant effect than Ridge.
# +
# import all the required libraries
from sklearn.linear_model import LinearRegression, Ridge, Lasso
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from yellowbrick.regressor import PredictionError, ResidualsPlot
import pandas as pd
# +
# read and print the dataset
df = pd.read_csv('advertising.csv')
df.head()
# +
# splitting test and train data
X=df.iloc[:,0:3]
y=df.iloc[:,3]
#Scaling The data
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X)
#Splitting the data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=101)
# +
# Linear Regression
model1 = LinearRegression()
visualizer = PredictionError(model1)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# -
# A prediction error plot shows the actual targets from the dataset against the predicted values generated by our model. This allows us to see how much variance is in the model.
visualizer = ResidualsPlot(model1)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# Residuals, in the context of regression models, are the difference between the observed value of the target variable (y) and the predicted value (ŷ), i.e. the error of the prediction. The residuals plot shows the difference between residuals on the vertical axis and the dependent variable on the horizontal axis.
# +
# Lasso Regression
model2 = Lasso()
visualizer = PredictionError(model2)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# -
model2 = Lasso()
visualizer = ResidualsPlot(model2)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# +
# Ridge Regression
model3 = Ridge()
visualizer = PredictionError(model3)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
# -
model3 = Ridge()
visualizer = ResidualsPlot(model3)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
|
Lab Projects/Visualizing Linear, Logistic and Ridge Regression .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from scipy.stats import gamma
import matplotlib.pyplot as plt
import geopandas as gpd
import numpy as np
import pandas as pd
import pyreadstat as prs
x = np.linspace(0,20,100)
plt.plot(x,gamma.pdf(x,6.5/0.62));
# Carregar mapa de zonas
gdf = gpd.read_file('Mapas/Shape/Zonas_2017_region.shp')
gdf.plot(color='white', edgecolor='black');
# Mostrar apenas cidade de São Paulo
gdf[gdf['NomeMunici'] == 'São Paulo'].plot(figsize=(5, 10),color='white', edgecolor='black');
# Pesquisa Origem-Destino 2017
df, meta = prs.read_sav('Banco de dados/OD_2017.sav')
print("Número de pessoas:", len(np.unique(df['id_pess']))) # acho que é isso
print("Número de domícilio:", len(np.unique(df['id_dom'])))
print("Número de famílias:", len(np.unique(df['id_fam'])))
print("Número de viagens:", len(df))
# + jupyter={"outputs_hidden": true}
# descrição das colunas
meta.column_names_to_labels
# -
# gerar proporção de entrevistados em cada zona
#gdf['prop'] = np.random.random(size=len(gdf))
zonas = np.array(df['zona'], dtype='int')
tally=np.zeros(max(zonas),dtype='int')
for x in zonas:
tally[x-1] += 1
plt.hist(tally,bins=range(1,518));
gdf['domicilio'] = tally
# Sanity checking
tally[81], len(df[df.zona == 82]) # no. de entrevistas com domicílio em Pinheiros
tally[297], len(df[df.zona == 298]) # no. de entrevistas com domicílio em bairro que não sei o nome mas aparece como populoso no mapa abaixo
# Gerar proporções em cada zona
gdf[gdf['NomeMunici'] == 'São Paulo'].plot(column='domicilio', cmap='coolwarm', legend=True,figsize=(10, 10));
plt.title('Domícilios');
# Desenha posição de entrevistados no mapa
gdf_p = gpd.GeoDataFrame(
df, geometry=gpd.points_from_xy(df.co_dom_x, df.co_dom_y))
ax = gdf.plot(figsize=(20, 10),color='white', edgecolor='black');
# We can now plot our ``GeoDataFrame``.
gdf_p.plot(ax=ax, color='red', markersize=1, alpha=0.1);
# Desenha posição de entrevistados no mapa com "jittering" apenas para a capital
latitude = df['co_dom_x']+np.random.normal(loc=0,scale=1000,size=len(df))
longitude = df['co_dom_y']++np.random.normal(loc=0,scale=1000,size=len(df))
gdf_p = gpd.GeoDataFrame(
df, geometry=gpd.points_from_xy(df.co_dom_x, df.co_dom_y))
ax = gdf[gdf['NomeMunici'] == 'São Paulo'].plot(figsize=(20, 10),color='white', edgecolor='black');
# We can now plot our ``GeoDataFrame``.
gdf_p[gdf_p.muni_dom == 36].plot(ax=ax, color='red', markersize=0.5, alpha=0.5);
origens = np.array(df['zona_o'][~df['zona_o'].isnull()], dtype='int')
#print(np.unique(origens))
tally=np.zeros(max(origens),dtype='int')
for x in origens:
tally[x-1] += 1
plt.hist(tally,bins=range(1,518));
gdf['origem'] = tally
# Gerar proporções em cada zona
gdf[gdf['NomeMunici'] == 'São Paulo'].plot(column='origem', cmap='coolwarm', legend=True,figsize=(10, 10));
plt.title('Origem');
destinos = np.array(df['zona_d'][~df['zona_d'].isnull()], dtype='int')
#print(np.unique(origens))
tally=np.zeros(max(destinos),dtype='int')
for x in origens:
tally[x-1] += 1
plt.hist(tally,bins=range(1,518));
gdf['destino'] = tally
# Gerar proporções em cada zona
gdf[gdf['NomeMunici'] == 'São Paulo'].plot(column='destino', cmap='coolwarm', legend=True,figsize=(10, 10));
plt.title('Destino');
df.muni_dom
df[['co_dom_x','co_dom_y']]
len(df)
|
notebooks/Geopandas SP/OD2017_Denis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# <img src="https://evilcoder.ru/wp-content/uploads/2017/09/TelegramMessenger.png" width=100>
# Всем привет! Цель сегодняшнего нашего занятия – написать своего первого телеграм-бота. Это сильно проще, чем кажется :)
#
# Нам потребуется чуть более удобная IDE, например, PyCharm – скачать можно [здесь](https://www.jetbrains.com/ru-ru/pycharm/), Вам нужна community edition.
#
# Будет писать бота, который хранит списки дел. То есть, для каждого пользователя, который начинает с ним диалог, бот сможет:
# * добавлять дело в список;
# * удалять дело из списка;
# * удалять все дела из списка;
# * присылать список дел.
#
# Результат наших сегодняшних стараний доступен по нику `@MLDS_planner_bot`. По любым вопросам можно писать мне `@anastasiyamaxx`.
# ## Получаем токен
# Первое, что надо сделать – найти в поисковой строке телеграма `@BotFather`, набрать команду `/start` и создать своего бота. Вас сначала попросят выбрать имя, потом имя пользователя (должно заканчиваться на "bot") и в конце выдадут токен, по которому мы будем подключаться к API телеграма.
#
# Также в `@BotFather` можно поставить аватарку и описание нашему боту, а также обновить токен, если Вы его случайно кому-то отдали (команда `/revoke`).
# ## Подключаем бота и пишем первую функцию
# * Открываем новый проект в PyCharm
# * Работать будем в файле `main.py` – если не пользовались раньше PyCharm, то можете глянуть первую неделю курсеры, которую мы рекомендовали в начале курса :)
# Для работы с API телеграма есть специальная библиотека – `telebot`. Вам нужно ее установить. Откройте вкладку терминал в PyCharm и напишите такую строчку:
#
# `pip install pyTelegramBotAPI==3.7.9`
#
# На скрине ниже выделила как открыть терминал:
#
# <img src=imgs/pyc_terminal.png width=700>
# Будьте аккуратны! Не используйте для текущего семинара `pip install telebot`.
# Отлично! Теперь мы готовы начинать:
import telebot
from telebot import types
# подключим токен нашего бота
bot = telebot.TeleBot("вставьте сюда свой токен")
# Вообще токены обычно хранят в отдельном файле с конфигурациями – можете его создать и импортировать токен оттуда.
# Следующий наш шаг – написать первую команду!
# напишем, что делать нашему боту при команде старт
@bot.message_handler(commands=['start'])
def send_keyboard(message, text="Привет, чем я могу тебе помочь?"):
keyboard = types.ReplyKeyboardMarkup(row_width=2) # наша клавиатура
itembtn1 = types.KeyboardButton('Добавить дело в список') # создадим кнопку
itembtn2 = types.KeyboardButton('Показать список дел')
itembtn3 = types.KeyboardButton('Удалить дело из списка')
itembtn4 = types.KeyboardButton("Удалить все дела из списка")
itembtn5 = types.KeyboardButton('Другое')
itembtn6 = types.KeyboardButton('Пока все!')
keyboard.add(itembtn1, itembtn2) # добавим кнопки 1 и 2 на первый ряд
keyboard.add(itembtn3, itembtn4, itembtn5, itembtn6) # добавим кнопки 3, 4, 5 на второй ряд
# но если кнопок слишком много, они пойдут на след ряд автоматически
# пришлем это все сообщением и запишем выбранный вариант
msg = bot.send_message(message.from_user.id,
text=text, reply_markup=keyboard)
# отправим этот вариант в функцию, которая его обработает
bot.register_next_step_handler(msg, callback_worker)
# Давайте разберемся, что здесь происходит.
#
# `@bot.message_handler(commands=['start'])` обозначает, когда выполнять функцию, которая написана под ней (`send_keyboard`) в нашем случае. Такая конструкция называется декоратор и использует механизм хендлеров. `commands=['start']` значит, что `send_keyboard` будет высылаться при команде `\start`.
#
# Аналогично можно написать `@bot.message_handler(commands=['start', 'help'])`, чтобы функция вызывалась при командах `\start` и `\help`. А `@bot.message_handler(content_types=["text"])` будет вызывать функцию под ней при получении любого текстового сообщения. Можно сделать его более кастомным: `@bot.message_handler(func=lambda message: message.text == "hi")
# ` будет работать, если сообщение было словом "hi". Можно настроить его для ответов на присланные пользователем файлы, фотографии и аудио – [см. документацию](https://core.telegram.org/bots/api#available-types).
#
# `types.ReplyKeyboardMarkup` – создает нам клавиатуру, а `types.KeyboardButton('Добавить дело в список')` – кнопку. `keyboard.add(itembtn1, itembtn2)` добавляет кнопки на клавиатуру.
#
# `msg = bot.send_message(message.from_user.id, text=text, reply_markup=keyboard)` присылает пользователю клавиатуру с сообщением "Привет, чем я могу тебе помочь?". `bot.send_message(message.from_user.id,` отвечает за то, чтобы прислать сообщение в нужный чат, `text=text,` задает текст сообщения (я его задала аргументом в функции, тк буду использовать эту функцию дальше в других случаях с другим текстом). `reply_markup=keyboard` прикрепляет клавиатуру к сообщению. Выбор пользователя сохраняем в переменную `msg`.
#
# Если Вы следуете этой инструкции код параллельно с семинаром, то можете запустить код – бот будет присылать клавиатуру (но больше ничего делать не будет, т.к. функцию-обработчик мы еще не написали). Добавьте только в конце строчку `bot.polling(none_stop=True)`, мы чуть позже разберем, что она делает.
#
# `bot.register_next_step_handler(msg, callback_worker)` – отправит выбор пользователя с клавиатуры в функцию `callback_worker`, которую мы позже напишем.
#
# Кстати, вы наверняка видели и другой тип клавиатуры – inline.
#
# <img src="https://core.telegram.org/file/811140659/1/RRJyulbtLBY/ea6163411c7eb4f4dc" width=300>
#
# Почитать о ней подробнее можно [здесь](https://core.telegram.org/bots/api/#inlinekeyboardmarkup), запускается она практически аналогично.
# ## Добавляем SQLite
# Нам нужно, чтобы для каждого обратившегося к ней пользователя наша система хранила список дел. Я предлагаю воспользоваться базой данных SQLite – она быстрая (т.к. обращается напрямую к файлам, без портов и сокетов), легко встраивается в приложения и проста в использовании. Вы можете попробовать подключить более продвинутые MySQL и PostgreSQL, но это потребует чуть больше времени.
#
# Можно и просто сохранять все в обычный файлик, но помните, что такой вариант масштабировать не получится – вспомните историю, [как в Великобритании потеряли 16 тысяч положительных тестов на коронавирус.](https://www.kommersant.ru/doc/4520501)
import sqlite3
# Устанавливать ее не нужно – она встроена в питон. Можем сразу начинать работать. Подключим ее:
# +
# подключаем базу данных
conn = sqlite3.connect('planner_hse.db')
# курсор для работы с таблицами
cursor = conn.cursor()
# -
# Если базы данных с таким названием раньше не было, то она создастся, а если есть, то откроется. Теперь давайте проверим, есть ли в ней нужная нам таблица:
try:
# sql запрос для создания таблицы
query = "CREATE TABLE \"planner\" (\"ID\" INTEGER UNIQUE, \"user_id\" INTEGER, \"plan\" TEXT, PRIMARY KEY (\"ID\"))"
# исполняем его –> ура, теперь у нас есть таблица, куда будем все сохранять!
cursor.execute(query)
except:
pass
# Этот код попытается создать таблицу под названием planner, в которой есть столбец ID, который должен содержать уникальные целые числа (это будет идентификатор записи – за это отвечает конец строки `, PRIMARY KEY (\"ID\")`). Также есть столбец "user_id", который может содержать только целые числа и столбец "plan", который может содержать текст.
#
# То, что мы записали в переменную query, называется SQL запросом. Для того, чтобы понимать все, что происходит в этом семинаре с SQL, Вам хватит прочтения [20-минутного введения](https://proglib.io/p/sql-for-20-minutes).
#
# После того, как мы написали код, мы нажимаем кнопку запуска. Аналогично здесь, запрос надо исполнить – за это отвечает `cursor.execute(query)`.
#
# Если таблицы раньше не было, то эти несколько строчек ее создадут в нашей базе данных 'planner_hse.db'. Если была, то вылетит ошибка (`table already exists`), мы перейдем в код под `except`. `pass` означает, что ничего делать не надо. То есть если таблица уже есть, то ничего не произойдет – ни создание таблицы заново, ни вылет ошибки.
#
# Последнее, что хочется отметить – названия таблицы и колонок в SQL запросе должны быть в кавычках. Чтобы запихнуть кавычки внутрь строки в питоне я использую экранирование – ставлю обратный слэш перед каждой кавычкой.
# ## Пишем функции нашему боту
# В начале занятия мы сделали 6 кнопок для нашего бота:
# 1. Добавить дело в список
# 2. Показать список дел
# 3. Удалить дело из списка
# 4. Удалить все дела из списка
# 5. Другое
# 6. Пока все!
#
# Давайте по очереди напишем функции для каждой из них.
# **Добавить дело в список**
# напишем функции для каждого случая
# эта добавляет строчку с планом в хранилище
def add_plan(msg):
with sqlite3.connect('planner_hse.db') as con:
cursor = con.cursor()
cursor.execute('INSERT INTO planner (user_id, plan) VALUES (?, ?)',
(msg.from_user.id, msg.text))
con.commit()
bot.send_message(msg.chat.id, 'Запомню :-)')
send_keyboard(msg, "Чем еще могу помочь?")
# ```with sqlite3.connect('planner_hse.db') as con:
# cursor = con.cursor()```
#
# Присоединимся к базе данных – внутри функций это лучше делать так, иначе будет ругаться, что мы используем коннекшны, созданные в разных запусках.
# `cursor.execute('INSERT INTO planner (user_id, plan) VALUES (?, ?)', (msg.from_user.id, msg.text)`
#
# С этим концептом мы уже знакомы – здесь исполняется SQL запрос. Мы вставляем в таблицу planner в столбцы user_id, plan значения, которые встанут на место вопросов – msg.from_user.id, msg.text (id пользователя и текст, соответственно).
#
# `con.commit()` сохраняет изменения в базе данных.
#
# `bot.send_message(msg.chat.id, 'Запомню :-)')` присылает сообщение. Если после этого ничего не прислать, то мы, получается, будем игнорировать пользователя. Давайте ему пришлем клавиатуру и спросим, что еще он бы хотел поделать: `send_keyboard(msg, "Чем еще могу помочь?")` (мы воспользовались функцией, которую написали в начале).
# **Показать список дел**
# +
# просто функция, которая делает нам красивые строки для отправки пользователю
def get_plans_string(tasks):
tasks_str = []
for val in list(enumerate(tasks)):
tasks_str.append(str(val[0] + 1) + ') ' + val[1][0] + '\n')
return ''.join(tasks_str)
# отправляем пользователю его планы
def show_plans(msg):
with sqlite3.connect('planner_hse.db') as con:
cursor = con.cursor()
cursor.execute('SELECT plan FROM planner WHERE user_id=={}'.format(msg.from_user.id))
tasks = get_plans_string(cursor.fetchall())
bot.send_message(msg.chat.id, tasks)
send_keyboard(msg, "Чем еще могу помочь?")
# -
# Разберем сначала вторую функцию. Мы снова присоеднились к базе данных, выполнили SQL запрос, который выбрал значения только из столбца plan из таблицы planner (`SELECT plan FROM planner`), где id пользователя равен id того, кто прислал нам сообщение (`WHERE user_id=={}'.format(msg.from_user.id)`).
#
# Чтобы получить результаты запроса используем `cursor.fetchall()`. Он вернет нам данные в формате `[(дело1,), (дело2,), (дело3,)]`. Напишем функцию помощник, которая превратит это в красивую запись в формате (ее мы разбирать не будем, т.к. тут все знакомое):
# ```1) дело1
# 2) дело2
# 3) дело3```
# Осталось отправить красивое сообщение, где будет эта строка и клавиатуру с нашим меню заново.
# **Удалить дело из списка**
# +
# выыделяет одно дело, которое пользователь хочет удалить
def delete_one_plan(msg):
markup = types.ReplyKeyboardMarkup(row_width=2)
with sqlite3.connect('planner_hse.db') as con:
cursor = con.cursor()
# достаем все задачи пользователя
cursor.execute('SELECT plan FROM planner WHERE user_id=={}'.format(msg.from_user.id))
# достанем результат запроса
tasks = cursor.fetchall()
for value in tasks:
markup.add(types.KeyboardButton(value[0]))
msg = bot.send_message(msg.from_user.id,
text = "Выбери одно дело из списка",
reply_markup=markup)
bot.register_next_step_handler(msg, delete_one_plan_)
# удаляет это дело
def delete_one_plan_(msg):
with sqlite3.connect('planner_hse.db') as con:
cursor = con.cursor()
cursor.execute('DELETE FROM planner WHERE user_id==? AND plan==?', (msg.from_user.id, msg.text))
bot.send_message(msg.chat.id, 'Ура, минус одна задача!')
send_keyboard(msg, "Чем еще могу помочь?")
# -
# Для того, чтобы удалить дело из списка, нам нужно реализовать такую последовательность действий:
# * Обратимся к базе данных и достанем все дела для данного пользователя (по аналогии с предыдущим пунктом)
# * Выведем все его дела на клавиатуру, спросим какое нужно удалить, сохраним выбор
# * Удалим его
#
# Все команды в первой функции нам уже знакомы, разве что кнопки добавляем теперь циклом по всем делам пользователя:
#
#
# ```
# tasks = cursor.fetchall()
# for value in tasks:
# markup.add(types.KeyboardButton(value[0]))
# ```
# Напомню, что tasks внутри выглядит примерно так: `[(дело1,), (дело2,), (дело3,)]`.
#
# То есть на каждой итерации value будет принимать значения типа кортеж с одной строкой внутри: `(дело1,),`. Поэтому мы берем `value[0]` (только текст) и добавляем его на кнопку `types.KeyboardButton(value[0]))`, а кнопку новым рядом на клавиатуру `markup.add(types.KeyboardButton(value[0]))`.
# Во второй функции мы также подключаемся к базе данных, добавляем условие, что plan должен быть равен msg.text (т.к. выбранная на клавиатуре кнопка присылается как текстовое сообщение от пользователя). Не забываем отправить сообщение, что дело удалено, и мы готовы помогать пользователю дальше.
# **Удалить все дела из списка**
# удаляет все планы для конкретного пользователя
def delete_all_plans(msg):
with sqlite3.connect('planner_hse.db') as con:
cursor = con.cursor()
cursor.execute('DELETE FROM planner WHERE user_id=={}'.format(msg.from_user.id))
con.commit()
bot.send_message(msg.chat.id, 'Удалены все дела. Хорошего отдыха!')
send_keyboard(msg, "Чем еще могу помочь?")
# Тут также все нам уже знакомо. `DELETE FROM planner` удаляет все строки таблицы, где выполняются условия, которые идут после `WHERE`.
# Самое сложное позади. Давайте теперь соберем это все воедино. Помните, как в начале занятия мы собирались написать функцию, которая все обработает?
#
# ```
# # отправим этот вариант в функцию, которая его обработает
# bot.register_next_step_handler(msg, callback_worker)
# ```
#
# Пришло ее время!
# привязываем функции к кнопкам на клавиатуре
def callback_worker(call):
if call.text == "Добавить дело в список":
msg = bot.send_message(call.chat.id, 'Давайте добавим дело! Напишите его в чат')
bot.register_next_step_handler(msg, add_plan)
elif call.text == "Показать список дел":
try:
show_plans(call)
except:
bot.send_message(call.chat.id, 'Здесь пусто. Можно отдыхать :-)')
send_keyboard(call, "Чем еще могу помочь?")
elif call.text == "Удалить дело из списка":
try:
delete_one_plan(call)
except:
bot.send_message(call.chat.id, 'Здесь пусто. Можно отдыхать :-)')
send_keyboard(call, "Чем еще могу помочь?")
elif call.text == "Удалить все дела из списка":
try:
delete_all_plans(call)
except:
bot.send_message(call.chat.id, 'Здесь пусто. Можно отдыхать :-)')
send_keyboard(call, "Чем еще могу помочь?")
elif call.text == "Другое":
bot.send_message(call.chat.id, 'Больше я пока ничего не умею :-(')
send_keyboard(call, "Чем еще могу помочь?")
elif call.text == "Пока все!":
bot.send_message(call.chat.id, 'Хорошего дня! Когда захотите продолжнить нажмите на команду /start')
# Простая if-else конструкция, ничего нового. В некоторые пункты я добавила конструкцию try-except на случай, если планов пользователя в нашей таблице нет (наши функции, написанные ранее, могут кидать ошибки на такое).
#
# Осталась лишь пара штрихов. Когда я кинула этот бот первым пользователям потестить, они начали писать туда текстовые сообщения или как-то еще ломать алгоритм. Например, если на вопрос "Чем я еще могу помочь?" с прикрепленной клавиатурой ответить не вариантом с клавиатуры, а другим сообщением, то бот замолкал. Давайте допишем функцию, которая на любое текстовое сообщение будет заново кидать клавиатуру:
@bot.message_handler(content_types=['text'])
def handle_docs_audio(message):
send_keyboard(message, text="Я не понимаю :-( Выберите один из пунктов меню:")
# И последняя строчка:
bot.polling(none_stop=True) # или bot.infinity_polling()
# Это значит, что наш бот постоянно спрашивает у телеграма нет ли для него новых сообщений. Если Вы добавите эту строчку в Ваш код в PyCharm и запустите код (зеленый треугольник в правом верхнем углу), то Ваш бот заработает! НО работать он будет только если Ваш код запущен – то есть при выключении компьютера или закрытии программы бот отвечать перестанет.
# ## Деплой
# Чтобы Ваш бот работал без постоянно запущенной программы, мы должны его где-то развернуть. Очень простой и приятный сервис для этого – [heroku](https://www.heroku.com). Для небольших по количеству пользователей ботов он будет работать бесплатно. Документация [здесь](https://devcenter.heroku.com/articles/getting-started-with-python).
#
# Для начала работы с ним нужно зарегистрироваться на сайте (не забудьте подтвердить почту). Вам также потребуется установить git, если не делали этого ранее, [вот ссылка](https://git-scm.com/downloads).
#
# А также установить heroku. На маке легко это сделать с помощью brew: `brew install heroku`. Для винды лучше скачать установщик [отсюда](https://devcenter.heroku.com/articles/heroku-cli).
# После того, как вы зарегистрировались на сайте heroku и скачали git, возвращайтесь в PyCharm. Откройте терминал в нем, напишите команду `heroku login`. После этого он предложит нажать любую кнопку для подтверждения Ваших намерений, Вы нажимаете, открывается окно в браузере, где надо залогиниться на сайте. Делайте это и возвращайтесь в PyCharm.
# Теперь нам нужно создать 2 файлика: Procfile и requirements.txt.
#
# Чтобы создать первый нажимаем в правом верхнем углу File, потом New. Должно появиться что-то как на картинке ниже:
#
# <img src=imgs/pyc_file.png width=300>
#
# Нужно выбрать первый вариант. Важно: нам нужно сделать файл без расширения.
#
# В этот файл вставляем такую строку: `worker: python main.py`
#
# Теперь создадим второй файл. Это сделать просто – откройте терминал в PyCharm опять и напишите `pip freeze > requirements.txt`.
# Возвращаемся в терминал. Надо ввести несколько команд:
#
# 1. `git init` – инициализирует гитхаб репозиторий
# 2. `heroku git:remote -a название_проекта` – создаст удаленный репозиторий и назовет приложение (если не работает, то создайте приложение вручную [тут](https://dashboard.heroku.com/apps) и вернитесь к этому шагу)
# 3. `git add .` – добавляет файлы
# 4. `git commit -am "make it better"` – делает коммит (можете менять подпись)
# 5. `git push heroku master` – отправляет коммит в репозиторий (после этого лучше подождать минутку-другую)
# 6. `heroku ps:scale worker=1` – запускает worker dyno (так heroku называет контейнеры, мы используем бесплатную версию, подробней [тут](https://www.heroku.com/dynos))
#
# Все, победа! Бот должен заработать (иногда нужно подождать пару минут).
#
# В дальнейшем, если Вы будете менять код и дополнять функционал бота, Вам нужно будет делать только пункты 3-6 из списка выше, чтобы обновить бот.
#
# Дашборды со всякой информацией по Вашему приложению можно смотреть тут: https://dashboard.heroku.com/apps
# ## Задание
#
# Если у Вас осталось время в конце занятия, допишите пару функций боту в раздел "Другое". Например, можно уточнить другую информацию о пользователе, прислать ему ссылки на последние новости и т.д.
# ## Дополнительное чтение
#
# Туториалов и уроков на тему ботов – тьма. В принципе можете брать любой на хабре (если любите читать) или ютубе (если любите смотреть). Но вот несколько, что показались мне полезными:
#
# 1. [Очень классный и большой набор уроков, телеграм боты от А до Я](https://mastergroosha.github.io/telegram-tutorial/)
# 2. [Как сделать бота с базой данных MySQL](https://www.youtube.com/playlist?list=PLvpsu6rFEzwUtzK6qf9wyDsAzM-sdtHTT)
# 3. [Милое вдохновение для грядущей домашки – бот, который присылает дешевые билеты в театр](https://habr.com/ru/post/445632/)
# 4. [Как деплоить с хероку – похоже на наш семинар, но есть пара неточностей](https://www.youtube.com/watch?v=B72-sZyiW10&lc=UgxmpAY651efoK6fGgd4AaABAg.9B7tj_dFDQq9B7vksC6WGf&ab_channel=%D0%92%D0%BB%D0%B0%D0%B4%D0%9D%D0%B5%D0%B2%D0%B5%D1%80%D0%BE%D0%B2)
|
class_6/bot_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Financial and technology articles taken from [webhose.io](https://webhose.io/datasets)
import pandas as pd
import json,urllib2
import glob
# %matplotlib inline
# ## Take a look at one JSON file
with open('data/news_0000001.json','r') as inFile:
d=json.loads(inFile.read())
print d.keys()
print d['text']
# ## Define a function to open a file and get the text
def getText(f):
with open(f,'r') as inFile:
d=json.loads(inFile.read())
return d['text']
# %time financeTexts=map(getText,glob.glob('../financial_news/data/news_*json'))
len(financeTexts)
# %time techTexts=map(getText,glob.glob('../tech_news/data/news*json'))
len(techTexts)
# ## Combine tech and financial news into one dataframe
df=pd.DataFrame(data={'text':financeTexts,'category':'finance'})
df=df.append(pd.DataFrame(data={'text':techTexts,'category':'tech'}))
df.head()
df.shape
df.head()
# ## Build up a pipeline
# +
from sklearn.feature_extraction.text import TfidfTransformer,CountVectorizer
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier,LogisticRegression
from sklearn.naive_bayes import MultinomialNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import f1_score
from sklearn import preprocessing
# -
# ## Binarise the category labels
lb = preprocessing.LabelBinarizer()
lb.fit(df['category'])
df['category_bin']=lb.transform(df['category'])
# ## Test Naive Bayes Classifier fr our baseline
steps=[('vectorise',CountVectorizer()),\
('transform',TfidfTransformer()),\
('clf',MultinomialNB())]
# Our pipeline has three steps
pipe=Pipeline(steps)
X_train, X_test, y_train, y_test=\
train_test_split(df['text'],df['category_bin'],test_size=0.25)
pipe.fit(X_train,y_train)
pred=pipe.predict(X_test)
print 'Accuracy = %.3f' % f1_score(y_test,pred)
# ## Write out model
import pickle
with open('model.out','w') as outFile:
pickle.dump(pipe,outFile)
lb
# ## Video 4.3
# ## Grid Search
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
pipe.named_steps.keys()
param_grid = dict(vectorise__stop_words=[None,'english'],\
vectorise__binary=[True,False],\
#vectorise__min_df=[1,5,10],\
#clf__class_weight=[None,'balanced'],\
#transform__norm=['l1','l2']
)
grid_search = GridSearchCV(pipe, param_grid=param_grid,\
scoring=make_scorer(f1_score),n_jobs=2)
# With n_jobs=1, takes 10.33
# With n_jobs=-1 takes YYYs
# %time res=grid_search.fit(df['text'],df['category_bin'])
res.best_params_
print 'Best score = %.3f' % res.best_score_
# ## Compare classifiers
CountVectorizer()
for clf in [SGDClassifier(),LogisticRegression(),RandomForestClassifier()]:
print clf.__class__
steps=[('vectorise',CountVectorizer()),('transform',TfidfTransformer()),\
('clf',clf)]
pipe=Pipeline(steps)
pipe.set_params(vectorise__decode_error='ignore')
grid_search = GridSearchCV(pipe, param_grid=param_grid,n_jobs=-1,\
scoring=make_scorer(f1_score))
res=grid_search.fit(df['text'],df['category_bin'])
print 'Best score = %.3f' % res.best_score_
print res.best_params_
print ''
|
Section 4/4.2_news_section_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/B-Lorentz/physics-bsc-visual/blob/main/15_Quantum_Statistics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="valPqXjUN0Jb"
import numpy as np
from matplotlib import pyplot as plt
from jax import numpy as jnp
from jax import grad, vmap, jit, random, lax, ops
from matplotlib.widgets import Slider, Button, RadioButtons
from ipywidgets import interact, widgets
# + [markdown] id="aGI0OrcbIxpT"
# # Quantum Statistics
#
# ### Ideal quantum gases: Bose-Einstein and Fermi-Dirac statistics
# + [markdown] id="gVpBpQjfJF9d"
# A key property of quantum mechanics is that identical particles are considered "the same": There are no labels attached. But our wavefunctions distinguish them merely by having variables specified in an ordered manner: $$\Psi(x_1, x_2, x_3,...)$$
# (Here $x$ can include spin or any other descriptors)
#
# To remedy this, the wavefunction must be either symmetric or anti-symmetric in the interchange of particles:
# - symmetric: $\Psi(x_1, x_2, x_3,...) = \Psi(x_2, x_1, x_3,...) = \Psi(x_2, x_3, x_1,...) = ...$
#
# - anti-symmtteric: $\Psi(x_1, x_2, x_3,...) = -\Psi(x_2, x_1, x_3,...) = \Psi(x_2, x_3, x_1,...) = ...$
#
# This way repeated interchange of two perticles is the identity as expected ($1 = 1\cdot1 = (-1)\cdot(-1)$), but since the probabilities are given by $|\Psi|^2$, the physical state is unchanged under permutation even in the antisymmetric case. ($\cdot(-1)$ is just a phase change by $\pi$ )
#
# It can be shown that whether the symmetric or the anti-symmetric case is in force depends only on the particle type. Particles whose wavefunction is symmetric to interchange are called bosons, and the anti-symmetric ones fermions.
#
# It is an additional important fact (the spin-statistics theorem) that bosons have integer spins and fermions half-integer spins. This is however not part of their definition, but a separate and notrivial result that can be arrived at from relativistc field theory.
# + [markdown] id="AHjiQW3QPH-x"
# If the particles are non-interacting (their Hamiltionian is the sum of the individual Hamiltionians) we can construct states with the required symmetry by linearly combining products of one-particle states:
#
# - For bosons all permutations are simply added together. Two particles: $\Psi(x_1, x_2) = \Psi_1(x_1)\Psi_2(x_2) + \Psi_1(x_2)\Psi_2(x_1)$
#
# - For fermions we multiply them by $-1^p$ where $p$ is the number of permutations needed from the natural ordering. Two particles: $\Psi(x_1, x_2) = \Psi_1(x_1)\Psi_2(x_2) - \Psi_1(x_2)\Psi_2(x_1)$. Therefore if $\Psi_1 \equiv \Psi_2$, the wavefunction is zero everywhere, prohibiting two fermions from ever assuming the same quantum state. THis is called the Pauli exclusion principle. (This generalized to $N$ particles where the wavefunction can be written asa determinat formed from the one-particle wavefunctions)
#
# A demonstration for two particles in a 1D infinite potential well:
# + colab={"base_uri": "https://localhost:8080/", "height": 600, "referenced_widgets": ["ac3244e8591749da8f879e75fa7d92bf", "f7bf55f4a04c4a46889b2f181f61f0a6", "ba43a0b180674e46abf01131cff5786b", "50b54f23445b4edd9b0b396295c9253c", "8da0f7235fa14c0f8ba1dbbef933747b", "cd2a6a1b0f0d4421b7c1e1e199a22ed2", "a153d859338345979c7ab30f384eed00", "4d136457dd8241469253a424ed81d3d6", "f71570ae9ff64db5b0589ab6f52c9f67", "8a50e4e36d9249509510e916847f2bf7"]} id="VuzAQdFlB1jN" outputId="9db75ea5-de66-452f-dcff-a2525c3bdecd"
x1 = x2 = np.linspace(0, 1, 100)
def ploti(name, f):
x1s, x2s = np.meshgrid(x1, x2)
plt.contourf(x1, x2, f(x1s, x2s), levels=50,cmap=plt.get_cmap("coolwarm"))
plt.plot([0,1],[0,1], "black")
#plt.plot(x1, psi1(x1))
plt.xlabel("$x_1$", fontsize=14)
plt.xlabel("$x_2$", fontsize=14)
plt.title(name, fontsize=16)
def interplot(n1, n2):
psi1 = lambda x: jnp.sin(n1*np.pi*x)
psi2 = lambda x: jnp.sin(n2*np.pi*x)
plt.figure(figsize=(16, 8))
plt.subplot(121)
ploti("Bozon", lambda x1, x2: psi1(x1)*psi2(x2) + psi1(x2)*psi2(x1))
plt.subplot(122)
ploti("Fermion", lambda x1, x2: psi1(x1)*psi2(x2) - psi1(x2)*psi2(x1))
interact(interplot, n1=widgets.IntSlider(1, 1, 10), n2=widgets.IntSlider(3, 1, 10))
# + [markdown] id="nUIWAnKaSHZ5"
# To do statistical physics with non-interaction identical particles - that is, to study an ideal gas made of fermions or bosons - we use the occupation representation: For each quantum state with energy $\epsilon_i$ we record only the number of particles $n_i$, and then compute the grand canonical partition function as $$\mathcal{Z} = \sum_{n_1}\sum_{n_2}...\prod_{k=1}^{\infty}e^{-\beta n_k(\epsilon_k -\mu)}$$
#
# For bosons $n_i\in\mathbb{N}$, while for fermions $n_i \in \{0, 1\}$, encoding the only difference between them, the Pauli exclusion.
#
# This can be computed by cleverly interchanging the sum and the product and solving the sum (which is trivial for the fermions and a geometric series for bosons, that converges if $\mu<\epsilon_0$).
#
# $\langle n_i \rangle$ van be computed by noticing that in:
# $$\langle n_i \rangle = \frac{1}{\mathcal{Z}}\sum_{n_1}\sum_{n_2}...n_i\prod_{k=1}^{\infty}e^{-\beta n_k(\epsilon_k -\mu)}$$ the respective contributions of all energy levels other than the $i$th are destroyed by the same appearing in $\mathcal{Z}$.
#
# The result is:
#
# - $\langle n_i \rangle = f_B(\epsilon_i) = \frac{1}{e^{\beta(\epsilon_i - \mu) }-1}$ for bosons.
#
# - $\langle n_i \rangle = f_F(\epsilon_i) = \frac{1}{e^{\beta(\epsilon_i - \mu) }+1}$ for fermions.
#
# The following interactive plots illustrate the shape of these two functions, and also show how there is nothing more ongoing than grand-canonical formalism in occupancy representation, as I show an approximate computation where I simply sample {n_i}-s (like (n_1=1, n_2=3, n_3=0 ...)) using a Monte-Carlo walk with grand canonical probabilities.
# + [markdown] id="9Kn4Gkg5YiFV"
# ### High temperature limit
#
# For a sytem with a fixed number ($N$) of particles (like a gas in a box, but not a solid with phonons or a box with photons) $\mu$ is given implicitly by:
# $$N=\sum_i f(\epsilon_i, \mu)= \sum_i \frac{1}{e^{\beta(\epsilon_i - \mu) }\pm1}$$
#
# If $T \to \infty \iff \beta\to0$
#
# therefore $\mu<<0 \implies e^{\beta(\epsilon_i - \mu)} >> 1$
#
# Bosons and fermions all comverge to the Boltzmann-distribution.
# + id="3eMBTqXVyq_7"
def MCn(Es, beta, mu, stat):
N_es = Es.shape[0]
efun = lambda ns: (ns*(Es-mu)).sum()
def mcf(key):
def upd(i, state):
ns, sum_ns, key = state
k1, k2, k3, k4 = random.split(key, 4)
inds = random.choice(k1, N_es, shape=(N_es//20,), replace=False)
ns2 = ns.at[inds].set(stat.proposer(k2, ns[inds]))
e0 = efun(ns)
e1 = efun(ns2)
dec = jnp.logical_or(e1<e0, jnp.exp(-beta*(e1-e0)) > random.uniform(k3, (1,)))
ns = lax.cond(dec[0], lambda x:ns2, lambda x:x, ns)
sum_ns = lax.cond(i%5 == 4, lambda x: x+ns, lambda x:x, sum_ns)
return (ns, sum_ns, k4)
k1, k2 = random.split(key)
init = (stat.proposer(k1, np.zeros(N_es, dtype=int)), np.zeros(N_es, dtype=int), k2)
res = lax.fori_loop(0, 1000, upd, init)[1]
return res
return mcf
class Stat:
def __init__(self, name, f, proposer):
self.name = name
self.f = f
self.proposer = proposer
fermi = Stat("F", lambda E, beta, mu : 1/(np.exp(beta*(E-mu))+1), lambda key, ns: random.choice(key, 2, shape=ns.shape ))
# + colab={"base_uri": "https://localhost:8080/", "height": 582, "referenced_widgets": ["e64aeb7818684bd394e86862c022e58f", "4a468a1292ee4150b2b7383633f7a41d", "43395a8d91ce4ace98b79345a1fd0900", "8184d90d9ae248dc9e8de1700afe3b92", "39f6e81d638740868fc7c37482158e09", "a37cb29c99e64ff3ba7686ff6afbca8a", "6b437758cf06477d89128017737df09f", "1c78a26099f4445b9ec331d039a09447", "bb360291e6194da580e0b1f15efeaff0", "682e455856a944b1a73096ffc84632ff"]} id="BUFYMO3pOGcW" outputId="ba38b4bf-04d2-4907-ef70-037852b745b8"
N_stat = 100
Es = np.linspace(0, 1.0, N_stat)
def plotbuilder(stat):
def plotter(beta, mu):
plt.figure(figsize=(10, 8))
ns = stat.f(Es, beta, mu)
plt.plot(Es, ns, label=f"$f_{stat.name}(\epsilon)$")
bns = np.exp(-beta*Es)
plt.plot(Es, bns*ns.max(), label="Boltzmann")
plt.plot([mu, mu], [0, ns.max()], "r", label=r"$\mu$")
plt.xlabel("$\epsilon$", fontsize=14)
plt.ylabel("<n>", fontsize=14)
plt.grid()
mcs = vmap(MCn(Es, beta, mu, stat), 0)(random.split(random.PRNGKey(127), 100))
res, resd = mcs.mean(0), mcs.std(0)
maxi = res.max()/ns.max()
#print(res.shape, res.min(), res.max())
plt.errorbar(Es, res/maxi, resd/maxi)
plt.title(r"$\beta =" +f"{beta:.2f}$")
plt.legend(fontsize=14)
return plotter
interact(plotbuilder(fermi), beta=widgets.FloatSlider(min=1.0, max=30, value=3.5), mu=widgets.FloatSlider(min=-1, max=1, value=0.1, step=0.05))
None
# + id="EMG4Cu69QT7y" colab={"base_uri": "https://localhost:8080/", "height": 582, "referenced_widgets": ["2256494e566f42979979e967a759d1ef", "7208ffde97c44a229a4c9fac74ccfd4e", "0fca330f12774b39be801fb07d32120b", "5375a037f0314ffbae89b6984d3ea63b", "d8a9bf9315ca426f81655981920071c2", "4fc0dc24eedc4ec9bff9e9d77f0322ea", "8c1aa535a7fe4cd3a85149c80f059360", "<KEY>", "2d777d06bc4843b8b23992b2534b5c7b", "1f59f6844d7544f5a6957ba88b8c13ac"]} outputId="22fabba7-3523-4d59-c5ea-15ea8ada4b8e"
bose = Stat("B", lambda E, beta, mu : 1/(np.exp(beta*(E-mu))-1), lambda key, ns: jnp.maximum((random.normal(key, ns.shape)*jnp.maximum(ns, 10)/2).astype(int)+ns, 0) )
interact(plotbuilder(bose), beta=widgets.FloatSlider(min=1.0, max=10, value=1.5), mu=widgets.FloatSlider(min=-1, max=-0.15, value=-0.05, step=0.05))
None
# + [markdown] id="K_ywlELOIvMa"
# ## Example: electrons in metals
#
# $$D(\epsilon) = 4\pi V (\frac{2 m}{h^2})^{3/2} \sqrt{e}$$
#
# at T=0:
#
# $$N = \int_0^{\epsilon_F}D(\epsilon)d\epsilon$$
#
# $$\epsilon_F = \frac{h^2}{2m}\left(\frac{3 N}{8\pi V}\right)^{\frac{2}{3}}$$
#
# $$E = \int_0^\infty\epsilon f(\epsilon) D(\epsilon)d \epsilon \approx \int_0^{\epsilon_F}\epsilon D(\epsilon)d\epsilon$$
#
# A Sommerfeld-series would give a better approximation
# + id="5XjJUuDpv8z2" colab={"base_uri": "https://localhost:8080/", "height": 550, "referenced_widgets": ["82a3a372e19a473a9a0f019c0effad38", "4b420b90f3224dff8693e79833af1a6b", "edf288d87f964a198970064191ac2e96", "b3ef513ed5cd48169afd5cdc1cafa544", "<KEY>", "73ca7c537d6e497cb3150d85c14b17e1", "aa6c3d6a10d047d3b51afa67b39c000e"]} outputId="dfeb18b2-559c-4be1-9869-fb1413210328"
def plotbuilder2(stat):
N_stat = 300
Es = np.linspace(0, 2.0, N_stat)
def plotter(beta):
mu=1.0
plt.figure(figsize=(10, 8))
ns = stat.f(Es, beta, mu)*np.sqrt(Es)
plt.plot(Es, ns, label="")
bns = np.exp(-beta*Es)
plt.plot([mu, mu], [0, ns.max()], "r", label=r"$\mu$")
plt.xlabel("$\epsilon$", fontsize=14)
plt.ylabel(r"$D(\epsilon)\cdot f(\epsilon)$", fontsize=14)
plt.grid()
plt.title(r"$\beta =" +f"{beta:.2f}$")
plt.legend(fontsize=14)
return plotter
interact(plotbuilder2(fermi), beta=widgets.FloatSlider(min=1.0, max=100, value=10))
None
# + id="HspwKgLUx6MV"
|
15_Quantum_Statistics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#RNN-à-partir-de-caractères" data-toc-modified-id="RNN-à-partir-de-caractères-1"><span class="toc-item-num">1 </span>RNN à partir de caractères</a></span></li><li><span><a href="#Imports" data-toc-modified-id="Imports-2"><span class="toc-item-num">2 </span>Imports</a></span></li><li><span><a href="#Préparation-des-données" data-toc-modified-id="Préparation-des-données-3"><span class="toc-item-num">3 </span>Préparation des données</a></span></li><li><span><a href="#Création-du-RNN" data-toc-modified-id="Création-du-RNN-4"><span class="toc-item-num">4 </span>Création du RNN</a></span></li><li><span><a href="#Création-de-votre-propre-modèle-RNN" data-toc-modified-id="Création-de-votre-propre-modèle-RNN-5"><span class="toc-item-num">5 </span>Création de votre propre modèle RNN</a></span></li><li><span><a href="#Training" data-toc-modified-id="Training-6"><span class="toc-item-num">6 </span>Training</a></span></li><li><span><a href="#Évaluation" data-toc-modified-id="Évaluation-7"><span class="toc-item-num">7 </span>Évaluation</a></span></li><li><span><a href="#Visualizing-memorization" data-toc-modified-id="Visualizing-memorization-8"><span class="toc-item-num">8 </span>Visualizing memorization</a></span></li></ul></div>
# -
# # RNN à partir de caractères
# Dans ce TP, nous allons définir notre propre RNN en créant une classe RNN qui hérite de l'objet nn.Module de PyTorch.
# # Imports
# +
from __future__ import unicode_literals, print_function, division
from io import open
import glob
import os
import random
import unicodedata
import string
import time
import math
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# -
import torch
import torch.nn as nn
from torch.autograd import Variable
import torch.autograd
import numpy as np
# # Préparation des données
# +
def findFiles(path): return glob.glob(path)
print(findFiles('data/names/*.txt'))
all_letters = string.ascii_letters + " .,;'"
n_letters = len(all_letters)
def unicodeToAscii(s):
return ''.join(
c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn'
and c in all_letters
)
print(unicodeToAscii('Ślusàrski'))
# Build the category_lines dictionary, a list of names per language
category_lines = {}
all_categories = []
# Read a file and split into lines
def readLines(filename):
lines = open(filename, encoding='utf-8').read().strip().split('\n')
return [unicodeToAscii(line) for line in lines]
for filename in findFiles('data/names/*.txt'):
category = os.path.splitext(os.path.basename(filename))[0]
all_categories.append(category)
lines = readLines(filename)
category_lines[category] = lines
n_categories = len(all_categories)
# -
print(category_lines['Italian'][:5])
# +
# Find letter index from all_letters, e.g. "a" = 0
def letterToIndex(letter):
return all_letters.find(letter)
# Just for demonstration, turn a letter into a <1 x n_letters> Tensor
def letterToTensor(letter):
tensor = torch.zeros(1, n_letters)
tensor[0][letterToIndex(letter)] = 1
return tensor
# Turn a line into a <line_length x 1 x n_letters>,
# or an array of one-hot letter vectors
def lineToTensor(line):
tensor = torch.zeros(len(line), 1, n_letters)
for li, letter in enumerate(line):
tensor[li][0][letterToIndex(letter)] = 1
return tensor
print(letterToTensor('J'))
print(lineToTensor('Jones').size())
# -
# # Création du RNN
# +
class RNN(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(RNN, self).__init__()
self.hidden_size = hidden_size
self.rnn = nn.RNN(input_size, hidden_size)
self.linear = nn.Linear(hidden_size, output_size)
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, input, hidden):
output, _ = self.rnn(input, hidden)
output = self.linear(output)
return self.softmax(output)
def initHidden(self):
return torch.zeros(1, self.hidden_size)
n_hidden = 128
rnn = RNN(n_letters, n_hidden, n_categories)
# -
input = letterToTensor('A')
input.unsqueeze_(1)
hidden =torch.zeros(1, 1, n_hidden)
print(input.size(), hidden.size())
output = rnn(input, hidden)
print(output.size())
# # Création de votre propre modèle RNN
# <img src="images/rnn_a_definir.png" width=300>
# +
class RNN(nn.Module):
def __init__(self, input_size, hidden_size, output_size):
super(RNN, self).__init__()
self.hidden_size = hidden_size
self.i2h = nn.Linear(input_size + hidden_size, hidden_size)
self.i2o = nn.Linear(input_size + hidden_size, output_size)
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, input, hidden):
combined = torch.cat((input, hidden), 1)
hidden = self.i2h(combined)
output = self.i2o(combined)
output = self.softmax(output)
return output, hidden
def initHidden(self):
return torch.zeros(1, self.hidden_size)
n_hidden = 128
rnn = RNN(n_letters, n_hidden, n_categories)
# -
input = letterToTensor('A')
hidden =torch.zeros(1, n_hidden)
print(input.size(), hidden.size())
output, next_hidden = rnn(input, hidden)
# +
input = lineToTensor('Albert')
hidden = torch.zeros(1, n_hidden)
output, next_hidden = rnn(input[0], hidden)
print(output)
# -
# # Training
# +
def categoryFromOutput(output):
top_n, top_i = output.topk(1)
category_i = top_i[0].item()
return all_categories[category_i], category_i
print(categoryFromOutput(output))
# +
def randomChoice(l):
return l[random.randint(0, len(l) - 1)]
def randomTrainingExample():
category = randomChoice(all_categories)
line = randomChoice(category_lines[category])
category_tensor = torch.tensor([all_categories.index(category)], dtype=torch.long)
line_tensor = lineToTensor(line)
return category, line, category_tensor, line_tensor
for i in range(10):
category, line, category_tensor, line_tensor = randomTrainingExample()
print('category =', category, '/ line =', line)
# -
criterion = nn.NLLLoss()
# +
learning_rate = 0.005 # If you set this too high, it might explode. If too low, it might not learn
def train(category_tensor, line_tensor):
hidden = rnn.initHidden()
rnn.zero_grad()
for i in range(line_tensor.size()[0]):
output, hidden = rnn(line_tensor[i], hidden)
loss = criterion(output, category_tensor)
loss.backward()
# Add parameters' gradients to their values, multiplied by learning rate
for p in rnn.parameters():
p.data.add_(-learning_rate, p.grad.data)
return output, loss.item()
# +
n_iters = 100000
print_every = 5000
plot_every = 1000
# Keep track of losses for plotting
current_loss = 0
all_losses = []
def timeSince(since):
now = time.time()
s = now - since
m = math.floor(s / 60)
s -= m * 60
return '%dm %ds' % (m, s)
start = time.time()
for iter in range(1, n_iters + 1):
category, line, category_tensor, line_tensor = randomTrainingExample()
output, loss = train(category_tensor, line_tensor)
current_loss += loss
# Print iter number, loss, name and guess
if iter % print_every == 0:
guess, guess_i = categoryFromOutput(output)
correct = '✓' if guess == category else '✗ (%s)' % category
print('%d %d%% (%s) %.4f %s / %s %s' % (iter, iter / n_iters * 100, timeSince(start), loss, line, guess, correct))
# Add current loss avg to list of losses
if iter % plot_every == 0:
all_losses.append(current_loss / plot_every)
current_loss = 0
# -
plt.figure()
plt.plot(all_losses)
plt.show()
# # Évaluation
# +
# Keep track of correct guesses in a confusion matrix
confusion = torch.zeros(n_categories, n_categories)
n_confusion = 10000
# Just return an output given a line
def evaluate(line_tensor):
hidden = rnn.initHidden()
for i in range(line_tensor.size()[0]):
output, hidden = rnn(line_tensor[i], hidden)
return output
# Go through a bunch of examples and record which are correctly guessed
for i in range(n_confusion):
category, line, category_tensor, line_tensor = randomTrainingExample()
output = evaluate(line_tensor)
guess, guess_i = categoryFromOutput(output)
category_i = all_categories.index(category)
confusion[category_i][guess_i] += 1
# Normalize by dividing every row by its sum
for i in range(n_categories):
confusion[i] = confusion[i] / confusion[i].sum()
# Set up plot
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(confusion.numpy())
fig.colorbar(cax)
# Set up axes
ax.set_xticklabels([''] + all_categories, rotation=90)
ax.set_yticklabels([''] + all_categories)
# Force label at every tick
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
# sphinx_gallery_thumbnail_number = 2
plt.show()
# +
def predict(input_line, n_predictions=3):
print('\n> %s' % input_line)
with torch.no_grad():
output = evaluate(lineToTensor(input_line))
# Get top N categories
topv, topi = output.topk(n_predictions, 1, True)
predictions = []
for i in range(n_predictions):
value = topv[0][i].item()
category_index = topi[0][i].item()
print('(%.2f) %s' % (value, all_categories[category_index]))
predictions.append([value, all_categories[category_index]])
predict('Dovesky')
predict('Jackson')
predict('Satoshi')
# -
# # Visualizing memorization
# +
# https://distill.pub/2019/memorization-in-rnns/
# nom = 'Dovesky'
# nom = 'Jackson'
# nom = 'Mohammed'
# nom = 'Sébastien'
nom = 'Dominique'
# nom='Thomas'
category_index = 5
line_tensor = lineToTensor(nom)
# print(line_tensor.size())
# print(line_tensor[0,0])
line_tensor.requires_grad_()
gradient_groundtruth_list = []
hidden = rnn.initHidden()
for i in range(line_tensor.size()[0]):
output, hidden = rnn(line_tensor[i], hidden)
g_groundtruth = torch.autograd.grad(output[0, category_index], line_tensor, retain_graph=True)[0].data
gradient_groundtruth_list.append(g_groundtruth)
# print(i, g.size(), g)
topv, topi = output.topk(1, 1, True)
category_index_predicted = topi[0][0].item()
print('pred: %i %s' %(category_index_predicted, all_categories[category_index_predicted]))
print('GT: %i %s' %(category_index, all_categories[category_index]))
gradient_pred_list = []
hidden = rnn.initHidden()
for i in range(line_tensor.size()[0]):
output, hidden = rnn(line_tensor[i], hidden)
g = torch.autograd.grad(output[0, category_index_predicted], line_tensor, retain_graph=True)[0].data
gradient_pred_list.append(g)
# On calcule la "connectivity"
# https://discuss.pytorch.org/t/newbie-getting-the-gradient-with-respect-to-the-input/12709/2
# g = torch.autograd.grad(output[0,category_index], line_tensor, retain_graph=True)[0].data
# print(g.size())
# g = torch.autograd.grad(outputs[:,0,category_index], line_tensor, retain_graph=True)[0].data
connectivity = np.zeros((line_tensor.size()[0],line_tensor.size()[0]))
for i in range(line_tensor.size()[0]):
char_index = letterToIndex(nom[i])
# print(i, g[i,0,char_index])
connectivity[:,i] = np.abs(gradient_groundtruth_list[i][:, 0, char_index].detach().cpu().numpy())
# print(nom[i], connectivity[:,i])
plt.figure(figsize=(8,6))
plt.imshow(connectivity)
plt.ylabel("Time step", fontsize=14)
toto = plt.yticks(ticks=range(len(nom)), fontsize=14)
toto = plt.xticks(ticks=range(len(nom)), labels=list(nom), fontsize=14)
# +
connectivity = np.zeros((line_tensor.size()[0],line_tensor.size()[0]))
for i in range(line_tensor.size()[0]):
char_index = letterToIndex(nom[i])
# print(i, g[i,0,char_index])
connectivity[:,i] = np.abs(gradient_pred_list[i][:, 0, char_index].detach().cpu().numpy())
# print(nom[i], connectivity[:,i])
plt.figure(figsize=(8,6))
plt.imshow(connectivity)
plt.ylabel("Time step", fontsize=14)
toto = plt.yticks(ticks=range(len(nom)), fontsize=14)
toto = plt.xticks(ticks=range(len(nom)), labels=list(nom), fontsize=14)
# +
# torch.autograd.grad?
# -
all_categories
|
tp2_names/TP2_classification_de_noms.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# <h1> 2. Creating a sampled dataset </h1>
#
# This notebook illustrates:
# <ol>
# <li> Sampling a BigQuery dataset to create datasets for ML
# <li> Preprocessing with Pandas
# </ol>
# + deletable=true editable=true
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
# + deletable=true editable=true
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# + deletable=true editable=true language="bash"
# if ! gsutil ls | grep -q gs://${BUCKET}/; then
# gsutil mb -l ${REGION} gs://${BUCKET}
# fi
# + [markdown] deletable=true editable=true
# <h2> Create ML dataset by sampling using BigQuery </h2>
# <p>
# Let's sample the BigQuery data to create smaller datasets.
# </p>
# + deletable=true editable=true
# Create SQL query using natality data after the year 2000
import google.datalab.bigquery as bq
query = """
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
ABS(FARM_FINGERPRINT(CONCAT(CAST(YEAR AS STRING), CAST(month AS STRING)))) AS hashmonth
FROM
publicdata.samples.natality
WHERE year > 2000
"""
# + [markdown] deletable=true editable=true
# There are only a limited number of years and months in the dataset. Let's see what the hashmonths are.
# + deletable=true editable=true
# Call BigQuery but GROUP BY the hashmonth and see number of records for each group to enable us to get the correct train and evaluation percentages
df = bq.Query("SELECT hashmonth, COUNT(weight_pounds) AS num_babies FROM (" + query + ") GROUP BY hashmonth").execute().result().to_dataframe()
print("There are {} unique hashmonths.".format(len(df)))
df.head()
# + [markdown] deletable=true editable=true
# Here's a way to get a well distributed portion of the data in such a way that the test and train sets do not overlap:
# + deletable=true editable=true
# Added the RAND() so that we can now subsample from each of the hashmonths to get approximately the record counts we want
trainQuery = "SELECT * FROM (" + query + ") WHERE MOD(hashmonth, 4) < 3 AND RAND() < 0.0005"
evalQuery = "SELECT * FROM (" + query + ") WHERE MOD(hashmonth, 4) = 3 AND RAND() < 0.0005"
traindf = bq.Query(trainQuery).execute().result().to_dataframe()
evaldf = bq.Query(evalQuery).execute().result().to_dataframe()
print("There are {} examples in the train dataset and {} in the eval dataset".format(len(traindf), len(evaldf)))
# + [markdown] deletable=true editable=true
# <h2> Preprocess data using Pandas </h2>
# <p>
# Let's add extra rows to simulate the lack of ultrasound. In the process, we'll also change the plurality column to be a string.
# + deletable=true editable=true
traindf.head()
# + [markdown] deletable=true editable=true
# Also notice that there are some very important numeric fields that are missing in some rows (the count in Pandas doesn't count missing data)
# + deletable=true editable=true
# Let's look at a small sample of the training data
traindf.describe()
# + deletable=true editable=true
# It is always crucial to clean raw data before using in ML, so we have a preprocessing step
import pandas as pd
def preprocess(df):
# clean up data we don't want to train on
# in other words, users will have to tell us the mother's age
# otherwise, our ML service won't work.
# these were chosen because they are such good predictors
# and because these are easy enough to collect
df = df[df.weight_pounds > 0]
df = df[df.mother_age > 0]
df = df[df.gestation_weeks > 0]
df = df[df.plurality > 0]
# modify plurality field to be a string
twins_etc = dict(zip([1,2,3,4,5],
['Single(1)', 'Twins(2)', 'Triplets(3)', 'Quadruplets(4)', 'Quintuplets(5)']))
df['plurality'].replace(twins_etc, inplace=True)
# now create extra rows to simulate lack of ultrasound
nous = df.copy(deep=True)
nous.loc[nous['plurality'] != 'Single(1)', 'plurality'] = 'Multiple(2+)'
nous['is_male'] = 'Unknown'
return pd.concat([df, nous])
# + deletable=true editable=true
traindf.head()# Let's see a small sample of the training data now after our preprocessing
traindf = preprocess(traindf)
evaldf = preprocess(evaldf)
traindf.head()
# + deletable=true editable=true
traindf.tail()
# + deletable=true editable=true
# Describe only does numeric columns, so you won't see plurality
traindf.describe()
# + [markdown] deletable=true editable=true
# <h2> Write out </h2>
# <p>
# In the final versions, we want to read from files, not Pandas dataframes. So, write the Pandas dataframes out as CSV files.
# Using CSV files gives us the advantage of shuffling during read. This is important for distributed training because some workers might be slower than others, and shuffling the data helps prevent the same data from being assigned to the slow workers.
#
# + deletable=true editable=true
traindf.to_csv('train.csv', index=False, header=False)
evaldf.to_csv('eval.csv', index=False, header=False)
# + deletable=true editable=true
# %bash
wc -l *.csv
head *.csv
tail *.csv
# + [markdown] deletable=true editable=true
# Copyright 2017-2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
# -
|
2_sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.cm as cm
import numpy as np
import pandas as pd
import random
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
# +
df = pd.read_csv('track_data.csv', index_col=0)
df.drop_duplicates(subset=['Track IDs'], keep='first', inplace=True)
df.head()
# +
X = df.iloc[:, 5:].values
y = df.iloc[:, 1].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20)
# +
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# -
classifier = RandomForestClassifier(n_estimators=20)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
print(accuracy_score(y_test, y_pred))
|
data/RandomForest_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# There's more to the world of deep learning than just dense layers. There are dozens of kinds of layers you might add to a model. (Try browsing through the [Keras docs](https://www.tensorflow.org/api_docs/python/tf/keras/layers/) for a sample!) Some are like dense layers and define connections between neurons, and others can do preprocessing or transformations of other sorts.
#
# In this lesson, we'll learn about a two kinds of special layers, not containing any neurons themselves, but that add some functionality that can sometimes benefit a model in various ways. Both are commonly used in modern architectures.
#
# # Dropout #
#
# The first of these is the "dropout layer", which can help correct overfitting.
#
# In the last lesson we talked about how overfitting is caused by the network learning spurious patterns in the training data. To recognize these spurious patterns a network will often rely on very a specific combinations of weight, a kind of "conspiracy" of weights. Being so specific, they tend to be fragile: remove one and the conspiracy falls apart.
#
# This is the idea behind **dropout**. To break up these conspiracies, we randomly *drop out* some fraction of a layer's input units every step of training, making it much harder for the network to learn those spurious patterns in the training data. Instead, it has to search for broad, general patterns, whose weight patterns tend to be more robust.
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/a86utxY.gif" width="600" alt="An animation of a network cycling through various random dropout configurations.">
# <figcaption style="textalign: center; font-style: italic"><center>Here, 50% dropout has been added between the two hidden layers.</center></figcaption>
# </figure>
#
# You could also think about dropout as creating a kind of *ensemble* of networks. The predictions will no longer be made by one big network, but instead by a committee of smaller networks. Individuals in the committee tend to make different kinds of mistakes, but be right at the same time, making the committee as a whole better than any individual. (If you're familiar with random forests as an ensemble of decision trees, it's the same idea.)
#
# ## Adding Dropout ##
#
# In Keras, the dropout rate argument `rate` defines what percentage of the input units to shut off. Put the `Dropout` layer just before the layer you want the dropout applied to:
#
# ```
# keras.Sequential([
# # ...
# layer.Dropout(rate=0.3), # apply 30% dropout to the next layer
# layer.Dense(16),
# # ...
# ])
# ```
#
# # Batch Normalization #
#
# The next special layer we'll look at performs "batch normalization" (or "batchnorm"), which can help correct training that is slow or unstable.
#
# With neural networks, it's generally a good idea to put all of your data on a common scale, perhaps with something like scikit-learn's [StandardScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) or [MinMaxScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html). The reason is that SGD will shift the network weights in proportion to how large an activation the data produces. Features that tend to produce activations of very different sizes can make for unstable training behavior.
#
# Now, if it's good to normalize the data before it goes into the network, maybe also normalizing inside the network would be better! In fact, we have a special kind of layer that can do this, the **batch normalization layer**. A batch normalization layer looks at each batch as it comes in, first normalizing the batch with its own mean and standard deviation, and then also putting the data on a new scale with two trainable rescaling parameters. Batchnorm, in effect, performs a kind of coordinated rescaling of its inputs.
#
# Most often, batchnorm is added as an aid to the optimization process (though it can sometimes also help prediction performance). Models with batchnorm tend to need fewer epochs to complete training. Moreover, batchnorm can also fix various problems that can cause the training to get "stuck". Consider adding batch normalization to your models, especially if you're having trouble during training.
#
# ## Adding Batch Normalization ##
#
# It seems that batch normalization can be used at almost any point in a network. You can put it after a layer...
#
# ```
# layers.Dense(16, activation='relu'),
# layers.BatchNormalization(),
# ```
#
# ... or between a layer and its activation function:
#
# ```
# layers.Dense(16),
# layers.BatchNormalization(),
# layers.Activation('relu'),
# ```
#
# And if you add it as the first layer of your network it can act as a kind of adaptive preprocessor, standing in for something like Sci-Kit Learn's `StandardScaler`.
#
# # Example - Using Dropout and Batch Normalization #
#
# Let's continue developing the *Red Wine* model. Now we'll increase the capacity even more, but add dropout to control overfitting and batch normalization to speed up optimization. This time, we'll also leave off standardizing the data, to demonstrate how batch normalization can stabalize the training.
# +
#$HIDE_INPUT$
# Setup plotting
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
# Set Matplotlib defaults
plt.rc('figure', autolayout=True)
plt.rc('axes', labelweight='bold', labelsize='large',
titleweight='bold', titlesize=18, titlepad=10)
import pandas as pd
red_wine = pd.read_csv('../input/dl-course-data/red-wine.csv')
# Create training and validation splits
df_train = red_wine.sample(frac=0.7, random_state=0)
df_valid = red_wine.drop(df_train.index)
# Split features and target
X_train = df_train.drop('quality', axis=1)
X_valid = df_valid.drop('quality', axis=1)
y_train = df_train['quality']
y_valid = df_valid['quality']
# -
# Whan adding dropout, you may need to increase the number of units in your `Dense` layers.
# +
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
layers.Dense(1024, activation='relu', input_shape=[11]),
layers.Dropout(0.3),
layers.BatchNormalization(),
layers.Dense(1024, activation='relu'),
layers.Dropout(0.3),
layers.BatchNormalization(),
layers.Dense(1024, activation='relu'),
layers.Dropout(0.3),
layers.BatchNormalization(),
layers.Dense(1),
])
# -
# There's nothing to change this time in how we set up the training.
# +
model.compile(
optimizer='adam',
loss='mae',
)
history = model.fit(
X_train, y_train,
validation_data=(X_valid, y_valid),
batch_size=256,
epochs=100,
verbose=0,
)
# Show the learning curves
history_df = pd.DataFrame(history.history)
history_df.loc[:, ['loss', 'val_loss']].plot();
# -
# You'll typically get better performance if you standardize your data before using it for training. That we were able to use the raw data at all, however, shows how effective batch normalization can be on more difficult datasets.
#
# # Your Turn #
#
# Move on to [**improve predictions**](#$NEXT_NOTEBOOK_URL$) on the *Spotify* dataset with dropout and see how batch normalization can help with difficult datasets.
|
notebooks/deep_learning_intro/raw/tut5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%HTML
<style>.container{width:100%;}</style>
# Dieses Notebook hängt vom Notebook `1-Splay-Trees.ipynb` ab.
# %run 1-Splay-Trees.ipynb
# Zusätzlich benötigen wir Funktionen aus den `collections` sowie aus den `functools`, die beide schon in Python-Distributionen enthalten sind.
import collections
import functools
# # Mengen auf Basis von Splay Trees
#
# Wir benutzen nun die Splay Trees, um Mengen (`set`) in Python mit Ordnung zu unterstützen.
#
# Dazu bauen wir die hier gelisteten Methoden nach ([Permalink zur Datei, aus der dies generiert wird](https://github.com/python/cpython/blob/3.7/Objects/setobject.c "R. <NAME>ettinger et al. (2019): cpython/Objects/setobject.c, GitHub")):
help(set)
# Wir definieren dafür die Klasse `OrderedSet`.
class OrderedSet:
pass
# ## Konstruktion
#
# Wenn wir das Konstruieren eines Sets aus einem `iterable` unterstützen wollen, begegnen wir dem Problem der Implementierung von Mengen modifizierbarer Objekte, zum Beispiel Mengen von Mengen.
#
# Sei $N$ eine Menge und $M$ ein modifizierbares Objekt, zum Beispiel eine Menge, und sei $M \in N$. Wie stellen wir sicher, dass $N$ geordnet bleibt, wenn $M$ geändert wird? Wir könnten verlangen, dass in $N$ ein Element, wann immer es preisgegeben wird, neu einsortiert werden muss. Abgesehen vom damit verbundenen Berechnungsaufwand ist dies aber nicht praktikabel, da auf $M$ auch Referenzen gehalten werden können, die nicht von $N$ abhängig sind, und somit $M$ geändert werden könnte, ohne dass $N$ davon erfährt. (Die $N$-seitige Überwachung aller gehaltenen Objekte wäre noch unpraktischer als das erneute Einsortieren.)
#
# Ein vergleichbares Problem ergibt sich übrigens auch bei den Hashtabellen in der Referenzimplementierung CPython. Hier kann keine Ordnungsbedingung verletzt werden, jedoch ist im Allgemeinen ein Objekt nach Änderung nicht mehr unter ihrem vorigen Hash zu finden. Der Python-Standard gibt daher vor, dass es strenggenommen keine Mengen von Mengen gibt – [es gibt nur Mengen von unmodifizierbaren Mengen](https://docs.python.org/3.7/library/stdtypes.html#set "Python Software Foundation (2019): The Python Standard Library/Built-in Types/set, Python Documentation"), den `frozenset`s. Ebenso gibt es beispielsweise keine Mengen von Listen.
#
# Wir wollen auch die `frozenset`s geordnet haben und werden sie parallel mitimplementieren.
class OrderedFrozenset:
pass
# Wir übernehmen `_arb_lt`, `_arb_eq` und `_arb_gt` von `Node`, sodass wir direkt Zugriff darauf haben.
OrderedSet._arb_lt = OrderedFrozenset._arb_lt = Node._arb_lt
OrderedSet._arb_eq = OrderedFrozenset._arb_eq = Node._arb_eq
OrderedSet._arb_gt = OrderedFrozenset._arb_gt = Node._arb_gt
# Konstruieren wir nun eine Menge aus einem `iterable`, so fügen wir die Elemente einzeln ein. Wir arbeiten direkt mit `Node`s, wrappen also nicht noch zusätzlich einen `SplayTree`. Wir überprüfen dabei die hinzuzufügenden Elemente darauf, ob sie in einer Menge existieren dürfen, indem wir überprüfen, ob sie eine `__hash__`-Methode haben.
# +
def __init__(self, iterable=[]):
if isinstance(self, OrderedFrozenset) and hasattr(self, "_tree"):
return # frozenset was already created
self._tree = None
iterator = iter(iterable)
try:
element = next(iterator)
if not element.__hash__:
raise TypeError(f"unhashable type: '{type(element).__name__}'")
self._tree = Node(element, None, None)
while True:
element = next(iterator)
if not element.__hash__:
raise TypeError("unhashable type: " +
f"'{type(element).__name__}'")
self._tree = self._tree.insert(element)
except StopIteration:
pass
OrderedSet.__init__ = OrderedFrozenset.__init__ = __init__
del __init__
# -
# ### Lexikographische Ordnung
#
# Wir müssen, wenn wir geordnete Mengen von geordneten Mengen unterstützen wollen, die totalen Ordnungen `<` und `>` sowie die Operation `==` zwischen geordneten Mengen definieren – diese Operatoren werden von `_arb_lt` und verwandten Methoden und somit mittelbar von `splay` benutzt. Wir müssen dabei leider das Verhalten von `set` und `frozenset` verändern; für diese drücken `<` und `>` die echte Ober- bzw. Teilmenge aus. Alternativ könnten wir in `_arb_lt` und verwandten Methoden vorher den Typ überprüfen und im Fall von `OrderedSet` und `OrderedFrozenset` nicht `<` und `>`, sondern andere Methoden verwenden. Die damit verbundenen Performanceeinbußen sind jedoch nicht vertretbar.
#
# Eine totale Ordnung, die sich anbietet, ist die lexikographische Ordnung. Hier wird durch die Mengen iteriert und das erste ungleiche Wertepaar entscheidet die Ungleichheit. Wir definieren dafür zunächst die Iteration `__iter__`, um durch die Elemente in der Menge iterieren zu können.
#
# Dies erreichen wir, indem wir den Baum wie bei einer *Inorder*-Ausgabe durchlaufen, wobei wir zur Ausgabe das Keyword `yield` benutzen. Die Implementierung ist iterativ und hält sich einen eigenen Stapel, da dieser in seiner Größe weniger begrenzt ist als der Aufrufstapel, der sich bei einer rekursiven Implementierung aufbauen würde. Für diesen Stapel benutzen wir [`deque`](https://docs.python.org/3.7/library/collections.html#collections.deque "Python Software Foundation (2020): The Python Standard Library/Data Types/collections/deque, Python Documentation") (double-ended queue) aus den `collections` der Standard Library, da sich diese für größere Stapel besser eignet als die Liste.
# +
def __iter__(self):
stack = collections.deque()
tree = self._tree
while stack or tree is not None:
if tree is not None:
stack.append(tree)
tree = tree.left
continue
tree = stack.pop()
yield tree.payload
tree = tree.right
OrderedSet.__iter__ = OrderedFrozenset.__iter__ = __iter__
del __iter__
# -
# Nun können wir (für `<`) die Methode `__lt__` mit lexikographischer Ordnung definieren. Wir geben bei Typinkompatibilität `NotImplemented` zurück, woraus von Python zur Laufzeit gegebenenfalls ein `TypeError` gemacht wird. Wir iterieren durch die geordneten Mengen und können uns bei der ersten Verschiedenheit entscheiden. Die leere Menge ist kleiner als jede Menge außer der leeren.
#
# Wir können dies auch formal notieren, wenn wir mit $\mathrm{head}$ das erste und mit $\mathrm{tail}$ alle übrigen Elemente der geordneten Menge bezeichnen. Bei allen formalen Definitionen sprechen wir vereinfachend nur vom $\mathrm{OrderedSet}$, nicht vom $\mathrm{OrderedFrozenset}$.
#
# $$\mathrm{\_\_lt\_\_}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{\_\_lt\_\_}(x, y) = \begin{cases}
# \mathrm{false} &\mathrm{falls}\ y = \{\}\\
# \mathrm{true} &\mathrm{falls}\ x = \{\} \land y \neq \{\}\\
# \mathrm{true} &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{\_\_lt\_\_}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
# +
def __lt__(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
return NotImplemented
x_iter, y_iter = iter(self), iter(other)
while True:
try:
y_item = next(y_iter)
except StopIteration:
return False # x is longer or equal
try:
x_item = next(x_iter)
except StopIteration:
return True # x is shorter
if self._arb_lt(x_item, y_item):
return True
if self._arb_gt(x_item, y_item):
return False
OrderedSet.__lt__ = OrderedFrozenset.__lt__ = __lt__
del __lt__
# -
# Ähnlich definieren wir `__gt__` für `>`. Formal heißt das:
#
# $$\mathrm{\_\_gt\_\_}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{\_\_gt\_\_}(x, y) = \begin{cases}
# \mathrm{false} &\mathrm{falls}\ x = \{\}\\
# \mathrm{true} &\mathrm{falls}\ x \neq \{\} \land y = \{\}\\
# \mathrm{true} &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{\_\_gt\_\_}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
# +
def __gt__(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
return NotImplemented
x_iter, y_iter = iter(self), iter(other)
while True:
try:
x_item = next(x_iter)
except StopIteration:
return False # x is shorter or equal
try:
y_item = next(y_iter)
except StopIteration:
return True # x is longer
if self._arb_gt(x_item, y_item):
return True
if self._arb_lt(x_item, y_item):
return False
OrderedSet.__gt__ = OrderedFrozenset.__gt__ = __gt__
del __gt__
# -
# Für `__eq__` (`==`) müssen wir nur Element für Element vergleichen. Wir verändern hier auch nicht das Verhalten im Vergleich zu den Vorbildern `set` und `frozenset`. Hier ist der Vergleich mit fremden Typen kein `NotImplemented`, sondern einfach die Ungleichheit. Wir können auch formal definieren, wobei wir mit $\dot{\lor}$ das *exklusive Oder* meinen:
#
# $$\mathrm{\_\_eq\_\_}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{\_\_eq\_\_}(x, y) = \begin{cases}
# \mathrm{true} &\mathrm{falls}\ x = \{\} \land y = \{\}\\
# \mathrm{false} &\mathrm{falls}\ x = \{\}\ \dot{\lor}\ y = \{\}\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) \neq \mathrm{head}(y)\\
# \mathrm{\_\_eq\_\_}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
# +
def __eq__(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
return False
x_iter, y_iter = iter(self), iter(other)
while True:
try:
x_item = next(x_iter)
except StopIteration:
try: # assert y is also exhausted
next(y_iter)
except StopIteration:
return True
return False
try:
y_item = next(y_iter)
except StopIteration:
return False # x was not exhausted
if not self._arb_eq(x_item, y_item):
return False
OrderedSet.__eq__ = OrderedFrozenset.__eq__ = __eq__
del __eq__
# -
# `__ne__` für `!=` ist damit schon implizit definiert.
#
# ### Hashen
#
# Der letzte Schritt, um `OrderedFrozenset`s in `OrderedSet`s haben zu können, ist die Unterstützung einer `__hash__`-Funktion für `OrderedFrozenset` (`OrderedSet` ist wegen seiner Veränderlichkeit bewusst **nicht** hashbar). Wir addieren dazu die Hashwerte aller Elemente, deren Hashbarkeit wir ja schon zur Zeit des Hinzufügens überprüft haben.
# +
def __hash__(self):
return sum(hash(el) for el in self)
OrderedFrozenset.__hash__ = __hash__
del __hash__
# -
# Um das Verhalten von Python abzubilden, werfen wir im Fall von `OrderedSet` einen `TypeError`, wenn versucht wird, eines zu hashen.
# +
def __hash__(self):
raise TypeError(f"unhashable type: '{type(self).__name__}'")
OrderedSet.__hash__ = __hash__
del __hash__
# -
# ## Operationen mit einem Element
#
# Wir implementieren im Folgenden die Methoden von `set` und `frozenset` für `OrderedSet` und `OrderedFrozenset`. Wir beginnen mit den Operationen mit einem einzelnen Element als Parameter.
#
# ### Hinzufügen
#
# Wir implementieren `add`, das Hinzufügen eines Elements, ähnlich wie beim `SplayTree`:
# +
def add(self, element):
if not element.__hash__:
raise TypeError(f"unhashable type: '{type(element).__name__}'")
if self._tree is None:
self._tree = Node(element, None, None)
else:
self._tree = self._tree.insert(element)
OrderedSet.add = add
del add
# -
# ### Entfernen
#
# Das Gegenstück zu `add` ist `remove`, das Entfernen eines Elementes.
# +
def remove(self, element):
if not element.__hash__:
raise TypeError(f"unhashable type: '{type(element).__name__}'")
if self._tree is None:
raise KeyError(element)
rc, self._tree = self._tree.remove(element)
if not rc:
raise KeyError(element)
OrderedSet.remove = remove
del remove
# -
# Wir implementieren auch `discard`, welches ähnlich zu `remove` ist, aber keinen Fehler verursacht, falls das Element nicht vorhanden ist.
# +
def discard(self, element):
if not element.__hash__:
raise TypeError(f"unhashable type: '{type(element).__name__}'")
if self._tree is not None:
_, self._tree = self._tree.remove(element)
OrderedSet.discard = discard
del discard
# -
# `add`, `remove` und `discard` sind **keine** Methoden von `OrderedFrozenset`, weil diese Operationen die Menge modifizieren!
#
# ### Finden
#
# Eine nichtmodifizierende Operation mit einem Element als Parameter ist `__contains__` (`in`), die wir ebenfalls für `Node` schon implementiert haben.
# +
def __contains__(self, element):
if not element.__hash__:
raise TypeError(f"unhashable type: '{type(element).__name__}'")
if self._tree is None:
return False
contains, self._tree = self._tree.contains(element)
return contains
OrderedSet.__contains__ = OrderedFrozenset.__contains__ = __contains__
del __contains__
# -
# ## Operationen ohne Parameter
#
# Wir implementieren im Folgenden Operationen auf die geordnete Menge, die überhaupt keine Parameter haben. Wir beginnen wieder mit solchen, die die Menge verändern.
#
# ### Kopieren
#
# Wir implementieren `copy`, welches eine Kopie der Menge zurückgibt. Da die Funktion so kurz ist, schreiben wir die Funktionen für die beiden Methoden einzeln, statt den Typ von `self` zu überprüfen.
OrderedSet.copy = lambda self: OrderedSet(el for el in self)
OrderedFrozenset.copy = lambda self: OrderedFrozenset(el for el in self)
# ### Entfernen eines beliebigen Elements
#
# Wir implementieren `pop`, welches ein beliebiges Element entfernt und zurückgibt. Wir entfernen dabei das Minimum des Baums.
# +
def pop(self):
if self._tree is None:
raise KeyError("pop from an empty set")
popped = self.minimum()
self.remove(popped)
return popped
OrderedSet.pop = pop
del pop
# -
# ### Leeren
#
# Wir implementieren auch `clear`, welches alle Elemente aus der Menge entfernt. Wir setzen einfach den Baum auf `None`.
# +
def clear(self):
self._tree = None
OrderedSet.clear = clear
del clear
# -
# ### Länge
#
# Wir kehren zurück zu den Operationen, die auch für `OrderedFrozenset` erlaubt sind, und implementieren `__len__`, welches die Länge von Mengen berechnet und dann mit `len(some_ordered_set)` benutzt werden kann.
# +
def __len__(self):
return sum(1 for i in self)
OrderedSet.__len__ = OrderedFrozenset.__len__ = __len__
del __len__
# -
# ### Darstellung
#
# Wir implementieren auch `__repr__`, welches eine String-Darstellung anbietet. Wir wollen zum Beispiel für die Elemente `a, b` die Darstellung `OrderedSet([a, b])` haben und implementieren:
# +
def __repr__(self):
if self._tree is None:
return f"{type(self).__name__}()"
return f"{type(self).__name__}({list(self)})"
OrderedSet.__repr__ = OrderedFrozenset.__repr__ = __repr__
del __repr__
# -
# ### Pickling
#
# `set` implementiert `__reduce__`, welches von der *pickle*-API aufgerufen wird. Diese API dient zur Serialisierung. Wir implementieren für diese API nicht `__reduce__`, sondern das simplere [`__getnewargs__`](https://docs.python.org/3/library/pickle.html#object.__getnewargs__ "Python Software Foundation (2020): The Python Standard Library/Data Persistence/object.__getnewargs__(), Python Documentation"), welches die Argumente, mit denen eine Rekonstruktion erreicht wird, zurückgibt. Wir geben den Mengeninhalt als Simpel aus einer Liste aus, sodass zum Beispiel für die Elemente `a, b` beim Deserialisieren `OrderedSet(([a, b],))` aufgerufen wird.
# +
def __getnewargs__(self):
return (list(self),)
OrderedSet.__getnewargs__ = OrderedFrozenset.__getnewargs__ = __getnewargs__
del __getnewargs__
# -
# ### Minimum und Maximum
#
# Wir implementieren noch die *zusätzliche* Methode `minimum`. Mit `__iter__` funktioniert zwar bereits das Built-in `min`, wir können dies mit einem Baum aber effizienter gestalten: Wir geben die Nutzlast des linkesten Knotens zurück. Für `Node` definieren wir
# +
def minimum(self):
while self.left is not None:
self = self.left
return self.payload
Node.minimum = minimum
del minimum
# -
# und für `OrderedSet` und `OrderedFrozenset`
# +
def minimum(self):
if self._tree is None:
raise ValueError("Set is empty")
return self._tree.minimum()
OrderedSet.minimum = OrderedFrozenset.minimum = minimum
del minimum
# -
# Wir implementieren auch `maximum`, bei der wir die Nutzlast des rechtesten Knotens zurückgeben.
# +
def maximum(self):
while self.right is not None:
self = self.right
return self.payload
Node.maximum = maximum
del maximum
# -
# und für die `Set`-Klassen
# +
def maximum(self):
if self._tree is None:
raise ValueError("Set is empty")
return self._tree.maximum()
OrderedSet.maximum = OrderedFrozenset.maximum = maximum
del maximum
# -
# ## Boolesche Operationen auf andere Mengen
#
# Wir implementieren im Folgenden Operationen auf andere Mengen, die einen Wahrheitswert als Ergebnis haben. `==` und `!=` hatten wir schon für die lexikographische Ordnung implementiert.
#
# ### Teilmenge
#
# Wir haben `issubset`, `__le__` (`<=`, in der Mengenlehre $\subseteq$) und `__lt__` (`<`, in der Mengenlehre $\subset$ bzw. $\subsetneq$) zu implementieren. `issubset` arbeitet wie `__le__`, erlaubt aber einen beliebigen Parameter als Operand, statt nur Mengen. Wir werden die echte Teilmenge nicht unter dem Namen `<` implementieren können, weil wir diesen Operator für die lexikographische Ordnung benutzen. Wir werden dafür in Anlehnung an `issubset` die Methode `is_proper_subset` implementieren, die *keine* Methode von `set` oder `frozenset` ist.
#
# Für die unechte Teilmenge implementieren wir die Hilfsfunktion `_subseteq`, welches von geordneten Mengen $x, y$ die Relation $x \subseteq y$ überprüft. So brauchen wir für die eigentlichen Methoden nur `_subseteq` mit den entsprechenden Parametern aufrufen. Dies erspart besonders für die Obermenge Arbeit.
#
# Hier gehen wir für jedes Element aus $x$ so lange in $y$ weiter, bis die Elemente gleich sind. In diesem Fall gehen wir in beiden Mengen ein Element weiter. Ist ein Element aus $y$ größer, so kam ein Element aus $x$ nicht in $y$ vor, und wir verneinen die Relation $\subseteq$. Ist $x$ erschöpft, so waren alle Elemente in $y$. Ist $y$ erschöpft, so war mindestens ein Element von $x$ nicht in $y$. Formal können wir das auch schreiben als
#
# $$\mathrm{\_subseteq}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{\_subseteq}(x, y) = \begin{cases}
# \mathrm{true} &\mathrm{falls}\ x = \{\}\\
# \mathrm{false} &\mathrm{falls}\ x \neq \{\} \land y = \{\}\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{\_subseteq}(x, \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{\_subseteq}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
# +
def _subseteq(self, x, y):
x_iter, y_iter = iter(x), iter(y)
while True:
try:
x_item = next(x_iter)
except StopIteration:
return True
while True:
try:
y_item = next(y_iter)
except StopIteration:
return False
if self._arb_lt(x_item, y_item):
return False
if self._arb_eq(x_item, y_item):
break
OrderedSet._subseteq = OrderedFrozenset._subseteq = _subseteq
del _subseteq
# -
# So können wir `issubset` implementieren, wo wir das andere Iterable gegebenenfalls in ein `OrderedFrozenset` bringen und so sortieren.
# +
def issubset(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedFrozenset(other)
return self._subseteq(self, other)
OrderedSet.issubset = OrderedFrozenset.issubset = issubset
del issubset
# -
# Bei `__le__` erlauben wir für den anderen Operanden nur `OrderedSet` und `OrderedFrozenset`.
# +
def __le__(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
return NotImplemented
return self._subseteq(self, other)
OrderedSet.__le__ = OrderedFrozenset.__le__ = __le__
del __le__
# -
# Für `is_proper_subset` und später `is_proper_superset` definieren wir `_subsetneq`, welches $x \subset y$ überprüft. Hier halten wir uns die auf `False` initialisierte Variable `proper_subset`, mit deren Hilfe wir sicherstellen, dass $x$ eine echte Teilmenge von $y$ ist. Sie wird auf `True` gesetzt, sobald ein Element aus $y$ kleiner ist als aus $x$. Ist $x$ erschöpft und `proper_subset` nicht `True`, so prüfen wir, ob noch Elemente in $y$ sind, dann ist ebenfalls $x \subset y$.
#
# Formal schreiben wir zusätzlich `_subsetneq_help` mit zusätzlichem Parameter $p$ (proper). Zu Beginn ist dieser $\mathrm{false}$.
#
# $$\mathrm{\_subsetneq}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{\_subsetneq}(x, y) = \mathrm{\_subsetneq\_help}(x, y, \mathrm{false})$$
# $$\mathrm{\_subsetneq\_help}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \times \mathbb{B} \to \mathbb{B}$$
# $$\mathrm{\_subsetneq\_help}(x, y, p) = \begin{cases}
# p &\mathrm{falls}\ y = \{\}\\
# \mathrm{true} &\mathrm{falls}\ x = \{\} \land y \neq \{\}\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{\_subsetneq\_help}(x, \mathrm{tail}(y), \mathrm{true}) &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{\_subsetneq\_help}(\mathrm{tail}(x), \mathrm{tail}(y), p) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
# +
def _subsetneq(self, x, y):
x_iter, y_iter = iter(x), iter(y)
proper_subset = False
while True:
try:
x_item = next(x_iter)
except StopIteration:
if not proper_subset:
try: # assert y is not exhausted
next(y_iter)
except StopIteration:
return False
return True
while True:
try:
y_item = next(y_iter)
except StopIteration:
return False
if self._arb_lt(x_item, y_item):
return False
if self._arb_gt(x_item, y_item):
proper_subset = True
else:
break
OrderedSet._subsetneq = OrderedFrozenset._subsetneq = _subsetneq
del _subsetneq
# -
# So können wir `is_proper_subset` ähnlich wie `issubset` implementieren.
# +
def is_proper_subset(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
return self._subsetneq(self, other)
OrderedSet.is_proper_subset = is_proper_subset
OrderedFrozenset.is_proper_subset = is_proper_subset
del is_proper_subset
# -
# ### Obermenge
#
# Wir können für die Obermenge jetzt `_subseteq` und `_subsetneq` wiederverwenden und müssen nur die Parameter ändern. Analog zur den Operationen für die Untermenge haben wir `issuperset`…
# +
def issuperset(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
return self._subseteq(other, self)
OrderedSet.issuperset = OrderedFrozenset.issuperset = issuperset
del issuperset
# -
# …`__ge__` (`>=`, in der Mengenlehre $\supseteq$)…
# +
def __ge__(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
return NotImplemented
return self._subseteq(other, self)
OrderedSet.__ge__ = OrderedFrozenset.__ge__ = __ge__
del __ge__
# -
# …und `is_proper_superset` statt `__gt__` (`>`, in der Mengenlehre $\supset$ bzw. $\supsetneq$).
# +
def is_proper_superset(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
return self._subsetneq(other, self)
OrderedSet.is_proper_superset = is_proper_superset
OrderedFrozenset.is_proper_superset = is_proper_superset
del is_proper_superset
# -
# ### Überprüfen, ob Mengen disjunkt sind
#
# Wir implementieren zuletzt `isdisjoint`, das überprüft, ob zwei Mengen disjunkt sind, das heißt, ob sie keine gemeinsamen Elemente haben. Wir iterieren parallel durch die Mengen und überprüfen, dass wir jedes Element in nur einer Menge wiederfinden. Formal heißt das:
#
# $$\mathrm{isdisjoint}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathbb{B}$$
# $$\mathrm{isdisjoint}(x, y) = \begin{cases}
# \mathrm{true} &\mathrm{falls}\ \{\} \in \{x, y\}\\
# \mathrm{false} &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)\\
# \mathrm{isdisjoint}(\mathrm{tail}(x), y) &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{isdisjoint}(x, \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)
# \end{cases}$$
# +
def isdisjoint(self, other):
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
x_iter, y_iter = iter(self), iter(other)
try:
y_item = next(y_iter) # we need a y_item ready for first comparison
except StopIteration:
return True
while True:
try:
x_item = next(x_iter)
except StopIteration:
return True
while True:
if self._arb_lt(x_item, y_item):
break
if self._arb_gt(x_item, y_item):
try:
y_item = next(y_iter)
except StopIteration:
return True
continue
return False
OrderedSet.isdisjoint = OrderedFrozenset.isdisjoint = isdisjoint
del isdisjoint
# -
# ## Verknüpfung von Mengen
#
# Zuletzt implementieren wir die Operationen auf andere Mengen, die eine Menge als Ergebnis haben. Wir beginnen mit den einfacheren Vereinigungs- und Differenzmengen.
#
# ### Vereinigungsmenge
#
# Bei Vereinigungsmengen ($\cup$) fügen wir einfach alle Elemente hinzu. Zunächst definieren wir die Methode `union`. Bei Bedarf wandeln wir die Arbeitsmenge wieder in ein `Frozenset` um.
# +
def union(self, *others):
union = OrderedSet(self)
for other in others:
for el in other:
union.add(el)
if isinstance(self, OrderedFrozenset):
frozen = OrderedFrozenset()
frozen._tree = union._tree
return frozen
return union
OrderedSet.union = OrderedFrozenset.union = union
del union
# -
# Für die Vereinigungsmenge gibt es auch `__or__` (`|`). Die beiden Methoden haben jedoch nicht ganz dasselbe Verhalten: `__or__` funktioniert nur mit Mengen, und operiert auf nur eine zusätzliche Menge; hingegen nimmt `union` beliebig viele iterable Sammlungen. Diese Überprüfungen benötigen wir später auch für `-`, `&` und `^`, weshalb wir eine Technik aus der funktionalen Programmierung benutzen, um uns Code zu ersparen, nämlich *Currying*. Dabei wird eine Funktion mit mehreren Argumenten als mehrere Funktionen mit je nur einem Argument behandelt. In unserem Fall schreiben wir eine generische Funktion mit den genannten Überprüfungen, die getrennt die Operation und auf der nächsten Ebene die Operanden nimmt – sie ist teilweise *gecurryt* und *uncurryt* umgekehrt wieder. Wir formulieren `_uncurry_reg_op` (regular operation), die die übergebene Funktion (z. B. `union`) in die Routine für den Operator (z. B. `|`) einsetzt.
# +
def _uncurry_reg_op(func):
def reg_op(self, other):
if isinstance(other, OrderedSet) \
or isinstance(other, OrderedFrozenset):
# this style is necessary when uncurrying a class function
return func(self, other)
else:
return NotImplemented
return reg_op
OrderedSet._uncurry_reg_op = OrderedFrozenset._uncurry_reg_op = _uncurry_reg_op
del _uncurry_reg_op
# -
# Nun können wir für beide Klassen `__or__` setzen.
OrderedSet.__or__ = OrderedSet._uncurry_reg_op(OrderedSet.union)
OrderedFrozenset.__or__ = OrderedFrozenset._uncurry_reg_op(
OrderedFrozenset.union)
# Wir benötigen auch `__ror__` (reverse or), bei dem das Objekt selbst (`self`) an zweiter Stelle steht. Wir definieren dafür analog `_uncurry_rev_op` (reverse operation). Zwar verhält sich `|` kommutativ, sodass wir auch `_uncurry_reg_op` verwenden könnten, dies ist aber für `-` nicht der Fall, und so definieren wir gleich die generische Variante für alle umgekehrten Operationen.
# +
def _uncurry_rev_op(func):
def rev_op(self, other):
if isinstance(other, OrderedSet) \
or isinstance(other, OrderedFrozenset):
return func(other, self)
else:
return NotImplemented
return rev_op
OrderedSet._uncurry_rev_op = OrderedFrozenset._uncurry_rev_op = _uncurry_rev_op
del _uncurry_rev_op
# -
# Das Setzen von `__ror__` verläuft analog.
OrderedSet.__ror__ = OrderedSet._uncurry_rev_op(OrderedSet.union)
OrderedFrozenset.__ror__ = OrderedFrozenset._uncurry_rev_op(
OrderedFrozenset.union)
# *In-place* ist die `union` entsprechende Operation `update`. Hier operieren wir direkt auf `self`. Diese Methode ordnen wir nicht `OrderedFrozenset` zu, da die Menge überschrieben wird.
# +
def update(self, *others):
for other in others:
for el in other:
self.add(el)
OrderedSet.update = update
del update
# -
# Als Operator haben wir `__ior__` (`|=`). Hier definieren wir noch `_uncurry_inp_op` (in-place operation), weil wir dieses Codemuster wieder für die folgenden Operationen benötigen.
# +
def _uncurry_inp_op(func):
def inp_op(self, other):
if isinstance(other, OrderedSet) \
or isinstance(other, OrderedFrozenset):
func(self, other)
return self
else:
return NotImplemented
return inp_op
OrderedSet._uncurry_inp_op = _uncurry_inp_op
del _uncurry_inp_op
# -
# So können wir `__ior__` setzen, natürlich nur für `OrderedSet`.
OrderedSet.__ior__ = OrderedSet._uncurry_inp_op(OrderedSet.update)
# ### Differenzmenge
#
# Es folgt die Differenzmenge ($\setminus$), für die wir einfach alle Elemente entfernen. Dafür haben wir `difference`.
# +
def difference(self, *others):
difference = OrderedSet(self)
for other in others:
for el in other:
if not el.__hash__:
raise TypeError(f"unhashable type: '{type(el).__name__}'")
difference.discard(el)
if isinstance(self, OrderedFrozenset):
frozen = OrderedFrozenset()
frozen._tree = difference._tree
return frozen
return difference
OrderedSet.difference = OrderedFrozenset.difference = difference
del difference
# -
# Als Operatoren haben wir `__sub__` (`-`) und `__rsub__` (reverse).
OrderedSet.__sub__ = OrderedSet._uncurry_reg_op(OrderedSet.difference)
OrderedFrozenset.__sub__ = OrderedFrozenset._uncurry_reg_op(
OrderedFrozenset.difference)
OrderedSet.__rsub__ = OrderedSet._uncurry_rev_op(OrderedSet.difference)
OrderedFrozenset.__rsub__ = OrderedFrozenset._uncurry_rev_op(
OrderedFrozenset.difference)
# In-place haben wir `difference_update`…
# +
def difference_update(self, *others):
for other in others:
for el in other:
if not el.__hash__:
raise TypeError(f"unhashable type: '{type(el).__name__}'")
self.discard(el)
OrderedSet.difference_update = difference_update
del difference_update
# -
# …und `__isub__` (`-=`).
OrderedSet.__isub__ = OrderedSet._uncurry_inp_op(
OrderedSet.difference_update)
# ### Schnittmenge
#
# Komplexer sind `intersection` und `__and__` (`&`). Wir definieren in der Methode zunächst die lokale binäre Relation `intersect`, die wir auch formal notieren. Wir iterieren durch die Mengen, wobei wir gleiche Elemente aufnehmen, und enden, wenn eine Menge ausgeschöpft ist.
#
# $$\mathrm{intersect}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathrm{OrderedSet}$$
# $$\mathrm{intersect}(x, y) = \begin{cases}
# \mathrm{OrderedSet}() &\mathrm{falls}\ \{\} \in \{x, y\}\\
# \mathrm{intersect}(\mathrm{tail}(x), y) &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{intersect}(x, \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{OrderedSet}([\mathrm{head}(x)]) \cup \mathrm{intersect}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
#
# Wir machen aus allen zu schneidenden Sammlungen `OrderedSet`s und *reduzieren* sie dann im funktionalen Sinne, das heißt, wir fassen sie anhand von `intersect` mithilfe von [`reduce`](https://docs.python.org/3.7/library/functools.html#functools.reduce "Python Software Foundation (2020): The Python Standard Library/Functional Programming Modules/functools/reduce, Python Documentation") aus den `functools` der Standard Library zusammen. Wir können das tun, weil sich der Schnitt von Mengen assoziativ verhält.
# +
def intersection(self, *others):
def intersect(x, y):
intersection = OrderedSet()
x_iter, y_iter = iter(x), iter(y)
try:
x_item, y_item = next(x_iter), next(y_iter)
while True:
if self._arb_lt(x_item, y_item):
x_item = next(x_iter)
continue
if self._arb_gt(x_item, y_item):
y_item = next(y_iter)
continue
intersection.add(x_item)
x_item, y_item = next(x_iter), next(y_iter)
except StopIteration:
return intersection
if not others:
return self.copy() # otherwise we're returning self
sets = [self] + [OrderedSet(other) if not isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset) else other
for other in others]
intersection = functools.reduce(intersect, sets)
if isinstance(self, OrderedFrozenset):
frozen = OrderedFrozenset()
frozen._tree = intersection._tree
return frozen
return intersection
OrderedSet.intersection = OrderedFrozenset.intersection = intersection
del intersection
# -
# Für die Operatoren `__and__` und `__rand__` können wir wieder die Uncurry-Funktionen benutzen.
OrderedSet.__and__ = OrderedSet._uncurry_reg_op(OrderedSet.intersection)
OrderedFrozenset.__and__ = OrderedFrozenset._uncurry_reg_op(
OrderedFrozenset.intersection)
OrderedSet.__rand__ = OrderedSet._uncurry_rev_op(OrderedSet.intersection)
OrderedFrozenset.__rand__ = OrderedFrozenset._uncurry_rev_op(
OrderedFrozenset.intersection)
# ### Schnittmenge in-place
#
# Verwandt mit diesen Methoden sind `intersection_update` und `__iand__` (`&=`), die die Schnittmenge *in-place* berechnen, das heißt, sie überschreiben die Menge direkt. Hier definieren wir `intersect_update`, welches ähnlich zu `intersect` ist, nur eben in-place funktioniert, sodass die Menge selbst nicht kopiert werden muss. Wir können nicht konventionell durch die Menge iterieren, wenn wir sie unterdessen verändern, sodass wir die Möglichkeit des Splayens ausnutzen: Beim Entfernen eines Knotens ist die neue Wurzel gerade das nächstgrößere Element. Gehen wir regulär weiter, so können wir einfach das Minimum aus dem rechten Teilbaum hochsplayen.
#
# Gerade weil diese Methoden die Menge überschreiben, sind sie keine Methoden von `OrderedFrozenset`.
# +
def intersection_update(self, *others):
def intersect_update(self, other):
self._tree = self._tree._splay(self.minimum())
other_iter = iter(other)
try:
other_item = next(other_iter)
while True:
if self._arb_lt(self._tree.payload, other_item):
if self._tree.right is None:
raise StopIteration
minimum = self._tree.left is None
self.remove(self._tree.payload) # equivalent to moving on
if minimum and self._tree is not None:
# removal leads to using right subtree
# instead of next element
self._tree = self._tree._splay(self.minimum())
continue
if self._arb_gt(self._tree.payload, other_item):
other_item = next(other_iter)
continue
if self._tree.right is None:
return
# equivalent to inserting and moving on
self._tree = self._tree._splay(self._tree.right.minimum())
other_item = next(other_iter)
except StopIteration:
# leave the rest
self._tree = self._tree.left
for other in others:
if self._tree is None:
return
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)) \
or id(self) == id(other): # don't iterate through self
other = OrderedSet(other)
intersect_update(self, other)
OrderedSet.intersection_update = intersection_update
del intersection_update
# -
# Als Operator haben wir `__iand__`.
OrderedSet.__iand__ = OrderedSet._uncurry_inp_op(
OrderedSet.intersection_update)
# ### Symmetrische Differenz
#
# Für die symmetrische Differenz ($\bigtriangleup$) brauchen wir out-of-place `symmetric_difference`, welches nur auf zwei Mengen definiert sein muss. Wir wiederholen das Verhalten der Schnittmenge, nur, dass gerade die Elemente reinkommen, die in der anderen Menge nicht vorhanden sind. Dies können wir auch formulieren als:
#
# $$\mathrm{symmetric\_difference}: \mathrm{OrderedSet} \times \mathrm{OrderedSet} \to \mathrm{OrderedSet}$$
# $$\mathrm{symmetric\_difference}(x, y) = \begin{cases}
# x \cup y &\mathrm{falls}\ \{\} \in \{x, y\}\\
# \mathrm{OrderedSet}([\mathrm{head}(x)]) \cup \mathrm{symmetric\_difference}(\mathrm{tail}(x), y) &\mathrm{falls}\ \mathrm{head}(x) < \mathrm{head}(y)\\
# \mathrm{OrderedSet}([\mathrm{head}(y)]) \cup \mathrm{symmetric\_difference}(x, \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) > \mathrm{head}(y)\\
# \mathrm{symmetric\_difference}(\mathrm{tail}(x), \mathrm{tail}(y)) &\mathrm{falls}\ \mathrm{head}(x) = \mathrm{head}(y)
# \end{cases}$$
#
# Um bei unserer iterativen Implementierung leichter mit der Frage umgehen zu können, welche Menge zuerst zu Ende war, halten wir uns die Ausnahme `OtherStopIteration`, die bedeutet, dass die andere Menge ($y$) zu Ende war.
# +
def symmetric_difference(self, other):
class OtherStopIteration(StopIteration):
pass
if self._tree is None:
return type(self)(other)
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
x_iter, y_iter = iter(self), iter(other)
x_item = next(x_iter)
try:
y_item = next(y_iter)
except StopIteration:
return self.copy()
symmetric_difference = OrderedSet()
try:
try:
while True:
if self._arb_lt(x_item, y_item):
symmetric_difference.add(x_item)
try:
x_item = next(x_iter)
except StopIteration:
symmetric_difference.add(y_item)
raise StopIteration
continue
if self._arb_gt(x_item, y_item):
symmetric_difference.add(y_item)
try:
y_item = next(y_iter)
except StopIteration:
symmetric_difference.add(x_item)
raise OtherStopIteration
continue
x_item = next(x_iter)
try:
y_item = next(y_iter)
except StopIteration:
symmetric_difference.add(x_item) # already updated
raise OtherStopIteration
except OtherStopIteration:
while True:
symmetric_difference.add(next(x_iter))
except StopIteration:
try:
while True:
symmetric_difference.add(next(y_iter))
except StopIteration:
pass
if isinstance(self, OrderedFrozenset):
frozen = OrderedFrozenset()
frozen._tree = symmetric_difference._tree
return frozen
return symmetric_difference
OrderedSet.symmetric_difference = symmetric_difference
OrderedFrozenset.symmetric_difference = symmetric_difference
del symmetric_difference
# -
# Als Operation haben wir `__xor__` (`^`) und `__rxor__`.
OrderedSet.__xor__ = OrderedSet._uncurry_reg_op(
OrderedSet.symmetric_difference)
OrderedFrozenset.__xor__ = OrderedFrozenset._uncurry_reg_op(
OrderedFrozenset.symmetric_difference)
OrderedSet.__rxor__ = OrderedSet._uncurry_rev_op(
OrderedSet.symmetric_difference)
OrderedFrozenset.__rxor__ = OrderedFrozenset._uncurry_rev_op(
OrderedFrozenset.symmetric_difference)
# ### Symmetrische Differenz in-place
#
# Wenn wir die symmetrische Differenz in-place, also überschreibend berechnen, benutzen wir stattdessen `SelfStopIteration`, da dies den Code verkürzt. Ansonsten gehen wir ähnlich vor. Wir nutzen wieder aus, dass beim Entfernen das nächste Element an der Wurzel steht.
# +
def symmetric_difference_update(self, other):
class SelfStopIteration(StopIteration):
pass
if self._tree is None:
return OrderedSet(other)
if id(self) == id(other):
self._tree = None
return # iteration through self is unstable
if not (isinstance(other, OrderedSet)
or isinstance(other, OrderedFrozenset)):
other = OrderedSet(other)
self._tree = self._tree._splay(self.minimum())
other_iter = iter(other)
try:
other_item = next(other_iter)
try:
while True:
if self._arb_lt(self._tree.payload, other_item):
if self._tree.right is None:
self.add(other_item)
raise SelfStopIteration
self._tree = self._tree._splay(self._tree.right.minimum())
continue
if self._arb_gt(self._tree.payload, other_item):
root = self._tree.payload
self.add(other_item)
self._tree = self._tree._splay(root)
other_item = next(other_iter)
continue
minimum = self._tree.left is None
maximum = self._tree.right is None
self.remove(self._tree.payload)
if self._tree is None or maximum:
raise SelfStopIteration
if minimum:
self._tree = self._tree._splay(self.minimum())
other_item = next(other_iter)
except SelfStopIteration:
while True:
self.add(next(other_iter))
except StopIteration:
return
OrderedSet.symmetric_difference_update = symmetric_difference_update
del symmetric_difference_update
# -
# Als Operation haben wir `__ixor__` (`^=`).
OrderedSet.__ixor__ = OrderedSet._uncurry_inp_op(
OrderedSet.symmetric_difference_update)
|
2-Ordered-Sets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
from IPython.core.display import HTML
css = open('style-table.css').read() + open('style-notebook.css').read()
HTML('<style>{}</style>'.format(css))
titles = pd.read_csv('data/titles.csv')
titles.head()
cast = pd.read_csv('data/cast.csv')
cast.head()
# +
# Define a year as a "Superman year"
# whose films feature more Superman characters than Batman.
# How many years in film history have been Superman years?
c = cast
c = c[(c.character == 'Superman') | (c.character == 'Batman')]
c = c.groupby(['year', 'character']).size()
c = c.unstack()
c = c.fillna(0)
c.head()
# -
d = c.Superman - c.Batman
print('Superman years:')
print(len(d[d > 0.0]))
# +
# How many years have been "Batman years",
# with more Batman characters than Superman characters?
print('Batman years:')
print(len(d[d < 0.0]))
# +
# Plot the number of actor roles each year
# and the number of actress roles each year
# over the history of film.
c = cast
c = c.groupby(['year', 'type']).size()
c = c.unstack('type')
c.plot()
# +
# Plot the number of actor roles each year
# and the number of actress roles each year,
# but this time as a kind='area' plot.
c = cast
c = c.groupby(['year', 'type']).size()
c = c.unstack('type')
c.plot(kind='area')
# +
# Plot the difference between the number of actor roles each year
# and the number of actress roles each year over the history of film.
c = cast
c = c.groupby(['year', 'type']).size()
c = c.unstack('type')
(c.actor - c.actress).plot()
# +
# Plot the fraction of roles that have been 'actor' roles
# each year in the history of film.
c = cast
c = c.groupby(['year', 'type']).size()
c = c.unstack('type')
(c.actor / (c.actor + c.actress)).plot(ylim=[0,1])
# +
# Plot the fraction of supporting (n=2) roles
# that have been 'actor' roles
# each year in the history of film.
c = cast
c = c[c.n == 2]
c = c.groupby(['year', 'type']).size()
c = c.unstack('type')
(c.actor / (c.actor + c.actress)).plot(ylim=[0,1])
# +
# Build a plot with a line for each rank n=1 through n=3,
# where the line shows what fraction of that rank's roles
# were 'actor' roles for each year in the history of film.
c = cast
c = c[c.n <= 3]
c = c.groupby(['year', 'type', 'n']).size()
c = c.unstack('type').fillna(0)
r = c.actor / (c.actor + c.actress)
r = r.unstack('n')
r.plot(ylim=[0,1])
# -
|
pycon-pandas-tutorial/Solutions-4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Overlap-Strelka2-and-Mutect2-calls-and-filter-variant-(VAF->-0.03)-for-CBTTC" data-toc-modified-id="Overlap-Strelka2-and-Mutect2-calls-and-filter-variant-(VAF->-0.03)-for-CBTTC-1">Overlap Strelka2 and Mutect2 calls and filter variant (VAF > 0.03) for CBTTC</a></span></li><li><span><a href="#Dependencies" data-toc-modified-id="Dependencies-2">Dependencies</a></span></li><li><span><a href="#Paths" data-toc-modified-id="Paths-3">Paths</a></span></li><li><span><a href="#Main" data-toc-modified-id="Main-4">Main</a></span><ul class="toc-item"><li><span><a href="#Overlap-variants-with-chr:start_pos" data-toc-modified-id="Overlap-variants-with-chr:start_pos-4.1">Overlap variants with chr:start_pos</a></span></li><li><span><a href="#Add-sample_id-from-clinical_df" data-toc-modified-id="Add-sample_id-from-clinical_df-4.2">Add sample_id from clinical_df</a></span></li></ul></li></ul></div>
# -
# # Overlap Strelka2 and Mutect2 calls and filter variant (VAF > 0.03) for CBTTC
# # Dependencies
library(maftools)
library(ggplot2)
# # Paths
manifestpath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Manifests/"
datapath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Data/"
plotpath <- "/Users/anabbi/OneDrive - UHN/Documents/IPD2/Plots/"
# # Main
strelka2_CBTTC <- read.maf(maf = paste0(datapath, "MAF_mat/CBTTC_recoded_mafs_KB_strelka2.maf"),
useAll = TRUE,
verbose = T,
vc_nonSyn = c("Frame_Shift_Del", "Frame_Shift_Ins",
"Nonsense_Mutation","Nonstop_Mutation",
"In_Frame_Del","In_Frame_Ins",
"Missense_Mutation"))
mutect2_CBTTC <- read.maf(maf = paste0(datapath, "MAF_mat/CBTTC_Mutect2_combined_kb.maf"),
useAll = TRUE,
verbose = T,
vc_nonSyn = c("Frame_Shift_Del", "Frame_Shift_Ins",
"Nonsense_Mutation","Nonstop_Mutation",
"In_Frame_Del","In_Frame_Ins",
"Missense_Mutation"))
strelka2 <- strelka2_CBTTC@data
mutect2 <- mutect2_CBTTC@data
head(strelka2)
# Find samples with both strelka2 and mutect2 calls
# +
strelkaIDs <- as.character(strelka2$Tumor_Sample_Barcode[!duplicated(strelka2$Tumor_Sample_Barcode)])
mutectIDs <- as.character(mutect2$Tumor_Sample_Barcode[!duplicated(mutect2$Tumor_Sample_Barcode)])
# -
length(strelkaIDs)
length(mutectIDs)
length(strelkaIDs[strelkaIDs %in% mutectIDs])
# ## Overlap variants with chr:start_pos
strelka2$chrpos <- paste(strelka2$Chromosome, strelka2$Start_Position,sep = ":")
mutect2$chrpos <- paste(mutect2$Chromosome, mutect2$Start_Position,sep = ":")
#samples in strelka2
strelkaIDs <- as.character(strelka2$Tumor_Sample_Barcode[!duplicated(strelka2$Tumor_Sample_Barcode)])
#samples in mutect2
mutectIDs <- as.character(mutect2$Tumor_Sample_Barcode[!duplicated(mutect2$Tumor_Sample_Barcode)])
#overlapping samples
overlapID <- strelkaIDs[strelkaIDs %in% mutectIDs]
strelka2_sample <- strelka2[strelka2$Tumor_Sample_Barcode == overlapID[1],]
mutect2_sample <- mutect2[mutect2$Tumor_Sample_Barcode == overlapID[1],]
overlap_sample <- merge(strelka2_sample[, c("chrpos", "Hugo_Symbol")], mutect2_sample,
by = "chrpos")
overlap_sample
# +
for(id in overlapID[2:length(overlapID)]){
strelka2_sample <- strelka2[strelka2$Tumor_Sample_Barcode == id,]
mutect2_sample <- mutect2[mutect2$Tumor_Sample_Barcode == id,]
message("number of strelka2 calls for: ", id)
print(nrow(strelka2_sample))
message("number of mutect2 calls for: ", id)
print(nrow(mutect2_sample))
overlap_sample1 <- merge(strelka2_sample[, c("chrpos", "Hugo_Symbol")],
mutect2_sample,
by = "chrpos")
overlap_sample <- rbind(overlap_sample,
overlap_sample1)
message("number of overlapping calls for: ", id)
print(nrow(overlap_sample1))
}
# -
length(as.character(overlap_sample$Tumor_Sample_Barcode[!duplicated(overlap_sample$Tumor_Sample_Barcode)]))
# Are there any samples with no overlap?
strelka2$Tumor_Sample_Barcode <- as.character(strelka2$Tumor_Sample_Barcode)
mutect2$Tumor_Sample_Barcode <- as.character(mutect2$Tumor_Sample_Barcode)
# +
for(id in overlapID[1:length(overlapID)]){
strelka2_sample <- strelka2[strelka2$Tumor_Sample_Barcode == id,]
mutect2_sample <- mutect2[mutect2$Tumor_Sample_Barcode == id,]
overlap_sample1 <- merge(strelka2_sample[, c("chrpos", "Hugo_Symbol")],
mutect2_sample,
by = "chrpos")
if(nrow(overlap_sample1) == 0){
print(mutect2_sample$Tumor_Sample_Barcode[!duplicated(mutect2_sample$Tumor_Sample_Barcode)])}
}
# -
# 64 cases had no mutations overlapping
#
# I ended up with 658 cases for which I had both rna and mutect2 results. using overlapped calls I will end up with 597 cases with both rna and mutation calls. See TMB_final_overlapped.
overlap_sample$Hugo_Symbol.x <- NULL
head(overlap_sample)
colnames(overlap_sample)[2] <- "Hugo_Symbol"
overlap_sample$t_VAF <- overlap_sample$t_alt_count / overlap_sample$t_depth
head(overlap_sample$t_VAF)
overlap_sample_VAF <- overlap_sample[ overlap_sample$t_VAF > 0.03,]
# +
dim(overlap_sample)
dim(overlap_sample_VAF)
# -
ggplot(data = overlap_sample) + geom_histogram(aes(x = t_VAF), bins = 100)
write.table(overlap_sample_VAF,
file = paste0(datapath,"snv/CBTTC_maf_overlapped_3VAF.maf"),
sep = "\t", quote = F, row.names = F)
# ## Add sample_id from clinical_df
mymaf <- read.maf(maf = paste0(datapath,"snv/CBTTC_maf_overlapped_3VAF.maf"),
useAll = TRUE,
verbose = T,
vc_nonSyn = c("Frame_Shift_Del", "Frame_Shift_Ins",
"Nonsense_Mutation","Nonstop_Mutation",
"In_Frame_Del","In_Frame_Ins",
"Missense_Mutation"))
CBTTC_mut <- getSampleSummary(mymaf)
dim(CBTTC_mut)
head(CBTTC_mut)
IDlist <- read.table(paste0(manifestpath,"CBTTC_KidsFirstIDs.txt"),
sep = "\t", header = T,na.strings = "Not Applicable")
head(IDlist)
CBTTC_mut$sample_id <- NA
CBTTC_mut$sample_id1 <- NA
CBTTC_mut$Tumor_Sample_Barcode <- as.character(CBTTC_mut$Tumor_Sample_Barcode)
IDlist$clinical_event <- as.character(IDlist$clinical_event)
IDlist$tumor_dna_bs_1 <- as.character(IDlist$tumor_dna_bs_1)
IDlist$tumor_dna_bs_2 <- as.character(IDlist$tumor_dna_bs_2)
# finding sampleIds from biospecimen1 and 2 and putting them together
for(i in 1:nrow(CBTTC_mut)){
CBTTC_mut$sample_id1[i] <- IDlist$clinical_event[match(CBTTC_mut$Tumor_Sample_Barcode[i],
IDlist$tumor_dna_bs_2)]
}
for(i in 1:nrow(CBTTC_mut)){
CBTTC_mut$sample_id[i] <- IDlist$clinical_event[match(CBTTC_mut$Tumor_Sample_Barcode[i],
IDlist$tumor_dna_bs_1)]
}
dim(CBTTC_mut)
CBTTC_mut$sample_id[!is.na(CBTTC_mut$sample_id1)] <- CBTTC_mut$sample_id1[!is.na(CBTTC_mut$sample_id1)]
CBTTC_mut$sample_id1 <- NULL
dim(CBTTC_mut[!is.na(CBTTC_mut$sample_id)])
# These are the cellline entries, other_aliquots and some samples that I do not have rna
CBTTC_mut[is.na(CBTTC_mut$sample_id)]
write.table(CBTTC_mut[!is.na(CBTTC_mut$sample_id)],
file = paste0(datapath,"snv/CBTTC_mafsummary_overlapped_3VAF.txt"),
sep = "\t", quote = F, row.names = F)
|
notebooks/CBTTC_snv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.patches import Rectangle
import numpy as np
from pprint import pprint as pp
import csv
from pathlib import Path
import itertools
from itertools import product, combinations, permutations
import random
from datetime import datetime, timedelta, date
from collections import OrderedDict, defaultdict, Counter, namedtuple
from pandas.io.json import json_normalize
import seaborn as sns
import matplotlib.dates as mdates
import re
import requests
from bs4 import BeautifulSoup
import statsmodels.api as sm
import calendar
import scipy
import json
from PIL import Image
from bs4 import BeautifulSoup as bs
from ast import literal_eval # use to eval a string as a list df['column'] = df['column'].apply(literal_eval)
from stockstats import StockDataFrame as sdf # dataframe wrapper for stack calculations
import math
import time
import sys
import string
import json
import sklearn
import urllib
import urllib3
import psutil
pd.set_option('display.max_columns', 700)
pd.set_option('display.max_rows', 100)
pd.set_option('display.min_rows', 10)
pd.set_option('display.expand_frame_repr', True)
# ## Synthetic Data:
# ### Use recursion to flatten the nested `dicts`
#
# - [Thinking Recursively in Python](https://realpython.com/python-thinking-recursively/)
# - [Flattening JSON objects in Python](https://towardsdatascience.com/flattening-json-objects-in-python-f5343c794b10)
# - [flatten](https://github.com/amirziai/flatten)
# - The `flatten_json` function, will be used to flatten `data`
# +
def flatten_json(nested_json: dict, exclude: list=[''], sep='_') -> dict:
"""
Flatten a list of nested dicts.
"""
out = dict()
def flatten(x: (list, dict, str), name: str='', exclude=exclude):
if type(x) is dict:
for a in x:
if a not in exclude:
flatten(x[a], f'{name}{a}{sep}')
elif type(x) is list:
i = 0
for a in x:
flatten(a, f'{name}{i}{sep}')
i += 1
else:
out[name[:-1]] = x
flatten(nested_json)
return out
# df = pd.DataFrame([flatten_json(x) for x in data])
# df = pd.DataFrame([flatten_json(x) for x in data[key]])
# -
np.random.seed(365)
date_0 = pd.date_range(pd.datetime.today(), periods=10).to_pydatetime().tolist()
date_0[:2]
date_1 = pd.date_range(end=pd.datetime.today(), periods=10).to_pydatetime().tolist()
date_1[:2]
date = pd.bdate_range(end=pd.datetime.today(), periods=10).to_pydatetime().tolist()
date[:2]
np.random.seed(365)
data = {'a': [np.random.randint(10) for _ in range(15)],
'b': [np.random.randint(10) for _ in range(15)],
'c': [np.random.randint(10) for _ in range(15)],
'd': [np.random.randint(10) for _ in range(15)],
'e': [np.random.randint(10) for _ in range(15)],
'date': pd.bdate_range(pd.datetime.today(), periods=15).tolist()}
random.seed(365)
data = {'groups': [random.choice(['1-5', '6-25', '26-100', '100-500', '500-1000', '>1000']) for _ in range(1000)],
'treatment': [random.choice(['Yes', 'No']) for _ in range(1000)]}
# +
# stock data
periods = '3600'
resp = requests.get('https://api.cryptowat.ch/markets/poloniex/ethusdt/ohlc', params={'periods': periods})
data = resp.json()
df = pd.DataFrame(data['result'][periods], columns=['date', 'open', 'high', 'low', 'close', 'volume', 'amount'])
df['date'] = pd.to_datetime(df['date'], unit='s')
stock = sdf.retype(df)
stock['macds']
# stock
# -
# p = Path('test.json') # if in current dir
p = Path.cwd() / 'test.json'
# p = Path.cwd() / 'data/nvdcve-1.1-2019.json/nvdcve-1.1-2019.json'
with p.open('r', encoding='utf-8') as f:
data = json.loads(f.read())
# Use with a list of dicts
with p.open("r") as f:
data = literal_eval(f.read())
df = json_normalize(data)
df = pd.read_clipboard(sep='\\s+')
df.to_clipboard(sep='\\s+', index=False)
df = pd.read_csv('test.csv')
# df = pd.read_csv('test.txt', sep='|', header=None, converters={2: eval}) # converters={'file_path_lists': eval}
df = pd.read_excel('test.xlsx')
df = pd.DataFrame(data)
# +
# These are the usual ipython objects, including this one you are creating
ipython_vars = ['In', 'Out', 'exit', 'quit', 'get_ipython', 'ipython_vars'] # list a variables
# Get a sorted list of the objects and their sizes
sorted([(x, sys.getsizeof(globals().get(x))) for x in dir() if not x.startswith('_') and x not in sys.modules and x not in ipython_vars], key=lambda x: x[1], reverse=True)
# -
# ## End Synthetic Data:
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
from keras.models import load_model
import cv2
import numpy as np
# Load trained deep learning model
model = load_model('face-mask-image-classification-with-keras.h5')
# Classifier to detect face-mask
face_det_classifier=cv2.CascadeClassifier('haarcascade_frontalface_default.xml')
# Capture video for face scanning
source=cv2.VideoCapture(0)
# Dictionaries caontaing details of wearing mask and color of rectangle arund face.
# Green for wearing and Red for not wearing face mask
labels_dict={0:'MASK',1:'NO MASK'}
color_dict={0:(0,255,0),1:(0,0,255)}
# +
while(True):
ret, img=source.read()
grayscale_img =cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
faces = face_det_classifier.detectMultiScale(grayscale_img,1.3,5)
for x,y,w,h in faces:
face_img=grayscale_img[y:y+w,x:x+w]
resized = cv2.resize(face_img,(100,100))
normalized=resized/255.0
reshaped=np.reshape(normalized,(1,100,100,1))
result=model.predict(reshaped)
label=np.argmax(result,axis=1)[0]
cv2.rectangle(img,(x,y),(x+w,y+h),color_dict[label],2)
cv2.rectangle(img,(x,y-40),(x+w,y),color_dict[label],-1)
cv2.putText(img, labels_dict[label], (x, y-10),cv2.FONT_HERSHEY_SIMPLEX,0.8,(255,255,255),2)
#If level = 1 then it means wearing No mask and 0 means wearing mask
if (label == 1):
img = cv2.imwrite('nomask.png',img)
# %run mail.py nomask.png
#messagebox.showwarning("Access Denied")
# #%run mail.py
# #%run alert_notification.py
# #%run alert_email.py
else:
pass
break
cv2.imshow('Live face scanning',img)
key=cv2.waitKey(1)
if(key==27):
break
cv2.destroyAllWindows()
source.release()
|
real-time-mask-detction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Murine bone-marrow derived macrophages
# https://data.broadinstitute.org/bbbc/BBBC020/
# ## Make a torch dataset
from segmentation.datasets import BroadDataset
# ### Show some images
# %matplotlib inline
import matplotlib.pyplot as plt
#base = '/Users/nicholassofroniew/Documents/BBBC/BBBC020_v1/BBBC020_v1-cells'
base = '/Users/nicholassofroniew/Documents/DATA/neurofinder/localcorr'
train_dataset = BroadDataset(base, 'train')
# +
fig = plt.figure()
for i in range(len(train_dataset)):
img, mask = train_dataset[i]
# print(i, sample['image'].shape)
ax = plt.subplot(2, 4, i + 1)
plt.tight_layout()
ax.set_title('Sample #{}'.format(i))
ax.axis('off')
plt.imshow(img)
ax = plt.subplot(2, 4, i + 5)
ax.set_title('Mask #{}'.format(i))
ax.axis('off')
plt.imshow(mask)
if i == 3:
plt.show()
break
# -
type(img)
from numpy import asarray
print(asarray(img).max())
print(asarray(mask).max())
# ## Define transforms
# +
from torchvision import transforms
import segmentation.transforms as extended_transforms
joint_transform = extended_transforms.Compose([
extended_transforms.RandomHorizontallyFlip(),
extended_transforms.RandomVerticallyFlip()
# extended_transforms.CenterCrop(512),
# extended_transforms.RandomRotate(45),
# extended_transforms.RandomWarp(5, 20)
])
input_transform = transforms.Compose(
[transforms.ToTensor(),
#transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
])
target_transform = extended_transforms.MaskToTensor()
# -
train_dataset = BroadDataset(base, 'train', joint_transform=joint_transform)
# +
fig = plt.figure()
for i in range(len(train_dataset)):
img, mask = train_dataset[i]
# print(i, sample['image'].shape)
ax = plt.subplot(2, 4, i + 1)
plt.tight_layout()
ax.set_title('Sample #{}'.format(i))
ax.axis('off')
plt.imshow(img)
ax = plt.subplot(2, 4, i + 5)
ax.set_title('Mask #{}'.format(i))
ax.axis('off')
plt.imshow(mask)
if i == 3:
plt.show()
break
# -
type(img)
from numpy import asarray
print(asarray(img).max())
print(asarray(mask).max())
train_dataset = BroadDataset(base, 'train', joint_transform=joint_transform, input_transform=input_transform, target_transform=target_transform)
from torch.utils.data import DataLoader
trainloader = DataLoader(train_dataset, batch_size=1,
shuffle=True, num_workers=2)
# ## Load network
from segmentation.model import UNet
net = UNet(1)
net.train()
# ### Define loss function and optimizer
from segmentation.utilities import mIoULoss, CrossEntropyLoss2d, BCELoss2d
from torch import nn
criterion = mIoULoss(size_average=False)
#criterion = nn.NLLLoss2d()
#criterion = nn.BCEWithLogitsLoss()
import torch.optim as optim
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
# ### Pass data throuhg network
import torch.nn.functional as F
from torch.autograd import Variable
img, mask = train_dataset[0]
tmp = Variable(mask).unsqueeze(0)
foo = net(Variable(img).unsqueeze(0)).squeeze(0)
print(tmp)
print(foo)
#print(bar)
loss = criterion(foo, tmp)
print(loss)
loss.backward()
# ### Train network
val_dataset = BroadDataset(base, 'train', input_transform=input_transform, target_transform=target_transform)
valloader = DataLoader(val_dataset, batch_size=1,
shuffle=False, num_workers=2)
from segmentation.main import train, validate
from os.path import isdir, join
from os import mkdir
# +
save_epoch = 1
epochs = 2
for epoch in range(epochs): # loop over the dataset multiple times
train(trainloader, net, criterion, optimizer, epoch, 18)
# save out model every n epochs
if save_epoch is not None:
if epoch % save_epoch == save_epoch-1:
snapshot_name = 'model-%04d' % epoch
save_path = join(base, 'models-local', snapshot_name)
if not isdir(save_path):
mkdir(save_path)
validate(valloader, net, criterion, optimizer, True, save_path)
# -
# ### Load and save model
import torch
from os.path import join
snapshot_name = 'model-2999/model'
# +
#torch.save(net.state_dict(), join(base, 'models-local', snapshot_name + '.pth'))
#torch.save(optimizer.state_dict(), join(base, 'models', 'opt_' + snapshot_name + '.pth'))
# -
netCompare = UNet(1)
netCompare.load_state_dict(torch.load(join(base, 'models', snapshot_name + '.pth'), map_location={'cuda:0': 'cpu'}))
#optimizer.load_state_dict(torch.load(join(base, 'models', 'opt_' + snapshot_name + '.pth'), map_location={'cuda:0': 'cpu'}))
netCompare.train();
# ### Look at perforance on validation set
from torch.autograd import Variable
img, mask = val_dataset[0]
output = netCompare(Variable(img).unsqueeze(0))
prediction = F.sigmoid(output)
ax = plt.subplot(1,3,1);
plt.imshow(255*img.numpy().transpose(1,2,0))
ax.axis('off');
ax = plt.subplot(1,3,2);
plt.imshow(mask.numpy())
ax.axis('off');
ax = plt.subplot(1,3,3);
plt.imshow(prediction.squeeze(0).squeeze(0).data.numpy())
ax.axis('off');
loss = criterion(output, Variable(mask).unsqueeze(0))
print(loss.data.numpy()[0])
from skimage.io import imsave
# +
correct = 0
total = 0
ind = 0
for data in valloader:
images, labels = data
outputs = netCompare(Variable(images))
loss = criterion(outputs, Variable(labels)).data.numpy()[0]
prediction = F.sigmoid(outputs)
predict = prediction.squeeze(0).squeeze(0).data.numpy()
#imsave(join(base, 'predictions','predict_%04d.tif' % ind), (255*predict).astype('uint8'), plugin='tifffile', photometric='minisblack')
print(loss)
total += labels.size(0)
correct += loss
ind += 1
print('Mean loss: %.2f %%' % (
correct / total))
# -
test_dataset = BroadDataset(base, 'test', input_transform=input_transform, target_transform=target_transform)
testloader = DataLoader(test_dataset, batch_size=1,
shuffle=False, num_workers=2)
from skimage.io import imsave
ind = 0
for data in testloader:
images, labels = data
outputs = netCompare(Variable(images))
prediction = F.sigmoid(outputs)
predict = prediction.squeeze(0).squeeze(0).data.numpy()
imsave(join(base, 'test-model-2999','predict_%04d.tif' % ind), (255*predict).astype('uint8'), plugin='tifffile', photometric='minisblack')
ind += 1
|
notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## iPyLeaflet
# [ipyleaflet](https://github.com/ellisonbg/ipyleaflet) is a bridge between jupyter notebooks and the [leaflet](http://leafletjs.com/) javascript library for drawing maps.
#
# ipyleaflet comes with a few examples notebooks (this notebook was derived from one) but very little documentation,
# for more documentation read the [Leaflet IPA](http://leafletjs.com/reference.html)
#
# For installation directions, see the README on [ipyleaflet](https://github.com/ellisonbg/ipyleaflet)
#
# [GeoJson](https://tools.ietf.org/html/rfc7946#section-3.1.7) is a good way to add data on top of an ipyleaflet map.
#
# For other backgrounds than open streets, you can alter the ipyleaflet.py file to point to one of [these map providers](https://leaflet-extras.github.io/leaflet-providers/preview/)
# + run_control={"frozen": false, "read_only": false}
from ipyleaflet import (
Map,
Marker,
TileLayer, ImageOverlay,
Polyline, Polygon, Rectangle, Circle, CircleMarker,
GeoJSON,
DrawControl
)
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## Visualizing the distribution of the observations
#
# ## Load the required libraries
# + run_control={"frozen": false, "read_only": false}
import pandas as pd
import numpy as np
import sklearn as sk
import urllib
import math
import pylab as plt
import sys
sys.path.append('./lib')
from leaflet import *
# +
data_dir = "../../Data/Weather/"
from pickle import load
with open(data_dir+'/PRCP_residuals_PCA.pickle','rb') as file:
Res_eig=load(file)
print Res_eig.keys()
#stations.pkl
with open(data_dir+'/stations.pkl','rb') as file:
Stations=load(file)
Stations.head()
# -
import pandas as pd
Eig=pd.DataFrame(Res_eig['eigen-vecs'].transpose())
Eig.head()
Eig['station']=Res_eig['stations']
Eig=Eig.set_index('station')
print Res_eig['stations'][:5]
Eig.head()
table=Eig.join(Stations,how='left')
table=table[['name','latitude','longitude','elevation',0,1,2,3,4,5,6,7]]
table.head(4)
import pylab as plt
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
# + [markdown] run_control={"frozen": false, "read_only": false}
# ## Map
# + code_folding=[14]
min_lat,max_lat,min_long,max_long = box = (42.1103, 42.6167, -72.6, -70.8)
center = [(min_lat+max_lat)/2, (min_long+max_long)/2]
zoom = 9
m = Map(default_tiles=TileLayer(opacity=0.6), center=center, zoom=zoom)
r = Rectangle(bounds=[[min_lat,min_long],[max_lat,max_long]], weight=5, fill_opacity=0.0)
m += r
lat_margin=(max_lat-min_lat)/4
long_margin=(max_long-min_long)/4
for index,row in table.iterrows():
_lat=row['latitude']
_long=row['longitude']
for col in range(4):
_coef=row[col]
if np.isnan(_coef):
continue
r=abs(_coef)/5
#print r
color=colors[col]
signs=[[+1,+1],[+1,-1],[-1,-1],[-1,+1]]
lat_sign,long_sign=signs[col]
triangle=[(_lat,_long),(_lat+lat_sign*r,_long),(_lat,_long+long_sign*r),(_lat,_long)]
if _coef<0:
poly= Polygon(locations=triangle, weight=0,
color=color, opacity=0, fill_opacity=0.7,
fill_color=color)
else:
poly= Polygon(locations=triangle, weight=2,
color=color, opacity=0.8, fill_opacity=0,
fill_color=color)
m +=poly
m
# + [markdown] run_control={"frozen": false, "read_only": false}
# ### excercises:
# * Add a legend that relates the colors to values.
# * Leaflet supports a variety of maps. See if you can get a topographical map as the background.
# + run_control={"frozen": false, "read_only": false}
# %matplotlib inline
pdf.plot.scatter(x='elevation',y='avg(coeff_1)');
# -
|
TestCode/5.5 DataOnMaps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tarea 11
#
# _Tarea 11_ de _<NAME>_
# para el curso de __Métodos Numéricos__
# impartido por _<NAME>_.
#
# Fecha limite de entrega __...__.
# +
import sys
import seaborn as sns
import scipy
import numpy as np
import matplotlib.pyplot as plt
from scipy.linalg import solve_triangular # Para backward y forward substitution
NOTEBOOK = True
# -
# ## Ejercicio 1
#
# Programar el __metodo de Runge-Kutta__ de orden 2 para resolver un problemade valor inicial:
# y(a) =y0;
#
# $$ (PVI)
# \begin{cases}
# y' = f(x, y) \quad x \in (a, b] \\
# y(a) = y_0
# \end{cases}$$
# ### Solucion numerica del PVI
#
# Escribir el codigo de la funcion que calcula la solucion numerica del PVI con __algoritmo de RungeKutta__ de segundo orden. La funcion debe recibir como argumentos:
# - un arreglo con los puntos de la particion uniforme $a=x_0< x_1< \dots < x_n=n$,
# - un valor inicial $y0$,
# - el numero de subdivisiones $n$ del intervalo $[a, b]$, y
# - el apuntador la funcion $f(x, y)$.
#
# Crear el arreglo para almacenar los valores $y_0, y_1, \dots , y_n$. Hacer $y_0=y0$, $h=x_1 - x_0$
# y para cada $i= 0,1, \dots, n-1$ calcular
# \begin{eqnarray*}
# K_1 &=& f(x_i, y_i), \\
# K_2 &=& f(x_i+h, yi_+h K_1), \\
# y_{i+1} &=& y_i+ 0.5h(K_1+K_2)
# \end{eqnarray*}
# La funci ́on debe devolver el arreglo con los valores $y_i$.
def runge_kutta(xs, y0, n, f):
""" Funcion que implementa el algoritmo deRunge-Kutta. """
ys = np.zeros(n+1)
ys[0] = y0
h = xs[1] - xs[0]
for i in range(n):
K1 = f(xs[i], ys[i])
K2 = f(xs[i] + h, ys[i] + h*K1)
ys[i+1] = ys[i] + 0.5*h*(K1 + K2)
return ys
# ### Linea de comandos
#
# Escriba el programa que reciba desde la lınea de comandos el valor $n$ que define el numero de divisiones del intervalo de solucion $[a, b]$. Programe la funcion $f$ que corresponde al problema de valor inicial:
#
# $$ (PVS)
# \begin{cases}
# y' = 4x^2 - 6x + \frac{y}{x} \quad x\in(1,6] \\
# y(1) = 4
# \end{cases}$$
#
# La solucion analıtica de este problema es $y(x) =-6x2+ 8x+ 2x3$. Genere una particion del intervalo $[1,6]$ con $x_k= 1 +hk$, para $k= 0,1, \dots, n$, con $h= (6−1)/n$, y calcule los valores de la solucion numerica $y_0, y_1, y_2, \dots, y_n$ del _PVI_ usando la funcion del inciso anterior.
#
# Programe la funcion que evalua la solucion analıtica $y(x)$ y haga que el programa calcule el maximo del error relativo:
# $$E_{max}= \max_{k=1,\dots,n} \frac{|y_k - y(x_k)|}{|y(x_k)|} $$
#
# Haga que el programa imprima los valores $E_{max}$
# definimos f
def f(x, y):
return 4*x**2-6*x+(y/x)
def y(x):
return -6*x**2 + 8*x + 2*x**3
def gui(n, f=f, y=y, a=1, b=6, y0=4):
""" Funcion para llamar a la funcion del ejercicio anterior con un
caso especifico
"""
part = np.linspace(a, b, n+1, True)
ys = runge_kutta(part, y0, n, f)
xs = part
e_max = max([abs(ys[k] - y(xs[k]))
/ abs(y(xs[k]))
for k in range(1, n+1)])
print(e_max)
# ### Prueba
#
# Prueba el programa con $n= 10$ y $n=100$, y escriba un comentario sobre los resultados obtenidos.
gui(10)
gui(50)
gui(100)
# Además de los solicitados, también probe el metodo para $n = 50$. Me sorprende lo rapido que decience el error con _relativamente_ pocas iteraciones
|
MN/Tareas/T11/tarea11.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PIL import Image
from PIL import ImageFilter
from IPython.display import Image as JubImage
import numpy
# NOTE: last try before it somehow worked was pip3 install opencv-python
#
# Sorts pictures in current directory into two subdirs, blurred and ok
# Found original here: https://photo.stackexchange.com/questions/20432/is-there-photo-analysis-software-which-will-pre-sort-images-by-identifying-poten
# Modified for my own purposes.
#
# stock python libs
import os, shutil, re, sys
# additional libs
#import cv2
from pathlib import Path
# DEFAULTS
FOCUS_THRESHOLD = 20
DIRECTORY = Path.cwd()
EXT = '.JPG'
HELP_MESSAGE = f"""
This tool assists with blur dectection on images. The follow parameters are available:
FOCUS_THRESHOLD => --focus <int> [0-100]
DIRECTORY => --dir <path> [i.e., ./dir1/dir2/dir3 or C:/dir1/dir2/dir3]
EXT => --ext <ext> [.jpg, jpeg, etc]
Defaults for each are the following => FOCUS_THRESHOLD: {FOCUS_THRESHOLD} | DIRECTORY: {DIRECTORY} | EXT: {EXT}
"""
arguments = len(sys.argv)
position = 1
while position < arguments:
if sys.argv[position] == '--focus':
FOCUS_THRESHOLD = sys.argv[position + 1]
if isinstance(int(FOCUS_THRESHOLD), int):
FOCUS_THRESHOLD = int(FOCUS_THRESHOLD)
else:
print('ERROR: --focus parameter did not meet int requirements. See "--help" for options.')
exit()
elif sys.argv[position] == '--ext':
EXT = sys.argv[position + 1]
elif sys.argv[position] == '--dir':
DIRECTORY = Path(sys.argv[position + 1])
elif sys.argv[position] == '--help':
print(HELP_MESSAGE)
exit()
elif sys.argv[position] == '--h':
print(HELP_MESSAGE)
exit()
position = position + 1
#elif sys.argv[position] and sys.argv[position] != re.match('^--', sys.argv[position]):
# print('\nERROR: INVALID ARGUMENT. See the --help information below.\n' + HELP_MESSAGE)
# exit()
position = position + 1
print(f"PARAMETERS => FOCUS_THRESHOLD: {FOCUS_THRESHOLD} | DIRECTORY: '{DIRECTORY}' | EXT: {EXT}. For options, use '--help'")
BLURRED_DIR = 'blurred'
OK_DIR = 'ok'
blur_count = 0
print(str(Path.cwd()) + '\\' + str(DIRECTORY))
files = [f for f in os.listdir(DIRECTORY) if f.endswith(EXT)]
try:
os.makedirs(BLURRED_DIR)
os.makedirs(OK_DIR)
except:
pass
next = 'y'
for infile in files:
while next == 'y':
print(f'Processing file {infile}... {DIRECTORY}\{infile}')
print(str(Path.cwd()) + '\\' + str(DIRECTORY) + '\\' + infile)
image = Image.open(infile)
#print(image.getdata())
imageWithEdges = image.filter(ImageFilter.FIND_EDGES)
#imageWithEdges.show()
print(list(imageWithEdges.getdata()))
next = input('y for next, n for stop')
|
working_sample/2017.12.31/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
import numpy as np
import math
import functools
fact = math.factorial
def ordered(n, r=False):
if not r:
r = n
return math.factorial(n)/math.factorial(n-r)
def combine(n, r=False):
if not r:
r = n
return math.factorial(n)/(math.factorial(r)*math.factorial(n-r))
def non_distinct(n, d_list):
return math.factorial(n)/functools.reduce(lambda a, b : a * math.factorial(b), d_list, 1)
def split(word):
return [char for char in word]
# + language="markdown"
# For the following exercises, assume that there are n ways an event A can happen, m ways an event B can happen, and
# that A and B are non-overlapping.
# 1. Use the Addition Principle of counting to explain
# how many ways event A or B can occur.
# ## answer: If event A can happen $m$ times and event B can happen $n$ times and they are not related, they can happen in $m+n$ ways
# 2. Use the Multiplication Principle of counting to
# explain how many ways event A and B can occur.
# Answer the following questions.
# ## answer: if event A occurs after event B, they can occur in $A\cdot B$ ways
# 3. When given two separate events, how do we know
# whether to apply the Addition Principle or the
# Multiplication Principle when calculating possible
# outcomes? What conjunctions may help to determine
# which operations to use?
# ## answer: Assuming event two events A and B, if B occurs after A and have a combined outcome, then the Multiplication Principle should be used. if The events have no common outcome, the the Addition Priciple should be used.
# 4. Describe how the permutation of n objects differs
# from the permutation of choosing r objects from a
# set of n objects. Include how each is calculated.
# ## answer:Choosing $r$ objects from a set of $n$ objects limits the number of permutations.
# 5. What is the term for the arrangement that selects r
# objects from a set of n objects when the order of the
# r objects is not important? What is the formula for
# calculating the number of possible outcomes for this
# type of arrangement?
# ## answer: It is refered to as $n$ taken $r$ at a time, and the formula is $_nP_r = \frac{n!}{(n-r)!}$
# + language="markdown"
# For the following exercises, determine whether to use the Addition Principle or the Multiplication Principle. Then
# perform the calculations.
#
# + language="markdown"
# 7. Let the set B = { −23, −16, −7, −2, 20, 36, 48, 72}.
# How many ways are there to choose a positive or an
# odd number from A?
# ### $n_{positive} = 4$ ; $n_{negative} = 4$
# ### $ P = 8$
#
# + language="markdown"
# 9. How many ways are there to pick a paint color from 5 shades of green, 4 shades of blue, or 7 shades of
# yellow?
# ### $n_{green} = 5$ ; $n_{blue} = 4$ ; $n_{yellow} = 7$
# ### $ P = 16$
# + language="markdown"
# 11. How many outcomes are possible from tossing a coin
# and rolling a 6-sided die
# ### $n_{coin} = 2$ ; $n_{dice} = 6$
# ### $P = 12$
#
# + language="markdown"
# 13. How many ways are there to construct a string of 3
# digits if numbers can be repeated?
# ### $n_{digits} = 10$
# ### $P = 10^3 = 1000$
#
#
# -
9**3
# + language="markdown"
# For the following exercises, compute the value of the expression.
#
# + language="markdown"
# 15. P(5, 2)
# ### $_5P_2 = 20$
# -
ordered(5, 2)
# + language="markdown"
# 17. P(3, 3)
# ### $_nP_r = \frac{n!}{(n-r)!}$
# ### $_3P_3 = 20$
#
# + language="markdown"
# 19. P(11, 5)
# ### $_nP_r = \frac{n!}{(n-r)!}$
# ### $_11P_5 = 55440$
#
# + language="markdown"
# 21. C(12, 4)
# ### $_nC_r = \frac{n!}{r!(n-r)!}$
# ### $_{12}P_4 = 239500800$
# + language="markdown"
# 23. C(7, 6)
# ### $_nC_r = \frac{n!}{r!(n-r)!}$
# ### $_{7}P_{6} = 7$
# + language="markdown"
# For the following exercises, find the number of subsets in each given set.
#
# + language="markdown"
# 25. {1, 2, 3, 4, 5, 6, 7, 8, 9, 10}
# ### 1024
#
# + language="markdown"
# 27. A set containing 5 distinct numbers, 4 distinct
# letters, and 3 distinct symbols
# ### 4096
#
# + language="markdown"
# 29. The set of two-digit numbers between 1 and 100
# containing the digit 0
# 10, 20, 30, 40, 50, 60, 70, 80, 90
# ### 512
# + language="markdown"
# For the following exercises, find the distinct number of arrangements.
#
# + language="markdown"
# 31. The letters in the word “academia”
#
# + language="markdown"
# 33. The symbols in the string \#,\#,\#,\@,\@,\$,\$,\$,\%,\%,\%,\%
# ### 277200
#
# + language="markdown"
# 35) The set ,S consists of 900,000,000 whole numbers, each being the same number of digits long. How many digits long is a number from S?(Hint: use the fact that a whole number cannot start with the digit 0).
# ### 1) $S_{size} = 900000000$
# ### 2) $9\cdot10^8 = 900000000$
# ### 3)
# + language="markdown"
# 37) Can C(n, r) ever equal P(n,r)? Explain,
# ### $\frac{n!}{r!(n-r)!} = \frac{n!}{(n-r)!}$
# ### the nominators are the same, so lets compare the denominators:
# ### $r!(n-r)! = (n-r)!$ assume $n\ne r$
# ### $r! = 1 \Rightarrow r = 0 or r = 1$
#
# + language="markdown"
# 39) How many arrangements can be made from the letters of the word "mountains" if all the vowels must form a string?
# ### Compute the permutations of the vowls as a string:
# ### \[o,u,a,i\]
# ### $_nP_r = \frac{n!}{(n-r)!}$
# ### $_4P_4 = \frac{4!}{(0)!} \Rightarrow 24$
# ### Compute the distinct permutations of the consonants and a string of vowlsvand multiply it with the permutation of the string of vowls \[m, n. t, n, s,vowls]:
# ### $\frac{n!}{n_1!\cdot...n_k!}$
# ### $\frac{6!}{2!}\cdot24 \Rightarrow 8640$
# # answer: there are 8,640 different arrangements from the letters of the word mountains if all the vowels must form a string
#
# + language="markdown"
# 41) A cell phone company offers 6 different voice packages and 8 different data packages. Of those, 3 packages include both voice and data. How man ways are there to choose either voice or data, but not both?
# ### $num_{voice packages} = 6$
# ### $num_{data packages} = 8$
# ### $num_{voice \cap data} = 3$
# ### There are 5 packages that only include data, and 3 packages that only include voice, added together there are 8 ways to choose either one but not both
# + language="markdown"
# 43) A wholesale T-shirt company offers sizes small, medium, large, and XL in organic or non-organic cotton and colors white, black, gray, blue and red. How many different T-shirts are there to choose from?
# ### Sizes: \[s, m, l, xl\]
# ### Cotton: \[organic, non\]
# ### Color: \[white, black, gray, blue, red\]
# ### There are 4 Size options, 2 cotton options, and 5 color options, multiplied together it produces 40 different t shirts.
#
# + language="markdown"
# 45) An art store has 4 brands of paint pens in 12 different colors and 3 types of ink. How many paint pens are there to choose from?
# ### brands:4
# ### Colors: 12
# ### ink Type: 3
# # answer: there are 144 different pens to choose from
#
# + language="markdown"
# 47)How many ways can a baseball coach arrange the order of 9 batters if there are 15 players on the team?
# ### $_nP_r = \frac{n!}{(n-r)!}$
# ### $_{15}P_P9 = \frac{15!}{(15-9)! = 1,816,214,400}$
# + language="markdown"
# 49) A motorcycle shop has 10 choppers, 6 bobbers and 5 cafe racers - different types of vintage motorcycles. How many ways can the shop choose 3 choppers, 5 bobbers and 2 cafe racers for a weekend showcase?
# ### choppers:10
# ### bobbers: 6
# ### cafe racers: 5
# ### $_nP_r = \frac{n!}{(n-r)!}$
# ### choppers: $_{10}P_3 = \frac{10!}{(7)!} \Rightarrow 120$
# ### bobbers: $_6P_5 = \frac{6!}{(1)!} \Rightarrow 6$
# ### cafe racers: $_5P_2 = \frac{5!}{(3)!} \Rightarrow 10$
# # answer: There are 7,200 ways for him to organize the bikes for the shocase
# + language="markdown"
# 51) Just-For-Kicks Sneaker Company offers an online customizing service. How any ways are there to design a custom pair of Just-For_kicks sneakers if a customer can choose from a basic shoe up to 11 customizable options?
# ### 2^11 = 2048
#
# + language="markdown"
# 53) Susan bought 20 plants to arrange along the border of her garden. How many distinct arrangements can she make if the plants are comprised of 6 tulips, 6 roses, and 8 daises?
# ### Tulips:6
# ### Roses: 6
# ### Daisies: 8
# ### arrangements: $\frac{20!}{6!\cdot6!\cdot8!} \Rightarrow 116,396,280$
# -
# math.factorial(10)
# l = list("###@@$$$%%%%")
# print (l)
# print(len(l))
# ordered(10, 3)*ordered(6, 5)*ordered(5, 2)
# print(fact(15))
# print(fact(6))
# print(fact(15)/fact(6))
#combine
# 3*4*12
non_distinct(20,[6, 6, 8])
# 2**11
|
Math/Precalculus/Chapter 11 Sequences,Probability and Counting/part05-exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Algorithms Exercise 3
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
# + nbgrader={}
from IPython.html.widgets import interact
# + [markdown] nbgrader={}
# ## Character counting and entropy
# + [markdown] nbgrader={}
# Write a function `char_probs` that takes a string and computes the probabilities of each character in the string:
#
# * First do a character count and store the result in a dictionary.
# * Then divide each character counts by the total number of character to compute the normalized probabilties.
# * Return the dictionary of characters (keys) and probabilities (values).
# + nbgrader={"checksum": "f11bac096ada913538c9a47721fb98a1", "solution": true}
def char_probs(s):
"""Find the probabilities of the unique characters in the string s.
Parameters
----------
s : str
A string of characters.
Returns
-------
probs : dict
A dictionary whose keys are the unique characters in s and whose values
are the probabilities of those characters.
"""
dic = {}
for x in range(len(s)):
if s[x] in dic.keys():
dic[s[x]] += 1.
else:
dic[s[x]] = 1.
for a in dic:
dic[a] = dic[a]/len(s)
return dic
# + deletable=false nbgrader={"checksum": "97f4091c66f9a555c766706bcf4a7681", "grade": true, "grade_id": "algorithmsex03a", "points": 4}
test1 = char_probs('aaaa')
assert np.allclose(test1['a'], 1.0)
test2 = char_probs('aabb')
assert np.allclose(test2['a'], 0.5)
assert np.allclose(test2['b'], 0.5)
test3 = char_probs('abcd')
assert np.allclose(test3['a'], 0.25)
assert np.allclose(test3['b'], 0.25)
assert np.allclose(test3['c'], 0.25)
assert np.allclose(test3['d'], 0.25)
# + [markdown] nbgrader={}
# The [entropy](http://en.wikipedia.org/wiki/Entropy_%28information_theory%29) is a quantiative measure of the disorder of a probability distribution. It is used extensively in Physics, Statistics, Machine Learning, Computer Science and Information Science. Given a set of probabilities $P_i$, the entropy is defined as:
#
# $$H = - \Sigma_i P_i \log_2(P_i)$$
#
# In this expression $\log_2$ is the base 2 log (`np.log2`), which is commonly used in information science. In Physics the natural log is often used in the definition of entropy.
#
# Write a funtion `entropy` that computes the entropy of a probability distribution. The probability distribution will be passed as a Python `dict`: the values in the `dict` will be the probabilities.
#
# To compute the entropy, you should:
#
# * First convert the values (probabilities) of the `dict` to a Numpy array of probabilities.
# * Then use other Numpy functions (`np.log2`, etc.) to compute the entropy.
# * Don't use any `for` or `while` loops in your code.
# + nbgrader={"checksum": "93e205f7727df5161387fa73af53718b", "solution": true}
def entropy(d):
x = np.zeros(len(d))
a=0
for b in d:
x[a]= d[b]
a+=1
c = 0
for e in x:
c += e*np.log2(e)
c *= -1
return c
# + deletable=false nbgrader={"checksum": "0499a53c730bb4fbb2cd81d7c34486da", "grade": true, "grade_id": "algorithmsex03b", "points": 4}
assert np.allclose(entropy({'a': 0.5, 'b': 0.5}), 1.0)
assert np.allclose(entropy({'a': 1.0}), 0.0)
# + [markdown] nbgrader={}
# Use IPython's `interact` function to create a user interface that allows you to type a string into a text box and see the entropy of the character probabilities of the string.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def ent_of_string(s):
print(entropy(char_probs(s)))
interact(ent_of_string, s='Type String Here');
# + deletable=false nbgrader={"checksum": "2eeb2ebb1993a6f046deec7ff81c4930", "grade": true, "grade_id": "algorithmsex03c", "points": 2}
assert True # use this for grading the pi digits histogram
|
assignments/midterm/AlgorithmsEx03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2 style = "color : Brown">Case Study - Sales Data </h2>
# All imports
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# ##### Sales and Profit data is read in dataframe "sales"
# +
# Read file
sales = pd.read_excel('sales.xlsx')
sales
# +
# Read file and set 1st two columns as index
sales = pd.read_excel('sales.xlsx', index_col = [0,1])
sales
# -
# <h4 style = "color : Sky blue"> Example - 1</h4>
#
# ##### Display first 3 land last 3 rows of the sales dataframe
#
sales.head() # Default - returns top 5 rows
sales.head(3)
sales.tail()
sales.tail(3)
# <h4 style = "color : Sky blue"> Example - 2</h4>
#
# ##### Display the information about the data stored in data frame
#
sales.info()
# ##### Display the statistical information about the data in dataframe
sales.describe()
sales[["Sales", "Profit"]].plot(kind= "box", subplots= True)
plt.show()
sales["Profit"]
|
Session+-+2_part_2-Teaching.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Forecasting Air Passenger using XGBoost
# +
import numpy as np
import pandas as pd
import os
import warnings
from copy import copy
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error, mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
import xgboost as xgb
# perosnal wrapper created for time-series plots
import ts_utils
# -
plt.rcParams['figure.figsize'] = 15, 6
plt.style.use('ggplot')
warnings.filterwarnings("ignore")
# ## Data preparation
DATADIR = '../data/air-passenger/'
data_path = os.path.join(DATADIR, 'AirPassengers.csv')
MODELDIR = '../checkpoints/air-passenger/xgb/model/'
df = pd.read_csv(data_path, usecols=[1], header=0, engine='python')
data = df.values
data = data.astype('float32')
plt.plot(data)
plt.title('#Air Passengers from 1949 to 1960')
plt.ylabel('#thousands of passengers')
plt.xlabel('Years')
plt.tight_layout()
plt.show()
scaled_data = np.log(data)
print(scaled_data[:5])
plt.plot(scaled_data)
plt.title('#Air Passengers from 1949 to 1960')
plt.ylabel('Scaled #passengers')
plt.xlabel('Years')
plt.tight_layout()
plt.show()
# ## Train test split
train, test = train_test_split(scaled_data, train_size=0.8 ,shuffle=False)
len(train), len(test)
X_train, y_train = ts_utils.prepare_data(train, time_step=1)
X_test, y_test = ts_utils.prepare_data(test, time_step=1)
print(X_train.shape)
print(X_test.shape)
# ## Parameter selection using Grid search
# +
# seed = 42
# grid_param = {
# 'max_depth': [3, 5],
# 'n_estimators': [100, 200],
# 'learning_rate': [0.01, 0.1]
# }
# +
# xgb_param = {'objective' :'reg:linear', 'silent' : 1, 'colsample_bytree' : 1.0}
# +
# xgb_grid = GridSearchCV(estimator=xgb.XGBRegressor(**xgb_param, seed=seed),
# param_grid=grid_param)
# +
# xgb_grid.fit(X_train, y_train)
# +
# print(f'Best estimator: {xgb_grid.best_estimator_}')
# +
# print(f'Best Paramters: {xgb_grid.best_params_}')
# +
# print(f'Best result: {xgb_grid.best_score_}')
# -
# ## Model fitting
params = {'objective' :'reg:linear',
'colsample_bytree' : 1,
'learning_rate' : 0.1,
'max_depth': 5,
'alpha' : 10,
'n_estimators' : 200}
model = xgb.XGBRegressor(**params)
eval_set = [(X_test, y_test)]
model.fit(X_train, y_train, early_stopping_rounds=10, eval_set=eval_set, verbose=False)
# ## Prediction and evaluation
# +
# prediction
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
# the results are in the form of scaled value, so inverse the transformation
y_train_pred_inv = np.exp(y_train_pred)
y_test_pred_inv = np.exp(y_test_pred)
# will be used for calculating MAE and RMSE
y_train_inv = np.exp(y_train)
y_test_inv = np.exp(y_test)
# MAE and RMSE calculation
train_rmse = np.sqrt(mean_squared_error(y_train_inv, y_train_pred_inv))
train_mae = mean_absolute_error(y_train_inv, y_train_pred_inv)
train_nrmse = train_rmse/np.std(y_train_inv)
test_rmse = np.sqrt(mean_squared_error(y_test_inv, y_test_pred_inv))
test_mae = mean_absolute_error(y_test_inv, y_test_pred_inv)
test_nrmse = test_rmse/np.std(y_test_inv)
print(f'Training NRMSE: {train_nrmse}')
print(f'Training MAE: {train_mae}')
print(f'Test NRMSE: {test_nrmse}')
print(f'Test MAE: {test_mae}')
# -
plt.plot(y_test_inv, label='test')
plt.plot(y_test_pred_inv, label='test predicted')
plt.xlabel('time')
plt.ylabel('#passengers')
plt.title('Actual vs predicted on test data using XGBoost', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
plt.plot(y_train_inv, label='actual')
plt.plot(y_train_pred_inv, label='predicted')
plt.ylabel('#passengers')
plt.xlabel('time')
plt.title('Actual vs Predicted on Training data', fontsize=14)
plt.legend()
plt.tight_layout()
plt.show()
actual_data = np.vstack((y_train_inv, y_test_inv))
actual_data.shape
# +
test_plot = np.empty_like(actual_data)
test_plot[:, :] = np.nan
test_plot[len(y_train_pred_inv):len(actual_data):, ] = y_test_pred_inv.reshape(-1, 1)
plt.plot(actual_data, label='actual')
plt.plot(y_train_pred_inv, color='blue', linestyle='-.', label='training prediction')
plt.plot(test_plot, color='green', linestyle='-', label='test prediction')
plt.xlabel('Years')
plt.ylabel('#thousand air passengers')
plt.title('Actual v/s Predicted using XGBoost', fontsize=15)
plt.legend()
plt.tight_layout()
plt.show()
# -
|
01.air_passenger/08.passenger-forecasting-XGBoost.ipynb
|