code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# Import Dependencies
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.cluster import KMeans
# # Data Prep
#Load CSV
crypto_df = pd.read_csv('crypto_data.csv')
crypto_df.head()
# Filter for currencies currently being traded
crypto_df = crypto_df[crypto_df['IsTrading'] == True]
crypto_df.head()
# Remove rows with null values
crypto_df = crypto_df.dropna()
crypto_df.head()
# Filter for cryptocurrencies that have been mined
crypto_df = crypto_df[crypto_df["TotalCoinsMined"]> 0]
crypto_df.head()
# Delete the CoinName from the dataframe
crypto_df.drop(columns='CoinName', axis=1, inplace= True)
crypto_df = crypto_df.drop(columns=['Unnamed: 0'])
crypto_df
# Convert Algorithm and ProofType
X = pd.get_dummies(crypto_df, columns=['Algorithm', 'ProofType'])
X
# ***Examine the number of rows and columns of your dataset now. How did they change?***
#
# The number of columns increased from 6 to 100, creating a column for each type.
# Standarize dataset
scaler = StandardScaler()
crypto_scaled = scaler.fit_transform(X)
# # Dimensionality Reduction
# Apply PCA
pca = PCA(n_components=.9)
crypto_pca = pca.fit_transform(crypto_scaled)
crypto_pca
# Transform to DataFrame
crypto_pca_df = pd.DataFrame(data=crypto_pca)
crypto_pca_df
# **How did it change?**
# The number of columns reduced from 99 to 74.
# Explained Variance
pca.explained_variance_ratio_
# Initialize t-SNE
tsne = TSNE(learning_rate=35)
# Reduce dimensionsa and check that dimensions have been reduced to two
tsne_features = tsne.fit_transform(crypto_pca_df)
tsne_features.shape
# +
# First column & Second column
x = tsne_features[:,0]
y = tsne_features[:,1]
# Plot the results
plt.scatter(x, y)
plt.show()
# -
# # Cluster Analysis with k-Means
# +
inertia = []
k = list(range(1, 11))
# Use a for-loop to determine the inertia for each k between 1-10
for i in k:
km = KMeans(n_clusters=i, random_state=42)
km.fit(crypto_pca_df)
inertia.append(km.inertia_)
# Create an elbow plot
elbow_data = {"k": k, "inertia": inertia}
df_elbow = pd.DataFrame(elbow_data)
plt.plot(df_elbow['k'], df_elbow['inertia'])
plt.xticks(range(1,11))
plt.xlabel('Number of Clusters')
plt.ylabel('Inertia')
plt.show()
# -
# **Recommendation**
#
# The best k value appears to be 6. An output of 6 clusters would be best to categorize the crytocurrencies.
| Cryptocurrency Clusters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="U2dD1VOYIKLy" colab_type="text"
# # Denoising mel-spectrograms with a residual dense network
# + [markdown] id="7_gkiuH1IKLz" colab_type="text"
# ## Setup
# + [markdown] id="sUADgWKkNKxb" colab_type="text"
# ### Environment
# + id="kk6zcnjMNJxe" colab_type="code" outputId="a2773f05-4f03-4a4d-fc4b-40cb899cce8f" colab={"base_uri": "https://localhost:8080/", "height": 119}
# !pip install -q ipython_secrets gsheet-keyring comet_ml
# + id="D2_1v8kP8cDG" colab_type="code" outputId="af4812f7-9209-4a12-db4c-6f257a4769eb" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %%writefile setup.sh
export CUDA_HOME=/usr/local/cuda-10.1
[[ -d ./apex ]] || git clone https://github.com/NVIDIA/apex && pip install -v --no-cache-dir --global-option="--cpp_ext" --global-option="--cuda_ext" ./apex
# + id="SpYIG8TN8dhL" colab_type="code" outputId="af30dedf-a069-4eae-fc86-7e00602df6dc" colab={"base_uri": "https://localhost:8080/", "height": 51}
# !sh setup.sh
# + [markdown] id="Vg4F4fwrIKL4" colab_type="text"
# ### Variables
# + id="IZECU7xxIKL4" colab_type="code" outputId="e0bb26d9-9290-4277-bfb8-a8971fb27ad1" colab={"base_uri": "https://localhost:8080/", "height": 34}
from ipython_secrets import get_secret
CODEBASE = "https://github.com/sdll/audio-denoising"
PROJECT = "Arbeit"
COMET_ML_API_KEY = get_secret("comet-{}".format(PROJECT))
GDRIVE_MOUNT_POINT = "/content/drive"
TRAIN_DATASET = "dataset/train"
VAL_DATASET = "dataset/val"
WORKSPACE= "sdll"
from google.colab import drive
import os
drive.mount(GDRIVE_MOUNT_POINT)
GOOGLE_DRIVE_ROOT = GDRIVE_MOUNT_POINT + "/" + list(filter(lambda x: x[0] != '.', os.listdir(GDRIVE_MOUNT_POINT)))[0]
# + [markdown] id="sZ1yH07oIL_l" colab_type="text"
# ### Data
# + id="o3BYoRWjI7Ok" colab_type="code" outputId="5ca31ec3-3160-43bb-a9d2-1eb4714bf4d4" colab={"base_uri": "https://localhost:8080/", "height": 119}
# !git clone $CODEBASE
# + id="vFGml0y6KZCj" colab_type="code" outputId="0191bc67-5e7b-453a-b71d-916eff5fd4a2" colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd audio-denoising
# + id="4JBGXUFxKgzT" colab_type="code" outputId="2b731512-28c1-45e4-c938-f32333ebc9c2" colab={"base_uri": "https://localhost:8080/", "height": 853}
# !mkdir -p dataset
![[ -f ./dataset/train.zip ]] || wget https://www.dropbox.com/s/n6nhp5e231rl0b5/train.zip?raw=1 -O dataset/train.zip
![[ -f ./dataset/test.zip ]] || wget https://www.dropbox.com/s/nt4q2n0esiboc1i/val.zip?raw=1 -O dataset/val.zip
# + id="T2yImuBiMecl" colab_type="code" outputId="a8448d35-595c-477a-92c5-c85a50757220" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %cd dataset
![[ -d train ]] || unzip -qq train.zip
![[ -d val ]] || unzip -qq val.zip
# %cd ..
# + id="MYGELWxWkFf8" colab_type="code" colab={}
![[ -d pytorch_ssim ]] || (git clone https://github.com/Po-Hsun-Su/pytorch-ssim && mv pytorch-ssim/pytorch_ssim . && rm -rf pytorch-ssim)
# + [markdown] id="-Terp4boIKL0" colab_type="text"
# ### Imports
# + id="_XdbUG3iIKL0" colab_type="code" outputId="4847c4a1-6c20-4ce4-ad9c-e13e1b1f0238" colab={"base_uri": "https://localhost:8080/", "height": 88}
# %cd /content/audio-denoising/
from comet_ml import Experiment
import argparse
from timeit import default_timer as timer
import numpy as np
import seaborn as sns
import torch
from matplotlib import pyplot as plt
from torch import nn, optim
from tqdm import tqdm as tqdm_base
from audio_denoising.data.loader import load
from audio_denoising.model.rdn import ResidualDenseNetwork as Model
from pytorch_ssim import ssim
def tqdm(*args, **kwargs):
if hasattr(tqdm_base, "_instances"):
for instance in list(tqdm_base._instances):
tqdm_base._decr_instances(instance)
return tqdm_base(*args, **kwargs)
# + id="AhEuYFXWQSQy" colab_type="code" colab={}
from apex import amp, optimizers
# + id="_BrFndbpNrAw" colab_type="code" colab={}
sns.set()
# + [markdown] id="GmOFsyYuw4TA" colab_type="text"
# ## Training
# + id="bMrCaY8zxFGT" colab_type="code" colab={}
def get_arg_parser():
parser = argparse.ArgumentParser()
parser.add_argument("--batch-size", type=int, default=8)
parser.add_argument("--growth-rate", type=int, default=16)
parser.add_argument("--kernel-size", type=int, default=3)
parser.add_argument("--lr", type=float, default=1e-3)
parser.add_argument("--num-blocks", type=int, default=20)
parser.add_argument("--num-channels", type=int, default=1)
parser.add_argument("--num-epochs", type=int, default=80)
parser.add_argument("--num-features", type=int, default=16)
parser.add_argument("--num-layers", type=int, default=6)
parser.add_argument("--seed", type=int, default=42)
return parser
def get_criterion():
return nn.MSELoss()
def get_optimizer(model, lr=1e-3):
return optimizers.FusedAdam(model.parameters(), lr)
# return optim.SGD(model.parameters(), lr)
def psnr(prediction, target, max_pixel=255.0):
return 10.0 * ((max_pixel ** 2) / ((prediction - target) ** 2).mean()).log10()
def train(
experiment,
loader,
model,
criterion,
optimizer,
args,
device=torch.device("cuda" if torch.cuda.is_available() else "cpu"),
scheduler=None,
validator=None, # (validate, val_loader)
verbose=True,
):
experiment.log_parameters(vars(args))
np.random.seed(args.seed)
# save the model weights with the best psnr
if validator:
best_psnr = 0.0
for epoch in tqdm(range(args.num_epochs), desc="Epoch", unit="epochs"):
with experiment.train():
model.train()
train_psnr = []
train_ssim = []
for clean_image, noisy_image in tqdm(
loader, desc="Train images", unit="batches"
):
image = noisy_image.to(device, dtype=torch.float)
gt_image = clean_image.to(device, dtype=torch.float)
noise = image - gt_image
prediction = model(image)
cleaned_image = image - prediction
loss = criterion(prediction, noise)
with amp.scale_loss(loss, optimizer) as scaled_loss:
scaled_loss.backward()
optimizer.step()
optimizer.zero_grad()
current_psnr = psnr(cleaned_image, gt_image).data.item()
current_ssim = ssim(cleaned_image, gt_image).data.item()
train_psnr.append(current_psnr)
train_ssim.append(current_ssim)
experiment.log_metric("psnr", current_psnr)
experiment.log_metric("ssim", current_ssim)
experiment.log_metric("loss", scaled_loss.data.item())
experiment.log_metric("mean_psnr", np.mean(train_psnr))
experiment.log_metric("mean_ssim", np.mean(train_ssim))
if validator:
validate, val_loader = validator
test_psnr, _, _ = validate(experiment, val_loader, model, device, verbose)
if scheduler:
scheduler.step(test_psnr)
if test_psnr > best_psnr:
best_psnr = test_psnr
filename_pth = GOOGLE_DRIVE_ROOT + '/audio_denoising_psnr_{:.4f}_epoch_{}_D_{}_C_{}_G_{}_G0_{}.pth'.format(
test_psnr, epoch, args.num_blocks, args.num_layers, args.growth_rate, args.num_features
)
torch.save(model.state_dict(), filename_pth)
return model
def validate(
experiment,
loader,
model,
device=torch.device("cuda" if torch.cuda.is_available() else "cpu"),
verbose=True,
):
with experiment.test():
model.eval()
test_psnr = []
test_ssim = []
test_prediction_times = []
for clean_image, noisy_image in tqdm(loader, desc="Val images", unit="batches"):
image = noisy_image.to(device, dtype=torch.float)
gt_image = clean_image.to(device, dtype=torch.float)
start = timer()
prediction = model(image)
end = timer()
cleaned_image = image - prediction
prediction_time = end - start
test_prediction_times.append(prediction_time)
experiment.log_metric("prediction_time", prediction_time)
current_psnr = psnr(cleaned_image, gt_image).data.item()
current_ssim = ssim(cleaned_image, gt_image).data.item()
test_psnr.append(current_psnr)
test_ssim.append(current_ssim)
test_psnr = np.mean(test_psnr)
test_ssim = np.mean(test_ssim)
test_prediction_time = np.mean(test_prediction_times)
experiment.log_metric("mean_psnr", test_psnr)
experiment.log_metric("mean_ssim", test_ssim)
experiment.log_metric("mean_prediction_time", test_prediction_time)
if verbose:
print(
"\nMean Test PSNR: {:.2f}\nMean Test SSIM: {:.2f}\nMean Prediction Time: {:.2f}".format(
test_psnr, test_ssim, test_prediction_time
)
)
return test_psnr, test_ssim, test_prediction_time
# + id="M6NanS3srrLo" colab_type="code" outputId="6786301c-80b8-4d16-a1ac-734d3088fae2" colab={"base_uri": "https://localhost:8080/", "height": 51}
experiment = Experiment(
api_key=COMET_ML_API_KEY,
project_name=PROJECT,
workspace=WORKSPACE,
auto_output_logging=None,
)
# + id="qR-cwjqpVf-l" colab_type="code" colab={}
args = get_arg_parser().parse_args(args=[])
train_loader = load(TRAIN_DATASET, batch_size=args.batch_size)
val_loader = load(VAL_DATASET, batch_size=1)
device=torch.device("cuda" if torch.cuda.is_available() else "cpu")
model = Model(args).to(device)
# + id="F5-x3BOAQ8KK" colab_type="code" colab={}
optimizer = get_optimizer(model, args.lr)
model, optimizer = amp.initialize(model, optimizer, opt_level="O2")
criterion = get_criterion()
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, mode='max', factor=0.75, patience=0, verbose=True)
# + id="xYR1sApR8jtb" colab_type="code" outputId="0ae7bc1d-6c38-498f-f0a0-9780080d5cae" colab={"base_uri": "https://localhost:8080/", "height": 1000}
train(
experiment,
train_loader,
model,
criterion,
optimizer,
args,
device,
scheduler=scheduler,
validator=(validate, val_loader)
)
# + [markdown] id="wL9EVJN5SB4O" colab_type="text"
# The Comet logs of this round of training can be found [here](https://www.comet.ml/sdll/arbeit/a40a28c88443436984eedc52913c3a09).
# + id="xSIeVW1Xym7Y" colab_type="code" colab={}
filename_pth = GOOGLE_DRIVE_ROOT + '/audio_denoising_psnr_{:.4f}_epoch_{}.pth'.format(
59.71, 35
)
torch.save(model.state_dict(), filename_pth)
# + [markdown] id="Y5R_Mf80sz9F" colab_type="text"
# ## Validation
# + id="0vrtuy39zB4M" colab_type="code" outputId="16ab9080-5a5a-4cda-c8eb-62ae79d63d8d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# ckpt = GOOGLE_DRIVE_ROOT + '/audio_denoising_psnr_59.7472_epoch_6.pth'
ckpt = GOOGLE_DRIVE_ROOT + "/audio_denoising_psnr_65.1736_epoch_15_D_20_C_6_G_16_G0_16.pth"
model.load_state_dict(torch.load(ckpt))
# + id="zu1qq-Wx4FVS" colab_type="code" outputId="6d866187-2877-48d9-9037-8bde23e8261b" colab={"base_uri": "https://localhost:8080/", "height": 102}
test_psnr, test_ssim, test_prediction_time = validate(experiment, val_loader, model, verbose=True)
# + id="p1Kgfrk452eQ" colab_type="code" outputId="fcbb2de8-c537-4ddc-ab05-57fbac9dff89" colab={"base_uri": "https://localhost:8080/", "height": 685}
# %matplotlib inline
from IPython.display import clear_output, display
import sys
import time
for clean_mel, noisy_mel in val_loader:
time.sleep(.25)
clean_mel = clean_mel.to("cpu", dtype=torch.float)
noisy_mel = noisy_mel.to("cpu", dtype=torch.float)
fig = plt.figure(figsize=(17, 8), dpi=100)
ax1 = fig.add_subplot(121)
ax2 = fig.add_subplot(122)
ax1.imshow(clean_mel.squeeze(), interpolation='nearest', aspect='auto')
ax1.set_title('Clean image')
ax2.imshow((noisy_mel - model(noisy_mel.to(device)).data.to("cpu")).squeeze(),
interpolation='nearest', aspect='auto')
ax2.set_title('Cleaned image')
display(fig)
clear_output(wait=True)
sys.stdout.flush()
break
# + [markdown] id="me4AFTr4RjtK" colab_type="text"
# ## SSIM Inference Analysis
# + id="M7S3t2LFRruf" colab_type="code" colab={}
from audio_denoising.data.loader import SpectogramDataset
clean_dataset = SpectogramDataset(VAL_DATASET + "/clean")
noisy_dataset = SpectogramDataset(VAL_DATASET + "/noisy")
# + id="UfrX9xM3SjXe" colab_type="code" colab={}
def compute_ssims(dataset):
ssims = []
for file_idx in tqdm(range(len(dataset)), unit="files"):
img = dataset[file_idx].unsqueeze(0).to(device, dtype=torch.float)
noise = model(img).to("cpu")
img = img.to("cpu")
ssims.append(ssim(img - noise, img).data.item())
return ssims
# + id="h5tm9yAKTetz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6dc9db93-ce08-4f4d-fca1-18d9785f676a"
clean_ssims = compute_ssims(clean_dataset)
# + id="QuHrDwXKVnPe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="ca784915-6d36-468f-9015-8e81a18fb3cd"
plt.figure(dpi=100)
sns.distplot(clean_ssims)
plt.show()
# + id="s42_lVlcUV6b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="07c2b41d-1bb8-4584-f8d0-7f029cab346d"
noisy_ssims = compute_ssims(noisy_dataset)
# + id="lNLT30wzVu_Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="62f1fc97-8c5a-4fee-8f19-9c0e9755af8a"
plt.figure(dpi=100)
sns.distplot(noisy_ssims)
plt.show()
# + id="A_vbWvymWF9M" colab_type="code" colab={}
clean_ssims = np.asarray(clean_ssims)
noisy_ssims = np.asarray(noisy_ssims)
threshold = 0.77
true_clean = np.sum(clean_ssims > threshold) / len(clean_ssims)
true_noisy = np.sum(noisy_ssims < threshold) / len(noisy_ssims)
# + id="RhPF0Ey3WlLy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f03a760f-258d-4555-bbaa-cf4753085f98"
print("True clean ratio: {}".format(true_clean))
print("True noisy ratio: {}".format(true_noisy))
# + [markdown] id="r5loyU7HOGru" colab_type="text"
# ## Scratchpad
# + id="2aECe_tpIKMB" colab_type="code" colab={}
dataset = load(TRAIN_DATASET, batch_size=1)
heights = []
for clean_mel, noisy_mel in dataset:
heights.append(clean_mel.shape[2])
# + id="bQ68YeMcNeqZ" colab_type="code" colab={}
plt.figure(dpi=120)
plt.title("Distribution of Train MEL Heights")
plt.xlabel("Height, px")
plt.ylabel("Frequency")
sns.distplot(heights)
plt.show()
# + id="C6aT3SMXTCm6" colab_type="code" colab={}
min(heights), max(heights)
# + id="fYgcbPhzqoAb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="decffacd-b0fb-427d-94c5-b93542a6df96"
# %cd /content/audio-denoising/
# !git pull
# + id="0BL6I0v-s7EE" colab_type="code" colab={}
# %%capture
args = get_arg_parser().parse_args(args=[])
train_loader = load(TRAIN_DATASET, batch_size=args.batch_size)
val_loader = load(VAL_DATASET, batch_size=1)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
experiment = Experiment(
api_key=COMET_ML_API_KEY,
project_name=PROJECT,
workspace=WORKSPACE,
auto_output_logging=None,
)
grid_search = {
"num_blocks": np.arange(3, 12, 3),
"num_layers": np.arange(6, 18, 3),
"num_features": np.arange(8, 32, 8),
"growth_rate": np.arange(8, 32, 8),
}
best = {
"num_blocks": None,
"num_layers": None,
"num_features": None,
"growth_rate": None,
}
args.num_epochs = 2
best_psnr = 0.0
for num_blocks in grid_search["num_blocks"]:
for num_layers in grid_search["num_layers"]:
for num_features in grid_search["num_features"]:
for growth_rate in grid_search["growth_rate"]:
setattr(args, "num_blocks", num_blocks)
setattr(args, "num_layers", num_layers)
setattr(args, "num_features", num_features)
setattr(args, "growth_rate", growth_rate)
model = Model(args).to(device)
optimizer = get_optimizer(model, args.lr)
model, optimizer = amp.initialize(model, optimizer, opt_level="O3")
criterion = get_criterion()
scheduler = optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode="max", factor=0.75, patience=0, verbose=True
)
train(
experiment,
[next(iter(train_loader))],
model,
criterion,
optimizer,
args,
device,
scheduler=scheduler,
)
test_psnr, _, _ = validate(
experiment, [next(iter(val_loader))], model, device, verbose=False
)
if test_psnr > best_psnr:
best_psnr = test_psnr
best["num_blocks"] = num_blocks
best["num_layers"] = num_layers
best["num_features"] = num_features
best["growth_rate"] = growth_rate
del model, optimizer, criterion, scheduler
torch.cuda.empty_cache()
experiment.end()
# + id="4AhNVJKWXMNX" colab_type="code" colab={}
print(best)
# + id="eCxMyxWG9pbH" colab_type="code" colab={}
while True:
pass
# + id="6-LRKqrc9qiZ" colab_type="code" colab={}
next(iter(val_loader))
# + id="fyiQ6otaV6Sa" colab_type="code" colab={}
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Detecting and Analyzing Faces
#
# Computer vision solutions often require an artificial intelligence (AI) solution to be able to detect, analyze, or identify human faces. or example, suppose the retail company Northwind Traders has decided to implement a "smart store", in which AI services monitor the store to identify customers requiring assistance, and direct employees to help them. One way to accomplish this is to perform facial detection and analysis - in other words, determine if there are any faces in the images, and if so analyze their features.
#
# 
#
# ## Use the Face cognitive service to detect faces
#
# Suppose the smart store system that Northwind Traders wants to create needs to be able to detect customers and analyze their facial features. In Microsoft Azure, you can use **Face**, part of Azure Cognitive Services to do this.
#
# ### Create a Cognitive Services Resource
#
# Let's start by creating a **Cognitive Services** resource in your Azure subscription.
#
# > **Note**: If you already have a Cognitive Services resource, just open its **Quick start** page in the Azure portal and copy its key and endpoint to the cell below. Otherwise, follow the steps below to create one.
#
# 1. In another browser tab, open the Azure portal at https://portal.azure.com, signing in with your Microsoft account.
# 2. Click the **+Create a resource** button, search for *Cognitive Services*, and create a **Cognitive Services** resource with the following settings:
# - **Name**: *Enter a unique name*.
# - **Subscription**: *Your Azure subscription*.
# - **Location**: *Choose any available region*:
# - **Pricing tier**: S0
# - **Resource group**: *Create a resource group with a unique name*.
# 3. Wait for deployment to complete. Then go to your cognitive services resource, and on the **Overview** page, click the link to manage the keys for the service. You will need the endpoint and keys to connect to your cognitive services resource from client applications.
#
# ### Get the Key and Endpoint for your Cognitive Services resource
#
# To use your cognitive services resource, client applications need its endpoint and authentication key:
#
# 1. In the Azure portal, on the **Keys and Endpoint** page for your cognitive service resource, copy the **Key1** for your resource and paste it in the code below, replacing **YOUR_COG_KEY**.
# 2. Copy the **endpoint** for your resource and and paste it in the code below, replacing **YOUR_COG_ENDPOINT**.
# 3. Run the code in the cell below by clicking the Run Cell <span>▷</span> button (at the top left of the cell).
# + gather={"logged": 1599693964655}
cog_key = 'YOUR_COG_KEY'
cog_endpoint = 'YOUR_COG_ENDPOINT'
print('Ready to use cognitive services at {} using key {}'.format(cog_endpoint, cog_key))
# -
# To use the Face service in your Cognitive Services resource, you'll need to install the Azure Cognitive Services Face package.
# + tags=[]
# ! pip install azure-cognitiveservices-vision-face
# -
# Now that you have a Cognitive Services resource and the SDK package installed, you can use the Face service to detect human faces in the store.
#
# Run the code cell below to see an example.
# + gather={"logged": 1599693970079}
from azure.cognitiveservices.vision.face import FaceClient
from msrest.authentication import CognitiveServicesCredentials
from python_code import faces
import os
# %matplotlib inline
# Create a face detection client.
face_client = FaceClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Open an image
image_path = os.path.join('data', 'face', 'store_cam2.jpg')
image_stream = open(image_path, "rb")
# Detect faces
detected_faces = face_client.face.detect_with_stream(image=image_stream)
# Display the faces (code in python_code/faces.py)
faces.show_faces(image_path, detected_faces)
# -
# Each detected face is assigned a unique ID, so your application can identify each individual face that was detected.
#
# Run the cell below to see the IDs for some more shopper faces.
# + gather={"logged": 1599693970447}
# Open an image
image_path = os.path.join('data', 'face', 'store_cam3.jpg')
image_stream = open(image_path, "rb")
# Detect faces
detected_faces = face_client.face.detect_with_stream(image=image_stream)
# Display the faces (code in python_code/faces.py)
faces.show_faces(image_path, detected_faces, show_id=True)
# -
# ## Analyze facial attributes
#
# Face can do much more than simply detect faces. It can also analyze facial features and expressions to suggest age and emotional state; For example, run the code below to analyze the facial attributes of a shopper.
# + gather={"logged": 1599693971321}
# Open an image
image_path = os.path.join('data', 'face', 'store_cam1.jpg')
image_stream = open(image_path, "rb")
# Detect faces and specified facial attributes
attributes = ['age', 'emotion']
detected_faces = face_client.face.detect_with_stream(image=image_stream, return_face_attributes=attributes)
# Display the faces and attributes (code in python_code/faces.py)
faces.show_face_attributes(image_path, detected_faces)
# -
# Based on the emotion scores detected for the customer in the image, the customer seems pretty happy with the shopping experience.
#
# ## Find similar faces
#
# The face IDs that are created for each detected face are used to individually identify face detections. You can use these IDs to compare a detected face to previously detected faces and find faces with similar features.
#
# For example, run the cell below to compare the shopper in one image with shoppers in another, and find a matching face.
# + gather={"logged": 1599693972555}
# Get the ID of the first face in image 1
image_1_path = os.path.join('data', 'face', 'store_cam3.jpg')
image_1_stream = open(image_1_path, "rb")
image_1_faces = face_client.face.detect_with_stream(image=image_1_stream)
face_1 = image_1_faces[0]
# Get the face IDs in a second image
image_2_path = os.path.join('data', 'face', 'store_cam2.jpg')
image_2_stream = open(image_2_path, "rb")
image_2_faces = face_client.face.detect_with_stream(image=image_2_stream)
image_2_face_ids = list(map(lambda face: face.face_id, image_2_faces))
# Find faces in image 2 that are similar to the one in image 1
similar_faces = face_client.face.find_similar(face_id=face_1.face_id, face_ids=image_2_face_ids)
# Show the face in image 1, and similar faces in image 2(code in python_code/face.py)
faces.show_similar_faces(image_1_path, face_1, image_2_path, image_2_faces, similar_faces)
# -
# ## Recognize faces
#
# So far you've seen that Face can detect faces and facial features, and can identify two faces that are similar to one another. You can take things a step further by inplementing a *facial recognition* solution in which you train Face to recognize a specific person's face. This can be useful in a variety of scenarios, such as automatically tagging photographs of friends in a social media application, or using facial recognition as part of a biometric identity verification system.
#
# To see how this works, let's suppose the Northwind Traders company wants to use facial recognition to ensure that only authorized employees in the IT department can access secure systems.
#
# We'll start by creating a *person group* to represent the authorized employees.
# + gather={"logged": 1599693973492}
group_id = 'employee_group_id'
try:
# Delete group if it already exists
face_client.person_group.delete(group_id)
except Exception as ex:
print(ex.message)
finally:
face_client.person_group.create(group_id, 'employees')
print ('Group created!')
# -
# Now that the *person group* exists, we can add a *person* for each employee we want to include in the group, and then register multiple photographs of each person so that Face can learn the distinct facial characetristics of each person. Ideally, the images should show the same person in different poses and with different facial expressions.
#
# We'll add a single employee called Wendell, and register three photographs of the employee.
# + gather={"logged": 1599693976898}
import matplotlib.pyplot as plt
from PIL import Image
import os
# %matplotlib inline
# Add a person (Wendell) to the group
wendell = face_client.person_group_person.create(group_id, 'Wendell')
# Get photo's of Wendell
folder = os.path.join('data', 'face', 'wendell')
wendell_pics = os.listdir(folder)
# Register the photos
i = 0
fig = plt.figure(figsize=(8, 8))
for pic in wendell_pics:
# Add each photo to person in person group
img_path = os.path.join(folder, pic)
img_stream = open(img_path, "rb")
face_client.person_group_person.add_face_from_stream(group_id, wendell.person_id, img_stream)
# Display each image
img = Image.open(img_path)
i +=1
a=fig.add_subplot(1,len(wendell_pics), i)
a.axis('off')
imgplot = plt.imshow(img)
plt.show()
# -
# With the person added, and photographs registered, we can now train Face to recognize each person.
# + gather={"logged": 1599693977046}
face_client.person_group.train(group_id)
print('Trained!')
# -
# Now, with the model trained, you can use it to identify recognized faces in an image.
# + gather={"logged": 1599693994820}
# Get the face IDs in a second image
image_path = os.path.join('data', 'face', 'employees.jpg')
image_stream = open(image_path, "rb")
image_faces = face_client.face.detect_with_stream(image=image_stream)
image_face_ids = list(map(lambda face: face.face_id, image_faces))
# Get recognized face names
face_names = {}
recognized_faces = face_client.face.identify(image_face_ids, group_id)
for face in recognized_faces:
person_name = face_client.person_group_person.get(group_id, face.candidates[0].person_id).name
face_names[face.face_id] = person_name
# show recognized faces
faces.show_recognized_faces(image_path, image_faces, face_names)
# -
# ## Learn More
#
# To learn more about the Face cognitive service, see the [Face documentation](https://docs.microsoft.com/azure/cognitive-services/face/)
#
| 01d - Face Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/realmistic/pysteps-basic-fin-analysis/blob/master/Part3_NLP_Sentiment_Analysis_for_Stocks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Uuue5TpY4jsq"
#
# + id="2qM0yT-b4jSk"
# + [markdown] id="vyNkVr2fklpA"
# **TO** READ "Stock News Sentiment Analysis with Python": https://towardsdatascience.com/stock-news-sentiment-analysis-with-python-193d4b4378d4
# https://finviz.com/quote.ashx?t=DAL&ty=c&ta=1&p=d
#
# Additional idea: use the specific set of articles from the website - that may be available over a longer period of time, rather than NewsAPI
#
# + [markdown] id="C7x6ECvRwpf8"
# **Article outline**
#
# 1) **Intro**: 'Why news and NLP?'
# The stock market can be influenced by the news: it can be any important thing about the company (eg new contract, new business line, strong manager hired , etc.), or financial results (quarterly and annual earnings, profits , etc.) compared with analysts expectations .
# In this article you will try to get automatically the list of news using the newsapi API, apply sentiment analysis, and compare the results with the stock/index prices (we covered the data piece for this in the prev email)
#
# 2) **News Api**: show the endpoints for get everything/ sources. Show how to get list of ids for only business English sources. Show the python wrapper and the functionality that is limited compared to the newsapi API (the idea is that you can get it with the requests library using the full functionality).
# It is not ideal though: you don't understand if the news are relevant to the stocks, you have only 1 month of history.
# Alternative to investigate:
# - Finviz https://finviz.com/quote.ashx?t=DAL&ty=c&ta=1&p=d
# - the article: https://towardsdatascience.com/stock-news-sentiment-analysis-with-python-193d4b4378d4
# You can have links of articles and their names that are highly relevant and hand-curated
#
# 3) **Sentiment analysis: introduce the library**
# - https://medium.com/analytics-vidhya/simplifying-social-media-sentiment-analysis-using-vader-in-python-f9e6ec6fc52f
# - the paper!!: http://comp.social.gatech.edu/papers/icwsm14.vader.hutto.pdf
# I like to cover not only compound, but also positive, neutral, and negative.
# We can also check what is more correlated with the graph movements.
#
# Show scores for several article names.
#
# 4)**The idea:** analyse one event with the news sentiment
#
# Get one request for BRK.B (Berkshire Hathaway Inc.) stock around 9th July
# - check the links list, descriptions, and short contents. Are the news similar, are they relevant to financials?
# We selected 9th July and large capitalisation stock brk.b because there are many news about it and there was a jump in stock’s price. We will try to find the evidence why did that happen In the news coverage.
#
# What does it show on the news about brk.b stock. Can it be debugged ? (Show which words Influenced on positive/negative sentiment)
# Compare the sentiment score vs. Stock rise and speculate what articles may influenced its growth
#
# 5) **Scale the analysis**: check daily news (about stocks) sentiment vs. growth of S&P500 index.
#
# - 5.1) option 1 : all news from the en newspapers about stocks (we get max number =100 news via python library ).
# - 5.2) option 2: we limit the sources only to 7 business related sources (it is 20-30 news about Stocks).
#
# We compare both results vs. Index movement and find that business news sentiment is visually more correlated with the move .
#
# + id="uU0n6aUcfpi2"
# GLOBAL PARAMS - for one event
SHOW_ALL_ARTICLES = True
# SHOW_ALL_ARTICLES = False
#the day of the event
# STARTD = '22-Jun-2020'
# STARTD = '19-Jun-2020'
STARTD = '9-Jul-2020'
#the search phrase for News API
# KEYWRD = 'Inovio Pharmaceuticals stock'
# KEYWRD = 'Apple Inc.'
KEYWRD = 'Berkshire Hathaway Inc.'
#tiker
# tkr = "INO"
# tkr = 'AAPL'
tkr = 'BRK-B'
# tkr = 'LMND'
# + [markdown] id="1nX5RbLaf--o"
# # 0) Imports
# + id="kT-G2EaJf8oy" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="ce59e096-5bbf-47f4-bf8f-b809fb123965"
import sys
import nltk
from nltk.sentiment.vader import SentimentIntensityAnalyzer
### Uncomment it when the script runs for the first time
nltk.download('vader_lexicon')
sia = SentimentIntensityAnalyzer()
# + id="S0HtUha3gBES" colab={"base_uri": "https://localhost:8080/", "height": 190} outputId="ba43b4ad-7eec-4d5d-ea41-681b70254401"
# !pip install newsapi-python
# + id="0TmesyHxgEQg" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="7b1aea09-3a3b-4082-a9d4-1bcf2ffb0cd1"
# !pip install yfinance
# + id="K-RxcldxgFZw"
from newsapi import NewsApiClient
#from newsapi.newsapi_client import NewsApiClient
from datetime import date, timedelta, datetime
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import yfinance as yf
# + id="0BBLz5CWW1IY"
# Show full output in Colab
# https://stackoverflow.com/questions/54692405/output-truncation-in-google-colab
pd.set_option('display.max_colwidth',1000)
# + [markdown] id="jhs5dhAYgyZV"
# # 1) News API BY KEYWORD - download some news on a search keyword for a specific date, sorted on relevancy for language =en, first 100 articles
# + [markdown] id="ANJCxVA9o_0G"
# ## 1.0) Define functions calling news api
# + [markdown] id="iQQl-ExUgNJw"
# We use this function to call one end point to filter proper sources
# + id="2hqZ9RfXbWoB"
# https://newsapi.org/docs/endpoints/sources
# https://github.com/mattlisiv/newsapi-python
def get_sources(category = None):
newsapi = NewsApiClient(api_key='<KEY>')
sources = newsapi.get_sources()
if category is not None:
rez = [source['id'] for source in sources['sources'] if source['category'] == category and source['language'] == 'en']
else:
rez = [source['id'] for source in sources['sources'] if source['language'] == 'en']
return rez
# + id="ChnQafKHgYE_" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2a0d89f2-261a-4968-8508-650c321bb899"
# 81 en sources
len(get_sources())
# + id="v5ak7ityc0eL" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="23a1486b-18c6-46ed-bec1-15b4559ab231"
# 7 business sources
get_sources('business')
# + id="ZZ9T9XP7tYk9" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c036592d-f52d-4790-c25b-02a9625a4d9b"
type(date(2020,1,1))
# + id="eu3uDCJhg_Nf"
# https://github.com/mattlisiv/newsapi-python/blob/master/newsapi/newsapi_client.py
# https://newsapi.org/docs/endpoints/everything
def get_articles_sentiments(keywrd, startd, sources_list = None, show_all_articles = False):
newsapi = NewsApiClient(api_key='<KEY>')
if type(startd) == str:
my_date = datetime.strptime(startd,'%d-%b-%Y')
else:
my_date = startd
# business_en_sources = get_sources('business','en')
if sources_list:
articles = newsapi.get_everything(q=keywrd,
from_param = my_date.isoformat(),
to = (my_date + timedelta(days = 1)).isoformat(),
language="en",
sources = ",".join(sources_list),
sort_by="relevancy",
page_size = 100)
else:
articles = newsapi.get_everything(q=keywrd,
from_param = my_date.isoformat(),
to = (my_date + timedelta(days = 1)).isoformat(),
language="en",
sort_by="relevancy",
page_size = 100)
article_content = ''
date_sentiments = {}
date_sentiments_list = []
seen = set()
for article in articles['articles']:
if str(article['title']) in seen:
continue
else:
seen.add(str(article['title']))
article_content = str(article['title']) + '. ' + str(article['description'])
sentiment = sia.polarity_scores(article_content)['compound']
date_sentiments.setdefault(my_date, []).append(sentiment)
date_sentiments_list.append((sentiment, article['url'],article['title'],article['description']))
date_sentiments_l = sorted(date_sentiments_list, key=lambda tup: tup[0],reverse=True)
sent_list = list(date_sentiments.values())[0]
return pd.DataFrame(date_sentiments_list, columns=['Sentiment','URL','Title','Description'])
# + [markdown] id="g7lXVsGvpFSA"
# ## 1.1) Test 2 versions for all news with q='stock': for ALL en sources
# + id="2aRvBEpMg_Hu" colab={"base_uri": "https://localhost:8080/"} outputId="89249098-d840-41e9-eafc-e0a2d8787a3c"
# Easy version when we don't filter the business source -- seems to be relevant though, but the description
# Get all sources in en
return_articles = get_articles_sentiments(keywrd= 'stock' ,startd = '21-Jul-2020',sources_list = None, show_all_articles= True)
return_articles.Sentiment.hist(bins=30,grid=False)
print(return_articles.Sentiment.mean())
print(return_articles.Sentiment.count())
print(return_articles.Description)
# + [markdown] id="Sw2_R7yFWP9z"
# Result: you can see 100 articles, with a lot of neutral sentiment,
# and it is skewed towards very positive
# + id="Pw6_ZeLlWdJ7" colab={"base_uri": "https://localhost:8080/"} outputId="f34f30b4-d003-4e21-c490-beb2d6de7dc9"
return_articles.sort_values(by='Sentiment', ascending=True)[['Sentiment','URL']].head(2)
# + id="pIo_Dn7_kWt6" colab={"base_uri": "https://localhost:8080/"} outputId="0bed3f76-fed9-4614-b6a0-0b4e5e4eab8d"
return_articles.sort_values(by='Sentiment', ascending=True)[['Sentiment','URL']].tail(2)
# + [markdown] id="FK9zsvzZXDqP"
# Top negative: https://www.reuters.com/article/india-nepal-palmoil-idUSL3N2ES1Y3
# Nepal stops buying (New Dehli Suspended 39 oil import...)
# + id="yF18iqTljXGl" colab={"base_uri": "https://localhost:8080/"} outputId="1fac94cc-b172-49ab-b5fc-e6c676f7598a"
return_articles.sort_values(by='Sentiment', ascending=True)[['Sentiment','URL']].head(2)
# + [markdown] id="-2pcsUKMkdwT"
# From the article above: "TOKYO, July 21 (Reuters) - Japanese stocks rose on Tuesday as signs of progress in developing a COVID-19 vaccine boosted investor confidence in the outlook for future economic growth."
# + [markdown] id="nhLhbO86o4d4"
# ## 1.2 Same as 1.1, but try only business sources : different articles covered?
# + id="S7R5z8OTZJUL" colab={"base_uri": "https://localhost:8080/"} outputId="4ed57fc5-ee21-4bee-b7e6-e6d50b71667d"
# Look at only business sources in en
sources = get_sources('business')
return_articles = get_articles_sentiments('stock','21-Jul-2020',sources_list=sources, show_all_articles=True)
return_articles.Sentiment.hist(bins=30,grid=False)
print(return_articles.Sentiment.mean())
print(return_articles.Sentiment.count())
print(return_articles.Description)
# + id="cLAH3sdRjbvN" colab={"base_uri": "https://localhost:8080/"} outputId="4e0decee-3fde-4711-d391-89105ff607d2"
return_articles = get_articles_sentiments('stock','20-Jul-2020', sources_list=sources, show_all_articles=True)
return_articles.Sentiment.hist(bins=30,grid=False)
print(return_articles.Sentiment.mean())
print(return_articles.Sentiment.count())
# + id="TDHS_N1DjjGu" colab={"base_uri": "https://localhost:8080/"} outputId="cf254ddd-73f1-48e3-b432-c5a4905126e6"
return_articles = get_articles_sentiments('stock','19-Jul-2020',show_all_articles=True)
return_articles.Sentiment.hist(bins=30,grid=False)
return_articles.Sentiment.mean()
# + [markdown] id="VlqW6NdwmhPs"
# # 2) Get sentiment for 1 month: q= 'stocks', 'en' language, with and without business category filter
# + id="ssp30DuOnTiH" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="6983a294-3523-45db-d9e9-1640b18c35f9"
from datetime import date
end_date = date.today()
start_date = date(year=end_date.year, month=end_date.month-1, day=end_date.day)
print('Start day = ', start_date)
print('End day = ', end_date)
current_day = start_date
business_sources = get_sources('business')
sentiment_all_score = []
sentiment_business_score = []
dates=[]
while current_day <= end_date:
dates.append(current_day)
sentiments_all = get_articles_sentiments(keywrd= 'stock' ,
# current_day.strftime('%b-%d-%Y'),
startd = current_day,
sources_list = None,
show_all_articles= True)
sentiment_all_score.append(sentiments_all.mean())
sentiments_business = get_articles_sentiments(keywrd= 'stock' ,
# startd = current_day.strftime('%b-%d-%Y'),
startd = current_day,
sources_list = business_sources,
show_all_articles= True)
sentiment_business_score.append(sentiments_business.mean())
current_day = current_day + timedelta(days=1)
# + id="2E05ed93uTEl"
sentiments = pd.DataFrame([dates,np.array(sentiment_all_score),np.array(sentiment_business_score)]).transpose()
# + id="-OUa7_1Sw9NN"
sentiments.columns=['Date','All_sources_sentiment','Business_sources_sentiment']
# + id="_re8cZ9hznlt"
sentiments['Date'] = pd.to_datetime(sentiments['Date'])
# + id="KYPEZowmz3b0"
sentiments['All_sources_sentiment'] = sentiments['All_sources_sentiment'].astype(float)
sentiments['Business_sources_sentiment'] = sentiments['Business_sources_sentiment'].astype(float)
# + id="SP_FINHQ0eBy" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="440c4b6f-aaa9-4228-cb24-6e46166b2b3d"
sentiments.info()
# + id="iuadYPYXyB09"
sentiments.set_index("Date", inplace=True)
# + id="C3mM9qKWyG0e" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="af02dbfd-69d4-4eaa-9964-a77b4f24f23a"
sentiments.head()
# + [markdown] id="yBPTCqNlXz9R"
# # 3) S&P 500 and Dow Jones index - reuse from the prev article
# + id="U2gyuhzYmXIk" colab={"base_uri": "https://localhost:8080/", "height": 285} outputId="8c2ff362-81c5-435c-8199-b3bf20ee52d3"
# https://pydata.github.io/pandas-datareader/remote_data.html#stooq-index-data
# FRED: Federal Reserve Economic Data // fred,stlouisfed.org
import pandas_datareader.data as pdr
# from datetime import datetime
from datetime import date
end = date.today()
# start = datetime(2020,1,1)
start = datetime(year=end.year, month=end.month-1, day=end.day)
# end = datetime(2020,6,29)
print(f'Period 1 year until today: {start} to {end} ')
# + id="P6qnzZH2YT4X" colab={"base_uri": "https://localhost:8080/", "height": 180} outputId="66264a24-3d49-4ad6-e9dc-7bcc34297449"
spx_index = pdr.get_data_stooq('^SPX', start, end)
dji_index = pdr.get_data_stooq('^DJI',start,end)
# + id="PtVdXXiLwCct"
spx_index.index
# + id="bhO5iaDm5m0i"
spx_index['Close'].plot(title='1 month price history for index S&P500 Index')
# + id="BFWagLvY5k0K"
dji_index['Close'].plot(title='1 month price history for index Dow Jones Index')
# + id="qmQCbeAXmEBY"
sentiments_vs_snp = sentiments.join(spx_index['Close']).dropna()
# spx_index['Close'].join(sentiments).head()
# plot(title='1 month price history for index S&P500')
# + id="segc6f012iRQ"
sentiments_vs_snp.rename(columns={'Close':'s&p500_close'}, inplace=True)
# + id="aJ_0Zim61OXi"
sentiments_vs_snp.head()
# + id="OTR-qUWv3D_4"
import matplotlib.pyplot as plt
import seaborn as sns
# https://stackoverflow.com/questions/31594549/how-do-i-change-the-figure-size-for-a-seaborn-plot
# https://stackoverflow.com/questions/47591650/second-y-axis-time-series-seaborn
sns.set(rc={'figure.figsize':(13.0,8.0)})
ax=sns.lineplot(data=sentiments_vs_snp['s&p500_close'], color="b",label='S&P500 Close price')
ax2 = plt.twinx()
sns.lineplot(data=sentiments_vs_snp["All_sources_sentiment"], color="g", ax=ax2, label='All sources sentiment')
# + id="MbmR9BIB5BEf"
# BUSINESS SENTIMENT LOOKS CLOSER!
sns.set(rc={'figure.figsize':(13.0,8.0)})
ax=sns.lineplot(data=sentiments_vs_snp['s&p500_close'], color="b", label='S&P500 Close price')
ax2 = plt.twinx()
sns.lineplot(data=sentiments_vs_snp["Business_sources_sentiment"], color="g", ax=ax2, label='Business_sources_sentiment')
# + [markdown] id="QMzciBR0zFiV"
# # 4.1) Individual cases - Shopify
# + id="7DlEZKRpzpMi" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="1a393290-541d-4dba-c1d8-a9ec95f992ea"
business_sources
# + id="vtoeYpjVzKE0" colab={"base_uri": "https://localhost:8080/", "height": 197} outputId="b69d7916-7b9b-46a5-d990-195abd5d7d4c"
startd='29-Jul-2020'
shopify_stock = get_articles_sentiments("Shopify", startd, sources_list = business_sources, show_all_articles = True)
# + id="9NST69Cvztrn"
shopify_stock
# + id="kV2oQeBkz0hS"
shopify_stock_all = get_articles_sentiments("Shopify", startd, sources_list = None, show_all_articles = True)
# + id="vn-ZMesO0EcY"
shopify_stock_all.Sentiment.mean()
# + id="PU4xDSW3z5TB"
shopify_stock_all
# + id="lTGEpickz73C"
# Previous day
startd='28-Jul-2020'
shopify_stock_all_prevDay = get_articles_sentiments("Shopify", startd, sources_list = None, show_all_articles = True)
# + id="0hnGG3LS0ZMZ"
shopify_stock_all_prevDay.Sentiment.mean()
# + id="C34S5qdO0cFn"
shopify_stock_all_prevDay
# + id="p3RUwVbB0eV3"
from datetime import date
end_date = date.today()
start_date = date(year=end_date.year, month=end_date.month-1, day=end_date.day)
print('Start day = ', start_date)
print('End day = ', end_date)
current_day = start_date
sentiment_all_score = []
dates=[]
while current_day <= end_date:
dates.append(current_day)
sentiments_all = get_articles_sentiments(keywrd= 'Shopify' ,
# current_day.strftime('%b-%d-%Y'),
startd = current_day,
sources_list = None,
show_all_articles= True)
sentiment_all_score.append(sentiments_all.mean())
current_day = current_day + timedelta(days=1)
# + id="kCMtlfLF1NLm"
sentiments_shopify = pd.DataFrame([dates,np.array(sentiment_all_score)]).transpose()
# + id="uBNDAjJK193b"
sentiments_shopify.columns
# + id="-QsWt32M1joA"
# sentiments_shopify.columns=['Date','All_sources_sentiment']
# sentiments_shopify['Date'] = pd.to_datetime(sentiments_shopify['Date'])
# sentiments_shopify['All_sources_sentiment'] = sentiments_shopify['All_sources_sentiment'].astype(float)
sentiments_shopify.set_index("Date", inplace=True)
# + id="_Khu7uy217kU"
sentiments_shopify.plot()
# + [markdown] id="7qJ7UY1Z9SWS"
# # 4.2) GAFA stock - 30 july earnings call :get news and sentiment for Google, Amazon, Facebook, Apple
| colab_notebooks/Part3_NLP_Sentiment_Analysis_for_Stocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="2HLGI04NAnoD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610174403034, "user_tz": -540, "elapsed": 8444, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="4a015557-2da4-4c99-dd34-1a42c83606a1" language="bash"
#
# # データのダウンロード
#
# mkdir dataset
# curl -Ss https://www.rondhuit.com/download/livedoor-news-data.tar.gz > dataset/dataset.tar.gz
# cd dataset
# tar -xvf dataset.tar.gz
# rm dataset.tar.gz
# cd ../
#
#
# # ツールのインストール
#
# pip install tensorflow_text
# + id="6JJL6h8BA5d0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610174405138, "user_tz": -540, "elapsed": 10524, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="f47efdaf-5e21-4509-c24b-6db0dd4ead4b"
'''
-----------------------------------------------------
データの前処理(今回は単語分割が不要です)
-----------------------------------------------------
'''
# 1. XMLからのテキスト抽出
import glob
import xml.etree.ElementTree as ET
def get_data(file_name, target):
data = list()
tree = ET.parse(file_name)
for doc in tree.getroot():
for element in doc:
if element.attrib["name"] == target:
data.append(element.text)
return data
titles, labels = list(), list()
for file_name in sorted(glob.glob("dataset/*.xml")):
titles.extend(get_data(file_name, target="title"))
labels.extend(get_data(file_name, target="cat"))
# 2. 訓練用/検証用/評価用に分割
import numpy as np
np.random.seed(seed=42)
def shuffle(list1, list2):
tmp = list(zip(list1, list2))
np.random.shuffle(tmp)
list1, list2 = zip(*tmp)
return list(list1), list(list2)
texts, labels = shuffle(titles, labels)
texts_train, labels_train = texts[:5000], labels[:5000]
texts_dev, labels_dev = texts[5000:6000], labels[5000:6000]
texts_test, labels_test = texts[6000:7000], labels[6000:7000]
# タイトルとカテゴリの確認
print("カテゴリ: %s" % labels_train[0])
print("タイトル: %s\n" % texts_train[0])
print("カテゴリ: %s" % labels_dev[0])
print("タイトル: %s\n" % texts_dev[0])
print("カテゴリ: %s" % labels_test[0])
print("タイトル: %s\n" % texts_test[0])
# + id="Nam4G0F8BkHN" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610174907459, "user_tz": -540, "elapsed": 512840, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="ed876fb2-ad4a-425e-c6cb-ab640d4a0204"
'''
-----------------------------------------------------
Universal Sentence Encoder
-----------------------------------------------------
'''
import tensorflow_hub as hub
from tensorflow_text import SentencepieceTokenizer
use = hub.load("https://tfhub.dev/google/universal-sentence-encoder-multilingual-large/3")
X_train = use(texts_train)
X_dev = use(texts_dev)
X_test = use(texts_test)
print(len(X_train), len(X_train[0]), X_train[0])
# + id="LwzfoMCgDHlZ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610174913952, "user_tz": -540, "elapsed": 519328, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="1abfde22-0596-4ea7-cbb1-f7c15d5ad8b6"
'''
-----------------------------------------------------
分類器の訓練
-----------------------------------------------------
'''
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# ラベルをIDに変換
label2id = dict()
for label in sorted(set(labels)):
label2id[label] = len(label2id)
y_train = [label2id[label] for label in labels_train]
y_dev = [label2id[label] for label in labels_dev]
y_test = [label2id[label] for label in labels_test]
# 分類器の訓練
best_c, best_score = 0, 0
for c in [0.1, 1, 10]:
classifier = LogisticRegression(C=c, max_iter=1000)
classifier.fit(X_train, y_train)
dev_acc = accuracy_score(y_dev, classifier.predict(X_dev))
if best_score < dev_acc:
best_score = dev_acc
best_c = c
print("Dev accuracy = %1.3f\tC = %s" % (dev_acc, str(c)))
print("Best parameter: C = %s" % str(best_c))
# + id="fFeUCoOoDKW5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610174917643, "user_tz": -540, "elapsed": 523014, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "03431199727064994008"}} outputId="2e039a0b-71f6-4371-9b71-79e1a09c1a2a"
'''
-----------------------------------------------------
評価
-----------------------------------------------------
'''
classifier = LogisticRegression(C=best_c, max_iter=1000)
classifier.fit(X_train, y_train)
test_acc = accuracy_score(y_test, classifier.predict(X_test))
print("Test accuracy = %1.3f" % test_acc)
| NLP_1/NLP3-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# I love reading The Onion. The brilliant snark of their universally caustic takes on politics and everyday life make me feel less alone in reacting to the utter bizarreness of modern life.
#
# It is a reflection of the weirdness of our reality that Onion headlines so frequently get mistaken for real ones. There are plenty of stories about celebrities, politicians, and other public figures getting fooled by and inciting outrage over stories that in hindsight seem like they would be too obviously absurd to be taken seriously. But are they? Could an algorithm tell the difference?
#
# In this project, I'll be tackling just that question.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Obtain
# First, we need a source of these headlines. As a seed, I'll be using a collection of headlines from The Onion and the Huffington Post. These were collected for sarcasm classification. I have some qualms with the appropriateness of this data for that purpose -- it is not entirely clear to me or I think to anyone that sarcasm and satire/parody are the same thing. Perhaps there is some intersection, but the line is fuzzy. Still, for *our* purposes, this is a good dataset.
sarcasm = pd.read_json('../data/raw/Sarcasm_Headlines_Dataset.json', lines=True)
sarcasm.head()
sarcasm.info()
sarcasm['is_sarcastic'].value_counts()
# This is promising. We have rows with the article link and lowercased headline, and whether a headline is 'sarcastic' or not. I don't believe that sarcastic is a good way to characterize this, so I'll just alter that for my own happiness.
sarcasm.columns = ['article_link', 'headline', 'is_parody']
sarcasm.head()
# I know from some previous exploration that this dataset includes some articles that are linked through the Huffington Post, but not from the Huffington Post. Some of these might be parody articles themselves, and so might be miscoded. Let's make sure.
sarcasm.loc[sarcasm['article_link'].str.contains('comhttp')]
len('https://www.huffingtonpost.com')
sarcasm.loc[sarcasm['article_link'].str.contains('comhttp'), 'article_link'] = sarcasm.loc[sarcasm['article_link'].str.contains('comhttp'), 'article_link'].str[30:]
sarcasm.loc[(sarcasm['article_link'].str.contains('onion')) & (sarcasm['is_parody'] == 0)]
sarcasm.iloc[19948, 2] = 1
parody = sarcasm[sarcasm['is_parody'] == 1]
type(parody)
parody.to_csv('../data/processed/parody.csv')
# # Word2Vec
from gensim.models import Word2Vec
from keras.preprocessing import text, sequence
from sklearn.model_selection import train_test_split
y = sarcasm['is_parody'].values
y
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation
from keras.layers import Bidirectional, GlobalMaxPool1D
from keras.models import Model
from keras import initializers, regularizers, constraints, optimizers, layers
X_train, X_test, y_train, y_test = train_test_split(sarcasm['headline'], y, test_size=.20)
type(X_train)
tokenizer = Tokenizer(num_words=20000)
tokenizer.fit_on_texts(list(X_train))
list_tokenized_train = tokenizer.texts_to_sequences(X_train)
list_tokenized_test = tokenizer.texts_to_sequences(X_test)
total_word_counts = [len(headline) for headline in list_tokenized_train]
fig, ax = plt.subplots(figsize=(9,9))
ax.hist(total_word_counts, bins = 40)
max(total_word_counts)
counter = pd.Series(total_word_counts)
counter.value_counts()
maxlen = 18
X_t = pad_sequences(list_tokenized_train, maxlen=maxlen)
X_te = pad_sequences(list_tokenized_test, maxlen=maxlen)
X_t.shape
tokens = tokenizer.sequences_to_texts(list_tokenized_train) + tokenizer.sequences_to_texts(list_tokenized_test)
tokens = [t.split(' ') for t in tokens]
tokens[:1]
# +
model = Word2Vec(tokens, size=100, window=5, min_count=1, workers=4)
model.train(tokens, total_examples=model.corpus_count, epochs=10)
# -
embeddings = model.wv
keras_weights = embeddings.get_keras_embedding()
embeddings.most_similar(positive=['obama', 'policy'])
from gensim.models import Word2Vec
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense, Input, LSTM, Embedding, Dropout, Activation, Conv1D, MaxPooling1D, GlobalMaxPooling1D
from keras.layers import Bidirectional, GlobalMaxPool1D
from keras.models import Model, Sequential
from keras import initializers, regularizers, constraints, optimizers, layers
model = Sequential()
model.add(keras_weights)
model.add(Conv1D(64, 3, activation='relu', padding='same'))
model.add(MaxPooling1D())
model.add(Conv1D(64, 3, activation='relu', padding='same'))
model.add(GlobalMaxPooling1D())
model.add(Dropout(0.5))
model.add(Dense(32, activation='relu', kernel_regularizer=regularizers.l2(1e-4)))
model.add(Dense(1, activation='sigmoid'))
adam = optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
model.compile(loss='binary_crossentropy', optimizer=adam, metrics=['accuracy'])
model.summary()
early_stopping = EarlyStopping(monitor='val_loss', min_delta=0.01, patience=4, verbose=1)
callbacks_list = [early_stopping]
batch_size = 256
num_epochs = 30
hist = model.fit(X_t, y_train, batch_size=batch_size, epochs=num_epochs, validation_split=0.1, shuffle=True, verbose=2)
y_pred = model.predict(X_te)
from sklearn.metrics import accuracy_score
def rounder(array):
new = []
for i in array:
if i >= .5:
new.append(1)
else:
new.append(0)
return np.array(new)
y_pred_bin = rounder(y_pred)
accuracy_score(y_test, y_pred_bin)
| notebooks/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# language: python
# name: python395jvsc74a57bd031f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# ---
import re
import pandas as pd
# +
path = '../data/sample_coco.txt'
lines = []
with open(path) as f:
lines = f.readlines()
# -
lines[:5]
class my_dictionary(dict):
def __init__(self):
self = dict()
def add(self, key, value):
self[key] = value
def datarestructure(columns,dataList):
ls = []
dataDict = my_dictionary()
for i in dataList:
a = i.replace(',\n','')
val = re.findall('[0-9]+', a)
for cnt,dat in zip(columns,val):
dataDict.add(cnt,dat)
ls.append(dataDict.copy())
return ls
columns = ['id', 'height', 'width', 'x', 'y', 'bbox_width', 'bbox_height']
data = pd.DataFrame(datarestructure(columns,dataList=lines))
data = data.astype(dtype=np.int64)
data.to_csv('./data/coco.csv',index=False)
data.head(10)
import numpy as np
from matplotlib import pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.cluster import KMeans
data['n_height'] = [x/x for x in data.height]
data['n_width'] = [x/x for x in data.width]
data['n_x'] = [x/y for x,y in zip(data.x,data.width)]
data['n_y'] = [x/y for x,y in zip(data.y,data.height)]
data['n_bbox_width'] = [x/y for x,y in zip(data.bbox_width,data.width)]
data['n_bbox_height'] = [x/y for x,y in zip(data.bbox_height,data.height)]
data.head()
plt.scatter(data['n_bbox_width'],data['n_bbox_height'])
X = data[['n_bbox_width','n_bbox_height']]
# +
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters=i, init='k-means++', max_iter=300, n_init=10, random_state=0)
kmeans.fit(X)
wcss.append(kmeans.inertia_)
plt.plot(range(1, 11), wcss)
plt.title('Elbow Method')
plt.xlabel('Number of clusters')
plt.ylabel('WCSS')
plt.show()
# -
kmeans = KMeans(n_clusters=6, init='k-means++', max_iter=300, n_init=10, random_state=0)
pred_y = kmeans.fit_predict(X)
plt.scatter(X['n_bbox_width'], X['n_bbox_height'])
plt.scatter(kmeans.cluster_centers_[:, 0], kmeans.cluster_centers_[:, 1], s=300, c='red')
plt.show()
X['n_bbox_width']
path
X.to_csv('../data/kmeans.csv',index=False)
| assignment_10/part_b/experiments/DataLoader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Molecular Determinants
# Instead of focusing on the crude raw data use bootstrapping to emphasize the real differences between increasing/decreasing and emergent. Given that drug perturbation range from a broad range of perturbation e.g. when looking at the feature chemical similarity almost the whole spectrum of similarities is covered but by using bootstrap one can focus on the mean differences and the variation of the mean.
#
# 1.) Load all features
# 2.) perform bootstrap analysis
# 3.) save results (+ plots)
# Load all important python modules
import numpy as np
from matplotlib import pylab as plt
import scipy.stats as stats
from scipy.stats import mannwhitneyu as mu
import seaborn as sns
import os
from math import pi
import math
from sympy import Symbol, solve, sqrt
import networkx as nx
# ### 1. Load features
# +
#Load features per drug pair e.g. if two drugs share a common transporter, mean PPI distance between their drug targets etc.
fp = open('../data/Molecular_Determinants/DrugPair_Feature_Overview.csv','r')
features = fp.readline().strip().split(',')[4:]
print ('Number of features: %d' %len(features))
#Define interaction types as well as colors for the final plots (uniform with previous color coding)
interactionTypes = ['NoInteraction','Interaction','Increasing','Decreasing','Emergent']
interaction_colors = {'Increasing':'#ACD900','Decreasing':'#F70020','Emergent':'#0096FF','Interaction':'#F8B301','NoInteraction':'grey'}
#Create a dictionary where the individual results per feature can be split into the 5 types of interactions e.g. ChemicalSimilarity: {Increasing:[], Decreasing: [] ... }
dic_feature_results = {}
for f in features:
dic_feature_results[f] = {}
for iT in interactionTypes:
dic_feature_results[f][iT] = []
#Go through the results of the drug pair feature file
for line in fp:
tmp = line.strip().split(',')
interactionType = tmp[3]
#add the results to the corresponding list
for f,i in zip(features, range(4,len(tmp))):
val = tmp[i]
#if the val is 'nan' ignore this row
if val != 'nan':
val = float(val)
#if interaction type == None, then NoInteraction
if interactionType == 'None':
dic_feature_results[f]['NoInteraction'].append(val)
#Else split into one of the posible other interaction types, keep only pure row e.g. only increasing/decreasing
else:
if interactionType == 'Increasing' or interactionType == 'Increasing;Increasing':
dic_feature_results[f]['Increasing'].append(val)
dic_feature_results[f]['Interaction'].append(val)
if interactionType == 'Decreasing' or interactionType == 'Decreasing;Decreasing':
dic_feature_results[f]['Decreasing'].append(val)
dic_feature_results[f]['Interaction'].append(val)
if interactionType == 'Emergent':
dic_feature_results[f]['Emergent'].append(val)
dic_feature_results[f]['Interaction'].append(val)
print ('Done loading data')
# -
# ### 2. Perform Statistical Analysis
# #### 2.1 Define functions for calculating bootstrapping and effect size
def bootstrapping(data, number_iterations=10000, bootstrap_sample_size = None):
'''
Function for bootstrapping
data = Data that needs to be bootsraped
number_iteration = how often should bootstrapping be perfomred
bootstrap_sample_size = sample size to draw for, if None then sample size = len(data) which is the typical procedure for bootstrapping
'''
#Define the bootstrap sample size
if bootstrap_sample_size == None:
bootstrap_sample_size = len(data)
#draw randomly from data to get an estimation of it's variation. Save both the mean per bootstrap run as well as the calculated std
bootstrap_samples_means = []
bootstrap_samples_stds = []
for i in range(0,number_iterations):
bootstrap_sample = np.random.choice(data,bootstrap_sample_size,replace=True)
bootstrap_samples_means.append(np.mean(bootstrap_sample))
bootstrap_samples_stds.append(np.std(bootstrap_sample))
#return the results
return bootstrap_samples_means, bootstrap_samples_stds
# +
def cohen_d(x, y):
'''
Cohen's D is a typical meassure of effect size including the standard deviation of both samples (compared to ZScore which only uses one)
'''
#Get length of the two samples
nx = len(x)
ny = len(y)
#Define degrees of freedom
dof = nx + ny - 2
#Calculate Cohen's D and return
return (np.mean(x) - np.mean(y)) / np.sqrt(
((nx - 1) * np.std(x, ddof=1) ** 2 + (ny - 1) * np.std(y, ddof=1) ** 2) / dof)
# -
# #### 2.2 Calculate Statistics for each feature
# 1. Binary features e.g. have overlap/no overlap use Fisher Exact test
# 2. Continues features e.g. PPI distance, use Mann Whitney U test
# +
#Define a significance threshold
alpha = 0.05
#define the output file (shows the results for all features)
fp_out = open('../results/Molecular_Determinants/ResultsOverview.csv','w')
fp_out.write('Feature,InteractionType1,InteractionType2,Mean1,Mean2,FisherTest,PVal,PercentChange/OddsRatio,CohenD,BootstrapSign\n')
#Go thorugh each feature
for f in features:
print f
#check if all values of the given features are either 1 or 0 => then use Fisher Exact test to determine significance
make_Fisher = False
if all(v == 0 or v ==1 for v in dic_feature_results[f].values()[0]):
make_Fisher = True
#Define and create the output folder for the Bootstrapping plots (if doesnt exist)
directory = os.path.dirname('../results/Molecular_Determinants/Bootstrapping/' + f + '/')
if not os.path.exists(directory):
os.makedirs(directory)
####
# CREATE a standard boxplot for the features (only rly makes sense for continues values - make for binary features still though)
bplot = sns.boxplot(data=[dic_feature_results[f]['NoInteraction'],dic_feature_results[f]['Interaction'],dic_feature_results[f]['Increasing'],dic_feature_results[f]['Decreasing'],dic_feature_results[f]['Emergent']],orient='h',showmeans = True, showfliers = False)
#Define labels and colors
interaction_types_2 = ['NoInteraction','Interaction','Increasing','Decreasing','Emergent']
interaction_colors_2 = ['grey','#F8B301','#ACD900','#F70020','#0096FF']
color_dict = dict(zip(interaction_types_2, interaction_colors_2))
for i in range(0,5):
mybox = bplot.artists[i]
mybox.set_facecolor(color_dict[interaction_types_2[i]])
#Add title and proper ticks
plt.title(f)
plt.yticks(range(0,5),['NoInteraction','NoInteraction','Increasing','Decreasing','Emergent'])
plt.ylabel('Interaction Type')
plt.tick_params(axis = 'y', which = 'major', labelsize = 5)
plt.xlabel('Amount')
plt.savefig(directory+'/Boxplot.pdf')
plt.close()
# END creating standard boxplot
#########
#####
# CREATE Bootstrap plot (histogram showing the results for the 5 interaction types)
#save the temporary bootstrap results for each interaction thype separately
bootstrap_results = {}
#Calculate bootstrap results for the 5 interaction types
for iT in interactionTypes:
#save mean and std as results
bootstrap_results[iT] = {'b_mean':[],'b_std':[]}
#get the actual data
data = dic_feature_results[f][iT]
#perform bootstrapping with standard bootstrapping rules
b_means, b_stds = bootstrapping(data,number_iterations=10000, bootstrap_sample_size=None)
#Save results
bootstrap_results[iT]['b_mean'] = b_means
bootstrap_results[iT]['b_std'] = b_stds
#Xreate a histogram
plt.hist(b_means,bins='auto', color = interaction_colors[iT], alpha=0.4)
plt.savefig(directory+'/BootstrapOVerview.pdf')
plt.close()
# END creating Bootsrap plot
#########
#####
# Comparison of mean results per interaction types (Interacting, Increasing, Decrasing, Emergent) compared to NO_INTERACTION
# Create a Histogram for NoInteraction and compare to 4 individual points (represented as lines)
plt.hist(bootstrap_results['NoInteraction']['b_mean'],bins='auto')
plt.axvline(np.mean(dic_feature_results[f]['Interaction']),color=interaction_colors['Interaction'])
plt.axvline(np.mean(dic_feature_results[f]['Increasing']),color=interaction_colors['Increasing'])
plt.axvline(np.mean(dic_feature_results[f]['Decreasing']),color=interaction_colors['Decreasing'])
plt.axvline(np.mean(dic_feature_results[f]['Emergent']),color=interaction_colors['Emergent'])
plt.savefig(directory+'/OldBootstrapPlot.pdf')
plt.close()
# END creating NoInteraction comparison plot
######
###
# COMPARE the bootstrap results between two interaction types to see if they are significantly different
# Go through all different pairs
for iT1 in interactionTypes:
for iT2 in interactionTypes:
if iT1 > iT2:
#Extract data
data1 = np.array(bootstrap_results[iT1]['b_mean'])
data2 = np.array(bootstrap_results[iT2]['b_mean'])
# Create a new distribution by substracting all the bootstrap results from each other
# If 0 is completely outside this distribution (outside 95 percentile) then significant difference
bootstrap_mean_diff = list(data1 - data2)
CI = (np.percentile(bootstrap_mean_diff,2.5), np.percentile(bootstrap_mean_diff,97.5))
bootstrapSign = (0 > CI[0] and 0 > CI[1]) or (0 < CI[0] and 0 < CI[1])
# Calculate corresponding Cohen's D
c_d = cohen_d(data1,data2)
# Calculate if two groups are significant different according to Fisher test (if binary data)
if make_Fisher:
group1_Overlap = sum(dic_feature_results[f][iT1])
group1_NonOverlap = len(dic_feature_results[f][iT1]) - group1_Overlap
group2_Overlap = sum(dic_feature_results[f][iT2])
group2_NonOverlap = len(dic_feature_results[f][iT2]) - group2_Overlap
effect, pval = stats.fisher_exact([[group1_Overlap, group1_NonOverlap], [group2_Overlap, group2_NonOverlap]])
# Else calulate according to Mann Whitney U
else:
pval = mu(dic_feature_results[f][iT1],dic_feature_results[f][iT2])[1]
effect = (np.mean(data1) - np.mean(data2))/np.mean(data2) * 100
# Create the difference bootstrap plot, with percentile and zero as markers; Add significance calculation to the title
plt.hist(bootstrap_mean_diff,bins='auto', color='grey')
plt.title(iT1 +'_' +iT2+': %.2f' %pval)
plt.axvline(CI[0])
plt.axvline(CI[1])
plt.axvline(0,c='red',ls='--')
#plt.show()
plt.savefig(directory+'/Bootstrap_'+iT1 +'_' +iT2+'.pdf')
plt.close()
# Save the results to the overview file
fp_out.write(f+','+iT1+','+iT2+','+str(np.mean(dic_feature_results[f][iT1]))+','+str(np.mean(dic_feature_results[f][iT2]))+','+str(make_Fisher)+','+str(pval)+','+str(effect)+','+str(c_d)+','+str(bootstrapSign)+'\n')
fp_out.close()
# -
# ### 3. Radar plots
# Additionally also create per feature radar plots, that are capable showing the disticnt moleular properties per interaction type
# #### 3.1 Define functions for the creation of radar plots
# +
def radiusAngle_ToCoordinates(r, phi):
'''
Transform the radius and angle into x and y coordinates. Depending on the quadrant in, the sin gives either the x
or y coordinate (and cos the other). As the angle is given between zero and 2pi, depending on the quadrant
adjusted so we can create triangles
:param r: radius of the point
:param phi: angle between 0 and 2pi
:return: x and y coordinate
'''
if phi <= pi / 2:
#print 'Upper Right'
x = math.sin(phi) * r
y = math.cos(phi) * r
quadr = 'UR'
elif phi <= pi:
#print 'Below Right'
phi = phi - (pi / 2)
x = math.cos(phi) * r
y = math.sin(phi) * r * (-1)
quadr = 'BR'
elif phi <= (3 * pi) / 2:
#print 'Below Left'
phi = phi - (pi)
x = math.sin(phi) * r * (-1)
y = math.cos(phi) * r * (-1)
quadr = 'BL'
else:
#print 'Upper Left'
phi = phi - (3 * pi / 2)
x = math.cos(phi) * r * (-1)
y = math.sin(phi) * r
quadr = 'UL'
return x, y, quadr
def Find_Intersection(rc, phi1, r1, phi2, r2):
'''
Find the intersection of a line drawn between two points (given by their radius and angle) with a circle centered
around zero and a given radius
:param rc: radius of the circle
:param phi1: first angle
:param r1: first radius
:param phi2: second angle
:param r2: second radius
:return: angle of the intersection (as radius has to be rc)
'''
# transform radius and angle into x and y coordinates (using sin/cos)
x1, y1, quadr1 = radiusAngle_ToCoordinates(r1, phi1)
x2, y2, quadr2 = radiusAngle_ToCoordinates(r2, phi2)
# Create Function to plot
# factor = (y2-y1)/(x2-x1)
# print 'y = %.3fx + %.3f' %(factor,-(factor*x1) + y1)
# Define the symbol to solve for
x = Symbol("x")
# Intersect the line with the circle
x_intersect = solve(((y2 - y1) * (x - x1)) / (x2 - x1) + y1 - sqrt(
rc * rc - x * x)) # take positive values of circle results (minus times plus = minus) // gives you all result for the positive circle (> 0)
# Go thre all POSITIVE VALUES (check if one of the angles is between the two original angles; intersection net to be between)
for x in x_intersect:
# Get the corresponding y coordinate
y_intersect = ((y2 - y1) * (x - x1)) / (x2 - x1) + y1
# calculate Phi
result_phi = math.acos(abs(x) / rc)
# Again adjust to quadrant
if x >= 0 and y_intersect >= 0:
#print 'Upper Right'
result = (pi / 2 - result_phi)
elif x >= 0 and y_intersect <= 0:
#print 'Lower Right'
result = (pi / 2 + result_phi)
elif x <= 0 and y_intersect <= 0:
#print 'Lower Left'
result = (((3 * pi) / 2) - result_phi)
else:
#print 'Upper Left'
result = (((3 * pi) / 2) + result_phi)
# if proper angle found return
if result > phi1 and result < phi2:
return result
# Define the symbol to solve for
x = Symbol("x")
# Intersect the line with the circle
x_intersect = solve(((y2 - y1) * (x - x1)) / (x2 - x1) + y1 + sqrt(
rc * rc - x * x)) # take negative values of circle results (minus times plus = minus)// gives you all result for the negative circle (< 0)
# Go thre all NEGATIVE VALUES (check if one of the angles is between the two original angles; intersection net to be between)
for x in x_intersect:
# Get the corresponding y coordinate
y_intersect = ((y2 - y1) * (x - x1)) / (x2 - x1) + y1
# calculate Phi
result_phi = math.acos(abs(x) / rc)
# Again adjust to quadrant
if x >= 0 and y_intersect >= 0:
#print 'Upper Right'
result = (pi / 2 - result_phi)
elif x >= 0 and y_intersect <= 0:
#print 'Lower Right'
result = (pi / 2 + result_phi)
elif x <= 0 and y_intersect <= 0:
#print 'Lower Left'
result = (((3 * pi) / 2) - result_phi)
else:
#print 'Upper Left'
result = (((3 * pi) / 2) + result_phi)
# if proper angle found return
if result > phi1 and result < phi2:
return result
return 'Nothing Found'
def my_SpiderPlot(categories, values, color, title,num='None', toNormalizeSmallest='None', toNormalizeBiggest="None"):
'''
Create a Spider Plot
:param categories: categories of the spiderplots (the individual factors)
:param values: actual values
:param color: the colorscheme (e.g. deactivating = red)
:param title: name of the spiederplot
:param num: in case of overlay (else just None for individual Spiderplots)
:return:
'''
if toNormalizeSmallest !="None":
#Normalize all values to a pre given value
nullValue = int(toNormalizeSmallest) - 3
newValues = [x + abs(toNormalizeSmallest) + 3 for x in values]
max_yticks = int(toNormalizeBiggest) + 1
else:
#Get the lowest value (e.g . -10), for ploting this will be zero; add three so the lowest value is NOT in the middle but a bit away
nullValue = int(min(values)) - 3
#Normalize all values, e.g. the -10 to zero, whereas the zero will be 10 in the plot
newValues = [x+abs(min(values))+3 for x in values]
#Define the max tick as max value plus one (for aesthetics)
max_yticks = int(max(values))+3
#get the negative ticks and positive ticks
negative_ticks = [str(x) for x in range(nullValue,0,1)]
positive_ticks = [str(x) for x in range(0,max_yticks+1,1)]
negative_ticks.extend(positive_ticks)
#print negative_ticks
#exit()
#Take only 8 tick marks
to_take = len(negative_ticks)/8
chosen_ticks = [negative_ticks[x] for x in range(0,len(negative_ticks),to_take)]
#take the normalized values to plot (i.e. the values where the -10 became the zero
values = newValues
#Find number of categories
N = len(categories)
# What will be the angle of each axis in the plot? (we divide the plot / number of variable)
# The total of 2pi (around 6.2) is divided into the amount of categories; In final plot it will be just from 0 till 2 in pi (factor 3.1415 is missing)
angles = [n / float(N) * 2 * pi for n in range(N)]
angles += angles[:1]
# Initialise the spider plot
if num != 'None':
ax = plt.subplot(1, 3, num+ 1, polar=True, )
else:
ax = plt.subplot(1, 1, 1, polar=True, )
# If you want the first axis to be on top:
ax.set_theta_offset(pi / 2)
ax.set_theta_direction(-1)
# Draw one axe per variable + add labels labels yet
#categories = [x.split('AllRandom')[0] for x in categories]
plt.xticks(angles[:-1], categories, color='grey', size=8)
# Draw ylabels
ax.set_rlabel_position(0)
#add last value, to close the circle
values.append(values[0])
#plot the line
ax.plot(angles, values, color=color, linewidth=2, linestyle='solid')
#ax.fill(angles, values, color=color, alpha=0.4)
#Go threw all the points, whenever there is a switch between a positive and a negative ZScore, the line
#intersects with the zero line, hence new color; use the find_intersection function to find probper intersection
i_was = 'Nowhere'
tmp = []
tmp_angles = []
to_save = []
prev_val = 0
prev_ang = 0
angles_to_save_cut = []
normal_angles = []
#Go thre all values and angles
for val,ang in zip(newValues,angles):
#Check if value is positive or negative
if val > abs(nullValue):
i_am = 'Positive'
else:
i_am = 'Negative'
#Check if there is a switch between positive and negative
if i_was != i_am and i_was != 'Nowhere':
#Define the radius of the circle (=y)
y = abs(nullValue)
#if the last line is between 3 quadrant and the origin (change 0.0 to 6.2831 = 2pi = full circle)
if prev_ang > 3.15 and ang == 0.0:
ang = 6.2831
#Find the actual intersection
result = Find_Intersection(y,prev_ang,prev_val,ang,val)
angles_to_save_cut.append(result)
#if more than one angle belongs to one section, before creating new tmp, add current to save
if len(tmp) >0:
to_save.append(tmp)
normal_angles.append(tmp_angles)
#start new tmp (= section of color)
tmp = [val]
tmp_angles = [ang]
#if still in same section just add angle and value
else:
tmp.append(val)
tmp_angles.append(ang)
#Remember previous location
i_was = i_am
prev_val = val
prev_ang = ang
#Final results of intersection parts (angles and values)
to_save.append(tmp)
normal_angles.append(tmp_angles)
#make a fine grained amount of angles (361 individual degrees), and close circle again
angles2 = [n / float(360) * 2 * pi for n in range(360)]
angles2 += angles2[:1]
#Define color scheme
'''
colorscheme = {'green':{0:'#acd900',1:'#a6c143',2:'#648a58',3:'#5c5e4c',4:'#acd900',5:'#a6c143',6:'#648a58',7:'#5c5e4c'},
'red': {0: '#f70020', 1: '#e66a22', 2: '#e79935', 3: '#dcb471', 4: '#f70020',5:'#e66a22',6:'#e79935',7:'#dcb471'},
'blue':{0: '#0096ff', 1: '#2bbfb8', 2: '#29a2ac', 3: '#4c7584', 4: '#0096ff', 5: '#2bbfb8',6:'#29a2ac',7:'#4c7584'},
'grey':{0:'#252525',1:'#636363',2:'#969696',3:'#cccccc',4:'#f7f7f7'}
}
'''
'''
colorscheme = {'green':{0:'#acd900',1:'#acd900',2:'#acd900',3:'#acd900',4:'#acd900',5:'#acd900',6:'#acd900',7:'#acd900'},
'red': {0: '#f70020', 1: '#f70020', 2: '#f70020', 3: '#f70020', 4: '#f70020',5:'#f70020',6:'#f70020',7:'#f70020'},
'blue':{0: '#0096ff', 1: '#0096ff', 2: '#0096ff', 3: '#0096ff', 4: '#0096ff', 5: '#0096ff',6:'#0096ff',7:'#0096ff'},
'grey':{0:'#252525',1:'#252525',2:'#252525',3:'#252525',4:'#252525'}
}
'''
colorscheme = {'green':{0:'#acd900',1:'#a6c143',2:'#acd900',3:'#a6c143',4:'#acd900',5:'#a6c143',6:'#acd900',7:'#a6c143'},
'red': {0: '#f70020', 1: '#e66a22', 2: '#f70020', 3: '#e66a22', 4: '#f70020',5:'#e66a22',6:'#f70020',7:'#e66a22'},
'blue':{0: '#0096ff', 1: '#2bbfb8', 2: '#0096ff', 3: '#2bbfb8', 4: '#0096ff', 5: '#2bbfb8',6:'#0096ff',7:'#2bbfb8'},
'grey':{0:'#252525',1:'#636363',2:'#252525',3:'#636363',4:'#252525'}
}
#If the first section is bigger than one immedieatly
nofirstcut = False
if len(to_save[0]) > 0:
angles_to_save_cut.insert(0,0)
nofirstcut = True
angles_to_save_cut += angles_to_save_cut[:1]
#fill the individual parts
for i in range(0,len(to_save)):
#save_cut[i] to savecut[i+1] define the whole area, + add all the angles between these two
to_fill_angles = [angles_to_save_cut[i]]
to_fill_Values = [abs(nullValue)]
to_fill_Values.extend(to_save[i])
to_fill_angles.extend(normal_angles[i])
to_fill_angles.append(angles_to_save_cut[i+1])
to_fill_Values.append(abs(nullValue))
#This part followes the zero line back to define where things should be filled
if angles_to_save_cut[i+1] > angles_to_save_cut[i]:
go_back = [x for x in angles2 if x < angles_to_save_cut[i+1] and x > angles_to_save_cut[i]]
go_back = go_back[::-1]
go_back.pop(0)
else:
go_back = [x for x in angles2 if x < angles_to_save_cut[i+1]]
go_back2 = [x for x in angles2 if x > angles_to_save_cut[i]]
go_back = go_back[::-1]
if 0 in go_back:
go_back.pop(0)
go_back2 = go_back2[::-1]
go_back.extend(go_back2)
#add here the previously go back angles and values (values is always the radius of the zero line)
to_fill_angles.extend(go_back)
to_fill_Values.extend([abs(nullValue)] * len(go_back))
#in case there is a not directly a first cut adjust color
if nofirstcut == True and i == len(to_save)-1:
ax.fill(to_fill_angles, to_fill_Values, color=colorscheme[color][0])
else:
ax.fill(to_fill_angles, to_fill_Values, color=colorscheme[color][i])
#Plot the zero zScore line plus and minus 2 (significance
plt.plot(angles2,[abs(nullValue)]*361, color = 'black')
plt.yticks(range(0,len(negative_ticks),to_take),chosen_ticks)
# Add a title
plt.title(title, size=11, color=color, y=1.1)
plt.setp( ax.get_yticklabels(), visible=False)
plt.setp( ax.get_xticklabels(), visible=False)
# -
# #### 3.2 Create actual radar plots
# Go through the results of the indivdiual interactions types and create radar plots. The radar plots show Cohen's D (effect size) difference between the individual interaction results e.g. Increasing, Decreasing, Emergent compared to the overall interaction results. In case all 3 interaction types have very similar results, then also all 3 interactions types result in Cohen's D close to zero. High Cohen's D indicate big variability between interaction types
# +
# Interaction types
selected_results = {'Increasing':{},'Decreasing':{},'Emergent':{},'Interaction':{}}
# Define what to compare the results to, here choosen overall interaction results
compare_to = 'Interaction'
# The 12 representative features chosen to cover all feature classes
selected_features = ['ChemicalSimilarity','Enzymes_Overlap','Transporters_Overlap','PPI_Mean_AB_All_Filtered',
'KeGG_Indirect_Overlap','GO_Component','GO_Function','GO_Process','Msig_ChemGen_Perturbation_Overlap',
'SideEffects_CLOUD_to_Offsides_Overlap','SideEffects_TwoSide_CLOUDs','Disease']
# Read the result file and save the corresponding results
fp = open('../results/Molecular_Determinants/ResultsOverview.csv','r')
fp.next()
for line in fp:
tmp = line.strip().split(',')
if tmp[0] in selected_features:
if tmp[1] == compare_to:
selected_results[tmp[2]][tmp[0]] = float(tmp[8]) * -1
# +
#Define colors
interaction_colors = {'Increasing':'green','Decreasing':'red','Emergent':'blue','Interaction':'grey'}
# Create a spider plot for each interaction type separately
for key in ['Increasing','Decreasing','Emergent']:
part = key
categories = []
values = []
for f in selected_features:
categories.append(f)
values.append(selected_results[key][f])
my_SpiderPlot(categories, values, interaction_colors[part], part,'None',-1,2) #-1 and 2 for compare to Interaction, or -11 and 6
plt.savefig('../results/Molecular_Determinants/SpiderPlots/'+part+'.pdf',format='pdf')
plt.close()
# Create one spider plot for all 3 interaction types together
for key in ['Increasing','Decreasing','Emergent']:
part = key
categories = []
values = []
for f in selected_features:
categories.append(f)
values.append(selected_results[key][f])
my_SpiderPlot(categories, values, interaction_colors[part], part,'None',-1,2) #-1 and 2 for compare to Interaction, or -11 and 6
#plt.show()
plt.savefig('../results/Molecular_Determinants/SpiderPlots/Combined.pdf',format='pdf')
plt.close()
# -
# ### 4 Create final overview plot
# Create a final overview plot that visualizes which feature, in which network compartment (e.g. Core, Periphery ...) for which interaction type (increasing, decreasing ...) significant is. Therefor calculate depending on the type of interaction the signficance as well as a foldchange/oddsRation to get an idea wheter the feature is rather depleeted or enriched.
# #### 4.1 Create binned results (per network layer)
# Similar as in previous parts here, split the results accordingly into the various parts. Add here the network layer so that each result is properly sorted for each network layer as well as interaction type
# +
# The 12 representative features chosen to cover all feature classes
selected_features = ['ChemicalSimilarity','Enzymes_Overlap','Transporters_Overlap','PPI_Mean_AB_All_Filtered',
'KeGG_Indirect_Overlap','GO_Component','GO_Function','GO_Process','Msig_ChemGen_Perturbation_Overlap',
'SideEffects_CLOUD_to_Offsides_Overlap','SideEffects_TwoSide_CLOUDs','Disease']
# Define size and colors for the final plot
interaction_colors = {'Increasing':'#ACD900','Decreasing':'#F70020','Emergent':'#0096FF','All':'black'}
interaction_sizes = {'Increasing':200,'Decreasing':200,'Emergent':200,'All':2000}
network_parts = ['Complete','Core','CoreToPeriphery','Periphery']
# Get the result file
fp = open('../data/Molecular_Determinants/DrugPair_Feature_Overview.csv','r')
# Dictionary that will contain the information which interactions belong to which network layer
network_part_interactions = {}
# add the individual network parts to the network_part_interactions (result dictionary)
for part in network_parts:
network_part_interactions[part] = []
network_part = nx.read_gml('../data/Molecular_Determinants/Networks/DPI_Network_'+part+'.gml')
network_part_interactions[part] = network_part
# List containing all features (i.e. features in DrugPair_Feature_Overview.csv = all investigated features )
features = fp.readline().strip().split(',')[4:]
# Dictionary that will contain the individual results, split for network layers as well as interaction types
network_part_values = {}
# go through all features
for f in range(0,len(features)):
# always start at row one (first row containing results, zero row = header)
fp.seek(0)
fp.next()
# only continue if the feature is one of the representative features
if features[f] not in selected_features:
continue
print features[f]
# add section for this feature to the result dictionary: network_part_values
network_part_values[features[f]] = {}
# as next level add the individual network parts as well as interaction types
for part in network_parts:
network_part_values[features[f]][part] = {'Increasing':[],'Decreasing':[],'Emergent':[]}
network_part_values[features[f]]['AllCLOUDS'] = []
network_part_values[features[f]]['NonInteracting'] = []
# now go through all results and add every feature result into the correct bin
for line in fp:
tmp = line.strip().split(',')
# do not include 'nan' values (e.g. if one drug has no targets then PPI mean distance = nan)
if tmp[f+4] == 'nan':
continue
interaction_found = False
#only include pure single edges e.g. do not include increasing/decreasing interactions
if tmp[3] == 'Increasing' or tmp[3] == 'Decreasing' or tmp[3] == 'Emergent' or tmp[3] == 'None':
# AllCLOUDs = all pairs (is always added)
network_part_values[features[f]]['AllCLOUDS'].append(float(tmp[f+4]))
######
# Add the result accordingly (which interaction type or network layer it belongs)
# This creates the actual final network_part_values dictionary that will be used in the next step to create the overview plot
for part in network_parts:
if network_part_interactions[part].has_edge(tmp[0],tmp[1]):
interaction_found = True
for key in network_part_interactions[part][tmp[0]][tmp[1]]:
network_part_values[features[f]][part][network_part_interactions[part][tmp[0]][tmp[1]][key]['Type']].append(float(tmp[f+4]))
if network_part_interactions[part].has_edge(tmp[1],tmp[0]):
interaction_found = True
for key in network_part_interactions[part][tmp[1]][tmp[0]]:
network_part_values[features[f]][part][network_part_interactions[part][tmp[1]][tmp[0]][key]['Type']].append(float(tmp[f+4]))
if interaction_found == False:
network_part_values[features[f]]['NonInteracting'].append(float(tmp[f+4]))
fp.close()
# -
# #### 4.2 Create actual overview plot
# Use the results stored in network_part_values to create an easy overview plot. Split the result into the individual network layers (= rows) and features (=columns). Each cell (row X column) will have 4 triangles: one black big triangle = All interaction types, and 3 smaller ones indicating the individual interaction type results (emergent, increasing and decreasing)
# +
# Create overview plot for SELECTED features
all_Features = selected_features
# Create overview plot for ALL features
#all_Features = features
# Get number of features
number_features = len(all_Features)
# Change size of the final plot accordingly to the number of features (more features = larger plot)
plt.figure(figsize=(number_features,4))
#plt.tight_layout()
feature_names = []
# position to start adding entries, with each features will be increased by 0.2
current_x = 0.8
# Go through all features
for f in all_Features:
print f
#add the feature name
feature_names.append(f)
#save NoInteraction values
no_interaction_values = network_part_values[f]['NonInteracting']
# similar as current_x, defines where to put the results on the y axis
y = 4.2
#Go through all network parts
for part in network_parts:
# AllInteractions is simply the union of the 3 different interaction types
AllInteractions = network_part_values[f][part]['Increasing'] + network_part_values[f][part]['Decreasing'] + network_part_values[f][part]['Emergent']
# Things to test include AllInteraction, Increasing, Decreasing and Emergent
things_to_test = {'All':AllInteractions,'Increasing':network_part_values[f][part]['Increasing'],'Decreasing':network_part_values[f][part]['Decreasing'],'Emergent':network_part_values[f][part]['Emergent']}
# Check wheter the feature is a continues feature (Mann Whitney U) or a binary feature (Fisher Exact test)
continues_features = True
if all(v == 0 or v ==1 for v in no_interaction_values):
continues_features = False
x = current_x
# Calculate the signficance accordingly
for subset in ['All','Increasing','Decreasing','Emergent']:
# If continues feature calculate significance according to Mann Whitney U
if continues_features:
direction = np.mean(things_to_test[subset]) > np.mean(no_interaction_values)
sign = mu(things_to_test[subset],no_interaction_values)[1] < 0.05
# If binary feature calculate significance according to Fisher Exact test
else:
real_Overlap = sum(things_to_test[subset])
real_NonOverlap = len(things_to_test[subset]) - real_Overlap
non_Interactions_Overlap = sum(no_interaction_values)
non_Interactions_NonOverlap = len(no_interaction_values) - sum(no_interaction_values)
oddsratio, pvalue = stats.fisher_exact([[real_Overlap, real_NonOverlap], [non_Interactions_Overlap, non_Interactions_NonOverlap]])
sign = pvalue < 0.05
direction = oddsratio > 1
# Depending on the fold change/ oddsRation define if the feature is rather depleeted or enriched (arrow down or arrow up)
if direction:
symbol = '^'
else:
symbol = 'v'
if sign:
color = interaction_colors[subset]
else:
color = 'grey'
# add the cell entry accordingly (color if significant, arrow according to depleetion or emergence)
x = x + 0.2
plt.scatter([x],[y],marker=symbol, s=interaction_sizes[subset], color=color)
y = y - 1
current_x = current_x + 1
# Create the output folder if it doesn't exist
directory = os.path.dirname('../results/Molecular_Determinants/')
if not os.path.exists(directory):
os.makedirs(directory)
# Create the final output (overview plot)
plt.ylim([0.6,4.8])
plt.xlim([0.3,number_features+1])
plt.yticks([1.2,2.2,3.2,4.2],['Periphery','CoreToPeriphery','Core','Complete'])
plt.xticks(range(1,number_features),feature_names, rotation='vertical')
plt.savefig('../results/Molecular_Determinants/Overviewplot.pdf', bbox_inches = "tight")
plt.close()
| code/13_Molecular_Determinants.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# > 1. **Necesity**: Build Model
# > 2. **Google**: How do you search for the solution?
# > 3. **Solution**: Find the `function()` that makes it happen
# ## Code Thinking
#
# > Which function computes the Model?
# > - `fit()`
# >
# > How could can you **import the function in Python**?
fit()
model.fit()
# `model = ?`
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier(max_depth=3)
model.fit()
# ### Separate Variables for the Model
#
# > Regarding their role:
# > 1. **Target Variable `y`**
# >
# > - [ ] What would you like **to predict**?
# >
# > Total number of accients? Or Alcohol?
# >
# > 2. **Explanatory Variable `X`**
# >
# > - [ ] Which variable will you use **to explain** the target?
df.drop(labels="survived")
df.drop(labels="survived", axis=1)
df.drop(columns='survived', labels='sex')
explanatory
import pandas as pd
df
a = pd.DataFrame({'raza': ['blanco', 'blanco', 'negro', 'asiatico']})
pd.get_dummies(a)
a
pd.get_dummies(a, drop_first=True)
df.isna().sum()
mask = df.isna().sum(axis=1) > 0
df[mask]
df
df = df.dropna()
df
df = pd.get_dummies(df, drop_first=True)
df
explanatory = df.drop(columns='survived')
explanatory
target = df.survived
target
# ### Finally `fit()` the Model
model.fit(X=explanatory, y=target)
# ## Calculate a Prediction with the Model
# > - `model.predict_proba()`
juan = df.sample(random_state=42)
juan
juan_pred = juan.drop(columns='survived')
juan_pred
model.predict(X=juan_pred)
model.predict_proba(X=juan_pred)
# ## Model Visualization
# > - `tree.plot_tree()`
# +
# tree?
# -
from sklearn import tree
model
type(model)
import matplotlib.pyplot as plt
explanatory
explanatory.columns
plt.figure(figsize=(20,10))
tree.plot_tree(decision_tree=model, feature_names=explanatory.columns, filled=True);
# ## Model Interpretation
# > Why `sex` is the most important column? What has to do with **EDA** (Exploratory Data Analysis)?
330
39/330
juan
juan_pred = juan.drop(columns='survived')
juan_pred
model.predict(X=juan_pred)
model.predict_proba(X=juan_pred)
# # Prediction vs Reality
# > How good is our model?
# ## Precision
# > - `model.score()`
model.score(X=explanatory, y=target)
dfsel = df[['survived']].copy()
pred = model.predict(X=explanatory)
dfsel['pred'] = pred
dfsel
dfsel.pred == dfsel.survived
(dfsel.pred == dfsel.survived).sum()
(dfsel.pred == dfsel.survived).sum()/714
(dfsel.pred == dfsel.survived).mean()
# ## Confusion Matrix
# > 1. **Sensitivity** (correct prediction on positive value, $y=1$)
# > 2. **Specificity** (correct prediction on negative value $y=0$).
# > - `plot_confusion_matrix()`
from sklearn.metrics import plot_confusion_matrix
# - imagina que tenemos 100 personas
# - 90 no tienen covid
# - 10 tienen covid
# - el modelo
# - predice todas las personas con no covid 100...
#
# - metricas
# - precision: 90%
# - la gente que tiene covid: 0% → sensibilidad
# - la gente que no tiene covid: 100% → especificidad
plot_confusion_matrix(estimator=model, X=explanatory, y_true=target);
sensibilidad = 205/(85 + 205)
sensibilidad
especificidad = 372/(372 + 52)
especificidad
from sklearn.metrics import classification_report
classification_report(y_true=dfsel.survived, y_pred=dfsel.pred)
a = classification_report(y_true=dfsel.survived, y_pred=dfsel.pred)
print(a)
# ## ROC Curve
# > A way to summarise all the metrics (score, sensitivity & specificity)
from sklearn.metrics import plot_roc_curve
plot_roc_curve(estimator=model, X=explanatory, y=target)
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(X=explanatory, y=target)
plot_roc_curve(estimator=lr, X=explanatory, y=target)
from sklearn.ensemble import RandomForestClassifier
lr = RandomForestClassifier()
lr.fit(X=explanatory, y=target)
plot_roc_curve(estimator=lr, X=explanatory, y=target)
| II Machine Learning & Deep Learning/#06. Decision Tree. A Supervised Classification Model/06session.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
The MIT License (MIT)
Copyright (c) 2021 NVIDIA
Permission is hereby granted, free of charge, to any person obtaining a copy of
this software and associated documentation files (the "Software"), to deal in
the Software without restriction, including without limitation the rights to
use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of
the Software, and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS
FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR
COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER
IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
"""
# This code example is very similar to c5e1_mnist_learning but the network is modified to use ReLU neruons in the hidden layer, softmax in the output layer, categorical crossentropy as loss function, Adam as optimizer, and a mini-batch size of 64. More context for this code example can be found in the section "Experiment: Tweaking Network and Learning Parameters" in Chapter 5 in the book Learning Deep Learning by <NAME> (ISBN: 9780137470358).
# +
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.utils import to_categorical
import numpy as np
import logging
tf.get_logger().setLevel(logging.ERROR)
EPOCHS = 20
BATCH_SIZE = 64
# Load training and test datasets.
mnist = keras.datasets.mnist
(train_images, train_labels), (test_images,
test_labels) = mnist.load_data()
# Standardize the data.
mean = np.mean(train_images)
stddev = np.std(train_images)
train_images = (train_images - mean) / stddev
test_images = (test_images - mean) / stddev
# One-hot encode labels.
train_labels = to_categorical(train_labels, num_classes=10)
test_labels = to_categorical(test_labels, num_classes=10)
# Create a Sequential model.
# 784 inputs.
# Two Dense (fully connected) layers with 25 and 10 neurons.
# relu as activation function for hidden layer and
# He normal initializer.
# Softmax as activation function for output layer
# and Glorot uniform initializer.
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(25, activation='relu',
kernel_initializer='he_normal',
bias_initializer='zeros'),
keras.layers.Dense(10, activation='softmax',
kernel_initializer='glorot_uniform',
bias_initializer='zeros')])
# Use Adam optimizer with default parameters.
# Categorical cross-entropy as loss function and
# report accuracy during training.
model.compile(loss='categorical_crossentropy',
optimizer = 'adam',
metrics =['accuracy'])
# Train the model for 20 epochs.
# Shuffle (randomize) order.
# Update weights after 64 examples (batch_size=64).
history = model.fit(train_images, train_labels,
validation_data=(test_images, test_labels),
epochs=EPOCHS, batch_size=BATCH_SIZE,
verbose=2, shuffle=True)
| tf_framework/c5e2_mnist_learning_conf5.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
// # Extending ImageJ: Data I/O
// This notebook demonstrates how to write an `IOPlugin`, which handles reading and/or writing of external data to and from Java data structures.
// %classpath config resolver imagej.public https://maven.imagej.net/content/groups/public
// %classpath add mvn net.imagej imagej 2.0.0-rc-71
ij = new net.imagej.ImageJ()
"ImageJ v${ij.getVersion()} is ready to go."
// ## Creating a data reader
// Suppose we have data stored in a file format, `.char-table`, which represents a table as a sequence of characters, along with metadata defining the number of rows and columns. We would like to write a plugin so that ImageJ can import these files via its _File ▶ Open_ command.
//
// The format of a `.char-table` file is key-value pairs, one per line. Valid keys are:
//
// * `cols` to define the column count
// * `rows` to define the row count
// * `data` to specify the actual character data: a sequence of $cols \times rows$ characters enumerated in row-major order, and bracketed by `<` and `>`.
//
// Let's start by writing out an example `.char-table` file, which we will use for testing:
import java.io.File
import org.scijava.util.DigestUtils
import org.scijava.util.FileUtils
data = """
cols = 7
rows = 7
data = <@@@@@@@@ @ @ @ @@ @ @ @@ @ @ @@ @ @ @@@ >
"""
desktop = System.getProperty("user.home") + "/Desktop/"
tablePath = desktop + "example.char-table"
outFile = new File(tablePath)
FileUtils.writeFile(outFile, DigestUtils.bytes(data))
"Wrote ${outFile.length()} bytes to '$outFile'"
// Without further ado, here is the `IOPlugin` implementation:
// +
import java.io.File
import org.scijava.io.AbstractIOPlugin
import org.scijava.io.IOPlugin
import org.scijava.plugin.Plugin
import org.scijava.table.DefaultGenericTable
import org.scijava.table.Table
import org.scijava.util.DigestUtils
import org.scijava.util.FileUtils
@Plugin(type = IOPlugin.class)
public class CharTableReader extends AbstractIOPlugin<Table> {
@Override
public Class<Table> getDataType() {
// This is the type of object produced by the reader.
return Table.class
}
@Override
public boolean supportsOpen(final String source) {
// Check whether the source is indeed a .char-table.
// This check can be as shallow or as deep as you want,
// but it is advised to keep it as fast as possible.
// As such, it is not recommended to actually open and
// interrogate the source unless you have no choice.
return source.toLowerCase().endsWith(".char-table")
}
@Override
public Table open(final String source) throws IOException {
// This is where we read the data from its source,
// and convert it into the destination data type.
// Read in the file.
String contents = DigestUtils.string(FileUtils.readFile(new File(source)))
// Parse the contents line by line.
int rows = 0, cols = 0
String data = null
for (line in contents.split("\n")) {
int equals = line.indexOf("=")
if (equals < 0) continue
String key = line.substring(0, equals).trim()
String val = line.substring(equals + 1).trim()
switch (key) {
case "rows":
rows = Integer.parseInt(val)
break
case "cols":
cols = Integer.parseInt(val)
break
case "data":
data = val
break
}
}
// Do some error checking.
if (rows <= 0) throw new IOException("Missing or invalid rows")
if (cols <= 0) throw new IOException("Missing or invalid cols")
if (data == null || !data.startsWith("<") || !data.endsWith(">")) {
throw new IOException("Missing or invalid data")
}
if (cols * rows != data.length() - 2) {
throw new IOException("Expected data length ${cols * rows} but was ${data.length() - 2}")
}
// Build the resultant table.
Table table = new DefaultGenericTable(cols, rows)
int index = 1
for (int r = 0; r < rows; r++) {
for (int c = 0; c < cols; c++) {
table.set(c, r, data.charAt(index++))
}
}
// HACK: Work around an SJJK bug when column headers are unspecified.
for (int c = 0; c < cols; c++) table.setColumnHeader(c, "")
return table
}
// HACK: Work around weird bug in Groovy(?).
// It is normally not needed to override this method here.
@Override
public Class<String> getType() { return String.class }
}
// Register the plugin with the existing SciJava context.
import org.scijava.plugin.PluginInfo
info = new PluginInfo(CharTableReader.class, IOPlugin.class)
ij.plugin().addPlugin(info)
info
// -
// Now that we have an `IOPlugin` registered to handle the reading of `.char-table` files, let's give it a spin on the example data we wrote earlier:
table = ij.io().open(tablePath)
ij.notebook().display((Object) table)
// ## Creating a data writer
// Similarly, `IOPlugin`s also extend ImageJ's capabilities when writing data to external sources. In the future, ImageJ will have a unified _File ▶ Save As..._ command which offers all available export options in a unified UI. But for the moment, the routine must be called programmatically via the `IOService`.
//
// Let's write an exporter for tables to another custom file format: `.ascii-table`. This format writes each table cell as a single readable ASCII character (32 - 126 inclusive); characters outside this range are written as `-` (45). All columns of a row are written on the same line, with a newline between each row. Of course, this can be a lossy export.
//
// Here is the exporter implementation for `.ascii-table`:
// +
import java.io.File
import org.scijava.Priority
import org.scijava.io.AbstractIOPlugin
import org.scijava.io.IOPlugin
import org.scijava.plugin.Plugin
import org.scijava.table.DefaultGenericTable
import org.scijava.table.Table
import org.scijava.util.DigestUtils
import org.scijava.util.FileUtils
@Plugin(type = IOPlugin.class)
public class AsciiTableWriter extends AbstractIOPlugin<Table> {
@Override
public Class<Table> getDataType() {
// This is the type of object exported by the writer.
return Table.class
}
@Override
public boolean supportsSave(final String destination) {
// Check whether the destination should be a .ascii-table.
// This is typically a format extension check.
return destination.toLowerCase().endsWith(".ascii-table")
}
@Override
public void save(final Table table, final String destination) throws IOException {
// This is where we write the data to its destination,
// converting it from the source data type.
// Define the default character.
byte other = (byte) '-'
// Build up the output bytes.
int cols = table.getColumnCount()
int rows = table.getRowCount()
byte[] bytes = new byte[(cols + 1) * rows]
int index = 0
for (int r = 0; r < rows; r++) {
for (int c = 0; c < cols; c++) {
Object cell = table.get(c, r)
String s = cell == null ? null : cell.toString()
if (s == null || s.length() == 0) {
bytes[index++] = other
continue
}
int v = s.charAt(0)
bytes[index++] = v >= 32 && v <= 126 ? (byte) v : other
}
bytes[index++] = '\n'
}
// Write out the file.
FileUtils.writeFile(new File(destination), bytes)
}
// HACK: Work around weird bug in Groovy(?).
// It is normally not needed to override this method here.
@Override
public Class<String> getType() { return String.class }
}
// Register the plugin with the existing SciJava context.
import org.scijava.plugin.PluginInfo
info = new PluginInfo(AsciiTableWriter.class, IOPlugin.class)
ij.plugin().addPlugin(info)
// HACK: Refresh the IOService. (This bug is fixed on scijava-common master.)
import org.scijava.util.ClassUtils
ClassUtils.setValue(ClassUtils.getField(org.scijava.plugin.AbstractSingletonService.class, "instances"), ij.io(), null)
info
// -
outPath = desktop + "fiji.ascii-table"
ij.io().save(table, desktop + "fiji.ascii-table")
outFile = new File(outPath)
"Wrote ${outFile.length()} bytes to '$outFile'"
// Check that it did what we wanted:
import org.scijava.util.DigestUtils
import org.scijava.util.FileUtils
DigestUtils.string(FileUtils.readFile(outFile))
// ## Supporting both reading and writing
//
// If you wish to support both reading and writing to/from the same format, you can include both `open` and `save` implementations in the same `IOPlugin`.
| notebooks/2-Extending-ImageJ/2-Input-Output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
@author: TomClifford
This file is an initial routine for preprocessing seismic data.
It reads a waveform, filters, removes response, demeans and detrends, finds SNR, FAS, and plots.
"""
#%% import libraries
import obspy
from obspy.clients.fdsn.mass_downloader import CircularDomain, \
Restrictions, MassDownloader
from obspy.io.xseed import Parser
from obspy.signal import PPSD
from obspy.signal import freqattributes
import os
from scipy.fft import fft, ifft, fftfreq
from scipy.integrate import cumtrapz
import numpy as np
from matplotlib import pyplot as plt
import pandas as pd
from response_spectrum import *
# +
# #%%paths
data_path = r"C:\Users\TomClifford\SlateGeotech\Duynefontyn PSHA - DuynefontynPSHA\05 - GMM\GMM_Scripts\preprocessing"
os.chdir(data_path)
#%% read waveform data into an obspy stream object, st
#origin time
origin_time = obspy.UTCDateTime(2016,10,18,6,25,33)
#https://docs.obspy.org/packages/autogen/obspy.core.stream.Stream.html
st = obspy.read(os.path.join(data_path, 'waveforms/*'))
print(st)
# +
# #%%remove response
inv = obspy.read_inventory(os.path.join(data_path, 'stations/*'))
# response_list = os.listdir(os.path.join(data_path, 'stations'))
#create empty stream object to add once response removed
st_r = obspy.core.stream.Stream()
#prefilter, remove response, and append new trace to stream
#tr is a waveform trace: https://docs.obspy.org/packages/autogen/obspy.core.trace.Trace.html
for tr in st:
#determine nyquist f
nyq = tr.stats.sampling_rate/2
#set prefilter according to nyquist
prefilter = [0.001, 0.005, nyq-5, nyq]
#find matching response
tr_response = tr.copy()
tr_response.remove_response(inventory=inv,
pre_filt = prefilter,
output = "ACC",
)
st_r.append(tr_response)
# print(tr_response)
st_rd = st_r.copy()
#https://docs.obspy.org/packages/autogen/obspy.core.trace.Trace.detrend.html
#demean
st_rd.detrend('demean')
#detrend
st_rd.detrend('linear')
#trim waveform
st_rd.trim(origin_time, origin_time+(1000/2))
for tr in st_rd:
print(tr)
tr.plot()
# #%%SNR
def snr(trace):
#input: obspy trace object
peak = trace.data.max()
rms = np.sqrt(np.mean(trace.data**2))
snr = peak/rms
return snr
#%% test snr
for tr in st_rd:
# tr.plot()
print(snr(tr))
# +
#%% FAS
def fas(tr):
#tr: obspy trace object
y = fft(tr.data)
yf = 2.0/tr.stats.npts*np.abs(y[:(tr.stats.npts//2)])
xf = fftfreq(tr.stats.npts, tr.stats.delta)[:(tr.stats.npts//2)]
return xf, yf
x, y = fas(st_rd[0])
#plot
fig, ax = plt.subplots()
ax.plot(x, y, lw=0.3, color='k')
ax.set_xlabel("Frequency [Hz]")
ax.set_ylabel("Amplitude")
plt.yscale('log')
plt.show()
#%% response spectra
#get response spectra from trace
#seems number of periods has to be same length as trace?
r = NewmarkBeta([tr.times(), tr.data/100], tr.stats.delta, np.logspace(.1, 10, len(tr.data)))#np.array([.1,1,10])) #convert to cm/s/s
#why returning period instead of time
plt.plot(r.response_spectrum['Period'], r.response_spectrum['Acceleration']) #so this is the waveform
#%% save trace amplitudes, times, and fourier spectra to excel
for tr in st:
print(tr)
trace_data = pd.DataFrame({'trace_amplitudes': tr.data,
'trace_time' : tr.times()
})
trace_fft = pd.DataFrame({'fftx': fas(tr)[0],
'ffty': fas(tr)[1]
})
trace_data.to_csv(os.path.join(data_path, 'raw_traces', str(tr.id)+'_data.csv'))
trace_fft.to_csv(os.path.join(data_path, 'raw_traces', str(tr.id)+'_fft.csv'))
#%% download data for event - not for final script
#M4.5 2016-10-18 06:25:33.160000
# origin_time = obspy.UTCDateTime(2016,10,18,6,25,33)
# domain = CircularDomain(latitude=-33.676667, longitude=18.431389,
# minradius=0.0, maxradius= 1000/111) #1000 km to deg
# restrictions = Restrictions(
# #5 minutes before, 30 minutes after origin
# starttime=origin_time - 5 * 60,
# endtime=origin_time + 30*60,
# network='*',
# station = '*',
# location='*',
# # channel='*', #allowing all channels downloads non-seismic data
# reject_channels_with_gaps=False,
# )
# mdl = MassDownloader(providers=['IRIS'])
# mdl.download(domain, restrictions, mseed_storage=data_path+"/waveforms",
# stationxml_storage=data_path+"/stations")
# #downloads 13 waveforms from 3 stations
| processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import mc_sim_usman as mcn
import numpy as np
import matplotlib.pyplot as plt
# special jupyter notebook command to make plots interactive
# %matplotlib notebook
coordinates, box_length = mcn.read_xyz('../../../lj_sample_configurations/lj_sample_config_periodic1.txt')
final_coordinates, time_average_gr, r_values = mcn.run_simulation_np(coordinates, box_length, 3, 0.9, 5000, engine='numpy')
print("Creating RDF plot at T=0.9")
rdf_fig = plt.figure(figsize=(6,6))
rdf_ax = rdf_fig.add_subplot()
rdf_ax.plot(r_values, time_average_gr,'-og')
rdf_ax.set_xlabel("r/$\sigma$")
rdf_ax.set_ylabel("g(r)")
plt.savefig("Time_avg_RDF_at_T_0p9.png", dpi=200)
| homework/day_5/mc_package_usman/Testing_functionality_and_time_avg_rdf_usman.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2. Améliorations - Le système solaire n'est pas parfaitement un plan :
#
# ## 2.1 Prise en compte de la dimension Z:
#
# Dans la première partie de ce rapport, on a utilisé le schéma de Verlet afin de simuler le système solaire en 2D, mais, il faut mentionner le fait que les planètes de système solaire ne sont pas parfaitement dans le plan de l'écliptique, il faudra alors ajouter la composante $Z$ aux paramètres de mouvements pour que la trajectoire de la sonde soit précise, d'où les modifications suivantes:
#
# Ps: les nouvelles modifications seront dans le dossier en **In_3dim**
#
# Il faut modifier la classe **objet** et les **fonctions auxiliaires** :
#
# +
import numpy as np
import matplotlib.pyplot as plt
class objet:
""" Classe représentant les objets qui influence par la gravitation
Attributs:
nom
masse: Kg
position (x, y, z): au
vitesse (v_x, v_y, v_z) : au/day
"""
nom = "objet"
masse = None
x0 = 0
y0 = 0
z0 = 0
vx0 = 0
vy0 = 0
vz0 = 0
#Listes des positions et vitesse
x = None
y = None
z = None
vx = None
vy = None
vz = None
def __init__(self, nom = "objet", masse = None, x0 = 0, y0 = 0, z0 = 0, vx0 = 0, vy0 = 0, vz0 = 0):
"""Constructeur de notre classe"""
self.nom = nom
self.masse = masse
self.x0 = x0
self.y0 = y0
self.z0 = z0
self.vx0 = vx0
self.vy0 = vy0
self.vz0 = vz0
# -
# **Variables Globales:**
# +
au = 1.49597870e11 #Unité astronomique
jour = 24*3600 #Un jour
G = 6.67408e-11 #Constante gravitationelle
# -
# **Modification des fonctions auxiliares :**
# +
#Definition de fonction fx(M,x,y,z) et fy(M,x,y,z)
def fx(M,x,y,z):
"""
Retourne l'acceleration gravitationnelle suivant x dû à un objet de masse M distants de l'objet étudié de x**2+y**2+z**2
"""
return -((G*M)/(x**2+y**2+z**2)**(3/2))*x*(jour**2/au**3)
def fy(M,x,y,z):
"""
Retourne l'acceleration gravitationnelle suivant y dû à un objet de masse M distants de l'objet étudié de x**2+y**2+z**2
"""
return -((G*M)/(x**2+y**2+z**2)**(3/2))*y*(jour**2/au**3)
def fz(M,x,y,z):
"""
Retourne l'acceleration gravitationnelle suivant z dû à un objet de masse M distants de l'objet étudié de x**2+y**2+z**2
"""
return -((G*M)/(x**2+y**2+z**2)**(3/2))*z*(jour**2/au**3)
def E(M, x, y, z, vx, vy, vz):
"""
Calculer l'energie massique d'un objet sous effet d'un seul objet de masse M
"""
return 0.5*(vx**2+vy**2+vz**2)*(au**2/jour**2)-(G*M)/(np.sqrt(x**2+y**2+z**2)*au)
E = np.vectorize(E) #Vectoriser une fonction est benefique en terme de performance et memoire
def pot(M, x, y, z):
"""
Retourne le potentiel massique d'un objet par rapport à un autre objet de masse M et distant de x**2+y**2+z**2
"""
return -(G*M)/(np.sqrt(x**2+y**2+z**2)*au)
pot = np.vectorize(pot)
# -
# Comme c'est fait dans la partie précédente, on va aussi généraliser les fonctions **acceleration** et **Energy** en 3D.
# +
def acceleration(bodies, i, j):
"""
Calculer l'acceleration relative à un objet bodies[i]
bodies: tous les objets
i: index of concerned body which undergoes the gravitation of other objects.
j: index of the step
"""
N = len(bodies)
ax = 0; ay = 0; az = 0 #L'acceleration
for ip in range(N):
#Chaque objet bodies[ip] applique une force de gravitation sur l'objet bodies[i]
if ip == i: #On veut que pas avoir le même objet bodies[ip]
continue
ax += fx(bodies[ip].masse, bodies[i].x[j]-bodies[ip].x[j], bodies[i].y[j]-bodies[ip].y[j], bodies[i].z[j]-bodies[ip].z[j])
ay += fy(bodies[ip].masse, bodies[i].x[j]-bodies[ip].x[j], bodies[i].y[j]-bodies[ip].y[j], bodies[i].z[j]-bodies[ip].z[j])
az += fz(bodies[ip].masse, bodies[i].x[j]-bodies[ip].x[j], bodies[i].y[j]-bodies[ip].y[j], bodies[i].z[j]-bodies[ip].z[j])
return (ax, ay, az)
def Energy(bodies, i):
"""
L'Energie massique d'un objet sous l'effet d'autres objet qui lui entoure.
"""
N = len(bodies)
potential = 0
for ip in range(N):
if ip == i:
continue
potential += pot(bodies[ip].masse, bodies[i].x-bodies[ip].x, bodies[i].y-bodies[ip].y, bodies[i].z-bodies[ip].z)
return 0.5*(au**2/jour**2)*(bodies[i].vx**2+bodies[i].vy**2+bodies[i].vz**2)+potential
# -
# **Il faut aussi modifier le fichier des conditions initiales :**
#
# NB: On prend les mêmes conditions initiales qu'auparavant mais en ajoutant les informations $Z$ et $V_z$.
# +
import os
os.chdir("/home/mh541/Desktop/Projet_Numerique/In_3dim") #Please change to the directory where 'initial_conditions_solarsystem.txt' is saved
data = np.genfromtxt("initial_conditions_solarsystem.txt", usecols=(1,2,3,4,5,6,7), skip_header=1) #On ne peut pas importer du texte avec genfromtxt à cause de problème d'encodage
names = np.loadtxt("names_solarsystem.txt", dtype = str, skiprows=1, usecols=(1,)) #Importation de texte (nomes des objets dans le systeme solaire)
# -
# **Il aussi rénitialiser les données :**
#
# +
bodies = np.array([objet() for i in range(10)]) #Creation d'une liste des objets (on a au total 9 objets: soleil et 9 planetes)
Nbr_obj = len(bodies) #Definition de nombres d'objets
#Definition des parametres de chaque objet
for i in range(Nbr_obj):
bodies[i].nom = names[i][2:-1] # [2:-1] pour supprimer les caracteres indesires
bodies[i].masse = data[i][0]
bodies[i].x0 = data[i][1]
bodies[i].y0 = data[i][2]
bodies[i].z0 = data[i][3]
bodies[i].vx0 = data[i][4]
bodies[i].vy0 = data[i][5]
bodies[i].vz0 = data[i][6]
# -
# ## 2.2 Implémentation de trajectoires des planètes de système solaire:
#
# Maintenant, comme on a adapté ce qui est fait dans la partie 1 au cas général, on peut implémenter les trajectoires des planètes de la même manière que la partie 1 avec le schéma de Verlet.
#
# PS: On fait l'hypothèse que le soleil reste fixe.
# +
#Definition de pas de temps
dt = 1 #step
T = int(365/dt)*10 # (Nombre de steps)<-> Periode d'integration T = int(365/dt)* nbr ans
#Intialisation des attributs x,y,z,vx,vy,vz de chaque objet bodies[i]
for i in range(Nbr_obj):
bodies[i].x = np.zeros(T); bodies[i].x[0] = bodies[i].x0
bodies[i].y = np.zeros(T); bodies[i].y[0] = bodies[i].y0
bodies[i].z = np.zeros(T); bodies[i].z[0] = bodies[i].z0
bodies[i].vx = np.zeros(T); bodies[i].vx[0] = bodies[i].vx0
bodies[i].vy = np.zeros(T); bodies[i].vy[0] = bodies[i].vy0
bodies[i].vz = np.zeros(T); bodies[i].vz[0] = bodies[i].vz0
#Definitions des v_demi.
vx_demi = np.zeros(Nbr_obj)
vy_demi = np.zeros(Nbr_obj)
vz_demi = np.zeros(Nbr_obj)
#Implementation de l'integrateur de Verlet pour chaque objet (sauf le soleil)
for j in range(T-1):
#Phase 1: Calcul de vitesses milieu et affectation des position a l'intant j+1
for i in range(1,Nbr_obj): #Modification des parametres pour chaque objet a l' instant j
#Récupération de l'acceleration
fx_j, fy_j, fz_j = acceleration(bodies, i, j) #L'acceleration au pas j relative à l'objet i
#Definition des variables de milieux
vx_demi[i] = bodies[i].vx[j] + (dt/2)*fx_j
vy_demi[i] = bodies[i].vy[j] + (dt/2)*fy_j
vz_demi[i] = bodies[i].vz[j] + (dt/2)*fz_j
# Affectation des positions à l'indice i+1
bodies[i].x[j+1] = bodies[i].x[j] + dt*vx_demi[i]
bodies[i].y[j+1] = bodies[i].y[j] + dt*vy_demi[i]
bodies[i].z[j+1] = bodies[i].z[j] + dt*vz_demi[i]
#Phase 2: Affectation des vitesse a l'instant j+1
for i in range(1,Nbr_obj):
#L'acceleration au pas i+1 relative à l'objet j
fx_jplus1, fy_jplus1, fz_jplus1 = acceleration(bodies, i, j+1) #Il faut faire cette étape après le calcul de postion à l'indice i+1
# Affectation des vitesses à l'indice j+1
bodies[i].vx[j+1] = vx_demi[i] + (dt/2)*fx_jplus1
bodies[i].vy[j+1] = vy_demi[i] + (dt/2)*fy_jplus1
bodies[i].vz[j+1] = vz_demi[i] + (dt/2)*fz_jplus1
#[End]----------------------------------------------------------------------------------------------------
# -
# **Plot des trajectoires :**
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe
for i in range(1,Nbr_obj):
plt.plot(bodies[i].x, bodies[i].y)
plt.xlabel("x (Au)")
plt.ylabel("y (Au)")
plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
plt.show()
# -
# **Estimation précision en regardant l'énergie mécanique:**
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe
Nrg = Energy(bodies, 1) #Cacul de l'energie d'un objet -> Changez le numero pour voir l'energie de chaque objet;
Nrg /= np.abs(Nrg[0]) #Pour Normaliser
#Plot de l'energie
t = np.linspace(1,T,T)*dt
ax.plot(t, Nrg)
# ax.plot(t[:365], Nrg[:365])
ax.set_xlabel("t (jour)")
ax.set_ylabel("E/$|E_0|$")
ax.get_yaxis().get_major_formatter().set_useOffset(False) #Disable scaling of values in plot wrt y-axis
#Affichage des résulats
print("Résultats : ")
print("Energie moyenne = " + str(np.mean(Nrg)) + ", Ecart_Type = " + str(np.std(Nrg)))
plt.show()
# -
# On remarque ici qu'on obtient une précision similaire à celle de l'approche en 2 dimensions, pour encore avoir de précision, il suffit juste de raffiner le pas d'intégration.
# # 3- Comparaison avec des trajectoires non couplées:
#
# Pour savoir s'il est nécessaire d'inclure les effets des autres planètes, on va comparer les trajectoires obtenues précédement avec celles de 9 planètes supposées indépendantes: c-à-d, on considère seulement la gravitation de soleil sur chaque planète.
#
# Dans le cas des trajectoires non couplées, on va considèrer la fonction **acceleration_sol** qui permet de calculer l'accéleration gravitationnelle dû au soleil sur chaque objet et aussi **Energy_sol** qui calcule l'énergie mécanique totale d'une planète sous effet de soleil seulement.
# +
def acceleration_sol(bodies, i, j): #avec effet de soleil seulement
"""
Calculer l'acceleration relative à un objet bodies[i]
bodies: tous les objets
i: indice de l'objet concerne qui subit la gravitation du soleil seulement
j: indice de pas
"""
ax = fx(bodies[0].masse, bodies[i].x[j], bodies[i].y[j], bodies[i].z[j])
ay = fy(bodies[0].masse, bodies[i].x[j], bodies[i].y[j], bodies[i].z[j])
az = fz(bodies[0].masse, bodies[i].x[j], bodies[i].y[j], bodies[i].z[j])
return (ax, ay, az)
def Energy_sol(bodies,i):
"""
L'Energie massique d'un objet sous l'effet de soleil seulement.
"""
potential = pot(bodies[0].masse, bodies[i].x, bodies[i].y, bodies[i].z)
return 0.5*(au**2/jour**2)*(bodies[i].vx**2+bodies[i].vy**2+bodies[i].vz**2)+potential
# -
# Maintenant, on a tout ce qu'il faut pour effectuer la comparaison entre les trajectoires couplées et non couplées. On a déja fait une implémentation des trajectoires couplées en haut, il suffit de sauvegarder les résultats obtenues afin de pouvoir rénitialiser les attributs de ** bodies ** et implémenter les trajectoires non couplées.
#
# La manière la plus simple de sauvegarder ces données, c'est d'affecter le tableau d'objets **bodies** à un nouveau tableau **bodies_**
# +
bodies_ = np.array([objet() for i in range(10)])
import copy
bodies_ = copy.deepcopy(bodies) #clonage de bodies sans faire attribuer la meme addresse a bodies_
# -
# **Implémentation des trajectoires non couplées :**
# +
#----------------------------------------------------------------------------------------------------------
# Simuler les trajectoires à l'aide de Verlet sans couplage
#-------------------------
#Definition de pas de temps
dt = 1 #step
T = int(365/dt)*10 # (Nombre de steps)<-> Periode d'integration
#Definition des tableaux
for i in range(Nbr_obj):
bodies[i].x = np.zeros(T); bodies[i].x[0] = bodies[i].x0
bodies[i].y = np.zeros(T); bodies[i].y[0] = bodies[i].y0
bodies[i].z = np.zeros(T); bodies[i].z[0] = bodies[i].z0
bodies[i].vx = np.zeros(T); bodies[i].vx[0] = bodies[i].vx0
bodies[i].vy = np.zeros(T); bodies[i].vy[0] = bodies[i].vy0
bodies[i].vz = np.zeros(T); bodies[i].vz[0] = bodies[i].vz0
#Def des v_demi pour chaque objet
vx_demi = np.zeros(Nbr_obj)
vy_demi = np.zeros(Nbr_obj)
vz_demi = np.zeros(Nbr_obj)
#Implementation de l'integrateur de Verlet pour chaque objet (sauf le soleil)
for j in range(T-1):
#Phase 1: Calcul de vitesses milieu et affectation des position a l'intant j+1
for i in range(1,Nbr_obj): #Modification des parametres pour chaque objet a un instant donne
#Récupération de l'acceleration sous l'effet de soleil seulement
fx_j, fy_j, fz_j = acceleration_sol(bodies, i, j) #L'acceleration au pas j relative à l'objet i
#Definition des variables de milieux
vx_demi[i] = bodies[i].vx[j] + (dt/2)*fx_j
vy_demi[i] = bodies[i].vy[j] + (dt/2)*fy_j
vz_demi[i] = bodies[i].vz[j] + (dt/2)*fz_j
# Affectation des positions à l'indice i+1
bodies[i].x[j+1] = bodies[i].x[j] + dt*vx_demi[i]
bodies[i].y[j+1] = bodies[i].y[j] + dt*vy_demi[i]
bodies[i].z[j+1] = bodies[i].z[j] + dt*vz_demi[i]
#Phase 2: Affectation des vitesse a l'instant j+1
for i in range(1,Nbr_obj):
#L'acceleration au pas i+1 relative à l'objet j
fx_jplus1, fy_jplus1, fz_jplus1 = acceleration_sol(bodies, i, j+1) #Il faut faire cette étape après le calcul de postion à l'indice i+1
bodies[i].vx[j+1] = vx_demi[i] + (dt/2)*fx_jplus1
bodies[i].vy[j+1] = vy_demi[i] + (dt/2)*fy_jplus1
bodies[i].vz[j+1] = vz_demi[i] + (dt/2)*fz_jplus1
#[End]----------------------------------------------------------------------------------------------------
# -
# **Plot d'énergie mécanique des deux cas (sans couplage et avec couplage) :**
# +
#----------------------------------------------------------------------------------------------------------
# Plot de l'energie, cas: avec couplage et sans couplage
#-------------------------
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe
#Calcul de l'energie
#Cas sans couplage
Nrg = Energy(bodies, 1) #Calcul de l'energie d'un objet -> Changez le numero pour voir l'energie de chaque objet;
Nrg /= np.abs(Nrg[0]) #Pour Normaliser
#Cas avec couplage
Nrg_ = Energy(bodies_, 1) #Cacul de l'energie d'un objet -> Changez le numero pour voir l'energie de chaque objet;
Nrg_ /= np.abs(Nrg_[0]) #Pour Normaliser
#Plot de l'energie
t = np.linspace(1,T,T)*dt
ax.plot(t, Nrg, label= "Cas sans Couplage")
ax.plot(t, Nrg_, label= "Cas avec Couplage")
ax.set_xlabel("t (jour)")
ax.set_ylabel("E/$|E_0|$")
ax.get_yaxis().get_major_formatter().set_useOffset(False) #Disable scaling of values in plot wrt y-axis
#Affichage des résulats
print("Résultats : ")
print("Energie moyenne = " + str(np.mean(Nrg)) + ", Ecart_Type = " + str(np.std(Nrg)))
plt.legend()
plt.show()
# -
# En termes d'énergie, on ne voit pas une différence significative entre les énergies de chaque objet dans le cas de couplage ou sans couplage, et les deux énergies ne se dévient pas trop de la valeur initial $E_0$, alors à priori pour vérifier s'il y a une différence entres les trajectoires obtenues, on va comparer les trajectoires des deux approches aux trajectoires réelles obtenues par observations astrométriques.
# ## 2.3 Comparaison avec Trajectoires Réelles:
#
# Dans cette partie, on va comparer les trajectoires réelles, couplées et non couplées, en prenant toujours comme date initiale la date prise dans la partie 1 de ce rapport: "2017-02-28" à "00:00 GMT". On fera la comparaison pour Venus, Terre et Jupyter.
#
# PS: les trajectoires réelles sont prises ici: http://vo.imcce.fr/webservices/miriade/?forms
#
# ### 2.3.1 Venus:
# **Plot des trajectoires suivant x et y:**
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[2].x, bodies[2].y, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[2].x, bodies_[2].y, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
x, y = np.genfromtxt("real_simulation_venus.txt", usecols=(2,3), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier En_3dim
ax.plot(x, y, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("x (Au)")
ax.set_ylabel("y (Au)")
plt.legend()
plt.show()
# -
# **Plot des trajectoires suivant y et z:**
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[2].y, bodies[2].z, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[2].y, bodies_[2].z, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
y, z = np.genfromtxt("real_simulation_venus.txt", usecols=(3,4), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier In_3dim
ax.plot(y, z, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("y (Au)")
ax.set_ylabel("z (Au)")
plt.legend()
plt.show()
# -
# ### 2.3.2 Terre:
# **Plot des trajectoires suivant x et y:**
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[3].x, bodies[3].y, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[3].x, bodies_[3].y, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
x, y = np.genfromtxt("real_simulation_earth.txt", usecols=(2,3), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier In_3dim
ax.plot(x, y, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("x (Au)")
ax.set_ylabel("y (Au)")
plt.legend()
plt.show()
# -
# **Plot des trajectoires suivant y et z**:
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[3].y, bodies[3].z, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[3].y, bodies_[3].z, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
y, z = np.genfromtxt("real_simulation_earth.txt", usecols=(3,4), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier In_3dim
ax.plot(y, z, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("y (Au)")
ax.set_ylabel("z (Au)")
plt.legend()
plt.show()
# -
# ### 2.3.2 Jupyter:
# **Plot des trajectoires suivant x et y**:
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[5].x, bodies[5].y, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[5].x, bodies_[5].y, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
x, y = np.genfromtxt("real_simulation_jupyter.txt", usecols=(2,3), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier In_3dim
ax.plot(x, y, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("x (Au)")
ax.set_ylabel("y (Au)")
plt.legend()
plt.show()
# -
# **Plot des trajectoires suivant y et z**:
# +
#Definition de figure
fig=plt.figure(figsize=(9, 6), dpi= 100, facecolor='w', edgecolor='k') #To modify the size of the figure
ax = fig.add_subplot(111) #definition de l'axe en 3d
#plt.gca().set_aspect('equal', adjustable='box') #equal ratios of x and y
ax.plot(bodies[5].y, bodies[5].z, label = "Trajectoire Non Couplée") #Cas sans couplage
ax.plot(bodies_[5].y, bodies_[5].z, label = "Trajectoire Couplée") #Cas couplage
#Import de la trajectoire de Jupyter
y, z = np.genfromtxt("real_simulation_jupyter.txt", usecols=(3,4), skip_header=1, unpack=True, delimiter=",") #Fichier dans le meme dossier In_3dim
ax.plot(y, z, label="Trajectoire Réelle") #Traj réelle
ax.set_xlabel("y (Au)")
ax.set_ylabel("z (Au)")
plt.legend()
plt.show()
# -
# D'après les comparaisons entre les différentes trajectoires (on pourra aussi tester la précision des trajectoires pour les autres planètes), les trajectoires sans couplages reproduit bien les trajectoires réelles, alors on peut se limiter aux trajectoires sans couplage comme trajectoires de référence afin d'accélerer les calculs, cette approximation sera largement suffisante si on se limite à des durées d'integration relativement courtes à l'échelle astronomique comme la période de voyage d'une sonde interplanétaire, donc dans la troisième partie, on pourra travailler avec des trajectoires non couplées afin de déterminer la trajectoire des sondes spatiales.
| Solar System in 3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Slicer 4.11
# language: python
# name: slicer-4.11
# ---
# +
import JupyterNotebooksLib as slicernb
this_notebook_name = "SpineSegmentationVisualTestFunctions"
# +
# place paths to scenes to test here
scenes_to_reconstruct = [
r"E:\Perk\Summer 2021\Scenes\Q001_04_tu_Segmented.mrb",
r"E:\Perk\Summer 2021\Scenes\Q002_04_tu_Segmented.mrb",
r"E:\Perk\Summer 2021\Scenes\Q003_04_tu_Segmented.mrb",
r"E:\Perk\Summer 2021\Scenes\Q004_04_tu_Segmented.mrb"
]
# all paths to models to study
models = [
r"E:\Perk\Summer 2021\Models\model_2019-05-31_21-22-03.h5",
r"E:\Perk\Summer 2021\Models\model_2019-06-01_13-47-36.h5"
]
# set to true to save reconstructed images as pngs
store_images = False
# folder to store images
image_output_path = r'E:\Perk\Summer 2021\SavedImages\Image_{}.png'
# folder to store AI segmentation as nrrd files
#volume_output_path = r'E:\Perk\Summer 2021\NotebookOutput\Volume_{}.nrrd'
# set to true to mute all diagnostic output except images
mute_output = True
# output segmentations as arrays
array_output = False
# Input ultrasound sequence names
input_browser_name = r"spine001"
input_image_name = r"Image_Image"
# Output will be saved using these names
output_browser_name = r"BonePredictionBrowser"
output_sequence_name = r"PredictionSequence"
output_image_name = r"PredictionImage"
# Save output volume(s) as nrrd files
volume_output = True
volume_output_path = r"E:\Perk\Summer 2021\NotebookOutput\Scene_{}_Model_{}.nrrd"
# Image processing parameters
# Erases the side of prediction images. 1.0 means the whole prediction is erased.
# Background should be the first component (i.e. y[:,:,:,0]) in the prediction output array.
clip_side_ratio = 0.3
apply_logarithmic_transformation = True
logarithmic_transformation_decimals = 4
# notebook output path
notebooks_save_path = r"E:\Perk\Summer 2021\SavedNotebooks"
# Volume rendering parameters
volRendWindow = 200
volRendLevel = 156
volRendColor1 = [0.1, 0.1, 0.0, 1]
volRendColor2 = [1.0, 0.9, 0.4, 1]
import datetime
save_timestamp = datetime.datetime.now().strftime('%Y-%m-%d_%H-%M-%S')
print("Save timestamp: {}".format(save_timestamp))
# +
import datetime
import os
import scipy.ndimage
import qt
import numpy as np
import tensorflow as tf
from tensorflow.keras.models import load_model
# from local_vars import root_folder
# +
parent_folder = os.path.dirname(os.path.abspath(os.curdir))
sys.path.append(parent_folder)
import Models.segmentation_unet as unet
import utils
# -
def setup(scene):
# loading scene
if not mute_output:
print("Loading scene from " + scene)
slicer.mrmlScene.Clear()
try:
slicer.util.loadScene(scene)
except:
# NOTE: for some reason loading a scene throws an error every time, but laods the scene just fine
if not mute_output:
print('Error thrown. Continuing.')
# changing transform hierarchy
image_image = slicer.util.getFirstNodeByName(input_image_name)
imageToTransd = slicer.util.getFirstNodeByName("ImageToReference")
image_image.SetAndObserveTransformNodeID(None)
image_image.SetAndObserveTransformNodeID(imageToTransd.GetID())
slicer.app.layoutManager().setLayout(slicer.vtkMRMLLayoutNode.SlicerLayoutOneUp3DView)
#hide skeleton model
skeleton = slicer.util.getFirstNodeByName("SkeletonModel")
if skeleton is not None:
skeleton.SetDisplayVisibility(False)
def load_model(model_fullname):
# Check if keras model file exists. Abort if not found. Load model otherwise.
if not os.path.exists(model_fullname):
raise Exception("Could not find model: " + model_fullname)
if not mute_output:
print("Loading model from: " + model_fullname)
model = tf.keras.models.load_model(model_fullname, compile=False)
return model
# model.summary()
def segment(model):
# Check input. Abort if browser or image doesn't exist.
#input_browser_node = slicer.util.getFirstNodeByName(input_browser_name, className='vtkMRMLSequenceBrowserNode')
input_browser_node = slicer.util.getNodesByClass(className='vtkMRMLSequenceBrowserNode')[0]
input_image_node = slicer.util.getFirstNodeByName(input_image_name, className="vtkMRMLScalarVolumeNode")
if input_browser_node is None:
logging.error("Could not find input browser node: {}".format(input_browser_node))
raise
if input_image_node is None:
logging.error("Could not find input image node: {}".format(input_image_name))
raise
# Create output image and browser for segmentation output.
output_browser_node = slicer.util.getFirstNodeByName(output_browser_name, className='vtkMRMLSequenceBrowserNode')
if output_browser_node is None:
output_browser_node = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceBrowserNode', output_browser_name)
output_sequence_node = slicer.util.getFirstNodeByName(output_sequence_name, className="vtkMRMLSequenceNode")
if output_sequence_node is None:
output_sequence_node = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceNode', output_sequence_name)
output_browser_node.AddSynchronizedSequenceNode(output_sequence_node)
output_image_node = slicer.util.getFirstNodeByName(output_image_name, className="vtkMRMLScalarVolumeNode")
# browser_logic = slicer.modules.sequences.logic()
if output_image_node is None:
volumes_logic = slicer.modules.volumes.logic()
output_image_node = volumes_logic.CloneVolume(slicer.mrmlScene, input_image_node, output_image_name)
browser_logic = slicer.modules.sequences.logic()
browser_logic.AddSynchronizedNode(output_sequence_node, output_image_node, output_browser_node)
output_browser_node.SetRecording(output_sequence_node, True)
# Add all input sequences to the output browser for being able to conveniently replay everything
proxy_collection = vtk.vtkCollection()
input_browser_node.GetAllProxyNodes(proxy_collection)
for i in range(proxy_collection.GetNumberOfItems()):
proxy_node = proxy_collection.GetItemAsObject(i)
output_sequence = slicer.mrmlScene.AddNewNodeByClass('vtkMRMLSequenceNode')
browser_logic.AddSynchronizedNode(output_sequence, proxy_node, output_browser_node)
output_browser_node.SetRecording(output_sequence, True)
# Iterate input sequence, compute segmentation for each frame, record output sequence.
num_items = input_browser_node.GetNumberOfItems()
n = num_items
input_browser_node.SelectFirstItem()
input_array = slicer.util.array(input_image_node.GetID())
slicer_to_model_scaling = model.layers[0].input_shape[0][1] / input_array.shape[1]
model_to_slicer_scaling = input_array.shape[1] / model.layers[0].input_shape[0][1]
if not mute_output:
print("Will segment {} images".format(n))
if array_output:
array_output_ultrasound = np.zeros((n, input_array.shape[1], input_array.shape[1]))
array_output_segmentation = np.zeros((n, input_array.shape[1], input_array.shape[1]), dtype=np.uint8)
model_output_size = model.layers[-1].output_shape[1]
num_output_components = model.layers[-1].output_shape[3]
mask_model = np.ones([model_output_size, model_output_size])
mask_model_background = np.zeros([model_output_size, model_output_size])
columns_to_mask = int(model_output_size / 2 * clip_side_ratio)
if not mute_output:
print("Will mask {} columns on both sides".format(columns_to_mask))
mask_model[:,:columns_to_mask] = 0
mask_model[:,-columns_to_mask:] = 0
mask_model_background[:,:columns_to_mask] = 1
mask_model_background[:,-columns_to_mask:] = 1
# Display mask
# import matplotlib
# matplotlib.use('WXAgg')
# from matplotlib import pyplot as plt
# plt.imshow(mask_model[:,:])
# plt.show()
start_timestamp = datetime.datetime.now()
if not mute_output:
print("Processing started at: {}".format(start_timestamp.strftime('%H-%M-%S')))
for i in range(n):
# if i > 10: # todo Just for debugging
# break
input_array = slicer.util.array(input_image_node.GetID())
if array_output:
array_output_ultrasound[i, :, :] = input_array[0, :, :]
resized_input_array = scipy.ndimage.zoom(input_array[0,:,:], slicer_to_model_scaling)
resized_input_array = np.flip(resized_input_array, axis=0)
resized_input_array = resized_input_array / resized_input_array.max() # Scaling intensity to 0-1
resized_input_array = np.expand_dims(resized_input_array, axis=0)
resized_input_array = np.expand_dims(resized_input_array, axis=3)
y = model.predict(resized_input_array)
if apply_logarithmic_transformation:
e = logarithmic_transformation_decimals
y = np.log10(np.clip(y, 10**(-e), 1.0)*(10**e))/e
y[0,:,:,:] = np.flip(y[0,:,:,:], axis=0)
for component in range(1, num_output_components):
y[0,:,:,component] = y[0,:,:,component] * mask_model[:,:]
y[0,:,:,0] = np.maximum(y[0,:,:,0], mask_model_background)
upscaled_output_array = scipy.ndimage.zoom(y[0,:,:,1], model_to_slicer_scaling)
upscaled_output_array = upscaled_output_array * 255
upscaled_output_array = np.clip(upscaled_output_array, 0, 255)
if array_output:
array_output_segmentation[i, :, :] = upscaled_output_array[:, :].astype(np.uint8)
# output_array = slicer.util.array(output_image_node.GetID())
# output_array[0, :, :] = upscaled_output_array[:, :].astype(np.uint8)
slicer.util.updateVolumeFromArray(output_image_node, upscaled_output_array.astype(np.uint8)[np.newaxis, ...])
output_browser_node.SaveProxyNodesState()
input_browser_node.SelectNextItem()
# If Slicer crashes during processing, try commenting this following line out and run this notebook again.
slicer.app.processEvents()
stop_timestamp = datetime.datetime.now()
if not mute_output:
print("Processing finished at: {}".format(stop_timestamp.strftime('%H-%M-%S')))
time_seconds = (stop_timestamp - start_timestamp).total_seconds()
if not mute_output:
print("Processed {} frames in {:.2f} seconds".format(n, time_seconds))
print("FPS = {:.2f}".format(n / time_seconds))
def reconstruct_volume():
# Find input volume (image) for volume reconstruction
predictionVolume = slicer.mrmlScene.GetFirstNodeByName("PredictionImage")
if predictionVolume is None:
raise Exception("PredictionImage not found in Slicer scene")
# Create output volume node
reconstructedVolume = slicer.mrmlScene.GetFirstNodeByName("AiVolume")
if reconstructedVolume is None:
reconstructedVolume = slicer.mrmlScene.AddNewNodeByClass("vtkMRMLScalarVolumeNode")
reconstructedVolume.SetName("AiVolume")
inputBrowserNode = slicer.mrmlScene.GetFirstNodeByName("BonePredictionBrowser")
if inputBrowserNode is None:
raise Exception("BonePredictionBrowser missing")
# Prepare volume reconstructor node
volumeReconstructor = slicer.mrmlScene.GetFirstNodeByName("AiVolumeReconstructor")
if volumeReconstructor is None:
volumeReconstructor = slicer.vtkMRMLVolumeReconstructionNode()
volumeReconstructor.SetName("AiVolumeReconstructor")
volumeReconstructor.SetLiveVolumeReconstruction(False)
volumeReconstructor.SetOptimizationMode(slicer.vtkMRMLVolumeReconstructionNode.FULL_OPTIMIZATION)
volumeReconstructor.SetCompoundingMode(slicer.vtkMRMLVolumeReconstructionNode.MAXIMUM_COMPOUNDING_MODE)
volumeReconstructor.SetInterpolationMode(slicer.vtkMRMLVolumeReconstructionNode.LINEAR_INTERPOLATION)
slicer.mrmlScene.AddNode(volumeReconstructor)
volumeReconstructor.SetAndObserveInputSequenceBrowserNode(inputBrowserNode)
volumeReconstructor.SetAndObserveOutputVolumeNode(reconstructedVolume)
volumeReconstructor.SetAndObserveInputVolumeNode(predictionVolume)
volumeReconstructionLogic = slicer.modules.volumereconstruction.logic()
# Volume reconstruction
volumeReconstructionLogic.ReconstructVolumeFromSequence(volumeReconstructor)
def render_volume():
# Volume rendering
# find input volume
aivolumeNode = slicer.mrmlScene.GetFirstNodeByName("AiVolume")
if aivolumeNode is None:
raise Exception("AiVolume node was never constructed")
# find or build ROI
annotationROINode = slicer.mrmlScene.GetFirstNodeByName("AnnotationROI")
if annotationROINode is None:
annotationROINode = slicer.vtkMRMLAnnotationROINode()
annotationROINode.SetName("AnnotationROI")
slicer.mrmlScene.AddNode(annotationROINode)
# annotationROINode.SetDisplayVisibility(False)
# find or build volume property
propertyPresetNode = slicer.mrmlScene.GetFirstNodeByName("volMR-Default")
if propertyPresetNode is None:
propertyPresetNode = slicer.vtkMRMLVolumePropertyNode()
propertyPresetNode.SetName("volMR-Default")
volumeRenderingLogic = slicer.modules.volumerendering.logic()
propertyPresetNode.Copy(volumeRenderingLogic.GetPresetByName('MR-Default'))
slicer.mrmlScene.AddNode(propertyPresetNode)
# build 3D renderer
volumeRenderingLogic = slicer.modules.volumerendering.logic()
displayNode = volumeRenderingLogic.GetFirstVolumeRenderingDisplayNode(aivolumeNode)
if displayNode is None:
displayNode = slicer.vtkMRMLGPURayCastVolumeRenderingDisplayNode()
displayNode.SetName("AiVolumeRenderer")
slicer.mrmlScene.AddNode(displayNode)
aivolumeNode.AddAndObserveDisplayNodeID(displayNode.GetID())
displayNode.SetAndObserveVolumePropertyNodeID(propertyPresetNode.GetID())
displayNode.SetAndObserveROINodeID(annotationROINode.GetID())
# Set up custom volume rendering parameters
lut = vtk.vtkLookupTable()
lut.SetNumberOfTableValues(2)
lut.Build()
lut.SetTableValue(0, volRendColor1)
lut.SetTableValue(1, volRendColor2)
volumeProperty = displayNode.GetVolumePropertyNode().GetVolumeProperty()
volumeRenderingLogic.SetThresholdToVolumeProp(
[0, 500],
[volRendLevel - volRendWindow / 2.0, volRendLevel + volRendWindow / 2.0],
volumeProperty,
True,
True)
upper = min(255, volRendLevel + volRendWindow / 2)
lower = max(0, volRendLevel - volRendWindow / 2)
volumeRenderingLogic.SetWindowLevelToVolumeProp(
[0, 255],
[upper - lower, lower + (upper - lower) / 2],
lut,
volumeProperty)
displayNode.SetVisibility(True)
# clean up
# Hide all ROI nodes
roiCollection = slicer.mrmlScene.GetNodesByClass('vtkMRMLAnnotationROINode')
for i in range(roiCollection.GetNumberOfItems()):
roiNode = roiCollection.GetItemAsObject(i)
roiNode.SetDisplayVisibility(False)
# reseting 3d camera
"""
threeDWidget = slicer.app.layoutManager().threeDWidget(0)
threeDView = threeDWidget.threeDView()
threeDView.resetFocalPoint()
"""
# hide ultrasound scan image
# TODO: make this a global variable? it is defined in two seperate functions
input_image_node = slicer.util.getFirstNodeByName(input_image_name, className="vtkMRMLScalarVolumeNode")
input_image_node.SetDisplayVisibility(False)
# +
sceneCount = 0
for scene in scenes_to_reconstruct:
modelCount = 0
for model in models:
setup(scene)
segment(load_model(model))
reconstruct_volume()
render_volume()
display(slicernb.ViewDisplay())
if store_images:
numbered_image_path = image_output_path.format(str(sceneCount))
print("Saving image at " + numbered_image_path)
renderWindow = slicer.app.layoutManager().threeDWidget(0).threeDView().renderWindow()
renderWindow.SetAlphaBitPlanes(1)
wti = vtk.vtkWindowToImageFilter()
wti.SetInputBufferTypeToRGBA()
wti.SetInput(renderWindow)
writer = vtk.vtkPNGWriter()
writer.SetFileName(numbered_image_path)
writer.SetInputConnection(wti.GetOutputPort())
writer.Write()
# volume output
if volume_output:
numbered_volume_path = volume_output_path.format(str(sceneCount),str(modelCount))
output_volume_node = slicer.util.getFirstNodeByName('AiVolume', className="vtkMRMLScalarVolumeNode");
slicer.util.saveNode(output_volume_node, numbered_volume_path)
modelCount += 1
sceneCount += 1
# +
# Save notebook so all output is archived by the next cell
from IPython.display import Javascript
script = '''
require(["base/js/namespace"],function(Jupyter) {
Jupyter.notebook.save_checkpoint();
});
'''
Javascript(script)
# +
import nbformat
from nbconvert import HTMLExporter
import json
notebook_path = slicernb.notebookPath()
with open(notebook_path, mode="r") as f:
file_json = json.load(f)
notebook_content = nbformat.reads(json.dumps(file_json), as_version=4)
html_exporter = HTMLExporter()
(body, resources) = html_exporter.from_notebook_node(notebook_content)
this_notebook_name = os.path.splitext(os.path.basename(notebook_path))[0]
save_file_name = this_notebook_name + "_" + save_timestamp + ".html"
notebook_fullpath = os.path.join(notebooks_save_path, save_file_name)
f = open(notebook_fullpath, 'wb')
f.write(body.encode())
f.close()
print("Notebook saved to: {}".format(notebook_fullpath))
| UltrasoundSegmentation/SpineSegmentationVisualTestFunctions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Matplotlib Assignment
# import pandas
import pandas as pd
import numpy as np
# import matplotlib
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (10,6)
df=pd.read_csv('company_sales_data.csv')
df
# ### Task 1: Read the `total_profit` of all months and display it using a line plot.
# 200 values from the interval <0,100>, equidistantly divided
# a line plot
plt.plot(df["month_number"],df["total_profit"],'red')
plt.show()
# ### Task 2: Read all the different product sales data and display it using a multiline plot.
plt.plot(df["month_number"], df["facecream"], label='facecream')
plt.plot(df["month_number"], df["facewash"], label='facewash')
plt.plot(df["month_number"], df["toothpaste"], label='toothpaste')
plt.plot(df["month_number"], df["bathingsoap"], label='bathingsoap')
plt.plot(df["month_number"], df["shampoo"], label='shampoo')
plt.plot(df["month_number"], df["moisturizer"], label='moisturizer')
plt.xlabel('Month')
plt.ylabel('Sales')
plt.title("Sales")
plt.legend()
plt.show()
# ### Task 3: Read the `total_profit` of each month and display it using a histogram to find out which profit ranges are the most common.
fig, ax = plt.subplots(tight_layout=True)
hist = ax.hist2d(df["month_number"], df["total_profit"])
| w3/w3-day_1/matplotlib_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Custom Image Data
# +
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
import torch.optim as optim
import os
import torchvision
from torch.utils.data import Dataset,DataLoader,random_split
from skimage import io
import torchvision.transforms as transforms
# -
| DataLoader_ImageFolder/.ipynb_checkpoints/custom_image_data_leaf_classification-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# # A deep dive into DataFrames.jl indexing
# # Part 1: indexing in DataFrames.jl by example
# ### <NAME>
# What are we going to cover:
# * `getindex`, a.k.a. `x[...]`
# * `setindex!`, a.k.a. `x[...] =`
# * `broadcast`, a.k.a. `fun.(x)`
# * `broadcat!`, a.k.a. `x .= ...`
#
# Indexable types that DataFrames.jl defines:
# * `DataFrame`
# * `SubDataFrame`
# * `DataFrameRow`
# * `DataFrameRows`
# * `DataFrameColumns`
# * `GroupedDataFrame`
# * `GroupKeys`
# * `GroupKey`
# * `StackedVector`
# * `RepeatedVector`
# ### Environment setup
using DataFrames
using CSV
using BenchmarkTools
using Dates
using Statistics
ENV["COLUMNS"] = 500 # allow output up to 500 characters wide not to be truncated when displayed
ENV["LINES"] = 15 # we do not need to see too many lines in the examples we work with
df = CSV.File("too-big/fh_5yrs.csv") |> DataFrame
# #### Warm up exercises
# *Get short description of columns in our data frame*
describe(df)
# (see https://github.com/JuliaData/DataFrames.jl/issues/2269 for a discussion of the design decisions here, feel free to comment there if you have an opinion)
# *Get information about exact types of the columns stored in the data frame*
# *Get names of columns as strings*
# *Get names of columns as `Symbol`s*
# ## `getindex`
# Get a single column as a whole without copying
unique([df.date,
df."date",
df[!, 1],
df[!, :date],
df[!, "date"]])
unique([getproperty(df, :date),
getproperty(df, "date"),
getindex(df, !, 1),
getindex(df, !, :date),
getindex(df,!, "date")])
# Get a single column as a whole with copying
unique([copy(df.date),
copy(df."date"),
df[:, 1],
df[:, :date],
df[:, "date"]])
# Let us compare the performance of various ways to get a column without copying
@btime $df.date
@btime $df."date"
@btime $df[!, 1]
@btime $df[!, :date]
@btime $df[!, "date"];
# #### Exercise
# Check the same but with copying
# Do you think it really matters in practice how fast is an access to column of a data frame?
# Let us check how lookup speed scales with the number of columns:
@time df_tmp = DataFrame(ones(1, 100_000))
@btime $df_tmp.x100000
@btime $df_tmp."x100000"
@btime $df_tmp[!, 100000];
# Get a single column, but take a subset of rows: you can either make a copy or get a view
df[1:2, :date]
view(df, 1:2, :date)
# this is the same as:
df.date[1:2]
view(df.date, 1:2)
# you can use `Not` for inverted selection
df[Not(3:end), :date]
# Get a single cell in a data frame: you can either get a value or a view
df[1, :date]
view(df, 1, :date)
# #### Exercise
# In what case you might want to use a view instead of getting a value?
# Check what is the consequence of running the following lines:
tmp_cell = view(df, 1, :date)
tmp_cell2 = getindex(df, 1, :date)
tmp_cell[] = Date("2222-07-02")
# Revert the change we have just made
# To conclude note that with `view` there is not difference between `!` and `:`:
@view df[!, 1]
@view df[:, 1]
# Summary:
#
# > passing a single column as an integer, `Symbol` or string drops one dimension of
# > a data frame and allows you to select or subset a column from it
# Multiple column selection options:
# * a vector of `Symbol` (does not have to be a subtype of `AbstractVector{Symbol}`);
# * a vector of `AbstractString` (does not have to be a subtype of `AbstractVector{<:AbstractString}`);
# * a vector of `Integer` other than `Bool` (does not have to be a subtype of `AbstractVector{<:Integer}`);
# * a vector of `Bool` that has to be a subtype of `AbstractVector{Bool}`;
# * a regular expression, which gets expanded to a vector of matching column names;
# * a `Not` expression;
# * an `All` or `Between` expression;
# * a colon literal :.
# The type of output depends on the row selecor:
# * if it is a single row you get a `DataFrameRow` (a dimension is dropped)
# * if it is a collection of rows you get a data frame
# Single row selection is always a view that is `DataFrameRow`:
df[1, [:date]]
@view df[1, [:date]]
# Multiple row selection is a `DataFrame` for `getindex`:
df[1:2, 1:2]
df2 = df[!, 1:2] # this does not copy columns
df2.date === df.date
# Using `view` creates a `SubDataFrame`
df3 = view(df, 1:2, 1:2)
typeof(df3)
# For `view` using `:` and `!` gives you the same result:
dump(view(df, !, :))
dump(view(df, :, :))
# Normally when you modify the parent of a `SubDataFrame` (or `DataFrameRow`) you may get an error when trying to access it:
df4 = DataFrame(reshape(1:12, 3, 4))
df4_v = view(df4, 1:2, 1:3)
select!(df4, 1)
df4_v
# A special case is when you use `:` as a column selection with a `view`. In this case the `SubDataFrame` and `DataFrameRow` always get updated with the changed columns:
df4 = DataFrame(reshape(1:12, 3, 4))
df4_v = view(df4, 1:2, :)
select!(df4, 1, :x2 => :newcol)
df4_v
# The reason for this behavior is that subsetting of a data frame by only rows (and taking all columns) is very common, and in this case we can create and index such views much faster. In particular `DataFrameRow`s produced by `eachrow` are efficient this way:
@btime mean(x -> x.open, eachrow(df))
@btime mean(i -> df[i, :open], 1:nrow(df))
# Of course, type-stable operation would be faster (but sometimes processing data row-wise is more convenient):
@btime mean(df.open)
# or, if your table is not very wide (so that you are not penalized by the compilation cost of `NamedTuple`) you can use:
@btime mean(x -> x.open, Tables.namedtupleiterator(df))
# Note though that `DataFrameRow` allows you to modify the source data frame, while iterating `NamedTuple`s is read-only (more on `setindex!` later).
df5 = copy(df)
# #### Exercise
# In `df5` find rows in which `:high` is less than `:low` and swap them.
# I give you the following column selectors. Can you tell the effect of each of them when trying to run `df[1:2, selector]`?
# Write the code that tests it.
selectors = [Between(1, 10), Between(:low, :high), [:low, :low], All(:low, :low), All(:low, :), All()]
# ### Indexing `GroupedDataFrame`
# A `GroupedDataFrame` is a view into a data frame which defines a key allowing a fast lookup (and in particular this key is then automatically used in split-apply-combine operations with `select`, `select!`, `transform`, `transform!` and `combine`).
gdf = groupby(df, :symbol)
gdf_keys = keys(gdf)
# As usual - indexing by a single value drops a dimension (you get a `SubDataFrame`)
gdf[1]
gdf_keys[1]
gdf[gdf_keys[1]]
gdf[(symbol="AAAU",)]
gdf[("AAAU",)]
# And indexing by a collection produces a subsetted `GroupedDataFrame`:
gdf[1:2]
gdf[tuple.(["AAAU", "AACG"])]
# ## setindex!
| indexing_part1_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import glob
import gensim
from sklearn.cluster import KMeans
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn import metrics
import tqdm
import nltk
import re
from operator import itemgetter
import time
from pprint import pprint
from bertopic import BERTopic
EPOCHS = 205
TOPICS = 5
CHUNK_SIZE = 1000
WORKERS = 7
EVAL_PERIOD = 10
ALPHA = 0.01
BETA = 0.9
# -
nltk.download('punkt')
nltk.download('wordnet')
nltk.download('stopwords')
nltk.download('omw-1.4')
def preprocess_text(sentence):
# Lowercase
sentence = sentence.lower()
# Remove all non-alphabets (punctuation, numbers, new-line characters and extra-spaces)
sentence = re.sub(r'[^a-zA-Z]+', ' ', sentence)
sentence = sentence.replace('\n', '')
# sentence = re.sub('\s\s+', ' ', sentence)
# Tokenize & remove stop-words
word_list = nltk.word_tokenize(sentence)
stopwords_list = nltk.corpus.stopwords.words('english')
stopwords_list.extend(['trump','realdonaldtrump','thank','trump','presid','america','american','fjv'])
word_list = [word for word in word_list if word not in stopwords_list]
# Remove very small words, length < 3, they don't contribute any useful information
word_list = [word for word in word_list if len(word) > 3]
# Stem & Lemmatize
porter_stemmer = nltk.stem.PorterStemmer()
lemmatizer = nltk.stem.WordNetLemmatizer()
word_list = [porter_stemmer.stem(word) for word in word_list]
word_list = [lemmatizer.lemmatize(word) for word in word_list]
sentence = ' '.join(word_list)
return sentence
# ## Load Data
# +
path = '../../data/twitter/raw/users/'
republicans_df = pd.concat([pd.read_csv(f) for f in glob.glob(path+'republicans/required/*.csv')])
democrats_df = pd.concat([pd.read_csv(f) for f in glob.glob(path+'democrats/required/*.csv')])
df = pd.concat([republicans_df, democrats_df], ignore_index=True)
# -
tqdm.tqdm.pandas()
df['tweet_tokenized'] = df['tweet'].progress_apply(lambda x:preprocess_text(str(x)))
performance_metrics = pd.DataFrame(columns=['feature-extraction','clustering-algo', 'run#', 'state', 'c_v','c_umass','topics','time'])
# ## TF-IDF
# +
documents = df['tweet_tokenized'].str.split()
dictionary = gensim.corpora.Dictionary(documents)
dictionary.filter_extremes(no_below=5, no_above=0.5, keep_n=20000)
tfidf_model = gensim.models.TfidfModel(dictionary=dictionary)
corpus = [dictionary.doc2bow(document) for document in documents]
corpus_tfidf = list(tfidf_model[corpus])
# -
# ### LDA
for run, state in zip(range(2, 6, 1), range(4, 12, 2)):
print('Run #', run)
start = time.time()
gensim_lda = gensim.models.ldamodel.LdaModel(corpus=corpus_tfidf, num_topics=TOPICS, id2word=dictionary, chunksize=CHUNK_SIZE, passes=EPOCHS,
eval_every = EVAL_PERIOD, per_word_topics=True, random_state=state, alpha=ALPHA, eta=BETA)
coherence_cv = gensim.models.CoherenceModel(model=gensim_lda, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
coherence_cumass = gensim.models.CoherenceModel(model=gensim_lda, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
topics = gensim_lda.print_topics()
stop = time.time()
performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'LDA', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
'topics':topics,'time':(stop-start)}, ignore_index=True)
# ### Parallel LDA
for run, state in zip(range(1, 6, 1), range(2, 12, 2)):
print('Run #', run)
start = time.time()
gensim_plda = gensim.models.ldamulticore.LdaMulticore(corpus=corpus_tfidf, num_topics=TOPICS, id2word=dictionary, chunksize=CHUNK_SIZE, workers=WORKERS, passes=EPOCHS,
eval_every = EVAL_PERIOD, per_word_topics=True, random_state=state, alpha=ALPHA, eta=BETA)
coherence_cv = gensim.models.CoherenceModel(model=gensim_plda, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
coherence_cumass = gensim.models.CoherenceModel(model=gensim_plda, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
topics = gensim_plda.print_topics()
stop = time.time()
performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'Parallel LDA', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
'topics':topics,'time':(stop-start)}, ignore_index=True)
# ### NMF
for run, state in zip(range(1, 6, 1), range(2, 12, 2)):
print('Run #', run)
start = time.time()
gensim_nmf = gensim.models.Nmf(corpus=corpus_tfidf, num_topics=TOPICS, id2word=dictionary, chunksize=CHUNK_SIZE, passes=EPOCHS, eval_every=EVAL_PERIOD, minimum_probability=0,
random_state=state, kappa=1)
coherence_cv = gensim.models.CoherenceModel(model=gensim_nmf, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
coherence_cumass = gensim.models.CoherenceModel(model=gensim_nmf, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
topics = gensim_nmf.print_topics()
stop = time.time()
performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'NMF', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
'topics':topics,'time':(stop-start)}, ignore_index=True)
# ### LSI
for run, state in zip(range(1, 6, 1), range(2, 12, 2)):
print('Run #', run)
start = time.time()
gensim_lsi = gensim.models.LsiModel(corpus=corpus_tfidf, num_topics=TOPICS, id2word=dictionary, chunksize=CHUNK_SIZE)
coherence_cv = gensim.models.CoherenceModel(model=gensim_lsi, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
coherence_cumass = gensim.models.CoherenceModel(model=gensim_lsi, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
topics = gensim_lsi.print_topics()
stop = time.time()
performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'LSI', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
'topics':topics,'time':(stop-start)}, ignore_index=True)
# ### BERTopic
# +
# for run, state in zip(range(1, 6, 1), range(2, 12, 2)):
# print('Run #', run)
# start = time.time()
# tfidf_embeddings = TfidfVectorizer(min_df=5).fit_transform(df['tweet_tokenized'].astype(str))
# topic_model = BERTopic(verbose=True, nr_topics=5)
# topics, probs = topic_model.fit_transform(df['tweet_tokenized'].astype(str), tfidf_embeddings)
# coherence_cv = gensim.models.CoherenceModel(model=topic_model, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
# coherence_cumass = gensim.models.CoherenceModel(model=topic_model, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
# topics = (topic_model.get_topic_info()[1:]['Name']).tolist()
# # print(topics)
# stop = time.time()
# performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'BERTopic', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
# 'topics':topics,'time':(stop-start)}, ignore_index=True)
# -
# ### HDP
for run, state in zip(range(1, 6, 1), range(2, 12, 2)):
print('Run #', run)
start = time.time()
gensim_hdp = gensim.models.hdpmodel.HdpModel(corpus=corpus_tfidf, id2word=dictionary, chunksize=CHUNK_SIZE, random_state=state, kappa=1, alpha=ALPHA)
coherence_cv = gensim.models.CoherenceModel(model=gensim_hdp, texts=documents, dictionary=dictionary, coherence='c_v').get_coherence()
coherence_cumass = gensim.models.CoherenceModel(model=gensim_hdp, texts=documents, dictionary=dictionary, coherence='u_mass').get_coherence()
topics = gensim_hdp.print_topics()
stop = time.time()
performance_metrics = performance_metrics.append({'feature-extraction':'tf-idf', 'clustering-algo':'HDP', 'run#':run, 'state':state,'c_v':coherence_cv,'c_umass':coherence_cumass,
'topics':topics,'time':(stop-start)}, ignore_index=True)
performance_metrics
mean_perf = performance_metrics.groupby('clustering-algo')[['c_v','c_umass','time']].mean()
mean_perf.to_csv('../../results/topic-modelling/mean-perf-topic-modelling.csv')
| code/topic-wise analysis/topic-modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jbell1991/profanity-filter-solving-scunthorpe-problem/blob/master/Profanity_Filter_and_Scunthorpe_Problem.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="oey1YuVbc3K2" colab_type="text"
# # The Problem to be Solved
#
# For my Lambda Labs project, I was tasked with creating a profanity filter for a children's reading app called Story Squad. Story Squad prompts kids to read a new story or chapter of a book every week. They then write their own creative story and draw a picture that branches off what they read. The stories are handwritten to promote creativity from the students. Their handwriting is read by the [Google Cloud Vision API](https://https://cloud.google.com/vision/?utm_source=google&utm_medium=cpc&utm_campaign=na-US-all-en-dr-bkws-all-all-trial-e-dr-1009135&utm_content=text-ad-none-any-DEV_c-CRE_291249276628-ADGP_Hybrid+%7C+AW+SEM+%7C+BKWS+%7C+US+%7C+en+%7C+EXA+~+ML/AI+~+Vision+API+~+Google+Cloud+Vision+Api-KWID_43700036257547156-kwd-475108777569&utm_term=KW_google%20cloud%20vision%20api-ST_Google+Cloud+Vision+Api&gclid=EAIaIQobChMI9PTkyvaB6wIVGey1Ch1p4gqpEAAYASAAEgJwjfD_BwE), which converts handwritten letters to typed text.
#
# Kids will be kids and from time to time inappropriate language might seep into the user experience. Parents want to be able to trust their children are safe using the app. However, we don't want to deny students entry if their story was falsely flagged for profanity. All stories are reviewed by human eyes to comply with the [Children's Online Privacy Protection Rule](https://www.ftc.gov/enforcement/rules/rulemaking-regulatory-reform-proceedings/childrens-online-privacy-protection-rule) ("COPPA"), but we could prioritize stories to be reviewed by a moderator by flagging ones with possible profane words. If a story is flagged, a moderator will review it before others and see if the flag is a true or false positive.
#
# # Options
#
# One option explored was uing python packages to find profanity. Profanity-filter worked well on most words, but still missed some individual words, missed phrases and did well to avoid flagging Scunthorpe like words.
#
# Another option was a package called profanity check, which uses machine learning and not an explicit list of words to censor. However, profanity-check also failed to catch certain phrases.
# + id="XF5-jV5GUWm4" colab_type="code" colab={}
# install profanity-filter
# !pip install profanity-filter
# + id="G9oGwpSVUerb" colab_type="code" colab={}
# install profanity-check
# !pip install profanity-check
# + id="4Z6crk4xVgrj" colab_type="code" colab={}
# imports
from json import loads, dumps
import pandas as pd
from profanity_filter import ProfanityFilter
from profanity_check import predict, predict_prob
# + id="qE_JD6JzTMz4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="cf024e9a-2ab8-47b8-9ff7-df3a4ef9f3d3"
# using profanity-filter package
pf = ProfanityFilter()
# doesn't work on certain inappropriate words and phrases
# but isn't triggerd by Scunthorpe Problem words
pf.censor("Shit piss fuck cunt cocksucker motherfucker tits fuck turd and twat grape scunthorpe shell")
# + id="KsX9gJaKUh3u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ff4f9c5c-85f7-4fb3-f898-8976d7336813"
# profanity filter doesn't work on certain inappropriate phrases
pf.censor("2 girls 1 cup")
# + id="xU0h5sfYU0Fr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16e13afd-6b01-4d19-e46f-51e9fb3b3867"
# profanity check doesn't work on inappropriate phrases either
from profanity_check import predict, predict_prob
predict(['2 girls 1 cup'])
# + [markdown] id="SZT5hMJnfjUt" colab_type="text"
# # Using a Custom List of Words
#
# The Story Squad stakeholder had a preference for flexibility in changing the list as slang changes. Also, there are words that are inappropriate for elementary children that would not be considered inappropriate for adults. These require a custom list of words.
# + id="_nWPwZvAU2sJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="3f561003-a157-4497-f74b-3d86e6b58b4b"
# load in bad words
df = pd.read_csv('bad_single.csv', usecols=[0], names=None)
print(df.shape)
df.head()
# + id="AznJYqYNV7Wi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="a1c31e33-c4dd-4c27-91ff-92364fa9f307"
# load in bad phrases
df2 = pd.read_csv('bad_phrases.csv', usecols=[0], names=None)
print(df2.shape)
df2.head()
# + id="_cEVTi_lXjZg" colab_type="code" colab={}
# convert to list
bad_words = df['Bad_words'].to_list()
bad_phrases = df2['Bad_phrases'].to_list()
# combine lists
bad_words_combined = bad_words + bad_phrases
# + id="HCH1U4RJYVUu" colab_type="code" colab={}
# flag True or False if string in transcriptions contains bad words from the list
transcriptions = {'images': ['The quick alabama hot pocket donkey punch fuck shit however against grape scunthorpe'],
'metadata': []}
def flag_bad_words(transcriptions):
# convert dict into str
parsed_string = dumps(transcriptions)
# determine if any words in the story are in the bad words list
res = any(word in parsed_string for word in bad_words_combined)
# return dictionary with True or False for backend to send to admin
if res == True:
dict = {'bad_words': [True]}
return transcriptions.update(dict)
else:
dict = {'bad_words': [False]}
return transcriptions.update(dict)
# + id="8hv2clvEYfRd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="c1be4242-1a8b-408f-dc49-9f4d6467103e"
# call function on transcriptions
flag_bad_words(transcriptions)
# show transcriptions
transcriptions
# + id="IoVCtOh6Yi05" colab_type="code" colab={}
def return_bad_words(transcriptions):
# convert dict to str
parsed_string = dumps(transcriptions)
# returns list of matching words
new_list = []
for word in bad_words_combined:
if word in parsed_string:
new_list.append(word)
# returns dictionary with list of matches
dict = {'possible_words' : new_list}
return transcriptions.update(dict)
# + id="tm68kqJwZRZB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="7a6d1e6f-2222-439b-fb95-3f774283f8a3"
# return possible bad words
# as you can see the Scunthorpe problem exists return bad words within other words
return_bad_words(transcriptions)
transcriptions
# + [markdown] id="CiIDY7Fifr2l" colab_type="text"
# # Solving the Scunthorpe Problem
# + [markdown] id="QHRvrHAkfvJH" colab_type="text"
# Another problem with flagging profanity is that some words contain bad words within them. For example, the word "hell" is in "shell" and while hell would be considered inappropriate for elementary students using the app, shell would not. The problem is well-documented as the [Scunthorpe Problem](https://https://en.wikipedia.org/wiki/Scunthorpe_problem#:~:text=The%20Scunthorpe%20problem%20is%20the,obscene%20or%20otherwise%20unacceptable%20meaning.). Scunthorpe is a town in England that contains a profane word. A human would not make the mistake, but you could see how a computer might censor users from the town trying to set up an account on the web.
#
# To fix the Scunthorpe problem rather than looking to see if a word from the bad words list is in the text, we need to find only exact matches.
# + id="PIMAtvHGaYk1" colab_type="code" colab={}
# redefine transcriptions
transcriptions = {'images': ['The quick alabama hot pocket donkey punch fuck shit however against grape scunthorpe'],
'metadata': []}
# + id="6q9K1h_Zact2" colab_type="code" colab={}
# Global variable to put caught words and phrase in
flagged_list = []
# Function that removes punctuation from story
def remove_punctuation(transcriptions):
parsed_string = dumps(transcriptions)
punctuations = '''[],!.'"\\?'''
for char in parsed_string:
if char in punctuations:
parsed_string = parsed_string.replace(char, '')
return parsed_string
# Function that looks for bad phrases in story
def return_bad_phrases(transcriptions):
# Convert dict to str using dumps to keep phrases in tact
parsed_string = dumps(transcriptions)
# Lowercase to match list of bad phrases
parsed_string = parsed_string.lower()
# Remove punctuation
parsed_string = remove_punctuation(parsed_string)
# Returns list of matching words and puts in flagged_list global variable
for word in bad_phrases:
if word in parsed_string:
flagged_list.append(word)
# Returns dictionary with list of matches
dict = {'possible_words' : flagged_list}
return transcriptions.update(dict)
# Function that looks for single bad words in story
def return_bad_words(transcriptions):
# Parsing out just the story string from dict to avoid conflicts
parsed_string = list(transcriptions.values())[0][0]
# Lowercase to match list of bad words
parsed_string = parsed_string.lower()
# Remove punctuation
parsed_string = remove_punctuation(parsed_string)
# Splitting into list of strings to detect exact matches
parsed_string = parsed_string.split()
# Finding matches and appending them to flagged_list
for word in bad_words:
if word in parsed_string:
flagged_list.append(word)
# Returns dictionary with list of matches
dict = {'possible_words' : flagged_list}
return transcriptions.update(dict)
# Checks to see if any words have been added to the flagged_list
def flag_bad_words(transcriptions):
if any(flagged_list):
dict = {'flagged' : [True]}
return transcriptions.update(dict)
else:
dict = {'flagged' : [False]}
return transcriptions.update(dict)
# + id="5MEBu0hyamt9" colab_type="code" colab={}
# call functions on transcriptions
return_bad_phrases(transcriptions)
return_bad_words(transcriptions)
# + id="bHscPsk8aral" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="f0043cda-02b7-4387-a1a7-076de2e3cb11"
# Scunthorpe Problem solved!
transcriptions
| Profanity_Filter_and_Scunthorpe_Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Load dependencies
import pandas as pd
import numpy as np
import sys
sys.path.insert(0, '../../statistics_helper')
from fraction_helper import *
pd.options.display.float_format = '{:,.3f}'.format
# # Estimating the fraction of fungi out of the biomass of soil microbes
# Our estimate for the fraction of fungi out of the biomass of soil microbes is based on a study by [<NAME> ](http://dx.doi.org/10.1016/j.soilbio.2008.08.017). Joergensen & Wichern survey the fraction of fungi out of the total microbial biomass using several independent methods. The data in Joergensen & Wichern contains measurements of the fraction of fungi out of the total biomass of soil microbes in four differennt soil types - arable soil, forest soil, grassland soil and litter. We rely on measurement collected in these four soil types using two independent methods - microscopy and measurement of cell wall components.
#
# Here is a sample of the data from Joergensen & Wichern:
# +
data = pd.read_excel('fungi_fraction_data.xlsx',skiprows=1)
data.head()
# -
# Our general methodology for calculating the fraction of fungi out of the biomass of soil microbes is the following. We calculate the geometric mean of all values reported from the same soil type using the same method. This gives us estimates for characteric fraction of fungi in each soil type for each method.
# +
def groupby_geo_frac_mean(input):
return frac_mean(input['Fraction'],weights=input['N'])
type_method_mean = data.groupby(['Method','Type']).apply(groupby_geo_frac_mean).unstack(level=0)
type_method_mean
# -
# We then calculate the geometric mean of the characteristic fractions from different soil types using the same method. This gives us a characteristic fraction of fungi for each of the two methods.
method_mean = type_method_mean.apply(frac_mean)
method_mean
# In the last stage, we calculate the geometric mean of the characteristic values from the two methods. We use the geometric mean as our best estimate for the fraction of fungi out of the total biomass of soil microbes.
best_estimate = frac_mean(method_mean)
print('Our best estimate for the fraction of fungi out of the total biomass of fungi is ≈' + '{:,.0f}%'.format(best_estimate*100))
# # Uncertainty analysis
#
# To calculate the uncertainty associated with the estimate for the fraction of fungi out of the total biomass of number of of bacteria and archaea, we first collect all available uncertainties and then take the largest value as our best projection for the uncertainty.
#
# **Variability of studies using the same method and done in the same soil type** <br>
# We calculate the 95% confidence confidence interval of the values reported by studies performed in the same soil type and using the same method.
#
# +
def groupby_frac_CI(input):
return frac_CI(input['Fraction'])
type_method_CI = data.groupby(['Method','Type']).apply(groupby_frac_CI).unstack(level=0)
type_method_CI
# -
# **Variability of fractions from different soil types measured using the same method** <br>
# We calculate the 95% confidence interval of the characteristic values from each soil type measured in the same method.
intra_method_CI = type_method_mean.apply(frac_CI)
intra_method_CI
# **Variability of fraction measured using different methods** <br>
# We calculate the 95% confidence interval of the characteristic values from each method.
inter_method_CI = frac_CI(method_mean)
print('The 95' + '%'+' confidence interval of the characteristic values from each method is ≈%.1f-fold' % inter_method_CI)
# We choose the highest uncertainty among the uncertianties we collected which is ≈3-fold, as our projection for the uncertainty of the fraction of fungi out of the total biomass of soil microbes.
# Our final parameters are:
# +
mul_CI = np.max([type_method_CI.values.flatten().max(),intra_method_CI.max(),inter_method_CI])
print('Fraction of fungi out of the total biomass of microbes:' +'{:.1f}%'.format(best_estimate*100))
print('Uncertainty associated with the estimate of the total biomass of soil microbes ≈%.1f-fold' % mul_CI)
old_results = pd.read_excel('../fungi_biomass_estimate.xlsx')
result = old_results.copy()
result.loc[1] = pd.Series({
'Parameter': 'Fraction of fungi ou out the total biomass of soil microbes',
'Value': '{0:.1f}'.format(best_estimate),
'Units': 'Unitless',
'Uncertainty': "{0:.1f}".format(mul_CI)
})
result.to_excel('../fungi_biomass_estimate.xlsx',index=False)
| fungi/fungi_fraction/fungi_fraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# Third-party
from astropy.io import fits
import astropy.coordinates as coord
import astropy.units as u
from astropy.table import Table
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
from pyia import GaiaData
# -
R0 = 8.3*u.kpc
gc_frame = coord.Galactocentric(z_sun=0*u.pc, galcen_distance=R0)
# +
# see FGK-select.ipynb
# stype = 'fgk'
stype = 'af'
if stype == 'af':
vmax = 1E2
hex_h = 150 # pc
elif stype == 'fgk':
vmax = 3e2
hex_h = 120
g = GaiaData('../data/{0}.fits'.format(stype))
c = g.skycoord
galcen = c.transform_to(gc_frame)
# -
gal = c.galactic
gal.set_representation_cls('cartesian')
# ---
#
# ## Hexagons
def get_hexagons(h):
a = np.sqrt(3)/2 * h # inner side
pas = dict() # keyed by "ring"
pas[0] = list()
pas[1] = list()
pas[2] = list()
pa0 = mpl.patches.RegularPolygon((0., 0.), numVertices=6,
radius=h, orientation=np.pi/2)
pas[0].append(pa0.get_verts())
for ang in np.arange(0, 360, 60)*u.deg:
# Ring 1
xy0 = 2*a * np.array([np.cos(ang+90*u.deg), np.sin(ang+90*u.deg)])
pa = mpl.patches.RegularPolygon(xy0, numVertices=6,
radius=h, orientation=np.pi/2)
pas[1].append(pa.get_verts())
# Ring 2
xy0 = 4*a * np.array([np.cos(ang+90*u.deg), np.sin(ang+90*u.deg)])
pa = mpl.patches.RegularPolygon(xy0, numVertices=6,
radius=h, orientation=np.pi/2)
pas[2].append(pa.get_verts())
xy0 = 3*h * np.array([np.cos(ang+120*u.deg), np.sin(ang+120*u.deg)])
pa = mpl.patches.RegularPolygon(xy0, numVertices=6,
radius=h, orientation=np.pi/2)
pas[2].append(pa.get_verts())
return pas
hexs = get_hexagons(hex_h)
# +
fig, ax = plt.subplots(1, 1, figsize=(6, 6))
for k in hexs.keys():
for pa in hexs[k]:
pa = mpl.patches.Polygon(pa, facecolor='none', edgecolor='#333333')
ax.add_patch(pa)
ax.plot(gal.u, gal.v,
marker=',', alpha=0.1, color='k',
linestyle='none', zorder=100)
lim = 1000
ax.set_xlim(-lim, lim)
ax.set_ylim(-lim, lim)
ax.set_xlabel('$x$ [pc]')
ax.set_ylabel('$y$ [pc]')
# -
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].hist(c.distance.value, bins=np.linspace(0, 500, 128));
axes[1].hist(gal.w.value, bins=np.linspace(-250, 250, 101));
# ---
# +
cyl = gal.transform_to(gc_frame)
cyl.set_representation_cls('cylindrical')
xyz = np.vstack((gal.u.to(u.pc).value,
gal.v.to(u.pc).value,
gal.w.to(u.pc).value)).T
UVW = np.vstack((cyl.d_rho.to(u.km/u.s).value,
(cyl.rho * cyl.d_phi).to(u.km/u.s, u.dimensionless_angles()).value,
cyl.d_z.to(u.km/u.s).value)).T
# -
# testing
hex_mask = mpl.patches.Path(hexs[0][0]).contains_points(xyz[:, :2])
from scipy.stats import binned_statistic
# +
fig, axes = plt.subplots(1, 2, figsize=(12, 6))
_rho = cyl.rho.value[hex_mask]
axes[0].plot(_rho, UVW[hex_mask, 1],
marker=',', linestyle='none')
# stat = binned_statistic(_rho, UVW[hex_mask, 1],
# bins=np.linspace(_rho.min(), _rho.max(), 16),
# statistic='mean')
# axes[0].plot(0.5*(stat.bin_edges[:-1]+stat.bin_edges[1:]), stat.statistic)
xx = np.linspace(8150, 8450, 15)
axes[0].plot(xx, 5/200 * (xx - 8300) - 185)
axes[0].set_ylim(-300, -100)
_phi = cyl.phi.wrap_at(2*np.pi*u.radian).radian[hex_mask]
axes[1].plot(_phi, UVW[hex_mask, 0],
marker=',', linestyle='none')
stat = binned_statistic(_phi, UVW[hex_mask, 0],
bins=np.linspace(_phi.min(), _phi.max(), 16),
statistic='mean')
axes[1].plot(0.5*(stat.bin_edges[:-1]+stat.bin_edges[1:]), stat.statistic)
axes[1].set_ylim(-10, 10)
# -
# Bovy: A = ~15, B = ~-11
# +
dVphi_dR = 5*u.km/u.s / (200*u.pc)
# dVR_dphi = -2*u.km/u.s / (0.02*u.radian)
dVR_dphi = np.polyfit(0.5*(stat.bin_edges[:-1]+stat.bin_edges[1:]),
stat.statistic, deg=1)[-1] * u.km/u.s / u.radian
_R = cyl.rho.value[hex_mask]
statR = binned_statistic(_R, UVW[hex_mask, 0],
bins=np.linspace(_R.min(), _R.max(), 16),
statistic='mean')
dVR_dR = np.polyfit(0.5*(statR.bin_edges[:-1]+statR.bin_edges[1:]),
statR.statistic, deg=1)[-1] * u.km/u.s / u.kpc
statphi = binned_statistic(_phi, UVW[hex_mask, 1],
bins=np.linspace(_phi.min(), _phi.max(), 16),
statistic='mean')
dVphi_dphi = np.polyfit(0.5*(statphi.bin_edges[:-1]+statphi.bin_edges[1:]),
statphi.statistic, deg=1)[-1] * u.km/u.s / u.radian
# - sign's on A,B because phi increases opposite way as Oort defines!
with u.set_enabled_equivalencies(u.dimensionless_angles()):
A = -0.5 * (np.mean(UVW[hex_mask, 1])*u.km/u.s / R0 - dVphi_dR - dVR_dphi/R0)
B = -0.5 * (-np.mean(UVW[hex_mask, 1])*u.km/u.s / R0 - dVphi_dR + dVR_dphi/R0)
C = 0.5 * (-np.mean(UVW[hex_mask, 0])*u.km/u.s / R0 + dVR_dR - dVphi_dphi/R0)
D = 0.5 * (np.mean(UVW[hex_mask, 0])*u.km/u.s / R0 + dVR_dR + dVphi_dphi/R0)
# -
A
B
C
D
| notebooks/Oorts-constants.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from utils.all import *
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.datasets import reuters
import numpy as np
(train_data, train_labels), (test_data, test_labels) = reuters.load_data(num_words=10000)
len(train_data), len(test_data)
train_data[0][:10]
word_index = reuters.get_word_index()
index_word = {v:k for k,v in word_index.items()}
index_word[-3] = '_PAD_'
index_word[-2] = '_START_'
index_word[-1] = '_UNK_'
' '.join([index_word.get(i-3, '?') for i in train_data[0]])
def vectorize_sequences(sequences, dimension=10000):
result = np.zeros([len(sequences), dimension])
for i, sequence in enumerate(sequences):
for s in sequence:
result[i,s] = 1.
return result
x_train, x_test = [vectorize_sequences(s) for s in [train_data, test_data]]
def to_one_hot(labels, dimension=46):
result = np.zeros([len(labels), dimension])
for i, label in enumerate(labels):
result[i, label] = 1.
return result
y_train, y_test = [to_one_hot(l) for l in [train_labels, test_labels]]
model = keras.Sequential([
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(46, activation='softmax')
])
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
x_val = x_train[:1000]
partial_x_train = x_train[1000:]
y_val = y_train[:1000]
partial_y_train = y_train[1000:]
partial_x_train.shape
history = model.fit(
partial_x_train,
partial_y_train,
batch_size=512,
epochs=20,
validation_data=[x_val, y_val]
)
plot_history(history)
# +
model = keras.Sequential([
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(46, activation='softmax')
])
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(
partial_x_train,
partial_y_train,
batch_size=512,
epochs=9,
validation_data=[x_val, y_val]
)
# -
a = np.unique(train_labels, return_counts=True)
for b in zip(a):
print(b)
values, counts = np.unique(train_labels, return_counts=True)
for value, count in zip(values, counts):
print(value, count)
# Predicting 3 every time would give us an accuracy of ~35% (3159/8982)
predictions = model.predict(x_test[:5])
tf.argmax(predictions, axis=1)
# if we don't want to 1-hot the labels ... `sparse_categorical_crossentropy`
# +
y_train, y_test = [np.array(l) for l in [train_labels, test_labels]]
y_val = y_train[:1000]
partial_y_train = y_train[1000:]
model = keras.Sequential([
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(64, activation='relu'),
keras.layers.Dense(46, activation='softmax')
])
model.compile(optimizer='rmsprop', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
history = model.fit(
partial_x_train,
partial_y_train,
batch_size=512,
epochs=2,
validation_data=[x_val, y_val]
)
| 04.2_routers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Language basics
# This chapter will start with a short tutorial to get you familiar with Python. You will quickly see the similarities with whatever programming language you already know. After this introduction we will start by formalizing things and naming them (**semantics**). As we discussed last week, using clear semantics is primordial to understand software documentation and to "ask questions the right way" in search engines.
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Language-basics" data-toc-modified-id="Language-basics-5"><span class="toc-item-num">5 </span>Language basics</a></span><ul class="toc-item"><li><span><a href="#An-entry-level-tutorial" data-toc-modified-id="An-entry-level-tutorial-5.1"><span class="toc-item-num">5.1 </span>An entry level tutorial</a></span><ul class="toc-item"><li><span><a href="#Python-as-a-Calculator" data-toc-modified-id="Python-as-a-Calculator-5.1.1"><span class="toc-item-num">5.1.1 </span>Python as a Calculator</a></span></li><li><span><a href="#Strings" data-toc-modified-id="Strings-5.1.2"><span class="toc-item-num">5.1.2 </span>Strings</a></span></li></ul></li><li><span><a href="#Basic-data-types" data-toc-modified-id="Basic-data-types-5.2"><span class="toc-item-num">5.2 </span>Basic data types</a></span><ul class="toc-item"><li><span><a href="#Asking-for-the-type-of-an-object" data-toc-modified-id="Asking-for-the-type-of-an-object-5.2.1"><span class="toc-item-num">5.2.1 </span>Asking for the type of an object</a></span></li><li><span><a href="#Numeric-types" data-toc-modified-id="Numeric-types-5.2.2"><span class="toc-item-num">5.2.2 </span>Numeric types</a></span></li><li><span><a href="#Booleans" data-toc-modified-id="Booleans-5.2.3"><span class="toc-item-num">5.2.3 </span>Booleans</a></span></li><li><span><a href="#Text" data-toc-modified-id="Text-5.2.4"><span class="toc-item-num">5.2.4 </span>Text</a></span></li><li><span><a href="#Sequence-types---list,-tuple,-range" data-toc-modified-id="Sequence-types---list,-tuple,-range-5.2.5"><span class="toc-item-num">5.2.5 </span>Sequence types - list, tuple, range</a></span></li><li><span><a href="#Sets" data-toc-modified-id="Sets-5.2.6"><span class="toc-item-num">5.2.6 </span>Sets</a></span></li><li><span><a href="#Mapping-types---dictionaries" data-toc-modified-id="Mapping-types---dictionaries-5.2.7"><span class="toc-item-num">5.2.7 </span>Mapping types - dictionaries</a></span></li><li><span><a href="#Semantics-parenthesis:-"literals"" data-toc-modified-id="Semantics-parenthesis:-"literals"-5.2.8"><span class="toc-item-num">5.2.8 </span>Semantics parenthesis: "literals"</a></span></li></ul></li><li><span><a href="#Control-flow" data-toc-modified-id="Control-flow-5.3"><span class="toc-item-num">5.3 </span>Control flow</a></span><ul class="toc-item"><li><span><a href="#First-steps-towards-programming" data-toc-modified-id="First-steps-towards-programming-5.3.1"><span class="toc-item-num">5.3.1 </span>First steps towards programming</a></span></li><li><span><a href="#The-if-statement" data-toc-modified-id="The-if-statement-5.3.2"><span class="toc-item-num">5.3.2 </span>The <code>if</code> statement</a></span></li><li><span><a href="#The-for-statement" data-toc-modified-id="The-for-statement-5.3.3"><span class="toc-item-num">5.3.3 </span>The <code>for</code> statement</a></span></li><li><span><a href="#The-break-and-continue-statements" data-toc-modified-id="The-break-and-continue-statements-5.3.4"><span class="toc-item-num">5.3.4 </span>The <code>break</code> and <code>continue</code> statements</a></span></li></ul></li><li><span><a href="#Defining--functions" data-toc-modified-id="Defining--functions-5.4"><span class="toc-item-num">5.4 </span>Defining functions</a></span><ul class="toc-item"><li><span><a href="#A-first-example" data-toc-modified-id="A-first-example-5.4.1"><span class="toc-item-num">5.4.1 </span>A first example</a></span></li><li><span><a href="#Positional-and-keyword-arguments" data-toc-modified-id="Positional-and-keyword-arguments-5.4.2"><span class="toc-item-num">5.4.2 </span>Positional and keyword arguments</a></span></li></ul></li><li><span><a href="#Importing-modules-and-functions" data-toc-modified-id="Importing-modules-and-functions-5.5"><span class="toc-item-num">5.5 </span>Importing modules and functions</a></span></li><li><span><a href="#Take-home-points" data-toc-modified-id="Take-home-points-5.6"><span class="toc-item-num">5.6 </span>Take home points</a></span></li><li><span><a href="#What's-next?" data-toc-modified-id="What's-next?-5.7"><span class="toc-item-num">5.7 </span>What's next?</a></span></li><li><span><a href="#License" data-toc-modified-id="License-5.8"><span class="toc-item-num">5.8 </span>License</a></span></li></ul></li></ul></div>
# -
# ## An entry level tutorial
# Let's start by following a simple tutorial together. You can simply read through the examples; however, I highly recommend to open an **ipython** interpreter or a **notebook** (see the climate lecture) to test the commands yourself as the tutorial goes on.
#
# In most online tutorials you will see ```>>>``` to represent the python prompt, but with ipython or this tutorial you will use the numerated prompt ```In [1]:```.
#
# **Copyright notice**: many of these examples and explanations are simply copy-pasted from the [official python tutorial](https://docs.python.org/3/tutorial/).
# ### Python as a Calculator
# The interpreter acts as a simple calculator: you can type an expression at it and it will write the value. Expression syntax is straightforward: the operators ``+``, ``-``, ``*`` and ``/`` work just like in most other languages:
2 + 2
50 - 5*6
8 / 5 # division always returns a floating point number
# Comments in Python start with the hash character, `#`, and extend to the end of the physical line. A comment may appear at the start of a line or following whitespace or code:
# this is the first comment
spam = 1 # and this is the second comment
# ... and now a third!
# Parentheses `()` can be used for grouping:
(50 - 5*6) / 4
# With Python, the `**` operator is used to calculate powers:
5 ** 2
# The equal sign (`=`) is used to assign a value to a variable. Afterwards, no result is displayed before the next interactive prompt:
width = 20
height = 5 * 9
width * height
# If a variable is not “defined” (assigned a value), trying to use it will give you an error:
n # try to access an undefined variable
# In interactive mode, the last printed expression is assigned to the variable `_`. This means that when you are using Python as a desk calculator, it is somewhat easier to continue calculations, for example:
tax = 12.5 / 100
price = 100.50
price * tax
price + _
# `_` should be treated as a read-only variable, to use in the interpreter only.
# ### Strings
# Besides numbers, Python can also manipulate strings, which can be expressed in several ways. They can be enclosed in single quotes (`'...'`) or double quotes (`"..."`) with the same result:
'spam eggs'
"spam eggs"
# The double quotes are useful if you need to use a single quote in a string:
"doesn't"
# Alternatively, `\` can be used to escape quotes:
'doesn\'t'
# If you don’t want characters prefaced by `\` to be interpreted as special characters, you can use raw strings by adding an `r` before the first quote. This is useful for Windows paths:
print('C:\some\name') # here \n means newline!
print(r'C:\some\name') # note the r before the quote
# Strings can be concatenated (glued together) with the `+` operator, and repeated with `*`:
("She's a " + 'witch! ') * 3
# Strings can be indexed (subscripted), with the first character having index 0:
word = 'Python'
word[0] # character in position 0
word[5] # character in position 5
# Indices may also be negative numbers, to start counting from the right:
word[-1] # last character
word[-2] # second-last character
# In addition to indexing, slicing is also supported. While indexing is used to obtain individual characters, slicing allows you to obtain a substring:
word[0:2] # characters from position 0 (included) to 2 (excluded)
word[2:5] # characters from position 2 (included) to 5 (excluded)
# Note how the start is always included, and the end always excluded. This makes sure that `s[:i] + s[i:]` is always equal to `s`:
word[:2] + word[2:]
# Attempting to use an index that is too large will result in an error:
word[42] # the word only has 6 characters
# However, out of range slice indexes are handled gracefully when used for slicing:
word[4:42]
word[42:]
# The **built-in** **function** `len()` returns the length of a string:
s = 'supercalifragilisticexpialidocious'
len(s)
# ## Basic data types
# Now that you are more familiar with the basics, let's start to name things "the right way". For example: an informal way to describe a programming language is to say that it "does things with stuff".
#
# These "stuff" are formally called "objects" in python. We will define objects more precisely towards the end of the lecture, but for now remember one important thing: **in python, everything is an object**. Yes, everything.
#
# Python objects have a **type** (synonym: [data type](https://en.wikipedia.org/wiki/Data_type)). In the previous tutorial, you used exclusively [built-in](https://docs.python.org/3/library/stdtypes.html) types. **Built-in data types** are directly available in the interpreter, as opposed to other data types which maybe obtained either by importing them (e.g. ``from collections import OrderedDict``) or by creating new data types yourselves.
# ### Asking for the type of an object
type(1)
a = 'Hello'
type(a)
# <img src="../img/logo_ex.png" align="left" style="width:1em; height:1em;"> **Exercise**: add a ``print`` call in the statement above to see the difference with ipython's simplified print. What is the type of ``type``, by the way?
# ### Numeric types
# There are three distinct numeric types: **integers** (``int``), **floating point numbers** (``float``), and **complex numbers** (``complex``). We will talk about these in more details in the numerics chapter.
# ### Booleans
# There is a built-in boolean data type (``bool``) useful to test for truth value. Examples:
type(True), type(False)
type(a == 'Hello')
3 < 5
# Note that there are other rules about testing for truth in python. This is quite convenient if you want to avoid doing operation on invalid or empty containers:
if '':
print('This should not happen')
# In Python, like in C, any non-zero integer value is true; zero is false:
if 1 and 2:
print('This will happen')
# Refer to the [docs](https://docs.python.org/3/library/stdtypes.html#truth-value-testing) for an exhaustive list of boolean operations and comparison operators.
# ### Text
# In python (and many other languages) text sequences are named **strings** (``str``), which can be of any length:
type('Français, 汉语') # unicode characters are no problem in Python
# Unlike some languages, there is no special type for characters:
for char in 'string':
# "char" is also a string of length 1
print(char, type(char))
# Since strings behave like **lists** in many ways, they are often classified together with the **sequence** types, as we will see below.
# Python strings cannot be changed - they are [immutable](https://en.wikipedia.org/wiki/Immutable_object). Therefore, assigning to an indexed position in the string results in an error:
word = 'Python'
word[0] = 'J'
# Python objects have **methods** attached to them. We will learn more about methods later, but here is an example:
word.upper() # the method .upper() converts all letters in a string to upper case
"She's a witch!".split(' ') # the .split() method divides strings using a separator
# ### Sequence types - list, tuple, range
# Python knows a number of sequence data types, used to group together other values. The most versatile is the list, which can be written as a list of comma-separated values (items) between square brackets. Lists might contain items of different types, but usually the items all have the same type.
squares = [1, 4, 9, 16, 25, 36, 49]
squares
# Lists can be indexed and sliced:
squares[0]
squares[-3:]
squares[0:7:2] # new slicing! From element 0 to 7 in steps of 2
squares[::-1] # new slicing! All elements in steps of -1, i.e. reverse
# **Careful!** Lists are not the equivalent of arrays in Matlab. One major difference being that the addition operator *concatenates* lists together (like strings), instead of adding the numbers elementwise like in Matlab:
squares + [64, 81, 100]
# Unlike strings, which are immutable, lists are a mutable type, i.e. it is possible to change their content:
cubes = [1, 8, 27, 65, 125] # something's wrong here
cubes[3] = 64
cubes
# Assignment to slices is also possible, and this can even change the size of the list:
letters = ['a', 'b', 'c', 'd', 'e', 'f', 'g']
letters[2:5] = ['C', 'D', 'E'] # replace some values
letters
letters[2:5] = [] # now remove them
letters
# The built-in function `len()` also applies to lists:
len(letters)
# It is possible to nest lists (create lists containing other lists), as it is possible to store different objects in lists. For example:
a = ['a', 'b', 'c']
n = [1, 2, 3]
x = [a, n, 3.14]
x
x[0][1]
# Lists also have methods attached to them (see [5.1 More on lists](https://docs.python.org/3/tutorial/datastructures.html#more-on-lists) for the most commonly used). For example:
alphabet = ['c', 'b', 'd']
alphabet.append('a') # add an element to the list
alphabet.sort() # sort it
alphabet
# Other sequence types include: **string, tuple, range**. Sequence types support a [common set of operations](https://docs.python.org/3/library/stdtypes.html#common-sequence-operations) and are therefore very similar:
l = [0, 1, 2]
t = (0, 1, 2)
r = range(3)
s = '123'
# Test if elements can be found in the sequence(s)
1 in l, 1 in t, 1 in r, '1' in s
# Ask for the length
len(l), len(t), len(r), len(s)
# Addition
print(l + l)
print(t + t)
print(s + s)
# The addition operator won't work for the range type though. Ranges are a little different than lists or strings:
r = range(2, 13, 2)
r # r is an object of type "range". It doesn't print all the values, just the interval and steps
list(r) # applying list() converts range objects to a list of values
# Ranges are usually used as loop counter or to generate other sequences. Ranges have a strong advantage over lists and tuples: their elements are generated *when they are needed*, not before. Ranges have therefore a very low memory consumption. See the following:
range(2**100) # no problem
list(range(2**100)) # trying to make a list of values out of it results in an error
# An ``OverflowError`` tells me that I'm trying to create an array too big to fit into memory.
# The "**tuple**" data type is probably a new concept for you, as tuples are quite specific to python. A tuple behaves *almost* like a list, but the major difference is that a tuple is **immutable**:
l[1] = 'ha!' # I can change an element of a list
l
t[1] = 'ha?' # But I cannot change an element of a tuple
# It is immutability which makes tuples useful, but for beginners this is not really obvious at the first sight. We will get back to tuples later in the lecture.
# ### Sets
# Sets are an unordered collection of **distinct** objects:
s1 = {'why', 1, 9}
s2 = {9, 'not'}
s1
# Let's compute the union of these two sets. We use the method ".union()" for this purpose:
s1.union(s2) # 9 was already in the set, however it is not doubled in the union
# Sets are useful for operations such as intersection, union, difference, and symmetric difference between sequences. You won't see much use for them in this semester, but remember that they exist.
# ### Mapping types - dictionaries
# A **mapping object** maps values (**keys**) to arbitrary objects (**values**): the most frequently used mapping object is called a **dictionary**. It is a collection of (key, value) pairs:
tel = {'jack': 4098, 'sape': 4139}
tel
tel['guido'] = 4127
tel
del tel['sape']
tel
# *Keys* can be of any immutable type: e.g. strings and numbers are often used as keys. The keys in a dictionary are all unique (they have to be):
d = {'a':1, 2:'b', 'c':1} # a, 2, and c are keys
d
# You can ask whether a (key, value) pair is available in a dict with the statement:
2 in d
# However, you cannot check appartenance by value, since the values are not necessarily unique:
1 in d
# Dictionaries are (together with lists) the **container** type you will use the most often.
#
# *Note: there are other mapping types in python, but they are all related to the original ``dict``. Examples include ``collections.OrderedDict``, which is a dictionary preserving the order in which the keys are entered.*
# <img src="../img/logo_ex.png" align="left" style="width:1em; height:1em;"> **Exercise**: can you think of examples of application of a ``dict``? Describe a couple of them!
# ### Semantics parenthesis: "literals"
# **Literals** are the fixed values of a programming language ("notations"). Some of them are pretty universal, like numbers or strings (``9``, ``3.14``, ``"Hi!"``, all literals) some are more language specific and belong to the language's syntax. Curly brackets ``{}`` for example are the literal representation of a ``dict``. The literal syntax has been added for convenience only:
d1 = dict(bird='parrot', plant='crocus') # one way to make a dict
d2 = {'bird':'parrot', 'plant':'crocus'} # another way to make a dict
d1 == d2
# Both `{}` and `dict()` are equivalent: using one or the other to construct your containers is a matter of taste, but in practice you will see the literal version more often.
# ## Control flow
# ### First steps towards programming
# Of course, we can use Python for more complicated tasks than adding two and two together. For instance, we can write an initial sub-sequence of the [Fibonacci series](https://en.wikipedia.org/wiki/Fibonacci_number) as follows:
# Fibonacci series:
# the sum of two previous elements defines the next
a, b = 0, 1
while a < 10:
print(a)
a, b = b, a+b
# This example introduces several new features.
# - The first line contains a multiple assignment: the variables a and b simultaneously get the new values 0 and 1. On the last line this is used again, demonstrating that the expressions on the right-hand side are all evaluated first before any of the assignments take place. The right-hand side expressions are evaluated from the left to the right.
# - The while loop executes as long as the condition (here: ``a < 10``) remains true. The standard comparison operators are written the same as in C: `<` (less than), `>` (greater than), `==` (equal to), `<=` (less than or equal to), `>=` (greater than or equal to) and `!=` (not equal to).
# - The body of the loop is **indented**: indentation is Python’s way of grouping statements, and not via brackets or ``begin .. end`` statements. Hate it or love it, this is how it is ;-). I learned to like this style a lot. **Note that each line within a basic block must be indented by the same amount.** Although the indentation could be anything (two spaces, three spaces, tabs...), the recommended way is to use **four spaces**.
# The print() function accepts multiple arguments:
i = 256*256
print('The value of i is', i)
# The keyword argument (see definition below) ``end`` can be used to avoid the newline after the output, or end the output with a different string:
a, b = 0, 1
while a < 1000:
print(a, end=',')
a, b = b, a+b
# ### The `if` statement
# Perhaps the most well-known statement type is the if statement:
x = 12
if x < 0:
x = 0
print('Negative changed to zero')
elif x == 0:
print('Zero')
elif x == 1:
print('Single')
else:
print('More')
# There can be zero or more `elif` parts, and the `else` part is optional. The keyword `elif` is short for "else if", and is useful to avoid excessive indentation.
# ### The `for` statement
# The `for` loops in python can be quite different than in other languages: **in python, one iterates over sequences, not indexes**. This is a feature I very much like for its readability:
words = ['She', 'is', 'a', 'witch']
for w in words:
print(w)
# The equivalent for loop with a counter is considered "unpythonic", i.e. not elegant.
#
# **Unpythonic:**
seq = ['This', 'is', 'very', 'unpythonic']
# Do not do this at home!
n = len(seq)
for i in range(n):
print(seq[i])
# **Pythonic**:
seq[-1] = 'pythonic'
for s in seq:
print(s)
# ``for i in range(xx)`` is *almost never* what you want to do in python. If you have several sequences you want to **iterate** over, then do:
squares = [1, 4, 9, 25]
for s, l in zip(seq, squares):
print(l, s)
# ### The `break` and `continue` statements
# The `break` statement breaks out of the innermost enclosing for or while loop:
for letter in 'Python':
if letter == 'h':
break
print('Current letter:', letter)
# The `continue statement` continues with the next iteration of the loop:
for num in range(2, 10):
if num % 2 == 0:
print("Found an even number", num)
continue
print("Found a number", num)
# ## Defining functions
# ### A first example
# +
def fib(n):
"""Print a Fibonacci series up to n."""
a, b = 0, 1
while a < n:
print(a, end=' ')
a, b = b, a+b
# Now call the function we just defined:
fib(2000)
# -
# The `def` statement introduces a function definition. It must be followed by the function name and the parenthesized list of formal parameters. The statements that form the body of the function start at the next line, and **must be indented**.
#
# The first statement of the function body can optionally be a string literal; this string literal is the function's documentation string, or docstring (more about docstrings later: in the meantime, make a habit out of it).
#
# A function definition introduces the function name in the current **scope** (we will learn about scopes soon). The value of the function name has a type that is recognized by the interpreter as a user-defined function. This value can be assigned to another name which can then also be used as a function. This serves as a general renaming mechanism:
fib
f = fib
f(100)
# Coming from other languages, you might object that `fib` is not a function but a procedure since it doesn't return a value. In fact, even functions without a return statement do return a value, albeit a rather boring one. This value is called `None` (it’s a built-in name). Writing the value `None` is normally suppressed by the interpreter if it would be the only value written. You can see it if you really want to by using `print()`:
fib(0) # shows nothing
print(fib(0)) # prints None
# It is simple to write a function that returns a list of the numbers of the Fibonacci series, instead of printing it:
# +
def fib2(n): # return Fibonacci series up to n
"""Return a list containing the Fibonacci series up to n."""
result = []
a, b = 0, 1
while a < n:
result.append(a)
a, b = b, a+b
return result
r = fib2(100) # call it
r # print the result
# -
# ### Positional and keyword arguments
# Functions have two types of arguments: **positional arguments** and **keyword arguments**.
#
# **keyword arguments** are preceded by an identifier (e.g. ``name=``) and are attributed a default value. They are therefore *optional*:
def f(arg1, arg2, kwarg1=None, kwarg2='Something'):
"""Some function with arguments."""
print(arg1, arg2, kwarg1, kwarg2)
f(1, 2) # no need to specify them - they are optional and have default values
f(1, 2, kwarg1=3.14, kwarg2='Yes') # but you can set them to a new value
f(1, 2, kwarg2='Yes', kwarg1=3.14) # and the order is not important!
# Unfortunately, it is also possible to set keyword arguments without naming them, in which case the order matters:
f(1, 2, 'Yes', 'No')
# I am not a big fan of this feature because it reduces the clarity of the code. I recommend to always use the ``kwarg=`` syntax. Others agree with me, and therefore python implemented a syntax to make calls like the above illegal:
# The * before the keyword arguments make them keyword arguments ONLY
def f(arg1, arg2, *, kwarg1=None, kwarg2='None'):
print(arg1, arg2, kwarg1, kwarg2)
f(1, 2, 'Yes', 'No') # This now raises an error
# **positional arguments** are named like this because their position matters, and unlike keyword arguments they don't have a default value and they are mandatory. Forgetting to set them results in an error:
f(1)
# ## Importing modules and functions
# Although python ships with some built-in functions available in the interpreter (e.g. `len()`, `print()`), it is by far not enough to do real world programming. Thankfully, python comes with a mechanism which allows us to access *much* more functionality:
import math
print(math)
print(math.pi)
# `math` is a **module**, and it has attributes (e.g. `pi`) and functions attached to it:
math.sin(math.pi / 4) # compute a sinus
# `math` is available in the python **standard library** (https://docs.python.org/3/library/): this means that it comes pre-installed together with python itself. Other modules can be installed (like `numpy` or `matplotlib`), but we won't need them for now.
#
# Modules often have a thematic grouping, i.e. `math`, `time`, `multiprocessing`. You will learn more about them in the next lecture.
# ## Take home points
# - in python, everything is an object
# - all objects have a data type: examples of data types include floats, strings, dicts, lists...
# - you can ask for the type of an object with the built-in function ``type()``
# - "built-in" means that a function or data type is available at the command prompt without import statement
# - objects also have methods attached to them, e.g. ``.upper()`` for strings, ``.append()`` for lists
# - lists and dicts are the container data types you will use most often
# - certain objects are immutable (strings, tuples), but others are mutable and can change their state (dicts, lists)
# - in python, indentation matters! This is how you define blocks of code. Keep your indentation consistent, with 4 spaces
# - in python, one iterates over sequences, not indexes
# - functions are defined with ``def``, and also rely on indentation to define blocks. They can have a `return` statement
# - there are two types or arguments in functions: positional (mandatory) and keyword (optional) arguments
# - the `import` statement opens a whole new world of possibilities: you can access other standard tools that are not available at the top-level prompt
# ## What's next?
# We learned the basic elements of the python syntax: to become fluent with this new language you will have to get familiar with all of the elements presented above. With time, you might want to get back to this chapter (or to the python reference documentation) to revisit what you've learned. I also highly recommend to follow the official python tutorial, sections 3 to 5.
# Back to the [table of contents](00-Introduction.ipynb#ctoc), or [jump to this week's assignment](06-Assignment-02.ipynb).
# ## License
# <a href="https://creativecommons.org/licenses/by/4.0/" target="_blank">
# <img align="left" src="https://mirrors.creativecommons.org/presskit/buttons/88x31/svg/by.svg"/>
# </a>
| notebooks/05-Language-Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Anatomía de un Gráfico
#
# Cada gráfico en `altair` es compuesto al describir un **mínimo** de tres elementos:
# * Datos
# * Marcadores
# * Codificaciones
#
# ***
# ## Datos
# `Altair` acepta __datasets__ de 3 maneras:
# * un `DataFrame` de `pandas`
# * un objeto de clase `Data` de `altair` u objetos relacionados (`InLineData`, `NamedData`, `UrlData`)
# * datos en formato `JSON` o `csv` de manera directa o una URL
# * un objeto con la propiedad `__geo_interface__` (`GeoDataFrame` de geopandas, geometrías de `shapely` y objetos GeoJSON)
#
# Nosotros trabajaremos con `DataFrames` de `pandas`.
#
# `Pandas` es una de las mejores opciones para trabajar con estructuras de datos en `python`. El nombre `pandas` proviene de __Panel Data__. `pandas` esta basada en `numpy` (de __Numeric Python__) la cual provee estructuras de datos (__arrays__ o _matrices_) las cuales `pandas` utiliza para crear __DataFrames__. Un __DataFrame__ es una estructura de datos en la cual se pueden guardar datos de distintos tipos (cadenas de caractéres (__strings__), __integers__, __floats__, y más) en cada columna. Es similar a una tabla o una planilla de Excel o Google Spreadsheets.
#
# Como es una estructura de `python` claro que su índice comienza en 0.
# ## Marcadores
#
# El marcador en un gráfico es la representación visual de tus datos. `Altair` ofrece los siguientes marcadores hasta el momento:
#
# | Marcador | Método | Descripción | Ejemplo |
# |-----------|-----------------|--------------------------------------------------|-------------------------------------------|
# | area | mark_area() | Un gráfico de area. | [Simple Stacked Area Chart](https://altair-viz.github.io/gallery/simple_stacked_area_chart.html#gallery-simple-stacked-area-chart)|
# | barra | mark_bar() | Un gráfico de barras. | [Simple Bar Chart](https://altair-viz.github.io/gallery/simple_bar_chart.html#gallery-simple-bar-chart)|
# | círculo | mark_circle() | Un diagrama de dispersión con círculos rellenos. | [One Dot Per Zipcode](https://altair-viz.github.io/gallery/one_dot_per_zipcode.html#gallery-one-dot-per-zipcode)|
# |_geofigura_| mark_geoshape() | Una fígura geográfica. | [Choropleth Map](https://altair-viz.github.io/gallery/choropleth.html#gallery-choropleth)|
# | imagen | mark_image() | Un gráfico de dispersión con imágenes como marcadores | [Image mark](https://altair-viz.github.io/user_guide/marks.html#user-guide-image-mark) |
# | línea | mark_line() | Un gráfico de líneas. | [Simple Line Chart](https://altair-viz.github.io/gallery/simple_line_chart.html#gallery-simple-line-chart)|
# | punto | mark_point() | Un diagrama de dispersión con formas de puntos configurables. | [Faceted Scatter Plot with Linked Brushing](https://altair-viz.github.io/gallery/scatter_linked_brush.html#gallery-scatter-linked-brush)|
# | rectángulo| mark_rect() | Un rectángulo relleno, usado para mapas de calor (_heatmaps_). | [Simple Heatmap](https://altair-viz.github.io/gallery/simple_heatmap.html#gallery-simple-heatmap)|
# | regla | mark_rule() |Una línea vertical u horizontal que abarca el eje.| [Candlestick Chart](https://altair-viz.github.io/gallery/candlestick_chart.html#gallery-candlestick-chart)|
# | cuadrado | mark_square() | Un diagrama de dispersián con cuadrados. | N/A |
# | texto | mark_text() | Un diagrama de dispersián con los puntos representados con texto. | [Simple Bar Chart with Labels](https://altair-viz.github.io/gallery/bar_chart_with_labels.html#gallery-bar-chart-with-labels)|
# | marca | mark_tick() | Una marca o línea horizontal o vertical. | [Strip Plot](https://altair-viz.github.io/gallery/strip_plot.html#gallery-strip-plot)|
# ## Codificaciones
#
# Un gráfico es una representación visual de tus datos. Es esencial conectar tu información a un elemento visual en el gráfico. En `altair` eso se le conoce como __encode__ o _codificar_ tus datos. Es el proceso de asignar valores (en este caso columnas de tu __DataFrame__) a elementos posicionales (como el eje X o Y) o propiedades de tu marcador (como el color o el tamaño).
# `Altair` es una biblioteca para crear gráficos altamente configurables asi que simplemente enlistar todas las codificaciones posibles sería una manera muy ineficáz de aprender. La mejor manera de aprender es haciendo.
# ### Tipos de Datos
#
# `altair` hace un buen trabajo deduciendo el tipo de datos con el que estas trabajando al igual que `pandas`. Pero también puedes especificar el tipo de datos en tus gráficos. `Altair` reconoce 4 tipos de datos:
#
# | Tipo de datos | Código | Descripción |
# |:-------------:|:------:|:------------------------------------------|
# | cuantitativo | `Q` | una cantidad continua y de números reales |
# | ordinal | `O` | una cantidad discreta y ordenada |
# | nominal | `N` | una cantidad discreta y desordenada |
# | temporal | `T` | un valor de tiempo o fecha |
# | geojson | `G` | una _geofigura_ en formato GeoJSON |
#
# <p style="text-align:center;"><img src="anatomia-de-un-grafico.png"></p>
import altair as alt
import pandas as pd
import seaborn as sns
data = sns.utils.load_dataset('penguins')
data.head()
alt.Chart(data).mark_tick().encode(x="culmen_length_mm", y = "island")
| notebooks/01-anatomia-de-un-grafico.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# <center><h2>Scale your pandas workflows by changing one line of code</h2>
#
# # Exercise 6: Executing on a cluster environment
#
# **GOAL**: Learn how to connect Modin to a Ray cluster and run pandas queries on a cluster.
#
# **NOTE**: Exercise 5 must be completed first, this exercise relies on the cluster created in Exercise 5.
# Modin performance scales as the number of nodes and cores increases. In this exercise, we will reproduce the data from the plot below using the 120GB [NYC Taxi dataset](https://s3.amazonaws.com/nyc-tlc/trip+data/yellow_tripdata_2015-01.csv) that was provided as part of our [modin-cluster.yaml script](https://github.com/modin-project/modin/blob/master/examples/tutorial/tutorial_notebooks/cluster/modin-cluster.yaml).
#
# 
# Don't change this cell!
import ray
ray.init(address="auto")
import modin.pandas as pd
from modin.config import NPartitions
if NPartitions.get() != 768:
print("This notebook was designed and tested for an 8 node Ray cluster. "
"Proceed at your own risk!")
# !du -h big_yellow.csv
# %%time
df = pd.read_csv("big_yellow.csv", quoting=3)
# %%time
count_result = df.count()
# print
count_result
# %%time
groupby_result = df.groupby("passenger_count").count()
# print
groupby_result
# %%time
apply_result = df.applymap(str)
# print
apply_result
ray.shutdown()
# ## Shutting down the cluster
#
# **You may have to change the path below**. If this does not work, log in to your
#
# Now that we have finished computation, we can shut down the cluster with `ray down`.
# !ray down modin-cluster.yaml
# ### This ends the cluster exercise
| examples/tutorial/tutorial_notebooks/cluster/exercise_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spiral Galaxies
#
# *<NAME>, Nov 2018*
# ## Contents
#
# - [Rotation curves](#rotcurves)
# - [Out-of-plane and radial motion](#othermotions)
# - [Visualizing epicycles](#visepi)
# - [Making spirals](#spirals)
# - [The winding problem](#winding)
# - [Ovals within ovals](#ovals)
# <a id='rotcurve'></a>
# ## Rotation curves
#
# ***TODO***
# +
# %matplotlib inline
import time
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# from IPython.display import Image, HTML, display, clear_output
# from matplotlib.offsetbox import AnchoredText
from ipywidgets import interact, interactive, fixed, interact_manual, Layout, Output
import ipywidgets as w
# make graph labels more readable
plt.rcParams.update({'font.size': 16})
# -
# <a id='othermotions'></a>
# ## Out-of-plane and radial motion
#
# Though stars follow an approximately circular, in-plane path around the galaxy following the minimum in the effective gravitational potential $\Phi_{eff}$, there are a variety of small-amplitude deviations. These are complicated in detail but can be approximated as simple harmonic motion.
#
# For vertical motions, the oscillation follows $\ddot{z} \approx -\nu^2 z$, with a minimum at the midplane and frequency $\nu$.
#
# For radial motions, the minimum is at $R_0$ where $\frac{d\Phi_{eff}}{dR}=0$. Defining $\rho(t) \equiv (R(t)-R_0)$, the oscillation is $\ddot{\rho} = -\kappa^2 \rho$. Here $\kappa$ is the frequency of oscillation, but because of some mathematical similarities to planetary motions in the Ptolemeic system it is often know as *epicycle frequency*.
#
# The frequencies are defined by the 2nd derivative of $\Phi_{eff}$, evaluated at the minimum:
#
# $$ \nu^2 = \frac{\partial^2\Phi_{eff}}{\partial z^2} \bigg|_{midplane} \qquad \kappa^2 = \frac{\partial^2\Phi_{eff}}{\partial R^2} \bigg|_{R_0} $$
#
# In both cases, the frequencies can be arbitrary and will in general give open orbits. Having an oscillation period which is in a simple integer relationship to the galactic orbit period is a special case: now the star will return to its original position, in a closed orbit.
# <a id='visepi'></a>
# ### Visualizing epicycles
#
# What would radial oscillations look like if viewed from above the galactic plane? Probably too small to notice, but we can exaggerate that.
# +
def plot_epicycles(ratio, amplitude):
"ratio: number of radial oscillations per galactic orbit"
nPoints = 100
thetas = np.linspace(0, 4*np.pi, 200)
rs = 1 + amplitude * np.sin(ratio*thetas)
xs = rs*np.cos(thetas)
ys = rs*np.sin(thetas)
plt.figure(figsize=(9, 9))
plt.plot(xs, ys)
plt.title('Epicycles with {} oscillations per orbit, amplitude {} $R_0$'.format(ratio, amplitude))
style = {'description_width': 'initial'} # to avoid the labels getting truncated
interact(plot_epicycles,
ratio = w.FloatSlider(description="Oscillations per orbit", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=1.0, max=8.0,
value=3),
amplitude = w.FloatSlider(description="Amplitude, units of $R_0$", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=0.01, max=0.3, step=0.01,
value=0.1));
# -
# <a id='spirals'></a>
# ## Making spirals
#
# The spirals are visually spectacular and the location of a lot of star formation, but how do they form and how do they persist for billions of years?
#
# We know that for stars within the disk, $\dot{\phi(R)} \ne constant$, so what is the effect of differential rotation?
# <a id='winding'></a>
# ### The winding problem
#
# Imagine we start with a straight line of stars across the galactic diameter, and let it evolve through time as the stars at their individual velocities.
# To implement this, we need an efficient way to rotate points through arbitrary angles in Cartesian coordinates. Some simple linear algebra will do the trick.
#
# The rotation matrix rotates a point in 2-D space anticlockwise about the origin by angle $\theta$:
# $$ \begin{pmatrix} \cos \theta & -\sin \theta \\ \sin \theta & \cos \theta \end{pmatrix}
# \begin{pmatrix} x_0 \\ y_0 \end{pmatrix} = \begin{pmatrix} x_1 \\ y_1 \end{pmatrix} $$
# Note that $| \mathbf{x_0} | = | \mathbf{x_1} |$, so the radius is unchanged.
def rotation(theta):
# theta: angle in degrees
# returns: 2x2 rotation matix as numpy array
theta_rad = theta*np.pi/180
return np.array([[np.cos(theta_rad), -np.sin(theta_rad)],[np.sin(theta_rad), np.cos(theta_rad)]])
# This can multiply $2 \times N$ arrays directly, providing the angle is the same throughout (see the Ovals demo below). However, the winding problem is all about differential rotation so the code gets a bit clumsier.
#
# As a simple model, assume that we ignore the core of the galaxy (the inner 10% of the radius), and outside that the linear velocity $v$ is constant. Then the angular velocity $\omega(r) = v/r$. The plot routine takes an angle (in degrees) for rotation of the outer edge.
#
# Reminder for the old Python 2 programmers: since version 3.5 Python has the @ operator for matrix multiplication. It is no longer necessary to call `numpy.matmul()` explicitly.
def plotWinding(phi_step):
nPoints = 1000
startx = np.linspace(0.1, 1, nPoints)
starty = np.zeros(nPoints)
startline = np.stack([startx, starty])
phis = phi_step*np.ones(nPoints)/startx
rots = rotation(phis)
newline = np.zeros((2,nPoints))
for i in range(nPoints):
newline[:,i] = rots[:,:,i] @ startline[:,i]
fig = plt.figure(figsize=(8,8))
plt.axis('equal')
plt.plot(newline[0,:], newline[1,:], 'b-')
plt.plot(-newline[0,:], -newline[1,:], 'r-')
style = {'description_width': 'initial'} # to avoid the labels getting truncated
interact(plotWinding,
phi_step = w.IntSlider(description="Degrees rotation", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=0, max=150,
value=0) );
# Clearly, in this model the spirals rapidly wind up tighter and tighter, on a timescale much shorter than the lifetime of a spiral galaxy. This doesn't match observations, so we need a different model.
# <a id='ovals'></a>
# ### Ovals within ovals
#
# We should probably think of the spiral arms as patterns rather than structures.
# +
def cartesianCircle(r = 1):
phis = np.linspace(0, 2*np.pi, 100)
x = r*np.cos(phis)
y = r*np.sin(phis)
return x, y
def squashCircle(circ, b):
# scale down y-axis by factor b, leaving x-axis unchanged
M = np.array([[1,0],[0,b]])
return M @ circ
def scale(c):
# scale by axes by a factor c
return np.array([[c,0],[0,c]])
# -
# Define a plot of `nOvals` ellipses, each a factor `dr` larger than the one inside and rotated `dtheta` degrees anticlockwise.
def plotOvals(nOvals=25, dtheta=5, dr=0.05, axratio=0.7):
circ = cartesianCircle()
ell = squashCircle(circ, axratio)
fig = plt.figure(figsize=(8,8))
plt.axis('equal')
plt.plot(ell[0,:], ell[1,:], '-', color='tab:gray')
for n in range(1,nOvals):
newell = rotation(dtheta*n) @ scale(1+dr*n) @ ell
plt.plot(newell[0,:], newell[1,:], '-', color='tab:gray')
style = {'description_width': 'initial'} # to avoid the labels getting truncated
interact(plotOvals,
nOvals = w.IntSlider(description="Ovals to plot", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=5, max=100,
value=25),
dtheta = w.FloatSlider(description="Rotation step (deg)", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=0.1, max=20,
value=5.0),
dr = w.FloatSlider(description="Scale step", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=0, max=0.5, step=0.01,
value=0.05),
axratio = w.FloatSlider(description="Axis ratio", style=style,
layout=Layout(width='80%'),
continuous_update=False,
min=0.5, max=0.9, step=0.01,
value=0.8)
);
# So we can make quite conspicuous spiral arms appear without explicitly drawing them.
| gravity/4 - SpiralGalaxies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="w4B3OoY-shn-" pycharm={}
# # Job Shop Scheduling
#
# Keywords: job shop, scheduling, cbc usage, neos usage, cplex, gdp, disjunctive programming, batch processes
# + [markdown] id="r4uB65VcPHBR" pycharm={}
# ## Imports
#
# The following cell specifies the solver to used in the subsequent calculations. Some of these problems can become quite larger, and therefore the `gurobi` solver has been set as a default. If you don't have the `gurobi` solver then adjust the code to use the `glpk` solver, but know the calculations may take longer (and the benchmark problem will not solve at all). If you do have the `gurobi` solver, edit the location of the executable to match the location on your computer.
# + executionInfo={"elapsed": 529, "status": "ok", "timestamp": 1603817900912, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="eY36JajosjKK" pycharm={}
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
import pandas as pd
import shutil
import sys
import os.path
if not shutil.which("pyomo"):
# !pip install -q pyomo
assert(shutil.which("pyomo"))
if not (shutil.which("cbc") or os.path.isfile("cbc")):
if "google.colab" in sys.modules:
# !apt-get install -y -qq coinor-cbc
else:
try:
# !conda install -c conda-forge coincbc
except:
pass
assert(shutil.which("cbc") or os.path.isfile("cbc"))
from pyomo.environ import *
from pyomo.gdp import *
# + [markdown] id="lffHCSpHshoF" pycharm={}
# ## Background
#
# A job shop consists of a set of distinct machines that process jobs. Each job is a series of tasks that require use of particular machines for known durations, and which must be completed in specified order. The job shop scheduling problem is to schedule the jobs on the machines to minimize the time necessary to process all jobs (i.e, the makespan) or some other metric of productivity. Job shop scheduling is one of the classic problems in Operations Research.
#
# Data consists of two tables. The first table is decomposition of the jobs into a series of tasks. Each task lists a job name, name of the required machine, and task duration. The second table list task pairs where the first task must be completed before the second task can be started. This formulation is quite general, but can also specify situations with no feasible solutions.
# + [markdown] id="N5nzDplWPHBV" pycharm={}
# ## Job shop example
#
# The following example of a job shop is from from <NAME>, <NAME>, <NAME>, "Applications of Optimization with Xpress-MP," Dash Optimization, 2000.
#
# In this example, there are three printed paper products that must pass through color printing presses in a particular order. The given data consists of a flowsheet showing the order in which each job passes through the color presses
#
# 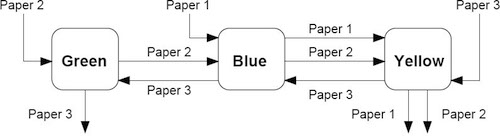
#
# and a table of data showing, in minutes, the amount of time each job requires on each machine.
#
# | Machine | Color | Paper 1 | Paper 2 | Paper 3 |
# | :-----: | :---: | :-----: | :-----: | :-----: |
# | 1 | Blue | 45 | 20 | 12 |
# | 2 | Green | - | 10 | 17 |
# | 3 | Yellow| 10 | 34 | 28 |
#
# What is the minimum amount of time (i.e, what is the makespan) for this set of jobs?
# + [markdown] id="6aYVOOd7PHBV" pycharm={}
# ## Task decomposition
#
# The first step in the analysis is to decompose the process into a series of tasks. Each task is a (job,machine) pair. Some tasks cannot start until a prerequisite task is completed.
#
# | Task (Job,Machine) | Duration | Prerequisite Task |
# | :----------------: | :------: | :---------------: |
# | (Paper 1, Blue) | 45 | - |
# | (Paper 1, Yellow) | 10 | (Paper 1,Blue) |
# | (Paper 2, Blue) | 20 | (Paper 2, Green) |
# | (Paper 2, Green) | 10 | - |
# | (Paper 2, Yellow) | 34 | (Paper 2, Blue) |
# | (Paper 3, Blue) | 12 | (Paper 3, Yellow) |
# | (Paper 3, Green) | 17 | (Paper 3, Blue) |
# | (Paper 3, Yellow) | 28 | - |
#
# We convert this to a JSON style representation where tasks are denoted by (Job,Machine) tuples in Python. The task data is stored in a Python dictionary indexed by (Job,Machine) tuples. The task data conists of a dictionary with duration ('dur') and (Job,Machine) pair for any prerequisite task.
# + executionInfo={"elapsed": 666, "status": "ok", "timestamp": 1603817901060, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="0k5vVxKIshoF" pycharm={}
TASKS = {
('Paper_1','Blue') : {'dur': 45, 'prec': None},
('Paper_1','Yellow') : {'dur': 10, 'prec': ('Paper_1','Blue')},
('Paper_2','Blue') : {'dur': 20, 'prec': ('Paper_2','Green')},
('Paper_2','Green') : {'dur': 10, 'prec': None},
('Paper_2','Yellow') : {'dur': 34, 'prec': ('Paper_2','Blue')},
('Paper_3','Blue') : {'dur': 12, 'prec': ('Paper_3','Yellow')},
('Paper_3','Green') : {'dur': 17, 'prec': ('Paper_3','Blue')},
('Paper_3','Yellow') : {'dur': 28, 'prec': None},
}
# + [markdown] id="F4nmNpnhshoI" pycharm={}
# ## Model formulation
#
# Each task is indexed by an ordered pair $(j,m)$ where $j$ is a job, and $m$ is a machine. Associated with each task is data describing the time needed to perform the task, and a preceeding task that must be completed before the index task can start.
#
# | Parameter | Description |
# | :-------- | :-----------|
# | $\text{dur}_{j,m}$ | Duration of task $(j,m)$ |
# | $\text{prec}_{j,m}$ | A task $(k,n) = \text{prec}_{j,m}$ that must be completed before task $(j,m)$|
#
# The choice of decision variables for this problem are key to modeling. We introduce $makespan$ as the time needed to complete all tasks. $makespan$ is a candidate objective function. Variable $start_{j,m}$ denotes the time when task $(j,m)$ begins.
#
# | Decision Variables | Description |
# | :-------- | :-----------|
# | $\text{makespan}$ | Completion of all jobs |
# | $\text{start}_{j,m}$ | Start time for task $(j,m)$ |
#
# The constraints include lower bounda on the start and an upper bound on the completion of each task $(j,m)$
#
# \begin{align}
# \text{start}_{j,m} & \geq 0\\
# \text{start}_{j,m}+\text{dur}_{j,m} & \leq \text{makespan}
# \end{align}
#
# Any preceeding tasks must be completed before task $(j,m)$ can start.
#
# \begin{align}
# \text{start}_{k,n}+\text{dur}_{k,n}\leq\text{start}_{j,m}\ \ \ \ \text{for } (k,n) =\text{prec}_{j,m}
# \end{align}
#
# Finally, for every task performed on machine $m$, there can be no overlap among those tasks. This leads to a set of pair-wise disjunctive constraints for each machine.
#
# \begin{align}
# \left[\text{start}_{j,m}+\text{dur}_{j,m} \leq \text{start}_{k,m}\right] \vee \left[\text{start}_{k,m}+\text{dur}_{k,m} \leq \text{start}_{j,m}\right]
# \end{align}
#
# avoids conflicts for use of the same machine.
# + [markdown] id="HdNWs72ZPHBY" pycharm={}
# ## Pyomo implementation
#
# The job shop scheduling problem is implemented below in Pyomo. The implementation consists of of a function JobShopModel(TASKS) that accepts a dictionary of tasks and returns a Pyomo model.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 658, "status": "ok", "timestamp": 1603817901061, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="2nJ72N-rshoI" outputId="6786b332-59e7-4c44-83e1-1341657c5942" pycharm={}
def jobshop_model(TASKS):
model = ConcreteModel()
# tasks is a two dimensional set of (j,m) constructed from the dictionary keys
model.TASKS = Set(initialize = TASKS.keys(), dimen=2)
# the set of jobs is constructed from a python set
model.JOBS = Set(initialize = list(set([j for (j,m) in model.TASKS])))
# set of machines is constructed from a python set
model.MACHINES = Set(initialize = list(set([m for (j,m) in model.TASKS])))
# the order of tasks is constructed as a cross-product of tasks and filtering
model.TASKORDER = Set(initialize = model.TASKS * model.TASKS, dimen=4,
filter = lambda model, j, m, k, n: (k,n) == TASKS[(j,m)]['prec'])
# the set of disjunctions is cross-product of jobs, jobs, and machines
model.DISJUNCTIONS = Set(initialize = model.JOBS * model.JOBS * model.MACHINES, dimen=3,
filter = lambda model, j, k, m: j < k and (j,m) in model.TASKS and (k,m) in model.TASKS)
# load duration data into a model parameter for later access
model.dur = Param(model.TASKS, initialize=lambda model, j, m: TASKS[(j,m)]['dur'])
# establish an upper bound on makespan
ub = sum([model.dur[j, m] for (j,m) in model.TASKS])
# create decision variables
model.makespan = Var(bounds=(0, ub))
model.start = Var(model.TASKS, bounds=(0, ub))
model.objective = Objective(expr = model.makespan, sense = minimize)
model.finish = Constraint(model.TASKS, rule=lambda model, j, m:
model.start[j,m] + model.dur[j,m] <= model.makespan)
model.preceding = Constraint(model.TASKORDER, rule=lambda model, j, m, k, n:
model.start[k,n] + model.dur[k,n] <= model.start[j,m])
model.disjunctions = Disjunction(model.DISJUNCTIONS, rule=lambda model,j,k,m:
[model.start[j,m] + model.dur[j,m] <= model.start[k,m],
model.start[k,m] + model.dur[k,m] <= model.start[j,m]])
TransformationFactory('gdp.hull').apply_to(model)
return model
jobshop_model(TASKS)
# + colab={"base_uri": "https://localhost:8080/", "height": 697} executionInfo={"elapsed": 1025, "status": "ok", "timestamp": 1603817901438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="VtQDkBmBPHBc" outputId="649f46f0-31f8-4113-e667-858fee72a315" pycharm={}
def jobshop_solve(model):
SolverFactory('cbc').solve(model)
results = [{'Job': j,
'Machine': m,
'Start': model.start[j, m](),
'Duration': model.dur[j,m],
'Finish': model.start[(j, m)]() + model.dur[j,m]}
for j,m in model.TASKS]
return results
def jobshop(TASKS):
return jobshop_solve(jobshop_model(TASKS))
results = jobshop(TASKS)
results
# + [markdown] id="7uDCIyqpshoO" pycharm={}
# ## Printing schedules
# + colab={"base_uri": "https://localhost:8080/", "height": 425} executionInfo={"elapsed": 1015, "status": "ok", "timestamp": 1603817901439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="ZmtO_2EeshoO" outputId="fecdce9c-ddca-4aad-d7fe-c3d8fcebde8a" pycharm={}
schedule = pd.DataFrame(results)
print('\nSchedule by Job')
print(schedule.sort_values(by=['Job','Start']).set_index(['Job', 'Machine']))
print('\nSchedule by Machine')
print(schedule.sort_values(by=['Machine','Start']).set_index(['Machine', 'Job']))
# + [markdown] id="EBvet1ACshoS" pycharm={}
# ## Visualizing Results with Gantt Charts
# + colab={"base_uri": "https://localhost:8080/", "height": 477} executionInfo={"elapsed": 1586, "status": "ok", "timestamp": 1603817902020, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="RtQ5NWPTshoU" outputId="bab2addd-0b90-45ef-d5eb-e65eb43d823f" pycharm={}
def visualize(results):
schedule = pd.DataFrame(results)
JOBS = sorted(list(schedule['Job'].unique()))
MACHINES = sorted(list(schedule['Machine'].unique()))
makespan = schedule['Finish'].max()
bar_style = {'alpha':1.0, 'lw':25, 'solid_capstyle':'butt'}
text_style = {'color':'white', 'weight':'bold', 'ha':'center', 'va':'center'}
colors = mpl.cm.Dark2.colors
schedule.sort_values(by=['Job', 'Start'])
schedule.set_index(['Job', 'Machine'], inplace=True)
fig, ax = plt.subplots(2,1, figsize=(12, 5+(len(JOBS)+len(MACHINES))/4))
for jdx, j in enumerate(JOBS, 1):
for mdx, m in enumerate(MACHINES, 1):
if (j,m) in schedule.index:
xs = schedule.loc[(j,m), 'Start']
xf = schedule.loc[(j,m), 'Finish']
ax[0].plot([xs, xf], [jdx]*2, c=colors[mdx%7], **bar_style)
ax[0].text((xs + xf)/2, jdx, m, **text_style)
ax[1].plot([xs, xf], [mdx]*2, c=colors[jdx%7], **bar_style)
ax[1].text((xs + xf)/2, mdx, j, **text_style)
ax[0].set_title('Job Schedule')
ax[0].set_ylabel('Job')
ax[1].set_title('Machine Schedule')
ax[1].set_ylabel('Machine')
for idx, s in enumerate([JOBS, MACHINES]):
ax[idx].set_ylim(0.5, len(s) + 0.5)
ax[idx].set_yticks(range(1, 1 + len(s)))
ax[idx].set_yticklabels(s)
ax[idx].text(makespan, ax[idx].get_ylim()[0]-0.2, "{0:0.1f}".format(makespan), ha='center', va='top')
ax[idx].plot([makespan]*2, ax[idx].get_ylim(), 'r--')
ax[idx].set_xlabel('Time')
ax[idx].grid(True)
fig.tight_layout()
visualize(results)
# + [markdown] id="JMWyAp2MshoW" pycharm={}
# ## Application to the scheduling of batch processes
#
# We will now turn our attention to the application of the job shop scheduling problem to the short term scheduling of batch processes. We illustrate these techniques using Example II from Dunn (2013).
#
# 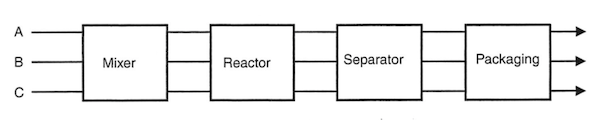
#
# | Recipe | Mixer | Reactor | Separator | Packaging |
# | :-----: | :---: | :-----: | :-------: | :-------: |
# | A | 1.0 | 5.0 | 4.0 | 1.5 |
# | B | - | - | 4.5 | 1.0 |
# | C | - | 3.0 | 5.0 | 1.5 |
# + [markdown] id="FllAPz4KshoX" pycharm={}
# ### Single product strategies
#
# Before going further, we create a function to streamline the generation of the TASKS dictionary.
# + colab={"base_uri": "https://localhost:8080/", "height": 459} executionInfo={"elapsed": 2006, "status": "ok", "timestamp": 1603817902449, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="xfurTCWyshoY" outputId="d027a1a6-e465-426e-b800-4c5ff3522443" pycharm={}
def recipe_to_tasks(jobs, machines, durations):
TASKS = {}
for j in jobs:
prec = (None,None)
for m,d in zip(machines,durations):
task = (j,m)
if prec == (None,None):
TASKS.update({(j,m): {'dur': d, 'prec': None}})
else:
TASKS.update({(j,m): {'dur': d, 'prec': prec}})
prec = task
return TASKS
recipeA = recipe_to_tasks('A', ['Mixer', 'Reactor', 'Separator', 'Packaging'], [1, 5, 4, 1.5])
visualize(jobshop(recipeA))
# + colab={"base_uri": "https://localhost:8080/", "height": 423} executionInfo={"elapsed": 2441, "status": "ok", "timestamp": 1603817902899, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="QndOwo19shoc" outputId="65bcc252-7e07-409c-ab29-23761ff73567" pycharm={}
recipeB = recipe_to_tasks('B', ['Separator', 'Packaging'], [4.5, 1])
visualize(jobshop(recipeB))
# + colab={"base_uri": "https://localhost:8080/", "height": 441} executionInfo={"elapsed": 2809, "status": "ok", "timestamp": 1603817903280, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="VR2QKxzRshof" outputId="a7ed824a-4176-462a-e613-7c3b46f6ea7c" pycharm={}
recipeC = recipe_to_tasks('C', ['Separator', 'Reactor', 'Packaging'], [5, 3, 1.5])
visualize(jobshop(recipeC))
# + [markdown] id="s1U47NNXshoj" pycharm={}
# ### Multiple Overlapping tasks
#
# Let's now consider an optimal scheduling problem where we are wish to make two batches of Product A.
# + colab={"base_uri": "https://localhost:8080/", "height": 530} executionInfo={"elapsed": 4166, "status": "ok", "timestamp": 1603817904648, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="EGU9wmLoshoj" outputId="04c36086-113c-42aa-de15-ad126e748291" pycharm={}
TASKS = recipe_to_tasks(['A1','A2','A3', 'A4'],['Mixer','Reactor','Separator','Packaging'],[1,5,4,1.5])
results = jobshop(TASKS)
visualize(results)
print("Makespan =", max([task['Finish'] for task in results]))
# + [markdown] id="j9-oR-tIshop" pycharm={}
# Earlier we found it tood 11.5 hours to produce one batch of product A. As we see here, we can produce a second batch with only 5.0 additional hours because some of the tasks overlap. The overlapping of tasks is the key to gaining efficiency in batch processing facilities.
#
# Let's next consider production of a single batch each of products A, B, and C.
# + colab={"base_uri": "https://localhost:8080/", "height": 665} executionInfo={"elapsed": 4593, "status": "ok", "timestamp": 1603817905087, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="mnntHP1jshoq" outputId="a657efa0-2122-48d5-f3c7-3dd19a55f5e9" pycharm={}
# update is used to append dictionaries
TASKS = recipeA
TASKS.update(recipeB)
TASKS.update(recipeC)
for k, v in TASKS.items():
print(k, v)
results = jobshop(TASKS)
visualize(results)
print("Makespan =", max([task['Finish'] for task in results]))
# + [markdown] id="daGJ6KE7shot" pycharm={}
# The individual production of A, B, and C required 11.5, 5.5, and 9.5 hours, respectively, for a total of 25.5 hours. As we see here, by scheduling the production simultaneously, we can get all three batches done in just 15 hours.
#
# As we see below, each additional set of three products takes an additionl 13 hours. So there is considerable efficiency gained by scheduling over longer intervals whenever possible.
# + colab={"base_uri": "https://localhost:8080/", "height": 566} executionInfo={"elapsed": 14311, "status": "ok", "timestamp": 1603817914815, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="tmmIFHETshov" outputId="189eb366-cb64-4da1-d5ad-0ee5317c8fd4" pycharm={}
TASKS = recipe_to_tasks(['A1','A2'],['Mixer','Reactor','Separator','Packaging'],[1,5,4,1.5])
TASKS.update(recipe_to_tasks(['B1','B2'],['Separator','Packaging'],[4.5,1]))
TASKS.update(recipe_to_tasks(['C1','C2'],['Separator','Reactor','Packaging'],[5,3,1.5]))
results = jobshop(TASKS)
visualize(results)
print("Makespan =", max([task['Finish'] for task in results]))
# + [markdown] id="68wdMOQ9sho0" pycharm={}
# ### Adding time for unit clean out
#
# A common feature of batch unit operations is a requirement that equipment be cleaned prior to reuse.
#
# In most cases the time needed for clean out would be specific to the equipment and product. But for the purposes this notebook, we implement can implement a simple clean out policy with a single non-negative parameter $t_{clean} \geq 0$ which, if specified, requires a period no less than $t_{clean}$ between the finish of one task and the start of another on every piece of equipment.
#
# This policy is implemented by modifying the usual disjunctive constraints to avoid machine conflicts to read
#
# \begin{align}
# \left[\text{start}_{j,m}+\text{dur}_{j,m} + t_{clean} \leq \text{start}_{k,m}\right] \vee \left[\text{start}_{k,m}+\text{dur}_{k,m} + t_{clean} \leq \text{start}_{j,m}\right]
# \end{align}
#
# For this purpose, we write a new JobShopModel_Clean
# + colab={"base_uri": "https://localhost:8080/", "height": 566} executionInfo={"elapsed": 23784, "status": "ok", "timestamp": 1603817924300, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="HjvMPceePHBz" outputId="b282c450-7024-47ec-faa2-b428ae65f24e"
def jobshop_model_clean(TASKS, tclean=0):
model = ConcreteModel()
# tasks is a two dimensional set of (j,m) constructed from the dictionary keys
model.TASKS = Set(initialize = TASKS.keys(), dimen=2)
# the set of jobs is constructed from a python set
model.JOBS = Set(initialize = list(set([j for (j,m) in model.TASKS])))
# set of machines is constructed from a python set
model.MACHINES = Set(initialize = list(set([m for (j,m) in model.TASKS])))
# the order of tasks is constructed as a cross-product of tasks and filtering
model.TASKORDER = Set(initialize = model.TASKS * model.TASKS, dimen=4,
filter = lambda model, j, m, k, n: (k,n) == TASKS[(j,m)]['prec'])
# the set of disjunctions is cross-product of jobs, jobs, and machines
model.DISJUNCTIONS = Set(initialize = model.JOBS * model.JOBS * model.MACHINES, dimen=3,
filter = lambda model, j, k, m: j < k and (j,m) in model.TASKS and (k,m) in model.TASKS)
# load duration data into a model parameter for later access
model.dur = Param(model.TASKS, initialize=lambda model, j, m: TASKS[(j,m)]['dur'])
# establish an upper bound on makespan
ub = sum([model.dur[j,m] for (j,m) in model.TASKS])
model.makespan = Var(bounds=(0, ub))
model.start = Var(model.TASKS, bounds=(0, ub))
model.objective = Objective(expr = model.makespan, sense = minimize)
model.finish = Constraint(model.TASKS, rule=lambda model, j, m:
model.start[j,m] + model.dur[j,m] <= model.makespan)
model.preceding = Constraint(model.TASKORDER, rule=lambda model, j, m, k, n:
model.start[k,n] + model.dur[k,n] <= model.start[j,m])
model.disjunctions = Disjunction(model.DISJUNCTIONS, rule=lambda model,j,k,m:
[model.start[j,m] + model.dur[j,m] + tclean <= model.start[k,m],
model.start[k,m] + model.dur[k,m] + tclean <= model.start[j,m]])
TransformationFactory('gdp.hull').apply_to(model)
return model
model = jobshop_model_clean(TASKS, tclean=0.5)
results = jobshop_solve(model)
visualize(results)
print("Makespan =", max([task['Finish'] for task in results]))
# + [markdown] id="HrHUYrQUsho7" pycharm={}
# ### Adding a zero wait policy
#
# One of the issues in the use of job shop scheduling for chemical process operations are situations where there it is not possible to store intermediate materials. If there is no way to store intermediates, either in the processing equipment or in external vessels, then a **zero-wait** policy may be required.
#
# A zero-wait policy requires subsequent processing machines to be available immediately upon completion of any task. To implement this policy, the usual precident sequencing constraint of a job shop scheduling problem, i.e.,
#
# \begin{align*}
# \text{start}_{k,n}+\text{Dur}_{k,n} \leq \text{start}_{j,m}\ \ \ \ \text{for } (k,n) =\text{Prec}_{j,m}
# \end{align*}
#
# is changed to
#
# \begin{align*}
# \text{start}_{k,n}+\text{Dur}_{k,n} = \text{start}_{j,m}\ \ \ \ \text{for } (k,n) =\text{Prec}_{j,m}\text{ and ZW is True}
# \end{align*}
#
# if the zero-wait policy is in effect.
#
# While this could be implemented on an equipment or product specific basis, here we add an optional ZW flag to the JobShop function that, by default, is set to False.
# + colab={"base_uri": "https://localhost:8080/", "height": 566} executionInfo={"elapsed": 25811, "status": "ok", "timestamp": 1603817926340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="owKSMRnHPHB2" outputId="495434fa-efb5-4111-8eba-6df5974387cb"
def jobshop_model_clean_zw(TASKS, tclean=0, ZW=False):
model = ConcreteModel()
# tasks is a two dimensional set of (j,m) constructed from the dictionary keys
model.TASKS = Set(initialize = TASKS.keys(), dimen=2)
# the set of jobs is constructed from a python set
model.JOBS = Set(initialize = list(set([j for (j,m) in model.TASKS])))
# set of machines is constructed from a python set
model.MACHINES = Set(initialize = list(set([m for (j,m) in model.TASKS])))
# the order of tasks is constructed as a cross-product of tasks and filtering
model.TASKORDER = Set(initialize = model.TASKS * model.TASKS, dimen=4,
filter = lambda model, j, m, k, n: (k,n) == TASKS[(j,m)]['prec'])
# the set of disjunctions is cross-product of jobs, jobs, and machines
model.DISJUNCTIONS = Set(initialize = model.JOBS * model.JOBS * model.MACHINES, dimen=3,
filter = lambda model, j, k, m: j < k and (j,m) in model.TASKS and (k,m) in model.TASKS)
# load duration data into a model parameter for later access
model.dur = Param(model.TASKS, initialize=lambda model, j, m: TASKS[(j,m)]['dur'])
# establish an upper bound on makespan
ub = sum([model.dur[j,m] for (j,m) in model.TASKS])
# to implement a zero-wait policy
model.bigM = Param(initialize=ub if ZW else 0)
model.makespan = Var(bounds=(0, ub))
model.start = Var(model.TASKS, bounds=(0, ub))
model.objective = Objective(expr = model.makespan, sense = minimize)
model.finish = Constraint(model.TASKS, rule=lambda model, j, m:
model.start[j,m] + model.dur[j,m] <= model.makespan)
def _preceding(model, j, m, k, n):
if ZW:
return model.start[k,n] + model.dur[k,n] == model.start[j,m]
else:
return model.start[k,n] + model.dur[k,n] <= model.start[j,m]
model.preceding = Constraint(model.TASKORDER, rule=_preceding)
model.disjunctions = Disjunction(model.DISJUNCTIONS, rule=lambda model,j,k,m:
[model.start[j,m] + model.dur[j,m] + tclean <= model.start[k,m],
model.start[k,m] + model.dur[k,m] + tclean <= model.start[j,m]])
TransformationFactory('gdp.hull').apply_to(model)
return model
model = jobshop_model_clean_zw(TASKS, tclean=0.5, ZW=True)
results = jobshop_solve(model)
visualize(results)
print("Makespan =", max([task['Finish'] for task in results]))
# + [markdown] id="AFy7ULi_shpA" pycharm={}
# ## Solving the LA19 benchmark problem with NEOS
#
# The file [`jobshop1.txt` contains 82 benchmark problems](http://people.brunel.ac.uk/~mastjjb/jeb/orlib/files/jobshop1.txt) from a well-known collection of job shop scheduling problems in the [OR-Library maintained by <NAME>](http://people.brunel.ac.uk/~mastjjb/jeb/info.html). The data format for each example consists of a single line for each job. The data on each line is a sequence of (machine number, time) pairs showing the order in which machines process each job.
#
# LA19 is a benchmark problem for job shop scheduling introduced by Lawrence in 1984, and a solution presented by Cook and Applegate in 1991. The following cell may take many minutes to hours to run, depending on the choice of solver and hardware. To run, uncomment the the last lines in the cell.
# + executionInfo={"elapsed": 25802, "status": "ok", "timestamp": 1603817926341, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V<KEY>", "userId": "09038942003589296665"}, "user_tz": 240} id="RAxBT2KeshpC" pycharm={}
data = """
2 44 3 5 5 58 4 97 0 9 7 84 8 77 9 96 1 58 6 89
4 15 7 31 1 87 8 57 0 77 3 85 2 81 5 39 9 73 6 21
9 82 6 22 4 10 3 70 1 49 0 40 8 34 2 48 7 80 5 71
1 91 2 17 7 62 5 75 8 47 4 11 3 7 6 72 9 35 0 55
6 71 1 90 3 75 0 64 2 94 8 15 4 12 7 67 9 20 5 50
7 70 5 93 8 77 2 29 4 58 6 93 3 68 1 57 9 7 0 52
6 87 1 63 4 26 5 6 2 82 3 27 7 56 8 48 9 36 0 95
0 36 5 15 8 41 9 78 3 76 6 84 4 30 7 76 2 36 1 8
5 88 2 81 3 13 6 82 4 54 7 13 8 29 9 40 1 78 0 75
9 88 4 54 6 64 7 32 0 52 2 6 8 54 5 82 3 6 1 26
"""
TASKS = {}
for job, line in enumerate(data.splitlines()[1:]):
nums = line.split()
prec = None
for m, dur in zip(nums[::2], nums[1::2]):
task = (f"J{job}",f"M{m}")
TASKS[task] = {'dur':int(dur), 'prec':prec}
prec = task
#pd.DataFrame(TASKS).T
# + [markdown] id="yUXuahgpPHB6"
# Depending on the choice of solver, this benchmark example may require from minutes to hours of computational effort on a laptop. An alternative to solving on a laptop is to submit the job to [NEOS](https://neos-server.org/neos/), a free internet-based service for solving optimization problems hosted by the University of Wisconsin and utilizing high performance servers at locations across the globe.
#
# The following cell shows how to solve a model using [CPLEX](https://www.ibm.com/analytics/cplex-optimizer), a high performance commericial solver, on NEOS. The solution may take several minutes, and depends on the current length of the NEOS job queue.
# + colab={"base_uri": "https://localhost:8080/", "height": 729} executionInfo={"elapsed": 89691, "status": "ok", "timestamp": 1603817990240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_n8V7bVINy02QRuRgOoMo11Ri7NKU3OUKdC1bkQ=s64", "userId": "09038942003589296665"}, "user_tz": 240} id="p-dA89fhPHB6" outputId="766aa8e7-3b6a-4547-ad95-34250b3df143"
def jobshop_solve_neos(model):
solver_manager = SolverManagerFactory('neos')
solver_manager.solve(model, opt='cplex')
results = [{'Job': j,
'Machine': m,
'Start': model.start[j, m](),
'Duration': model.dur[j,m],
'Finish': model.start[(j, m)]() + model.dur[j,m]}
for j,m in model.TASKS]
return results
model = jobshop_model(TASKS)
results = jobshop_solve_neos(model)
visualize(results)
# + [markdown] id="mINloyXxPHB8"
# ## References
#
# * <NAME>, and <NAME>. ["A computational study of the job-shop scheduling problem."](https://doi.org/10.1287/ijoc.3.2.149) ORSA Journal on computing 3, no. 2 (1991): 149-156. [pdf available](http://www.math.uwaterloo.ca/~bico/papers/jobshop.pdf)
# * Beasley, <NAME>. ["OR-Library: distributing test problems by electronic mail."](https://www.jstor.org/stable/pdf/2582903.pdf?casa_token=RWUXQ6e2VngAAAAA:<KEY>) Journal of the operational research society 41, no. 11 (1990): 1069-1072. [OR-Library](http://people.brunel.ac.uk/~mastjjb/jeb/info.html)
# * <NAME>, <NAME>, and <NAME>. ["Applications of optimization with Xpress-MP."](http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.69.9634&rep=rep1&type=pdf) contract (1999): 00034.
# * Manne, <NAME>. ["On the job-shop scheduling problem."](https://doi.org/10.1287/opre.8.2.219) Operations Research 8, no. 2 (1960): 219-223.
# + [markdown] id="9ubGM7r1PHB8"
# ## Exercises
# + [markdown] id="gagPe62ZPHB9"
# ### Task specific cleanout
#
# Clean out operations are often slow and time consuming. Further more, the time required to perform a clean out frequently depends on the type of machine, and the task performed by the machine. For this exercise, create a data format to include task-specific clean out times, and model the job shop model to accomodate this additional informaton.
# + [markdown] id="g21rYpmhPHB9"
# ### Computational impact of a zero-wait policy
#
# Repeat the benchmark problem calculation, but with a zero-wait policy. Does the execution time increase or descrease as a consequence of specifying zero-wait?
| notebooks/03/03.03-Job-Shop-Scheduling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.1 64-bit (''venv'': venv)'
# name: python3
# ---
# # Imports
import os
import pickle
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import confusion_matrix
# # Test Models on MLS
# ## Load test data
# +
features = 'Features_F0_MFCCs'
dataset = 'CETUC'
project_root = os.path.dirname(os.path.dirname(os.getcwd()))
test_MLS = pd.read_csv(os.path.join(project_root, 'data', 'MLS_split',f'{features}_data.csv'))
if features == 'Features':
X_test = test_MLS[['nobs', 'mean', 'skew', 'kurtosis', 'median', 'mode', 'std', 'low', 'peak', 'q25', 'q75', 'iqr']].copy()
Y_test = test_MLS[['Gender']].copy()#.values.ravel()
elif features == 'MFCCs':
X_test = test_MLS[['MFCC_1', 'MFCC_2', 'MFCC_3', 'MFCC_4', 'MFCC_5', 'MFCC_6', 'MFCC_7', 'MFCC_8', 'MFCC_9', 'MFCC_10',
'MFCC_11', 'MFCC_12', 'MFCC_13', 'MFCC_14', 'MFCC_15', 'MFCC_16', 'MFCC_17', 'MFCC_18', 'MFCC_19', 'MFCC_20']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
elif features == 'Features_MFCCs':
X_test = test_MLS[['nobs', 'mean', 'skew', 'kurtosis', 'median', 'mode', 'std', 'low', 'peak', 'q25', 'q75', 'iqr',
'MFCC_1', 'MFCC_2', 'MFCC_3', 'MFCC_4', 'MFCC_5', 'MFCC_6', 'MFCC_7', 'MFCC_8', 'MFCC_9', 'MFCC_10',
'MFCC_11', 'MFCC_12', 'MFCC_13', 'MFCC_14', 'MFCC_15', 'MFCC_16', 'MFCC_17', 'MFCC_18', 'MFCC_19', 'MFCC_20']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
elif features == 'F0':
X_test = test_MLS[['nobs_pitch', 'mean_pitch', 'skew_pitch', 'kurtosis_pitch', 'median_pitch', 'mode_pitch', 'std_pitch', 'low_pitch', 'peak_pitch', 'q25_pitch', 'q75_pitch', 'iqr_pitch']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
elif features == 'F0_MFCCs':
X_test = test_MLS[['nobs_pitch', 'mean_pitch', 'skew_pitch', 'kurtosis_pitch', 'median_pitch', 'mode_pitch', 'std_pitch', 'low_pitch', 'peak_pitch', 'q25_pitch', 'q75_pitch', 'iqr_pitch',
'MFCC_1', 'MFCC_2', 'MFCC_3', 'MFCC_4', 'MFCC_5', 'MFCC_6', 'MFCC_7', 'MFCC_8', 'MFCC_9', 'MFCC_10',
'MFCC_11', 'MFCC_12', 'MFCC_13', 'MFCC_14', 'MFCC_15', 'MFCC_16', 'MFCC_17', 'MFCC_18', 'MFCC_19', 'MFCC_20']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
elif features == 'Features_F0':
X_test = test_MLS[['nobs', 'mean', 'skew', 'kurtosis', 'median', 'mode', 'std', 'low', 'peak', 'q25', 'q75', 'iqr',
'nobs_pitch', 'mean_pitch', 'skew_pitch', 'kurtosis_pitch', 'median_pitch', 'mode_pitch', 'std_pitch', 'low_pitch', 'peak_pitch', 'q25_pitch', 'q75_pitch', 'iqr_pitch']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
elif features == 'Features_F0_MFCCs':
X_test = test_MLS[['nobs', 'mean', 'skew', 'kurtosis', 'median', 'mode', 'std', 'low', 'peak', 'q25', 'q75', 'iqr',
'nobs_pitch', 'mean_pitch', 'skew_pitch', 'kurtosis_pitch', 'median_pitch', 'mode_pitch', 'std_pitch', 'low_pitch', 'peak_pitch', 'q25_pitch', 'q75_pitch', 'iqr_pitch',
'MFCC_1', 'MFCC_2', 'MFCC_3', 'MFCC_4', 'MFCC_5', 'MFCC_6', 'MFCC_7', 'MFCC_8', 'MFCC_9', 'MFCC_10',
'MFCC_11', 'MFCC_12', 'MFCC_13', 'MFCC_14', 'MFCC_15', 'MFCC_16', 'MFCC_17', 'MFCC_18', 'MFCC_19', 'MFCC_20']].copy()
Y_test = test_MLS[['Gender']].copy().values.ravel()
# -
X_test
# scaler = StandardScaler()
scaler = pickle.load(open(os.path.join(project_root, 'models', dataset, features, 'scaler.pkl'), 'rb'))
X_test = pd.DataFrame(scaler.transform(X_test), columns=X_test.columns)
# ## Load Models
# +
filename = os.path.join(project_root, 'models', dataset, features, 'DecisionTree.sav')
tree = pickle.load(open(filename, 'rb'))
print("\nDecision Tree")
print("Accuracy on test set: {:.3f}".format(tree.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, tree.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
filename = os.path.join(project_root, 'models', dataset, features, 'RandomForest.sav')
forest = pickle.load(open(filename, 'rb'))
print("\nRandom Forests")
print("Accuracy on test set: {:.3f}".format(forest.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, forest.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
filename = os.path.join(project_root, 'models', dataset, features, 'GradientBoosting.sav')
gbrt = pickle.load(open(filename, 'rb'))
print("\nGradient Boosting")
print("Accuracy on test set: {:.3f}".format(gbrt.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, gbrt.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
# +
filename = os.path.join(project_root, 'models', dataset, features, 'LogisticRegression.sav')
lgr = pickle.load(open(filename, 'rb'))
print("\nLogisticRegression")
print("Accuracy on test set: {:.3f}".format(lgr.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, lgr.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
filename = os.path.join(project_root, 'models', dataset, features, 'SVM.sav')
svm = pickle.load(open(filename, 'rb'))
print("\nSupport Vector Machine")
print("Accuracy on test set: {:.3f}".format(svm.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, svm.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
filename = os.path.join(project_root, 'models', dataset, features, 'MLP.sav')
mlp = pickle.load(open(filename, 'rb'))
print("\nMultilayer Perceptron")
print("Accuracy on test set: {:.3f}".format(mlp.score(X_test, Y_test)))
cm = confusion_matrix(Y_test, mlp.predict(X_test), labels=[1, 0])
precision = cm[0][0]/(cm[0][0]+cm[1][0])
recall = cm[0][0]/(cm[0][0]+cm[0][1])
# print(f"Confusion Matrix:\n {cm}")
print(f"Precision on test set: {precision:.3f}")
print(f"Recall on test set: {recall:.3f}")
print(f"F1-score on test set: {(2 * (precision * recall) / (precision + recall)):.3f}")
# -
| notebooks/MLS_split/MLS_split_test_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="zodPJNT3TsLv" colab_type="code" outputId="4094e0d7-b0bf-4f1c-ebce-20401fc79eb1" colab={"base_uri": "https://localhost:8080/", "height": 88}
# make sure we have the alpha version of tensorflow 2.0 ready to go in this
# Collab instance
# !pip install -q tensorflow-gpu==2.0.0-alpha0
# + id="08ShAZk6iMcH" colab_type="code" outputId="d0c0930b-f766-4fc3-fba5-e97513377802" colab={"base_uri": "https://localhost:8080/", "height": 160}
# manually download the translation data file
# !curl -O http://www.manythings.org/anki/deu-eng.zip
# !mkdir deu-eng
# %cd /content/deu-eng
# !unzip ../deu-eng.zip
# %cd /content
# + id="Ml_aL6AEeDa1" colab_type="code" colab={}
import tensorflow as tf # tensorflow of course
tf.debugging.set_log_device_placement(True)
import io # io to read in our file
import re # we use regexp to replace unwated characters in the text
import unicodedata # input text might be unicode, we use this to convert to ascii
from sklearn.model_selection import train_test_split # we use this to split into training and test
import time
import os
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
# + id="aAjTDKZnh4Xy" colab_type="code" colab={}
# helper function to convert unicode characters to ascii, replaces ü with u etc
def unicode_to_ascii(w):
return ''.join(c for c in unicodedata.normalize('NFD', w) if unicodedata.category(c) != 'Mn')
# convert the incoming text to ascii, surround punctuation marks with spaces,
# remove multiple spaces and surround sentences with <start> <end> markers
def process_text(text):
w = unicode_to_ascii(text.lower().strip())
w = re.sub(r"([?.!,¿])", r" \1 ", w)
w = re.sub(r'[" "]+', " ", w)
w = re.sub(r'[^a-zA-Z?.!,¿]+', ' ', w)
w = w.rstrip().strip()
return '<start> ' + w + ' <end>'
# input and target language sentences are separated by a tab, split and process
def load_dataset(path, num_examples=None):
lines = io.open(path, encoding='UTF-8').read().strip().split('\n')
if num_examples == None:
print("Loading {} examples from the dataset.".format(len(lines)))
return zip(*[[process_text(w) for w in l.split('\t')] for l in lines[:num_examples]])
# + id="VQhjzvCwrOJz" colab_type="code" colab={}
# we use Keras' preprocessing library to embed the text
# no magic is happening here, this just tokenizes the text by splitting on spaces
# then it creates a dictionary by counting word frequencies
# afterwards the words are mapped to integers (ordered by frequency) that we can
# then use in our model
def tokenize(text):
tokenizer = tf.keras.preprocessing.text.Tokenizer(filters='')
tokenizer.fit_on_texts(text)
sequences = tokenizer.texts_to_sequences(text)
sequences = tf.keras.preprocessing.sequence.pad_sequences(sequences, padding='post')
return sequences, tokenizer
# load the dataset and tokenize
def prepare_dataset(path, num_examples=None):
target_lang, input_lang = load_dataset(path, num_examples)
input_tensor, input_lang_tokenizer = tokenize(input_lang)
target_tensor, target_lang_tokenizer = tokenize(target_lang)
return input_tensor, target_tensor, input_lang_tokenizer, target_lang_tokenizer
# helper function to check length of elements in tensor (sentences)
def max_length(tensor):
return max([len(t) for t in tensor])
# + id="-J3HebZrvpko" colab_type="code" colab={}
input_tensor, target_tensor, input_lang_tokenizer, target_lang_tokenizer = prepare_dataset('/content/deu-eng/deu.txt',10000)
# + id="o7WHIUvJVeyF" colab_type="code" outputId="3298d290-319a-4957-c99b-4220b8533ce7" colab={"base_uri": "https://localhost:8080/", "height": 71}
max_length_target, max_length_input = max_length(target_tensor), max_length(input_tensor)
print('Max length input: {}\nMax length target: {}\n'.format(max_length_input, max_length_target))
# + id="iYA5gzClWriD" colab_type="code" outputId="bc616287-e5ee-4ba5-eca2-44534ccaffe3" colab={"base_uri": "https://localhost:8080/", "height": 106}
# split into training and test set
input_tensor_train, input_tensor_val, target_tensor_train, target_tensor_val = train_test_split(input_tensor, target_tensor, test_size=0.2)
print('Input train #: {}\nTarget train #: {}\nInput val #: {}\nTarget val #: {}\n'.format(len(input_tensor_train), len(target_tensor_train), len(input_tensor_val), len(target_tensor_val)))
# + id="ZSEzvvSKWx4v" colab_type="code" outputId="4c0f6318-88bb-4454-cad9-69dabc155441" colab={"base_uri": "https://localhost:8080/", "height": 553}
# look up and print the actual words for a sequence of embedding integers
def wordify(tokenizer, tensor):
for t in tensor:
if t != 0:
print('{} ----> {}\n'.format(t, tokenizer.index_word[t]))
wordify(input_lang_tokenizer, input_tensor_train[0])
wordify(target_lang_tokenizer, target_tensor_train[0])
# + id="HnPONjyHbaE2" colab_type="code" outputId="7ad496d0-aa1a-4bf8-856b-59c14e776d8f" colab={"base_uri": "https://localhost:8080/", "height": 35}
# next we create a tf.Dataset so tensorflow can consume our prepared data
BATCH_SIZE = 16
steps_per_epoch = len(input_tensor_train)//BATCH_SIZE
dataset = tf.data.Dataset.from_tensor_slices((input_tensor_train, target_tensor_train)).shuffle(len(input_tensor_train))
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
vocab_input_size = len(input_lang_tokenizer.word_index)+1
vocab_target_size = len(target_lang_tokenizer.word_index)+1
EMBEDDING_DIM = 256
ENCODER_UNITS = 1024
vocab_input_size, vocab_target_size
# + id="NL_XohxRcrmp" colab_type="code" outputId="3cf8eb25-2944-403d-a5b9-9ad779fb8a0d" colab={"base_uri": "https://localhost:8080/", "height": 35}
a, b = next(iter(dataset))
a.shape, b.shape
# + id="ho0u2-uHc5lg" colab_type="code" colab={}
# at this point our dataset is ready to be consumed by our model ... which we have to build next
# first we start with the encoder part,
# this is a set of GRUs that encode the input sequence
# + id="n3u4Jlk2-1pY" colab_type="code" colab={}
class Encoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_size):
super(Encoder, self).__init__()
self.batch_size = batch_size
self.enc_units = enc_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.enc_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state=hidden)
return output, state
def initialize_hidden_state(self):
return tf.zeros((self.batch_size, self.enc_units))
# + id="wqQe8u4DG_Zg" colab_type="code" colab={}
encoder = Encoder(vocab_input_size, EMBEDDING_DIM, ENCODER_UNITS, BATCH_SIZE)
sample_hidden = encoder.initialize_hidden_state()
sample_input_batch, sample_target_batch = next(iter(dataset))
sample_output, sample_hidden = encoder(sample_input_batch, sample_hidden)
# + id="hwJOkxXbHUW6" colab_type="code" outputId="1c4b0b82-82cd-4818-e89e-547adacf910c" colab={"base_uri": "https://localhost:8080/", "height": 35}
sample_input_batch.shape, sample_output.shape
# + id="xM0_FP2qHZx6" colab_type="code" colab={}
class AttentionModel(tf.keras.Model):
def __init__(self, units):
super(AttentionModel,self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, query, values):
hidden_expanded = tf.expand_dims(query, 1) # turn (a,b) tensor into (a,1,b)
score = self.V(tf.nn.tanh(self.W1(values) + self.W2(hidden_expanded)))
attention_weights = tf.nn.softmax(score, axis=1)
context_vector = attention_weights * values
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
# + id="6didJgfnEidi" colab_type="code" colab={}
attention_layer = AttentionModel(10)
attention_result, attention_weights = attention_layer(sample_hidden, sample_output)
# + id="drqv00YOEyKO" colab_type="code" outputId="b0c9ee13-0d49-41c5-a16d-6356c601c4f9" colab={"base_uri": "https://localhost:8080/", "height": 35}
attention_result.shape, attention_weights.shape
# + id="F-WzV_5fE_V2" colab_type="code" colab={}
class Decoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_size):
super(Decoder, self).__init__()
self.batch_size = batch_size
self.dec_units = dec_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.dec_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc = tf.keras.layers.Dense(vocab_size)
self.attention = AttentionModel(self.dec_units)
def call(self, x, hidden, enc_output):
context_vector, attention_weights = self.attention(hidden, enc_output)
x = self.embedding(x)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
output, state = self.gru(x)
output = tf.reshape(output, (-1, output.shape[2]))
x = self.fc(output)
return x, state, attention_weights
# + id="vOLFLFLKls-9" colab_type="code" colab={}
# + id="zDSUiMx4NXmx" colab_type="code" colab={}
decoder = Decoder(vocab_input_size, EMBEDDING_DIM, ENCODER_UNITS, BATCH_SIZE)
# + id="aev4lvuZRDa6" colab_type="code" colab={}
sample_decoder_output, _, _ = decoder(tf.random.uniform((BATCH_SIZE,1)), sample_hidden, sample_output)
# + id="aQvsnNwdRPMF" colab_type="code" outputId="3b5cabbd-679e-4f4c-8b3c-0144060b762c" colab={"base_uri": "https://localhost:8080/", "height": 35}
sample_decoder_output.shape
# + id="J0s69cG8RSiW" colab_type="code" outputId="c53781e2-3f19-47f6-a7ec-51fba6047dd8" colab={"base_uri": "https://localhost:8080/", "height": 35}
tf.math.logical_not(tf.math.equal(tf.constant([0,1,0]),0))
# + id="6SJsZ9zmRjmT" colab_type="code" colab={}
# now we can define an optimizer and the loss function
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True,
reduction='none')
def loss_func(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
lossy = loss_object(real, pred)
mask = tf.cast(mask, dtype=lossy.dtype)
lossy *= mask
return tf.reduce_mean(lossy)
CHECKPOINT_DIR = './training_checkpoints'
checkpoint_prefix = os.path.join(CHECKPOINT_DIR,'ckpt')
checkpoint = tf.train.Checkpoint(optimizer=optimizer,
encoder=encoder,
decoder=decoder)
# + id="pCeeo7lPUO4E" colab_type="code" colab={}
# now we define the training loop
@tf.function
def train_step(inp, targ, enc_hidden):
loss = 0
with tf.GradientTape() as tape:
enc_output, enc_hidden = encoder(inp, enc_hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([target_lang_tokenizer.word_index['<start>']]*BATCH_SIZE,1)
for t in range(1, targ.shape[1]):
predictions, dec_hidden, _ = decoder(dec_input,
dec_hidden,
enc_output)
loss += loss_func(targ[:, t], predictions)
dec_input = tf.expand_dims(targ[:, t], 1)
batch_loss = (loss / int(targ.shape[1]))
variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(gradients,variables))
return batch_loss
# + id="tx1il5tMVamG" colab_type="code" outputId="88de789a-07c1-4109-d5ea-7e5d223dd096" colab={"base_uri": "https://localhost:8080/", "height": 1446}
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
enc_hidden = encoder.initialize_hidden_state()
total_loss = 0
for (batch, (inp, targ)) in enumerate(dataset.take(steps_per_epoch)):
batch_loss = train_step(inp, targ, enc_hidden)
total_loss += batch_loss
if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch+1,
batch,
batch_loss.numpy()))
if (epoch + 1) % 2 == 0:
checkpoint.save(file_prefix=checkpoint_prefix)
print('Epoch {} Loss {:.4f}'.format(epoch+1, total_loss/steps_per_epoch))
print('Time taken for 1 epoch {} sec\n'.format(time.time()-start))
# + id="l8jRh5UIVcEI" colab_type="code" colab={}
def evaluate(sentence):
attention_plot = np.zeros((max_length_target, max_length_input))
sentence = process_text(sentence)
inputs = [input_lang_tokenizer.word_index[i] for i in sentence.split(' ')]
inputs = tf.keras.preprocessing.sequence.pad_sequences([inputs],
maxlen=max_length_input,
padding='post')
inputs = tf.convert_to_tensor(inputs)
result = ''
hidden = [tf.zeros((1, ENCODER_UNITS))]
enc_out, enc_hidden = encoder(inputs, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([target_lang_tokenizer.word_index['<start>']], 0)
for t in range(max_length_target):
predictions, dec_hidden, attention_weights = decoder(dec_input,
dec_hidden,
enc_out)
attention_weights = tf.reshape(attention_weights, (-1,))
attention_plot[t] = attention_weights.numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result += target_lang_tokenizer.index_word[predicted_id] + ' '
if target_lang_tokenizer.index_word[predicted_id] == '<end>':
return result, sentence, attention_plot
dec_input = tf.expand_dims([predicted_id], 0)
return result, sentence, attention_plot
# + id="giDRqhxTeork" colab_type="code" colab={}
def plot_attention(attention, sentence, predicted_sentence):
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1,1,1)
ax.matshow(attention, cmap='viridis')
fontdict = {'fontsize': 14}
ax.set_xticklabels([''] + sentence, fontdict=fontdict, rotation=90)
ax.set_yticklabels([''] + predicted_sentence, fontdict=fontdict)
plt.show()
# + id="KuJ-kThog-Zs" colab_type="code" colab={}
def translate(sentence):
result, sentence, attention_plot = evaluate(sentence)
print('Input: {}'.format(sentence))
print('Predicted translation: {}'.format(result))
attention_plot = attention_plot[:len(result.split(' ')),
:len(sentence.split(' '))]
plot_attention(attention_plot, sentence.split(' '), result.split(' '))
# + id="-Vh6zUfahao7" colab_type="code" outputId="861cca39-a806-485a-ed43-32eeaffacddb" colab={"base_uri": "https://localhost:8080/", "height": 677}
translate(u'wir lachen')
# + id="I2rkAhuihfeo" colab_type="code" colab={}
# + id="R1XMyDbckTWR" colab_type="code" colab={}
# + id="hM7BXCEYkr2i" colab_type="code" colab={}
# + id="5Jf48b96kt9z" colab_type="code" colab={}
| nlp/Seq2Seq_translation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This code is used to test how does kpca perform on non-linear clustering using generated samples instead of on x-ray images
# +
from sklearn.datasets import make_circles
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
import warnings
# +
# build the kernel, total 3 functions
import numpy as np
class kernel:
def __init__(self, gamma = 1, sigma = 1, d_anova = 1, d_poly = 2, d_power = 1, alpha = 1, c = 0):
self.gamma = gamma
self.sigma = sigma
self.d_anova = d_anova
self.alpha = alpha
self.c = c
self.d_poly = d_poly
self.d_power = d_power
def rbf(self, x, y):
"""
k(x, y) = exp(- gamma * ||x-y||^2)
Hiperparametros: gamma
"""
return np.exp(- self.gamma * (np.linalg.norm(x-y)**2))
def polynomial(self, x, y):
"""
k(x, y) = (alpha * <x, y> + c)^d
Hiperparametros: alpha, c, d_poly
"""
return (self.alpha * (x.T@y) + self.c)**self.d_poly
def sigmoid(self, x, y):
"""
k(x, y) = tanh( alpha * <x, y> + c)
Hiperparametros: alpha, c
"""
return np.tanh(self.alpha * (x.T@y) + self.c)
# +
#build the kpca
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
import warnings
class KPCA:
def __init__(self, X, kernel, d):
"""
KPCA object
Parameters
----------
X: dxn matrix
kernel: kernel function from kernel class
d: number of principal components to be chosen
"""
self.X = X
self.kernel = kernel
self.d = d
def _is_pos_semidef(self, x):
return np.all(x >= 0)
def __kernel_matrix(self):
"""
Compute kernel matrix
Output:
K: nxn matrix
"""
K = []
r, c = self.X.shape
for fil in range(c):
k_aux = []
for col in range(c):
k_aux.append(self.kernel(self.X[:, fil], self.X[:, col]))
K.append(k_aux)
K = np.array(K)
# Centering K
ones = np.ones(K.shape)/c
K = K - ones@K - K@ones + ones@K@ones
return K
def __descomp(self):
"""
Decomposition of K
Output:
tuplas_eig: List of ordered tuples by singular
values; (singular_value, eigenvector)
"""
self.K = self.__kernel_matrix()
eigval, eigvec = np.linalg.eig(self.K)
if not self._is_pos_semidef(eigval):
warnings.warn("La matriz K no es semidefinida positiva")
# Normalize eigenvectors and compute singular values of K
tuplas_eig = [(np.sqrt(eigval[i]), eigvec[:,i]/np.sqrt(eigval[i]) ) for i in range(len(eigval))]
tuplas_eig.sort(key=lambda x: x[0], reverse=True)
return tuplas_eig
def project(self):
"""
Compute scores
Output:
scores: T = sigma * V_d^t
"""
self.tuplas_eig = self.__descomp()
tuplas_eig_dim = self.tuplas_eig[:self.d]
self.sigma = np.diag([i[0] for i in tuplas_eig_dim])
self.v = np.array([list(j[1]) for j in tuplas_eig_dim]).T
self.sigma = np.real_if_close(self.sigma, tol=1)
self.v = np.real_if_close(self.v, tol=1)
self.scores = self.sigma @ self.v.T
return self.scores
def plot_singular_values(self, grid = True):
eig_plot = [np.real_if_close(e, tol=1) for (e, _) in self.tuplas_eig if e > 0.01]
plt.style.use('seaborn-whitegrid')
fig = plt.figure(figsize=(15,7.5))
plt.plot(list(range(1, len(eig_plot) + 1)), eig_plot)
plt.grid(grid)
plt.title('Valores singulares de la matriz $K$ distintos de 0')
plt.ylabel('$\sigma^2$')
plt.show()
def plot_scores_2d(self, colors, grid = True, dim_1 = 1, dim_2 = 2):
if self.d < 2:
warnings.warn("No hay suficientes componentes prinicpales")
return
plt.style.use('seaborn-whitegrid')
fig = plt.figure(figsize=(15,10))
plt.axhline(c = 'black', alpha = 0.2)
plt.axvline(c = 'black', alpha = 0.2)
plt.scatter(self.scores[dim_1 - 1,:], self.scores[dim_2 - 1,:], c = colors)
plt.grid(grid)
plt.title('KPCA Space')
plt.xlabel('${}^a$ componente principal en el espacio $\phi(X)$'.format(dim_1))
plt.ylabel('${}^a$ componente principal en el espacio $\phi(X)$'.format(dim_2))
plt.show()
def plot_scores_3d(self, colors, grid = True, dim_1 = 1, dim_2 = 2, dim_3 = 3):
if self.d < 3:
warnings.warn("No hay suficientes componentes prinicpales")
return
plt.style.use('seaborn-whitegrid')
fig = plt.figure(figsize=(15,10))
ax = fig.add_subplot(111, projection="3d")
ax.scatter(self.scores[dim_1 - 1,:], self.scores[dim_2 - 1,:], self.scores[dim_3 - 1,:], c = colors)
plt.grid(grid)
ax.axis('on')
plt.title('KPCA Space')
ax.set_xlabel('${}^a$ componente principal en el espacio $\phi(X)$'.format(dim_1))
ax.set_ylabel('${}^a$ componente principal en el espacio $\phi(X)$'.format(dim_2))
ax.set_zlabel('${}^a$ componente principal en el espacio $\phi(X)$'.format(dim_3))
plt.show()
def plot_density(self, labels, dim=1, grid = False):
plt.style.use('seaborn-whitegrid')
fig = plt.figure(figsize=(15,5))
for ele in np.unique(labels):
sns.distplot(self.scores[dim - 1,:][np.where(labels == ele)], hist = False,
kde = True, kde_kws = {'linewidth': 3}, label = ele)
plt.grid(grid)
plt.legend()
plt.title('Distribuciones en la ${}^a$ componente principal'.format(dim))
plt.show()
# -
# create a sample code
X, Y = make_circles(n_samples=200, noise =0.1, factor= 0.0002)
colors = ['#FFB300' if e==1 else '#E64A19' for e in Y]
plt.style.use('seaborn-whitegrid')
fig = plt.figure(figsize=(15,10))
plt.axhline(c = 'black', alpha = 0.2)
plt.axvline(c = 'black', alpha = 0.2)
plt.scatter(X[:,0], X[:,1], c = colors)
plt.grid(False)
plt.show()
# +
# apply KPCA
X = X.T # X must be dxn
#uncomment this for gaussian kernel
#k = kernel(gamma = 25).rbf
#uncomment this for polynomial kernal
#k = kernel(alpha=4,c=1,d_poly=2).polynomial
#uncomment this for sigmoid kernel
k = kernel(alpha=0.2,c=1).sigmoid
kpca = KPCA(X, k, 3)
scores = kpca.project()
# -
T = kpca.scores #Matrix of scores
K = kpca.K #Kernel matrix
V = kpca.v #Matrix of eigenvectors
S = kpca.sigma #Diagonal matrix of (real) singular values
kpca.plot_singular_values(grid = False)
kpca.plot_density(labels = Y, dim=1, grid = False)
kpca.plot_scores_2d(colors = colors, grid = False, dim_1 = 1, dim_2 = 2)
kpca.plot_scores_3d(colors = colors, dim_1 = 1, dim_2 = 2, dim_3 = 3)
| kpca_sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Notebook for reading and cheking model TM5_AP3-INSITU
# +
import pyaerocom as pya
pya.change_verbosity('critical')
MODEL_ID = 'TM5_AP3-INSITU'
# -
reader = pya.io.ReadGridded(MODEL_ID)
print(reader)
# #### Variable 'abs550dryaer'
abs550aer = reader.read('abs550dryaer')
print(abs550aer)
# #### Variable 'ec550dryaer'
ec550dryaer = reader.read('ec550dryaer')
print(ec550dryaer)
| sorted_out/Mod_TM5_AP3-INSITU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-4.2 Intro Python
# ## Working with Files
# 4.1 File import in Jupyter Notebooks
# 4.1 File **`open()`** and **`.read()`**
# 4.2 **File Read as a list with `.readlines()`**
# 4.2 **File Closing to free resources with `.close()`**
# 4.3 Remove characters using **`.strip()`**
# 4.3 File Read a line at a time with **`.readline()`**
# 4.4 File **`.write()`** with **`.seek()`**
# 4.4 File append mode
#
# -----
#
# ><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
# 4.1 Import files in Jupyter Notebooks using the curl command
# 4.1 **`open()`** and **`.read()`** local files in memory
# 4.1 **`.read(`)** a specific number of characters
# 4.2 **Use `.readlines()` to read text from files as a list of lines**
# 4.2 **Use `.close` to free system resources**
# 4.3 Use **`.readline()`** to read data from file a line at a time
# 4.3 Use **`.strip()`** to remove new line characters
# 4.4 **`.write()`** data to a new local file
# 4.4 Use **`.seek()`** to set file read or write location
# 4.4 Use file append mode
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
# ## `.readlines()`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/0cd43d02-5eac-40b5-ba2d-97f078415ddd/Unit2_Section4.2a-Readlines-Open_Text_as_List.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/0cd43d02-5eac-40b5-ba2d-97f078415ddd/Unit2_Section4.2a-Readlines-Open_Text_as_List.vtt","srclang":"en","kind":"subtitles","label":"english"}])
#
# ### File read as a list with .readlines()
# converts the lines of a file into a **list** of strings
#
# ```python
# poem_lines = poem1.readlines()
# ```
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# [ ] Run to download file to notebook
# !curl https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/poem1.txt -o poem1.txt
# +
# [ ] review and run example
# open address to file
poem1 = open('poem1.txt', 'r')
# readlines and print as a list
poem_lines = poem1.readlines()
poem_lines
# -
# [ ] review and run example
for line in poem_lines:
print(line)
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
#
# ## `.readlines()`
# ### open the cities file as a list
# 1. **Import a list of cities using curl**
# a. git the list from https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/cities
# b. name the list cities.txt
# 2. **Open cities.txt in read mode using a variable: cities_file**
# 3. **Read cities_file as a list variable: cities_lines using `.readlines()`**
# 4. **Print each line of cities_lines by iterating the list**
# +
# [ ] import cities
# ! curl https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/cities -o cities.txt
# +
# [ ] open cities.txt as cities_file and read the file as a list: cities_lines
cities1 = open('cities.txt','r')
cities_lines = cities1.readlines()
cities_lines
# +
# [ ] use list iteration to print each city in cities_lines list
for line in cities_lines:
print(line)
# -
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
#
# ## working with lists from .readlines()
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/ed9b1523-6d69-462c-b18c-01e5423c1e52/Unit2_Section4.2b-Readlines-Remove_Characters.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/ed9b1523-6d69-462c-b18c-01e5423c1e52/Unit2_Section4.2b-Readlines-Remove_Characters.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### remove newline characters from lists created using .readlines()
# ```python
# for line in poem_lines:
# poem_lines[count] = line[:-1]
# count += 1
# ```
# **`line[:-1]`** sets the end point at the last character of the string, the result is the **`'\n'`** (newline) character is omitted
#
# | list item | list item contents |
# |-----|-----|
# | poem_lines[0] | 'Loops I repeat\n' |
# | poem_lines[1] | 'loops\n' |
# | poem_lines[2] | 'loops\n' |
# | poem_lines[3] | 'I repeat\n' |
# |... | ... |
# ###
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# This example assumes that poem1.txt has been imported in 1st example above
# [ ] review and run examples
# [ ] re-open file and read file as a list of strings
poem1 = open('poem1.txt', 'r')
poem_lines = poem1.readlines()
print(poem_lines)
# [ ] print each list item
for line in poem_lines:
print(line)
# +
# [ ] remove the last character of each list item, which is "\n"
count = 0
for line in poem_lines:
poem_lines[count] = line[:-1]
count += 1
print(poem_lines)
# -
# [ ] print each list item
for line in poem_lines:
print(line)
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
#
# ## remove newline characters from cities lists created using .readlines()
# - This task assumes that cites.txt has been imported in Task 1 above
# - In task 1, the cities were printed with a blank line between each city - this task removes the blank lines
#
# +
# [ ] re-open file and read file as a list of strings
# [ ] open cities.txt as cities_file and read the file as a list: cities_lines
cities1 = open('cities.txt','r')
cities_lines = cities1.readlines()
cities_lines
# +
# [ ] remove the last character, "\n", of each cities_lines list item
count = 0
for line in cities_lines:
cities_lines[count] = line[:-1]
count += 1
print(cities_lines)
# -
# [ ] print each list item in cities_lines
for line in cities_lines:
print(line)
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
#
# ## `.close()`
# []( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/50a925e8-25e2-4bfa-936b-e2d181af36f0/Unit2_Section4.2c-File_Close_Method.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/50a925e8-25e2-4bfa-936b-e2d181af36f0/Unit2_Section4.2c-File_Close_Method.vtt","srclang":"en","kind":"subtitles","label":"english"}])
# ### File .close() method frees resources
# flie.close() method removes the reference created from file open() function
#
# ```python
# poem1.close()
# ```
# #
# <font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
# This example assumes that poem1.txt has been imported in 1st example above
# [ ] review and run example: open and readlines of poem1.txt
poem1 = open('poem1.txt', 'r')
# +
# [ ] review and run example: readlines breaks if file is no longer open
poem_lines = poem1.readlines()
print(poem_lines)
# -
# [ ] review and run example: Close poem1
poem1.close()
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
# ## File .close()
# write each item in it's own cell
# - open cities.txt as cities_file
# - read the lines as cities_lines
# - print the cities that **start with the letter "D" or greater**
# - close cities_file
# - test that file is closed
# +
# [ ] open cities.txt as cities_file
cities_file = open('cities.txt', 'r')
# +
# [ ] read the lines as cities_lines
cities_lines = cities_file.readlines()
# +
# [ ] print the cities that start with the letter "D" or greater
for city in cities_lines:
if city.lower() >= "d":
print(city)
# +
# [ ] test that file is closed
cities_file.close()
for city in cities_lines:
if city.lower() >= "d":
print(city)
# +
# [ ] close cities_file
cities_file.close()
# -
# #
# <font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
# ## readlines() poem2
# write each item in its own cell
# - import https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/poem2.txt as poem2.txt
# - open poem2.txt as poem2_file in read mode
# - create a list of strings, called poem2_lines, from each line of poem2_text (use **.readlines()**)
# - remove the newline character for each list item in poem2_lines
# - print the poem2 lines in reverse order
# +
# [ ] import https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/poem2.txt as poem2.txt
# !curl https://raw.githubusercontent.com/MicrosoftLearning/intropython/master/poem2.txt -o poem2.txt
# +
# [ ] open poem2.txt as poem2_text in read mode
poem2 = open('poem2.txt', 'r')
# +
# [ ] create a list of strings, called poem2_lines, from each line of poem2_text
poem2_lines = poem2.readlines()
# +
# [ ] remove the newline character for each list item in poem2_lines
count = 0
for line in poem2_lines:
poem2_lines[count] = line[:-1]
count += 1
print(poem2_lines)
# +
# [ ] print the poem2 lines in reverse order
poem2_lines.reverse()
for line in poem2_lines:
print(line)
# -
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Python Fundamentals/Module_4_2_Python_Fundamentals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import pandas as pd
import numpy as np
import glob # to find all files in folder
from datetime import datetime
from datetime import date, time
from dateutil.parser import parse
import seaborn as sns
import matplotlib.pyplot as plt
import itertools
# %matplotlib inline
sns.set_context('notebook')
pd.options.mode.chained_assignment = None # default='warn'
import requests
from bs4 import BeautifulSoup
# -
# # Collect the data
# ## Get the select options
reportModel = 133685247
full_form_url = 'http://isa.epfl.ch/imoniteur_ISAP/!GEDPUBLICREPORTS.filter?ww_i_reportModel=133685247'
r = requests.get(full_form_url)
soup = BeautifulSoup(r.text, 'html.parser')
# get the name of the dropdown menus
select = soup.find_all('select')
select_name = [s.attrs['name'] for s in select]
select_name
select_fields = [soup.find('select',{'name': name}) for name in select_name]
# the html for each <select> field
# find the value for the informatique section
unite_acad_options = select_fields[0].find_all('option')
#unite_acad_options
unite_acad_informatique ={opt['value']: opt.text for opt in unite_acad_options if opt.text == 'Informatique'}
unite_acad_informatique
# periode academic
#select_fields[1].find_all('option')
period_acad = {opt['value']: opt.text for opt in select_fields[1].find_all('option') if opt['value'] != 'null' and int(opt.text.split('-')[0]) >= 2007}
period_acad
# get all the pedagogic periods
option = select_fields[2].find_all('option')
period_pedago = {opt['value']: opt.text for opt in option if opt.text != '' }
period_pedago
option = select_fields[3].find_all('option')
hiverEte = {opt['value']: opt.text for opt in option if opt['value'] != 'null'}
hiverEte
# ## Collect
# arguments are tuples (key, 'description') eg: ('2936286': "Semestre d'automne")
def collect_dataframe(t_unite_acad, t_periode_acad, t_periode_pedago, t_hiver_ete, final_headers):
print("collect_dataframe: input: "+str(t_unite_acad)+" & "+str(t_periode_acad)+" &"+str(t_periode_pedago)+" & "+str(t_hiver_ete))
#Send request
params = {
'ww_x_GPS': -1,
'ww_i_reportModel': reportModel,
'ww_i_reportModelXsl': 133685270,
'ww_x_UNITE_ACAD': t_unite_acad[0],
'ww_x_PERIODE_ACAD': t_periode_acad[0],
'ww_x_PERIODE_PEDAGO': t_periode_pedago[0],
'ww_x_HIVERETE': t_hiver_ete[0]
}
url = 'http://isa.epfl.ch/imoniteur_ISAP/!GEDPUBLICREPORTS.html'
r = requests.get(url, params=params)
soupe = BeautifulSoup(r.text, 'html.parser')
# get all the tr tags
tr_tags = soupe.find_all('tr')
#Temporary dictionary that will collect all the entry of the dataframe
data = {}
# there may be several tables.
current_table = 't1'
# for each tr tag, determine if it is a table title, a header (ignore those) or a student row
for tr in tr_tags:
th = tr.find_all('th')
if(len(th) == 1): #this is a table title
current_table = th[0].text.split('\n')[0]
data[current_table] = []
#print('current table: '+str(current_table))
elif(len(th) > 1): #this is the header row (ignore because is always the same)
#print('headers: '+str([t.text for t in th]))
pass
else:
# this is a student
td_tags = tr.find_all('td')
student = [td.text.replace('\xa0', ' ') for td in td_tags[:-1]] #drop last td because it is always empty
# add the desired columns
student.append(current_table)
student.append(t_periode_acad[1])
student.append(t_periode_pedago[1])
#print('student: '+str(student))
data[current_table].append(student)
# return all different dataframes
dframes = [pd.DataFrame(data[k], columns=final_headers) for k in list(data.keys())]
return dframes
# make one request for all permutations of (unite_acad, periode_acad, periode_pedago). We ignore hiverete because it is redundant.
perm_list = list(itertools.product(list(unite_acad_informatique.items()), list(period_acad.items()), list(period_pedago.items())))
# get the data
header = ['Civilité', 'Nom_Prénom', 'Orientation_Bachelor', 'Orientation_Master', 'Spécialisation', 'Filière_opt.', 'Mineur', 'Statut', 'Type_Echange', 'Ecole_Echange', 'No_Sciper', 'title', 'periode_acad', 'periode_pedago']
dframes = []
for (ua, pa, pp) in perm_list:
res = collect_dataframe(ua, pa, pp,('null', 'null'), header)
[dframes.append(df) for df in res]
#concatenate
all_data = pd.concat([df for df in dframes])
# +
# write to file
#all_data.to_csv('all_data.csv')
# -
len(all_data)
# Note that the Mineur semestre X are always empty. That is why they dont appear here.
all_data['periode_pedago'].unique()
| HW02-Data_from_the_Web/collect_and_persist_all_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Specifying a Custom Anchor Picker
# In this notebook we show how to write your own anchor picker class, and demonstrate the effect different anchor pickers can have on the resulting approximate nearest neighbor graph.
# +
# import some useful packages
import numpy as np
from numba import njit
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from annchor import Annchor, BruteForce,compare_neighbor_graphs
from tqdm.auto import tqdm as tq
# -
# Specify some ANNchor params that we will keep constant
n_anchors = 10
p_work = 0.05
# +
# Create and visualise a test data set
X,y,centers = make_blobs(centers=10,n_samples=1000,random_state=42,return_centers=True)
fig,ax = plt.subplots(figsize=(7,7))
ax.scatter(X[:,0],X[:,1],marker='.',c='k',alpha=0.5)
#ax.scatter(centers[:,0],centers[:,1],marker='o',c='r',s=50)
plt.show()
# +
# define our metric (just euclidean distance) and get exact k-NN graph
@njit()
def d(x,y):
return np.linalg.norm(x-y)
bruteforce = BruteForce(X,d)
bruteforce.fit()
# -
# Run Annchor with default anchor picker (maxmin) and determine its accuracy
annchor = Annchor(X,d,
n_anchors=n_anchors,
p_work=p_work)
annchor.fit()
n_errors = compare_neighbor_graphs(bruteforce.neighbor_graph,annchor.neighbor_graph,n_neighbors=15)
print('# errors (maxmin) = %d' % n_errors)
# +
# Here we create an anchor picker class which uses a set of pre-chosen points in R2.
class ExternalAnchorPicker:
def __init__(self,A):
# Initialise our anchor picker
self.A=A # Init our class with the list of pre-chosen points (A)
self.is_anchor_safe=False # If your anchors do not come from the set X
# then is_anchor_safe should be False
def get_anchors(self, ann: 'Annchor'):
# This is the main bulk of the class.
# get_anchors should find the anchor points, and work out the distances to
# each point in the data set.
# define some shorthand variables
nx = ann.nx
na = ann.n_anchors
# set random seed for reproducability
np.random.seed(ann.random_seed)
# D stores distances to anchor points
# note: at this point D is shape (n_anchors, nx),
# but we transpose this after calculations.
D = np.zeros((na, nx)) + np.infty
# This bit wraps our loop in tqdm for progress bars if we choose to be 'verbose'
if ann.verbose:
v = lambda f:tq(f)
else:
v = lambda f:f
# loop over our data set and calculate distance to anchor points
for i in v(range(na)):
D[i] = np.array([ann.f(x,self.A[i]) for x in ann.X])
# Returns 3-tuple (A,D,n_evals)
# A = array of indices of anchor points if they are in our data set, otherwise empty array
# D = array of distances to anchor points
# n_evals = number of calls to the metric
return np.array([]), D.T, na*nx
# +
# Let's pick a ring of points surrounding our data and use those as the anchors
theta = np.linspace(0,np.pi*2,11)[:-1]
ring = np.vstack([15*np.cos(theta),15*np.sin(theta)]).T
ring_anchor_picker = ExternalAnchorPicker(ring)
annchor_ring = Annchor(X,d,
n_anchors=n_anchors,
anchor_picker=ring_anchor_picker,
p_work=p_work)
annchor_ring.fit()
n_errors = compare_neighbor_graphs(bruteforce.neighbor_graph,
annchor_ring.neighbor_graph,
n_neighbors=15)
print('# errors (ring) = %d' % n_errors)
# -
# Now let's try picking the centers of our data clusters and using those as the anchors
center_anchor_picker = ExternalAnchorPicker(centers)
annchor_center = Annchor(X,d,
n_anchors=n_anchors,
anchor_picker=center_anchor_picker,
p_work=p_work)
annchor_center.fit()
n_errors = compare_neighbor_graphs(bruteforce.neighbor_graph,
annchor_center.neighbor_graph,
n_neighbors=15)
print('# errors (centers) = %d' % n_errors)
# Quickly visualise the difference in locations of our anchor points
fig,ax = plt.subplots(figsize=(7,7))
ax.scatter(X[:,0],X[:,1],marker='.',c='k',alpha=0.5,label='data')
ax.scatter(ring[:,0],ring[:,1],marker='o',s=50,label='ring')
ax.scatter(centers[:,0],centers[:,1],marker='d',s=50,label='center')
ax.scatter(X[annchor.A][:,0],X[annchor.A][:,1],marker='s',s=50,label='maxmin')
ax.legend()
plt.show()
# +
# Let's do a bunch of runs to see how everything averages out over the random
# this could take a while - go grab a cup of tea (or lower n_runs...)
n_runs = 100
def test(anchor_picker, seed,p_work=0.1):
ann = Annchor(X,d,
n_anchors=n_anchors,
anchor_picker=anchor_picker,
p_work=p_work,
random_seed=seed)
ann.fit()
return compare_neighbor_graphs(bruteforce.neighbor_graph,
ann.neighbor_graph,
n_neighbors=15)
from annchor.pickers import MaxMinAnchorPicker
cs = [test(ExternalAnchorPicker(centers),i,p_work=0.05) for i in tq(range(n_runs))]
rs = [test(ExternalAnchorPicker(ring),i,p_work=0.05) for i in tq(range(n_runs))]
ms = [test(MaxMinAnchorPicker(),i,p_work=0.05) for i in tq(range(n_runs))]
# +
# Visualise the distribution of errors for the different anchor pickers
fig,ax = plt.subplots(figsize=(15,5))
ax.bar(np.arange(49)-0.2,np.histogram(ms,bins= np.arange(50))[0],width=0.2,label = 'maxmin')
ax.bar(np.arange(49),np.histogram(cs,bins= np.arange(50))[0],width=0.2,label = 'centers')
ax.bar(np.arange(49)+0.2,np.histogram(rs,bins= np.arange(50))[0],width=0.2,label = 'ring')
ax.legend()
ax.set_xticks(np.arange(50))
ax.set_ylabel('Count')
ax.set_xlabel('Number of Errors')
plt.show()
# +
# It is worth noting that increasing p_work mitigates the choice of anchor picker
n_runs = 10
cs = [test(ExternalAnchorPicker(centers),i,p_work=0.1) for i in tq(range(n_runs))]
rs = [test(ExternalAnchorPicker(ring),i,p_work=0.1) for i in tq(range(n_runs))]
ms = [test(MaxMinAnchorPicker(),i,p_work=0.1) for i in tq(range(n_runs))]
# +
fig,ax = plt.subplots(figsize=(15,5))
ax.bar(np.arange(9)-0.2,np.histogram(ms,bins= np.arange(10))[0],width=0.2,label = 'maxmin')
ax.bar(np.arange(9),np.histogram(cs,bins= np.arange(10))[0],width=0.2,label = 'centers')
ax.bar(np.arange(9)+0.2,np.histogram(rs,bins= np.arange(10))[0],width=0.2,label = 'ring')
ax.legend()
ax.set_xticks(np.arange(10))
ax.set_ylabel('Count')
ax.set_xlabel('Number of Errors')
plt.show()
# -
| Examples/CustomAnchorPicker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mrp
# language: python
# name: mrp
# ---
try:
__IPYTHON__
USING_IPYTHON = True
except NameError:
USING_IPYTHON = False
# #### Argparse
import argparse
ap = argparse.ArgumentParser()
ap.add_argument('mrp_data_dir', help='')
ap.add_argument('--train-sub-dir', default='training', help='')
ap.add_argument('--companion-sub-dir', default='./mrp-companion/2019/companion')
ap.add_argument('--mrp-file-extension', default='.mrp')
ap.add_argument('--companion-file-extension', default='.conllu')
ap.add_argument('--graphviz-file-template', default='http://localhost:8000/files/proj29_ds1/home/slai/mrp/graphviz/{}/{}.mrp/{}.png')
arg_string = """
./data/
"""
arguments = [arg for arg_line in arg_string.split(r'\\n') for arg in arg_line.split()]
if USING_IPYTHON:
args = ap.parse_args(arguments)
else:
args = ap.parse_args()
args
# #### Library imports
# +
import json
import logging
import os
import pprint
import string
from collections import Counter
from collections import defaultdict
from tqdm import tqdm
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
# -
# #### ipython notebook specific imports
if USING_IPYTHON:
# matplotlib config
# %matplotlib inline
logging.basicConfig(level=logging.INFO, handlers=[logging.StreamHandler()])
logger = logging.getLogger(__name__) # pylint: disable=invalid-name
logger.setLevel(logging.INFO)
# ### Constants
UNKWOWN = 'UNKWOWN'
# ### Load data
train_dir = os.path.join(args.mrp_data_dir, args.train_sub_dir)
frameworks = [sub_dir for sub_dir in os.listdir(train_dir) if os.path.isdir(os.path.join(train_dir, sub_dir))]
frameworks
framework2dataset2mrp_jsons = {}
for framework in tqdm(frameworks, desc='frameworks'):
dataset2mrp_jsons = {}
framework_dir = os.path.join(train_dir, framework)
dataset_names = os.listdir(framework_dir)
for dataset_name in tqdm(dataset_names, desc='dataset_name'):
mrp_jsons = []
if not dataset_name.endswith(args.mrp_file_extension):
continue
with open(os.path.join(framework_dir, dataset_name)) as rf:
for line in rf:
mrp_json = json.loads(line.strip())
if framework == 'ucca' and 'nodes' in mrp_json and 'input' in mrp_json:
input_text = mrp_json['input']
nodes = mrp_json['nodes']
for i, node in enumerate(nodes):
if 'anchors' not in node:
continue
text_segments = []
for anchor in node['anchors']:
text_segments.append(input_text[anchor.get('from', -1): anchor.get('to', -1)])
mrp_json['nodes'][i]['label'] = ''.join(text_segments)
mrp_jsons.append(mrp_json)
dataset_name = dataset_name.split('.')[0]
dataset2mrp_jsons[dataset_name] = mrp_jsons
framework2dataset2mrp_jsons[framework] = dataset2mrp_jsons
for framework in framework2dataset2mrp_jsons:
logger.info(framework)
logger.info(list(framework2dataset2mrp_jsons[framework].keys()))
# ### Data Preprocessing companion
dataset2cid2parse = {}
for framework in os.listdir(args.companion_sub_dir):
framework_dir = os.path.join(args.companion_sub_dir, framework)
if not os.path.isdir(framework_dir):
continue
for dataset in tqdm(os.listdir(framework_dir), desc='dataset'):
if not dataset.endswith(args.companion_file_extension):
continue
dataset_name = dataset.split('.')[0].rstrip(string.digits)
cid2parse = {}
with open(os.path.join(framework_dir, dataset)) as rf:
parse = []
for line in rf:
line = line.strip()
if not line:
cid2parse[cid] = parse
parse = []
cid = ''
elif line.startswith('#'):
cid = line[1:]
else:
parse.append(line.split('\t'))
dataset2cid2parse[dataset_name] = cid2parse
dataset2cid2parse.keys()
dataset = 'xinhua'
framework = 'amr'
dataset2cid2parse[dataset][framework2dataset2mrp_jsons[framework][dataset][1]['id']]
framework2dataset2mrp_jsons[framework][dataset][1]
# ### Companion statistics
list(dataset2cid2parse['wsj'].keys())[:10]
verb_lemma2org2dataset_cids = defaultdict(lambda: defaultdict(list))
for dataset, cid2parse in dataset2cid2parse.items():
for cid, parse in cid2parse.items():
for word_record in parse:
word_index, org, lemma, pos, *_ = word_record
if pos == 'VERB':
verb_lemma2org2dataset_cids[lemma][org].append((dataset, cid))
logger.info('No. of unique verb lemma: {}'.format(len(verb_lemma2org2dataset_cids)))
for verb_lemma, org2dataset_cids in verb_lemma2org2dataset_cids.items():
sorted_org_count = sorted([
(org, len(dataset_cids))
for org, dataset_cids in org2dataset_cids.items()
], key=lambda x: x[1], reverse=True)
print(verb_lemma, sorted_org_count[:10])
verb_lemma = 'be'
org = 'is'
cid_index = 1
def view_parse(verb_lemma, org, cid_index):
dataset, cid = verb_lemma2org2dataset_cids[verb_lemma][org][cid_index]
for framework, dataset2mrp_jsons in framework2dataset2mrp_jsons.items():
if dataset in dataset2mrp_jsons:
logger.info(pprint.pformat(dataset2mrp_jsons[dataset][cid_index]))
graphviz_file_name = dataset2mrp_jsons[dataset][cid_index].get('id')
if graphviz_file_name:
logger.info((framework, dataset, graphviz_file_name))
logger.info(args.graphviz_file_template.format(framework, dataset, graphviz_file_name))
return dataset2cid2parse[dataset][cid]
dataset
dataset
view_parse(verb_lemma, org, cid_index)
| src/2019-06-14-data-preprocessing-and-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('gapminder.csv')
df.head()
X= df.drop(['Region'],1)
y = df['Region']
from sklearn.metrics import accuracy_score,classification_report,confusion_matrix
from sklearn.model_selection import train_test_split as tts,GridSearchCV, cross_val_score
from sklearn.ensemble import RandomForestClassifier, BaggingClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
X_train,X_test,y_train,y_test = tts(X,y,random_state=42, test_size=0.25)
knn = KNeighborsClassifier()
dte = DecisionTreeClassifier()
knn
br = BaggingClassifier(base_estimator=dte,
n_estimators=100,
bootstrap=True,
oob_score=True)
br.fit(X_train,y_train)
y_pred = br.predict(X_test)
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
rfe = RandomForestClassifier()
rfe
rfe.fit(X_train,y_train)
y_pred1 = rfe.predict(X_test)
print(classification_report(y_test,y_pred1))
print(accuracy_score(y_test,y_pred1))
import numpy as np
param = {
'n_estimators' : np.arange(100,300,50),
'max_depth' : np.arange(4,12,1),
'max_features' : np.arange(4,8,1),
# 'criterion': ['gini','entropy'],
# 'min_samples_split': np.arange(0.01,0.05,0.01)
}
rfe_cv = GridSearchCV(estimator=rfe,param_grid=param,cv=3)
rfe_cv.fit(X,y)
rfe_cv.best_estimator_
rfe_cv.best_score_
rfe_cv.best_estimator_.feature_importances_
from sklearn.ensemble import VotingClassifier
knn = KNeighborsClassifier()
dte =DecisionTreeClassifier()
rfe= RandomForestClassifier()
vc = VotingClassifier(estimators= [('knn',knn),('dt',dte),('rf',rfe)],
voting='soft',
weights=None,
# n_jobs=None,
# flatten_transform=True,
)
vc.fit(X_train,y_train)
y_pred2 = vc.predict(X_test)
print(classification_report(y_test,y_pred2))
print(accuracy_score(y_test,y_pred2))
| C17_Ensemble Learning/C17_Ensemble Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="td1va_ndHUED"
# ###Imports
# + id="aBYix7CoHQhc" executionInfo={"status": "ok", "timestamp": 1618403611730, "user_tz": -120, "elapsed": 2313, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
import numpy as np
import pandas as pd
from keras.utils import np_utils
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Dense
# + [markdown] id="seDXVMv7HdIR"
# ###Load csv
# + id="Bt5n0kYFObOr" executionInfo={"status": "ok", "timestamp": 1618403611733, "user_tz": -120, "elapsed": 2304, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
COLUMN_NAMES = ['SepalLength', 'SepalWidth', 'PetalLength', 'PetalWidth', 'Species']
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="Zmhd0n9WHYiP" executionInfo={"status": "ok", "timestamp": 1618403611737, "user_tz": -120, "elapsed": 2297, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}} outputId="4a77c588-c569-4493-99c9-3b372f9ddf14"
dataset = pd.read_csv("/content/drive/MyDrive/Škola/NS/cvicenie_8/iris_training.csv", names=COLUMN_NAMES, header=0)
dataset.head()
# + [markdown] id="Bx1OOmSxH-Ch"
# ###Create train data
# + id="Njb4hGkuHk4S" executionInfo={"status": "ok", "timestamp": 1618403611739, "user_tz": -120, "elapsed": 2283, "user": {"displayName": "\u013dudov\u0<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
trainX = dataset.iloc[:,0:4].values
# + id="S6ELuYTzHrqg" executionInfo={"status": "ok", "timestamp": 1618403611742, "user_tz": -120, "elapsed": 2282, "user": {"displayName": "\u013dudov\u0<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
trainY = dataset.iloc[:,4].values
# + colab={"base_uri": "https://localhost:8080/"} id="ygUv-EsBH689" executionInfo={"status": "ok", "timestamp": 1618403612367, "user_tz": -120, "elapsed": 2901, "user": {"displayName": "\u013dudov\u<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}} outputId="b7c18429-625c-47dc-ff08-218f5c35514e"
kategoricka_premenna = np_utils.to_categorical(trainY)
kategoricka_premenna
# + [markdown] id="cOzWf609SVHI"
# ### Sekvenčný model
# + id="kXAAnpexNx2M" executionInfo={"status": "ok", "timestamp": 1618403612369, "user_tz": -120, "elapsed": 2897, "user": {"displayName": "\u013dudov\u<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
moj_keras_model = Sequential()
# + id="itdg5-eZSTkC" executionInfo={"status": "ok", "timestamp": 1618403612372, "user_tz": -120, "elapsed": 2892, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
# first layer
moj_keras_model.add(Dense(10, input_dim=4, activation='sigmoid'))
# + id="k1kPpvYdS1dG" executionInfo={"status": "ok", "timestamp": 1618403612373, "user_tz": -120, "elapsed": 2888, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
# second layer
moj_keras_model.add(Dense(10, activation='sigmoid'))
# + id="CMpd0iBHTJQ-" executionInfo={"status": "ok", "timestamp": 1618403612374, "user_tz": -120, "elapsed": 2882, "user": {"displayName": "\u013dudov\u00edt Laca", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
moj_keras_model.add(Dense(3))
# + id="CJXPaO29T-zx" executionInfo={"status": "ok", "timestamp": 1618403612376, "user_tz": -120, "elapsed": 2881, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
# compile model
# optimizer - (best adam) another = sgd
moj_keras_model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
# + [markdown] id="4ETtO8tBTdlT"
# ### Train model
# + colab={"base_uri": "https://localhost:8080/"} id="nO45ZriaTXJz" executionInfo={"status": "ok", "timestamp": 1618403612736, "user_tz": -120, "elapsed": 3237, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}} outputId="3ae85a52-ef59-4453-c567-e89d4474ea21"
moj_keras_model.fit(trainX, kategoricka_premenna)
# + colab={"base_uri": "https://localhost:8080/"} id="YE7fwfcZT67R" executionInfo={"status": "ok", "timestamp": 1618403612742, "user_tz": -120, "elapsed": 3239, "user": {"displayName": "\u013dudov\u00<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}} outputId="5084bf5b-67f8-49a4-b536-5adbaf7e9241"
moj_keras_model
# + id="dg9FiaRAUw5Y" executionInfo={"status": "ok", "timestamp": 1618403612744, "user_tz": -120, "elapsed": 3230, "user": {"displayName": "\u013dudov\u0<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg_8RtneW7PgTTuiCnx0VUNQNyApN5rCT7UJZW89w=s64", "userId": "17075019361640864910"}}
# https://keras.io/api/layers/activations/
| cvicenie_8/Keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import shutil
import urllib3
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import datetime
from scipy import stats
pd.set_option('display.max_columns', 500)
# -
# Based on: machine-learning/examples/bbengfort/traffic/Traffic.ipynb
#
# Pulled data for one building and modeled electricity usage
# #DATA LOADING
data = pd.read_csv("0008_0806.csv",
index_col="DATE_TIME",
parse_dates=True
)
data.head(24)
# +
def plot_numeric_features(df=data, cols=["NKBTUPERSQFT", "AWND", "CLDD", "HTDD", "SNOW", "TAVG", "TMAX", "TMIN", "WDF2", "WDF5", "WSF2", "WSF5"]):
#"YEARBUILT", "AWND", "CLDD", "HTDD", "SNOW", "TAVG", "TMAX", "TMIN", "WDF2", "WDF5", "WSF2", "WSF5", "KBTUPERSQFT"
fig, axes = plt.subplots(nrows=len(cols), sharex=True, figsize=(20,20))
for col, ax in zip(cols, axes) :
df[col].plot(ax=ax)
ax.set_title(col)
if ax == axes[-1]:
ax.set_xlabel("date")
# fig.tight_layout()
return ax
# Plot features in date range
_ = plot_numeric_features(data.loc["2018-01-01":"2019-12-01"])
# +
#def plot_categorical_features(df=data, cols=['WARD2','RES']):
# fig, axes = plt.subplots(nrows=len(cols), sharex=True, figsize=(9,18))
# for col, ax in zip(cols, axes):
# sns.barplot(x='KBTUPERSQFT', y=col, data=df, ax=ax)
# ax.set_title(col)
# if ax == axes[-1]:
# ax.set_xlabel('KBTUPERSQFT')
# fig.tight_layout()
# return ax
#_ = plot_categorical_features(df)
# +
from sklearn.model_selection import train_test_split as tts
TARGET = "EKBTUPERSQFT"
COLS = ["NKBTUPERSQFT","DATE_TIME","YEARBUILT", "CLDD", "HTDD", "SNOW","WDF2", "WSF2"]
def make_sklearn_data(df=data, splits=True, train_size=0.8, target=TARGET, cols=COLS):
# Add the datetime column back to the main dataframe
df = df.reset_index()
X, y = df[cols], df[target]
if splits:
return tts(X, y, train_size=train_size)
return X, y
# +
from sklearn.base import BaseEstimator, TransformerMixin
class CyclicEncoder(BaseEstimator, TransformerMixin):
def __init__(self, date_extract="month"):
if date_extract not in {"minute", "hour", "week", "month", "year"}:
raise ValueError(f"specify correct date component to extract, not {date_extract}")
self.date_extract = date_extract
def get_date_component(self, x):
if self.date_extract == "month":
return x.dt.month
elif self.date_extract == "year":
return x.dt.year
else:
raise NotImplementedError(f"{self.date_extract} date component not implemented yet")
def fit(self, X, y=None):
self.cycle_max_ = self.get_date_component(X).max()
return self
def transform(self, X, y=None):
cols = []
names = []
x = self.get_date_component(X)
xn = 2 * np.pi * x / self.cycle_max_
cols.append(np.cos(xn))
names.append(f"{X.name}_cos")
cols.append(np.sin(xn))
names.append(f"{X.name}_sin")
return pd.DataFrame(np.asarray(cols).T, columns=names)
ce = CyclicEncoder().fit_transform(data.reset_index()["DATE_TIME"])
ce.plot(x="DATE_TIME_cos", y="DATE_TIME_sin", kind="scatter")
# +
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import FeatureUnion, Pipeline
from sklearn.preprocessing import OneHotEncoder
extraction = [('column_selection', ColumnTransformer([('time_components', FeatureUnion([('month', CyclicEncoder(date_extract="month")),('year', CyclicEncoder(date_extract="year")),]), 'DATE_TIME'),], remainder="passthrough")),]
X, y = make_sklearn_data(splits=False)
Pipeline(extraction).fit_transform(X).shape
# +
from sklearn.ensemble import RandomForestRegressor
X_train, X_test, y_train, y_test = make_sklearn_data()
extraction.append(("clf", RandomForestRegressor()))
model = Pipeline(extraction)
model.fit(X_train, y_train)
model.score(X_test, y_test)
# -
# Based on: demos/20201031.ipynb
# Regression on KBTU for the numeric columns
data.shape
# +
cols=["NKBTUPERSQFT", "AWND", "CLDD", "HTDD", "SNOW", "TAVG", "WDF2", "WDF5", "WSF2", "WSF5"]
X = data[cols]
y = data["EKBTUPERSQFT"]
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True)
model.fit(X, y)
# +
def describe_model(model):
print(model.get_params())
print()
for attr in dir(model):
if attr.endswith("_") and not attr.startswith("_"):
print(f"{attr} {getattr(model, attr)}")
describe_model(model)
# +
from sklearn.model_selection import train_test_split as tts
X_train, X_test, y_train, y_test = tts(X, y, train_size=0.8)
# -
lrm = LinearRegression(normalize=True).fit(X_train, y_train)
lrm.score(X_test, y_test)
# +
from sklearn.ensemble import RandomForestRegressor
rfm = RandomForestRegressor(n_estimators=10, max_depth=3).fit(X_train, y_train)
describe_model(rfm)
rfm.score(X_test, y_test)
# -
| machine_learning/.ipynb_checkpoints/Single_building_model_test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # English Wikipedia page views, 2008 - 2017
#
# ### <NAME>
#
# The goal of this project is to construct, analyze, and publish a dataset of monthly traffic on English Wikipedia from July 1 2008 through September 30 2017 and create a visulization of the dataset. Following are the steps for data acquisition, data processing and data analysis.
# The module will then output:
# 1) 5 source data files in JSON format.
# 2) 1 final data file in CSV format.
# 3) 1 .png image of the visualization.
# ### Import Necessary Libraries
#import libraries
import requests
import json
from datetime import datetime
from matplotlib.dates import DateFormatter
import pandas as pd
import numpy as np
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Step 1 Data acquisition
# Collect data from two different API endpoints, the Pageviews API (mobile-web, mobile-app, desktop) and the Pagecounts API (mobile-site, desktop).
# And then save the raw results into 5 separate JSON source data files (one file per API query) before continuing to step 2.
# +
# Collect data from Pageviews API for mobile-web traffic data from Jul 2015 to Oct 2017.
endpoint = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{access}/{agent}/{granularity}/{start}/{end}'
headers={'User-Agent' : 'https://github.com/anqiwang0827', 'From' : '<EMAIL>'}
params = {'project' : 'en.wikipedia.org',
'access' : 'mobile-web',
'agent' : 'user',
'granularity' : 'monthly',
'start' : '2015070100',
'end' : '2017100100'
}
pageview_mobileweb_call = requests.get(endpoint.format(**params))
pageview_mobileweb = pageview_mobileweb_call.json()
pageviews_mobileweb_201507_201709 = json.dumps(pageview_mobileweb)
# Save raw data results as Json file pageviews_mobile-web_201507-201709.json
with open('pageviews_mobile-web_201507-201709.json', 'w') as file:
json.dump(pageviews_mobileweb_201507_201709, file)
# +
# Collect data from Pageviews API for mobile-app traffic data from Jul 2015 to Oct 2017.
endpoint = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{access}/{agent}/{granularity}/{start}/{end}'
headers={'User-Agent' : 'https://github.com/anqiwang0827', 'From' : '<EMAIL>'}
params = {'project' : 'en.wikipedia.org',
'access' : 'mobile-app',
'agent' : 'user',
'granularity' : 'monthly',
'start' : '2015070100',
'end' : '2017100100'
}
pageview_mobileapp_call = requests.get(endpoint.format(**params))
pageview_mobileapp = pageview_mobileapp_call.json()
pageviews_mobileapp_201507_201709 = json.dumps(pageview_mobileapp)
# Save raw data results as Json file pageviews_mobile-app_201507-201709.json
with open('pageviews_mobile-app_201507-201709.json', 'w') as file2:
json.dump(pageviews_mobileapp_201507_201709, file2)
# +
# Collect data from Pageviews API for desktop traffic data from Jul 2015 to Oct 2017.
endpoint = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{access}/{agent}/{granularity}/{start}/{end}'
headers={'User-Agent' : 'https://github.com/anqiwang0827', 'From' : '<EMAIL>'}
params = {'project' : 'en.wikipedia.org',
'access' : 'desktop',
'agent' : 'user',
'granularity' : 'monthly',
'start' : '2015070100',
'end' : '2017100100'
}
pageview_desktop_call = requests.get(endpoint.format(**params))
pageview_desktop = pageview_desktop_call.json()
pageviews_desktop_201507_201709 = json.dumps(pageview_desktop)
# Save raw data results as Json file pageviews_desktop_201507-201709.json
with open('pageviews_desktop_201507-201709.json', 'w') as file3:
json.dump(pageviews_desktop_201507_201709, file3)
# +
# Collect data from Pagecounts API for mobile site traffic data from Oct 2014 to Jul 2017.
endpoint = 'https://wikimedia.org/api/rest_v1/metrics/legacy/pagecounts/aggregate/{project}/{access-site}/{granularity}/{start}/{end}'
headers={'User-Agent' : 'https://github.com/anqiwang0827', 'From' : '<EMAIL>'}
params = {'project' : 'en.wikipedia.org',
'access-site' : 'mobile-site',
'granularity' : 'monthly',
'start' : '2014100100',
'end' : '2016080100'
}
pagecounts_mobile_site_call = requests.get(endpoint.format(**params))
pagecounts_mobile_site = pagecounts_mobile_site_call.json()
pagecounts_mobile_site_201410_201607 = json.dumps(pagecounts_mobile_site)
# Save raw data results as Json file pagecounts_mobile-site_201410-201607.json
with open('pagecounts_mobile-site_201410-201607.json', 'w') as file4:
json.dump(pagecounts_mobile_site_201410_201607, file4)
# +
# Collect data from Pagecounts API for desktop site traffic data from jan 2008 to Jul 2017.
endpoint = 'https://wikimedia.org/api/rest_v1/metrics/legacy/pagecounts/aggregate/{project}/{access-site}/{granularity}/{start}/{end}'
headers={'User-Agent' : 'https://github.com/anqiwang0827', 'From' : '<EMAIL>'}
params = {'project' : 'en.wikipedia.org',
'access-site' : 'desktop-site',
'granularity' : 'monthly',
'start' : '2008010100',
'end' : '2016080100'
}
pagecounts_desktop_site_call = requests.get(endpoint.format(**params))
pagecounts_desktop_site = pagecounts_desktop_site_call.json()
pagecounts_desktop_site_200801_201607 = json.dumps(pagecounts_desktop_site)
# Save raw data results as Json file pagecounts_desktop-site_200801-201607.json
with open('pagecounts_desktop-site_200801-201607.json', 'w') as file5:
json.dump(pagecounts_desktop_site_200801_201607, file5)
# -
# ## Step 2 Data processing
# The Step 2 includes series of processing steps on these data files needed in order to prepare them for analysis. At the end of this step, a single CSV-formatted data file will be saved as en_wikipedia_traffic_200801_201709.csv.
#
# The CSV file combined all the data from previous step with the following headers:
#
# Column (Value)
# year (YYYY)
# month (MM)
# pagecount_all_views (num_views)
# pagecount_desktop_views (num_views)
# pagecount_mobile_views (num_views)
# pageview_all_views (num_views)
# pageview_desktop_views (num_views)
# pageview_mobile_views (num_views)
#
# get items in api raw data
df_pv_mw = pd.DataFrame(pageview_mobileweb)
df_pv_ma = pd.DataFrame(pageview_mobileapp)
df_pv_dk = pd.DataFrame(pageview_desktop)
df_pc_ms = pd.DataFrame(pagecounts_mobile_site)
df_pc_ds = pd.DataFrame(pagecounts_desktop_site)
df_pv_mw_items = df_pv_mw['items']
df_pv_ma_items = df_pv_ma['items']
df_pv_dk_items = df_pv_dk['items']
df_pc_ms_items = df_pc_ms['items']
df_pc_ds_items = df_pc_ds['items']
# +
# get timestamp and views from pageview API
Temp1_pv_mw = []
Temp2_pv_mw = []
Temp3_pv_mw = []
for x in range(0,len(df_pv_mw_items)):
Temp1_pv_mw.append(df_pv_mw['items'][x])
Temp2_pv_mw.append(Temp1_pv_mw[x]['views'])
Temp3_pv_mw.append(Temp1_pv_mw[x]['timestamp'])
Temp1_pv_ma = []
Temp2_pv_ma = []
Temp3_pv_ma = []
for x in range(0,len(df_pv_ma_items)):
Temp1_pv_ma.append(df_pv_ma['items'][x])
Temp2_pv_ma.append(Temp1_pv_ma[x]['views'])
Temp3_pv_ma.append(Temp1_pv_ma[x]['timestamp'])
Temp1_pv_dk = []
Temp2_pv_dk = []
Temp3_pv_dk = []
for x in range(0,len(df_pv_dk_items)):
Temp1_pv_dk.append(df_pv_dk['items'][x])
Temp2_pv_dk.append(Temp1_pv_dk[x]['views'])
Temp3_pv_dk.append(Temp1_pv_dk[x]['timestamp'])
# -
# create pageview dataframe, add mobile traffic together
pageview_list = pd.DataFrame(
{'Date':Temp3_pv_mw,
'pageview_all_views': [Temp2_pv_mw[i] + Temp2_pv_ma[i] + Temp2_pv_dk[i] for i in range(len(Temp2_pv_ma))],
'pageview_desktop_views': Temp2_pv_dk,
'pageview_mobile_views': [Temp2_pv_mw[i] + Temp2_pv_ma[i] for i in range(len(Temp2_pv_ma))]
})
# +
# get timestamp and counts from pagecount API
Temp1_pc_ms = []
Temp2_pc_ms = []
Temp3_pc_ms = []
for x in range(0,len(df_pc_ms_items)):
Temp1_pc_ms.append(df_pc_ms['items'][x])
Temp2_pc_ms.append(Temp1_pc_ms[x]['count'])
Temp3_pc_ms.append(Temp1_pc_ms[x]['timestamp'])
Temp1_pc_ds = []
Temp2_pc_ds = []
Temp3_pc_ds = []
for x in range(0,len(df_pc_ds_items)):
Temp1_pc_ds.append(df_pc_ds['items'][x])
Temp2_pc_ds.append(Temp1_pc_ds[x]['count'])
Temp3_pc_ds.append(Temp1_pc_ds[x]['timestamp'])
# -
# create pagecount dataframe
pagecount_list_ms = pd.DataFrame(
{'Date':Temp3_pc_ms,
'pagecount_mobile_views': Temp2_pc_ms
})
pagecount_list_ds = pd.DataFrame(
{'Date':Temp3_pc_ds,
'pagecount_desktop_views': Temp2_pc_ds
})
pagecount_list = pagecount_list_ms.merge(pagecount_list_ds, left_on='Date', right_on='Date', how='outer')
pagecount_list['pagecount_all_views'] = pagecount_list['pagecount_mobile_views']+pagecount_list['pagecount_desktop_views']
# combine pagecount and pageview into one dataframe
en_wikipedia_traffic = pagecount_list.merge(pageview_list, left_on='Date', right_on='Date', how='outer')
en_wikipedia_traffic = en_wikipedia_traffic.sort(['Date'])
en_wikipedia_traffic['DateTime']= en_wikipedia_traffic['Date'].apply(lambda x: pd.to_datetime(str(x), format='%Y%m%d%H'))
# split Date to Year and Month
en_wikipedia_traffic_200801_201709 = en_wikipedia_traffic
en_wikipedia_traffic_200801_201709['year']=en_wikipedia_traffic_200801_201709['Date'].str[:4]
en_wikipedia_traffic_200801_201709['month']=en_wikipedia_traffic_200801_201709['Date'].str[4:6]
en_wikipedia_traffic_200801_201709 = en_wikipedia_traffic_200801_201709[['year', 'month', 'pagecount_all_views', 'pagecount_desktop_views',
'pagecount_mobile_views','pageview_all_views','pageview_desktop_views','pageview_mobile_views']]
en_wikipedia_traffic_200801_201709 = en_wikipedia_traffic_200801_201709.fillna(0)
#write to csv
en_wikipedia_traffic_200801_201709.to_csv('en_wikipedia_traffic_200801_201709.csv', sep=',', index = False)
# ## Step 3 Data Processing
# After creating one final dataset in Step 2. Step 3 will generate a visualization graph. The visualization will plot three traffic metrics: mobile traffic, desktop traffic, and all traffic (mobile + desktop) from both Pageviews API and Pagecounts API from Jan 2008 to Sep 2017.
# Create the plot
fig, ax = plt.subplots(figsize=(15,4))
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pagecount_all_views'], 'k--', color='black')
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pageview_all_views'], 'k', color='black')
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pagecount_mobile_views'], 'k--', color='blue')
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pagecount_desktop_views'], 'k--',color='green')
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pageview_mobile_views'], 'k', color='blue')
ax.plot_date(en_wikipedia_traffic['DateTime'], en_wikipedia_traffic['pageview_desktop_views'], 'k', color='green')
plt.suptitle('Page View on Wikipidia *e10', fontsize=20)
plt.xlabel('Year', fontsize=18)
plt.ylabel('Views', fontsize=16)
plt.grid(True)
black_patch = mpatches.Patch(color='black', label='total')
green_patch = mpatches.Patch(color='green', label='main site')
blue_patch = mpatches.Patch(color='blue', label='mobiel site')
plt.legend(handles=[green_patch, blue_patch,black_patch], loc='upper left')
ax.xaxis.set_major_formatter(DateFormatter('%Y'))
# Save the plot
fig.savefig('Pave View on Wikipidia.png')
| ProgrammingStep/hcds-a1-data-curation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Function-cos" data-toc-modified-id="Function-cos-1"><span class="toc-item-num">1 </span>Function cos</a></div><div class="lev2 toc-item"><a href="#Synopse" data-toc-modified-id="Synopse-11"><span class="toc-item-num">1.1 </span>Synopse</a></div><div class="lev2 toc-item"><a href="#Description" data-toc-modified-id="Description-12"><span class="toc-item-num">1.2 </span>Description</a></div><div class="lev2 toc-item"><a href="#Examples" data-toc-modified-id="Examples-13"><span class="toc-item-num">1.3 </span>Examples</a></div><div class="lev3 toc-item"><a href="#Example-1" data-toc-modified-id="Example-1-131"><span class="toc-item-num">1.3.1 </span>Example 1</a></div><div class="lev2 toc-item"><a href="#Equation" data-toc-modified-id="Equation-14"><span class="toc-item-num">1.4 </span>Equation</a></div><div class="lev2 toc-item"><a href="#See-Also" data-toc-modified-id="See-Also-15"><span class="toc-item-num">1.5 </span>See Also</a></div>
# + [markdown] deletable=true editable=true
# # Function cos
#
# ## Synopse
#
# Create a cosine wave image.
#
# - **f = iacos(s, t, theta, phi)**
#
# - **f**: Image.
#
#
# - **s**: Image. size: [rows cols].
# - **t**: Image. Period: in pixels.
# - **theta**: Double. spatial direction of the wave, in radians. 0 is a wave on the horizontal direction.
# - **phi**: Double. Phase
# + deletable=true editable=true
import numpy as np
def cos(s, t, theta, phi):
r, c = np.indices(s)
tc = t / np.cos(theta)
tr = t / np.sin(theta)
f = np.cos(2*np.pi*(r/tr + c/tc) + phi)
return f
# -
# ## Description
#
# Generate a cosine wave image of size s with amplitude 1, period T, phase phi and wave direction of theta. The output image is a double array.
# ## Examples
testing = (__name__ == "__main__")
if testing:
# ! jupyter nbconvert --to python cos.ipynb
import numpy as np
import sys,os
ia898path = os.path.abspath('../../')
if ia898path not in sys.path:
sys.path.append(ia898path)
import ia898.src as ia
# + [markdown] deletable=true editable=true
# ### Example 1
# + deletable=true editable=true
if testing:
f = ia.cos([128,256], 100, np.pi/3, 0)
ia.adshow(ia.normalize(f, [0,255]))
# + [markdown] deletable=true editable=true
# ## Equation
#
# $$ \begin{matrix}
# f(r,c) & = & cos( 2\pi (\frac{1}{T_r}r + \frac{1}{T_c}c) + \phi) \\
# f(r,c) & = & cos( 2\pi (\frac{v}{H}r + \frac{u}{W}c) + \phi) \\
# T_r & = & \frac{T}{sin(\theta)} \\
# T_c & = & \frac{T}{cos(\theta)} \\
# u & = & \frac{W}{T_c} \\
# v & = & \frac{H}{T_r}
# \end{matrix} $$
#
# - $\theta$ is the direction of the cosine wave.
# - $T$ is the wave period, in number of pixels.
# - $T_r$ and $T_c$ are the period or wave length in the vertical and horizontal directions, respectively, in number of pixels.
# - $H$ and $W$ are the number of image rows and columns, respectively.
# - $v$ and $u$ are the normalized frequency in the horizontal and vertical directions, respectively, in cycles per image dimension.
# + [markdown] deletable=true editable=true
# ## See Also
#
# - `iacosdemo iacosdemo` -- Illustrate discrete cosine wave and its DFT showing its periodic nature.
# + deletable=true editable=true
if testing:
print('testing cos')
print(repr(np.floor(0.5 + 127*(ia.cos([7,10], 3, np.pi/4, 0)+1))) == repr(np.array(
[[ 254., 138., 2., 93., 246., 182., 18., 52., 223., 219.],
[ 138., 2., 93., 246., 182., 18., 52., 223., 219., 48.],
[ 2., 93., 246., 182., 18., 52., 223., 219., 48., 21.],
[ 93., 246., 182., 18., 52., 223., 219., 48., 21., 187.],
[ 246., 182., 18., 52., 223., 219., 48., 21., 187., 244.],
[ 182., 18., 52., 223., 219., 48., 21., 187., 244., 88.],
[ 18., 52., 223., 219., 48., 21., 187., 244., 88., 3.]])))
# -
| src/cos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
Jan_file = 'JC-202001-citibike-tripdata.csv'
Jan_df = pd.read_csv(Jan_file)
#Jan_df.head()
Jan_df.columns
# check missing data in the dataset
Jan_df.info()
Feb_file = 'JC-202002-citibike-tripdata.csv'
Feb_df = pd.read_csv(Feb_file)
#Feb_df.head()
March_file = 'JC-202003-citibike-tripdata.csv'
March_df = pd.read_csv(March_file)
#March_df.head()
April_file = 'JC-202004-citibike-tripdata.csv'
April_df = pd.read_csv(April_file)
#April_df.head()
May_file = 'JC-202005-citibike-tripdata.csv'
May_df = pd.read_csv(May_file)
#May_df.head()
June_file = 'JC-202006-citibike-tripdata.csv'
June_df = pd.read_csv(June_file)
#June_df.head()
July_file = 'JC-202007-citibike-tripdata.csv'
July_df = pd.read_csv(July_file)
#July_df.head()
Aug_file = 'JC-202008-citibike-tripdata.csv'
Aug_df = pd.read_csv(Aug_file)
#Aug_df.head()
Sep_file = 'JC-202009-citibike-tripdata.csv'
Sep_df = pd.read_csv(Sep_file)
#Sep_df.head()
Oct_file = 'JC-202010-citibike-tripdata.csv'
Oct_df = pd.read_csv(Oct_file)
#Oct_df.head()
Nov_file = 'JC-202011-citibike-tripdata.csv'
Nov_df = pd.read_csv(Nov_file)
#Nov_df.head()
Dec_file = 'JC-202012-citibike-tripdata.csv'
Dec_df = pd.read_csv(Dec_file)
#Dec_df.head()
frames = [Jan_df,Feb_df,March_df,April_df,May_df,June_df,July_df,Aug_df,Sep_df,Oct_df,Nov_df,Dec_df]
combined_bike_df = pd.concat(frames)
combined_bike_df
combined_bike_df.head()
combined_bike_df.to_csv("citi_bike.csv", index=False)
| Tableau Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
def f1(a,b,h):
part_1 = ((a**3)/(-3) + a + 2/3)
part_2 = abs((-b**3)/3 + b + (a**3)/3 - a + (a-b)*h)
part_3 = ((b**3)/3 - b + 2/3)
return part_1 + part_2 + part_3
f1(a=-0.9,b=0.9,h=0.9)
def f0(x):
return -1*x**2 + 1
f0(0.9)
f1(a=-0.9,b=0.9,h=0.18999999999999995)
| CS/CSC413/HW01/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Pandas 数据分析连续(第二部分)
#
#
# <font color = red>注意:代码的输出,是我用标准答案run好的,不要run题目的chunk,输出会消失!另开一个代码块输入你的答案!!!</font>
# ## 抽样
# +
# 创建随机数df
import pandas as pd
import numpy as np
df = pd.read_csv('./Iris.csv')
# -
df
# ### 简单抽样
# 随机抽取五行
df.sample(n=5)
# 每一次run这一chunk,都不会变
df.sample(n=5,random_state = 1)
df.Species.unique()
# 加权抽样
sample = df.sample(n = 20,weights = [20]*50+[50]*50+[30]*50,random_state=1024)
sample
sample.Species.value_counts()
# `value_counts()` 可以统计categorical数据的对应类别的size
# 常用指数:⭐⭐⭐
sample.Species.value_counts()/sample.shape[0] # 求百分比
# 如何更直观的看它们的分布呢?
sample.Species.value_counts().plot(kind = 'bar',
title = 'count - Species',
rot = 0,
figsize = [8,4])
# <div class="alert alert-block alert-success"><b>Step 1</b>:
# <font color = red>第一题:df这个数据行太多了,我们用不到,所以在每一类别里,随机抽选5条样本吧!seed = 1</font>
#
#
# 1. df新增一列,命名为`Species_encoding`,当Species为setosa,则值为1,versicolor,则值为2,virginica,值为3(不能用if elif)【2分】
# 2. 根据每个`Species`进行抽样,在每一类别里,随机抽选5条样本,命名为`iris_sample`。【3分】
# </div>
#
# +
# 不要run,在下面的代码块写答案
# -
# ## 分组【groupby】上
#
# 那么如何根据类别对数据分组?
#
# groupby函数可以帮我们解决问题!
#
# 什么是groupby? 可以总结为3个步骤:
#
# 1. Splitting
#
# 根据分组标准,把数据拆分。
#
#
# 2. Applying
#
# 分组之后,目的是通过
#
# - 聚合函数
# - 转换函数
# - 筛选函数
#
# 对组内数据进行操作(有实例)
#
# 3. Combining
#
#
# 今天这一期,我们只讲【Splitting】的部分!
# <div class="alert alert-block alert-warning">
# <b>注意:</b>
# 没有完成上一题目,无法向下进行哦,请务必完成上题,得到 iris_sample
# </div>
iris_sample.groupby('Species') # 会得到一个DataFrameGroupBy对象
sample_group = iris_sample.groupby('Species')
list(sample_group) # 用list查看
# .ngroups 查看有几组
sample_group.ngroups
# .size() 查看组内容量
sample_group.size()
# <div class="alert alert-block alert-success"><b>Step 2</b>:
#
# 第二题:基于`sample_group`
#
#
# 1. 返回一个df,叫first,为每个group的第一行数据合并【1分】
# 2. 返回一个df,叫last,为每个group的最后一行数据合并 【1分】
# </div>
# +
# first
# -
# +
# last
# -
# 根据分组的名称取出对应数据
sample_group.get_group('Iris-setosa')
# <div class="alert alert-block alert-success"><b>Step 3</b>:
#
# 第三题:基于`sample_group`【提示:filter】
#
#
# 请问那组数据的`SepalWidthCm`的最大值大于3.2,并且`PetalWidthCm`的最小值小于1【2分】
#
# </div>
# ## 小结
#
# 今天就分享到这里,总结一下!
#
# 1. 抽样以及具体参数用法
# 2. groupby的Splitting以及group的基本manipulation
| porto-seguro-safe-driver-prediction/Phase1/Numpy&Pandas/Pandas/Exercises/Pandas_basis_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.0 64-bit (''api_book'': venv)'
# language: python
# name: python3
# ---
# # Data types
#
# ## Strong and dynamic typing
#
# In programming, a **strong typed** language is one that guarantees that the types of all variables are known at compile time. This means that the compiler can check the types of all variables at compile time and will not allow the program to run if the types are incorrect.
#
# A **dynamic typed** language is one that allows the types of variables to be checked at run time. This means that the compiler can not check the types of all variables at compile time, but the program will run if the types are correct.
#
# Python is a dynamic typed language. For example Python allows one to add an integer and a floating point number, but adding an integer to a string produces an error. The gain in flexibility of compiling everything at runtime add big flexibility but hinders performance in some cases.
#
# In order to reduce the number of errors and make your code as bulletproof as possible, it is essential to understand data types and use them them correctly.
# # Data types in Python
#
# A data type or simply type is an attribute of data which tells the compiler or interpreter how the programmer intends to use the data {cite}`wiki:data_type`.
#
# The most common data types in Python are:
#
# * Integer (int)
# * Floating-point number (float)
# * String (str)
# * Boolean (bool)
# * DateTime
#
# To check for a specific type of data in Python, one can use the built in function **type**.
# +
# Importing the datetime library which holds the datetime type
import datetime
# Defining the variables
a = 1.0
b = "1"
c = 1
d = True
e = datetime.datetime.now()
# Checking the types
print(f"Type of {a} is: {type(a)}")
print(f"Type of {b} is: {type(b)}")
print(f"Type of {c} is: {type(c)}")
print(f"Type of {d} is: {type(d)}")
print(f"Type of {e} is: {type(e)}")
# -
# Each data type takes up different space in computer memory.
# +
# Importing the needed package
import sys
# Checking the size of objects
print(f"Size of the float object: {sys.getsizeof(a)} bytes")
print(f"Size of the str object: {sys.getsizeof(b)} bytes")
print(f"Size of the int object: {sys.getsizeof(c)} bytes")
print(f"Size of the boolean object: {sys.getsizeof(d)} bytes")
print(f"Size of the boolean object: {sys.getsizeof(e)} bytes")
# -
# # Functionalities of various data types
# Every Python data type has its own attributes and methods. You can read all of them following the official Python documentation:
#
# https://docs.python.org/3/library/datatypes.html
#
# ## String data type
#
# String data type is probably the most popular data type in terms of methods used. To read the full list of string methods available:
#
# https://docs.python.org/3/library/stdtypes.html#str
#
# Some examples:
# +
# Defining a string
string = "<NAME>"
print(f"Original string: {string}")
# Capitalizing the string
print(f"Capitalized string: {string.capitalize()}")
# All calps
print(f"All caps string: {string.upper()}")
# Checking if the string ends with a specific character
print(f"Does the string end with 'rld'?: {string.endswith('rld')}")
# Checking if the string starts with a specific character
print(f"Does the string starts with 'hell'?: {string.startswith('hell')}")
# Spliting the string into substrings; If no splitting char is defined, it will split by whitespace
print(f"Spliting the string into a list: {string.split()}")
# -
# ## Datetime data type
#
# To read the full list of available datetime methods and other documentation visit:
#
# https://docs.python.org/3/library/datetime.html
#
#
# A datetime object is a single object containing all the information from a date object and a time object.
#
# Like a date object, datetime assumes the current Gregorian calendar extended in both directions; like a time object, datetime assumes there are exactly 3600*24 seconds in every day.
#
# Some examples:
#
# +
# Creating a datetime object
dt = datetime.datetime.now()
print(f"The created datetime object: {dt}")
# Getting the year from the datetime object
print(f"The year from the datetime object: {dt.year}")
# Getting the month from the datetime object
print(f"The month from the datetime object: {dt.month}")
# Getting the day from the datetime object
print(f"The day from the datetime object: {dt.day}")
# Extracting the date from the datetime object
print(f"The date part: {dt.date()}")
# Converting to string (year - month - day hour:minute)
print(f"The datetime object as a string: {dt.strftime('%Y-%m-%d %H:%M')}")
# -
# ## Float data type
#
# To read the full list of available float methods and other documentation visit:
#
# https://www.geeksforgeeks.org/python-float-type-and-its-methods/
#
# Some examples:
# +
# Defining the float data type
float_number = 67.5
print(f"The float number: {float_number}")
# Is it an integer?
print(f"Is the float number an integer? (no decimal part): {float_number.is_integer()}")
# Spliting the float into a ratio of two numbers
print(f"Two integers whose ratio produces the original float number: {float_number.as_integer_ratio()}")
# Hexadeciaml representation of the float number
print(f"Hexadecimal representation of the float number: {float_number.hex()}")
| api-book/_build/html/_sources/chapter-3-python/data-types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GbVJ2cozoRWW"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
#Importing the relevant libraries for the project
# + id="eg2pSY7guffm"
def path_finder(date):
#this function will find the day of the week according to the day year and month
#the purpose is to be able to find what day has more activity.
day_name = ['Lunes','Martes','Miercoles','Jueves','Viernes','Sabado','Domingo']
date = str(date)
day = datetime.datetime.strptime(date, '%Y-%m-%d').weekday()
day_found = day_name[day]
return day_found
def twitter_cleaner(mes):
#this function will clean completely the dataset given by twitter when you download by tweet.
dataset = pd.read_csv('{}.csv'.format(mes)) #Reading the file
mes = str(mes)
dataset = pd.DataFrame(dataset) #Making sure the dataset is on a pandas DataFrame object type
dataset['hora']=pd.to_datetime(dataset['hora']) # Changing feature content type
dataset.drop(['ID del Tweet',
'Enlace permanente de Tweet', 'impresiones promocionado',
'interacciones promocionado',
'tasa de interacción promocionado',
'retweets promocionado',
'respuestas promocionado',
'me gusta promocionado',
'clics de perfil de usuario promocionado',
'clics en URL promocionado',
'clics de etiquetas promocionado',
'ampliaciones de detalles promocionado',
'clics en enlaces permanentes promocionado',
'Se abre la aplicación promocionado',
'descargas de app promocionado',
'seguimientos promocionado',
'enviar Tweet por correo electrónico promocionado',
'marcar teléfono promocionado',
'visualizaciones multimedia promocionado',
'interacciones con el contenido multimedia promocionado',
'Se abre la aplicación', 'descargas de app',
'enviar Tweet por correo electrónico', 'marcar teléfono', 'seguimientos', 'clics en enlaces permanentes'], axis=1, inplace=True) # Dropping not relevant columns
dataset.columns = ['Texto del Tweet', 'Hora', 'impresiones', 'interacciones',
'tasa de interacción en %', 'retweets', 'respuestas', 'me gusta',
'clics de perfil de usuario', 'clics en URL', 'clics de etiquetas',
'ampliaciones de detalles', 'visualizaciones multimedia',
'interacciones con el contenido multimedia']#changing the name of some columns
dataset['tasa de interacción en %'] = dataset['tasa de interacción en %'].apply(lambda x: x*100) #Engineering the columns so we can see it on percentage
dataset['tasa de interacción en %'] = np.round(dataset['tasa de interacción en %'],2) #A simple rounding, too many decimals.
list_to_integer = [
'impresiones', 'interacciones','retweets', 'respuestas', 'me gusta',
'clics de perfil de usuario', 'clics en URL', 'clics de etiquetas',
'ampliaciones de detalles'] #Im too lazy to change those columns to integer one by one
for name in list_to_integer:
dataset[name] = np.int64(dataset[name])#changing the column to integer
dataset['Fecha'] = dataset['Hora'].apply(str).copy()
for i in range(0, int(dataset['Fecha'].shape[0])):
dataset['Fecha'][i] = dataset['Fecha'][i][:10] #Fechas means date, and the actual kind of date I want is only with Year, month and day, not with those weird numbers it had.
dataset['Dia'] = dataset['Fecha'].copy()
dataset['Dia'] = dataset['Dia'].apply(lambda x: path_finder(x))#Implementing the path finder function previously described
dataset['Fecha'] =pd.to_datetime(dataset['Fecha'])
#dataset['Fecha'] = dataset['Fecha'].apply(pd.to_datetime)
#dataset['hora'] = pd.to_datetime(dataset['hora'], format='%H%M')
dataset['Hora'] = dataset['Hora'].apply(str) #Changing hora feature to string it was datetime
e = list(dataset['Hora'].values)
for i in range(0, len(e)):#This for will engineer the hora feature so the values will be two integers like 14, 14 hours of course.
e[i] = e[i][11:13]
dataset['Hora'] = e
dataset['Hora'] = np.int64(dataset['Hora'])
#dataset['Hora'] = pd.Timestamp(dataset['Hora'], unit='h')
dataset = dataset[['Texto del Tweet',
'Fecha',
'Dia',
'Hora',
'impresiones',
'interacciones',
'tasa de interacción en %',
'retweets',
'respuestas',
'me gusta',
'clics de perfil de usuario',
'clics en URL',
'clics de etiquetas',
'ampliaciones de detalles',
'visualizaciones multimedia',
'interacciones con el contenido multimedia']] #making sure the features I want will be in the final result file
dataset.to_csv('{}_clean.csv'.format(mes), encoding='utf-8', index=False) #Saving the engineering.
dataset['impresiones'].describe().to_csv('{}_described_clean.csv'.format(mes), encoding='utf-8', index=True) #A quick statistical description for the numeric values and saving it.
def fb_cleaner_scope(name): #Short function to engineer the date column for facebook content data
df = pd.read_csv('{}.csv'.format(name), encoding='latin-1')
df['Fecha'] = df['Fecha'].apply(lambda x: x[0:10])
df.to_csv('{}_clean.csv'.format(name), encoding='utf-8', index=False)
def fb_cleaner_content(name): #Short function to drop not relevant columns and engineering the hour columns, and saving it immediately
df = pd.read_csv('{}.csv'.format(name), encoding='latin-1')
df = df.drop(columns=['Resultados','Costo por resultado'])
df['Hora de publicación'] = df['Hora de publicación'].apply(lambda x: x[0:10])
df.to_csv('{}_clean.csv'.format(name), encoding='utf-8', index=False)
df['Alcance'].describe().to_csv('{}_described.csv'.format(name))
# + colab={"base_uri": "https://localhost:8080/"} id="UEPjwWQRp5LW" outputId="febdae47-bbfd-4497-afb6-449b890e4aef"
fb_cleaner_content('Contenido feb final fb')
twitter_cleaner('twitter feb')
fb_cleaner_scope('Tendencias Feb fb')
| Libertarian_cleaning_github_version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: septum
# language: python
# name: septum_detection
# ---
# # This notebook could be used to download and pre-process the ACDC RVIP data and labels and slice them into 2D
# +
# ------------------------------------------define logging and working directory
from ProjectRoot import change_wd_to_project_root
change_wd_to_project_root()
from src.utils.Tensorflow_helper import choose_gpu_by_id
# ------------------------------------------jupyter magic config
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# ------------------------------------------ import helpers
# Notebook_imports should import glob, os, and many other standard libs
from src.utils.Notebook_imports import *
# load helper function to slice 3d_volumes into 2d_slices
from src.data.Dataset import ensure_dir, create_2d_slices_from_3d_volume_files
from ipyfilechooser import FileChooser
# -
# # Load CMR and mask file names
# Download and unpack the raw data
# small helper
def clean_import(dir_path):
import shutil
try:
shutil.rmtree(dir_path)
except OSError as e:
print("Error: %s : %s" % (dir_path, e.strerror))
print('Dont worry, irectory will be created.')
ensure_dir(dir_path)
# Please change only the data_root var
dataroot_path = FileChooser()
dataroot_path.title = '<b>Choose a data root, data will be downloaded and extracted in this dir</b>'
display(dataroot_path)
path_to_original_acdc_files = FileChooser()
path_to_original_acdc_files.title = '<b>Choose the path to the original "acdc" root-folder with the patient sub-folders</b>'
display(path_to_original_acdc_files)
# define a folder for the acdc cmr and masks, make sure not to use an existing folder
data_root = dataroot_path.selected
import_path = os.path.join(data_root, 'import')
ensure_dir(data_root)
clean_import(import_path)
# + jupyter={"outputs_hidden": true} tags=[]
# download cleaned rvip 3D cmr and masks
# !wget https://heibox.uni-heidelberg.de/f/8776d7311ec84723aacf/?dl=1 -P {import_path}
print('downloaded')
# unzip and replace
zip_file = glob.glob(os.path.join(import_path,'index.html?dl=*'))[0]
# !unzip -o {zip_file} -d {data_root}
# clean temp import older
clean_import(import_path)
# + tags=[]
# remove old and download new cv-dataframe
# !rm {data_root}df_kfold.csv -f
# !wget https://heibox.uni-heidelberg.de/f/03f57e89dc8b46668144/?dl=1 -P {import_path}
print('downloaded')
# unzip and replace
zip_file = glob.glob(os.path.join(import_path,'index.html?dl=*'))[0]
# !unzip -o {zip_file} -d {data_root}
# clean temp import older
clean_import(import_path)
# -
# # Download original acdc dataset
# + jupyter={"outputs_hidden": true} tags=[]
# !wget https://heibox.uni-heidelberg.de/f/45dc4e1c44754e3c95cb/?dl=1 -P {import_path}
print('downloaded')
# unzip and replace
zip_file = glob.glob(os.path.join(import_path,'index.html?dl=*'))[0]
# !unzip -o {zip_file} -d {data_root}
# clean temp import older
clean_import(import_path)
# -
# # Check the folder structure in data_root
# io == interobserver
# pp == 100 patients x phases xrvip/cmr = 400 files
# !tree -L 1 {data_root}
# # Reuse an existing ACDC directory
#
# --> copy the 3D CMR files from the original ACDC download folder
path_to_acdc_original = path_to_original_acdc_files.selected
print('collect 3D CMR from: {}'.format(path_to_acdc_original))
#searches in all patient folders for any 3D CMR (2 frames per patient) as nifti
images = sorted(glob.glob(os.path.join(path_to_acdc_original, '*/*frame[0-9][0-9].nii.gz')))
print('images: {}'.format(len(images)))
# # Collect 3D CMR and labels/masks in sorted order
#
# --> make sure both lists are of equal length
# quality check of the image and mask names, find names with wrong names
# give input and output path here
input_path = os.path.join(data_root, 'pp')
export_path = os.path.join(data_root, '2D')
#images = sorted(glob.glob(os.path.join(input_path, '*frame[0-9][0-9].nii')))
masks = sorted(glob.glob(os.path.join(input_path, '*frame[0-9][0-9]_rvip.nrrd'))) #searches in all first level folders for any mask as nrrd
print('images: {}'.format(len(images)))
print('masks: {}'.format(len(masks)))
assert(len(images) == len(masks)), 'len(images)-> {} != len(masks)-> {} '.format(len(images), len(masks))
# in the optimal case there should be as many images as masks. If not, some of the annotations might have been saved with a wrong name.
# + tags=[]
# Slice the 3D vol in 2D slices
ensure_dir(export_path)
[create_2d_slices_from_3d_volume_files(img_f=img,mask_f=msk, export_path=export_path) for img,msk in zip(images,masks)]
# -
| notebooks/Dataset/Prepare_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys, os
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from scipy.stats import bayes_mvs as bayesest
import time
sys.path.insert(0, '../../PyEcoLib')
from simulator import Simulator
# %matplotlib inline
# -
mean_size = 3 # micron
doubling_time = 18 #min
tmax = 180 #min
sample_time = 2 #min
div_steps = 10
ncells = 5000
gr = np.log(2)/doubling_time
if not os.path.exists('./figures'):
os.makedirs('./figures')
if not os.path.exists('./data'):
os.makedirs('./data')
sim = Simulator(ncells=ncells, gr = gr, sb=mean_size, steps = div_steps, lamb = 2)
start = time.time()
tmax=10*doubling_time
sim.divstrat(tmax = tmax, sample_time = 0.1*doubling_time, nameDSM = "./data/dataDSM.csv")
print('It took', np.int(time.time()-start), 'seconds.')
start = time.time()
tmax=10*doubling_time
sim.szdyn(tmax = tmax, sample_time= 0.1*doubling_time, nameCRM = "./data/dataCRM.csv")
print('It took', np.int(time.time()-start), 'seconds.')
start = time.time()
sim.szdynFSP(tmax = tmax, nameFSP = "./data/dataFSP.csv")
print('It took', np.int(time.time()-start), 'seconds.')
# +
data2=pd.read_csv("./data/dataDSM.csv")
data2=data2[data2.time>5*doubling_time]
quantnumber=5
pvadd2=data2
CV2d=[]
delt=[]
sb=[]
errcv2d=[]
errdelt=[]
errsb=[]
for i in range(quantnumber):
lperv0=np.percentile(pvadd2.S_b,i*100/quantnumber)
hperv0=np.percentile(pvadd2.S_b,(i+1)*100/quantnumber)
quanta1=pvadd2[pvadd2.S_b>lperv0]
quanta2=quanta1[quanta1.S_b<hperv0]
mean_cntr, var_cntr, std_cntr = bayesest((quanta2.S_d-quanta2.S_b)/np.mean(pvadd2.S_d-pvadd2.S_b),alpha=0.95)
meanv0_cntr, varv0_cntr, stdv0_cntr = bayesest(quanta2.S_b/np.mean(pvadd2.S_b),alpha=0.95)
CV2d.append(var_cntr[0]/mean_cntr[0]**2)
delt.append(mean_cntr[0])
sb.append(meanv0_cntr[0])
errv=(var_cntr[1][1]-var_cntr[0])/mean_cntr[0]**2+2*(mean_cntr[1][1]-mean_cntr[0])*var_cntr[0]/mean_cntr[0]**3
errcv2d.append(errv)
errdelt.append(mean_cntr[1][1]-mean_cntr[0])
errsb.append(meanv0_cntr[1][1]-meanv0_cntr[0])
# -
start = time.time()
sbar=np.linspace(0.5,1.5,100)*mean_size
cv2sz=[]
deltsz=[]
for i in sbar:
Adder,cv2=sim.SdStat(i)
cv2sz.append(cv2)
deltsz.append(Adder)
print('It took', np.int(time.time()-start), 'seconds.')
# +
data2=pd.read_csv("./data/dataDSM.csv")
mn=mean_size
data2=data2[data2.time>3*doubling_time]
fig, ax = plt.subplots(1,2, figsize=(12,4))
ax[0].scatter(data2.S_b/mn,(data2.S_d-data2.S_b)/mn,s=2)
ax[0].errorbar(np.array(sb),np.array(delt),xerr=errsb,yerr=errdelt, fmt='o',mec='k',capsize=5,markersize='8',elinewidth=3,c='#0075BD')
ax[1].errorbar(np.array(sb),CV2d,xerr=errsb,yerr=errcv2d, fmt='o',mec='k',capsize=5,markersize='8',elinewidth=3,c='#0075BD')
ax[1].set_ylim([0,0.2])
ax[0].set_xlabel("$s_b/\overline{s_b}$",size=20)
ax[1].set_xlabel("$s_b/\overline{s_b}$",size=20)
ax[0].set_ylabel("$\Delta/\overline{s_b}$",size=15)
ax[1].set_ylabel("$C_V^2(\Delta)$",size=15)
for l in [0,1]:
ax[l].set_xlim([0.2,2])
ax[l].grid()
ax[l].tick_params(axis='x', labelsize=15)
ax[l].tick_params(axis='y', labelsize=15)
for axis in ['bottom','left']:
ax[l].spines[axis].set_linewidth(2)
ax[l].tick_params(axis='both', width=2,length=6)
for axis in ['top','right']:
ax[l].spines[axis].set_linewidth(0)
ax[l].tick_params(axis='both', width=0,length=6)
ax[0].plot(np.array(sbar)/mean_size, np.array(deltsz)/mean_size, lw=2,c='k',label="$\lambda=2$")
ax[1].plot(np.array(sbar)/mean_size, cv2sz, lw=2,c='k')
plt.savefig('./figures/div_strategy_sizerlike.eps',bbox_inches='tight')
plt.savefig('./figures/div_strategy_sizerlike.svg',bbox_inches='tight')
plt.savefig('./figures/div_strategy_sizerlike.png',bbox_inches='tight')
# +
data1=pd.read_csv("./data/dataCRM.csv")
timearray=data1.time.unique()
mnszarray=[]
cvszarray=[]
errcv2sz=[]
errmnsz=[]
df=data1
del df['time']
for m in range(len(df)):
szs=df.loc[m, :].values.tolist()
mean_cntr, var_cntr, std_cntr = bayesest(szs,alpha=0.95)
mnszarray.append(np.mean(szs))
errmnsz.append(mean_cntr[1][1]-mean_cntr[0])
cvszarray.append(var_cntr[0]/mean_cntr[0]**2)
errv=(var_cntr[1][1]-var_cntr[0])/mean_cntr[0]**2+2*(mean_cntr[1][1]-mean_cntr[0])*var_cntr[0]/mean_cntr[0]**3
errcv2sz.append(errv)
# +
fig, ax = plt.subplots(1,2, figsize=(12,4))
ax[0].plot(np.array(timearray)/doubling_time,np.array(mnszarray))
ax[0].fill_between(np.array(timearray)/doubling_time,np.array(mnszarray)-np.array(errmnsz),np.array(mnszarray)+np.array(errmnsz),
alpha=1, edgecolor='#4db8ff', facecolor='#4db8ff',linewidth=0,label="SSA")
ax[1].plot(np.array(timearray)/doubling_time,np.array(cvszarray))
ax[1].fill_between(np.array(timearray)/doubling_time,np.array(cvszarray)-np.array(errcv2sz),np.array(cvszarray)+np.array(errcv2sz),
alpha=1, edgecolor='#4db8ff', facecolor='#4db8ff',linewidth=0)
ax[0].set_ylabel("$s$ $(\mu m)$",size=20)
ax[1].set_ylabel("$C_V^2(s)$",size=20)
ax[0].set_xlabel(r"$t/\tau$",size=20)
ax[1].set_xlabel(r"$t/\tau$",size=20)
for l in [0,1]:
ax[l].set_xlim([0,7])
taqui=np.arange(0,8,step=1)
ax[l].set_xticks(np.array(taqui))
ax[l].grid()
ax[l].tick_params(axis='x', labelsize=15)
ax[l].tick_params(axis='y', labelsize=15)
for axis in ['bottom','left']:
ax[l].spines[axis].set_linewidth(2)
ax[l].tick_params(axis='both', width=2,length=6)
for axis in ['top','right']:
ax[l].spines[axis].set_linewidth(0)
ax[l].tick_params(axis='both', width=0,length=6)
plt.subplots_adjust(hspace=0.3,wspace=0.3)
data=pd.read_csv("./data/dataFSP.csv")
ax[0].plot(data.time/doubling_time,data.Meansize,ls='--',c='g',label="Numeric")
ax[1].plot(data.time/doubling_time,data.VarSize/data.Meansize**2,ls='--',c='g')
plt.savefig('./figures/size_statisticssizer.svg',bbox_inches='tight')
plt.savefig('./figures/size_statisticssizer.png',bbox_inches='tight')
# -
| development/examples/SizerLike/Sizerlike.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pylab as plot
from astropy.io import ascii,fits
from scipy import interpolate
import grb_catalogs_copy
from BurstCube.LocSim.Detector import *
from BurstCube.LocSim.Spacecraft import *
from astropy.coordinates import SkyCoord
from astropy import units as u
from scipy.optimize import curve_fit
import math
from astropy.table import Table
import pandas as pd
## code to use when reading in GBM effective area in order to get data into the desired format
def getGBMdata(gbmfile=None):
"""Reads the GBM NaI effective area file and returns a numpy array
with two columns ``energy`` and ``aeff``.
Parameters
----------
gbmfile : string
Name of file that contains the GBM data.
Returns
----------
gbmdata : array
numpy array with two columns ``energy`` and ``aeff``
"""
return np.genfromtxt(gbmfile,skip_header=2,names=('energy', 'aeff'))
# +
## bit of useful code for interpolating in log space
def loginterpol(x,y,x1):
f=interpolate.interp1d(np.log10(x),np.log10(y),bounds_error=False,fill_value="extrapolate",kind='linear')
y1=10**f(np.log10(x1))
return y1
def loginterpol2d(x,y,z,x1,y1):
wz=np.where(z==0)[0]
zz=z
zz[wz]=1.
f=interpolate.interp2d(x,y,np.log10(zz),bounds_error=False,fill_value="extrapolate",kind='linear')
z1=10**f(x1,y1)
# +
#read in GBM Trigger Catalog
trigfit=fits.open('gbmtrigcat.fits')
trig=trigfit[1].data
#read in GBM Burst Catalog
gbmfit=fits.open('gbmgrbcat_copy.fits')
gbm=gbmfit[1].data
# -
## generate random positions on the sky with equal area probability
def random_sky(n=1):
u=np.random.rand(n)
v=np.random.rand(n)
phi=2*np.pi*u
theta=np.arccos(2*v-1.)
dec=-np.degrees(theta-np.pi/2.)
ra=np.degrees(np.pi*2-phi)
return ra,dec
#function to match GRBs in the Trigger catalog to those in the grb catalog so that we can create an array of the grbs in both
#We will use the trigger timescale found in the trigger catalog
def match_catalogs_name(name1,name2):
ind_dict = dict((q,e) for e,q in enumerate(name1))
inter = set(ind_dict).intersection(name2)
m1 = [ind_dict[element] for element in inter]
print(np.shape(m1))
ind_dict = dict((q,e) for e,q in enumerate(name2))
inter = set(ind_dict).intersection(name1)
m2 = [ind_dict[element] for element in inter]
print(np.shape(m2))
return m1,m2
# +
#ordering the trig and gbm catalog so that they are in the same order
so=np.argsort(np.array(trig['NAME']))
trig=trig[so]
so=np.argsort(np.array(gbm['NAME']))
gbm=gbm[so]
#creating array of grbs that are found in both catalogs
m1, m2 = match_catalogs_name(trig['NAME'],gbm['NAME'])
#defining our two samples of bursts that are found in both catalogs so that we can utilize them further down
trigbursts = trig[m1]
gbmbursts = gbm[m2]
print(gbmbursts['NAME'])
# -
## read in the GBM Aeff
aeff_gbm = getGBMdata('/home/alyson/NASA/Simulation/BurstCube/Users/ajoens/gbm_effective_area.dat')
## read in BurstCube Aeff for various BC configurations
file='/home/alyson/NASA/Simulation/BurstCube/Users/jracusin/BC_eff_area_curves.ecsv'
bcaeffs=ascii.read(file,format='ecsv')
## separate GBM short & long GRBs
w=np.where(gbmbursts['FLUX_1024']>0)
gbmbursts=gbmbursts[w]
s=np.where((gbmbursts['T90'] <= 2.)&((gbmbursts['PFLX_SPECTRUM_STOP']-gbmbursts['PFLX_SPECTRUM_START'])>0))[0]
l=np.where(gbmbursts['T90'] > 2.)[0]
m=np.where(gbmbursts['PFLX_BEST_FITTING_MODEL'][s] == ' ')
# +
## grab short GRBs with peak spectral info & plot all of the Aeff curves
bceng=bcaeffs['keV']
bcengdiff=bceng[1:]-bceng[0:-1]
w=np.where(bcengdiff<0)[0]
nsims=len(w)
w=np.append(-1,w)#,len(eng))
for i in range(nsims):
plot.plot(bcaeffs['keV'][w[i]+1:w[i+1]+1],bcaeffs['aeff'][w[i]+1:w[i+1]+1])
plot.xscale('log')
plot.yscale('log')
plot.xlabel('Energy (keV)')
plot.ylabel(r'Effective Area (cm$^2$)')
plot.plot(aeff_gbm['energy'],aeff_gbm['aeff'])
i=0
gbmae=loginterpol(aeff_gbm['energy'],aeff_gbm['aeff'],bceng[w[i]+1:w[i+1]+1])
plot.plot(bceng[w[i]+1:w[i+1]+1],gbmae)
plot.show()
# -
## grab energies from those curves and create an array of the energies
E=np.array(bceng[w[i]+1:w[i+1]+1])
print(E)
# +
#Integrating the best fit spectrum for each GRB in the energy range of 50-300 KeV to get max. observed photon flux.
#Doing the same but also folding in the effective area in order to get count rate.
#This will give us the photon flux in units of ph/cm^2/s.
mo=gbmbursts['PFLX_BEST_FITTING_MODEL'][s]
bcpf=np.zeros(len(s))
pf=np.zeros(len(s))
gbmcr=np.zeros(len(s))
bccr=np.zeros(len(s))
outE=np.logspace(np.log10(50),np.log10(300),100) # returns numbers spaced evenly on a log scale
for i in range(len(s)):
for j in range(nsims):
E=np.array(bceng[w[j]+1:w[j+1]+1])
AeffBC=loginterpol(E,bcaeffs['aeff'][w[j]+1:w[j+1]+1],outE)
AeffGBM=loginterpol(aeff_gbm['energy'],aeff_gbm['aeff'],outE) #eng[w[j]+1:w[j+1]+1])
Aratio=(AeffBC/AeffGBM)
# this should give us an array of the maximum observed photon flux for GBM
if mo[i]=='PFLX_PLAW':
gbmcr[i]=np.trapz(gbmbursts['PFLX_PLAW_AMPL'][s[i]]*grb_catalogs_copy.pl(outE,gbmbursts['PFLX_PLAW_INDEX'][s[i]])*AeffGBM,outE)
pf[i]=np.trapz(gbmbursts['PFLX_PLAW_AMPL'][s[i]]*grb_catalogs_copy.pl(outE,gbmbursts['PFLX_PLAW_INDEX'][s[i]]),outE)
bccr[i]=np.trapz(gbmbursts['PFLX_PLAW_AMPL'][s[i]]*grb_catalogs_copy.pl(outE,gbmbursts['PFLX_PLAW_INDEX'][s[i]])*AeffGBM*Aratio,outE)
bcpf[i]=np.trapz(gbmbursts['PFLX_PLAW_AMPL'][s[i]]*grb_catalogs_copy.pl(outE,gbmbursts['PFLX_PLAW_INDEX'][s[i]])*Aratio,outE)
if mo[i]=='PFLX_COMP':
gbmcr[i]=np.trapz(gbmbursts['PFLX_COMP_AMPL'][s[i]]*grb_catalogs_copy.comp(outE,gbmbursts['PFLX_COMP_INDEX'][s[i]],gbmbursts['PFLX_COMP_EPEAK'][s[i]])*AeffGBM,outE)
pf[i]=np.trapz(gbmbursts['PFLX_COMP_AMPL'][s[i]]*grb_catalogs_copy.comp(outE,gbmbursts['PFLX_COMP_INDEX'][s[i]],gbmbursts['PFLX_COMP_EPEAK'][s[i]]),outE)
bccr[i]=np.trapz(gbmbursts['PFLX_COMP_AMPL'][s[i]]*grb_catalogs_copy.comp(outE,gbmbursts['PFLX_COMP_INDEX'][s[i]],gbmbursts['PFLX_COMP_EPEAK'][s[i]])*AeffGBM*Aratio,outE)
bcpf[i]=np.trapz(gbmbursts['PFLX_COMP_AMPL'][s[i]]*grb_catalogs_copy.comp(outE,gbmbursts['PFLX_COMP_INDEX'][s[i]],gbmbursts['PFLX_COMP_EPEAK'][s[i]])*Aratio,outE)
if mo[i]=='PFLX_BAND':
gbmcr[i]=np.trapz(gbmbursts['PFLX_BAND_AMPL'][s[i]]*grb_catalogs_copy.band(outE,gbmbursts['PFLX_BAND_ALPHA'][s[i]],gbmbursts['PFLX_BAND_EPEAK'][s[i]],gbmbursts['PFLX_BAND_BETA'][s[i]])*AeffGBM,outE)
pf[i]=np.trapz(gbmbursts['PFLX_BAND_AMPL'][s[i]]*grb_catalogs_copy.band(outE,gbmbursts['PFLX_BAND_ALPHA'][s[i]],gbmbursts['PFLX_BAND_EPEAK'][s[i]],gbmbursts['PFLX_BAND_BETA'][s[i]]),outE)
bccr[i]=np.trapz(gbmbursts['PFLX_BAND_AMPL'][s[i]]*grb_catalogs_copy.band(outE,gbmbursts['PFLX_BAND_ALPHA'][s[i]],gbmbursts['PFLX_BAND_EPEAK'][s[i]],gbmbursts['PFLX_BAND_BETA'][s[i]])*AeffGBM*Aratio,outE)
bcpf[i]=np.trapz(gbmbursts['PFLX_BAND_AMPL'][s[i]]*grb_catalogs_copy.band(outE,gbmbursts['PFLX_BAND_ALPHA'][s[i]],gbmbursts['PFLX_BAND_EPEAK'][s[i]],gbmbursts['PFLX_BAND_BETA'][s[i]])*Aratio,outE)
if mo[i]=='PFLX_SBPL':
gbmcr[i]=np.trapz(gbmbursts['PFLX_SBPL_AMPL'][s[i]]*grb_catalogs_copy.sbpl(outE,gbmbursts['PFLX_SBPL_INDX1'][s[i]],gbmbursts['PFLX_SBPL_BRKEN'][s[i]],gbm['PFLX_SBPL_INDX2'][s[i]])*AeffGBM,outE)
pf[i]=np.trapz(gbmbursts['PFLX_SBPL_AMPL'][s[i]]*grb_catalogs_copy.sbpl(outE,gbmbursts['PFLX_SBPL_INDX1'][s[i]],gbmbursts['PFLX_SBPL_BRKEN'][s[i]],gbm['PFLX_SBPL_INDX2'][s[i]]),outE)
bccr[i]=np.trapz(gbmbursts['PFLX_SBPL_AMPL'][s[i]]*grb_catalogs_copy.sbpl(outE,gbmbursts['PFLX_SBPL_INDX1'][s[i]],gbmbursts['PFLX_SBPL_BRKEN'][s[i]],gbm['PFLX_SBPL_INDX2'][s[i]])*AeffGBM*Aratio,outE)
bcpf[i]=np.trapz(gbmbursts['PFLX_SBPL_AMPL'][s[i]]*grb_catalogs_copy.sbpl(outE,gbmbursts['PFLX_SBPL_INDX1'][s[i]],gbmbursts['PFLX_SBPL_BRKEN'][s[i]],gbm['PFLX_SBPL_INDX2'][s[i]])*Aratio,outE)
# -
#plot Batse[64] against pf to see if they are the same
flux=gbmbursts['FLUX_BATSE_64'][s]
# +
#define probability
#p = np.array((np.arange(pf.shape[0])+1)**(-1.0))
p = np.array((np.arange(pf.shape[0])+1.05)**(-0.5))
p=p/sum(p)
#randomly sample from the array of photon fluxes found above using our probability function found above so we draw more low flux bursts
#creating our "intrinsic" sample
r=np.random.choice(pf.shape[0], 1200, replace=True, p=p)
simgbmpfsample = np.array(pf[r])
simgbmcr = np.array(gbmcr[r])
simbcpfsample = np.array(bcpf[r])
simbccr = np.array(bccr[r])
# -
#examining our probability distribution to be sure it is performing the eay we intend it to
print(min(p),max(p))
plot.hist(p)
# +
## setup GBM
gbm_pointings = {'01': ('45:54:0','20:36:0'),
'02': ('45:6:0','45:18:0'),
'03': ('58:24:0','90:12:0'),
'04': ('314:54:0','45:12:0'),
'05': ('303:12:0','90:18:0'),
'06': ('3:24:0','89:48:0'),
'07': ('224:54:0','20:24:0'),
'08': ('224:36:0','46:12:0'),
'09': ('236:36:0','90:0:0'),
'10': ('135:12:0','45:36:0'),
'11': ('123:42:0','90:24:0'),
'12': ('183:42:0','90:18:0')}
fermi = Spacecraft(gbm_pointings,window=0.1)
res = 250
rr,dd = np.meshgrid(np.linspace(0,360,res,endpoint=False),np.linspace(-90,90,res))
exposure_positions = np.vstack([rr.ravel(),dd.ravel()])
gbm_exposures = np.array([[ detector.exposure(position[0],position[1]) for position in exposure_positions.T]
for detector in fermi.detectors])
# +
## setup BurstCube
pointings = {'01': ('0:0:0','45:0:0'),
'02': ('90:0:0','45:0:0'),
'03': ('180:0:0','45:0:0'),
'04': ('270:0:0','45:0:0')}
burstcube = Spacecraft(pointings,window=0.1)
res = 250
rr,dd = np.meshgrid(np.linspace(0,360,res,endpoint=False),np.linspace(-90,90,res))
exposure_positions = np.vstack([rr.ravel(),dd.ravel()])
exposures = np.array([[ detector.exposure(position[0],position[1]) for position in exposure_positions.T]
for detector in burstcube.detectors])
# -
#using SkyCoord to convert coordinates to degrees and solve for distances.
def separation(ra1,dec1,ra2,dec2):
c=SkyCoord(ra=ra1*u.deg,dec=dec1*u.deg)
d=SkyCoord(ra=ra2*u.deg,dec=dec2*u.deg)
dist=c.separation(d)
dist=dist.value
return dist
# +
# now that GBM and BurstCube's pointings are set up we will throw GRBs at it and determine the exposure for each GRB.
#generate GRBs and throw them at GBM
def throw_grbs(fermi,minflux,maxflux):
nsims=int(np.round(len(simgbmpfsample)))
ra,dec=random_sky(nsims)
ra=np.array(ra)-180
dec=np.array(dec)
#GBM and BurstCube exposures for each random GRB.
randgbmexposures = np.array([[detector.exposure(ra[i],dec[i]) for i in range(nsims)] for detector in fermi.detectors])
randbcexposures = np.array([[detector.exposure(ra[i],dec[i]) for i in range(nsims)] for detector in burstcube.detectors])
#Order randgbmexposures into descending order
for column in randgbmexposures.T:
newrandgbm = -np.sort(-randgbmexposures.T)
gbmexposures = np.transpose(newrandgbm)
for col in randbcexposures.T:
newrandbc = -np.sort(-randbcexposures.T)
bcexposures = np.transpose(newrandbc)
#Select the second highest exposure value.
#We will use this to ensure the second highest exposure detector has a sig >4.5
secondhighestgbm = gbmexposures[1,:]
secondhighestbc = bcexposures[1,:]
return gbmexposures, bcexposures, secondhighestgbm, secondhighestbc, randgbmexposures, randbcexposures
# -
#define the peak flux interval using the trigger catalog
msinterval = trigbursts['Trigger_Timescale'][s]
interval = msinterval/1000
# +
#flux=simpf this is in ph/sec
flux=simgbmpfsample
minflux=min(flux)
maxflux=max(flux)
gbmexposures, bcexposures, secondhighestgbm, secondhighestbc, randgbmexposures, randbcexposures = throw_grbs(fermi,minflux,maxflux)
# -
#Solve for the number of detected counts which will equal our source photons
sourcegbm = simgbmcr*secondhighestgbm*interval[r]
sourcebc = simbccr*secondhighestbc*interval[r]
# +
#Assuming a background count rate. units: cts/s
bckgrd=300
#scale the background count rate for the second highest detector
scaledgbmbckgrd = bckgrd*secondhighestgbm*interval[r]
scaledbcbckgrd = bckgrd*secondhighestbc*interval[r]
# +
#creating an array of zeros that I can manipulate to create an array of detected GRBs
detectgbm = np.zeros(len(simgbmpfsample))
detectbc = np.zeros(len(simbcpfsample))
#calculate the significance of the second highest exposure detector. If the significance is greater than 4.5 sigma than the burst is detectable.
for u in range(len(simgbmpfsample)):
sig = sourcegbm[u] / (math.sqrt(sourcegbm[u] + scaledgbmbckgrd[u]))
if sig > 4.5:
detectgbm[u] = 1.0
else:
detectgbm[u] = 0.0
for j in range(len(simbcpfsample)):
sig = sourcebc[j] / (math.sqrt(sourcebc[j] + scaledbcbckgrd[j]))
if sig > 4.5:
detectbc[j] = 1.0
else:
detectbc[j] = 0.0
# +
#Creating plot of peak flux versus counts for real and simulated GBM
w=np.where(pf>0)[0]
wg = np.where(simgbmcr*detectgbm>0)[0]
wbc = np.where(simbccr*detectbc>0)[0]
fig=plot.figure(figsize=(20,5))
plot.subplot(1,2,1)
plot.hist(gbmcr[w],label='real GBM',bins=np.logspace(1,6,40),color='orange')
plot.hist(simgbmcr[wg],label='Simulated GBM',bins=np.logspace(1,6,40),alpha=0.7,color='blue')
plot.hist(simbccr[wbc],label='Simulated BurstCube',bins=np.logspace(1,6,40),alpha=0.7,color='green')
plot.xscale('log')
plot.legend()
plot.subplot(1,2,2)
#plot.hist(flux,label='All',bins=np.logspace(-1,2,40),color='green')
#pf has been gathered from the GBM catalog
plot.hist(pf[w],label='real GBM',bins=np.logspace(-1,4,40),color='orange')
# this is the simulated GBM
plot.hist(simgbmpfsample[wg],label='Simulated GBM',bins=np.logspace(-1,4,40),alpha=0.7,color='blue')
plot.hist(simbcpfsample[wbc],label='Simulated BurstCube',bins=np.logspace(-1,4,40),alpha=0.7,color='green')
#plot.hist(flux[w],label='BC',bins=np.logspace(-1,2,40),alpha=0.7,color='red')
plot.xscale('log')
plot.legend()
plot.show()
# +
#solve for the detection fraction of BurstCube and Simulated GBM
detgbm = np.where(detectgbm == 1)[0]
ratiogbm = len(detgbm) / len(detectgbm)
print(ratiogbm)
detbc = np.where(detectbc == 1)[0]
ratiobc = len(detbc) / len(detectbc)
print(ratiobc)
#number of bursts BurstCube will see a year
bcbursts = ratiobc/ratiogbm *40
print(bcbursts)
| ajoens/BurstCube_grbrates_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # Geometry and Linear Algebraic Operations
# :label:`sec_geometry-linear-algebraic-ops`
#
# In :numref:`sec_linear-algebra`, we encountered the basics of linear algebra
# and saw how it could be used to express common operations for transforming our data.
# Linear algebra is one of the key mathematical pillars
# underlying much of the work that we do in deep learning
# and in machine learning more broadly.
# While :numref:`sec_linear-algebra` contained enough machinery
# to communicate the mechanics of modern deep learning models,
# there is a lot more to the subject.
# In this section, we will go deeper,
# highlighting some geometric interpretations of linear algebra operations,
# and introducing a few fundamental concepts, including of eigenvalues and eigenvectors.
#
# ## Geometry of Vectors
# First, we need to discuss the two common geometric interpretations of vectors,
# as either points or directions in space.
# Fundamentally, a vector is a list of numbers such as the Python list below.
#
# + origin_pos=1 tab=["tensorflow"]
v = [1, 7, 0, 1]
# + [markdown] origin_pos=2
# Mathematicians most often write this as either a *column* or *row* vector, which is to say either as
#
# $$
# \mathbf{x} = \begin{bmatrix}1\\7\\0\\1\end{bmatrix},
# $$
#
# or
#
# $$
# \mathbf{x}^\top = \begin{bmatrix}1 & 7 & 0 & 1\end{bmatrix}.
# $$
#
# These often have different interpretations,
# where data examples are column vectors
# and weights used to form weighted sums are row vectors.
# However, it can be beneficial to be flexible.
# As we have described in :numref:`sec_linear-algebra`,
# though a single vector's default orientation is a column vector,
# for any matrix representing a tabular dataset,
# treating each data example as a row vector
# in the matrix
# is more conventional.
#
# Given a vector, the first interpretation
# that we should give it is as a point in space.
# In two or three dimensions, we can visualize these points
# by using the components of the vectors to define
# the location of the points in space compared
# to a fixed reference called the *origin*. This can be seen in :numref:`fig_grid`.
#
# 
# :label:`fig_grid`
#
# This geometric point of view allows us to consider the problem on a more abstract level.
# No longer faced with some insurmountable seeming problem
# like classifying pictures as either cats or dogs,
# we can start considering tasks abstractly
# as collections of points in space and picturing the task
# as discovering how to separate two distinct clusters of points.
#
# In parallel, there is a second point of view
# that people often take of vectors: as directions in space.
# Not only can we think of the vector $\mathbf{v} = [3,2]^\top$
# as the location $3$ units to the right and $2$ units up from the origin,
# we can also think of it as the direction itself
# to take $3$ steps to the right and $2$ steps up.
# In this way, we consider all the vectors in figure :numref:`fig_arrow` the same.
#
# 
# :label:`fig_arrow`
#
# One of the benefits of this shift is that
# we can make visual sense of the act of vector addition.
# In particular, we follow the directions given by one vector,
# and then follow the directions given by the other, as is seen in :numref:`fig_add-vec`.
#
# 
# :label:`fig_add-vec`
#
# Vector subtraction has a similar interpretation.
# By considering the identity that $\mathbf{u} = \mathbf{v} + (\mathbf{u}-\mathbf{v})$,
# we see that the vector $\mathbf{u}-\mathbf{v}$ is the direction
# that takes us from the point $\mathbf{v}$ to the point $\mathbf{u}$.
#
#
# ## Dot Products and Angles
# As we saw in :numref:`sec_linear-algebra`,
# if we take two column vectors $\mathbf{u}$ and $\mathbf{v}$,
# we can form their dot product by computing:
#
# $$\mathbf{u}^\top\mathbf{v} = \sum_i u_i\cdot v_i.$$
# :eqlabel:`eq_dot_def`
#
# Because :eqref:`eq_dot_def` is symmetric, we will mirror the notation
# of classical multiplication and write
#
# $$
# \mathbf{u}\cdot\mathbf{v} = \mathbf{u}^\top\mathbf{v} = \mathbf{v}^\top\mathbf{u},
# $$
#
# to highlight the fact that exchanging the order of the vectors will yield the same answer.
#
# The dot product :eqref:`eq_dot_def` also admits a geometric interpretation: it is closely related to the angle between two vectors. Consider the angle shown in :numref:`fig_angle`.
#
# 
# :label:`fig_angle`
#
# To start, let us consider two specific vectors:
#
# $$
# \mathbf{v} = (r,0) \; \text{and} \; \mathbf{w} = (s\cos(\theta), s \sin(\theta)).
# $$
#
# The vector $\mathbf{v}$ is length $r$ and runs parallel to the $x$-axis,
# and the vector $\mathbf{w}$ is of length $s$ and at angle $\theta$ with the $x$-axis.
# If we compute the dot product of these two vectors, we see that
#
# $$
# \mathbf{v}\cdot\mathbf{w} = rs\cos(\theta) = \|\mathbf{v}\|\|\mathbf{w}\|\cos(\theta).
# $$
#
# With some simple algebraic manipulation, we can rearrange terms to obtain
#
# $$
# \theta = \arccos\left(\frac{\mathbf{v}\cdot\mathbf{w}}{\|\mathbf{v}\|\|\mathbf{w}\|}\right).
# $$
#
# In short, for these two specific vectors,
# the dot product combined with the norms tell us the angle between the two vectors. This same fact is true in general. We will not derive the expression here, however,
# if we consider writing $\|\mathbf{v} - \mathbf{w}\|^2$ in two ways:
# one with the dot product, and the other geometrically using the law of cosines,
# we can obtain the full relationship.
# Indeed, for any two vectors $\mathbf{v}$ and $\mathbf{w}$,
# the angle between the two vectors is
#
# $$\theta = \arccos\left(\frac{\mathbf{v}\cdot\mathbf{w}}{\|\mathbf{v}\|\|\mathbf{w}\|}\right).$$
# :eqlabel:`eq_angle_forumla`
#
# This is a nice result since nothing in the computation references two-dimensions.
# Indeed, we can use this in three or three million dimensions without issue.
#
# As a simple example, let us see how to compute the angle between a pair of vectors:
#
# + origin_pos=5 tab=["tensorflow"]
# %matplotlib inline
import tensorflow as tf
from IPython import display
from d2l import tensorflow as d2l
def angle(v, w):
return tf.acos(tf.tensordot(v, w, axes=1) / (tf.norm(v) * tf.norm(w)))
angle(tf.constant([0, 1, 2], dtype=tf.float32), tf.constant([2.0, 3, 4]))
# + [markdown] origin_pos=6
# We will not use it right now, but it is useful to know
# that we will refer to vectors for which the angle is $\pi/2$
# (or equivalently $90^{\circ}$) as being *orthogonal*.
# By examining the equation above, we see that this happens when $\theta = \pi/2$,
# which is the same thing as $\cos(\theta) = 0$.
# The only way this can happen is if the dot product itself is zero,
# and two vectors are orthogonal if and only if $\mathbf{v}\cdot\mathbf{w} = 0$.
# This will prove to be a helpful formula when understanding objects geometrically.
#
# It is reasonable to ask: why is computing the angle useful?
# The answer comes in the kind of invariance we expect data to have.
# Consider an image, and a duplicate image,
# where every pixel value is the same but $10\%$ the brightness.
# The values of the individual pixels are in general far from the original values.
# Thus, if one computed the distance between the original image and the darker one,
# the distance can be large.
# However, for most ML applications, the *content* is the same---it is still
# an image of a cat as far as a cat/dog classifier is concerned.
# However, if we consider the angle, it is not hard to see
# that for any vector $\mathbf{v}$, the angle
# between $\mathbf{v}$ and $0.1\cdot\mathbf{v}$ is zero.
# This corresponds to the fact that scaling vectors
# keeps the same direction and just changes the length.
# The angle considers the darker image identical.
#
# Examples like this are everywhere.
# In text, we might want the topic being discussed
# to not change if we write twice as long of document that says the same thing.
# For some encoding (such as counting the number of occurrences of words in some vocabulary), this corresponds to a doubling of the vector encoding the document,
# so again we can use the angle.
#
# ### Cosine Similarity
# In ML contexts where the angle is employed
# to measure the closeness of two vectors,
# practitioners adopt the term *cosine similarity*
# to refer to the portion
# $$
# \cos(\theta) = \frac{\mathbf{v}\cdot\mathbf{w}}{\|\mathbf{v}\|\|\mathbf{w}\|}.
# $$
#
# The cosine takes a maximum value of $1$
# when the two vectors point in the same direction,
# a minimum value of $-1$ when they point in opposite directions,
# and a value of $0$ when the two vectors are orthogonal.
# Note that if the components of high-dimensional vectors
# are sampled randomly with mean $0$,
# their cosine will nearly always be close to $0$.
#
#
# ## Hyperplanes
#
# In addition to working with vectors, another key object
# that you must understand to go far in linear algebra
# is the *hyperplane*, a generalization to higher dimensions
# of a line (two dimensions) or of a plane (three dimensions).
# In an $d$-dimensional vector space, a hyperplane has $d-1$ dimensions
# and divides the space into two half-spaces.
#
# Let us start with an example.
# Suppose that we have a column vector $\mathbf{w}=[2,1]^\top$. We want to know, "what are the points $\mathbf{v}$ with $\mathbf{w}\cdot\mathbf{v} = 1$?"
# By recalling the connection between dot products and angles above :eqref:`eq_angle_forumla`,
# we can see that this is equivalent to
# $$
# \|\mathbf{v}\|\|\mathbf{w}\|\cos(\theta) = 1 \; \iff \; \|\mathbf{v}\|\cos(\theta) = \frac{1}{\|\mathbf{w}\|} = \frac{1}{\sqrt{5}}.
# $$
#
# 
# :label:`fig_vector-project`
#
# If we consider the geometric meaning of this expression,
# we see that this is equivalent to saying
# that the length of the projection of $\mathbf{v}$
# onto the direction of $\mathbf{w}$ is exactly $1/\|\mathbf{w}\|$, as is shown in :numref:`fig_vector-project`.
# The set of all points where this is true is a line
# at right angles to the vector $\mathbf{w}$.
# If we wanted, we could find the equation for this line
# and see that it is $2x + y = 1$ or equivalently $y = 1 - 2x$.
#
# If we now look at what happens when we ask about the set of points with
# $\mathbf{w}\cdot\mathbf{v} > 1$ or $\mathbf{w}\cdot\mathbf{v} < 1$,
# we can see that these are cases where the projections
# are longer or shorter than $1/\|\mathbf{w}\|$, respectively.
# Thus, those two inequalities define either side of the line.
# In this way, we have found a way to cut our space into two halves,
# where all the points on one side have dot product below a threshold,
# and the other side above as we see in :numref:`fig_space-division`.
#
# 
# :label:`fig_space-division`
#
# The story in higher dimension is much the same.
# If we now take $\mathbf{w} = [1,2,3]^\top$
# and ask about the points in three dimensions with $\mathbf{w}\cdot\mathbf{v} = 1$,
# we obtain a plane at right angles to the given vector $\mathbf{w}$.
# The two inequalities again define the two sides of the plane as is shown in :numref:`fig_higher-division`.
#
# 
# :label:`fig_higher-division`
#
# While our ability to visualize runs out at this point,
# nothing stops us from doing this in tens, hundreds, or billions of dimensions.
# This occurs often when thinking about machine learned models.
# For instance, we can understand linear classification models
# like those from :numref:`sec_softmax`,
# as methods to find hyperplanes that separate the different target classes.
# In this context, such hyperplanes are often referred to as *decision planes*.
# The majority of deep learned classification models end
# with a linear layer fed into a softmax,
# so one can interpret the role of the deep neural network
# to be to find a non-linear embedding such that the target classes
# can be separated cleanly by hyperplanes.
#
# To give a hand-built example, notice that we can produce a reasonable model
# to classify tiny images of t-shirts and trousers from the Fashion MNIST dataset
# (seen in :numref:`sec_fashion_mnist`)
# by just taking the vector between their means to define the decision plane
# and eyeball a crude threshold. First we will load the data and compute the averages.
#
# + origin_pos=9 tab=["tensorflow"]
# Load in the dataset
((train_images, train_labels), (
test_images, test_labels)) = tf.keras.datasets.fashion_mnist.load_data()
X_train_0 = tf.cast(tf.stack(train_images[[i for i, label in enumerate(
train_labels) if label == 0]] * 256), dtype=tf.float32)
X_train_1 = tf.cast(tf.stack(train_images[[i for i, label in enumerate(
train_labels) if label == 1]] * 256), dtype=tf.float32)
X_test = tf.cast(tf.stack(test_images[[i for i, label in enumerate(
test_labels) if label == 0]] * 256), dtype=tf.float32)
y_test = tf.cast(tf.stack(test_images[[i for i, label in enumerate(
test_labels) if label == 1]] * 256), dtype=tf.float32)
# Compute averages
ave_0 = tf.reduce_mean(X_train_0, axis=0)
ave_1 = tf.reduce_mean(X_train_1, axis=0)
# + [markdown] origin_pos=10
# It can be informative to examine these averages in detail, so let us plot what they look like. In this case, we see that the average indeed resembles a blurry image of a t-shirt.
#
# + origin_pos=12 tab=["tensorflow"]
# Plot average t-shirt
d2l.set_figsize()
d2l.plt.imshow(tf.reshape(ave_0, (28, 28)), cmap='Greys')
d2l.plt.show()
# + [markdown] origin_pos=13
# In the second case, we again see that the average resembles a blurry image of trousers.
#
# + origin_pos=15 tab=["tensorflow"]
# Plot average trousers
d2l.plt.imshow(tf.reshape(ave_1, (28, 28)), cmap='Greys')
d2l.plt.show()
# + [markdown] origin_pos=16
# In a fully machine learned solution, we would learn the threshold from the dataset. In this case, I simply eyeballed a threshold that looked good on the training data by hand.
#
# + origin_pos=19 tab=["tensorflow"]
# Print test set accuracy with eyeballed threshold
w = tf.transpose(ave_1 - ave_0)
predictions = tf.reduce_sum(X_test * tf.nest.flatten(w), axis=0) > -1500000
# Accuracy
tf.reduce_mean(
tf.cast(tf.cast(predictions, y_test.dtype) == y_test, tf.float32))
# + [markdown] origin_pos=20
# ## Geometry of Linear Transformations
#
# Through :numref:`sec_linear-algebra` and the above discussions,
# we have a solid understanding of the geometry of vectors, lengths, and angles.
# However, there is one important object we have omitted discussing,
# and that is a geometric understanding of linear transformations represented by matrices. Fully internalizing what matrices can do to transform data
# between two potentially different high dimensional spaces takes significant practice,
# and is beyond the scope of this appendix.
# However, we can start building up intuition in two dimensions.
#
# Suppose that we have some matrix:
#
# $$
# \mathbf{A} = \begin{bmatrix}
# a & b \\ c & d
# \end{bmatrix}.
# $$
#
# If we want to apply this to an arbitrary vector
# $\mathbf{v} = [x, y]^\top$,
# we multiply and see that
#
# $$
# \begin{aligned}
# \mathbf{A}\mathbf{v} & = \begin{bmatrix}a & b \\ c & d\end{bmatrix}\begin{bmatrix}x \\ y\end{bmatrix} \\
# & = \begin{bmatrix}ax+by\\ cx+dy\end{bmatrix} \\
# & = x\begin{bmatrix}a \\ c\end{bmatrix} + y\begin{bmatrix}b \\d\end{bmatrix} \\
# & = x\left\{\mathbf{A}\begin{bmatrix}1\\0\end{bmatrix}\right\} + y\left\{\mathbf{A}\begin{bmatrix}0\\1\end{bmatrix}\right\}.
# \end{aligned}
# $$
#
# This may seem like an odd computation,
# where something clear became somewhat impenetrable.
# However, it tells us that we can write the way
# that a matrix transforms *any* vector
# in terms of how it transforms *two specific vectors*:
# $[1,0]^\top$ and $[0,1]^\top$.
# This is worth considering for a moment.
# We have essentially reduced an infinite problem
# (what happens to any pair of real numbers)
# to a finite one (what happens to these specific vectors).
# These vectors are an example a *basis*,
# where we can write any vector in our space
# as a weighted sum of these *basis vectors*.
#
# Let us draw what happens when we use the specific matrix
#
# $$
# \mathbf{A} = \begin{bmatrix}
# 1 & 2 \\
# -1 & 3
# \end{bmatrix}.
# $$
#
# If we look at the specific vector $\mathbf{v} = [2, -1]^\top$,
# we see this is $2\cdot[1,0]^\top + -1\cdot[0,1]^\top$,
# and thus we know that the matrix $A$ will send this to
# $2(\mathbf{A}[1,0]^\top) + -1(\mathbf{A}[0,1])^\top = 2[1, -1]^\top - [2,3]^\top = [0, -5]^\top$.
# If we follow this logic through carefully,
# say by considering the grid of all integer pairs of points,
# we see that what happens is that the matrix multiplication
# can skew, rotate, and scale the grid,
# but the grid structure must remain as you see in :numref:`fig_grid-transform`.
#
# 
# :label:`fig_grid-transform`
#
# This is the most important intuitive point
# to internalize about linear transformations represented by matrices.
# Matrices are incapable of distorting some parts of space differently than others.
# All they can do is take the original coordinates on our space
# and skew, rotate, and scale them.
#
# Some distortions can be severe. For instance the matrix
#
# $$
# \mathbf{B} = \begin{bmatrix}
# 2 & -1 \\ 4 & -2
# \end{bmatrix},
# $$
#
# compresses the entire two-dimensional plane down to a single line.
# Identifying and working with such transformations are the topic of a later section,
# but geometrically we can see that this is fundamentally different
# from the types of transformations we saw above.
# For instance, the result from matrix $\mathbf{A}$ can be "bent back" to the original grid. The results from matrix $\mathbf{B}$ cannot
# because we will never know where the vector $[1,2]^\top$ came from---was
# it $[1,1]^\top$ or $[0, -1]^\top$?
#
# While this picture was for a $2\times2$ matrix,
# nothing prevents us from taking the lessons learned into higher dimensions.
# If we take similar basis vectors like $[1,0, \ldots,0]$
# and see where our matrix sends them,
# we can start to get a feeling for how the matrix multiplication
# distorts the entire space in whatever dimension space we are dealing with.
#
# ## Linear Dependence
#
# Consider again the matrix
#
# $$
# \mathbf{B} = \begin{bmatrix}
# 2 & -1 \\ 4 & -2
# \end{bmatrix}.
# $$
#
# This compresses the entire plane down to live on the single line $y = 2x$.
# The question now arises: is there some way we can detect this
# just looking at the matrix itself?
# The answer is that indeed we can.
# Let us take $\mathbf{b}_1 = [2,4]^\top$ and $\mathbf{b}_2 = [-1, -2]^\top$
# be the two columns of $\mathbf{B}$.
# Remember that we can write everything transformed by the matrix $\mathbf{B}$
# as a weighted sum of the columns of the matrix:
# like $a_1\mathbf{b}_1 + a_2\mathbf{b}_2$.
# We call this a *linear combination*.
# The fact that $\mathbf{b}_1 = -2\cdot\mathbf{b}_2$
# means that we can write any linear combination of those two columns
# entirely in terms of say $\mathbf{b}_2$ since
#
# $$
# a_1\mathbf{b}_1 + a_2\mathbf{b}_2 = -2a_1\mathbf{b}_2 + a_2\mathbf{b}_2 = (a_2-2a_1)\mathbf{b}_2.
# $$
#
# This means that one of the columns is, in a sense, redundant
# because it does not define a unique direction in space.
# This should not surprise us too much
# since we already saw that this matrix
# collapses the entire plane down into a single line.
# Moreover, we see that the linear dependence
# $\mathbf{b}_1 = -2\cdot\mathbf{b}_2$ captures this.
# To make this more symmetrical between the two vectors, we will write this as
#
# $$
# \mathbf{b}_1 + 2\cdot\mathbf{b}_2 = 0.
# $$
#
# In general, we will say that a collection of vectors
# $\mathbf{v}_1, \ldots, \mathbf{v}_k$ are *linearly dependent*
# if there exist coefficients $a_1, \ldots, a_k$ *not all equal to zero* so that
#
# $$
# \sum_{i=1}^k a_i\mathbf{v_i} = 0.
# $$
#
# In this case, we can solve for one of the vectors
# in terms of some combination of the others,
# and effectively render it redundant.
# Thus, a linear dependence in the columns of a matrix
# is a witness to the fact that our matrix
# is compressing the space down to some lower dimension.
# If there is no linear dependence we say the vectors are *linearly independent*.
# If the columns of a matrix are linearly independent,
# no compression occurs and the operation can be undone.
#
# ## Rank
#
# If we have a general $n\times m$ matrix,
# it is reasonable to ask what dimension space the matrix maps into.
# A concept known as the *rank* will be our answer.
# In the previous section, we noted that a linear dependence
# bears witness to compression of space into a lower dimension
# and so we will be able to use this to define the notion of rank.
# In particular, the rank of a matrix $\mathbf{A}$
# is the largest number of linearly independent columns
# amongst all subsets of columns. For example, the matrix
#
# $$
# \mathbf{B} = \begin{bmatrix}
# 2 & 4 \\ -1 & -2
# \end{bmatrix},
# $$
#
# has $\mathrm{rank}(B)=1$, since the two columns are linearly dependent,
# but either column by itself is not linearly dependent.
# For a more challenging example, we can consider
#
# $$
# \mathbf{C} = \begin{bmatrix}
# 1& 3 & 0 & -1 & 0 \\
# -1 & 0 & 1 & 1 & -1 \\
# 0 & 3 & 1 & 0 & -1 \\
# 2 & 3 & -1 & -2 & 1
# \end{bmatrix},
# $$
#
# and show that $\mathbf{C}$ has rank two since, for instance,
# the first two columns are linearly independent,
# however any of the four collections of three columns are dependent.
#
# This procedure, as described, is very inefficient.
# It requires looking at every subset of the columns of our given matrix,
# and thus is potentially exponential in the number of columns.
# Later we will see a more computationally efficient way
# to compute the rank of a matrix, but for now,
# this is sufficient to see that the concept
# is well defined and understand the meaning.
#
# ## Invertibility
#
# We have seen above that multiplication by a matrix with linearly dependent columns
# cannot be undone, i.e., there is no inverse operation that can always recover the input. However, multiplication by a full-rank matrix
# (i.e., some $\mathbf{A}$ that is $n \times n$ matrix with rank $n$),
# we should always be able to undo it. Consider the matrix
#
# $$
# \mathbf{I} = \begin{bmatrix}
# 1 & 0 & \cdots & 0 \\
# 0 & 1 & \cdots & 0 \\
# \vdots & \vdots & \ddots & \vdots \\
# 0 & 0 & \cdots & 1
# \end{bmatrix}.
# $$
#
# which is the matrix with ones along the diagonal, and zeros elsewhere.
# We call this the *identity* matrix.
# It is the matrix which leaves our data unchanged when applied.
# To find a matrix which undoes what our matrix $\mathbf{A}$ has done,
# we want to find a matrix $\mathbf{A}^{-1}$ such that
#
# $$
# \mathbf{A}^{-1}\mathbf{A} = \mathbf{A}\mathbf{A}^{-1} = \mathbf{I}.
# $$
#
# If we look at this as a system, we have $n \times n$ unknowns
# (the entries of $\mathbf{A}^{-1}$) and $n \times n$ equations
# (the equality that needs to hold between every entry of the product $\mathbf{A}^{-1}\mathbf{A}$ and every entry of $\mathbf{I}$)
# so we should generically expect a solution to exist.
# Indeed, in the next section we will see a quantity called the *determinant*,
# which has the property that as long as the determinant is not zero, we can find a solution. We call such a matrix $\mathbf{A}^{-1}$ the *inverse* matrix.
# As an example, if $\mathbf{A}$ is the general $2 \times 2$ matrix
#
# $$
# \mathbf{A} = \begin{bmatrix}
# a & b \\
# c & d
# \end{bmatrix},
# $$
#
# then we can see that the inverse is
#
# $$
# \frac{1}{ad-bc} \begin{bmatrix}
# d & -b \\
# -c & a
# \end{bmatrix}.
# $$
#
# We can test to see this by seeing that multiplying
# by the inverse given by the formula above works in practice.
#
# + origin_pos=23 tab=["tensorflow"]
M = tf.constant([[1, 2], [1, 4]], dtype=tf.float32)
M_inv = tf.constant([[2, -1], [-0.5, 0.5]])
tf.matmul(M_inv, M)
# + [markdown] origin_pos=24
# ### Numerical Issues
# While the inverse of a matrix is useful in theory,
# we must say that most of the time we do not wish
# to *use* the matrix inverse to solve a problem in practice.
# In general, there are far more numerically stable algorithms
# for solving linear equations like
#
# $$
# \mathbf{A}\mathbf{x} = \mathbf{b},
# $$
#
# than computing the inverse and multiplying to get
#
# $$
# \mathbf{x} = \mathbf{A}^{-1}\mathbf{b}.
# $$
#
# Just as division by a small number can lead to numerical instability,
# so can inversion of a matrix which is close to having low rank.
#
# Moreover, it is common that the matrix $\mathbf{A}$ is *sparse*,
# which is to say that it contains only a small number of non-zero values.
# If we were to explore examples, we would see
# that this does not mean the inverse is sparse.
# Even if $\mathbf{A}$ was a $1$ million by $1$ million matrix
# with only $5$ million non-zero entries
# (and thus we need only store those $5$ million),
# the inverse will typically have almost every entry non-negative,
# requiring us to store all $1\text{M}^2$ entries---that is $1$ trillion entries!
#
# While we do not have time to dive all the way into the thorny numerical issues
# frequently encountered when working with linear algebra,
# we want to provide you with some intuition about when to proceed with caution,
# and generally avoiding inversion in practice is a good rule of thumb.
#
# ## Determinant
# The geometric view of linear algebra gives an intuitive way
# to interpret a fundamental quantity known as the *determinant*.
# Consider the grid image from before, but now with a highlighted region (:numref:`fig_grid-filled`).
#
# 
# :label:`fig_grid-filled`
#
# Look at the highlighted square. This is a square with edges given
# by $(0, 1)$ and $(1, 0)$ and thus it has area one.
# After $\mathbf{A}$ transforms this square,
# we see that it becomes a parallelogram.
# There is no reason this parallelogram should have the same area
# that we started with, and indeed in the specific case shown here of
#
# $$
# \mathbf{A} = \begin{bmatrix}
# 1 & 2 \\
# -1 & 3
# \end{bmatrix},
# $$
#
# it is an exercise in coordinate geometry to compute
# the area of this parallelogram and obtain that the area is $5$.
#
# In general, if we have a matrix
#
# $$
# \mathbf{A} = \begin{bmatrix}
# a & b \\
# c & d
# \end{bmatrix},
# $$
#
# we can see with some computation that the area
# of the resulting parallelogram is $ad-bc$.
# This area is referred to as the *determinant*.
#
# Let us check this quickly with some example code.
#
# + origin_pos=27 tab=["tensorflow"]
tf.linalg.det(tf.constant([[1, -1], [2, 3]], dtype=tf.float32))
# + [markdown] origin_pos=28
# The eagle-eyed amongst us will notice
# that this expression can be zero or even negative.
# For the negative term, this is a matter of convention
# taken generally in mathematics:
# if the matrix flips the figure,
# we say the area is negated.
# Let us see now that when the determinant is zero, we learn more.
#
# Let us consider
#
# $$
# \mathbf{B} = \begin{bmatrix}
# 2 & 4 \\ -1 & -2
# \end{bmatrix}.
# $$
#
# If we compute the determinant of this matrix,
# we get $2\cdot(-2 ) - 4\cdot(-1) = 0$.
# Given our understanding above, this makes sense.
# $\mathbf{B}$ compresses the square from the original image
# down to a line segment, which has zero area.
# And indeed, being compressed into a lower dimensional space
# is the only way to have zero area after the transformation.
# Thus we see the following result is true:
# a matrix $A$ is invertible if and only if
# the determinant is not equal to zero.
#
# As a final comment, imagine that we have any figure drawn on the plane.
# Thinking like computer scientists, we can decompose
# that figure into a collection of little squares
# so that the area of the figure is in essence
# just the number of squares in the decomposition.
# If we now transform that figure by a matrix,
# we send each of these squares to parallelograms,
# each one of which has area given by the determinant.
# We see that for any figure, the determinant gives the (signed) number
# that a matrix scales the area of any figure.
#
# Computing determinants for larger matrices can be laborious,
# but the intuition is the same.
# The determinant remains the factor
# that $n\times n$ matrices scale $n$-dimensional volumes.
#
# ## Tensors and Common Linear Algebra Operations
#
# In :numref:`sec_linear-algebra` the concept of tensors was introduced.
# In this section, we will dive more deeply into tensor contractions
# (the tensor equivalent of matrix multiplication),
# and see how it can provide a unified view
# on a number of matrix and vector operations.
#
# With matrices and vectors we knew how to multiply them to transform data.
# We need to have a similar definition for tensors if they are to be useful to us.
# Think about matrix multiplication:
#
# $$
# \mathbf{C} = \mathbf{A}\mathbf{B},
# $$
#
# or equivalently
#
# $$ c_{i, j} = \sum_{k} a_{i, k}b_{k, j}.$$
#
# This pattern is one we can repeat for tensors.
# For tensors, there is no one case of what
# to sum over that can be universally chosen,
# so we need specify exactly which indices we want to sum over.
# For instance we could consider
#
# $$
# y_{il} = \sum_{jk} x_{ijkl}a_{jk}.
# $$
#
# Such a transformation is called a *tensor contraction*.
# It can represent a far more flexible family of transformations
# that matrix multiplication alone.
#
# As a often-used notational simplification,
# we can notice that the sum is over exactly those indices
# that occur more than once in the expression,
# thus people often work with *Einstein notation*,
# where the summation is implicitly taken over all repeated indices.
# This gives the compact expression:
#
# $$
# y_{il} = x_{ijkl}a_{jk}.
# $$
#
# ### Common Examples from Linear Algebra
#
# Let us see how many of the linear algebraic definitions
# we have seen before can be expressed in this compressed tensor notation:
#
# * $\mathbf{v} \cdot \mathbf{w} = \sum_i v_iw_i$
# * $\|\mathbf{v}\|_2^{2} = \sum_i v_iv_i$
# * $(\mathbf{A}\mathbf{v})_i = \sum_j a_{ij}v_j$
# * $(\mathbf{A}\mathbf{B})_{ik} = \sum_j a_{ij}b_{jk}$
# * $\mathrm{tr}(\mathbf{A}) = \sum_i a_{ii}$
#
# In this way, we can replace a myriad of specialized notations with short tensor expressions.
#
# ### Expressing in Code
# Tensors may flexibly be operated on in code as well.
# As seen in :numref:`sec_linear-algebra`,
# we can create tensors as is shown below.
#
# + origin_pos=31 tab=["tensorflow"]
# Define tensors
B = tf.constant([[[1, 2, 3], [4, 5, 6]], [[7, 8, 9], [10, 11, 12]]])
A = tf.constant([[1, 2], [3, 4]])
v = tf.constant([1, 2])
# Print out the shapes
A.shape, B.shape, v.shape
# + [markdown] origin_pos=32
# Einstein summation has been implemented directly.
# The indices that occurs in the Einstein summation can be passed as a string,
# followed by the tensors that are being acted upon.
# For instance, to implement matrix multiplication,
# we can consider the Einstein summation seen above
# ($\mathbf{A}\mathbf{v} = a_{ij}v_j$)
# and strip out the indices themselves to get the implementation:
#
# + origin_pos=35 tab=["tensorflow"]
# Reimplement matrix multiplication
tf.einsum("ij, j -> i", A, v), tf.matmul(A, tf.reshape(v, (2, 1)))
# + [markdown] origin_pos=36
# This is a highly flexible notation.
# For instance if we want to compute
# what would be traditionally written as
#
# $$
# c_{kl} = \sum_{ij} \mathbf{b}_{ijk}\mathbf{a}_{il}v_j.
# $$
#
# it can be implemented via Einstein summation as:
#
# + origin_pos=39 tab=["tensorflow"]
tf.einsum("ijk, il, j -> kl", B, A, v)
# + [markdown] origin_pos=40
# This notation is readable and efficient for humans,
# however bulky if for whatever reason
# we need to generate a tensor contraction programmatically.
# For this reason, `einsum` provides an alternative notation
# by providing integer indices for each tensor.
# For example, the same tensor contraction can also be written as:
#
# + origin_pos=43 tab=["tensorflow"]
# TensorFlow doesn't support this type of notation.
# + [markdown] origin_pos=44
# Either notation allows for concise and efficient representation of tensor contractions in code.
#
# ## Summary
# * Vectors can be interpreted geometrically as either points or directions in space.
# * Dot products define the notion of angle to arbitrarily high-dimensional spaces.
# * Hyperplanes are high-dimensional generalizations of lines and planes. They can be used to define decision planes that are often used as the last step in a classification task.
# * Matrix multiplication can be geometrically interpreted as uniform distortions of the underlying coordinates. They represent a very restricted, but mathematically clean, way to transform vectors.
# * Linear dependence is a way to tell when a collection of vectors are in a lower dimensional space than we would expect (say you have $3$ vectors living in a $2$-dimensional space). The rank of a matrix is the size of the largest subset of its columns that are linearly independent.
# * When a matrix's inverse is defined, matrix inversion allows us to find another matrix that undoes the action of the first. Matrix inversion is useful in theory, but requires care in practice owing to numerical instability.
# * Determinants allow us to measure how much a matrix expands or contracts a space. A nonzero determinant implies an invertible (non-singular) matrix and a zero-valued determinant means that the matrix is non-invertible (singular).
# * Tensor contractions and Einstein summation provide for a neat and clean notation for expressing many of the computations that are seen in machine learning.
#
# ## Exercises
# 1. What is the angle between
# $$
# \vec v_1 = \begin{bmatrix}
# 1 \\ 0 \\ -1 \\ 2
# \end{bmatrix}, \qquad \vec v_2 = \begin{bmatrix}
# 3 \\ 1 \\ 0 \\ 1
# # \end{bmatrix}?
# $$
# 2. True or false: $\begin{bmatrix}1 & 2\\0&1\end{bmatrix}$ and $\begin{bmatrix}1 & -2\\0&1\end{bmatrix}$ are inverses of one another?
# 3. Suppose that we draw a shape in the plane with area $100\mathrm{m}^2$. What is the area after transforming the figure by the matrix
# $$
# \begin{bmatrix}
# 2 & 3\\
# 1 & 2
# \end{bmatrix}.
# $$
# 4. Which of the following sets of vectors are linearly independent?
# * $\left\{\begin{pmatrix}1\\0\\-1\end{pmatrix}, \begin{pmatrix}2\\1\\-1\end{pmatrix}, \begin{pmatrix}3\\1\\1\end{pmatrix}\right\}$
# * $\left\{\begin{pmatrix}3\\1\\1\end{pmatrix}, \begin{pmatrix}1\\1\\1\end{pmatrix}, \begin{pmatrix}0\\0\\0\end{pmatrix}\right\}$
# * $\left\{\begin{pmatrix}1\\1\\0\end{pmatrix}, \begin{pmatrix}0\\1\\-1\end{pmatrix}, \begin{pmatrix}1\\0\\1\end{pmatrix}\right\}$
# 5. Suppose that you have a matrix written as $A = \begin{bmatrix}c\\d\end{bmatrix}\cdot\begin{bmatrix}a & b\end{bmatrix}$ for some choice of values $a, b, c$, and $d$. True or false: the determinant of such a matrix is always $0$?
# 6. The vectors $e_1 = \begin{bmatrix}1\\0\end{bmatrix}$ and $e_2 = \begin{bmatrix}0\\1\end{bmatrix}$ are orthogonal. What is the condition on a matrix $A$ so that $Ae_1$ and $Ae_2$ are orthogonal?
# 7. How can you write $\mathrm{tr}(\mathbf{A}^4)$ in Einstein notation for an arbitrary matrix $A$?
#
# + [markdown] origin_pos=47 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1085)
#
| d2l/tensorflow/chapter_appendix-mathematics-for-deep-learning/geometry-linear-algebraic-ops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:farallon-fall-2020]
# language: python
# name: conda-env-farallon-fall-2020-py
# ---
# # Data Exploration
# +
# Modules
import warnings
warnings.simplefilter('ignore') # filter some warning messages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import xarray as xr
import seaborn as sns
import datetime as dt
import fsspec
import s3fs
import scipy.stats as stats
# make datasets display nicely
xr.set_options(display_style="html")
#magic fncts #put static images of your plot embedded in the notebook
# %matplotlib inline
plt.rcParams['figure.figsize'] = 12, 6
# %config InlineBackend.figure_format = 'retina'
# -
# # Mole Crab Data
# +
### Read and explore mole crab data
crabs = pd.read_csv('./MoleCrab_abundance_annual.csv',index_col=2)
crabs.head()
# -
crabs = pd.read_csv('./MoleCrab_abundance_annual.csv',index_col=2)
crabs.tail()
### Plot a distribution of abundance
sns.distplot(crabs['Abundance (psm)'])
# +
### Plot annual data
plt.bar(crabs.index,crabs['Abundance (psm)'])
plt.xticks([*range(2000,2020,2)])
plt.grid()
plt.show()
# +
### Read Sea Surface Data Available in zar
file_location = 's3://mur-sst/zarr'
ikey = fsspec.get_mapper(file_location, anon=True)
ds_sst = xr.open_zarr(ikey,consolidated=True)
#ds_sst
# +
### Read data that matches crab data, in time and location
sst_timeseries = ds_sst['analysed_sst'].sel(time = slice('2003-01-01','2016-12-31'),
lat = 37.76,
lon = -124.5
).load()
### Plot and explore it
sst_timeseries.plot()
sst_timeseries
# +
### Average data annually
sst_annual = sst_timeseries.groupby('time.year').mean('time',keep_attrs=True)
sst_annual.plot()
# -
# ### Add temperature data to the crab data frame
#
# crabs = crabs[crabs.index>=2003]
#
# tmp = pd.DataFrame(data=sst_annual.data - 273.15, columns={'SST'}, index=[*range(2003,2018)])
#
# crabs['SST'] = tmp
#
# ### Take a look
#
# crabs
# ### Scatter Plot
#
# plt.figure(figsize=(8,6))
#
# plt.plot(crabs['SST'],crabs['Abundance (psm)'],'*')
#
# ### Correlation
# stats.pearsonr(crabs['SST'], crabs['Abundance (psm)'])
# ### Time series plot
#
# plt.figure(figsize=(8,6))
#
# plt.plot(crabs['SST'],crabs['Abundance (psm)'],'*')
#
# fig, ax1 = plt.subplots()
#
# color = 'tab:red'
#
# ax1.set_ylabel('SST', color='tab:red')
#
# ax1.plot(crabs.index, crabs['SST'], color='tab:red')
#
# ax1.tick_params(axis='y', labelcolor='tab:red')
#
# ax2 = ax1.twinx() # instantiate a second axes that shares the same x-axis
#
# color = 'tab:blue'
#
# ax2.set_ylabel('sin', color=color) # we already handled the x-label with ax1
#
# ax2.plot(t, data2, color=color)
#
# ax2.tick_params(axis='y', labelcolor=color)
#
# fig.tight_layout()
#
| notebooks/CalAcademy/.ipynb_checkpoints/CalAcademy_explore_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
from __future__ import print_function
import numpy as np
import numpy
import PIL
from PIL import Image
np.random.seed(1337) # for reproducibility
import random
from keras.datasets import mnist
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Input, Lambda
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.layers import Flatten
from keras.optimizers import RMSprop
from keras import backend as K
from keras.layers import Concatenate, Dense, LSTM, Input, concatenate
def euclidean_distance(x, y):
return K.sqrt(K.sum(K.square(x - y), axis=1, keepdims=True))
def eucl_dist_output_shape(shapes):
shape1, shape2 = shapes
return (shape1[0], 1)
def conc(vects):
x, y = vects
conc1 = concatenate([x,y])
return conc1
def conc_shape(shapes):
shape1, shape2 = shapes
return (shape1[0],64)
def contrastive_loss(y_true, y_pred):
'''Contrastive loss from Hadsell-et-al.'06
http://yann.lecun.com/exdb/publis/pdf/hadsell-chopra-lecun-06.pdf
'''
x = y_pred[:,0:128]
y = y_pred[:,128:268]
y_pred1 = euclidean_distance(x,y)
p = x
q = y
p = K.clip(p, K.epsilon(), 1)
q = K.clip(q, K.epsilon(), 1)
#y_true1 = y_true[:,0]
#y_true1 = K.reshape(y_true1,(-1,))
#print(y_true1)
#tr_same = y_true[:,1]
#tr_same = K.reshape(tr_same, (-1,))
y_true1 = y_true
tr_same = K.round(y_true/3)
margin = 1
test = 0.001*K.sum(p*K.abs(K.log(p)-K.log(q)), axis=1)
return K.mean((1-tr_same)*(y_true1 * K.square(y_pred1) + (1 - y_true1) * K.square(K.maximum(margin - y_pred1, 0)))
+ (tr_same)*test)
def triplet_loss(y_true, y_pred):
x = y_pred[:,0:32]
y = y_pred[:,32:64]
y_pred1 = euclidean_distance(x,y)
y_true1 = y_true
margin = 1
return K.mean(y_true1 * K.square(y_pred1) + (1 - y_true1) * K.square(K.maximum(margin - y_pred1, 0)))
def coral_loss(y_true, y_pred):
x = y_pred[:,0:32]
y = y_pred[:,32:64]
n = 32.0
mul1 = K.dot(K.transpose(x),x)
one = x*0+1
mul2 = K.dot(K.transpose(one), x)
sub = K.dot(K.transpose(mul2), mul2)
source = (mul1 - (sub)/n)/(n-1)
source = K.abs(source)
source = K.clip(source, K.epsilon(),10000)
source1 = K.log(source)
mul11 = K.dot(K.transpose(y),y)
mul21 = K.dot(K.transpose(one), y)
sub1 = K.dot(K.transpose(mul2), mul2)
n = float(n)
target = (mul11 - (sub1)/n)/(n-1)
target = K.abs(target)
target = K.clip(target, K.epsilon(),10000)
target1 = K.log(target)
return (K.sum(K.dot((source1-target1),(source1-target1)))/(4*32*32.0))
def create_pairs(x, digit_indices):
'''Positive and negative pair creation.
Alternates between positive and negative pairs.
'''
pairs = []
labels = []
n = min([len(digit_indices[d]) for d in range(10)]) - 1
for d in range(10):
for i in range(n):
z1, z2 = digit_indices[d][i], digit_indices[d][i + 1]
pairs += [[x[z1], x[z2]]]
inc = random.randrange(1, 10)
dn = (d + inc) % 10
z1, z2 = digit_indices[d][i], digit_indices[dn][i]
pairs += [[x[z1], x[z2]]]
labels += [1, 0]
return np.array(pairs), np.array(labels)
def create_addi_pairs(x, y):
pairs = []
labels = []
for i in range(0,10):
k1 = k1 = random.randrange(0,x.shape[0])
for j in range(0,5):
k2 = random.randrange(0, y.shape[0])
pairs+= [[x[k1],y[k2]]]
labels += [3]
return np.array(pairs), np.array(labels)
def create_base_network():
'''Base network to be shared (eq. to feature extraction).
'''
seq = Sequential()
seq.add(Dense(16, input_shape=(10,), activation='relu'))
seq.add(Dense(32, activation='relu'))
seq.add(Dense(32, activation='relu'))
return seq
def compute_accuracy(predictions, labels):
'''Compute classification accuracy with a fixed threshold on distances.
'''
return labels[predictions.ravel() < 0.5].mean()
# -
import scipy.io
mat = scipy.io.loadmat('/home/aniruddha/Documents/data_10feature.mat')
arr = mat['TR1_10feature']
arr = np.array(arr)
arr = arr.reshape(-1)
print(arr.shape)
X_train = []
for i in range(0,14):
for j in range(0,arr[i].shape[0]):
X_train.append(arr[i][j])
X_train = np.array(X_train)
print(X_train.shape)
y_train = []
for i in range(0,arr.shape[0]):
for j in range(0,arr[i].shape[0]):
y_train.append(i)
y_train = np.array(y_train)
print(y_train.shape)
print(y_train[1])
arr1 = mat['TS1_10feature']
arr1 = np.array(arr1)
arr1 = arr1.reshape(-1)
print(arr1.shape)
X_test = []
for i in range(0,14):
for j in range(0,arr1[i].shape[0]):
X_test.append(arr1[i][j])
X_test = np.array(X_test)
print(X_test.shape)
y_test = []
for i in range(0,arr1.shape[0]):
for j in range(0,arr1[i].shape[0]):
y_test.append(i)
y_test = np.array(y_test)
print(y_test.shape)
print(y_test[1])
arr2 = mat['TS2_10feature']
arr2 = np.array(arr2)
arr2 = arr2.reshape(-1)
print(arr2.shape)
X_test1 = []
for i in range(0,14):
for j in range(0,arr2[i].shape[0]):
X_test1.append(arr2[i][j])
X_test1 = np.array(X_test1)
print(X_test1.shape)
y_test1 = []
for i in range(0,arr2.shape[0]):
for j in range(0,arr2[i].shape[0]):
y_test1.append(i)
y_test1 = np.array(y_test1)
print(y_test1.shape)
print(y_test1[1])
print(X_train.max())
print(X_test.max())
print(X_test1.max())
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_test1 = X_test1.astype('float32')
X_train = X_train/10000
X_test = X_test/10000
X_test1 = X_test1/10000
print(X_train.max())
print(X_test.max())
print(X_test1.max())
# +
# create training+test positive and negative pairs
digit_indices = [np.where(y_train == i)[0] for i in range(10)]
tr_pairs, tr_y = create_pairs(X_train, digit_indices)
digit_indices = [np.where(y_test == i)[0] for i in range(10)]
te_pairs, te_y = create_pairs(X_test, digit_indices)
# -
tr1_pairs, tr1_y = create_addi_pairs(X_train, X_test1)
print(tr_pairs.shape)
print(te_pairs.shape)
print(tr1_pairs.shape)
# +
# network definition
input_dim=X_train.shape[1:]
base_network = create_base_network()
input_a = Input(shape=input_dim)
input_b = Input(shape=input_dim)
#input_a=K.reshape(input_a,(28,28,1))
#input_b=K.reshape(input_b,(28,28,1))
# because we re-use the same instance `base_network`,
# the weights of the network
# will be shared across the two branches
print(input_b.shape)
# +
processed_a = base_network(input_a)
processed_b = base_network(input_b)
print(processed_a.shape)
distance = Lambda(conc, output_shape=conc_shape)([processed_a, processed_b])
print(distance.shape)
model = Model(input=[input_a, input_b], output=distance)
# -
test_model = Model(input = input_a, output = processed_a)
# +
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
SVG(model_to_dot(model).create(prog='dot', format='svg'))
# +
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
SVG(model_to_dot(base_network).create(prog='dot', format='svg'))
# -
# train
rms = RMSprop()
for i in range(0,50):
model.compile(loss=triplet_loss, optimizer=rms)
model.fit([tr_pairs[:,0], tr_pairs[:, 1]], tr_y, validation_data=([te_pairs[:, 0], te_pairs[:, 1]], te_y), batch_size=32, nb_epoch=1)
model.compile(loss=coral_loss, optimizer=rms)
model.fit([tr1_pairs[:,0], tr1_pairs[:, 1]], tr1_y, batch_size=32, nb_epoch=1)
# +
# %matplotlib inline
from time import time
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import offsetbox
from sklearn import manifold, datasets, decomposition, ensemble, discriminant_analysis, random_projection
# +
def plot_embedding(mu, Y, title=None):
num_class = 50 # data points per class
# x_min, x_max = np.min(mu, 0), np.max(mu, 0)
# mu = (mu - x_min) / (x_max - x_min)
# classes = [0, 1, 2, 3, 4, 6, 7, 8, 9, 10, 11, 12, 15, 16, 18, 19,
# 20, 21, 22, 25, 26, 27, 28, 29, 30, 31, 32, 34, 35, 36, 37, 39, 40, 42, 43, 44, 45, 46, 48, 49]
classes = [0,1,2,3,4,5,6,7,8,9,10,11,12,13]
data = [[] for i in classes]
for i, y in enumerate(Y):
data[classes.index(y)].append(np.array(mu[i]))
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'olive', 'orange', 'mediumpurple','pink','grey','mediumgrey','brown']
l = [i for i in range(14)]
alphas = 0.3 * np.ones(14)
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_aspect(1)
font_size = 13
for i in range(13):
temp = np.array(data[i])
l[i] = plt.scatter(temp[:num_class, 0], temp[:num_class, 1], s = 5, c = colors[i], edgecolors = 'face', alpha=alphas[i])
leg = plt.legend((l[0],l[1],l[2],l[3],l[4],l[5],l[6],l[7],l[8],l[9],l[10],l[11],l[12],l[13]),
('0','1','2','3','4','5','6','7','8','9','10','11','12','13'), loc='center left', bbox_to_anchor=(1, 0.5), fontsize=font_size)
leg.get_frame().set_linewidth(0.0)
plt.xticks(fontsize=font_size)
plt.yticks(fontsize=font_size)
# -
from keras.utils import np_utils
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
y_test1 = np_utils.to_categorical(y_test1)
num_classes = 14
print(y_train.shape)
print(y_test.shape)
print(y_test1.shape)
num_pixels = 32
# define baseline model
def baseline_model1():
# create model
model = Sequential()
model.add(Dense(32, input_dim=num_pixels, kernel_initializer='normal', activation='relu'))
model.add(Dense(16, kernel_initializer='normal', activation='relu'))
model.add(Dense(num_classes, kernel_initializer='normal', activation='softmax'))
# Compile model
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
processed_train = test_model.predict(X_train)
processed_test = test_model.predict(X_test)
processed_test1 = test_model.predict(X_test1)
print(processed_train.shape)
# +
# build the model
model1 = baseline_model1()
# Fit the model
model1.fit(processed_train, y_train, validation_data=(processed_test, y_test), epochs=5000, batch_size=128, verbose=1)
# Final evaluation of the model
scores_train = model1.evaluate(processed_train, y_train, verbose=1)
scores_test = model1.evaluate(processed_test, y_test, verbose=1)
# +
# 5000 epochs
# 32 output dim
# 50 epoch of model1
# 0.01*logcoral+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 32 output dim
# 50 epoch of model1
# coral + cpntrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 32 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 32 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
# 100 samples
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 5 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 10 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 16 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 16 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
# 100 samples
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 32 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
# +
# 5000 epochs
# 64 output dim
# 50 epoch of model1
# logcoral+mean+contrastive
print('* Accuracy on training set: %0.2f%%' % (100 * scores_train[1]))
print('* Accuracy on test set: %0.2f%%' % (100 * scores_test[1]))
scores_test_target=model1.evaluate(processed_test1, y_test1, verbose=1)
print('* Accuracy on test target set: %0.2f%%' % (100 * scores_test_target[1]))
| (LogCoral)Siamese_Train-TR1+TR2(100images)_Test-TS2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wW4AtO3MNYH3" colab_type="text"
# # OMX Validator
#
# The OMX validator is a [Jupyter notebook](https://jupyter.org/) hosted in Google Colaboratory. It is an interactive Python environment that validates OMX matrices using the [openmatrix](https://github.com/osPlanning/omx-python) library. The validator has been tested with the [example](https://github.com/osPlanning/omx/blob/master/example.omx?raw=true) omx file. OMX files can also be inspected with the [OMX Viewer](https://github.com/osPlanning/omx/wiki/OMX-Viewer).
#
# + [markdown] id="Ojb_3G0iJdxO" colab_type="text"
# # Upload Files
#
# Select local files for upload via the Table of Contents + Files + Upload. Run the code cell below to list the OMX files uploaded by clicking on the [ ] play button. While testing, you may need to reset the UI control, which you can do via Runtime + Restart runtime. The Files control can be accessed by clicking the File button.
#
# 
#
#
#
# + id="Jx_VaZZdGYon" colab_type="code" outputId="d6f3696f-7b5f-4631-b1b7-b50b2e4ab4a8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !ls -d *omx
# + [markdown] id="IFGg33IoELVR" colab_type="text"
# # Validator Functions
#
# This step installs the openmatrix package from [pypi.org](https://pypi.org/project/OpenMatrix/), which includes the OMX validation functions and command line tool. Run the code cell to install the package.
# + id="7rvtFHB8EVPz" colab_type="code" outputId="606854d5-4750-4de7-aa33-da7a7ece1528" colab={"base_uri": "https://localhost:8080/", "height": 173}
# !pip install openmatrix
# + [markdown] id="2xBorP6FJi_j" colab_type="text"
# # Validate
#
# This section validates the OMX file against the [Specification](https://github.com/osPlanning/omx/wiki/Specification) by using the omx-validate command line tool. Specify the file to validate in the command line call below and then run the code cell.
#
# The following checks are run and an overall Pass or Fail is returned at the end.
# 1. Has OMX_VERSION attribute set to 0.2
# 2. Has SHAPE array attribute set to two item integer array
# 1. Has data group for matrices
# 1. Matrix shape matches file shape
# 1. Uses common data types (float or int) for matrices
# 1. Matrices chunked for faster I/O
# 2. Uses zlib compression and level 1 if compression used
# 2. Has NA attribute if desired (but not required)
# 2. Has lookup group for labels/indexes if desired (but not required)
# 2. Lookup length matches shape
# 1. Uses common data types (int or str) for lookups
# 2. Has Lookup DIM attribute of 0 (row) or 1 (column) if desired (but not required)
# + id="a8VncE6qKKLC" colab_type="code" outputId="305b785c-679c-4b0a-addc-f4e54c6b107c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !omx-validate example.omx
| omx_validate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp-2020] *
# language: python
# name: conda-env-nlp-2020-py
# ---
# # TP PLAGIO
# ## Librerias:
import docx2txt
import pdftotext
import filetype
import os
import nltk
from nltk import word_tokenize
from nltk import sent_tokenize
from nltk import re
from nltk.corpus import stopwords
import nltk.stem
from sklearn.feature_extraction.text import CountVectorizer
import matplotlib.pyplot as plt; plt.rcdefaults()
import numpy as np
import matplotlib.pyplot as plt
# ## Funciones
# +
def pdf_a_txt(pdf):
with open(pdf, "rb") as f:
pdf_convertido = pdftotext.PDF(f)
txt = ("\n\n".join(pdf_convertido))
return txt
def doc_a_txt(docx):
return (docx2txt.process(docx))
def conversion_de_archivo(ruta_de_archivo):
kind = filetype.guess(ruta_de_archivo)
tipo = kind.extension
if(tipo == "pdf"):
return pdf_a_txt(ruta_de_archivo)
else:
return doc_a_txt(ruta_de_archivo)
def remover_stopwords(texto):
return [word for word in texto if word not in stopwords.words('spanish')]
def procesar_archivos(ruta_arch,ruta_arch2):
print("El archivo con el que se lo compara es:")
print(os.path.basename(ruta_arch2)+'\n')
archivo = conversion_de_archivo(ruta_arch)
archivo2 = conversion_de_archivo(ruta_arch2)
archivo_tok = word_tokenize(archivo)
archivo2_tok = word_tokenize(archivo2)
archivo_sent=sent_tokenize(archivo)
archivo2_sent=sent_tokenize(archivo2)
archivo_tokenizado = [word.lower() for word in archivo_tok if re.search("\w",word)]
archivo2_tokenizado = [word.lower() for word in archivo2_tok if re.search("\w",word)]
archivo_tok_sin_stops = remover_stopwords(archivo_tokenizado)
archivo2_tok_sin_stops = remover_stopwords(archivo2_tokenizado)
#print(taggear(archivo_tok_sin_stops))
list_de_score = comparacion_con_trigramas(archivo_tok_sin_stops,archivo2_tok_sin_stops)
print("Comparacion por lcs:")
score_lcs = comparacion_por_longest_common_subsequence(archivo,archivo2)
print('\n'+'\n')
graficar(list_de_score[0],list_de_score[1],score_lcs)
if ruta_arch not in lista_de_archivos:
guardar_archivo(ruta_arch)
def lemattizar(archivo):
stemmer = nltk.stem.SnowballStemmer('spanish')
stem_vectorizer = StemmedCountVectorizer(min_df=1, stop_words='spanish')
stem_analyze = stem_vectorizer.build_analyzer()
Y = stem_analyze(archivo)
for tok in Y:
print(tok)
class StemmedCountVectorizer(CountVectorizer):
def build_analyzer(self):
analyzer = super(StemmedCountVectorizer, self).build_analyzer()
return lambda doc: (english_stemmer.stem(w) for w in analyzer(doc))
def taggear(archivo_tok):
archivo_taggeado = nltk.pos_tag(archivo_tok)
return archivo_taggeado
def comparacion_con_trigramas(arch1,arch2):
s = 0
trigrams_arch1 = []
for i in range(len(arch1)-2):
t=(arch1[i],arch1[i+1],arch1[i+2])
trigrams_arch1.append(t)
trigrams_arch2 = []
for i in range(len(arch2)-2):
t=(arch2[i],arch2[i+1],arch2[i+2])
trigrams_arch2.append(t)
if t in trigrams_arch1:
s+=1
#print(s)
print("Comparacion por jaccard:")
score_jaccard = comparacion_por_jaccard(s,trigrams_arch1,trigrams_arch2)
print("Comparacion por containment measure:")
score_containment = comparacion_por_containment_measure(s,trigrams_arch2)
return [score_containment,score_jaccard]
def comparacion_por_jaccard(s,trigrams_arch1,trigrams_arch2):
jaccard_coefficient = s/(len(trigrams_arch1)+len(trigrams_arch2))
print(jaccard_coefficient)
medidor_de_comparacion(jaccard_coefficient)
print('\n')
return(jaccard_coefficient)
def comparacion_por_containment_measure(s,trigrams_arch2):
containment = s/len(trigrams_arch2)
print(containment)
medidor_de_comparacion(containment)
print('\n')
return(containment)
def lcs(l1,l2):
s1=word_tokenize(l1)
s2=word_tokenize(l2)
# storing the dp values
dp = [[None]*(len(s1)+1) for i in range(len(s2)+1)]
for i in range(len(s2)+1):
for j in range(len(s1)+1):
if i == 0 or j == 0:
dp[i][j] = 0
elif s2[i-1] == s1[j-1]:
dp[i][j] = dp[i-1][j-1]+1
else:
dp[i][j] = max(dp[i-1][j] , dp[i][j-1])
return dp[len(s2)][len(s1)]
def comparacion_por_longest_common_subsequence(arch1,arch2):
sent_o = sent_tokenize(arch1)
sent_p = sent_tokenize(arch2)
tokens_p = word_tokenize(arch2)
#maximum length of LCS for a sentence in suspicious text
max_lcs=0
sum_lcs=0
for i in sent_p:
for j in sent_o:
l=lcs(i,j)
max_lcs=max(max_lcs,l)
sum_lcs+=max_lcs
max_lcs=0
score=sum_lcs/len(tokens_p)
print(score)
medidor_de_comparacion(score)
print('\n')
return(score)
def guardar_archivo(ruta_archivo):
lista_de_archivos.append(ruta_archivo)
def procesar_archivo(ruta_archivo):
print("El nombre del archivo a procesar es:")
print(os.path.basename(ruta_archivo)+'\n')
for a in lista_de_archivos:
if(a != ruta_archivo):
procesar_archivos(ruta_archivo,a)
def medidor_de_comparacion(score):
if score <= 0.2:
print("no existe plagio")
elif 0.2 <= score and score <= 0.4:
print("Poca probabilidad de plagio")
elif 0.4 <= score and score <= 0.6:
print("Ligera probabilidad de plagio")
elif score >= 0.6:
print("Alta probabilidad de plagio")
def graficar(score_containment,score_jaccard,score_lcs):
objects = ('score_containment','score jaccard','score lcs')
y_pos = np.arange(len(objects))
performance = [score_containment,score_jaccard,score_lcs]
plt.bar(y_pos, performance, align='center', alpha=0.5)
plt.xticks(y_pos, objects)
plt.ylabel('Score')
plt.title('Metodos de medición')
plt.show()
# +
global lista_de_archivos
lista_de_archivos = []
ruta_archivo = '/Users/Usuario/Desktop/dataset/Marketing - TP 0.docx'
ruta_archivo2 = '/Users/Usuario/Desktop/dataset/Economia de experiencia.pdf'
ruta_archivo3 = '/Users/Usuario/Desktop/dataset/Economia de experiencia (1).pdf'
ruta_archivo4 = '/Users/Usuario/Desktop/dataset/K5071 - <NAME> - TP N°5 Rifkin (1).pdf'
ruta_archivo5 = '/Users/Usuario/Desktop/dataset/Marketing - TP 1.docx'
ruta_archivo6 = '/Users/Usuario/Desktop/dataset/Marketing - TP 2.docx'
guardar_archivo(ruta_archivo)
guardar_archivo(ruta_archivo2)
guardar_archivo(ruta_archivo3)
guardar_archivo(ruta_archivo4)
guardar_archivo(ruta_archivo5)
guardar_archivo(ruta_archivo6)
#procesar_archivos(ruta_archivo2,ruta_archivo3)
procesar_archivo(ruta_archivo2)
# -
| tp-plagio-nlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7EcHxfZ6rB9x"
# # Predição de chuva da Australia
# Link: https://www.kaggle.com/jsphyg/weather-dataset-rattle-package
#
# + [markdown] id="RT5CJdmErXtg"
# ## Dependências
# + id="5fE2c5Qh3myT" outputId="8d3ba9e3-268f-4213-94e4-f01a67c67d1e" colab={"base_uri": "https://localhost:8080/"}
# !pip install category_encoders
# + id="OK-edKhwrWCo" outputId="e6e90a0a-ad9d-41be-c5cb-1c90e5c258f1" colab={"base_uri": "https://localhost:8080/"}
import pandas as pd
import numpy as np
import missingno as msno
from sklearn.model_selection import train_test_split
import category_encoders as ce
from tensorflow.keras import layers
from tensorflow.keras import activations
import tensorflow as tf
print(tf.__version__)
# + id="iqaFr6L-qnmv"
data = pd.read_csv('weatherAUS.csv')
# + id="zqx_wMMxAy11"
# + [markdown] id="ypkeRH0Zspa8"
# ## Análise inicial dos dados
# + id="jiHWHRuosr7H" outputId="dab3fea5-7c15-4f24-8887-2590335458bc" colab={"base_uri": "https://localhost:8080/", "height": 245}
print('shape', data.shape)
data.head()
# + id="W8NmroBVsyz-" outputId="cc872479-137f-4a9d-b299-58abe5c0775b" colab={"base_uri": "https://localhost:8080/", "height": 712}
msno.matrix(data)
# + id="_p9oCOWFwPik" outputId="44cf658c-65d7-44a2-a2e5-6a9b5e68ab4d" colab={"base_uri": "https://localhost:8080/"}
data.info()
# + id="V970ZxLHwRk_" outputId="daf17070-ec99-4b4a-b2bd-9297a5c69ce5" colab={"base_uri": "https://localhost:8080/", "height": 320}
data.describe()
# + [markdown] id="fYLINEY2uWCr"
# ## Tratando valores nulos
# + id="JSecQi9GuzgM"
#@title drop da coluna RISK_MM, como fala na descrição do dataset
data.drop(['RISK_MM'], axis=1, inplace=True)
# + id="r-AHmuIIxR6P"
#@title median imputation nos dados numéricos faltantes
# colunas com dados numéricos
numerical = [col for col in data.columns if data[col].dtypes != 'O']
# imputação com mediana
for col in numerical:
col_median=data[col].median()
data[col].fillna(col_median, inplace=True)
# + id="6wYvRnqQxcpV"
#@title most frequent imputation nos dados categóricos faltantes
# colunas com dados categóricos
categorical = [col for col in data.columns if data[col].dtypes == 'O']
# most frequent imputation
for col in categorical:
data[col].fillna(data[col].mode()[0], inplace=True)
# + id="Cx2Opzkeybqv" outputId="ac4b5db7-a9a9-4b99-c9f4-b32caac4e236" colab={"base_uri": "https://localhost:8080/", "height": 712}
#@title resultado
msno.matrix(data)
# + [markdown] id="rZSW4k-n08_C"
# ## Tratando outliers
# + id="2L9WwAKp0_Oz"
def max_value(df3, variable, top):
return np.where(df3[variable]>top, top, df3[variable])
data['Rainfall'] = max_value(data, 'Rainfall', 3.2)
data['Evaporation'] = max_value(data, 'Evaporation', 21.8)
data['WindSpeed9am'] = max_value(data, 'WindSpeed9am', 55)
data['WindSpeed3pm'] = max_value(data, 'WindSpeed3pm', 57)
# + [markdown] id="UOL4ZDhN1yF4"
# ## Encoding variáveis categóricas
# + id="oP7VgaAm13AW" outputId="86adeb38-0ecc-4184-a1be-41cedb0595b1" colab={"base_uri": "https://localhost:8080/"}
# encode RainToday variable
encoder = ce.BinaryEncoder(cols=['RainToday'])
data = encoder.fit_transform(data)
# + id="IajZpHYo3_I6"
data = pd.concat([data[numerical], data[['RainToday_0', 'RainToday_1']],
pd.get_dummies(data.Location),
pd.get_dummies(data.WindGustDir),
pd.get_dummies(data.WindDir9am),
pd.get_dummies(data.WindDir3pm),
data['RainTomorrow'].map({'No':0, 'Yes':1}).astype(int)], axis=1)
# + id="Yn-sxItR4H0C" outputId="d0b362bc-72d1-425b-9401-c6dc8a646440" colab={"base_uri": "https://localhost:8080/", "height": 444}
data
# + [markdown] id="-1D1WILP1gzR"
# ## Treinamento
# + id="X0H6QicDycsC"
# 75/25 train test split
data_x = data.drop(['RainTomorrow'], axis=1)
data_y = data[['RainTomorrow']]
X_train, X_test, y_train, y_test = train_test_split(data_x, data_y, test_size=0.25, random_state=42)
# + id="rppI0o34zW0L" outputId="abda2b38-ff5e-4043-92ea-86363f105c5e" colab={"base_uri": "https://localhost:8080/"}
# Instantiate a simple classification model
model = tf.keras.Sequential([
layers.Dense(256, activation=tf.nn.relu),
layers.Dense(256, activation=tf.nn.relu),
layers.Dense(1, activation=tf.nn.sigmoid)
])
# Instantiate a logistic loss function that expects integer targets.
loss = tf.keras.losses.BinaryCrossentropy()
# Instantiate an accuracy metric.
accuracy = tf.keras.metrics.BinaryAccuracy(
name="binary_accuracy", dtype=None, threshold=0.5
)
# Instantiate an optimizer.
optimizer = tf.keras.optimizers.Adam()
model.compile(optimizer=optimizer, loss=loss, metrics=[accuracy])
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=30, batch_size=64)
# + id="puiAX93F11Fm" outputId="30d93967-5d79-4af6-c22a-08919becf1af" colab={"base_uri": "https://localhost:8080/"}
model.predict(X_test)
# + [markdown] id="EVbkO-9kA8ow"
# ## Melhorias
# + id="tLKwJ54b9MQE"
# train, test, cv with 0.6, 0.2, 0.2 ratios
x, x_test, y, y_test = train_test_split(data_x, data_y, test_size=0.2, train_size=0.8)
x_train, x_cv, y_train, y_cv = train_test_split(x,y,test_size = 0.25, train_size =0.75)
# + id="buM0xKCYadLn" outputId="22633a94-e262-4cc6-eced-1924b074df61" colab={"base_uri": "https://localhost:8080/"}
model.fit(x_train, y_train, validation_data=(x_cv, y_cv), epochs=30, batch_size=64)
# + id="p9Ny1NNXwVEh"
y_tested = y_test.to_numpy()
classes = y_pred > 0.5
# + id="TPacSYRIfYej" outputId="6ddccc25-67c2-48ee-dbd7-2ba412b7e0ba" colab={"base_uri": "https://localhost:8080/"}
y_pred = model.predict(x_test)
m = tf.keras.metrics.BinaryAccuracy(name='binary_accuracy', dtype=None, threshold=0.5)
m.update_state(classes,y_tested)
m.result().numpy()
# + [markdown] id="H0zCLL_6yEW0"
# ## Regularização com L2
#
# + id="Jbh3Ny1nw-lI"
# Instantiate a simple classification model
model = tf.keras.Sequential([
layers.Dense(256, activation=tf.nn.relu),
layers.Dense(256, activation=tf.nn.relu,kernel_regularizer=tf.keras.regularizers.l2(l=0.01)),# Regulazer with L2
layers.Dense(1, activation=tf.nn.sigmoid)
])
# Instantiate a logistic loss function that expects integer targets.
loss = tf.keras.losses.BinaryCrossentropy()
# Instantiate an accuracy metric.
accuracy = tf.keras.metrics.BinaryAccuracy(
name="binary_accuracy", dtype=None, threshold=0.5
)
# Instantiate an optimizer.
optimizer = tf.keras.optimizers.Adam()
model.compile(optimizer=optimizer, loss=loss, metrics=[accuracy])
# + id="OuY9oK5XzgYz" outputId="73aaf40c-0f8c-4007-ce74-c3da9ffa24e9" colab={"base_uri": "https://localhost:8080/"}
model.fit(x_train, y_train, validation_data=(x_cv, y_cv), epochs=30, batch_size=64)
# + id="b-OhYC9nzo0B" outputId="83947690-ad3e-40b9-e75c-67595924a320" colab={"base_uri": "https://localhost:8080/"}
y_tested = y_test.to_numpy()
classes = y_pred > 0.5
y_pred = model.predict(x_test)
m = tf.keras.metrics.BinaryAccuracy(name='binary_accuracy', dtype=None, threshold=0.5)
m.update_state(classes,y_tested)
m.result().numpy()
| Projeto_01_Deep_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="2bc8967d-d32d-355c-021d-9e47f25be04b"
# <h1> Introduction </h1>
#
# <p> The intention of this notebook is to utilize tensorflow to build a neural network that helps to predict default likelihood, and to visualize some of the insights generated from the study. This kernel will evolve over time as I continue to add features and study the Lending Club data </p>
# + [markdown] _cell_guid="48701902-6fc5-a857-4cc3-426c6f6d6563"
# <h3> Dependencies </h3>
#
# <p> Below the data and some external libraries are imported to begin the process </p>
# + _cell_guid="a7785cb6-61fd-0a89-7d67-9de44fb26ef2"
# #%matplotlib inline
import numpy as np
import pandas as pd
import itertools
from sklearn import preprocessing
import matplotlib.pyplot as plt
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
from tensorflow.contrib.learn.python.learn import metric_spec
from tensorflow.contrib.learn.python.learn.estimators import _sklearn
from tensorflow.contrib.learn.python.learn.estimators import estimator
from tensorflow.contrib.learn.python.learn.estimators import model_fn
from tensorflow.python.framework import ops
from tensorflow.python.saved_model import loader
from tensorflow.python.saved_model import tag_constants
from tensorflow.python.util import compat
tf.logging.set_verbosity(tf.logging.FATAL)
df = pd.read_csv("../input/loan.csv", low_memory=False)
# + [markdown] _cell_guid="7237e9f9-d392-2c0a-b555-0d58b1a50ceb"
# <h3> Creating the Target Label </h3>
#
# <p> From a prior notebook, I examined the 'loan_status' column. The cell below creates a column with binary value 0 for loans not in default, and binary value 1 for loans in default.
# + _cell_guid="ccf2aab3-dcf1-7c6d-0654-3e9a280147cb"
df['Default_Binary'] = int(0)
for index, value in df.loan_status.iteritems():
if value == 'Default':
df.set_value(index,'Default_Binary',int(1))
if value == 'Charged Off':
df.set_value(index, 'Default_Binary',int(1))
if value == 'Late (31-120 days)':
df.set_value(index, 'Default_Binary',int(1))
if value == 'Late (16-30 days)':
df.set_value(index, 'Default_Binary',int(1))
if value == 'Does not meet the credit policy. Status:Charged Off':
df.set_value(index, 'Default_Binary',int(1))
# + [markdown] _cell_guid="66a153fd-76b9-02d7-326e-05ecaa02efeb"
# <h3> Creating a category feature for "Loan Purpose" </h3>
#
# <p> Below I create a new column for loan purpose, and assign each type of loan purpose an integer value. </p>
# + _cell_guid="92d1330a-4919-1ad1-95c7-3380fc7e13d6"
df['Purpose_Cat'] = int(0)
for index, value in df.purpose.iteritems():
if value == 'debt_consolidation':
df.set_value(index,'Purpose_Cat',int(1))
if value == 'credit_card':
df.set_value(index, 'Purpose_Cat',int(2))
if value == 'home_improvement':
df.set_value(index, 'Purpose_Cat',int(3))
if value == 'other':
df.set_value(index, 'Purpose_Cat',int(4))
if value == 'major_purchase':
df.set_value(index,'Purpose_Cat',int(5))
if value == 'small_business':
df.set_value(index, 'Purpose_Cat',int(6))
if value == 'car':
df.set_value(index, 'Purpose_Cat',int(7))
if value == 'medical':
df.set_value(index, 'Purpose_Cat',int(8))
if value == 'moving':
df.set_value(index, 'Purpose_Cat',int(9))
if value == 'vacation':
df.set_value(index,'Purpose_Cat',int(10))
if value == 'house':
df.set_value(index, 'Purpose_Cat',int(11))
if value == 'wedding':
df.set_value(index, 'Purpose_Cat',int(12))
if value == 'renewable_energy':
df.set_value(index, 'Purpose_Cat',int(13))
if value == 'educational':
df.set_value(index, 'Purpose_Cat',int(14))
# + [markdown] _cell_guid="76da5ae6-6ba6-ac54-7552-1729cce271fd"
# <h3> Scaling Interest Rates </h3>
#
# <p> Below I scale the interest rate for each loan to a value between 0 and 1 </p>
# + _cell_guid="d3976cc7-71a4-1f53-8d6f-b05359c2d52b"
x = np.array(df.int_rate.values).reshape(-1,1)
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df['int_rate_scaled'] = pd.DataFrame(x_scaled)
print (df.int_rate_scaled[0:5])
# + [markdown] _cell_guid="0c0f2a88-1caa-14f3-35b8-2a2ea6e73c04"
# <h3> Scaling Loan Amount </h3>
#
# <p> Below I scale the amount funded for each loan to a value between 0 and 1 </p>
# + _cell_guid="ed034055-7d37-7931-771f-6da895809b1a"
x = np.array(df.funded_amnt.values).reshape(-1,1)
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df['funded_amnt_scaled'] = pd.DataFrame(x_scaled)
print (df.funded_amnt_scaled[0:5])
# + [markdown] _cell_guid="c9ff9618-349b-014e-4b74-73a90f67d357"
# <h3> Setting up the Neural Network </h3>
#
# <p> Below I split the data into a training, testing, and prediction set </p>
# <p> After that, I assign the feature and target columns, and create the function that will be used to pass the data into the model </p>
# + _cell_guid="d8e3775c-2f81-8b3a-35b1-13c998555c87"
training_set = df[0:500000] # Train on first 500k rows
testing_set = df[500001:849999] # Test on next 350k rows
prediction_set = df[850000:] # Predict on final 37k rows
COLUMNS = ['Purpose_Cat','funded_amnt_scaled','int_rate_scaled','Default_Binary']
FEATURES = ['Purpose_Cat','funded_amnt_scaled','int_rate_scaled']
LABEL = 'Default_Binary'
def input_fn(data_set):
feature_cols = {k: tf.constant(data_set[k].values) for k in FEATURES}
labels = tf.constant(data_set[LABEL].values)
return feature_cols, labels
# + [markdown] _cell_guid="64d960f5-71af-a957-64ed-8e8e0bda34cc"
# <h3> Fitting The Model </h3>
# + _cell_guid="f3cb8715-98e0-5baf-2a09-cbd8aca9e81c"
feature_cols = [tf.contrib.layers.real_valued_column(k)
for k in FEATURES]
#config = tf.contrib.learn.RunConfig(keep_checkpoint_max=1) ######## DO NOT DELETE
regressor = tf.contrib.learn.DNNRegressor(
feature_columns=feature_cols, hidden_units=[10, 20, 10], )
regressor.fit(input_fn=lambda: input_fn(training_set), steps=251)
# + [markdown] _cell_guid="c6226ab1-fa2b-1a7b-0e52-72da67dfb289"
# <h3> Evaluating the Model </h3>
# + _cell_guid="2f0e6c5f-5b97-279e-086f-6fda35cf0161"
# Score accuracy
ev = regressor.evaluate(input_fn=lambda: input_fn(testing_set), steps=10)
loss_score = ev["loss"]
print("Loss: {0:f}".format(loss_score))
# + [markdown] _cell_guid="d69b7b45-28e4-e108-8e13-5eb4aace72ac"
# <h3> Predicting on new data </h3>
# + _cell_guid="4411bcb7-d7ad-1bdb-6871-a8779fc39530"
y = regressor.predict(input_fn=lambda: input_fn(prediction_set))
predictions = list(itertools.islice(y, 37379))
# + [markdown] _cell_guid="21c01a6a-e787-f7f0-0b1e-f9a461c24553"
# <h3> Visualize Predictions Relative To Interest Rates </h3>
# + _cell_guid="e8f808ed-bd4f-6894-72c3-1dfb7e0df76c"
plt.plot(prediction_set.int_rate_scaled, predictions, 'ro')
plt.ylabel("Model Prediction Value")
plt.xlabel("Interest Rate of Loan (Scaled between 0-1)")
plt.show()
# + [markdown] _cell_guid="49d9523a-641e-562c-85dd-aeee3bbbb8a8"
# <h3> Visualize Predictions Relative to Loan Size </h3>
# + _cell_guid="29581fa9-499c-5377-d62f-686ce6ca8942"
plt.plot(prediction_set.funded_amnt_scaled, predictions, 'ro')
plt.ylabel("Model Prediction Value")
plt.xlabel("Funded Amount of Loan (Scaled between 0-1)")
plt.show()
# + [markdown] _cell_guid="f6b0be60-e5c3-bf1d-ccaf-d14663f21bfb"
# <h3> Visualize Predictions Relative to Loan Purpose </h3>
# + _cell_guid="f42d09f7-5f4a-c8d3-3c9f-0dd2eb4f1797"
plt.plot(prediction_set.Purpose_Cat, predictions, 'ro')
plt.ylabel("Default Prediction Value")
plt.xlabel("Loan Purpose")
plt.title("DNN Regressor Predicting Default By Loan Purpose")
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 8
fig_size[1] = 8
plt.rcParams["figure.figsize"] = fig_size
labels = ['Debt Consolidation', 'Credit Card', 'Home Improvement', 'Other',
'Major Purchase', 'Small Business', 'Car', 'Medical',
'Moving', 'Vacation', 'House', 'Wedding',
'Renewable Energy']
plt.xticks([1,2,3,4,5,6,7,8,9,10,11,12,13,14], labels, rotation='vertical')
plt.show()
# + _cell_guid="c87fba7a-8073-0859-957b-4a4250472e36"
| downloaded_kernels/loan_data/kernel_152.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering Astronomical Sources
#
# The objective of this hands-on activity is to cluster a set of candidate sources from the Zwicky Transient Facility's (ZTF) image subtraction pipeline. All candidate features and postage stamps were extracted from ZTF's public alert stream.
#
# The goal of this exercise is to become familiar with the ZTF data, the examination of some of its features, and running sklearn's [KMeans](http://scikit-learn.org/stable/modules/generated/sklearn.cluster.KMeans.html) algorithm on 2 or more features. Here are the steps we will take:
#
# 1. Load data
# 2. Plot Features 'elong' and 'chipsf'
# 3. Run KMeans on 2 Features
# 4. Feature Scaling
# 4. Evaluation Results Quantitatively
# 5. Evaluate Results by Examining Postage Stamps
# 6. Clustering in a Dimensionally-Reduced Space
#
# ### 0a. Imports
#
# These are all the imports that will be used in this notebook. All should be available in the DSFP conda environment.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import glob
import os
from time import time
from matplotlib.pyplot import imshow
from matplotlib.image import imread
from sklearn.cluster import KMeans
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn import metrics
from sklearn.metrics.pairwise import euclidean_distances
# ### 0b. Data Location
#
# You will need the following files:
# - [dsfp_ztf_meta.npy](https://northwestern.box.com/s/ssnzcfjp4xggu7q2jgrmgrduy4de48w1)
# - [dsfp_ztf_feats.npy](https://northwestern.box.com/s/2b4bdtqkv9v8b9ooginyx0wkmgnyi8nc)
# - [dsfp_ztf_png_stamps.tar.gz](https://northwestern.box.com/s/pcx3hks09qwbi2tn1nokbmy8tkzk4btv)
#
# You will need to unzip and unpack this last file (a "tarball") called `dsfp_ztf_png_stamps.tar.gz`. Run the following commands in the same directory as this notebook to unpack everything (note - some operating systems automatically unzip downloaded files):
#
# - gunzip dsfp_ztf_png_stamps.tar.gz
# - tar -xvf dsfp_ztf_png_stamps.tar
#
# You should now have a directory in your current working directory (cwd) called dsfp_ztf_png_stamps.
#
# Please specify the following file locations:
#
F_META = # complete
F_FEATS = # complete
D_STAMPS = # complete
#
# ## 1. Load Data
#
# We are ready to get started! :) Start by loading the data and confirming that feats has the same number of columns as COL_NAMES. Please note that the last columns is a class label with values {0, 1}, where 0=bogus, and 1=real. Today we are doing unsupervised learning, but some clustering evaluation methods use labels to quantitatively measure the quality of the clustering result.
# +
meta = np.load(F_META)
feats = np.load(F_FEATS)
COL_NAMES = ['diffmaglim', 'magpsf', 'sigmapsf', 'chipsf', 'magap', 'sigmagap',
'distnr', 'magnr', 'sigmagnr', 'chinr', 'sharpnr', 'sky',
'magdiff', 'fwhm', 'classtar', 'mindtoedge', 'magfromlim', 'seeratio',
'aimage', 'bimage', 'aimagerat', 'bimagerat', 'elong', 'nneg',
'nbad', 'ssdistnr', 'ssmagnr', 'sumrat', 'magapbig', 'sigmagapbig',
'ndethist', 'ncovhist', 'jdstarthist', 'jdendhist', 'scorr', 'label']
# INSTRUCTION: Verify that feats has the same number of columns as COL_NAMES
#
# -
# ## 2. Plot Features
#
# We will perform K-means clustering using two features: 'chipsf' and 'elong'. Chipsf is the uncertainty associated with performing PSF-fit photometry. The higher the chi values, the more uncertainty associated with the source's PSF fit. Elong is a measure of how elongated the source is. A transient point source should have a spherical point spread function. An elongated point source may be a sign of a problem with image subtraction.
#
# Extract features chipsf and along from the data. Scatter plot them together, and also plot their histograms.
#
# #### Question: What do you notice about these features?
#
# +
featnames_to_select = ['chipsf', 'elong']
# Extract the Correct Features
#
featidxs_to_select_indices = [ COL_NAMES.index(x) for x in featnames_to_select]
feats_selected = feats[:,featidxs_to_select_indices]
# Scatter Plot the Two Features
#
def plot_scatter(dat, xlabel, ylabel, xscale='linear', yscale='linear'):
plt.plot(dat[:,0], dat[:,1], 'k.')
plt.xlabel(xlabel)
plt.ylabel(ylabel)
plt.xscale(xscale)
plt.yscale(yscale)
plt.show()
# Scatter Plot the Two Features
#
def plot_histogram(dat, bins, title, xscale='linear', yscale='linear'):
plt.hist(dat, bins)
plt.xscale(xscale)
plt.yscale(yscale)
plt.title(title)
plt.show()
# INSTRUCTION: Scatter Plot the Data
#
# INSTRUCTION: Plot the Histograms for both features. Hint, it may be helpful to plot some features on a log scale.
#
# -
# ## 3. KMeans Using Two Features
#
# We rarely ever cluster only two features from a dataset. However, the advantage of doing so is that we can readily visualize two-dimensional data. Let's start off by clustering features elong and chipsf with KMeans. The plotKMeans function below implements a visualization of KMean's partitioning that was used in sklearn's [KMean's demo](http://scikit-learn.org/stable/auto_examples/cluster/plot_kmeans_digits.html).
#
# #### Question: What do you think about the quality of the clusterings produced?
# +
def runKMeans(dat, n_clusters=2, seed=0):
return KMeans(n_clusters, random_state=seed).fit(dat)
def plotKMeans(kmeans_res, reduced_dat, xlabel, ylabel, xscale='linear', yscale='linear'):
# Plot the decision boundary. For that, we will assign a color to each
h = .02 # point in the mesh [x_min, x_max]x[y_min, y_max].
x_min, x_max = reduced_dat[:, 0].min() - 1, reduced_dat[:, 0].max() + 1
y_min, y_max = reduced_dat[:, 1].min() - 1, reduced_dat[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# Obtain labels for each point in mesh. Use last trained model.
Z = kmeans_res.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.imshow(Z, interpolation='nearest',
extent=(xx.min(), xx.max(), yy.min(), yy.max()),
cmap=plt.cm.Paired,
aspect='auto', origin='lower')
plt.plot(reduced_dat[:,0], reduced_dat[:,1], 'k.')
plt.scatter(kmeans_res.cluster_centers_[:, 0], kmeans_res.cluster_centers_[:, 1],
marker='x', s=169, linewidths=3,
color='w', zorder=10)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
plt.xscale(xscale)
plt.yscale(yscale)
plt.show()
# INSTRUCTION: Use the runKMeans and plotKMeans functions to cluster the data (feats_selected)
# with several values of k.
# -
# # 4. Feature Scaling
#
# We just discovered that distance metrics can be sensitive to the scale of your data (e.g., some features span large numeric ranges, but others don't). For machine learning methods that calculate similiarty between feature vectors, it is important to normalize data within a standard range such as (0, 1) or with z-score normalization (scaling to unit mean and variance). Fortunately, sklearn also makes this quite easy. Please review sklearn's [preprocessing](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.preprocessing) module options, specifically StandardScaler which corresponds to z-score normalization and MinMaxScaler. Please implement one.
#
# After your data has been scaled, scatter plot your rescaled features, and run KMeans with the transformed data. Compare the results on the transformed data with those above.
# +
# INSTRUCTION: Re-scale your data using either the MinMaxScaler or StandardScaler from sklearn
#
# INSTRUCTION: Scatter plot your rescaled data
#
# INSTRUCTION: Retry KMeans with the same values of k used above.
#
# -
# # 5. Quantitative Cluster Evaluation
#
# So far, we've been visually verifying our clusters. Let's use quantitative methods to verify our results.
#
# The following is a score that does not require labels:
# - inertia: "Sum of squared distances of samples to their closest cluster center."
# - Silhouette coefficient: Measures minimal inertia in ratio to distance to next nearest cluster. The score is higher are clusters become more compact and well-separated.
#
# The following scores do require labels, and are documented [here](http://scikit-learn.org/stable/modules/clustering.html#clustering-evaluation).
#
# - ARI, AMI measure the similarity between ground_truth labels and predicted_labels. ARI measure similarity, and AMI measures in terms of mutual information. Random assignments score close to 0, correct assignments close to 1.
# - homogeneity: purity of the cluster (did all cluster members have the same label?). Scores in [0,1] where 0 is bad.
# - completeness: did all labels cluster together in a single cluster? Scores in [0,1] where 0 is bad.
#
# +
sample_size = 300
def bench_k_means(estimator, name, data, labels):
t0 = time()
estimator.fit(data)
print('%-9s\t%.2fs\t%i\t%.3f\t%.3f\t%.3f\t%.3f\t%.3f\t%.3f'
% (name, (time() - t0), estimator.inertia_,
metrics.homogeneity_score(labels, estimator.labels_),
metrics.completeness_score(labels, estimator.labels_),
metrics.v_measure_score(labels, estimator.labels_),
metrics.adjusted_rand_score(labels, estimator.labels_),
metrics.adjusted_mutual_info_score(labels, estimator.labels_),
metrics.silhouette_score(data, estimator.labels_,
metric='euclidean',
sample_size=sample_size)))
labels = feats[:,-1]
print(82 * '_')
print('init\t\ttime\tinertia\thomo\tcompl\tv-meas\tARI\tAMI\tsilhouette')
# INSTRUCTIONS: Use the bench_k_means method to compare your clustering results
#
# -
# # 6. Cluster Evaluation by Visual Inspection
#
# ## This time with postage stamps!
#
# It can be tempting to let yourself be guided by metrics alone, and the metrics are useful guideposts that can help determine whether you're moving in the right direction. However, the goal of clustering is to reveal structure in your dataset. Fortunately, because the features were extracted from sources that were extracted from images, we can view the cutouts from each source to visually verify whether our clusters contain homogeneous objects.
#
# The display methods below give you an opportunity to display random candidates from each cluster, or the candidates that are closest to the cluster center.
# +
def display_stamps(candids, fig_title):
# display five across
num_per_row = 5
for i, candid in enumerate(candids):
f_stamp = glob.glob(os.path.join(D_STAMPS, 'candid{}*.png'.format(candid)))[0] # there should only be one file returned!
if (i % num_per_row) == 0:
fig = plt.figure(figsize=(18, 3))
fig.suptitle(fig_title)
ax = fig.add_subplot(1, num_per_row, i%num_per_row + 1)
ax.set_axis_off()
ax.set_title(candid)
stamp = imread(f_stamp)
imshow(stamp)
return
def closest_to_centroid(centroid, cluster_feats, cluster_candids):
dists = euclidean_distances(cluster_feats, centroid.reshape(1, -1))[:,0]
closest_indices = np.argsort(dists)[:10]
return cluster_candids[closest_indices]
def show_cluster_stamps(kmeans_res, displayMode='closest', num_to_display=10):
# spits out a random selection of stamps from each cluster
for i in range(kmeans_res.n_clusters):
centroid = kmeans_res.cluster_centers_[i, :]
mask = kmeans_res.labels_ == i
cluster_candids = meta[mask]['candid']
cluster_feats = feats_selected_scaled[mask]
if displayMode == 'near_centroid':
selected_candids = closest_to_centroid(centroid, cluster_feats, cluster_candids)
if displayMode == 'random':
np.random.shuffle(cluster_candids)
selected_candids = cluster_candids[:num_to_display]
display_stamps(selected_candids, 'Cluster {}'.format(i))
# INSTRUCTION: Use the show_cluster_stamps method to display cutouts associated with each cluster.
# Do you see similar objects in each cluster?
#
# -
# # 7. Clustering in a Dimensionally-Reduced Space
#
# Given the tools seen above, starting clustering more than 2 features at a time. This work is free-form. I'll start you off with some suggested features. After plotting the feature distributions, you may choose to down-select further.
#
# Because we're now working with more than 2 features, use PCA to project the feature space onto its first two principal components. You may use the methods above to run KMeans in that reduced feature space and evaluate your results.
# +
featnames_to_select = ['chipsf', 'elong', 'diffmaglim', 'magpsf', 'sigmapsf',
'chipsf', 'magap', 'sigmagap', 'sky', 'magdiff', 'fwhm',
'mindtoedge', 'magfromlim', 'seeratio', 'aimage', 'bimage',
'aimagerat', 'bimagerat', 'elong', 'nneg', 'nbad', 'sumrat', 'magapbig', 'sigmagapbig']
# INSTRUCTION: Visualize these features. Discard any you consider to be problematic.
# INSTRUCTION: Filter the feature space
# INSTRUCTION: Run PCA on this feature space to reduce it to 2 principal components
# INSTRUCTION: Run KMeans on this 2-dimensional PCA space, and evaluate your results both quantatively and qualitatively.
| Sessions/Session07/Day2/Clustering-Astronomical-Sources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 32-bit
# language: python
# name: python3
# ---
# +
import pip
def import_or_install(package):
try:
__import__(package)
except ImportError:
pip.main(['install', package])
import_or_install('importlib')
import_or_install('sys')
import_or_install('os')
import_or_install('pandas')
import_or_install('importlib')
import_or_install("numpy")
import_or_install("matplotlib")
import_or_install("sklearn")
import_or_install("pandas_market_calendars")
import_or_install("scipy")
import_or_install("datetime")
import_or_install("dateutil")
import_or_install("schedule")
import_or_install("patsy")
import_or_install("stldecompose==0.0.5")
import_or_install("statsmodels==0.10.2")
import_or_install("PyYAML==5.4.1")
import importlib
import sys
import os,sys
sys.path.insert(1, os.path.join(os.getcwd() , '..'))
# +
import importlib
import numpy as np
import pandas as pd
import TradingStrategy as ts
import Common.ApiClient as ac
import MA.ExponentialMovingAverageStrategy as ema
import MA.SimpleMovingAverageStrategy as sma
import PaperTrader as pTrader
pd.options.mode.chained_assignment = None
importlib.reload(ts)
importlib.reload(ac)
importlib.reload(ema)
importlib.reload(sma)
# -
# # Please provide your API Key and Secret Key below before you execute
Api_Key =''
Secret_Key=''
endpoint='https://paper-api.alpaca.markets'
client = ac.ApiClient(api_key_Id=Api_Key,api_key_secret=Secret_Key)
# # EMA Based Machine Learning Model
# +
for ticker in ["FB","MSFT","AMZN","AMD","GOOG"]:
df= client. get_closing_price(ticker,255)
ema_instance = ema.ExponentialMovingAverageStrategy(df=df,ticker=ticker)
test, pred_label=ema_instance.generate_train_model(ticker=ticker)
print(f'Buy and hold strategy returns for the backtest: {(test["daily_profit"].sum()*100).round(3)}%')
print(f'EMA based model strategy returns for the backtest: {(test["strategy_profit"].sum()*100).round(3)}%')
# -
total = 0.0
from STL.StlMl import STL_strategy
for s,ticker in enumerate([ "MSFT","AAPL","GM","GOOG","TSLA"]):
df= client.get_closing_price(ticker,255)
df.index=pd.to_datetime(df.index,utc=True)
stl = STL_strategy(ticker,df,'close',10,3)
total += stl.backtest()
print('totl profit:{:.2f}%'.format(total*100))
# # Running Paper trading
# please uncomment the code below for running the trade job
p = pTrader.PaperTrader( API_KEY_ID=Api_Key,SECRET_KEY=Secret_Key,model='ema')
p.run_trading()
| src/program.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (system-wide)
# language: python
# name: python3
# ---
# +
import sys
import csv
import warnings
import numpy as np
import seaborn as sns
from scipy import stats
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
if not sys.warnoptions:
warnings.simplefilter("ignore")
# -
# # Example 1
# - 120 children participated in the study, with 60 children assigned to the treatment (trt = 1), and 60 assigned to control (trt = 0).
#
#
# - There were some issues with noncompliance, and not every student in the treatment group received the treatment (rectrt = 1).
#
#
# - The outcomes of interest are height, weight and IQ scores in 2012.
# - In addition to the outcome variables, you have also been given pre-treatment scores in 2011 as well as a variety of demographic covariates (eg., living in urban or suburban).
# - The health outcome scores have been normalized and so are in standard deviation units. All of the variables in the dataset are labeled.
#
# # 1. Upload data
#Locate file
# !ls
#Upload file
file = 'rct_data.dta'
df = pd.read_stata(file)
df.head(3)
#Check variables
df.info()
# # 2. EDA
# ## 2.1 Descriptive statistics
df.describe()
# ## 2.2 Plot distribution of treated vs. non-treated
def summary_plot_2periods(boxplot, colnames):
fig, axes = plt.subplots(len(colnames) // 2, 2, figsize=(12, 8))
k = 0
for i in range(len(colnames) // 2):
for j in range(2):
if boxplot:
df.boxplot(column=colnames[k], by='trt', ax=axes[(i, j)])
else:
sns.violinplot('trt', colnames[k], data=df, ax=axes[(i, j)])
k += 1
colnames = ['height_2012', 'height_2011', 'weight_2012', 'weight_2011', 'IQ_2012', 'IQ_2011', 'urban', 'suburban']
summary_plot_2periods(boxplot=True, colnames=colnames)
# # 3. Preprocess data
# 120 children participated in the study, with 60 children assigned to the treatment (trt = 1), and 60 assigned to control (trt = 0). There were some issues with noncompliance, and not every student in the treatment group received the treatment (rectrt = 1).
#
# - The outcomes of interest are height, weight and IQ scores in 2012.
# - Y = height, weight, IQ
#
#
# - Pre-treatment scores in 2011 as well as a variety of demographic covariates (eg., living in urban or suburban).
# - X 2011, suburban vs. urban
# - The health outcome scores have been normalized and so are in standard deviation units.
# - All of the variables in the dataset are labeled.
# # 3.1 Constructing vectors of Y (label), and X(regressors)
#
# - Iterate data to create 3 seperate models
# Y = height, weight, IQ
# +
file = 'rct_data.dta' #DTA filename
df = pd.read_stata(file) #Import data from the top to be save
y_list = ['height_2012', 'weight_2012', 'IQ_2012'] #Y variables
lagged_DV = [sub.replace('2012', '2011') for sub in y_list] #Lagged Y-vars
for idx, (DV, lagged) in enumerate(zip(y_list, lagged_DV)): #Iterate over y_list and lagged_DV
x = df[['trt' ,'urban', str(lagged)]] #Set independent variables + each respective lagged var
x = sm.add_constant(x) #Add constant to indepdent variables
y = df[[DV]] #Y-variable
print('Model '+str(idx+1)+': y='+str(y.columns.values), 'x='+str(x.columns.values)) #Print each model that will be used
# -
# # 4. Baseline regressions
# ## 4.1 Exploring non-compliant
# ### 4.1.1 Approaches to non-compliant
# 1. `Per protocal:` Examine only participants who were compliant
# 2. `As treated:` Examined as they were treated
# 3. `Intention to treat:` Examined as they were assigned
# ### 4.1.2 Exploring non-compliars in data set
mask = (df.trt==1) & (df.rectrt==0)
df.loc[mask]
# ### 4.1.2 Create Non-compliant dummy
df['Non_compliant'] = pd.get_dummies(mask, drop_first=True)
pct_noncomplier = df['Non_compliant'].mean()
print('{:.2f}% of treated were non-complient'.format(pct_noncomplier*100))
# ## 4.2 Is RCT balanced?
# ### 4.2.1 T-test on means
# +
means = ['height_2011', 'weight_2011', 'IQ_2011', 'urban', 'suburban']
for mean in means:
non_comp = df.loc[df['trt']==1, mean].values
complier = df.loc[df['trt']==0, mean].values
#print(stats.ttest_ind(non, complier))
tval, pval = stats.ttest_ind(non_comp, complier)
if pval>.01:
print('Fail to reject, P-val:'+str(round(pval,2))+'\n For '+ str(mean)+ ' we cannot conclude there is a difference between treated and un-treated means')
# -
# ## 4.3 Baseline regression
# ### Intention to treat (ITT) regressions:
# `Intention to treat:` Examined as they were assigned
y_list = ['height_2012', 'weight_2012', 'IQ_2012'] #Y variables
lagged_DV = [sub.replace('2012', '2011') for sub in y_list] #Lagged Y-vars
for idx, (DV, lagged) in enumerate(zip(y_list, lagged_DV)): #Iterate over y_list
x = df[['trt' ,'urban', str(lagged)]]
x = sm.add_constant(x)
y = df[[DV]]
baseline = sm.OLS(y, x.values).fit() #OLS
print(baseline.summary())#Print summary of OLS
'''Code to save a csv file for regressions'''
#f = open(str(DV)+'_csvfile.csv','w')
#f.write(baseline.summary().as_csv())
#f.close()
'''Code to show scatter plots of fitted values'''
#plt.scatter(baseline.fittedvalues, baseline.resid) #Plot residuals
#plt.axhline(0, linestyle=':', color='orange') #Add line
#plt.title('Model '+str(idx+1)+ ' Residuals') #Title
#plt.show() #Show plot
print('\n \n \n')#Space between tables
# ## 4.4 Two-stage least-square regressions (Instrumental variable)
# ## Late regression
#
# - [Using the "Linear Models" Library](https://bashtage.github.io/linearmodels/doc/iv/index.html)
# - [Examples of 2SLS in linear models](https://bashtage.github.io/linearmodels/doc/iv/examples/basic-examples.html)
# - [More examples](https://bashtage.github.io/linearmodels/doc/iv/examples/advanced-examples.html)
from linearmodels.iv import IV2SLS
# + hideCode=true hideOutput=true
instrument = df[['rectrt']]
endogenous = df[['trt']]
y_list = ['height_2012', 'weight_2012', 'IQ_2012'] #Y variables
lagged_DV = [sub.replace('2012', '2011') for sub in y_list] #Lagged Y-vars
instrument = df[['trt']]
endogenous = df[['rectrt']]
for idx, (DV, lagged) in enumerate(zip(y_list, lagged_DV)): #Iterate over y_list
x = df[['urban', str(lagged)]]
x = sm.add_constant(x)
y = df[[DV]]
IV = IV2SLS(y, x, endogenous, instrument).fit()
print(IV)
print('\n \n \n')#Space between tables
| 2sls/RCT_2sls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="XHAFPHWmfFoB" colab={"base_uri": "https://localhost:8080/"} outputId="7872eab5-6fc9-47df-d390-70921b9fe24d"
#Lista
animais = [1,2,3]
animais
# + id="5v-w_ftdga0J" colab={"base_uri": "https://localhost:8080/"} outputId="98b9c2bd-10a4-46e8-f38f-0ab2976c25d7"
animais = ["cachorro", "gato", 12345, 6.5]
animais
# + id="bJkOFrGfgrKR" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4558a8c7-09ae-4e78-eb4b-9d93eab4968e"
animais[0]
# + id="M-2mXu3mgvuS" colab={"base_uri": "https://localhost:8080/"} outputId="46b8c981-21a5-4783-eacd-3712848a6829"
animais[3]
# + id="RDCW4hs_gxyT"
animais[0] = "papagaio"
# + id="QJrP-830gzvh" colab={"base_uri": "https://localhost:8080/"} outputId="0b69d63c-7901-40c0-eccd-177dfc68bf4e"
animais
# + id="3QFbAI6Og0wP"
animais.remove('gato')
# + id="b8g1bs42g31p" colab={"base_uri": "https://localhost:8080/"} outputId="d4950ab2-005a-4edf-bb4b-d510dffeeb46"
animais
# + id="xfu4p_g3g7OS" colab={"base_uri": "https://localhost:8080/"} outputId="17ae3057-e164-45fa-bd30-0f6c3d06b94d"
len(animais)
# + id="0HedcjoFg-5m" colab={"base_uri": "https://localhost:8080/"} outputId="614f4a3a-4dcd-4c62-b0e1-2617cef5ff7d"
"gato" in animais
# + id="VDZ6nZU2hDdf"
lista = [500, 30, 300, 80, 10]
# + id="VjrYRfHJhGhL" colab={"base_uri": "https://localhost:8080/"} outputId="ab4b36e0-0656-4466-e0e7-96f9ca95f470"
max(lista)
# + id="jkeXKWQPhJRr" colab={"base_uri": "https://localhost:8080/"} outputId="5149af70-ab31-4da4-d2dc-1a69a7e91314"
min(lista)
# + id="20T_hCwAhMsU"
animais.append(["leão", "Cachorro"])
# + id="WgWWMNJXhQdB" colab={"base_uri": "https://localhost:8080/"} outputId="ce66aaa9-5463-4b7f-872d-ed2fd01de6f9"
animais
# + id="Cl-M69f5hSsD"
animais.extend(["cobra", 6])
# + id="_scJEZbWMFBW" colab={"base_uri": "https://localhost:8080/"} outputId="e2f4d95a-f996-48a0-86d7-2d0339f28fe8"
animais
# + id="kNn0DiyxheHC" colab={"base_uri": "https://localhost:8080/"} outputId="c667c89c-c65d-4138-a925-5210739782d3"
animais.count("leão")
# + id="VgoLBevzhinC"
lista.sort()
# + id="mPJTKqXMhrDB" colab={"base_uri": "https://localhost:8080/"} outputId="8a733c6a-7050-41e6-9a6c-855944bbc47f"
lista
# + id="Qrv07mzWocDT"
tp = ("Banana", "Maçã", 10, 50)
# + id="r7ZMMLn8pt7R" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="572b458f-fd85-451e-bd81-9839c2239566"
tp[0]
# + id="jKRnh-DbpzBj" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="42e0032b-f3a6-42a1-91c4-1056f8efd098"
tp[0] = "Laranja"
# + id="a-KaNhePE0nb" colab={"base_uri": "https://localhost:8080/"} outputId="1c634037-b768-4760-8e44-49e610051951"
tp.count("Maçã")
# + id="ntXTi7MaFAX7" colab={"base_uri": "https://localhost:8080/"} outputId="d06c6428-673a-4daf-a166-34b8c654fdda"
tp[0:2]
# + id="43OFknPUFO68"
#Dicionario
dc = {"Maçã":20, "Banana":10, "Laranja":15, "Uva":5}
# + id="dCH9PefxGBdZ" colab={"base_uri": "https://localhost:8080/"} outputId="efc34393-5bfe-4b10-f233-1bd344c745e6"
dc
# + id="iyGxfz72FxNa" colab={"base_uri": "https://localhost:8080/"} outputId="3ec7e792-38ca-4f63-e7cb-0c7752d7404a"
dc["Maçã"]
# + id="6WwkohdpF5Cq" colab={"base_uri": "https://localhost:8080/"} outputId="ba64ebde-833e-4b31-985b-c0dbb354ffb8"
dc["Maçã"] = 25
dc
# + id="_50MV-XaGFae" colab={"base_uri": "https://localhost:8080/"} outputId="ef69e32f-c561-4a95-e432-3a89e99e2d3a"
dc.keys()
# + id="AHLXfikMGRS4" colab={"base_uri": "https://localhost:8080/"} outputId="6edb9720-9fe8-4009-e10b-bbc784ddd027"
dc.values()
# + id="co2yQ4IrGX8n" colab={"base_uri": "https://localhost:8080/"} outputId="06d49295-b27d-4706-9946-e537a7edb3a2"
dc.setdefault("Limão", 22)
# + id="M9_v4jAZGyz6" colab={"base_uri": "https://localhost:8080/"} outputId="10f6fdad-a94e-454b-f9e8-a5ebb766df17"
dc
# + id="l2-4mE1LG0CC"
| 2_Estrutura_dados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: zerospeech
# language: python
# name: zerospeech
# ---
# +
from utils.ZR_utils import (get_matches_df, post_disc, plot_match_stats, change_post_disc_thr,
get_nodes_df, get_clusters_list, get_matches_all)
from utils.eval import evaluate
from utils.helper_fncs import save_obj, load_obj
from utils.db_utils import gold_fragments_df_for_signer
from utils.feature_utils import (op_100_meancenter)
import glob
import pandas as pd
import numpy as np
import os
# +
import matplotlib.pyplot as plt
label_dict = {'coverage': 'Coverage',
'ned':'NED',
'purity': 'Purity',
'avg_purity': 'Average Purity',
'clus_purity':'Cluster Purity',
'grouping_P': 'Grouping Precision',
'grouping_R': 'Grouping Recall',
'grouping_F': 'Grouping F-score',
'clus_purity_inv': 'Inverse Cluster Purity'}
def slct_res(res,sel):
tmp = []
for i, r in enumerate(res):
if i not in sel: tmp.append(r)
return tmp
def plot_curve(ax, results, ax_dict, marker, label, annot_freq=2):
# results: list of score dicts
data = np.zeros((len(results), 2))
annots = []
for i,score in enumerate(results):
data[i,0] = score[ax_dict['x']]
data[i,1] = score[ax_dict['y']]
annots.append(score['dtw'])
ax.plot(data[:,0], data[:,1], marker, label=label)
ax.set_xlabel(label_dict[ax_dict['x']])
ax.set_ylabel(label_dict[ax_dict['y']])
for i,ann in enumerate(annots):
if (i%annot_freq != 0) or (annot_freq==-1): continue
ax.annotate(str(ann),
xy=(results[i][ax_dict['x']], results[i][ax_dict['y']]),
xycoords='data')
# +
exp_root = '/home/korhan/Desktop/zerospeech2017/track2/src/ZRTools/exp/'
exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer07'
#exp_ = 'Signer01_Signer08_c3_right_PCA50_10_3_02_7_05_055_11_06_03'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer01'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer04_Signer05_Signer0950_10_35_02_7_05_06_9_06_032'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer01_Signer0850_10_4_02_7_05_06_7_06_032'
#exp_ = 'Signer07op10050_10_5_02_7_05_065_13_06_04'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanhSigner0350_10_4_02_7_05_06_9_06_03'
#exp_ = 'Signer03_Signer07_c3_right_PCA50_10_3_02_7_05_055_11_06_03'
exp_ = 'Signer03_c3_right_PCA50_10_3_02_7_05_06_13_06_04'
#exp_ = 'op100_Signer03_Signer0750_10_4_02_7_05_06_9_06_03'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer03_Signer0750_10_4_02_7_05_06_9_06_032'
#exp_ = 'op100_Signer08'
#exp_ = 'corAE_op100_s4s5_64_iter0_self_tanh_Signer0450_10_4_02_7_05_06_7_06_032'
signer_id = ['Signer0'+ str(i) for i in [8]]
#signer_id = ['Signer0'+ str(i) for i in [3,7]]
signer_id = ['Signer0'+ str(i) for i in [3]]
print(signer_id)
def full_exp_path(exp_):
exp_root = '/home/korhan/Desktop/zerospeech2017/track2/src/ZRTools/exp/'
try:
name_root = glob.glob(exp_root + 'zrroot0_{}*/exp/*'.format(exp_))[0]
exp_name = '/'.join(name_root.split('/')[-3:])[:-6]
except:
name_root = glob.glob(exp_root + '{}*/'.format(exp_))[0]
exp_name = glob.glob(exp_root + '{}*/'.format(exp_))[0].split('/')[-2][:-6]
return name_root, exp_name
print(exp_name)
# -
matches_df = get_matches_all(exp_name)
matches_df.head()
plot_match_stats(matches_df)
# +
results = []
#for dtw in np.arange(0.89,0.98,0.01):
for dtw in [0.91]:
print(dtw)
results_path = post_disc(exp_name, dtw)
" get nodes "
nodes_df = get_nodes_df(exp_name)
" get clusters "
clusters_list = get_clusters_list(exp_name)
if len(clusters_list)==0: continue
scores = evaluate(None, signer_id, matches_df, nodes_df, clusters_list,
group=3, interp=False, boundary_th=0.5, fast_compute=True, purify=True)
scores['dtw'] = dtw
scores['exp'] = exp_name
results.append(scores)
# -
scores
results
resname = 'train_set'
final = load_obj(name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves')
final['AE (Signer04)2'] = results
final = dict()
resname = 'train_set'
final['AE (Signer04 Signer05 Signer09)'] = results
save_obj(obj=final, name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves' )
resname = 'test_set'
final = load_obj(name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves')
results
results.extend(final['AE (Signer01)'])
final['AE (Signer01)'] = results
fig, axes = plt.subplots(1,1,squeeze=False)
for ky, res in final.items():
plot_curve(axes[0,0], res, {'x': 'coverage', 'y': 'clus_purity'}, label=ky, annot_freq=6)
plt.legend()
fig, axes = plt.subplots(1,1,squeeze=False)
for ky, res in final.items():
plot_curve(axes[0,0], res, {'x': 'coverage', 'y': 'clus_purity'}, label=ky, annot_freq=6)
plt.legend()
res
resname = 'train_set'
train = load_obj(name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves')
train.keys()
# +
# %matplotlib inline
tmp = train
ax_dict = {'x': 'coverage', 'y': 'clus_purity'}
fig, axes = plt.subplots(1,1,squeeze=False, figsize=(5,5))
ky = 'AE (Signer04)'
res = tmp[ky]
lb = 'AutoEncoder (4)'
plot_curve(axes[0,0], slct_res(res[:],[]), ax_dict, '--x', label=lb, annot_freq=1)
ky = 'AE (Signer04)2'
res = tmp[ky]
lb = 'AutoEncoder (4)2'
plot_curve(axes[0,0], slct_res(res[:],[]), ax_dict, '--x', label=lb, annot_freq=1)
"""ky = 'AE (Signer05)'
res = tmp[ky]
lb = 'AutoEncoder (5)'
plot_curve(axes[0,0], slct_res(res[:],[]), ax_dict, '--x', label=lb, annot_freq=1)
"""
plt.legend()
# -
resname = 'test_set'
test = load_obj(name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves')
test.keys()
# +
# %matplotlib inline
tmp = test
ax_dict = {'x': 'coverage', 'y': 'clus_purity'}
fig, axes = plt.subplots(1,1,squeeze=False, figsize=(5,5))
ky = 'DH (Signer01,Signer08)'
res = tmp[ky]
lb = 'DeepHand (test)'
plot_curve(axes[0,0], res[:-2], ax_dict, '-x', label=lb, annot_freq=-1)
ky = 'AE (Signer01,Signer08)'
res = tmp[ky]
lb = 'AutoEncoder (test)'
plot_curve(axes[0,0], slct_res(res[2:-4],[]), ax_dict, '--x', label=lb, annot_freq=-1)
ky = 'AE (Signer08)'
res = tmp[ky]
lb = 'AutoEncoder (S08)'
plot_curve(axes[0,0], slct_res(res[2:],[]), ax_dict, '--x', label=lb, annot_freq=-1)
ky = 'DH (Signer08)'
res = tmp[ky]
lb = 'DeepHand (S08)'
plot_curve(axes[0,0], slct_res(res[:],[]), ax_dict, '-x', label=lb, annot_freq=-1)
ky = 'DH (Signer01)'
res = tmp[ky]
lb = 'DeepHand (S01)'
plot_curve(axes[0,0], slct_res(res[:],[]), ax_dict, '-x', label=lb, annot_freq=-1)
ky = 'OP (Signer01,Signer08)'
res = tmp[ky]
lb = 'OpenPose (test)'
plot_curve(axes[0,0], res[6:], ax_dict, '-.x', label=lb, annot_freq=-1)
plt.legend()
# -
resname = 'dev_set'
dev = load_obj(name='grid_post_disc_' + resname, path='/home/korhan/Dropbox/tez/files/ZR_dtw_curves')
dev.keys()
# +
tmp = dev
ax_dict = {'x': 'coverage', 'y': 'ned'}
fig, axes = plt.subplots(1,1,squeeze=False, figsize=(5,5))
ky = 'DH (Signer03,Signer07)'
res = tmp[ky]
lb = 'DeepHand (dev)'
plot_curve(axes[0,0], res[4:-3], ax_dict, '-x', label=lb, annot_freq=-1)
ky = 'DH (Signer01,Signer08)'
res = test[ky]
lb = 'DeepHand (test)'
plot_curve(axes[0,0], res[1:-2], ax_dict, '-x', label=lb, annot_freq=-1)
"""ky = 'AE (Signer03,Signer07)'
res = tmp[ky]
lb = 'AutoEncoder (dev)'
plot_curve(axes[0,0], res[4:-2], ax_dict, '--x', label=lb, annot_freq=-1)
"""
"""ky = 'OP (Signer03,Signer07)'
res = tmp[ky]
lb = 'OpenPose (dev)'
plot_curve(axes[0,0], res[6:], ax_dict, '-.x', label=lb, annot_freq=-1)
"""
plt.legend()
# +
cost_thr = 0.20
cost_val = matches_df.score.sort_values(ascending=False).values[round(len(matches_df) * cost_thr)]
print(cost_val)
# +
#cost_val = 0.90
results_path = post_disc(exp_name, cost_val)
" get nodes "
nodes_df = get_nodes_df(exp_name)
" get clusters "
clusters_list = get_clusters_list(exp_name)
scores = evaluate(None, signer_id, matches_df, nodes_df, clusters_list,
group=3, interp=False, boundary_th=0.5, fast_compute=False)
scores['exp'] = exp_name
# -
scores
# +
import pandas as pd
select = ['exp','n_clus','n_node','coverage',
'matching_P','matching_R','matching_F',
'grouping_P','grouping_R','grouping_F',
'clus_purity', 'clus_purity_inv']
pd.DataFrame([scores], columns=scores.keys())[select]
# -
nodes_df
# +
from utils.db_utils import nodes_with_info, gold_fragments_df_for_signer, nodes_with_types
clusters_set = set()
for row in clusters_list: clusters_set |= set(row)
all_centroids = list(clusters_set)
# select only nodes that belong to a cluster (dedups?)
nodes_df = nodes_df.loc[all_centroids]
gold_fragments = gold_fragments_df_for_signer(signer_id)
nodes_df = nodes_with_info(nodes_df=nodes_df, gold_fragments=gold_fragments)
nodes_df = nodes_with_types(nodes_df=nodes_df, gold_fragments=gold_fragments)
for clus in clusters_list:
n_signer_different = len(nodes_df.signer_id[clus].unique())
if n_signer_different>1:
print(clus,n_signer_different)
# -
for clus in clusters_list:
n_signer_different = len(nodes_df.signer_id[clus].unique())
if n_signer_different>1:
print(clus,n_signer_different)
nodes_df
nodes
change_post_disc_thr(olapthr=0.9, dedupthr=0.2, durthr=5, rhothr=1000, min_edge_w=0)
# +
# 0.9 0.2 0
# -
#results = []
for olap in [ 0.3]:
for dedup in [0.5]:
for th in [600, 800]:
change_post_disc_thr(olapthr=olap, dedupthr=dedup, durthr=5, rhothr=1000, min_edge_w=th)
print('olap {}, dedup {}'.format(olap, dedup))
results_path = post_disc(exp_name, dtw)
" get nodes "
nodes_df = get_nodes_df(exp_name)
" get clusters "
clusters_list = get_clusters_list(exp_name)
if len(clusters_list)==0: continue
scores = evaluate(None, signer_id, matches_df, nodes_df, clusters_list,
group=3, interp=False, boundary_th=0.5, fast_compute=True)
scores['dtw'] = dtw
scores['exp'] = exp_name
scores['olap'] = olap
scores['dedup'] = dedup
scores['th'] = th
results.append(scores)
results[0].keys()
pd.DataFrame.from_records(results)[['mean_len','n_node','n_clus','ned','coverage','olap','dedup','th']]
| post_disc_result_prepare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# +
# #%matplotlib notebook
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# -
# Read the mouse data and the study results. Load in csv file.
mouse_metadata_df= pd.read_csv(mouse_metadata_path)
mouse_metadata_df
# Read the mouse data and the study results. Load in csv file.
study_results_df = pd.read_csv(study_results_path)
study_results_df
# Combine the data into a single dataset
combined_results_df=pd.merge(mouse_metadata_df,study_results_df,how="outer",on="Mouse ID")
combined_results_df
# Checking the number of mice in the DataFrame.
# mice_instances_combined=combined_results_df["Mouse ID"].count()
# mice_instances_combined
mouse_metadata_df.count()
# +
## DUPLICATE MOUSE IDENTIFIED ##
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
duplicate_rows=combined_results_df[combined_results_df.duplicated()]
duplicate_rows
# +
## Optional: Get all the data for the duplicate mouse ID. ##
duplicate_rows=combined_results_df[combined_results_df.duplicated(keep=False)]
print("All Duplicate Rows based on all data columns is :")
print(duplicate_rows)
# -
# Checking the number of mice in the clean DataFrame before dropping duplicate records.
combined_results_df.count()
# +
## REMOVE THE DUPLICATE MOUSE/MICE ##
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
#### <NAME> DAY 2 -01 ####
#clean_combined_results_df=combined_results_df.drop_duplicates(keep='first')
#print('Duplicate records dropped :\n', clean_combined_results_df)
clean_combined_results_df=combined_results_df.drop_duplicates(inplace=True)
#print(clean_combined_results_df)
# -
# Test to validate that the duplicate record is dropped from the dataset.
duplicate_rows=combined_results_df[combined_results_df.duplicated(keep=False)]
print("All Duplicate Rows based on all data columns is :")
print(duplicate_rows)
# Checking the number of mice in the clean DataFrame.
combined_results_df.count()
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# This method is the most straighforward, creating multiple series and putting them all together at the end.
# For Tumor Volume only use necessary columns
tumor_volume_df=combined_results_df.loc[:,["Drug Regimen","Mouse ID","Timepoint","Tumor Volume (mm3)"]]
tumor_volume_df
# -
# Generate a summary statistics table
drug_regimen_df=tumor_volume_df.groupby(["Drug Regimen"])
drug_regimen_df.describe()
# +
## DRUG REGIMEN VS. TUMOR VOLUME & TIMEPOINT SUMMARY STATISTICS TABLE ##
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
tumor_volume_statistics_df=tumor_volume_df.groupby(["Drug Regimen","Timepoint"]).agg({"Tumor Volume (mm3)":["mean","median","var","std","sem"]})
tumor_volume_statistics_df
# -
## DRUG REGIMEN VS. TUMOR VOLUME SUMMARY STATISTICS TABLE ##
tumor_volume_summary=pd.DataFrame(tumor_volume_df.groupby("Drug Regimen").count())
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
tumor_volume_summary=tumor_volume_df.groupby(["Drug Regimen"]).agg({"Tumor Volume (mm3)":["mean","median","var","std","sem"]})
#tumor_volume_summary2=tumor_volume_summary[["Mouse ID", "Mean", "Median", "Variance","Standard Deviation","SEM"]]
#tumor_volume_summary=tumor_volume_summary2.rename(columns={"Mouse ID":"Treatments"})
tumor_volume_summary
# +
## DRUG REGIMEN VS. TUMOR VOLUME SUMMARY STATISTICS TABLE OUTPUT ##
#Use groupby to create summary stats by drug regime, add results into columns in summarystats
tumor_volume_summary_output=pd.DataFrame(tumor_volume_df.groupby("Drug Regimen").count())
tumor_volume_summary_output["Mean"] = pd.DataFrame(tumor_volume_df.groupby("Drug Regimen")["Tumor Volume (mm3)"].mean())
tumor_volume_summary_output["Median"] = pd.DataFrame(tumor_volume_df.groupby("Drug Regimen")["Tumor Volume (mm3)"].median())
tumor_volume_summary_output["Variance"] = pd.DataFrame(tumor_volume_df.groupby("Drug Regimen")["Tumor Volume (mm3)"].var())
tumor_volume_summary_output["Standard Deviation"] = pd.DataFrame(tumor_volume_df.groupby("Drug Regimen")["Tumor Volume (mm3)"].std())
tumor_volume_summary_output["SEM"] = pd.DataFrame(tumor_volume_df.groupby("Drug Regimen")["Tumor Volume (mm3)"].sem())
#Clean up columns and rename count column
tumor_volume_summary_output = tumor_volume_summary_output[["Mouse ID", "Mean", "Median", "Variance","Standard Deviation","SEM"]]
tumor_volume_summary_output = tumor_volume_summary_output.rename(columns={"Mouse ID":"Treatments"})
tumor_volume_summary_output
# -
# ## Bar Plots
bar_pandas_plot=combined_results_df
# +
# Generate a bar plot showing the number of mice per time point for each treatment throughout the course of the study using pandas.
drug_regimen_timepoints_df=combined_results_df.groupby(["Drug Regimen"])
#drug_regimen_timepoints_df.head()
mice_count_df=drug_regimen_timepoints_df['Mouse ID'].count()
#mice_count_df
# Set x and y limits
#x_axis=np.arange(len(datapoints))
#tick_locations=[value for value in x_axis]
#plt.xlim(-0.75, len(x_axis)-.25)
# Chart the data
chart_mice_per_drugregimen_timepoint = mice_count_df.plot(kind="bar", title="Drug Regimen Mice Count Per Timepoint",color='b',legend=False)
#chart_mice_per_drugregimen_timepoint = drug_regimen_timepoints_df.plot(kind="bar", title="Drug Regimen Mice Count Per Timepoint")
chart_mice_per_drugregimen_timepoint.set_xlabel("Drug Regimen")
chart_mice_per_drugregimen_timepoint.set_ylabel("Count of Mice Per Timepoint")
plt.show()
plt.tight_layout()
#bar_plot_data=combined_results_df[["Drug Regimen"]]
#bar_plot_data
# -
# Generate a bar plot showing the number of mice per time point for each treatment throughout the course of the study using pyplot.
# ## Pie Plots
#gender=combined_results_df.groupby('Sex')
gender_counts=combined_results_df["Sex"].value_counts()
gender_counts
# Generate a pie plot showing the distribution of female versus male mice using pandas
#combined_results_df.groupby('Sex')["Mouse ID"].nunique().plot(kind='pie',title="Drug Regimen Gender Distribution",autopct='%1.1f%%',shadow=True, startangle=25)
combined_results_df.groupby('Sex')["Mouse ID"].nunique().plot(kind='pie',title="Drug Regimen Gender Distribution",autopct='%1.1f%%',shadow=True, startangle=25)
# +
# Generate a pie plot showing the distribution of female versus male mice using pyplot
#gender_counts.plot(kind='pie',title="Drug Regimen Gender Distribution",autopct='%1.1f%%',shadow=True,startangle=205)
#plt.title("Drug Regimen Gender Distribution")
plt.pie(gender_counts,autopct='%1.1f%%',shadow=True,startangle=205)
#plt.axis("equal")
#plt.show()
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens. Calculate the IQR and quantitatively determine if there are any potential outliers.
# Grab just data for the 4 smallest mean tumor volume regimens
filtered_df = combined_results_df.loc[(combined_results_df["Drug Regimen"] == "Capomulin") | (combined_results_df["Drug Regimen"] == "Ramicane") | (combined_results_df["Drug Regimen"] == "Ceftamin") | (combined_results_df["Drug Regimen"] == "Propriva"), :]
# Sort by Timpepoints based on the latest values
filtered_df = filtered_df.sort_values("Timepoint", ascending = False)
# Dropping duplicates, keeping first value, should be the latest timepoint per mouse
filtered_df = filtered_df.drop_duplicates(subset="Mouse ID", keep='first')
# Determine quartiles
quartiles = filtered_df['Tumor Volume (mm3)'].quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
# Determine upper and lower bounds
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
# Print a filtered dataframe of any outliers
outliers_df = filtered_df.loc[(filtered_df['Tumor Volume (mm3)'] > upper_bound) | (filtered_df['Tumor Volume (mm3)' ] < lower_bound), :]
outliers_df
# Not finding any outliers.
# -
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
Tumor_Volume = filtered_df['Tumor Volume (mm3)']
fig1, ax1 = plt.subplots()
ax1.set_title('Tumor Volume of Mice')
ax1.set_ylabel('Tumor Volume')
ax1.boxplot(Tumor_Volume)
plt.show()
# ## Line and Scatter Plots
# +
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
# Filter original data for just the Capomulin Drug Regime
Capomulin_df = combined_results_df.loc[(combined_results_df["Drug Regimen"] == "Capomulin"),:]
# Set variables to hold relevant data
timepoint = Capomulin_df["Timepoint"]
tumor_volume = Capomulin_df["Tumor Volume (mm3)"]
# Plot the tumor volume for various mice
tumor_volume_line = plt.plot(timepoint, tumor_volume)
# Show the chart, add labels
plt.xlabel('Timepoint')
plt.ylabel('Tumor Volume')
plt.title('Tumor Volume over Time for Capomulin Mice')
plt.show()
# +
# Generate a scatter plot of mouse weight versus average tumor volume for the Capomulin regimen
# Pull values for x and y values
mouse_weight = Capomulin_df.groupby(Capomulin_df["Mouse ID"])["Weight (g)"].mean()
tumor_volume = Capomulin_df.groupby(Capomulin_df["Mouse ID"])["Tumor Volume (mm3)"].mean()
# Create Scatter Plot with values calculated above
plt.scatter(mouse_weight,tumor_volume)
plt.xlabel("Weight of Mouse")
plt.ylabel("Tumor Volume")
plt.show()
# -
# ## Correlation and Regression
# +
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
# Pull values for x and y values
mouse_weight = Capomulin_df.groupby(Capomulin_df["Mouse ID"])["Weight (g)"].mean()
tumor_volume = Capomulin_df.groupby(Capomulin_df["Mouse ID"])["Tumor Volume (mm3)"].mean()
# Perform a linear regression on year versus violent crime rate
slope, int, r, p, std_err = st.linregress(mouse_weight, tumor_volume)
# Create equation of line to calculate predicted violent crime rate
fit = slope * mouse_weight + int
# Plot the linear model on top of scatter plot
plt.scatter(mouse_weight,tumor_volume)
plt.xlabel("Weight of Mouse")
plt.ylabel("Tumor Volume")
plt.plot(mouse_weight,fit,"--")
plt.xticks(mouse_weight, rotation=90)
plt.show()
# Caculate correlation coefficient
corr = round(st.pearsonr(mouse_weight,tumor_volume)[0],2)
print(f'The correlation between weight and tumor value is {corr}')
# -
| .ipynb_checkpoints/pymaceuticals_starter_Rob-checkpoint.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// ### Permutation Test
#include <iostream>
#include <cstdio>
#include <vector>
#include <algorithm>
using namespace std;
int permutation(vector<int>& vec, int r)
{
reverse(vec.begin()+r, vec.end());
return next_permutation(vec.begin(), vec.end());
}
// +
vector<int> vec;
vector<int>::iterator iter;
int n = 6;
for(int i=0; i<n; i++)
{
vec.push_back(i);
}
// -
int r = 3;
while(1)
{
for(int i=0; i<r; i++)
{
cout <<vec[i] << " ";
}
cout << endl;
if(!permutation(vec, r))
break;
}
// #### int형 1차원 벡터와 r(nPr 중 뽑아낼 개수 r임)을 입력하면 모든 경우의 수를 dst에 입력한다.
void perm(vector<int>& vec, vector<vector<int>>& dst, int r)
{
while(1)
{
vector<int> indice;
for(int i=0; i<r; i++)
{
indice.push_back(vec[i]);
}
dst.push_back(indice);
if(!permutation(vec, r))
break;
}
}
vector<int> test;
for(int i=0; i<5; i++)
{
test.push_back(i+1);
}
vector<vector<int>> perm_vec;
perm(test, perm_vec, 3);
cout << perm_vec.size();
for(int i=0; i< perm_vec.size(); i++)
{
for(int j=0; j<perm_vec[i].size(); j++)
{
cout << perm_vec[i][j] << " ";
}
cout << endl;
}
for(auto row : perm_vec)
{
for(auto elem : row)
{
cout << elem << " ";
}
cout << endl;
}
| notebook/permutation.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0-DEV
# language: julia
# name: julia-1.7
# ---
# +
# soft scope in REPL, Jupyter, etc.
n = 10
s = 0
for k in 1:n
s += k^3
end
@show s;
# +
# hard scope
module O1
n = 10
s = 0
for k in 1:n
s += k^3
end
@show s
end;
# +
# hard scope
module O2
n = 10
s = 0
for k in 1:n
global s += k^3
end
@show s
end;
# +
# hard scope
module O3
function f(n)
s = 0
for k in 1:n
s += k^3
end
s
end
end
@show O3.f(10);
# +
# hard scope
module O4
let
n = 10
s = 0
for k in 1:n
s += k^3
end
@show s
end
end;
# +
# hard scope
module O5
let
n = 10
s = 0
for k in 1:n
s += k^3
end
s
end
print("outside of let block:\n")
@show s
end;
# +
# hard scope
module O6
s = let
n = 10
s = 0
for k in 1:n
s += k^3
end
s
end
print("\noutside of let block:\n")
@show s
end;
# -
| 0001/soft and hard scopes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Machine Learning Engineer Nanodegree
# ## Supervised Learning
# ## Project: Building a Student Intervention System
# Welcome to the second project of the Machine Learning Engineer Nanodegree! In this notebook, some template code has already been provided for you, and it will be your job to implement the additional functionality necessary to successfully complete this project. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `'TODO'` statement. Please be sure to read the instructions carefully!
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ### Question 1 - Classification vs. Regression
# *Your goal for this project is to identify students who might need early intervention before they fail to graduate. Which type of supervised learning problem is this, classification or regression? Why?*
# **Answer: ** This is going to be a **classification** problem as we are classifying students into one of the two classes, need intervention or not.
# ## Exploring the Data
# Run the code cell below to load necessary Python libraries and load the student data. Note that the last column from this dataset, `'passed'`, will be our target label (whether the student graduated or didn't graduate). All other columns are features about each student.
# +
# Import libraries
import numpy as np
import pandas as pd
from time import time
from sklearn.metrics import f1_score
# Read student data
student_data = pd.read_csv("student-data.csv")
print "Student data read successfully!"
# -
# ### Implementation: Data Exploration
# Let's begin by investigating the dataset to determine how many students we have information on, and learn about the graduation rate among these students. In the code cell below, you will need to compute the following:
# - The total number of students, `n_students`.
# - The total number of features for each student, `n_features`.
# - The number of those students who passed, `n_passed`.
# - The number of those students who failed, `n_failed`.
# - The graduation rate of the class, `grad_rate`, in percent (%).
#
# +
# TODO: Calculate number of students
n_students = len(student_data.index)
# TODO: Calculate number of features
n_features = len(student_data.columns) - 1 # to account for the additional target label "passed"
# TODO: Calculate passing students
n_passed = student_data.passed.str.contains('yes').sum()
# TODO: Calculate failing students
n_failed = student_data.passed.str.contains('no').sum()
# TODO: Calculate graduation rate
grad_rate = (float(n_passed)/float(n_students))*100
# Print the results
print "Total number of students: {}".format(n_students)
print "Number of features: {}".format(n_features)
print "Number of students who passed: {}".format(n_passed)
print "Number of students who failed: {}".format(n_failed)
print "Graduation rate of the class: {:.2f}%".format(grad_rate)
# -
# ## Preparing the Data
# In this section, we will prepare the data for modeling, training and testing.
#
# ### Identify feature and target columns
# It is often the case that the data you obtain contains non-numeric features. This can be a problem, as most machine learning algorithms expect numeric data to perform computations with.
#
# Run the code cell below to separate the student data into feature and target columns to see if any features are non-numeric.
# +
# Extract feature columns
feature_cols = list(student_data.columns[:-1])
# Extract target column 'passed'
target_col = student_data.columns[-1]
# Show the list of columns
print "Feature columns:\n{}".format(feature_cols)
print "\nTarget column: {}".format(target_col)
# Separate the data into feature data and target data (X_all and y_all, respectively)
X_all = student_data[feature_cols]
y_all = student_data[target_col]
# Show the feature information by printing the first five rows
print "\nFeature values:"
print X_all.head()
# -
# ### Preprocess Feature Columns
#
# As you can see, there are several non-numeric columns that need to be converted! Many of them are simply `yes`/`no`, e.g. `internet`. These can be reasonably converted into `1`/`0` (binary) values.
#
# Other columns, like `Mjob` and `Fjob`, have more than two values, and are known as _categorical variables_. The recommended way to handle such a column is to create as many columns as possible values (e.g. `Fjob_teacher`, `Fjob_other`, `Fjob_services`, etc.), and assign a `1` to one of them and `0` to all others.
#
# These generated columns are sometimes called _dummy variables_, and we will use the [`pandas.get_dummies()`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html?highlight=get_dummies#pandas.get_dummies) function to perform this transformation. Run the code cell below to perform the preprocessing routine discussed in this section.
# +
def preprocess_features(X):
''' Preprocesses the student data and converts non-numeric binary variables into
binary (0/1) variables. Converts categorical variables into dummy variables. '''
# Initialize new output DataFrame
output = pd.DataFrame(index = X.index)
# Investigate each feature column for the data
for col, col_data in X.iteritems():
# If data type is non-numeric, replace all yes/no values with 1/0
if col_data.dtype == object:
col_data = col_data.replace(['yes', 'no'], [1, 0])
# If data type is categorical, convert to dummy variables
if col_data.dtype == object:
# Example: 'school' => 'school_GP' and 'school_MS'
col_data = pd.get_dummies(col_data, prefix = col)
# Collect the revised columns
output = output.join(col_data)
return output
X_all = preprocess_features(X_all)
print "Processed feature columns ({} total features):\n{}".format(len(X_all.columns), list(X_all.columns))
# -
# ### Implementation: Training and Testing Data Split
# So far, we have converted all _categorical_ features into numeric values. For the next step, we split the data (both features and corresponding labels) into training and test sets. In the following code cell below, you will need to implement the following:
# - Randomly shuffle and split the data (`X_all`, `y_all`) into training and testing subsets.
# - Use 300 training points (approximately 75%) and 95 testing points (approximately 25%).
# - Set a `random_state` for the function(s) you use, if provided.
# - Store the results in `X_train`, `X_test`, `y_train`, and `y_test`.
# +
# TODO: Import any additional functionality you may need here
from sklearn.cross_validation import train_test_split
# TODO: Set the number of training points
num_train = 300
# Set the number of testing points
num_test = X_all.shape[0] - num_train
# TODO: Shuffle and split the dataset into the number of training and testing points above
X_train, X_test, y_train, y_test = train_test_split(
X_all,
y_all,
train_size=num_train,
test_size=num_test,
random_state=42)
# Show the results of the split
print "Training set has {} samples.".format(X_train.shape[0])
print "Testing set has {} samples.".format(X_test.shape[0])
# -
# ## Training and Evaluating Models
# In this section, you will choose 3 supervised learning models that are appropriate for this problem and available in `scikit-learn`. You will first discuss the reasoning behind choosing these three models by considering what you know about the data and each model's strengths and weaknesses. You will then fit the model to varying sizes of training data (100 data points, 200 data points, and 300 data points) and measure the F<sub>1</sub> score. You will need to produce three tables (one for each model) that shows the training set size, training time, prediction time, F<sub>1</sub> score on the training set, and F<sub>1</sub> score on the testing set.
#
# **The following supervised learning models are currently available in** [`scikit-learn`](http://scikit-learn.org/stable/supervised_learning.html) **that you may choose from:**
# - Gaussian Naive Bayes (GaussianNB)
# - Decision Trees
# - Ensemble Methods (Bagging, AdaBoost, Random Forest, Gradient Boosting)
# - K-Nearest Neighbors (KNeighbors)
# - Stochastic Gradient Descent (SGDC)
# - Support Vector Machines (SVM)
# - Logistic Regression
# ### Question 2 - Model Application
# *List three supervised learning models that are appropriate for this problem. For each model chosen*
# - Describe one real-world application in industry where the model can be applied. *(You may need to do a small bit of research for this — give references!)*
# - What are the strengths of the model; when does it perform well?
# - What are the weaknesses of the model; when does it perform poorly?
# - What makes this model a good candidate for the problem, given what you know about the data?
# **Answer: ** The models that I chose for the problem are: Support Vector Machines (SVM), Decision Trees(DT) and Naive Bayes (NB).I chose these models based on their success in the scientific literature as being trustworthy for classification problems and also for small data sets with good performance.
# ### 1. Support Vector Machines
# - General Applications: Highly competitive performance in numerous real-world applications, such as bioinformatics, text mining, face recognition, and image processing.
# - Strengths: SVM is very effective in high-dimensional spaces, and in situations when we have a non-linear separation problem. With SVM, we have the possibility to apply new kernels which increases the flexibility for our decision boundaries, leading to a better classification performance.
# - Weaknesses: One major disadvantage of the SVM is the choice of the kernel and also it is somewhat slower as compared to some other models like Decision trees or Naive bayes.
# - Why I chose this model: I chose this model because we are working with a small number of data samples and SVMs work well with small data sets and applying different kernels provide flexibility for our decision boundaries thus leading to a better classification performance.
#
# ### 2. Decision Trees
# - General Applications: Decision trees have been widely used in the field of medical research and practice. Recent uses of automatic induction of decision trees can be found in diagnosis, cardiology, psychiatry, gastroenterology, for detecting microcalcifications in mammography, to analyze Sudden Infant Death(SID) syndrome and for diagnosing thyroid disorders.
# - Strengths: The advantages of a decision trees are that nonlinear relationships between parameters do not affect our performance metrics and they give us faster prediction as compared to some other models like SVMs.
# - Weaknesses: They do not work well if we have smooth boundaries. i.e they work best when we have discontinuous piece wise constant model. If we truly have a linear target function, decision trees are not the best.
# - Why I chose this model: I chose this model because the decision tree provides a good performance metric for non-linear data. So, if our student data turns out to be non-linear, then decision trees would prove to be a better option.
#
# ### 3. Naive Bayes
# - General Applications: Naive Bayes methods can be used to mark an email as spam or not spam, to check a piece of text expressing positive emotions, or negative emotions and also in face recognition softwares.
# - Strengths: The advantages of naive bayes is that a NB classifier will converge quicker than discriminative models like logistic regression, so we need less training data.
# - Weaknesses: The same conditional independence assumption can be a disadvantage when we have no occurrences of a class label and a feature value together, what will give us a zero frequency-based value probability that affects any posterior probability estimate.
# - Why I chose this model: I chose this model because if the NB conditional independence assumption actually holds, the Naive Bayes classifier will converge quicker than discriminative models like logistic regression, so we'll need less training data which is in accordance with the number of data samples given to us.
# ### Setup
# Run the code cell below to initialize three helper functions which you can use for training and testing the three supervised learning models you've chosen above. The functions are as follows:
# - `train_classifier` - takes as input a classifier and training data and fits the classifier to the data.
# - `predict_labels` - takes as input a fit classifier, features, and a target labeling and makes predictions using the F<sub>1</sub> score.
# - `train_predict` - takes as input a classifier, and the training and testing data, and performs `train_clasifier` and `predict_labels`.
# - This function will report the F<sub>1</sub> score for both the training and testing data separately.
# +
def train_classifier(clf, X_train, y_train):
''' Fits a classifier to the training data. '''
# Start the clock, train the classifier, then stop the clock
start = time()
clf.fit(X_train, y_train)
end = time()
# Print the results
print "Trained model in {:.4f} seconds".format(end - start)
def predict_labels(clf, features, target):
''' Makes predictions using a fit classifier based on F1 score. '''
# Start the clock, make predictions, then stop the clock
start = time()
y_pred = clf.predict(features)
end = time()
# Print and return results
print "Made predictions in {:.4f} seconds.".format(end - start)
return f1_score(target.values, y_pred, pos_label='yes')
def train_predict(clf, X_train, y_train, X_test, y_test):
''' Train and predict using a classifer based on F1 score. '''
# Indicate the classifier and the training set size
print "Training a {} using a training set size of {}. . .".format(clf.__class__.__name__, len(X_train))
# Train the classifier
train_classifier(clf, X_train, y_train)
# Print the results of prediction for both training and testing
print "F1 score for training set: {:.4f}.".format(predict_labels(clf, X_train, y_train))
print "F1 score for test set: {:.4f}.\n\n".format(predict_labels(clf, X_test, y_test))
# -
# ### Implementation: Model Performance Metrics
# With the predefined functions above, you will now import the three supervised learning models of your choice and run the `train_predict` function for each one. Remember that you will need to train and predict on each classifier for three different training set sizes: 100, 200, and 300. Hence, you should expect to have 9 different outputs below — 3 for each model using the varying training set sizes. In the following code cell, you will need to implement the following:
# - Import the three supervised learning models you've discussed in the previous section.
# - Initialize the three models and store them in `clf_A`, `clf_B`, and `clf_C`.
# - Use a `random_state` for each model you use, if provided.
# - **Note:** Use the default settings for each model — you will tune one specific model in a later section.
# - Create the different training set sizes to be used to train each model.
# - *Do not reshuffle and resplit the data! The new training points should be drawn from `X_train` and `y_train`.*
# - Fit each model with each training set size and make predictions on the test set (9 in total).
# **Note:** Three tables are provided after the following code cell which can be used to store your results.
# +
# TODO: Import the three supervised learning models from sklearn
from sklearn import tree
from sklearn import svm
from sklearn.naive_bayes import GaussianNB
# TODO: Initialize the three models
clf_A = tree.DecisionTreeClassifier(random_state=42)
clf_B = svm.SVC(random_state=42)
clf_C = GaussianNB()
# TODO: Execute the 'train_predict' function for each classifier and each training set size
for size in [100, 200, 300]:
train_predict(clf_A, X_train[:size], y_train[:size], X_test, y_test)
for size in [100, 200, 300]:
train_predict(clf_B, X_train[:size], y_train[:size], X_test, y_test)
for size in [100, 200, 300]:
train_predict(clf_C, X_train[:size], y_train[:size], X_test, y_test)
# -
# ### Tabular Results
# Edit the cell below to see how a table can be designed in [Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet#tables). You can record your results from above in the tables provided.
# ** Classifer 1 - Decision Trees**
#
# | Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
# | :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
# | 100 | 0.0020 | 0.0000 | 1.0000 | 0.6552 |
# | 200 | 0.0020 | 0.0010 | 1.0000 | 0.7500 |
# | 300 | 0.0030 | 0.0010 | 1.0000 | 0.6613 |
#
# ** Classifer 2 - SVM**
#
# | Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
# | :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
# | 100 | 0.0060 | 0.0030 | 0.8777 | 0.7746 |
# | 200 | 0.0070 | 0.0040 | 0.8679 | 0.7815 |
# | 300 | 0.0110 | 0.0080 | 0.8761 | 0.7838 |
#
# ** Classifer 3 - Naive Bayes**
#
# | Training Set Size | Training Time | Prediction Time (test) | F1 Score (train) | F1 Score (test) |
# | :---------------: | :---------------------: | :--------------------: | :--------------: | :-------------: |
# | 100 | 0.0010 | 0.0010 | 0.8467 | 0.8029 |
# | 200 | 0.0020 | 0.0000 | 0.8406 | 0.7244 |
# | 300 | 0.0020 | 0.0000 | 0.8038 | 0.7634 |
# ## Choosing the Best Model
# In this final section, you will choose from the three supervised learning models the *best* model to use on the student data. You will then perform a grid search optimization for the model over the entire training set (`X_train` and `y_train`) by tuning at least one parameter to improve upon the untuned model's F<sub>1</sub> score.
# ### Question 3 - Choosing the Best Model
# *Based on the experiments you performed earlier, in one to two paragraphs, explain to the board of supervisors what single model you chose as the best model. Which model is generally the most appropriate based on the available data, limited resources, cost, and performance?*
# **Answer: ** From the results of the three models evaluated, and from the mean of their metrics results, for the F1 score, the method that had the highest prediction rate on the test set was the Support Vector Machines with an average F1 score of (0.7799) when compared with the average of the other two candidate models Decision Trees (0.6589) and Naive Bayes (0.7635).Although the training and prediction performance is a bit slower as compared to other methods,we can choose SVM as the best model for this case as with a better selection of features SVM compensates for its slower performance.A reassessment of the features can reduce their number to only those features that are really important to the prediction process, thus increasing it's performance.We can see from the performance metrics that even for 200 data samples the SVM model gives better performance metric on test data(0.7815) as compared to the rest two thus revealing that SVM is a better choice when we have a limited number of available data.
# ### Question 4 - Model in Layman's Terms
# *In one to two paragraphs, explain to the board of directors in layman's terms how the final model chosen is supposed to work. Be sure that you are describing the major qualities of the model, such as how the model is trained and how the model makes a prediction. Avoid using advanced mathematical or technical jargon, such as describing equations or discussing the algorithm implementation.*
# **Answer: ** The model chosen to classify students likely to suffer intervention or not, is the Support Vector Machines(SVMs). The model uses the data from the previous students (age, sex, family, etc.) to make predictions on the new students' data. We can understand the working of SVMs as let's suppose we try to fit a sheet of paper in between two clouds of points, the margin could then be described as us trying to place this sheet of paper as dead center between the two clouds of points as possible (so as to avoid touching either cloud).Also we can use different kernel trick(to deal with non-linearity) which can be described as the similarity measure by which we determine whether two points are close or far.So,this model will undergo a training process with the data of these students and will create a margin between a set of students who failed and those who managed to pass.After establishing this margin that separates our class well for the model, we introduce the model to the new data from the newly arrived students. So, the model will be able to predict depending on the side of the margin the student falls if he or she would pass or not.
# ### Implementation: Model Tuning
# Fine tune the chosen model. Use grid search (`GridSearchCV`) with at least one important parameter tuned with at least 3 different values. You will need to use the entire training set for this. In the code cell below, you will need to implement the following:
# - Import [`sklearn.grid_search.GridSearchCV`](http://scikit-learn.org/0.17/modules/generated/sklearn.grid_search.GridSearchCV.html) and [`sklearn.metrics.make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html).
# - Create a dictionary of parameters you wish to tune for the chosen model.
# - Example: `parameters = {'parameter' : [list of values]}`.
# - Initialize the classifier you've chosen and store it in `clf`.
# - Create the F<sub>1</sub> scoring function using `make_scorer` and store it in `f1_scorer`.
# - Set the `pos_label` parameter to the correct value!
# - Perform grid search on the classifier `clf` using `f1_scorer` as the scoring method, and store it in `grid_obj`.
# - Fit the grid search object to the training data (`X_train`, `y_train`), and store it in `grid_obj`.
# +
# TODO: Import 'GridSearchCV' and 'make_scorer'
from sklearn import svm, grid_search, datasets
from sklearn.metrics import make_scorer
from sklearn.svm import SVC
# TODO: Create the parameters list you wish to tune
parameters = [{'C': [1, 10, 1000],
'kernel': ['linear']},
{'C': [1, 10, 1000],
'gamma': [0.001, 0.0001],
'kernel': ['rbf']}]
# TODO: Initialize the classifier
clf = SVC()
# TODO: Make an f1 scoring function using 'make_scorer'
f1_scorer = make_scorer(f1_score, pos_label='yes')
# TODO: Perform grid search on the classifier using the f1_scorer as the scoring method
grid_obj = grid_search.GridSearchCV(clf,param_grid = parameters, scoring = f1_scorer)
# TODO: Fit the grid search object to the training data and find the optimal parameters
grid_obj.fit(X_train, y_train)
# Get the estimator
clf = grid_obj.best_estimator_
# Report the final F1 score for training and testing after parameter tuning
print "Tuned model has a training F1 score of {:.4f}.".format(predict_labels(clf, X_train, y_train))
print "Tuned model has a testing F1 score of {:.4f}.".format(predict_labels(clf, X_test, y_test))
# -
# ### Question 5 - Final F<sub>1</sub> Score
# *What is the final model's F<sub>1</sub> score for training and testing? How does that score compare to the untuned model?*
# **Answer: ** The final model's F1 score for training set is **0.8423** and for testing set is **0.7838** which is better as compared to the mean of the F1 score for the testing set(0.7799) and training set(0.8739) on untuned model.Therefore, tuning improved the F1 score, but by only 0.008 for testing set and 0.06 for training set.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
# **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| student_intervention/student_intervention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hoezithet
# language: python
# name: hoezithet
# ---
import bokeh
from bokeh.plotting import figure, show
from bokeh.io import output_notebook, save
from bokeh.embed import json_item
import json
from pathlib import Path
output_notebook()
# +
import numpy as np
from hoezithet import graphs
from bokeh.models import Label, Arrow, NormalHead, HoverTool, CustomJSHover
import json
def f(x): return -x**2 + 9
xs = np.arange(-10, 10, 0.1)
ys = [f(x) for x in xs]
p = graphs.get_plot()
p.title.text = 'Tekens van functiewaarden'
del p.tools[0] # Don't show hover;
p.quad(-12, -3, 0, -12, fill_color=graphs.RED, fill_alpha=0.5, line_alpha=0, name='neg')
p.quad(-3, 3, 12, 0, fill_color=graphs.GREEN, fill_alpha=0.5, line_alpha=0, name='pos')
p.quad(3, 12, 0, -12, fill_color=graphs.RED, fill_alpha=0.5, line_alpha=0, name='neg')
def get_tooltips(text: str):
return ('<div style="font-family: \'Quicksand\'; font-size: 14pt;'
'color:#555555;">'
f'{text}'
'</div>')
p.add_tools(HoverTool(tooltips=get_tooltips(f'<b><span style="color: {graphs.RED}">Negatieve</span></b> functiewaarden'), names=['neg'], mode='vline'))
p.add_tools(HoverTool(tooltips=get_tooltips(f'<b><span style="color: {graphs.GREEN}">Positieve</span></b> functiewaarden'),
names=['pos'],
mode='vline',
attachment='below'))
p.add_tools(HoverTool(tooltips=get_tooltips(f'Functiewaarde <b><span style="color: {graphs.BLUE}">nul</span></b>'), names=['zero']))
p.line(xs, ys, line_color=graphs.BLUE, line_width=5)
p.circle([-3, 3], [0, 0], radius=.1, line_width=10, color=graphs.BLUE, name='zero')
# show(p)
item = json.dumps(json_item(p))
Path('plt/tekenschema.json').write_text(item)
# -
| content/lessen/wiskunde/functies/tekenschema/tekenschema.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## PolynomialFeatures調査
# - 動作を調べ、ラベル付けを行う
import pandas as pd
import numpy as np
from sklearn.preprocessing import PolynomialFeatures
import sympy
# ## ダミーデータ作り
# 1番目からn番目までの素数をnp.arrayで返す関数
def primes(n=0):
ret_primes = list()
for i in np.arange(n):
ret_primes.append(sympy.prime(i+1))
return ret_primes
# +
# コラム名リストからコラムに対してユニークな素数が割り付けられたデータフレームを返す関数
def generate_df_prime_from_column(columns_original = np.array(["a","b","c"])):
data_original = np.array(primes(len(columns_original)))
return pd.DataFrame(data=data_original[np.newaxis,:],columns=columns_original,index=["original"])
# テスト
display(generate_df_prime_from_column())
# -
# ## PolynomialFeaturesの動作を調べる
# ### PolynomialFeatures
# - degree : integer
# The degree of the polynomial features. Default = 2.
# - interaction_only : boolean, default = False
# If true, only interaction features are produced: features that are products of at most degree distinct input features (so not x[1] ** 2, x[0] * x[2] ** 3, etc.).
# - include_bias : boolean
# If True (default), then include a bias column, the feature in which all polynomial powers are zero (i.e. a column of ones - acts as an intercept term in a linear model).
# - order : str in {‘C’, ‘F’}, default ‘C’
# Order of output array in the dense case. ‘F’ order is faster to compute, but may slow down subsequent estimators.
#
# from https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html
# ### データフレームとしてPolynomialFeatureの結果を返す関数
# コラム名もa×bなど、何と何を掛けたものであるかが分かるようにしている
def investigate_PolynomialFeatures(poly=PolynomialFeatures(2),columns_from=["a","b","c"],power=False):
df_from = generate_df_prime_from_column(columns_from)
columns_from = df_from.columns
data_from = df_from.values
data_poly = poly.fit_transform(df_from)
# columnをもう一度作り直す
columns_poly = list()
for i in np.arange(data_poly.shape[1]):
if (data_poly[0][i] == 1):
columns_poly.append("bias")
else:
prime_dict=sympy.factorint(data_poly[0][i])
keys = list(prime_dict.keys())
column_str = ""
if power:
# 累乗で書ける部分は累乗で書く(例:a^2)
for j in np.arange(len(keys)):
column_str += columns_from[list(data_from[0]).index(keys[j])]
if prime_dict[keys[j]] > 1:
column_str += "^" + str(prime_dict[keys[j]])
if (j < len(keys)-1):
column_str += "×"
else:
# 単純に×で項目をつなげていく(例:a×a×b)
for j in np.arange(len(keys)):
for k in np.arange(prime_dict[keys[j]]):
column_str += columns_from[list(data_from[0]).index(keys[j])]
if (j < len(keys)-1) | (k < prime_dict[keys[j]]-1):
column_str += "×"
columns_poly.append(column_str)
return pd.DataFrame(data=data_poly,columns=columns_poly,index=["poly"])
# ### 調査結果
# - バイアス→係数1個→係数2個→・・・の順
# - 係数2個の中では、a×(a→b→c)→b×(b→c)→c×cの順
# - interaction_only = True (デフォルトはFalse)の時は同じ係数が2個以上登場しないもののみとなる
# +
print("degree=2のとき")
display(investigate_PolynomialFeatures())
print("")
print("degree=2, interaction_only=Trueのとき")
display(investigate_PolynomialFeatures(poly=PolynomialFeatures(degree=2,interaction_only=True)))
print("")
print("degree=2, interaction_only=True, include_bias=Falseのとき")
display(investigate_PolynomialFeatures(poly=PolynomialFeatures(degree=2,interaction_only=True,include_bias=False)))
print("")
print("degree=3, interaction_only=False, include_bias=Falseのとき")
display(investigate_PolynomialFeatures(poly=PolynomialFeatures(degree=3,include_bias=False),power=True))
print("")
print("degree=3, interaction_only=True, include_bias=Falseのとき")
display(investigate_PolynomialFeatures(poly=PolynomialFeatures(degree=3,interaction_only=True,include_bias=False)))
# -
| 001_poly/PolynominalFeatures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is an introduction to the fastbt framework.
#
# Fastbt is built upon the assumption that you enter and hold all your positions for a specific time period and exit at the end of the period or when stop loss is triggered. For more information, see the philosophy and the rationale pages.
#
# I would take a top-down approach where I would run a strategy with the default parameters and then adjust the parameters one by one. You could always inspect the results by exporting them to a Excel file.
#
# +
# Import the necessary libraries
import pandas as pd
from fastbt.rapid import backtest, metrics
# Option to display all the columns
pd.options.display.max_columns = 50
# Load the data from a csv file
# We would work with this data throughout this notebook
df = pd.read_csv('data/data.csv', parse_dates=['timestamp'])
# -
# > **fastbt** requires only a dataframe to throw some results. You could get started with just one line of code
result = backtest(data=df)
# So, what did it do? Let's see the results and the metrics
print(metrics(result))
result.head()
# You run a backtest with the default parameters.
#
# By default, the strategy buys the top 5 securities with the lowest price at the open price and sells them at the same price with a capital of 100000. This looks dumb but lets look at the result dataframe. In addition to the columns in the original dataframe, it has added a few more columns. The important ones to look out for are
#
# * price - price at which the position is to be entered
# * stop_loss - price at which stop loss would be triggered
# * buy - the actual buy price
# * sell - the actual sell price
# * qty - the quantity of securities bought
#
# fastbt always returns a dataframe with these columns. qty is calculated by dividing the total capital by the number of securities (all securities are equally weighted for this purpose) and by the price. Since we have specified no stop loss securities are bought and sold at the same price.
#
# Now, let's try adding a stop loss and see the results. Stop loss is the percentage to be kept as stop loss from the price. Just specify it as an integer and it would be converted into percentage. So if you need a 0.5 percentage stop loss, use `stop_loss=0.5`
# ### A note on the backtest function
#
# > **backtest** is a function with multiple arguments. You could look at the all the arguments and their default values by typing `backtest?` in your console.
#
# > Its recommended to call the backtest function with keyword arguments instead of positional arguments.
#
# ```python
# backtest(data=df, capital=100000, leverage=1)
# ```
result = backtest(data=df, stop_loss=3)
print(metrics(result))
result.head()
# Yeps! your strategy has resulted in a massive loss. Let's try to make a sense of how price is determined.
#
# Taking the first case **NTPC**, you expect to buy it at a price of 177.05 and exit the position with a stop loss of 3%, 171.75. Since the buy price is equal to open price, the buy order is considered executed. Since the stop loss of 171.75 is not hit during the period, you exit the position at the close price. See the rationale page for a more detailed explanation of how prices are determined.
#
# You know this is a BUY order since the stop loss is less than the price. It is a SELL order if the stop loss is greater than the price. So let's go short instead of long.
result = backtest(data=df, stop_loss=3, order='S')
metrics(result)
# **Hurray! you made a profit**
#
#
# Now, let's take a look at the result dataframe
result.head()
# ## A simple digression
# Notice that the results are asymmetric. The long results didn't replicate exactly as the short results. We made a loss of 12116 with the long strategy but we only made a profit of 5516 with the short strategy. So the strategies aren't exact mirror of each other. This is due to the fact that we introduced stop loss. Let's try without stop loss; you would due to this by making stop loss a big number.
# +
result_one = metrics(backtest(data=df, stop_loss=200, order='B'))
result_two = metrics(backtest(data=df, stop_loss=200, order='S'))
result_one['profit'], result_two['profit']
# -
# Now, let's change price. Price is simply a formula string based on existing columns in your original dataframe. You can use any column along with common mathematical and logical operators (all numexpr expressions are valid). **Your formula should just be a single expression and should not contain any functions**. The following are all valid formulas
#
# * open + 0.05
# * prevhigh * 1.01
# * (prevhigh+prevlow)/2
result = backtest(data=df, stop_loss=3,
price='(open+prevclose)/2', order='S')
print(metrics(result))
result.head()
# Now try adding a few conditions to filter down the results. Conditions are similar to the price string with the only difference being they must evaluate to a boolean expression. Also, conditions should be specified as a list of conditions even if there is a single conditions. Conditions are always AND to each other. So securities would be picked only if all the conditions are fulfilled.
#
# The following conditions are valid
# * open > 100
# * open > prevclose
# * open < prevhigh
#
# Conditions are always AND to each other. So securities would be picked only if all the conditions are fulfilled.
#
# Henceforth, I would use a dictionary for all the arguments
params = {
'data': df,
'stop_loss': 3,
'price': '(open+prevclose)/2',
'order': 'S',
'conditions': [
'open > 220'
]
}
result = backtest(**params)
print(metrics(result))
result.head()
# By default, 5 symbols are returned for each day and they are sorted by the price column in ascending order. Use the limit, sort_mode and sort_by arguments to change them. To sort in descending order, pass sort_mode equals False
# +
params = {
'data': df,
'stop_loss': 3,
'price': '(open+prevclose)/2',
'order': 'S',
'limit': 3,
'sort_mode': False,
'sort_by': 'prevdayret'
}
result = backtest(**params)
print(metrics(result))
result.head()
# -
# Impressive, we have started barebones and developed a profitable strategy in a few steps. Let's add commission and slippage to see what would be our net profit. You must pass commission and slippage as a percentage of price.
#
# Note
# > Commission is applied to both BUY and SELL while slippage is applied only on one side since one of the orders is considered a LIMIT order. To apply slippage for both BUY and SELL, double the slippage percentage.
params = {
'data': df,
'stop_loss': 3,
'price': '(open+prevclose)/2',
'order': 'S',
'limit': 3,
'sort_mode': False,
'sort_by': 'prevdayret',
'commission': 0.02,
'slippage': 0.03
}
result = backtest(**params)
print(metrics(result))
# The above is identical to
# ```python
# result = backtest(data=df, stop_loss=3, price='(open+prevclose)/2',
# order='S', commission=0.02, slippage=0.03,
# limit=3, sort_mode=False, sort_by='prevdayret')
# ```
# Yeps! Most of our profits are wiped away. So let's see a couple of charts
# %pylab inline
(result.groupby('timestamp').net_profit.sum().cumsum()+100000).plot()
result.groupby('timestamp').sum()[['profit', 'net_profit']].boxplot()
# The graphics are minimal. I recommend the excellent pyfolio package for visualizing your returns. pyfolio accepts only returns as its input along with timestamp. So to convert our dataframe, just groupby timestamp, sum the daily profits and divide by your capital.
#
# ```python
# result.groupby('timestamp').net_profit.sum()/capital
# ```
# Now, let's try a simple simulation by varying the stoploss with the above structure and plot the results.
# +
params = {
'data': df,
'stop_loss': 3,
'price': '(open+prevclose)/2',
'order': 'S',
'limit': 3,
'sort_mode': False,
'sort_by': 'prevdayret',
'commission': 0.02,
'slippage': 0.03
}
stoploss = [1,1.5,2,2.5,3,3.5,4,4.5,5]
result_array = []
for s in stoploss:
params.update({'stop_loss': s})
r = backtest(**params)
m = metrics(r)
m['stoploss'] = s
result_array.append(m)
pd.DataFrame(result_array).plot.bar(x='stoploss', y='net_profit',
title='Stop Loss Effect on Profit')
# -
# ## Notes
# I have left out a three important arguments; universe, columns and strategy.
#
# Universe is the stock universe to get symbols from if you are connecting to a SQL database. Pass a list of symbols, a sql alchemy connection string and the table name to fetch data from a database. You could also use the start and end dates to limit your queries. If you specify both data and the above arguments, data takes precedence and the rest are ignored.
#
# Columns are columns you could add to your dataframe in an expressive text format instead of adding them as code. The format is identical to the `batch_process` function of the `DataSource` object.
#
# You could pass a custom strategy altogether to suit your needs. In such a case, the data returned is upto the point of adding columns and applying conditions and these results are grouped by the timestamp column. You could then apply your strategy to this dataframe.
#
# For running bigger simulations, use the multi_args function from utils. Make sure you save your results by including them in your code in case of long running simulations.
#
# The backtest function is just a wrapper to other 5 functions that run behind the scenes.
| examples/Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# As we discovered in the [Introduction](Introduction.ipynb), HoloViews allows plotting a variety of data types. Here we will use the sample data module and load the pandas and dask hvPlot API:
import numpy as np
import hvplot.pandas # noqa
import hvplot.dask # noqa
# As we learned The hvPlot API closely mirrors the [Pandas plotting API](https://pandas.pydata.org/pandas-docs/stable/visualization.html), but instead of generating static images when used in a notebook, it uses HoloViews to generate either static or dynamically streaming Bokeh plots. Static plots can be used in any context, while streaming plots require a live [Jupyter notebook](http://jupyter.org), a deployed [Bokeh Server app](https://bokeh.pydata.org/en/latest/docs/user_guide/server.html), or a deployed [Panel](https://panel.pyviz.org) app.
#
# HoloViews provides an extensive, very rich set of objects along with a powerful set of operations to apply, as you can find out in the [HoloViews User Guide](http://holoviews.org/user_guide/index.html). But here we will focus on the most essential mechanisms needed to make your data visualizable, without having to worry about the mechanics going on behind the scenes.
#
# We will be focusing on two different datasets:
#
# - A small CSV file of US crime data, broken down by state
# - A larger Parquet-format file of airline flight data
#
# The ``hvplot.sample_data`` module makes these datasets Intake data catalogue, which we can load either using pandas:
# +
from hvplot.sample_data import us_crime, airline_flights
crime = us_crime.read()
print(type(crime))
crime.head()
# -
# Or using dask as a ``dask.DataFrame``:
flights = airline_flights.to_dask().persist()
print(type(flights))
flights.head()
# ## The plot interface
# The ``dask.dataframe.DataFrame.hvplot``, ``pandas.DataFrame.hvplot`` and ``intake.DataSource.plot`` interfaces (and Series equivalents) from HvPlot provide a powerful high-level API to generate complex plots. The ``.hvplot`` API can be called directly or used as a namespace to generate specific plot types.
# ### The plot method
# The most explicit way to use the plotting API is to specify the names of columns to plot on the ``x``- and ``y``-axis respectively:
crime.hvplot.line(x='Year', y='Violent Crime rate')
# As you'll see in more detail below, you can choose which kind of plot you want to use for the data:
crime.hvplot(x='Year', y='Violent Crime rate', kind='scatter')
# To group the data by one or more additional columns, specify an additional ``by`` variable. As an example here we will plot the departure delay ('depdelay') as a function of 'distance', grouping the data by the 'carrier'. There are many available carriers, so we will select only two of them so that the plot is readable:
flight_subset = flights[flights.carrier.isin([b'OH', b'F9'])]
flight_subset.hvplot(x='distance', y='depdelay', by='carrier', kind='scatter', alpha=0.2, persist=True)
# Here we have specified the `x` axis explicitly, which can be omitted if the Pandas index column is already the desired x axis. Similarly, here we specified the `y` axis; by default all of the non-index columns would be plotted (which would be a lot of data in this case). If you don't specify the 'y' axis, it will have a default label named 'value', but you can then provide a y axis label explicitly using the ``value_label`` option.
#
# Putting all of this together we will plot violent crime, robbery, and burglary rates on the y-axis, specifying 'Year' as the x, and relabel the y-axis to display the 'Rate'.
crime.hvplot(x='Year', y=['Violent Crime rate', 'Robbery rate', 'Burglary rate'],
value_label='Rate (per 100k people)')
# ### The hvplot namespace
# Instead of using the ``kind`` argument to the plot call, we can use the ``hvplot`` namespace, which lets us easily discover the range of plot types that are supported. Use tab completion to explore the available plot types:
#
# ```python
# crime.hvplot.<TAB>
# ```
#
# Plot types available include:
#
# * <a href="#Area">``.area()``</a>: Plots a area chart similar to a line chart except for filling the area under the curve and optionally stacking
# * <a href="#Bars">``.bar()``</a>: Plots a bar chart that can be stacked or grouped
# * <a href="#Bivariate">``.bivariate()``</a>: Plots 2D density of a set of points
# * <a href="#Box-Whisker-Plots">``.box()``</a>: Plots a box-whisker chart comparing the distribution of one or more variables
# * <a href="#HeatMap">``.heatmap()``</a>: Plots a heatmap to visualizing a variable across two independent dimensions
# * <a href="#HexBins">``.hexbins()``</a>: Plots hex bins
# * <a href="#Histogram">``.histogram()``</a>: Plots the distribution of one or histograms as a set of bins
# * <a href="#KDE">``.kde()``</a>: Plots the kernel density estimate of one or more variables.
# * <a href="#The-plot-method">``.line()``</a>: Plots a line chart (such as for a time series)
# * <a href="#Scatter">``.scatter()``</a>: Plots a scatter chart comparing two variables
# * <a href="#Step">``.step()``</a>: Plots a step chart akin to a line plot
# * <a href="#Tables">``.table()``</a>: Generates a SlickGrid DataTable
# * <a href="#Violin-Plots">``.violin()``</a>: Plots a violin plot comparing the distribution of one or more variables using the kernel density estimate
# #### Area
#
# Like most other plot types the ``area`` chart supports the three ways of defining a plot outlined above. An area chart is most useful when plotting multiple variables in a stacked chart. This can be achieve by specifying ``x``, ``y``, and ``by`` columns or using the ``columns`` and ``index``/``use_index`` (equivalent to ``x``) options:
crime.hvplot.area(x='Year', y=['Robbery', 'Aggravated assault'])
# We can also explicitly set ``stacked`` to False and define an ``alpha`` value to compare the values directly:
crime.hvplot.area(x='Year', y=['Aggravated assault', 'Robbery'], stacked=False, alpha=0.4)
# Another use for an area plot is to visualize the spread of a value. For instance using the flights dataset we may want to see the spread in mean delay values across carriers. For that purpose we compute the mean delay by day and carrier and then the min/max mean delay for across all carriers. Since the output of ``hvplot`` is just a regular holoviews object, we can use the overlay operator (\*) to place the plots on top of each other.
# +
delay_min_max = flights.groupby(['day', 'carrier'])['carrier_delay'].mean().groupby('day').agg([np.min, np.max])
delay_mean = flights.groupby('day')['carrier_delay'].mean()
delay_min_max.hvplot.area(x='day', y='amin', y2='amax', alpha=0.2) * delay_mean.hvplot()
# -
# #### Bars
#
# In the simplest case we can use ``.hvplot.bar`` to plot ``x`` against ``y``. We'll use ``rot=90`` to rotate the tick labels on the x-axis making the years easier to read:
crime.hvplot.bar(x='Year', y='Violent Crime rate', rot=90)
# If we want to compare multiple columns instead we can set ``y`` to a list of columns. Using the ``stacked`` option we can then compare the column values more easily:
crime.hvplot.bar(x='Year', y=['Violent crime total', 'Property crime total'],
stacked=True, rot=90, width=800, legend='top_left')
# #### Scatter
#
# The scatter plot supports many of the same features as the other chart types we have seen so far but can also be colored by another variable using the ``c`` option.
crime.hvplot.scatter(x='Violent Crime rate', y='Burglary rate', c='Year')
# Anytime that color is being used to represent a dimension, the ``cmap`` option can be used to control the colormap that is used to represent that dimension. Additionally, the colorbar can be disabled using ``colorbar=False``.
# #### Step
#
# A step chart is very similar to a line chart but instead of linearly interpolating between samples the step chart visualizes discrete steps. The point at which to step can be controlled via the ``where`` keyword allowing `'pre'`, `'mid'` (default) and `'post'` values:
crime.hvplot.step(x='Year', y=['Robbery', 'Aggravated assault'])
# #### HexBins
#
# You can create hexagonal bin plots with the ``hexbin`` method. Hexbin plots can be a useful alternative to scatter plots if your data are too dense to plot each point individually. Since these data are not regularly distributed, we'll use the ``logz`` option to map z-axis (color) to a log scale colorbar.
flights.hvplot.hexbin(x='airtime', y='arrdelay', width=600, height=500, logz=True)
# #### Bivariate
#
# You can create a 2D density plot with the ``bivariate`` method. Bivariate plots can be a useful alternative to scatter plots if your data are too dense to plot each point individually.
crime.hvplot.bivariate(x='Violent Crime rate', y='Burglary rate', width=600, height=500)
# #### HeatMap
#
# A ``HeatMap`` lets us view the relationship between three variables, so we specify the 'x' and 'y' variables and an additional 'C' variable. Additionally we can define a ``reduce_function`` that computes the values for each bin from the samples that fall into it. Here we plot the 'depdelay' (i.e. departure delay) for each day of the month and carrier in the dataset:
flights.compute().hvplot.heatmap(x='day', y='carrier', C='depdelay', reduce_function=np.mean, colorbar=True)
# #### Tables
#
# Unlike all other plot types, a table only supports one signature: either all columns are plotted, or a subset of columns can be selected by defining the ``columns`` explicitly:
crime.hvplot.table(columns=['Year', 'Population', 'Violent Crime rate'], width=400)
# ### Distributions
#
# Plotting distributions differs slightly from other plots since they plot only one variable in the simple case rather than plotting two or more variables against each other. Therefore when plotting these plot types no ``index`` or ``x`` value needs to be supplied. Instead:
#
# 1. Declare a single ``y`` variable, e.g. ``source.plot.hist(variable)``, or
# 2. Declare a ``y`` variable and ``by`` variable, e.g. ``source.plot.hist(variable, by='Group')``, or
# 3. Declare columns or plot all columns, e.g. ``source.plot.hist()`` or ``source.plot.hist(columns=['A', 'B', 'C'])``
#
# #### Histogram
#
# The Histogram is the simplest example of a distribution; often we simply plot the distribution of a single variable, in this case the 'Violent Crime rate'. Additionally we can define a range over which to compute the histogram and the number of bins using the ``bin_range`` and ``bins`` arguments respectively:
crime.hvplot.hist(y='Violent Crime rate')
# Or we can plot the distribution of multiple columns:
columns = ['Violent Crime rate', 'Property crime rate', 'Burglary rate']
crime.hvplot.hist(y=columns, bins=50, alpha=0.5, legend='top', height=400)
# We can also group the data by another variable. Here we'll use ``subplots`` to split each carrier out into its own plot:
flight_subset = flights[flights.carrier.isin([b'AA', b'US', b'OH'])]
flight_subset.hvplot.hist('depdelay', by='carrier', bins=20, bin_range=(-20, 100), width=300, subplots=True)
# #### KDE (density)
#
# You can also create density plots using ``hvplot.kde()`` or ``hvplot.density()``:
crime.hvplot.kde(y='Violent Crime rate')
# Comparing the distribution of multiple columns is also possible:
columns=['Violent Crime rate', 'Property crime rate', 'Burglary rate']
crime.hvplot.kde(y=columns, alpha=0.5, value_label='Rate', legend='top_right')
# The ``hvplot.kde`` also supports the ``by`` keyword:
flight_subset = flights[flights.carrier.isin([b'AA', b'US', b'OH'])]
flight_subset.hvplot.kde('depdelay', by='carrier', xlim=(-20, 70), width=300, subplots=True)
# #### Box-Whisker Plots
#
# Just like the other distribution-based plot types, the box-whisker plot supports plotting a single column:
crime.hvplot.box(y='Violent Crime rate')
# It also supports multiple columns and the same options as seen previously (``legend``, ``invert``, ``value_label``):
columns=['Burglary rate', 'Larceny-theft rate', 'Motor vehicle theft rate',
'Property crime rate', 'Violent Crime rate']
crime.hvplot.box(y=columns, group_label='Crime', legend=False, value_label='Rate (per 100k)', invert=True)
# Lastly, it also supports using the ``by`` keyword to split the data into multiple subsets:
flight_subset = flights[flights.carrier.isin([b'AA', b'US', b'OH'])]
flight_subset.hvplot.box('depdelay', by='carrier', ylim=(-10, 70))
# ## Composing Plots
#
# One of the core strengths of HoloViews is the ease of composing
# different plots. Individual plots can be composed using the ``*`` and
# ``+`` operators, which overlay and compose plots into layouts
# respectively. For more information on composing objects, see the
# HoloViews [User Guide](http://holoviews.org/user_guide/Composing_Elements.html).
#
# By using these operators we can combine multiple plots into composite plots. A simple example is overlaying two plot types:
crime.hvplot(x='Year', y='Violent Crime rate') * crime.hvplot.scatter(x='Year', y='Violent Crime rate', c='k')
# We can also lay out different plots and tables together:
(crime.hvplot.bar(x='Year', y='Violent Crime rate', rot=90, width=550) +
crime.hvplot.table(['Year', 'Population', 'Violent Crime rate'], width=420))
# ## Large data
#
# The previous examples summarized the fairly large airline dataset using statistical plot types that aggregate the data into a feasible subset for plotting. We can instead aggregate the data directly into the viewable image using [datashader](http://datashader.org), which provides a rendering of the entire set of raw data available (as far as the resolution of the screen allows). Here we plot the 'airtime' against the 'distance':
flights.hvplot.scatter(x='distance', y='airtime', datashade=True)
# ## Groupby
#
# Thanks to the ability of HoloViews to explore a parameter space with a set of widgets we can apply a groupby along a particular column or dimension. For example we can view the distribution of departure delays by carrier grouped by day, allowing the user to choose which day to display:
flights.hvplot.violin(y='depdelay', by='carrier', groupby='dayofweek', ylim=(-20, 60), height=500)
# This user guide merely provided an overview over the available plot types; to see a detailed description on how to customize plots see the [Customization](Customization.ipynb) user guide.
| examples/user_guide/Plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Description
# ## HR Analytics: Job Change of Data Scientists - Predict who will move to a new job
#
# ### Context and Content
# A company which is active in Big Data and Data Science wants to hire data scientists among people who successfully pass some courses which conduct by the company. Many people signup for their training. Company wants to know which of these candidates are really wants to work for the company after training or looking for a new employment because it helps to reduce the cost and time as well as the quality of training or planning the courses and categorization of candidates. Information related to demographics, education, experience are in hands from candidates signup and enrollment.
#
# This dataset designed to understand the factors that lead a person to leave current job for HR researches too. By model(s) that uses the current credentials,demographics,experience data you will predict the probability of a candidate to look for a new job or will work for the company, as well as interpreting affected factors on employee decision.
#
# ### Inspiration
#
# 1. Predict the probability of a candidate will work for the company
#
#
# 2. Interpret model(s) such a way that illustrate which features affect candidate decision
#
#
# ### Features
#
# - enrollee_id : Unique ID for candidate
#
# - city: City code
#
# - city_ development _index : Developement index of the city (scaled)
#
# - gender: Gender of candidate
#
# - relevent_experience: Relevant experience of candidate
#
# - enrolled_university: Type of University course enrolled if any
#
# - education_level: Education level of candidate
#
# - major_discipline :Education major discipline of candidate
#
# - experience: Candidate total experience in years
#
# - company_size: No of employees in current employer's company
#
# - company_type : Type of current employer
#
# - lastnewjob: Difference in years between previous job and current job
#
# - training_hours: training hours completed
#
# - target: 0 – Not looking for job change, 1 – Looking for a job change
#
# Reference:
# https://www.kaggle.com/arashnic/hr-analytics-job-change-of-data-scientists?select=aug_test.csv
import numpy as np
import pandas as pd
# train data
data = pd.read_csv("resources/aug_train.csv")
data
data.info()
data.describe()
# ## Observations:
#
# 1. Total Number Features = 13
# 2. Number of Categorical Features = 10
# 3. Number of Numerical Features = 3
# 4. Number of records = 19158
# 5. 75% of target is 0 which means data is imbalanced- More data on class 0 (not look for a job) than class 1 (look for a job)
# # Data Preprocessing
#
# ## Feature selection:
#
# - Eliminate insignificant features: enrollee_id
new_data = data.drop(columns = 'enrollee_id')
# ## Missing values:
#
# ### Methods applied to handle missing values:
#
# As all missing values in this data are categoical, we can either drop the corresponding records or treat missing data as just another category named Unknown. Note: there are also other ways like "Developing model to predict missing values" which we have not considered.
#
# If we drop all records that include NaN, more than half of the rows will be eliminated. So we decided to drop NaN values for some columns (that we think their impact in the outcome is more) and use Unknown category for the rest.
#
# We drop records with NaN values in the following features: education_level, major_discipline, experience, last_new_job
#
# And use Unknown category for the rest
#
new_data.isna().sum()
# +
# drop NaN values for some columns
data_wo_nan = new_data.dropna(subset=['education_level','major_discipline', 'experience', 'last_new_job'])
# Replace other NaN with Unknown value
data_wo_nan = data_wo_nan.replace(np.nan,'Unknown')
# check missing values again
data_wo_nan.isna().sum()
# -
# ## Categorical Columns:
# - check the unique values for each column so we can decide better how to convert them to numerical ones
#Select categorical columns
cat_columns = data_wo_nan.select_dtypes(include='object')
print("Unique values of city: \n",cat_columns.city.unique())
print("\nUnique values of gender: \n",cat_columns.gender.unique())
print("\nUnique values of relevent_experience: \n",cat_columns.relevent_experience.unique())
print("\nUnique values of enrolled_university: \n",cat_columns.enrolled_university.unique())
print("\nUnique values of education_level: \n",cat_columns.education_level.unique())
print("\nUnique values of major_discipline: \n",cat_columns.major_discipline.unique())
print("\nUnique values of experience: \n",cat_columns.experience.unique())
print("\nUnique values of company_size: \n",cat_columns.company_size.unique())
print("\nUnique values of company_type: \n",cat_columns.company_type.unique())
print("\nUnique values of last_new_job: \n",cat_columns.last_new_job.unique())
# ### observations
# - relevent_experience is a boolian column we can convert it to 0 for no experience and 1 for have experience
# - education_level and company_size are ordinal categories.
# - experience and last_new_job can be converted to numeric values directly.
# - company_type, enrolled_university, gender, major_discipline are nominal categories and we can use get_dummies for them.
# - We can also use one_hot_encoder for city feature which is also a nominal categoriy but the drawback is the number of unique city values are a lot and it will add a lot of features to our space if we use one_hot_encoder.
# +
proc_data = data_wo_nan.copy()
# relevent_experience replace with 0 and 1, 1 for having experience and 0 for no experience
proc_data['relevent_experience'] = proc_data['relevent_experience'].replace(['Has relevent experience','No relevent experience'],[1,0])
# manually assign ordinal numbers to education_level and company_size
# for graduate level I will give 1 and for master 2 and for phd 3. Graduate level can be equals to masters and phd but usually people with phd would not represent themselves as graduate.
# any graduate level certificate can be considered as graduate so I will assign a lower number to graduate than masters.
# for company_size unknown will get 0.
proc_data['education_level'] = proc_data['education_level'].replace(['Graduate','Masters','Phd'],[1,2,3])
proc_data['company_size'] = proc_data['company_size'].replace(['Unknown','<10', '10/49','50-99', '100-500','500-999','1000-4999','5000-9999','10000+'] ,range(0,9))
# convert experience and last_new_job to numeric values
proc_data['experience'] = proc_data['experience'].str.replace('>','').str.replace('<','')
proc_data['experience'] = pd.to_numeric(proc_data['experience'])
proc_data['last_new_job'] = proc_data['last_new_job'].str.replace('>','')
proc_data['last_new_job'] = proc_data['last_new_job'].replace('never',0)
proc_data['last_new_job'] = pd.to_numeric(proc_data['last_new_job'])
final_data = pd.get_dummies(proc_data, columns = ['company_type', 'enrolled_university', 'gender', 'major_discipline','city'])
final_data
# -
final_data.info()
# ## Preprocessing function
# preprocessing function
def preprocessing_data(df: pd.DataFrame):
data = df.copy()
# drop NaN values for some columns
data = data.dropna(subset=['education_level','major_discipline', 'experience', 'last_new_job'])
# Replace other NaN with Unknown value
data = data.replace(np.nan,'Unknown')
# relevent_experience replace with 0 and 1, 1 for having experience and 0 for no experience
data['relevent_experience'] = data['relevent_experience'].replace(['Has relevent experience','No relevent experience'],[1,0])
# manually assign ordinal numbers to education_level and company_size
# for graduate level I will give 1 and for master 2 and for phd 3. Graduate level can be equals to masters and phd but usually people with phd would not represent themselves as graduate.
# any graduate level certificate can be considered as graduate so I will assign a lower number to graduate than masters.
# for company_size unknown will get 0.
data['education_level'] = data['education_level'].replace(['Graduate','Masters','Phd'],[1,2,3])
data['company_size'] = data['company_size'].replace(['Unknown','<10', '10/49','50-99', '100-500','500-999','1000-4999','5000-9999','10000+'] ,range(0,9))
# convert experience and last_new_job to numeric values
data['experience'] = data['experience'].str.replace('>','').str.replace('<','')
data['experience'] = pd.to_numeric(data['experience'])
data['last_new_job'] = data['last_new_job'].str.replace('>','')
data['last_new_job'] = data['last_new_job'].replace('never',0)
data['last_new_job'] = pd.to_numeric(data['last_new_job'])
data = pd.get_dummies(proc_data, columns = ['company_type', 'enrolled_university', 'gender', 'major_discipline','city'])
return(data)
raw_data = pd.read_csv("resources/aug_train.csv")
processed_data = preprocessing_data(raw_data)
processed_data
| DataPreprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Imports, utils and setup
from IPython.display import display
import pandas as pd
import plotly.express as px
from transformers import pipeline
# +
def sum_model_trainable_parameters(pytorch_model) -> int:
return sum(p.numel() for p in pytorch_model.parameters())
def show_model_summary(model_name, model, model_size_mb):
params_count = sum_model_trainable_parameters(model)
return pd.Series(dict(
parameters=f'{params_count:,}',
size=f'{model_size_mb} MB'
), name=model_name).to_frame()
# -
# # Generating fluent text
# ## Preparation
#
# Model site: https://huggingface.co/gpt2
# +
from transformers import pipeline
text_generation_model_name = 'gpt2'
text_generation_pipeline = pipeline('text-generation',
model=text_generation_model_name)
text_generation_model_size_mb = 523
# -
show_model_summary(text_generation_model_name,
text_generation_pipeline.model,
text_generation_model_size_mb)
# ## Demo
# +
from transformers import pipeline, set_seed
from pprint import pprint
set_seed(1)
text_generation_pipeline("Once upon the time",
max_length=25,
num_return_sequences=2)
# +
set_seed(2)
text_generation_pipeline("Once upon the time",
max_length=25,
num_return_sequences=2)
# -
# # Question answering
#
# Model site: https://huggingface.co/deepset/roberta-base-squad2
#
# ## Preparation
qa_model_name = 'deepset/roberta-base-squad2'
qa_pipeline = pipeline('question-answering', qa_model_name)
qa_model_size_mb = 473
show_model_summary(qa_model_name, qa_pipeline.model, qa_model_size_mb)
def answer_question(question, context, qa_pipeline):
return qa_pipeline(dict(
question=question,
context=context
))
australia_context = """Australia, officially the Commonwealth of Australia,
is a sovereign country comprising the mainland of the Australian continent,
the island of Tasmania, and numerous smaller islands.
It is the largest country by area in Oceania and the world's sixth-largest country.
Australia's population of nearly 26 million, in an area of 7,617,930 square
kilometres (2,941,300 sq mi),[14] is highly urbanised and heavily concentrated on
the eastern seaboard. Canberra is the nation's capital, while the largest city is
Sydney, and other major metropolitan areas include Melbourne, Brisbane, Perth,
and Adelaide."""
question_variants = [
'How many citizens Australia has?',
'How many people live in Australia?',
'For how many people is Australia home?',
]
# ### Demo
for question in question_variants:
result = answer_question(question=question,
context=australia_context,
qa_pipeline=qa_pipeline)
answer = result['answer']
confidence_score = round(result['score']*100)
print(f'{question} {answer}. Confidence: {confidence_score}%')
# #### Answer is a string between 'start' and 'end' positions in a question
result
australia_context[ result['start'] : result['end'] ]
# # Summarization
# ## Generate summary
# ### Preparation
summarize_model_name="facebook/bart-large-cnn"
summarize_pipeline = pipeline("summarization", model=summarize_model_name)
summarize_model_size_mb=1510
show_model_summary(summarize_model_name, summarize_pipeline.model, summarize_model_size_mb)
# ### Demo
australia_context = """Australia, officially the Commonwealth of Australia,
is a sovereign country comprising the mainland of the Australian continent,
the island of a
Tasmania, and numerous smaller islands.
It is the largest country by area in Oceania and the world's sixth-largest country.
Australia's population of nearly 26 million, in an area of 7,617,930 square
kilometres (2,941,300 sq mi),[14] is highly urbanised and heavily concentrated on
the eastern seaboard. Canberra is the nation's capital, while the largest city is
Sydney, and other major metropolitan areas include Melbourne, Brisbane, Perth,
and Adelaide."""
summarize_pipeline(australia_context, max_length=50, min_length=30, do_sample=False)
# ## Generate title for article
#
# ### Preparation
headline_generator_model_name = "Michau/t5-base-en-generate-headline"
headline_generator_pipeline = pipeline('text2text-generation',
model=headline_generator_model_name)
headline_generator_model_size_mb = 850
show_model_summary(headline_generator_model_name,
headline_generator_pipeline.model,
headline_generator_model_size_mb)
# ### Demo
article = """
To make observations in the infrared spectrum, JWST must be kept under 50 K
(−223.2 °C; −369.7 °F); otherwise, infrared radiation from the telescope itself
would overwhelm its instruments. It therefore uses a large sunshield to block light
and heat from the Sun, Earth, and Moon, and its position near the Sun-Earth L2 keeps
all three bodies on the same side of the spacecraft at all times.[56] Its halo orbit
around the L2 point avoids the shadow of the Earth and Moon, maintaining a constant
environment for the sunshield and solar arrays.[54] The shielding maintains a stable
temperature for the structures on the dark side, which is critical to maintaining precise
alignment of the primary mirror segments in space.[18]. The five-layer sunshield,
each layer as thin as a human hair,[57] is constructed from Kapton E, a commercially
available polyimide film from DuPont, with membranes specially coated with aluminum on
both sides and a layer of doped silicon on the Sun-facing side of the two hottest layers
to reflect the Sun's heat back into space.[18] Accidental tears of the delicate film
structure during testing in 2018 were among the factors delaying the project.[58]
The sunshield was designed to be folded twelve times so that it fit within the Ariane
5 rocket's payload fairing, which is 4.57 m (15.0 ft) in diameter, and 16.19 m (53.1 ft)
long. The shield's fully deployed dimensions were planned as 14.162 m × 21.197 m
(46.46 ft × 69.54 ft). The sunshield was hand-assembled at ManTech (NeXolve) in
Huntsville, Alabama, before it was delivered to Northrop Grumman in Redondo Beach,
California, for testing.[59]. Because of the sunshield, JWST does not have an unlimited
field of regard at any given time. The telescope can see 40 percent of the sky from one
position and can see all of the sky over a period of six months,[60] the amount of time
it takes to complete half its orbit around the Sun."""
headline_generator_pipeline(article)
# # Text classification
# ## Sentiment analysis
# Model site: https://huggingface.co/distilbert-base-uncased-finetuned-sst-2-english
# ### Preparation
sentiment_model_name = 'distilbert-base-uncased-finetuned-sst-2-english'
sentiment_pipeline = pipeline('sentiment-analysis', model=sentiment_model_name)
sentiment_model_size_mb = 255
show_model_summary(sentiment_model_name, sentiment_pipeline.model, sentiment_model_size_mb)
# ### Demo
#
# Watch how well sacrasm is detected
sentiment_pipeline('I love being served hot food')
sentiment_pipeline('I love being served cold food')
# ## Emotion classification
# ### Preparation
emotion_model = 'bhadresh-savani/distilbert-base-uncased-emotion'
emotion_pipeline = pipeline('sentiment-analysis', model=emotion_model,
return_all_scores=True)
emotion_model_size_mb = 255
show_model_summary(emotion_model, emotion_pipeline.model,
emotion_model_size_mb)
# ### Demo
emotion_pipeline('Its something great being human.')
emotion_pipeline("I didn't expect that at all. It must be a joke...")
# ## Zero shot classification
# Classify by any label
# ### Preparation
# +
from transformers import pipeline
zero_shot_model_name = "facebook/bart-large-mnli"
zero_shot_classifier = pipeline("zero-shot-classification",
model=zero_shot_model_name)
zero_shot_model_size_mb = 1520
# -
show_model_summary(zero_shot_model_name, zero_shot_classifier.model, zero_shot_model_size_mb)
# ### Demo
# +
australia_description = """Australia, officially the Commonwealth of Australia,
is a sovereign country comprising the mainland of the Australian continent,
the island of Tasmania, and numerous smaller islands."""
gradient_description = """In vector calculus, the gradient of a scalar-valued differentiable function f of several
variables is the vector field (or vector-valued function) whose value at a point p is the
vector[a] whose components are the partial derivatives"""
plankton_description = """Plankton are the diverse collection of organisms found in water
(or air) that are unable to propel themselves against a current (or wind). The individual
organisms constituting plankton are called plankters."""
brave_new_world_description = """Brave New World is a dystopian social science fiction novel by
<NAME>, written in 1931 and published in 1932. Largely set in a futuristic World State, whose citizens are
environmentally engineered into an intelligence-based social hierarchy."""
classes = ['geography', 'math', 'biology', 'literature']
texts_to_classify = [australia_description, gradient_description,
plankton_description, brave_new_world_description]
zero_shot_predictions = zero_shot_classifier(texts_to_classify,
classes)
# +
import pandas as pd
index = []
scores = []
for prediction in zero_shot_predictions:
index.append(prediction['sequence'])
scores_rounded = [round(s,3) for s in prediction['scores']]
scores_dict = dict(zip(prediction['labels'], scores_rounded))
scores.append(scores_dict)
pd.DataFrame(scores, index=index, columns=classes)
# -
# ## Zero shot vs sentiment specific model
classify_sentiment_texts = [
'I love being served hot food',
'I love being served cold food'
]
sentiment_model_predictions = sentiment_pipeline(classify_sentiment_texts,
return_all_scores=True)
sentiment_model_predictions
labels = list(sentiment_pipeline.model.config.label2id.keys())
labels
zeroshot_model_predictions = zero_shot_classifier(classify_sentiment_texts, labels)
zeroshot_model_predictions
# # Conversational
# https://huggingface.co/microsoft/DialoGPT-small
# ## Preparation
conversational_model_model_name = 'microsoft/DialoGPT-large'
conversational_pipeline = pipeline('conversational', model=conversational_model_model_name)
conversational_model_size_mb = 1630
show_model_summary(conversational_model_model_name,
conversational_pipeline.model,
conversational_model_size_mb)
from transformers import Conversation
# ## Demo
conversation_1 = Conversation("Can I add fries?")
conversational_pipeline(conversation_1)
conversation_1.add_user_input("How much is it?")
conversational_pipeline(conversation_1)
conversation_1.add_user_input("Ok, so I will order two")
conversational_pipeline(conversation_1)
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 3 Practice Assessment - Mission Planning
# **This is an ungraded practice assignment**. In this notebook, you will be given an opportunity to implement Dijkstra's search algorithm on a road network in Berkeley, California. You will then modify that algorithm using a distance heuristic to perform A* search. You will then get a chance to compare your shortest path to the mapping library's path. If they match, congratulations!
#
# **In this notebook, you will:**
# * Implement Dijkstra's search algorithm on a road network graph.
# * Implement the A* search algorithm using a Euclidean heuristic on a road network graph.
#
# For most exercises, you are provided with a suggested outline. You are encouraged to diverge from the outline if you think there is a better, more efficient way to solve a problem.
# Launch the Jupyter Notebook to begin!
# We recommend that you refer to the solution only after you finish this practice exercise.
# This practice assessment will give you a chance to cement your knowledge that you gained through the Module 3 videos by giving you hands-on experience using real map data. In this assessment, we will be relying on the [OSMNX library](https://osmnx.readthedocs.io/en/stable/) to generate Python graphs from Open Street Map (OSM) data. These graphs will be represented using the [NetworkX library](https://networkx.github.io/documentation/stable/). Both of these links are to the documentation, which you will find useful in this assessment.
import osmnx as ox
import networkx as nx
import queue
import math
import priority_dict
# For this assessment, we're going to be focusing on planning in Berkeley, California, between the two nodes given below. After running the code up to and includeing the box below, you should see the output of the shortest path between the two points. Your goal is to get the same output yourself when you implement Dijkstra's and A*.
# +
map_graph = ox.graph_from_place('Berkeley, California', network_type='drive')
origin = ox.get_nearest_node(map_graph, (37.8743, -122.277))
destination = list(map_graph.nodes())[-1]
shortest_path = nx.shortest_path(map_graph, origin, destination, weight='length')
fig, ax = ox.plot_graph_route(map_graph, shortest_path)
# -
# ## Dijkstra's Search
# First, let's focus on Dijkstra's algorithm. As a refresher, we've included the pseudocode from the Module 3 lessons below.
#
# 
#
# This function will be implemented by you below in `dijkstras_search()`. We have included a helper function `get_path()` that will assist in retrieving the path from the dictionary of predecessors once the goal is found.
#
# To perform Dijkstra's search, we require a priority queue (or a min heap), which is defined as the `priority_dict` class. This class is accessed just as a standard dictionary is, except it orders the keys by their value. We can use the vertices as the keys to our priority queue, and their distance from the start as their value. For example, to set the distance of vertex `v` to the variable `dist`, we can do `open_queue[v] = dist`. To get the smallest value in the priority queue, we can use `priority_dict.pop_smallest()`. This returns a tuple of the vertex key and it's distance from the origin.
#
# The main input to the search is the `graph`, an OSMNX graph representation of the road network. The vertices are stored as keys, and as such the origin is given as input as `origin_key` and the goal is given as `goal_key`. To get the outgoing edges of a given vertex `u`, we can use `graph.out_edges([u], data=True)`. The return value of this is a list of tuples, each of which represent an outgoing edge. The second element of each tuple is the outgoing vertex at the other end of the edge. You can iterate over this list of tuples using:
#
# `for edge in graph.out_edges([u], data=True):`
#
# and can get the outgoing vertex of these edges by accessing the 2nd element of the tuple:
#
# `edge[1]`.
#
# To get the weight of this edge, you can access the data stored in the 3rd element of the tuple:
#
# `length = edge[2]['length']`.
#
# For more details, you can refer to the NetworkX documentation [here](https://networkx.github.io/documentation/networkx-2.3/reference/classes/generated/networkx.DiGraph.out_edges.html?highlight=out_edges#networkx.DiGraph.out_edges).
#
# Your goal now is to find the shortest path in the graph from the origin to the goal using Dijkstra's search. Make sure to store the optimal predecessors of each vertex in the `predecessors` dictionary, so you can retrieve the optimal path once you find the goal node in your search. Good luck!
# For a given graph, origin vertex key, and goal vertex key,
# computes the shortest path in the graph from the origin vertex
# to the goal vertex using Dijkstra's algorithm.
# Returns the shortest path as a list of vertex keys.
def dijkstras_search(origin_key, goal_key, graph):
# The priority queue of open vertices we've reached.
# Keys are the vertex keys, vals are the distances.
open_queue = priority_dict.priority_dict({})
# The dictionary of closed vertices we've processed.
closed_dict = {}
# The dictionary of predecessors for each vertex.
predecessors = {}
# Add the origin to the open queue.
open_queue[origin_key] = 0.0
# Iterate through the open queue, until we find the goal.
# Each time, perform a Dijkstra's update on the queue.
# TODO: Implement the Dijstra update loop.
goal_found = False
while (open_queue):
u, uCost = open_queue.pop_smallest()
if u == goal_key:
return get_path(origin_key, goal_key, predecessors)
for edge in graph.out_edges([u], data=True):
v = edge[1]
if v in closed_dict:
continue
uvCost = edge[2]['length']
cost = uCost + uvCost
if v in open_queue:
if cost < open_queue[v]:
open_queue[v] = cost
predecessors[v] = u
else:
open_queue[v] = cost
predecessors[v] = u
closed_dict[u] = uCost
# If we get through entire priority queue without finding the goal,
# something is wrong.
if not goal_found:
raise ValueError("Goal not found in search.")
# Construct the path from the predecessors dictionary.
return get_path(origin_key, goal_key, predecessors)
# This function follows the predecessor
# backpointers and generates the equivalent path from the
# origin as a list of vertex keys.
def get_path(origin_key, goal_key, predecessors):
key = goal_key
path = [goal_key]
while (key != origin_key):
key = predecessors[key]
path.insert(0, key)
return path
# Once these two functions have been implemented, run the box below to see if your output matches that of the library function above. If it doesn't, you've made a mistake with your implementation.
path = dijkstras_search(origin, destination, map_graph)
fig, ax = ox.plot_graph_route(map_graph, path)
# ## A* Search
# Next, we will use a distance heuristic to implement A* search for our map search problem. Since we are using real map data here, we will need to convert the data to a format which we can use for distance computation. Each data point has a latitude and longitude associated with it, which we then have to convert into (x, y, z) coordinates on the earth (which we will assume to be a sphere with radius 6371 km). We can then take the straight line distance between these two points as an approximation for the distance between them. Over small distances, this approximation is accurate. This is implemented in the `distance_heuristic()` function below.
# Computes the Euclidean distance between two vertices.
# Assume that the earth is a sphere with radius 6371 km.
def distance_heuristic(state_key, goal_key, node_data):
n1 = node_data[state_key]
n2 = node_data[goal_key]
# Get the longitude and latitude for each vertex.
long1 = n1['x']*math.pi/180.0
lat1 = n1['y']*math.pi/180.0
long2 = n2['x']*math.pi/180.0
lat2 = n2['y']*math.pi/180.0
# Use a spherical approximation of the earth for
# estimating the distance between two points.
r = 6371000
x1 = r*math.cos(lat1)*math.cos(long1)
y1 = r*math.cos(lat1)*math.sin(long1)
z1 = r*math.sin(lat1)
x2 = r*math.cos(lat2)*math.cos(long2)
y2 = r*math.cos(lat2)*math.sin(long2)
z2 = r*math.sin(lat2)
d = ((x2-x1)**2 + (y2-y1)**2 + (z2-z1)**2)**0.5
return d
# Now, we can use our distance heuristic to perform A* search on our map. As a refresher, we've included the A* pseudocode from Module 3 below.
# 
# This function will be implemented in the `a_star_search()` function below. As with Dijkstra's search, you should make use of the `get_path()` helper function above. As before, you should find the shortest path from the origin to the goal in the graph, but this time you should use A* with the distance heuristic given above. Good luck!
# For a given graph, origin vertex key, and goal vertex key,
# computes the shortest path in the graph from the origin vertex
# to the goal vertex using A* search.
# Returns the shortest path as a list of vertex keys.
def a_star_search(origin_key, goal_key, graph):
# The priority queue of open vertices we've reached.
# Keys are the vertex keys, vals are the accumulated
# distances plus the heuristic estimates of the distance
# to go.
open_queue = priority_dict.priority_dict({})
# The dictionary of closed vertices we've processed.
closed_dict = {}
# The dictionary of predecessors for each vertex.
predecessors = {}
# The dictionary that stores the best cost to reach each
# vertex found so far.
costs = {}
# Get the spatial data for each vertex as a dictionary.
node_data = graph.nodes(True)
# Add the origin to the open queue and the costs dictionary.
costs[origin_key] = 0.0
open_queue[origin_key] = distance_heuristic(origin_key, goal_key, node_data)
# Iterate through the open queue, until we find the goal.
# Each time, perform an A* update on the queue.
# TODO: Implement the A* update loop.
goal_found = False
while (open_queue):
u, _ = open_queue.pop_smallest()
uCost = costs[u]
if u == goal_key:
return get_path(origin_key, goal_key, predecessors)
for edge in graph.out_edges([u], data=True):
v = edge[1]
if v in closed_dict:
continue
uvCost = edge[2]['length']
cost = uCost + uvCost
h = distance_heuristic(v, goal_key, node_data)
if v in open_queue:
if cost + h < open_queue[v]:
open_queue[v] = cost + h
costs[v] = cost
predecessors[v] = u
else:
open_queue[v] = cost + h
costs[v] = cost
predecessors[v] = u
closed_dict[u] = uCost
# If we get through entire priority queue without finding the goal,
# something is wrong.
if not goal_found:
raise ValueError("Goal not found in search.")
# Construct the path from the predecessors dictionary.
return get_path(origin_key, goal_key, predecessors)
# Once this function has been implemented, run the box below to see if your output matches that of the library function at the start of the notebook. If it doesn't, you've made a mistake with your implementation.
path = a_star_search(origin, destination, map_graph)
fig, ax = ox.plot_graph_route(map_graph, path)
# Congratulations! You've now implemented two important mission planning algorithms on real map data.
| 4_motion_planning_for_self_driving_cars/Module3_Practice_Assessment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matching
#
# ## What is Regression Doing After All?
#
# As we've seen so far, regression does an amazing job at controlling for additional variables when we do a test vs control comparison. If we have independence, \\((Y_0, Y_1)\perp T | X\\), then regression can identify the ATE by controlling for X. The way regression does this is kind of magical. To get some intuition about it, let's remember the case when all variables X are dummy variables. If that is the case, regression partitions the data into the dummy cells and computes the mean difference between test and control. This difference in means keeps the Xs constant, since we are doing it in a fixed cell of X dummy. It is as if we were doing \\(E[Y|T=1] - E[Y|T=0] | X=x\\), where \\(x\\) is a dummy cell (all dummies set to 1, for example). Regression then combines the estimate in each of the cells to produce a final ATE. The way it does this is by applying weights to the cell proportional to the variance of the treatment on that group.
# +
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
from matplotlib import style
from matplotlib import pyplot as plt
import statsmodels.formula.api as smf
import graphviz as gr
# %matplotlib inline
style.use("fivethirtyeight")
# -
# To give an example, let's suppose I'm trying to estimate the effect of a drug and I have 6 men and 4 women. My response variable is days hospitalised and I hope my drug can lower that. On men, the true causal effect is -3, so the drug lowers the stay period by 3 days. On women, it is -2. To make matters more interesting, men are much more affected by this illness and stay longer at the hospital. They also get much more of the drug. Only 1 out of the 6 men does not get the drug. On the other hand, women are more resistant to this illness, so they stay less at the hospital. 50% of the women get the drug.
drug_example = pd.DataFrame(dict(
sex= ["M","M","M","M","M","M", "W","W","W","W"],
drug=[1,1,1,1,1,0, 1,0,1,0],
days=[5,5,5,5,5,8, 2,4,2,4]
))
# Note that simple comparison of treatment and control yields a negatively biased effect, that is, the drug seems less effective than it truly is. This is expected, since we've omitted the sex confounder. In this case, the estimated ATE is smaller than the true one because men get more of the drug and are more affected by the illness.
drug_example.query("drug==1")["days"].mean() - drug_example.query("drug==0")["days"].mean()
# Since the true effect for man is -3 and the true effect for woman is -2, the ATE should be
#
# $
# ATE=\dfrac{(-3*6) + (-2*4)}{10}=-2.6
# $
#
# This estimate is done by 1) partitioning the data into confounder cells, in this case, man and women, 2) estimating the effect on each cell and 3) combining the estimate with a weighted average, where the weight is the sample size of the cell or covariate group. If we had exactly the same size of man and woman in the data, the ATE estimate would be right in the middle of the ATE of the 2 groups, -2.5. Since there are more men than women in our dataset, the ATE estimate is a little bit closer to the man's ATE. This is called a non-parametric estimate, since it places no assumption on how the data was generated.
#
# If we control for sex using regression, we will add the assumption of linearity. Regression will also partition the data into man and woman and estimate the effect on both of these groups. So far, so good. However, when it comes to combining the effect on each group, it does not weigh them by the sample size. Instead, regression uses weights that are proportional to the variance of the treatment in that group. In our case, the variance of the treatment in men is smaller than in women, since only one man is in the control group. To be exact, the variance of T for man is \\(0.139=1/6*(1 - 1/6)\\) and for women is \\(0.25=2/4*(1 - 2/4)\\). So regression will give a higher weight to women in our example and the ATE will be a bit closer to the women's ATE of -2.
smf.ols('days ~ drug + C(sex)', data=drug_example).fit().summary().tables[1]
# This result is more intuitive with dummy variables, but, in its own weird way, regression also keeps continuous variables constant while estimating the effect. Also with continuous variables, the ATE will point in the direction where covariates have more variance.
#
# So we've seen that regression has its idiosyncrasies. It is linear, parametric, likes high variance features... This can be good or bad, depending on the context. Because of this, it's important to be aware of other techniques we can use to control for confounders. Not only are they an extra tool in your causal tool belt, but understanding different ways to deal with confounding expands our understanding of the problem. For this reason, I present you now the **Subclassification Estimator!**
#
#
# ## The Subclassification Estimator
#
# 
#
# If there is some causal effect we want to estimate, like the effect of job training on earnings, **and** the treatment is not randomly assigned, we need to watch out for confounders. It could be that only more motivated people do the training and they would have higher earnings regardless of the training. We need to estimate the effect of the training program within small groups of individuals that are roughly the same in motivation level and any other confounders we may have.
#
# More generally, if there is some causal effect we want to estimate, but it is hard to do so because of confounding of some variables X, what we need to do is make the treatment vs control comparison within small groups where X is the same. If we have conditional independence \\((Y_0, Y_1)\perp T | X\\), then we can write the ATE as follows.
#
# $
# ATE = \int(E[Y|X, T=1] - E[Y|X, T=0])dP(x)
# $
#
# What this integral does is it goes through all the space of the distribution of features X, computes the difference in means for all those tiny spaces and combines everything into the ATE. Another way to see this is to think about a discrete set of features. In this case, we can say that the features X takes on K different cells \\(\{X_1, X_2, ..., X_k\}\\) and what we are doing is computing the treatment effect in each cell and combining them into the ATE. In this discrete case, converting the integral to a sum, we can derive the subclassifications estimator
#
#
# $
# \hat{ATE} = \sum^K_{i=0}(\bar{Y}_{k1} - \bar{Y}_{k0}) * \dfrac{N_k}{N}
# $
#
# where the bar represent the mean of the outcome on the treated, \\(Y_{k1}\\), and treated, \\(Y_{k0}\\), cell k and \\(N_{k}\\) is the number of observations in that same cell. As you can see, we are computing a local ATE for each cell and combining them using a weighted average, where the weights are the sample size of the cell. In our medicine example above, this would be the first estimate, which gave us −2.6.
#
# ## Matching Estimator
#
# 
#
# The subclassification estimator isn't used much in practice (we will see why shortly, it is because of the curse of dimensionality) but it gives us a nice intuition of what a causal inference estimator should do, how it should control for confounders. This allows us to explore other kinds of estimators, such as the Matching Estimator.
#
# The idea is very similar. Since some sort of confounder X makes it so that treated and untreated are not initially comparable, I can make them so by **matching each treated unit with a similar untreated unit**. It is like I'm finding an untreated twin for every treated unit. By making such comparisons, treated and untreated become again comparable.
#
# As an example, let's suppose we are trying to estimate the effect of a trainee program on earnings. Here is what the trainees looks like
trainee = pd.read_csv("./data/trainees.csv")
trainee.query("trainees==1")
# And here are the non-trainees:
trainee.query("trainees==0")
# If I do a simple comparison in means, we get that the trainees earn less money than those that didn't go through the program.
trainee.query("trainees==1")["earnings"].mean() - trainee.query("trainees==0")["earnings"].mean()
# However, if we look at the table above, we notice that trainees are much younger than non trainees, which indicates that age is probably a confounder. Let's use matching on age to try to correct that. We will take unit 1 from the treated and pair it with unit 27, since both are 28 years old. Unit 2 we will pair it with unit 34, unit 3 with unit 37, unit 4 we will pair it with unit 35... When it comes to unit 5, we need to find someone with age 29 from the non treated, but that is unit 37, which is already paired. This is not a problem, since we can use the same unit multiple times. If more than 1 unit is a match, we can choose randomly between them.
#
# This is what the matched dataset looks like for the first 7 units
# +
# make dataset where no one has the same age
unique_on_age = (trainee
.query("trainees==0")
.drop_duplicates("age"))
matches = (trainee
.query("trainees==1")
.merge(unique_on_age, on="age", how="left", suffixes=("_t_1", "_t_0"))
.assign(t1_minuts_t0 = lambda d: d["earnings_t_1"] - d["earnings_t_0"]))
matches.head(7)
# -
# Notice how the last column has the difference in earnings between the treated and its matched untreated unit. If we take the mean of this last column we get the ATET estimate while controlling for age. Notice how the estimate is now very positive, compared to the previous one where we used a simple difference in means.
matches["t1_minuts_t0"].mean()
# But this was a very contrived example, just to introduce matching. In reality, we usually have more than one feature and units don't match perfectly. In this case, we have to define some measurement of proximity to compare how units are close to each other. One common metric for this is the euclidean norm \\(||X_i - X_j||\\). This difference, however, is not invariant to the scale of the features. This means that features like age, that take values on the tenths, will be much less important when computing this norm compared to features like income, which take the order of hundreds. For this reason, before applying the norm, we need to scale the features so that they are on roughly the same scale.
#
# Having defined a distance measure, we can now define the match as the nearest neighbour to that sample we wish to match. In math terms, we can write the matching estimator the following way
#
# $
# \hat{ATE} = \frac{1}{N} \sum^N_{i=0} (2T_i - 1)\big(Y_i - Y_{jm}(i)\big)
# $
#
# Where \\(Y_{jm}(i)\\) is the sample from the other treatment group which is most similar to \\(Y_i\\). We do this \\(2T_i - 1\\) to match both ways: treated with controls and controls with the treatment.
#
# To test this estimator, let's consider a medicine example. Once again, we want to find the effect of a medication on days until recovery. Unfortunately, this effect is confounded by severity, sex and age. We have reasons to believe that patients with more severe conditions have a higher chance of receiving the medicine.
med = pd.read_csv("./data/medicine_impact_recovery.csv")
med.head()
# If we look at a simple difference in means, \\(E[Y|T=1]-E[Y|T=0]\\), we get that the treatment takes, on average, 16.9 more days to recover than the untreated. This is probably due to confounding, since we don't expect the medicine to cause harm to the patient.
med.query("medication==1")["recovery"].mean() - med.query("medication==0")["recovery"].mean()
# To correct for this bias, we will control for X using matching. First, we need to remember to scale our features, otherwise, features like age will have higher importance than features like severity when we compute the distance between points. To do so, we can standardise the features.
# +
# scale features
X = ["severity", "age", "sex"]
y = "recovery"
med = med.assign(**{f: (med[f] - med[f].mean())/med[f].std() for f in X})
med.head()
# -
# Now, to the matching itself. Instead of coding a matching function, we will use the K nearest neighbour algorithm from [Sklearn](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsRegressor.html). This algorithm makes predictions by finding the nearest data point in an estimation or training set.
#
# For matching, we will need 2 of those. One, `mt0`, will store the untreated points and will find matches in the untreated when asked to do so. The other, `mt1`, will store the treated point and will find matches in the treated when asked to do so. After this fitting step, we can use these KNN models to make predictions, which will be our matches.
# +
from sklearn.neighbors import KNeighborsRegressor
treated = med.query("medication==1")
untreated = med.query("medication==0")
mt0 = KNeighborsRegressor(n_neighbors=1).fit(untreated[X], untreated[y])
mt1 = KNeighborsRegressor(n_neighbors=1).fit(treated[X], treated[y])
predicted = pd.concat([
# find matches for the treated looking at the untreated knn model
treated.assign(match=mt0.predict(treated[X])),
# find matches for the untreated looking at the treated knn model
untreated.assign(match=mt1.predict(untreated[X]))
])
predicted.head()
# -
# With the matches, we can now apply the matching estimator formula
#
# $
# \hat{ATE} = \frac{1}{N} \sum^N_{i=0} (2T_i - 1)\big(Y_i - Y_{jm}(i)\big)
# $
np.mean((2*predicted["medication"] - 1)*(predicted["recovery"] - predicted["match"]))
# Using this sort of matching, we can see that the effect of the medicine is not positive anymore. This means that, controlling for X, the medicine reduces the recovery time by about 1 day, on average. This is already a huge improvement on top of the biased estimate that predicted a 16.9 increase in recovery time.
#
# However, we can still do better.
#
# ## Matching Bias
#
# It turns out the matching estimator as we've designed above is biased. To see this, let's consider the ATET estimator, instead of the ATE, just because it is simpler to write. The intuition will apply to the ATE as well.
#
# $
# \hat{ATET} = \frac{1}{N_1}\sum (Y_i - Y_j(i))
# $
#
# where \\(N_1\\) is the number of treated individuals and \\(Y_j(i)\\) is the untreated match of treated unit i. To check for bias, what we do is hope we can apply the Central Limit Theorem so that this down there converges to a normal distribution with mean zero.
#
# $
# \sqrt{N_1}(\hat{ATET} - ATET)
# $
#
# However, this doesn't alway happen. If we define the mean outcome for the untreated given X, \\(\mu_0(x)=E[Y|X=x, T=0]\\), we will have that (btw, I've omitted the proof for that because it's a little beyond the point here).
#
# $
# E[\sqrt{N_1}(\hat{ATET} - ATET)] = E[\sqrt{N_1}(\mu_0(X_i) - \mu_0(X_j(i)))]
# $
#
# Now, \\(\mu_0(X_i) - \mu_0(X_j(i))\\) is not so simple to understand, so let's look at it more carefully. \\(\mu_0(X_i)\\) is the outcome Y value of a treated unit i had it not been treated. So, it is the counterfactual outcome \\(Y_0\\) for unit i. \\(\mu_0(X_j(i))\\) is the outcome of the untreated unit j that is the match of unit i. So, it is also the \\(Y_0\\) , but for unit j now. Only this time, it is a factual outcome, because j is in the non treated group. Now, because j and i are only similar, but not the same, this will likely not be zero. In other words, \\(X_i \approx X_j \\). So, \\(Y_{0i} \approx Y_{0j} \\).
#
# As we increase the sample size, there will be more units to match, so the difference between unit i and its match j will also get smaller. But this difference converges to zero slowly. As a result \\(E[\sqrt{N_1}(\mu_0(X_i) - \mu_0(X_j(i)))]\\) may not converge to zero, because the \\(\sqrt{N_1}\\) grows faster than \\((\mu_0(X_i) - \mu_0(X_j(i)))\\) diminishes.
#
# Bias arises when the matching discrepancies are huge. Fortunately, we know how to correct it. Each observation contributes \\((\mu_0(X_i) - \mu_0(X_j(i)))\\) to the bias so all we need to do is subtract this quantity from each matching comparison in our estimator. To do so, we can replace \\(\mu_0(X_j(i))\\) with some sort of estimate of this quantity \\(\hat{\mu_0}(X_j(i))\\), which can be obtained with models like linear regression. This updates the ATET estimator to the following equation
#
# $
# \hat{ATET} = \frac{1}{N_1}\sum \big((Y_i - Y_{j(i)}) - (\hat{\mu_0}(X_i) - \hat{\mu_0}(X_{j(i)}))\big)
# $
#
# where \\(\hat{\mu_0}(x)\\) is some estimative of \\(E[Y|X, T=0]\\), like a linear regression fitted only on the untreated sample.
# +
from sklearn.linear_model import LinearRegression
# fit the linear regression model to estimate mu_0(x)
ols0 = LinearRegression().fit(untreated[X], untreated[y])
ols1 = LinearRegression().fit(treated[X], treated[y])
# find the units that match to the treated
treated_match_index = mt0.kneighbors(treated[X], n_neighbors=1)[1].ravel()
# find the units that match to the untreatd
untreated_match_index = mt1.kneighbors(untreated[X], n_neighbors=1)[1].ravel()
predicted = pd.concat([
(treated
# find the Y match on the other group
.assign(match=mt0.predict(treated[X]))
# build the bias correction term
.assign(bias_correct=ols0.predict(treated[X]) - ols0.predict(untreated.iloc[treated_match_index][X]))),
(untreated
.assign(match=mt1.predict(untreated[X]))
.assign(bias_correct=ols1.predict(untreated[X]) - ols1.predict(treated.iloc[untreated_match_index][X])))
])
predicted.head()
# -
# One immediate question that arises is: doesn't this defeat the point of matching? If I have to run a linear regression anyway, why don't I use only that, instead of this complicated model. That's a fair point, so I should take some time to answer it.
#
# 
#
# First of all, this linear regression that we are fitting doesn't extrapolate on the treatment dimension to get the treatment effect. Instead, its purpose is just to correct bias. Linear regression here is local, in the sense that it doesn't try to see how the treated would be if it looked like the untreated. It does none of that extrapolation. This is left to the matching part. The meat of the estimator is still the matching component. The point I want to make here is that OLS is secondary to this estimator.
#
# The second point is that matching is a non-parametric estimator. It doesn't assume linearity or any kind of parametric model. As such, it is more flexible than linear regression and can work in situations where linear regression will not, namely, those where non linearity is very strong.
#
# Does this mean that you should only use matching? Well, that's a tough question. <NAME> makes a case that yes, you should. It's more flexible and, once you have the code, equally simple to run. I'm not entirely convinced by that. For once, Abadie spent a lot of time studying and developing the estimator (yes, he is one of the scientists that contributes to matching being what it is), so he obviously is personally invested in the method. Second, there is something about linear regression's simplicity that you don't see in matching. The partial derivative math of "holding everything else constant" is much easier to grasp with linear regression than with matching. But that's just my preference. To be honest, there is no clear answer to this question. Anyway, back to our example.
#
# With the bias correction formula, I get the following ATE estimation.
np.mean((2*predicted["medication"] - 1)*((predicted["recovery"] - predicted["match"])-predicted["bias_correct"]))
# Of course, we also need to place a confidence interval around this measurement, but enough of math theory now. In practice, we can simply use someone else's code and just import a matching estimator. Here is one from the library [causalinference](https://github.com/laurencium/causalinference).
# +
from causalinference import CausalModel
cm = CausalModel(
Y=med["recovery"].values,
D=med["medication"].values,
X=med[["severity", "age", "sex"]].values
)
cm.est_via_matching(matches=1, bias_adj=True)
print(cm.estimates)
# -
# Finally, we can say with confidence that our medicine does indeed lower the time someone spends at the hospital. The ATE estimate is just a little bit lower than mine, so probably my code is not perfect, so here is another reason to import someone else's code instead of building it your own.
#
# Before we close this topic, I just wanted to address the cause of bias in matching a little bit more. We saw that matching is biased when the unit and its match are not so similar. But what causes them to be so different?
#
#
# ## The Curse of Dimensionality
#
# As it turns out, the answer is quite simple and intuitive. It is easy to find people that mach on a few characteristics, like sex. But if we add more characteristics, like age, income, city of birth and so on, it becomes harder and harder to find matches. In more general terms, the more features we have, the higher will be the distance between units and their matches.
#
# This is not something that hurts only the matching estimator. It ties back to the subclassification estimator we saw earlier. Early on, in that contrived medicine example where with man and woman, it was quite easy to build the subclassification estimator. That was because we only had 2 cells: man and woman. But what would happen if we had more? Let's say we have 2 continuous features like age and income and we manage to discretise them into 5 buckets each. This will give us 25 cells, or \\(5^2\\). And what if we had 10 covariates with 3 buckets each? Doesn't seem like a lot right? Well, this would give us 59049 cells, or \\(3^{10}\\). It's easy to see how this can blow out of proportion pretty quickly. This is a phenomena pervasive in all data science, which is called the **The Curse of Dimensionality**!!!
#
# 
# Image Source: https://deepai.org/machine-learning-glossary-and-terms/curse-of-dimensionality
#
# Despite its scary and pretentious name, this only means that the number of data points required to fill a feature space grows exponentially with the number of features, or dimensions. So, if it takes X data points to fill the space of, say, 3 feature spaces, it takes exponentially more points to fill in the space of 4 features.
#
# In the context of the subclassification estimator, the curse of dimensionality means that it will suffer if we have lots of features. Lots of features imply multiple cells in X. If there are multiple cells, some of them will have very few data. Some of them might even have only treated or only control, so it won't be possible to estimate the ATE there, which would break our estimator. In the matching context, this means that the feature space will be very space and units will be very far from each other. This will increase the distance between matches and cause bias problems.
#
# As for linear regression, it actually handles this problem quite well. What it does is project all the features X into a single one, the Y dimension. It then makes treatment and control comparison on that projection. So, in some way, linear regression performs some sort of dimensionality reduction to estimate the ATE. It's quite elegant.
#
# Most causal models also have some way to deal with the curse of dimensionality. I won't keep repeating myself, but it is something you should keep in mind when looking at them. For instance, when we deal with propensity scores in the following section, try to see how it solves this problem.
#
# ## Key Ideas
#
# We've started this section understanding what linear regression does and how it can help us identify causal relationships. Namely, we understood that regression can be seen as partitioning the dataset into cells, computing the ATE in each cell and then combining the cell's ATE into a single ATE for the entire dataset.
#
# From there, we've derived a very general causal inference estimator with subclassification. We saw how that estimator is not very useful in practice but it gave us some interesting insights on how to tackle the problem of causal inference estimation. That gave us the opportunity to talk about the matching estimator.
#
# Matching controls for the confounders by looking at each treated unit and finding an untreated pair that is very similar to it and similarly for the untreated units. We saw how to implement this method using the KNN algorithm and also how to debiase it using regression. Finally, we discussed the difference between matching and linear regression. We saw how matching is a non parametric estimator that doesn't rely on linearity the way linear regression does.
#
# Finally, we've delved into the problem of high dimensional datasets and we saw how causal inference methods can suffer from it.
#
#
# ## References
#
# I like to think of this entire book as a tribute to <NAME>, <NAME> and <NAME> for their amazing Econometrics class. Most of the ideas here are taken from their classes at the American Economic Association. Watching them is what is keeping me sane during this tough year of 2020.
# * [Cross-Section Econometrics](https://www.aeaweb.org/conference/cont-ed/2017-webcasts)
# * [Mastering Mostly Harmless Econometrics](https://www.aeaweb.org/conference/cont-ed/2020-webcasts)
#
# I'll also like to reference the amazing books from Angrist. They have shown me that Econometrics, or 'Metrics as they call it, is not only extremely useful but also profoundly fun.
#
# * [Mostly Harmless Econometrics](https://www.mostlyharmlesseconometrics.com/)
# * [Mastering 'Metrics](https://www.masteringmetrics.com/)
#
# My final reference is <NAME> and <NAME>' book. It has been my trustworthy companion in the most thorny causal questions I had to answer.
#
# * [Causal Inference Book](https://www.hsph.harvard.edu/miguel-hernan/causal-inference-book/)
#
# 
#
# ## Contribute
#
# Causal Inference for the Brave and True is an open-source material on causal inference, the statistics of science. It uses only free software, based in Python. Its goal is to be accessible monetarily and intellectually.
# If you found this book valuable and you want to support it, please go to [Patreon](https://www.patreon.com/causal_inference_for_the_brave_and_true). If you are not ready to contribute financially, you can also help by fixing typos, suggesting edits or giving feedback on passages you didn't understand. Just go to the book's repository and [open an issue](https://github.com/matheusfacure/python-causality-handbook/issues). Finally, if you liked this content, please share it with others who might find it useful and give it a [star on GitHub](https://github.com/matheusfacure/python-causality-handbook/stargazers).
| causal-inference-for-the-brave-and-true/10-Matching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1) Print Hello World
print('hello world')
# # 2) comments
# ### 2.1) comments
# +
# note that print is a function
print('hello world')
# Code tells you how, comments should tell you why.
# -
# ### 2.2) Multiline Comments
"""
If I really hate pressing `enter` and
typing all those hash marks, I could
just do this instead
"""
# ### 2.3) Docstring
# +
def somefuction():
"""Depending on where they sit in your program,
they could turn into docstrings, which are pieces of documentation
that are associated with a function or method.
If you slip one of these bad boys right after a function definition,
then what you intended to be a comment will become associated
with that object."""
# -
# # 3) Literal Constants
5 #is a literal constant
1.23 #is a literal constant
'This is a string' #is a literal constant
"It's is a string" #is a literal constant
# also notice how to use apostrophe in a string
# ## 3.1) Numbers
# Two types of numbers:
# * __integers__ : The whole numbers like: 2, 33885, 405930442859, etc.
# * __float__ : The decimals/fractions like: 3.23 and 3.45E-2 = 0.0345. The `E` notation indicates powers of 10
# ## 3.2) Strings
# ### 3.2.1) using single quotes
a = 'string'
a
# ### 3.2.2) using double qoutes
b = "what's up?" #preferred
b
# ### 3.2.3) using triple qoutes
# multiline comments (2 ways like multiline comments)
c = """sdf
dfd"""
d = '''as
fg
ggg
'''
e = 'This is the first line\nThis is the second line'
c
d
e
# without newline (\n)
f = "This is the first sentence. \
This is the second sentence."
f
# ### 3.3.1) format method
# The format method is used when wish to subsititue variable values in a string.
# +
n = 3
price = 1.5
st = 'the price of {0} apples is {1}'.format(n,price)
st
# -
st2 = 'the price of {} apples is {}'.format(n,price)
st2
print('order matters in this case')
st3 = 'the price of {} apples is {}'.format(price,n)
st3
print('order matters in this case')
st4 = 'the price of {n} apples is {price}'.format(price = price,n = n)
st4
# decimal (.) precision of 3 for float '0.333'
print('{0:.3f}'.format(1.0/3))
# fill with underscores (_) with the text centered
# (^) to 11 width '___hello___'
print('{0:_^11}'.format('hello'))
# keyword-based 'Swaroop wrote A Byte of Python'
print('{name} wrote {book}'.format(name='Swaroop', book='A Byte of Python'))
# ### 3.3.2) string concatenation (ugly and error prone)
"the price of "+ str(n) + " apples is "+ str(price)
# ### 3.3.3) fStrings method
f'the price of {n} apples is {price}' #preferred
# # 4) Print function
# Print always ends with an invisible "new line" character ( \n ) so that repeated
# calls to print will all print on a separate line each. To prevent this newline character from being printed, you can specify that it
# should end with a blank:
print('a', end='')
print('b', end='')
print('a', end=' ')
print('b', end=' ')
print('c')
# # 5) Escape Sequences
'What's your name?'
# _You can see the apostrophe cannot be used in the avove case. To make this possible we need escape Sequence `\`_
'What\'s your name?'
"What's your name?" # same output as above
'Male\\Female'
# __Some often used Escape Sequence__
#
# * tab: `\t`
#
# * newline: `\n`
# # 6) Raw Strings
# If you need to specify some strings where no special processing such as escape sequences are handled, then what you need is to
# specify a raw string by prefixing r or R to the string. An example is:
r"Newlines are indicated by \n"
R'hi\tlol'
'hi\tlol'
print('hi\tlol')
print(R'hi\tlol')
# Always use raw strings when dealing with regular expressions. Otherwise, a lot of backwhacking may be required. For
# example, backreferences can be referred to as `\\1` or r`\1` .
# # 6) Variable / Identifier
# Using just literal constants can soon become boring - we need some way of storing any information and manipulate them as well.
# This is where variables come into the picture. Variables are exactly what the name implies - their value can vary, i.e., you can
# store anything using a variable. Variables are just parts of your computer's memory where you store some information. Unlike
# literal constants, you need some method of accessing these variables and hence you give them names.
# ### 6.2) Identifier Naming
# Variables are examples of identifiers. Identifiers are names given to identify something. There are some rules you have to follow
# for naming identifiers:
#
# The first character of the identifier must be a letter of the alphabet (uppercase ASCII or lowercase ASCII or Unicode
# character) or an underscore ( _ ).
#
# The rest of the identifier name can consist of letters (uppercase ASCII or lowercase ASCII or Unicode character),
# underscores ( _ ) or digits (0-9).
#
# Identifier names are case-sensitive. For example, myname and myName are not the same. Note the lowercase n in the
# former and the uppercase N in the latter.
#
# Examples of valid identifier names are i , name_2_3 . Examples of invalid identifier names are 2things , this is
# spaced out , my-name and >a1b2_c3
# # 7) Data Types
#
# Variables can hold values of different types called data types. The basic types are numbers and strings, which we have already
# discussed. In later chapters, we will see how to create our own types using classes.
# # 8) Object
#
# Remember, Python refers to anything used in a program as an object. This is meant in the generic sense. Instead of saying "the
# something"', we say "the object".
# Note for Object Oriented Programming users:
# Python is strongly object-oriented in the sense that everything is an object including numbers, strings and functions.
#
# # 9) Logical And Physical Line
# A physical line is what you see when you write the program. A logical line is what Python sees as a single statement. Python
# implicitly assumes that each physical line corresponds to a logical line.
# Implicitly, Python encourages the use of a single statement per line which makes code more readable.
# If you want to specify more than one logical line on a single physical line, then you have to explicitly specify this using a
# semicolon ( ; ) which indicates the end of a logical line/statement.
# one logical line & one physical line:
print('hi, there, you are the best!')
# two logical line & one physical line:
print('hi'); print('you are beutiful')
# *__Recommended__* use is to stick to writing a maximum of a single logical line on each single physical line. The idea
# is that you should never use the semicolon.
# There is one kind of situation where this concept is really useful: if you have a long line of code, you can break it into multiple
# physical lines by using the backslash. This is referred to as __explicit line joining__:
s = 'This is a string. \
This continues the string.'
print(s)
i = \
5
i
# above is same as
i = 5
# Sometimes, there is an implicit assumption where you don't need to use a backslash. This is the case where the logical line has a
# starting parentheses, starting square brackets or a starting curly braces but not an ending one. This is called __implicit line joining__ (when using _list_).
# # 10) Indentation
# Whitespace is important in Python. Actually, __whitespace__ at the __beginning of the line is important__. This is called _indentation_.
# Leading whitespace (spaces and tabs) at the beginning of the logical line is used to determine the indentation level of the logical
# line, which in turn is used to determine the grouping of statements.
#
# This means that statements which go together __must__ have the same indentation. Each such set of statements is called a __block__.
#
# We will see examples of how blocks are important in later chapters.
# One thing you should remember is that wrong indentation can give rise to errors. For example:
i = 5
# Error below! Notice a single space at the start of the line
print('Value is', i)
print('I repeat, the value is', i)
# __you cannot arbitrarily start new
# blocks of statements__ (except for the default main block which you have been using all along, of course). Cases where you can use new blocks will be detailed in later chapters such as the control flow.
#
#
#
# __The official Python language recommendation is Use four spaces for indentation.__
# # Mutability in python
# Also check this:
# * https://medium.com/datadriveninvestor/mutable-and-immutable-python-2093deeac8d9
# * https://www.pitt.edu/~naraehan/python3/mutability.htm
| 1_pythonBasic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WQhuvSIVoh0_" colab_type="text"
# # Data Exploration
#
# Just for personal usage, read google drive
# + id="AoreAl2qomRu" colab_type="code" outputId="d79e152c-51a7-4966-c7fe-29fbfd71c797" executionInfo={"status": "ok", "timestamp": 1576665271313, "user_tz": 300, "elapsed": 758, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="KhKvwBWhlLq3" colab_type="text"
# set path
# + id="fmKotplKoh1A" colab_type="code" outputId="bc273fc4-a6bb-4adb-876c-3e93abfe140b" executionInfo={"status": "ok", "timestamp": 1576665271724, "user_tz": 300, "elapsed": 198, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 70}
# %cd /content/gdrive/My\ Drive/
# %cd cs435
import json
import os
import pandas as pd
from pathlib import Path
pd.set_option('max_colwidth',300)
from pprint import pprint
current_path = os.getcwd()
print(current_path)
#path is /content/gdrive/My Drive/cs435
# + [markdown] id="Jd-RMj1Noh1j" colab_type="text"
# ## Exploring The Full Dataset
#
# + [markdown] id="Qe5A6mi6oh1j" colab_type="text"
# The preprocessed data re stored in [json lines](http://jsonlines.org/) format. First, we can get a list of all these files for further inspection:
# Reading python file from this location, change to any location for reading.
# For full dataset, go to share link https://colab.research.google.com/drive/1s9DMUhYz0fhDDUAk1tQvfKcY3_W51LsW or Download directly from CodeSearchNet
# + id="vfAqKTHhoh1k" colab_type="code" colab={}
python_files = sorted(Path('/content/gdrive/My Drive/cs435/data/python/').glob('**/*.gz'))
# For full dataset, go to share link https://colab.research.google.com/drive/1s9DMUhYz0fhDDUAk1tQvfKcY3_W51LsW
# or Download directly from CodeSearchNet
#java_files = sorted(Path('/content/gdrive/My Drive/cs435/data/java/').glob('**/*.gz'))
#go_files = sorted(Path('/content/gdrive/My Drive/cs435/data/go/').glob('**/*.gz'))
#php_files = sorted(Path('/content/gdrive/My Drive/cs435/data/php/').glob('**/*.gz'))
#javascript_files = sorted(Path('/content/gdrive/My Drive/cs435/data/javascript/').glob('**/*.gz'))
#ruby_files = sorted(Path('/content/gdrive/My Drive/cs435/data/ruby/').glob('**/*.gz'))
#all_files = python_files + go_files + java_files + php_files + javascript_files + ruby_files
# + [markdown] id="lGgrLMRpoh1q" colab_type="text"
# To make analysis of this dataset easier, we can load all of the data into a pandas dataframe:
# + id="-o4STan8oh1r" colab_type="code" colab={}
columns_long_list = ['repo', 'path', 'url', 'code',
'code_tokens', 'docstring', 'docstring_tokens',
'language', 'partition']
columns_short_list = ['code_tokens', 'docstring_tokens',
'language', 'partition']
def jsonl_list_to_dataframe(file_list, columns=columns_long_list):
"""Load a list of jsonl.gz files into a pandas DataFrame."""
return pd.concat([pd.read_json(f,
orient='records',
compression='gzip',
lines=True)[columns]
for f in file_list], sort=False)
# + [markdown] id="SSu1qBnJoh1z" colab_type="text"
# Two columns that will be heavily used in this dataset are `code_tokens` and `docstring_tokens`, which represent a parallel corpus that can be used for interesting tasks like information retrieval (for example trying to retrieve a codesnippet using the docstring.). You can find more information regarding the definition of the above columns in the README of this repo.
#
# Next, we will read in all of the data for a limited subset of these columns into memory so we can compute summary statistics. **Warning:** This step takes ~ 5 minutes.
# + id="1CdFJsL-oh10" colab_type="code" outputId="89e7c927-81fc-499d-dd7a-b90c2040471e" executionInfo={"status": "ok", "timestamp": 1576665322331, "user_tz": 300, "elapsed": 41621, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 720}
#all_df = jsonl_list_to_dataframe(all_files, columns_short_list)
#all_df = top10_jsonl_list_to_dataframe(all_files, columns_short_list)
pydf = jsonl_list_to_dataframe(python_files) #We can change this to other language based on the code type of prediction
all_df = pydf
pydf.head()
# + [markdown] id="_c47Pn24oh13" colab_type="text"
# ## Summary Statistics
# + [markdown] id="zU61Iqv3oh13" colab_type="text"
# ### Row Counts
#
# By Partition
# + id="SL1do7cNoh14" colab_type="code" outputId="53b4d582-0bff-4de8-c768-2223b90ddc33" executionInfo={"status": "ok", "timestamp": 1576651026230, "user_tz": 300, "elapsed": 52080, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
all_df.partition.value_counts()
# + [markdown] id="knV0aX89oh17" colab_type="text"
# By Language
# + id="AERtjXl7oh18" colab_type="code" outputId="07ed8380-1a24-40ea-f05b-4f1d984cddc6" executionInfo={"status": "ok", "timestamp": 1576651026479, "user_tz": 300, "elapsed": 52320, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
all_df.language.value_counts()
# + [markdown] id="TRePE0Tjoh1_" colab_type="text"
# By Partition & Language
# + id="fccVb729oh2A" colab_type="code" outputId="45f9a8b9-d84a-4b48-df96-be7f10f77628" executionInfo={"status": "ok", "timestamp": 1576651026480, "user_tz": 300, "elapsed": 52317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 106}
all_df.groupby(['partition', 'language'])['code_tokens'].count()
# + [markdown] id="F1pc9ZX2oh2C" colab_type="text"
# ### Token Lengths By Language
# + id="u8fNZ0yvoh2D" colab_type="code" colab={}
all_df['code_len'] = all_df.code_tokens.apply(lambda x: len(x))
all_df['query_len'] = all_df.docstring_tokens.apply(lambda x: len(x))
# + [markdown] id="LT08jXJsoh2K" colab_type="text"
# #### Code Length Percentile By Language
#
# For example, the 80th percentile length for python tokens is 72
# + id="lJdbipuToh2N" colab_type="code" outputId="315624af-0bec-4113-c75c-6b72d9a512f1" executionInfo={"status": "ok", "timestamp": 1576651027262, "user_tz": 300, "elapsed": 53086, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 231}
code_len_summary = all_df.groupby('language')['code_len'].quantile([.5, .7, .8, .9, .95])
display(pd.DataFrame(code_len_summary))
# + [markdown] id="92WMdvfsoh2R" colab_type="text"
# #### Query Length Percentile By Language
#
# For example, the 80th percentile length for python tokens is 19
# + id="u-kculoloh2S" colab_type="code" outputId="d0cea699-3340-43dc-bb12-ae4639b20944" executionInfo={"status": "ok", "timestamp": 1576651027616, "user_tz": 300, "elapsed": 53436, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 231}
query_len_summary = all_df.groupby('language')['query_len'].quantile([.5, .7, .8, .9, .95])
display(pd.DataFrame(query_len_summary))
# + [markdown] id="NiyS7Dqtoh2W" colab_type="text"
# #### Query Length All Languages
# + id="9oKHGltHoh2Y" colab_type="code" outputId="12ae4a5f-bcea-48c4-db95-ca3b02c807ad" executionInfo={"status": "ok", "timestamp": 1576651027617, "user_tz": 300, "elapsed": 53428, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 201}
query_len_summary = all_df['query_len'].quantile([.5, .7, .8, .9, .95])
display(pd.DataFrame(query_len_summary))
# + [markdown] id="FN_13DxTCa3U" colab_type="text"
# # Train the Model
#
# + [markdown] id="1vLPS2OvC3HN" colab_type="text"
# Load in all the libraries needed
#
# + id="W08n3bZeoh2d" colab_type="code" outputId="f346de93-14f2-4c8c-e4f8-2943f8c165ff" executionInfo={"status": "ok", "timestamp": 1576665322333, "user_tz": 300, "elapsed": 34810, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 720}
from fastai import *
from fastai.vision import *
from fastai.text import *
from fastai.callbacks import *
import numpy as np
import pandas as pd
import collections
import warnings
#no UserWarning display
warnings.simplefilter("ignore", UserWarning)
#inspect into python dataframe
pydf.head()
# + [markdown] id="3eQm5VIaDGL7" colab_type="text"
# Convert all code and docstring to lower case for better training and use 10% of training data
#
# + id="lMX4lb3PAeu1" colab_type="code" colab={}
pydf['code']=pydf['code'].apply(lambda x:x.lower())
pydf['docstring']=pydf['docstring'].apply(lambda x:x.lower())
pydf.head()
pydf_train_df=pydf.loc[pydf['partition'] == 'train']
# + [markdown] id="WqLnvHKJllNm" colab_type="text"
# Install Sentencepiece
# + id="Q-ieLOYaW1B0" colab_type="code" outputId="b4a53541-462a-4ec2-e3fd-7c2b29d6d171" executionInfo={"status": "ok", "timestamp": 1576665328054, "user_tz": 300, "elapsed": 5683, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
pip install sentencepiece
# + [markdown] id="c8bmi18nDTJq" colab_type="text"
# Train the sentencepiece model for prepocessing using the training dataframe
#
#
#
# + id="qQxUiWDv9K3O" colab_type="code" outputId="ee6c8d51-c120-4921-dbf2-9125a261814b" executionInfo={"status": "ok", "timestamp": 1576655984618, "user_tz": 300, "elapsed": 387008, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE<KEY>", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
import sentencepiece as spm
train_txt = "train.txt"
with open(train_txt, "w+", encoding='UTF-8') as fp:
for file in pydf_train_df.code:
fp.write(file+"\n")
spm.SentencePieceTrainer.Train('--input=train.txt \
--model_prefix=train_py --vocab_size=5000 --model_type=bpe')
# + [markdown] id="Hqm69VgIDcPA" colab_type="text"
# Generate databunch using sentencepiece as preprocessor for AWT_LSTM model and Transformer
# + id="pPRtjd5Vaxa-" colab_type="code" colab={}
def conv_to_ds(df, data_path, model_name, bs = 64):
# convert dataframe to TextList
return (TextList
.from_df(df, data_path,
processor = SPProcessor(
sp_model = model_name+".model",
sp_vocab = model_name+".vocab"
))
.split_none()
.label_for_lm()
.databunch(bs = bs)
)
def gen_lm_data(df_trn, df_val, data_path, model_name, bs = 64, sample = 1):
# generate data bunch for training
db_trn = conv_to_ds(df_trn, data_path, model_name)
db_val = conv_to_ds(df_val, data_path, model_name)
data = TextLMDataBunch.create(
train_ds = db_trn.train_ds, valid_ds = db_val.train_ds, path = data_path, bs = bs
)
data.label_list = db_trn.label_list
data.label_list.valid = db_val.label_list.train
return data
# + [markdown] id="3dxl5ELnOgQK" colab_type="text"
# ## Basic training of AWT_LSTM model
# + [markdown] id="X2Vemd8rOe0N" colab_type="text"
# AWT_LSTM model to generate codes after certain words.
#
#
# + id="-C8UBLxROKQM" colab_type="code" outputId="7100ac58-7233-4ef1-e906-d5c48d607ea0" executionInfo={"status": "ok", "timestamp": 1576665406694, "user_tz": 300, "elapsed": 84291, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 140}
#generate validation for model
pydf_valid_df_10 = pydf_train_df.sample(frac=0.1, random_state=0)
pydf_train_10_90 = pydf_train_df.drop(pydf_valid_df_10.index)
py_datalm = gen_lm_data(pydf_train_10_90, pydf_valid_df_10, current_path, "train_py")
#create model learn and fit model
learn = language_model_learner(py_datalm, AWD_LSTM, drop_mult=0.5)
#First choos default with smaller slice to improve accuracy
learn.fit_one_cycle(1, 1e-2)
learn.unfreeze()
#increase the accuracy everytime with e^-1 improved
#can try with smaller slice to increase accuracy but may perform bad on test. Usually 1e-3 is the best
learn.fit_one_cycle(1, 1e-3)
# + [markdown] id="8DBAJ4e3lt8F" colab_type="text"
# Generate codes that start with "xxbos". Use preprocessor model to decode the prediction
# + id="G0BhyHP4OV2a" colab_type="code" outputId="ad561034-3503-4992-eec0-1bc695f6e195" executionInfo={"status": "ok", "timestamp": 1576665605175, "user_tz": 300, "elapsed": 1777, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 73}
sp1 = spm.SentencePieceProcessor()
sp1.Load("train_py.model")
#original prediction
print(learn.predict("xxbos", n_words=300))
#decoded prediction
xxbos_predict_AWT = sp1.decode_pieces(list(learn.predict("xxbos", n_words=300).split(" ")))
print(xxbos_predict_AWT)
# + [markdown] id="gfycq6baPVZJ" colab_type="text"
# ## Basic training of Transformer model
# + [markdown] id="LDzgTNN9RVzE" colab_type="text"
# Train the basic transformer model with fine-tuned hyperparameter 2e-5 (with early stopping callback)
# + id="tSJrP4oKN7rX" colab_type="code" outputId="48a81b6e-3605-4846-a2bf-a4c2577d6916" executionInfo={"status": "ok", "timestamp": 1576666048968, "user_tz": 300, "elapsed": 266519, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 140}
#generate validation for model
pydf_valid_df_10 = pydf_train_df.sample(frac=0.1, random_state=0)
pydf_train_10_90 = pydf_train_df.drop(pydf_valid_df_10.index)
py_datalm = gen_lm_data(pydf_train_10_90, pydf_valid_df_10, current_path, "train_py")
#train with default Transformer,drop_mult applied to avoid overfit
learn = language_model_learner(py_datalm, Transformer,drop_mult=0.5)
#use similar result from above, 1e-3 is the best here
learn.fit_one_cycle(1, 1e-3)
#accuracy increased
learn.unfreeze()
learn.fit_one_cycle(1, 2e-5)
#can add callback to the model
#n = len(py_datalm.train_ds.vocab.itos)
#model = Transformer(n, n, n_layers=6, n_heads=8, d_model=256, d_head=32, d_inner=1024)
#learn = Learner(py_datalm, model, metrics=[accuracy],
# callback_fns=[partial(EarlyStoppingCallback, monitor='accuracy', min_delta=0.01, patience=3)])
# + id="C6ulAsMzPHBn" colab_type="code" outputId="f053e194-f9b2-4ac9-c30a-4b69e62b8161" executionInfo={"status": "ok", "timestamp": 1576666053580, "user_tz": 300, "elapsed": 3702, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 73}
#original prediction
print(learn.predict("xxbos", n_words=100))
#decoded prediction
xxbos_predict_Trans = sp1.decode_pieces(list(learn.predict("xxbos", n_words=100).split(" ")))
print(xxbos_predict_Trans)
# + [markdown] id="IQlEyn-OpFO0" colab_type="text"
# ## Seq2Seq and Transformer
# + [markdown] id="L0UxVNWRpVJY" colab_type="text"
# A better language translation model, from docstrings to codes
#
# + [markdown] id="hIUkuvZNCpST" colab_type="text"
# ### Prepocessing
# + [markdown] id="669ld5pZPc-r" colab_type="text"
# #### Seq2Seq and Sentencepiece preprocessor
#
# + [markdown] id="KiG4CQGfMH2J" colab_type="text"
# Train the sentencepiece model for prepocessing the docstring
# + id="ZPZv4KE6MCZL" colab_type="code" outputId="5f04731c-d722-47e2-cfb1-f909b03510b7" executionInfo={"status": "ok", "timestamp": 1576659003458, "user_tz": 300, "elapsed": 50933, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
train_docstring_txt = "train_docstring.txt"
with open(train_txt, "w+", encoding='UTF-8') as fp:
for file in pydf_train_df.docstring:
fp.write(file+"\n")
spm.SentencePieceTrainer.Train('--input=train.txt \
--model_prefix=train_docstring_py --vocab_size=2000 --model_type=bpe')
# + [markdown] id="o3bYifX0Do_g" colab_type="text"
# Firt,we will need to collate inputs and targets in a batch: they have different lengths so we need to add padding to make the sequence length the same. (More details can be found in fast.ai Seq2Seq Translation part)
# + id="drXhUbvhoIYp" colab_type="code" colab={}
def seq2seq_collate(samples, pad_idx=1, pad_first=True, backwards=False):
"Function that collect samples and adds padding. Flips token order if needed"
samples = to_data(samples)
max_len_x,max_len_y = max([len(s[0]) for s in samples]),max([len(s[1]) for s in samples])
res_x = torch.zeros(len(samples), max_len_x).long() + pad_idx
res_y = torch.zeros(len(samples), max_len_y).long() + pad_idx
if backwards: pad_first = not pad_first
for i,s in enumerate(samples):
if pad_first:
res_x[i,-len(s[0]):],res_y[i,-len(s[1]):] = LongTensor(s[0]),LongTensor(s[1])
else:
res_x[i, :len(s[0])],res_y[i, :len(s[1])] = LongTensor(s[0]),LongTensor(s[1])
if backwards: res_x,res_y = res_x.flip(1),res_y.flip(1)
return res_x, res_y
# + [markdown] id="nuBMXR2RD4jM" colab_type="text"
# Then we create a special DataBunch that uses this collate function.
#
#
# + id="1JEL-n9qoLyv" colab_type="code" colab={}
class Seq2SeqDataBunch(TextDataBunch):
"Create a `TextDataBunch` suitable for training an RNN classifier."
@classmethod
def create(cls, train_ds, valid_ds, test_ds=None, path='.', bs=32, val_bs=None, pad_idx=1,
dl_tfms=None, pad_first=False, device=None, no_check=False, backwards=False, **dl_kwargs):
"Function that transform the `datasets` in a `DataBunch` for classification. Passes `**dl_kwargs` on to `DataLoader()`"
datasets = cls._init_ds(train_ds, valid_ds, test_ds)
val_bs = ifnone(val_bs, bs)
collate_fn = partial(seq2seq_collate, pad_idx=pad_idx, pad_first=pad_first, backwards=backwards)
train_sampler = SortishSampler(datasets[0].x, key=lambda t: len(datasets[0][t][0].data), bs=bs//2)
train_dl = DataLoader(datasets[0], batch_size=bs, sampler=train_sampler, drop_last=True, **dl_kwargs)
dataloaders = [train_dl]
for ds in datasets[1:]:
lengths = [len(t) for t in ds.x.items]
sampler = SortSampler(ds.x, key=lengths.__getitem__)
dataloaders.append(DataLoader(ds, batch_size=val_bs, sampler=sampler, **dl_kwargs))
return cls(*dataloaders, path=path, device=device, collate_fn=collate_fn, no_check=no_check)
# + [markdown] id="boYOZk69D-mx" colab_type="text"
#
# And a subclass of TextList that will use this DataBunch class in the call .databunch and will use TextList to label (since our targets are other texts).
# + id="WRqNC-hfoZGq" colab_type="code" colab={}
class Seq2SeqTextList(TextList):
_bunch = Seq2SeqDataBunch
_label_cls = TextList
# + [markdown] id="w46OoeCfEDAz" colab_type="text"
# Thats all we need to use the data block API!
#
#
# + id="skZ8WDe3obNB" colab_type="code" outputId="6ba26437-f08a-45c3-9071-9c2663137c0c" executionInfo={"status": "ok", "timestamp": 1576659679479, "user_tz": 300, "elapsed": 268775, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 17}
src = Seq2SeqTextList.from_df(pydf_train_df, path = current_path, cols='docstring', processor = SPProcessor(
sp_model = "train_docstring_py.model",
sp_vocab = "train_docstring_py.vocab"
)).split_by_rand_pct(seed=42).label_from_df(cols='code', label_cls=TextList, processor = SPProcessor(
sp_model = "train_py.model",
sp_vocab = "train_py.vocab"
))
# + [markdown] id="CZ86KQZPEEiu" colab_type="text"
# We remove the items where one of the target is more than 300 tokens long or docstring longer than 100. To make prediction more precise
# + id="huwqa04tA5T8" colab_type="code" outputId="95fa6c8e-2353-488e-cb0f-640f2ae7c2ec" executionInfo={"status": "ok", "timestamp": 1576659698155, "user_tz": 300, "elapsed": 1166, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
src = src.filter_by_func(lambda x,y: len(x) > 300 or len(y) > 100)
len(src.train) + len(src.valid)
# + [markdown] id="4pau0SzVEJ_d" colab_type="text"
# Save the data for later load in
# + id="I41mX7BWtDSB" colab_type="code" outputId="807a38d0-bf01-40db-f788-d1c6f241b032" executionInfo={"status": "ok", "timestamp": 1576659729398, "user_tz": 300, "elapsed": 26213, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 339}
data = src.databunch()
data.save()
data
# + [markdown] id="xE-O5W2am6cr" colab_type="text"
# show the batch result
# + id="qWeDkg6ctyVO" colab_type="code" outputId="5db027a3-3974-4dd2-8087-d088022be402" executionInfo={"status": "ok", "timestamp": 1576659795219, "user_tz": 300, "elapsed": 17240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 614}
data = load_data(current_path)
data.show_batch()
# + [markdown] id="MV9jfGYKEtME" colab_type="text"
# ### Training
#
#
# + [markdown] id="IAsB-aCeEPEB" colab_type="text"
# BLEU metric, detailed doc can be searched in bleu in fast.ai library
#
# + id="eZnZes7U9bt9" colab_type="code" colab={}
class NGram():
def __init__(self, ngram, max_n=5000): self.ngram,self.max_n = ngram,max_n
def __eq__(self, other):
if len(self.ngram) != len(other.ngram): return False
return np.all(np.array(self.ngram) == np.array(other.ngram))
def __hash__(self): return int(sum([o * self.max_n**i for i,o in enumerate(self.ngram)]))
def get_grams(x, n, max_n=5000):
return x if n==1 else [NGram(x[i:i+n], max_n=max_n) for i in range(len(x)-n+1)]
def get_correct_ngrams(pred, targ, n, max_n=5000):
pred_grams,targ_grams = get_grams(pred, n, max_n=max_n),get_grams(targ, n, max_n=max_n)
pred_cnt,targ_cnt = Counter(pred_grams),Counter(targ_grams)
return sum([min(c, targ_cnt[g]) for g,c in pred_cnt.items()]),len(pred_grams)
class CorpusBLEU(Callback):
def __init__(self, vocab_sz):
self.vocab_sz = vocab_sz
self.name = 'bleu'
def on_epoch_begin(self, **kwargs):
self.pred_len,self.targ_len,self.corrects,self.counts = 0,0,[0]*4,[0]*4
def on_batch_end(self, last_output, last_target, **kwargs):
last_output = last_output.argmax(dim=-1)
for pred,targ in zip(last_output.cpu().numpy(),last_target.cpu().numpy()):
self.pred_len += len(pred)
self.targ_len += len(targ)
for i in range(4):
c,t = get_correct_ngrams(pred, targ, i+1, max_n=self.vocab_sz)
self.corrects[i] += c
self.counts[i] += t
def on_epoch_end(self, last_metrics, **kwargs):
precs = [c/t for c,t in zip(self.corrects,self.counts)]
len_penalty = exp(1 - self.targ_len/self.pred_len) if self.pred_len < self.targ_len else 1
bleu = len_penalty * ((precs[0]*precs[1]*precs[2]*precs[3]) ** 0.25)
return add_metrics(last_metrics, bleu)
# + [markdown] id="XqnW_JnbEmZX" colab_type="text"
# We add a transform to the dataloader that shifts the targets right and adds a padding at the beginning.
# + id="uLm1sCOCCcMH" colab_type="code" colab={}
v = data.vocab
v.stoi['xxpad']
def shift_tfm(b):
x,y = b
y = F.pad(y, (1, 0), value=1)
return [x,y[:,:-1]], y[:,1:]
data.add_tfm(shift_tfm)
# + [markdown] id="GsMr5sklEybk" colab_type="text"
# Transformer class, detailed doc can be found in fast.ai library:
# https://github.com/fastai/course-nlp/blob/master/8-translation-transformer.ipynb
# + id="Fj1kmVrRDM09" colab_type="code" outputId="f93f636f-aec1-41d4-a96e-dd4b4156ff5e" executionInfo={"status": "ok", "timestamp": 1576659832106, "user_tz": 300, "elapsed": 1072, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/<KEY>", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
class PositionalEncoding(nn.Module):
"Encode the position with a sinusoid."
def __init__(self, d):
super().__init__()
self.register_buffer('freq', 1 / (10000 ** (torch.arange(0., d, 2.)/d)))
def forward(self, pos):
inp = torch.ger(pos, self.freq)
enc = torch.cat([inp.sin(), inp.cos()], dim=-1)
return enc
tst_encoding = PositionalEncoding(20)
res = tst_encoding(torch.arange(0,100).float())
_, ax = plt.subplots(1,1)
for i in range(1,5): ax.plot(res[:,i])
class TransformerEmbedding(nn.Module):
"Embedding + positional encoding + dropout"
def __init__(self, vocab_sz, emb_sz, inp_p=0.):
super().__init__()
self.emb_sz = emb_sz
self.embed = embedding(vocab_sz, emb_sz)
self.pos_enc = PositionalEncoding(emb_sz)
self.drop = nn.Dropout(inp_p)
def forward(self, inp):
pos = torch.arange(0, inp.size(1), device=inp.device).float()
return self.drop(self.embed(inp) * math.sqrt(self.emb_sz) + self.pos_enc(pos))
def feed_forward(d_model, d_ff, ff_p=0., double_drop=True):
layers = [nn.Linear(d_model, d_ff), nn.ReLU()]
if double_drop: layers.append(nn.Dropout(ff_p))
return SequentialEx(*layers, nn.Linear(d_ff, d_model), nn.Dropout(ff_p), MergeLayer(), nn.LayerNorm(d_model))
class MultiHeadAttention(nn.Module):
def __init__(self, n_heads, d_model, d_head=None, p=0., bias=True, scale=True):
super().__init__()
d_head = ifnone(d_head, d_model//n_heads)
self.n_heads,self.d_head,self.scale = n_heads,d_head,scale
self.q_wgt,self.k_wgt,self.v_wgt = [nn.Linear(
d_model, n_heads * d_head, bias=bias) for o in range(3)]
self.out = nn.Linear(n_heads * d_head, d_model, bias=bias)
self.drop_att,self.drop_res = nn.Dropout(p),nn.Dropout(p)
self.ln = nn.LayerNorm(d_model)
def forward(self, q, kv, mask=None):
return self.ln(q + self.drop_res(self.out(self._apply_attention(q, kv, mask=mask))))
def create_attn_mat(self, x, layer, bs):
return layer(x).view(bs, x.size(1), self.n_heads, self.d_head
).permute(0, 2, 1, 3)
def _apply_attention(self, q, kv, mask=None):
bs,seq_len = q.size(0),q.size(1)
wq,wk,wv = map(lambda o: self.create_attn_mat(*o,bs),
zip((q,kv,kv),(self.q_wgt,self.k_wgt,self.v_wgt)))
attn_score = wq @ wk.transpose(2,3)
if self.scale: attn_score /= math.sqrt(self.d_head)
if mask is not None:
attn_score = attn_score.float().masked_fill(mask, -float('inf')).type_as(attn_score)
attn_prob = self.drop_att(F.softmax(attn_score, dim=-1))
attn_vec = attn_prob @ wv
return attn_vec.permute(0, 2, 1, 3).contiguous().view(bs, seq_len, -1)
def get_output_mask(inp, pad_idx=1):
return torch.triu(inp.new_ones(inp.size(1),inp.size(1)), diagonal=1)[None,None].byte()
# return ((inp == pad_idx)[:,None,:,None].long() + torch.triu(inp.new_ones(inp.size(1),inp.size(1)), diagonal=1)[None,None] != 0)
class EncoderBlock(nn.Module):
"Encoder block of a Transformer model."
#Can't use Sequential directly cause more than one input...
def __init__(self, n_heads, d_model, d_head, d_inner, p=0., bias=True, scale=True, double_drop=True):
super().__init__()
self.mha = MultiHeadAttention(n_heads, d_model, d_head, p=p, bias=bias, scale=scale)
self.ff = feed_forward(d_model, d_inner, ff_p=p, double_drop=double_drop)
def forward(self, x, mask=None): return self.ff(self.mha(x, x, mask=mask))
class DecoderBlock(nn.Module):
"Decoder block of a Transformer model."
#Can't use Sequential directly cause more than one input...
def __init__(self, n_heads, d_model, d_head, d_inner, p=0., bias=True, scale=True, double_drop=True):
super().__init__()
self.mha1 = MultiHeadAttention(n_heads, d_model, d_head, p=p, bias=bias, scale=scale)
self.mha2 = MultiHeadAttention(n_heads, d_model, d_head, p=p, bias=bias, scale=scale)
self.ff = feed_forward(d_model, d_inner, ff_p=p, double_drop=double_drop)
def forward(self, x, enc, mask_out=None): return self.ff(self.mha2(self.mha1(x, x, mask_out), enc))
class Transformer(Module):
def __init__(self, inp_vsz, out_vsz, n_layers=6, n_heads=8, d_model=256, d_head=32,
d_inner=1024, p=0.1, bias=True, scale=True, double_drop=True, pad_idx=1):
self.enc_emb = TransformerEmbedding(inp_vsz, d_model, p)
self.dec_emb = TransformerEmbedding(out_vsz, d_model, 0.)
args = (n_heads, d_model, d_head, d_inner, p, bias, scale, double_drop)
self.encoder = nn.ModuleList([EncoderBlock(*args) for _ in range(n_layers)])
self.decoder = nn.ModuleList([DecoderBlock(*args) for _ in range(n_layers)])
self.out = nn.Linear(d_model, out_vsz)
self.out.weight = self.dec_emb.embed.weight
self.pad_idx = pad_idx
def forward(self, inp, out):
mask_out = get_output_mask(out, self.pad_idx)
enc,out = self.enc_emb(inp),self.dec_emb(out)
enc = compose(self.encoder)(enc)
out = compose(self.decoder)(out, enc, mask_out)
return self.out(out)
# + [markdown] id="EGyWk9SMOMFp" colab_type="text"
# Train the transformer model using default setting and metrics with using accuracy and BLEU to see how model goes, with loss function of cross entropy analysis and apply early stopping to make sure the model still works well for test data
# + id="1CXzMk89mn0h" colab_type="code" colab={}
n_x_vocab,n_y_vocab = len(data.train_ds.x.vocab.itos), len(data.train_ds.y.vocab.itos)
#defalut parameters for Transformer model
model = Transformer(n_x_vocab, n_y_vocab, n_layers=6, n_heads=8, d_model=256, d_head=32,
d_inner=1024)
#here we do cross entropy analysis, you can also do LabelSmoothingCrossEntropy for higher BLEU scores
learn = Learner(data,model,metrics=[accuracy, CorpusBLEU(n_y_vocab)], loss_func = CrossEntropyFlat(),
callback_fns=[partial(EarlyStoppingCallback, monitor='accuracy', min_delta=0.01, patience=3)])
# + [markdown] id="Wb_joMxhOhJ7" colab_type="text"
# Train the model for four cycle. 5e-4 as slice and division fater 5 is the best parameter fine-tuned for language model, detailed in doc https://github.com/fastai/course-nlp/blob/master/8-translation-transformer.ipynb. Here we use four cycles for better performance. Approximately taking ~30 mins
# + id="9-T6w6NHnIZ7" colab_type="code" outputId="bdfbfe43-2c44-4675-f9a1-7e067c5cb158" executionInfo={"status": "ok", "timestamp": 1576661550105, "user_tz": 300, "elapsed": 1712221, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 170}
learn.fit_one_cycle(4, 5e-4, div_factor=5)
# + [markdown] id="fk61bqyWOn84" colab_type="text"
# Get prediction of the learn model stored in learn.data.train_ds.y
# + id="QH4DrzXl7Iih" colab_type="code" colab={}
def get_predictions(learn, ds_type=DatasetType.Valid):
learn.model.eval()
inputs, targets, outputs = [],[],[]
with torch.no_grad():
for xb,yb in progress_bar(learn.dl(ds_type)):
out = learn.model(*xb)
for x,y,z in zip(xb[0],xb[1],out):
inputs.append(learn.data.train_ds.x.reconstruct(x))
targets.append(learn.data.train_ds.y.reconstruct(y))
outputs.append(learn.data.train_ds.y.reconstruct(z.argmax(1)))
return inputs, targets, outputs
# + [markdown] id="q9EGUgonrOJQ" colab_type="text"
# Compare input tokens, target code and real output code of
# the model, based on index
# + id="VFsKd8Mv7Yg3" colab_type="code" outputId="caa097e8-1f4a-4bd8-9de2-5b6516c9b4c5" executionInfo={"status": "ok", "timestamp": 1576663383554, "user_tz": 300, "elapsed": 126794, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 37}
inputs, targets, outputs = get_predictions(learn)
# + [markdown] id="6LkArH3UdE1g" colab_type="text"
# Load the two sentencePiece processors to decode the inputs, targets and outputs for predictions
# + id="Kv10CHzOX6c7" colab_type="code" outputId="b5148b3d-5884-40ab-952f-754bb13f72f6" executionInfo={"status": "ok", "timestamp": 1576662769863, "user_tz": 300, "elapsed": 264, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
sp1 = spm.SentencePieceProcessor()
sp1.Load("train_py.model")
sp2 = spm.SentencePieceProcessor()
sp2.Load("train_docstring_py.model")
# + [markdown] id="_W-dktBUdS3V" colab_type="text"
# Make Text object into list and decode it with corresponding preprocessor. Print the result
# + id="d5-SFM90YeSR" colab_type="code" colab={}
def print_result(inputt, targett, outputt):
i = sp2.decode_pieces(list(inputt.text.split(" ")))
t = sp1.decode_pieces(list(targett.text.split(" ")))
o = sp1.decode_pieces(list(outputt.text.split(" ")))
print("Input docstring: "+i)
print("Target code snippets: "+t)
print("Output code snippets: "+o)
# + [markdown] id="J0YAlTyjLn0n" colab_type="text"
# We can see here that we actually get a incredibly good prediction based on the input docstring. Target and our models' output are nearly the same, for the followin three show cases (you can do more tests by just changing the index)
# + id="Z5Ruin2CYIgm" colab_type="code" outputId="99265916-eb37-4fec-8732-550623ae96cb" executionInfo={"status": "ok", "timestamp": 1576663027114, "user_tz": 300, "elapsed": 588, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
print_result(inputs[10],targets[10],outputs[10])
# + id="h2QI7M23FQXm" colab_type="code" outputId="370c2252-4407-4ea6-86f5-8ff9a5f885f1" executionInfo={"status": "ok", "timestamp": 1576663100318, "user_tz": 300, "elapsed": 335, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
print_result(inputs[1002],targets[1002],outputs[1002])
# + id="oneaznnQWlMj" colab_type="code" outputId="b55252d0-2034-4752-8d87-ce749cf804c7" executionInfo={"status": "ok", "timestamp": 1576663103412, "user_tz": 300, "elapsed": 346, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBeYLkFQX0rmIjxn7RbCQdOA8NbZZ0B0yu9dHdr=s64", "userId": "06966794032599656492"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
print_result(inputs[4002],targets[4002],outputs[4002])
| main/ArtificialCodeGen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Extracting tabular data from OCR-processed PDFs with Python
#
# This notebook explores a method of extracting tabular data from OCR-processed PDF files using Python. We will use two key software libraries: [poppler](https://poppler.freedesktop.org/) and [pdftabextract](https://pypi.python.org/pypi/pdftabextract/). Poppler will be used to extract the images from the PDF files and pdftabextract will be used to extract the tabular data. We will then anaylse the successfulness of this method, how feasible it is to perform on a large scale and what might be done to improve it.
#
# The dataset that we will use contains samples of digitised material from the [India Office Medical Archives](https://www.bl.uk/collection-guides/india-office-medical-archive-collections). The dataset was created by <NAME> (2017) and is available for [download](https://data.bl.uk/indiaofficemedicalarchives/ioma3.html) via data.bl.uk. The printed material in this dataset contains tabular data on medical topography, which we will attempt to extract for further analysis. An example of a digitised image from the collection is presented below.
#
# 
# ## Prerequisites
#
# This tutorial assumes that you already have [Python](https://www.python.org/) installed and have some familiarity with running Python scripts.
#
# With that in mind, we will start by installing the required software libraries via [PyPi](https://pypi.python.org/pypi). Open up a command-line interface and run the code below. There is no need to include the exclamation mark at the start of the code block, which is used to install the libraries within this notebook.
# !pip install requests tqdm pdftabextract numpy pandas opencv-python --quiet
# ## Import the libraries
#
# We can now start writing our Python script.
#
# Using a text editor, create a new file and save it as `run.py`, then enter the following.
# +
import os
import re
import requests
import tqdm
import zipfile
import cv2
import pandas
import numpy as np
import pdftabextract
from pdftabextract import imgproc
from pdftabextract.geom import pt
from pdftabextract.common import read_xml, parse_pages, all_a_in_b, save_page_grids
from pdftabextract.common import DIRECTION_VERTICAL, ROTATION, SKEW_X, SKEW_Y
from pdftabextract.textboxes import rotate_textboxes, deskew_textboxes
from pdftabextract.textboxes import border_positions_from_texts, split_texts_by_positions, join_texts
from pdftabextract.clustering import calc_cluster_centers_1d, find_clusters_1d_break_dist, zip_clusters_and_values
from pdftabextract.extract import make_grid_from_positions, fit_texts_into_grid, datatable_to_dataframe
from math import radians, degrees
# -
# # Declare some common variables
#
# There are some common variables that will be used in various places throughout the tutorial. We will declare these below the imports for easier reference. The comments above each variable indicate their purpose.
# +
# The directory to which we will download our dataset.
DATA_DIR = '../data'
# An HTTP header that we add to identify our application over a network.
USER_AGENT = 'bl-digischol-notebooks'
# The name of the dataset collection
COLLECTION = 'indiaofficemedicalarchives'
# The name of the dataset
DATASET = 'ioma-samples-small'
# -
# ## Prepare the dataset
#
# We now need to download our dataset and extract the files is contains. The code block below will handle this programmatically. Copy the code into your Python script, save the file, then open up a command-line interface, navigate to the location of your script and run the following:
#
# ```
# python run.py
# ```
#
# Assuming the dataset does not already exist in the correct location it will be downloaded and the files extracted. For more details about how the process works, see [Downloading datasets with Python](downloading_datasets_with_python.ipynb).
# +
def create_data_dir(directory):
if not os.path.exists(directory):
os.mkdir(directory)
def download_dataset(collection, dataset, directory, user_agent):
url = 'https://data.bl.uk/{0}/{1}.zip'.format(collection, dataset)
download_fn = url.split('/')[-1]
download_path = os.path.join(directory, download_fn)
if not os.path.exists(download_path):
headers = {'User-agent': user_agent}
r = requests.get(url, stream=True, headers=headers)
total_length = int(r.headers.get('Content-Length'))
total_size = (total_length/1024) + 1
with open(download_path, 'wb') as f:
for chunk in tqdm.tqdm(r.iter_content(chunk_size=1024),
total=total_size,
desc='Downloading',
unit='kb',
unit_scale=True,
miniters=1,
leave=False):
if chunk:
f.write(chunk)
def extract_dataset(dataset, collection, data_dir):
fn = '{}.zip'.format(dataset)
in_path = os.path.join(data_dir, fn)
out_path = os.path.join(data_dir, collection)
with zipfile.ZipFile(in_path) as archive:
unextracted = [name for name in archive.namelist()
if not os.path.exists(os.path.join(out_path, name))]
if unextracted:
for i in tqdm.tqdm(range(len(unextracted)),
desc='Extracting',
unit='file',
leave=False):
archive.extract(unextracted[i], path=out_path)
create_data_dir(DATA_DIR)
download_dataset(COLLECTION, DATASET, DATA_DIR, USER_AGENT)
extract_dataset(DATASET, COLLECTION, DATA_DIR)
# -
# ## Extract the images from the PDF files
#
# As a first step to accessing OCR data embedded in the image files we need to extract the images from our PDF files and convert the OCR data to XML. To do this we use a PDF rendering library called [poppler](https://github.com/davidben/poppler), which is available in most Linux distributions.
#
# Note that if running this notebook on a Windows machine, the latest binaries for poppler can be downloaded [here](http://blog.alivate.com.au/poppler-windows/). For Mac, install [Homebrew](https://brew.sh/), open a terminal and run `brew install poppler`.
# +
def extract_images(data_dir, collection):
base_dir = os.path.join(data_dir, collection)
pdfs = [fn for fn in os.listdir(base_dir) if fn.endswith('.pdf')]
xmls = ['{}.xml'.format(os.path.splitext(pdf)[-2]) for pdf in pdfs]
unconverted = [fn for fn in xmls if not os.path.exists(os.path.join(base_dir, fn))]
if unconverted:
for fn in tqdm.tqdm(unconverted, desc='Converting', unit='file', leave=False):
pdf_path = os.path.join(base_dir, '{}.pdf'.format(os.path.splitext(fn)[-2]))
xml_path = os.path.join(base_dir, fn)
script = 'C:/users/amendes/downloads/poppler-0.51/bin/pdftohtml -c -q -hidden -xml "{0}" "{1}"'.format(pdf_path, xml_path)
# !{script}
extract_images(DATA_DIR, COLLECTION)
# -
# We should now have an XML file containing our OCR data, along with a JPEG image for each page of the PDF.
# ## Detect lines in the images
#
# The following code is largely taken from the online tutorial [Data Mining OCR PDFs — Using pdftabextract to Liberate Tabular Data from Scanned Documents](https://datascience.blog.wzb.eu/2017/02/16/data-mining-ocr-pdfs-using-pdftabextract-to-liberate-tabular-data-from-scanned-documents/) (<NAME>, 2017), which includes a detailed explanation about the programatic process below.
#
# Essentially, we identify the lines in each image, attempt to fix any rotation or skewing, cluster similar lines, identify any grids and output the data contanied within those grids to a set of tables. The result should be the conversion of data from printed to electronic tables.
# helper function to save an image
def save_image_w_lines(iproc_obj, imgfilebasename):
img_lines = iproc_obj.draw_lines(orig_img_as_background=True)
img_lines_file = os.path.join('%s-lines-orig.png' % imgfilebasename)
cv2.imwrite(img_lines_file, img_lines)
def detect_lines(iproc_obj, img_path, metadata):
# calculate the scaling of the image file in relation to the text boxes
page_scaling_x = iproc_obj.img_w / metadata['width']
page_scaling_y = iproc_obj.img_h / metadata['height']
# detect the lines
lines_hough = iproc_obj.detect_lines(canny_kernel_size=3, canny_low_thresh=50, canny_high_thresh=150,
hough_rho_res=1,
hough_theta_res=np.pi/500,
hough_votes_thresh=int(round(0.2 * iproc_obj.img_w)))
# save the image with detected lines
save_image_w_lines(iproc_obj, img_base_fn)
# ## Fix rotation or skew
#
# If the tables are rotated or skewed we'll have a hard time extracting the data from them. The following code fixes this...
def fix_rotation_and_skew(iproc_obj, img_base_fn, metadata, xml, xmltree):
rot_or_skew_type, rot_or_skew_radians = iproc_obj.find_rotation_or_skew(radians(0.5),
radians(1),
omit_on_rot_thresh=radians(0.5))
# rotate back or deskew text boxes
needs_fix = True
if rot_or_skew_type == ROTATION:
rotate_textboxes(metadata, -rot_or_skew_radians, pt(0, 0))
elif rot_or_skew_type in (SKEW_X, SKEW_Y):
deskew_textboxes(metadata, -rot_or_skew_radians, rot_or_skew_type, pt(0, 0))
else:
needs_fix = False
if needs_fix:
# rotate back or deskew detected lines
lines_hough = iproc_obj.apply_found_rotation_or_skew(rot_or_skew_type,
-rot_or_skew_radians)
save_image_w_lines(iproc_obj, img_base_fn + '-repaired')
out_base_fn = xml[:xml.rindex('.')]
repaired_xmlfile = os.path.join(out_base_fn + '.repaired.xml')
xmltree.write(repaired_xmlfile)
def cluster_lines(iproc_obj, img_base_fn, metadata):
MIN_COL_WIDTH = 60 # minimum width of a column in pixels
# cluster the detected *vertical* lines using find_clusters_1d_break_dist as simple clustering function
page_scaling_x = iproc_obj.img_w / metadata['width']
vertical_clusters = iproc_obj.find_clusters(imgproc.DIRECTION_VERTICAL, find_clusters_1d_break_dist,
remove_empty_cluster_sections_use_texts=metadata['texts'],
remove_empty_cluster_sections_n_texts_ratio=0.1,
remove_empty_cluster_sections_scaling=page_scaling_x,
dist_thresh=MIN_COL_WIDTH/2)
# draw the clusters
try:
img_w_clusters = iproc_obj.draw_line_clusters(imgproc.DIRECTION_VERTICAL, vertical_clusters)
except Exception as e:
print('ERROR:' + e.message)
return
save_img_file = os.path.join('%s-vertical-clusters.png' % img_base_fn)
cv2.imwrite(save_img_file, img_w_clusters)
return vertical_clusters
def find_row_positions(iproc_obj, metadata, vertical_clusters):
page_scaling_x = iproc_obj.img_w / metadata['width']
page_colpos = np.array(calc_cluster_centers_1d(vertical_clusters)) / page_scaling_x
# right border of the second column
col2_rightborder = page_colpos[2]
# calculate median text box height
median_text_height = np.median([t['height'] for t in metadata['texts']])
# get all texts in the first two columns with a "usual" textbox height
# we will only use these text boxes in order to determine the line positions because they are more "stable"
# otherwise, especially the right side of the column header can lead to problems detecting the first table row
text_height_deviation_thresh = median_text_height / 2
texts_cols_1_2 = [t for t in metadata['texts']
if t['right'] <= col2_rightborder
and abs(t['height'] - median_text_height) <= text_height_deviation_thresh]
# get all textboxes' top and bottom border positions
borders_y = border_positions_from_texts(texts_cols_1_2, DIRECTION_VERTICAL)
# break into clusters using half of the median text height as break distance
clusters_y = find_clusters_1d_break_dist(borders_y, dist_thresh=median_text_height/2)
clusters_w_vals = zip_clusters_and_values(clusters_y, borders_y)
# for each cluster, calculate the median as center
pos_y = calc_cluster_centers_1d(clusters_w_vals)
pos_y.append(metadata['height'])
print('number of line positions:', len(pos_y))
# a (possibly malformed) population number + space + start of city name
pttrn_table_row_beginning = re.compile(r'^[\d Oo][\d Oo]{2,} +[A-ZÄÖÜ]')
# 1. try to find the top row of the table
texts_cols_1_2_per_line = split_texts_by_positions(texts_cols_1_2, pos_y, DIRECTION_VERTICAL,
alignment='middle',
enrich_with_positions=True)
# go through the texts line per line
for line_texts, (line_top, line_bottom) in texts_cols_1_2_per_line:
line_str = join_texts(line_texts)
if pttrn_table_row_beginning.match(line_str): # check if the line content matches the given pattern
top_y = line_top
break
else:
top_y = 0
# hints for a footer text box
words_in_footer = ('anzeige', 'annahme', 'ala')
# 2. try to find the bottom row of the table
min_footer_text_height = median_text_height * 1.5
min_footer_y_pos = metadata['height'] * 0.7
# get all texts in the lower 30% of the page that have are at least 50% bigger than the median textbox height
bottom_texts = [t for t in metadata['texts']
if t['top'] >= min_footer_y_pos and t['height'] >= min_footer_text_height]
bottom_texts_per_line = split_texts_by_positions(bottom_texts,
pos_y + [metadata['height']], # always down to the end of the page
DIRECTION_VERTICAL,
alignment='middle',
enrich_with_positions=True)
# go through the texts at the bottom line per line
page_span = page_colpos[-1] - page_colpos[0]
min_footer_text_width = page_span * 0.8
for line_texts, (line_top, line_bottom) in bottom_texts_per_line:
line_str = join_texts(line_texts)
has_wide_footer_text = any(t['width'] >= min_footer_text_width for t in line_texts)
# check if there's at least one wide text or if all of the required words for a footer match
if has_wide_footer_text or all_a_in_b(words_in_footer, line_str):
bottom_y = line_top
break
else:
bottom_y = metadata['height']
page_rowpos = [y for y in pos_y if top_y <= y <= bottom_y]
return page_colpos, page_rowpos
def create_grid(page_colpos, page_rowpos, page_num, img_base_fn):
grid = make_grid_from_positions(page_colpos, page_rowpos)
if not grid:
return
n_rows = len(grid)
n_cols = len(grid[0])
page_grids_file = '{0}.pagegrids_p{1}.json'.format(img_base_fn, page_num)
save_page_grids({page_num: grid}, page_grids_file)
return grid
# +
def extract_tables(data_dir, collection):
base_dir = os.path.join(data_dir, collection)
xmls = [fn for fn in os.listdir(base_dir) if os.path.splitext(fn)[-1] == '.xml']
data = []
for xml in xmls:
xml_path = os.path.join(base_dir, xml)
xmltree, xmlroot = read_xml(xml_path)
pages = parse_pages(xmlroot)
for page_num, page in tqdm.tqdm(enumerate(pages),
desc='Processing',
unit='page',
unit_scale=True,
miniters=1,
leave=False):
if page_num != 27:
continue
metadata = pages[page_num]
img_path = metadata['image']
img_base_fn = img_path[:img_path.rindex('.')]
print img_base_fn
iproc_obj = imgproc.ImageProc(img_path)
detect_lines(iproc_obj, img_base_fn, metadata)
fix_rotation_and_skew(iproc_obj, img_base_fn, metadata, xml, xmltree)
vertical_clusters = cluster_lines(iproc_obj, img_base_fn, metadata)
page_colpos, page_rowpos = find_row_positions(iproc_obj, metadata, vertical_clusters)
grid = create_grid(page_colpos, page_rowpos, page_num, img_base_fn)
if grid:
datatable = fit_texts_into_grid(metadata['texts'], grid)
df = datatable_to_dataframe(datatable)
print df
extract_tables(DATA_DIR, COLLECTION)
# -
#
# ## Bibliography
#
# <NAME>. (2017) *Data Mining OCR PDFs — Using pdftabextract to Liberate Tabular Data from Scanned Documents*, WZB Science Blog, February 16, 2017, https://datascience.blog.wzb.eu/2017/02/16/data-mining-ocr-pdfs-using-pdftabextract-to-liberate-tabular-data-from-scanned-documents/
#
# <NAME>. (2017) *India Office Medical Archives samples (small)*. British Library. https://doi.org/10.21250/ioma3
| notebooks/extracting_tabular_data_with_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="IeTl5GZz7dAx" colab_type="code" colab={}
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm as tqdm
# %matplotlib inline
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import random
from torch.utils.data import Dataset, DataLoader
# + id="TIGGAsNe-0nt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="ac3a8846-62f8-4e45-a43f-9b716afb8d4f"
from google.colab import drive
drive.mount('/content/drive')
# + id="QPDTZIA97dA8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="b8cf34c4-7745-425f-e5da-3574cc01091e"
x11 = np.random.uniform(low=[0,0], high = [0.5,0.5],size =(15,2) )
x12 = np.random.uniform(low=[0.5,0.5], high = [1,1],size =(15,2) )
x2 = np.random.uniform(low = [0,1.5] , high = [1,2.5],size=(30,2))
x3 = np.random.uniform(low = [0,3] , high = [1,4],size=(30,2))
x4 = np.random.uniform(low = [2,0] , high = [3,1],size=(30,2))
x5 = np.random.uniform(low = [2,1.5] , high = [3,2.5],size=(30,2))
x6 = np.random.uniform(low = [2,3] , high = [3,4],size=(30,2))
x7 = np.random.uniform(low = [4,0] , high = [5,1],size=(30,2))
x8 = np.random.uniform(low = [4,1.5] , high = [5,2.5],size=(30,2))
x9 = np.random.uniform(low = [4,3] , high = [5,4],size=(30,2))
plt.scatter(x11[:,0],x11[:,1])
plt.scatter(x12[:,0],x12[:,1])
plt.scatter(x2[:,0],x2[:,1])
plt.scatter(x3[:,0],x3[:,1])
plt.scatter(x4[:,0],x4[:,1])
plt.scatter(x5[:,0],x5[:,1])
plt.scatter(x6[:,0],x6[:,1])
plt.scatter(x7[:,0],x7[:,1])
plt.scatter(x8[:,0],x8[:,1])
plt.scatter(x9[:,0],x9[:,1])
# + id="SFmyt1B57dBG" colab_type="code" colab={}
y11 = np.zeros(15)
y12 = np.ones(15)
Y2_ = []
for i in range(8):
idx = np.random.randint(0,30,size=15)
y2 = np.ones(30)
y2[idx] = 0
Y2_.append(y2)
Y2_ = np.concatenate(Y2_,axis=0)
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
# + id="0HNClf_b7dBN" colab_type="code" colab={}
X_train = np.concatenate((x11,x12,x2,x3,x4,x5,x6,x7,x8,x9))
Y_train = np.concatenate((y11,y12,Y2_))
# + id="t7hbwnrb7dBZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4d127662-55ae-40bc-8c1e-254418b3cc9a"
X_train.shape,Y_train.shape
# + id="1IJoDagt7dBe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="9f50b23f-5547-434f-8d2c-0dc8fde3ec40"
plt.scatter(X_train[Y_train==0,0],X_train[Y_train==0,1],label = "class_0")
plt.scatter(X_train[Y_train==1,0],X_train[Y_train==1,1],label = "class_1")
plt.legend()
# + id="IZUnbahp7dBk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="a73fc59b-cb71-4ef7-dcb0-5f104d7a75dc"
x11 = np.random.uniform(low=[0,0], high = [0.5,0.5],size =(15,2) )
x12 = np.random.uniform(low=[0.5,0.5], high = [1,1],size =(15,2) )
x2 = np.random.uniform(low = [0,1.5] , high = [1,2.5],size=(30,2))
x3 = np.random.uniform(low = [0,3] , high = [1,4],size=(30,2))
x4 = np.random.uniform(low = [2,0] , high = [3,1],size=(30,2))
x5 = np.random.uniform(low = [2,1.5] , high = [3,2.5],size=(30,2))
x6 = np.random.uniform(low = [2,3] , high = [3,4],size=(30,2))
x7 = np.random.uniform(low = [4,0] , high = [5,1],size=(30,2))
x8 = np.random.uniform(low = [4,1.5] , high = [5,2.5],size=(30,2))
x9 = np.random.uniform(low = [4,3] , high = [5,4],size=(30,2))
plt.scatter(x11[:,0],x11[:,1])
plt.scatter(x12[:,0],x12[:,1])
plt.scatter(x2[:,0],x2[:,1])
plt.scatter(x3[:,0],x3[:,1])
plt.scatter(x4[:,0],x4[:,1])
plt.scatter(x5[:,0],x5[:,1])
plt.scatter(x6[:,0],x6[:,1])
plt.scatter(x7[:,0],x7[:,1])
plt.scatter(x8[:,0],x8[:,1])
plt.scatter(x9[:,0],x9[:,1])
yt11 = np.zeros(15)
yt12 = np.ones(15)
Yt2_ = []
for i in range(8):
idx = np.random.randint(0,30,size=15)
yt2 = np.ones(30)
yt2[idx] = 0
Yt2_.append(yt2)
Yt2_ = np.concatenate(Yt2_,axis=0)
Y_test = np.concatenate((yt11,yt12,Yt2_))
X_test = np.concatenate((x11,x12,x2,x3,x4,x5,x6,x7,x8,x9))
# + id="MuDuqKzi7dBt" colab_type="code" colab={}
class Grid_data(Dataset):
def __init__(self,x,y):
self.x = torch.Tensor(x)
self.y = torch.Tensor(y).type(torch.LongTensor)
def __len__(self):
return len(self.x)
def __getitem__(self,idx):
self.dx = self.x[idx,:]
self.dy = self.y[idx]
self.dx = self.dx
self.dy = self.dy
return self.dx, self.dy
# + id="1GJ4sl0U7dBy" colab_type="code" colab={}
trainset = Grid_data(X_train,Y_train)
# + id="cHVvhJ5o7dB2" colab_type="code" colab={}
trainloader = DataLoader(trainset,batch_size=30,shuffle = False)
# + id="piqsayfW7dB8" colab_type="code" colab={}
inputs,label = iter(trainloader).next()
# + id="eINcPHl87dCA" colab_type="code" colab={}
class Net(nn.Module):
def __init__(self):
super(Net,self).__init__()
self.linear1 = nn.Linear(2,24)
self.linear2 = nn.Linear(24,64)
self.linear3 = nn.Linear(64,128)
self.linear4 = nn.Linear(128,256)
self.linear5 = nn.Linear(256,128)
self.linear6 = nn.Linear(128,64)
self.linear7 = nn.Linear(64,32)
self.linear8 = nn.Linear(32,16)
self.linear9 = nn.Linear(16,2)
def forward(self,x):
x = F.relu(self.linear1(x))
x = F.relu(self.linear2(x))
x = F.relu(self.linear3(x))
x = F.relu(self.linear4(x))
x = F.relu(self.linear5(x))
x = F.relu(self.linear6(x))
x = F.relu(self.linear7(x))
x = F.relu(self.linear8(x))
x = self.linear9(x)
return x
# + id="z66CySFG7dCF" colab_type="code" colab={}
net = Net()
# net(inputs)
# + id="zsRUIIjf7dCJ" colab_type="code" colab={}
net = net.to("cuda")
# + id="3zLOvLuT7dCM" colab_type="code" colab={}
criterion = nn.CrossEntropyLoss()
lr=0.001
optimizer = optim.Adam(net.parameters(), lr)#, momentum=0.9)
# + id="qTTpwkpH7dCP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0ca913ae-70ec-494d-f00d-7dd063e458bf"
loss_curi = []
epochs_nos= 10000
for epoch in range(epochs_nos): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 3 == 2: # print every 50 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss/2 ))
ep_lossi.append(running_loss/2) # loss per minibatch
running_loss = 0.0
if(np.mean(ep_lossi)<= 0.3):
epochs_nos = epoch
break
loss_curi.append(np.mean(ep_lossi)) #loss per epoch
# if (epoch%5 == 0):
# _,actis= inc(inputs)
# acti.append(actis)
print('Finished Training')
# + id="sV4LeKuW7dCU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4fdfb7d9-a8c3-499f-9595-910fd44fe876"
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
out.append(labels.cpu().numpy())
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 60000 train images: %d %%' % (100 * correct / total))
train_acc = 100 * correct / total
# + id="oOhAovWY7dCX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3df2891d-8b94-4ef0-95cf-2adac8eec1a1"
total,correct
# + id="IIbabwdC7dCa" colab_type="code" colab={}
out = np.concatenate(out,axis=0)
pred = np.concatenate(pred,axis=0)
# + id="9trYeVn37dCe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="c503e661-08ef-4bfe-a86a-3f46dba2851c"
out[:30]
# + id="VwJLLQOC7dCj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="af6633c1-9592-4ae4-e0e3-f96a6812ce77"
pred[:30]
# + id="--AqzJs77dCn" colab_type="code" colab={}
X_axis,Y_axis = np.meshgrid(np.arange(0,5,0.01),np.arange(0,5,0.01))
X_mesh = np.concatenate( (X_axis.reshape((-1,1)), Y_axis.reshape(-1,1)), axis=1 )
Y_mesh = np.zeros(X_mesh.shape[0])
# + id="dfzrda_N7dCp" colab_type="code" colab={}
mesh_set = Grid_data(X_mesh,Y_mesh)
meshloader = DataLoader(mesh_set,batch_size=1000,shuffle=False)
# + id="L1DtTf227dCt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c4b04598-e8d0-4100-9294-7bfc9a098106"
total = 0
mesh_pred = []
with torch.no_grad():
for data in meshloader:
images, _ = data
images = images.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
mesh_pred.append(predicted.cpu().numpy())
total += labels.size(0)
print("finished")
# + id="YFuxZ_mP7dCw" colab_type="code" colab={}
mesh_pred = np.concatenate(mesh_pred,axis=0)
# mesh_pred = mesh_pred.reshape(X.shape)
# + id="dgBR0eX_7dCz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="a4f017c0-0c83-4e6f-f169-6e67aec43c6c"
# plt.axis('equal')
plt.scatter(X_mesh[:,0],X_mesh[:,1],c= mesh_pred, cmap = 'RdGy' )
plt.scatter(X_train[Y_train==1,0], X_train[Y_train==1,1] ,c="red")
plt.scatter(X_train[Y_train==0,0], X_train[Y_train==0,1],c= "green" )
# + id="mAXCZTE_7dC1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a2960906-d896-4d44-e5ce-7c1695263896"
X_test.shape
# + id="yWR6ZHfk7dC4" colab_type="code" colab={}
testset = Grid_data(X_test,Y_test)
testloader = DataLoader(testset,batch_size=10,shuffle=False)
# + id="wBlYxTjq7dC7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="208a631a-33c4-4909-802f-a34220c9e1f7"
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
out.append(labels.cpu().numpy())
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %d %%' % (270, 100 * correct / total))
# + id="uWN0I0Va7dC-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="1a5caf15-0837-4f6c-c066-b8b022226f9d"
plt.scatter(X_mesh[:,0],X_mesh[:,1],c= mesh_pred, cmap = "Greys")
plt.scatter(X_test[Y_test==1,0], X_test[Y_test==1,1] ,c="Blue")
plt.scatter(X_test[Y_test==0,0], X_test[Y_test==0,1],c= "green" )
# + id="GVyOS5Ln7dDB" colab_type="code" colab={}
torch.save(net.state_dict(),"/content/drive/My Drive/Research/confounded_noise/weights/model_"+str(epochs_nos)+"_lr_"+str(lr)+"_acc_"+str(train_acc)+".pkl")
# + id="m6H4h1aMEISj" colab_type="code" colab={}
| 6_confounded_noise/linear_separable_at_corner/confounded_noise_acc_90.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
# 3. NLP in Practice
#
# ___
# # Part of Speech Basics
# The challenge of correctly identifying parts of speech is summed up nicely in the [spaCy docs](https://spacy.io/usage/linguistic-features):
# <div class="alert alert-info" style="margin: 20px">Processing raw text intelligently is difficult: most words are rare, and it's common for words that look completely different to mean almost the same thing. The same words in a different order can mean something completely different. Even splitting text into useful word-like units can be difficult in many languages. While it's possible to solve some problems starting from only the raw characters, it's usually better to use linguistic knowledge to add useful information. That's exactly what spaCy is designed to do: you put in raw text, and get back a **Doc** object, that comes with a variety of annotations.</div>
# In this section we'll take a closer look at coarse POS tags (noun, verb, adjective) and fine-grained tags (plural noun, past-tense verb, superlative adjective).
# Perform standard imports
import spacy
nlp = spacy.load('en_core_web_sm')
# Create a simple Doc object
doc = nlp(u"The quick brown fox jumped over the lazy dog's back.")
# ## View token tags
# Recall that you can obtain a particular token by its index position.
# * To view the coarse POS tag use `token.pos_`
# * To view the fine-grained tag use `token.tag_`
# * To view the description of either type of tag use `spacy.explain(tag)`
#
# <div class="alert alert-success">Note that `token.pos` and `token.tag` return integer hash values; by adding the underscores we get the text equivalent that lives in **doc.vocab**.</div>
# Print the full text:
print(doc.text)
# Print the fifth word and associated tags:
print(doc[4].text, doc[4].pos_, doc[4].tag_, spacy.explain(doc[4].tag_))
# We can apply this technique to the entire Doc object:
for token in doc:
print(f'{token.text:{10}} {token.pos_:{8}} {token.tag_:{6}} {spacy.explain(token.tag_)}')
# ## Coarse-grained Part-of-speech Tags
# Every token is assigned a POS Tag from the following list:
#
#
# <table><tr><th>POS</th><th>DESCRIPTION</th><th>EXAMPLES</th></tr>
#
# <tr><td>ADJ</td><td>adjective</td><td>*big, old, green, incomprehensible, first*</td></tr>
# <tr><td>ADP</td><td>adposition</td><td>*in, to, during*</td></tr>
# <tr><td>ADV</td><td>adverb</td><td>*very, tomorrow, down, where, there*</td></tr>
# <tr><td>AUX</td><td>auxiliary</td><td>*is, has (done), will (do), should (do)*</td></tr>
# <tr><td>CONJ</td><td>conjunction</td><td>*and, or, but*</td></tr>
# <tr><td>CCONJ</td><td>coordinating conjunction</td><td>*and, or, but*</td></tr>
# <tr><td>DET</td><td>determiner</td><td>*a, an, the*</td></tr>
# <tr><td>INTJ</td><td>interjection</td><td>*psst, ouch, bravo, hello*</td></tr>
# <tr><td>NOUN</td><td>noun</td><td>*girl, cat, tree, air, beauty*</td></tr>
# <tr><td>NUM</td><td>numeral</td><td>*1, 2017, one, seventy-seven, IV, MMXIV*</td></tr>
# <tr><td>PART</td><td>particle</td><td>*'s, not,*</td></tr>
# <tr><td>PRON</td><td>pronoun</td><td>*I, you, he, she, myself, themselves, somebody*</td></tr>
# <tr><td>PROPN</td><td>proper noun</td><td>*Mary, John, London, NATO, HBO*</td></tr>
# <tr><td>PUNCT</td><td>punctuation</td><td>*., (, ), ?*</td></tr>
# <tr><td>SCONJ</td><td>subordinating conjunction</td><td>*if, while, that*</td></tr>
# <tr><td>SYM</td><td>symbol</td><td>*$, %, §, ©, +, −, ×, ÷, =, :), 😝*</td></tr>
# <tr><td>VERB</td><td>verb</td><td>*run, runs, running, eat, ate, eating*</td></tr>
# <tr><td>X</td><td>other</td><td>*sfpksdpsxmsa*</td></tr>
# <tr><td>SPACE</td><td>space</td></tr>
# ___
# ## Fine-grained Part-of-speech Tags
# Tokens are subsequently given a fine-grained tag as determined by morphology:
# <table>
# <tr><th>POS</th><th>Description</th><th>Fine-grained Tag</th><th>Description</th><th>Morphology</th></tr>
# <tr><td>ADJ</td><td>adjective</td><td>AFX</td><td>affix</td><td>Hyph=yes</td></tr>
# <tr><td>ADJ</td><td></td><td>JJ</td><td>adjective</td><td>Degree=pos</td></tr>
# <tr><td>ADJ</td><td></td><td>JJR</td><td>adjective, comparative</td><td>Degree=comp</td></tr>
# <tr><td>ADJ</td><td></td><td>JJS</td><td>adjective, superlative</td><td>Degree=sup</td></tr>
# <tr><td>ADJ</td><td></td><td>PDT</td><td>predeterminer</td><td>AdjType=pdt PronType=prn</td></tr>
# <tr><td>ADJ</td><td></td><td>PRP\$</td><td>pronoun, possessive</td><td>PronType=prs Poss=yes</td></tr>
# <tr><td>ADJ</td><td></td><td>WDT</td><td>wh-determiner</td><td>PronType=int rel</td></tr>
# <tr><td>ADJ</td><td></td><td>WP\$</td><td>wh-pronoun, possessive</td><td>Poss=yes PronType=int rel</td></tr>
# <tr><td>ADP</td><td>adposition</td><td>IN</td><td>conjunction, subordinating or preposition</td><td></td></tr>
# <tr><td>ADV</td><td>adverb</td><td>EX</td><td>existential there</td><td>AdvType=ex</td></tr>
# <tr><td>ADV</td><td></td><td>RB</td><td>adverb</td><td>Degree=pos</td></tr>
# <tr><td>ADV</td><td></td><td>RBR</td><td>adverb, comparative</td><td>Degree=comp</td></tr>
# <tr><td>ADV</td><td></td><td>RBS</td><td>adverb, superlative</td><td>Degree=sup</td></tr>
# <tr><td>ADV</td><td></td><td>WRB</td><td>wh-adverb</td><td>PronType=int rel</td></tr>
# <tr><td>CONJ</td><td>conjunction</td><td>CC</td><td>conjunction, coordinating</td><td>ConjType=coor</td></tr>
# <tr><td>DET</td><td>determiner</td><td>DT</td><td>determiner</td><td></td></tr>
# <tr><td>INTJ</td><td>interjection</td><td>UH</td><td>interjection</td><td></td></tr>
# <tr><td>NOUN</td><td>noun</td><td>NN</td><td>noun, singular or mass</td><td>Number=sing</td></tr>
# <tr><td>NOUN</td><td></td><td>NNS</td><td>noun, plural</td><td>Number=plur</td></tr>
# <tr><td>NOUN</td><td></td><td>WP</td><td>wh-pronoun, personal</td><td>PronType=int rel</td></tr>
# <tr><td>NUM</td><td>numeral</td><td>CD</td><td>cardinal number</td><td>NumType=card</td></tr>
# <tr><td>PART</td><td>particle</td><td>POS</td><td>possessive ending</td><td>Poss=yes</td></tr>
# <tr><td>PART</td><td></td><td>RP</td><td>adverb, particle</td><td></td></tr>
# <tr><td>PART</td><td></td><td>TO</td><td>infinitival to</td><td>PartType=inf VerbForm=inf</td></tr>
# <tr><td>PRON</td><td>pronoun</td><td>PRP</td><td>pronoun, personal</td><td>PronType=prs</td></tr>
# <tr><td>PROPN</td><td>proper noun</td><td>NNP</td><td>noun, proper singular</td><td>NounType=prop Number=sign</td></tr>
# <tr><td>PROPN</td><td></td><td>NNPS</td><td>noun, proper plural</td><td>NounType=prop Number=plur</td></tr>
# <tr><td>PUNCT</td><td>punctuation</td><td>-LRB-</td><td>left round bracket</td><td>PunctType=brck PunctSide=ini</td></tr>
# <tr><td>PUNCT</td><td></td><td>-RRB-</td><td>right round bracket</td><td>PunctType=brck PunctSide=fin</td></tr>
# <tr><td>PUNCT</td><td></td><td>,</td><td>punctuation mark, comma</td><td>PunctType=comm</td></tr>
# <tr><td>PUNCT</td><td></td><td>:</td><td>punctuation mark, colon or ellipsis</td><td></td></tr>
# <tr><td>PUNCT</td><td></td><td>.</td><td>punctuation mark, sentence closer</td><td>PunctType=peri</td></tr>
# <tr><td>PUNCT</td><td></td><td>''</td><td>closing quotation mark</td><td>PunctType=quot PunctSide=fin</td></tr>
# <tr><td>PUNCT</td><td></td><td>""</td><td>closing quotation mark</td><td>PunctType=quot PunctSide=fin</td></tr>
# <tr><td>PUNCT</td><td></td><td>``</td><td>opening quotation mark</td><td>PunctType=quot PunctSide=ini</td></tr>
# <tr><td>PUNCT</td><td></td><td>HYPH</td><td>punctuation mark, hyphen</td><td>PunctType=dash</td></tr>
# <tr><td>PUNCT</td><td></td><td>LS</td><td>list item marker</td><td>NumType=ord</td></tr>
# <tr><td>PUNCT</td><td></td><td>NFP</td><td>superfluous punctuation</td><td></td></tr>
# <tr><td>SYM</td><td>symbol</td><td>#</td><td>symbol, number sign</td><td>SymType=numbersign</td></tr>
# <tr><td>SYM</td><td></td><td>\$</td><td>symbol, currency</td><td>SymType=currency</td></tr>
# <tr><td>SYM</td><td></td><td>SYM</td><td>symbol</td><td></td></tr>
# <tr><td>VERB</td><td>verb</td><td>BES</td><td>auxiliary "be"</td><td></td></tr>
# <tr><td>VERB</td><td></td><td>HVS</td><td>forms of "have"</td><td></td></tr>
# <tr><td>VERB</td><td></td><td>MD</td><td>verb, modal auxiliary</td><td>VerbType=mod</td></tr>
# <tr><td>VERB</td><td></td><td>VB</td><td>verb, base form</td><td>VerbForm=inf</td></tr>
# <tr><td>VERB</td><td></td><td>VBD</td><td>verb, past tense</td><td>VerbForm=fin Tense=past</td></tr>
# <tr><td>VERB</td><td></td><td>VBG</td><td>verb, gerund or present participle</td><td>VerbForm=part Tense=pres Aspect=prog</td></tr>
# <tr><td>VERB</td><td></td><td>VBN</td><td>verb, past participle</td><td>VerbForm=part Tense=past Aspect=perf</td></tr>
# <tr><td>VERB</td><td></td><td>VBP</td><td>verb, non-3rd person singular present</td><td>VerbForm=fin Tense=pres</td></tr>
# <tr><td>VERB</td><td></td><td>VBZ</td><td>verb, 3rd person singular present</td><td>VerbForm=fin Tense=pres Number=sing Person=3</td></tr>
# <tr><td>X</td><td>other</td><td>ADD</td><td>email</td><td></td></tr>
# <tr><td>X</td><td></td><td>FW</td><td>foreign word</td><td>Foreign=yes</td></tr>
# <tr><td>X</td><td></td><td>GW</td><td>additional word in multi-word expression</td><td></td></tr>
# <tr><td>X</td><td></td><td>XX</td><td>unknown</td><td></td></tr>
# <tr><td>SPACE</td><td>space</td><td>_SP</td><td>space</td><td></td></tr>
# <tr><td></td><td></td><td>NIL</td><td>missing tag</td><td></td></tr>
# </table>
# For a current list of tags for all languages visit https://spacy.io/api/annotation#pos-tagging
# ## Working with POS Tags
# In the English language, the same string of characters can have different meanings, even within the same sentence. For this reason, morphology is important. **spaCy** uses machine learning algorithms to best predict the use of a token in a sentence. Is *"I read books on NLP"* present or past tense? Is *wind* a verb or a noun?
# +
doc = nlp(u'I read books on NLP.')
r = doc[1]
print(f'{r.text:{10}} {r.pos_:{8}} {r.tag_:{6}} {spacy.explain(r.tag_)}')
# +
doc = nlp(u'I read a book on NLP.')
r = doc[1]
print(f'{r.text:{10}} {r.pos_:{8}} {r.tag_:{6}} {spacy.explain(r.tag_)}')
# -
# In the first example, with no other cues to work from, spaCy assumed that ***read*** was present tense.<br>In the second example the present tense form would be ***I am reading a book***, so spaCy assigned the past tense.
# ## Counting POS Tags
# The `Doc.count_by()` method accepts a specific token attribute as its argument, and returns a frequency count of the given attribute as a dictionary object. Keys in the dictionary are the integer values of the given attribute ID, and values are the frequency. Counts of zero are not included.
# +
doc = nlp(u"The quick brown fox jumped over the lazy dog's back.")
# Count the frequencies of different coarse-grained POS tags:
POS_counts = doc.count_by(spacy.attrs.POS)
POS_counts
# -
# This isn't very helpful until you decode the attribute ID:
doc.vocab[83].text
# ### Create a frequency list of POS tags from the entire document
# Since `POS_counts` returns a dictionary, we can obtain a list of keys with `POS_counts.items()`.<br>By sorting the list we have access to the tag and its count, in order.
for k,v in sorted(POS_counts.items()):
print(f'{k}. {doc.vocab[k].text:{5}}: {v}')
# +
# Count the different fine-grained tags:
TAG_counts = doc.count_by(spacy.attrs.TAG)
for k,v in sorted(TAG_counts.items()):
print(f'{k}. {doc.vocab[k].text:{4}}: {v}')
# -
# <div class="alert alert-success">**Why did the ID numbers get so big?** In spaCy, certain text values are hardcoded into `Doc.vocab` and take up the first several hundred ID numbers. Strings like 'NOUN' and 'VERB' are used frequently by internal operations. Others, like fine-grained tags, are assigned hash values as needed.</div>
# <div class="alert alert-success">**Why don't SPACE tags appear?** In spaCy, only strings of spaces (two or more) are assigned tokens. Single spaces are not.</div>
# +
# Count the different dependencies:
DEP_counts = doc.count_by(spacy.attrs.DEP)
for k,v in sorted(DEP_counts.items()):
print(f'{k}. {doc.vocab[k].text:{4}}: {v}')
# -
# Here we've shown `spacy.attrs.POS`, `spacy.attrs.TAG` and `spacy.attrs.DEP`.<br>Refer back to the **Vocabulary and Matching** lecture from the previous section for a table of **Other token attributes**.
# ___
# ## Fine-grained POS Tag Examples
# These are some grammatical examples (shown in **bold**) of specific fine-grained tags. We've removed punctuation and rarely used tags:
# <table>
# <tr><th>POS</th><th>TAG</th><th>DESCRIPTION</th><th>EXAMPLE</th></tr>
# <tr><td>ADJ</td><td>AFX</td><td>affix</td><td>The Flintstones were a **pre**-historic family.</td></tr>
# <tr><td>ADJ</td><td>JJ</td><td>adjective</td><td>This is a **good** sentence.</td></tr>
# <tr><td>ADJ</td><td>JJR</td><td>adjective, comparative</td><td>This is a **better** sentence.</td></tr>
# <tr><td>ADJ</td><td>JJS</td><td>adjective, superlative</td><td>This is the **best** sentence.</td></tr>
# <tr><td>ADJ</td><td>PDT</td><td>predeterminer</td><td>Waking up is **half** the battle.</td></tr>
# <tr><td>ADJ</td><td>PRP\$</td><td>pronoun, possessive</td><td>**His** arm hurts.</td></tr>
# <tr><td>ADJ</td><td>WDT</td><td>wh-determiner</td><td>It's blue, **which** is odd.</td></tr>
# <tr><td>ADJ</td><td>WP\$</td><td>wh-pronoun, possessive</td><td>We don't know **whose** it is.</td></tr>
# <tr><td>ADP</td><td>IN</td><td>conjunction, subordinating or preposition</td><td>It arrived **in** a box.</td></tr>
# <tr><td>ADV</td><td>EX</td><td>existential there</td><td>**There** is cake.</td></tr>
# <tr><td>ADV</td><td>RB</td><td>adverb</td><td>He ran **quickly**.</td></tr>
# <tr><td>ADV</td><td>RBR</td><td>adverb, comparative</td><td>He ran **quicker**.</td></tr>
# <tr><td>ADV</td><td>RBS</td><td>adverb, superlative</td><td>He ran **fastest**.</td></tr>
# <tr><td>ADV</td><td>WRB</td><td>wh-adverb</td><td>**When** was that?</td></tr>
# <tr><td>CONJ</td><td>CC</td><td>conjunction, coordinating</td><td>The balloon popped **and** everyone jumped.</td></tr>
# <tr><td>DET</td><td>DT</td><td>determiner</td><td>**This** is **a** sentence.</td></tr>
# <tr><td>INTJ</td><td>UH</td><td>interjection</td><td>**Um**, I don't know.</td></tr>
# <tr><td>NOUN</td><td>NN</td><td>noun, singular or mass</td><td>This is a **sentence**.</td></tr>
# <tr><td>NOUN</td><td>NNS</td><td>noun, plural</td><td>These are **words**.</td></tr>
# <tr><td>NOUN</td><td>WP</td><td>wh-pronoun, personal</td><td>**Who** was that?</td></tr>
# <tr><td>NUM</td><td>CD</td><td>cardinal number</td><td>I want **three** things.</td></tr>
# <tr><td>PART</td><td>POS</td><td>possessive ending</td><td>Fred**'s** name is short.</td></tr>
# <tr><td>PART</td><td>RP</td><td>adverb, particle</td><td>Put it **back**!</td></tr>
# <tr><td>PART</td><td>TO</td><td>infinitival to</td><td>I want **to** go.</td></tr>
# <tr><td>PRON</td><td>PRP</td><td>pronoun, personal</td><td>**I** want **you** to go.</td></tr>
# <tr><td>PROPN</td><td>NNP</td><td>noun, proper singular</td><td>**Kilroy** was here.</td></tr>
# <tr><td>PROPN</td><td>NNPS</td><td>noun, proper plural</td><td>The **Flintstones** were a pre-historic family.</td></tr>
# <tr><td>VERB</td><td>MD</td><td>verb, modal auxiliary</td><td>This **could** work.</td></tr>
# <tr><td>VERB</td><td>VB</td><td>verb, base form</td><td>I want to **go**.</td></tr>
# <tr><td>VERB</td><td>VBD</td><td>verb, past tense</td><td>This **was** a sentence.</td></tr>
# <tr><td>VERB</td><td>VBG</td><td>verb, gerund or present participle</td><td>I am **going**.</td></tr>
# <tr><td>VERB</td><td>VBN</td><td>verb, past participle</td><td>The treasure was **lost**.</td></tr>
# <tr><td>VERB</td><td>VBP</td><td>verb, non-3rd person singular present</td><td>I **want** to go.</td></tr>
# <tr><td>VERB</td><td>VBZ</td><td>verb, 3rd person singular present</td><td>He **wants** to go.</td></tr>
# </table>
# ### Up Next: Visualizing POS
| notebooks/6_POS-Basics.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; # 2 データを用いた抽象化の構築
;;
;; 概要の詳細はテキストを参照してください。
;; ここでは重要と思われる内容について説明を記載します。
;;
;; 第1章の⼿続きは全て単純な数値データを処理するものでした。
;; 一般的にコンピュータで解決したい問題の多くは、
;; 単純なデータでは不十分であり、
;; 複雑な現象をモデル化するには複雑なデータ構造が必要です。
;; この章では、このような問題に対応すべく、複雑なデータ構造を見ていきます。
;;
;; プログラミング言語には、
;; 既存のデータオブジェクトを組み合わせて **複合データ(compound data)**
;; を作る手段が用意されています(テキストでは糊と表現)。
;; なぜ、複合データが必要かというと、
;; 複合手続きが必要でる理由と同じで、
;; プログラムを設計する概念レベルを引き上げ、
;; 設計のモジュール性を⾼め、⾔語の表現⼒を強くしたいからです。
;;
;; - **複合データオブジェクト**
;; - **データ抽象化**
;; ここで例として、2つの整数、分子と分母で表現した有理数を考えます。
;; 1つ目の有理数を$x=\frac{n_x}{m_x}$、
;; 2つ目の有理数を$y=\frac{n_y}{m_y}$
;; とすると、
;; 2つの有理数の和と積は、
;; $x + y = \frac{m_y n_x + m_x n_y}{m_x m_y}$
;; $x \cdot y = \frac{n_x n_y}{m_x m_y}$
;; となります。
;; ここで上記の有理数の線形結合 $ax+by$ を考えた場合、
;; $n_x,m_x,n_y,m_y$でプログラムを記述しようとすると、
;; どの変数がどの有理数に対応しているか覚えておく必要があり、
;; また線形結合の計算式も複雑になり、煩雑になってしまいます。
;; プログラミング上でも、$ax+by$ として表現できたほうが簡潔であり、分かりやすいです。
;; 手続きの引数や変数も$n_x,m_x,n_y,m_y$ではなく、$x,y$として考えたいところです。
;; これがデータ抽象化の考え方であり、$x,y$のそれぞれが複合データになります。
;; (プログラムで具体的に表現された複合データ=複合データオブジェクト)
;; もっと言うとデータ抽象化とは、
;; プログラムの中のデータオブジェクトをどうやって表すかを扱う部分と、
;; データオブジェクトをどうやって使うかを扱う部分とを分離して設計することです。
;;
;; - **抽象化の壁** ・・・「図2.1」参照
;; 複合データの糊としては、いろいろな方法があります。
;; その例として、特別なデータ操作(cons)をまったく使わないでも、
;; ⼿続きだけを使って複合データを作ることができるということを見てきます。
;; [時田所感]
;; 後述される「⼿続きだけを使って複合データを作ることができる」については興味深いです。
;; 関数型言語の特徴、ファーストクラスの地位というのを、もっともよく体現しているように思えるからです。
;;
;; - **クロージャ**
;; クロージャ(閉包)とはデータオブジェクトを組み合わせるのに使う糊が、
;; 基本データオブジェクトだけでなく、
;; 複合データオブジェクトも組み合わせられるようになっていなければいけないということです。
;; (複合データオブジェクトも組わせる際、さらに別のやり方があるわけでなく、
;; 基本データオブジェクトを組み合わせる方法と同じ方法が使えるということ)
;; 「2.2 階層データと閉包性」の注釈6参照。
;; ・・・一般的に言われているクロージャと違うと説明されています。
;; https://ja.wikipedia.org/wiki/%E3%82%AF%E3%83%AD%E3%83%BC%E3%82%B8%E3%83%A3#%E3%82%AF%E3%83%AD%E3%83%BC%E3%82%B8%E3%83%A3%E3%82%92%E6%8C%81%E3%81%A4%E3%83%97%E3%83%AD%E3%82%B0%E3%83%A9%E3%83%9F%E3%83%B3%E3%82%B0%E8%A8%80%E8%AA%9E
;;
;; - **標準インターフェイス** ・・・「2.2.3 標準インターフェイスとしての列 」、「2.2.4 例:図形⾔語」参照
;;
;; - **記号式** ・・・「2.3 記号データ」参照
;;
;; - **ジェネリック演算** ・・・「2.5.1 ジェネリック算術演算」
;;
;; - **データ主導プログラミング**
;; データ主導プログラミングとは、データ表現を独⽴して設計し、それらを加法的(additively)に
;; (つまり、修正なしに)組み合わせられるようにするプログラミングのテクニック・手法です。
;; ## 2.1 データ抽象化入門
;;
;; - **データ抽象化**
;; ・・・説明済み
;;
;; データ抽象化によって、
;; 複合データオブジェクトがどう使われるかというところを、
;; それがより基本的なデータオブジェクトによってどのように
;; 構築されているかといった細かいところから
;; 分離することが可能になります。
;;
;; データ抽象化の基本的な考え⽅は、
;; 複合データオブジェクトを使うようなプログラムを構築する際に、
;; "抽象データ"を扱うようにするということです。
;;
;; "具体的な"データ表現は、
;; そのデータを使うプログラムとは独⽴に定義され、
;; これら⼆つのプログラムをつなぐインターフェイスとして、
;; ⼀組の⼿続きで、コンストラクタとセレクタがあります。
;;
;; - **コンストラクタ**
;; オブジェクト指向言語のコンストラクタに相当します。
;; 既存のデータオブジェクトを使って複合データオブジェクトを作る手続きのことです。
;;
;; - **セレクタ**
;; オブジェクト指向言語のgetterに相当します。
;; コンストラクタを使って作った複合データオブジェクトから、
;; 個別のデータオブジェクトを取り出す手続きのことです。
;; ### 2.1.1 例:有理数の数値演算
;;
;; ここでは、有理数を使って数値演算を行う場合を考えます。
;; 分⼦と分⺟から有理数を構築する⽅法(コンストラクタ)はすでに持っていると仮定します。
;; (これを **希望的思考** といいます)
;;
;; 有理数が与えられたときに、その分⼦と分⺟を抽出する⽅法(セレクタ)もあるとします。
;;
;; - (make-rat ⟨n⟩⟨d⟩)は、分⼦が整数⟨n⟩で分⺟が整数⟨d⟩である有理数を返す。[コンストラクタ]
;; - (numer ⟨x⟩)は、有理数⟨x⟩の分⼦を返す。[セレクタ]
;; - (denom ⟨x⟩)は、有理数⟨x⟩の分⺟を返す。[セレクタ]
; 有理数の加算
(define (add-rat x y)
(make-rat (+ (* (numer x) (denom y))
(* (numer y) (denom x)))
(* (denom x) (denom y))))
; 有理数の減算
(define (sub-rat x y)
(make-rat (- (* (numer x) (denom y))
(* (numer y) (denom x)))
(* (denom x) (denom y))))
; 有理数の乗算
(define (mul-rat x y)
(make-rat (* (numer x) (numer y))
(* (denom x) (denom y))))
; 有理数の除算
(define (div-rat x y)
(make-rat (* (numer x) (denom y))
(* (denom x) (numer y))))
; 2つの有理数が等しいか
(define (equal-rat? x y)
(= (* (numer x) (denom y))
(* (numer y) (denom x))))
;; #### ペア
;; Schemeは、ペアの構築として、cons手続きが用意されています。
;; また、ペアからデータを取り出すための手続きとして、
;;
;; - car手続き・・・ペアの1つ目の要素を取り出す。
;; - cdr手続き・・・ペアの2つ目の要素を取り出す。
;;
;; があります。
;; これらが概要にあった糊になります。
;; ペアによって構築されるデータオブジェクトは**リスト構造**(list-structured)と呼ばれます。
; ペアの構築
(define x (cons 1 2))
; カーと呼び、ペア(リスト)の先頭を返す。
(car x)
; クダーと呼び、ペアの後尾を返す。
; リストの場合は先頭を除いたリストを返す。
(cdr x)
(define x (cons 1 2))
(define y (cons 3 4))
; ペアを組み合わせて、更にペアを構築することができる。
(define z (cons x y))
z
(car z)
(cdr z)
(car (car z))
(cdr (car z))
(car (cdr z))
(cdr (cdr z))
; リストは2.2に説明あり
(define x (list 1 2 3))
(car x)
(cdr x)
;; #### 有理数を表現する
;;
;; ペアを使って、有理数を分子と分母の2つの整数で表します。
; コンストラクタ(2つの正数を渡して有理数を構成する)
(define (make-rat n d) (cons n d))
; セレクタ(分子を返す)
(define (numer x) (car x))
; セレクタ(分母を返す)
(define (denom x) (cdr x))
;; この本では有理数のコンストラクタとセレクタを
;; 以下のように実装しないことに注意してください。
;; このようにすると手続きの呼び出しは効率よくなりますが、
;; デバッグがしにくくなるためです。
;;
;; (define make-rat cons)
;; (define numer car)
;; (define denom cdr)
; 有理数の表示
(define (print-rat x)
(newline)
(display (numer x))
(display "/")
(display (denom x)))
(define one-half (make-rat 1 2))
(print-rat one-half)
(define one-third (make-rat 1 3))
(print-rat (add-rat one-half one-third))
(print-rat (mul-rat one-half one-third))
(print-rat (add-rat one-third one-third))
(print-rat (make-rat 2 -4))
;; 最後の例からわかるように、
;; この有理数計算の実装は有理数を既約のものに簡約しません。
;; これを直すには、make-ratを修正します。1.2.5節で扱ったような、
;; ⼆つの整数の最⼤公約数を返すgcd⼿続きがあれば、
;; ペアを構築する前にgcdを使って分⼦と分⺟を既約にできます。
;; この修正はコンストラクタmake-ratの変更だけで完了し、
;; 実際の演算を実装する(add-ratやmul-ratといった)⼿続きはどれも変更する必要はりません。
; 分子と分母の既約
(define (make-rat n d)
(let ((g (gcd n d)))
(cons (/ n g) (/ d g))
)
)
(define (gcd a b)
(if (= b 0) a
(gcd b (remainder a b))
)
)
(print-rat (add-rat one-third one-third))
| texts/2.1.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda-prediction_utils] *
# language: python
# name: conda-env-anaconda-prediction_utils-py
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import glob
import os
import matplotlib.pyplot as plt
import shutil
from prediction_utils.util import df_dict_concat, yaml_read, yaml_write
project_dir = "/share/pi/nigam/projects/spfohl/cohorts/admissions/mimic_omop/"
os.listdir(os.path.join(project_dir, 'experiments'))
experiment_name = 'baseline_tuning_fold_1_10'
baseline_files = glob.glob(
os.path.join(
project_dir,
'experiments',
experiment_name,
'**',
'result_df_training_eval.parquet'
),
recursive=True
)
baseline_df_dict = {
tuple(file_name.split('/'))[-4:-1]: pd.read_parquet(file_name)
for file_name in baseline_files
}
baseline_df = df_dict_concat(baseline_df_dict,
['task', 'config_filename', 'fold']
)
baseline_df.head()
# +
mean_performance = (
pd.DataFrame(
baseline_df
.query('metric == "loss" & phase == "val"')
.groupby(['config_filename', 'task'])
.agg(performance=('performance', 'mean'))
.reset_index()
)
)
best_model = (
mean_performance
.groupby('task')
.agg(performance=('performance','min'))
.merge(mean_performance)
)
# -
best_model_config_df = best_model[['config_filename', 'task']]
best_model_performance = baseline_df.merge(best_model_config_df)
best_model_performance.groupby(['task', 'config_filename', 'metric', 'phase', 'epoch']).agg('mean')
best_model_performance[['task', 'config_filename']].drop_duplicates()
best_model_config_df
best_model
base_config_path = os.path.join(project_dir, 'experiments', experiment_name, 'config')
# retrain_experiment_name = 'baseline_best'
selected_config_path = os.path.join(project_dir, 'experiments', experiment_name, 'config', 'selected_models')
# Write to a new directory
for i, row in best_model_config_df.iterrows():
the_config = yaml_read(os.path.join(base_config_path, row.task, row.config_filename))
print(row.task)
print(the_config)
the_config['label_col'] = row.task
os.makedirs(os.path.join(selected_config_path, row.task), exist_ok=True)
yaml_write(the_config, os.path.join(selected_config_path, row.task, row.config_filename))
| fairness_benchmark/notebooks/get_best_model_mimic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py2)
# language: python
# name: py2
# ---
""" Load some libs """
""" python 2 lib using networkx """
import matplotlib.pyplot as plt
import networkx as nx
import random
import math
import pandas as pd
import statsmodels.api as sm
import glob
import os
import numpy as np
from PIL import Image
from helpers import *
import pickle
import time
#random.seed(100)
#tic = time.time()
""" Let's see what WEPPS we have """
folders = sorted(glob.glob('WEPP_FILES/*'))
print folders
"""Let's get a single WEPP db"""
chosen_db = folders[14]
#get the date as int
quarter_dict = {'Q1':0.0, 'Q2':0.25,'Q3':0.5,'Q4':0.75}
DATE = int(chosen_db.split('/')[1].split('-')[0])+quarter_dict[chosen_db.split('/')[1].split('-')[1].upper()]
print DATE
""" let's load in our WEPP db"""
wepp_df = get_wepp(chosen_db)
print wepp_df
# +
""" fix all the wepp stuff, fix categories, interpolate dates, add all the columns """
def prep_wepp(wepp_df):
# merge with ISO, country budgets and load factors
print '~~~~~~ GENERATING DF ~~~~~~~'
print 'loading df...'
df_iso = pd.read_csv('country_ISO_update.csv')
fuel_class = 'fuel_classification_database.dta'
df_fuel_class = pd.io.stata.read_stata(fuel_class)
heat_rates_xls = 'Heat rates_v3.xls'
df_heatrates = pd.read_excel(heat_rates_xls, sheet_name='CSV_output')
df_load_factor = pd.io.stata.read_stata('load_factor_database.dta')
print 'loaded dfs: '
print 'merging dfs and filling missing years...'
#df_fuel_load = pd.merge(df_fuel_class, df_load_factor, on='fuel_class')
#print df_iso
#print df_fuel_class
#print df_heatrates
#print df_load_factor
#print list(wepp_df)
#print wepp_df['FUEL']
df_fuel_class.rename(columns = {'fuel': 'FUEL'}, inplace = True)
#fix fuel classes
wepp_df = wepp_df.merge(df_fuel_class, on='FUEL', how='left')
df_wepp_em_fact = pd.read_csv('wepp_em_fact.csv')
#merge emissions factors
wepp_df = wepp_df.merge(df_wepp_em_fact, left_on='FUEL', right_on='fuel', how='left')
#prepare lookup indexer
wepp_df['FORMAT_HR'] = wepp_df.apply(lambda row: format_hr(row), axis=1)
#standardise statuses
wepp_df.loc[wepp_df.STATUS=='DEF', 'STATUS'] = 'PLN'
wepp_df.loc[wepp_df.STATUS=='DEL', 'STATUS'] = 'CON'
wepp_df.loc[wepp_df.STATUS=='UNK', 'STATUS'] = 'PLN'
wepp_df.loc[wepp_df.STATUS=='DAC', 'STATUS'] = 'STN'
#print list(df_iso)
#add ISO
wepp_df = wepp_df.merge(df_iso[['Caps','ISO','Region']], left_on='COUNTRY', right_on='Caps', how='left')
#fill in missing years
all_training = wepp_df[['YEAR','fuel_class','STATUS','Region','FORMAT_HR']]
all_training['fuel_class'] = all_training['fuel_class'].astype('category')
all_training['STATUS'] = all_training['STATUS'].astype('category')
all_training['Region'] = all_training['Region'].astype('category')
all_training['FORMAT_HR'] = all_training['FORMAT_HR'].astype('category')
all_training = pd.get_dummies(all_training[['YEAR','fuel_class','STATUS','Region','FORMAT_HR']], columns = ['fuel_class','STATUS','Region','FORMAT_HR'])
year_train_X = all_training[all_training.YEAR.notnull()].drop('YEAR', axis=1)
year_train_Y = all_training.loc[all_training.YEAR.notnull(),'YEAR']
year_train_X = sm.add_constant(year_train_X)
test_data = all_training.loc[all_training.YEAR.isnull()].drop('YEAR', axis=1)
test_data = sm.add_constant(test_data)
est = sm.OLS(year_train_Y, year_train_X)
est = est.fit()
wepp_df['YEAR_EST_FLAG'] = 0
wepp_df.loc[wepp_df.YEAR.isnull(),'YEAR_EST_FLAG'] = 1
wepp_df.loc[wepp_df.YEAR.isnull(),'YEAR'] = est.predict(test_data)
#get heatrates
wepp_df = wepp_df.merge(df_heatrates, left_on='FORMAT_HR', right_on='unique_id', how='left')
wepp_df['HEATRATE'] = wepp_df.apply(lambda row: get_hr(row), axis=1)
drop_cols = [col for col in list(wepp_df) if isinstance(col,int)]
wepp_df.drop(drop_cols, axis=1, inplace=True)
#get CO2 int, CCCE
wepp_df = wepp_df.merge(df_load_factor, on='fuel_class', how='left')
wepp_df['YEARS_LEFT'] = np.where(wepp_df['STATUS']=='OPR', wepp_df['YEAR']+40-2017, 0)
wepp_df.YEARS_LEFT.clip(lower=0.0, inplace=True) #set min years left to 0
print 'dfs merged and interped: '
print 'calculating carbon and MWs...'
wepp_df['CO2_INT'] = wepp_df['em_fact'] /2.205 * wepp_df['HEATRATE'] / 1000
wepp_df['CCCE'] = 8760 * wepp_df['MW'] * wepp_df['YEARS_LEFT'] * wepp_df['load_factor'] * wepp_df['CO2_INT'] /1000 #tonnes
#wepp_df.sort_values('CCCE', inplace=True)
#print wepp_df
#print list(wepp_df)
#print wepp_df.CCCE
#print all_countries
#exit()
#sort WEPP
wepp_df.sort_values('CCCE', inplace=True, ascending=False)
wepp_df['green']=wepp_df.fuel_class.isin(['SUN','BIOGAS','WASTE','BIOOIL','WIND','BIOMASS','GEOTHERMAL'])
wepp_df['green_MW'] = wepp_df.MW*wepp_df.green
wepp_df['blue']=wepp_df.fuel_class.isin(['WATER','NUCLEAR'])
wepp_df['blue_MW'] = wepp_df.MW*wepp_df.blue
return wepp_df
# +
""" let's get the main matrix"""
df_in = prep_wepp(wepp_df)#, threshold=0.0, threshold_column='mw')
print list(df_in)
#dataframe is m_companies by n_countries with additional columns for total MW and CCCE
# -
print df_in[df_in.RETIRE>0.0].RETIRE.unique()
def back_cast(wepp_df,YEAR,threshold=2000, threshold_column='mw'):
#get countries
all_countries = wepp_df.ISO.unique()
#re-opr if RETIRE >= YEAR -> STATUS=OPR
wepp_df[wepp_df.RETIRE>=YEAR].STATUS='OPR'
#drop plants if year
wepp_df = wepp_df.drop(wepp_df[wepp_df.YEAR>YEAR].index)
#plot companies by CCCE
company_df = pd.DataFrame(wepp_df.CCCE.groupby(wepp_df['COMPANY']).sum())
company_df.sort_values('CCCE', inplace=True, ascending=False)
#print 'company_df'
#print company_df
for country in all_countries:
#for CCCE
#company_df[country] = wepp_df.loc[wepp_df.ISO==country,'CCCE'].groupby(wepp_df['COMPANY']).sum()
#for MW
company_df[country] = wepp_df.loc[wepp_df.ISO==country,'MW'].groupby(wepp_df['COMPANY']).sum()
company_df[str(country)+'_green'] = wepp_df.loc[wepp_df.ISO==country,'green_MW'].groupby(wepp_df['COMPANY']).sum()
company_df[str(country)+'_blue'] = wepp_df.loc[wepp_df.ISO==country,'blue_MW'].groupby(wepp_df['COMPANY']).sum()
#checksum calculation
#print list(company_df)
#company_df.to_csv('test_nan.csv')
company_df.drop(labels=[np.nan,'nan_green'], axis=1, inplace=True)
iso_col = [h for h in list(company_df) if len(h)<3]
company_df['checksum'] = company_df[iso_col].sum(axis=1)
company_df.sort_values('checksum', inplace=True, ascending=False)
#print company_df
company_df.fillna(0.0, inplace=True)
#see how many edges we've got
#company_df['edges'] = company_df.count(axis=1)
#print company_df.edges.mean() #mean about 1.4 - nice.
if threshold_column=='mw':
print 'MW'
company_subset_df = company_df[company_df.checksum>threshold]
elif threshold_column=='ccce':
company_subset_df = company_df[company_df.CCCE>=threshold]
all_ccce = company_df.CCCE.sum()
print 'all CCCE', all_ccce
print 'por_CCCE',company_subset_df.CCCE.sum()/float(all_ccce)
print 'calculated carbon and MWs: '
return company_subset_df
# +
DATE='2008.XX'
df = back_cast(df_in,2008.0,threshold=0.0, threshold_column='mw')
print df
# -
print DATE
print list(df)
df.to_csv('./matrix_csvs/'+str(DATE)+'_master_MW.csv', encoding='utf-8')
""" drop columns to just leave country columns """
df.dropna(axis = 1, how='all', inplace=True)
df.drop(['CCCE','checksum'], inplace=True, axis='columns')
all_companies = list(df.index)
""" load country centroids """
df_centroids = pd.read_csv('country_centroids.csv').set_index('country')
#print df_centroids.get_value('TH','latitude')
# +
""" Generate Adjacency Matrices for Green, Blue, and Total (Bipartite: countries adjacent to companies)"""
green_cols = [h for h in list(df) if 'green' in h]
blue_cols = [h for h in list(df) if 'blue' in h]
ISO_cols = [h for h in list(df) if len(h)<3]
ADJ_bipartite_g = df[green_cols].fillna(0.0) #df with companies as index, ISOs as columns, green MWs as data
ADJ_bipartite_b = df[blue_cols].fillna(0.0) #df with companies as index, ISOs as columns, blue MWs as data
ADJ_bipartite_all = df[ISO_cols].fillna(0.0)
green_dict = {}
blue_dict = {}
for h in list(ADJ_bipartite_g):
green_dict[h]=h[0:2]
for h in list(ADJ_bipartite_b):
blue_dict[h]=h[0:2]
ADJ_bipartite_g.rename(green_dict,axis='columns', inplace=True)
ADJ_bipartite_b.rename(blue_dict,axis='columns', inplace=True)
print (list(ADJ_bipartite_g))
print (list(ADJ_bipartite_b))
print (list(ADJ_bipartite_all))
ADJ_bipartite_g.to_csv('./matrix_csvs/'+str(DATE)+'_green_MW.csv', encoding='utf-8')
ADJ_bipartite_b.to_csv('./matrix_csvs/'+str(DATE)+'_blue_MW.csv', encoding='utf-8')
ADJ_bipartite_all.to_csv('./matrix_csvs/'+str(DATE)+'_all_MW.csv', encoding='utf-8')
print ADJ_bipartite_all
# -
por_g = ADJ_bipartite_g.sum()/ADJ_bipartite_all.sum()
print sorted(por_g)
plt.plot(sorted(np.log(por_g)))
plt.show()
# +
""" generate some country nodes"""
nodes = []
#Transpose countries to index, companies to columns
ADJ_bipartite_g = ADJ_bipartite_g.T
ADJ_bipartite_b = ADJ_bipartite_b.T
ADJ_bipartite_all = ADJ_bipartite_all.T
print 'Generating country nodes ...'
for j in ADJ_bipartite_all.index.values:
#g and b just get portion of green and blue for the node properties
g = int(ADJ_bipartite_g.loc[ADJ_bipartite_g.index==j].sum(axis=1)/ADJ_bipartite_all.loc[ADJ_bipartite_all.index==j].sum(axis=1)*255.0)
b = int(ADJ_bipartite_b.loc[ADJ_bipartite_b.index==j].sum(axis=1)/ADJ_bipartite_all.loc[ADJ_bipartite_all.index==j].sum(axis=1)*255.0)
#set country position to actual country centroid, set color to RGB with g and b, set size to f(total_MW)
nodes.append(
(j,{
'type': 'country',
'pos':np.array([df_centroids.get_value(j,'longitude'),df_centroids.get_value(j,'latitude')]),
'n_color':"#{0:02x}{1:02x}{2:02x}".format(clamp(0), clamp(g), clamp(b)),
'n_alpha':1.0,
'n_size':(math.exp(math.log(ADJ_bipartite_all.loc[ADJ_bipartite_all.index==j].sum(axis=1),2.0)/3))}
))
print (nodes)
# +
""" generate some country-country edges """
edges=[]
#Adjacency matrices in MW
ADJ_bipartite_g_reduced = ADJ_bipartite_g.loc[(ADJ_bipartite_g != 0.0).any(axis=1), (ADJ_bipartite_g != 0.0).any(axis=0)] #MW
ADJ_bipartite_b_reduced = ADJ_bipartite_b.loc[(ADJ_bipartite_b != 0.0).any(axis=1), (ADJ_bipartite_b != 0.0).any(axis=0)] #MW
#ADJ_country_all =ADJ_bipartite_all.dot(ADJ_bipartite_all.T>0.0) #MW
#print ADJ_country_g.sum(axis=1)
#indices = list(ADJ_ISO.index)
#adjacency matrices in edge algo
#Generage country-country adjacentcy matrices, normalise
#drop sum==0 columns
ADJ_country_g = (ADJ_bipartite_g_reduced/ADJ_bipartite_g_reduced.sum()).dot(ADJ_bipartite_g_reduced.T/(ADJ_bipartite_g_reduced.T.sum()))
ADJ_country_b = (ADJ_bipartite_b_reduced/ADJ_bipartite_b_reduced.sum()).dot(ADJ_bipartite_b_reduced.T/(ADJ_bipartite_b_reduced.T.sum()))
ADJ_country_all = (ADJ_bipartite_all/ADJ_bipartite_all.sum()).dot(ADJ_bipartite_all.T/(ADJ_bipartite_all.T.sum()))
ADJ_country_g.to_csv('./matrix_csvs/'+str(DATE)+'_green_projection.csv', encoding='utf-8')
ADJ_country_b.to_csv('./matrix_csvs/'+str(DATE)+'_blue_projection.csv', encoding='utf-8')
ADJ_country_all.to_csv('./matrix_csvs/'+str(DATE)+'_all_projection.csv', encoding='utf-8')
print ADJ_country_g
#for k1,k2s in ADJ_ISO.to_dict(orient='index').items():
# for k2,v in k2s.items():
# if v>0.0:
# print (k1,k2,v)
#print ADJ_blue
#print ADJ_green
#df_ISO_pairs = [ (k1,k2,v) for k2,v in k2s.items() for k1,k2s in ADJ_ISO.to_dict(orient='index').items()]
df_all_pairs = [ {k:v for k,v in m.items() if v>0.0} for m in ADJ_country_all.to_dict(orient='rows')]
df_all_pairs = dict(zip(ADJ_country_all.index.values,df_all_pairs))
df_green_pairs = [ {k:v for k,v in m.items() if v>0.0} for m in ADJ_country_g.to_dict(orient='rows')]
df_green_pairs = dict(zip(ADJ_country_g.index.values,df_green_pairs))
df_blue_pairs = [ {k:v for k,v in m.items() if v>0.0} for m in ADJ_country_b.to_dict(orient='rows')]
df_blue_pairs = dict(zip(ADJ_country_b.index.values,df_blue_pairs))
#print df_ISO_pairs
for k,v in df_all_pairs.items():
v.pop(k,None)
for k2,v2 in v.items():
try:
g = int(df_green_pairs[k][k2] / float(v2) * 255)
except:
g = 0
try:
b = int(df_blue_pairs[k][k2] / float(v2) * 255)
except:
b = 0
edges.append(
(k,k2,{
'weight':np.log10(v2),
'type':'country',
'e_color': "#{0:02x}{1:02x}{2:02x}".format(clamp(0), clamp(g), clamp(b)),
}))
#print edges
# +
""" prep data for analysis """
m = len(ADJ_country_all)
print m
#print ADJ_country_g
#ADJ_country_g.values[[np.arange(m)]*2]=0
#ADJ_country_b.values[[np.arange(m)]*2]=0
ADJ_country_all.values[[np.arange(m)]*2]=0
POR_GREEN = ADJ_bipartite_g.T.sum()/ADJ_bipartite_all.T.sum()
#print ADJ_I_algo
print POR_GREEN
INFLUENCE = ADJ_country_all.T.dot(POR_GREEN) #transpose for inbound influence = general connection * green por
COUNTRY_1HOT = pd.get_dummies(INFLUENCE.index).set_index(INFLUENCE.index)
#print COUNTRY_1HOT
print INFLUENCE
DATA_OUT = POR_GREEN.to_frame(name='POR_GREEN').join(INFLUENCE.to_frame(name='INFLUENCE'))
DATA_OUT = DATA_OUT.join(COUNTRY_1HOT)
DATA_OUT['DATE']=DATE
print DATA_OUT
DATA_OUT.to_csv('./matrix_csvs/'+str(DATE)+'_por_green_influence.csv', encoding='utf-8')
#pickle.dump(DATA_OUT,open('data_pickles/data_'+str(DATE)+'.pickle','wb'))
# +
""" show the country-projection graph """
G = nx.Graph()
G.add_nodes_from(nodes)
G.add_edges_from(edges)
pos = nx.get_node_attributes(G,'pos')
xs = [pos[k][0] for k,v in pos.iteritems()]
ys = [pos[k][1] for k,v in pos.iteritems()]
n_sizes = nx.get_node_attributes(G,'n_size')
n_colors = nx.get_node_attributes(G,'n_color')
n_alphas = nx.get_node_attributes(G,'n_alpha')
n_type = nx.get_node_attributes(G,'type')
fixed_nodes = [k for k in G.nodes() if n_type[k]=='country']
company_nodes = [k for k in G.nodes() if n_type[k]=='company']
n_cou_sizes = [n_sizes[k] for k in fixed_nodes]
n_cou_colors = [n_colors[k] for k in fixed_nodes]
n_cou_alphas = [n_alphas[k] for k in fixed_nodes]
n_com_sizes = [n_sizes[k] for k in company_nodes]
n_com_colors = [n_colors[k] for k in company_nodes]
n_com_alphas = [n_alphas[k] for k in company_nodes]
e_colors = nx.get_edge_attributes(G,'e_color')
e_colors = [G[u][v]['e_color'] for u,v in G.edges()]
e_colors_d = dict([((u,v),G[u][v]['e_color']) for u,v in G.edges()])
e_type = [((u,v),G[u][v]['type']) for u,v in G.edges()]
e_anchors = [l[0] for l in e_type if l[1]=='country']
e_anchors_colors = [e_colors_d[l] for l in e_anchors]
e_companies = [l[0] for l in e_type if l[1]=='company']
e_companies_colors = [e_colors_d[l] for l in e_companies]
fig, ax = plt.subplots(figsize=(16,9 ), dpi=100)
x_min = min(xs)-10
x_max = max(xs)+10
y_min = min(ys)-10
y_max = max(ys)+10
y_lim = [y_min,y_max]
x_lim = [x_min,x_max]
ax.set_ylim(y_lim)
ax.set_xlim(x_lim)
ax.set_position([0,0,1,1])
#print e_anchors_colors
#print e_anchors
nx.draw_networkx_edges(G, pos, edgelist =e_anchors, alpha=0.2, ax=ax, edge_color=e_anchors_colors)#, nodelist=[ncenter], alpha=0.4)
#nx.draw_networkx_edges(G, pos, edgelist =e_companies, alpha=0.2, ax=ax, edge_color=e_companies_colors)#, nodelist=[ncenter], alpha=0.4)
nodes_ax = nx.draw_networkx_nodes(G, pos,ax=ax, nodelist=fixed_nodes, node_size = n_cou_sizes, node_color = n_cou_colors,alpha= n_cou_alphas)#, nodelist=list(p.keys()),
#node_size=80,
#node_color=list(p.values()),
#cmap=plt.cm.Reds_r)
#nodes_ax = nx.draw_networkx_nodes(G, pos,ax=ax, nodelist=company_nodes, node_size = n_com_sizes, node_color = n_com_colors,alpha= n_com_alphas)#, nodelist=list(p.keys()),
nodes_ax.set_edgecolor('w')
print 'Generating graphic: '
plt.axis('off')
plt.show()
plt.savefig('output.png')
# -
def generate_network_graph(nodes, edges, spring_iters=0):
print '~~~~~~~ GENEARTING GRAPHIC ~~~~~~'
print 'Getting graph attributes...'
G = nx.Graph()
G.add_nodes_from(nodes)
G.add_edges_from(edges)
pos = nx.get_node_attributes(G,'pos')
pos_prev = pickle.load( open('pos_2017_Q3.pickle','rb'))
for k,v in pos.items():
if k in pos_prev.keys():
pos[k] = pos_prev[k]
#pos = nx.kamada_kawai_layout(G, k=10, weight='weight', pos=pos, center=[50,50])
#for n in G:
# print n
# print n[1]['type'], n_alpha(n), n_color(n), n_size(n)
#print G.nodes()
#print pos
n_sizes = nx.get_node_attributes(G,'n_size')
n_colors = nx.get_node_attributes(G,'n_color')
n_alphas = nx.get_node_attributes(G,'n_alpha')
n_type = nx.get_node_attributes(G,'type')
fixed_nodes = [k for k in G.nodes() if n_type[k]=='country']
company_nodes = [k for k in G.nodes() if n_type[k]=='company']
n_cou_sizes = [n_sizes[k] for k in fixed_nodes]
n_cou_colors = [n_colors[k] for k in fixed_nodes]
n_cou_alphas = [n_alphas[k] for k in fixed_nodes]
n_com_sizes = [n_sizes[k] for k in company_nodes]
n_com_colors = [n_colors[k] for k in company_nodes]
n_com_alphas = [n_alphas[k] for k in company_nodes]
e_colors = nx.get_edge_attributes(G,'e_color')
e_colors = [G[u][v]['e_color'] for u,v in G.edges()]
e_colors_d = dict([((u,v),G[u][v]['e_color']) for u,v in G.edges()])
e_type = [((u,v),G[u][v]['type']) for u,v in G.edges()]
e_anchors = [l[0] for l in e_type if l[1]=='anchor']
e_anchors_colors = [e_colors_d[l] for l in e_anchors]
e_companies = [l[0] for l in e_type if l[1]=='company']
e_companies_colors = [e_colors_d[l] for l in e_companies]
#print G.edges()
#print e_colors
#print len(e_colors)
#print len(G.edges())
#print e_type
#print fixed_nodes
#raw_input('-->')
#print n_sizes
#print n_colors
#print n_alphas
print 'Getting graph attributes: '
print 'Generating force layout...'
pos = nx.spring_layout(G, k=50, iterations=spring_iters, pos=pos, fixed=fixed_nodes, center=[0,0])
pickle.dump(pos,open('pos_2017_Q4.pickle','wb'))
print 'Generating force layout: '
print 'Generating graphic...'
fig, ax = plt.subplots(figsize=(16,9 ), dpi=100)
xs = [pos[k][0] for k,v in pos.iteritems()]
ys = [pos[k][1] for k,v in pos.iteritems()]
x_min = min(xs)-10
x_max = max(xs)+10
y_min = min(ys)-10
y_max = max(ys)+10
y_lim = [y_min,y_max]
x_lim = [x_min,x_max]
ax.set_ylim(y_lim)
ax.set_xlim(x_lim)
ax.set_position([0,0,1,1])
#nx.draw_networkx_edges(G, pos, edgelist =e_anchors, alpha=0.2, ax=ax, edge_color=e_anchors_colors)#, nodelist=[ncenter], alpha=0.4)
nx.draw_networkx_edges(G, pos, edgelist =e_companies, alpha=0.2, ax=ax, edge_color=e_companies_colors)#, nodelist=[ncenter], alpha=0.4)
#nodes_ax = nx.draw_networkx_nodes(G, pos,ax=ax, nodelist=fixed_nodes, node_size = n_cou_sizes, node_color = n_cou_colors,alpha= n_cou_alphas)#, nodelist=list(p.keys()),
#node_size=80,
#node_color=list(p.values()),
#cmap=plt.cm.Reds_r)
nodes_ax = nx.draw_networkx_nodes(G, pos,ax=ax, nodelist=company_nodes, node_size = n_com_sizes, node_color = n_com_colors,alpha= n_com_alphas)#, nodelist=list(p.keys()),
#node_size=80,
#node_color=list(p.values()),
#cmap=plt.cm.Reds_r)
nodes_ax.set_edgecolor('w')
"""
for k,v in n_type.iteritems():
if v == 'country':
try:
#print k,pos[k]
im = Image.open(os.path.join('flags_round','ISO', k+'.png'))
im.thumbnail((16,16), Image.ANTIALIAS)
im = np.array(im).astype(np.float)/255
#print im.shape
#print (pos[k][0] - x_lim[0])/103.*16*100-8, (pos[k][1]-y_lim[0])/float(y_lim[1]-y_lim[0])*9*100-8
fig.figimage(im, xo=(pos[k][0] - x_lim[0])/float(x_lim[1]-x_lim[0])*16*100-8,yo=(pos[k][1]-y_lim[0])/float(y_lim[1]-y_lim[0])*9*100-8, zorder=3) #offset in pixels.... pix per inch.
#fig.figimage(im, xo= -pos[k][0],yo=pos[k][1])
except:
print 'broken country', k
pass
"""
#plt.xlim(-0.05, 1.05)
#plt.ylim(-0.05, 1.05)
print 'Generating graphic: '
plt.axis('off')
#plt.show()
plt.savefig('output.png')
"""
G = nx.random_geometric_graph(200, 0.125)
# position is stored as node attribute data for random_geometric_graph
pos = nx.get_node_attributes(G, 'pos')
# find node near center (0.5,0.5)
dmin = 1
ncenter = 0
for n in pos:
x, y = pos[n]
d = (x - 0.5)**2 + (y - 0.5)**2
if d < dmin:
ncenter = n
dmin = d
pos = nx.spring_layout(G, iterations=50, pos=pos, center=[0.5,0.5])
# color by path length from node near center
p = dict(nx.single_source_shortest_path_length(G, ncenter))
plt.figure(figsize=(12, 12))
nx.draw_networkx_edges(G, pos, nodelist=[ncenter], alpha=0.4)
nx.draw_networkx_nodes(G, pos, nodelist=list(p.keys()),
node_size=80,
node_color=list(p.values()),
cmap=plt.cm.Reds_r)
#plt.xlim(-0.05, 1.05)
#plt.ylim(-0.05, 1.05)
plt.axis('off')
plt.show()
"""
company_subset_df = calculate_CCCE(wepp_df)
nodes, edges, ADJ_green, ADJ_blue, ADJ_ISO = gen_country_projection(company_subset_df)
generate_network_graph(nodes, edges)
print ADJ_green
| synthesize_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="8c80a83f7fb1ab87aabbdad97d11627fb0380e8f"
# # Convolutional Neural Networks (CNN)
#
#
# * [Loading the Data Set](#1)
# * [Normalization, Reshape and Label Encoding ](#2)
# * [Train Test Split](#3)
# * [Convolutional Neural Network](#4)
# * [What is Convolution Operation?](#5)
# * [Same Padding](#6)
# * [Max Pooling](#7)
# * [Flattening](#8)
# * [Full Connection](#9)
# * [Implementing with Keras](#10)
# * [Create Model](#11)
# * [Define Optimizer](#12)
# * [Compile Model](#13)
# * [Epochs and Batch Size](#14)
# * [Data Augmentation](#15)
# * [Fit the Model](#16)
# * [Evaluate the Model](#17)
# * [Deep Learning Tutorial for Beginners](https://www.kaggle.com/kanncaa1/deep-learning-tutorial-for-beginners)
# * [Artificial Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/pytorch-tutorial-for-deep-learning-lovers)
# * [Convolutional Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/pytorch-tutorial-for-deep-learning-lovers)
# * [Recurrent Neural Network with Pytorch](https://www.kaggle.com/kanncaa1/recurrent-neural-network-with-pytorch)
# * [Conclusion](#18)
#
# -
# https://www.kaggle.com/kanncaa1/convolutional-neural-network-cnn-tutorial
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
# import warnings
import warnings
# filter warnings
warnings.filterwarnings('ignore')
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
# Any results you write to the current directory are saved as output.
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# <a id="1"></a>
# ## Loading the Data Set
# * In this part we load and visualize the data.
# + _uuid="6884db4361a209f639cd6f3bf15231d271ecf563"
# read train
train = pd.read_csv("train.csv.zip")
print(train.shape)
train.head()
# + _uuid="d7ab3a0db0c87aef2bb65fae5f564ff4b1260a96"
# read test
test= pd.read_csv("test.csv.zip")
print(test.shape)
test.head()
# + _uuid="318d94f6e935a6d8210d0ac03707dfa6e3946475"
# put labels into y_train variable
Y_train = train["label"]
# Drop 'label' column
X_train = train.drop(labels = ["label"],axis = 1)
# + _uuid="8902b0312e6c047596cf27ebba554a68b82604b2"
# visualize number of digits classes
plt.figure(figsize=(15,7))
g = sns.countplot(Y_train, palette="icefire")
plt.title("Number of digit classes")
Y_train.value_counts()
# + _uuid="4c9d3a1467cf82a3dff04967a9846906f9758ed4"
# plot some samples
img = X_train.iloc[0].to_numpy()
img = img.reshape((28,28))
plt.imshow(img,cmap='gray')
plt.title(train.iloc[0,0])
plt.axis("off")
plt.show()
# + _uuid="d6b119ecbe774a45656d1d157f9b33f38adf8e96"
# plot some samples
img = X_train.iloc[3].to_numpy()
img = img.reshape((28,28))
plt.imshow(img,cmap='gray')
plt.title(train.iloc[3,0])
plt.axis("off")
plt.show()
# + [markdown] _uuid="74e341f4845b42101182eda3c990e3dc4dc64dff"
# <a id="2"></a>
# ## Normalization, Reshape and Label Encoding
# * Normalization
# * We perform a grayscale normalization to reduce the effect of illumination's differences.
# * If we perform normalization, CNN works faster.
# * Reshape
# * Train and test images (28 x 28)
# * We reshape all data to 28x28x1 3D matrices.
# * Keras needs an extra dimension in the end which correspond to channels. Our images are gray scaled so it use only one channel.
# * Label Encoding
# * Encode labels to one hot vectors
# * 2 => [0,0,1,0,0,0,0,0,0,0]
# * 4 => [0,0,0,0,1,0,0,0,0,0]
# + _uuid="f39e537f15757b5da3363c138a33e50bd78bbf49"
# Normalize the data
X_train = X_train / 255.0
test = test / 255.0
print("x_train shape: ",X_train.shape)
print("test shape: ",test.shape)
# + _uuid="d524f6de086928158a76c54d1685a92c1802e230"
# Reshape
X_train = X_train.values.reshape(-1,28,28,1)
test = test.values.reshape(-1,28,28,1)
print("x_train shape: ",X_train.shape)
print("test shape: ",test.shape)
# + _uuid="d15d35ca439dce194a96f4442c7a1c085ce24d28"
# Label Encoding
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
Y_train = to_categorical(Y_train, num_classes = 10)
# + [markdown] _uuid="111d4a1e66d823363f890f892bba39d692479663"
# <a id="3"></a>
# ## Train Test Split
# * We split the data into train and test sets.
# * test size is 10%.
# * train size is 90%.
# + _uuid="882d9dfb1bee94b2ec1e83911e3e334994d74f4b"
# Split the train and the validation set for the fitting
from sklearn.model_selection import train_test_split
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size = 0.1, random_state=2)
print("x_train shape",X_train.shape)
print("x_test shape",X_val.shape)
print("y_train shape",Y_train.shape)
print("y_test shape",Y_val.shape)
# + _uuid="1cdc57350e1c251ec583093ebba18c51a537f55f"
# Some examples
plt.imshow(X_train[2][:,:,0],cmap='gray')
plt.show()
# + [markdown] _uuid="caf4b9f7fcae153ac1d4c0246b1defc9d9d30776"
# <a id="4"></a>
# ## Convolutional Neural Network
# * CNN is used for image classification, object detection
# * <a href="https://ibb.co/kV1j9p"><img src="https://preview.ibb.co/nRkBpp/gec2.jpg" alt="gec2" border="0"></a>
# + [markdown] _uuid="682cb6f5144794038c0bef1dc0814dde0f7fe6c0"
# <a id="5"></a>
# ### What is Convolution Operation?
# * We have some image and feature detector(3*3)
# * Feature detector does not need to be 3 by 3 matrix. It can be 5 by 5 or 7 by 7.
# * Feature detector = kernel = filter
# * Feauture detector detects features like edges or convex shapes. Example, if out input is dog, feature detector can detect features like ear or tail of the dog.
# * feature map = conv(input image, feature detector). Element wise multiplication of matrices.
# * feature map = convolved feature
# * Stride = navigating in input image.
# * We reduce the size of image. This is important bc code runs faster. However, we lost information.
# * We create multiple feature maps bc we use multiple feature detectors(filters).
# * Lets look at gimp. Edge detect: [0,10,0],[10,-4,10],[0,10,0]
# * <a href="https://imgbb.com/"><img src="https://image.ibb.co/m4FQC9/gec.jpg" alt="gec" border="0"></a>
# * After having convolution layer we use ReLU to break up linearity. Increase nonlinearity. Because images are non linear.
# * <a href="https://ibb.co/mVZih9"><img src="https://preview.ibb.co/gbcQvU/RELU.jpg" alt="RELU" border="0"></a>
# + [markdown] _uuid="3af8fb49243719ad70016b64cd4dff6e62c413d9"
# <a id="6"></a>
# ### Same Padding
# * As we keep applying conv layers, the size of the volume will decrease faster than we would like. In the early layers of our network, we want to preserve as much information about the original input volume so that we can extract those low level features.
# * input size and output size are same.
# * <a href="https://ibb.co/jUPkUp"><img src="https://preview.ibb.co/noH5Up/padding.jpg" alt="padding" border="0"></a>
# + [markdown] _uuid="6694b8ea366f974d6bd055a2915a5dc06f7a96bb"
# <a id="7"></a>
# ### Max Pooling
# * It makes down-sampling or sub-sampling (Reduces the number of parameters)
# * It makes the detection of features invariant to scale or orientation changes.
# * It reduce the amount of parameters and computation in the network, and hence to also control overfitting.
# * <a href="https://ibb.co/ckTjN9"><img src="https://preview.ibb.co/gsNYFU/maxpool.jpg" alt="maxpool" border="0"></a>
# + [markdown] _uuid="c2fb971fc473c9104a27975242a36f8e9183742c"
# <a id="8"></a>
# ### Flattening
# * <a href="https://imgbb.com/"><img src="https://image.ibb.co/c7eVvU/flattenigng.jpg" alt="flattenigng" border="0"></a>
# + [markdown] _uuid="f437407cf18fdfacf8626baa5e76ed5d64be1a11"
# <a id="9"></a>
# ### Full Connection
# * Neurons in a fully connected layer have connections to all activations in the previous layer
# * Artificial Neural Network
# * <a href="https://ibb.co/hsS14p"><img src="https://preview.ibb.co/evzsAU/fullyc.jpg" alt="fullyc" border="0"></a>
# + [markdown] _uuid="7b290eb3f3b111d6841f8e4093277be1bde05078"
# <a id="10"></a>
# ## Implementing with Keras
# + [markdown] _uuid="fd9feb4c4dadbb8a5cc8a32b5ef582d41b5c0698"
# <a id="11"></a>
# ### Create Model
# * conv => max pool => dropout => conv => max pool => dropout => fully connected (2 layer)
# * Dropout: Dropout is a technique where randomly selected neurons are ignored during training
# * <a href="https://ibb.co/jGcvVU"><img src="https://preview.ibb.co/e7yPPp/dropout.jpg" alt="dropout" border="0"></a>
# + _uuid="c441d7b3852cee5d3636272d4da2f96b169f81ac"
#
from sklearn.metrics import confusion_matrix
import itertools
from keras.utils.np_utils import to_categorical # convert to one-hot-encoding
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
from keras.optimizers import RMSprop,Adam
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ReduceLROnPlateau
model = Sequential()
#
model.add(Conv2D(filters = 8, kernel_size = (5,5),padding = 'Same',
activation ='relu', input_shape = (28,28,1)))
model.add(MaxPool2D(pool_size=(2,2)))
model.add(Dropout(0.25))
#
model.add(Conv2D(filters = 16, kernel_size = (3,3),padding = 'Same',
activation ='relu'))
model.add(MaxPool2D(pool_size=(2,2), strides=(2,2)))
model.add(Dropout(0.25))
# fully connected
model.add(Flatten())
model.add(Dense(256, activation = "relu"))
model.add(Dropout(0.5))
model.add(Dense(10, activation = "softmax"))
# + [markdown] _uuid="9a432df491777ca0019db6f6b972581f2c5bd9f9"
# <a id="12"></a>
# ### Define Optimizer
# * Adam optimizer: Change the learning rate
#
# + _uuid="607a02b42636e3115a1ac7a8edcadf61cf5ea1b0"
# Define the optimizer
optimizer = Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
# + [markdown] _uuid="91bf542a979c5b191c534876186bf31e70ec7f06"
# <a id="13"></a>
# ### Compile Model
# * categorical crossentropy
# * We make binary cross entropy at previous parts and in machine learning tutorial
# * At this time we use categorical crossentropy. That means that we have multi class.
# * <a href="https://ibb.co/jm1bpp"><img src="https://preview.ibb.co/nN3ZaU/cce.jpg" alt="cce" border="0"></a>
#
# + _uuid="0d1eefc68470b4cdcec04c2570651da3d97676d0"
# Compile the model
model.compile(optimizer = optimizer , loss = "categorical_crossentropy", metrics=["accuracy"])
# + [markdown] _uuid="d5d8fea4cbb9b53cf1bb5089357a33dc179ee981"
# <a id="14"></a>
# ### Epochs and Batch Size
# * Say you have a dataset of 10 examples (or samples). You have a **batch size** of 2, and you've specified you want the algorithm to run for 3 **epochs**. Therefore, in each epoch, you have 5 **batches** (10/2 = 5). Each batch gets passed through the algorithm, therefore you have 5 iterations **per epoch**.
# * reference: https://stackoverflow.com/questions/4752626/epoch-vs-iteration-when-training-neural-networks
# + _uuid="a237feb5e53ecbc8799101cb6e699877faafde77"
epochs = 10 # for better result increase the epochs
batch_size = 250
# + [markdown] _uuid="e87c60e45759e0c52c2b72ee562fdb964c8008d5"
# <a id="15"></a>
# ### Data Augmentation
# * To avoid overfitting problem, we need to expand artificially our handwritten digit dataset
# * Alter the training data with small transformations to reproduce the variations of digit.
# * For example, the number is not centered The scale is not the same (some who write with big/small numbers) The image is rotated.
# * <a href="https://ibb.co/k24CUp"><img src="https://preview.ibb.co/nMxXUp/augment.jpg" alt="augment" border="0"></a>
#
#
# + _uuid="c339fa7d06e9b73a519ca661bed19de482707d1f"
# data augmentation
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # dimesion reduction
rotation_range=5, # randomly rotate images in the range 5 degrees
zoom_range = 0.1, # Randomly zoom image 10%
width_shift_range=0.1, # randomly shift images horizontally 10%
height_shift_range=0.1, # randomly shift images vertically 10%
horizontal_flip=False, # randomly flip images
vertical_flip=False) # randomly flip images
datagen.fit(X_train)
# + [markdown] _uuid="e42e43f1b3b9e6da0d99dafff8f4a4b514a07f4c"
# <a id="16"></a>
# ### Fit the model
# + _uuid="4b2957bb8976a25cdbbbdc3110d68c5035a9773c"
# Fit the model
history = model.fit_generator(datagen.flow(X_train,Y_train, batch_size=batch_size),
epochs = epochs, validation_data = (X_val,Y_val), steps_per_epoch=X_train.shape[0] // batch_size)
# + [markdown] _uuid="debd5424728e11aa30a8513ac7b4f7377193a2da"
# <a id="17"></a>
# ### Evaluate the model
# * Test Loss visualization
# * Confusion matrix
#
# + _uuid="180a06f7ae01e69117c6c8258411cfe1b9b7b991"
# Plot the loss and accuracy curves for training and validation
plt.plot(history.history['val_loss'], color='b', label="validation loss")
plt.title("Test Loss")
plt.xlabel("Number of Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
# + _uuid="6586e37bd470db822086e191a90388e7175d504f"
# confusion matrix
import seaborn as sns
# Predict the values from the validation dataset
Y_pred = model.predict(X_val)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(Y_val,axis = 1)
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
f,ax = plt.subplots(figsize=(8, 8))
sns.heatmap(confusion_mtx, annot=True, linewidths=0.01,cmap="Greens",linecolor="gray", fmt= '.1f',ax=ax)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plt.title("Confusion Matrix")
plt.show()
# + [markdown] _uuid="9409bc6a7e905d5887c8eec8d056c9f015bf4878"
# <a id="18"></a>
# ## Conclusion
# * http://scs.ryerson.ca/~aharley/vis/conv/flat.html
# * HW
# * If you have any question I will be very happy to hear it.
| 4-Machine_Learning/3-Deep Learning/2-Redes Neuronales Convolucionales/CNN MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="EfHKyp9TAH1i"
# # Classification Metrics
#
# In the last notebook we fit binary classifier to predict whether patients were diabetic or not. We used accuracy as a measure of how well the model performed, but accuracy isn't everything. In this notebook, we will look at alternatives to accuracy that can be much more useful in machine learning.
#
# ## Alternative metrics for binary classifiers
#
# Accuracy seems like a sensible metric to evaluate (and to a certain extent it is), but you need to be careful about drawing too many conclusions from the accuracy of a classifier. Remember that it's simply a measure of how many cases were predicted correctly. Suppose only 3% of the population is diabetic. You could create a classifier that always just predicts 0, and it would be 97% accurate - but not terribly helpful in identifying patients with diabetes!
#
# Fortunately, there are some other metrics that reveal a little more about how our model is performing. Scikit-Learn includes the ability to create a *classification report* that provides more insight than raw accuracy alone.
#
# To get started, run the next cell to load our data and train our model like last time.
# + colab={"base_uri": "https://localhost:8080/"} id="QYsyjz4FANgb" executionInfo={"status": "ok", "timestamp": 1646197289155, "user_tz": -345, "elapsed": 1629, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="18e79c68-71a0-47cf-d382-3ca76835a15c"
import pandas as pd
from matplotlib import pyplot as plt
# %matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
# load the training dataset
# !wget https://raw.githubusercontent.com/MicrosoftDocs/mslearn-introduction-to-machine-learning/main/Data/ml-basics/diabetes.csv
diabetes = pd.read_csv('diabetes.csv')
# Separate features and labels
features = ['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']
label = 'Diabetic'
X, y = diabetes[features].values, diabetes[label].values
# Split data 70%-30% into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
print ('Training cases: %d\nTest cases: %d' % (X_train.shape[0], X_test.shape[0]))
# Train the model
from sklearn.linear_model import LogisticRegression
# Set regularization rate
reg = 0.01
# train a logistic regression model on the training set
model = LogisticRegression(C=1/reg, solver="liblinear").fit(X_train, y_train)
predictions = model.predict(X_test)
print('Predicted labels: ', predictions)
print('Actual labels: ' ,y_test)
print('Accuracy: ', accuracy_score(y_test, predictions))
# + [markdown] id="62jZ4Od5ASnI"
# One of the simplest places to start is a classification report. Run the next cell to see a range of alternatives ways to assess our model
# + colab={"base_uri": "https://localhost:8080/"} id="ygMtoOHLAP90" executionInfo={"status": "ok", "timestamp": 1646197303881, "user_tz": -345, "elapsed": 478, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="efd965b1-a741-4500-fdb5-789ecf119057"
from sklearn. metrics import classification_report
print(classification_report(y_test, predictions))
# + [markdown] id="rpv5aSNEAYmJ"
# The classification report includes the following metrics for each class (0 and 1)
#
# > note that the header row may not line up with the values!
#
# * *Precision*: Of the predictions the model made for this class, what proportion were correct?
# * *Recall*: Out of all of the instances of this class in the test dataset, how many did the model identify?
# * *F1-Score*: An average metric that takes both precision and recall into account.
# * *Support*: How many instances of this class are there in the test dataset?
#
# The classification report also includes averages for these metrics, including a weighted average that allows for the imbalance in the number of cases of each class.
#
# Because this is a *binary* classification problem, the ***1*** class is considered *positive* and its precision and recall are particularly interesting - these in effect answer the questions:
#
# - Of all the patients the model predicted are diabetic, how many are actually diabetic?
# - Of all the patients that are actually diabetic, how many did the model identify?
#
# You can retrieve these values on their own by using the **precision_score** and **recall_score** metrics in scikit-learn (which by default assume a binary classification model).
# + colab={"base_uri": "https://localhost:8080/"} id="YXzziIdFAWQi" executionInfo={"status": "ok", "timestamp": 1646197323855, "user_tz": -345, "elapsed": 449, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="1eade71f-ab62-47c7-9e32-dd1494bb2586"
from sklearn.metrics import precision_score, recall_score
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
# + [markdown] id="BxJoZiyxAdi4"
# The precision and recall metrics are derived from four possible prediction outcomes:
# * *True Positives*: The predicted label and the actual label are both 1.
# * *False Positives*: The predicted label is 1, but the actual label is 0.
# * *False Negatives*: The predicted label is 0, but the actual label is 1.
# * *True Negatives*: The predicted label and the actual label are both 0.
#
# These metrics are generally tabulated for the test set and shown together as a *confusion matrix*, which takes the following form:
#
# <table style="border: 1px solid black;">
# <tr style="border: 1px solid black;">
# <td style="border: 1px solid black;color: black;" bgcolor="lightgray">TN</td><td style="border: 1px solid black;color: black;" bgcolor="white">FP</td>
# </tr>
# <tr style="border: 1px solid black;">
# <td style="border: 1px solid black;color: black;" bgcolor="white">FN</td><td style="border: 1px solid black;color: black;" bgcolor="lightgray">TP</td>
# </tr>
# </table>
#
# Note that the correct (*true*) predictions form a diagonal line from top left to bottom right - these figures should be significantly higher than the *false* predictions if the model is any good.
#
# In Python, you can use the **sklearn.metrics.confusion_matrix** function to find these values for a trained classifier:
# + colab={"base_uri": "https://localhost:8080/"} id="HNqdzZlYAbHx" executionInfo={"status": "ok", "timestamp": 1646197345363, "user_tz": -345, "elapsed": 494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="6c446dce-d04a-427d-a8ff-80ab9319bf09"
from sklearn.metrics import confusion_matrix
# Print the confusion matrix
cm = confusion_matrix(y_test, predictions)
print (cm)
# + [markdown] id="5K7J5Q8JAiTu"
# Until now, we've considered the predictions from the model as being either 1 or 0 class labels. Actually, things are a little more complex than that. Statistical machine learning algorithms, like logistic regression, are based on *probability*; so what actually gets predicted by a binary classifier is the probability that the label is true (**P(y)**) and the probability that the label is false (1 - **P(y)**). A threshold value of 0.5 is used to decide whether the predicted label is a 1 (*P(y) > 0.5*) or a 0 (*P(y) <= 0.5*). You can use the **predict_proba** method to see the probability pairs for each case:
# + colab={"base_uri": "https://localhost:8080/"} id="kVvxdBwUAgXZ" executionInfo={"status": "ok", "timestamp": 1646197362255, "user_tz": -345, "elapsed": 599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="226f351e-6dac-49bd-9a8e-5641e258b66e"
y_scores = model.predict_proba(X_test)
print(y_scores)
# + [markdown] id="IWygJnA9Ameb"
# The decision to score a prediction as a 1 or a 0 depends on the threshold to which the predicted probabilities are compared. If we were to change the threshold, it would affect the predictions; and therefore change the metrics in the confusion matrix. A common way to evaluate a classifier is to examine the *true positive rate* (which is another name for recall) and the *false positive rate* for a range of possible thresholds. These rates are then plotted against all possible thresholds to form a chart known as a *received operator characteristic (ROC) chart*, like this:
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="1BynGthiAkkE" executionInfo={"status": "ok", "timestamp": 1646197380793, "user_tz": -345, "elapsed": 640, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="e543ed60-3aba-4c25-920e-c8539896c847"
from sklearn.metrics import roc_curve
from sklearn.metrics import confusion_matrix
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# + [markdown] id="GzgVhmaRArDr"
# The ROC chart shows the curve of the true and false positive rates for different threshold values between 0 and 1. A perfect classifier would have a curve that goes straight up the left side and straight across the top. The diagonal line across the chart represents the probability of predicting correctly with a 50/50 random prediction; so you obviously want the curve to be higher than that (or your model is no better than simply guessing!).
#
# The area under the curve (AUC) is a value between 0 and 1 that quantifies the overall performance of the model. The closer to 1 this value is, the better the model. Once again, scikit-Learn includes a function to calculate this metric.
# + colab={"base_uri": "https://localhost:8080/"} id="cYzzvT-pAo8D" executionInfo={"status": "ok", "timestamp": 1646197398908, "user_tz": -345, "elapsed": 617, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="70e33db7-4939-4f09-8970-22359ed5ad05"
from sklearn.metrics import roc_auc_score
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
# + [markdown] id="54Eaw0EWAvh0"
# ### Perform preprocessing in a pipeline
#
# In this case, the ROC curve and its AUC indicate that the model performs better than a random guess which is not bad considering we performed very little preprocessing of the data.
#
# In practice, it's common to perform some preprocessing of the data to make it easier for the algorithm to fit a model to it. There's a huge range of preprocessing transformations you can perform to get your data ready for modeling, but we'll limit ourselves to a few common techniques:
#
# - Scaling numeric features so they're on the same scale. This prevents features with large values from producing coefficients that disproportionately affect the predictions.
# - Encoding categorical variables. For example, by using a *one hot encoding* technique you can create individual binary (true/false) features for each possible category value.
#
# To apply these preprocessing transformations, we'll make use of a Scikit-Learn feature named *pipelines*. These enable us to define a set of preprocessing steps that end with an algorithm. You can then fit the entire pipeline to the data, so that the model encapsulates all of the preprocessing steps as well as the regression algorithm. This is useful, because when we want to use the model to predict values from new data, we need to apply the same transformations (based on the same statistical distributions and category encodings used with the training data).
#
# >**Note**: The term *pipeline* is used extensively in machine learning, often to mean very different things! In this context, we're using it to refer to pipeline objects in Scikit-Learn, but you may see it used elsewhere to mean something else.
#
# + colab={"base_uri": "https://localhost:8080/"} id="4ZR-Dmr_AtXC" executionInfo={"status": "ok", "timestamp": 1646197418723, "user_tz": -345, "elapsed": 475, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="8c6a7f72-ffa8-493e-c97b-da264b17cc45"
# Train the model
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.linear_model import LogisticRegression
import numpy as np
# Define preprocessing for numeric columns (normalize them so they're on the same scale)
numeric_features = [0,1,2,3,4,5,6]
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
# Define preprocessing for categorical features (encode the Age column)
categorical_features = [7]
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
# Combine preprocessing steps
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, numeric_features),
('cat', categorical_transformer, categorical_features)])
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', LogisticRegression(C=1/reg, solver="liblinear"))])
# fit the pipeline to train a logistic regression model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# + [markdown] id="bC5Mm-0iA1KE"
# The pipeline encapsulates the preprocessing steps as well as model training.
#
# Let's use the model trained by this pipeline to predict labels for our test set, and compare the performance metrics with the basic model we created previously.
# + colab={"base_uri": "https://localhost:8080/", "height": 543} id="K7NJObu8AyNq" executionInfo={"status": "ok", "timestamp": 1646197440229, "user_tz": -345, "elapsed": 548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="f0b56ecb-9314-4d96-8894-df259b0ac2c1"
# Get predictions from test data
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
# Get evaluation metrics
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('AUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# + [markdown] id="rnIpah4SA6Ct"
# The results look a little better, so clearly preprocessing the data has made a difference.
#
# ### Try a different algorithm
#
# Now let's try a different algorithm. Previously we used a logistic regression algorithm, which is a *linear* algorithm. There are many kinds of classification algorithm we could try, including:
#
# - **Support Vector Machine algorithms**: Algorithms that define a *hyperplane* that separates classes.
# - **Tree-based algorithms**: Algorithms that build a decision tree to reach a prediction
# - **Ensemble algorithms**: Algorithms that combine the outputs of multiple base algorithms to improve generalizability.
#
# This time, We'll use the same preprocessing steps as before, but we'll train the model using an *ensemble* algorithm named *Random Forest* that combines the outputs of multiple random decision trees (for more details, see the [Scikit-Learn documentation](https://scikit-learn.org/stable/modules/ensemble.html#forests-of-randomized-trees)).
# + colab={"base_uri": "https://localhost:8080/"} id="yv8HP20vA3hp" executionInfo={"status": "ok", "timestamp": 1646197468427, "user_tz": -345, "elapsed": 5632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="d3e43086-aa88-469f-bd81-a9cf95e97add"
from sklearn.ensemble import RandomForestClassifier
# Create preprocessing and training pipeline
pipeline = Pipeline(steps=[('preprocessor', preprocessor),
('logregressor', RandomForestClassifier(n_estimators=100))])
# fit the pipeline to train a random forest model on the training set
model = pipeline.fit(X_train, (y_train))
print (model)
# + [markdown] id="_FYVhbukA_fX"
# Let's look at the performance metrics for the new model.
# + colab={"base_uri": "https://localhost:8080/", "height": 560} id="3oAx2Kg2A9Q4" executionInfo={"status": "ok", "timestamp": 1646197484449, "user_tz": -345, "elapsed": 2259, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="bb0d1911-845d-46f0-fc71-1cbc00d13325"
predictions = model.predict(X_test)
y_scores = model.predict_proba(X_test)
cm = confusion_matrix(y_test, predictions)
print ('Confusion Matrix:\n',cm, '\n')
print('Accuracy:', accuracy_score(y_test, predictions))
print("Overall Precision:",precision_score(y_test, predictions))
print("Overall Recall:",recall_score(y_test, predictions))
auc = roc_auc_score(y_test,y_scores[:,1])
print('\nAUC: ' + str(auc))
# calculate ROC curve
fpr, tpr, thresholds = roc_curve(y_test, y_scores[:,1])
# plot ROC curve
fig = plt.figure(figsize=(6, 6))
# Plot the diagonal 50% line
plt.plot([0, 1], [0, 1], 'k--')
# Plot the FPR and TPR achieved by our model
plt.plot(fpr, tpr)
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curve')
plt.show()
# + [markdown] id="b2_DLwDVBEcf"
# That looks better!
#
# ### Use the Model for Inferencing
# Now that we have a reasonably useful trained model, we can save it for use later to predict labels for new data:
# + colab={"base_uri": "https://localhost:8080/"} id="O5GavP8XBB0c" executionInfo={"status": "ok", "timestamp": 1646197504437, "user_tz": -345, "elapsed": 411, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="dc8396ff-67ca-4dea-f84b-888c46ae19cb"
import joblib
# Save the model as a pickle file
filename = './diabetes_model.pkl'
joblib.dump(model, filename)
# + [markdown] id="Ji5u4MtEBJjP"
# When we have some new observations for which the label is unknown, we can load the model and use it to predict values for the unknown label:
# + colab={"base_uri": "https://localhost:8080/"} id="fY-2SAw1BHOU" executionInfo={"status": "ok", "timestamp": 1646197523426, "user_tz": -345, "elapsed": 429, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjLLRQteQA-BSVM11RQI0Jpgt7v0nw6hWzW4zvL=s64", "userId": "07019866810236427547"}} outputId="c631991c-bf70-463d-b377-5b0f4a5816a5"
# Load the model from the file
model = joblib.load(filename)
# predict on a new sample
# The model accepts an array of feature arrays (so you can predict the classes of multiple patients in a single call)
# We'll create an array with a single array of features, representing one patient
X_new = np.array([[2,180,74,24,21,23.9091702,1.488172308,22]])
print ('New sample: {}'.format(list(X_new[0])))
# Get a prediction
pred = model.predict(X_new)
# The model returns an array of predictions - one for each set of features submitted
# In our case, we only submitted one patient, so our prediction is the first one in the resulting array.
print('Predicted class is {}'.format(pred[0]))
# + [markdown] id="9HHzWOInBN-a"
# ## Summary
#
# In this notebook, we looked at a range of metrics for binary classification and tried a few algorithms beyond logistic regression. We will move onto more complex classification problems in the following notebook.
| 10. Perform classification with alternative metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part 1 of the Assignment - Creating the dataframe
# ### Importing libraries and extracting table
# +
import pandas as pd
# Webpage url
url = 'https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M'
# Extract tables
dfs = pd.read_html(url)
# print number of tables
print(len(dfs))
# Get first table which is the table of interest
df = dfs[0]
# -
# ### Extract Required columns into a df
# Extract required columns
df2 = df[['Postal Code','Borough','Neighbourhood']]
# ### Ignore cells with a borough that is Not assigned
# get rid of rows with Borough value 'Not assigned'
df2 = df2[df2.Borough != 'Not assigned'].reset_index(drop=True)
# ### If a cell has a borough but a Not assigned neighborhood, then the neighborhood will be the same as the borough
mask = df2['Neighbourhood'] == "Not assigned"
df2.loc[mask,'Neighbourhood'] = df2.loc[mask, 'Borough']
# ### print number of rows of the df
# + pycharm={"name": "#%%\n"}
print(df2.shape[0])
# -
# ### Display dataframe
df2.head(12)
# ## Part 2 of the assignement - obtaining latitudes and longitudes
# ### read csv file with longitude an latitude details
df_lng_lat = pd.read_csv('Geospatial_Coordinates.csv')
df_lng_lat.head()
# ### Merge two dataframes with the common column latitude and longitude
df_merged = df2.merge(df_lng_lat, on="Postal Code", how = 'left')
df_merged.head()
print(df_merged.shape[0])
# ## Part 3 of the assignment - Explore and cluster the neighborhoods in Toronto.
# ### Extracting boroughs that contain the word Toronto
df_merged = df_merged[df_merged['Borough'].str.contains("Toronto")]
df_merged.head()
# ### Create a map of Toronto with neighborhoods superimposed on top.
# +
import numpy as np # library to handle data in a vectorized manner
import pandas as pd # library for data analsysis
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import json # library to handle JSON files
# # !conda install -c conda-forge geopy --yes # uncomment this line if you haven't completed the Foursquare API lab
# from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
import requests # library to handle requests
from pandas.io.json import json_normalize # tranform JSON file into a pandas dataframe
# Matplotlib and associated plotting modules
import matplotlib.cm as cm
import matplotlib.colors as colors
# import k-means from clustering stage
from sklearn.cluster import KMeans
# #!conda install -c conda-forge folium=0.5.0 --yes # uncomment this line if you haven't completed the Foursquare API lab
import folium # map rendering library
print('Libraries imported.')
# +
latitude = 43.651070
longitude = -79.347015
# create map of New York using latitude and longitude values
map_Toronto = folium.Map(location=[latitude, longitude], zoom_start=11)
# add markers to map
for lat, lng, borough, neighborhood in zip(df_merged['Latitude'], df_merged['Longitude'], df_merged['Borough'], df_merged['Neighbourhood']):
label = '{}, {}'.format(neighborhood, borough)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_Toronto)
map_Toronto
# -
# ### Define Foursquare Credentials and Version
# +
CLIENT_ID = 'GURYN0HXLCV2RLRBQZSKURSEVN5ZVZTB14HYM5DKEON3KGSW' # your Foursquare ID
CLIENT_SECRET = '<KEY>' # your Foursquare Secret
VERSION = '20180605' # Foursquare API version
LIMIT = 100 # A default Foursquare API limit value
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
# -
# ### Explore Neighborhoods in Toronto
def getNearbyVenues(names, latitudes, longitudes, radius=500):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}&query=coffee'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
Toronto_venues = getNearbyVenues(names=df_merged['Neighbourhood'],
latitudes=df_merged['Latitude'],
longitudes=df_merged['Longitude']
)
# ### Cluster the neighborhoods
# +
# set number of clusters
kclusters = 5
# toronto_grouped_clustering = df_merged.drop('Neighbourhood', 1)
toronto_grouped_clustering = df_merged.drop(['Neighbourhood', 'Borough', 'Postal Code'], axis=1)
# run k-means clustering
kmeans = KMeans(n_clusters=kclusters, random_state=0).fit(toronto_grouped_clustering)
# check cluster labels generated for each row in the dataframe
kmeans.labels_[0:10]
print(len(kmeans.labels_))
print(toronto_grouped_clustering.shape[0])
# -
# add clustering labels
df_merged.insert(0, 'Cluster Labels', kmeans.labels_)
df_merged.head()
# ### Display clusters on map
# +
# create map
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
# set color scheme for the clusters
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
# add markers to the map
markers_colors = []
for lat, lon, poi, cluster in zip(df_merged['Latitude'], df_merged['Longitude'], df_merged['Neighbourhood'], df_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
# -
| Segmenting and Clustering Neighborhoods in Toronto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### df_loc (Canada)
#
# We will use census Distribution Areas as proxies for neighborhoods for cities in Canada. In previous work where the Forward Sortation Areas (first three characters of the postal code) were used as neighborhood proxies, the sizes of many areas were quite large (several kilometers across) and therefore are likely internally non-homogeneous from a features perspective at the walking-distance (500 m) length scale. To convert to neighborhood names we can look up the associated census tract as seen on [this](https://en.wikipedia.org/wiki/Demographics_of_Toronto_neighbourhoods) Wikipedia page.
#
# File lda_000b16g_e.gml was downloaded from the [Statistics Canada: Boundary Files](https://www12.statcan.gc.ca/census-recensement/2011/geo/bound-limit/bound-limit-2016-eng.cfm) site.
#
# Exploring the gml file and computing the area and centroid of the distribution areas can be done using the [osgeo library](https://pcjericks.github.io/py-gdalogr-cookbook/geometry.html#quarter-polygon-and-create-centroids).
# File T1901EN.CSV was downloaded from the [Canadian Census Data](https://www12.statcan.gc.ca/census-recensement/2016/dp-pd/hlt-fst/pd-pl/comprehensive.cfm) site
# Exploring the gml file and computing the area and centroid of the distribution areas can be done using the [osgeo library](https://pcjericks.github.io/py-gdalogr-cookbook/geometry.html#quarter-polygon-and-create-centroids).
#
# '''python
#
# # Exporting to geojson
# from osgeo import ogr
#
# # Create test polygon
# ring = ogr.Geometry(ogr.wkbLinearRing)
# ring.AddPoint(1179091.1646903288, 712782.8838459781)
# ring.AddPoint(1161053.0218226474, 667456.2684348812)
# ring.AddPoint(1214704.933941905, 641092.8288590391)
# ring.AddPoint(1228580.428455506, 682719.3123998424)
# ring.AddPoint(1218405.0658121984, 721108.1805541387)
# ring.AddPoint(1179091.1646903288, 712782.8838459781)
# poly = ogr.Geometry(ogr.wkbPolygon)
# poly.AddGeometry(ring)
#
# geojson = poly.ExportToJson()
# print geojson
#
#
# '''python
# # Get centroid
# import ogr
#
# # Given a test polygon
# poly_Wkt= "POLYGON((-107.42631019589980212 40.11971708125970082,-107.42455436683293613 40.12061219666851741,-107.42020981542387403 40.12004414402532859,-107.41789122063043749 40.12149008687303819,-107.41419947746419439 40.11811617239460048,-107.41915181585792993 40.11761695654455906,-107.41998470913324581 40.11894245264452508,-107.42203317637793702 40.1184088144647788,-107.42430674991324224 40.1174448122981957,-107.42430674991324224 40.1174448122981957,-107.42631019589980212 40.11971708125970082))"
# geom_poly = ogr.CreateGeometryFromWkt(poly_Wkt)
#
#
# geom_poly_envelope = geom_poly.GetEnvelope()
# minX = geom_poly_envelope[0]
# minY = geom_poly_envelope[2]
# maxX = geom_poly_envelope[1]
# maxY = geom_poly_envelope[3]
#
# '''
# coord0----coord1----coord2
# | | |
# coord3----coord4----coord5
# | | |
# coord6----coord7----coord8
# '''
# coord0 = minX, maxY
# coord1 = minX+(maxX-minX)/2, maxY
# coord2 = maxX, maxY
# coord3 = minX, minY+(maxY-minY)/2
# coord4 = minX+(maxX-minX)/2, minY+(maxY-minY)/2
# coord5 = maxX, minY+(maxY-minY)/2
# coord6 = minX, minY
# coord7 = minX+(maxX-minX)/2, minY
# coord8 = maxX, minY
#
# ringTopLeft = ogr.Geometry(ogr.wkbLinearRing)
# ringTopLeft.AddPoint_2D(*coord0)
# ringTopLeft.AddPoint_2D(*coord1)
# ringTopLeft.AddPoint_2D(*coord4)
# ringTopLeft.AddPoint_2D(*coord3)
# ringTopLeft.AddPoint_2D(*coord0)
# polyTopLeft = ogr.Geometry(ogr.wkbPolygon)
# polyTopLeft.AddGeometry(ringTopLeft)
#
#
# quaterPolyTopLeft = polyTopLeft.Intersection(geom_poly)
# centroidTopLeft = quaterPolyTopLeft.Centroid()
# '''
#
# '''python
# # Create geometry from gml
# from osgeo import ogr
#
# gml = """<gml:Point xmlns:gml="http://www.opengis.net/gml"><gml:coordinates>108420.33,753808.59</gml:coordinates></gml:Point>"""
# point = ogr.CreateGeometryFromGML(gml)
# print "%d,%d" % (point.GetX(), point.GetY())
# '''
# Note: must install gdal module
# +
# %%time
from osgeo import ogr
with open('lda_000b16g_e.gml','r') as f:
geo_gml = f.read()
# -
len(geo_gml)
g = ogr.CreateGeometryFromGML(geo_gml)
print(g)
# +
from osgeo import ogr
source = ogr.Open('lda_000b16g_e.gml')
# -
type(source)
dir(source)
s2=source.GetLayer(0)
s2
len(s2)
s4 = s2[0]
s4
s4.GetGeometryRef()
print(s4.GetGeometryRef())
print(s2.GetSpatialRef())
print(s4.GetGeometryRef().GetSpatialReference())
# +
from osgeo import ogr
from osgeo import osr
inref = s4.GetGeometryRef().GetSpatialReference()
outref = osr.SpatialReference()
outref.ImportFromEPSG(4326)
transform = osr.CoordinateTransformation(inref, outref)
point = s4.GetGeometryRef().Centroid() # ogr.CreateGeometryFromWkt("POINT (1120351.57 741921.42)")
point.Transform(transform)
print(point.ExportToWkt())
print(point)
print(point.GetX(), point.GetY())
# -
s4.GetGeometryRef().Centroid()
print(s4.GetGeometryRef().Centroid())
s4.GetGeometryRef().Centroid().ExportToJson()
s4.GetGeometryRef().Centroid().ExportToWkt()
s4.GetGeometryRef().Area()
s2.GetLayerDefn()
# +
layerDefinition = s2.GetLayerDefn()
for i in range(layerDefinition.GetFieldCount()):
print(layerDefinition.GetFieldDefn(i).GetName(),f" \t",s2.GetFeature(0).GetFieldAsString(s2.GetFeature(0).GetFieldIndex(layerDefinition.GetFieldDefn(i).GetName())))
# -
# See the feature name definitions [here](https://www150.statcan.gc.ca/n1/pub/92-160-g/2016002/tbl/tbl_4.13-eng.htm).
# But this is for 2011.
#
# The meaning of prefixes for NAME (N), UID (U), PUID (PU), TYPE (T), and CODE (C):
# * DA U Dissemination Area unique identifier (composed of the 2-digit province/territory unique identifier followed by the 2-digit census division code and the 4-digit dissemination area code)
# * PR U,N Province or territory
# * CD U,N,T Census Division
# * CCS U,N Census Consolidated Subdivision
# * CSD U,N,T Census Subdivision
# * ER U,N Economic Region
# * SAC T,C Statistical Area Classification: Part of are a component of a census metropolitan area, a census agglomeration, a census metropolitan influenced zone or the territories?
# * CMA U,PU,N,T Census Metropolitan Area or Census Agglomeration name, PU Uniquely identifies the provincial or territorial part of a census metropolitan area or census agglomeration (composed of the 2-digit province or territory unique identifier followed by the 3-digit census metropolitan area or census agglomeration unique identifier)
# * CT U,N Census Tract within census metropolitan area/census agglomeration
# * ADA U Aggregate dissemination area unique identifier
#
s5 = s2.GetName()
s5
s2[1016].GetField("DAUID")
# +
s6 = s2[1010]
s6.GetFieldCount()
for i in range(s6.GetFieldCount()):
print(s6.GetFieldDefn(i).GetName())
# -
for feature in s2:
print(feature.GetField("CSDNAME"))
for feature in s2:
geom = feature.GetGeometryRef()
print geom.Centroid().ExportToWkt()
| exploration_osgeo.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: jp-Babel (Node.js)
// language: babel
// name: babel
// ---
import * as Utils from 'causal-net.utils';
import * as Log from 'causal-net.log';
import * as Preprocessing from 'causal-net.preprocessing';
import { causalNetCore } from 'causal-net.core';
import * as fs from 'fs';
var R = causalNetCore.CoreFunction;
var T = causalNetCore.CoreTensor;
var { imagePreprocessing } = Preprocessing;
var { PNG } = Utils;
imagePreprocessing
var image = [];
var filePath = '../datasets/MNIST_dataset/data.png';
(async ()=>{
function readImage(filePath){
return new Promise((resolve, reject)=>{
let pngParser = new PNG();
pngParser.on('parsed', function(){
console.log(this.data.length);
resolve(this.data);
})
fs.createReadStream(filePath).pipe(pngParser)
});
}
image = await readImage(filePath);
})();
var splitedImage = imagePreprocessing.imageSplit(28*28*4, image)
imagePreprocessing
| notebooks/mnist.visualization.node.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia faststart
# language: julia
# name: julia-1.5
# ---
# # Michaelis–Menten fitting
#
# Inhibited enzyme reactions often follow what are known as _Michaelis–Menten_ kinetics, in which a reaction rate $v$ follows a law of the form
#
# $$v(x) = \frac{V x}{K_m + x},$$
#
#
# where $x$ is the concentration of a substrate. The real values $V$ and $K_m$ are parameters that are free to fit to data. For this example we cook up some artificial data with $V=2$ and $K_m=1/2$.
using FundamentalsNumericalComputation
m = 25;
x = LinRange(0.05,6,m)
yf = @. 2*x/(0.5+x) # exactly on the curve
y = @. yf + 0.15*cos(2*exp(x/16)*x); # noise added
scatter(x,y,label="actual data",xlabel="x",ylabel="v",leg=:bottomright)
plot!(x,yf,l=:dash,label="nominal data")
# The idea is to pretend that we know nothing of the origins of this data and use nonlinear least squares on the misfit to find the parameters in the theoretical model function $v(x)$. Note in the Jacobian that the derivatives are _not_ with respect to $x$, but with respect to the two parameters, which are contained in the vector `c`.
# +
function misfit(c)
V,Km = c # rename components for clarity
return @. V*x/(Km+x) - y
end
function misfitjac(c)
V,Km = c # rename components for clarity
J = zeros(m,2)
J[:,1] = @. x/(Km+x) # d/d(V)
J[:,2] = @. -V*x/(Km+x)^2 # d/d(Km)
return J
end
# +
c1 = [1, 0.75]
c = FNC.newtonsys(misfit,misfitjac,c1)
@show V,Km = c[end] # final values
# -
# The final values are close to the noise-free values of $V=2$, $K_m=0.5$ that we used to generate the data. We can calculate the amount of misfit at the end, although it's not completely clear what a "good" value would be. Graphically, the model looks reasonable.
model = x -> V*x/(Km+x)
residual = @. model(x) - y
@show final_misfit_norm = norm(residual);
plot!(model,0,6,label="MM fit" )
# For this model, we also have the option of linearizing the fit process. Rewrite the model as $1/v= (\alpha/x)+\beta$ for the new parameters $\alpha=K_m/V$ and $\beta=1/V$. This corresponds to the misfit function whose entries are
#
# $$f_i(\alpha,\beta) = \alpha \cdot \frac{1}{x_i} + \beta - \frac{1}{y_i},$$
#
# for $i=1,\ldots,m$. Although the misfit is nonlinear in $x$ and $y$, it's linear in the unknown parameters $\alpha$ and $\beta$, and so can be posed and solved as a linear least-squares problem.
A = [ x.^(-1) x.^0 ]
u = 1 ./ y
z = A\u
α,β = z
# The two fits are different, because they do not optimize the same quantities.
linmodel = x -> 1 / (β + α/x)
@show final_misfit_linearized = norm(linmodel.(x)-y);
plot!(linmodel,0,6,label="linearized fit")
# The truly nonlinear fit is clearly better in this case. It optimizes a residual for the original measured quantity rather than a transformed one we picked for algorithmic convenience.
| book/nonlineqn/demos/nlsq-MM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="oA6GZzYgRBsu"
# <img src="https://www.kaunokolegija.lt/kk_wp_content/uploads/sites/5/2020/05/kaunas-university-of-applied-sciences.png" width="300"/>
#
# ------
# + [markdown] id="6QBqbb8BRFzu"
# # Recurrent Neural Networks
#
# A case study of univariate time series analysis.
#
# ### Practical Session
#
# <br/> Prof. Dr. <NAME>
# <br/> email: <EMAIL>
# <br/>last update: June 25, 2021
#
#
# --------
# + [markdown] id="TdhSL4xTROUn"
# ## Contents
#
# 1. [Challenge](#challenge)
# 2. [Download the data](#download-the-data)
# 3. [Visualize the stock price history](#visualize-the-stock-price-history)
# 4. [Data transforms and preprocessing](#data-transforms-and-preprocessing)
# 5. [Build the RNN](#build-the-rnn)
# 6. [Train the RNN](#train-the-rnn)
# 7. [Improve the RNN](#improve-the-rnn)
# 8. [Fine tune the RNN](#fine-tune-the-rnn)
# + [markdown] id="CU-lVwMmRTNK"
# ## Challenge <a name="challenge"></a>
#
# <img src="https://miro.medium.com/max/3504/1*NpT5pyemQQsGEHXbfS51Zw.png" width="600" align="left"/>
#
# + [markdown] id="WehN_1mKRhu4"
# Given a 5-year history of any stock traded in NASDAQ predict the stock prices for the period of the recent-most month that are not included in the historical data.
#
# To address this challenge we will employ [**univariate time series analysis**](www.homepages.ucl.ac.uk/~uctpsc0/Teaching/GR03/TS1.pdf) with [**recurrent neural networks**](https://stanford.edu/~shervine/teaching/cs-230/cheatsheet-recurrent-neural-networks).
# + [markdown] id="mReQATWhIv6d"
# ## Download the data <a name="download-the-data"></a>
# + id="TGl4xmaGRYCc"
# importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="hvLG67JMRlJf" colab={"base_uri": "https://localhost:8080/"} outputId="3d4af5bf-8824-49d3-d6ed-921fb271dc54"
# install yahoo-finance
# !pip install yfinance
# + id="gBW1a1O3RnKx" colab={"base_uri": "https://localhost:8080/"} outputId="740f576b-9995-4335-e767-e20f47acc255"
# # copy some custom code files
# !wget https://raw.githubusercontent.com/georgiosouzounis/deep-learning-lectures/main/code/NASDAQ_io.py
# + id="CB8DpuO6_sp1"
# import necessary functions
from NASDAQ_io import * #getStockTickerSymbols, searchBySymbol, getStockPriceHistory, getDateTime
# + id="jalyBe1iRs2G"
# get the companies listed in NASDAQ
companies = getStockTickerSymbols()
# + id="Nn2ULPRPRvGq" colab={"base_uri": "https://localhost:8080/", "height": 511} outputId="16d9f5c5-0608-47b7-bb3b-3be0ca56cd78"
# view the companies
companies
# + id="j0FSCohxRv7z"
# Let us go for Tesla
symbol = 'TSLA'
# + id="1YTwxW9v_A3E"
# check if the symbol exists; if it doesn't the dataframe will be empty
df = searchBySymbol(companies, symbol)
# + id="BK4aLl4m99D5"
# set the strat and end date for our training data
start_date = getDateTime(2016,1,1)
end_date = getDateTime(2021,5,31)
# + id="EGy3NceYR2yM" colab={"base_uri": "https://localhost:8080/"} outputId="73e768ac-a4ae-4c7b-9952-d5a2bd380315"
# get the stock history
stock_history = getStockPriceHistory(df, start_date, end_date)
# + id="f1N84CzJrbhq" colab={"base_uri": "https://localhost:8080/", "height": 455} outputId="a5c24a1d-eaff-4f1b-9321-03f380da6762"
# view the contents of the dataframe
stock_history
# + [markdown] id="Qwetf-Nt5eVR"
# ### a quick visualization example
# + colab={"base_uri": "https://localhost:8080/", "height": 679} id="RV-gww_E4nHT" outputId="7252e26a-ae96-4b27-9743-cb8603317582"
# let us plot a quick chart to see the history of the stock in the timeframe specified
stock_history['Close'].plot(figsize=(16, 12))
# + colab={"base_uri": "https://localhost:8080/", "height": 687} id="KonITMNU5B4q" outputId="075a237f-1501-4523-ebdb-d639a40ab5d8"
# use truncate() to zoom into a specific date range
stock_history.truncate(before='2020-03-01')['Close'].plot(figsize=(16, 12))
# + id="SxMdovBWR8sB"
# The Date is set as the dataframe index. This is not very elegant!
# Use the reset_index() function to make this the first column in
# your dataframe and reset the index
stock_history.reset_index(inplace=True)
# + id="8Qj9FFRvTiIm" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="898161aa-4ec0-4864-9a87-e5255976c2e1"
# confirm the change
stock_history
# + [markdown] id="Nk4-N88VooQp"
# ## Visualize the stock price history <a name="visualize-the-stock-price-history"></a>
#
# The visualization code is a modified version of the original, preseneted by [Trifunovic Uros](https://trifunovic-uros.medium.com/) in [Medium.com](https://medium.com/analytics-vidhya/visualizing-historical-stock-price-and-volume-from-scratch-46029b2c5ef9) on Mar. 23, 2021.
#
# Start by getting a copy of the code in the local path
# + id="NmUA0JEMSCB2" colab={"base_uri": "https://localhost:8080/"} outputId="aea26703-9cb3-4ede-86e9-c91eb8f59dca"
# !wget https://raw.githubusercontent.com/georgiosouzounis/deep-learning-lectures/main/code/stock_price_chart.py
# + id="r9MafdA7SV8j"
# import the get_charts() function
from stock_price_chart import get_charts
# + id="nyWd_ja3Sqgu" colab={"base_uri": "https://localhost:8080/", "height": 728} outputId="57842b90-1ebe-4688-daac-0c2d57210ef3"
# plot the contents of our dataframe
get_charts(stock_history)
# + [markdown] id="e2TSkYFIqr2v"
# ## Data transforms and preprocessing <a name="data-transforms-and-preprocessing"></a>
#
# Next we need to tidy-up our data and transform it according to the needs of our
# project
# ### Data clean-up
# + id="b2gyDdYxSsmU"
# extract the relevant data, i.e. `Open` values
# the .values makes this a vector numpy array
training_set = stock_history.iloc[:, 1:2].values
# + id="_BxncqBArqgg" colab={"base_uri": "https://localhost:8080/"} outputId="3827a014-1395-45de-933b-2bcb06499a24"
training_set
# + [markdown] id="yND4Bs9JtgT1"
# ### Feature Scaling
#
# Next we need to rescale our data to the range from 0.0 to 1.0.
#
# Feature scaling is essential as discussed if the Features lecture and needs to be applied to both the training and test sets.
#
# It is computed using the ScikitLearn library [MinMaxScaler()](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html#sklearn.preprocessing.MinMaxScaler) which transforms the selected feature by scaling it to a given range. If more than one, this estimator scales and translates each feature individually such that it is in the given range on the training set, i.e. between zero and one.
# + id="Nue4vBM1tUI4"
# import the MinMaxScaler
from sklearn.preprocessing import MinMaxScaler
# + id="HZE8mnR8t46g"
# create a scaler instance to rescale all data to the range of 0.0 to 1.0
sc = MinMaxScaler(feature_range = (0, 1))
# + id="BTtu_cbxt8Ac"
# create the actual training set of scaled values
training_set_scaled = sc.fit_transform(training_set)
# + id="zWe9ghvIt_4-" colab={"base_uri": "https://localhost:8080/"} outputId="da2377b8-8834-4d8c-b560-4af127754307"
# confirm feature scaling
training_set_scaled
# + [markdown] id="Hg8Q4j1guJG0"
# ### Splitting the training set to dependent and independent variables
#
# The stock prices are stored in a 2D numpy array containing a single column of data.
#
# From this array we need to define our feature vectors. Each feature vector starts with the stock price a day later compared to its predecessor. It is of fixed length, i.e it consists of a fixed number of consecutive stock prices. The stock price after the last one inserted in each feature vector can be considered as our dependent variable, i.e. the N stock prices before today can be used to predict today's price. N is the number of timesteps.
# + [markdown] id="kytPJkBTuKuA"
# <img src="https://drive.google.com/uc?id=1bckuLGZCeLUzNA-xJCGOODzC-4n2U-If"/>
# + id="cTsXaCAmuEv0"
# define the number of timesteps
timesteps = 60 # 60 stock prices or 3 months worth of data assuming 20 working days a month
# each row is to contain the last 60 stock prices before the reference date
X_train = []
# each entry (this is a 1D vector) is to contain teh stock price at the reference date
y_train = []
# + id="kXPTJ8JJySvL" colab={"base_uri": "https://localhost:8080/"} outputId="a0b95fa5-d51b-4d45-984c-ac96bf2f1d32"
training_set_scaled.shape
# + id="Zm_90YUWyXBG"
# we start from day 60 because that is the first instance allowing us to
# go back 60 days; first day is 0
for i in range(timesteps, training_set_scaled.shape[0]):
# 0 is the column ID, the only column in this case.
# put the last 60 days values in one row of X_train
X_train.append(training_set_scaled[i-60:i, 0])
y_train.append(training_set_scaled[i, 0])
# + id="LhPpPZjPyolg"
# convert these to numpy arrays
X_train, y_train = np.array(X_train), np.array(y_train)
# + id="TjckpEXzyySa" colab={"base_uri": "https://localhost:8080/"} outputId="15449315-cde7-4698-fbec-b99965ed8f46"
# the X_train 2D numpy array has 60 rows less compared to the original dataset
X_train.shape
# + [markdown] id="Bw7IIoQ30vPF"
# ### Reshaping the Matrix
# + [markdown] id="POHKyNR-00yA"
# We need to add a new matrix dimension to accommodate the indicator (predictor).
#
# NumPy matrices are tensors (3D) and essentially we need to specify that our matrix consists of **60 days** (dimension x) times **total days in data set** (dimension y) times **1 value per matrix cell (scalar)** (dimension z). If we were to include the value of a different stock with the the past 60 days of Tesla, we would need to change the length of the 3d dimension to 2. RNN training tables are 3D!!! Read: [Reshaping NumPy Array | Numpy Array Reshape Examples](https://backtobazics.com/python/python-reshaping-numpy-array-examples/)
# + id="xtCooPLO1_rX"
# Reshaping the data matrix, we retain the 2 original dimensions and add a third of depth=1
X_train = np.reshape(X_train, (X_train.shape[0], X_train.shape[1], 1))
# + [markdown] id="_WEbnMrwKJ1z"
# ## Build the RNN <a name="build-the-rnn"></a>
# + [markdown] id="z_Jy9wfEKMV0"
# ### RNN initialization
#
# - Import the sequential model from the Keras API;
# - Import the Dense layer template from the Keras API;
# - Import the LSTM model from the Keras API
# - Create an instance of the sequential model called regressor because we want to predict a continuous value
# + id="svBnHDIH2qO1"
# Importing the Keras libraries and packages
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from keras.layers import Dropout
# + id="hKGJjzmIKR3f"
# Initialising the RNN as a sequence of layers
regressor = Sequential()
# + [markdown] id="wKebpAqcKXDo"
# ### Add First Layer
#
# We first add an object of the LSTM class!
#
# - The first argument is the number of units or LSTM memory cells. Include many neurons to address the high dimensionality of the problem; say 50 neurons!
# - Second arg: return sequences = true; stacked LSTM !
# - Third arg: input 3D shape: observations vs time steps vs number of indicators
# + id="UPrMlppDKTXK"
# Adding the input layer and the LSTM layer
regressor.add(LSTM(units = 50, return_sequences = True, input_shape = (X_train.shape[1], 1)))
# + id="k9PvyarKKauB"
# the argument is the dropout rate to ignore in the layers (20%),
# i.e. 50 units * 20% = 10 units will be dropped each time
regressor.add(Dropout(0.2))
# + [markdown] id="YwPyVM1UKgP9"
# ### Add More Layers
#
# We can add more LSTM layers but along with Dropout regularization to make sure we avoid overfitting!
#
# We don’t need to add the shape of the layer again because it is recognized automatically from the number of input units.
#
# The last layer does not return a sequence but connected directly to a fully connected output layer.
# + id="8Y0olBBdKcDk"
# Adding a second LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
# + id="T9xORNaLKj1v"
# Adding a third LSTM layer and some Dropout regularisation
regressor.add(LSTM(units = 50, return_sequences = True))
regressor.add(Dropout(0.2))
# + id="xd676oPKKlMC"
# Adding a fourth LSTM layer and some Dropout regularisation
# we removed the return_sequences because we no longer return a
# sequence but a value instead
regressor.add(LSTM(units = 50))
regressor.add(Dropout(0.2))
# + [markdown] id="tEbq5tG8KpDI"
# ### Add Output Layer & Compile
#
# The output has 1 dimension , i.e. one value to be predicted thus or output fully connected layer has dimensionality = 1.
#
# - **Optimizer**: rmsprop is recommended in the Keras documentation. The Adam optimizer is also a powerful choice.
# - **Loss function**: regression problems take the mean square error as most common
# + id="zpiJ1X0AKmb9"
# Adding the output layer
regressor.add(Dense(units = 1))
# + id="WjEnNuUmKtfq"
# Compiling the RNN
regressor.compile(optimizer = 'adam', loss = 'mean_squared_error')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="K_pjAjiw05jc" outputId="9e39f1e6-bb2f-4038-c2eb-489a7638d62d"
# review the network architecture
from keras.utils.vis_utils import plot_model
plot_model(regressor, to_file='model_plot.png', show_shapes=True, show_layer_names=True)
# + [markdown] id="D9hK_3Q-Kx4g"
# ## Train the RNN <a name="train-the-rnn"></a>
# + [markdown] id="CBVpsF0EK4nB"
# ### Fit the RNN to the Training set
#
# We now want to train our RNN using the data in our **Training Set X** and **predictors in y** (ground truth in this case). Parameters that can be specified are the:
#
# - **Batch size**: update the cell weights not on every stock price on every batch_size values;
# - **Number of epochs**: how many iterations to be used, i.e. number of forward and backward propagations for the update of the weights
# + id="RlW3HDbIK29t" colab={"base_uri": "https://localhost:8080/"} outputId="d244a1f0-e7b2-48b6-9bec-3aa046acfefa"
# Fitting the RNN to the Training set
regressor.fit(X_train, y_train, epochs = 100, batch_size = 32)
# + [markdown] id="h8gZFgBAOW4P"
# ## Deploy the RNN <a name="deploy-the-rnn"></a>
# + [markdown] id="eYueYBfdK_AB"
# ### Computing Predictions
#
# Get the test set in a new dataframe. That will be one month of stock prices after the end of the preior used for training.
#
# There are 20 (19 to 21) financial days in one month, weekends excluded!
# + id="Ob4psLLNK7yb"
# Getting the real stock price
# first set the date range (make sure it does not overlap with our training data period)
start_date = getDateTime(2021,6,1)
end_date = getDateTime(2021,6,30)
# + id="x2wS-EywNSva" colab={"base_uri": "https://localhost:8080/"} outputId="e541da5d-b98f-4989-ea66-373512bd88d1"
# get the relevant stock proce history
dataset_test = getStockPriceHistory(df, start_date, end_date)
dataset_test.reset_index(inplace=True)
# + id="oLHkDE4dLDcn" colab={"base_uri": "https://localhost:8080/"} outputId="b57aa8ae-a348-427e-f388-dc901b413758"
# extract the relevant columns
real_stock_price = dataset_test.iloc[:, 1:2].values
real_stock_price.size
# + id="XZpRcm_TLFYD" colab={"base_uri": "https://localhost:8080/"} outputId="77ba56d5-9af2-4f6a-d311-4c71e1675441"
# verify your data
real_stock_price
# + [markdown] id="fSFS-8O8LHOY"
# To predict the stock price value for each day in this date range, we need the values in the last 60 days.
#
# To obtain this **history** we need to combine both the training and test sets in one.
#
# If we were to use the training_set and test_set we would need to use the scaler but that would change the actual test values. Thus concatenate the original data frames!
# + id="W5Y88AHkLJj0"
# axis = 0 means concatenate the lines (i.e. vertical axis)
dataset_total = pd.concat((stock_history['Open'], dataset_test['Open']), axis = 0)
# + id="9TusBCXYLQxV" colab={"base_uri": "https://localhost:8080/"} outputId="efe43327-142d-4995-998f-40015c76edc9"
dataset_total.size
# + id="QJoti2VsLT5G"
# the difference in the length of the first two gives us
# the first day in the new date range, and we need to go back 60 days to get the necessary range
inputs = dataset_total[len(dataset_total) - len(dataset_test) - timesteps:].values
# + id="Tw2NzN_6LUsk" colab={"base_uri": "https://localhost:8080/"} outputId="91abadc5-0a2d-4f05-9b57-af675d9b0fd4"
inputs.size
# + id="IOntslnMLWco"
# we did not use iloc from panda so lets reshape the numpy array for
# compatibility: i.e. all the values from input lines to be stacked in one
# column. The -1 means that the numpy has no knowledge of how the
# values were stored in lines. The 1 means we want to them in one
# column.
inputs = inputs.reshape(-1,1)
# apply the feature scaler
inputs = sc.transform(inputs)
# + [markdown] id="8Y4adPmSLZVj"
# 1. For each price in test date range we need the **immediate 60 values** before it.
# 2. We have 21 prices in June;
# 3. We need a numpy 3D array of 60 prices (columns) times 21 days (rows) times 1 dependent variable
# 4. We don’t need y_test. That is what we are trying to compute!
# + id="kdlX_S6NLbmE"
# Getting the predicted stock price of 2017
X_test = []
# + id="vrLyeEzzLeV_"
# the first 60 from inputs are from training set; start
# from 60 and get the extra 21, i.e. up to 81
for i in range(timesteps, inputs.size):
X_test.append(inputs[i-timesteps:i, 0])
# + id="TEBuVmlZLf2g"
X_test = np.array(X_test) # not 3D structure yet
# + id="mGi3PTs_Li_S"
# create a 3D structure
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
# + id="q6rVxW66LjGK"
# source this input to our regressor model
predicted_stock_price = regressor.predict(X_test)
# + id="Ft8H1ifSLmtm" colab={"base_uri": "https://localhost:8080/"} outputId="c2fdcd2a-9230-435f-c722-df49e70b3073"
# need to inverse the scaling to get meaningful predicted stock price # outputs
predicted_stock_price = sc.inverse_transform(predicted_stock_price)
predicted_stock_price.size
# + [markdown] id="6CzDUW3CLoUK"
# ### Result Visualization
# + id="aE-WgHRPLsa_" colab={"base_uri": "https://localhost:8080/", "height": 621} outputId="d3a616e7-de45-46ef-ed20-f63078601913"
# Visualising the results
plt.plot(real_stock_price, color = 'red', label = 'Real Tesla Stock Price')
plt.plot(predicted_stock_price, color = 'blue', label = 'Predicted Tesla Stock Price')
plt.title('Tesla Stock Price Prediction')
plt.xlabel('Time')
plt.ylabel('Tesla Stock Price')
plt.legend()
plt.show()
# + [markdown] id="527YCNE_LyMD"
# The blue line shows the trend of the stock for the month of January 2017.
#
# Some observations:
#
# - The prediction lags behind the actual price curve because the model cannot react to fast non-linear changes. Spikes are examples of fast non-linear changes
# - Model reacts pretty well to smooth changes
#
#
# + [markdown] id="tN3c8G77L48A"
# ### Compute the RMSE
#
# If we need to compute the RMSE for our Stock Price Prediction problem, we use the real stock price and predicted stock price as shown.
#
# Then consider dividing this RMSE by the range of the Google Stock Price values of January 2017 to get a relative error, as opposed to an absolute error.
# + id="8SQnOtDgLv50"
#import the libraries
import math
from sklearn.metrics import mean_squared_error
# + id="XoGDYOUUL8tb" colab={"base_uri": "https://localhost:8080/"} outputId="499baf80-7704-4431-d2ec-966ad543adb3"
rmse = math.sqrt( mean_squared_error( real_stock_price[0:21,:], predicted_stock_price))
rmse
# + [markdown] id="j6srtrdLL-ui"
# ## Improve the RNN <a name="improve-the-rnn"></a>
# + [markdown] id="JoRyqGZVMDh2"
# The new data need to be placed in the same order/format as in the case of the training/test sets.
#
# 1. Getting more training data: we trained our model on the past 5 years of the Google Stock Price but it would be even better to train it on the past 10 years.
#
# 2. Increasing the number of time steps: the model remembered the stock price from the 60 previous financial days to predict the stock price of the next day. That’s because we chose a number of 60 time steps (3 months). You could try to increase the number of time steps, by choosing for example 120 time steps (6 months).
#
# 3. Adding some other indicators: if you have the financial instinct that the stock price of some other companies might be correlated to the one of Google, you could add this other stock price as a new indicator in the training data.
#
# 4. Adding more LSTM layers: we built a RNN with four LSTM layers but you could try with even more.
#
# 5. Adding more neurons in the LSTM layers: we highlighted the fact that we needed a high number of neurons in the LSTM layers to respond better to the complexity of the problem and we chose to include 50 neurons in each of our 4 LSTM layers. You could try an architecture with even more neurons in each of the 4 (or more) LSTM layers.
# + [markdown] id="_OJqZPvSMHup"
# ## Fine tune the RNN <a name="fine-tune-the-rnn"></a>
#
# Parameter Tuning on the RNN model: we are dealing with a Regression problem because we predict a continuous outcome.
#
# Tip: replace: scoring = 'accuracy' by scoring = 'neg_mean_squared_error' in the GridSearchCV class parameters as we did in the ANN case.
# + id="jh4hCygu8Y-5"
| recurrent_neural_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # References
# The 5 measure factors:
# - Throughput
# - Efficiency
# - Latency
# - ACcuracy
# - Memory Usage
#
# TensorRT Developer guide p3
# 1. Export model to format parsable by TensorRT: ONNX, Caffe, UFF.
# 2. Choose optimization options
# 3. TensorRT create an optimized inference runtime engine.
# 4. The engine can be serialized / loaded again.
# 5. The engine can run inference
#
# TensorRT devloper guide p7
# TensorRT takes network def, perform optis, generate runtime engine.
#
# Optis are:
# - Eliminator of unused layers and no-ops
# - Layer & tensor fusion
# - Kernel Auto-tuning
# - Modify precision of weights
#
# TensorRT developer guide p8
# TensorRT core interface:
# - Network Definition
# - Builder
# - Engine
#
# TensorRT developer guide p9
# C++ API steps
#
# 1. Define a network
# - Manually with API
# - Using a parser of file ONNX, Caffe or UFF
#
# 2. Build an engine
# - From the network def, optimize it
# - From a file, desarialize the engine
#
# 3. (Optional) Serialize the engine
#
# 4. Perform Inference
# 4.1. Create execution context
# 4.2. Prepare input and output buffers
# 4.3. enquele kernels on a CUDA stream
# 4.4. use sync methods such as other CUDA stream to get results
#
# TensorRT developer guide chapter 2
# Create custom layers:
#
# Extend tensorRT functionality: creating a custom layer with implem.
#
# 1. Create class that herits from a plugin base class
# 2. Create class that herits from IPluginCreator
# 1. Use the plugin registery to get the plugin creator
# 1. Create plugin using plugin name and metada
# 1. Add the plugin into the current network definition
#
# TensorRT developer guide chapter 4
# Mixed precision
#
# Types: fp32, fp16, quantized int8.
# You can set the precision for any layer. It actually set the prefered types for input and outputs.
# It may insert reformat ops if necessary.
# These are only preferences, TensorRT chose it only if it leads to higher performance (some layers may be faster in higher precision). It may also not exist such a layer implem for this precision.
# You can also set the prefered type for the whole network.
#
# For int8 quanrization, TensorRT need a scale factor.
# Only supports Symmetric quantization: scale calculated with absolute maximum dynamic range values.
# The dyanmic range can be set manually for each tensor, or it can be computed using calibration dataset.
#
# Explicit precision network: precision of all tensors and layers known, usefull to export pre-quantized models with scaling already established.
#
# TensorRT developer guide chapter 5
# Optimization profile: describe range of dimensions for each network input, and the dimensions the autotuner should use.
# Usefull when having network with dynamic shapes (only resolved at run-time).
#
# TensorRT developer guide section 7.2
# Loops
#
# Loops are defined wih 4 layer kinds:
#
# - ITripLimitLayer: number of iterations
# - IIteratorLayer: iterate over a tensor
# - IRecurrenceLayer: recurrent definition
# - ILoopOutputLayer: Output value of the loop
#
# The loop body can only have a restricted kind of layers, and nested loops.
#
# TensorRT developer guide chapter 8
# Quantization
#
# TensorRT has quantization and dequantization operations.
# 1. Train a model using QAT (Quantized Aware Training) with TensorFlow
# 2. Export the model to quantized ONNX
# 3. Export the mort to TensorRT using explicit precision mode
#
# TensorRT developer guide chapter 9
# Graph surgeon API:
#
# Manipulate TF grapg. Search, Add, Edit, Remove nodes.
# You can mark some nodes as plugins for custom layers implementations.
#
# TensorRT developer guide section 12.1.5
| refs/tensorrt/references.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Polynomial Regression
# **Importing Necessary Files**
#importing libraries
import numpy as np
import pandas as pd
import sklearn.preprocessing
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set()
# **Task 1: Import the 'Boston.csv' data file**
# write code here
data =pd.read_csv('Boston.csv')
df=data.copy()
# **Task 2: Make a copy of data in 'df'**
# +
# Write cod
# -
# **Task 3: Display top 5 rows of data**
# Write code here
df.head()
columnss = ['Crime Rate','Residential Proportion','non-retail business acres/Town','Charles River',
'NO2 concentration','Average Rooms/Dwelling.','Prior Built Units Proportion','Distance to Employment Centres',
'Radial Highways Distance','ValueProperty/tax rate','Teacher/town','blacks/town','Lower Status Percent']
# ### Creating Independent and Dependent Variables
# **Task 4: Create Independent and Dependent variables in 'X' and 'Y' and print their respective shapes.**
# Write code here
X =df.drop(['median home price'],axis=1)
Y =df[['median home price']]
print(X.shape)
print(Y.shape)
# ### Normalizing Data
# **Task 5: Normalize the data by applying MinMax Scaling on X.**
# Write code here
from sklearn.preprocessing import MinMaxScaler
min_max=MinMaxScaler()
x_min_max=min_max.fit_transform(X)
X=pd.DataFrame(x_min_max,columns=columnss)
X.head()
# ### Splitting Data
# **Task 6: Split the data into training and testing set and print their shapes.**
# Write code here
from sklearn.model_selection import train_test_split
xtrain ,xtest, ytrain, ytest =train_test_split(X,Y,test_size=0.30,random_state=25,shuffle=True)
print(xtrain.shape,xtest.shape)
print(ytrain.shape,ytest.shape)
# ### Applying Linear Regression
# **Task 7: Apply Linear Regression on the data.**
#Write the code here
from sklearn.linear_model import LinearRegression
model = LinearRegression()
# Apply the fit() function on the model
#Write code here
model.fit(xtrain, ytrain)
# Get the predicted values on training data set in train_pred
#Write code here
train_pred = model.predict(xtrain)
# Get the predicted values on test data set in test_pred
#Write code here
test_pred= model.predict(xtest)
# ### Finding R2 Score
# **Task 8: Find out r2 Score for training and testing data.**
#Write code here for testing data
from sklearn.metrics import r2_score
r2_test_lr=r2_score(ytest,test_pred)
#Write code here for training data
r2_train_lr=r2_score(ytrain,model.predict(xtrain))
print('R2 score for testing:',r2_test_lr)
print('R2 score for training:',r2_train_lr )
# **Ploting Residual Plot**
# !pip install yellowbrick
# +
from yellowbrick.regressor import ResidualsPlot
plt.figure(figsize=(15,6))
visualizer = ResidualsPlot(model)
visualizer.fit(xtrain.values, ytrain.values)
visualizer.score(xtest.values, ytest.values)
visualizer.poof()
# -
# # Applying Polynomial Regression
from sklearn.preprocessing import PolynomialFeatures
poly_features = PolynomialFeatures(degree=2)
# transform the features to higher degree features.
X_train_poly = poly_features.fit_transform(xtrain)
# fit the transformed features to Linear Regression
poly_model = LinearRegression()
poly_model.fit(X_train_poly, ytrain)
# predicting on training data-set
y_train_predicted = poly_model.predict(X_train_poly)
# predicting on test data-set
X_test_poly=poly_features.fit_transform(xtest)
y_test_predict = poly_model.predict(X_test_poly)
# evaluating the model on training dataset
r2_train = r2_score(ytrain, y_train_predicted)
# evaluating the model on test dataset
r2_test = r2_score(ytest, y_test_predict)
print ('The r2 score for training set is: ',r2_train)
print ('The r2 score for testing set is: ',r2_test)
# # Model Complexity vs Performance
from sklearn.metrics import mean_squared_error
mse_train=[]
mse_test=[]
r2_train=[]
r2_test=[]
for i in range(1,8):
poly_reg = PolynomialFeatures(degree=i)
X_tr_poly,X_tst_poly= poly_reg.fit_transform(xtrain),poly_reg.fit_transform(xtest)
poly = LinearRegression()
poly.fit(X_tr_poly, ytrain)
y_tr_predicted,y_tst_predict = poly.predict(X_tr_poly),poly.predict(X_tst_poly)
r2_train.append(r2_score(ytrain, y_tr_predicted))
r2_test.append(r2_score(ytest, y_tst_predict))
mse_train.append(mean_squared_error(ytrain, y_tr_predicted))
mse_test.append(mean_squared_error(ytest, y_tst_predict))
# ### Comparison of R2 Score
plt.figure(figsize=(18,5))
sns.set_context('poster')
#plt.subplot(1,2,1)
sns.lineplot(x=list(range(1,8)), y=r2_train, label='Training')
#plt.subplot(1,2,2)
sns.lineplot(x=list(range(1,8)), y=r2_test, label='Testing')
plt.ylim(0.7,1.0)
list(zip(r2_train,r2_test))
# ### Comparison of Mean Squared Error
plt.figure(figsize=(15,4))
#plt.subplot(1,2,1)
sns.lineplot(x=list(range(1,8)), y=mse_train, label='Training')
#plt.subplot(1,2,2)
sns.lineplot(x=list(range(1,8)), y=mse_test, label='Testing')
plt.ylim(0,25) # y axis limit
plt.show();
| Linear Regression/polynomial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 8 - Using Complex Types to Analyse Unstructured or JSON Data
# My challenge of today is to go beyond processing well structured data, which complies to a schema and where all values are clearly seperated into typed columns. Today I want to analyse the stock descriptions in the retail data set, which come as unstructured text. This is my use case to investigate Sparks complex datataypes like arrays and maps. Next to that, I want to get familiar with the processing of semi-structured data like JSON.
# +
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession\
.builder\
.getOrCreate()
retailDF = spark.read\
.option("header", "true")\
.option("inferSchema", "true")\
.format("csv")\
.load("./data/retail-data/by-day/*.csv")
# -
# There are two questions, I want to investigate regarding the description data:
# * What is the average number of words in the Description per StockCode?
# * Which are the most frequently used words?
#
# ## Data Preparation
# The granularity of my analysis is StockCode and not individual invoice items. So to prevent StockCode duplicates, I tailor the data set to get a DataFrame containing distinct StockCodes and their description.
# +
distinctDF = retailDF.select(
"StockCode",
"Description").distinct()
distinctDF.orderBy("StockCode").show(10, truncate=False)
# -
# Apparently the null value problem, I investigated yesterday, occures again. Rows having null values in any column are uselesss for my analysis, so I want to remove them.
# +
cleanedDF = distinctDF.dropna(how="any")
cleanedDF.orderBy("StockCode").show(10, truncate=False)
# -
# ## Arrays
# Next I've to do is to split up the text strings into arrays of words. The words in the descriptions are seperated by blanks, so I define this as split seperator. The result looks like Python lists but in contrast to lists, all array elements must have the same data type.
# +
from pyspark.sql.functions import split
splittedDF = cleanedDF.select(
"StockCode",
split("Description", " ").alias("word_list")
)
splittedDF.show(10, truncate=False)
# -
# Like with normal Python lists I can grab specific elements, i.e. words from my word lists, by referencing their index starting with 0 for the first element. So to get the second word in each description, I need to refer to index 1.
# +
from pyspark.sql.functions import col
splittedDF.select("StockCode", col("word_list")[1]).show(10)
# -
# Interesting to note that InvoiceNo 21249 seems to have a double blank after the first word. Maybe a typo in a free-text field? Anyway, I dont to count words, not blanks, so I have to removing them later. First, I want to double check, if this is a more general or single-case issue.
#
# I can easily check wether or not a word list contains specific key words by using the `array_contains()` function. For my analysis, I want to identify rows having empty words in the list, which I'dont want to count.
# +
from pyspark.sql.functions import array_contains
splittedDF.select(
"StockCode",
"word_list",
array_contains("word_list", "").alias("empty strings inside")
).show(10, truncate=False)
# -
# So now lets let's clean up the word lists and remove any empty words.
# +
from pyspark.sql.functions import array_remove
cleanedWordListDF = splittedDF.select(
"StockCode",
array_remove("word_list", "").alias("word_list")
)
cleanedWordListDF.show(10, truncate=False)
# -
# Did it work?
cleanedWordListDF.select(
"StockCode",
"word_list",
array_contains("word_list", "").alias("empty strings inside")
).show(10, truncate=False)
# yes, it did.
#
# Back to my questions. Now, after having cleaned up the data the number of words per stock description is simply the array length which is provided by the `size()` function.
# +
from pyspark.sql.functions import size
cleanedWordListDF.select(
"StockCode",
size("word_list").alias("num_of_words")
).show(10)
# +
from pyspark.sql.functions import avg
avgDF = cleanedWordListDF.select(
avg(
size("word_list")
).alias("avg_num_of_words")
)
avgDF.show(10)
# -
# So the answer to my first question is that stock descriptions are quite short, just about four words in average.
#
# Pyspark module [pyspark.sql.functions](https://spark.apache.org/docs/latest/api/python/pyspark.sql.html#module-pyspark.sql.functions) provides further array related functions, which I just list here for later reference:
#
# * **array()** - creates a new array column from a list of columns or column expressions that have the **same data type**
# * **array_distinct(col)** - Collection function: removes duplicate values from the array
# * **array_except(col1, col2)** - Collection function: returns an array of the elements in col1 but not in col2, without duplicates
# * **array_intersect(col1, col2)** - Collection function: returns an array of the elements in the intersection of col1 and col2, without duplicates
# * **array_join()**
# * **array_max()** - Collection function: returns the maximum value of the array
# * **array_min()** - Collection function: returns the maximum value of the array
# * **array_position()** - Collection function: Locates the position of the first occurrence of the given value in the given array
# * **array_repeat(col, count)** - Collection function: creates an array containing a column repeated count times
# * **array_sort(col)** - Collection function: sorts the input array in ascending order
# * **array_union(col1, col2)** - Collection function: returns an array of the elements in the union of col1 and col2, without duplicates
# * **arrays_overlap(a1, a2)** - Collection function: returns true if the arrays contain any common non-null element
# * **arrays_zip()** - Collection function: Returns a merged array of structs in which the N-th struct contains all N-th values of input arrays
#
# ## Explode
# Two answer my secondf question, it would be easier for me having all words in in column instead of spread across many lists. To turn array elements into rows, I need to apply the `explode()` function. As the name of the function indicates, this can heavily increase the number of rows and the values of all remaining columns get duplicated.
# +
from pyspark.sql.functions import explode
explodedDF = cleanedWordListDF.select(
"StockCode",
explode("word_list").alias("words")
)
explodedDF.orderBy("StockCode").show(20)
# -
# The anser to my second question is simply a count of rows per word sorted in descending order.
# +
from pyspark.sql.functions import desc, count, lit
explodedDF\
.groupBy("words")\
.agg(count(lit(1)).alias("word_count"))\
.orderBy(desc("word_count"))\
.show(10)
# -
# Pink stocks seems to be quite popular.
#
# ## Maps
# For handling data in key:value structure, Spark provides another complex datatype: *maps*.
#
# My testdata does not provide key:value structured data. So first, I will transform my existing data into maps and second, I can investigate, how to handle key:value source data as an input to my ETL dataprocessing.
#
# ### Creating Maps
# +
dfFlight = spark.read\
.option("inferSchema", "true")\
.option("header", "true")\
.csv("./data/flight-data/2015-summary.csv")
from pyspark.sql.functions import lit, struct, array, col
from pyspark.sql.types import StringType
arrDF = dfFlight.select(
array(
lit("destination"),
lit("origin"),
lit("count")
).alias("key"),
array(
"DEST_COUNTRY_NAME",
"ORIGIN_COUNTRY_NAME",
col("count").cast(StringType())
).alias("value")
)
arrDF.show(10, truncate=False)
# +
from pyspark.sql.functions import map_from_arrays
mapDF = arrDF.select(
map_from_arrays("key", "value").alias("data_map")
)
mapDF.show(10, truncate=False)
# -
mapDF.select(col("data_map")["destination"]).show(10)
mapDF.select(col("data_map")["origin"]).show(10)
# +
from pyspark.sql.functions import map_keys
mapDF.select(
map_keys("data_map")
).show(10, truncate=False)
# +
from pyspark.sql.functions import map_values
mapDF.select(
map_values("data_map")
).show(10, truncate=False)
# -
# The data I've processed so far looks at least semi-structured because the keys and values all appear in identical order. So there is still an implicit schema because all rows match to the same pattern:
#
# destination -> descVal, origin -> origValue, count -> cntVal
#
# What would happen, if rows have keys and values in different order? Because my testdata does not provide examples for this, I create a DataFrame manuall with synthetic data in multiple orders.
# +
unstructuredDF = spark.createDataFrame(
[
(["destination", "origin", "count"], ["United States", "Germany", "10"],),
(["count", "origin", "destination"], ["25", "France", "Spain"],),
(["count", "destination", "origin"], ["75", "Italy", "Spain"],)
],
["key", "value"]
)
unstructuredDF.show(truncate=False)
# +
mapDF2 = unstructuredDF.select(
map_from_arrays("key", "value").alias("data_map")
)
mapDF2.show(truncate=False)
# -
mapDF2.select(col("data_map")["origin"]).show(10)
# Luckily the odering doesn't matter because I reference the values by keys and not by positions. Maps are more like dictionaries than lists or arrays.
#
# ### Turning Maps into DataFrames
#
# So with my self-created map I can now investigate how to handle such data as input for my ETL process which finally will write data in tabular form into a file or database table. So as intermediate step, I will have to align more or less ordered *key:value* pairs with the schema of a `DataFrame`.
#
# Can the `explode()` function help again?
mapDF2.select(explode("data_map")).show(10)
# Well, yes and no. Yes, `explode()` accepts both arrays as well as maps as an argument. No, because now I've lost the information, which three rows belong together. Additionally my intention was to gain three columns, one for each key value, and not just two. For maps referencing by key is always a better approach than referencing by position.
mapDF2.select(
col("data_map")["destination"].alias("destination"),
col("data_map")["origin"].alias("origin"),
col("data_map")["count"].alias("count")
).show(10)
# So with Spark handling nearly unstructured data records of key:value pairs in different orders is not a big problem.
#
# Pyspark module <a href=https://spark.apache.org/docs/latest/api/python/pyspark.sql.html#module-pyspark.sql.functions>pyspark.sql.functions</a> provides further map related functions, which I also just list here for later reference:
#
# * **map_concat()** - Returns the union of all the given maps
# * **map_from_entries()** - Collection function: Returns a map created from the given array of entries
#
# ### Turning Arrays or Maps into JSON
# A nice Spark feature is the `to_json()` function which converts StructType, ArrayType or MapType data into JSON. This can be relevant for me if I have to call a REST API which expects JSON documents as paylod.
# +
from pyspark.sql.functions import to_json
mapDF2.select(to_json("data_map")).show(10, truncate=False)
# -
mapDF2.select(to_json("data_map")).printSchema()
# ## Processing Semi-structured JSON data
# As I've learned on day 3, reading data from JSON file and transforming it into a DataFreame is quite simple. Just for repetition:
# +
jsonDF = spark.read\
.option("inferSchema", "true")\
.format("json")\
.load("./data/flight-data/2015-summary.json")\
jsonDF.printSchema()
# -
jsonDF.show(10)
# But what have I to to in case of having tabular data where only one column contains JSON strings? To check this out first I create same testdata.
df = spark.createDataFrame(
[
(123, "DUS", '{"destinations" : ["FRA", "MUC", "TXL"], "airlines" : ["LH", "EW", "RY"]}'),
(456, "FRA", '{"destinations" : ["CDG", "MUC", "JFK"], "airlines" : ["AF", "LH", "DL"]}'),
(789, "MUC", '{"destinations" : ["FRA", "ZUC", "DUS"], "airlines" : ["EW", "LH", "EW"]}')
],
["key", "airport", "dest"]
)
df.show(truncate=False)
# ### Navigation along JSON Paths
# Each row in the "dest" column contains a valid JSON document. Now I can use the `get_json_object()` function to access the values inside of the JSON documents by specifiying the path from the root element (represented by `$`) down the nesting hierarchie to the specific JSON obect I want to extract.
#
# path: `$.key_level1.key_level_2....key_level_n`
#
# Since in my DataFrame the objects "destinations", and "airlines" have value lists, I have to specify the list index to get one singular value per row.
# +
from pyspark.sql.functions import get_json_object
df.select(
"key",
"airport",
get_json_object("dest", '$.destinations[2]').alias("destination"),
get_json_object("dest", '$.airlines[1]').alias("airline"),
).show(truncate=False)
# -
# If I omitt the list index, I'll get the entire value list in my result DataFrame.
df.select(
"key",
"airport",
get_json_object("dest", '$.destinations').alias("destination"),
get_json_object("dest", '$.airlines').alias("airline"),
).show(truncate=False)
# There is a similar function `json_tuple()` but I'm not sure if it provides any benefits to me, because:
# 1. I cannot use it if the JSON document has more than one level of nesting, and
# 1. I cannot refer to single list elements
from pyspark.sql.functions import json_tuple
df.select("key",
"airport",
json_tuple("dest", "destinations", "airlines").alias("destination", "airline"),
).show(truncate=False)
# ### Turning JSON to Map based on Schema
# Finally, like I can read from JSON files using an explicit schema definition, I can also apply `from_json()` on DataFrame columns containing JSON by using a schema. Depending on the schema definition `from_json()` will return StructType, ArrayType or MapType. Actually I could perform a conversion round-trip from StructType, ArrayType or MapType -> `to_json()` -> {Json} -> `from_json()` -> StructType, ArrayType or MapType.
#
# I convert the Json.
# +
from pyspark.sql.types import *
from pyspark.sql.functions import from_json
jsonSchema = MapType(
StringType(),
ArrayType(StringType(), True),
True
)
mappedDF = df.select("key",
"airport",
from_json("dest", jsonSchema).alias("json_data")
)
mappedDF.show(truncate=False)
# -
# Now I can navigate on the Map structure to extract single values similar to navigating the JSON path using `get_json_object()`, e.g. grabbing the third element of the destinations lists.
mappedDF.select(
"key",
"airport",
col("json_data")["destinations"][2]
).show()
# The question is: what is the benefit of taking these extra effort, defining a schema and converting JSON to Map? In my opinion this leads to cleaner code and a better design, because:
# 1. now the JSON structure, a mexpecting is explicitly documented in the code by the schema instead of implicitly assumed
# 1. the Map structure is a unifying abstraction of any key:value data, regardles of the source format, e.g. CSV file, JSON documents or key-value database tables
| day-008_complex_types_analysing_unstructured_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# YouTube per-CPU TGID residency analysis
# =======================
# This is a run of experiments/run_youtube.py with the cgroups module enabled.
# This notebook parses and plots the trace.html
# +
# #!/usr/bin/env python
# %pylab inline
import trappy
from trace import Trace
import itertools
import logging
import pandas as pd
import numpy as np
import os
from conf import LisaLogging
LisaLogging.setup()
logging.info('#### Setup FTrace')
# path_to_html = "/home/joelaf/repo/lisa-aosp/external/lisa/results/UiBench_default/trace.html"
# path_to_html = "/home/joelaf/repo/lisa-aosp/external/lisa/ipynb/residency/trace-residency.html"
path_to_html = "/home/joelaf/repo/lisa-aosp/external/lisa/results/YouTube_cgroups/trace.html"
# +
def plot_cgroups(df):
# Bug in matplot lib causes plotting issues when residency is < 1
df = df.apply(lambda x: x*10)
plt.style.use('ggplot')
colors = plt.rcParams['axes.color_cycle']
fig, axes = plt.subplots(nrows=5, ncols=2, figsize=(12,30))
for ax, col in zip(axes.flat, df.columns):
ax.pie(df[col], labels=df.index, autopct='%.2f', colors=colors)
ax.set(ylabel='', title=col, aspect='equal')
axes[0, 0].legend(bbox_to_anchor=(0, 0.5))
# fig.savefig('your_file.png') # Or whichever format you'd like
plt.show()
tr = Trace(None, path_to_html,
cgroup_info = {
'cgroups': ['foreground', 'background', 'system-background', 'top-app', 'rt'],
'controller_ids': { 4: 'cpuset', 2: 'schedtune' }
},
events=[ 'sched_switch', 'cgroup_attach_task_devlib', 'cgroup_attach_task', 'sched_process_fork' ],
normalize_time=False)
# -
# Total amount of time spent per TGID
# ===========================
cdf = tr.data_frame.cpu_residencies('tgid_comm', 'sched_switch_cgroup')
# Due to the noise in the pie plots due to large number of TGIDs, drop any that
# don't meet the bar (run on CPUs for a total of atleast 0.7s)
cdf = cdf[~(cdf < 0.7).all(1)]
cdf
# Plot per-CPU breakdown without considering idle time
# ------------------------------------------------------------
# +
# Drop NaN
ncdf = cdf[pd.isnull(cdf.index) != True]
plot_cgroups(ncdf)
# -
# Plot per-CPU breakdown WITH considering idle time (yellow slice)
# ------------------------------------------------------------
# %pylab inline
plot_cgroups(cdf)
| ipynb/residency/task_residencies_youtube-tgid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Analisi dati delle zone arancioni degli ultimi 14 giorni disponibili
# I dati elaborati sono quelli presenti nel file * dpc-covid19-ita-regioni.json * nella directory * dati-json *.
# **Elaborazione dei dati:**
# * estrazione del dataset
# * trovo la data massima contenuta nel dataset
# * calcolo la data di riferimento per trovare i dati dei 14 giorni precedenti
# * estraggo dei sotto dataset con i dati delle regioni rilevati per l'analisi in questione relativi al periodo temporale in esame
# +
import pandas as pd
pd.plotting.register_matplotlib_converters()
dataset = pd.read_json('../dati-json/dpc-covid19-ita-regioni.json')
dataset['data'] = pd.to_datetime(dataset['data'])
dataset.set_index('data', inplace=True)
max_date = dataset.index.max()
# +
from datetime import timedelta
ref_date = max_date - timedelta(days=14)
# +
regioni = ['Abruzzo', 'Campania', 'P.A. Bolzano', 'Toscana', 'Valle d\'Aosta']
data_filter = (dataset.index > ref_date) & (dataset.denominazione_regione.isin(regioni))
filtered_set = dataset[data_filter].loc[:, ['denominazione_regione', 'ricoverati_con_sintomi', 'terapia_intensiva', 'totale_ospedalizzati', 'isolamento_domiciliare', 'totale_positivi', 'variazione_totale_positivi', 'nuovi_positivi', 'dimessi_guariti', 'deceduti', 'casi_da_sospetto_diagnostico', 'casi_da_screening', 'totale_casi', 'tamponi', 'casi_testati']]
# -
subsets = []
for r in regioni:
subsets.append((r, filtered_set[filtered_set.denominazione_regione == r].sort_index()))
# Configurazione preferenze di base per i grafici.
# +
import matplotlib.pyplot as plt
plt.rc('xtick', labelsize=15)
plt.rc('ytick', labelsize=15)
default_figsize = (24, 6)
default_titlesize = 20
default_padding = 8
# -
# ## Andamento ricoverati con sintomi
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['ricoverati_con_sintomi'].values, label=r)
plt.title('Ricoverati con sintomi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento occupazione delle terapie intensive
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['terapia_intensiva'].values, label=r)
plt.title('Terapia intensiva', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento totale dei pazienti ospedalizzati
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['totale_ospedalizzati'].values, label=r)
plt.title('Totale ospedalizzati', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento persone in isolamento domiciliare
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['isolamento_domiciliare'].values, label=r)
plt.title('Isolamento domiciliare', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento totale attualmente positivi
# Calcolati con la formula: $ totale\_ospedalizzati + isolamento\_domiciliare $
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['totale_positivi'].values, label=r)
plt.title('Totale positivi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento della variazione totale positivi
# Calcolati con la formula: $ totale\_positivi\ giorno\ corrente - totale\_positivi\ giorno\ precedente $
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['variazione_totale_positivi'].values, label=r)
plt.title('Variazione totale positivi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento nuovi positivi
# Calcolati con la formula: $ totale\_casi\ giorno\ corrente - totale\_casi\ giorno\ precedente $
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['nuovi_positivi'].values, label=r)
plt.title('Nuovi positivi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento dimessi guariti
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['dimessi_guariti'].values, label=r)
plt.title('Dimessi guariti', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento deceduti
# Il dato è cumulativo.
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['deceduti'].values, label=r)
plt.title('Deceduti', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Casi da sospetto diagnostico
#
# Casi positivi al tampone emersi da attività clinica.
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['casi_da_sospetto_diagnostico'].values, label=r)
plt.title('Casi da sospesso diangostico', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Casi da screening
#
# Casi positivi emersi da indagini e test, pianificati a livello nazionale o regionale.
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['casi_da_screening'].values, label=r)
plt.title('Casi da screening', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento totale casi
#
# Totale casi positivi.
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['totale_casi'].values, label=r)
plt.title('Totale casi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento tamponi
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['tamponi'].values, label=r)
plt.title('Tamponi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Andamento casi testati
plt.figure(figsize=default_figsize)
for (r, s) in subsets:
plt.plot(s.index, s['casi_testati'].values, label=r)
plt.title('Casi testati', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Analisi variazione dei dati forniti in modo cumulativo
diff_col = ['ricoverati_con_sintomi', 'terapia_intensiva', 'totale_ospedalizzati', 'isolamento_domiciliare', 'dimessi_guariti', 'deceduti', 'casi_da_sospetto_diagnostico', 'casi_da_screening', 'tamponi', 'casi_testati']
diff_subsets = []
for (r, s) in subsets:
diff_subsets.append((r, s.loc[:, diff_col].diff()))
# ## Variazione ricoverati con sintomi
# Calcolata con la formula: $ ricoverati\_con\_sintomi\ giorno\ X - ricoverati\_con\_sintomi\ giorno\ X-1 $
plt.figure(figsize=default_figsize)
for (r, s) in diff_subsets:
plt.plot(s.index, s['ricoverati_con_sintomi'].values, label=r)
plt.title('Variazione ricoverati con sintomi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Variazione terapie intensive
# Calcolata con la formula: $ terapia\_intensiva\ giorno\ X - terapia\_intensiva\ giorno\ X-1 $
plt.figure(figsize=default_figsize)
for (r, s) in diff_subsets:
plt.plot(s.index, s['terapia_intensiva'].values, label=r)
plt.title('Variazione terapia intensiva', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Variazione totale ospedalizzati
# Calcolata con la formula: $ totale\_ospedalizzati\ giorno\ X - totale\_ospedalizzati\ giorno\ X-1 $
plt.figure(figsize=default_figsize)
for (r, s) in diff_subsets:
plt.plot(s.index, s['totale_ospedalizzati'].values, label=r)
plt.title('Variazione totale ospedalizzati', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Variazione deceduti
# Calcolata con la formula: $ deceduti\ giorno\ X - deceduti\ giorno\ X-1 $
plt.figure(figsize=default_figsize)
for (r, s) in diff_subsets:
plt.plot(s.index, s['deceduti'].values, label=r)
plt.title('Variazione deceduti', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
# ## Variazione tamponi
# Calcolata con la formula: $ tamponi\ giorno\ X - tamponi\ giorno\ X-1 $
plt.figure(figsize=default_figsize)
for (r, s) in diff_subsets:
plt.plot(s.index, s['tamponi'].values, label=r)
plt.title('Variazione tamponi', fontsize=default_titlesize, pad=default_padding)
plt.legend()
plt.show()
| notebook/analisi_zone_arancioni_14_giorni.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import csv
csv_file = r"C:\Users\skm72\Desktop\git\Homework\pandas-challenge\HeroesOfPymoli\Resources\purchase_data.csv"
game_df=pd.read_csv(csv_file)
game_df.head()
# -
#Finding the total unique "SN"
total_players = len(game_df["SN"].value_counts())
total_players_df = pd.DataFrame([{"Total Players" : total_players}])
total_players_df
# +
#Finding the amount of unique items the average price and total amount of purchases and revenue
unique_items = len(game_df["Item Name"].unique())
average_price = game_df["Price"].mean()
total_purchases = game_df["Purchase ID"].count()
total_revenue = game_df["Price"].sum()
#Creating a dataframe for the values
purchasing_analysis_total = pd.DataFrame([{"Number of Unique Items" : unique_items,
"Average Price" : average_price,
"Number of Purchases" : total_purchases,
"Total Revenue" : total_revenue}])
#Formatting the average price and total revenue to look cleaner
purchasing_analysis_total["Average Price"] = purchasing_analysis_total["Average Price"].map("${:,.2f}".format)
purchasing_analysis_total["Total Revenue"] = purchasing_analysis_total["Total Revenue"].map("${:,.2f}".format)
purchasing_analysis_total
# +
#Finding the gender counts based on the unique "SN"
gender_types = game_df.groupby("Gender")
total_gender_counts = gender_types.nunique()["SN"]
gender_percentage = (total_gender_counts / total_players) * 100
#Creating a dataframe for the values
gender_types_df = pd.DataFrame({"Percentage of Players" : gender_percentage,
"Total Count" : total_gender_counts})
#Formatting the dataframe to make it look cleaner
gender_types_df.index.name = None
gender_types_df = gender_types_df.sort_values(["Total Count"], ascending = False)
gender_types_df["Percentage of Players"] = gender_types_df["Percentage of Players"].map("{:,.2f}%".format)
gender_types_df
# +
#Finding the average of purchase based on the gender
gender_count = gender_types["Purchase ID"].count()
average_purchase_gender = gender_types["Price"].mean()
average_total_gender = gender_types["Price"].sum()
average_person = average_total_gender/total_gender_counts
#Creating a dataframe for the values
purchasing_analysis_gender = pd.DataFrame({"Purchase Count" : gender_count,
"Average Purchase Price" : average_purchase_gender,
"Average Purchase Value" : average_total_gender,
"Avg Total Purchase per Person" : average_person})
#Formatting the dataframe to make it look cleaner
purchasing_analysis_gender.index.name = "Gender"
purchasing_analysis_gender["Average Purchase Price"] = purchasing_analysis_gender["Average Purchase Price"].map("${:,.2f}".format)
purchasing_analysis_gender["Average Purchase Value"] = purchasing_analysis_gender["Average Purchase Value"].map("${:,.2f}".format)
purchasing_analysis_gender["Avg Total Purchase per Person"] = purchasing_analysis_gender["Avg Total Purchase per Person"].map("${:,.2f}".format)
purchasing_analysis_gender
# +
#Creating the bins and the group names
bins = [0, 9.9, 14.9, 19.9, 24.9, 29.9, 34.9, 39.9, 150]
group_names = ["<10", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", "40+"]
#Putting the data into their bins based on age
game_df["Age Group"] = pd.cut(game_df["Age"], bins, labels = group_names, include_lowest = True)
#Finding the total number in each age group based on the unique "SN"
age_group = game_df.groupby(["Age Group"])
age_count = age_group["SN"].nunique()
age_percentage = (age_count/total_players) * 100
#Creating a dataframe for the values
age_group_df = pd.DataFrame({"Total Count": age_count, "Percentage of Players": age_percentage})
#Formating the dataframe to make it look cleaner
age_group_df.index.name = None
age_group_df["Percentage of Players"] = age_group_df["Percentage of Players"].map("{:,.2f}%".format)
age_group_df
# -
#Finding the average of purchase based on the gender
age_count_total = age_group["Purchase ID"].count()
| HeroesOfPymoli/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recording the macro
# 
#
# delvopment tool -> recording macro
# **in case of making the name of macro,
# the function name which is like inner function,number will not be acceptable
#
# # making the button for implementing the macro
# it would be good to make the button for macro to the location which is not same row or column as the raw data.
# also in case of saving the file with macro, the type of file should be "xlsm" not "xlsx".
#
| VBA/Day02.making the macro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import altair as alt
assays_long = pd.read_csv('results.csv')
assays_long['outcome'].unique()
# # Proportion stacked bar plot:
# +
###Count up conditions using groupby:
numActive = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: (grouped_df['outcome'].str.contains('Active').any())).values.sum()
numInactive = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: (grouped_df['outcome'].str.contains('Inactive').any())).values.sum()
numBoth = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: ((grouped_df['outcome'].str.contains('Inactive').any()) &
(grouped_df['outcome'].str.contains('Active').any()))).values.sum()
numNone = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: (~(grouped_df['outcome'].str.contains('Inactive').any()) &
~(grouped_df['outcome'].str.contains('Active').any()))).values.sum()
# -
###Or count up conditions the boring way:
y=0
n=0
b=0
neither=0
instance_ids = assays_long.drop_duplicates(['lig_chemblid', 'pref_name'])['lig_chemblid']
pref_names = assays_long.drop_duplicates(['lig_chemblid', 'pref_name'])['pref_name']
for i, name in zip(instance_ids, pref_names):
mask = (assays_long['pref_name']==name) & (assays_long['lig_chemblid']==i)
group = assays_long[mask]
outcomes = list(group['outcome'])
if ('Active' in outcomes) and ('Inactive' in outcomes):
b+=1
elif ('Active' in outcomes):
y+=1
elif ('Inactive' in outcomes):
n+=1
else:
neither+=1
assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: (~(grouped_df['outcome'].str.contains('Inactive').any()) &
~(grouped_df['outcome'].str.contains('Active').any()))).values.sum()
#Verify both ways are the same:
print(f'Active: {y}')
print(f'Inactive: {n}')
print(f'Both: {b}')
print(f'None: {neither}')
#Both ways are the same:
print(f'Active: {numActive-numBoth}')
print(f'Inactive: {numInactive-numBoth}')
print(f'Both: {numBoth}')
print(f'None: {numNone}')
# +
df = pd.DataFrame(columns=['results','activity', 'number'])
df.loc[0]=['Results', 'Active', numActive]
df.loc[1]=['Results', 'Inactive', numInactive]
df.loc[2]=['Results', 'Both', numBoth]
###Remember - inconclusive or unspecified must be added to all the predictions that had NO records.
#without the 'unknown' records:
#df.loc[3]=['Results', 'Unknown', numNone ]
#including all absent records:
number_of_absent_records = 10000 - sum([df.iloc[0]['number'], df.iloc[1]['number']])
df.loc[3] = ['Results', 'Unknown', number_of_absent_records]
df['proportion'] = df['number']/df['number'].sum()
df
# +
chart_one = alt.Chart(df).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q', title='All assays'),#,sort='descending'),
y=alt.Y('results', title=''),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
)#.add_selection(
# selector
#)
chart_two = alt.Chart(df.iloc[:3]).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q', title='Assays with activity evidence'),#,sort='descending'),
y=alt.Y('results', title=''),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
)
#.add_selection(
# selector
#)
chart_one&chart_two
# +
####Interactive version:
# brush = alt.selection(type='interval', encodings=['x'])
# base = alt.Chart(df).transform_joinaggregate(
# Total='sum(number)'
# ).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
# #x='number',
# x='PercentOfTotal:Q',
# y='results',
# color='activity')
# upper = base.encode(
# alt.X('PercentOfTotal:Q', scale=alt.Scale(domain=brush)))
# lower = base.add_selection(brush)
# lower&upper
# -
# # Target type barplot:
#
# +
out = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(
lambda grouped_df: (~(grouped_df['outcome'].str.contains('Inactive').any()) &
(grouped_df['outcome'].str.contains('Active').any())))
positive_targets=[i[1] for i, v in zip(out.index, out.values) if v==True]
all_pos_targets = pd.DataFrame(out[out==True]).reset_index().drop([0],axis=1)
pos_targets_gb= all_pos_targets.groupby(['pref_name']).count().reset_index().sort_values(by='lig_chemblid', ascending=False)
pos_targets_gb.columns=['pref_name', 'count']
# -
# # All together now:
# +
chart_one = alt.Chart(df).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q'),#,sort='descending'),
y=alt.Y('results', title='',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
).properties(height=30, width=550)
#.add_selection(
# selector
#).properties(height=30, width=550)
chart_two = alt.Chart(df.iloc[:3]).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q', title='Hi'),#,sort='descending'),
y=alt.Y('results', title='',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
).properties(height=30, width=550)
#.add_selection(
# selector
#).properties(height=30, width=550)
chart = alt.Chart(all_pos_targets).mark_bar()
chart = chart.encode(
x = alt.X('pref_name', sort='-y', title='Target preferred name'),
y = alt.Y('count()'),
).properties(height=200,width=700)
(chart_one&chart_two&chart
).configure_axisBottom(labelAngle=-35).configure_header(
titleFontSize=40,
labelFontSize=40
).configure_title()
# +
def parse_group(group):
inactive = group['outcome'].str.contains('Inactive').any()
active = group['outcome'].str.contains('Active').any()
if inactive and active:
return 'Both'
elif active and ~inactive:
return 'Active'
elif inactive and ~active:
return 'Inactive'
else:
return False
out = assays_long.groupby(['lig_chemblid', 'pref_name']).apply(parse_group)
out_df= pd.DataFrame(out[out!=False]).reset_index().drop('lig_chemblid',axis=1)
out_df.columns=['pref_name', 'activity']
# -
out_df
# +
chart_one = alt.Chart(df).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q', title='Proportion'),#,sort='descending'),
y=alt.Y('results', title='All records',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
).properties(height=30, width=750)
#.add_selection(
# selector
#).properties(height=30, width=550)
chart_two = alt.Chart(df.iloc[:3]).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
x=alt.X('PercentOfTotal:Q', title='Proportion'),#,sort='descending'),
y=alt.Y('results', title='Records with evidence',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title='Activity type'),
order=alt.Order('activity', sort='ascending')
).properties(height=30, width=750)
#.add_selection(
# selector
#).properties(height=30, width=550)
ch = alt.Chart(out_df).mark_bar()
ch = ch.encode(
x = alt.X('pref_name', sort='-y', title='Target preferred name'),
y = alt.Y('count()'),
color=alt.Color('activity'),
order=alt.Order('activity', sort='ascending'),
).properties(height=200,width=750)
# ch = ch.transform_aggregate(
# count='count()',
# groupby=['pref_name']
# ).transform_filter(
# 'datum.count >= 5'
# ).mark_bar().encode(
# alt.X('pref_name:N', sort='-y', title='Target preferred name'),
# alt.Y('count:Q'),
# color=alt.Color('activity'),
# order=alt.Order('activity', sort='ascending'),
# ).properties(height=200,width=750)
(chart_one&chart_two&ch
).configure_axisBottom(labelAngle=-35).configure_header(
titleFontSize=40,
labelFontSize=40
).configure_axisY(
titleAngle=0,
titleAlign="left",
titleY=-10,
titleX=-10,
titleFontSize=16,
)
# +
chart_one = alt.Chart(df).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
y=alt.Y('PercentOfTotal:Q', title='All assays'),#,sort='descending'),
x=alt.X('results', title='',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title=''),
order=alt.Order('activity', sort='ascending')
).properties(height=500, width=30)
chart_two = alt.Chart(df.iloc[:3]).transform_joinaggregate(
Total='sum(number)'
).transform_calculate(PercentOfTotal="datum.number / datum.Total").mark_bar().encode(
y=alt.Y('PercentOfTotal:Q', title='Existing evidence',),#,sort='descending'),
x=alt.X('results', title='',axis=alt.Axis(labels=False)),
color=alt.Color('activity', title=''),
order=alt.Order('activity', sort='ascending')
).properties(height=500, width=30)
ch = alt.Chart(out_df).mark_bar()
ch = ch.encode(
y = alt.Y('pref_name', sort='-x', title='Target occurrence'),
x = alt.X('count()', title='Count'),
color=alt.Color('activity'),
order=alt.Order('activity', sort='ascending'),
).properties(height=500,width=300)
(chart_one|chart_two|ch
).configure_axisBottom(labelAngle=-35).configure_header(
titleFontSize=40,
labelFontSize=40
).configure_axisY(
titleAngle=0,
titleAlign="left",
titleY=-10,
titleX=-20,
titleFontSize=16,
).configure_legend(orient='bottom',labelFontSize=22)
#top right' is not one of ['none', 'left', 'right', 'top',
#'bottom', 'top-left', 'top-right', 'bottom-left', 'bottom-right']
# +
temp = out_df.groupby('pref_name').filter(lambda x : len(x)>2)
ch = alt.Chart(temp).mark_bar()
# ch = ch.encode(
# x = alt.X('pref_name', sort='-y', title='Target preferred name'),
# y = alt.Y('count()'),
# color=alt.Color('activity'),
# order=alt.Order('activity', sort='ascending'),
# )
ch = ch.encode(
y = alt.Y('pref_name', sort='-x', title='Target preferred name'),
x = alt.X('count()', title='Count'),
color=alt.Color('activity'),
order=alt.Order('activity', sort='ascending'),
).properties(height=500,width=300)
ch.properties(height=200,width=750).configure_axisY(
titleFontSize=13,
).configure_axisX(
titleFontSize=13
).configure_axisBottom(labelAngle=-35).configure_header(
titleFontSize=40,
labelFontSize=40
).configure_axisY(
titleAngle=0,
titleAlign="left",
titleY=-10,
titleX=-20,
titleFontSize=16,
).configure_legend(orient='bottom',labelFontSize=22)
# ch = ch.transform_aggregate(
# count='count()',
# groupby=['pref_name']
# ).transform_filter(
# 'datum.count >= 5'
# ).mark_bar().encode(
# alt.X('pref_name:N', sort='-y', title='Target preferred name'),
# alt.Y('count:Q'),
# color=alt.Color('activity'),
# order=alt.Order('activity', sort='ascending'),
# ).properties(height=200,width=750)
ch
# -
gb = out_df.groupby('pref_name')
out_df.groupby('pref_name').filter(lambda x : len(x)>3)
gb.count().reset_index()[(gb.count()>2)['activity']]
(gb.count()>2)['activity']
out_df
| 4_pubchem_validation/figure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Manage data processing
# Architecture of the code :
# - sbttimport module : extract data from srt files to pandas dataframe
# - dataprocess module : processing data for NLP workflow
# - insightviz : visualize the data
import main
orch = main.Orchestrator(lang="eng", nb=5)
orch.run()
orch.show_topics()
| chernobylapp/Manager.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rubybea/Linear-Algebra_CHE_2nd-Sem-2021-2022/blob/main/Python_Fundamental.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="tEkV_WhI_c79"
# # Welcome to Python Fundamentals
# In this module, we are going to establish or review our skills in Python programming. In this notebook we are going to cover:
# * Variables and Data Types
# * Operations
# * Input and Output Operations
# * Logic Control
# * Iterables
# * Functions
# + [markdown] id="h8TFvuwnchfe"
# ## Variable and Data Types
#
#
#
#
# + id="MzFQdtW1crCk"
x = 1
a,b = 0, -1
# + colab={"base_uri": "https://localhost:8080/"} id="gQCIZ-_Kdaxa" outputId="671e9b49-f93c-4d0d-b1d8-742959b485c2"
type(x)
# + colab={"base_uri": "https://localhost:8080/"} id="OwRRM3_Id82v" outputId="ad5270c0-f312-4386-f477-8f837151be7a"
y = 1,0
type(y)
# + colab={"base_uri": "https://localhost:8080/"} id="1CQR7sKfeFh8" outputId="a8671cbd-6ca8-4660-93e6-a4d2c470863f"
x = float(x)
type(x)
# + colab={"base_uri": "https://localhost:8080/"} id="zvL7vk7rfATz" outputId="e5873ac0-f97f-4930-9caa-1cd9c3436848"
s,t,u ="0", "1", "one"
type(s)
# + colab={"base_uri": "https://localhost:8080/"} id="4enpr9zCfnLJ" outputId="72b30154-6cd4-477c-8ec0-0270210e5aaa"
s_int = int(s)
s_int
# + [markdown] id="psZwl-QWf7AP"
# ##Operations
#
# + [markdown] id="c3GffArWgO44"
# ### Arithmetic
#
# + id="5KWTOMFGgSXC"
a,b,c,d = 2.0, -0.5, 0, -32
# + colab={"base_uri": "https://localhost:8080/"} id="3uWMeZtwgk62" outputId="1bfb4da9-c444-40ca-c1c3-14c96f710f4c"
### Addition
S = a+b
S
# + colab={"base_uri": "https://localhost:8080/"} id="yJtxjUJugk90" outputId="5c4885c4-6dcd-444c-c9ec-305ff084c8ee"
### Subtraction
D = b-d
D
# + colab={"base_uri": "https://localhost:8080/"} id="L0-bOVV5g29D" outputId="dadc1418-848a-4f62-8cc6-f54d4b6671d2"
### Multiplication
P = a*d
P
# + colab={"base_uri": "https://localhost:8080/"} id="EtDWv78Cg3Im" outputId="638f3386-844e-4048-e2fe-4e002f869b62"
### Division
Q = c/d
Q
# + colab={"base_uri": "https://localhost:8080/"} id="9GrW9cHvhQZB" outputId="4e25317d-abc7-4565-f03c-d950fff11b23"
### Floor Division
Fq = a//b
Fq
# + colab={"base_uri": "https://localhost:8080/"} id="NK26ID5FhQi-" outputId="b371c897-e4a3-45e8-b0eb-d5208026cdfa"
### Exponentiation
E = a**b
E
# + colab={"base_uri": "https://localhost:8080/"} id="NmFRqsSthRBq" outputId="f761f9e8-89d7-4656-c330-b1a42a90122f"
### Modulo
mod = d%a
mod
# + [markdown] id="3q1pDGr1jezz"
# ## Assingment Operations
# + id="TWpKnHWIjjX0"
G, H, J, K = 0, 100, 2, 2
# + colab={"base_uri": "https://localhost:8080/"} id="tSRdlJkgjsJk" outputId="728e52e5-e4dc-45a7-eff9-0f5b694b7919"
G += a
G
# + colab={"base_uri": "https://localhost:8080/"} id="Un-rcl2zjsR9" outputId="8db039ab-074a-4c2e-c7cf-2aa7e54f3a70"
H -= d
H
# + colab={"base_uri": "https://localhost:8080/"} id="HDJ36I1tjsaL" outputId="98ddd976-712e-4874-8b8d-d3bf73fdcd73"
J *= 2
J
# + colab={"base_uri": "https://localhost:8080/"} id="BN2x0M7fjsbp" outputId="48a2e0b1-8f67-4c5c-fcb6-946b3637e577"
K **= 3
K
# + [markdown] id="Zz9zV6jIkpnw"
# ## Comparators
# + id="OZLFxwf1kwnp"
res_1, res_2, res_3 = 1, 2.0, "1"
true_val = 1.0
# + colab={"base_uri": "https://localhost:8080/"} id="bMk4w6bmlC3D" outputId="de495ebe-39e6-4d85-8fee-a321c0bfa769"
## Equality
res_1 == true_val
# + colab={"base_uri": "https://localhost:8080/"} id="idHHpffPlDBa" outputId="7b24df0f-180d-4367-fdef-a6bb512ab14e"
## Non-equality
res_2 != true_val
# + colab={"base_uri": "https://localhost:8080/"} id="MhtV9KsflDDI" outputId="20f019e0-bcf4-4569-b836-675ca4609d9c"
## Inequality
t1 = res_1 > res_2
t2 = res_1 < res_2/2
t3 = res_1 >= res_2/2
t4 = res_1 <= res_2
t1
# + [markdown] id="bCR1wAz0wiqL"
#
# + [markdown] id="Qdvm7RwDmGDn"
# ## Logical
# + colab={"base_uri": "https://localhost:8080/"} id="n8K-ubjVlDKr" outputId="8bf1ad9e-950d-406a-dd69-e7b68666d69c"
res_1 == true_val
# + colab={"base_uri": "https://localhost:8080/"} outputId="c9e6967b-9b66-413f-dd81-75a32c569cbb" id="hnfF4--hmP4o"
res_1 is true_val
# + colab={"base_uri": "https://localhost:8080/"} outputId="3397b52e-31c7-48d5-f74b-02508f770403" id="08AYnNokmX52"
res_1 is not true_val
# + colab={"base_uri": "https://localhost:8080/"} id="OmkdwgQElDSP" outputId="3a9e24e3-ac0c-46c9-a480-37961febdff7"
p, q = True, False
conj = p and q
conj
# + colab={"base_uri": "https://localhost:8080/"} outputId="d5c1afcf-aa78-431d-c25f-1f9676294b47" id="nfYdX4bFmrob"
p, q = True, False
disj = p or q
disj
# + colab={"base_uri": "https://localhost:8080/"} outputId="5652df8a-88f7-495b-84d0-91b0e165e2b3" id="HViV4VC5m7hC"
p, q = True, False
nand = not(p and q)
nand
# + colab={"base_uri": "https://localhost:8080/"} outputId="ed0f20d4-eff5-437f-d136-7a68be8fad80" id="zcIUEdjmnR1j"
p, q = True, False
xor = (not p and q) or (p and not q)
xor
# + [markdown] id="r2xAqjHsnq4M"
# ## 1/0
# + colab={"base_uri": "https://localhost:8080/"} id="kNCINtmAnfZN" outputId="b9df13d0-c23b-4dae-8e1e-3f112ab5fb05"
print ("Hello World")
# + id="vle_289XxMwj"
cnt = 1
# + colab={"base_uri": "https://localhost:8080/"} id="BXz_MqzxnffS" outputId="1aa414ed-4421-49db-b1b7-3e15977ffd08"
string = "Hello World"
print(string, ", Current run count is:", cnt)
cnt +=1
# + colab={"base_uri": "https://localhost:8080/"} id="fUBOw3wGoaUh" outputId="0f4e8cf4-d71a-4730-f5ef-bf051ccf35c2"
print(f"{string}, Current count is {cnt}")
# + colab={"base_uri": "https://localhost:8080/"} id="zMzQv3vTozTg" outputId="84d0477c-a3e6-457a-f688-13852c4363bb"
sem_grade = 82.24356457461234
name = "cath"
print("Hello {}, your semestral grade is: {}".format(name, sem_grade))
# + colab={"base_uri": "https://localhost:8080/"} id="XB_gw9KPq970" outputId="03229b3b-5a72-4672-98e2-8200085c3d67"
w_pg, w_mg, w_fg = 0.3, 0.3, 0.4
print("The weights of your semestral grades are:\
\n\t{:.2%} for Prelims\
\n\t{:.2%} for Midterms, and\
\n\t{:.2%} for Finals, ".format(w_pg, w_mg, w_fg))
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="aUS3mjBYr6VC" outputId="c9bb5db2-6635-41fd-eca5-9ede4383e02d"
x = input("enter a number: ")
x
# + colab={"base_uri": "https://localhost:8080/"} id="aQOf29eODbQD" outputId="5f3a974b-265e-44d3-abb9-baacae371542"
name = input("<NAME>: ")
pg = input("Enter prelim grade: ")
mg = input("Enter midterm grade: ")
fg = input("Enter finals grade: ")
sem_grade = None
print("Hello {}, your semestral grade is: {}". format (name, sem_grade))
# + [markdown] id="jph46P8qysqR"
# # Looping Statements
# + [markdown] id="05QhoJmDGucA"
# ## While
# + colab={"base_uri": "https://localhost:8080/"} id="rSZTHMtiy2sJ" outputId="d7bb2ead-5e97-4923-8c43-32d45383a940"
## while loops
i, j = 0, 10
while(i<=j):
print(f"{i}\t|\t{j}")
i+=1
# + [markdown] id="2JOZbAVDyxIV"
# ## For
# + colab={"base_uri": "https://localhost:8080/"} id="eQ3MLC4qGvl9" outputId="1300fdb8-0b3c-4c89-e18a-d5ccef2e3891"
# for(int i=0; i<10; i++){
# printf(i)
# }
i=0
for i in range(11):
print(i)
# + colab={"base_uri": "https://localhost:8080/"} id="F9JrogbLHr6u" outputId="6fd53646-55ea-4b02-c799-78249bdcb6e4"
playlist = ["Crazier", "Bahay-Kubo", "Happier"]
print('Now Playing:\n')
for song in playlist:
print(song)
# + [markdown] id="Zybs--40IVZ-"
# # Flow Control
# + [markdown] id="f9ZQ4Ntu0C4D"
# ## Conditions Statemnents
# + colab={"base_uri": "https://localhost:8080/"} id="TLcNpJT-IXQ8" outputId="b7503ca9-e1ce-4acb-99df-13d5fb94ca88"
numeral1, numeral2 = 12, 12
if(numeral1 == numeral2):
print("Yey")
elif(numeral1>numeral2):
print("Hoho")
else:
print("AWW")
print("Hip hip")
# + [markdown] id="7Wv05jJwJmaM"
# ## Functions
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="q8fVoZ3fJn-Q" outputId="898fc4de-cab9-41be-a735-5ea7182c109b"
[ ] # void DeleteUser(int userid){
# delete(userid);
# }
def delete_user (userid):
print("Successfully deleted user: {}". format(userid))
def delete_all_users ():
print("Successfully deleted all users")
userid = 202011844
delete_user(202011844)
delete_all_users()
# + colab={"base_uri": "https://localhost:8080/"} id="Vc-U4TFtMKCU" outputId="0f07f306-f54b-4011-c2a7-11571bc246a2"
def add(addend1, addend2):
print("I know how to add addend1 and addend2")
return addend1 + addend2
def power_of_base2(exponent):
return 2**exponent
addend1 = 5
addend2 = 10
exponent = 5
#add(addend1, addend2)
power_of_base2(exponent)
# + [markdown] id="tTVBSDf8rkR7"
# ## Grade Calculator
# + [markdown] id="vzndaBMu2Vzo"
# Create a grade calculator that computes for the semestral grade of a course. Students could type their names, the name of the course, then their prelim, midterm, and final grade.
# The program should print the semestral grade in 2 decimal points and should display the following emojis depending on the situation:
# happy - when grade is greater thann 70.00
# laughing - wen grade is exactly 70.00
# sad - when grade is below 70.00
# ...
# happy, lol, sad - "\U0001F600", "\U0001F606", "\U0001F62D"
# + colab={"base_uri": "https://localhost:8080/"} id="4iMJmogNefXn" outputId="d01728cc-671d-4cd9-b630-60e8e7afeda3"
w_pg, w_mg, w_fg = 0.3, 0.3, 0.4
name = input("Enter your name: ")
course = input("Enter your course: ")
pg = float(input("Enter prelim grade: "))
mg = float(input("Enter midterm grade: "))
fg = float(input("Enter final grade: "))
sem_grade = (pg*w_pg)+(mg*w_mg)+(fg*w_fg)
print("Hello {} from {}, your semetral grade is: {}" .format(name, course, round(sem_grade, 2)))
if(sem_grade > 70.00):
print("\U0001f600")
elif(sem_grade == 70.00):
print("\U0001F606")
else:
print("\U0001F620")
| Python_Fundamental.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # lamda
def test(f, a, b):
print(f(a, b))
fun = lambda x,y:x**2+y
test(fun,2,3)
# # 程序作用域
def f(x):
a = 7
print(a + x)
a = 5
f(3)
print(a)
# +
# Error
# -
a=3
print(a ** b)#print NameError
# ### 布尔表达式
# +
x = None
if x:
print("x is true")
else:
print("x is false")
x = 0
if x:
print("x is true")
else:
print("x is false")
x = []
if x:
print("x is true")
else:
print("x is false")
x = {}
if x:
print("x is true")
else:
print("x is false")
x = {1}
if x:
print("x is true")
else:
print("x is false")
# -
# ## String
# 使用dir 列出所有str的内建函数 并查找up开头的函数
[nameWithUp for nameWithUp in dir(str) if nameWithUp.startswith('up') ]
SingASong = "The sun is rising. The sunlight is shinning on the wind"
SingASong.upper()
# ### upper split join find encode decode
SingASong.split(' ')
'[]'.join(SingASong.split(' '))
SingASong.find("The")
# encode
type("你好".encode('utf-8'))
"你好".encode('utf-8')
# encode list of python ,can refer to https://docs.python.org/2.4/lib/standard-encodings.html
("你好".encode('utf-8')).decode('utf_8') # utf-8 , utf_8 are both work here
# help(bytes.decode)
# #### 转义字符
# \ooo表示8进制, \xxx表示16进制
print('\101\t\x41');print('\141\t\x61')
print('\102\n\x41');print('\142\n\x62')
help(bytes.decode)
# ### 列表
# #### 列表解析
# [expression for expr1 in sequence1
# for expr2 in sequence2
# ...
# for exprn in sequencen
# if condition]
[x**2 for x in range(10) for y in range(10) if x**2<50]
[(x+1,y+1) for x in range(2) for y in range(2)]
# ##### 列表的浅拷贝
#浅拷贝只完成列表x的第一层元素的拷贝,第二层元素(这里是[3,4])没有拷贝,拷贝的操作会产生新的对象
x = [1,2,[3,4]]
y = x.copy()
y[0],y[2][0]=9,9
x,y #x[2][0]也发生了改变
x = [1,2,[3,4]]
y = x.copy()
y[0],y[2]=9,[9,4]#此处是重新指向新的对象
x,y
# ##### 列表深拷贝
import copy
# help(copy)
# help(copy.deepcopy)
y=copy.deepcopy(x)
y[0],y[2][0]=8,8
x,y
# ### 输入输出
#如何输入得到一个均为数值型的列表
# lst = eval(input('enter a list'))
lst = list(input('enter numbers'))
lst = list(eval(input('enter numbers')))
# help(list)
lst
#eval 会只能分析输入的数值类型, 'abc',33 分别会转成 字符串和数值
x,y = eval(input('enter two params'))
x,y
# print(eval('',22,['p','y','thon']))
# print(eval('124857'))
print(eval('ab'))
#如何将循环输出的数据放在同一行
for i in range(4):
print(i,i+1,i+2,sep=',',end=' ')
# #### 函数式编程
# ##### map 函数
#map 函数使用
lst=[11,22,33]
list(map(str,lst))#map 将lst每个元素作为参数传入str函数得到的返回值作为新的元素
#map 函数 对列表元素转化大写
lst = ['fade','away','show']
list(map(upper,lst))
list(map(lambda x:x.upper(),lst))
# ##### 函数式编程应用2000个5(55555...55555)除以84的余数
from functools import reduce;reduce(lambda x,y:(x*10+y)%84 ,([5])*2000 )
dir(__builtins__)
| basic/ch01.ipynb |
# ---
# title: "Iterate An Ifelse Over A List"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "Iterate an ifelse over a list in Python."
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Create some data
# +
word_list = ['Egypt', 'Watching', 'Eleanor']
vowels = ['A', 'E', 'I', 'O', 'U']
# -
# ## Create a for loop
# for each item in the word_list,
for word in word_list:
# if any word starts with e, where e is vowels,
if any([word.startswith(e) for e in vowels]):
# then print is valid,
print('Is valid')
# if not,
else:
# print invalid
print('Invalid')
| docs/python/basics/iterate_ifelse_over_list.ipynb |