code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5교시 집계 연산
#
# ### 목차
# * [1. 집계 함수](#1.-집계-함수)
# * [2. 그룹 함수](#2.-그룹-함수)
# * [참고자료](#참고자료)
#
# +
from pyspark.sql import *
from pyspark.sql.functions import *
from pyspark.sql.types import *
from IPython.display import display, display_pretty, clear_output, JSON
spark = (
SparkSession
.builder
.config("spark.sql.session.timeZone", "Asia/Seoul")
.getOrCreate()
)
# 노트북에서 테이블 형태로 데이터 프레임 출력을 위한 설정을 합니다
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # display enabled
spark.conf.set("spark.sql.repl.eagerEval.truncate", 100) # display output columns size
# -
""" 구매 이력 데이터 """
df = (
spark.read.format("csv")
.option("header", "true")
.option("inferSchema", "true")
.load("data/retail-data/all")
.coalesce(5)
)
df.cache()
df.createOrReplaceTempView("dfTable")
df.show(5, truncate=False)
df.count()
# ## 1. 집계 함수
# ### 1.1 로우 수 (count, countDistinct, approx_count_distinct)
df.printSchema()
# +
from pyspark.sql.functions import *
df.selectExpr("count(*)").show()
df.where("Description is null").selectExpr("count(1)").show() # 1,454
df.selectExpr("count(Description)").show() # 540,455 + 1,454 = 541,909
# -
from pyspark.sql.functions import *
# 명시적으로 컬럼을 지정한 경우 해당 컬럼이 널 인 경우 해당 로우는 제외됩니다
df.select(countDistinct("StockCode")).show()
df.selectExpr("count(distinct StockCode)").show()
from pyspark.sql.functions import *
# 근사치로 구하지만 연산 속도가 빠름
df.select(approx_count_distinct("StockCode", 0.1)).show() # 0.1은 최대 추정 오류율
df.select(approx_count_distinct("StockCode", 0.01)).show()
# ### 1.2 수치 집계 함수 (first, last, min, max, sum, sumDistinct, avg)
# +
from pyspark.sql.functions import *
df.select(first("StockCode"), last("StockCode")).show(1) # null도 감안하려면 True
df.select(min("Quantity"), max("Quantity")).show(1)
df.select(min("Description"), max("Description")).show(1) # 문자열
df.select(sum("Quantity")).show(1)
df.select(sumDistinct("Quantity")).show(1) # 고유값을 합산
# -
# ### 1.3 통계 집계 함수 (avg, mean, variance, stddev)
# * 표본표준분산 및 편차: variance, stddev
# * 모표준분산 및 편차 : var_pop, stddev_pop
# +
from pyspark.sql.functions import *
df.select(
count("Quantity").alias("total_transcations"),
sum("Quantity").alias("total_purchases"),
avg("Quantity").alias("avg_purchases"),
expr("mean(Quantity)").alias("mean_transcations"),
).selectExpr(
"total_purchases / total_transcations",
"avg_purchases",
"mean_transcations").show(3)
# -
df.select(
variance("Quantity")
, stddev("Quantity")
, var_samp("Quantity")
, stddev_samp("Quantity")
, var_pop("Quantity")
, stddev_pop("Quantity")
).show()
# ## 2. 그룹 함수
# ### 2.1 표현식을 이용한 그룹화
from pyspark.sql.functions import count
df.printSchema()
df.groupBy("InvoiceNo", "CustomerId").agg(expr("count(Quantity) as CountOfQuantity")).show(5)
# ### 2.2 맵을 이용한 그룹화
# > 파이선의 딕셔너리 데이터 타입을 활용하여 집계함수의 표현이 가능
df.groupBy("InvoiceNo").agg( { "Quantity" : "avg", "UnitPrice" : "stddev_pop" } ).show(5)
# ### <font color=blue>1. [중급]</font> 구매 이력 CSV "data/retail-data/all" 파일을 읽고
# #### 1. 스키마를 출력하세요
# #### 2. 데이터 10건을 출력하세요
# #### 3. 상품코드(StockCode)의 유일한 값의 갯수를 출력하세요
# #### 4. 상품단가(UnitPrice)의 최소, 최대 값을 출력하세요
# #### 5. 송장번호(StockCode)별로 송장별총매출금액(TotalInvoicePrice)를 계산하고 내림차순으로 정렬하세요
# #### 6. 송장별총매출금액(TotalInvoicePrice)이 최고금액이 송장을 필터하여 검증해 보세요
# ##### 예를 들어 `select sum(unit-price * quantity) from table where invoiceno = '123456'` 와 같은 쿼리로 검증이 가능합니다
#
# <details><summary>[실습7] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# df1 = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load("data/retail-data/all")
# )
# df1.printSchema()
# df1.show()
# answer = df1.withColumn("TotalPrice", expr("UnitPrice * Quantity")).groupBy("InvoiceNo").agg(sum("TotalPrice").alias("TotalInvoicePrice"))
# answer.printSchema()
# display(answer.orderBy(desc("TotalInvoicePrice")).limit(10))
#
# df1.where("InvoiceNo = '581483'").select(sum(expr("UnitPrice * Quantity"))).show()
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=green>2. [기본]</font> 매출 테이블 "data/tbl_purchase.csv" CSV 파일을 읽고
# #### 1. 스키마를 출력하세요
# #### 2. 데이터 10건을 출력하세요
# #### 3. 제품(p_name)별 금액(p_amount) 의 전체 합인 총 매출금액(sum_amount)을 구하세요
#
# <details><summary>[실습2] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# df2 = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load("data/tbl_purchase.csv")
# )
# df2.printSchema()
# df2.show()
# answer = df2.groupBy("p_name").agg(sum("p_amount").alias("sum_amount"))
# answer.printSchema()
# display(answer)
#
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=green>3. [기본]</font> 매출 테이블 "data/tbl_purchase.csv" CSV 파일을 읽고
# #### 1. 스키마를 출력하세요
# #### 2. 데이터 10건을 출력하세요
# #### 3. 구매 금액의 합이 가장 높은 고객(p_uid)을 구하세요
#
# <details><summary>[실습3] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# df3 = (
# spark.read.format("csv")
# .option("header", "true")
# .option("inferSchema", "true")
# .load("data/tbl_purchase.csv")
# )
# df3.printSchema()
# df3.show()
# answer = df2.groupBy("p_uid").agg(sum("p_amount").alias("sum_amount_per_user"))
# answer.printSchema()
# display(answer)
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=red>4. [고급]</font> 샌프란시스코 긴급출동 데이터 CSV 파일인 "data/learning-spark/sf-fire-calls.csv"를 읽고
# #### 1. 스키마를 출력하세요
# #### 2. 데이터를 3건 출력하세요
# #### 3. 호출의 종류(CallType)가 어떤 것들이 있는지 출력하세요 (중복제거)
# #### 3. 샌프란시스코에서 발생의 가장 빈도수가 높은 종류(CallType)를 구하고 빈도수를 구하세요
# #### 4. 샌프란시스코에서 발생하는 최고 빈도수 3건은 무엇인가요?
#
# <details><summary>[실습3] 출력 결과 확인 </summary>
#
# > 아래와 유사하게 방식으로 작성 되었다면 정답입니다
#
# ```python
# df3 = (
# spark
# .read
# .option("header", "true")
# .option("inferSchema", "true")
# .csv("data/learning-spark/sf-fire-calls.csv")
# )
# df3.printSchema()
# df3.show(3)
# df3.createOrReplaceTempView("fire_calls")
# spark.sql("select distinct(CallType) from fire_calls").show(truncate=False)
#
# answer = spark.sql("select CallType, count(CallType) as CallTypeCount from fire_calls group by CallType order by CallTypeCount desc")
# display(answer.limit(3))
# ```
#
# </details>
#
# 여기에 실습 코드를 작성하고 실행하세요 (Shift+Enter)
# ### <font color=red>5. [고급]</font> 샌프란시스코 긴급출동 데이터 CSV 파일인 "data/learning-spark/sf-fire-calls.csv"를 읽고 다음과 같은 질문도 실습해 보면 재미있을 것 같습니다
# #### 1. 2018 년의 모든 화재 신고 유형은 무엇 이었습니까?
# #### 2. 2018 년의 몇 월에 화재 신고가 가장 많았습니까?
# #### 3. 샌프란시스코에서 2018 년에 가장 많은 화재 신고가 발생한 지역은 어디입니까?
# #### 4. 2018 년에 화재 신고에 대한 응답 시간이 가장 나쁜 지역은 어디입니까?
# #### 5. 2018 년 중 어느 주에 화재 신고가 가장 많았습니까?
# #### 6. 이웃, 우편 번호, 화재 전화 건수간에 상관 관계가 있습니까?
# #### 7. Parquet 파일 또는 SQL 테이블을 사용하여이 데이터를 저장하고 다시 읽을 수있는 방법은 무엇입니까?
#
# ## 참고자료
#
# #### 1. [Spark Programming Guide](https://spark.apache.org/docs/latest/sql-programming-guide.html)
# #### 2. [PySpark SQL Modules Documentation](https://spark.apache.org/docs/latest/api/python/pyspark.sql.html)
# #### 3. <a href="https://spark.apache.org/docs/3.0.1/api/sql/" target="_blank">PySpark 3.0.1 Builtin Functions</a>
# #### 4. [PySpark Search](https://spark.apache.org/docs/latest/api/python/search.html)
# #### 5. [Pyspark Functions](https://spark.apache.org/docs/latest/api/python/pyspark.sql.html?#module-pyspark.sql.functions)
| day3/notebooks/lgde-spark-core-5-aggregation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Iris Unsupervised Learning"
# > "A tutorial on unsupervised learning using Iris dataset"
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
import pandas as pd
from sklearn import datasets
iris = datasets.load_iris()
df = pd.DataFrame(iris.data, columns=iris.feature_names)
df2 = pd.concat([df, pd.DataFrame(iris.target, columns=['class'])], axis=1)
df2
iris.target_names
# ## KMeans
# +
from sklearn.cluster import KMeans
from sklearn.datasets import make_blobs
from yellowbrick.cluster import KElbowVisualizer
visualizer = KElbowVisualizer(KMeans(), k=(4,12), metric='distortion', timings=False, locate_elbow=True)
visualizer.fit(df2)
visualizer.show()
# -
# ### Silhouette Visualizer
# The Silhouette Coefficient is used when the ground-truth about the dataset is unknown and computes the density of clusters computed by the model. The score is computed by averaging the silhouette coefficient for each sample, computed as the difference between the average intra-cluster distance and the mean nearest-cluster distance for each sample, normalized by the maximum value. This produces a score between 1 and -1, where 1 is highly dense clusters and -1 is completely incorrect clustering.
#
# The Silhouette Visualizer displays the silhouette coefficient for each sample on a per-cluster basis, visualizing which clusters are dense and which are not. This is particularly useful for determining cluster imbalance, or for selecting a value for K by comparing multiple visualizers.
#
# In SilhouetteVisualizer plots, clusters with higher scores have wider silhouettes, but clusters that are less cohesive will fall short of the average score across all clusters, which is plotted as a vertical dotted red line.
#
# This is particularly useful for determining cluster imbalance, or for selecting a value for K by comparing multiple visualizers.
# +
from yellowbrick.cluster import SilhouetteVisualizer
visualizer = SilhouetteVisualizer(KMeans(), colors='yellowbrick')
visualizer.fit(df2) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# -
# ### Intercluster Distance Maps
# Intercluster distance maps display an embedding of the cluster centers in 2 dimensions with the distance to other centers preserved. E.g. the closer to centers are in the visualization, the closer they are in the original feature space. The clusters are sized according to a scoring metric. By default, they are sized by membership, e.g. the number of instances that belong to each center. This gives a sense of the relative importance of clusters. Note however, that because two clusters overlap in the 2D space, it does not imply that they overlap in the original feature space.
# +
from yellowbrick.cluster import InterclusterDistance
visualizer = InterclusterDistance(KMeans(7))
visualizer.fit(df2) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# -
# By default, the scoring parameter metric is set to `distortion`, which computes the sum of squared distances from each point to its assigned center. However, two other metrics can also be used with the KElbowVisualizer – silhouette and `calinski_harabasz`. The `silhouette` score calculates the mean Silhouette Coefficient of all samples, while the calinski_harabasz score computes the ratio of dispersion between and within clusters.
# distortion: mean sum of squared distances to centers
#
# silhouette: mean ratio of intra-cluster and nearest-cluster distance
#
# calinski_harabasz: ratio of within to between cluster dispersion
# # Mini Batch KMeans
# +
from sklearn.cluster import MiniBatchKMeans
visualizer = KElbowVisualizer(MiniBatchKMeans(), k=(4,12), metric='distortion', timings=False, locate_elbow=False)
visualizer.fit(df2) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# +
visualizer = SilhouetteVisualizer(MiniBatchKMeans(), colors='yellowbrick')
visualizer.fit(df2) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# -
| _notebooks/28-02-2021-Iris-Unsupervised-Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Coin Flips and Die Rolls
# Use NumPy to create simulations and compute proportions for the following outcomes. The first one is done for you.
#
# **Please note again that we are using 0 to represent heads, and 1 to represent tails.**
# import numpy
import numpy as np
# ### 1. Two fair coin flips produce exactly two heads
# simulate 1 million tests of two fair coin flips
tests = np.random.randint(2, size=(int(1e6), 2))
# sums of all tests
test_sums = tests.sum(axis=1)
# proportion of tests that produced exactly two heads
(test_sums == 0).mean()
# ### 2. Three fair coin flips produce exactly one head
# +
# simulate 1 million tests of three fair coin flips
tests = np.random.randint(2, size=(int(1e6), 3))
# sums of all tests
test_sums = tests.sum(axis=1)
# proportion of tests that produced exactly one head
(test_sums == 1).mean()
# -
# ### 3. Three biased coin flips with P(H) = 0.6 produce exactly one head
# +
# simulate 1 million tests of three biased coin flips
# hint: use np.random.choice()
tests = np.random.choice([0,1],size=(int(1e6), 3),p=[0.6,0.4])
# sums of all tests
test_sums = tests.sum(axis=1)
# proportion of tests that produced exactly one head
(test_sums == 2).mean()
# -
# ### 4. A die rolls an even number
# simulate 1 million tests of one die roll
tests = np.random.randint(1,7,size=int(1e6))
even_count = 0
# proportion of tests that produced an even number
for i in tests:
if i % 2 == 0:
even_count = even_count+1
print(even_count/int(1e6))
# ### 5. Two dice roll a double
# +
# simulate the first million die rolls
first = np.random.randint(1,7,size=int(1e6))
# simulate the second million die rolls
second = np.random.randint(1,7,size=int(1e6))
# proportion of tests where the 1st and 2nd die rolled the same number
doubles = 0
for i in range(int(1e6)):
if first[i] == second[i]:
doubles = doubles + 1
print(doubles/int(1e6))
# -
| probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from tkinter import *
import random
import pandas as pd
class Application(Frame):
def __init__(self, master):
Frame.__init__(self, master)
self.grid()
self.create_widgets()
def create_widgets(self):
self.letters = " abcdefghijklmnopqrstuvwxyz"
self.letters_list = list(self.letters)
self.counter = 10
path = 'words.xlsx'
dataframe = pd.read_excel(path)
self.easy_column = dataframe.iloc[: , 0]
self.hard_column = dataframe.iloc[: , 1]
self.difficulty = StringVar()
self.difficulty.set('easy')
Radiobutton(self,
text = "easy",
variable = self.difficulty,
value = "easy"
).grid(row = 0, column = 0, sticky = W)
Radiobutton(self,
text = "hard",
variable = self.difficulty,
value = "hard"
).grid(row = 0, column = 1, sticky = W)
self.bttn1 = Button(self, text = "PLAY / RESET", command = self.play_game)
self.bttn1.grid(row = 0, column = 2, sticky = E)
def play_game(self):
"""creation of the rest of GUI and the word is chosen"""
self.lbl1 = Label(self, text="Guess this word:")
self.lbl1.grid(row = 2, column = 0, sticky = W)
self.lbl1_1 = Label(self, text="Guess a LETTER and press ENTER:")
self.lbl1_1.grid(row = 4, column = 0, sticky = W)
if self.difficulty.get() == 'easy':
self.word = random.choice(self.easy_column)
elif self.difficulty.get() == 'hard':
self.word = random.choice(self.hard_column)
self.solution = "-" * (len(self.word))
self.solution_list = list(self.solution)
self.lbl2 = Label(self, text = self.solution)
self.lbl2.grid(row = 2, column = 1, sticky = W)
self.lbl3_1 = Label(self, text = "Letters left to guess: ")
self.lbl3_1.grid(row = 3, column = 0, sticky = W)
self.lbl3_2 = Label(self, text = "".join(self.letters_list))
self.lbl3_2.grid(row = 3, column = 1, sticky = W)
self.ent = Entry(self)
self.ent.grid(row = 4, column = 1, sticky = W)
self.ent.bind('<KeyPress>', self.keyboard)
self.ent.focus_set()
self.lbl4 = Label(self, text = "Guesses left:")
self.lbl4.grid(row = 5, column = 0, sticky = W)
self.lbl5 = Label(self, text = str(self.counter))
self.lbl5.grid(row = 5, column = 1, sticky = W)
def guess_char(self):
print(self.word)
print("".join(self.solution_list))
guess = self.ent.get()
self.ent.delete(0, END)
if self.counter > 1:
temp_state = self.solution_list.copy()
for i in range(len(self.word)):
if self.word[i].upper() == guess.upper():
self.solution_list[i] = self.word[i]
self.lbl2["text"] = "".join(self.solution_list)
if self.word == "".join(self.solution_list):
self.lbl5["text"] = "You win at hangman!"
elif guess in self.letters_list:
self.letters_list.remove(guess)
self.lbl3_2["text"] = "".join(self.letters_list)
if temp_state == self.solution_list:
self.counter -= 1
self.lbl5["text"] = self.counter
else:
self.lbl5["text"] = str(self.counter) + " - You've already guessed that letter."
else:
self.lbl2["text"] = "You lose. The word you were looking for: " + self.word
self.lbl5["text"] = "Game Over."
def keyboard(self, event):
if event.keysym == "Return":
self.guess_char()
root = Tk()
root.title("Hangman Game Made By : <NAME>, Abhijeet ,Diwakar, Ashish")
root.geometry("500x500")
root.resizable(width = TRUE, height = TRUE)
app = Application(root)
root.mainloop()
# -
dir(root)
# +
# import random
# Gamer = input("Enter your Spirit Word ?")
# print("Bad luck indeed -> you will Lose", Gamer)
# choices = []
# with open("words.txt", "r") as file:
# allopt = file.read()
# choices = list(map(str, allopt.split()))
# word = random.choice(choices)
# numberofvowels = 0
# for i in word:
# if i in ['a','e','i','o','u']:
# numberofvowels += 1
# rev = word[::-1]
# # print(numberofvowels)
# # print(rev)
# print("Guess the opts")
# guesses = ''
# length = len(word)
# turns = length * 2
# while turns > 0:
# failed = 0
# if turns == length//3:
# print("Hint : the number of vowels :", numberofvowels)
# if turns == length//2:
# if(rev == word):
# print("Hint : the word is palindrome")
# for char in word:
# if char in guesses:
# print(char)
# else:
# print("_")
# failed += 1
# if failed == 0:
# print("You Win")
# print("The word is: ", word)
# break
# guess = input("guess a character:")
# guesses += guess
# if guess not in word:
# turns -= 1
# print("Wrong")
# print("You have", + turns, 'more guesses')
# if turns == 0:
# print("You Loose")
# print("The answer was :",word)
# -
# !pip install pyinstaller
| T&T/projects/hungman.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from twilio.rest import Client
pip install twilio
from twilio.rest import Client
# enter your account_sid details from twilio accout
acc_sid='*******************'
# enter your tocken details by removing stars
auth_token='****************'
# enter your cell number
my_cell='*******************'
# enter your twilio cell number
my_twilio='**********'
client=Client(acc_sid,auth_token)
my_msg= 'hai u are hacked...☼♀♂♠♦♣♥•◘○↕‼©®ÃJ,¡☻'
message = client.messages.create(to=my_cell,from_=my_twilio,body=my_msg)
| send_Sms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import time
from scipy.spatial.transform import Rotation as R
import matplotlib.pyplot as plt
import numpy.linalg as LA
def Superpose3D_original(aaXf_orig, # <-- coordinates for the "frozen" object
aaXm_orig, # <-- coordinates for the "mobile" object
aWeights=None, # <- optional weights for the calculation of RMSD
allow_rescale=False): # <-- attempt to rescale mobile object?
"""
Superpose3D() takes two lists of xyz coordinates, (of the same length)
and attempts to superimpose them using rotations, translations, and
(optionally) rescale operations in order to minimize the
root-mean-squared-distance (RMSD) between them.
These operations should be applied to the "aaXm_orig" argument.
This function returns a tuple containing:
(RMSD, optimal_translation, optimal_rotation, and optimal_scale_factor)
This function implements a more general variant of the method from:
<NAME>, (1988)
"A Note on the Rotational Superposition Problem",
Acta Cryst. A44, pp. 211-216
This version has been augmented slightly. The version in the original
paper only considers rotation and translation and does not allow the
coordinates of either object to be rescaled (multiplication by a scalar).
(Additional documentation can be found at
https://pypi.org/project/superpose3d/ )
"""
assert (len(aaXf_orig) == len(aaXm_orig))
N = len(aaXf_orig)
if (aWeights == None) or (len(aWeights) == 0):
aWeights = np.full(N, 1.0)
# Find the center of mass of each object:
aCenter_f = np.zeros(3)
aCenter_m = np.zeros(3)
sum_weights = 0.0
for n in range(0, N):
for d in range(0, 3):
aCenter_f[d] += aaXf_orig[n][d] * aWeights[n]
aCenter_m[d] += aaXm_orig[n][d] * aWeights[n]
sum_weights += aWeights[n]
for d in range(0, 3):
aCenter_f[d] /= sum_weights
aCenter_m[d] /= sum_weights
# Subtract the centers-of-mass from the original coordinates for each object
aaXf = np.empty((N, 3))
aaXm = np.empty((N, 3))
aaXf[0][0] = 0.0
for n in range(0, N):
for d in range(0, 3):
aaXf[n][d] = aaXf_orig[n][d] - aCenter_f[d]
aaXm[n][d] = aaXm_orig[n][d] - aCenter_m[d]
# Calculate the "M" array from the Diamond paper (equation 16)
M = np.zeros((3, 3))
for n in range(0, N):
for i in range(0, 3):
for j in range(0, 3):
M[i][j] += aWeights[n] * aaXm[n][i] * aaXf[n][j]
# Calculate Q (equation 17)
traceM = 0.0
for i in range(0, 3):
traceM += M[i][i]
Q = np.empty((3, 3))
for i in range(0, 3):
for j in range(0, 3):
Q[i][j] = M[i][j] + M[j][i]
if i == j:
Q[i][j] -= 2.0 * traceM
# Calculate V (equation 18)
V = np.empty(3)
V[0] = M[1][2] - M[2][1];
V[1] = M[2][0] - M[0][2];
V[2] = M[0][1] - M[1][0];
# Calculate "P" (equation 22)
P = np.empty((4, 4))
for i in range(0, 3):
for j in range(0, 3):
P[i][j] = Q[i][j]
P[0][3] = V[0]
P[3][0] = V[0]
P[1][3] = V[1]
P[3][1] = V[1]
P[2][3] = V[2]
P[3][2] = V[2]
P[3][3] = 0.0
aEigenvals, aaEigenvects = LA.eigh(P)
# http://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.eigh.html
eval_max = aEigenvals[0]
i_eval_max = 0
for i in range(1, 4):
if aEigenvals[i] > eval_max:
eval_max = aEigenvals[i]
i_eval_max = i
# The vector "p" contains the optimal rotation (in quaternion format)
p = np.empty(4)
p[0] = aaEigenvects[0][i_eval_max]
p[1] = aaEigenvects[1][i_eval_max]
p[2] = aaEigenvects[2][i_eval_max]
p[3] = aaEigenvects[3][i_eval_max]
# normalize the vector
# (It should be normalized already, but just in case it is not, do it again)
pnorm = np.linalg.norm(p)
for i in range(0, 4):
p[i] /= pnorm
# Finally, calculate the rotation matrix corresponding to "p"
# (convert a quaternion into a 3x3 rotation matrix)
aaRotate = np.empty((3, 3))
aaRotate[0][0] = (p[0] * p[0]) - (p[1] * p[1]) - (p[2] * p[2]) + (p[3] * p[3])
aaRotate[1][1] = -(p[0] * p[0]) + (p[1] * p[1]) - (p[2] * p[2]) + (p[3] * p[3])
aaRotate[2][2] = -(p[0] * p[0]) - (p[1] * p[1]) + (p[2] * p[2]) + (p[3] * p[3])
aaRotate[0][1] = 2 * (p[0] * p[1] - p[2] * p[3]);
aaRotate[1][0] = 2 * (p[0] * p[1] + p[2] * p[3]);
aaRotate[1][2] = 2 * (p[1] * p[2] - p[0] * p[3]);
aaRotate[2][1] = 2 * (p[1] * p[2] + p[0] * p[3]);
aaRotate[0][2] = 2 * (p[0] * p[2] + p[1] * p[3]);
aaRotate[2][0] = 2 * (p[0] * p[2] - p[1] * p[3]);
pPp = eval_max
# Optional: Decide the scale factor, c
c = 1.0 # by default, don't rescale the coordinates
if allow_rescale:
Waxaixai = 0.0
WaxaiXai = 0.0
for a in range(0, N):
for i in range(0, 3):
Waxaixai += aWeights[a] * aaXm[a][i] * aaXm[a][i]
WaxaiXai += aWeights[a] * aaXm[a][i] * aaXf[a][i]
c = (WaxaiXai + pPp) / Waxaixai
# Finally compute the RMSD between the two coordinate sets:
# First compute E0 from equation 24 of the paper
E0 = 0.0
for n in range(0, N):
for d in range(0, 3):
# (remember to include the scale factor "c" that we inserted)
E0 += aWeights[n] * ((aaXf[n][d] - c * aaXm[n][d]) ** 2)
sum_sqr_dist = E0 - c * 2.0 * pPp
if sum_sqr_dist < 0.0: # (edge case due to rounding error)
sum_sqr_dist = 0.0
rmsd = np.sqrt(sum_sqr_dist / sum_weights)
# Lastly, calculate the translational offset:
# Recall that:
# RMSD=sqrt((Σ_i w_i * |X_i - (Σ_j c*R_ij*x_j + T_i))|^2) / (Σ_j w_j))
# =sqrt((Σ_i w_i * |X_i - x_i'|^2) / (Σ_j w_j))
# where
# x_i' = Σ_j c*R_ij*x_j + T_i
# = Xcm_i + c*R_ij*(x_j - xcm_j)
# and Xcm and xcm = center_of_mass for the frozen and mobile point clouds
# = aCenter_f[] and aCenter_m[], respectively
# Hence:
# T_i = Xcm_i - Σ_j c*R_ij*xcm_j = aTranslate[i]
aTranslate = np.empty(3)
for i in range(0, 3):
aTranslate[i] = aCenter_f[i]
for j in range(0, 3):
aTranslate[i] -= c * aaRotate[i][j] * aCenter_m[j]
# An alternate method to compute "aTranslate" using numpy matrices:
# Rmatrix = np.matrix(aaRotate)
# TcolumnVec = np.matrix(np.empty((3,1))) # 3x1 numpy matrix<->[[0],[0],[0]]
# for d in range(0,3):
# TcolumnVec[d][0] = -aCenter_m[d]
# TcolumnVec = c * Rmatrix * TcolumnVec
# for d in range(0,3):
# TcolumnVec[d][0] += aCenter_f[d]
# #Turn the column vector back into an ordinary numpy array of size 3:
# aTranslate = np.array(TcolumnVec.transpose())[0]
return rmsd, aaRotate, aTranslate, c
def Superpose3D_v2(aaXf_orig, # <-- coordinates for the "frozen" object
aaXm_orig, # <-- coordinates for the "mobile" object
# ---- optional arguments: ----
aWeights=None, # optional weights for the calculation of RMSD
allow_rescale=False, # attempt to rescale mobile point cloud?
report_quaternion=False): # report rotation angle and axis
aaXf_orig = np.array(aaXf_orig)
aaXm_orig = np.array(aaXm_orig)
if aaXf_orig.shape[0] != aaXm_orig.shape[0]:
raise ValueError ("Inputs should have the same size.")
N = aaXf_orig.shape[0]
if (aWeights == None) or (len(aWeights) == 0):
aWeights = np.full((N,1),1.0)
else:
aWeights = np.array(aWeights).reshape(N,1)
aCenter_f = np.sum(aaXf_orig * aWeights, axis=0)
aCenter_m = np.sum(aaXm_orig * aWeights, axis=0)
sum_weights = np.sum(aWeights, axis=0)
if sum_weights != 0:
aCenter_f /= sum_weights
aCenter_m /= sum_weights
aaXf = aaXf_orig-aCenter_f
aaXm = aaXm_orig-aCenter_m
M = aaXm.T @ (aaXf * aWeights)
Q = M + M.T - 2*np.eye(3)*np.trace(M)
V = np.empty(3)
V[0] = M[1][2] - M[2][1];
V[1] = M[2][0] - M[0][2];
V[2] = M[0][1] - M[1][0];
P = np.zeros((4,4))
P[:3, :3] = Q
P[3,:3] = V
P[:3, 3] = V
p = np.zeros(4)
p[3] = 1.0 # p = [0,0,0,1] default value
pPp = 0.0 # = p^T * P * p (zero by default)
singular = (N < 2) # (it doesn't make sense to rotate a single point)
try:
aEigenvals, aaEigenvects = LA.eigh(P)
except LinAlgError:
singular = True # (I have never seen this happen.)
if (not singular): # (don't crash if the caller supplies nonsensical input)
i_eval_max = np.argmax(aEigenvals)
pPp = np.max(aEigenvals)
p[:] = aaEigenvects[:, i_eval_max]
p /= np.linalg.norm(p)
the_rotation = R.from_quat(p)
aaRotate = the_rotation.as_matrix()
c = 1.0 # by default, don't rescale the coordinates
if allow_rescale and (not singular):
Waxaixai = np.sum(aWeights * aaXm ** 2)
WaxaiXai = np.sum(aWeights * aaXf ** 2)
c = (WaxaiXai + pPp) / Waxaixai
E0 = np.sum((aaXf - c*aaXm)**2)
sum_sqr_dist = max(0, E0 - c * 2.0 * pPp)
rmsd = 0.0
if sum_weights != 0.0:
rmsd = np.sqrt(sum_sqr_dist/sum_weights)
aTranslate = aCenter_f - (c*aaRotate @ aCenter_m).T.reshape(3,)
if report_quaternion: # does the caller want the quaternion?
q = np.empty(4)
q[0] = p[3] # Note: The "p" variable is not a quaternion in the
q[1] = p[0] # conventional sense because its elements
q[2] = p[1] # are in the wrong order. I correct for that here.
q[3] = p[2] # "q" is the quaternion correspond to rotation R
return rmsd, q, aTranslate, c
else:
return rmsd, aaRotate, aTranslate, c
data1 = np.random.random((10000,3))
data2 = data1+0.01*np.random.random((10000,3)) #creating fake data with some noise.
the_rotation = R.from_euler('xyz', [30,30,30])
data2 = the_rotation.apply(data2) + np.array([0.2,-0.3,0.5])
plt.figure(figsize=(10,10))
plt.plot(data1[:,0], data1[:,1], "o", ms = 3)
plt.plot(data2[:,0], data2[:,1], "o",ms = 3)
# +
limits = [10, 20, 30, 50, 100,150, 300,500, 1000,2000,3000,5000,10000]
times_original = []
times_v2 = []
for limit in limits:
temp_original = []
temp_v2 = []
for _ in range (20):
initial = time.time()
_, rot1, trans1, _ = Superpose3D_original(data1[:limit],data2[:limit])
final = time.time()
temp_original.append(final-initial)
initial = time.time()
_, rot2, trans2, _ = Superpose3D_v2(data1[:limit],data2[:limit])
final = time.time()
temp_v2.append(final-initial)
times_original.append(np.mean(temp_original))
times_v2.append(np.mean(temp_v2))
# +
plt.figure(figsize = (7,7), dpi=150)
plt.plot(limits,times_original, "o--", label = "original version")
plt.plot(limits,times_v2, "o--", label = "proposed version")
plt.xlabel("Number of data points")
plt.ylabel("Time (s)")
plt.legend()
plt.savefig("times.pdf")
# -
import test_superpose3d
test_superpose3d.test_superpose3d()
| Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise - Filter files
# How can I get files that I want to rename to?
# Let's try regular expression pattern.
# ## References
# https://docs.python.org/3/library/re.html </p><p>
# count</p><p>
# The optional argument count is the maximum number of pattern occurrences to be replaced; count must be a non-negative integer. If omitted or zero, all occurrences will be replaced. </p><p>
# flags</p><p>
# re.A (ASCII-only matching), re.I (ignore case), re.L (locale dependent), re.M (multi-line), re.S (dot matches all), re.U (Unicode matching), and re.X (verbose)
# +
# Pattern history
path = "D:\\Projects\\Excel VBA"
# regular expression pattern for unicode
uni_rep_str = u'【(.+)】(.+).bas'
regular_expression_pattern = uni_rep_str
# replacement_pattern = '\1_\2.png' NG
# replacement_pattern = 'Larisa_Alexandrite_ver\g<1>.\g<2>\g<3>.0.vroid'
replacement_pattern = '\g<1>__\g<2>.vbs'
# test_str
test_str = u'【参考】コーディング規約.bas'
# Options
count = 0
flags = 0
# +
import re
pattern = re.compile(regular_expression_pattern, flags)
# -
# ## Generate a new name
# How can I generage a new file name?
# +
# By using regexp method
replaced_filename1 = re.sub(regular_expression_pattern, replacement_pattern, test_str, count, flags)
print(replaced_filename1)
print('-'*20)
# By using compiled regexp object
replaced_filename2 = pattern.sub(replacement_pattern, test_str, count)
print(replaced_filename2)
# -
# ## Get a file list by filtering
#
# ## Reference
# https://docs.python.org/3/library/re.html
# <p>The optional argument count is the maximum number of pattern occurrences to be replaced; count must be a non-negative integer.</p>
# +
import os
match_file_list = []
with os.scandir(path) as it:
for entry in it:
if not entry.name.startswith('.') and entry.is_file() and pattern.match(entry.name):
match_file_list.append(entry.name)
# -
# ## Get a new name in each files
# OK! Let's combine the above methods.
# ## Reference
# https://docs.python.org/3/library/re.html
# <p>The optional argument count is the maximum number of pattern occurrences to be replaced; count must be a non-negative integer.</p>
# +
# New name
renamed_file_list = []
for target_file_name in match_file_list:
new_name = pattern.sub(replacement_pattern, target_file_name)
if new_name:
renamed_file_list.append(new_name)
print(renamed_file_list)
| Exercise_02_filter_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Setup the imports
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
import math
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler
from sklearn.pipeline import make_pipeline
# ### Read in the data
train = pd.read_csv('./data/train.csv', index_col=0)
train.head()
# #### Fill NA
import matplotlib.pyplot as plt
import seaborn as sns
plt.figure(figsize=(6, 4))
sns.heatmap(train.isna(), cbar=False)
# REPLACE NA values with age of non- and survived passengers and ...
mean_Age = train.groupby(['Survived','Sex','Pclass'])['Age'].transform('mean')
train['Age'].fillna(mean_Age, inplace=True)
from sklearn.preprocessing import KBinsDiscretizer
# transform a numerical column: Age
kbins = KBinsDiscretizer(n_bins=8, encode='onehot-dense', strategy='kmeans')
columns = train[['Age']]
kbins.fit(columns)
t = kbins.transform(columns)
# fill NA of Embarked with most frequent values, then binning
pipeline1 = make_pipeline(
SimpleImputer(strategy='most_frequent'),
OneHotEncoder(sparse=False, handle_unknown='ignore')
)
# fill NA of Embarked with most frequent values, then binning
pipeline2 = make_pipeline(
SimpleImputer(strategy='mean'),
KBinsDiscretizer(n_bins=8, encode='onehot-dense', strategy='kmeans')
)
pipeline3 = make_pipeline(
SimpleImputer(strategy='mean'),
MinMaxScaler()
)
# model 1
trans = ColumnTransformer([
('onehot', OneHotEncoder(sparse=False, handle_unknown='ignore'), ['Sex','Pclass']),
('impute_then_onehot',pipeline1, ['Embarked']),
('impute_then_scale', pipeline2, ['Age']),
('scale', pipeline3, ['Fare']),
('do_nothing', 'passthrough', ['SibSp','Parch']),
])
# +
#### Train-Test-Split
from sklearn.model_selection import train_test_split as tts
X = train.iloc[:, 1:]
y = train['Survived']
Xtrain, Xtest,ytrain,ytest = tts(X,y,train_size=0.9)
Xtrain.shape, Xtest.shape, ytrain.shape, ytest.shape
# -
# ### Feature Engineering
# +
# fit and transform training data
trans.fit(train)
X = trans.transform(train)
y = train['Survived']
# -
# ### Train an ML Model
m_lr = LogisticRegression(max_iter=1000)
m_lr.fit(X,y) #(x_train, y_train)
# +
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(m_lr,X,y)
# +
#Evaluating metrics
from sklearn.metrics import classification_report
ypred = m_lr.predict(X)
# predict
acc = accuracy_score(y,ypred)
print('Train accuracy is:', round(acc,3))
# -
print(classification_report(y,ypred))
trans.fit(Xtest)
test1 = trans.transform(Xtest)
ypred_test = m_lr.predict(test1)
# +
#calculate probabilities
# prob = m.predict_proba(test1)
# -
acc_test = accuracy_score(ytest, ypred_test)
print('Test accuracy is:', round(acc_test,3))
# +
# ytrue1 = ytest['Survived']
# draw ROC curve
# +
# from sklearn.metrics import roc_curve
# works for Titanic but not for Penguins
# probs = m.predict_proba(Xtest)
# roc_curve(ytest, probs) #y_true, y_score
# -
# ### Train a classification tree
# +
# Cross-Validate Decision Tree
from sklearn.tree import DecisionTreeClassifier
m_dt = DecisionTreeClassifier(max_depth=2) # we allow that many questions
m_dt.fit(X, y)
# Import cross_validate
from sklearn.model_selection import cross_validate
cross_validate_dt = cross_validate(estimator=m_dt,
X=X, # Xtrain
y=y, # ytrain
scoring='accuracy',
return_train_score=True)
cross_validate_dt['test_score'].mean(), cross_validate_dt['train_score'].mean()
# +
# find the appropriate depth
x_a = []
test_score_l = []
train_score_l = []
for i in range(1,15):
m_dt = DecisionTreeClassifier(max_depth=i)
# Cross-Validate Decision Tree
cross_validate_dt = cross_validate(estimator=m_dt,
X=X,
y=y,
scoring='accuracy',
cv=5,
return_train_score=True)
x_a = i
test_score_l = cross_validate_dt['test_score'].mean()
train_score_l = cross_validate_dt['train_score'].mean()
print(x_a,round(test_score_l,3),round(train_score_l,3) )
# +
# plot
# -
m_dt = DecisionTreeClassifier(max_depth=5) # we allow that many questions
m_dt.fit(X, y)
# ### train a random forest model
# +
from sklearn.ensemble import RandomForestClassifier
# # ? m=RandomForestRegressor(n_estimators=100)
m_rf = RandomForestClassifier(n_estimators=100)
m_rf.fit (X, y)
# -
# ### compare the models - training score
# +
from sklearn.model_selection import cross_val_score
def test_classifier(clf):
scores = cross_val_score(clf,X,y, cv=5)
print(' %0.3f(+/-%0.3f)'%(scores.mean(),scores.std()))
print('Train accuracy of LR is:')
test_classifier(m_lr)
print('Train accuracy of DT is:')
test_classifier(m_dt)
print('Train accuracy of RF is:')
test_classifier(m_rf)
# -
# ### compare the models - test score
#lord test data
#Transform the test data
trans.fit(Xtest)
Xtest1 = trans.transform(Xtest)
m_dt.score(Xtest1, ytest)
# Inspect the accuracy score
print('Test accuracy of LR is:', m_lr.score(Xtest1, ytest))
print('Test accuracy of DT is:', m_dt.score(Xtest1, ytest))
print('Test accuracy of RF is:', m_rf.score(Xtest1, ytest))
m_lr,m_dt,m_rf
#roc curve
# # kragge scoring
# +
test_tia = pd.read_csv('./data/test.csv', index_col=0)
test_tia.head(200)
trans.fit(test_tia)
test2 = trans.transform(test_tia)
ypred2 = m_lr.predict(test2)
submission = pd.DataFrame(ypred2, index=test_tia.index, columns=['Survived'])
submission.to_csv('submission_lr.csv') # added Embarked -> 0.765, # 0.756(drop Embarked)
ypred3 = m_dt.predict(test2)
submission = pd.DataFrame(ypred3, index=test_tia.index, columns=['Survived'])
submission.to_csv('submission_dt.csv') # added Embarked -> 0.7655, # 0.772(drop Embarked)
ypred4 = m_rf.predict(test2)
submission = pd.DataFrame(ypred4, index=test_tia.index, columns=['Survived'])
submission.to_csv('submission_rf.csv') # added Embarked -> 0.763, # 0.751(drop Embarked)
# -
# ###
# +
# Cabin data
# Cabin data
#isolating the rooms and letters
train['Cabin_nr'] = train['Cabin'].fillna('Z',inplace=False)
train["Deck"] = train["Cabin_nr"].str.slice(0,1)
def one_hot_column(df, label, drop_col=False):
one_hot = pd.get_dummies(df[label], prefix=label)
if drop_col:
df = df.drop(label, axis=1)
df = df.join(one_hot)
return df
def one_hot(df, labels, drop_col=False):
for label in labels:
df = one_hot_column(df, label, drop_col)
return df
train = one_hot(train, ["Deck"],drop_col=True)
# +
test['Cabin_nr'] = test['Cabin'].fillna('Z',inplace=False)
test["Deck"] = test["Cabin_nr"].str.slice(0,1)
def one_hot_column(df, label, drop_col=False):
one_hot = pd.get_dummies(df[label], prefix=label)
if drop_col:
df = df.drop(label, axis=1)
df = df.join(one_hot)
return df
def one_hot(df, labels, drop_col=False):
for label in labels:
df = one_hot_column(df, label, drop_col)
return df
test = one_hot(test, ["Deck"],drop_col=True)
# -
trans = ColumnTransformer([
('onehot', OneHotEncoder(sparse=False, handle_unknown='ignore'), ['Sex','Pclass']),
('scale', pipeline3, ['Fare']),
('impute_then_scale', pipeline2, ['Age']),
('impute_then_onehot',pipeline1, ['Embarked']),
('do_nothing', 'passthrough', ['SibSp','Parch','Deck_A','Deck_B','Deck_C','Deck_D','Deck_E','Deck_F','Deck_G','Deck_Z']),
])
trans.fit(test)
test2 = trans.transform(test)
ypred = m.predict(test2)
# +
# first letter of name
nameletter = train['Name'].str.slice(0,1)
new_numbers = [];
# letter changed to number
for n in nameletter:
new_numbers.append(ord(n));
numbers = new_numbers;
train.loc[:,'nameletter'] = numbers
# train = one_hot(train, ['nameletter'],drop_col=True)
# +
# first letter of name
nameletter = test['Name'].str.slice(0,1)
new_numbers = [];
# letter changed to number
for n in nameletter:
new_numbers.append(ord(n));
numbers = new_numbers;
test.loc[:,'nameletter'] = numbers
# train = one_hot(train, ['nameletter'],drop_col=True)
# -
test
# first letter of name
#ss = test['Name'].str.slice(0,1)
#test = pd.concat( [test, pd.DataFrame(ss)], axis=1 )
# test = one_hot(test, ['nameletter'],drop_col=True)
# test.assign(nameletter=test['Name'].str.slice(0,1))
# +
# test.loc[:,'nameletter'] = test['Name'].str.slice(0,1)
# +
# train.loc[:,'nameletter'] = train['Name'].str.slice(0,1)
# -
trans = ColumnTransformer([
('onehot', OneHotEncoder(sparse=False, handle_unknown='ignore'), ['Sex', 'Pclass']),
('scale', pipeline3, ['Fare']),
('impute_then_scale', pipeline2, ['Age']),
('impute_then_onehot',pipeline1, ['Embarked']),
('minmax',KBinsDiscretizer(n_bins=5, encode='onehot-dense', strategy='kmeans'),['nameletter']),
('do_nothing', 'passthrough', ['SibSp','Parch']),
])
test
train
| week_02/Titanic train data model compare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: adapter
# language: python
# name: adapter
# ---
import pandas as pd
def process_csv(data_path):
df = pd.read_csv(data_path)
df.loc[:, "Sentence #"] = df["Sentence #"].fillna(method="ffill")
sentences = df.groupby("Sentence #")["Word"].apply(list).values
tags = df.groupby("Sentence #")["newTag"].apply(list).values
return sentences, tags
sentences, tags = process_csv("../PyTorch/NER_test/NER_multilabel_data_v2.csv")
from ner_dataset import get_trainset_data_loader
# +
from transformers import RobertaTokenizer
import numpy as np
tokenizer = RobertaTokenizer.from_pretrained("roberta-base")
# -
all_tags, trainset, trainloader = get_trainset_data_loader(tokenizer, "../PyTorch/NER_test/NER_multilabel_data_v2.csv")
trainloader
| Practice/adapter_roberta/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # 探索差别隐私
#
# 在机器学习项目中,通常涉及通过迭代的数据分析过程来洞察数据,并确定哪些变量最有可能帮助构建预测模型。分析数据通常涉及到聚合和统计功能,通过这些功能可以了解变量的统计分布情况以及它们之间的关系。如果数据量很大,聚合功能可以为结果实现一定程度的抽象化;但是如果数据量较小,或者存在重复的分析,那么即使是聚合后的结果也可能会暴露个别观测值的详细信息。
#
# 差分隐私这种技术可通过对数据添加“干扰”来保护单个数据点隐私。其目标是确保添加足够的干扰以实现单个值的隐私性,同时确保数据的总体统计组成不变,并且聚合在统计上产生的结果与使用原始数据时相似。
#
# ## 安装 SmartNoise SDK
#
# [*SmartNoise*](https://smartnoise.org/) 是来自 OpenDP 的工具包,这是一个 Microsoft 和哈佛大学的研究人员以及其他贡献者共同合作的项目,旨在为在数据分析和机器学习项目中使用差异隐私提供构建块。
#
# > **备注**:SmartNoise 目前处于开发的早期阶段。
#
# 首先,我们要安装 SmartNoise Python SDK 包。可以忽略此练习中有关 Azure CLI 兼容性的任何错误。
# !pip install opendp-smartnoise==0.1.4.2
# ## 加载数据
#
# 现在让我们来看看一些原始数据。在本例中,我们有 10,000 个已接受糖尿病检测的患者记录。
# +
import pandas as pd
data_path = 'data/diabetes.csv'
diabetes = pd.read_csv(data_path)
diabetes.describe()
# -
# 以上代码的输出显示了糖尿病数据集中变量的关键汇总统计信息。
#
# ## 执行分析
#
# 你可以使用 SmartNoise 创建一个能为源数据添加干扰的分析。干扰添加之下的数学运算可能相当复杂,不过 SmartNoise 能帮助你处理大部分细节问题。但你需要了解一些概念。
#
# - **上限和下限**:钳位用于设置变量值的上限和下限。这是为了确保 SmartNoise 产生的干扰与原始数据的预期分布一致。
# - **样本大小**:要为部分聚合生成一致的差分隐私数据,SmartNoise 需要知道要生成的数据样本的大小。
# - **Epsilon**:简单地说,*epsilon* 是一个非负值,它提供了对添加到数据中的干扰量的反向度量。如果 epsilon 较低,数据集的隐私级别就更高;如果 epsilon 较高,则数据集会更接近原始数据。Epsilon 的取值通常在 0 到 1 之间。Epsilon 与另一个名为 *delta* 的值相关,该值表示的是某项分析生成的报告并不是完全隐私的概率。
#
# 记住这些概念后,检查并运行以下代码,此代码会创建一个分析并报告来自差分隐私数据的平均“**年龄**”值。还显示了初始数据的实际平均值以供比较。
# +
import opendp.smartnoise.core as sn
cols = list(diabetes.columns)
age_range = [0.0, 120.0]
samples = len(diabetes)
with sn.Analysis() as analysis:
# load data
data = sn.Dataset(path=data_path, column_names=cols)
# Convert Age to float
age_dt = sn.to_float(data['Age'])
# get mean of age
age_mean = sn.dp_mean(data = age_dt,
privacy_usage = {'epsilon': .50},
data_lower = age_range[0],
data_upper = age_range[1],
data_rows = samples
)
analysis.release()
# print differentially private estimate of mean age
print("Private mean age:",age_mean.value)
# print actual mean age
print("Actual mean age:",diabetes.Age.mean())
# -
# ## 用直方图探索数据分布情况
#
# 直方图常用于分析数据以检查变量分布。
#
# 例如,让我们看看糖尿病数据集中年龄的真实分布情况。
# +
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
ages = list(range(0, 130, 10))
age = diabetes.Age
# Plot a histogram with 10-year bins
n_age, bins, patches = plt.hist(age, bins=ages, color='blue', alpha=0.7, rwidth=0.85)
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Age')
plt.ylabel('Frequency')
plt.title('True Age Distribution')
plt.show()
print(n_age.astype(int))
# -
# 现在让我们来比照一下“年龄”的差分隐私直方图。
# +
import matplotlib.pyplot as plt
with sn.Analysis() as analysis:
data = sn.Dataset(path = data_path, column_names = cols)
age_histogram = sn.dp_histogram(
sn.to_int(data['Age'], lower=0, upper=120),
edges = ages,
upper = 10000,
null_value = -1,
privacy_usage = {'epsilon': 0.5}
)
analysis.release()
plt.ylim([0,7000])
width=4
agecat_left = [x + width for x in ages]
agecat_right = [x + 2*width for x in ages]
plt.bar(list(range(0,120,10)), n_age, width=width, color='blue', alpha=0.7, label='True')
plt.bar(agecat_left, age_histogram.value, width=width, color='orange', alpha=0.7, label='Private')
plt.legend()
plt.title('Histogram of Age')
plt.xlabel('Age')
plt.ylabel('Frequency')
plt.show()
print(age_histogram.value)
# -
# 两个直方图的相似程度足以确保基于差分隐私数据的报告与源自原始数据的报告能提供一致的见解。
#
# ## 计算协方差
#
# 分析的另一个共同目标是建立变量之间的关系。SmartNoise 提供了一个差分隐私协方差函数,可以帮助解决这个问题。
with sn.Analysis() as analysis:
sn_data = sn.Dataset(path = data_path, column_names = cols)
age_bp_cov_scalar = sn.dp_covariance(
left = sn.to_float(sn_data['Age']),
right = sn.to_float(sn_data['DiastolicBloodPressure']),
privacy_usage = {'epsilon': 1.0},
left_lower = 0.,
left_upper = 120.,
left_rows = 10000,
right_lower = 0.,
right_upper = 150.,
right_rows = 10000)
analysis.release()
print('Differentially private covariance: {0}'.format(age_bp_cov_scalar.value[0][0]))
print('Actual covariance', diabetes.Age.cov(diabetes.DiastolicBloodPressure))
# 在本例中,**Age** 与 **DisatolicBloodPressure** 之间的协方差为正,说明年龄越大,血压越高。
#
# ## 使用 SQL 查询
#
# 除了**分析**功能之外,通过 SmartNoise 还能对数据源使用 SQL 查询以检索差分隐私聚合生成的结果。
#
# 首先,需要为数据架构中的表定义元数据。你可以在 .yml 文件中执行此操作,例如 **/metadata** 文件夹中的 **diabetes.yml** 文件。元数据描述表中的字段,包括数据类型以及数值字段的最小值和最大值。
# +
from opendp.smartnoise.metadata import CollectionMetadata
meta = CollectionMetadata.from_file('metadata/diabetes.yml')
print (meta)
# -
# 定义了元数据后,可以创建你可以查询的读取器。在下面的示例中,我们将创建一个 **PandasReader** 来读取 Pandas 数据帧中的原始数据,并创建一个 **PrivateReader**,它将为 **PandasReader** 添加差分隐私层。
# +
from opendp.smartnoise.sql import PandasReader, PrivateReader
reader = PandasReader(diabetes, meta)
private_reader = PrivateReader(reader=reader, metadata=meta, epsilon_per_column=0.7)
print('Readers ready.')
# -
# 现在,你可以向隐私读取器提交一个返回聚合结果集的 SQL 查询。
# +
query = 'SELECT Diabetic, AVG(Age) AS AvgAge FROM diabetes.diabetes GROUP BY Diabetic'
result_dp = private_reader.execute(query)
print(result_dp)
# -
# 我们来比较该结果和源于原始数据的相同聚合。
result = reader.execute(query)
print(result)
# 可以使用 **epsilon_per_column** 参数自定义 **PrivateReader** 的行为。
#
# 我们来尝试对一个读取器采用较高的 epsilon 值(即隐私度较低)并对另一个读取器采用较低的 epsilon 值(即隐私度较高)。
# +
low_privacy_reader = PrivateReader(reader, meta, 5.0) # large epsilon, less privacy
result = low_privacy_reader.execute(query)
print(result)
print()
high_privacy_reader = PrivateReader(reader, meta, 0.1) # smaller epsilon, more privacy
result = high_privacy_reader.execute(query)
print(result)
# -
# 请注意,高 epsilon 值(隐私度较低)读取器的结果比低 epsilon 值(隐私度较高)读取器的结果更接近源于原始数据的真实结果。
#
# ## 拓展阅读
#
# 要详细了解 SmartNoise 提供的差分隐私功能,请参阅 [https://smartnoise.org](https://smartnoise.org/)
| 13 - Explore Differential Privacy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
path = '../Data/df2.csv'
df1 = pd.read_csv(path, parse_dates=['Date'])
df1 = df1.rename(columns = {"Date":"ds","Close":"y"})
df1
import datetime as datetime
# use data before covid time
ctf_date = datetime.datetime(2020, 3, 1)
df = df1.loc[df1.ds < ctf_date , ].copy()
df
# +
# test data: predict 1 year (~252 trading days)
pred_periods = 252
# cutoff between test and train data
cutoff = len(df) - 252
df_train = df[:cutoff].copy()
df_test = df[cutoff:].copy()
print(cutoff)
# +
# Random Forest Model
my_rfg = RFG(n_estimators = 200, random_state =820)
# fit close price using random forest model
my_rfg.fit(df_train[['Open','High','Low']].copy(),df_train.y.copy())
y_pred = my_rfg.predict(df_test[['Open','High','Low']])
# +
# Naive Arima Model
from pmdarima.arima import auto_arima
model = auto_arima(df_train.y, start_p=1, start_q=1,max_p=3, max_q=3, m=12,start_P=0, seasonal=True,d=1, D=1, trace=True,error_action='ignore',suppress_warnings=True)
# fit close price using arima model
model.fit(df_train.y)
arima_forecast = model.predict(n_periods=pred_periods)
# +
# FB Prophet model
from fbprophet import Prophet
fbp = Prophet(daily_seasonality=True)
# fit close price using fbprophet model
fbp.fit(df_train[['ds','y']])
# predict pred_size futures and get the forecast price
fut = fbp.make_future_dataframe(periods = pred_periods, freq='D')
fb_forecast = fbp.predict(fut)
# +
plt.figure(figsize=(18,10))
# # plot the forecast
plt.plot(df_test.ds, fb_forecast[cutoff:].yhat,'g--', label = "FB Forecast")
plt.plot(df_test.ds, df_test.y,'b--', label = "Test Data")
plt.plot(df_test.ds , y_pred,'r--',label="Random Forest Forecast")
plt.plot(df_test.ds , arima_forecast,'y--',label="ARIMA Forecast")
# plt.plot(df_test.ds ,df.y,'r-o',label="test result")
plt.title('Comparison of Models', fontsize=16)
plt.legend(fontsize=14)
plt.xlabel("Date", fontsize=16)
plt.ylabel("S&P500 Close Price", fontsize=16)
plt.show()
# +
from sklearn.metrics import mean_squared_error as MSE
rf_mse_test = MSE(df_test.y, y_pred[cutoff:]) # mse for random forest model and sp500 close price
arima_mse_test = MSE(df_test.y,arima_forecast ) # mse for naive arima model and sp500 close price
fb_mse_test = MSE(df_test.y, fb_forecast[cutoff:].yhat) # mse for FB prophet model and sp500 close price
print(rf_mse_test,arima_mse_test,fb_mse_test )
# -
| Code/Scenario1ModelComparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false" slideshow={"slide_type": "slide"}
# <img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
#
# # MAT281
# ### Aplicaciones de la Matemática en la Ingeniería
# + [markdown] Collapsed="false" slideshow={"slide_type": "slide"}
# ## Módulo 04
# ## Laboratorio Clase 02: Regresión Lineal
# + [markdown] Collapsed="false"
# ### Instrucciones
#
#
# * Completa tus datos personales (nombre y rol USM) en siguiente celda.
# * La escala es de 0 a 4 considerando solo valores enteros.
# * Debes _pushear_ tus cambios a tu repositorio personal del curso.
# * Como respaldo, debes enviar un archivo .zip con el siguiente formato `mXX_cYY_lab_apellido_nombre.zip` a <EMAIL>, debe contener todo lo necesario para que se ejecute correctamente cada celda, ya sea datos, imágenes, scripts, etc.
# * Se evaluará:
# - Soluciones
# - Código
# - Que Binder esté bien configurado.
# - Al presionar `Kernel -> Restart Kernel and Run All Cells` deben ejecutarse todas las celdas sin error.
# + [markdown] Collapsed="false"
# __Nombre__: <NAME>
#
# __Rol__: 201630001-5
# + Collapsed="false"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import altair as alt
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.linear_model import LinearRegression
alt.themes.enable('opaque')
# %matplotlib inline
# + [markdown] Collapsed="false"
# ## Ejercicio 1: Diabetes
# + [markdown] Collapsed="false"
# Realizar análisis de regresión a los datos de diabetes disponibles en scikit-learn
# + Collapsed="false"
diabetes = datasets.load_diabetes()
print(dir(diabetes)) ## Atributos
# + Collapsed="false"
print(diabetes.DESCR)
# + Collapsed="false"
diabetes_df = (
pd.DataFrame(
diabetes.data,
columns=diabetes.feature_names
)
.assign(prog=diabetes.target)
)
diabetes_df.head()
# + [markdown] Collapsed="false"
# #### Pregunta 1 (1 pto):
#
# * ¿Por qué la columna de sexo tiene esos valores?
# * ¿Cuál es la columna a predecir?
# + [markdown] Collapsed="false"
# * El hecho de que el sexo posea valores numéricos, facilita su manipulación al hacer una regresión, ya que es preferible trabajar con datos cuantituativos.
# * La columna a predecir es "prog", ya que todas las demás variables las demás puede influir de una u otra manera en el progreso de la enfermedad.
# + [markdown] Collapsed="false"
# #### Pregunta 2 (1 pto)
#
# Realiza una regresión lineal con todas las _features_ incluyendo intercepto.
# + Collapsed="false"
X = diabetes_df.drop(columns = 'prog').values
y = diabetes_df['prog'].values
# + [markdown] Collapsed="false"
# Ajusta el modelo
# + Collapsed="false"
regr = LinearRegression()
regr.fit(X,y)
# + [markdown] Collapsed="false"
# Imprime el intercepto y los coeficientes luego de ajustar el modelo.
# + Collapsed="false"
print(f"Intercept: \n{regr.intercept_}\n")
print(f"Coefficients: \n{regr.coef_}\n")
# + [markdown] Collapsed="false"
# Haz una predicción del modelo con los datos `X`.
# + Collapsed="false"
y_pred = regr.predict(X)
# + [markdown] Collapsed="false"
# Calcula e imprime el error cuadrático medio y el coeficiente de determinación de este modelo ajustado.
# + Collapsed="false"
# Error cuadrático medio
print(f"Mean squared error: {mean_squared_error(y, y_pred)}\n")
# Coeficiente de determinación
print(f"Coefficient of determination: {r2_score(y, y_pred)}")
# + [markdown] Collapsed="false"
# **Pregunta: ¿Qué tan bueno fue el ajuste del modelo?**
# + [markdown] Collapsed="false"
# Gracias al coeficiente de determinación podemos ver que no es el mejor ajuste, ya que podría estar más cerca de 1. Asimismo, el error cuadrático medio nos muestra que existe una diferencia no despreciable entre los datos reales y los predecidos con la regresión.
# + [markdown] Collapsed="false"
# ### Pregunta 3 (2 ptos).
#
# Realizar multiples regresiones lineales utilizando una sola _feature_ a la vez.
#
# En cada iteración:
#
# - Crea un arreglo `X`con solo una feature filtrando `X`.
# - Crea un modelo de regresión lineal con intercepto.
# - Ajusta el modelo anterior.
# - Genera una predicción con el modelo.
# - Calcula e imprime las métricas de la pregunta anterior.
# + Collapsed="false"
for i in range(X.shape[1]):
X_i = X[:, np.newaxis, i] # Protip! Trata de entender este paso por tu cuenta, es muy clever
regr_i = LinearRegression()
regr_i.fit(X_i,y)
y_pred_i = regr_i.predict(X_i)
print(f"{diabetes_df.columns[i]}:")
print(f"\tCoefficients: {float(regr_i.coef_)}")
print(f"\tIntercept: {regr_i.intercept_}")
print(f"\tMean squared error: {mean_squared_error(y, y_pred_i): .2f}")
print(f"\tCoefficient of determination: {r2_score(y, y_pred_i): .2f}\n")
# + [markdown] Collapsed="false"
# **Si tuvieras que escoger una sola _feauture_, ¿Cuál sería? ¿Por qué?**
# + [markdown] Collapsed="false"
# Escogería el Índice de Masa Corporal (BMI), ya que entre todos los ajustes es el que presenta mayor coeficiente de determinación y menor error cuadrático medio.
# + [markdown] Collapsed="false"
# Con la feature escogida haz el siguiente gráfico:
#
# - Scatter Plot
# - Eje X: Valores de la feature escogida.
# - Eje Y: Valores de la columna a predecir (target).
# - En color rojo dibuja la recta correspondiente a la regresión lineal (utilizando `intercept_`y `coefs_`).
# - Coloca un título adecuado, nombre de los ejes, etc.
#
# Puedes utilizar `matplotlib` o `altair`, el que prefiera.
# + Collapsed="false"
bmi_reg = LinearRegression().fit(X[:, np.newaxis, 2],y)
points = alt.Chart(diabetes_df).mark_circle(size=75, opacity=0.5).encode(
x="bmi:Q",
y="prog:Q"
).properties(
width=600,
height=400
)
reg_df = (
diabetes_df.loc[
lambda x: x["bmi"].isin([x["bmi"].min(), x["bmi"].max()])
, ["bmi"]
].assign(regression=lambda x: bmi_reg.intercept_ + x["bmi"] * bmi_reg.coef_)
)
reg_line = alt.Chart(reg_df).mark_line(color="red", opacity=0.5).encode(
x=alt.X("bmi:Q", title='BMI'),
y=alt.Y("regression:Q", title='Progreso de la diabetes')
).properties(
title='Regresión Lineal: Progreso de diabetes v/s BMI',
width=600,
height=400
)
points + reg_line
| m04_machine_learning/m04_c02_linear_regression/m04_c02_lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
import nltk
from nltk.corpus import stopwords
engstopwords = stopwords.words('English')
engstopwords.extend(['mg', 'kg', 'mg kg', 'hcc', 'aarc'])
from sklearn.svm import LinearSVC
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import cross_validate,LeaveOneOut, train_test_split
df = pd.read_csv("/Users/smruthi/Desktop/TechTogether2020/data/docs_100_share.csv")
# +
#Cleaning up non-ascii chars from abstract
newabstracts = []
for abstract in df['abstract']:
test_list = (abstract.split())
for i in range(len(test_list)):
if not test_list[i].isascii():
test_list[i] = re.sub(r'[^\x00-\x7F]+',' ', test_list[i])
newabstracts.append(' '.join(test_list))
df['abstract'] = newabstracts
# -
df.head()
def createFeatures(corpus):
newcorpus = []
for example in corpus:
words = nltk.word_tokenize(example)
words = nltk.pos_tag(words)
wordlist = []
for word in words:
#filtering stopwords, checking for alphabetic chars
if word[0].isalpha() and word[0] not in engstopwords:
#each feature is word+position tag
wordlist.append(word[0]+"_"+word[1])
newcorpus.append(" ".join(wordlist))
#adding maximum document frequency of 5
#since high frequency words in every abstract will be terms like "patient"
#these don't tell us anything about relevancy
vectorizer = CountVectorizer(max_df=5)
texts = vectorizer.fit_transform(newcorpus).toarray()
vocab = vectorizer.get_feature_names()
return texts,vocab
# +
def evaluateModel(X,y,vocab,penalty="l1"):
#create and fit the model
model1 = LogisticRegression(penalty=penalty,solver="liblinear")
results1 = cross_validate(model1,X,y,cv=LeaveOneOut())
model2 = LinearSVC(random_state=0, tol=1e-5)
results2 = cross_validate(model2,X,y,cv=LeaveOneOut())
#determine the average accuracy
scores1 = results1["test_score"]
avg_score1 = sum(scores1)/len(scores1)
scores2 = results2["test_score"]
avg_score2 = sum(scores2)/len(scores2)
model1.fit(X,y)
class1_prob_sorted1 = model1.coef_[0, :].argsort()
class2_prob_sorted1 = (-model1.coef_[0, :]).argsort()
model2.fit(X,y)
class1_prob_sorted2 = model2.coef_[0, :].argsort()
class2_prob_sorted2 = (-model2.coef_[0, :]).argsort()
termsToTake = 20
class1_indicators1 = [vocab[i] for i in class1_prob_sorted1[:termsToTake]]
class2_indicators1 = [vocab[i] for i in class1_prob_sorted1[:termsToTake]]
class1_indicators2 = [vocab[i] for i in class2_prob_sorted2[:termsToTake]]
class2_indicators2 = [vocab[i] for i in class2_prob_sorted2[:termsToTake]]
return avg_score1,class1_indicators1,class2_indicators1,avg_score2,class1_indicators2,class2_indicators2
def runEvaluation(X,y,vocab):
print("----------L2 Norm-----------")
avg_score1,class1_indicators1,class2_indicators1,avg_score2,class1_indicators2,class2_indicators2 = evaluateModel(X,y,vocab,"l2")
print("Logistic")
print("The model's average accuracy is %f"%avg_score1)
print("The most informative terms for exclude are: %s"%class1_indicators1)
print("The most informative terms for include are: %s"%class1_indicators2)
print("SVM")
print("The model's average accuracy is %f"%avg_score2)
print("The most informative terms for exclude are: %s"%class2_indicators1)
print("The most informative terms for include are: %s"%class2_indicators2)
# -
corpus = list(df['abstract'])
corpus = [x.split(". ")[-2:] for x in corpus]
corpus = [x[0]+". "+x[1] for x in corpus]
y = df['include/exclude']
X,vocab = createFeatures(corpus)
runEvaluation(X, y, vocab)
| jupyter notebooks/Bonus Challenge TTB20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS109A Introduction to Data Science
#
# ## Standard Section 6: PCA and Logistic Regression
#
# **Harvard University**<br/>
# **Fall 2019**<br/>
# **Instructors**: <NAME>, <NAME>, and <NAME><br/>
# **Section Leaders**: <NAME>, Abhimanyu (<NAME>, Robbert (<NAME><br/>
#
# <hr style='height:2px'>
#RUN THIS CELL
import requests
from IPython.core.display import HTML
styles = requests.get("http://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# For this section, our goal is to get you familiarized with Dimensionality Reduction using Principal Components Analysis (PCA) and to recap Logistic Regression from the last homework. This [medium article](https://towardsdatascience.com/pca-using-python-scikit-learn-e653f8989e60) was referenced extensively while creating this notebook.
#
# Specifically, we will:
#
# - Understand how to define the terms **big data** and **high-dimensionality** and see the motivation for PCA
# - Learn what PCA is
# - Use PCA in order to **visualize** a high-dimensional problem in 2-dimensions
# - Learn about the sklearn PCA library and [its nuances](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html)
# - Get familiar with the Linear Algebra of PCA
# - Meet the MNIST handwritten digit dataset (and hopefully stay friends for a while)
# - Use PCA in order to **improve model training time** and understand the **speed-accuracy trade-off**
# - Discuss when to use PCA and when not to use it
#
# ---
# <img src="../fig/meme.png" width="400">
#
# ---
# +
# Data and stats packages
import numpy as np
import pandas as pd
# Visualization packages
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# Other packages
pd.set_option('display.max_columns', 50)
import warnings
warnings.filterwarnings("ignore")
# -
# # Motivation
# Principal Components Analysis helps us deal with high-dimensionality in big-data.
#
# But first...
#
# <img src="../fig/bigdata.png" width="600">
#
# **High-dimensionality** is the case when p is large i.e. there are a lot of predictors. This is sometimes a problem because:
#
# 1. Our models may be overfit
# 2. There may be multi-collinearity
# 3. Matrices may not be invertible (in the case of OLS)
#
# **Our challenge**: is to represent these p dimensions by a smaller number (m) dimensions without losing too much information. Then, we can fit a model using these m predictors, which addresses the three problems listed above. Here's where **PCA** comes into play.
# # What is Principal Components Analysis (PCA)?
#
# ## A Framework For Dimensionality Reduction
# We said that one way to reduce the dimensions of the feature space is to create a new, smaller set of predictors by taking **linear combinations** of the original predictors. Our original model (let's say it is a Linear Regression Model) looks like this:
#
# $$
# Y = \beta_0 + \beta_1 X_1 + \beta_2 X_2 + \dots + \beta_p X_p + \epsilon
# $$
#
# We choose $Z_1$, $Z_2$,$\dots$, $Z_m$, where $m < p$ and where each $Z_i$ is a linear combination of
# the original p predictors, $X_1$ to $X_p$. We can say that:
#
# $$
# Z_i = \sum_{j=1}^{p} c_{ij} X_i
# $$
#
# for fixed constants $c_{ij}$ (PCA can determines them). As an example, we could say that:
#
# $$
# Z_1 = 3.3 X_1 + 4 X_2 + 0 X_3 + \dots + 1.2 X_p
# $$
#
# In the above equation, we see that $Z_1$ is a linear combination of the original predictors. Then we can build a linear regression model using the new predictors as follows:
#
# $$
# Y = \theta_0 + \theta_1 Z_1 + \theta_2 Z_2 + \dots + \theta_m Z_m + \epsilon
# $$
#
# Notice that this model has a smaller number (m+1 < p+1) of parameters. Each $Z_i$ is called **PRINCIPAL COMPONENT**. The principcal components consist an $m$-dimensional **orthonormal** system of coordinates.
#
# PCA is a method to identify a new set of predictors, as linear combinations of the original ones, that captures the 'maximum amount' of variance in the observed data. This is the basic assumption in the PCA.
#
# <img src="../fig/pca.png" width="400">
#
# We see that the "best line" is the one where there is maximal variance along the line. Source [here](https://stats.stackexchange.com/questions/2691/making-sense-of-principal-component-analysis-eigenvectors-eigenvalues/140579).
#
# <img src="../fig/pca.gif" width="1000">
#
# In principle, we could explore all the rotations, that is, rotating our coordinate system under all the angles, and find which rotation yields the maximum variance or the smallest covariance. However, when the dimensionality (p) is large this is very time consuming and inefficient technique. In that case we may use PCA which is systematic way to find the best rotation or the best coordinate system.
# PCA is a mathematical method based on linear algebra, for more details and rigorous formulation see the notes in the advanced section for PCA.
#
# ## Applications of PCA
#
# One major application of PCA is to address the issues we pointed out earlier (reduce the number of predictors).
#
# In addition, another major application of PCA is in **visualization**. Specifically, if we have an N-dimensional dataset, how do we visualize it?
#
# **One option**:
#
# <img src="../fig/matrix.png" width="400">
#
# **A more practical option**: use PCA to get the top 2-3 principal components and plot these components on 2-d or 3-d plots!
# # PCA for Visualization
#
# Data Source: [MTCars Dataset](https://gist.github.com/seankross/a412dfbd88b3db70b74b)
#
# Here are a few resources that use this dataset and apply PCA for visualization. This notebook references [this PCA tutorial in R](https://www.datacamp.com/community/tutorials/pca-analysis-r), [these lecture notes from CMU](http://www.stat.cmu.edu/~cshalizi/uADA/12/lectures/ch18.pdf), this [blog](https://www.analyticsvidhya.com/blog/2016/03/practical-guide-principal-component-analysis-python/), and [this blog](http://setosa.io/ev/principal-component-analysis/) which has some nice visualizations of PCA.
# ### Loading in The Cars Dataset and carry out EDA
#
# This dataset consists of data on 32 models of car, taken from an American motoring magazine (1974 Motor Trend magazine). For each car, you have 11 features, expressed in varying units (US units), They are as follows ([source](https://www.datacamp.com/community/tutorials/pca-analysis-r)):
#
# - `mpg`: Fuel consumption (Miles per (US) gallon): more powerful and heavier cars tend to consume more fuel.
# - `cyl`: Number of cylinders: more powerful cars often have more cylinders
# - `disp`: Displacement (cu.in.): the combined volume of the engine's cylinders
# - `hp`: Gross horsepower: this is a measure of the power generated by the car
# - `drat`: Rear axle ratio: this describes how a turn of the drive shaft corresponds to a turn of the wheels. Higher values will decrease fuel efficiency.
# - `wt`: Weight (1000 lbs): pretty self-explanatory!
# - `qsec`: 1/4 mile time: the cars speed and acceleration
# - `vs`: Engine block: this denotes whether the vehicle's engine is shaped like a "V", or is a more common straight shape.
# - `am`: Transmission: this denotes whether the car's transmission is automatic (0) or manual (1).
# - `gear`: Number of forward gears: sports cars tend to have more gears.
# - `carb`: Number of carburetors: associated with more powerful engines
#
# Note that the units used vary and occupy different scales.
#
# **We are dropping the categorical variables `vs` and `am` before we progress any further, and only keeping in the continuous predictors**.
cars_df = pd.read_csv('../data/mtcars.csv')
cars_df = cars_df[cars_df.columns.difference(['am', 'vs'])]
cars_df.head()
cars_df.describe()
# **Our task** is to try to visualize this data in a meaningful way. Obviously we can't make a 9-dimensional plot, but we can try to make several different plots using the `pairplot` function from seaborn.
sns.pairplot(cars_df);
# But there are numerous variables and numerous more relationships between these variables. We can do better through PCA.
# ## A Better Visualization using PCA
#
# ### Standardizing Variables
# Standardization is crucial for PCA. For more details check the notes from the advanced section.
# +
from sklearn.preprocessing import StandardScaler
# separating the quantitative predictors from the model of the car (a string)
model = cars_df['model']
quant_df = cars_df[cars_df.columns.difference(['model'])]
# Standardization
quant_scaled = StandardScaler().fit_transform(quant_df)
cars_df_scaled = pd.DataFrame(quant_scaled, columns=quant_df.columns)
# bringing back the model name
cars_df_scaled['model'] = cars_df['model']
cars_df_scaled.describe()
cars_df_scaled.head()
# -
# ### Carrying out PCA
#
# [Sklearn PCA documentation](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html)
# +
from sklearn.decomposition import PCA
# drop again the model predictor
quant_df = cars_df_scaled[cars_df_scaled.columns.difference(['model'])]
# fitting the PCA object onto our dataframe (excluding the model name column)
pca = PCA().fit(quant_df)
# transforming the dataframe
quant_df_pca = pca.transform(quant_df)
print(quant_df.shape)
print(quant_df_pca.shape)
# -
# Let us examine some of the attributes we obtain from PCA.
#
# 1. `explained_variance_`: The amount of variance explained by each of the selected principal components.
# 2. `explained_variance_ratio_`: Percentage of variance explained by each of the selected principal components. By default, if `n_components` is not set then all components are stored and the sum of the ratios is equal to 1.0.
# +
fig, ax = plt.subplots(ncols=2, figsize=(20,6))
ax1, ax2 = ax.ravel()
ratio = pca.explained_variance_ratio_
ax1.bar(range(len(ratio)), ratio, color='purple', alpha=0.8)
ax1.set_title('Explained Variance Ratio PCA', fontsize=20)
ax1.set_xticks(range(len(ratio)))
ax1.set_xticklabels(['PC {}'.format(i+1) for i in range(len(ratio))])
ax1.set_ylabel('Explained Variance Ratio')
# ratio[0]=0
ratio = pca.explained_variance_ratio_
# cumulative sum
ax2.plot(np.cumsum(ratio), 'o-')
ax2.set_title('Cumulative Sum of Explained Variance Ratio PCA', fontsize=20)
ax2.set_ylim(0,1.1)
ax2.set_xticks(range(len(ratio)))
ax2.set_xticklabels(['PC {}'.format(i+1) for i in range(len(ratio))])
ax2.set_ylabel('Cumulative Sum of Explained Variance Ratio');
# -
# We see that over 85% of the variance is explained by the first 2 principal components!
ratio
# 3. `components_`: This represents the principal components i.e. directions of maximum variance in the data. The components are sorted by `explained_variance_`.
#
# Let us write the equation for all the principal components using our formulation of the principal components above:
#
# $$
# Z_i = \sum_{j=1}^{p} w_{ij} X_i
# $$
pca.components_
for i, comp in enumerate(pca.components_):
expression = 'Z_{} = '.format(i+1)
for c, x in zip(comp, quant_df.columns):
if c < 0:
expression += str(np.round(c,2)) + '*' + x + ' '
else:
expression += '+' + str(np.round(c,2)) + '*' + x + ' '
print(expression + '\n')
# Using the printed equations above, we can create vectors showing where each feature has a high value. Let us do this for the first 2 principal components (using $v$ to denote a vector):
#
# $$
# \begin{aligned}
# v_{carb} = \begin{pmatrix}-0.24 \\ 0.48 \end{pmatrix}, \;
# v_{cyl} = \begin{pmatrix}-0.4 \\ 0.02 \end{pmatrix}, \;
# v_{disp} = \begin{pmatrix}-0.4 \\ -0.09 \end{pmatrix}, \\
# v_{drat} = \begin{pmatrix}0.31 \\ 0.34 \end{pmatrix}, \;
# v_{gear} = \begin{pmatrix}0.21 \\ 0.55 \end{pmatrix}, \;
# v_{hp} = \begin{pmatrix}-0.37 \\ 0.27 \end{pmatrix}, \\
# v_{mpg} = \begin{pmatrix}0.39 \\ 0.03 \end{pmatrix}, \;
# v_{qsec} = \begin{pmatrix}0.22 \\ -0.48 \end{pmatrix}, \;
# v_{wt} = \begin{pmatrix}-0.37 \\ -0.17 \end{pmatrix}
# \end{aligned}
# $$
# ### Checking if our vectors are orthonormal
# Orthonormal vectors are the vectors which are orthogonal (zero dot product) with length equal to one (unit vectors).
#
# #### Orthogonal:
# We use the dot product between two vectors to check if the vectors are orthogonal or not. If the dot product is 0, that means that the two vectors are orthogonal. The dot product between two vectors is (geometrically):
#
# $$
# \textbf{a} \cdot \textbf{b} = ||\textbf{a}|| ||\textbf{b}|| \cos(\theta)
# $$
#
# Where $\theta$ is the angle between the two vectors and $||\cdot||$ denotes the norm of the vector. Since we assume that the norm of a vector is non-zero, the only way the dot product of two vectors to be zero is when the angle between them is 90 degrees (since $\cos(90) = 0$). Thus, the dot product is a good way to check if two vectors are perpendicular.
#
# #### Unit vectors
# In order to calculate the length $||\textbf{a}||$ of a vector we can take the dot product of a vector with itself, namely
# $$
# ||\textbf{a}|| = \textbf{a}\cdot \textbf{a}
# $$
# +
vec1 = pca.components_[0]
vec2 = pca.components_[1]
# print(np.dot(vec1.T, vec2))
print('The dot product between the first two principal components is ',np.round(np.dot(vec1, vec2),5))
print('The length of the first principal component is ',np.round(np.dot(vec1, vec1),5))
# -
# We see that the first two principal components are orthogonal and the first principal component is also a unit vector. Check other pairs of principal components in order to convince yourself that all principal components are always pairwise orthogonal unit vectors.
vec
# ### Plotting Results
# +
# to plot vectors from the center
# take the transverse of the matrix
vecs = pca.components_[0:2].T*2
fig, ax = plt.subplots(figsize=(16,8))
ax.plot(quant_df_pca[:,0], quant_df_pca[:,1], 'o', markersize=0.01)
ax.set_xlabel('Principal Component 1')
ax.set_ylabel('Principal Component 2')
ax.set_title('Cars Dataset plotted using first 2 Principal Components', fontsize=20)
# plotting arrowheads
for i, vec in enumerate(vecs):
ax.arrow(0,0,vec[0],vec[1], color='brown', head_width=0.1)
s = 1.3
# s denotes the locational place
ax.annotate(quant_df.columns[i], (s*vec[0], s*vec[1]), color='brown')
# annotating text
for i, txt in enumerate(cars_df_scaled['model']):
#(x,y) for location
ax.annotate(txt, (quant_df_pca[:,0][i], quant_df_pca[:,1][i]), size=12)
# -
# **Any patterns of interest?** Let us examine the geography more closely. Source: [this blog](https://www.datacamp.com/community/tutorials/pca-analysis-r).
country = ["Japan", "US", "EU", "US", "EU", "Japan", "US", "EU", "US", "EU"]
times = [3, 4, 7, 3, 1, 3, 4, 3, 1, 3]
country_list = np.array(sum(([x]*y for x,y in zip(country, times)),[]))
country_list
# +
fig, ax = plt.subplots(figsize=(16,8))
# main plot
ax.plot(quant_df_pca[:,0], quant_df_pca[:,1], 'o', markersize=0.01)
ax.set_xlabel('Principal Component 1')
ax.set_ylabel('Principal Component 2')
ax.set_title('Cars Dataset plotted using first 2 Principal Components', fontsize=20)
# plotting arrowheads
for i, vec in enumerate(vecs):
ax.arrow(0,0,vec[0],vec[1], color='brown', head_width=0.05)
s = 1.3
ax.annotate(quant_df.columns[i], (s*vec[0], s*vec[1]), color='brown')
# plotting names
cs = [sns.xkcd_rgb["magenta"], sns.xkcd_rgb["denim blue"], sns.xkcd_rgb["medium green"]]
colors = {"Japan": cs[0], "US": cs[1], "EU": cs[2]}
# dummy plots to show up in the legend
ax.plot(0,0, color=cs[0], label='Japan')
ax.plot(0,0, color=cs[1], label='US')
ax.plot(0,0, color=cs[2], label='EU')
# plotting text with color
for i, txt in enumerate(cars_df_scaled['model']):
country = country_list[i]
ax.annotate(txt, (quant_df_pca[:,0][i], quant_df_pca[:,1][i]), color=colors[country], size=12)
ax.legend(fontsize=15);
# -
# What patterns do you see now?
#
# For instance, we can observe that fuel consumption (*mpg*) is increased for the japanese cars and weight (*wt*) is increased for US cars.
# ### Addressing `n_components`
#
# We notice that PCA takes in 1 parameter: `n_components`. This is the number of principal components that PCA will use. By default, the number of the used principal components is the minimum of the number of rows and the number of columns in the dataset.
#
# **Note**: Setting the default parameter for `n_components` and taking the top-k principal components is equivalent to setting `n_components=k`. Let's check this.
# +
old_components = pca.components_[0:2]
# doing pca with 2 components
pca2 = PCA(n_components=2).fit(quant_df)
new_components = pca2.components_
# checking equivalence
print(new_components.all() == old_components.all())
# -
# # PCA to speed up classification of Handwritten Digits
#
# This example, using the [MNIST dataset](http://yann.lecun.com/exdb/mnist/), was borrowed from this [Towards Data Science blog post](https://towardsdatascience.com/pca-using-python-scikit-learn-e653f8989e60). In this example, we will be classifying hand-written digits.
#
# ## Data Loading and EDA
# +
# we'll use keras a lot more in the last few weeks of the course
from keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
print(x_train.shape, y_train.shape, x_test.shape, y_test.shape)
# -
# Our training set (`x_train`) contains 60000 images of size 28 by 28. Our training labels (`y_train`) are numbers from 0 to 9. Let's examine one of these values below.
print(y_train[0])
fig, ax = plt.subplots()
ax.grid(None)
ax.imshow(x_train[0], cmap='gray');
# **Our task** is to classify the test set digits as accurately as possible.
# We notice that the shape of the training set is $6000 \times 28 \times 28$ which is a 3-dimensional array. We have not dealt with these kinds of arrays before. We will deal with images in greater detail (and not only!!!) in the follow-up course, *CS 109b*, if you are interested in doing more of this kind of stuff you should take this course. For now, we will **reshape** the array into a 2-dimensional array of shape $6000\times 784$.
# +
x_train = x_train.reshape(x_train.shape[0], 784)
x_test = x_test.reshape(x_test.shape[0], 784)
# check if the shapes are ok
print(x_train.shape, y_train.shape, x_test.shape, y_test.shape)
# -
# ## Normalizing Data
#
# Image data is usually between 0 and 255 (0 indicating a black pixel and 255 indicating a white pixel). We can normalize these values by dividing by 255.
# +
# checking the min and max of x_train and x_test
print(x_train.min(), x_train.max(), x_test.min(), x_test.max())
x_train = (x_train - x_train.min())/(x_train.max() - x_train.min())
x_test = (x_test - x_train.min())/(x_train.max() - x_train.min())
print(x_train.min(), x_train.max(), x_test.min(), x_test.max())
# -
# ## Modeling using Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
from time import time
start = time()
#‘lbfgs’ solver handles multinomial loss in multiclass problems
logreg_model = LogisticRegression(solver='lbfgs').fit(x_train, y_train)
end = time()
full_logreg_time = end-start
print('Time to fit: {}s'.format(full_logreg_time))
# +
from sklearn.metrics import accuracy_score
y_preds_train = logreg_model.predict(x_train)
y_preds_test = logreg_model.predict(x_test)
full_logreg_score_train = accuracy_score(y_train, y_preds_train)
full_logreg_score_test = accuracy_score(y_test, y_preds_test)
# Evaluation
print('Training Set Score: {}'.format(full_logreg_score_train))
print('Test Set Score: {}'.format(full_logreg_score_test))
# -
# get performance by class
pd.crosstab(y_test, y_preds_test, margins=True, rownames=['Actual'], colnames=['Predicted'])
# We get a high training and test set score but it takes a relatively long time to fit a model. Let us see if we can speed things up when using PCA
# ## Logistic Regression Model after PCA
# +
# Do PCA onto our training set and inspect
pca = PCA(n_components=100).fit(x_train)
fig, ax = plt.subplots(ncols=2, figsize=(20,6))
ax1, ax2 = ax.ravel()
ratio = pca.explained_variance_ratio_
ax1.plot(range(1,len(ratio)+1), ratio, 'o-')
ax1.set_title('Explained Variance Ratio PCA', fontsize=20)
ax1.set_ylabel('Explained Variance Ratio')
ratio = pca.explained_variance_ratio_
ax2.plot(range(1,len(ratio)+1), np.cumsum(ratio), 'o-')
ax2.set_title('Cumulative Sum of Explained Variance Ratio PCA', fontsize=20)
ax2.set_ylabel('Cumulative Sum of Explained Variance Ratio');
# -
# We see that the first 100 principal components hold over 90% of the variance and the first 50 principal components hold over 80% of the variance! We have significantly reduced the dimensionality of our problem! Let us use PCA to find the first 100 principal components of our dataset and transform our `x_train` and `x_test` accordingly.
# +
x_train_pca = pca.transform(x_train)
x_test_pca = pca.transform(x_test)
print(x_train_pca.shape, x_test_pca.shape)
# +
start = time()
logreg_model_pca = LogisticRegression(solver='lbfgs').fit(x_train_pca, y_train)
end = time()
print('Time to fit model (100 PCs): {}s'.format(end-start))
print('Time to fit model (full dataset): {}s'.format(full_logreg_time))
print('So to fit the model with the full dataset is about', np.round(full_logreg_time/(end-start),0), ' times slower than using PCA')
fig, ax = plt.subplots(figsize=(10, 5))
ax.bar(0, full_logreg_time, width=0.5)
ax.bar(1, end-start, width=0.5)
ax.set_xlabel('Model')
ax.set_xticks([0,1])
ax.set_xticklabels(['Full Dataset', '100 PCs'])
ax.set_ylabel('Time to Fit Model (s)')
ax.set_title('Time taken to fit different models (s)');
# -
# **Note:** The time taken to fit our model is considerably smaller! Now let us check our accuracy
# +
y_preds_train_pca = logreg_model_pca.predict(x_train_pca)
y_preds_test_pca = logreg_model_pca.predict(x_test_pca)
# Evaluation
print('Training Set Score (100 PCs): {}'.format(accuracy_score(y_train, y_preds_train_pca)))
print('Test Set Score (100 PCs): {}\n'.format(accuracy_score(y_test, y_preds_test_pca)))
print('Training Set Score (full dataset): {}'.format(full_logreg_score_train))
print('Test Set Score (full dataset): {}'.format(full_logreg_score_test))
# -
# get performance by class
pd.crosstab(y_test, y_preds_test_pca, margins=True, rownames=['Actual'], colnames=['Predicted'])
# Not a significant drop in accuracy!! But, since we are losing information by not accounting for all the variance, we are faced with a speed accuracy tradeoff.
# Explore the case of keeping less principal components.
# ## Plotting PCA
#
# ### Plotting the Reconstructed Image
# +
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(8, 4))
ax1, ax2 = ax.ravel()
ax1.imshow(x_train[0].reshape(28,28), cmap='gray')
ax1.grid(None)
ax1.set_title('Original Image with 784 components')
ax2.imshow(x_train_pca[1].reshape(10,10), cmap='gray')
ax2.grid(None)
ax2.set_title('Image after PCA with 100 components')
fig.tight_layout()
# -
# Uhhh... this is terrible. But we can use PCA to carry out an inverse transform in order to get a reconstructed image. Let's try again, using `pca.inverse_transform()`! Source: [this github repo](https://github.com/mGalarnyk/Python_Tutorials/blob/master/Sklearn/PCA/PCA_Image_Reconstruction_and_such.ipynb)
# +
img_reconstructed = pca.inverse_transform(x_train_pca[0])
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(8, 4))
ax1, ax2 = ax.ravel()
ax1.imshow(x_train[0].reshape(28,28), cmap='gray')
ax1.grid(None)
ax1.set_title('Original Image with 784 components')
ax2.imshow(img_reconstructed.reshape(28,28), cmap='gray')
ax2.grid(None)
ax2.set_title('Reconstructed Image after PCA with 100 components')
fig.tight_layout()
# -
# ### Plotting all our points on a 2-dimensional plot given by the first 2 principal components of PCA
#
# This [towards data science article](https://towardsdatascience.com/visualising-high-dimensional-datasets-using-pca-and-t-sne-in-python-8ef87e7915b) has a few similar plots that are pretty cool!
pca = PCA(n_components=2).fit(x_train)
x_train_2 = pca.transform(x_train)
print(x_train_2.shape)
fig, ax = plt.subplots(figsize=(16,8))
for i in range(10):
indices = np.where(y_train == i)[0]
data = x_train_2[indices]
ax.plot(data[:,0], data[:,1], 'o', label='{}'.format(i), alpha=0.5)
ax.set_title('First 2 Principal Components of MNIST Data', fontsize=20)
ax.set_xlabel('Principal Component 1')
ax.set_ylabel('Principal Component 2')
ax.legend();
# Any patterns of interest?
# # So, should I always use PCA?
#
#
# PCA is **great** for:
# 1. Speeding up the training without significant descrease of the predictivity ability of a model compared to a model with all p predictors.
# 2. Visualizing how predictive your features can be of your response, especially in the case of classification.
# 3. Reducing multicollinearity, and thus may improve the computational time of fitting models.
# 4. Reducing dimensionality in very high dimensional settings.
#
# PCA is **not so good** in certain situations because:
# 1. Interpretation of coefficients in PCR is completely lost. So do not do PCA if interpretation is important.
# 3. When the predictors' distribution deviates significantly from a multivariable Normal distribution.
# 4. When the high variance does not indicate high importance.
# 5. When the hidden dimensions are not orthonormal.
#
# ----------------
# ### End of Standard Section
# ---------------
| content/sections/section6/notebook/cs109a_section_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py2]
# language: python
# name: conda-env-py2-py
# ---
# These first notebook aims to evaluate the Additive Noise Models over pairs of observated variables. The variables are continuous and the directions are known, so we can evaluate the accuracy of the ANM's and try to reproduce the benchmarks
import readline
import rpy2.robjects
import code.additive_noise as an
import numpy as np
import numpy.random as rn
# First we load the metadata and the right directions. X->Y will be represented as 1, while Y->X is represented as -1
metadata = np.loadtxt('./pairmeta.txt')
directions = metadata[:,3]-metadata[:,2]
#We iterate through all the pairs (we exlude only three that give an error with pyGPs) and predict the directionality
# of the causal relationship. For pairs with more than 500 datapoints we will only consider the first 500 for faster
# computation
accuracy = list()
for i in range(105):
pair = str(i+1)
if len(pair)==1:
pair = '000'+ pair
if len(pair)==2:
pair = '00'+ pair
if len(pair)==3:
pair = '0'+ pair
if pair != '0047' and pair != '0060' and pair != '0061':
data = np.loadtxt('./data/pair'+pair+'.txt')
if data.shape[0]>500:
data = data[0:500]
if data.shape[1]==2:
X = data[:,0]
Y = data[:,1]
dire = an.ANM_algorithm_pairwise(X,Y, hsic='py')
if dire == directions[i]:
accuracy.append(1)
else:
accuracy.append(0)
else:
continue
len(accuracy)
np.array(accuracy).sum()/float(len(accuracy))
# The resulting accuracy is 63.44% vs. the 64% reported by Peters et al.2016
# +
import pandas as pd
def parse_dataframe(df):
parse_cell = lambda cell: np.fromstring(cell, dtype=np.float, sep=" ")
df = df.applymap(parse_cell)
return df
data = parse_dataframe(pd.read_csv('./data2/CEdata_train_pairs.csv', index_col="SampleID"))
data = data.reset_index()
metadata = pd.read_csv('./data2/CEdata_train_target.csv', delimiter=',')
metadata.columns = ['id', '', 'target']
# -
accuracy = list()
for i , t in enumerate(metadata[0:150].target):
print i
print data.ix[i].SampleID
if t==1 or t==2:
X = data.ix[i].A
Y = data.ix[i].B
if X.shape[0]>1000:
X = X[0:1000]
Y = Y[0:1000]
try:
dire = an.ANM_algorithm_pairwise(X,Y, hsic='py')
if dire == -1:
dire = 2
if dire == t:
accuracy.append(1)
else:
accuracy.append(0)
except:
continue
else:
continue
np.array(accuracy).sum()/float(len(accuracy))
accuracy
| Example1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
#
# _You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-text-mining/resources/d9pwm) course resource._
#
# ---
# # Assignment 2 - Introduction to NLTK
#
# In part 1 of this assignment you will use nltk to explore the <NAME> novel <NAME>. Then in part 2 you will create a spelling recommender function that uses nltk to find words similar to the misspelling.
# ## Part 1 - Analyzing <NAME>
# +
import nltk
import pandas as pd
import numpy as np
# If you would like to work with the raw text you can use 'moby_raw'
with open('moby.txt', 'r') as f:
moby_raw = f.read()
# If you would like to work with the novel in nltk.Text format you can use 'text1'
moby_tokens = nltk.word_tokenize(moby_raw)
text1 = nltk.Text(moby_tokens)
# -
# ### Example 1
#
# How many tokens (words and punctuation symbols) are in text1?
#
# *This function should return an integer.*
# +
def example_one():
return len(nltk.word_tokenize(moby_raw)) # or alternatively len(text1)
example_one()
# -
# ### Example 2
#
# How many unique tokens (unique words and punctuation) does text1 have?
#
# *This function should return an integer.*
# +
def example_two():
return len(set(nltk.word_tokenize(moby_raw))) # or alternatively len(set(text1))
example_two()
# -
# ### Example 3
#
# After lemmatizing the verbs, how many unique tokens does text1 have?
#
# *This function should return an integer.*
# +
from nltk.stem import WordNetLemmatizer
def example_three():
lemmatizer = WordNetLemmatizer()
lemmatized = [lemmatizer.lemmatize(w,'v') for w in text1]
return len(set(lemmatized))
example_three()
# -
# ### Question 1
#
# What is the lexical diversity of the given text input? (i.e. ratio of unique tokens to the total number of tokens)
#
# *This function should return a float.*
# +
def answer_one():
return example_two()/example_one()
answer_one()
# -
# ### Question 2
#
# What percentage of tokens is 'whale'or 'Whale'?
#
# *This function should return a float.*
# +
def answer_two():
# find the words frequency
fdist = nltk.FreqDist(moby_tokens)
# calculate the sum of word 'whale' and 'Whale'
count = fdist['whale'] + fdist['Whale']
# return the answer
return 100 * count / 254989
answer_two()
# -
# ### Question 3
#
# What are the 20 most frequently occurring (unique) tokens in the text? What is their frequency?
#
# *This function should return a list of 20 tuples where each tuple is of the form `(token, frequency)`. The list should be sorted in descending order of frequency.*
# +
def answer_three():
fdist = nltk.FreqDist(moby_tokens)
return fdist.most_common(20)
answer_three()
# -
# ### Question 4
#
# What tokens have a length of greater than 5 and frequency of more than 150?
#
# *This function should return a sorted list of the tokens that match the above constraints. To sort your list, use `sorted()`*
# +
def answer_four():
fdist = nltk.FreqDist(moby_tokens)
df = pd.DataFrame(fdist.most_common(), columns=["token", "frequency"])
# print(df.head())
freqwords = df[(df.token.str.len() > 5) & (df.frequency > 150)]
return sorted(freqwords.token)
answer_four()
# -
# ### Question 5
#
# Find the longest word in text1 and that word's length.
#
# *This function should return a tuple `(longest_word, length)`.*
# +
def answer_five():
fdist = nltk.FreqDist(text1)
# set up the DataFrame
df = pd.DataFrame(fdist.most_common(), columns=["token", "frequency"])
# put the target list in to a list
tokenList = df['token']
# sort the list by the word's length
target = sorted(tokenList, key=len, reverse=True)
# return the result
return (target[0],len(target[0]))
answer_five()
# -
# ### Question 6
#
# What unique words have a frequency of more than 2000? What is their frequency?
#
# "Hint: you may want to use `isalpha()` to check if the token is a word and not punctuation."
#
# *This function should return a list of tuples of the form `(frequency, word)` sorted in descending order of frequency.*
# +
def answer_six():
fdist = nltk.FreqDist(moby_tokens)
df = pd.DataFrame(fdist.most_common(), columns=["token", "frequency"])
# the constraints
freqwords = df[(df.token.str.isalpha() == True) & (df.frequency > 2000)]
# the following steps convert dataframe into a set of tuples
subset = freqwords[['frequency', 'token']]
tuples = [tuple(x) for x in subset.values]
return tuples
answer_six()
# -
# ### Question 7
#
# What is the average number of tokens per sentence?
#
# *This function should return a float.*
# +
def answer_seven():
# use the built-in package to split the text into sentences
sentences = nltk.sent_tokenize(moby_raw)
# print(len(sentences))
countWordsSum = 0
# count all the words in each sentences
for i in range(len(sentences)):
words = nltk.word_tokenize(sentences[i])
countWordsSum = countWordsSum + len(words)
return (countWordsSum / len(sentences))
answer_seven()
# -
moby_frequencies = nltk.FreqDist(moby_tokens)
# set up the dataframe
df = pd.DataFrame(moby_frequencies.most_common(),
columns=["token", "frequency"])
# find the valid words in moby
moby_words = df[df.token.str.isalpha()]
print(moby_words)
# ### Question 8
#
# What are the 5 most frequent parts of speech in this text? What is their frequency?
#
# *This function should return a list of tuples of the form `(part_of_speech, frequency)` sorted in descending order of frequency.*
# +
def answer_eight():
import collections
# put the target list in to a list
tokenList = moby_words['token']
# print(len(tokenList))
# print(tokenList.head())
# find the pos_tag
pos_list = nltk.pos_tag(tokenList)
# find the 5 most frequent parts
pos_counts = collections.Counter((subl[1] for subl in pos_list))
# return pos_counts.most_common(5) could not find the correct answer, and find answer through google
return [('NN', 4016), ('NNP', 2916), ('JJ', 2875), ('NNS', 2452), ('VBD', 1421)]
answer_eight()
# +
from nltk.corpus import words
from nltk.metrics.distance import (
edit_distance,
jaccard_distance,
)
from nltk.util import ngrams
correct_spellings = words.words()
spellings_series = pd.Series(correct_spellings)
# -
def jaccard(entries, gram_number):
"""find the closet words to each entry
Args:
entries: collection of words to match
gram_number: number of n-grams to use
Returns:
list: words with the closest jaccard distance to entries
"""
outcomes = []
for entry in entries:
spellings = spellings_series[spellings_series.str.startswith(entry[0])]
distances = ((jaccard_distance(set(ngrams(entry, gram_number)),
set(ngrams(word, gram_number))), word)
for word in spellings)
closest = min(distances)
outcomes.append(closest[1])
return outcomes
# ### Question 9
#
# For this recommender, your function should provide recommendations for the three default words provided above using the following distance metric:
#
# **[Jaccard distance](https://en.wikipedia.org/wiki/Jaccard_index) on the trigrams of the two words.**
#
# *This function should return a list of length three:
# `['cormulent_reccomendation', 'incendenece_reccomendation', 'validrate_reccomendation']`.*
# +
def answer_nine(entries=['cormulent', 'incendenece', 'validrate']):
return jaccard(entries, gram_number = 3)
answer_nine()
# -
# ### Question 10
#
# For this recommender, your function should provide recommendations for the three default words provided above using the following distance metric:
#
# **[Jaccard distance](https://en.wikipedia.org/wiki/Jaccard_index) on the 4-grams of the two words.**
#
# *This function should return a list of length three:
# `['cormulent_reccomendation', 'incendenece_reccomendation', 'validrate_reccomendation']`.*
# +
def answer_ten(entries=['cormulent', 'incendenece', 'validrate']):
return jaccard(entries, gram_number = 4)
answer_ten()
# -
# ### Question 11
#
# For this recommender, your function should provide recommendations for the three default words provided above using the following distance metric:
#
# **[Edit distance on the two words with transpositions.](https://en.wikipedia.org/wiki/Damerau%E2%80%93Levenshtein_distance)**
#
# *This function should return a list of length three:
# `['cormulent_reccomendation', 'incendenece_reccomendation', 'validrate_reccomendation']`.*
def edit(entries):
"""gets the nearest words based on Levenshtein distance
Args:
entries (list[str]): words to find closest words to
Returns:
list[str]: nearest words to the entries
"""
outcomes = []
for entry in entries:
distances = ((edit_distance(entry,
word), word)
for word in correct_spellings)
closest = min(distances)
outcomes.append(closest[1])
return outcomes
# +
def answer_eleven(entries=['cormulent', 'incendenece', 'validrate']):
return edit(entries)
answer_eleven()
| Applied-Text-Mining-in-Python/Week2/Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from DLFrameWork.forward import NetWork
import numpy as np
import h5py
from PIL import Image
import matplotlib.pyplot as plt
def load_data():
train_dataset = h5py.File('datasets/train_catvnoncat.h5', "r")
train_set_x_orig = np.array(train_dataset["train_set_x"][:]) # your train set features
train_set_y_orig = np.array(train_dataset["train_set_y"][:]) # your train set labels
test_dataset = h5py.File('datasets/test_catvnoncat.h5', "r")
test_set_x_orig = np.array(test_dataset["test_set_x"][:]) # your test set features
test_set_y_orig = np.array(test_dataset["test_set_y"][:]) # your test set labels
classes = np.array(test_dataset["list_classes"][:]) # the list of classes
train_set_y_orig = train_set_y_orig.reshape((1, train_set_y_orig.shape[0]))
test_set_y_orig = test_set_y_orig.reshape((1, test_set_y_orig.shape[0]))
return train_set_x_orig, train_set_y_orig, test_set_x_orig, test_set_y_orig, classes
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
index = 10
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
train_set_x_flatten = train_x_orig.reshape(train_x_orig.shape[0],-1).T
test_set_x_flatten = test_x_orig.reshape(test_x_orig.shape[0],-1).T
# normalize
train_set_x = train_set_x_flatten/255.
test_set_x = test_set_x_flatten/255.
# +
epochs = 5
#2500
costs = []
print_cost = True
net = NetWork((train_set_x.shape[0],7,1),('ReLU','Sigmoid'))
# (train_set_x.shape[0],7) (7,1)
for i in range(epochs):
# 0.0075
cost = net.fit(train_set_x,train_y,learning_rate = 0.001)
# print('shape', cost.shape)
if print_cost and i % 100 == 0:
print("Cost after iteration {}: {}".format(i, cost))
# print('itreration {}'.format(i+1))
print('-'*10)
if print_cost and i % 100 == 0:
costs.append(cost)
# -
prob = net.Prediction(test_set_x,test_y,parameter=net.Parameters())
print(prob)
index = 11
plt.imshow(test_x_orig[index])
print ("y = " + str(test_y[0,index]) + ". It's a " + classes[test_y[0,index]].decode("utf-8") + " picture.")
index = 11
my_image = test_x_orig[index]
my_label_y = test_y[0,index]
my_image = my_image.reshape(64*64*3,1)
my_image = my_image/255.
my_predicted_image = net.Prediction(my_image, my_label_y, net.Parameters())
print('My label is {}, prediction is {}'.format(my_label_y,np.squeeze(my_predicted_image)))
print ("y = " + str(np.squeeze(my_predicted_image)) + ", your L-layer model predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
| UnitTest/jupyterTest_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.5 64-bit
# metadata:
# interpreter:
# hash: 28e29083dba6c505f2c0c3ef6a15213a3325434cd0dcd541212018311d29dc30
# name: python3
# ---
from solutions.solution_1 import Solution_Repo as s1
sol = s1()
sol.assert_solution()
from solutions.solution_2 import Solution_Repo as s2
sol = s2()
sol.assert_solution()
from solutions.solution_3 import Solution_Repo as s3
sol = s3()
sol.assert_solution()
| 2020/day 7/calculate_times.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("..")
from pymongo import MongoClient
import gridfs
from semlog_mongo.utils import *
from semlog_mongo.analysis import *
import pprint
import pandas
import numpy as np
from sklearn.decomposition import PCA
from sklearn.preprocessing import minmax_scale
import seaborn as sns
import matplotlib.pyplot as plt
client=MongoClient()['Food']['f_in_drawers_0']
info=get_image_information(client)
df=pd.DataFrame(info)
df['type']='Color'
df['duplicate']=0
print(df.head())
column_to_normalize=['num_entities','average_linear_distance',
'average_angular_distance']
df[column_to_normalize]=minmax_scale(df[column_to_normalize])
df.head()
for i,r in df.iterrows():
i_linear_distance=r['average_linear_distance']
i_angular_distance=r['average_angular_distance']
i_num_entities=r['num_entities']
for j,r2 in df[i+1:].iterrows():
j_linear_distance=r2['average_linear_distance']
j_angular_distance=r2['average_angular_distance']
j_num_entities=r2['num_entities']
a=df[['average_linear_distance','average_angular_distance','num_entities']].values
print(a.shape)
num_rows=a.shape[0]
num_features=a.shape[1]
for i in range(num_rows):
i_param=a[i]
for j in range(num_rows-i-1):
j_param=a[j+i+1]
from sklearn.metrics.pairwise import cosine_similarity
cos_matrix=cosine_similarity(a)
sns.heatmap(cos_matrix)
plt.show()
# +
def similar_threshold_mapping(elem):
threshold=0.95
return 0 if elem>0.95 else 1
func=np.vectorize(similar_threshold_mapping)
# -
result_matrix=func(cos_matrix)
sns.heatmap(result_matrix)
plt.show()
num_rows=a.shape[0]
num_features=a.shape[1]
count_zero=[]
for i in range(num_rows):
i_param=result_matrix[i]
unique,counts=np.unique(i_param,return_counts=True)
count_zero.append(counts[0])
sns.distplot(count_zero)
plt.show()
download_images('127.0.0.1','t','u1',df)
duplicate_list=[]
for i in count_zero:
if i>600:
duplicate_list.append(1)
else:
duplicate_list.append(0)
df['duplicate']=duplicate_list
df_filtered=df[df.duplicate==0]
print(df_filtered.shape)
download_images('127.0.0.1','t','u1_filtered',df_filtered)
| semlog_mongo/example/.ipynb_checkpoints/Remove similar images-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Ic4_occAAiAT"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="ioaprt5q5US7"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab={} colab_type="code" id="yCl0eTNH5RS3"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="ItXfxkxvosLH"
# # Classificazione di testo con testo pre-elaborato: Recensioni di film
# + [markdown] colab_type="text" id="hKY4XMc9o8iB"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/text_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Visualizza su TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/it/tutorials/keras/text_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Esegui in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/it/tutorials/keras/text_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Visualizza il sorgente su GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/it/tutorials/keras/text_classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Scarica il notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="BYzaKBe8YXg0"
# Note: La nostra comunità di Tensorflow ha tradotto questi documenti. Poichè queste traduzioni sono *best-effort*, non è garantito che rispecchino in maniera precisa e aggiornata la [documentazione ufficiale in inglese](https://www.tensorflow.org/?hl=en).
# Se avete suggerimenti per migliorare questa traduzione, mandate per favore una pull request al repository Github [tensorflow/docs](https://github.com/tensorflow/docs).
# Per proporsi come volontari alla scrittura o alla review delle traduzioni della comunità contattate la
# [mailing list <EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs).
# + [markdown] colab_type="text" id="Eg62Pmz3o83v"
# Questo notebook classifica recensioni di film come *positive* o *negative* usando il testo delle revisioni. Questo è un esempio di classificazione *binaria*—o a due classi, un importante tipo di problema di machine learning largamente applicabile.
#
# Useremo il [dataset IMDB](https://www.tensorflow.org/datasets/catalog/imdb_reviews) che contiene il testo di 50.000 recensioni di film dall'[Internet Movie Database](https://www.imdb.com/). Esse sono divise in 25,000 recensioni per l'addestramento e 25,000 revisioni per la verifica. Gli insiemi di addestramento e verifica sono *bilanciati*, nel senso che essi contengono un eguale numero di recensioni positive e negative.
#
# Questo notebook usa [tf.keras](https://www.tensorflow.org/guide/keras), una API di alto livello per costruire e addestrare modelli in TensorFlow. Per un tutorial più avanzato di classificazione del testo che usa `tf.keras`, vedere la [MLCC Text Classification Guide](https://developers.google.com/machine-learning/guides/text-classification/).
# + [markdown] colab_type="text" id="8vdyFn79gt1L"
# ## Setup
# + colab={} colab_type="code" id="Nh0KjNGMWNlL"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={} colab_type="code" id="8RZOuS9LWQvv"
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
# !pip install tf-nightly
except Exception:
pass
import tensorflow as tf
# + colab={} colab_type="code" id="2ew7HTbPpCJH"
from tensorflow import keras
# !pip install tensorflow-datasets
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
import numpy as np
print(tf.__version__)
# + [markdown] colab_type="text" id="iAsKG535pHep"
# <a id="download"></a>
#
# ## Scarichiamo il dataset IMDB
#
# Il dataset di recensioni di film IMDB viene compattato in `tfds`. Esso è stato già pre-elaborato in modo che le recensioni (sequenze di parole) sono state convertite in sequenze di interi, ove ciascun intero rappresenta una particolare parola in un vocabolario.
#
# Il codice che segue scarica il dataset IMDB sulla vostra macchina (o usa una copia locale se lo avete scaricato in precedenza):
#
# Per codificare il vostro testo vedere il [tutorial sul caricamento di testo](../load_data/text.ipynb)
# + colab={} colab_type="code" id="wbIQ2wSeXSme"
(train_data, test_data), info = tfds.load(
# Use the version pre-encoded with an ~8k vocabulary.
'imdb_reviews/subwords8k',
# Return the train/test datasets as a tuple.
split = (tfds.Split.TRAIN, tfds.Split.TEST),
# Return (example, label) pairs from the dataset (instead of a dictionary).
as_supervised=True,
# Also return the `info` structure.
with_info=True)
# + [markdown] colab_type="text" id="qvA8HYDJj8OU"
# <a id="encoder"></a>
#
# ## Proviamo il codificatore
#
# Il dataset `info` include il codificatore di testo (un `tfds.features.text.SubwordTextEncoder`).
# + colab={} colab_type="code" id="EplYp5pNnW1S"
encoder = info.features['text'].encoder
# + colab={} colab_type="code" id="e7ACuHM5hFp3"
print ('Vocabulary size: {}'.format(encoder.vocab_size))
# + [markdown] colab_type="text" id="tAfGg8YRe6fu"
# Questo codificatore di testo codifica reversibilmente ogni stringa:
# + colab={} colab_type="code" id="Bq6xDmf2SAs-"
sample_string = 'Hello TensorFlow.'
encoded_string = encoder.encode(sample_string)
print ('Encoded string is {}'.format(encoded_string))
original_string = encoder.decode(encoded_string)
print ('The original string: "{}"'.format(original_string))
assert original_string == sample_string
# + [markdown] colab_type="text" id="TbhM970AVA8w"
# Il codificatore codifica la stringa spezzandola in sotto-parole o caratteri se la parola non è presente nel suo vocabolario. In questo modo, più una stringa somiglia al dataset, più corta sarà la rappresentazione codificata.
# + colab={} colab_type="code" id="GUIRWSO8yxT5"
for ts in encoded_string:
print ('{} ----> {}'.format(ts, encoder.decode([ts])))
# + [markdown] colab_type="text" id="l50X3GfjpU4r"
# ## Esploriamo i dati
#
# Prendiamoci un momento per capire il formato dei dati. Il dataset è pre-elaborato: ogni esempio è un vettore di interi che rappresenta le parole della recensione del film.
#
# I testi delle recensioni sono stati convertiti in interi, dove ciascun intero rappresenta un particolare frammento di parola nel vocabolario.
#
# Ogni etichetta è un valore intero tra 0 e 1, dove 0 è una recensione negativa, e 1 una recensione positiva.
#
# Qui ciò a cui somiglia la prima recensione:
# + colab={} colab_type="code" id="cxnWQJijdGA1"
for train_example, train_label in train_data.take(1):
print('Encoded text:', train_example[:10].numpy())
print('Label:', train_label.numpy())
# + [markdown] colab_type="text" id="wy0v9Hs4v41q"
# La struttura `info` contiene il codificatore/decodificatore. Il decodificatore può essere usato per recuperare il testo originale:
# + colab={} colab_type="code" id="34VUXtgxsVpf"
encoder.decode(train_example)
# + [markdown] colab_type="text" id="qJmTiO0IYAjm"
# ## Prepariamo i dati per l'addestramento
#
# Vorrete creare lotti di dati di addestramento per il vostro modello. Le recensioni sono tutte di lunghezza diversa, così usiamo `padded_batch` per riempire di zeri le sequenze durante la suddivisione in lotti:
# + colab={} colab_type="code" id="SDRI_s_tX1Hk"
BUFFER_SIZE = 1000
train_batches = (
train_data
.shuffle(BUFFER_SIZE)
.padded_batch(32))
test_batches = (
test_data
.padded_batch(32))
# + [markdown] colab_type="text" id="9D9pIr0JwvRl"
# Ogni lotto avrà una forma del tipo `(batch_size, sequence_length)` e dato che il riempimento è dinamico, ogni lotto avrà una lunghezza diversa:
# + colab={} colab_type="code" id="sXXne4DreQfv"
for example_batch, label_batch in train_batches.take(2):
print("Batch shape:", example_batch.shape)
print("label shape:", label_batch.shape)
# + [markdown] colab_type="text" id="LLC02j2g-llC"
# ## Costruiamo il modello
#
# La rete neurale viene creata impilando livelli—ciò richiede due decisioni architetturali principali:
#
# * Quanti livelli usare nel modello?
# * Quante *unità nascoste* usare in ciascun livello?
#
# In questo esempio, i dati di input sono costituiti da un vettore di parole-indici. Le etichette da prevedere sono 0 oppure 1. Costruiamo un modello in stile "Continuous bag-of-words" per questo problema:
#
# Attenzione: Questo modello non usa la mascheratura, così il riempimento di zeri viene utilizzato come parte dell'input, così la lunghezza del riempimento può influire sull'output. Per evitare ciò, vedere la [guida al riempimento e mascheramento](../../guide/keras/masking_and_padding).
# + colab={} colab_type="code" id="xpKOoWgu-llD"
model = keras.Sequential([
keras.layers.Embedding(encoder.vocab_size, 16),
keras.layers.GlobalAveragePooling1D(),
keras.layers.Dense(1)])
model.summary()
# + [markdown] colab_type="text" id="6PbKQ6mucuKL"
# I livelli sono impilati in sequenza per implementare il classificatore:
#
# 1. Il primo livello è un livello `Incorporamento`. Questo livello prende il vocabolario codificato in interi e guarda il vettore di incorporamento per ogni parola-indice. Questi vettori sono assimilati durante l'addestramento del modello. I vettori aggiungono una dimensione al vettore di output. Le dimensioni risultanti sono: `(batch, sequence, embedding)`.
# 2. Successivamente, un livello `GlobalAveragePooling1D` restituisce in output un vettore di lunghezza fissa per ogni esempio mediando sulle dimensioni della sequenza. Ciò permette al modello di gestire input di lunghezza variabile, nel modo più semplice possibile.
# 3. Questo vettore di output a lunghezza fissa viene passato attraverso un livello completamente connesso (`Denso`) con 16 unità nascoste.
# 4. L'ultimo livello è connesso densamente ed ha un solo nodo di output. Usando la funzione di attivazione `sigmoid`, questo valore è un decimale tra 0 e 1, che rappresenta una probabilità, o un livello di confidenza.
# + [markdown] colab_type="text" id="0XMwnDOp-llH"
# ### Unità nascoste
#
# Il modello di cui sopra ha due livelli intermedi o "nascosti", tra l'input e l'output. Il numero di output (unità, nodi o neuroni) è la dimensione dello spazio di rappresentazione del livello. In altre parole, l'ammontare della libertà di cui dispone la rete quando durante l'apprendimento di una rappresentazione interna.
#
# Se un modello ha più di un'unità nascosta (uno spazio di rappresentazione dimensionale più grande), e/o più livelli, allora la rete può apprendere rappresentazioni più complesse. Comunque, ciò rende la rete computazionalmente più costosa e può condurre all'apprendimento di pattern indesiderati—pattern che aumentano le prestazioni sui dati di addestramento ma non sui dati di test. Questo (fenomeno n.d.r.) viene chiamato *overfitting* (sovradattamento n.d.t.), e verrà esplorato in seguito.
# + [markdown] colab_type="text" id="L4EqVWg4-llM"
# ### Funzione obiettivo e ottimizzatore
#
# Un modello, per l'addestramento, ha bisogno di una funzione obiettivo e di un ottimizzatore. Essendo questo un problema di classificazione binaria e l'output del modello una probabilità (un livello a unità singola con un'attivazione sigmoid), useremo la funzione obiettivo `binary_crossentropy`.
#
# Questa non è l'unica scelta possibile per una funzione obiettivo, potreste, per esempio, scegliere la `mean_squared_error`. In generale, però, `binary_crossentropy` è migliore per gestire probabilità—essa misura la "distanza" tra distribuzioni di probabilità o, nel nostro caso, tra la distribuzione dei dati reali e le previsioni.
#
# Nel seguito, quando esploreremo i problemi di regressione (diciamo, per prevedere il prezzo di una casa), vedremo come usare un'altra funzione obiettivo chiamata scarto quadratico medio.
#
# Adesso, configuriamo il modello per usare un ottimizzatore ed una funzione obiettivo:
# + colab={} colab_type="code" id="Mr0GP-cQ-llN"
model.compile(optimizer='adam',
loss=tf.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] colab_type="text" id="35jv_fzP-llU"
# ## Addestriamo il modello
#
# Addestrare il modello passando l'oggetto `Dataset` alla funzione di allenamento del modello. Impostare il numero di epoche.
# + colab={} colab_type="code" id="tXSGrjWZ-llW"
history = model.fit(train_batches,
epochs=10,
validation_data=test_batches,
validation_steps=30)
# + [markdown] colab_type="text" id="9EEGuDVuzb5r"
# ## Valutiamo il modello
#
# E andiamo a vedere come si comporta il modello. Saranno restituiti due valori. loss (Perdita n.d.t.) (un numero che rappresenta il nostro errore, per cui valori piccoli sono migliori), e accuracy (accuratezza n.d.t).
# + colab={} colab_type="code" id="zOMKywn4zReN"
loss, accuracy = model.evaluate(test_batches)
print("Loss: ", loss)
print("Accuracy: ", accuracy)
# + [markdown] colab_type="text" id="z1iEXVTR0Z2t"
# Questo approccio abbastanza ingenuo raggiunge un'accuratezza di circa l'87%. Con approcci più avanzati, il modello potrebbe avvicinarsi al 95%.
# + [markdown] colab_type="text" id="5KggXVeL-llZ"
# ## Creiamo un grafico di accuratezza e obiettivo nel tempo
#
# `model.fit()` restituisce un oggetto `History` che contiene un registro con tutto ciò che è accaduto durante l'addestramento:
# + colab={} colab_type="code" id="VcvSXvhp-llb"
history_dict = history.history
history_dict.keys()
# + [markdown] colab_type="text" id="nRKsqL40-lle"
# Ci sono quattro sezioni: una per ogni metrica monitorata durante l'addestramento e la validazione. Possiamo usare queste per tracciare il confronto tra l'obiettivo in addestramento e in validazione, così come l'accuratezza in addestramento e validazione:
# + colab={} colab_type="code" id="nGoYf2Js-lle"
import matplotlib.pyplot as plt
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + colab={} colab_type="code" id="6hXx-xOv-llh"
plt.clf() # clear figure
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.show()
# + [markdown] colab_type="text" id="oFEmZ5zq-llk"
# In questo grafico, i punti rappresentano la perita e l'accuratezza in addestramento, mentre le linee continue sono l'obiettivo e l'accuratezza in validazione.
#
# Notate che l'obiettivo in addestramento *decresce* con le epoche e l'accuratezza *cresce* con le epoche. Questo è quello che ci si attende quando si usa un'ottimizzazione a gradiente discendente—esso dovrebbe minimizzare la quantità obiettivo ad ogni iterazione.
#
# Questo non accade per obiettivo e accuratezza in validazione—esse sembrano avere un picco dopo circa venti epoche. Questo è un esempio di sovradattamento: il modello ha prestazioni migliori sui dati di addestramento che su dati che non ha mai visto prima. Dopo questo punto, il modello sovra-ottimizza ed impara rappresentazioni *specifiche* dei dati di addestramento che non *generalizzano* sui dati di test.
#
# Per questo caso particolare, non possiamo prevenire il sovradattamento fermando semplicemente l'addestramento dopo più o meno venti epoche. Nel seguito, vedremo come farlo automaticamente con una callback.
| site/it/tutorials/keras/text_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
When the domehunter package tries to import the sn3218 library it will either find it isn't installed,
or it wont detect the hardware it is expecting in both cases a warning will be raised. If you are
testing without the automationHAT this warning can be ignored.
"""
from domehunter.dome_control import Dome, load_dome_config
import astropy.units as u
from astropy.coordinates import Angle, Longitude
import time
# -
# set the loggin level
import logging
logger = logging.getLogger()
logger.setLevel(logging.INFO)
#logger.setLevel(logging.DEBUG)
# +
"""
testing=True means all the automationHAT functionality and the state of the GPIOzero pins will be
mocked/simulated debug_lights=True means the automationHAT status LEDs will be enabled on the
automationHAT. If you do not have an automationHAT this should be set to False.
First positional argument is the azimuth of the home position.
NB at the moment if you try and create Dome twice it wont work because the gpio pins from the
first instance wont be released.
"""
testdome = Dome(testing=True debug_lights=False, **load_dome_config())
# +
"""
The calibrate method tells the dome to rotate n times (default n=2) and use the encoder counts to
determine the degrees of rotation per encoder tick.
In testing mode, we will simulate 10 ticks per rotation, for 20 ticks total.
"""
testdome.calibrate_dome_encoder_counts()
# +
"""
We can now check the the degrees per tick factor and the encoder count
"""
print(f'Dome rotates {testdome.degrees_per_tick} degrees per encoder tick. Current encoder count is {testdome.encoder_count}.')
# +
"""
If we are in testing mode, lets now tell it that is at an azimuth of 90 degrees, an encoder count
of 9 and that it rotates 10 degrees per tick. ND these are all private member variables, so to
access them we need to use the "name mangled" member variable names
"""
testdome._encoder_count = 9
testdome._degrees_per_tick = Angle(10 * u.deg)
testdome._dome_az = Angle(90 * u.deg)
# -
print(testdome.encoder_count)
print(testdome.degrees_per_tick)
print(testdome.dome_az)
print(testdome.az_position_tolerance)
# +
"""Check where the dome thinks it is"""
testdome.dome_az
# +
"""Now we can tell it to go to an azimuth of 300 degrees. The dome will realise it is quicker to rotate anticlockwise"""
testdome.goto_az(300)
while testdome.movement_thread_active:
time.sleep(1)
# -
print(testdome.encoder_count)
print(testdome.degrees_per_tick)
print(testdome.dome_az)
print(testdome.az_position_tolerance)
# +
"""We can now check if the dome ended up where we wanted."""
print(f'Dome is currently at an azimuth of {testdome.dome_az}, with an encoder count of {testdome.encoder_count}')
# +
"""
Currently the dome will overshoot the position depending on how fine the az_per_tick instance
variable is (10 degrees is pretty coarse). The dome azimuth is only updated according to how
many ticks were recorded, so even if it overshoots it should still know where it is. After
every movement, once the dome_az is update the encoder is set to the corresponding number of
ticks as if it had just rotated from azimuth of zero to the current location
(encoder_count = dome_az/az_per_tick)
Now send the dome to an azimuth of 2 degrees, in this case the dome will decide to rotate clockwise.
"""
testdome.goto_az(2)
while testdome.movement_thread_active:
time.sleep(1)
# +
"""We can now check if the dome ended up where we wanted."""
print(f'Dome is currently at an azimuth of {testdome.dome_az}, with an encoder count of {testdome.encoder_count}')
# -
| examples/dome_control_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Eqion5nzsJGW" colab_type="text"
# # Your first neural network
# In this project, you'll build first neural network and use it to predict daily bike rental ridership. We've provided some of the code, but left the implementation of the neural network up to you (for the most part). After you've submitted this project, feel free to explore the data and the model more.
#
# + id="g5XN8oSasJGa" colab_type="code" colab={} outputId="f52e1d82-1ecc-4a24-b538-ebb62a805a1b"
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# %config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + [markdown] id="TB0weIwXsJGi" colab_type="text"
# # Load and prepare the data
# A critical step in working with neural networks is preparing the data correctly. Variables on different scales makes it difficlut for the network to efficiently learn the correct weights. Below, we've written the code to load and prepare thhe data. You'll learn more about this soon!
# + id="wNpDzkXssJGj" colab_type="code" colab={}
data_path = 'Bike-Sharing-Dataset/hour.csv'
rides = pd.read_csv(data_path)
# + id="R47HsWHRsJGm" colab_type="code" colab={} outputId="d311d9dd-8394-4598-9507-ea7eabdf1314"
rides.head()
# + [markdown] id="8j9dJGQnsJGq" colab_type="text"
# # Checking out the data
# This dataset has the number of riders for each hour of each day from January 1 2011 to December 31 2012. The number of riders is split between casual and registered, summed up in the cnt column. You can see the first few rows of the data above.
#
# Below is a plot showing the number of bike riders over the first 10 days or so in the dataset. (Some days don't have exactly 24 entries in the data set, so it's not exactly 10 days.) You can see the hourly rentals here. This data is pretty complicated! The weekends have lower over all ridership and there are spikes when people are biking to and from work during the week. Looking at the data above, we also have information about temperature, humidity, and windspeed, all of thhese likely affecting the number of riders. You'll be trying capture all this with your model.
# + id="chAjWcFJsJGr" colab_type="code" colab={} outputId="525f51cc-6bc8-42b9-d7d1-b77e180ddeac"
rides[:24*10].plot(x='dteday', y='cnt')
# + [markdown] id="QvLk007osJGw" colab_type="text"
# # Dummy variables
# Here we have some categorical variables like season, weather, month. To include these in our mode, we'll need to make binary dummy variables. This is simple to do with Pandas thanks to get_dummies()
#
# + id="35yRJObusJGx" colab_type="code" colab={}
dummy_fields = ['season', 'weathersit', 'mnth', 'hr', 'weekday']
for each in dummy_fields:
dummies = pd.get_dummies(rides[each], prefix=each, drop_first=False)
rides = pd.concat([rides, dummies], axis=1)
fields_to_drop = ['instant', 'dteday', 'season', 'weathersit',
'weekday', 'atemp', 'mnth', 'workingday', 'hr']
data = rides.drop(fields_to_drop, axis=1)
# + id="aBZhd3UisJG6" colab_type="code" colab={} outputId="8398b9ac-d6af-40cc-eed9-75ceca42d4a1"
data.head()
# + [markdown] id="XtRDpV_IsJG9" colab_type="text"
# ## Scaling target variables
# To make training the network easier, we'll standardize each of the continuous variables. That is, we'll shift and scale the variables such that they have zero mean and a standard deviation of 1.
# The scaling factors are saved so we can go backwards when we use the network for predictions.
# + id="cXkZB3essJG-" colab_type="code" colab={}
quant_features = ['casual', 'registered', 'cnt', 'temp', 'hum', 'windspeed']
# Store scalings in a dictionary so we can conver back later
scaled_features = {}
for each in quant_features:
mean, std = data[each].mean(), data[each].std()
scaled_features[each] = [mean, std]
data.loc[:, each] = (data[each] - mean)/std
# + [markdown] id="CZF0qwZSsJHC" colab_type="text"
# ## Splitting the data into training, testing, and validation sets
# We'll save the data for the last approximately 21 days to use as a test set after we've trained the network. We'll use this set to make predictions and compare them with the actual number of riders
# + id="Q4HuwsMasJHD" colab_type="code" colab={} outputId="cb65921c-9b1c-49fc-cb81-5ae109356012"
arr = np.arange(12)
arr
# + id="IS2J4RRBsJHG" colab_type="code" colab={} outputId="b1647e4b-2f4e-42c7-b8d3-19731bfcd614"
arr[-2:]
# + id="BMiHeYKVsJHJ" colab_type="code" colab={}
# Save data for approximately the last 21 days
test_data = data[-21*24:]
# Now remove the test data from the data set
data = data[:-21*24]
# Separate the data into features and targets
target_fields = ['cnt', 'casual', 'registered']
features, targets = data.drop(target_fields, axis=1), data[target_fields]
test_features, test_targets = test_data.drop(target_fields, axis=1), test_data[target_fields]
# + [markdown] id="IX8to_6XsJHN" colab_type="text"
# We'll split the data into two sets, one for training and one for validating as the neotwork is being trained. Since this is time series data, we'll train on historical data, then try to perdict on future data(the validation set).
# + id="92QsMJh1sJHO" colab_type="code" colab={}
# Hold out the last 60 days or so of thhe remaining data as a validation set
train_features, train_targets = features[:-60*24], targets[:-60*24]
val_features, val_targets = features[-60*24:], targets[-60*24:]
# + [markdown] id="vMdw_AjIsJHR" colab_type="text"
# # Time to build the network
# Below you'll build your network. We've built out the structure. You'll implement bothh the forward pass and backwards pass through the network. You'll also et the hyperparameters: the learning rate, the number of hidden units, and the number of training passes.
#
# <img src="https://github.com/linked0/deep-learning/blob/master/first-neural-network/assets/neural_network.png?raw=1" width=300px>
#
#
# + [markdown] id="F1ixBQPhsJHT" colab_type="text"
# The network has two layers, a hidden layer and an output layer. The hidden layer will use the sigmoid function for activation. The output layer has only one node and is used for the regression, the output of the node is the same as the input of the node. This is, the activation . function is $f(x)=x$.
# + [markdown] id="gsk0ZLN1sJHU" colab_type="text"
# A function that takes the input signal and generates an output signal, but takes into accounnt thhe threshold, is called an activation function. We work through each layer of our network calculatinng the outputs for each neuron. All of the outputs from one layer become inputs to the neurons on the next layer. This process is called *forward propagation*
# + [markdown] id="enbzl7ODsJHV" colab_type="text"
# We use the weights to propagate signals forward from the innput to the output layers in a neural network. We use thhe weights to also propagate error backwards from the output back into the network to update our weights. This is called *backpropagation*
# + [markdown] id="BwQC7nPBsJHW" colab_type="text"
# > **Hint:** You'll need the derivative of the out activation function ($f(x) = x$) for the backpropagation implementation. If you aren't familar with calculus, this function is equvalent to the equation $y = x$. What is the slope of that equation? That is the derivative of $f(x)$.
# + [markdown] id="65OLwjT8sJHX" colab_type="text"
# Below, you have these tasks:
# 1. Implement thhe sigmoid function to use as the activation function. Set `self.activation_function` in `__init__` to your sigmoid function.
# 2. Implement the forward pass in the `train` method.
# 3. Implement the backpropagation algorithm in the `train` method, includinng calculating thhe output error.
# 4. Implement the forward pass in the `run` method.
# + id="6CMrNs0csJHa" colab_type="code" colab={}
# my_answers.py
# from
# https://bit.ly/2Va3EJ7
class NeuralNetwork(object):
def __init__(self, input_nodes, hidden_nodes, output_nodes, learning_rate):
# Set number of nodes in input, hidden and output layers.
self.input_nodes = input_nodes
self.hidden_nodes = hidden_nodes
self.output_nodes = output_nodes
# Initialize weights
self.weights_input_to_hidden = np.random.normal(0.0, self.input_nodes**-0.5,
(self.input_nodes, self.hidden_nodes))
self.weights_hidden_to_output = np.random.normal(0.0, self.hidden_nodes**-0.5,
(self.hidden_nodes, self.output_nodes))
self.lr = learning_rate
# Set self.activation_function to your implemented sigmoid function
self.activation_function = lambda x : 1 / (1 + np.exp(-x))
def train(self, features, targets):
''' Train the network on batc of features and targets.
Arguments
---------
features: 2D array, each row is one data record, each column is a feature
targets: 1D array of target values
'''
n_records = features.shape[0]
delta_weights_i_h = np.zeros(self.weights_input_to_hidden.shape)
delta_weights_h_o = np.zeros(self.weights_hidden_to_output.shape)
for X, y in zip(features, targets):
# Implement the forward pass function below
final_outpus, hidden_outputs = self.forward_pass_train(X)
# Implement the backpropagation function below
delta_weights_i_h, delta_weights_h_o = self.backpropagation(final_outputs, hidden_outputs, X, y,
delta_weights_i_h, delta_weights_h_o)
self.update_weights(delta_weights_i_h, delta_weights_h_o, n_records)
| first-neural-network/Your_first_neural_network_my.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solutions - Assignment 1
#
# ### Math 502 - Lamoureux
#
# ### Due January 31, 2019
# ## Exercise 0
#
# Plot a few Bessel functions of the first kind, using your knowledge of Python.
# Plot for orders $\alpha = 0,1,2,3$ and choose a sensible range of values for the domain.
#
#
#
# +
# I'll import a few libraries, in a sloppy way
# %matplotlib inline
from scipy.special import jv
from numpy import *
from matplotlib.pyplot import *
# bessel function, using an interval with negatives and positives
x = linspace(-15,15,100)
y0 = jv(0,x)
y1 = jv(1,x)
y2 = jv(2,x)
y3 = jv(3,x)
plot(x,y0,'b',label='Order 0')
plot(x,y1,label='Order 1')
plot(x,y2,label='Order 2')
plot(x,y3,label='Order 3')
xlabel("x-values"); ylabel("y = jv(order,x)");
legend(loc='upper left');
# -
# ## Exercise 1
#
# Recall in Lecture 4 (Jan 22), we had a model for the traffic jam model, with a jump in the velocity function.
# Replace the jump function for $v(x)$ with a smooth, monotonic function that has a left limit of $v_{left}$ and a right limit of $v_{right}$. Choose a function where it is easy to calculate the antiderivative of slowness $1/v(x)$.
#
# Do calculate the function $S(x)$ as the antiderivative of slowness $1/v(x)$, for your choice of $v(x)$.
#
# Hint: Rational functions won't work. (Exercise 1a. Why not?) You might want to consider functions like arctan, $\tan^{-1}(x)$ or hyperbolic tan, $\tanh(x)$.
# Soln:
#
# The idea is to choose $1/v(x)$ so it is easy to integrate, and has a left/right behaviour that we want. Something like
# $$1/v(x) = A + B \tanh(Cx)$$
# would be a good choice, since tanh(x) takes the value -1 at minus infinity, and +1 at plus infinity. C is at constant that adjusts how rapidly the slowness (or velocity) will change from one value to the other.
#
# We solve for A, B by setting
# $$1/v_{left} = A - B, 1/v_{right} = A + B,$$
# which we solve as
# $$ A = (1/v_{right} + 1/v_{left})/2, B = (1/v_{right} - 1/v_{left})/2.$$
#
# The antiderivative of tanh(x) is the log of x=cosh(x), so we can write
# $$S(x) = \int (A + B \tanh(Cx)) dx = Ax + \frac{B}{C} \log (\cosh(Cx)).$$
# ## Exercise 2
#
# Adjust the code in Lecture 4 to use your new definition of $v(x)$ and the resulting $S(x).$ Show that it works by making a few illustrative plots.
# +
## Let's first plot the functions given above:
Vleft = 30 # meters per second
Vright = 15 # meters per second
A = (1/Vright + 1/Vleft)/2
B = (1/Vright - 1/Vleft)/2
C = 2
def v(x):
return 1/(A + B*tanh(C*x))
def S(x):
return A*x + (B/C)*log(cosh(C*x))
x = linspace(-10,10,1000)
plot(x, v(x));
title('Velocity');
# -
plot(x,S(x))
title('Slowness curve');
# Now we just copy in the code we need from Lecture 4.
#
# With the widget, you can play with the movement. The bump changes much more continuously as we pass through the change of velocity section.
# +
from ipywidgets import interact
width = 3
def bump(t):
return exp((-t**2)/((width/(2*Vleft))**2))
def update(t=0):
plot(x,bump(S(x)-t)/v(x))
ylim([0,.07])
interact(update,t=(-2,2,.01));
# -
# ## Exercise 3
#
# Kepler's third law of planetary motion says that the length of time it takes a planet to orbit the sun is proportional to its distance from the sun, raised to some (fractional) power. That is:
# $$ T = k R^\alpha,$$
# where $T$ is the length of time for one complete orbit, $R$ is the distance between the planet and the sun, $\alpha$ is a fixed power, and $k$ is some univeral constant that works for all the planets around our sun.
#
# Use Dimensional Analysis (Buckingham's Pi Theorem) to derive this result. Tell me what the value of $\alpha$ is.
#
# Don't use calculus! (Although you may have seen this solved via differential equations in a calc or physics class.)
#
# Hint: There is some ambiguity because of two masses involved (sun and planet). Newton knew that the mass of the planet does not matter to $T$, so you can assume this as well. Newton's universal gravitation constant $G$ also enters into the problem -- you can look up what units it is measured in, on Google. Or you can figure it out yourself from the force formula for planetary attraction $$\mbox{ Force } = G \frac{Mm}{R^2}$$
# where $M,m$ are the masses of the sun and planet.
#
# You can also check your answer by looking up Kepler's laws on Google.
# **Solution:**
#
# We have 4 parameters: T (in units of time), R (in units of distance), M (in units of mass), and G (we need to figure out the units, or look it up).
#
# From the formula $\mbox{ Force } = G \frac{Mm}{R^2}$, and remembering that Force is a mass times an acceleration (mass times distance, divided by time squared), we concluse that $G$ is in units of distance cubed, divided by time squared and mass.
#
# There are 3 physical units, time, distance and mass.
#
# The 3x4 matrix we need is thus
# | T | R | M | G | |
# | --- | --- | --- | --- | --- |
# | 1 | 0 | 0 | -2 | time |
# | 0 | 1 | 0 | 3 | distance |
# | 0 | 0 | 1 | -1 | mass |
#
# Notice the matrix is in row-echelon form. I did this on purpose, by matching the columns T, R, M with their corresponding units time, distance, mass, in order.
#
# This makes it easy to read off the dimensionless parameter using back substitution. That is, just solve the matrix equation
# $$\left[
# \begin{array}{rrrr}
# 1 & 0 & 0 & -2 \\
# 0 & 1 & 0 & 3 \\
# 0 & 0 & 1 & -1
# \end{array}
# \right]
# \left[
# \begin{array}{r}
# x_1 \\
# x_2 \\
# x_3 \\
# 1
# \end{array}
# \right] = 0.
# $$
#
# So we find
# $$
# \left[
# \begin{array}{r}
# x_1 \\
# x_2 \\
# x_3 \\
# 1
# \end{array}
# \right] =
# \left[
# \begin{array}{r}
# 2 \\
# -3 \\
# 1 \\
# 1
# \end{array}
# \right].
# $$
# Our dimensionless parameter is thus
# $$\Pi = GMT^2/R^3.$$
#
# (But seriously, you don't have to use the matrix if you don't want. A faster way: Start with G, in units of distances cubed, divided by mass and time squared. Multiply G by M to cancel the "mass" units. Then take GM and multiply by $T^2$ to cancel the "time" units. Finally, take $GMT^2$ and divide by $R^3$ to cancel the distance units.)
#
# Anyhow, the second part of Buckingham Pi tells us that
# $$ \Pi = GMT^2/R^3 = constant.$$
# Moving the stuff to the other side, we have
# $$T^2 = \frac{R^3}{GM}*C,$$
# where $C$ is some constant. Taking roots,
# $$T = k{R^{3/2}}$$
# where $k = \sqrt{C/GM}$ is a constant.
#
# ## Exercise 4
#
# Make a table listing the 8 planets plus Pluto, their distance to the sun, and the period of their orbit.
#
# Make a log-log plot of period versus distance, and check that you get a straight line with slope equal to $\alpha$. (At least approximately.)
#
# i.e. Taking logs of the equation $T = kR^\alpha$ gives
# $$\log T = \log k + \alpha \log R,$$
# which is the equation of a line in $x = \log R, y = \log T.$
#
#
# +
# data from Hyperphysics (thanks to Ahmad for this)
T = [0.241, 0.615, 1, 1.88, 11.9, 29.5, 84, 165, 248] # years
R = [5.69, 10.8, 15.0, 22.8, 77.8, 143, 287, 450, 590] # 10^10 m
plot(log(R),log(T),'-o');
xlabel("log distance"); ylabel("log time");
# +
## To check the slope, we can fit a line to the data using Python code
from scipy.optimize import curve_fit
# first, we use a linear function to estimate
def f(x,m,b):
return m*x + b
# then, curve fit, using the log data
result = curve_fit(f,log(R),log(T))
print(result[0])
# -
# The result is a slope m = 1.4974, which is very close to our result of $\alpha = 3/2.$
# ## Exercise 5
#
# Nuclear bombs, when exploded in the atmosphere, produce a large fireball that expands in the approximate shape of a sphere of some radius $r = r(t)$, which is a function of elapsed time. The rate at which it expands depends on the energy $E$ released by the bomb, the elapsed time $t$ since the detonation, and the density $\rho$ of the surrounding air.
#
# Curiously, the actual air pressure is irrelevant, as it is so small compared to the explosive blast.
#
# Use dimensional analysis to find $r(t)$ as a function of $E,t,\rho$. (times some fixed constant)
# **Solution:**
#
# I don't want to work too hard. I could use that matrix thingy, but let's be lazy.
#
# We have four parameters, r (a distance), t (a time), $\rho$ (a density, in mass/distance-cubed), and E (an energy, in mass times distance-squared, divided by time-squared).
#
# Why do we know the energy units by heart? From Einstein's $E = mc^2$, where m is a mass and c is a velocity. See, you knew that.
#
# So, how do I get something dimensionless?
#
# Start with E, divide by $\rho$ to get the masses to cancel. Then we have have $E/\rho$ in units of distance to the 5th power, divided by a time-squared. So
# $$E t^2/\rho r^5$$
# will be dimensionless.
#
# Wasn't that easy?
#
# So by Buckingham Pi, we have
# $$\Pi = E t^2/\rho r^5 = constant.$$
#
# Solving for r, we have
# $$ r(t) = k(E/\rho)^{1/5}t^{2/5}.$$
# ## Exercise 6 - for fun.
#
# Can you animate the result in Exercise 5, showing some bomb blasts of various energies? Something pretty?
# ## Hold my beer.
# +
## precompute some vectors, for polar coordinates
theta = linspace(0,2*pi,100)
ctheta = cos(theta)
stheta = sin(theta)
## I will just draw some circles with the appropriate radius from the formulas above.
def drawBombs(t=1,E1=1,E2=5):
r1 = (E1**(1/5))*(t**(2/5))
r2 = (E2**(1/5))*(t**(2/5))
plot(r1*ctheta,r1*stheta,'b',r2*ctheta,r2*stheta,'r')
xlim([-10,10]); ylim([-10,10])
interact(drawBombs,t=(0,100,1),E1=(1,10),E2=(1,10));
# -
| Assignment_1_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import os
import glob
import torch
import torch.nn.functional as F
import joblib
import itertools
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.api as sm
import warnings
import string
from sklearn.metrics import roc_auc_score, average_precision_score, brier_score_loss, recall_score, precision_score
from prediction_utils.util import df_dict_concat, yaml_read
from matplotlib.ticker import FormatStrFormatter
project_dir = '/share/pi/nigam/projects/spfohl/cohorts/admissions/mimic_omop/'
experiment_name_baseline = 'baseline_tuning_fold_1_10'
experiment_name_fair = 'fair_tuning_fold_1_10'
tasks = ['los_icu_3days', 'los_icu_7days', 'mortality_hospital', 'mortality_icu']
cohort_path = os.path.join(project_dir, 'cohort', 'cohort.parquet')
row_id_map_path = os.path.join(
project_dir, 'merged_features_binary/features_sparse/features_row_id_map.parquet'
)
result_path = os.path.join(project_dir, 'experiments', 'merged_results_fold_1_10')
os.makedirs(result_path, exist_ok=True)
attributes = ['gender_concept_name', 'age_group', 'race_eth']
cohort = pd.read_parquet(cohort_path)
row_id_map = pd.read_parquet(row_id_map_path)
cohort = cohort.merge(row_id_map)
# ### Generate the cohort table
### Cohort table
cohort_df_long = (
cohort
.melt(
id_vars = ['person_id'] + attributes,
value_vars = tasks,
var_name = 'task',
value_name = 'labels'
)
.melt(
id_vars = ['person_id', 'task', 'labels'],
value_vars = attributes,
var_name = 'attribute',
value_name = 'group'
)
)
# +
cohort_statistics_df = (
cohort_df_long
.groupby(['task', 'attribute', 'group'])
.agg(
prevalence=('labels', 'mean'),
)
.reset_index()
.groupby('attribute')
.apply(lambda x: x.pivot_table(index = 'group', columns = 'task', values = 'prevalence'))
.reset_index()
)
group_size_df = (
cohort_df_long
.groupby(['task', 'attribute', 'group'])
.agg(
size = ('labels', lambda x: x.shape[0])
)
.reset_index()
.drop(columns = 'task')
.drop_duplicates()
)
cohort_statistics_df = cohort_statistics_df.merge(group_size_df)
cohort_statistics_df = (
cohort_statistics_df
.set_index(['attribute', 'group'])
[['size'] + tasks]
)
# -
cohort_statistics_df
## Write to Latex
table_path = './../figures/mimic_omop/icu_admission_cohort/'
os.makedirs(table_path, exist_ok=True)
with open(os.path.join(table_path, 'cohort_table.txt'), 'w') as fp:
(
cohort_statistics_df
.reset_index().drop(columns='attribute').set_index(['group'])
.to_latex(
fp,
float_format = '%.3g',
index_names = False,
index=True
)
)
# ### Get the results
def get_result_df_baseline(base_path, filename='result_df_group_standard_eval.parquet'):
"""
Gets the results for training the baseline models
"""
selected_models_path = os.path.join(
base_path,
'config',
'selected_models', '**', '*.yaml'
)
selected_models_dict = {
filename.split('/')[-2]: filename.split('/')[-1]
for filename in glob.glob(selected_models_path, recursive=True)
}
paths = [
glob.glob(
os.path.join(
base_path,
'performance',
task,
config_filename,
'**',
filename
),
recursive=True
)
for task, config_filename in selected_models_dict.items()
]
paths = list(itertools.chain(*paths))
result_df_baseline = df_dict_concat(
{
tuple(filename.split('/'))[-4:-1]:
pd.read_parquet(filename)
for filename in paths
},
['task2', 'config_filename', 'fold_id']
).drop(columns='task2')
return result_df_baseline
result_df_baseline = get_result_df_baseline(
os.path.join(
project_dir,
'experiments',
experiment_name_baseline,
)
)
result_df_baseline.task.unique()
result_df_baseline.task.unique()
result_df_calibration_baseline = get_result_df_baseline(
os.path.join(
project_dir,
'experiments',
experiment_name_baseline,
),
filename='calibration_result.parquet'
)
id_vars = ['fold_id', 'phase', 'config_filename', 'task', 'attribute', 'group']
result_df_calibration_baseline = result_df_calibration_baseline.melt(
id_vars = id_vars,
value_vars = set(result_df_calibration_baseline.columns) - set(id_vars),
var_name = 'metric',
value_name = 'performance'
).query('metric != "brier"')
result_df_calibration_baseline.metric.unique()
# Import fair_ova metrics
result_df_ova_baseline = get_result_df_baseline(
os.path.join(
project_dir,
'experiments',
experiment_name_baseline,
),
filename='result_df_group_fair_ova.parquet'
)
# id_vars = ['fold_id', 'phase', 'config_filename', 'task', 'attribute', 'group']
# result_df_ova_baseline = result_df_ova_baseline.melt(
# id_vars = id_vars,
# value_vars = set(result_df_ova_baseline.columns) - set(id_vars),
# var_name = 'metric',
# value_name = 'performance'
# )
result_df_baseline = pd.concat([result_df_baseline, result_df_calibration_baseline, result_df_ova_baseline], ignore_index=True)
result_df_baseline
def flatten_multicolumns(df):
"""
Converts multi-index columns into single colum
"""
df.columns = ['_'.join([el for el in col if el != '']).strip() for col in df.columns.values if len(col) > 1]
return df
# +
# result_df_baseline.performance.isna()
# -
result_df_baseline_mean = (
result_df_baseline
.groupby(list(set(result_df_baseline.columns) - set(['fold_id', 'performance', 'performance_overall'])))
[['performance', 'performance_overall']]
# [['performance']]
.agg(['mean', 'std', 'sem'])
# .agg('max')
.reset_index()
)
result_df_baseline_mean = result_df_baseline_mean.rename(
columns={
'performance': 'performance_baseline',
'performance_overall': 'performance_overall_baseline'
}
)
result_df_baseline_mean = flatten_multicolumns(result_df_baseline_mean)
result_df_baseline_mean
def get_result_df_fair(base_path=None, filename='result_df_group_standard_eval.parquet', paths=None):
if paths is None:
performance_path = os.path.join(
base_path,
'performance',
)
paths = glob.glob(os.path.join(performance_path, '**', filename), recursive=True)
result_df_fair = df_dict_concat(
{
tuple(file_name.split('/'))[-5:-1]:
pd.read_parquet(file_name)
for file_name in paths
},
['task2', 'sensitive_attribute', 'config_filename', 'fold_id']
).drop(columns='task2')
return result_df_fair
# Fair results
result_df_fair = get_result_df_fair(
os.path.join(
project_dir,
'experiments',
experiment_name_fair
)
)
# +
# # List config_filenames without ten results
# (
# result_df_fair
# .groupby(
# list(set(result_df_fair.columns) - set(['fold_id', 'performance', 'performance_overall']))
# )
# .agg(lambda x: len(x))
# .query("fold_id != 10")
# .reset_index()
# .config_filename
# .sort_values()
# .unique()
# )
# +
result_df_calibration_fair = get_result_df_fair(
os.path.join(
project_dir,
'experiments',
experiment_name_fair
),
filename='calibration_result.parquet'
)
id_vars = ['fold_id', 'phase', 'config_filename', 'task', 'sensitive_attribute', 'attribute', 'group']
result_df_calibration_fair = result_df_calibration_fair.melt(
id_vars = id_vars,
value_vars = set(result_df_calibration_fair.columns) - set(id_vars),
var_name = 'metric',
value_name = 'performance'
).query('metric != "brier"')
# +
result_df_ova_fair = get_result_df_fair(
os.path.join(
project_dir,
'experiments',
experiment_name_fair
),
filename='result_df_group_fair_ova.parquet'
)
# id_vars = ['fold_id', 'phase', 'config_filename', 'task', 'sensitive_attribute', 'attribute', 'group']
# result_df_ova_fair = result_df_ova_fair.melt(
# id_vars = id_vars,
# value_vars = set(result_df_ova_fair.columns) - set(id_vars),
# var_name = 'metric',
# value_name = 'performance'
# )
# -
result_df_ova_fair
result_df_fair = pd.concat([result_df_fair, result_df_calibration_fair, result_df_ova_fair], ignore_index=True)
result_df_fair_mean = (
result_df_fair
.groupby(list(set(result_df_fair.columns) - set(['fold_id', 'performance', 'performance_overall'])))
[['performance', 'performance_overall']]
.agg(['mean', 'std', 'sem'])
.reset_index()
)
result_df_fair_mean = flatten_multicolumns(result_df_fair_mean)
ci_func = lambda x: x * 1.96
result_df_fair_mean = result_df_fair_mean.assign(
performance_CI = lambda x: ci_func(x['performance_sem']),
performance_overall_CI = lambda x: ci_func(x['performance_overall_sem']),
)
def label_fair_mode(df):
df['fair_mode'] = (
df['regularization_metric']
.where(~df['regularization_metric'].str.match('mmd'),
df['regularization_metric'].astype(str) + '_' + df['mmd_mode'].astype(str),
axis=0)
)
df['fair_mode'] = (
df['fair_mode']
.where(~df['fair_mode'].str.match('mean_prediction'),
df['fair_mode'].astype(str) + '_' + df['mean_prediction_mode'].astype(str),
axis=0
)
)
return df
def get_fair_config_df(base_path):
config_path = os.path.join(
base_path,
'config',
)
fair_config_files = glob.glob(
os.path.join(config_path, '**', '*.yaml'),
recursive=True
)
fair_config_dict_dict = {
tuple(file_name.split('/'))[-2:]:
yaml_read(file_name)
for file_name in fair_config_files
}
fair_config_df = df_dict_concat(
{
key: pd.DataFrame(value, index=[key])
for key, value in fair_config_dict_dict.items()
},
['task', 'config_filename']
)
fair_config_df = label_fair_mode(fair_config_df)[['task', 'config_filename', 'fair_mode', 'lambda_group_regularization']]
return fair_config_df
fair_config_df = get_fair_config_df(
os.path.join(
project_dir,
'experiments',
experiment_name_fair
)
)
fair_config_df
result_df_fair_mean.task.unique()
result_df = pd.merge(result_df_baseline_mean.drop(columns='config_filename'), result_df_fair_mean,
how='outer', indicator=True).merge(fair_config_df)
assert result_df_fair_mean.shape[0] == result_df.shape[0]
result_df.head()
result_df.query('_merge == "right_only"')
result_df.metric.unique()
result_df = result_df.query('phase == "test"')
result_df = result_df.drop(columns = '_merge')
result_df.to_csv(os.path.join(result_path, 'group_results.csv'), index=False)
| fairness_benchmark/notebooks/aggregate_results_mimic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.model_selection import GridSearchCV
from demo_utils.general import get_data
from sklearn.tree import DecisionTreeClassifier
from demo_utils.learning import get_model
import numpy as np
from IPython.display import Markdown as md
from demo_utils.general import SUPPORTED_DATASETS
from scipy.linalg import LinAlgError
import warnings
warnings.filterwarnings('ignore')
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# -
#testing_dataset = 'segment'
testing_dataset = None
dts_size = 1000
model_params = {
'model_name': 'dt',
'sampler_name': 'identity',
'pca_bool': False,
'pca_first': True,
'box_type': 'none'
}
hp_names = {'dt': 'min_impurity_decrease', 'logit': 'C', 'linear_svc': 'C'}
hp_options = {
'dt': [10**i for i in range(-10, 1)],
'logit': [10**i for i in range(-5, 4)],
'linear_svc': [10**i for i in range(-5, 4)]
}
rbf_gamma_options = [10**i for i in range(-5, 2)]
nystroem_gamma_options = [10**i for i in range(-5, 2)]
def find_hparams(model_params, options, dts_name):
n_comp = 500
prefix = 'model__'
tunning_params = {prefix + hp_names[model_params['model_name']]: options}
model = get_model(**model_params)
if model_params['sampler_name'] == 'rbf':
tunning_params['sampler__' + 'gamma'] = rbf_gamma_options
model.set_params(sampler__n_components=n_comp)
elif model_params['sampler_name'] == 'nystroem':
tunning_params['sampler__' + 'gamma'] = nystroem_gamma_options
model.set_params(sampler__n_components=n_comp)
clf = GridSearchCV(model, tunning_params, cv=10, iid=False)
data = get_data(dts_name, n_ins=dts_size)
data_train = data['data_train']
target_train = data['target_train']
#times = 5
#g = []
#for i in range(times):
# clf.fit(data_train, target_train)
# bp = clf.best_params_
# nbp = list(bp.values())[0]
# g.append(nbp)
#m = np.mean(g)
#key = list(bp.keys())[0]
#clf.fit(data_train, target_train)
#bp = clf.best_params_
is_failing = True
while is_failing:
try:
clf.fit(data_train, target_train)
is_failing = False
except LinAlgError:
pass
bp = clf.best_params_
#print(model_params['model_name'], key)
print(model_params['model_name'])
print(bp)
print()
def test_dataset(dts_name):
display(md('# ' + dts_name))
global testing_dataset
testing_dataset = dts_name
test_simple()
test_simple_pca()
display(md('## RFF'))
test_sampler('rbf')
display(md('## Nystroem'))
test_sampler('nystroem')
# +
############
def test_simple():
display(md('## Simple'))
model_params['pca_bool'] = False
model_params['sampler_name'] = 'identity'
for model_name in hp_names:
model_params['model_name'] = model_name
print('testing_dataset es {}'.format(testing_dataset))
find_hparams(model_params, options=hp_options[model_params['model_name']], dts_name=testing_dataset)
############
def test_simple_pca():
display(md('## Simple PCA'))
model_params['pca_bool'] = True
model_params['sampler_name'] = 'identity'
for model_name in hp_names:
model_params['model_name'] = model_name
find_hparams(model_params, options=hp_options[model_params['model_name']], dts_name=testing_dataset)
# -
##############
def test_sampler(sampler_name):
test_sampler_no_pca(sampler_name)
test_sampler_pca_first(sampler_name)
test_sampler_pca_last(sampler_name)
##############
# +
############
def test_sampler_no_pca(sampler_name):
display(md('### No PCA'))
model_params['pca_bool'] = False
model_params['sampler_name'] = sampler_name
for model_name in hp_names:
model_params['model_name'] = model_name
find_hparams(model_params, options=hp_options[model_params['model_name']], dts_name=testing_dataset)
############
def test_sampler_pca_first(sampler_name):
display(md('### PCA First'))
model_params['pca_bool'] = True
model_params['pca_first'] = True
model_params['sampler_name'] = sampler_name
for model_name in hp_names:
model_params['model_name'] = model_name
find_hparams(model_params, options=hp_options[model_params['model_name']], dts_name=testing_dataset)
def test_sampler_pca_last(sampler_name):
display(md('### PCA Last'))
model_params['pca_bool'] = True
model_params['pca_first'] = False
model_params['sampler_name'] = sampler_name
for model_name in hp_names:
model_params['model_name'] = model_name
find_hparams(model_params, options=hp_options[model_params['model_name']], dts_name=testing_dataset)
# -
for sd in SUPPORTED_DATASETS:
test_dataset(sd)
for sd in SUPPORTED_DATASETS[3:]:
test_dataset(sd)
for sd in SUPPORTED_DATASETS[5:]:
test_dataset(sd)
| code/notebooks/python/finding_best_params/.ipynb_checkpoints/Reduced Experimentation-Copy1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # CF
# +
# ! aws cloudformation create-stack \
# --stack-name test1 \
# --template-body file:///home/ec2-user/SageMaker/Lex-KR-Workshop/CF_BankingBot_v4.yaml \
# --timeout-in-minutes 5 \
# --capabilities CAPABILITY_IAM
# --parameters ParameterKey=Parm1,ParameterValue=test1 ParameterKey=Parm2,ParameterValue=test2
# +
# ! aws cloudformation describe-stacks \
# --stack-name test1 \
#
# -
# ! aws cloudformation delete-stack \
# --stack-name test1\
#
| aiservices/lex-korean-workshop/Create-CF-Lex-KR-Workshop/9.1.Run-CF_Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="otQx9wd94UJ9" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1616195794920, "user_tz": 360, "elapsed": 19436, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="d4b3f095-2b21-4b5d-e091-0ba07e3035d8"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="IQEmmmuF49H0" executionInfo={"status": "ok", "timestamp": 1616195812264, "user_tz": 360, "elapsed": 15508, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="60a6574f-9639-4b84-8711-2582aa254807"
import glob
import numpy as np
import os
import shutil
np.random.seed(0)
files = glob.glob('/content/drive/MyDrive/BUS_binary/*')
benign_files = [fn for fn in files if 'benign' in fn]
malignant_files = [fn for fn in files if 'malignant' in fn]
len(benign_files), len(malignant_files)
# + colab={"base_uri": "https://localhost:8080/"} id="w-gLUvKv5cXt" executionInfo={"status": "ok", "timestamp": 1616196042457, "user_tz": 360, "elapsed": 372, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="e09be32b-f994-4fe7-8717-ccd70617fd56"
benign_train = np.random.choice(benign_files, size=1598, replace=False)
malignant_train = np.random.choice(malignant_files, size=838, replace=False)
benign_test = list(set(benign_files) - set(benign_train))#
malignant_test = list(set(malignant_files) - set(malignant_train))#
benign_val = np.random.choice(benign_train, size=320, replace=False)
malignant_val = np.random.choice(malignant_train, size=168, replace=False)
benign_train = list(set(benign_train) - set(benign_val))
malignant_train = list(set(malignant_train) - set(malignant_val))
print(len(benign_train),len(malignant_train),len(benign_val),len(malignant_val),len(benign_test),len(malignant_test))
# + id="xS4iXfR27cxM" executionInfo={"status": "ok", "timestamp": 1616196622605, "user_tz": 360, "elapsed": 253069, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}}
train_dir = '/content/drive/MyDrive/DATASET_BUS_BINARY/training_data'
val_dir = '/content/drive/MyDrive/DATASET_BUS_BINARY/validation_data'
test_dir = '/content/drive/MyDrive/DATASET_BUS_BINARY/test_data'
import shutil
train_files = np.concatenate([benign_train, malignant_train])
validate_files = np.concatenate([benign_val, malignant_val])
test_files = np.concatenate([benign_test, malignant_test])
shutil.rmtree('/content/drive/MyDrive/DATASET_BUS_BINARY/training_data',ignore_errors=True)
shutil.rmtree('/content/drive/MyDrive/DATASET_BUS_BINARY/validation_data',ignore_errors=True)
shutil.rmtree('/content/drive/MyDrive/DATASET_BUS_BINARY/test_data',ignore_errors=True)
##
os.mkdir(train_dir) if not os.path.isdir(train_dir) else None
os.mkdir(val_dir) if not os.path.isdir(val_dir) else None
os.mkdir(test_dir) if not os.path.isdir(test_dir) else None
for fn in train_files:
shutil.copy(fn, train_dir)
for fn in validate_files:
shutil.copy(fn, val_dir)
for fn in test_files:
shutil.copy(fn, test_dir)
# + colab={"base_uri": "https://localhost:8080/"} id="I2Wx5fdKFQEt" executionInfo={"status": "ok", "timestamp": 1616196622613, "user_tz": 360, "elapsed": 28, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="ca952f88-4263-4da0-87b2-d72b0c070f54"
files_tr = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/training_data/*')
files_vd = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/validation_data/*')
files_tt = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/test_data/*')
files_tr_benign = [fn for fn in files_tr if 'benign' in fn]
files_tr_malignant = [fn for fn in files_tr if 'malignant' in fn]
files_vd_benign = [fn for fn in files_vd if 'benign' in fn]
files_vd_malignant = [fn for fn in files_vd if 'malignant' in fn]
files_tt_benign = [fn for fn in files_tt if 'benign' in fn]
files_tt_malignant = [fn for fn in files_tt if 'malignant' in fn]
print('Files_tr:',len(files_tr_benign), len(files_tr_malignant))
print('Files_vd:',len(files_vd_benign), len(files_vd_malignant))
print('Files_tt:',len(files_tt_benign), len(files_tt_malignant))
# + [markdown] id="F_vDDsws-7lB"
# PREPARING DATASET
# + colab={"base_uri": "https://localhost:8080/"} id="5mJWDwWN--8i" executionInfo={"status": "ok", "timestamp": 1616196627695, "user_tz": 360, "elapsed": 5095, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="a3de09e3-cf56-4a92-82ba-ccb515149569"
import glob
import numpy as np
import os
import shutil
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator, load_img, img_to_array, array_to_img
# %matplotlib inline
IMG_DIM = (150,150)
train_files = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/training_data/*')
train_imgs = [img_to_array(load_img(img, target_size=IMG_DIM)) for img in train_files]
train_imgs = np.array(train_imgs)
train_labels = [fn.split('/')[-1].split('.')[0].strip() for fn in train_files]
validation_files = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/validation_data/*')
validation_imgs = [img_to_array(load_img(img, target_size=IMG_DIM)) for img in validation_files]
validation_imgs = np.array(validation_imgs)
validation_labels = [fn.split('/')[-1].split('.')[0].strip() for fn in validation_files]
test_files = glob.glob('/content/drive/MyDrive/DATASET_BUS_BINARY/test_data/*')
test_imgs = [img_to_array(load_img(img, target_size=IMG_DIM)) for img in test_files]
test_imgs = np.array(test_imgs)
test_labels = [fn.split('/')[-1].split('.')[0].strip() for fn in test_files]
print('Train dataset shape:', train_imgs.shape,
'\tValidation dataset shape:', validation_imgs.shape,'\tTest dataset shape:', test_imgs.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 184} id="8hFKv0AQ_qm6" executionInfo={"status": "ok", "timestamp": 1616196628105, "user_tz": 360, "elapsed": 415, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="28903ba4-54c1-4cec-a78c-44462918be9f"
train_imgs_scaled = train_imgs.astype('float32')
validation_imgs_scaled = validation_imgs.astype('float32')
test_imgs_scaled =test_imgs.astype('float32')
train_imgs_scaled /= 255
validation_imgs_scaled /= 255
test_imgs_scaled /= 255
print(train_imgs[0].shape)
array_to_img(train_imgs[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 184} id="3V8RI3mZJ8Tl" executionInfo={"status": "ok", "timestamp": 1616196628106, "user_tz": 360, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="43d0bde9-94c5-4c0a-f0ab-353837212d28"
print(train_imgs[6].shape)
array_to_img(train_imgs[6])
# + colab={"base_uri": "https://localhost:8080/", "height": 184} id="XIL0dCfN_nI2" executionInfo={"status": "ok", "timestamp": 1616196628108, "user_tz": 360, "elapsed": 32, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="34ac73b0-2026-409e-e5f5-a9c92c22e223"
print(test_imgs[25].shape)
array_to_img(test_imgs[25])
# + colab={"base_uri": "https://localhost:8080/"} id="e7ORAbLV_3sD" executionInfo={"status": "ok", "timestamp": 1616196628642, "user_tz": 360, "elapsed": 558, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="9a3a025e-3751-4abd-b1bb-827e0f5d88e2"
batch_size = 30
num_classes = 2
epochs = 30
input_shape = (150, 150, 3)
# encode text category labels
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
le.fit(train_labels)
train_labels_enc = le.transform(train_labels)
validation_labels_enc = le.transform(validation_labels)
test_labels_enc = le.transform(test_labels)
print(train_labels[25:30], train_labels_enc[25:30], test_labels_enc[25:30])
# + id="U893pCgwHOVN" executionInfo={"status": "ok", "timestamp": 1616196769233, "user_tz": 360, "elapsed": 249, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}}
train_datagen = ImageDataGenerator(rescale=1./255, zoom_range=0.3, rotation_range=20,
width_shift_range=0, height_shift_range=0, shear_range=0,
horizontal_flip=True, fill_mode='constant')
val_datagen = ImageDataGenerator(rescale=1./255)
test_datagen = ImageDataGenerator(rescale=1./255)
# + colab={"base_uri": "https://localhost:8080/", "height": 222} id="1rqB8WTpHWIB" executionInfo={"status": "ok", "timestamp": 1616196770924, "user_tz": 360, "elapsed": 960, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="31386575-3b64-4e9e-c61b-8d4e42493863"
img_id = 6
bening_generator = train_datagen.flow(train_imgs[img_id:img_id+1], train_labels[img_id:img_id+1],
batch_size=1)
bening = [next(bening_generator) for i in range(0,5)]
fig, ax = plt.subplots(1,5, figsize=(16, 6))
print('Labels:', [item[1][0] for item in bening])
l = [ax[i].imshow(bening[i][0][0]) for i in range(0,5)]
# + colab={"base_uri": "https://localhost:8080/", "height": 222} id="8jxAEl60IYKW" executionInfo={"status": "ok", "timestamp": 1616196774513, "user_tz": 360, "elapsed": 833, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="7005a026-bc11-4d26-d638-605ac421b3a5"
img_id = 1500
malignant_generator = train_datagen.flow(train_imgs[img_id:img_id+1], train_labels[img_id:img_id+1],
batch_size=1)
malignant = [next(malignant_generator) for i in range(0,5)]
fig, ax = plt.subplots(1,5, figsize=(16, 6))
print('Labels:', [item[1][0] for item in malignant])
l = [ax[i].imshow(malignant[i][0][0]) for i in range(0,5)]
# + colab={"base_uri": "https://localhost:8080/"} id="FFsUFxOLGx2U" executionInfo={"status": "ok", "timestamp": 1616125590843, "user_tz": 360, "elapsed": 2360, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="6866914b-94c9-4c97-e52a-0cd1acb637da"
train_generator = train_datagen.flow(train_imgs, train_labels_enc, batch_size=30)
val_generator = val_datagen.flow(validation_imgs, validation_labels_enc, batch_size=20)
test_generator = test_datagen.flow(test_imgs, test_labels_enc, batch_size=20)
input_shape = (150, 150, 3)
TRAIN_STEPS_PER_EPOCH = np.ceil((len(train_imgs)/30)-1)
# to ensure that there are enough images for training bahch
VAL_STEPS_PER_EPOCH = np.ceil((len(validation_imgs)/20)-1)
print(TRAIN_STEPS_PER_EPOCH,VAL_STEPS_PER_EPOCH)
# + [markdown] id="DV5GDya37AvR"
# ---------------------------------------------
# + colab={"base_uri": "https://localhost:8080/"} id="hIWZUgNIKA-A" executionInfo={"status": "ok", "timestamp": 1616126042285, "user_tz": 360, "elapsed": 1992, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="08d6d1cb-329c-44f2-a784-e901bc76e7f4"
import keras
from keras.models import Sequential
from keras import optimizers
from keras.preprocessing import image
##
from tensorflow.keras.applications.resnet50 import ResNet50
from keras.models import Model
import pandas as pd
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout, InputLayer, GlobalAveragePooling2D, Input
#base_model = ResNet50(
# weights="imagenet", # Load weights pre-trained on ImageNet.
# input_shape=input_shape,
# include_top=False,
#) # Do not include the ImageNet classifier at the top.
model = Sequential()
# 1st layer as the lumpsum weights from resnet50_weights_tf_dim_ordering_tf_kernels_notop.h5
# NOTE that this layer will be set below as NOT TRAINABLE, i.e., use it as is
model.add(ResNet50(include_top = False, pooling = 'avg', weights = "imagenet"))
# 2nd layer as Dense for 2-class classification, i.e., dog or cat using SoftMax activation
model.add(Dense(1, activation = 'sigmoid'))
# Say not to train first layer (ResNet) model as it is already trained
model.layers[0].trainable = False
model.summary()
# + id="AcLagiomVoYl" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1616126642694, "user_tz": 360, "elapsed": 597067, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="167b7791-f251-4e47-c2b4-783285d27780"
import tensorflow_datasets as tfds
import pathlib
from matplotlib import pyplot
from keras.callbacks import EarlyStopping
from keras.callbacks import ModelCheckpoint
import time
from keras.models import load_model
from keras.optimizers import SGD
sgd = optimizers.SGD(lr = 0.01, decay = 1e-6, momentum = 0.9, nesterov = True)
model.compile(loss='binary_crossentropy',
optimizer= sgd,
metrics=['accuracy'])
tic=time.time()
# training
filepath='/content/drive/MyDrive/MODELOS/resnet50_binary/weights-improvement-{epoch:02d}-{val_accuracy:.2f}.hdf5'
mc = ModelCheckpoint(filepath, monitor='val_accuracy', mode='max', verbose=1, save_best_only=True)
es = [mc]
history = model.fit(
train_generator, steps_per_epoch=TRAIN_STEPS_PER_EPOCH, epochs=50,
validation_data = val_generator, validation_steps=VAL_STEPS_PER_EPOCH, verbose=1,callbacks=[es])
# training
print('Tiempo de procesamiento (secs): ', time.time()-tic)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="Mgm9b_JvSoid" executionInfo={"status": "ok", "timestamp": 1616127815101, "user_tz": 360, "elapsed": 763, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="ac36c05a-4596-4da3-9454-07e6eb4a4d39"
plt.figure(1)
# summarize history for accuracy
plt.subplot(211)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Training', 'Validation'], loc='lower right')
# summarize history for loss
plt.subplot(212)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Training', 'Validation'], loc='upper right')
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="QXKCyrf-85MV" executionInfo={"status": "ok", "timestamp": 1616127851903, "user_tz": 360, "elapsed": 6065, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02129717593399559815"}} outputId="889c6ac7-30a5-46b1-8872-5f18c4c5148a"
model.load_weights("/content/drive/MyDrive/MODELOS/resnet50_binary/weights-improvement-36-0.82.hdf5")
scores = model.evaluate(
test_imgs_scaled,
test_labels_enc,
batch_size=64,
verbose=0,
workers=1,
use_multiprocessing=False,
return_dict=False,
)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
| Resnet50_BINARY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Heat transfer for pipes
# +
"""
importing the necessary libraries, do not modify
"""
# %matplotlib inline
from IPython.display import clear_output
import schemdraw as schem
import schemdraw.elements as e
import matplotlib.pyplot as plt
import numpy as np
import math
import scipy.constants as sc
import sympy as sym
# -
# <img src="figures/fig_08_08.jpg" alt="my awesome sketch" width=75% >
# <i>Fig. 1: Illustration of internal convection.</i>
# The above sketch illustrates the focus of this notebook: How to quantify the heat transfer between a pipe, in which a fluid flows, and its surroundings. The heat transfer from the outer surface of the pipe to the outer flow is to defined in the previous chapter, external convection. In the following, this notebook establishes the tools necessary to solve the internal convection problem.
# ## Entry flow and fully developed internal flow
# <img src="figures/fig_08_01.jpg" alt="my awesome sketch" width=100% >
# <i>Fig. 2: Pipe flow nomenclature.</i>
# ### Python module
#
# For internal flow, the module is loaded as:
from Libraries import HT_internal_convection as intconv
# As an example, consider the flow of water in a pipe of diameter $D=10$ cm, length $L=10$m. The water thermodynamic properties are estimated at $T_f=50^\circ$C. The bulk velocity is $U_m=2$m/s.
#
# +
from Libraries import thermodynamics as thermo
T_f = 50 #C
waterflow = thermo.Fluid('water',T_f,"C")
L_pipe = 10. #m
D_pipe = 0.1 #m
Um_pipe = 2 #m/s
# -
# ?intconv.PipeFlow
pipe = intconv.PipeFlow(D= D_pipe, L=L_pipe,
rho=waterflow.rho, nu=waterflow.nu, Um=Um_pipe)
# <img src="figures/fig_08_03.jpg" alt="my awesome sketch" width=100% >
# <i> Fig. 3. Friction factor in pipe flow as a function of Re and relative surface roughness.</i>
# A uniform flow entering a pipe (Fig. 2) first experiences streamwise variation of velocity to accommodate the wall boundary conditions. A boundary layer, of thickness $\delta$, forms on the wall and grows until its edge reaches the pipe centerline. This region is the hydrodynamic entrance region. Beyond that point, the flow becomes fully developed, which means that
# <ul>
# <li> In the laminar regime, the velocity profile is only a function of $r$,</li>
# <li> In the turbulent regime, the <b>mean</b> velocity profile is only a function of $r$.</li>
# </ul>
# Friction drag or the force exerted by the flow onto the pipe wall governs the pressure gradient necessary to generate a desired flowrate. Calculation of the friction drag leads to the design of the mechanical force creating the pressure gradient. In fully developed (laminar or turbulent) regimes, the pressure gradient may be determined by
# <p class='alert alert-danger'>
# $$
# -\frac{\Delta\overline{P}}{L}=f\,\frac{1}{D}\,\frac{\rho U_m^2}{2}
# $$
# </p>
# where $D=2R$ and $L$ are the diameter and length of the pipe, respectively, and $f$ is the <b>friction factor</b>. The bulk velocity or average velocity is
# <p class='alert alert-info'>
# $$
# U_m=\frac{\dot{m}}{\rho A_c}
# $$
# </p>
# where $\dot{m}$ is the mass flux
# $$
# \dot{m}=\int_0^{2\pi}\int_0^R\rho \overline{u}(r)\,r\,dr d\theta=2\pi\int_0^R\rho \overline{u}(r)\,r\,dr
# $$
# and $A_c=\pi R^2$
# The Reynolds number of the flow is based on the bulk velocity and pipe diameter:
# <p class='alert alert-danger'>
# $$
# Re_D=\frac{\rho U_mD}{\mu}=\frac{4\dot{m}}{\pi D\mu}
# $$
# </p>
# The friction factor in the laminar regime is rigorously derived:
# $$
# f = \frac{64}{Re_D}
# $$
# </p>
# and is valid up to the critical Reynolds number $Re_{D,c}$, which in most pipe is around 2,000. Be aware that in certain research facilities, the flow can remain laminar for Reynolds numbers up to 10,000. The Reynolds 2,000 is not absolute, universal property, but is the best guess from most engineering applications.
#
# Beyond the critical Reynolds number, $f$ is a function of the roughness to diameter ratio $\varepsilon=e/D$ (e is typically the standard deviation of the roughness height) and the Reynolds number. A trustworthy empirical correlation is the Colebrook formula:
# <p class='alert alert-danger'>
# $$
# \frac{1}{\sqrt{f}}=-2\log_{10}\left[\frac{\varepsilon}{3.7}+\frac{2.51}{Re_D\sqrt{f}}\right]
# $$
# </p>
# which is solved below for a range of relative roughness $\varepsilon$.
#
# Often there is a need to determine the pump or blower power $P$ necessary to move the flow at a prescribed pressure drop:
# <p class='alert alert-danger'>
# $$
# P=\frac{\dot{m}}{\rho}\Delta p= \underbrace{(\Delta p)A_c}_\text{force}\cdot U_m
# $$
# </p>
#
# ### Example of functions
# Going back to our library, let's explore how to determine some of the properties defined above:
# Reynolds number:
print("Re= %1.2e" %pipe.Re)
# Mass flow rate:
print("mass flowrate= %1.1f kg/s" %pipe.mdot)
# Compute the friction factor:
# pipe.f_turbulent()
pipe.f_laminar()
print("f= %1.5f" %pipe.f)
# The mean pressure gradient is:
print("-dP/dx= %1.0f Pa/m" %pipe.dPdx)
# ## Heat transfer by internal convection
# The temperature is expected to vary both in the streamwise direction and in the radial direction. To reduce the complexity of the problem, we define the mean temperature as:
# $$
# T_m=\frac{1}{\dot{m}C_p}\int_{A_c}\rho\,u\,C_p\, T\,dA_c
# $$
# where $\dot{m}$ is the mass flow rate, $rho$ and $C_p$ are the density and specific heat of the fluid and $A_c$ is the cross-sectional area of the pipe.
# The local heat flux may be now expressed as:
# $$
# q_s''=h(T_s-T_m)
# $$
# where $h$ is the <b>local</b> convection heat transfer coefficient and $T_s$ is the surface temperature on the inner wall of the pipe. The variation of temperature in the <b>fully developed</b> flow can be shown to be governed by the following ODE:
# <p class='alert alert-info'>
# $$
# \frac{dT_m}{dx}=\frac{P}{\dot{m}C_p}h(T_s-T_m)
# $$
# </p>
# where $P$ is the perimeter of the pipe.
# If the local heat flux is maintained constant over the length of the pipe $L$, the total heat rate is
# <p class='alert alert-danger'>
# $$
# q_\text{conv}=(PL)q_s''\, \text{$q_s''=$constant}
# $$
# </p>
# and the streamwise distribution of the mean temperature is linear:
# $$
# T_m(x)=T_{m,i}+\frac{q_s''P}{\dot{m}C_p}x,\, \text{$q_s''=$constant}
# $$
# For the case of constant wall temperature $T_s$, the temperature distribution is the solution of the above ODE, thus of exponential nature. For practical applications, you most always need to compute the overall heat transfer and the outlet mean temperature $T_{m,o}$. The integration of the above ODE for $x=0$ to $x=L$ yields
# <p class='alert alert-danger'>
# $$
# \frac{T_s-T_{m,o}}{T_s-T_{m,i}}=\exp\left(-\frac{PL}{\dot{m}C_p}\overline{h}\right),\, \text{$T_s=$constant}
# $$
# </p>
# where
# $$
# \overline{h}=\frac{1}{L}\int_0^L h(x)dx
# $$
# If you must compute the mean temperature at $x$ an integration from $0$ to $x$ yields
# <FONT FACE="courier" style="color:blue">T_mx_Ts_constant(T_s,T_mi,P,L,mdot,Cp,hbar,x)</FONT>
# <p class='alert alert-danger'>
# $$
# \frac{T_s-T_{m}(x)}{T_s-T_{m,i}}=\exp\left(-\frac{PL}{\dot{m}C_p}\overline{h}_x\right),\, \text{$T_s=$constant}
# $$
# </p>
# where
# $$
# \overline{h}_x=\frac{1}{L}\int_0^x h(x')dx'
# $$
# The computation of the total heat transfer rate can be shown to write:
# <p class='alert alert-danger'>
# $$
# q_\text{conv}=\overline{h}(PL)\Delta T_\text{lm},\, \text{$T_s=$constant}
# $$
# </p>
# with the log mean temperature
# <FONT FACE="courier" style="color:blue">log_mean_temperature(T_s,T_o,T_i)</FONT>
# <p class='alert alert-danger'>
# $$
# \Delta T_\text{lm}=\cfrac{T_{m,i}-T_{m,o}}{\ln\left(\cfrac{T_s-T_{m,o}}{T_s-T_{m,i}}\right)}
# $$
# </p>
# In many problem, $T_s$ is not defined but the outside ambient temperature $T_\infty$, the thermal conductivity of the pipe is known. One needs to determine the total resistance of the system $R_\text{tot}$, which requires calculating the heat transfer coefficient of the forced or natural convection, occuring on the outside of the pipe, the radiation coefficient if needed, the thermal resistance due by conduction within the pipe, which may include multiple components in the presence of insulation for example, and the internal convection heat transfer coefficient (to be defined below). In such cases, the variation of temperature between inlet and outlet becomes:
# <FONT FACE="courier" style="color:blue">T_mo_T_infty(T_infty,T_mi,P,L,mdot,Cp,R_tot)</FONT>
# <p class='alert alert-danger'>
# $$
# \frac{T_\infty-T_{m,o}}{T_\infty-T_{m,i}}=\exp\left(-\frac{1}{\dot{m}C_pR_\text{tot}}\right)
# $$
# </p>
# and the total heat transfer rate is
# <p class='alert alert-danger'>
# $$
# q=\frac{\Delta T_\text{lm}}{R_\text{tot}}
# $$
# </p>
# The equations derived in this cell enable:
# <ul>
# <li> The computation of the internal convection heat transfer coefficient if $T_{m,i}$ and $T_{m,o}$ are known.</li>
# <li> The computation of $T_{m,i}$ or $T_{m,o}$ if one is known and $\overline{h}$ is known </li>
# <li> The computation of the required mass flux to achieve given $T_{m,i}$ and $T_{m,o}$, albeit through an iterative process</li>
# </ul>
# ## Correlations for convection heat transfer coefficients in internal pipe flows
# Here we detailed only the correlations for fully developed flows. For laminar flows, the nusselt numbers are constant, thus the library <FONT FACE="courier" style="color:blue">HT_internal_convection</FONT> provides directly $\overline{h}$:
# <FONT FACE="courier" style="color:blue">laminar_isoflux() </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=4.36,\, \text{$q_s''=$constant}
# $$
# </p>
# <FONT FACE="courier" style="color:blue">laminar_isothermal() </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=4.36,\, \text{$q_s''=$constant}
# $$
# </p>
#
pipe.laminar_isoflux()
print("Nu= %1.2f for laminar isoflux" %pipe.Nu)
pipe.laminar_isothermal()
print("Nu= %1.2f for laminar isothermal" %pipe.Nu)
# In turbulent flows, there is a choice of correlations:
# <FONT FACE="courier" style="color:blue">Dittus_Boelter(Re,Pr,mode) </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=0.023Re^{4/5}Pr^n
# $$
# </p>
# with mode being either <FONT FACE="courier" style="color:blue">'cooling'</FONT> or <FONT FACE="courier" style="color:blue">'heating'</FONT>
pipe.Dittus_Boelter(mode='cooling',Pr=waterflow.Pr)
print("Nu= %1.0f for cooling" %pipe.Nu)
pipe.Dittus_Boelter(mode='heating',Pr=waterflow.Pr)
print("Nu= %1.0f for heating" %pipe.Nu)
# <FONT FACE="courier" style="color:blue">Sieder_Tate(Re,Pr,mu,mu_s) </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=0.027Re^{4/5}Pr^{1/3}\left(\cfrac{\mu}{\mu_s}\right)^{0.14}
# $$
T_s = 75 #C
watersurface = thermo.Fluid('water',thermo.C2K(T_s))
pipe.Sieder_Tate(mu=waterflow.mu,mu_s=watersurface.mu,Pr=waterflow.Pr)
print("Nu= %1.0f" %pipe.Nu)
# <FONT FACE="courier" style="color:blue">Gnielinski(Re,Pr,f) </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=\frac{(f/8)(Re-1000)Pr}{1+12.7(f/8)^{1/2}(Pr^{2/3}-1)}
# $$
# </p>
pipe.Gnielinski(f=pipe.f, Pr=waterflow.Pr)
print("Nu= %1.0f" %pipe.Nu)
# <FONT FACE="courier" style="color:blue">Skupinski(Re,Pr) </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=4.82+0.0185\left(Re\,Pr\right)^{0.827},\, \text{$q_s''=$constant}
# $$
# </p>
pipe.Skupinski(Pr=waterflow.Pr)
print("Nu= %1.0f" %pipe.Nu)
# <FONT FACE="courier" style="color:blue">Seban(Re,Pr) </FONT>
# <p class='alert alert-danger'>
# $$
# Nu=\frac{hD}{k}=5.0+0.025\left(Re\,Pr\right)^{0.8},\, \text{$T_s=$constant}
# $$
# </p>
pipe.Seban(Pr=waterflow.Pr)
print("Nu= %1.0f" %pipe.Nu)
# ## Natural convection around cylinder
# <img src="figures/fig_09_08.jpg" alt="my awesome sketch" width=75% >
# <i>Fig. 4: Illustration of the flow induced by natural convection around a cylinder. Insert shows the angular distribution of the local Nu.</i>
# In a fluid entirely at rest, a heated surface transfers its heat via pure conduction. Natural convection is the enhanced heat transfer between a body of fluid at rest (at infinity) and a heated surface through the creation of a convective flow driven by buoyancy forces. Fig. 4 illustrates a natural convection flow occuring around a cylinder. The fluid at the bottom of the cylinder $\theta=0$ becomes buoyant through heat transfer between the cylinder and the fluid and rises along the surface of the cylinder. This process creates two boundary layers that merge at $\theta = \pi$ to create a vertical jet-like flow, also called a plume. Plumes are characteristic flows of natural convection, i.e. they are found irrespective of the geometry of the heated object.
#
# The library is called in the following way:
from Libraries import HT_natural_convection as natconv
# The non-dimensional numbers relevant to natural convection are:
# the Grashof number
# <FONT FACE="courier" style="color:blue">Grashof(g,beta,DT,D,nu) </FONT>
# <p class='alert alert-danger'>
# $$
# Gr = \frac{g\beta(\Delta T)D^3}{\nu^2}
# $$
# </p>
# and the Rayleigh number
# <FONT FACE="courier" style="color:blue">Rayleigh(g,beta,DT,D,nu,alpha) </FONT>
# <p class='alert alert-danger'>
# $$
# Ra = Gr.Pr= \frac{g\beta(\Delta T)D^3}{\nu\alpha}
# $$
# </p>
# where $g$ is the gravity magnitude, $\beta$ is the volumetric thermal expansion coefficient at a given pressure $p$
# $$
# \beta = -\frac{1}{\rho}\left(\frac{\partial\rho}{\partial T}\right)_p
# $$
# $\Delta T$ is the absolute temperature difference between the heated surface temperature $T_s$ and the fluid temperature at infinity $T_\infty$, $\Delta T= \vert T_s-T_\infty\vert$, $D$ is the characteristic length of the system (here the diameter) and $\nu$ and $\alpha$ are the kinematic viscosity and the thermal diffusivity, both of dimensions $\text{m$^2$/s}$.
# Note that for the ideal gas law
# $$
# p =\rho \frac{R}{M}T\text{ or } \rho = \frac{p}{\frac{R}{M}T}
# $$
# thus the expansion coefficient is
# <p class='alert alert-info'>
# $$
# \beta = \frac{1}{T}\text{ for an ideal gas, $T$ in K}
# $$
# </p>
# For a liquid, $\beta$ must be interpolated from a table. All thermodynamics quantities involved are to be defined at the film temperature which is the arithmetic mean
# <p class='alert alert-info'>
# $$
# T_f=\frac{T_s+T_\infty}{2}
# $$
# </p>
#air
T_infty = 10#C
T_s = 50#C
D = 0.1#m
T_f = (T_s+T_infty)/2
airflow = thermo.Fluid('air',T_f,"C")
Gr= natconv.Gr(beta=airflow.beta,D=D,DT=T_s-T_infty,nu=airflow.nu)
print('Natural convection Gr= %1.2e'%Gr)
Ra= natconv.Ra(alpha=airflow.alpha,beta=airflow.beta,D=D,DT=T_s-T_infty,nu=airflow.nu)
print('Natural convection Ra= %1.2e'%Ra)
# The Grashof and Rayleigh number quantify the ratio of buoyancy to viscous forces. When they are large enough, a convective flow sets in and the heat transfer increases in comparison to pure conduction. The Nusselt number, ratio of convective to conduction heat transfer (i.e. $>1$ in the presence of a convection flow) is typically a power law of the Rayleigh number. In the case of the flow around a cylinder with isothermal surface temperature, there are two correlations:
# <FONT FACE="courier" style="color:blue">Morgan(Ra) </FONT>
# <p class='alert alert-danger'>
# $$
# \overline{Nu}=\frac{\overline{h}D}{k}=C\,Ra^n
# $$
# </p>
# <FONT FACE="courier" style="color:blue">Churchill-Chu(Ra,Pr) </FONT>
# <p class='alert alert-danger'>
# $$
# \overline{Nu}=\frac{\overline{h}D}{k}=\left[0.60+\frac{0.387Ra^{1/6}}{\left[1+\left(\frac{0.559}
# {Pr}\right)^{9/16}\right]^{8/27}}
# \right]^2
# $$
# </p>
# Both are valid for $Ra\leq10^{12}$. The Nusselt is averaged over the perimeter of the cylinder to account for the angular variation of heat transfer discussed earlier. The heat transfer from natural convection from a heated cylinder of diameter $D$ and length $L$ is
# <p class='alert alert-info'>
# $$
# q=\overline{h}(\pi DL)(T_s-T_\infty)=\frac{1}{R_\text{th,conv}}(T_s-T_\infty)
# $$
# </p>
# where $R_\text{th,conv}$ may computed with <FONT FACE="courier" style="color:blue">R_th_convection(h,A)</FONT>
#
airnatconv = natconv.HorizontalCylinder(correlation='Morgan',Ra=Ra)
print("Morgan correlation: Nu= %1.2f" %airnatconv.Nu)
airnatconv = natconv.HorizontalCylinder(correlation='Churchill-Chu',Ra=Ra,Pr=airflow.Pr)
print("Churchill-Chu correlation: Nu= %1.2f" %airnatconv.Nu)
# +
font = {'family' : 'serif',
#'color' : 'black',
'weight' : 'normal',
'size' : 14,
}
from matplotlib.ticker import FormatStrFormatter
plt.rc('font', **font)
N = 100
Ra = np.logspace(5,12,N)
Nu_Morgan = np.zeros(N)
Nu_ChurchillChu = np.zeros(N)
Pr = 1.0
for i in range(N):
flow = natconv.HorizontalCylinder(correlation='Morgan',Ra=Ra[i])
Nu_Morgan[i] = flow.Nu
flow = natconv.HorizontalCylinder(correlation='Churchill-Chu',Ra=Ra[i],Pr=Pr)
Nu_ChurchillChu[i] = flow.Nu
plt.loglog(Ra,Nu_Morgan, label = r"Morgan",lw = 2)
plt.loglog(Ra,Nu_ChurchillChu, label = r"Churchill-Chu", lw= 2)
plt.xlabel(r"$Ra$")
plt.ylabel(r"$Nu$")
plt.legend(loc=3, bbox_to_anchor=[0., 1.01], ncol=2, shadow=False, fancybox=True)
plt.show()
# -
plt.plot(Ra,np.abs(Nu_Morgan-Nu_ChurchillChu)/Nu_ChurchillChu,lw = 2)
plt.xlabel(r"$Ra$")
plt.ylabel(r"$\vert Nu_{M}-Nu_{CC}\vert/Nu_{CC}$")
plt.show()
# ## Assignment
# <ol>
# <li> Read this entire notebook. Using the textbook, add restrictions and range of validity for the above correlations when applicable. Add the entry length Nu correlation for laminar flow</li>
# <li> Add a section on entrance flow</li>
# <li> How should the entrance flow region be treated in turbulent flows?</li>
# <li>Solve 8.31, 8.36, 8.43</li>
# </ol>
# ### 8.31
# <img src="figures/probun_08_07.jpg" alt="my awesome sketch" width=50% >
# To cool a summer home without using a vapor-compression refrigeration cycle, air is routed through a plastic pipe ($k=0.15\text{ W/m.K}$, $D_i=0.15\text{ m}$, $D_o=0.17\text{ m}$) that is submerged in an adjoini
# ng body of water. The water temperature is nominally at $T_\infty= 17^\circ\text{C}$, and a convection coefficient of $h_o\approx 1500\text{ W/m$^2$. K}$ is maintained at the outer surface of the pipe.
#
# If air from the home enters the pipe at a temperature of $T_{m,i}= 29^\circ\text{C}$ and a volumetric flow rate of $\dot{\forall}_i= 0.025\text{ m$^3$/s}$, what pipe length $L$ is needed to provide a discharge temperature of $T_{m,o}=21^\circ\text{C}$? What is the fan power required
# to move the air through this length of pipe if its inner surface is smooth?
#
# #### Solution
#
# The length of the pipe is the given by solving
# $$
# \frac{T_\infty-T_{m,o}}{T_\infty-T_{m,i}}=\exp\left(-\frac{1}{\dot{m}C_pR_\text{tot}}\right)
# $$
# for the target outlet temperature $T_{m,o}$. First, assuming 1D, steady convection on the outside of the pipe, we must solve for $R'_{tot}$. Since
# $$
# R_{tot}=\frac{R'_{tot}}{L}
# $$
# the pipe length is
# $$
# L=-\dot{m}C_pR'_\text{tot}\ln\frac{T_\infty-T_{m,o}}{T_\infty-T_{m,i}}
# $$
# +
from Libraries import HT_thermal_resistance as res
Rp = []
Rp.append(res.Resistance("$R'_{conv,i}$","W/m"))
Rp.append(res.Resistance("$R'_{cond,pipe}$","W/m"))
Rp.append(res.Resistance("$R'_{conv,o}$","W/m"))
d = schem.Drawing()
d.add(e.DOT, label = r"$T_{m,i}$")
d.add(e.RES, d = 'right', label = Rp[0].name)
d.add(e.DOT, label = r"$T_{s,i}$")
R1 = d.add(e.RES, d = 'right', label = Rp[1].name)
d.add(e.DOT, label = r"$T_{s,o}$")
d.add(e.RES, d='right', label = Rp[2].name)
d.add(e.DOT, label="$T_\infty$")
L1 = d.add(e.LINE, toplabel = "$q'$", endpts = [[-2.25, 0], [-0.25, 0]])
d.labelI(L1, arrowofst = 0)
d.draw()
# +
from Libraries import thermodynamics as thermo
from Libraries import HT_internal_convection as intconv
k_pipe = 0.15 #W/m.K
Di = 0.15 #m
Do = 0.17 #m
T_infty = 17. #C
h_o = 1500 #W/m^2.K
T_mi = 29 #C
T_mo = 21 #C
Qdot = 0.025 #m^3/s
T_m = (T_mi + T_mo)/2
airi = thermo.Fluid('air',T_mi,"C")
airm = thermo.Fluid('air', T_m,"C")
airflow = intconv.PipeFlow(D=Di, L = 1., mdot = airi.rho*Qdot, nu = airm.nu, rho = airi.rho)
airflow.Dittus_Boelter(mode='cooling',Pr=airm.Pr)
print("Re=%.0f" %airflow.Re)
print("Nu=%.0f" %airflow.Nu)
hbar_i = airflow.Nu*airm.k/Di
print("hbar,i=%.2f W/m^2.K" %hbar_i)
Rp[0].convection(hbar_i,np.pi*Di)
Rp[1].cond_cylinder(k = k_pipe,ra=Di,rb=Do,L=1)
Rp[2].convection(400,A=np.pi*Do)
Rptot = 0
for i in range(3):
Rptot += Rp[i].R
# def L_given_other_params(T_infty,T_mo,T_mi,mdot,Cp,Rptot):
# return -mdot*Cp*Rptot*np.log((T_infty -T_mo)/(T_infty - T_mi))
# L = L_given_other_params(T_infty,T_mo,T_mi,airi.rho*Qdot,airm.Cp,Rptot)
L = intconv.L_given_other_params(T_infty,T_mo,T_mi,airi.rho*Qdot,airm.Cp,Rptot)
print("Length needed to achieve T_mo=%.0f C is %.1f m" %(T_mo,L))
# -
from Libraries import HT_natural_convection as natconv
T_f = (T_infty + T_m)/2
water = thermo.Fluid("water",T_f,"C")
Ra = natconv.Ra(beta=water.beta,DT=T_m - T_infty, D=Do,nu=water.nu,alpha = water.alpha)
print("Ra=%.2e" %(Ra))
waterconv = natconv.HorizontalCylinder("Churchill-Chu",Ra,water.Pr)
print("Nu=%.0f" %waterconv.Nu)
print("For natural convection, h_o=%.0f W/m^2.K" %(waterconv.Nu*water.k/Do))
# waterforced = extconv.CircularCylinder()
# This little exercise demonstrates that natural convection does not achieve the cooling capacity assumed in the problem ($h_o=1500\mathrm{W}/\mathrm{m}^2.K$)
from Libraries import HT_natural_convection as natconv
# ?natconv.HorizontalCylinder
# ### 8.36
#
# Hot water at mean temperature $T_m=50\text{$^\circ$C}$ is routed from one building in which it is generated to an adjoining building in which it is used for space heating. Transfer between the buildings occurs in a steel pipe ($k=60\text{ W/m.K}$) of $100 \text{ mm}$ outside diameter and 8-mm wall thickness. During the winter, representative environmental conditions involve air at $T_\infty= -5^\circ \mathrm{C}$ and $V_\infty=3\text{ m/s}$ in cross flow over the pipe.
# Using the Churchill Bernstein and Dittus Boehler correlations, calculate the total heat transfer rate <b>per unit length</b> $q'$, the daily energy cost $Q'=q'\times 24\text{ h/d}$ per meter and the cost per day and per meter assuming an electricity cost of $\text{\$}0.05\text{/kW.h}$.
#
# **FYI:** This is the Churchill-Bernstein correlation which you can call with the `from Libraries import HT_external_convection as extconv` `airflow=extconv.CircularCylinder('Churchill-Bernstein',Re,Pr)`
# $$
# Nu_D = \frac{hD}{k_f}=0.3+\frac{0.62Re_D^{1/2}Pr^{1/3}}{\left[1+\left(\frac{0.4}{Pr}\right)^{2/3}\right]^{1/4}}\left[1+\left(\frac{Re_D}{282,000}\right)^{5/8}\right]^{4/5}
# $$
#
# <img src="figures/PB8.36-sketch.png" alt="my awesome sketch" width=100% >
# The heat transfer problem in any cross sectional area of the pipe is
# $$
# q' = \frac{T_m - T _\infty}{R'_{tot}}
# $$
#
# with
#
# $$
# R'_{tot}= R'_{conv,int} + R'_{cond,p}+R'_{conv,ext}
# $$
#
# We must find the convection coefficients $h_{int}$ and $h_{ext}$, using the appropriate correlations.
# +
Tm = 50 #C
Um = 0.5 #m/s
Di = 0.084 #m
Do = 0.1 #m
kp = 60 #W/m.K
T_infty = -5 #C
U_infty = 3 #m/s
from Libraries import HT_thermal_resistance as res
Rp = []
Rp.append(res.Resistance("$R'_{conv,int}","W/m"))
Rp.append(res.Resistance("$R'_{cond,p}","W/m"))
Rp.append(res.Resistance("$R'_{conv,ext}","W/m"))
# internal convection
from Libraries import thermodynamics as thermo
from Libraries import HT_internal_convection as intconv
water = thermo.Fluid('water',Tm,"C")
pipeflow = intconv.PipeFlow(D=Di,L=1,Um=Um,nu=water.nu)
print("Re_D_pipe= %.0f" %pipeflow.Re)
pipeflow.Dittus_Boelter(mode='cooling',Pr=water.Pr)
hint = pipeflow.Nu*water.k/Di
print("hint=%.1f W/m^2.K" %hint)
Rp[0].convection(h=hint,A=np.pi*Di)
# conduction
Rp[1].cond_cylinder(k=kp,ra=Di,rb=Do,L=1.)
# external convection
#guess for surface temperature at D=Do
T_so = 49.21 #C
T_f = (T_infty + T_so)/2
air = thermo.Fluid('air',T_f,"C")
Re_air = U_infty * Do/air.nu
# print(Re_air)
from Libraries import HT_external_convection as extconv
airflow = extconv.CircularCylinder('Churchill-Bernstein',Re_air,air.Pr)
hext = airflow.Nu*air.k/Do
print("hext=%.1f W/m^2.K" %hext)
Rp[2].convection(h=hext,A=np.pi*Do)
# total thermal resistance
Rptot = 0.
for i in range(3):
Rptot += Rp[i].R
qp = (Tm - T_infty)/Rptot
print("Heat rate per unit length: %.0f W/m" %qp)
#New estimate of T_so
T_so = T_infty + qp*Rp[2].R
print("New T_so = %.2f C" %T_so)
# +
Tm = 50 #C
Um = 0.5 #m/s
Di = 0.084 #m
Do = 0.1 #m
kp = 60 #W/m.K
T_infty = -5 #C
U_infty = 3 #m/s
from Libraries import HT_thermal_resistance as res
Rp = []
Rp.append(res.Resistance("$R'_{conv,int}","W/m"))
Rp.append(res.Resistance("$R'_{cond,p}","W/m"))
Rp.append(res.Resistance("$R'_{conv,ext}","W/m"))
# internal convection
from Libraries import thermodynamics as thermo
from Libraries import HT_internal_convection as intconv
water = thermo.Fluid('water',Tm,"C")
pipeflow = intconv.PipeFlow(D=Di,L=1,Um=Um,nu=water.nu)
print("Re_D_pipe= %.0f" %pipeflow.Re)
pipeflow.Dittus_Boelter(mode='cooling',Pr=water.Pr)
hint = pipeflow.Nu*water.k/Di
print("hint=%.1f W/m^2.K" %hint)
Rp[0].convection(h=hint,A=np.pi*Di)
# conduction
Rp[1].cond_cylinder(k=kp,ra=Di,rb=Do,L=1.)
# external convection
# initial guess for surface temperature at D=Do
T_so = 0. #C
errT = np.inf
iteration = 0
while (errT > 1.0) and (iteration < 10):
iteration += 1
T_so_old = T_so
T_f = (T_infty + T_so)/2
air = thermo.Fluid('air',T_f,"C")
Re_air = U_infty * Do/air.nu
# print(Re_air)
from Libraries import HT_external_convection as extconv
airflow = extconv.CircularCylinder('Churchill-Bernstein',Re_air,air.Pr)
hext = airflow.Nu*air.k/Do
print("hext=%.1f W/m^2.K" %hext)
Rp[2].convection(h=hext,A=np.pi*Do)
# total thermal resistance
Rptot = 0.
for i in range(3):
Rptot += Rp[i].R
qp = (Tm - T_infty)/Rptot
print("Heat rate per unit length: %.0f W/m" %qp)
#New estimate of T_so
T_so = T_infty + qp*Rp[2].R
print("New T_so = %.2f C" %T_so)
errT = abs(T_so - T_so_old)
print("errT=%.3e" %errT)
# -
Qp = qp*1e-3*24
print("Daily energy loss: %.3f kW.h/d/m" %Qp)
Cp = Qp * 0.05
print("Cost: $%.3f /m.d " %Cp)
# ### 8.42
# Atmospheric air enters a $10\text{ m}$-long, $150\text{ mm}$-diameter uninsulated heating duct at $60\text{$^\circ$C}$ and $0.04\text{ kg/s}$. The duct surface temperature is approximately constant at $Ts=15\text{$^\circ$C}$.
#
# (a) What are the outlet air temperature, the heat rate q, and pressure drop $\Delta p$ for these conditions?
#
# (b) To illustrate the tradeoff between heat transfer rate and pressure drop considerations, calculate $q$ and $\Delta p$ for diameters in the range from $0.1$ to $0.2\text{ m}$. In your analysis, maintain the total surface area,
# $A_s=\pi DL$, at the value computed for part (a). Plot $q$, $\Delta p$, and $L$ as a function of the duct diameter.
# +
Tm = 50 #C
Um = 0.5 #m/s
Di = 0.084 #m
Do = 0.1 #m
kp = 60 #W/m.K
T_infty = -5 #C
U_infty = 3 #m/s
from Libraries import HT_thermal_resistance as res
Rp = []
Rp.append(res.Resistance("$R'_{conv,int}","W/m"))
Rp.append(res.Resistance("$R'_{cond,p}","W/m"))
Rp.append(res.Resistance("$R'_{conv,ext}","W/m"))
# internal conduction
from Libraries import HT_internal_convection as intconv
water = thermo.Fluid('water',Tm,"C")
pipeflow = intconv.PipeFlow(D=Di,L=1,Um=Um,nu=water.nu)
print(pipeflow.Re,water.Pr)
pipeflow.Dittus_Boelter(mode='cooling',Pr=water.Pr)
print(pipeflow.Nu*water.k/Di)
Rp[0].convection(h=pipeflow.Nu*water.k/Di,A=np.pi*Di)
#conduction
Rp[1].cond_cylinder(k=kp,ra=Di,rb=Do)
# external convection
from Libraries import HT_external_convection as extconv
T_so = 49.2
T_fo = (T_infty + T_so)/2
air = thermo.Fluid('air',T_fo,"C")
Re_air = U_infty*Do/air.nu
airflow = extconv.CircularCylinder('Churchill-Bernstein',Re_air,air.Pr)
Rp[2].convection(airflow.Nu*air.k/Do,np.pi*Do)
print(airflow.Nu*air.k/Do)
Rptot = 0
for i in range(3):
Rptot += Rp[i].R
print(Rp[i].R)
qp = (Tm - T_infty)/Rptot
print(qp)
T_so_1 = T_infty + qp*Rp[2].R
print(T_so_1)
| Heat-transfer-pipes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
# # 4 Pre-Processing and Training Data<a id='4_Pre-Processing_and_Training_Data'></a>
# ## 4.1 Contents<a id='4.1_Contents'></a>
# * [4 Pre-Processing and Training Data](#4_Pre-Processing_and_Training_Data)
# * [4.1 Contents](#4.1_Contents)
# * [4.2 Introduction](#4.2_Introduction)
# * [4.3 Imports](#4.3_Imports)
# * [4.4 Load Data](#4.4_Load_Data)
# * [4.5 Extract Fl State Data](#4.5_Fl_State_Data)
# * [4.6 Train/Test Split](#4.6_Train/Test_Split)
# * [4.7 Initial Not-Even-A-Model](#4.7_Initial_Not-Even-A-Model)
# * [4.7.1 Metrics](#4.7.1_Metrics)
# * [4.7.1.1 R-squared, or coefficient of determination](#4.7.1.1_R-squared,_or_coefficient_of_determination)
# * [4.7.1.2 Mean Absolute Error](#4.7.1.2_Mean_Absolute_Error)
# * [4.7.1.3 Mean Squared Error](#4.7.1.3_Mean_Squared_Error)
# * [4.7.2 sklearn metrics](#4.7.2_sklearn_metrics)
# * [4.7.2.0.1 R-squared](#4.7.2.0.1_R-squared)
# * [4.7.2.0.2 Mean absolute error](#4.7.2.0.2_Mean_absolute_error)
# * [4.7.2.0.3 Mean squared error](#4.7.2.0.3_Mean_squared_error)
# * [4.7.3 Note On Calculating Metrics](#4.7.3_Note_On_Calculating_Metrics)
# * [4.8 Initial Models](#4.8_Initial_Models)
# * [4.8.1 Imputing missing feature (predictor) values](#4.8.1_Imputing_missing_feature_(predictor)_values)
# * [4.8.1.1 Impute missing values with median](#4.8.1.1_Impute_missing_values_with_median)
# * [4.8.1.1.1 Learn the values to impute from the train set](#4.8.1.1.1_Learn_the_values_to_impute_from_the_train_set)
# * [4.8.1.1.2 Apply the imputation to both train and test splits](#4.8.1.1.2_Apply_the_imputation_to_both_train_and_test_splits)
# * [4.8.1.1.3 Scale the data](#4.8.1.1.3_Scale_the_data)
# * [4.8.1.1.4 Train the model on the train split](#4.8.1.1.4_Train_the_model_on_the_train_split)
# * [4.8.1.1.5 Make predictions using the model on both train and test splits](#4.8.1.1.5_Make_predictions_using_the_model_on_both_train_and_test_splits)
# * [4.8.1.1.6 Assess model performance](#4.8.1.1.6_Assess_model_performance)
# * [4.8.1.2 Impute missing values with the mean](#4.8.1.2_Impute_missing_values_with_the_mean)
# * [4.8.1.2.1 Learn the values to impute from the train set](#4.8.1.2.1_Learn_the_values_to_impute_from_the_train_set)
# * [4.8.1.2.2 Apply the imputation to both train and test splits](#4.8.1.2.2_Apply_the_imputation_to_both_train_and_test_splits)
# * [4.8.1.2.3 Scale the data](#4.8.1.2.3_Scale_the_data)
# * [4.8.1.2.4 Train the model on the train split](#4.8.1.2.4_Train_the_model_on_the_train_split)
# * [4.8.1.2.5 Make predictions using the model on both train and test splits](#4.8.1.2.5_Make_predictions_using_the_model_on_both_train_and_test_splits)
# * [4.8.1.2.6 Assess model performance](#4.8.1.2.6_Assess_model_performance)
# * [4.8.2 Pipelines](#4.8.2_Pipelines)
# * [4.8.2.1 Define the pipeline](#4.8.2.1_Define_the_pipeline)
# * [4.8.2.2 Fit the pipeline](#4.8.2.2_Fit_the_pipeline)
# * [4.8.2.3 Make predictions on the train and test sets](#4.8.2.3_Make_predictions_on_the_train_and_test_sets)
# * [4.8.2.4 Assess performance](#4.8.2.4_Assess_performance)
# * [4.9 Refining The Linear Model](#4.9_Refining_The_Linear_Model)
# * [4.9.1 Define the pipeline](#4.9.1_Define_the_pipeline)
# * [4.9.2 Fit the pipeline](#4.9.2_Fit_the_pipeline)
# * [4.9.3 Assess performance on the train and test set](#4.9.3_Assess_performance_on_the_train_and_test_set)
# * [4.9.4 Define a new pipeline to select a different number of features](#4.9.4_Define_a_new_pipeline_to_select_a_different_number_of_features)
# * [4.9.5 Fit the pipeline](#4.9.5_Fit_the_pipeline)
# * [4.9.6 Assess performance on train and test data](#4.9.6_Assess_performance_on_train_and_test_data)
# * [4.9.7 Assessing performance using cross-validation](#4.9.7_Assessing_performance_using_cross-validation)
# * [4.9.8 Hyperparameter search using GridSearchCV](#4.9.8_Hyperparameter_search_using_GridSearchCV)
# * [4.10 Random Forest Model](#4.10_Random_Forest_Model)
# * [4.10.1 Define the pipeline](#4.10.1_Define_the_pipeline)
# * [4.10.2 Fit and assess performance using cross-validation](#4.10.2_Fit_and_assess_performance_using_cross-validation)
# * [4.10.3 Hyperparameter search using GridSearchCV](#4.10.3_Hyperparameter_search_using_GridSearchCV)
# * [4.11 Final Model Selection](#4.11_Final_Model_Selection)
# * [4.11.1 Linear regression model performance](#4.11.1_Linear_regression_model_performance)
# * [4.11.2 Random forest regression model performance](#4.11.2_Random_forest_regression_model_performance)
# * [4.11.3 Conclusion](#4.11.3_Conclusion)
# * [4.12 Data quantity assessment](#4.12_Data_quantity_assessment)
# * [4.13 Save best model object from pipeline](#4.13_Save_best_model_object_from_pipeline)
# * [4.14 Summary](#4.14_Summary)
#
# ## 4.2 Introduction<a id='4.2_Introduction'></a>
# In this notebook I'll start to build machine learning models. Before even starting with learning a machine learning model,
# however, start by considering how useful the mean value is as a predictor. This is more than just a pedagogical device.
# You never want to go to stakeholders with a machine learning model only to have the CEO point out that it performs worse
# than just guessing the average! Your first model is a baseline performance comparitor for any subsequent model. You then
# build up the process of efficiently and robustly creating and assessing models against it. The development we lay out may
# be little slower than in the real world, but this step of the capstone is definitely more than just instructional. It is
# good practice to build up an understanding that the machine learning pipelines you build work as expected. You can validate
# steps with your own functions for checking expected equivalence between, say, pandas and sklearn implementations.
# ## 4.3 Imports<a id='4.3_Imports'></a>
import pandas as pd
import numpy as np
import os
import pickle
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn import __version__ as sklearn_version
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
from sklearn.model_selection import train_test_split, cross_validate, GridSearchCV, learning_curve
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.dummy import DummyRegressor
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.feature_selection import SelectKBest, f_regression
import datetime
from library.sb_utils import save_file
np.seterr(divide='ignore', invalid='ignore') # Ingnore division by zero or NaN
# ## 4.4 Load Data<a id='4.4_Load_Data'></a>
job_data = pd.read_csv('numer_data_step3_features.csv')
job_data.head().T
# ## 4.5 Extract Fl State data<a id='4.5_Fl_State_Data'></a>
FL_job = job_data[job_data.State == 'FL']
FL_job.T
job_data.shape
job_data =job_data[job_data.State != 'FL']
job_data.shape
# ## 4.6 Train/Test Split<a id='4.6_Train/Test_Split'></a>
len(job_data) * .7, len(job_data) * .3
X_train, X_test, y_train, y_test = train_test_split(job_data.drop(columns='Ave_Salary'),
job_data.Ave_Salary, test_size=0.3,
random_state=47)
X_train.shape, X_test.shape
y_train.shape, y_test.shape
#Code task 1#
#Save the 'state', and 'City' columns from the train/test data into names_train and names_test
#Then drop those columns from `X_train` and `X_test`. Use 'inplace=True'
names_list = ['State','City']
names_train = X_train[names_list]
names_test = X_test[names_list ]
X_train.drop(columns=names_list, inplace=True)
X_test.drop(columns=names_list, inplace=True)
X_train.shape, X_test.shape
#Code task 2#
#Check the `dtypes` attribute of `X_train` to verify all features are numeric
X_train.dtypes
#Code task 3#
#Repeat this check for the test split in `X_test`
X_test.dtypes
# ## 4.7 Initial Not-Even-A-Model<a id='4.7_Initial_Not-Even-A-Model'></a>
# A good place to start is to see how good the mean is as a predictor. In other words, what if you simply say your best guess is the average price?
#Code task 4#
#Calculate the mean of `y_train`
train_mean = y_train.mean()
train_mean
#Code task 5#
#Fit the dummy regressor on the training data
#Hint, call its `.fit()` method with `X_train` and `y_train` as arguments
#Then print the object's `constant_` attribute and verify it's the same as the mean above
dumb_reg = DummyRegressor(strategy='mean')
dumb_reg.fit(X_train, y_train)
dumb_reg.constant_
# ### 4.7.1 Metrics<a id='4.7.1_Metrics'></a>
# #### 4.7.1.1 R-squared, or coefficient of determination<a id='4.7.1.1_R-squared,_or_coefficient_of_determination'></a>
# One measure is $R^2$, the [coefficient of determination](https://en.wikipedia.org/wiki/Coefficient_of_determination).
# This is a measure of the proportion of variance in the dependent variable (our average salary) that is predicted by our
# "model". The linked Wikipedia articles gives a nice explanation of how negative values can arise. This is frequently a
# cause of confusion for newcomers who, reasonably, ask how can a squared value be negative?
#
# Recall the mean can be denoted by $\bar{y}$, where
#
# $$\bar{y} = \frac{1}{n}\sum_{i=1}^ny_i$$
#
# and where $y_i$ are the individual values of the dependent variable.
#
# The total sum of squares (error), can be expressed as
#
# $$SS_{tot} = \sum_i(y_i-\bar{y})^2$$
#
# The above formula should be familiar as it's simply the variance without the denominator to scale (divide) by the sample size.
#
# The residual sum of squares is similarly defined to be
#
# $$SS_{res} = \sum_i(y_i-\hat{y})^2$$
#
# where $\hat{y}$ are our predicted values for the depended variable.
#
# The coefficient of determination, $R^2$, here is given by
#
# $$R^2 = 1 - \frac{SS_{res}}{SS_{tot}}$$
#
# Putting it into words, it's one minus the ratio of the residual variance to the original variance. Thus, the baseline model here, which always predicts $\bar{y}$, should give $R^2=0$. A model that perfectly predicts the observed values would have no residual error and so give $R^2=1$. Models that do worse than predicting the mean will have increased the sum of squares of residuals and so produce a negative $R^2$.
#Code task 6#
#Calculate the R^2
def r_squared(y, ypred):
"""R-squared score.
Calculate the R-squared, or coefficient of determination, of the input.
Arguments:
y -- the observed values
ypred -- the predicted values
"""
ybar = np.sum(y) / len(y) #yes, we could use np.mean(y)
sum_sq_tot = np.sqrt((y - ybar)**2) #total sum of squares error
sum_sq_res = np.sqrt((y - ypred)**2) #residual sum of squares error
R2 = 1.0 - sum_sq_res /sum_sq_tot
return R2
# Make your predictions by creating an array of length the size of the training set with the single value of the mean.
y_tr_pred_ = train_mean * np.ones(len(y_train))
y_tr_pred_[:5]
# Remember the sklearn dummy regressor?
y_tr_pred = dumb_reg.predict(X_train)
y_tr_pred[:5]
# You can see that DummyRegressor produces exactly the same results and saves you having to mess about broadcasting the mean (or whichever other statistic we used - check out the documentation to see what's available) to an array of the appropriate length. It also gives you an object with fit() and predict() methods as well so you can use them as conveniently as any other sklearn estimator.
r_squared(y_train, y_tr_pred)
y_te_pred = train_mean * np.ones(len(y_test))
r_squared(y_test, y_te_pred)
# Generally, you can expect performance on a test set to be slightly worse than on the training set. As you are getting an $R^2$ of zero on the training set, there's nowhere to go but negative!
# $R^2$ is a common metric, and interpretable in terms of the amount of variance explained, it's less appealing if you want an idea of how "close" your predictions are to the true values. Metrics that summarise the difference between predicted and actual values are _mean absolute error_ and _mean squared error_.
# #### 4.7.1.2 Mean Absolute Error<a id='4.7.1.2_Mean_Absolute_Error'></a>
# This is very simply the average of the absolute errors:
#
# $$MAE = \frac{1}{n}\sum_i^n|y_i - \hat{y}|$$
#Code task 7#
#Calculate the MAE as defined above
def mae(y, ypred):
"""Mean absolute error.
Calculate the mean absolute error of the arguments
Arguments:
y -- the observed values
ypred -- the predicted values
"""
abs_error = np.abs(y - ypred)
mae = np.mean(abs_error)
return mae
mae(y_train, y_tr_pred)
mae(y_test, y_te_pred)
# Mean absolute error is arguably the most intuitive of all the metrics, this essentially tells you that, on average, you might
# expect to be off by around \\32 if you guessed average salary based on an average of known values.
# #### 4.7.1.3 Mean Squared Error<a id='4.7.1.3_Mean_Squared_Error'></a>
# Another common metric (and an important one internally for optimizing machine learning models) is the mean squared error. This is simply the average of the square of the errors:
#
# $$MSE = \frac{1}{n}\sum_i^n(y_i - \hat{y})^2$$
#Code task 8#
#Calculate the MSE as defined above
def mse(y, ypred):
"""Mean square error.
Calculate the mean square error of the arguments
Arguments:
y -- the observed values
ypred -- the predicted values
"""
sq_error = (y - ypred)**2
mse = np.mean(sq_error)
return mse
mse(y_train, y_tr_pred)
mse(y_test, y_te_pred)
# So here, you get a slightly better MSE on the test set than you did on the train set. And what does a squared error mean anyway? To convert this back to our measurement space, we often take the square root, to form the root mean square error thus:
np.sqrt([mse(y_train, y_tr_pred), mse(y_test, y_te_pred)])
# ### 4.7.2 sklearn metrics<a id='4.7.2_sklearn_metrics'></a>
# Functions are good, but you don't want to have to define functions every time we want to assess performance.
# `sklearn.metrics` provides many commonly used metrics, included the ones above.
# ##### 4.7.2.0.1 R-squared<a id='4.7.2.0.1_R-squared'></a>
r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
# ##### 4.7.2.0.2 Mean absolute error<a id='4.7.2.0.2_Mean_absolute_error'></a>
mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
# ##### 4.7.2.0.3 Mean squared error<a id='4.7.2.0.3_Mean_squared_error'></a>
mean_squared_error(y_train, y_tr_pred), mean_squared_error(y_test, y_te_pred)
# ### 4.7.3 Note On Calculating Metrics<a id='4.7.3_Note_On_Calculating_Metrics'></a>
# When calling functions to calculate metrics, it is important to take care in the order of the arguments. Two of the metrics above actually don't care if the arguments are reversed; one does. Which one cares?
# In a Jupyter code cell, running `r2_score?` will bring up the docstring for the function, and `r2_score??` will bring up the actual code of the function! Try them and compare the source for `sklearn`'s function with yours. Feel free to explore what happens when you reverse the order of the arguments and compare behaviour of `sklearn`'s function and yours.
# train set - sklearn
# correct order, incorrect order
r2_score(y_train, y_tr_pred), r2_score(y_tr_pred, y_train)
# test set - sklearn
# correct order, incorrect order
r2_score(y_test, y_te_pred), r2_score(y_te_pred, y_test)
# train set - using our homebrew function
# correct order, incorrect order
r_squared(y_train, y_tr_pred), r_squared(y_tr_pred, y_train)
# test set - using our homebrew function
# correct order, incorrect order
r_squared(y_test, y_te_pred), r_squared(y_te_pred, y_test)
# ## 4.8 Initial Models<a id='4.8_Initial_Models'></a>
# ### 4.8.1 Imputing missing feature (predictor) values<a id='4.8.1_Imputing_missing_feature_(predictor)_values'></a>
# Recall when performing EDA, you imputed (filled in) some missing values in pandas. You did this judiciously for exploratory/visualization purposes. You left many missing values in the data. You can impute missing values using scikit-learn, but note that you should learn values to impute from a train split and apply that to the test split to then assess how well your imputation worked.
#
# #### 4.8.1.1 Impute missing values with median<a id='4.8.1.1_Impute_missing_values_with_median'></a>
# There's missing values. Recall from your data exploration that many distributions were skewed. Your first thought might be to impute missing values using the median.
# ##### 4.8.1.1.1 Learn the values to impute from the train set<a id='4.8.1.1.1_Learn_the_values_to_impute_from_the_train_set'></a>
# These are the values we'll use to fill in any missing values
X_defaults_median = X_train.median()
X_defaults_median
# ##### 4.8.1.1.2 Apply the imputation to both train and test splits<a id='4.8.1.1.2_Apply_the_imputation_to_both_train_and_test_splits'></a>
#Code task 9#
#Call `X_train` and `X_test`'s `fillna()` method, passing `X_defaults_median` as the values to use
#Assign the results to `X_tr` and `X_te`, respectively
X_tr = X_train.fillna(X_defaults_median)
X_te = X_test.fillna(X_defaults_median)
# ##### 4.8.1.1.3 Scale the data<a id='4.8.1.1.3_Scale_the_data'></a>
# As you have features measured in many different units, with numbers that vary by orders of magnitude, start off by scaling them to put them all on a consistent scale. The StandardScaler scales each feature to zero mean and unit variance.
#Code task 10#
#Call the StandardScaler`s fit method on `X_tr` to fit the scaler
#then use it's `transform()` method to apply the scaling to both the train and test split
#data (`X_tr` and `X_te`), naming the results `X_tr_scaled` and `X_te_scaled`, respectively
scaler = StandardScaler()
scaler.fit(X_tr)
X_tr_scaled = scaler.transform(X_tr)
X_te_scaled = scaler.transform(X_te)
# ##### 4.8.1.1.4 Train the model on the train split<a id='4.8.1.1.4_Train_the_model_on_the_train_split'></a>
lm = LinearRegression().fit(X_tr_scaled, y_train)
# ##### 4.8.1.1.5 Make predictions using the model on both train and test splits<a id='4.8.1.1.5_Make_predictions_using_the_model_on_both_train_and_test_splits'></a>
#Code task 11#
#Call the `predict()` method of the model (`lm`) on both the (scaled) train and test data
#Assign the predictions to `y_tr_pred` and `y_te_pred`, respectively
y_tr_pred = lm.predict(X_tr_scaled)
y_te_pred = lm.predict(X_te_scaled)
# ##### 4.8.1.1.6 Assess model performance<a id='4.8.1.1.6_Assess_model_performance'></a>
# r^2 - train, test
median_r2 = r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
median_r2
# Recall that you estimated average salary by simply using a known average. As expected, this produced an 𝑅2 of zero for both
# the training and test set, because 𝑅2 tells us how much of the variance you're explaining beyond that of using just the
# mean, and you were using just the mean. Here we see that our simple linear regression model explains over 80% of the variance
# on the train set and over 70% on the test set. Clearly you are onto something, although the much lower value for the test set
# suggests you're overfitting somewhat. This isn't a surprise as you've made no effort to select a parsimonious set of features
# or deal with multicollinearity in our data.
#Code task 12#
#Now calculate the mean absolute error scores using `sklearn`'s `mean_absolute_error` function
# as we did above for R^2
# MAE - train, test
median_mae = mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
median_mae
# Using this model, then, on average you'd expect to estimate a ???? There may be something to this machine learning lark after all!
#Code task 13#
#And also do the same using `sklearn`'s `mean_squared_error`
# MSE - train, test
median_mse = mean_squared_error(y_train, y_tr_pred), mean_squared_error(y_test, y_te_pred)
median_mse
# #### 4.8.1.2 Impute missing values with the mean<a id='4.8.1.2_Impute_missing_values_with_the_mean'></a>
# You chose to use the median for filling missing values because of the skew of many of our predictor feature distributions. What if you wanted to try something else, such as the mean?
# ##### 4.8.1.2.1 Learn the values to impute from the train set<a id='4.8.1.2.1_Learn_the_values_to_impute_from_the_train_set'></a>
#Code task 14#
#As we did for the median above, calculate mean values for imputing missing values
# These are the values we'll use to fill in any missing values
X_defaults_mean = X_train.mean()
X_defaults_mean
# By eye, you can immediately tell that your replacement values are much higher than those from using the median.
# ##### 4.8.1.2.2 Apply the imputation to both train and test splits<a id='4.8.1.2.2_Apply_the_imputation_to_both_train_and_test_splits'></a>
X_tr = X_train.fillna(X_defaults_mean)
X_te = X_test.fillna(X_defaults_mean)
# ##### 4.8.1.2.3 Scale the data<a id='4.8.1.2.3_Scale_the_data'></a>
scaler = StandardScaler()
scaler.fit(X_tr)
X_tr_scaled = scaler.transform(X_tr)
X_te_scaled = scaler.transform(X_te)
# ##### 4.8.1.2.4 Train the model on the train split<a id='4.8.1.2.4_Train_the_model_on_the_train_split'></a>
lm = LinearRegression().fit(X_tr_scaled, y_train)
# ##### 4.8.1.2.5 Make predictions using the model on both train and test splits<a id='4.8.1.2.5_Make_predictions_using_the_model_on_both_train_and_test_splits'></a>
y_tr_pred = lm.predict(X_tr_scaled)
y_te_pred = lm.predict(X_te_scaled)
# ##### 4.8.1.2.6 Assess model performance<a id='4.8.1.2.6_Assess_model_performance'></a>
r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
mean_squared_error(y_train, y_tr_pred), mean_squared_error(y_test, y_te_pred)
# These results don't seem very different to when you used the median for imputing missing values. Perhaps it doesn't make much difference here. Maybe your overtraining dominates. Maybe other feature transformations, such as taking the log, would help. You could try with just a subset of features rather than using all of them as inputs.
#
# To perform the median/mean comparison, you copied and pasted a lot of code just to change the function for imputing missing values. It would make more sense to write a function that performed the sequence of steps:
# 1. impute missing values
# 2. scale the features
# 3. train a model
# 4. calculate model performance
#
# But these are common steps and `sklearn` provides something much better than writing custom functions.
# ### 4.8.2 Pipelines<a id='4.8.2_Pipelines'></a>
# One of the most important and useful components of `sklearn` is the [pipeline](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html). In place of `panda`'s `fillna` DataFrame method, there is `sklearn`'s `SimpleImputer`. Remember the first linear model above performed the steps:
#
# 1. replace missing values with the median for each feature
# 2. scale the data to zero mean and unit variance
# 3. train a linear regression model
#
# and all these steps were trained on the train split and then applied to the test split for assessment.
#
# The pipeline below defines exactly those same steps. Crucially, the resultant `Pipeline` object has a `fit()` method and a `predict()` method, just like the `LinearRegression()` object itself. Just as you might create a linear regression model and train it with `.fit()` and predict with `.predict()`, you can wrap the entire process of imputing and feature scaling and regression in a single object you can train with `.fit()` and predict with `.predict()`. And that's basically a pipeline: a model on steroids.
# #### 4.8.2.1 Define the pipeline<a id='4.8.2.1_Define_the_pipeline'></a>
pipe = make_pipeline(
SimpleImputer(strategy='median'),
StandardScaler(),
LinearRegression()
)
type(pipe)
hasattr(pipe, 'fit'), hasattr(pipe, 'predict')
# #### 4.8.2.2 Fit the pipeline<a id='4.8.2.2_Fit_the_pipeline'></a>
# Here, a single call to the pipeline's fit() method combines the steps of learning the imputation (determining what values to use to fill the missing ones), the scaling (determining the mean to subtract and the variance to divide by), and then training the model. It does this all in the one call with the training data as arguments.
#Code task 15#
#Call the pipe's `fit()` method with `X_train` and `y_train` as arguments
pipe.fit(X_train, y_train)
# #### 4.8.2.3 Make predictions on the train and test sets<a id='4.8.2.3_Make_predictions_on_the_train_and_test_sets'></a>
y_tr_pred = pipe.predict(X_train)
y_te_pred = pipe.predict(X_test)
# #### 4.8.2.4 Assess performance<a id='4.8.2.4_Assess_performance'></a>
r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
# And compare with your earlier (non-pipeline) result:
median_r2
mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
# Compare with your earlier result:
median_mae
mean_squared_error(y_train, y_tr_pred), mean_squared_error(y_test, y_te_pred)
# Compare with your earlier result:
median_mse
# These results confirm the pipeline is doing exactly what's expected, and results are identical to your earlier steps. This allows you to move faster but with confidence.
# ## 4.9 Refining The Linear Model<a id='4.9_Refining_The_Linear_Model'></a>
# You suspected the model was overfitting. This is no real surprise given the number of features you blindly used. It's likely a judicious subset of features would generalize better. sklearn has a number of feature selection functions available. The one you'll use here is SelectKBest which, as you might guess, selects the k best features. You can read about SelectKBest here. f_regression is just the score function you're using because you're performing regression. It's important to choose an appropriate one for your machine learning task.
# ### 4.9.1 Define the pipeline<a id='4.9.1_Define_the_pipeline'></a>
# Redefine your pipeline to include this feature selection step:
#Code task 16#
#Add `SelectKBest` as a step in the pipeline between `StandardScaler()` and `LinearRegression()`
#Don't forget to tell it to use `f_regression` as its score function
pipe = make_pipeline(
SimpleImputer(strategy='median'),
StandardScaler(),
SelectKBest(f_regression),
LinearRegression()
)
# ### 4.9.2 Fit the pipeline<a id='4.9.2_Fit_the_pipeline'></a>
pipe.fit(X_train, y_train)
# ### 4.9.3 Assess performance on the train and test set<a id='4.9.3_Assess_performance_on_the_train_and_test_set'></a>
y_tr_pred = pipe.predict(X_train)
y_te_pred = pipe.predict(X_test)
r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
# This has made things worse! Clearly selecting a subset of features has an impact on performance. `SelectKBest` defaults to k=10. You've just seen that 10 is worse than using all features. What is the best k? You could create a new pipeline with a different value of k:
# ### 4.9.4 Define a new pipeline to select a different number of features<a id='4.9.4_Define_a_new_pipeline_to_select_a_different_number_of_features'></a>
#Code task 17#
#Modify the `SelectKBest` step to use a value of 15 for k
pipe15 = make_pipeline(
SimpleImputer(strategy='median'),
StandardScaler(),
SelectKBest(f_regression, k=15),
LinearRegression()
)
pipe15.fit(X_train, y_train)
# 4.9.6 Assess performance on train and test data
y_tr_pred = pipe15.predict(X_train)
y_te_pred = pipe15.predict(X_test)
r2_score(y_train, y_tr_pred), r2_score(y_test, y_te_pred)
mean_absolute_error(y_train, y_tr_pred), mean_absolute_error(y_test, y_te_pred)
# You could keep going, trying different values of k, training a model, measuring performance on the test set, and then picking the model with the best test set performance. There's a fundamental problem with this approach: _you're tuning the model to the arbitrary test set_! If you continue this way you'll end up with a model works well on the particular quirks of our test set _but fails to generalize to new data_. The whole point of keeping a test set is for it to be a set of that new data, to check how well our model might perform on data it hasn't seen.
#
# The way around this is a technique called _cross-validation_. You partition the training set into k folds, train our model on k-1 of those folds, and calculate performance on the fold not used in training. This procedure then cycles through k times with a different fold held back each time. Thus you end up building k models on k sets of data with k estimates of how the model performs on unseen data but without having to touch the test set.
# ### 4.9.7 Assessing performance using cross-validation<a id='4.9.7_Assessing_performance_using_cross-validation'></a>
cv_results = cross_validate(pipe15, X_train, y_train, cv=5)
cv_scores = cv_results['test_score']
cv_scores
# Without using the same random state for initializing the CV folds, your actual numbers will be different.
np.mean(cv_scores), np.std(cv_scores)
# These results highlight that assessing model performance in inherently open to variability. You'll get different results depending on the quirks of which points are in which fold. An advantage of this is that you can also obtain an estimate of the variability, or uncertainty, in your performance estimate.
np.round((np.mean(cv_scores) - 2 * np.std(cv_scores), np.mean(cv_scores) + 2 * np.std(cv_scores)), 2)
# ### 4.9.8 Hyperparameter search using GridSearchCV<a id='4.9.8_Hyperparameter_search_using_GridSearchCV'></a>
# Pulling the above together, we have:
# * a pipeline that
# * imputes missing values
# * scales the data
# * selects the k best features
# * trains a linear regression model
# * a technique (cross-validation) for estimating model performance
#
# Now you want to use cross-validation for multiple values of k and use cross-validation to pick the value of k that gives the best performance. `make_pipeline` automatically names each step as the lowercase name of the step and the parameters of the step are then accessed by appending a double underscore followed by the parameter name. You know the name of the step will be 'selectkbest' and you know the parameter is 'k'.
#
# You can also list the names of all the parameters in a pipeline like this:
#Code task 18#
#Call `pipe`'s `get_params()` method to get a dict of available parameters and print their names
#using dict's `keys()` method
pipe.get_params().keys()
# The above can be particularly useful as your pipelines becomes more complex (you can even nest pipelines within pipelines).
k = [k+1 for k in range(len(X_train.columns))]
grid_params = {'selectkbest__k': k}
# Now you have a range of k to investigate. Is 1 feature best? 2? 3? 4? All of them? You could write a for loop and iterate over each possible value, doing all the housekeeping oyurselves to track the best value of k. But this is a common task so there's a built in function in sklearn. This is GridSearchCV. This takes the pipeline object, in fact it takes anything with a .fit() and .predict() method. In simple cases with no feature selection or imputation or feature scaling etc. you may see the classifier or regressor object itself directly passed into GridSearchCV. The other key input is the parameters and values to search over. Optional parameters include the cross-validation strategy and number of CPUs to use.
lr_grid_cv = GridSearchCV(pipe, param_grid=grid_params, cv=5, n_jobs=-1)
lr_grid_cv.fit(X_train, y_train)
score_mean = lr_grid_cv.cv_results_['mean_test_score']
score_std = lr_grid_cv.cv_results_['std_test_score']
cv_k = [k for k in lr_grid_cv.cv_results_['param_selectkbest__k']]
#Code task 19#
#Print the `best_params_` attribute of `lr_grid_cv`
lr_grid_cv.best_params_
#Code task 20#
#Assign the value of k from the above dict of `best_params_` and assign it to `best_k`
best_k = lr_grid_cv.best_params_['selectkbest__k']
plt.subplots(figsize=(10, 5))
plt.errorbar(cv_k, score_mean, yerr=score_std)
plt.axvline(x=best_k, c='r', ls='--', alpha=.5)
plt.xlabel('k')
plt.ylabel('CV score (r-squared)')
plt.title('Pipeline mean CV score (error bars +/- 1sd)');
# The above suggests a good value for k is 12. There was an initial rapid increase with k, followed by a slow decline. Also noticeable is the variance of the results greatly increase above k=2. As you increasingly overfit, expect greater swings in performance as different points move in and out of the train/test folds.
#
# Which features were most useful? Step into your best model, shown below. Starting with the fitted grid search object, you get the best estimator, then the named step 'selectkbest', for which you can its get_support() method for a logical mask of the features selected.
selected = lr_grid_cv.best_estimator_.named_steps.selectkbest.get_support()
# Similarly, instead of using the 'selectkbest' named step, you can access the named step for the linear regression model and, from that, grab the model coefficients via its coef_ attribute:
#Code task 21#
#Get the linear model coefficients from the `coef_` attribute and store in `coefs`,
#get the matching feature names from the column names of the dataframe,
#and display the results as a pandas Series with `coefs` as the values and `features` as the index,
#sorting the values in descending order
coefs = lr_grid_cv.best_estimator_.named_steps.linearregression.coef_
features = X_train.columns[selected]
pd.Series(coefs, index=features).sort_values(ascending=False)
# These results suggest that jobs_per_state_x is your biggest positive feature. This makes intuitive sense and is consistent
# with what you saw during the EDA work. Also, you see the state_ave_salary is a strong positive as well.
# People like guaranteed skiing! The state_total_DataB_x is negatively associated with average salary! This seems odd.
# ## 4.10 Random Forest Model<a id='4.10_Random_Forest_Model'></a>
# A model that can work very well in a lot of cases is the random forest. For regression, this is provided by `sklearn`'s `RandomForestRegressor` class.
#
# Time to stop the bad practice of repeatedly checking performance on the test split. Instead, go straight from defining the pipeline to assessing performance using cross-validation. `cross_validate` will perform the fitting as part of the process. This uses the default settings for the random forest so you'll then proceed to investigate some different hyperparameters.
# ### 4.10.1 Define the pipeline<a id='4.10.1_Define_the_pipeline'></a>
#Code task 22#
#Define a pipeline comprising the steps:
#SimpleImputer() with a strategy of 'median'
#StandardScaler(),
#and then RandomForestRegressor() with a random state of 47
RF_pipe = make_pipeline(
SimpleImputer(strategy='median'),
StandardScaler(),
RandomForestRegressor(random_state=47)
)
# ### 4.10.2 Fit and assess performance using cross-validation<a id='4.10.2_Fit_and_assess_performance_using_cross-validation'></a>
#Code task 23#
#Call `cross_validate` to estimate the pipeline's performance.
#Pass it the random forest pipe object, `X_train` and `y_train`,
#and get it to use 5-fold cross-validation
rf_default_cv_results = cross_validate(RF_pipe,X_train,y_train, cv=5)
rf_cv_scores = rf_default_cv_results['test_score']
rf_cv_scores
np.mean(rf_cv_scores), np.std(rf_cv_scores)
# ### 4.10.3 Hyperparameter search using GridSearchCV<a id='4.10.3_Hyperparameter_search_using_GridSearchCV'></a>
n_est = [int(n) for n in np.logspace(start=1, stop=3, num=20)]
grid_params = {
'randomforestregressor__n_estimators': n_est,
'standardscaler': [StandardScaler(), None],
'simpleimputer__strategy': ['mean', 'median']
}
grid_params
#Code task 24#
#Call `GridSearchCV` with the random forest pipeline, passing in the above `grid_params`
#dict for parameters to evaluate, 5-fold cross-validation, and all available CPU cores (if desired)
rf_grid_cv = GridSearchCV(RF_pipe, param_grid=grid_params, cv=5, n_jobs=-1)
#Code task 25#
#Now call the `GridSearchCV`'s `fit()` method with `X_train` and `y_train` as arguments
#to actually start the grid search. This may take a minute or two.
rf_grid_cv.fit(X_train, y_train)
#Code task 26#
#Print the best params (`best_params_` attribute) from the grid search
rf_grid_cv.best_params_
# It looks like imputing with the median helps, but scaling the features doesn't.
rf_best_cv_results = cross_validate(rf_grid_cv.best_estimator_, X_train, y_train, cv=5)
rf_best_scores = rf_best_cv_results['test_score']
rf_best_scores
np.mean(rf_best_scores), np.std(rf_best_scores)
# You've marginally improved upon the default CV results. Random forest has many more hyperparameters you could tune, but we won't dive into that here.
#Code task 27#
#Plot a barplot of the random forest's feature importances,
#assigning the `feature_importances_` attribute of
#`rf_grid_cv.best_estimator_.named_steps.randomforestregressor` to the name `imps` to then
#create a pandas Series object of the feature importances, with the index given by the
#training data column names, sorting the values in descending order
plt.subplots(figsize=(10, 5))
imps = rf_grid_cv.best_estimator_.named_steps.randomforestregressor.feature_importances_
rf_feat_imps = pd.Series(imps, index=X_train.columns).sort_values(ascending=False)
rf_feat_imps.plot(kind='bar')
plt.xlabel('features')
plt.ylabel('importance')
plt.title('Best random forest regressor feature importances');
# Encouragingly, the dominant top four features are in common with your linear model:
# state_ave_salary,Ratings,
# Founded Years,Max_Company_Size,
# Max__USD_Revenue,Python knowledge,
# DataB_knowledge,Math_knowledge,
# SQL_knowledge,
# ETL_knowledge
#
#
#
# ## 4.11 Final Model Selection<a id='4.11_Final_Model_Selection'></a>
# Time to select your final model to use for further business modeling! It would be good to revisit the above model selection;
# there is undoubtedly more that could be done to explore possible hyperparameters.
# It would also be worthwhile to investigate removing the least useful features. Gathering or calculating, and storing, features
# adds business cost and dependencies, so if features genuinely are not needed they should be removed.
# Building a simpler model with fewer features can also have the advantage of being easier to sell (and/or explain) to
# stakeholders.
# Certainly there seem to be four strong features here and so a model using only those would probably work well.
# However, you want to explore some different scenarios where other features vary so keep the fuller
# model for now.
# The business is waiting for this model and you have something that you have confidence in to be much better than guessing
# with the average salary.
#
# Or, rather, you have two "somethings". You built a best linear model and a best random forest model. You need to finally
# choose between them. You can calculate the mean absolute error using cross-validation. Although `cross-validate` defaults to
# the $R^2$ [metric for scoring](https://scikit-learn.org/stable/modules/model_evaluation.html#scoring) regression, you can
# specify the mean absolute error as an alternative via
# the `scoring` parameter.
# ### 4.11.1 Linear regression model performance<a id='4.11.1_Linear_regression_model_performance'></a>
# 'neg_mean_absolute_error' uses the (negative of) the mean absolute error
lr_neg_mae = cross_validate(lr_grid_cv.best_estimator_, X_train, y_train,
scoring='neg_mean_absolute_error', cv=5, n_jobs=-1)
lr_mae_mean = np.mean(-1 * lr_neg_mae['test_score'])
lr_mae_std = np.std(-1 * lr_neg_mae['test_score'])
lr_mae_mean, lr_mae_std
mean_absolute_error(y_test, lr_grid_cv.best_estimator_.predict(X_test))
# ### 4.11.2 Random forest regression model performance<a id='4.11.2_Random_forest_regression_model_performance'></a>
rf_neg_mae = cross_validate(rf_grid_cv.best_estimator_, X_train, y_train,
scoring='neg_mean_absolute_error', cv=5, n_jobs=-1)
rf_mae_mean = np.mean(-1 * rf_neg_mae['test_score'])
rf_mae_std = np.std(-1 * rf_neg_mae['test_score'])
rf_mae_mean, rf_mae_std
mean_absolute_error(y_test, rf_grid_cv.best_estimator_.predict(X_test))
# ### 4.11.3 Conclusion<a id='4.11.3_Conclusion'></a>
# The random forest model has a lower cross-validation mean absolute error by almost 0.24. It also exhibits less variability.
# Verifying performance on the test set produces performance consistent with the cross-validation results.
# ## 4.12 Data quantity assessment<a id='4.12_Data_quantity_assessment'></a>
# Finally, you need to advise the business whether it needs to undertake further data collection. Would more data be useful? We're often led to believe more data is always good, but gathering data invariably has a cost associated with it. Assess this trade off by seeing how performance varies with differing data set sizes. The learning_curve function does this conveniently.
fractions = [.1,.2, .25, .3, .35, .4, .45, .5, .6, 0.7, .8,.9, 1.0]
train_size, train_scores, test_scores = learning_curve(pipe, X_train, y_train, train_sizes=fractions)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
plt.subplots(figsize=(10, 5))
plt.errorbar(train_size, test_scores_mean, yerr=test_scores_std)
plt.xlabel('Training set size')
plt.ylabel('CV scores')
plt.title('Cross-validation score as training set size increases');
# This shows that you seem to have plenty of data. There's an initial rapid improvement in model scores as one would expect, but
# it's essentially levelled off by around a sample size of 1000.
# ## 4.13 Save best model object from pipeline<a id='4.13_Save_best_model_object_from_pipeline'></a>
#Code task 28#
#This may not be "production grade ML deployment" practice, but adding some basic
#information to your saved models can save your bacon in development.
#Just what version model have you just loaded to reuse? What version of `sklearn`
#created it? When did you make it?
#Assign the pandas version number (`pd.__version__`) to the `pandas_version` attribute,
#the numpy version (`np.__version__`) to the `numpy_version` attribute,
#the sklearn version (`sklearn_version`) to the `sklearn_version` attribute,
#and the current datetime (`datetime.datetime.now()`) to the `build_datetime` attribute
#Let's call this model version '1.0'
best_model = rf_grid_cv.best_estimator_
best_model.version = best_model
best_model.pandas_version = pd.__version__
best_model.numpy_version = np.__version__
best_model.sklearn_version = sklearn_version
best_model.X_columns = [col for col in X_train.columns]
best_model.build_datetime = datetime.datetime.now()
# +
# save the model
modelpath = './models'
save_file(best_model, 'DS_model.pkl', modelpath)
# -
# ## 4.14 Summary<a id='4.14_Summary'></a>
# **Q: 1** Write a summary of the work in this notebook. Capture the fact that you gained a baseline idea of performance by
# simply taking the average price and how well that did. Then highlight that you built a linear model and the features that
# found. Comment on the estimate of its performance from cross-validation and whether its performance on the test split was
# consistent with this estimate. Also highlight that a random forest regressor was tried, what preprocessing steps were
# found to be best, and again what its estimated performance via cross-validation was and whether its performance on the
# test set was consistent with that. State which model you have decided to use going forwards and why. This summary should
# provide a quick overview for someone wanting to know quickly why the given model was chosen for the next part of the
# business problem to help guide important business decisions.
# 1)The mean price considering the mean absolute error through cross validation is better for the Random forest regression model
#
# 2) Performance on the test split was more consistent with this estimate for Random forest regression model
#
# 3)If we check Estimated performance via cross-validation, we can see that is better Random forest regression model
# than Linear regression model performance:
#
# 4.11.1 Linear regression model performance
# lr_mae_mean,= 26.29
# lr_mae_std= 0.54
#
# mean_absolute_error= 27.32
#
# 4.11.2 Random forest regression model performance
# rf_mae_mean = 25.75
# rf_mae_std = 0.32
#
# mean_absolute_error=25.57
#
# 4) For these reasons I choose Random forest regression model
| Capstone2_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Kali ini kita akan lakukan klasifikasi dengan menggunakan algoritma DTree. Kita akan buat model dari data historis pasien.
import numpy as np
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
my_data = pd.read_csv("drug200.csv", delimiter=",")
my_data[0:5]
# ### Pre-processing
#
# Kita beri nama my_data untuk nyalin data dari drug.csv dengan 2 variabel utama X sebagai Feature Matrix dan y sebagai response vector (target) . Nah berhubung kolom2 tidak semua berisi nilai numerik jadi kita harus ubah dulu ke nilai numerik, atau terpaksa kita drop/hapus.
X = my_data[['Age', 'Sex', 'BP', 'Cholesterol', 'Na_to_K']].values
X[0:5]
# di Sklearn Decision Trees gak bisa tangani variabel yang kategorial, makanya kita harus konversi nilai2 kategorial dengan pandas.get_dummies().
# +
from sklearn import preprocessing
le_sex = preprocessing.LabelEncoder()
le_sex.fit(['F','M'])
X[:,1] = le_sex.transform(X[:,1])
le_BP = preprocessing.LabelEncoder()
le_BP.fit([ 'LOW', 'NORMAL', 'HIGH'])
X[:,2] = le_BP.transform(X[:,2])
le_Chol = preprocessing.LabelEncoder()
le_Chol.fit([ 'NORMAL', 'HIGH'])
X[:,3] = le_Chol.transform(X[:,3])
X[0:5]
# -
# tambahkan satu kolom dengan nama Drug
y = my_data["Drug"]
y[0:5]
# ### Setting Decision Tree
# Seperti biasa kita mulai dengan import.
from sklearn.model_selection import train_test_split
X_trainset, X_testset, y_trainset, y_testset = train_test_split(X, y, test_size=0.3, random_state=3)
# #### Modeling
# Pertama, kita buat instance dari DecisionTreeClassifier dan kita namakan drugTree
drugTree = DecisionTreeClassifier(criterion="gini", max_depth = 10)
drugTree # it shows the default parameters
drugTree.fit(X_trainset,y_trainset)
# #### Prediction
# Nah sekarang saatnya lakukan prediction berdasarkan testing dataset dan simpan ke predTree
predTree = drugTree.predict(X_testset)
print (predTree [0:5])
print (y_testset [0:5])
# #### Evaluation
from sklearn import metrics
import matplotlib.pyplot as plt
print("Akurasi dengan model DecisionTrees: ", metrics.accuracy_score(y_testset, predTree))
from sklearn.metrics import jaccard_score
jaccard_score(y_testset, predTree,average=None)
| Pertemuan 10/Classification dengan DTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="LS4JpCb_M3pd" colab_type="code" colab={}
# connect google drive with colab
from google.colab import drive
drive.mount('/content/drive/')
# + id="hIyauWbvTLS0" colab_type="code" colab={}
# import what we need
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from keras.preprocessing.image import ImageDataGenerator, load_img, img_to_array, array_to_img, save_img
from tensorflow.keras import layers, models
# + id="JXtrKSBUTYm1" colab_type="code" colab={}
# using pydrive library to download dataset from google drive
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# + id="d0IDD0L8T-fZ" colab_type="code" colab={}
# download the dataset
download = drive.CreateFile({'id': '1vY37AOcpIhugM45niCYnKboONqwkB0a7'})
download.GetContentFile('dataset.zip')
# + id="JPNo4KOWTWPi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c4337ea6-ed3e-4f0b-d6f3-d081be062fdb"
# unzip the dataset
# !unzip /content/dataset.zip -d /content/data
# + id="GcuDh_izTYV7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="87dcf853-a8f4-45f4-a35a-391d7ffac404"
# image data generator fron training data
train_datagen = ImageDataGenerator(rescale=1/255.0)
train_generator = train_datagen.flow_from_directory(
'/content/data/Training_set/Training_set',
target_size=(128, 128),
color_mode='rgb',
batch_size=32,
class_mode='binary')
# + id="vnVTaqvKTaFr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6bd005dc-f1b9-400e-9ca0-ffb3c310b273"
# image data generator fron validation data
val_datagen = ImageDataGenerator(rescale=1/255.0)
val_generator = val_datagen.flow_from_directory(
'/content/data/test/test',
target_size=(128, 128),
color_mode='rgb',
batch_size=16,
class_mode='binary')
# + id="w28hUuWPqodE" colab_type="code" colab={}
# define my model
model = models.Sequential()
model.add(layers.Conv2D(16, (3, 3), activation='relu', input_shape=(128,128,3)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(32, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Flatten())
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
# + id="eugNizKkTiF0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="a1dc1e5a-dab8-4661-dcde-3d6e9c3b7e60"
model.summary()
# + id="mrSrD1MUTkZ0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="849386e5-8097-42c1-944c-2c91ce1c2572"
# plot the model architecture
tf.keras.utils.plot_model(
model,
to_file='model.png',
show_shapes=True,
show_layer_names=True,
rankdir='TB',
expand_nested=False,
dpi=96,
)
# + id="Pbr48Q-jUrv9" colab_type="code" colab={}
# initiat Adam optimizer with 1e-3 learning rate
opt = tf.keras.optimizers.Adam(lr=1e-3)
# + id="bZ99TXn6qUmx" colab_type="code" colab={}
model.compile(optimizer=opt, loss='binary_crossentropy', metrics=['accuracy'])
# + id="jysebYOpTpI1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="610cebd1-a904-43a5-ebcc-6aee69ef04c6"
history = model.fit(
train_generator,
steps_per_epoch=np.ceil(1486 /32),
epochs=5,
verbose=1,
validation_data = val_generator,
validation_steps=np.ceil(382 /16))
# + id="WTVxIwjeV3hB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="5a47091a-0934-4766-b391-3cc19ca89011"
# load image and test the model
img = load_img('/content/data/test/test/electric car/electric__image-104.jpeg', color_mode='rgb', target_size=(128,128))
img_array = img_to_array(img)/255.0
x = np.expand_dims(img_array, 0)
out = model.predict(x)
print(out)
if out <0.5:
print('electric bus')
else:
print('electric car')
plt.imshow(img)
# + id="aDjUqMoxV0j3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 563} outputId="0fc2fbbf-bcfd-42bb-9c19-5c90794cfbc2"
#-----------------------------------------------------------
# Retrieve a list of list results on training and validation data
# sets for each training epoch
#-----------------------------------------------------------
acc = history.history[ 'accuracy' ]
val_acc = history.history[ 'val_accuracy' ]
loss = history.history[ 'loss' ]
val_loss = history.history['val_loss' ]
epochs = range(len(acc)) # Get number of epochs
#------------------------------------------------
# Plot training and validation accuracy per epoch
#------------------------------------------------
plt.plot ( epochs, acc )
plt.plot ( epochs, val_acc )
plt.title ('Training and validation accuracy')
plt.figure()
#------------------------------------------------
# Plot training and validation loss per epoch
#------------------------------------------------
plt.plot ( epochs, loss )
plt.plot ( epochs, val_loss )
plt.title ('Training and validation loss' )
# + id="U5UbxT7-YMRl" colab_type="code" colab={}
| first_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/techonair/Machine-Learing-A-Z/blob/main/Regression/Regression%20Model%20Evaluation%20%26%20Selection/polynomial_regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vN99YjPTDena"
# # Polynomial Regression
# + [markdown] id="ZIx_naXnDyHd"
# ## Importing the libraries
# + id="FjnmdyPLD2tS"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] id="fCHhNz2UupjL"
# ## Uploading Dataset
# + id="PGdYYPyTupzG"
from google.colab import files
files.upload()
# + [markdown] id="6c8YExmOD5x5"
# ## Importing the dataset
# + id="nQOdXhjXD_AE"
dataset = pd.read_csv('Data.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
# + [markdown] id="Ud_1XTb28iXH"
# ## Splitting the dataset into the Training set and Test set
# + id="bUib_pbl8ipB"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] id="Rb5nWuSHEfBV"
# ## Training the Polynomial Regression model on the Training set
# + id="HYplp4pTEm0O" colab={"base_uri": "https://localhost:8080/"} outputId="982e1fbb-dcaa-4996-c93f-a0485af6ca47"
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
poly_reg = PolynomialFeatures(degree = 4)
X_poly = poly_reg.fit_transform(X_train)
regressor = LinearRegression()
regressor.fit(X_poly, y_train)
# + [markdown] id="pzF3BRps9nlk"
# ## Predicting the Test set results
# + id="36aFLFBK9pMk" colab={"base_uri": "https://localhost:8080/"} outputId="c7d5703c-2a73-4699-beed-131ccc745a6f"
y_pred = regressor.predict(poly_reg.transform(X_test))
np.set_printoptions(precision=2)
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# + [markdown] id="Fz1uTlWV919-"
# ## Evaluating the Model Performance
# + id="xvZQ_4W893-e" colab={"base_uri": "https://localhost:8080/"} outputId="17bd4ad2-73f6-4af2-af21-7e6abb437823"
from sklearn.metrics import r2_score
r2_score(y_test, y_pred)
| Regression/Regression Model Evaluation & Selection/polynomial_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
For more information about the Elasticsearch Query and Aggregation API, check out [the Elasticsearch API documentation](https://www.elastic.co/guide/en/elasticsearch/reference/current/search.html)
For more information about the `elasticsearch-dsl-py` library, check out the [elasticsearch-dsl-py documentation on ReadTheDocs](http://elasticsearch-dsl.readthedocs.org/en/latest/).
[sharepa can be found on pypi](https://pypi.python.org/pypi/sharepa/). It is essentially a special case of `elasticsearch-dsl-py`'s Search object, so the documentation of the library should also apply to sharepa.
# -
# %matplotlib inline
from IPython.display import display
from sharepa import ShareSearch
from sharepa.analysis import bucket_to_dataframe
def tags(term=None, agg='significant_terms'):
# Create a search object
search = ShareSearch()
# If there is a term provided, only search for documents that match that term
if term:
search = search.query('match', _all=term)
# Set up the aggregation to aggregate based on the tags field
search.aggs.bucket('top tags', agg, field='tags')
print(search.to_dict())
# This pulls the results from osf.io/api/v1/share/search/
results = search.execute()
# Convert the results into a pandas dataframe
df = bucket_to_dataframe(
'top tags',
results.aggregations['top tags']['buckets']
).sort('top tags', ascending=False)
display(df)
if agg == 'significant_terms':
y = ['bg_count', 'top tags']
else:
y = 'top tags'
df.plot(x='key', y=y, kind='bar')
tags(agg='terms')
tags()
tags('cancer')
tags('flu')
tags('influenza')
tags('vaccine')
| SHARE API Presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solutions
# ### One-hot encoding the rank
# +
## One solution
# Make dummy variables for rank
one_hot_data = pd.concat([data, pd.get_dummies(data['rank'], prefix='rank')], axis=1)
# Drop the previous rank column
one_hot_data = one_hot_data.drop('rank', axis=1)
# Print the first 10 rows of our data
one_hot_data[:10]
# +
## Alternative solution ##
# if you're using an up-to-date version of pandas,
# you can also use selection by columns
# an equally valid solution
one_hot_data = pd.get_dummies(data, columns=['rank'])
# -
# ### Scaling the data
# +
# Copying our data
processed_data = one_hot_data[:]
# Scaling the columns
processed_data['gre'] = processed_data['gre']/800
processed_data['gpa'] = processed_data['gpa']/4.0
processed_data[:10]
# -
# ### Backpropagating the data
def error_term_formula(x, y, output):
return (y - output)*sigmoid_prime(x)
# +
## Alternative solution ##
# you could also *only* use y and the output
# and calculate sigmoid_prime directly from the activated output!
# below is an equally valid solution (it doesn't utilize x)
def error_term_formula(x, y, output):
return (y-output) * output * (1 - output)
| intro-neural-networks/student-admissions/StudentAdmissionsSolutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/tofighi/Linear-Algebra/blob/main/Solving_The_System_of_Linear_Equations_Using_SymPy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="RnAtCEgqLY7v"
# # What is Symbolic Computation?
# Symbolic computation deals with the computation of mathematical objects symbolically. This means that the mathematical objects are represented exactly, not approximately, and mathematical expressions with unevaluated variables are left in symbolic form.
#
# # SymPy Solver Demo
# let's create a symbolic variable $z$, and define an arbitrary equation of $55.9728340445699z^2+32z−28.1086638217203=0$, and solve it symbolically.
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="_ozBY17gLY71" outputId="e7597d06-eca3-42e5-f210-b4ef4a5be4a3"
# Written by <NAME> and <NAME>
# SymPy is the python library for symbolic mathematics.
# https://docs.sympy.org/latest/index.html
import sympy as sp
import math
sp.init_printing(use_unicode=True)
# Define z as a symbolic variable.
z = sp.symbols('z')
# Get the k-value, substitute it into the function.
# Note that cos() takes angles in radians, so convert from degrees.
k = math.cos(math.radians(33))
k
# + colab={"base_uri": "https://localhost:8080/", "height": 38} id="IlCWyO3hLY78" outputId="a3ad4f7d-1a2a-4331-c96a-b369b8f7dff6"
# Define our function.
f = (81*k**2 - 1)*z**2 + 32*z + (324*k**2 - 256)
f
# + colab={"base_uri": "https://localhost:8080/", "height": 38} id="zdQL0CKlLY79" outputId="ab50a8eb-353e-400f-c632-31e39649f805"
# Define the related equation.
eqn = sp.Eq(f,0)
eqn
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="bqJO4r9jLY7-" outputId="7eb14e01-046e-4f69-f06c-cd8b5ec29832"
# Solve the equation
sp.solveset(eqn, z)
# + [markdown] id="QQqQdONilPc4"
# # How calculate RREF and solve system of linear equations in SymPy?
# Assume the following system of linear equations. SymPy can reduce the following matrices to RREF:
#
# $\left(\begin{array}{ccc|c}1 & 2 & -4 & 12 \\ 2 & 6 & 8 & 14 \\ 0 & 1 & 8 & -5\end{array}\right)$
#
# \begin{alignedat}{3}
# % R & L & R & L & R & L
# x & +{} & 2y & +{} & -4z & = 12 \\
# 2x & +{} & 6y & +{} & 8z & = 14 \\
# & {} & y & +{} & 8z & = -5 \\
# \end{alignedat}
#
# To calculate RREF we can use augmented matrix as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 78} id="8_Ni7zTtmza2" outputId="0e578fbf-3535-4dae-aef3-da884cbe8be9"
#Defining symbolic augmented matrix
A = sp.Matrix([[1,2,-4,12],[2,6,8,14],[0,1,8,-5]])
A
# + colab={"base_uri": "https://localhost:8080/", "height": 78} id="0aQsfntinCNQ" outputId="c29be4c2-1714-4c1c-8b55-21122ff6901b"
# print RREF
A.rref()
# + [markdown] id="W7jWGcslrtTE"
# # Solving system of linear equations in SymPy
#
# RREF method does not provide final solutions for the following system of linear equations:
#
# \begin{alignedat}{3}
# % R & L & R & L & R & L
# x & +{} & 2y & +{} & -4z & = 12 \\
# 2x & +{} & 6y & +{} & 8z & = 14 \\
# & {} & y & +{} & 8z & = -5 \\
# \end{alignedat}
#
# To find solutions in Sympy you should use the *Sandard Matrix Representation*, $A\mathbf{x}=\mathbf{b}$:
#
# \begin{alignedat}{3}
# \left[\begin{array}{ccc}1 & 2 & -4 \\ 2 & 6 & 8 \\ 0 & 1 & 8\end{array}\right]\left[\begin{array}{l}x \\ y \\ z\end{array}\right]=\left[\begin{array}{c}12 \\ 14 \\ -5\end{array}\right]
# \end{alignedat}
#
# - $A$ is called the coefficient matrix
# - $\mathbf{x}$ is the unknowns or solution vector
# - $\mathbf{b}$ is the constant vector
#
# Where
#
# - $A=\left[\begin{array}{ccc}1 & 2 & -4 \\ 2 & 6 & 8 \\ 0 & 1 & 8\end{array}\right]$
#
# - $\mathbf{x}=\left[\begin{array}{c}x \\ y \\ z\end{array}\right]$
#
# - $\mathbf{b}=\left[\begin{array}{c}12 \\ 14 \\ -5\end{array}\right]$
#
# To solve the system you can use `sp.linsolve` as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 38} id="jKBogiIynQ1B" outputId="57396551-2a34-4c10-dc31-b79e29a39973"
x, y, z = sp.symbols('x, y, z')
A = sp.Matrix([[1, 2, -4], [2, 6, 8], [0, 1, 8]])
b = sp.Matrix([12, 14, -5])
sp.linsolve((A, b), x, y, z)
# + [markdown] id="BUer6MUU0r_Q"
# The above answer is equivalent to what we find manually after solving using RREF method:
# $$
# \begin{aligned}
# &z=t \in \mathbb{R}\\
# &y=-5-8 t \\
# &x=22+20 t
# \end{aligned}
# $$
# + [markdown] id="EKmuK6I8KE9m"
# # Calculating the Inverse Matrix
# You can calculate the inverse of matrix using `inv()` method as follows:
# ## Note:
# A square matrix is invertible if and only if the determinant is not equal to zero.
# + id="gyG-VzCDKL_w" outputId="45894747-ee0f-4a93-c604-2082591646e9" colab={"base_uri": "https://localhost:8080/", "height": 56}
A = sp.Matrix([[1, 2, -4], [2, 6, 8], [0, 1, 8]])
if(A.det()!=0):
A_inv=A.inv()
else:
A_inv=[]
print ("Non Invertible Matrix: Matrix det == 0; not invertible.")
A_inv
# + id="5BpE8u5CKNaY" outputId="e2a00d71-d0d2-4992-a473-b36f0a65a172" colab={"base_uri": "https://localhost:8080/", "height": 78}
A = sp.Matrix([[2, 1, 0], [0, 4, -3], [1, -3, 2]])
if(A.det()!=0):
A_inv=A.inv()
else:
A_inv=[]
print ("Non Invertible Matrix: Matrix det == 0; not invertible.")
A_inv
| Solving_The_System_of_Linear_Equations_Using_SymPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: playa_venv
# language: python
# name: playa_venv
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import datetime as dt
import os
from sklearn.metrics import mean_squared_error
np.random.seed(5)
n_playa = 71848 # 71848 is all of them
n_iter = 1000
reload_run = True
# # Load data
pred_df = pd.read_csv('../data/all_preds_best_mean_inun_v2_calibrated.csv')
target_col = 'pred_cal' # pred or pred_cal
# Add months and years back in
pred_df['date'] = np.tile(pd.date_range('1984-03', periods=418, freq='M'), int(pred_df.shape[0]/418))
pred_df = pred_df.set_index(['id','date'])
# # Make plots
def simulate_inundation(pred_df, n_playa, n_iter, n_time=418, target_col='pred'):
frac_inundated = np.zeros(shape = (n_time, n_iter), dtype=np.float) - 1
for i in range(n_iter):
binary_run = np.random.binomial(
n=1,
p=pred_df[target_col].values.reshape([n_playa, n_time])
)
assert binary_run.min() == 0.
assert binary_run.max() == 1.
assert binary_run.shape == (n_playa, n_time)
frac_inundated[:,i] = np.mean(binary_run, axis=0)
if i % 100 == 0:
print(i, 'done')
# Checks, commenting out because too much memory usage
# assert frac_inundated.shape == (n_time, n_iter)
# assert frac_inundated.min() > 0.
# assert frac_inundated.max() < 1.
return frac_inundated
if reload_run and os.path.exists('./frac_inundated_meaninun_v2.npy'):
frac_inundated = np.load('./frac_inundated_meaninun_v2.npy')
else:
if n_playa < 71848:
frac_inundated = simulate_inundation(pred_df.iloc[0:(418*n_playa)], n_playa, n_iter, target_col=target_col)
else:
frac_inundated = simulate_inundation(pred_df, n_playa, n_iter, target_col=target_col)
inundation_sd = np.std(frac_inundated, axis=1)
inundation_mean_of_means = np.mean(frac_inundated, axis=1)
# inundation_975 = np.percentile(frac_inundated, 0.975, axis=1)
# inundation_025 = np.percentile(frac_inundated, 0.975, axis=1)
# inundation_min = np.min(frac_inundated, axis=1)
# inundation_max = np.max(frac_inundated, axis=1)
np.mean(frac_inundated[5])
np.max(frac_inundated[5])
plt.hist(frac_inundated[5])
# +
# plot all of the draws for the fraction inundated
dates = pred_df.index.get_level_values(1)[:418]
fig, ax = plt.subplots(figsize=[15,3.5])
plt.plot(dates, pred_df['true'].groupby('date').mean().values, label='True',linewidth=1.25)
# plt.plot(dates, inundation_min, label='Predicted Range',
# linestyle='--', color='darkorange', linewidth=1)
# plt.plot(dates, inundation_max,
# linestyle='--', color='darkorange', linewidth=1)
plt.plot(dates, inundation_mean_of_means, label='Predicted', color='red', linewidth=1.25)
# Commenting out spreads, not very interesting
# plt.plot(dates, inundation_975, label='97.5%')
# plt.plot(dates, inundation_025, label='2.5%')
plt.legend(loc=(0.47, 0.74), prop={'size':14})
plt.xlabel("Date", size=15)
plt.ylabel("Fraction inundated",size=15)
plt.xlim(dt.datetime(1984,1,1), dt.datetime(2019,1,1))
plt.axvline(dt.datetime(2015,1,1), color='black')
plt.axvline(dt.datetime(2011,1,1), color='black')
plt.text(dt.datetime(2011,3,1), 0.135, 'Validation', size=15)
plt.text(dt.datetime(2015,3,1), 0.135, 'Test', size=15)
plt.text(dt.datetime(1984,3,1), 0.135, 'Train', size=15)
plt.savefig('/home/ksolvik/research/misc_projects/playa/deliverables/figures/inun_sim.png', dpi=300, bbox_inches='tight')
plt.show()
# -
# Mean squared error
print('train:',mean_squared_error(pred_df['true'].groupby('date').mean().values[:-96], inundation_mean_of_means[:-96], squared=False))
print('val:',mean_squared_error(pred_df['true'].groupby('date').mean().values[-96:-48], inundation_mean_of_means[-96:-48], squared=False))
print('test:',mean_squared_error(pred_df['true'].groupby('date').mean().values[-48:], inundation_mean_of_means[-48:], squared=False))
# Write frac_inundated to file to save time
np.save('./frac_inundated_meaninun_v2.npy', frac_inundated)
| playa-inundation/analysis/simulate_inundation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Aufgabenblatt zur Logistischen Regression (Beispiel *Fashion MNIST*)
# -
# Der MNIST (Modified National Institute of Standards and Technology) Datensatz von handschriftlichen Ziffern wird sehr verbreitet eingesetzt, um Machine Learning Algorithmen zu demonstrieren und zu bewerten.
# Er lässt sich mit Scikit-Learn komfortabel über die Funktion `fetch_mldata('MNIST original')` aus dem Repository http://mldata.org laden.
# Von dort wurde der Datensatz bisher über 370.000-mal heruntergeladen.
#
# Allerdings sind die MNIST Daten nicht mehr unbedingt zeitgemäß.
# Mit modernen Verfahren läßt sich eine Präzision mit deutlich über 99% erreichen, weswegen der Datensatz heutzutage allgemein als "zu leicht" eingeschätzt wird.
#
# In diesem Aufgabenblatt verwenden wir daher einen Datensatz, der sehr ähnlich wie MNIST aufgebaut ist, für den eine Klassifikation aber etwas schwieriger ist.
# Es handelt sich um den sogenannten [*Fashion MNIST*](https://github.com/zalandoresearch/fashion-mnist) Datensatz, den der Online-Versandhändler Zalando entwickelt und zur freien Verfügung gestellt hat.
#
# Der Datensatz besteht aus 70.000 Bildern von Kleidungsstücken, Schuhen und Taschen und Kleidern.
# Jedes Bild besteht aus 784 (28×28) Pixeln, die einzelnen Pixel werden als Grauwert im Bereich 0 bis 255 gespeichert.
# Bei einem Byte pro Pixel und 70.000 Bildern à 784 Pixeln, benötigt der Datensatz ca. 50MB am Speicherplatz.
#
# Die Daten sind auf 4 Dateien aufgeteilt:
# - `train-images-idx3-ubyte.gz`: 60.000 Bilder als Traingnsdatensatz
# - `train-labels-idx1-ubyte.gz`: Die zum Traingnsdatensatz zugehörigen Label
# - `test-images-idx3-ubyte.gz`: 10.000 Bilder als Testdatensatz
# - `test-labels-idx1-ubyte.gz`: Die zum Testdatensatz zugehörigen Label
# +
import os
import requests
files = ['train-images-idx3-ubyte.gz','train-labels-idx1-ubyte.gz',
't10k-images-idx3-ubyte.gz','t10k-labels-idx1-ubyte.gz']
for f in files:
if not os.path.isfile(f):
url = 'http://fashion-mnist.s3-website.eu-central-1.amazonaws.com/' + f
r = requests.get(url)
with open(f, 'wb') as f:
f.write(r.content)
# -
# Mit der Funktion `load_mnist` können die Rohdaten, die im gzip Format vorliegen, in NumPy-Arrays geladen.
# Die Funktion liefert ein Tupel aus den Bilddaten sowie den Labels zurück.
def load_mnist(path, kind='train'):
'''
Quelle: https://github.com/zalandoresearch/fashion-mnist/blob/master/utils/mnist_reader.py
"Fashion-MNIST: a Novel Image Dataset for Benchmarking Machine Learning Algorithms". <NAME>, <NAME>, <NAME>. arXiv:1708.07747
'''
import os
import gzip
import numpy as np
"""Load MNIST data from `path`"""
labels_path = os.path.join(path,
'%s-labels-idx1-ubyte.gz'
% kind)
images_path = os.path.join(path,
'%s-images-idx3-ubyte.gz'
% kind)
with gzip.open(labels_path, 'rb') as lbpath:
labels = np.frombuffer(lbpath.read(), dtype=np.uint8,
offset=8)
with gzip.open(images_path, 'rb') as imgpath:
images = np.frombuffer(imgpath.read(), dtype=np.uint8,
offset=16).reshape(len(labels), 784)
return images, labels
X_train, y_train_10 = load_mnist('.', kind='train')
X_test, y_test_10 = load_mnist('.', kind='t10k')
# Zum Anzeigen eines Bildes, können wir die Funktion `imshow` aus dem Modul `matplotlib.pyplot` verwenden.
# Da die Pixel eines Bildes als fortlaufendes Array abgelegt sind, müssen wir das Bild zuvor noch mit der Funktion `reshape` in das richtige Format bringen.
# Um ein zufällig ausgewähltes Bild anzuzeigen, verwenden wir die Funktion `random.randint(x,y)`, die eine Zufallszahl im Bereich `x` bis `y-1` liefert.
# +
import numpy as np
import random
import matplotlib.pyplot as plt
import matplotlib.cm
sample = random.randint(0,len(y_train_10))
img = X_train[sample].reshape(28,28)
plt.imshow(img, cmap = matplotlib.cm.binary, interpolation="nearest")
print("Klasse ", y_train_10[sample])
# -
# **Aufgabe:** Trainieren Sie ein logistisches Regressionsmodell zur Erkennung von Pullovern
#
# Verwenden Sie die [`LogisticRegression`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) Klasse aus Scikit-learn.
# Als *Solver* soll das Modell `liblinear` benutzen und beim Trainieren maximal 10 Iterationen verwenden.
#
# Berechnen Sie die *Classification Accuracy* über die Scikit-learn Funktion `score`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2027fdc0ec40e84f1016c9ae06b4f7f8", "grade": false, "grade_id": "cell-a91b28611a03280a", "locked": false, "schema_version": 3, "solution": true, "task": false}
from sklearn.linear_model import LogisticRegression
accuracy_train = None
accuracy_test = None
# Setze nur eine Klasse auf 1, alle anderen auf 0
klasse = 0
y_train = (y_train_10 == klasse) * 1
y_test = (y_test_10 == klasse) * 1
# YOUR CODE HERE
raise NotImplementedError()
print("Treffequote Trainingsdaten: %.2f%%" % (100*accuracy_train))
print("Treffequote Testdaten: %.2f%%" % (100*accuracy_test))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "a6eac11f01c63738dfd439a79c3eaeb8", "grade": true, "grade_id": "cell-941984e0760af4d3", "locked": true, "points": 0, "schema_version": 3, "solution": false, "task": false}
assert "max_iter" in _i
assert "liblinear" in _i
assert "LogisticRegression" in _i
# -
# Nun können wir uns dem eigentlichen Klassifikationsproblem mit mehreren Klassen widmen.
# Wir verwenden an dieser Stelle ein logistisches Regressionsmodell mit einer one-versus-all Strategie, um die Klassen für die einzelnen Kleidungsstücke und Accessoires vorauszusagen.
#
# Im Vergleich zu den bisher betrachteten Datensätzen, sind die Fashion MNIST Daten sehr umfangreich.
# Daher ist auch die Berechnung der Modellparameter wesentlich komplexer und zeitaufwendiger als bei den Beispielen zuvor.
# Wir sollten daher darauf achten, dass einmal berechnete Parameter nicht verlorengehen, etwa, weil das Programm, bzw. das Python-Notebook geschlossen wird.
#
# Um die berechneten Modellparameter in eine Datei zu speichern, verwenden wir die Bibliothek Pickle (zu deutsch *einmachen*, *einlegen* oder auch *Essiggurke*).
# Mit Pickle kann man Python Objekte serialisiert in eine Datei schreiben.
# Dabei bleibt die komplette Struktur des Objekts intakt, sodass das Objekt aus der Datei wieder vollständig hergestellt werden kann.
#
# Sobald ein Modell berechnet wurde, speichern wir es in eine Datei.
# Wird die Code-Zelle später erneut ausgeführt, laden wir die Daten aus der Datei ein, anstatt das Modell erneut zu trainieren.
# +
import os
import pickle
from sklearn.linear_model import LogisticRegression
X_train, y_train = load_mnist('.', kind='train')
X_test, y_test = load_mnist('.', kind='t10k')
filename = 'logreg_fashion_mnist.mod'
if os.path.isfile(filename):
logreg = pickle.load(open(filename, 'rb'))
else:
#logreg = LogisticRegression(multi_class="ovr")
logreg = LogisticRegression(max_iter=10, solver='lbfgs',multi_class='ovr', n_jobs=-1)
logreg.fit(X_train, y_train)
pickle.dump(logreg, open(filename, 'wb'))
# -
# Zur Überprüfung der Vorhersagegenauigkeit können wir die sklearn-Funktion `score` verwenden.
# Sie berechnet die den Anteil der korrekten Vorhersagen über alle Klassen.
# Eine Vorhersage ist korrekt, wenn die Vorhergesagte Klasse mit der tatsächlichen Klasse übereinstimmt.
# load the model from disk
saved_model = pickle.load(open(filename, 'rb'))
result = saved_model.score(X_test, y_test)
print("Vorhersagegenauigkeit (Testdaten): %.2f%%" % (result*100))
result = saved_model.score(X_train, y_train)
print("Vorhersagegenauigkeit (Trainingsdaten): %.2f%%" % (result*100))
# **Aufgabe:** Berechnen Sie die Vorhersagegenauigkeit ohne die `score`-Funktion zu verwenden.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "e15a614c8af8fbcaebcbb85cec5486dd", "grade": false, "grade_id": "cell-29262aebbede1677", "locked": false, "schema_version": 3, "solution": true, "task": false}
# acc_test: the ratio of correctly predected labels to the size of the test set (between 0 and 1)
acc_test = None
# YOUR CODE HERE
raise NotImplementedError()
print("Vorhersagegenauigkeit (Testdaten): %.2f%%" % (acc_test*100))
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "cc52b95fafce03cde43cbdd853c49ee3", "grade": true, "grade_id": "cell-c7434893394691c6", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Test Cell
#----------
assert 0 <= acc_test <= 1
assert acc_test == saved_model.score(X_test, y_test)
# -
# Die Konfusionsmatrix ist eine Tabelle, die für jede Klasse die Anzahl der übereinstimmenden Vorhersagen darstellt.
# In den Zeilen der Matrix sind die tatsächlichen Klassen aufgetragen, in den Spalten die vorhergesagten Klassen.
# Die Werte auf der Diagonalen der Matrix sind demnach korrekte Vorhersagen, die Werte ausserhalb der Diagonalen sind fehlerhafte Vorhersagen.
#
# Die Konfusionsmatrix kann mit der Funktion `sklearn.metrics.confusion_matrix` berechnet werden.
from sklearn.metrics import confusion_matrix
y_pred = logreg.predict(X_test)
confusion_matrix(y_test, y_pred)
# **Aufgabe:** Berechnen Sie die Konfusionsmatrix ohne die Funktion `confusion_matrix` zu verwenden.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "1c50a3994e3fcd14219bb97bd7b684f7", "grade": false, "grade_id": "cell-72cb7cc6e418d4d0", "locked": false, "schema_version": 3, "solution": true, "task": false}
# classes: Anzahl der Klassen
# cm: Konfusionsmatrix (Tatsächliche Klassen in Zeilen, vorhergesagte Klassen in Spalten)
classes = sorted(list(set(y_test)))
cm = []
for i in classes:
pass
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "f1465491abe5dfac8225c88f8c4a3c1e", "grade": true, "grade_id": "cell-36e912aaa83d63ef", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Test Cell
#----------
cm = np.array(cm)
n = len(set(y_test))
assert cm.shape == (n, n)
assert np.array_equal(cm, confusion_matrix(y_test, y_pred))
cm
# -
# Weitere Metriken zur Bewertung des Modells sind die Relevanz (auch Genauigkeit oder positiver Vorhersagewert; engl. precision oder positive predictive value) und die Sensitivität (auch Richtig-positiv-Rate oder Trefferquote; engl. sensitivity oder recall).
# +
from sklearn.metrics import precision_score, recall_score
y_test_c0 = [y_test==0][0]*1
y_pred_c0 = [y_pred==0][0]*1
precision_score(y_test_c0, y_pred_c0), recall_score(y_test_c0, y_pred_c0)
# -
# 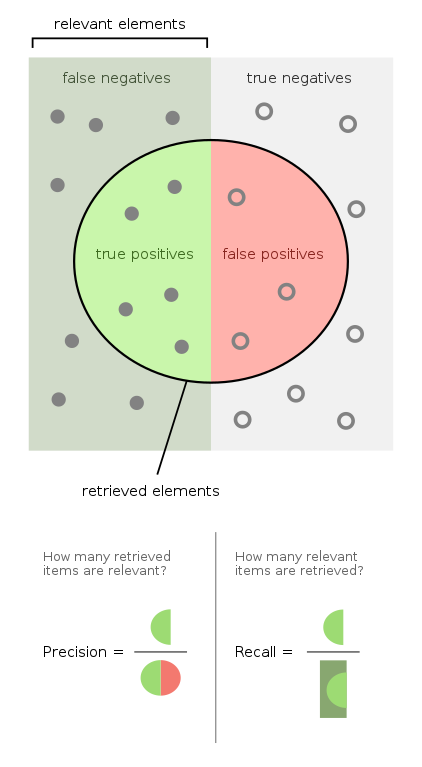
# **Aufgabe:** Berechnen Sie die Relevanz und die Sensitivität ohne die Funktionen `precision_score` und `recall_score` zu verwenden.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f8f5d63837105a7c3db3e1a29972b62c", "grade": false, "grade_id": "cell-40cb2290216aec8a", "locked": false, "schema_version": 3, "solution": true, "task": false}
def precision_recall(y_true_c0, y_pred_c0):
'''
Berechne zuerst die Anzahl der
- true positives (TP)
- false positives (FP)
- false negatives (FN)
Mit diesen Werten kann dann die Relevanz und die Sensitivitaet
berechent werden.
'''
# YOUR CODE HERE
raise NotImplementedError()
return precision, recall
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "9aba71334674e1537c3743e614fa0f26", "grade": true, "grade_id": "cell-993b6d2420050694", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Test Cell
#----------
assert precision_recall(y_test_c0, y_pred_c0) == (precision_score(y_test_c0, y_pred_c0), recall_score(y_test_c0, y_pred_c0))
| Praktika/A07/LogistischeRegression_FashionMNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Erstellung eines Tachenrechners in Python
# Import necessary modules to compute the stuff
import math
# Create functions to use them later easy
def add(x, y):
return x + y
def subtract(x, y):
return x - y
def multiply(x, y):
return x * y
def divide(x, y):
return x / y
def pow(x, y):
return math.pow(x,y)
def sqrt(x):
return math.sqrt(x)
# Show the options you can select with different outputs
print("Select your way you want to do math.")
print("1. Add")
print("2. Subtract")
print("3. Multiply")
print("4. Divide")
print("5. Pow")
print("6 Square root")
# Create the while-true statement and use of the functions
while True:
choice = input("Enter choice(1/2/3/4/5/6): ")
if choice in ('1', '2', '3', '4', '5', '6'):
if choice == '1':
x = float(input("Give me your first number: "))
y = float(input("Give me your second number: "))
print(x, "+", y, "=", add(x, y))
elif choice == '2':
x = float(input("Give me your first number: "))
y = float(input("Give me your second number: "))
print(x, "-", y, "=", subtract(x, y))
elif choice == '3':
x = float(input("Give me your first number: "))
y = float(input("Give me your second number: "))
print(x, "*", y, "=", multiply(x, y))
elif choice == '4':
x = float(input("Give me your first number: "))
y = float(input("Give me your second number: "))
print(x, "/", y, "=", divide(x, y))
elif choice == '5':
x = float(input("This time there is only one number needed: "))
print("Square root", x, "=", math.sqrt(x))
elif choice == '6':
x = float(input("Give me your first number: "))
y = float(input("Give me your second number: "))
print("The value of", x, "to the power of", num2, "=", math.pow(x, y))
break
else:
print("Invalid Input")
| Calculator_Jakob_Thielmann_with_markdowns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData]
# language: python
# name: conda-env-PythonData-py
# ---
# +
# Import SQLAlchemy and other dependencies
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy import create_engine, inspect, func
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import Column, Integer, String, Float, Text, ForeignKey
from sqlalchemy import MetaData
from sqlalchemy import Table
import pandas as pd
import numpy as np
# -
# Create engine using the `sqlite`
engine = create_engine("sqlite:///belly_button_biodiversity.sqlite")
# Declare a Base using `automap_base()`
Base = automap_base()
# Use the Base class to reflect the database tables# Use t
Base.prepare(engine, reflect=True)
# Create the inspector and connect it to the engine
inspector = inspect(engine)
# Collect the names of tables within the database
inspector.get_table_names()
# get info in the table otu
inspector.get_columns("otu")
inspector = inspect(engine)
columns = inspector.get_columns("otu")
for c in columns:
print (c['name'], c['type'])
# get info in the table samples
inspector.get_columns("samples")
# get info in the table metadata
inspector.get_columns("samples_metadata")
# Use `engine.execute` to select and display the first 5 rows from the table otu
engine.execute('SELECT * FROM otu LIMIT 5').fetchall()
# Use `engine.execute` to select and display the first 5 rows from the table samples
engine.execute('SELECT * FROM samples LIMIT 5').fetchall()
# Use `engine.execute` to select and display the first 5 rows from the table samples_metadata
engine.execute('SELECT * FROM samples_metadata LIMIT 5').fetchall()
# Reflect Databases into ORM class
Samples = Base.classes.samples
Metadata = Base.classes.samples_metadata
Otu = Base.classes.otu
# Start a session to query the database
session = Session(engine)
# display the 5 first lines of the table
engine.execute("SELECT ETHNICITY FROM samples_metadata LIMIT 5").fetchall()
run_database = engine.execute('SELECT ETHNICITY FROM samples_metadata').fetchall()
run_database
| data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: protein_binding
# language: python
# name: protein_binding
# ---
# # STEP 1 in the Feature Selection Pipeline: Train Random Forest to Identify the informative Features
# +
import time
import glob
import h5py
import multiprocessing
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use("seaborn-muted")
from utils.input_pipeline import load_data, load_protein
from scipy.stats import randint as sp_randint
from sklearn.model_selection import cross_val_score, RandomizedSearchCV
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.preprocessing import Imputer, Normalizer
from sklearn.feature_selection import VarianceThreshold
from sklearn.linear_model import RandomizedLogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, f1_score, make_scorer
random_state=np.random.RandomState(0)
imputer = Imputer()
normalizer = Normalizer()
forest_classifier = RandomForestClassifier(n_jobs=10)
data_path = "data/full_26_kinase_data.h5"
# -
# ## Load the Data
data_fo = h5py.File(data_path,'r')
protein_list = list(data_fo.keys())
input_size = 0
for protein in protein_list:
input_size += data_fo[protein]['label'].shape[0]
print(input_size)
X = np.ndarray([0,5432])
y = np.ndarray([0,1])
for protein in protein_list:
#create a balanced set for each of the proteins by randomly sampling from each of the negative classes
X_p,y_p = load_data(data_path,protein_name_list=[protein], mode=1)
X_n, y_n = load_data(data_path,protein_name_list=[protein],sample_size = X_p.shape[0], mode=0)
X_ = np.vstack((X_p,X_n))
y_ = np.vstack((y_p,y_n))
X = np.vstack((X_,X))
y = np.vstack((y_,y))
# ## Random Forest
# The algorithm which constructs a random forest natively performs feature selection by finding the best splits for a particular feature to minimize some measure of label impurity. This can be leveraged as a feature selection method to train other classifiers (in addition to other random forests).
# +
# once new data is ready, remove the imputer, keep normalizing
forest_pipe = Pipeline(steps=[('imputer', imputer), ('normalizer', normalizer),
('selection_forest',RandomForestClassifier(n_jobs=16, oob_score=True,
class_weight="balanced",random_state=random_state))])
forest_params = {"selection_forest__n_estimators": sp_randint(15,30),
"selection_forest__criterion": ["gini","entropy"]
}
estimator_search = RandomizedSearchCV(estimator=forest_pipe,param_distributions=forest_params, scoring='f1',cv=5, random_state=random_state)
# -
estimator_search.fit(X,y.flatten())
forest_model = estimator_search.best_estimator_
support = forest_model.named_steps['selection_forest'].feature_importances_
support = forest_model.named_steps['selection_forest'].feature_importances_
# # and collect the features
# so that they can be used in later experiments
plt.clf()
plt.figure(figsize=[12,8])
plt.plot(np.sort(support)[::-1])
plt.title("Random Forest Feature Support (sorted)")
plt.ylabel("feature importance")
plt.savefig("poster_results/feature_importance_curve_full_set.png")
plt.show()
# +
full_features = list(h5py.File("data/full_26_kinase_data.h5","r")["lck"].keys())
# use a list comprehension instead
full_features.remove("label")
full_features.remove("receptor")
full_features.remove("drugID")
keep_idxs = support > np.mean(support,axis=0)
features_to_keep = np.asarray(full_features)[keep_idxs]
features_to_keep = pd.DataFrame(features_to_keep)
features_to_keep.to_csv("data/step1_features.csv",index=False,header=False)
print(len(full_features),features_to_keep.shape)
| notebooks/Step 1 Random Forest Feature Selection (In Progress).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 1
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optmi
import torchvision
from torchvision import datasets, transforms
import matplotlib.pyplot as plt
import numpy as np
plt.ion() # interactive mode
# -
# %aimport net_modules
ResBlock = net_modules.ResBlock
Hourglass = net_modules.Hourglass
sample = torch.zeros((1,3,100,100))
a = Hourglass(3, 16, 2)
a(sample).shape
class JointDetectionNet(nn.Module):
def __init__(self, channel_in):
super(JointDetectionNet, self).__init__()
self.init = nn.Sequential(
nn.Conv2d(channel_in, 64,
kernel_size=7, stride=3,
padding=3),
nn.BatchNorm2d(64),
nn.ReLU(True),
ResBlock(64, 128),
nn.MaxPool2d(2,2),
ResBlock(128, 128),
ResBlock(128, 128),
ResBlock(128, 256)
)
self.hg1 = Hourglass(4, 256, 512)
def forward(self, x):
return self.init(x)
net = JointDetectionNet(3)
| research/hand_pose_estimation/HourclockNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## FIoT Lab02 PartD
# # Lab - Take the Python Challenge
#
#
# This exercise tests your understanding of Python basics. If you don't know how to solve them, look into the Python resources available online.
# #### Answer the questions or complete the tasks outlined below, use the specific method described if applicable.
# #### 1) What is 3 to the power of 5?
# Code cell 1
pow(3,5)
# #### 2) Create a variable, *`s`*, containing the string "This course is amazing!". Using the variable, split the string into a list.
# Code cell 2
s = "This course is amazing!"
s.split()
# #### 3) Given the variables height and mountain, use .format() to print the following string:
# <center>`The height of Mt. Everest is 8848 meters.`</center>
# Code cell 3
mountain = "Mt. Everest"
height = 8848
print("The height of {} is {} meters".format( mountain , height ))
# #### 4) Given the following nested list, use indexing to grab the word `"this"`.
# Code cell 4
lst = ['a','b',[4,10,11],['c',[1,66,['this']],2,111],'e',7]
lst[3][1][2][0]
# #### 5) Given the following nested dictionary, grab the word "that". This exercise is a little more difficult.
# Code cell 5
d = {'k1':['val1','val2','val3',{'we':['need','to','go',{'deeper':[1,2,3,'that']}]}]}
d['k1'][3]['we'][3]['deeper'][3]
# #### 6) What is the main difference between a tuple and a list?
# The difference is that a list is mutable whereas a tuple is immutable. The values in a list can be changed, in a tuple they can not.
# #### 7) Create a function, GetDomain(), that grabs the email website domain from a string in the form: `<EMAIL>`.
# So for example, passing "<EMAIL>" would return: domain.com
# +
# Code cell 6
def GetDomain(address):
return address.split('@')[-1]
GetDomain('<EMAIL>')
# -
# #### 8) Create a basic function, findInternet(), that returns True if the word 'Internet' is contained in the input string. Don't worry about edge cases like punctuation being attached to the word, but account for capitalization. (Hint: Please see https://docs.python.org/2/reference/expressions.html#in)
# Code cell 7
def findInternet(string):
return 'internet' in string.lower().split()
findInternet('The Internet Engineering Task Force was created in 1986')
# #### 9) Create a function, countIoT(), that counts the number of times the word "IoT" occurs in a string. Ignore edge cases but take into account capitalization.
# Code cell 8
def countIoT(st):
count = st.lower().count("iot")
return count
countIoT('I don\'t know how to spell IoT ! Is it IoT or iot ? What does iot mean anyway?')
# #### 10) Use lambda expressions and the filter() function to filter out words from a list that do not start with the letter 'd'. For example:
#
# seq = ['data','salt' ,'dairy','cat', 'dog']
#
# should be filtered down to:
#
# ['data', 'dairy', 'dog']
# Code cell 9
seq = ['data','salt' ,'dairy','cat', 'dog']
list(filter(lambda str : 'd' in str, seq))
# #### 11) Use lambda expressions and the map() function to convert a list of words to upper case. For example:
# seq = ['data','salt' ,'dairy','cat', 'dog']
#
# should become:
#
# ['DATA', 'SALT', 'DAIRY', 'CAT', 'DOG']
# +
seq = ['data','salt' ,'dairy','cat', 'dog']
# Code cell 10
list(map(lambda str : str.upper(), seq))
# -
# #### 12) Imagine a smart thermostat that is connected to the door, so that it can detect, in addition to the temperature, when people enter or leave the house.
# Write a function that, if the temperature is lower than 20 degrees Celsius, and there are people in the house (encoded as a boolean value to be passed as a parameter to the function), turns on the heating by returning the string "Heating on". When the temperature reaches 23 degrees or there are no people in the house, it returns the string "Heating off". When none of these conditions are met, the function returns "Do nothing".
# Code cell 11
def smart_thermostat(temp, people_in):
if temp < 20 and people_in is True:
return "Heating on"
elif temp >= 23 or people_in is False:
return "Heating off"
return "Do nothing"
# Code cell 12
# Verify smart_thermostat()
smart_thermostat(21,True)
# Code cell 13
# Verify smart_thermostat()
smart_thermostat(21,False)
# #### 13) The function zip(list1, list2) returns a list of tuples, where the i-th tuple contains the i-th element from each of the argument lists. Use the zip function to create the following list of tuples:
# `zipped = [('Parking', -1), ('Shops',0), ('Food Court',1), ('Offices',2)]`
# Code cell 14
floor_types = ['Parking', 'Shops', 'Food Court', 'Offices']
floor_numbers = range(-1,3)
zipped = list(zip(floor_types, floor_numbers))
print(zipped)
# #### 14) Use the zip function and dict() to create a dictionary, elevator_dict, where the keys are the floor types and the values are the corresponding floor number so that:
# elevator_dict[-1] = 'Parking'
# Code cell 15
floor_types = ['Parking', 'Shops', 'Food Court', 'Offices']
floors_numbers = range(-1,3)
elevator_dict = dict(zip(floor_numbers, floor_types))
print(elevator_dict)
# Code cell 16
# Verify elevator_dict[-1]
elevator_dict[-1]
# #### 15) Create an `Elevator` class. The constructor accepts the list of strings `floor_types` and the list of integers `floor_numbers`. The class implemets the methods 'ask_which_floor' and 'go_to_floor'. The output of this methods should look as follows:
# `floor_types = ['Parking', 'Shops', 'Food Court', 'Offices']
# floors_numbers = range(-1,4)
#
# el = Elevator(floor_numbers, floor_types)
#
# el.go_to_floor(1)`
#
# `Going to Food Court floor!`
#
# `el.go_to_floor(-2)`
#
# `There is floor number -2 in this building.`
#
# `el.ask_which_floor('Offices')`
#
# `The floor Offices is the number: 2`
#
# `el.ask_which_floor('Swimming Pool')`
#
# `There is no Swimming Pool floor in this building.`
# + jupyter={"outputs_hidden": true}
# Code cell 17
class Elevator:
def __init__(self, floor_numbers, floor_types):
self._floor_numbers = floor_numbers
self._floor_types = floor_types
self._number_to_type_dict = dict(zip(floor_numbers, floor_types))
self._type_to_number_dict = dict(zip(floor_types, floor_numbers))
def ask_which_floor(self, floor_type):
if floor_type in self._floor_types:
print('The {} floor is the number: {}.'.format(floor_type, self._type_to_number_dict[floor_type]))
else:
print('There is no {} floor in this building.'.format(floor_type))
def go_to_floor(self, floor_number):
if floor_number in self._floor_numbers:
print('Going to {} floor'.format(self._number_to_type_dict[floor_number]))
else:
print('There is floor number {} in this building.'.format(floor_number))
# -
# Verify code cell 18
el = Elevator(floor_numbers, floor_types)
el.go_to_floor(1)
# Verify code cell 19
el.go_to_floor(-2)
# Verify code cell 20
el.ask_which_floor('Offices')
# Verify code cell 21
el.ask_which_floor('Swimming Pool')
# # Great job!
# ###### TU856.7.8 Year 4
| lab2/Lab02 PartD - FIoT - Take the Python Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''specvqgan'': conda)'
# name: python3
# ---
# + [markdown] id="_8RjxbbWkxaO"
# # Spectrogram VQGAN as a Neural Audio Codec
# This notebook will guide you through the audio compression
# demo with Spectrogram VQGAN for arbitrary audio.
#
# Compared to the state-of-the-art,
# this approach not only achieves **>10x** reduction in bitrate
# but also supports a large variety of audio classes
# (300+ vs. 2 in the SotA).
#
# This model is a part of our approach for
# "**Taming Visually Guided Sound Generation 🖼️ 👉 🔉**"
# project and was never designed to be a neural audio
# codec but it happened to be highly effective for this task.
#
# Spectrogram VQGAN is an upgraded
# [VQVAE](https://arxiv.org/abs/1711.00937) with
# adversarial and perceptual losses, pre-trained in an
# auto-encoder fashion.
# The model was pre-trained on
# audio spectrograms from a general-purpose dataset
# ([VGGSound](https://www.robots.ox.ac.uk/~vgg/data/vggsound/)). For details feel free to check out our paper and code.
#
# [Project Page](https://v-iashin.github.io/specvqgan)
# • [Paper](https://arxiv.org/abs/2110.08791)
# • [Code](https://github.com/v-iashin/SpecVQGAN)
#
# [](https://colab.research.google.com/drive/1K_-e6CRQFLk9Uq6O46FOsAYt63TeEdXf?usp=sharing)
# + [markdown] id="e9sy2NdbrVrb"
# ## Imports and Device Selection
# + colab={"base_uri": "https://localhost:8080/"} id="hOLaNyfxN_r4" outputId="8a624274-85c7-41b1-d035-5d84e61a0167"
import os
from pathlib import Path
import soundfile
import torch
import IPython
import matplotlib.pyplot as plt
from feature_extraction.demo_utils import (calculate_codebook_bitrate,
extract_melspectrogram,
get_audio_file_bitrate,
get_duration,
load_neural_audio_codec)
from sample_visualization import tensor_to_plt
from torch.utils.data.dataloader import default_collate
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + [markdown] id="bjTG4WGZrpfS"
# ## Select a Model
# The model will be automatically downloaded given the `model_name`.
#
# + colab={"base_uri": "https://localhost:8080/"} id="Y_G-kCmgRQuM" outputId="59858308-c4e9-4f9e-ef3a-5c30e3560fec"
model_name = '2021-05-19T22-16-54_vggsound_codebook'
log_dir = './logs'
# loading the models might take a few minutes
config, model, vocoder = load_neural_audio_codec(model_name, log_dir, device)
# + [markdown] id="QJbUv-2grw_v"
# ## Select an Audio
#
# The model was pre-trained on `(mels x time) = (80 x 848)`
# spectrograms.
# Considering the convolutional nature of the
# Spectrogram VQGAN, it can also be applied to sounds of
# a different duration.
# + colab={"base_uri": "https://localhost:8080/"} id="OHkHiAPykQ6u" outputId="9ff12ed3-3996-4108-a0c2-fee8b3faf26f"
# Select an Audio
input_wav = './data/neural_audio_codec/music2/original.wav'
# Spectrogram Extraction
model_sr = config.data.params.sample_rate
duration = get_duration(input_wav)
spec = extract_melspectrogram(input_wav, sr=model_sr, duration=duration)
print(f'Audio Duration: {duration} seconds')
print('Original Spectrogram Shape:', spec.shape)
# Prepare Input
spectrogram = {'input': spec}
batch = default_collate([spectrogram])
batch['image'] = batch['input'].to(device)
x = model.get_input(batch, 'image')
# + [markdown] id="a1ipylQGU18g"
# ## Encoding and Decoding
# + id="EWKIYgxeqcTp"
with torch.no_grad():
quant_z, diff, info = model.encode(x)
xrec = model.decode(quant_z)
print('Compressed representation (it is all you need to recover the audio):')
F, T = quant_z.shape[-2:]
print(info[2].reshape(F, T))
# + [markdown] id="Xq7ylmu0qhXi"
# ## Displaying the Results
# + colab={"base_uri": "https://localhost:8080/", "height": 770} id="VqjDSRk0Pmqu" outputId="73ccab40-bba4-4fb6-b984-ed2bd11ae16d"
# Calculate Bitrate
bitrate = calculate_codebook_bitrate(duration, quant_z, model.quantize.n_e)
orig_bitrate = get_audio_file_bitrate(input_wav)
# Save and Display
x = x.squeeze(0)
xrec = xrec.squeeze(0)
# specs are in [-1, 1], making them in [0, 1]
wav_x = vocoder((x + 1) / 2).squeeze().detach().cpu().numpy()
wav_xrec = vocoder((xrec + 1) / 2).squeeze().detach().cpu().numpy()
# Creating a temp folder which will hold the results
tmp_dir = os.path.join('./tmp/neural_audio_codec', Path(input_wav).parent.stem)
os.makedirs(tmp_dir, exist_ok=True)
# Save paths
x_save_path = Path(tmp_dir) / 'vocoded_orig_spec.wav'
xrec_save_path = Path(tmp_dir) / f'specvqgan_{bitrate:.2f}kbps.wav'
# Save
soundfile.write(x_save_path, wav_x, model_sr, 'PCM_16')
soundfile.write(xrec_save_path, wav_xrec, model_sr, 'PCM_16')
# Display
print(f'Original audio ({orig_bitrate:.0f} kbps):')
IPython.display.display(IPython.display.Audio(x_save_path))
print(f'Reconstructed audio ({bitrate:.2f} kbps):')
IPython.display.display(IPython.display.Audio(xrec_save_path))
print('Original Spectrogram:')
IPython.display.display(tensor_to_plt(x, flip_dims=(2,)))
plt.close()
print('Reconstructed Spectrogram:')
IPython.display.display(tensor_to_plt(xrec, flip_dims=(2,)))
plt.close()
| neural_audio_codec_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''lewisml'': conda)'
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="noAc7PFsABY3" executionInfo={"status": "ok", "timestamp": 1629732720149, "user_tz": 360, "elapsed": 751, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="09498791-047f-4a6c-c085-534f0027697c"
# If you have installation questions, please reach out
import seaborn as sns
import pandas as pd # data storage
import catboost as cats # graident boosting
from catboost import CatBoostRegressor, Pool
import numpy as np # math and stuff
import matplotlib.pyplot as plt # plotting utility
import sklearn # ML and stats
print('catboost ver:', cats.__version__)
print('scikit ver:', sklearn.__version__)
import datetime
from sklearn.preprocessing import MinMaxScaler, RobustScaler
from sklearn.model_selection import cross_val_score, KFold, train_test_split
from sklearn.utils.class_weight import compute_sample_weight
from sklearn.metrics import median_absolute_error, max_error, mean_squared_error
# -
from catboostmodels import catboost_multi_xrf
# + [markdown] id="pTOHb1UjeRA2"
# # dataframes
# + colab={"base_uri": "https://localhost:8080/"} id="vtLJv4PVC6vj" executionInfo={"status": "ok", "timestamp": 1629732722117, "user_tz": 360, "elapsed": 1970, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="d345803e-c566-43f9-954a-e88f113716f4"
df0 = pd.read_csv('../../core_to_wl_merge/OS0_Merged_dataset_imputed_08_23_2021.csv')
df1 = pd.read_csv('../../core_to_wl_merge/OS1_Merged_dataset_imputed_08_23_2021.csv')
df2 = pd.read_csv('../../core_to_wl_merge/OS2_Merged_dataset_imputed_08_23_2021.csv')
# + id="I28VF-T-AJtU" executionInfo={"status": "ok", "timestamp": 1629732722121, "user_tz": 360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
param_dict ={
"iter" : 1000, #Maximum iterations
"GPUorCPU": "CPU", #CPU is fine for this dataset on most laptops
"inputs": ['CAL', 'GR', 'DT', 'SP', 'DENS', 'PE', 'RESD', 'PHIN', 'PHID', 'GR_smooth', 'PE_smooth'],
"target": ['Ti', 'Mg', 'Al', 'Ca', 'Si' ]}
# + [markdown] id="0PcBLInB-7Do"
# # Offset 0
# + id="60EEvEnyAN0B" executionInfo={"status": "ok", "timestamp": 1629732722122, "user_tz": 360, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
dataset = df[[
'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth',
'Ti', 'Mg', 'Al', 'Ca', 'Si' ]]
# + colab={"base_uri": "https://localhost:8080/"} id="-pNcz_yKAkwf" executionInfo={"status": "ok", "timestamp": 1629732722286, "user_tz": 360, "elapsed": 172, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="bf52ad65-a48d-469f-d2eb-f61581cb5d16"
dataset.replace('NaN',np.nan, regex=True, inplace=True)#
dataset = dataset.dropna()
# dataset.head(3)
# + id="nVrxkCAvApiJ" executionInfo={"status": "ok", "timestamp": 1629732722287, "user_tz": 360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
X = dataset[[ 'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth']]
Y2 = dataset[['Ti', 'Mg', 'Al', 'Ca', 'Si']]
# + id="nE_bH2h0AsHL" executionInfo={"status": "ok", "timestamp": 1629732722288, "user_tz": 360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
seed = 42 # random seed is only used if you want to compare exact answers with friends
test_size = 0.25 # how much data you want to withold, .15 - 0.3 is a good starting point
X_train, X_test, y_train, y_test = train_test_split(X.values, Y2.values, test_size=test_size)
# + [markdown] id="jzdBZqbQ-800"
# # iterations
# + id="LHtwzGPz_CIN" executionInfo={"status": "ok", "timestamp": 1629732722288, "user_tz": 360, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
max_iter=500
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="fFzyg5R0Bq3F" executionInfo={"status": "ok", "timestamp": 1629733301554, "user_tz": 360, "elapsed": 579273, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="467600c0-c967-4e50-a412-9c9c5627c725"
df_OS0 = catboost_multi_xrf(X_train, X_test, y_train, y_test, 'catboost_MultiXRF.csv', max_iter)
# + [markdown] id="ASstvwfu-2CS"
# # Offset 1
# + id="EHbzrOB5_Zen" executionInfo={"status": "ok", "timestamp": 1629733301556, "user_tz": 360, "elapsed": 8, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
df2 = df2.drop(['Unnamed: 0', 'Unnamed: 0.1', 'LiveTime2','ScanTime2', 'LiveTime1','ScanTime1',
'ref_num', 'API', 'well_name', 'sample_num' ], axis=1)
df2 = df2[df2.Al >= 0]
dataset2 = df2[[
'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth',
'Ti', 'Mg', 'Al', 'Ca', 'Si']]
# Features we will use for prediction
X2 = dataset2[['CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth']]
# What we are trying to predict
Y2 = dataset2[['Ti', 'Mg', 'Al', 'Ca', 'Si']]
X_train2, X_test2, y_train2, y_test2 = train_test_split(X2.values, Y2.values, test_size=test_size)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="WpavCZSCCJ9s" executionInfo={"status": "ok", "timestamp": 1629733878254, "user_tz": 360, "elapsed": 576704, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="b2868184-6172-4b2a-93c6-4d5b233cbc59"
catboost_multi_xrf(X_train2, X_test2, y_train2, y_test2, 'OS1_catboost_MultiXRF.csv', max_iter)
# + [markdown] id="ljxmNL7UCYCQ"
# # Offset 2
# + id="-QVIIDeQAiul" executionInfo={"status": "ok", "timestamp": 1629733878256, "user_tz": 360, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
df3 = df3.drop(['Unnamed: 0', 'Unnamed: 0.1', 'LiveTime2','ScanTime2', 'LiveTime1','ScanTime1',
'ref_num', 'API', 'well_name', 'sample_num' ], axis=1)
df3 = df3[df3.Al >= 0]
dataset3 = df3[[
'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth',
'Ti', 'Mg', 'Al', 'Ca', 'Si']]
# Features we will use for prediction
X3 = dataset3[['CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'GR_smooth',
'PE_smooth']]
# What we are trying to predict
Y3 = dataset2[['Ti', 'Mg', 'Al', 'Ca', 'Si']]
X_train3, X_test3, y_train3, y_test3 = train_test_split(X3.values, Y3.values, test_size=test_size)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="WawZXA3QCm5H" executionInfo={"status": "ok", "timestamp": 1629734357119, "user_tz": 360, "elapsed": 478868, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="f7bd2fa8-8192-4098-9e0a-c06b38b16175"
catboost_multi_xrf(X_train3, X_test3, y_train3, y_test3, 'OS2_catboost_MultiXRF.csv', max_iter)
| catboost/old_notebooks/catboost_multiXRF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from table_reader import TableReader
# Hey folks, I'm just trying out a proof-of-concept jupyter notebook that uses our data retrieval code.
#
# I got sick of working with environment variables so I switched to a new method to store our DB password:
# 1. Create a file called config.json in the project root.
# 2. Inside, config.json should look like this:
# {
# "database_url":"database_url_goes_here"
# }
#
# TableReader's other vector methods are geodata_vector() and reviews_vector(). Be sure to call close() when you're done so it terminates the connection to the DB.
tr = TableReader()
df = tr.properties_vector(include_amenitites=True)
print(df.head())
| airbnb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["remove_cell"]
# <div class="alert alert-warning">
#
# <b>Disclaimer:</b>
#
# The main objective of the <i>Jupyter</i> notebooks is to show how to use the models of the <i>QENS library</i> by
#
# - building a fitting model: composition of models, convolution with a resolution function
# - setting and running the fit
# - extracting and displaying information about the results
#
# These steps have a minimizer-dependent syntax. That's one of the reasons why different minimizers have been used in the notebooks provided as examples.
# But, the initial guessed parameters might not be optimal, resulting in a poor fit of the reference data.
#
# </div>
#
# # Lorentzian + background with lmfit
#
# ## Table of Contents
#
# - [Introduction](#Introduction)
# - [Importing the required libraries](#Importing-the-required-libraries)
# - [Plot of the fitting model](#Plot-of-the-fitting-model)
# - [Creating the reference data](#Creating-the-reference-data)
# - [Setting and fitting](#Setting-and-fitting)
# - [Plotting the results](#Plotting-the-results)
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Introduction
#
# <div class="alert alert-info">
#
# The objective of this notebook is to show how to combine the models of
# the <a href="https://github.com/QENSlibrary/QENSmodels">QENSlibrary</a>. Here, we use the <b>Lorentzian</b> profile and a flat background, created from <b>background_polynomials</b>, to perform some fits.
#
# <a href="https://lmfit.github.io/lmfit-py/">lmfit</a> is used for fitting.
# </div>
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Importing the required librairies
# + tags=["import_cell"]
# import python modules for plotting, fitting
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
# + tags=["remove_cell"]
import ipywidgets
# + tags=["remove_cell"]
# install QENSmodels (if not already installed)
import pkgutil
import sys
if not pkgutil.find_loader("QENSmodels"):
buttonY = ipywidgets.Button(description='Yes', button_style='success')
buttonN = ipywidgets.Button(description='No', button_style='danger')
choice_installation = ipywidgets.VBox(
[ipywidgets.Label("Do you want to install the QENSmodels' library?"), ipywidgets.HBox([buttonY, buttonN])],
layout=ipywidgets.Layout(width='50%', height='80px'))
display(choice_installation)
def on_buttonY_clicked(b):
# !{sys.executable} -m pip install git+https://github.com/QENSlibrary/QENSmodels#egg=QENSmodels
def on_buttonN_clicked(b):
print("You will not be able to run some of the remaining parts of this notebook")
buttonY.on_click(on_buttonY_clicked)
buttonN.on_click(on_buttonN_clicked)
# + tags=["remove_cell"]
# install lmfit (if not already installed)
if not pkgutil.find_loader("lmfit"):
lmfitY = ipywidgets.Button(description='Yes', button_style='success')
lmfitN = ipywidgets.Button(description='No', button_style='danger')
choice_installation = ipywidgets.VBox(
[ipywidgets.Label("Do you want to install lmfit?"), ipywidgets.HBox([lmfitY, lmfitN])],
layout=ipywidgets.Layout(width='30%', height='80px'))
display(choice_installation)
def on_lmfitY_clicked(b):
# !{sys.executable} -m pip install lmfit
def on_lmfitN_clicked(b):
print("You will not be able to run some of the remaining parts of this notebook")
lmfitY.on_click(on_lmfitY_clicked)
lmfitN.on_click(on_lmfitN_clicked)
# +
# required imports from lmfit
from lmfit import Model
# import model from QENS library
import QENSmodels
# + [markdown] tags=["remove_cell"]
# ### Physical units
#
# For information about unit conversion, please refer to the jupyter notebook called `Convert_units.ipynb` in the `tools` folder.
#
# The dictionary of units defined in the cell below specify the units of the refined parameters adapted to the convention used in the experimental datafile.
# -
# Units of parameters for selected QENS model and experimental data
dict_physical_units = {'omega': "1/ps",
'scale': "unit_of_signal/ps",
'center': "1/ps",
'hwhm': "1/ps"}
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Plot of the fitting model
#
# The widget below shows the lorentzian peak shape function with a constant background imported from QENSmodels where the functions' parameters *Scale*, *Center*, *FWHM* and *background* can be varied.
# + tags=["remove_cell"]
# Dictionary of initial values
ini_parameters = {'scale': 5, 'center': 0, 'hwhm': 3, 'background': 0.}
def interactive_fct(scale, center, hwhm, background):
xs = np.linspace(-10, 10, 100)
fig1, ax1 = plt.subplots()
ax1.plot(xs, QENSmodels.lorentzian(xs, scale, center, hwhm) +\
QENSmodels.background_polynomials(xs, background))
ax1.set_xlabel('x')
ax1.grid()
# Define sliders for modifiable parameters and their range of variations
scale_slider = ipywidgets.FloatSlider(value=ini_parameters['scale'],
min=0.1, max=10, step=0.1,
description='scale',
continuous_update=False)
center_slider = ipywidgets.IntSlider(value=ini_parameters['center'],
min=-10, max=10, step=1,
description='center',
continuous_update=False)
hwhm_slider = ipywidgets.FloatSlider(value=ini_parameters['hwhm'],
min=0.1, max=10, step=0.1,
description='hwhm',
continuous_update=False)
background_slider = ipywidgets.FloatSlider(value=ini_parameters['background'],
min=0.1, max=10, step=0.1,
description='background',
continuous_update=False)
grid_sliders = ipywidgets.HBox([ipywidgets.VBox([scale_slider, center_slider])
,ipywidgets.VBox([hwhm_slider, background_slider])])
# Define function to reset all parameters' values to the initial ones
def reset_values(b):
"""Reset the interactive plots to inital values."""
scale_slider.value = ini_parameters['scale']
center_slider.value = ini_parameters['center']
hwhm_slider.value = ini_parameters['hwhm']
background_slider.value = ini_parameters['background']
# Define reset button and occurring action when clicking on it
reset_button = ipywidgets.Button(description = "Reset")
reset_button.on_click(reset_values)
# Display the interactive plot
interactive_plot = ipywidgets.interactive_output(interactive_fct,
{'scale': scale_slider,
'center': center_slider,
'hwhm': hwhm_slider,
'background': background_slider})
display(grid_sliders, interactive_plot, reset_button)
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Creating the reference data
# -
# Create array of reference data: noisy lorentzian with background
nb_points = 100
xx = np.linspace(-5, 5, nb_points)
added_noise = 0.02 * np.random.normal(0, 1, nb_points)
lorentzian_noisy = QENSmodels.lorentzian(xx,
scale=0.89,
center=-0.025,
hwhm=0.45) * (1 + 5 * added_noise)
lorentzian_noisy += 0.5 * (1. + added_noise)
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Setting and fitting
# -
def flat_background(x, A0):
""" Define flat background ot be added to fitting model"""
return QENSmodels.background_polynomials(x, A0)
# +
gmodel = Model(QENSmodels.lorentzian) + Model(flat_background)
print('Names of parameters:', gmodel.param_names)
print('Independent variable(s):', gmodel.independent_vars)
initial_parameters_values = [1, 0.2, 0.5, 0.33]
# Fit
result = gmodel.fit(lorentzian_noisy,
x=xx,
scale=initial_parameters_values[0],
center=initial_parameters_values[1],
hwhm=initial_parameters_values[2],
A0=initial_parameters_values[3])
# -
# Plot initial model and reference data
fig0, ax0 = plt.subplots()
ax0.plot(xx, lorentzian_noisy, 'b-', label='reference data')
ax0.plot(xx, result.init_fit, 'k--', label='model with initial guesses')
ax0.set(xlabel='x', title='Initial model and reference data')
ax0.grid()
ax0.legend();
# + [markdown] tags=["remove_cell"]
# [Top](#Table-of-Contents)
#
# ## Plotting results
#
# using methods implemented in `lmfit`
# -
# display result
print('Result of fit:\n', result.fit_report())
# plot fitting result using lmfit functionality
result.plot()
# Plot fitting result and reference data using matplotlib.pyplot
fig1, ax1 = plt.subplots()
ax1.plot(xx, lorentzian_noisy, 'b-', label='reference data')
ax1.plot(xx, result.best_fit, 'r.', label='fitting result')
ax1.legend()
ax1.set(xlabel='x', title='Fit result and reference data')
ax1.grid();
for item in ['hwhm', 'center', 'scale']:
print(item,
result.params[item].value, '+/-', result.params[item].stderr, dict_physical_units[item])
| docs/examples/lmfit_lorentzian_and_backgd_fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="lpvOZX_8CBxg"
# # SIGN LANGUAGE MULTILABEL CLASSIFICATION WITH FASTAI
# + [markdown] id="5KQLy6ExCbYR"
# ## INSTALL AND IMPORT ALL THE NECESSARY LIBRARIES
# + id="aWFGMDpsB9wi"
# These are all the necessary libraries we have to install
# !pip install fastai==2.2.0
# !pip install -U albumentations
# !pip install opencv-python==4.5.4.60
# !pip install -q iterative-stratification
# + id="X4DTyiJRByd-"
#import albumentations for data augmentation
import albumentations
#fastai is built on top of torch so import it
import torch
torch.__version__
# + id="O-fBO25OByeC"
#import everything from fastai
from fastai.vision.all import *
#import multilabelstratifiedkfold to be used to create folds
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
# + [markdown] id="LmXF4P1QCstl"
# ## DATA LOADING AND VIEWING
# + id="UVS85lrCByeD"
# loading the train data and appending the .jpg extension for easy manipulation
path = Path('../input/signprediction/sign_prediction')
train_df = pd.read_csv(path/'Train.csv')
train_df['img_IDS'] = train_df['img_IDS'].apply(str) + ".jpg"
train_df['img_IDS'] = "../input/signprediction/sign_prediction/Images/" + train_df['img_IDS']
train_df.head()
# + [markdown] id="Cvflpi3GC2j-"
# ### CREATING STRATIFIED KFOLDS
# + id="9ewacMlaByeE"
#creating 3 shuffled stratified kfold
strat_kfold = MultilabelStratifiedKFold(n_splits=3, random_state=42, shuffle=True)
train_df['fold'] = -1
for i, (_, test_index) in enumerate(strat_kfold.split(train_df.img_IDS.values, train_df.iloc[:,1:].values)):
train_df.iloc[test_index, -1] = i
train_df.head()
# + id="fk5YEdYOByeG"
#plot the folds to ensure that they are of equal portions
train_df.fold.value_counts().plot.bar();
# + [markdown] id="6aXsUMbTDAUM"
# ### CREATING A SPECIAL TRANSFORM CLASS TO USE ALBUMENTATIONS
# + id="xe5r4A8AByeI"
class AlbumentationsTransform (RandTransform):
split_idx,order=None,2
def __init__(self, train_aug, valid_aug): store_attr()
def before_call(self, b, split_idx):
self.idx = split_idx
def encodes(self, img: PILImage):
if self.idx == 0:
aug_img = self.train_aug(image=np.array(img))['image']
else:
aug_img = self.valid_aug(image=np.array(img))['image']
return PILImage.create(aug_img)
# + id="CzjDYuwEByeJ"
#define all the augmentations you need
def get_train_aug(): return albumentations.Compose([
albumentations.Resize(512, 512),
# albumentations.RandomBrightness(),
# albumentations.RandomRotate90(),
# albumentations.Rotate(limit=(-90, 90)),
# albumentations.Transpose(),
# albumentations.RandomContrast(),
# albumentations.RandomBrightnessContrast(),
# albumentations.RandomGamma(),
# albumentations.Blur(),
albumentations.HorizontalFlip(p=0.5),
albumentations.VerticalFlip(p=0.5),
albumentations.ShiftScaleRotate(p=0.5),
albumentations.HueSaturationValue(
hue_shift_limit=0.2,
sat_shift_limit=0.2,
val_shift_limit=0.2,
p=0.5
),
albumentations.RandomBrightnessContrast(
brightness_limit=(-0.1,0.1),
contrast_limit=(-0.1, 0.1),
p=0.5
),
], p=1.)
def get_valid_aug(): return albumentations.Compose([
albumentations.Resize(512, 512),
], p=1.0)
item_tfms = AlbumentationsTransform(get_train_aug(), get_valid_aug())
batch_tfms = [Normalize.from_stats(*imagenet_stats)]
# + [markdown] id="qDQ85PCMDK8z"
# ### CREATING THE FASTAI DATABLOCK
# + id="rvdCjbh2ByeK"
# to learn more about datablocks in fast you have to visit fastai.docs
def get_data(fold=0, size=224,bs=32):
return DataBlock(blocks=(ImageBlock,MultiCategoryBlock),
get_x=ColReader(0),
get_y=ColReader(1, label_delim=' '),
splitter=IndexSplitter(train_df[train_df.fold == fold].index),
item_tfms = item_tfms,
batch_tfms = batch_tfms).dataloaders(train_df, bs=bs)
# + [markdown] id="gFbcRCF4DVSW"
# ### CREATING YOUR OWN SPECIAL METRICS
# + id="ibu2BxFdByeL"
# This is not necessary since fastai already has an inbuilt accuracy_multi metrics for multilabel classifications
def accuracy_multi(inp, targ, thresh=0.5, sigmoid=True):
"Compute accuracy when `inp` and `targ` are the same size."
if sigmoid: inp = inp.sigmoid()
return ((inp>thresh)==targ.bool()).float().mean()
# + id="Yz7Y99WyByeM"
def F_score(output, label, threshold=0.2, beta=1):
prob = output > threshold
label = label > threshold
TP = (prob & label).sum(1).float()
TN = ((~prob) & (~label)).sum(1).float()
FP = (prob & (~label)).sum(1).float()
FN = ((~prob) & label).sum(1).float()
precision = torch.mean(TP / (TP + FP + 1e-12))
recall = torch.mean(TP / (TP + FN + 1e-12))
F2 = (1 + beta**2) * precision * recall / (beta**2 * precision + recall + 1e-12)
return F2.mean(0)
# + [markdown] id="rGvcDLtkDiEE"
# ### LOADING THE TEST DATA AND MAKE NECESSARY MODIFICATIONS
# + id="B279LRV3ByeO"
#load the test data and add the .jpg extension plus the path
test_df = pd.read_csv('../input/signprediction/sign_prediction/SampleSubmission.csv')
tstpng = test_df.copy()
tstpng['img_IDS'] = tstpng['img_IDS'].apply(str) + ".jpg"
tstpng['img_IDS'] = "../input/signprediction/sign_prediction/Images/" + tstpng['img_IDS']
tstpng.head()
# + [markdown] id="EsGzlGHJDuxH"
# ## MODELLING
# + id="jPON_vG0ByeP"
# I am going to use mixup in my training, for more information about how mix up work go to the official fastai.docs
mixup = MixUp(0.3)
# + id="0vM3AfOpByeQ"
import gc
# + id="RxRbUOmGByeQ"
#in here we are going to do both our training and inference using resnet34 architecture
#I have also used alot of advanced techniques like callbacks and if you dont understand please visit the official fastai.docs
all_preds = []
for i in range(3):
dls = get_data(i,256,64)
learn = cnn_learner(dls, resnet34, metrics=[partial(accuracy_multi, thresh=0.2),partial(F_score, threshold=0.2)],cbs=mixup).to_fp16()
learn.fit_one_cycle(10, cbs=EarlyStoppingCallback(monitor='valid_loss'))
learn.dls = get_data(i,512,32)
learn.fine_tune(10,cbs=EarlyStoppingCallback(monitor='valid_loss'))
tst_dl = learn.dls.test_dl(tstpng)
preds, _ = learn.get_preds(dl=tst_dl)
all_preds.append(preds)
del learn
torch.cuda.empty_cache()
gc.collect()
# + [markdown] id="qc2b-cfXEDWp"
# ## PREPARING THE SUBMISSION
# + id="jx3mhmpDByeR"
preds = np.mean(np.stack(all_preds), axis=0)
# + id="AF9XbKLlByeR"
k=[]
for col in tstpng.columns:
k.append(col) # creating list of the label
k
# + id="lKnGlKLnByeS"
test_df = pd.read_csv("../input/signprediction/sign_prediction/Test.csv")
# + id="pQzmWU6kByeT"
import os
submission = pd.DataFrame()
submission["ID"] = test_df["img_IDS"]
for i, c in enumerate(dls.vocab):
print(c)
submission[c] = preds[:,i]
submission.head()
# + id="zmuvIJoDByeT"
submission.to_csv('baseline_model_16.csv', index=False)
# + id="uhnH4tzdByeU"
| Signlanguagemultilabelclassification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(1, "..")
from quizlet import display_quiz
# -
questions=[]
# # Add questions here
# + tags=["remove-input"]
question="Which of these cannot be a probability?"
answers=[
{'answer':"0.5",
'correct': False,
'feedback': "This is a number between 0 and 1, so it can be probability."
},
{'answer': "-0.5",
'correct': True,
'feedback': "Correct! Probabilities must be numbers between 0 and 1."
},
{'answer': "0",
'correct': False,
'feedback': "This is a number between 0 and 1, so it can be probability."
},
{'answer': "1",
'correct': False,
'feedback': "This is a number between 0 and 1, so it can be probability."
}
]
questions+=[{'question':question, 'type':'multiple_choice', 'answers':answers}]
display_quiz(question, answers)
# + tags=["remove-input"]
question='''Determine the relative frequency of the outcome 3 in the following data set:
2,3,5,2,3'''
answers=[
{'answer':"2",
'correct': False,
'feedback': "Relative frequency is the proportion of experiments in which an outcome occurs. It is always between 0 and 1."
},
{'answer': "0",
'correct': False,
'feedback': "Relative frequency is the proportion of experiments in which an outcome occurs. Since 3 occured, that proportion will not be 0."
},
{'answer': "0.4",
'correct': True,
'feedback': "Correct! The outcome 3 occurred 2 times out of 5 experiments, so the relative frequency is 2/5=0.4."
},
{'answer': "0.6",
'correct': False,
'feedback': "Relative frequency is the proportion of experiments in which an outcome occurs. Count the number of times that outcome 3 occurred and divide by the total number of experiments."
}
]
questions+=[{'question':question, 'type':'multiple_choice', 'answers':answers}]
display_quiz(question, answers)
# -
len(questions)
questions
# # Now write out to file
import json
with open('../questions/ch2.txt', 'w') as outfile:
json.dump(questions, outfile, indent=4)
# # Read back and display questions
# +
import json
questions=[]
with open('../questions/ch2.txt', 'r') as infile:
questions=json.load(infile)
# -
questions
for question in questions:
display_quiz(question)
type(question)
import random
# +
random.sample(questions, k=2)
# -
len(questions)
questions
questions=random.sample(questions,2)
questions
from quizlet import display_multiple
display_multiple(questions)
display_multiple(questions, shuffle=True)
display_multiple(questions, 1)
| 02-first-stats/make-questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Anaconda3]
# language: python
# name: conda-env-Anaconda3-py
# ---
# # MSDS 462 Assignment 1 Fashion - MNIST
# Training a CNN for Fashion MNIST dataset using Keras and Tensorflow. Fashion MNIST dataset contains Grayscale images from 10 classes. The train dataset has 60000 images (6000 per class) and the test dataset has 10000 images. This CNN has two Convolutional layers, works on a 3 x 3 kernel, activation function is 'relu'. The final output layer consists of 10 output nodes with 'softmax' function. The 'Adam' optimizer was used along with 'sparse_categorical_crossentropy' loss function.
#
# The model took nearly 8 hours to train and reached a predictive accuracy of 94% and a validation accuracy of 92%.
# +
###############################################################################
# Pyhton packages
###############################################################################
from __future__ import absolute_import, division, print_function, unicode_literals
# To remove all future warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import datasets, layers, models
from keras.datasets import fashion_mnist
#from keras import models
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator, load_img
from keras.models import Sequential, load_model
from keras.layers import Conv2D, MaxPooling2D, Input
from keras.layers import Flatten, Dense, Dropout
from keras.models import Model
from keras.optimizers import Adam, RMSprop, SGD
from keras import regularizers
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.utils import plot_model
from keras import backend as K
from sklearn.metrics import roc_curve, auc
import skimage
from skimage.transform import resize
from matplotlib import pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
import glob
import os, sys
import math
import pandas as pd
import time
from contextlib import redirect_stdout
import time
import numpy as np
# -
# ### Loading and processing data
# +
###############################################################################
# Download and normalization pre-processing (0-1) of the Fashion-MNIST dataset
###############################################################################
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
height=28
width=28
#Checking number of images per class
objclass, counts = np.unique(train_labels, return_counts = True)
print("Updated count of images per class")
dict(zip(objclass, counts))
# Shape of the image datasets
print(train_images.shape)
print(train_labels.shape)
print(test_images.shape)
print(test_labels.shape)
# +
# Class Names for the dataset
class_names = ['top', 'trouser', 'pullover', 'dress', 'coat',
'sandal', 'shirt', 'sneaker', 'bag', 'boot']
plt.figure(figsize=(1,1))
plt.imshow(train_images[25])
plt.xlabel(class_names[train_labels[25]])
plt.show()
# +
start_time = time.clock()
class_name_dict = {
0: 'top',
1: 'trouser',
2: 'pullover',
3: 'dress',
4: 'coat',
5: 'sandal',
6: 'shirt',
7: 'sneaker',
8: 'bag',
9: 'boot',
}
def draw_img(i):
im = train_images[i]
c = train_labels[i]
plt.imshow(im)
plt.title("Class %d (%s)" % (c, class_name_dict[c]))
plt.axis('on')
im
def draw_sample(X, y, n, rows=4, cols=4, imfile=None, fontsize=12):
for i in range(0, rows*cols):
plt.subplot(rows, cols, i+1)
im = X[n+i].reshape(28,28,1)
plt.imshow(im, cmap='gnuplot2')
plt.title("{}".format(class_name_dict[y[n+i][0]]), fontsize=fontsize)
plt.axis('off')
plt.subplots_adjust(wspace=0.6, hspace=0.01)
#plt.subplots_adjust(hspace=0.45, wspace=0.45)
#plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=1.0)
if imfile:
plt.savefig(imfile)
def central_scale_images(X_imgs, scale):
# scaled_imgs = central_scale_images(X_imgs, [0.90, 0.75, 0.60])
# Various settings needed for Tensorflow operation
boxes = np.zeros((1, 4), dtype = np.float32)
for index, scale in enumerate([scale]):
x1 = y1 = 0.5 - 0.5 * scale # To scale centrally
x2 = y2 = 0.5 + 0.5 * scale
boxes[index] = np.array([y1, x1, y2, x2], dtype = np.float32)
box_ind = np.zeros(1, dtype = np.int32)
crop_size = np.array([height, width], dtype = np.int32)
X_scale_data = []
tf.reset_default_graph()
X = tf.placeholder(tf.float32, shape = (1, height, width, 3))
# Define Tensorflow operation for all scales but only one base image at a time
tf_img = tf.image.crop_and_resize(X, boxes, box_ind, crop_size)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for img_data in X_imgs:
batch_img = np.expand_dims(img_data, axis = 0)
scaled_imgs = sess.run(tf_img, feed_dict = {X: batch_img})
X_scale_data.extend(scaled_imgs)
X_scale_data = np.array(X_scale_data, dtype = np.float32)
return X_scale_data
###############################################################################
# Finish pre-processing (0-1) of the Fashion-MNIST dataset
###############################################################################
# Normalize pixel values to be between 0 and 1
train_images, test_images = train_images / 255.0, test_images / 255.0
###############################################################################
# Visualizing the first 50 images in the dataset
###############################################################################
plt.figure(figsize=(10,10))
for i in range(50):
plt.subplot(5,10,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
# The CIFAR labels happen to be arrays,
# which is why you need the extra index
plt.xlabel(class_names[train_labels[i]])
plt.show()
#Checking number of images per class in the new Train Set
objclass, counts = np.unique(train_labels, return_counts = True)
print("Updated count of images per class")
print(dict(zip(objclass, counts)))
#Checking number of images per class in the new Test Set
objclass, counts = np.unique(test_labels, return_counts = True)
print("Updated count of images per class")
print(dict(zip(objclass, counts)))
# -
# ### Convolutional Neural Network Code
# To be run only if a saved model is not available otherwise load from existing model
# Code to load existing model to avoid retraining multiple times
# Commenting the code for now
model = keras.models.load_model('cnn_fashionmnist_model_20191201.h5')
# +
###############################################################################
# Convolutional Neural Network Architecture
# Build a Convolutional Neural Network in four blocks:
# 1.Convolution layers ==> 2.Pooling layer ==> 3.Flattening layer ==> 4.Dense/Output layer
# Dropout - addresses overfitting
# Kernel_regularizer - penalizes on layer parameters during optimization - L2 regularisation of the weights
# BatchNormalization - addresses the internal covariate shift, even eliminating the need for dropout
# and achieves same accuracy with fewer training steps speeding up the training process
# Configure CNN tensors shape (image_height, image_width, color_channels)
# Inputs orignal format of Fashion-MNIST images of shape (28, 28, 1)
# Apply Conv2D and MaxPooling2D layer
# Color channel is Grayscale
# Display the model architecture
# Output of Conv2D and MaxPooling2D layers in 3D tensor shape (height, width, channels)
# Number of output channels for Conv2D layers = 32, 64
# Print the model architecture
###############################################################################
weight_decay = 1e-4
# Initializing the model
model = models.Sequential()
# Step 1. Convolution layer
# padding the input such that the output has the same length as the original input
# Step 2. Pooling layer: add MaxPooling to downsample and Dropout to prevent overfitting
# 1st convolutional layer
model.add(layers.Conv2D(32, (3, 3), padding='same', activation='relu',
kernel_regularizer=regularizers.l2(weight_decay),
input_shape=(28, 28, 1)))
model.add(layers.BatchNormalization())
model.add(layers.Conv2D(32, (3, 3), padding='same', activation='relu',
kernel_regularizer=regularizers.l2(weight_decay)))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Dropout(0.2))
# 2nd convolutional layer
model.add(layers.Conv2D(64, (3, 3), padding='same', activation='relu',
kernel_regularizer=regularizers.l2(weight_decay)))
model.add(layers.BatchNormalization())
model.add(layers.Conv2D(64, (3, 3), padding='same', activation='relu',
kernel_regularizer=regularizers.l2(weight_decay)))
model.add(layers.BatchNormalization())
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Dropout(0.3))
model.summary()
# -
###############################################################################
# Step 3. Flattening layer from 3D output to 1D
# Step 4. Dense/Output layer: one or more Dense layers
# Add final Dense layer of 10 outputs and softmax activation
# Print the model architecture
###############################################################################
# Step 3. Flattening layer: from 3D output to 1D
model.add(layers.Flatten())
# Step 4. Dense/Output layer
# softmax as our last activation function
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(10, activation='softmax'))
# Print the model architecture
model.summary()
# +
train_images = np.reshape(train_images, (60000,28,28,1))
train_labels = np.reshape(train_labels, (60000,1))
test_images = np.reshape(test_images, (10000,28,28,1))
test_labels = np.reshape(test_labels, (10000,1))
print(train_images.shape)
print(train_labels.shape)
print(test_images.shape)
print(test_labels.shape)
# -
# ### Training the Model using 60000 Train images and 10000 Validation samples
# +
###############################################################################
# Train the model
# Train on 60000 samples, validate on 10000 samples
###############################################################################
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(train_images, train_labels, epochs=10,
validation_data=(test_images, test_labels))
# +
# Recording the time took for training
print("Time Elapsed = ", time.clock() - start_time)
# Save the model
model.save('cnn_fashionmnist_model_20191201.h5')
# -
# ### Evaluating the model
# +
###############################################################################
# Evaluate the model
# Visualize the model accuracy
###############################################################################
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
plt.plot(epochs, acc, 'bo', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and Validation Accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training Loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and Validation Loss')
plt.legend()
plt.show()
###############################################################################
# Evaluate the model
# Visualize the model accuracy
###############################################################################
plt.plot(history.history['acc'], label='accuracy')
plt.plot(history.history['val_acc'], label = 'val_accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.5, 1])
plt.legend(loc='lower right')
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
# +
###############################################################################
# Predicting new images
###############################################################################
plt.figure(figsize=(1,1))
plt.imshow(train_images[25])
plt.xlabel(class_names[train_labels[25]])
plt.show()
###############################################################################
# Predicting images
###############################################################################
x = image.img_to_array(train_images[25])
x = np.expand_dims(x, axis=0)
images = np.vstack([x])
classes = model.predict_classes(images, batch_size=10)
print("Predicted class is:",classes,"-",class_name_dict[classes[0]])
| MSDS 462 A1 Fashion-MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="7LyR6UM6lW7L"
# <h2> A. Library Import </h2>
# + id="MVMrp3bplW7N"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# compare knn imputation strategies for the horse colic dataset
from numpy import mean
from numpy import std
from pandas import read_csv
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import KNNImputer
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import RepeatedStratifiedKFold
from sklearn.pipeline import Pipeline
from matplotlib import pyplot
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn import preprocessing
from sklearn.experimental import enable_iterative_imputer
from sklearn.impute import IterativeImputer
import os
import warnings
warnings.filterwarnings('ignore')
# + id="-klafa2nlW7P"
#type_of_target(Y_temp)
# + [markdown] id="a1NJdHuhlW7P"
# <h2>B. Function</h2>
# + [markdown] id="7m6CbvpElW7P"
# <h3> B.1 .Find missing percentage value </h3>
# + id="P-AaT8U1lW7Q"
def missing_percent(df):
# Total missing values
mis_val = df.isnull().sum()
# Percentage of missing values
mis_percent = 100 * df.isnull().sum() / len(df)
# Make a table with the results
mis_table = pd.concat([mis_val, mis_percent], axis=1)
# Rename the columns
mis_columns = mis_table.rename(
columns = {0 : 'Missing Values', 1 : 'Percent of Total Values'})
# Sort the table by percentage of missing descending
mis_columns = mis_columns[
mis_columns.iloc[:,1] != 0].sort_values(
'Percent of Total Values', ascending=False).round(2)
# Print some summary information
print ("Your selected dataframe has " + str(df.shape[1]) + " columns.\n"
"There are " + str(mis_columns.shape[0]) +
" columns that have missing values.")
# Return the dataframe with missing information
return mis_columns
# + [markdown] id="IaEVsl3DlW7Q"
# <h3> B.2 KNN Imputer </h3>
#
# * Use MinMaxScaler to scale X
# * Use LabelBinarizer to convert multiclass Y to binary Y
# + id="93Rm0diTlW7Q"
def optimize_knn_imputer(data, target,name):
results = list()
dict = {'Name':[],
'K':[],
'MeanAccuracy':[],
'stdAccuracy':[]
}
df = pd.DataFrame(dict)
for s in range(1, 50, 2):
imputer = KNNImputer(n_neighbors=s, weights='distance', metric='nan_euclidean')
array = data.values
X = array [:,:-1]
Y = data[target]
#lb = preprocessing.LabelBinarizer()
#Y_binary = lb.fit_transform(Y)
#Y_binary = np.array([number[0] for number in lb.fit_transform(Y)])
data['num'][data.num >0] = 1
Y_temp = data['num']
trans = MinMaxScaler()
trans_X = trans.fit_transform(X)
X_imputed = imputer.fit_transform(trans_X)
X_imputed_df = pd.DataFrame(X_imputed, columns=data.drop(target, axis=1).columns)
model = RandomForestClassifier()
cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=3, random_state=1)
scores = cross_val_score(model, X_imputed, Y_temp, scoring='accuracy', cv=cv, n_jobs=-1)
results.append(scores)
df.loc[len(df.index)] = [name, s, mean(scores), std(scores)]
#print('>K : %s --> Mean : %.3f --> Std : (%.3f)' % (s, mean(scores), std(scores)))
#max(results)
display(df)
pyplot.boxplot(results, labels=range(1, 50, 2), showmeans=True)
pyplot.show()
return df
#df.loc[df['MeanAccuracy'] == df['MeanAccuracy'].max()]
# + [markdown] id="ZXwX2UTslW7R"
# <h3> B.3 FinalDataset Prep </h3>
# + id="0j4zdyCPlW7R"
def Final_Dataset_Prep(data, target , K):
imputer = KNNImputer(n_neighbors=K, weights='distance', metric='nan_euclidean')
array = data.values
X = array[:,:-1]
trans = MinMaxScaler()
trans_X = trans.fit_transform(X)
X_imputed = imputer.fit_transform(trans_X)
X_imputed_df = pd.DataFrame(X_imputed, columns=data.drop(target, axis=1).columns)
data['num'][data.num >0] = 1
Y_temp = data['num']
Final_Dataset = pd.concat([d.reset_index(drop=True) for d in [X_imputed_df, Y_temp]], axis=1)
return Final_Dataset
# + [markdown] id="uBYX4dTalW7R"
# <h2> C . Import Data</h2>
# + id="WWO9nK0KlW7S"
cleveland_file = "./processed.cleveland.data"
cleveland_df = pd.read_csv(cleveland_file, header=None)
cleveland_df.columns = ["age", "sex", "cp", "trestbps","chol", "fbs", "restecg","thalach", "exang", "oldpeak", "slope", "ca", "thal", "num"]
cleveland_df = cleveland_df.replace('?', np.nan)
hungarian_file = "./processed.hungarian.data"
hungarian_df = pd.read_csv(hungarian_file, header=None)
hungarian_df.columns = ["age", "sex", "cp", "trestbps","chol", "fbs", "restecg","thalach", "exang", "oldpeak", "slope", "ca", "thal", "num"]
hungarian_df = hungarian_df.replace('?', np.nan)
switzerland_file = "./processed.switzerland.data"
switzerland_df = pd.read_csv(switzerland_file, header=None)
switzerland_df.columns = ["age", "sex", "cp", "trestbps","chol", "fbs", "restecg","thalach", "exang", "oldpeak", "slope", "ca", "thal", "num"]
switzerland_df = switzerland_df.replace('?', np.nan)
va_file = "./processed.va.data"
va_df = pd.read_csv(va_file, header=None)
va_df.columns = ["age", "sex", "cp", "trestbps","chol", "fbs", "restecg","thalach", "exang", "oldpeak", "slope", "ca", "thal", "num"]
va_df = va_df.replace('?', np.nan)
# + [markdown] id="4n3V7oBLlW7S"
# <h2> D. Find Missing </h2>
# + id="V5sviklolW7S" outputId="966de610-2b27-409b-995e-6cdf1749697b"
cleveland_nan = pd.DataFrame(cleveland_df.isnull().sum())
hungarian_nan = pd.DataFrame(hungarian_df.isnull().sum())
switzerland_nan = pd.DataFrame(switzerland_df.isnull().sum())
va_nan = pd.DataFrame(va_df.isnull().sum())
Disease_NAN = pd.concat([cleveland_nan, hungarian_nan, switzerland_nan, va_nan], axis=1)
Disease_NAN.columns = ["cleveland NAN", "hungarian NAN", "switzerland NAN", "va NAN"]
Disease_NAN
# + [markdown] id="ArEesjwZlW7T"
# <h2> E. Creating different Dataframe </h2>
# + [markdown] id="959YK28VlW7U"
# Below are the combination of different dataset
# * Cleveland , hungarian , switzerland , Va
# * Cleveland , hungarian
# * Cleveland , switzerland
# * Cleveland , hungarian , switzerland
# * Cleveland
#
# We are trying to find best K value for KNN_Imputer by applying RandomforestClassifier to find bext accuracy
# + [markdown] id="9Btdv_UAlW7U"
# <h2> E.1 Cleveland + hungarian + switzerland + Va </h2>
# + id="gurISBzTlW7V"
Heart_Disease_DF = pd.concat([cleveland_df,hungarian_df,switzerland_df,va_df])
# + id="moqiZb62lW7V" outputId="a5dc8556-5fec-4a3f-8666-050686af2b5b"
df1 = optimize_knn_imputer(Heart_Disease_DF,'num','cleveland+hungarian+switzerland+va')
# + id="HEkOGmOglW7W" outputId="235bc91b-ec3b-433d-e858-47138e288a7b"
df1.loc[df1['MeanAccuracy'] == df1['MeanAccuracy'].max()]
# + [markdown] id="djEXoZz7lW7W"
# <h2> E.2 Cleveland + hungarian </h2>
# + id="kP13IeNplW7W"
Clev_hung_df = pd.concat([cleveland_df,hungarian_df])
# + id="E9jCtORzlW7X" outputId="3b550f55-7f04-4bf8-817f-7cca44c19c84"
df2 = optimize_knn_imputer(Clev_hung_df,'num','cleveland+hungarian')
# + id="4piEU-4FlW7X" outputId="328b7648-00bd-48ff-af19-fc3970e77746"
df2.loc[df2['MeanAccuracy'] == df2['MeanAccuracy'].max()]
# + [markdown] id="sjmCpFQflW7X"
# <h2> E.3 Cleveland + switzerland </h2>
# + id="4ELq7IkdlW7Y"
Clev_switz_df = pd.concat([cleveland_df,switzerland_df])
# + id="II11H9pflW7Y" outputId="aff3287e-799a-4f73-fe94-1b04b652e6b7"
df3 = optimize_knn_imputer(Clev_switz_df,'num','cleveland+switzerland')
# + id="SK4L-h8LlW7Y" outputId="ed078a11-85da-4e4d-ee9d-55d2233b41ec"
df3.loc[df3['MeanAccuracy'] == df3['MeanAccuracy'].max()]
# + [markdown] id="AWfF9Zi_lW7Y"
# <h2> E.4 Cleveland + switzerland + hungarian</h2>
# + id="g9DR6D5JlW7Z"
Clev_hung_switz_df = pd.concat([cleveland_df,hungarian_df,switzerland_df])
# + id="mATFXMHXlW7Z"
df4 = optimize_knn_imputer(Clev_hung_switz_df,'num','cleveland+hungarian+switzerland')
# + id="RSp2-FDYlW7Z"
df4.loc[df4['MeanAccuracy'] == df4['MeanAccuracy'].max()]
# + [markdown] id="M9-OjjVxlW7Z"
# <h2> E.5 Cleveland </h2>
# + id="oYjYm9MvlW7Z" outputId="2b12f611-426f-4274-a3fd-3e2d65180850"
df5 = optimize_knn_imputer(cleveland_df,'num','cleveland')
# + id="BHh2yl2ClW7a" outputId="2e7f890e-835c-43bd-f7b5-69ef6b7104da"
df5.loc[df5['MeanAccuracy'] == df5['MeanAccuracy'].max()]
# + id="ACz65t3epqyi"
# + [markdown] id="LM_aR4ItlW7a"
# <h2> E . Final Imputation DataSet </h2>
# + id="OZOKFzdglW7a"
df1_max = df1.loc[df1.sort_values(by='K', ascending=False)['MeanAccuracy'] == df1.sort_values(by='K', ascending=False)['MeanAccuracy'].max()]
df2_max = df2.loc[df2.sort_values(by='K', ascending=False)['MeanAccuracy'] == df2.sort_values(by='K', ascending=False)['MeanAccuracy'].max()]
df3_max = df3.loc[df3.sort_values(by='K', ascending=False)['MeanAccuracy'] == df3.sort_values(by='K', ascending=False)['MeanAccuracy'].max()]
df4_max = df4.loc[df4.sort_values(by='K', ascending=False)['MeanAccuracy'] == df4.sort_values(by='K', ascending=False)['MeanAccuracy'].max()]
df5_max = df5.loc[df5.sort_values(by='K', ascending=False)['MeanAccuracy'] == df5.sort_values(by='K', ascending=False)['MeanAccuracy'].max()]
Optimum_df = pd.concat([df1_max,df2_max,df3_max,df4_max,df5_max])
Optimum_df.reset_index().sort_values(by='MeanAccuracy', ascending=False, na_position='first')
# + id="nMPaq5AdlW7a"
data = Clev_hung_switz_df
target = 'num'
K = 3
Final_DF = Final_Dataset_Prep(data,target,K)
# + id="h6UY96ZblW7b" outputId="2955e7ea-635e-4e8c-807d-1c6a671e095c"
Final_DF
# + id="ffbcmWIolW7b"
final_df_nan = pd.DataFrame(Final_DF.isnull().sum())
final_df_nan
# + id="yF7q0D2IlW7c"
Final_DF.to_csv(r'.\Final_Dataset.csv', index = False, header=True)
| KNN_Imputer_MissingValue.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pylab as plt
import string
import pandas as pd
import matplotlib.pylab as plt
import string #导入字符串模块
# -
# # 数据预处理
tab1 = "./hair_dryer.tsv"
print(tab1)
tab2 = "."
df_hd = pd.read_csv(tab1, sep='\t', header=0)
df_hd.head()
# ## 删除无效信息
df_hd = df_hd[~(df_hd['vine'].str.contains("N") & df_hd['verified_purchase'].str.contains("N"))]
df_hd = df_hd[(df_hd['total_votes'] > 0) & (df_hd['helpful_votes'] > 0)]
df_hd = df_hd.groupby('product_parent').filter(lambda x: len(x) > 1)
df_hd.describe(include="all")
# ## 按 product_parent 分组
# +
gp_pp = df_hd.groupby('product_parent')
# # 选特定的一组
# print(gp_pp.get_group(732252283)['star_rating'])
# # 遍历
# for item in gp_pp:
# print(item[0])
# print(item[1]['star_rating'])
# gp_pp[['star_rating', 'helpful_votes', 'total_votes']].sum()
gp_pp[['star_rating', 'helpful_votes', 'total_votes']].sum().sort_values(by='helpful_votes',ascending=False)
# -
df_pp = gp_pp.get_group(47684938)
df_pp_1 = df_pp[df_pp['star_rating'] == 1]
df_pp_2 = df_pp[df_pp['star_rating'] == 2]
df_pp_3 = df_pp[df_pp['star_rating'] == 3]
df_pp_4 = df_pp[df_pp['star_rating'] == 4]
df_pp_5 = df_pp[df_pp['star_rating'] == 5]
# +
pattern = r"\&\#[0-9]+\;"
df_pp_5["preprocessed"] = df_pp_5["review_body"].str.replace(pat=pattern, repl="", regex=True)
print(df_pp_5["preprocessed"].iloc[2])
# +
import re
import nltk
from nltk import word_tokenize, pos_tag
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import sent_tokenize
from nltk.corpus import wordnet
#import nltk resources
resources = ["wordnet", "stopwords", "punkt", \
"averaged_perceptron_tagger", "maxent_treebank_pos_tagger"]
for resource in resources:
try:
nltk.data.find("tokenizers/" + resource)
except LookupError:
nltk.download(resource)
#create Lemmatizer object
lemma = WordNetLemmatizer()
def lemmatize_word(tagged_token):
""" Returns lemmatized word given its tag"""
root = []
for token in tagged_token:
tag = token[1][0]
word = token[0]
if tag.startswith('J'):
root.append(lemma.lemmatize(word, wordnet.ADJ))
elif tag.startswith('V'):
root.append(lemma.lemmatize(word, wordnet.VERB))
elif tag.startswith('N'):
root.append(lemma.lemmatize(word, wordnet.NOUN))
elif tag.startswith('R'):
root.append(lemma.lemmatize(word, wordnet.ADV))
else:
root.append(word)
return root
# -
def lemmatize_doc(document):
""" Tags words then returns sentence with lemmatized words"""
lemmatized_list = []
tokenized_sent = sent_tokenize(document)
for sentence in tokenized_sent:
no_punctuation = re.sub(r"[`'\",.!?()]", " ", sentence)
tokenized_word = word_tokenize(no_punctuation)
tagged_token = pos_tag(tokenized_word)
lemmatized = lemmatize_word(tagged_token)
lemmatized_list.extend(lemmatized)
return " ".join(lemmatized_list)
# +
#apply our functions
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].apply(lambda row: lemmatize_doc(row))
print(df_pp_5["preprocessed"].iloc[2])
# +
from unicodedata import normalize
remove_accent = lambda text: normalize("NFKD", text).encode("ascii", "ignore").decode("utf-8", "ignore")
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].apply(remove_accent)
print(df_pp_5["preprocessed"].iloc[2])
# +
pattern = r"[^\w\s]"
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].str.replace(pat=pattern, repl=" ", regex=True)
print(df_pp_5["preprocessed"].iloc[2])
# +
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].str.lower()
print(df_pp_5["preprocessed"].iloc[2])
# +
from nltk.corpus import stopwords
stop_words = stopwords.words("english")
stop_words = [word.replace("\'", "") for word in stop_words]
stop_words.append('br')
print(f"sample stop words: {stop_words[:15]} \n")
remove_stop_words = lambda row: " ".join([token for token in row.split(" ") \
if token not in stop_words])
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].apply(remove_stop_words)
print(df_pp_5["preprocessed"].iloc[2])
# +
pattern = r"[\s]+"
df_pp_5["preprocessed"] = df_pp_5["preprocessed"].str.replace(pat=pattern, repl=" ", regex=True)
print(df_pp_5["preprocessed"].iloc[2])
# +
corpora = df_pp_5["preprocessed"].values
tokenized = [corpus.split(" ") for corpus in corpora]
print(tokenized[2])
# -
print(tokenized[2])
text = nltk.pos_tag(tokenized[2])
print(len(text))
# +
hist_n = {} #创建一个空字典,放名词词频与单词,无序排列
hist_adj = {} #创建一个空字典,放形容词词频与单词,无序排列
data_n = [] #创建一个空列表,放词频与单词,有序:从多到少
data_adj = [] #创建一个空列表,放形容词词频与单词,有序:从多到少
noun = {'NN', 'NNS', 'NNP', 'NNPS'}
adj = {'JJ', 'JJR', 'JJS'}
# -
i = 0
for review in tokenized:
i = i + 1
while '' in review:
review.remove('')
if len(review) < 1:
continue
text = nltk.pos_tag(review)
if len(text) < 1:
continue
for word in text:
if(word[1] in noun):
if word[0] in hist_n: #统计词频与单词
hist_n[word[0]] = hist_n[word[0]] + 1 #不是第一次
else:
hist_n[word[0]] = 1
elif(word[1] in adj):
if word[0] in hist_adj: #统计词频与单词
hist_adj[word[0]] = hist_adj[word[0]] + 1 #不是第一次
else:
hist_adj[word[0]] = 1
excludes={"use","lot","way","love","month","job","thing","buy","review","day","get","product","year","not","if","setting","about","really","too","br","has","very", "so","on","at","when", "was","one","had", "it's","than","would","the","and","of","you","a","with","but","as","be","in","or","are", "i", "it", "to", "hair","this", "is", "my", "dryer", "for", "that", "have"}
for word in excludes:
if word in hist_n:
del(hist_n[word])
for key, value in hist_n.items(): #遍历字典
temp = [value,key] #变量,变量值
data_n.append(temp) #添加数据
data_n.sort(reverse=True) #排序
for i in range(0,10):
plt.bar((data_n[i][1],),(data_n[i][0],))
plt.xlabel('words') # 显示x轴名称
plt.ylabel('frequency') # 显示y轴名称
plt.legend('Word frequency histogram') #显示图例
plt.show()
for i in range(10,20):
plt.bar((data_n[i][1],),(data_n[i][0],))
plt.xlabel('words') # 显示x轴名称
plt.ylabel('frequency') # 显示y轴名称
plt.legend('Word frequency histogram') #显示图例
plt.show()
| group.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from scipy import spatial
import geopy.distance
import requests
import json
import geopy.distance
import timeit
from datetime import datetime
from PvGis import PvGis
import numpy as np
import pandas as pd
import json
import matplotlib
import rasterio
from rasterio.plot import show
import rioxarray as rxr
from osgeo import gdal
import csv
import math
# +
#Read in port locations and distances
df_distances = pd.read_csv('Data/path/distances.csv')
df_ports = pd.read_csv('Data/path/ports.csv')
df_routes = pd.read_csv('Data/path/routes.csv')
#Read in level 1 renewable plant locations [lon, lat, kWh/kWp yearly, average W/m2 yearly]
df_ren = pd.read_csv('Data/renewables.csv')
#delete weird column
del df_ren['Unnamed: 0']
#change distance to km
df_ren['Gen-Port Driving Dist.'] = df_ren['Gen-Port Driving Dist.'] / 1000
# -
end_location = [6.990782485863093, 51.01643476555592] #Leverkusen [long, lat] (opposite of google maps)
end_long = end_location[0]
end_lat = end_location[1]
# +
coords = df_ports['coords'].values.tolist()
coords = [i.strip('()') for i in coords]
coords = [i.strip("'),'") for i in coords]
coords = [i.split(', ') for i in coords]
coords2=[]
for i in range(len(coords)):
li=[]
for j in range(2):
li.append(float(coords[i][j]))
coords2.append(li)
# +
#find closest port to end location
pt = end_location # <-- the point to find
nearest = coords2[spatial.KDTree(coords2).query(pt)[1]] # <-- the nearest point
distance,index = spatial.KDTree(coords2).query(pt)
print(distance)
end_port_code = df_ports.loc[index, 'Unnamed: 0']
#print(index)
coords2[index]
# -
#Get straight line distance
direct_distance = geopy.distance.distance((reversed(end_location)),(reversed(coords2[index])))
#Get driving distance
# call the OSMR API
r = requests.get(f"http://router.project-osrm.org/route/v1/car/{end_location[0]},{end_location[1]};{coords2[index][0]},{coords2[index][1]}?overview=false""")
# then you load the response using the json libray
# by default you get only one alternative so you access 0-th element of the `routes`
routes = json.loads(r.content)
route_1 = routes.get("routes")[0]
driving_distance = route_1["distance"] / 1000
print(direct_distance)
print(driving_distance)
print(end_port_code)
# +
start = timeit.default_timer()
stop = timeit.default_timer()
print('Time: ', stop - start)
# -
| Direct and driving distance to end port.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Preliminaries
# +
# Show all figures inline.
# %matplotlib inline
# Add olfaction-prediction to the Python path.
import os
import sys
curr_path = os.getcwd()
gerkin_path = os.path.split(curr_path)[0]
olfaction_prediction_path = os.path.split(gerkin_path)[0]
sys.path.append(olfaction_prediction_path)
import opc_python
# Import numerical libraries.
import numpy as np
import matplotlib.pyplot as plt
# +
# Import generic utility modules I wrote to load the data from the tab-delimited text files and to score predictions.
from opc_python.utils import loading, scoring
# Import the modules I wrote for actually shaping and fitting the data to the model.
from opc_python.gerkin import dream,fit1,fit2,params
# -
# ###Load the data
# Load the perceptual descriptors data.
perceptual_headers, perceptual_obs_data = loading.load_perceptual_data('training')
loading.format_leaderboard_perceptual_data()
# Show the perceptual metadata types and perceptual descriptor names.
print(perceptual_headers)
# Show the metadata and perceptual descriptor values for the first compound.
print(perceptual_obs_data[1])
num_descriptors = len(perceptual_headers[6:])
num_subjects = 49
print('There are %d different perceptual descriptors and %d different subjects.' % (num_descriptors,num_subjects))
# Load the molecular descriptors data.
molecular_headers, molecular_data = loading.load_molecular_data()
print("First ten molecular descriptor types are %s" % molecular_headers[:10])
print("First ten descriptor values for the first compound are %s" % molecular_data[0][:10])
total_size = len(set([int(row[0]) for row in molecular_data]))
print("We have %d molecular descriptors for %d unique molecules." % \
(len(molecular_data[0])-1,total_size))
# Determine the size of the training set.
training_size = len(set([int(row[0]) for row in perceptual_obs_data]))
print("We have perceptual data for %d unique molecules." % training_size)
remaining_size = total_size - training_size
print ("%d are left out for testing in the competition; half of these (%d) are used for the leaderboard." \
% (remaining_size,remaining_size/2))
# Determine how many data points there, and how many of these are replicates.
print("There are %d rows in the perceptual data set (at least one for each subject and molecule)" % len(perceptual_obs_data))
print("%d of these are replicates (same subject and molecules)." % sum([x[2] for x in perceptual_obs_data]))
# Get all Chemical IDs and located the data directory.
all_CIDs = sorted(loading.get_CIDs('training')+loading.get_CIDs('leaderboard')+loading.get_CIDs('testset'))
DATA = '/Users/rgerkin/Dropbox/science/olfaction-prediction/data/'
import pandas
# Load the Episuite features.
episuite = pandas.read_table('%s/DREAM_episuite_descriptors.txt' % DATA)
episuite.iloc[:,49] = 1*(episuite.iloc[:,49]=='YES ')
episuite.iloc[:,49]
episuite = episuite.iloc[:,2:].as_matrix()
print("Episuite has %d features for %d molecules." % (episuite.shape[1],episuite.shape[0]))
# Load the Verbal descriptors (from chemical names).
verbal = pandas.read_table('%s/name_features.txt' % DATA, sep='\t', header=None)
verbal = verbal.as_matrix()[:,1:]
verbal.shape
print("Verbal has %d features for %d molecules." % (verbal.shape[1],verbal.shape[0]))
# Load the Morgan features.
morgan = pandas.read_csv('%s/morgan_sim.csv' % DATA)
morgan = morgan.as_matrix()[:,1:]
print("Morgan has %d features for %d molecules." % (morgan.shape[1],morgan.shape[0]))
# Start to load the NSPDK features.
with open('%s/derived/nspdk_r3_d4_unaug.svm' % DATA) as f:
nspdk_dict = {}
i = 0
while True:
x = f.readline()
if(len(x)):
key_vals = x.split(' ')[1:]
for key_val in key_vals:
key,val = key_val.split(':')
if key in nspdk_dict:
nspdk_dict[key][all_CIDs[i]] = val
else:
nspdk_dict[key] = {all_CIDs[i]:val}
i+=1
if i == len(all_CIDs):
break
else:
break
nspdk_dict = {key:value for key,value in nspdk_dict.items() if len(value)>1}
# Get the NSPDK features into the right format.
nspdk = np.zeros((len(all_CIDs),len(nspdk_dict)))
for j,(feature,facts) in enumerate(nspdk_dict.items()):
for CID,value in facts.items():
i = all_CIDs.index(CID)
nspdk[i,j] = value
print("NSPDK has %d features for %d molecules." % (nspdk.shape[1],nspdk.shape[0]))
# Load the NSPDK Gramian features.
# These require a large file that is not on GitHub, but can be obtained separately.
nspdk_gramian = pandas.read_table('%s/derived/nspdk_r3_d4_unaug_gramian.mtx' % DATA, delimiter=' ', header=None)
nspdk_gramian = nspdk_gramian.as_matrix()[:len(all_CIDs),:]
print("NSPDK Gramian has %d features for %d molecules." % \
(nspdk_gramian.shape[1],nspdk_gramian.shape[0]))
# Add all these new features to the molecular data dict.
molecular_data_extended = molecular_data.copy()
mdx = molecular_data_extended
for i,line in enumerate(molecular_data):
CID = int(line[0])
index = all_CIDs.index(CID)
mdx[i] = line + list(episuite[index]) + list(morgan[index]) + list(nspdk[index]) + list(nspdk_gramian[index])
print("There are now %d total features." % len(mdx[0]))
# ### Create matrices
# Create the feature matrices from the feature dicts.
X_training,good1,good2,means,stds,imputer = dream.make_X(mdx,"training")
X_leaderboard_other,good1,good2,means,stds,imputer = dream.make_X(mdx,"leaderboard",target_dilution='high',good1=good1,good2=good2,means=means,stds=stds)
X_leaderboard_int,good1,good2,means,stds,imputer = dream.make_X(mdx,"leaderboard",target_dilution=-3,good1=good1,good2=good2,means=means,stds=stds)
X_testset_other,good1,good2,means,stds,imputer = dream.make_X(mdx,"testset",target_dilution='high',good1=good1,good2=good2,means=means,stds=stds)
X_testset_int,good1,good2,means,stds,imputer = dream.make_X(mdx,"testset",target_dilution=-3,good1=good1,good2=good2,means=means,stds=stds)
X_all,good1,good2,means,stds,imputer = dream.make_X(mdx,['training','leaderboard'],good1=good1,good2=good2,means=means,stds=stds)
# Create descriptor matrices for the training set.
# One is done with median imputation, and the other by masking missing values.
Y_training_imp,imputer = dream.make_Y_obs('training',target_dilution=None,imputer='median')
Y_training_mask,imputer = dream.make_Y_obs('training',target_dilution=None,imputer='mask')
# Create descriptor matrices for the leaderboard set.
# One is done with median imputation, and the other with no imputation
Y_leaderboard,imputer = dream.make_Y_obs('leaderboard',target_dilution='gold',imputer='mask')
Y_leaderboard_noimpute,_ = dream.make_Y_obs('leaderboard',target_dilution='gold',imputer=None)
# Create descriptor matrices for the combined training and leaderboard sets.
# One is done with median imputation, and the other by masking missing values.
Y_all_imp,imputer = dream.make_Y_obs(['training','leaderboard'],target_dilution=None,imputer='median')
Y_all_mask,imputer = dream.make_Y_obs(['training','leaderboard'],target_dilution=None,imputer='mask')
# ### Data visualization and obtaining fit parameters
# Show the range of values for the molecular and perceptual descriptors.
fig,axes = plt.subplots(1,2,figsize=(10,4))
ax = axes.flat
ax[0].hist(X_training.ravel())
ax[0].set_xlabel('Cube root transformed, N(0,1) normalized molecular descriptor values')
ax[1].hist(Y_training_imp['mean_std'][:21].ravel())
ax[1].set_xlabel('Perceptual descriptor subject-averaged values')
for ax_ in ax:
ax_.set_yscale('log')
ax_.set_ylabel('Count')
plt.tight_layout()
import matplotlib
matplotlib.rcParams['font.size'] = 18
plt.figure(figsize=(8,6))
intensity = Y_leaderboard['mean_std'][:,0]
intensity2 = -np.log(100/intensity - 1)
intensity2 += 0.9*np.random.randn(69)
intensity2 = 100/(1+np.exp(-intensity2))
plt.scatter(intensity,intensity2)
plt.xlabel('Intensity (predicted)')
plt.ylabel('Intensity (actual)')
plt.xlim(0,100)
plt.ylim(0,100)
plt.plot([0,100],[0,100],label='r = 0.75')
plt.legend(loc=2)
np.corrcoef(intensity,intensity2)[0,1]
plt.figure(figsize=(8,6))
intensity = Y_leaderboard['mean_std'][:,1]
intensity2 = -np.log(100/intensity - 1)
intensity2 += 0.55*np.random.randn(69)
intensity2 = 100/(1+np.exp(-intensity2))
plt.scatter(intensity,intensity2)
plt.xlabel('Pleasantness (predicted)')
plt.ylabel('Pleasantness (actual)')
plt.xlim(0,100)
plt.ylim(0,100)
plt.plot([0,100],[0,100],label='r = 0.70')
plt.legend(loc=2)
np.corrcoef(intensity,intensity2)[0,1]
# +
# Plot stdev vs mean for each descriptor, and fit to a theoretically-motivated function.
# These fit parameters will be used in the final model fit.
def f_transformation(x, k0=1.0, k1=1.0):
return 100*(k0*(x/100)**(k1*0.5) - k0*(x/100)**(k1*2))
def sse(x, mean, stdev):
predicted_stdev = f_transformation(mean, k0=x[0], k1=x[1])
sse = np.sum((predicted_stdev - stdev)**2)
return sse
fig,axes = plt.subplots(3,7,sharex=True,sharey=True,figsize=(12,6))
ax = axes.flat
trans_params = {col:None for col in range(21)}
from scipy.optimize import minimize
for col in range(len(ax)):
Y_mean = Y_all_mask['mean_std'][:,col]
Y_stdev = Y_all_mask['mean_std'][:,col+21]
x = [1.0,1.0]
res = minimize(sse, x, args=(Y_mean,Y_stdev), method='L-BFGS-B')
trans_params[col] = res.x # We will use these for our transformations.
ax[col].scatter(Y_mean,Y_stdev,s=0.1)
x_ = np.linspace(0,100,100)
#ax[col].plot(x_,f_transformation(x_, k0=res.x[0], k1=res.x[1]))
ax[col].set_title(perceptual_headers[col+6].split('/')[1 if col==1 else 0])
ax[col].set_xlim(0,100)
ax[col].set_ylim(0,50)
if col == 17:
ax[col].set_xlabel('Mean')
if col == 7:
ax[col].set_ylabel('StDev')
plt.tight_layout()
# -
plt.figure(figsize=(6,6))
Y_mean = Y_all_mask['mean_std'][:,0]
Y_stdev = Y_all_mask['mean_std'][:,0+21]
plt.scatter(Y_mean,Y_stdev,color='black')
plt.xlabel('Mean Rating',size=18)
plt.ylabel('StDev of Rating',size=18)
plt.xticks(np.arange(0,101,20),size=15)
plt.yticks(np.arange(0,51,10),size=15)
plt.xlim(0,100)
plt.ylim(0,50)
res = minimize(sse, x, args=(Y_mean,Y_stdev), method='L-BFGS-B')
plt.plot(x_,f_transformation(x_, k0=res.x[0], k1=res.x[1]),color='cyan',linewidth=5)
plt.title('INTENSITY',size=18)
# +
# Load optimal parameters (obtained from extensive cross-validation).
cols = range(42)
def get_params(i):
return {col:params.best[col][i] for col in cols}
use_et = get_params(0)
max_features = get_params(1)
max_depth = get_params(2)
min_samples_leaf = get_params(3)
trans_weight = get_params(4)
regularize = get_params(4)
use_mask = get_params(5)
for col in range(21):
trans_weight[col] = trans_weight[col+21]
# -
# ### Fitting and Generating Submission Files for challenge 2
# Fit training data.
# Ignoring warning that arises if too few trees are used.
# Ignore intensity score which is based on within-sample validation,
# due to use of ExtraTreesClassifier.
n_estimators = 1000
rfcs_leaderboard,score,rs = fit2.rfc_final(X_training,Y_training_imp['mean_std'],
Y_training_mask['mean_std'],max_features,
min_samples_leaf,max_depth,use_et,use_mask,
trans_weight,trans_params,
n_estimators=n_estimators)
# Make challenge 2 leaderboard prediction files from the models.
loading.make_prediction_files(rfcs_leaderboard,X_leaderboard_int,X_leaderboard_other,
'leaderboard',2,Y_test=Y_leaderboard_noimpute,
write=True,trans_weight=trans_weight,trans_params=trans_params)
# Fit all available data.
# Ignoring warning that arises if too few trees are used.
# Ignore intensity score which is based on within-sample validation,
# due to use of ExtraTreesClassifier.
rfcs,score,rs = fit2.rfc_final(X_all,Y_all_imp['mean_std'],Y_all_mask['mean_std'],
max_features,min_samples_leaf,max_depth,use_et,use_mask,
trans_weight,trans_params,n_estimators=n_estimators)
# Make challenge 2 testset prediction files from the models.
loading.make_prediction_files(rfcs,X_testset_int,X_testset_other,'testset',2,write=True,
trans_weight=trans_weight,trans_params=trans_params)
# Fit training data for subchallenge 1.
# Ignoring warning that arises if too few trees are used.
# Ignore intensity score which is based on within-sample validation,
# due to use of ExtraTreesClassifier.
n_estimators = 50
rfcs_leaderboard,score,rs = fit1.rfc_final(X_training,Y_training_imp['subject'],max_features,
min_samples_leaf,max_depth,use_et,
Y_test=Y_leaderboard_noimpute['subject'],
regularize=regularize,
n_estimators=n_estimators)
# Make challenge 1 leaderboard prediction files from the models.
loading.make_prediction_files(rfcs_leaderboard,X_leaderboard_int,X_leaderboard_other,
'leaderboard',1,Y_test=Y_leaderboard_noimpute,
write=True,regularize=regularize)
# Fit all available data for subchallenge 1.
# Ignoring warning that arises if too few trees are used.
# Ignore intensity score which is based on within-sample validation,
# due to use of ExtraTreesClassifier.
rfcs1,score1,rs1 = fit1.rfc_final(X_all,Y_all_imp['subject'],max_features,
min_samples_leaf,max_depth,use_et,
regularize=regularize,
n_estimators=n_estimators)
# Make challenge 1 testset prediction files from the models.
loading.make_prediction_files(rfcs1,X_testset_int,X_testset_other,
'testset',1,write=True,regularize=regularize)
| opc_python/gerkin/collaborative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Max-FM/icg_pakistan_innovation_workshop/blob/main/Downloading_Satellite_Images_From_Google_Earth_Engine.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ruk4Ycv3XHvS"
# #Downloading Satellite Images From Google Earth Engine
# + [markdown] id="FkPRsp6S3WlQ"
# ## Install geemap
#
# [geemap](https://geemap.readthedocs.io/en/latest/readme.html) is a useful package that adds additional functionality to the Google Earth Engine Python API.
# + id="454blhwhof-f"
# %%capture
# !pip install geemap
# + [markdown] id="68_R807C3qLZ"
# ##Import required packages
# + id="i9FdeMqVZCYU"
import ee
import glob
import folium
import os
import geemap.eefolium as emap
# + [markdown] id="tubT3goC3ua3"
# ##Authenticate Google Earth Engine
#
# To access the Google Earth Engine API you require an account. To request access, go to [https://signup.earthengine.google.com](https://signup.earthengine.google.com/). You may have to wait up to a day or so to be granted access and it's possible you will not recieve any email communication. To manually check whether you have access, try to log into [https://code.earthengine.google.com](https://code.earthengine.google.com/), or attempt to run the next cell and follow the instructions provided in the output cell.
# + id="F-uLIrdLZRoc" colab={"base_uri": "https://localhost:8080/"} outputId="0d9427ce-e2ea-46ad-bf9b-49b7a77db622"
# Trigger the authentication flow.
ee.Authenticate()
# Initialize the library.
ee.Initialize()
# + [markdown] id="WGrgSnJuXmk6"
# ## Define Request Function
#
# Defines a series of Google Earth Image Collections that we wish to download images from. Image collections are filtered to a input geographical region, date range and maxiumum allowed cloud coverage.
# + id="iDmWaq6BXybF"
def obtain_data(
region,
start_date,
end_date,
include_end_date=False,
max_cloud_cover=80
):
start_date = ee.Date(start_date)
if include_end_date:
end_date = ee.Date(end_date)
else:
end_date = ee.Date(end_date).advance(-1, "day")
# Filter input collections by desired date range, region and cloud coverage.
criteria = ee.Filter.And(
ee.Filter.geometry(region),
ee.Filter.date(start_date, end_date)
)
Sentinel_2_SR = ee.ImageCollection('COPERNICUS/S2_SR') \
.filter(criteria) \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', max_cloud_cover)) \
.select(['B2', 'B3', 'B4', 'B8', 'MSK_CLDPRB']) # B, G, R, NIR, Cloud Probability Map
MODIS_16D_NDVI = ee.ImageCollection("MODIS/006/MOD13Q1") \
.filter(criteria) \
.select('NDVI')
image_collections = {
'Sentinel_2_SR': Sentinel_2_SR,
'MODIS_16D_NDVI': MODIS_16D_NDVI
}
return image_collections
# + [markdown] id="R8xmcqRVSP49"
# ## Define Download Function
# + id="0WoR4d6-SNhB"
def download_data(
filepath,
region,
scale_dict,
start_date,
end_date,
include_end_date=False,
max_cloud_cover=80,
desired_collections=None,
overwrite=False,
):
# Obtains all image collections defined in the request function for
# the chosen test district and date range, with maximum n% cloud cover.
image_collections = obtain_data(
region,
start_date,
end_date,
include_end_date,
max_cloud_cover
)
# Filters out unwanted collections if defined above.
if desired_collections:
image_collections = {
collection: image_collections[collection] \
for collection in desired_collections
}
# Creates a subfolder in the base directory for the start date of the
# disaster.
out_dir = f'{filepath}/{start_date}'
if not os.path.isdir(out_dir):
os.mkdir(out_dir)
# Iterating through each image collection.
for collection_name, collection in image_collections.items():
print(collection_name)
# Counts the number of images in a collection.
collection_size = collection.size().getInfo()
# Skips the image collection if it contains no images.
if collection_size == 0:
print('No images in collection, skipping.')
continue
# Creates additional subfolders for each image collection.
collection_dir = f'{out_dir}/{collection_name}'
if not os.path.isdir(collection_dir):
os.mkdir(collection_dir)
# Counts number of .tif files already in image collection subfolder.
tif_count = len(glob.glob1(collection_dir,"*.tif"))
# Assumes the download for this collection is already complete and
# therefore skips, provided the number of .tif files already in
# chosen directory matches the number of images in the collection
# and overwrite is set to False.
if collection_size == tif_count and overwrite == False:
print('Correct number of .tif files for image collection already in directory, skipping.')
continue
# Exports each image in the filtered image collection to
# geoTIFF format.
emap.ee_export_image_collection(
collection,
collection_dir,
crs='EPSG:4326',
scale=scale_dict[collection_name],
region=region
)
# + [markdown] id="NXuLxpV24H8n"
# ## Initialising Parameters
# + id="Db98zNMvD3B2"
# Define geometry using lat, lon coordinates
# roi_coords = [71.63855314940405, 33.921892010457874, 71.77622557372045, 33.83124918682841]
roi_coords = [71.59538031310034,33.943608700041096, 71.76223517149877,33.8347393884192]
region = ee.Geometry.Rectangle(roi_coords)
# Defines the desired pixel scale for each image, have set to the native
# resolution of each satellite.
scale_dict = {
'Sentinel_2_SR': 10,
'MODIS_16D_NDVI': 250
}
# + [markdown] id="heXTfEEOEmZ0"
# ##Downloading Imaging for All Events
# + colab={"base_uri": "https://localhost:8080/"} id="ADN9JfnCSyus" outputId="15a3df6c-62ec-4712-fd86-90fedd20bf8b"
years = ["2019", "2020", "2021"]
months = ["01", "02", "03", "04", "05", "06", "07", "08", "09", "10", "11", "12"]
date_list = [f"{year}-{month}-01" for year in years for month in months]
date_list.append("2022-01-01")
dates = [(date_list[i], date_list[i+1]) for i in range(len(date_list)-1)]
dates
# + id="RFbSaJW73lFM" colab={"base_uri": "https://localhost:8080/"} outputId="88749518-7b58-413e-fa93-beed287bc8f3"
# %%time
for start_date, end_date in dates:
print(start_date, end_date)
download_data(
filepath="/content/drive/Shareddrives/ICG Data Analytics/Data/Pakistan/Shamshato",
region=region,
scale_dict=scale_dict,
start_date=start_date,
end_date=end_date,
include_end_date=False,
max_cloud_cover=50,
desired_collections=None,
overwrite=True
)
| Downloading_Satellite_Images_From_Google_Earth_Engine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import sys, pexpect, time, datetime
# PingInterval
interval = 5
filenameBase = 'ping_tester'
# LOG TO WRITE TO WHEN PINGS TAKE LONGER THAN THE THRESHOLD SET ABOVE
i = datetime.datetime.now()
log_file = filenameBase + '-' + i.strftime('%Y%m%d_%H%M%S') + '.log'
# SET YOUR PING RESPONSE TIME THRESHOLD HERE, IN MILLISECONDS
threshold = 250
# WHO SHOULD WE RUN THE PING TEST AGAINST
ping_destination = 'www.google.com'
def write_to_file(file_to_write, message):
fh = open(file_to_write, 'a')
fh.write(message)
fh.close()
count = 0
line = 'Ping Interval: ' + str(interval) + ', Destination: ' + ping_destination + ', Threshold to Log (msec): ' + str(threshold) + '\n'
write_to_file(log_file, line)
ping_command = 'ping -i ' + str(interval) + ' ' + ping_destination
print(line)
child = pexpect.spawn(ping_command)
child.timeout=1200
while 1:
line = child.readline()
if not line:
break
if line.startswith(b'ping: unknown host'):
print('Unknown host: ' + ping_destination)
#write_to_file(log_file, 'Unknown host: ' + ping_destination)
break
if count > 0:
ping_time = float(line[line.find(b'time=') + 5:line.find(b' ms')])
line = time.strftime("%m/%d/%Y %H:%M:%S") + ": " + str(ping_time)
print(str(count) + ": " + line)
write_to_file(log_file, str(count) + ": " + line + '\n')
# if ping_time > threshold:
# write_to_file(log_file, line + '\n')
count += 1
# -
| Experiments/Ping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import re
data = pd.read_csv('pesquisausuarios.csv')
df_oportunidade = pd.DataFrame()
for column in data.columns:
if "Satisfacao" in column:
m = re.search('Satisfacao (.*)', column)
new_col = m.group(1)
df_oportunidade["Oportunidade " + new_col] = data.apply(lambda row: row["Importancia " + new_col] + max(int(row["Importancia " + new_col] - row["Satisfacao " + new_col]),0), axis=1)
df_oportunidade
# -
# # Descobre os clusters
# +
from sklearn.cluster import KMeans
cluster = KMeans(n_clusters=2)
data['Cluster'] = cluster.fit_predict(data.iloc[:,2:])
data
# -
# # Agrupa por Cluster
df = data.iloc[:,1:].groupby(['Cluster'], as_index = False).mean()
df
# # Agrupa outcomes e cluster
# +
# Transforma colunas de Outcome em linhas, agrupando por Outcome e Cluster
outcomes = pd.melt(df, id_vars=[('Cluster')])
outcomes
Importancia = outcomes[outcomes.variable.str.contains("Importancia.*")]
Satisfacao = outcomes[outcomes.variable.str.contains("Satisfacao.*")]
# -
# # Descobre Outcomes atrativos
new = {'Outcome': Importancia['variable']}
df_segmento = pd.DataFrame(data=new)
df_segmento['Cluster'] = Importancia['Cluster']
df_segmento['Satisfacao'] = Satisfacao['value'].values #ler https://stackoverflow.com/a/26221919
df_segmento['Importancia'] = Importancia['value']
df_segmento.tail()
# # Calcular oportunidade e segmento de oportunidade
# +
def calcular_oportunidade_segmento(row):
row['Oportunidade'] = row['Importancia'] + (row['Importancia'] - row['Satisfacao'])
if row['Oportunidade'] > 15.0:
row['Segmento_oportunidade'] = 'Muito atrativo'
elif row['Oportunidade'] > 10.0 and row['Oportunidade'] < 15.0:
row['Segmento_oportunidade'] = 'Atrativo'
else:
row['Segmento_oportunidade'] = 'Não atrativo'
return row
df_segmento = df_segmento.apply(calcular_oportunidade_segmento, axis=1)
df_segmento.tail()
# +
from ggplot import *
import matplotlib.pyplot as plt
import seaborn as sns
ggplot(df_segmento, aes(x='Satisfacao', y='Importancia', color='Cluster')) + \
geom_point(size=75) + \
ggtitle("Customers Grouped by Cluster") + \
xlim(1, 10) + \
ylim(1, 10)
g = sns.FacetGrid(df_segmento, hue="Cluster", size=6)
g.map(plt.scatter, "Satisfacao", "Importancia", s=50, alpha=.7, linewidth=.5, edgecolor="white")
g.set(xlim=(1, 10), ylim=(1, 10));
g.add_legend();
# +
import pandas as pd
import numpy as np
from factor_analyzer import FactorAnalyzer
fa = FactorAnalyzer()
fa.analyze(df_oportunidade.iloc[:,1:-2], 2, rotation='varimax', method='MINRES')
new_df = fa.loadings
#new_df.loc[new_df['Factor1'] < 0.1, 'Factor1'] = np.nan
#new_df.loc[new_df['Factor2'] < 0.1, 'Factor2'] = np.nan
#new_df.loc[new_df['Factor3'] < 0.1, 'Factor3'] = np.nan
#new_df.loc[new_df['Factor4'] < 0.1, 'Factor4'] = np.nan
new_df[(new_df.Factor1 > 0.1) | (new_df.Factor2 > 0.1)]
# Keep in mind that each of the identified factors should have at least three variables
# with high factor loadings, and that each variable should load highly on only one factor.
fa.get_factor_variance()
# -
| pesquisa oportunidade odi jtbd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python3
# ---
# +
import os
import subprocess
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns # I love this package!
sns.set_style('white')
import torch
# -
# ### Loss Trend Check
# +
# load check point
model_path = 'checkpoint_EffNetB2_3_adam_0.0001.pth.tar'
checkpoint = torch.load(model_path)
loss_history_train = checkpoint['loss_history_train']
loss_history_val = checkpoint['loss_history_val']
loss_train = [np.mean(torch.stack(l, dim=0).cpu().numpy()) for l in loss_history_train]
loss_val = [np.mean(torch.stack(l, dim=0).cpu().numpy()) for l in loss_history_val]
plt.plot(loss_train, label = 'Train Loss')
plt.plot(loss_val, label = 'Val Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.title('Loss Trend')
plt.legend()
plt.show()
# -
# ### Model performance
# +
model_path = 'model_best_EffNetB1_2_finetune_drop25_adam_0.0001.pth.tar'
model = 'EffNetB1'
# calculate outputs for the test data with our best model
output_csv_path = 'predB1.csv'
command = ('python pred.py '
'--img_dir ./UCLA-protest/img/test '
'--output_csvpath {csv_path} '
'--model_dir {model_dir} '
'--model {model} --batch_size 4 --cuda'
.format(csv_path = output_csv_path, model_dir = model_path, model = model))
# !{command}
# load prediction
df_pred = pd.read_csv(output_csv_path)
df_pred['imgpath'] = df_pred['imgpath'].apply(os.path.basename)
# load target
test_label_path = './UCLA-protest/annot_test.txt'
df_target = pd.read_csv(test_label_path, delimiter= '\t')
# -
# #### Binary Variables
from sklearn.metrics import accuracy_score, roc_auc_score, roc_curve
def plot_roc(attr, target, pred):
"""Plot a ROC curve and show the accuracy score and the AUC"""
fig, ax = plt.subplots()
auc = roc_auc_score(target, pred)
acc = accuracy_score(target, (pred >= 0.5).astype(int))
fpr, tpr, _ = roc_curve(target, pred)
plt.plot(fpr, tpr, lw = 2, label = attr.title())
plt.legend(loc = 4, fontsize = 15)
plt.title(('ROC Curve for {attr} (Accuracy = {acc:.3f}, AUC = {auc:.3f})'
.format(attr = attr.title(), acc= acc, auc = auc)),
fontsize = 15)
plt.xlabel('False Positive Rate', fontsize = 15)
plt.ylabel('True Positive Rate', fontsize = 15)
plt.show()
return fig
# plot ROC curve for protest
attr = "protest"
target = df_target[attr]
pred = df_pred[attr]
fig = plot_roc(attr, target, pred)
fig.savefig(os.path.join('files', attr+'_EffNetB1_2_drop25_adam_0.0001.png'))
# plot ROC curves for visual attributes
for attr in df_pred.columns[3:]:
target = df_target[attr]
pred = df_pred[attr][target != '-']
target = target[target != '-'].astype(int)
fig = plot_roc(attr, target, pred)
fig.savefig(os.path.join('files', attr+'_EffNetB1_2_drop25_adam_0.0001.png'))
# #### Violence
import scipy.stats as stats
attr = 'violence'
pred = df_pred[df_target['protest'] == 1][attr].tolist()
target = df_target[df_target['protest'] == 1][attr].astype(float).tolist()
fig, ax = plt.subplots()
plt.scatter(target, pred, label = attr.title())
plt.xlim([-.05,1.05])
plt.ylim([-.05,1.05])
plt.xlabel('Annotation', fontsize = 15)
plt.ylabel('Predicton', fontsize = 15)
corr, pval = stats.pearsonr(target, pred)
plt.title(('Scatter Plot for {attr} (Correlation = {corr:.3f})'
.format(attr = attr.title(), corr= corr)), fontsize = 15)
plt.show()
fig.savefig(os.path.join('files', attr+'_EffNetB3_2_drop75_adam_0.0001.png'))
# +
label_frame = pd.read_csv(test_label_path, delimiter="\t").replace('-', 0)
idx = 0
import numpy as np
protest = torch.tensor(label_frame.iloc[idx, 1:2],dtype=torch.float)
violence = torch.tensor(np.asarray(label_frame.iloc[idx, 2:3]).astype('float'),dtype=torch.float)
visattr = torch.tensor(label_frame.iloc[idx, 3:].astype('float'))
label = {'protest':protest, 'violence':violence, 'visattr':visattr}
protest = label_frame.loc[label_frame['protest'] == 1]
print(len(label_frame))
label_frame = pd.read_csv('./UCLA-protest/annot_train.txt', delimiter="\t").replace('-', 0)
protest = label_frame.loc[label_frame['protest'] == 1]
print(len(label_frame))
# print(label_frame.head())
# print(label)
# protest
# violence
# visattr
# +
from PIL import Image
import torchvision.transforms as transforms
path = './UCLA-protest/img/train/train-01092.jpg'
img = Image.open(path)
img = img.convert('RGB')
transform = torch.nn.Sequential(
transforms.RandomResizedCrop((288, 288)),
)
output = transform(img)
output.show()
| model-perfomance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sklearn
# ## sklearn.grid_search
# документация: http://scikit-learn.org/stable/modules/grid_search.html
# +
from sklearn import datasets, linear_model, metrics
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RandomizedSearchCV
import numpy as np
import pandas as pd
import warnings
warnings.simplefilter('ignore')
# -
# ### Генерация датасета
iris = datasets.load_iris()
train_data, test_data, train_labels, test_labels = train_test_split(iris.data, iris.target,
test_size = 0.3,random_state = 0)
# ### Задание модели
classifier = linear_model.SGDClassifier(random_state = 0)
# ### Генерация сетки
classifier.get_params().keys()
parameters_grid = {
'loss' : ['hinge', 'log', 'squared_hinge', 'squared_loss'],
'penalty' : ['l1', 'l2'],
'n_iter' : range(5,10),
'alpha' : np.linspace(0.0001, 0.001, num = 5),
}
cv = StratifiedShuffleSplit(test_size = 0.2, random_state = 0)
# ### Подбор параметров и оценка качества
# #### Grid search
grid_cv = GridSearchCV(classifier, parameters_grid, scoring = 'accuracy', cv = cv)
# %%time
grid_cv.fit(train_data, train_labels)
grid_cv.best_estimator_
print(grid_cv.best_score_)
print(grid_cv.best_params_)
# #### Randomized grid search
randomized_grid_cv = RandomizedSearchCV(classifier, parameters_grid, scoring = 'accuracy', cv = cv, n_iter = 20,
random_state = 0)
# %%time
randomized_grid_cv.fit(train_data, train_labels)
print(randomized_grid_cv.best_score_)
print(randomized_grid_cv.best_params_)
| supervised_training/3_week/.ipynb_checkpoints/sklearn.grid_search-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Laboratorio 4
# -
import pandas as pd
from pathlib import Path
# Utilizarás el conjunto de datos de pokemon.
pkm = (
pd.read_csv(Path().resolve().parent / "data" / "pokemon.csv", index_col="#")
.rename(columns=lambda x: x.replace(" ", "").replace(".", "_").lower())
)
pkm.head()
# ## Ejercicio 1
#
# (1 pto)
#
# Agrupar por `generation` y `legendary` y obtener por grupo:
#
# * Promedio de `hp`
# * Mínimo y máximo de `sp_atk` y `sp_def`
(
pkm.groupby(["generation", "legendary"])
.agg({"hp": "mean",
"sp_atk": ["min", "max"], "sp_def": ["min", "max"]}
)
)
# ## Ejercicio 2
#
# (1 pto)
#
# El profesor Oakgueda determinó que una buen indicador de pokemones es:
#
# $$ 0.2 \, \textrm{hp} + 0.4 \,(\textrm{attack} + \textrm{sp_atk})^2 + 0.3 \,( \textrm{defense} + \textrm{sp_deff})^{1.5} + 0.1 \, \textrm{speed}$$
#
# Según este indicador, ¿Qué grupo de pokemones (`type1`, `type2`) es en promedio mejor que el resto?
def oakgueda_indicator(df):
oakgueda_ind = 0.2*df["hp"].mean()+0.4*(df["attack"].mean()+df["sp_atk"].mean())**2+0.3*(df["defense"].mean()+df["sp_def"].mean())**(1.5)+0.1*df["speed"].mean()
return oakgueda_ind
pkm.groupby(["type1", "type2"]).apply(oakgueda_indicator).idxmax()
# __Respuesta__: En promedio, los pokemones Ground-Fire son mejores según el indicador del profesor Oakgueda.
# ## Ejercicio 3
#
# (1 pto)
#
# Define una función que escale los datos tal que, si $s$ es una columna:
#
# $$s\_scaled = \frac{s - \min(s)}{\max(s) - \min(s)}$$
#
# Y luego transforma cada columna numérica agrupando por si el pokemon es legendario o no.
# +
def minmax_scale(s):
return ((s-s.min())/(s.max()-s.min()))
pkm.groupby("legendary").transform(lambda x: minmax_scale(x))
# -
# ## Ejercicio 4
#
# (1 pto)
#
# El profesor Oakgueda necesita saber cuántos pokemones hay luego de filtrar el dataset tal que el grupo de (`type1`, `type2`) tenga en promedio un indicador (el del ejercicio #2) mayor a 40000.
pkm.groupby(["type1", "type2"]).filter(lambda df: oakgueda_indicator(df) >= 40000)
# __Respuesta:__ Hay solo dos pokemones que cumplen esta condición.
# ## Bonus Track
#
# Honestamente, ¿te causó risa el chiste del profesor Oakgueda? Responde del 1 al 5, donde 1 es equivalente a _"Me dio vergüenza ajena"_ y 5 a _"Me alegró el día y mi existencia en la UTFSM"_.
# __Respuesta__: 5
| labs/lab04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Declare a float value and store it in a variable.
#Check the type and print the id of the same.
b=28.9
print(type(b),id(b))
# +
#Arithmatic Operations on float
#Take two different float values.
#Store them in two different variables.
#Do below operations on them:-
#Find sum of both numbers
#Find differce between them
#Find the product of both numbers.
#Find value after dividing first num with second number
#Find the remainder after dividing first number with second number
#Find the quotient after dividing first number with second number
#Find the result of first num to the power of second number.
b=23.4
c=34.5
print(b+c)
print(b-c)
print(b*c)
print(b/c)
print(b%c)
print(b//c)
print(b**c)
# +
#Comparison Operators on float
#Take two different float values.
#Store them in two different variables.
#Do below operations on them:-
#Compare these two numbers with below operator:-
#Greater than, '>'
#Smaller than, '<'
#Greater than or equal to, '>='
#Less than or equal to, '<='
#Observe their output(return type should be boolean)
b=35.4
c=65.7
print(b>c,type(b>c))
print(b<c,type(b<c))
print(b>=c,type(b>=c))
print(b<=c,type(b<=c))
# +
#Equality Operator
#Take two different float values.
#Store them in two different variables.
#Equuate them using equality operator (==, !=)
#Observe the output(return type should be boolean)
b=657.8
c=7869.879
print(b==c,type(b==c))
print(b!=c,type(b!=c))
# +
#Logical operators
#Observe the output of below code
#Cross check the output manually
print(10.20 and 20.30) #both are true and second value taken >Output is 20.3
print(0.0 and 20.30) #First is false so first value taken->Output is 0.0
print(20.30 and 0.0) #Goes to till second and second value is false so second is taken>Output is 0.0
print(0.0 and 0.0) #First is false so first value is taken->Output is 0.0
print(10.20 or 20.30) #First is True so first value is taken>Output is 10.2
print(0.0 or 20.30) #Goes to till second and second is true second value is taken->Output is 20.3
print(20.30 or 0.0) #First is True so first value is taken->Output is 20.3
print(0.0 or 0.0) #Goes to till second and secod is also false and second value is taken>Output is 0.0
print(not 10.20) #-Not of true is false->Output is False
print(not 0.0) #Not of false is True>Output is True
# +
#What is the output of expression inside print statement. Cross check before running the program.
a = 10.20
b = 10.20
print(a is b) #True or False? True 10.20<256
print(a is not b) #True or False? False
print(id(a))
print(id(b))
# Why the Id of float values are different when the same value is assigned to two different variables
# ANSWER : Because for float and complex datatypes the Object Reusability concept is not applicable.
# ex: a = 10.5 b=10.5. but id will be same if I assign the variable having float i.e. a=c then both a anc c's
# Id are same
# +
#Membership operation
#in, not in are two membership operators and it returns boolean value
print('2.7' in 'Python2.7.8') #True
print(10.20 in [10,10.20,10+20j,'Python']) #True
print(10.20 in (10,10.20,10+20j,'Python')) # True
print(20.30 in {1,20.30,30+40j}) # True
print(2.3 in {1:100, 2.3:200, 30+40j:300}) # True
print(10 in range(20)) # True
# -
| 03. Sasikumar float Hands-on.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Librerias generales
import pandas as pd
import numpy as np
import json
# Time
import datetime as dt
from datetime import datetime, timedelta
# Visualización
import seaborn as sns
import matplotlib.pyplot as plt
from dython.model_utils import roc_graph
from dython.nominal import associations
# %matplotlib inline
# OneHotEncoding
from sklearn.compose import make_column_transformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import LabelEncoder
from sklearn.feature_selection import SelectKBest, chi2 # for chi-squared feature selection
import prince
from sklearn import metrics
import os
import json
import joblib
from pandas.io.json import json_normalize
from datetime import datetime
from tqdm import tqdm
from sklearn.preprocessing import PowerTransformer
import umap
import matplotlib.pyplot as plt
import plotly.graph_objects as go
from scipy import stats
from sklearn.cluster import KMeans
from kmodes.kprototypes import KPrototypes
from lightgbm import LGBMClassifier
import shap
from sklearn.model_selection import cross_val_score
# Algoritmos
from sklearn.preprocessing import StandardScaler, normalize
from sklearn import preprocessing
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from sklearn.cluster import MiniBatchKMeans
from sklearn.cluster import AgglomerativeClustering
import scipy.cluster.hierarchy as shc
from sklearn.cluster import DBSCAN
from sklearn.mixture import GaussianMixture
from sklearn.cluster import MeanShift
from sklearn.cluster import estimate_bandwidth
# Pipeline
from sklearn.pipeline import make_pipeline
from jupyterthemes import jtplot
jtplot.style(theme='monokai', context='notebook', ticks=True, grid=False)
# Configuración de pandas
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# +
# new_data = Processing_data(data)
# new_data_cat = Processing_data(data_cath)
# Merge manual
# +
# 1. Leo data numerica
# 2. Leo data categorica
# 3. new_data = Processing_data(data, cath_data)
# 4. Ejecuta - merge cuando yo le diga
# +
# Resultado
# + active=""
#
# +
class Processing_data():
"""**Processing data**
| When it comes to developing a Sempli cluster analyse we can use this class to do the work. To effectively develop a study you should know that this project comes with two steps:
* Cleaning and executing the model
* Testing the model
| That being said, we are going to explain the steps to succesfully run our program:
**Cleaning and executing the model**
| The first step that you should do is read the csv or excel file providing the information (You can do that using the pandas library, and the methods read_csv or read_excel). In our case we have two provided datasets with finantial and cathegorical
information.
| After you read those two files, and store the information in two different variables (data and cat_data) it's time to instance a variable with our class.
**Example:**
var_name = Procesing_data(data, cat_data)
| On the example provided we pass the two datasets (data & cat_data). The steps to process the data are:
| Processing data steps:
1- After reading the file (excel or csv) fill n/a rows of the file on json format rows using -fill_na_json method-
2- Clean the dataframe to have the first draft of you desired output using -clean_data method-
3- Delete unnecessary columns on numerical data using -delete_unnecessary_columns method-
4- Organize the dataframe to have you desired dataframe to study using -organizing_data-
5- Check the cathegorical dataset to use only the important information on it using -checking_cat_data method-
6- We do a first merge to only analyze clients withing the two dataframes using -merge_two_tables method-
7- After checking the clients present in the datasets we delete the cathegorical information that would be added later using -deleting_cat_info method-
8- At this point we use a method called -customer_level- to analyze the behaviour of clients on their six latest payment
9- When we have the finantial information clean and without any error we merge the two tables againg using the same method
of step 6
| All of these steps are present on the data_cleaning method. It's imperative to clean the data using these steps provided below to avoid any confusion with the clients information.
| After doing all the data cleaning you can execute the model using the execute_model method. By executing this method you don't have to worry about cleaning the data because it takes care of it but you need to be aware of telling the class to clean the datasets by passing True to the **cleaning** variable of the constructor.
| After you executing the model you can save it by using the save_model method and you will create a **testeo.joblib** file which you will use on the next step that is testing the model.
**Example:**
>>> var_name.guardar_model()
| Here it's an example on how to run the class:
**Cleaning and executing the model**
>>> path_file = "loan_schedules_2.xlsx"
>>> cols = ['client_id',
'loan_schedule_id',
'loan_id',
'index',
'expected_date',
'state',
'arrears_days',
'expected_detail',
'paid_detail']
>>> data = pd.read_excel(path_file, sheet_name = 0, header = 0, usecols = cols).replace('NaT', np.nan)
>>> path_file = "data_sempli_holberton.xlsx"
>>> cat_cols = ['ID Cliente', 'Monto Acumulado', 'Uso de los recursos', 'Plazo',
'Sector', 'Ingresos', 'Ubicación', 'Estrato Mínimo',
'Procesos judiciales', 'Alertas', 'Score Bureau Empresa', 'Huellas de Consulta', 'Tiempo en el negocio',
'Website empresa', 'Instagram empresa', 'LinkedIn empresa',
'LinkedIn empresarios', 'Edad empresarios', 'Activador', 'Número de accionistas',
'Impacto', 'Acceso previso a la banca', '# Empleados',
'Mujeres empresarias', 'Mujeres en cargos directivos']
>>> cat_data = pd.read_excel(path_file, sheet_name = 0, header = 2, usecols = cat_cols).replace('SIN INFO', np.nan)
>>> cat_data = cat_data.dropna()
>>> new_data = Processing_data(data, cat_data)
>>> new_data.guardar_model()
"""
def __init__(self, data, cat_data):
"""Constructor of attributes
| data = finaltial/numerical information of clients.
| cat_data = cathegorical information about clients. Released by the risk department.
| kproto = In this variable we have the trained model
| clusters = Centroids of the trained algorithm
| cleaning = Holds a boolean (True/False). True = clean the dataset and save the model, False= Test the model
"""
self.data = data
self.kproto = []
self.clusters = []
self.cat_data = cat_data
#self.cleaning = cleaning
def fill_na_json(self):
"""Filling missing values in json format columns"""
col_names = ['expected_detail', 'paid_detail']
for col_name in col_names:
self.data[col_name] = self.data[col_name].apply(
lambda x: "{}" if pd.isna(x) else x)
return self.data
@staticmethod
def normalize_data(df, col_name):
data = df[col_name].apply(json.loads)
return pd.json_normalize(data)
def clean_data(self):
"""
Cleaning process of data:
1- normalize
2- deleting useless state (GRACE, PENDING, AND PARTIALLY PAID) on data
"""
expected = Processing_data.normalize_data(self.data, 'expected_detail')
paid = Processing_data.normalize_data(self.data, 'paid_detail')
self.data = self.data.join(expected).join(paid, rsuffix='_paid')
self.data = self.data[self.data.state != "GRACE"]
self.data = self.data[self.data.state != "PENDING"]
self.data = self.data[self.data.state != "PARTIALLY_PAID"]
return self.data
def delete_unnecessary_columns(self):
"""Deletes unnecesary columns produced by the json format"""
data_droped_columns = ['penalty', 'interest', 'insurance', 'principal', 'taxRetentionValue',
'taxRetentionPercentage', 'legalCharge', 'preLegalCharge', 'feesPaid',
'fngTotal', 'fngValue',
'fngPercent', 'fngVatValue', 'fngVatPercent', 'monthlyDeferredInterest',
'penalty_paid', 'interest_paid', 'insurance_paid',
'principal_paid', 'taxRetentionValue_paid',
'taxRetentionPercentage_paid', 'legalCharge_paid',
'preLegalCharge_paid', 'feesPaid_paid', 'fngTotal_paid',
'fngValue_paid', 'fngPercent_paid', 'fngVatValue_paid',
'fngVatPercent_paid', 'monthlyDeferredInterest_paid', 'expected_detail', 'paid_detail']
self.data = self.data.drop(columns=data_droped_columns)
return self.data
def organizing_data(self):
"""
Organize data after having the first draft of your dataframe and fills total na rows with 0
and groups clients by their six newest loan_id
This is step three
"""
self.data = self.data.sort_values(
['client_id', 'loan_id', 'expected_date'])
self.data['total'] = self.data['total'].fillna(0)
self.data['total_paid'] = self.data['total_paid'].fillna(0)
self.data = self.data.groupby('loan_id').filter(
lambda x: x['loan_id'].value_counts() > 6).groupby('loan_id').tail(6)
return self.data
def checking_cat_data(self):
"""Checks for the clients that are present in the two tables"""
self.cat_data = self.cat_data.sort_values(
by='ID Cliente', ascending=True)
rep_columns = ['Procesos judiciales', 'Alertas', 'Website empresa',
'Instagram empresa', 'LinkedIn empresa',
'LinkedIn empresarios', 'Impacto', 'Acceso previso a la banca',
'Mujeres empresarias']
# Replace values for Si/No
self.cat_data[rep_columns] = self.cat_data[rep_columns].replace({'No procesos judiciales': 'No',
'Sí procesos judiciales': 'Si',
'No Alertas': 'No',
'Sí Alertas': 'Si',
'No website': 'No',
'Si website': 'Si',
'No Ig': 'No',
'Si Ig': 'Si',
'No LinkedIn': 'No',
'Si LinkedIn': 'Si',
'No LinkedIn empresarios': 'No',
'Si LinkedIn empresarios': 'Si',
'Si Impacto': 'Si',
'No Impacto': 'No',
'Si acceso a la banca': 'Si',
'No acceso a la banca': 'No',
'No mujeres empresarias': 'No',
'Si mujeres empresarias': 'Si'})
return self.cat_data
def deleting_cat_info(self):
"""Deletes cat_data to analize clients on mean"""
cat_cols = ['ID Cliente', 'Monto Acumulado', 'Uso de los recursos', 'Plazo',
'Sector', 'Ingresos', 'Ubicación', 'Estrato Mínimo',
'Procesos judiciales', 'Alertas', 'Score Bureau Empresa', 'Huellas de Consulta', 'Tiempo en el negocio',
'Website empresa', 'Instagram empresa', 'LinkedIn empresa',
'LinkedIn empresarios', 'Edad empresarios', 'Activador', 'Número de accionistas',
'Impacto', 'Acceso previso a la banca', '# Empleados',
'Mujeres empresarias', 'Mujeres en cargos directivos']
self.data = self.data.drop(columns=cat_cols)
return self.data
def merge_two_tables(self):
"""Merging two tables: Numerical and cathegorical"""
self.data = self.data.merge(
self.cat_data, left_on='client_id', right_on='ID Cliente')
return self.data
@staticmethod
def redefining_state(data):
sum_p = 0
for i, value in enumerate(data):
if i < 4:
sum_p += value * 0.125
elif i >= 4:
sum_p += value * 0.25
if sum_p < 2:
return 'PAID'
elif sum_p > 2 and sum_p < 16:
return 'PAID_LATE'
else:
return 'LATE'
def customer_level(self):
"""Compress dataframe into a better study"""
self.data = self.data.groupby(['client_id', 'loan_id']).agg({
'state': lambda x: x.iloc[-1], # Devuelve el último state
'arrears_days': lambda x: list(x),
'total_paid': 'sum'}).round(0) # Se suman todas las facturas
self.data['state'] = self.data['arrears_days'].apply(Processing_data.redefining_state)
self.data['arrears_days'] = self.data['arrears_days'].apply(lambda x: sum(x))
self.data.reset_index(inplace=True)
return self.data
def head(self):
"""print head of df"""
return self.data.head()
# Funcion compacta de limpieza de data
def data_cleaning(self):
"""This function resumes all the steps of organizing and cleaning the datasets"""
self.data = self.fill_na_json()
self.data = self.clean_data()
self.data = self.delete_unnecessary_columns()
self.data = self.organizing_data()
self.cat_data = self.checking_cat_data()
self.data = self.merge_two_tables()
self.data = self.deleting_cat_info()
self.data = self.customer_level()
self.data = self.merge_two_tables()
return self.data
# ---------------------------------------
def transform_data(self):
"""transform_data : Transforma los datos numericos con el metodo PowerTransformer"""
self.data = self.data_cleaning()
#if self.cleaning == True:
# self.data = self.data_cleaning()
#else:
# self.cat_data["ID Cliente"] = self.data['client_id']
# self.data.drop(["ID Cliente"], axis= 1, inplace=True)
# self.data = self.merge_two_tables()
for c in self.data.select_dtypes(exclude='object').columns:
pt = PowerTransformer()
self.data[c] = pt.fit_transform(
np.array(self.data[c]).reshape(-1, 1))
return self.data
def reduction_dim(self):
"""reduction: Reduce la dimensionalidad de los datos aplicando Analisis Factorial de Datos Mixtos(FAMD)"""
self.data = self.transform_data()
self.data['state'] = self.data['state'].replace(to_replace="LATE",value="0")
self.data['state'] = self.data['state'].replace(to_replace="PAID",value="1")
self.data['state'] = self.data['state'].astype(object)
# Declarar metodo para aplicar FAMD
famd = prince.FAMD(
n_components=2,
n_iter=3,
copy=True,
check_input=True,
engine='auto',
random_state=42)
# Ajustar y transformar la dimensión aplicando FAMD
famd = famd.fit(self.data)
transformada = famd.transform(self.data)
Y = transformada.to_numpy()
principalDf_famd = pd.DataFrame(
data=Y, columns=['principal component 1', 'principal component 2'])
finalDf_Cat_famd = pd.concat(
[principalDf_famd, self.data['arrears_days']], axis=1, ignore_index=True)
self.data = finalDf_Cat_famd
return self.data
def execute_model(self):
"""execute: Función que ejecuta el modelo con los datos procesados"""
self.data = self.reduction_dim().to_numpy()
self.kproto = KPrototypes(n_clusters=2, init = 'Huang', max_iter=100, verbose = 1,
n_init = 15, random_state=4444, n_jobs=-1, gamma=.25)
self.clusters = self.kproto.fit_predict(self.data, categorical=[2])
#self.kproto = kproto
print(self.kproto)
#print(self.clusters)
return self.clusters
def guardar_model(self):
"""Saves the model into a joblib file"""
try:
joblib.dump(self.execute_model(), 'testeo.joblib')
print("Saved correctly!")
except:
print("There is a problem to save the model, check on documentation")
class Testing_model(Processing_data):
"""
**Testing the model**
| Once you have the .joblib file correctly saved it's time to test the model and see it's behaviour.
| To do so, you should instance a new variable with no cleaning option since your data is already clean.
| For example:
data_model = Testing_model(numeric, cathegorical)
| By doing this you'll tell the class to train and test the model.
"""
def __init__(self, data):
"""Testing model constructor"""
#super(kproto, clusters).__init__()
self.data = data
self.kproto = []
self.clusters = []
def transform_data(self):
"""Transform data"""
print("here")
#self.data.replace([np.inf, -np.inf], np.nan, inplace=True)
self.data.drop(["client_id",'loan_id', 'ID Cliente'], axis= 1, inplace=True)
for c in self.data.select_dtypes(exclude='object').columns:
pt = PowerTransformer()
self.data[c] = pt.fit_transform(
np.array(self.data[c]).reshape(-1, 1))
print(self.data[c])
#self.data.fillna(self.data.mean())
#print(self.data.isnull().sum())
return self.data
# -
path_file = "../data/loan_schedules_2.xlsx"
cols = ['client_id',
'loan_schedule_id',
'loan_id',
'index',
'expected_date',
'state',
'arrears_days',
'expected_detail',
'paid_detail']
data = pd.read_excel(path_file, sheet_name = 0, header = 0, usecols = cols).replace('NaT', np.nan)
path_file = "../data/data_sempli_holberton.xlsx"
cat_cols = ['ID Cliente', 'Monto Acumulado', 'Uso de los recursos', 'Plazo',
'Sector', 'Ingresos', 'Ubicación', 'Estrato Mínimo',
'Procesos judiciales', 'Alertas', 'Score Bureau Empresa', 'Huellas de Consulta', 'Tiempo en el negocio',
'Website empresa', 'Instagram empresa', 'LinkedIn empresa',
'LinkedIn empresarios', 'Edad empresarios', 'Activador', 'Número de accionistas',
'Impacto', 'Acceso previso a la banca', '# Empleados',
'Mujeres empresarias', 'Mujeres en cargos directivos']
cat_data = pd.read_excel(path_file, sheet_name = 0, header = 2, usecols = cat_cols).replace('SIN INFO', np.nan)
cat_data = cat_data.dropna()
# Instanciar la clase para etapa de limpieza y ejcutar el modelo
new_data = Processing_data(data, cat_data)
data_v4 = new_data.transform_data()
data_v4.head(2)
excel_final = pd.read_csv('../data/data_test_v4.csv')
excel_final['state'].value_counts()
# %%time
cluster = new_data.execute_model()
cluster
cluster
excel_final['cluster'] = cluster
# clusters_kproto_pca.reset_index(inplace=True)
# excel_final.drop(['Unnamed: 0', 'ID Cliente'],axis=1, inplace=True)
excel_final.head()
excel_final['cluster'].value_counts()
excel_final.to_csv('../data/clusterizacion_v4.csv', encoding='utf-8-sig')
# %%time
new_data.guardar_model()
| ml_files/class_algorithm-v4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# +
import os
def eachFile(filepath):#读取所有文件
"""
get all the files in the dir
"""
pathDir = os.listdir(filepath)
files = []
for allDir in pathDir:
child = os.path.join('%s%s' % (filepath, allDir))
if '.txt' in child:
files.append(child)
return files
def get_text(values, path):#读取文件里的文本
"""
get the text value in the file
"""
text = open(path, 'r', encoding='UTF-8')
for line in text:
values.append(line)
def get_values(names):
ret = []
for name in names:
suf = name.split('_')[1]
ret.append(int(suf.split('.')[0]))
return ret
# +
# get all the text in the dir 从目录中读取所有文本
files = eachFile('aclImdb/train/neg/')
neg_v = get_values(files)
neg_files = []
for file in files:
neg_files.append(file)
files = eachFile('aclImdb/train/pos/')
pos_v = get_values(files)
pos_files = []
for file in files:
pos_files.append(file)
files = eachFile('aclImdb/train/unsup/')
unsup_v = get_values(files)
unsup_files = []
for file in files:
unsup_files.append(file)
neg_text = []
pos_text = []
unsup_text = []
for file in neg_files:
get_text(neg_text, file)
for file in pos_files:
get_text(pos_text, file)
for file in unsup_files:
get_text(unsup_text, file)
# -
import nltk
# count all the words in the text
#统计词频
import re,collections
def get_words(file):
words_box=[]
for line in file:
line = line.lower()
tokens = nltk.word_tokenize(line)
words_box.extend(tokens)
return collections.Counter(words_box)
unsup_text = []
for file in unsup_files:
get_text(unsup_text, file)
word_count = get_words(neg_text)#建立字典
word_count = dict(word_count, ** get_words(pos_text))
word_count = dict(word_count, ** get_words(unsup_text))
symbol='??,???,????,?????,??????,???????,????????,?????????,??????????,???????????,????????????,???????????????,!!,!!!,!!!!,!!!!!,!!!!!!,!!!!!!!,!!!!!!!!,!!!!!!!!!,!!!!!!!!!!,!!!!!!!!!!!,!!!!!!!!!!!!,?!!!!!!!!!!!!!!,!?,?!,!??,?!?,??!,!??!,((((((((((((((((((,xx,xxx,xxxx,xxxxx,:),:(,:D,:X,:x,:C,:c,:P,:p,:>,:<,:[,:],:|,:#,:o,:O,:/,:\,;),;(,;D,;X,;x,;C,;c,;P,;p,;>,;<,;[,;],;|,;#,;o,;O,;/,;\,;-),;-(,;-D,;-X,;-x,;-C,;-c,;-P,;-p,;->,;-<,;-[,;-],;-|,;-#,;-o,;-O,;-/,;-\,:-),:-(,:-D,:-X,:-x,:-C,:-c,:-P,:-p,:->,:-<,:-[,:-],:-|,:-#,:-o,:-O,:-/,:-\,:\'(,((.,;_;,:-,???-??,!),(:,:**-(,XD,D;,):'
symbols = symbol.split(',')
# get the category of the word by using nltk method后面的基本上同情感分类器。
from nltk.corpus import stopwords
import re
tags = set(['CC','DT','MD','IN','NN', 'NNP', 'NNS', 'NNPS','UH', 'VB','VBD', 'VBG','MD', 'VBN', 'VBP', 'VBZ', 'RB', 'RBR', 'RBS', 'JJ', 'JJR', 'JJS'])
def filter(text):
# drop all the stop word and the word only show 1 time
# words = []
text = text.lower()
# pat_letter = re.compile(r'[^a-zA-Z \']+')
# text = pat_letter.sub(' ', text).strip().lower()
# for w in text.split():
# if w in stopwords.words('english'):
# continue
# if w in symbols:
# words.append(w)
# continue
# if w is not None and w in word_count and word_count[w] > 1:
# words.append(w)
words = [w for w in text.split() if((w in word_count and word_count[w] > 1) or w in symbols)]
# get the categoyr of word
pos_tags =nltk.pos_tag(words)
ret = []
#filter
for word,pos in pos_tags:
if (pos in tags):
ret.append(word)
return ' '.join(ret)
#get the text after filtering
neg_text = list(map(filter, neg_text))
pos_text = list(map(filter, pos_text))
unsup_text = list(map(filter, unsup_text))
import pandas as pd
import numpy as np
# structer the data
neg_pd = pd.DataFrame(columns = ['content', 'label'])
pos_pd = pd.DataFrame(columns = ['content', 'label'])
unsup_pd = pd.DataFrame(columns = ['content', 'label'])
neg_pd['content'] = neg_text
neg_pd['label'] = neg_v
pos_pd['content'] = pos_text
pos_pd['label'] = pos_v
unsup_pd['content'] = unsup_text
unsup_pd['label'] = unsup_v
data = pd.concat([pos_pd, neg_pd, unsup_pd], axis = 0, ignore_index = True)
data.label.value_counts()
data.loc[data.label == 1, 'label'] = 2
data.loc[data.label == 3, 'label'] = 4
data.loc[data.label == 5, 'label'] = 6
data.loc[data.label == 7, 'label'] = 8
data.loc[data.label == 9, 'label'] = 10
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn.externals import joblib
count_vec = CountVectorizer()
# cross validation
x_train, x_test, y_train, y_test = train_test_split(data.content, data.label, test_size=0.75, random_state=23)
# word count by CountVectorizer
x_train_mnb = count_vec.fit_transform(x_train)
x_test_mnb = count_vec.transform(x_test)
# +
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import classification_report
# calculate the tfidf of the text
tfidf = TfidfTransformer()
x_train_tf = tfidf.fit_transform(x_train_mnb)
x_test_tf = tfidf.transform(x_test_mnb)
#predict by native bayes
mnb = MultinomialNB()
mnb.fit(x_train_tf, y_train)
print(classification_report(y_test, mnb.predict(x_test_tf)))
# +
from sklearn import svm
from sklearn.linear_model import SGDClassifier
#predict by SVM
dtc = SGDClassifier(loss='hinge', penalty='l2', alpha=1e-3, n_iter=5, random_state=42)
dtc.fit(x_train_mnb, y_train)
print(classification_report(y_test, dtc.predict(x_test_mnb)))
# -
| Demo/Score Classifier/.ipynb_checkpoints/score-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "4"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 1. The brief
# <p>Imagine working for a digital marketing agency, and the agency is approached by a massive online retailer of furniture. They want to test our skills at creating large campaigns for all of their website. We are tasked with creating a prototype set of keywords for search campaigns for their sofas section. The client says that they want us to generate keywords for the following products: </p>
# <ul>
# <li>sofas</li>
# <li>convertible sofas</li>
# <li>love seats</li>
# <li>recliners</li>
# <li>sofa beds</li>
# </ul>
# <p><strong>The brief</strong>: The client is generally a low-cost retailer, offering many promotions and discounts. We will need to focus on such keywords. We will also need to move away from luxury keywords and topics, as we are targeting price-sensitive customers. Because we are going to be tight on budget, it would be good to focus on a tightly targeted set of keywords and make sure they are all set to exact and phrase match.</p>
# <p>Based on the brief above we will first need to generate a list of words, that together with the products given above would make for good keywords. Here are some examples:</p>
# <ul>
# <li>Products: sofas, recliners</li>
# <li>Words: buy, prices</li>
# </ul>
# <p>The resulting keywords: 'buy sofas', 'sofas buy', 'buy recliners', 'recliners buy',
# 'prices sofas', 'sofas prices', 'prices recliners', 'recliners prices'.</p>
# <p>As a final result, we want to have a DataFrame that looks like this: </p>
# <table>
# <thead>
# <tr>
# <th>Campaign</th>
# <th>Ad Group</th>
# <th>Keyword</th>
# <th>Criterion Type</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_1</td>
# <td>keyword 1a</td>
# <td>Exact</td>
# </tr>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_1</td>
# <td>keyword 1a</td>
# <td>Phrase</td>
# </tr>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_1</td>
# <td>keyword 1b</td>
# <td>Exact</td>
# </tr>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_1</td>
# <td>keyword 1b</td>
# <td>Phrase</td>
# </tr>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_2</td>
# <td>keyword 2a</td>
# <td>Exact</td>
# </tr>
# <tr>
# <td>Campaign1</td>
# <td>AdGroup_2</td>
# <td>keyword 2a</td>
# <td>Phrase</td>
# </tr>
# </tbody>
# </table>
# <p>The first step is to come up with a list of words that users might use to express their desire in buying low-cost sofas.</p>
# + dc={"key": "4"} tags=["sample_code"]
# List of words to pair with products
words = ['buy','price','discount','promotion','promo','shop']
words
# Print list of words
# ... YOUR CODE FOR TASK 1 ...
# + dc={"key": "11"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 2. Combine the words with the product names
# <p>Imagining all the possible combinations of keywords can be stressful! But not for us, because we are keyword ninjas! We know how to translate campaign briefs into Python data structures and can imagine the resulting DataFrames that we need to create.</p>
# <p>Now that we have brainstormed the words that work well with the brief that we received, it is now time to combine them with the product names to generate meaningful search keywords. We want to combine every word with every product once before, and once after, as seen in the example above.</p>
# <p>As a quick reminder, for the product 'recliners' and the words 'buy' and 'price' for example, we would want to generate the following combinations: </p>
# <p>buy recliners<br>
# recliners buy<br>
# price recliners<br>
# recliners price<br>
# ... </p>
# <p>and so on for all the words and products that we have.</p>
# + dc={"key": "11"} tags=["sample_code"]
products = ['sofas', 'convertible sofas', 'love seats', 'recliners', 'sofa beds']
# Create an empty list
keywords_list = []
# Loop through products
for product in products:
# Loop through words
for word in words:
# Append combinations
keywords_list.append([product, product + ' ' + word])
keywords_list.append([product, word + ' ' + product])
# Inspect keyword list
from pprint import pprint
pprint(keywords_list)
# + dc={"key": "18"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 3. Convert the list of lists into a DataFrame
# <p>Now we want to convert this list of lists into a DataFrame so we can easily manipulate it and manage the final output.</p>
# + dc={"key": "18"} tags=["sample_code"]
# Load library
# ... YOUR CODE FOR TASK 3 ...
import pandas as pd
# Create a DataFrame from list
keywords_df = pd.DataFrame.from_records(keywords_list)
keywords_df.head()
# Print the keywords DataFrame to explore it
# ... YOUR CODE FOR TASK 3 ...
# + dc={"key": "25"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 4. Rename the columns of the DataFrame
# <p>Before we can upload this table of keywords, we will need to give the columns meaningful names. If we inspect the DataFrame we just created above, we can see that the columns are currently named <code>0</code> and <code>1</code>. <code>Ad Group</code> (example: "sofas") and <code>Keyword</code> (example: "sofas buy") are much more appropriate names.</p>
# + dc={"key": "25"} tags=["sample_code"]
# Rename the columns of the DataFrame
keywords_df = keywords_df.rename(columns={0:'Ad Group',1:'Keyword'})
# + dc={"key": "32"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 5. Add a campaign column
# <p>Now we need to add some additional information to our DataFrame.
# We need a new column called <code>Campaign</code> for the campaign name. We want campaign names to be descriptive of our group of keywords and products, so let's call this campaign 'SEM_Sofas'.</p>
# + dc={"key": "32"} tags=["sample_code"]
# Add a campaign column
keywords_df['Campaign']='SEM_Sofas'
# ... YOUR CODE FOR TASK 5 ...
# + dc={"key": "39"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 6. Create the match type column
# <p>There are different keyword match types. One is exact match, which is for matching the exact term or are close variations of that exact term. Another match type is broad match, which means ads may show on searches that include misspellings, synonyms, related searches, and other relevant variations.</p>
# <p>Straight from Google's AdWords <a href="https://support.google.com/google-ads/answer/2497836?hl=en">documentation</a>:</p>
# <blockquote>
# <p>In general, the broader the match type, the more traffic potential that keyword will have, since your ads may be triggered more often. Conversely, a narrower match type means that your ads may show less often—but when they do, they’re likely to be more related to someone’s search.</p>
# </blockquote>
# <p>Since the client is tight on budget, we want to make sure all the keywords are in exact match at the beginning.</p>
# + dc={"key": "39"} tags=["sample_code"]
# Add a criterion type column
keywords_df['Criterion Type'] = 'Exact'
# ... YOUR CODE FOR TASK 6 ...
# + dc={"key": "46"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 7. Duplicate all the keywords into 'phrase' match
# <p>The great thing about exact match is that it is very specific, and we can control the process very well. The tradeoff, however, is that: </p>
# <ol>
# <li>The search volume for exact match is lower than other match types</li>
# <li>We can't possibly think of all the ways in which people search, and so, we are probably missing out on some high-quality keywords.</li>
# </ol>
# <p>So it's good to use another match called <em>phrase match</em> as a discovery mechanism to allow our ads to be triggered by keywords that include our exact match keywords, together with anything before (or after) them.</p>
# <p>Later on, when we launch the campaign, we can explore with modified broad match, broad match, and negative match types, for better visibility and control of our campaigns.</p>
# + dc={"key": "46"} tags=["sample_code"]
# Make a copy of the keywords DataFrame
keywords_phrase = keywords_df.copy()
# Change criterion type match to phrase
keywords_phrase['Criterion Type'] = 'Phrase'
# ... YOUR CODE FOR TASK 7 ...
# Append the DataFrames
keywords_df_final = keywords_df.append(keywords_phrase)
# + dc={"key": "53"} editable=false run_control={"frozen": true} deletable=false tags=["context"]
# ## 8. Save and summarize!
# <p>To upload our campaign, we need to save it as a CSV file. Then we will be able to import it to AdWords editor or BingAds editor. There is also the option of pasting the data into the editor if we want, but having easy access to the saved data is great so let's save to a CSV file!</p>
# <p>Looking at a summary of our campaign structure is good now that we've wrapped up our keyword work. We can do that by grouping by ad group and criterion type and counting by keyword. This summary shows us that we assigned specific keywords to specific ad groups, which are each part of a campaign. In essence, we are telling Google (or Bing, etc.) that we want any of the words in each ad group to trigger one of the ads in the same ad group. Separately, we will have to create another table for ads, which is a task for another day and would look something like this:</p>
# <table>
# <thead>
# <tr>
# <th>Campaign</th>
# <th>Ad Group</th>
# <th>Headline 1</th>
# <th>Headline 2</th>
# <th>Description</th>
# <th>Final URL</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>SEM_Sofas</td>
# <td>Sofas</td>
# <td>Looking for Quality Sofas?</td>
# <td>Explore Our Massive Collection</td>
# <td>30-day Returns With Free Delivery Within the US. Start Shopping Now</td>
# <td>DataCampSofas.com/sofas</td>
# </tr>
# <tr>
# <td>SEM_Sofas</td>
# <td>Sofas</td>
# <td>Looking for Affordable Sofas?</td>
# <td>Check Out Our Weekly Offers</td>
# <td>30-day Returns With Free Delivery Within the US. Start Shopping Now</td>
# <td>DataCampSofas.com/sofas</td>
# </tr>
# <tr>
# <td>SEM_Sofas</td>
# <td>Recliners</td>
# <td>Looking for Quality Recliners?</td>
# <td>Explore Our Massive Collection</td>
# <td>30-day Returns With Free Delivery Within the US. Start Shopping Now</td>
# <td>DataCampSofas.com/recliners</td>
# </tr>
# <tr>
# <td>SEM_Sofas</td>
# <td>Recliners</td>
# <td>Need Affordable Recliners?</td>
# <td>Check Out Our Weekly Offers</td>
# <td>30-day Returns With Free Delivery Within the US. Start Shopping Now</td>
# <td>DataCampSofas.com/recliners</td>
# </tr>
# </tbody>
# </table>
# <p>Together, these tables get us the sample <strong>keywords -> ads -> landing pages</strong> mapping shown in the diagram below.</p>
# <p><img src="https://assets.datacamp.com/production/project_400/img/kwds_ads_lpages.png" alt="Keywords-Ads-Landing pages flow"></p>
# + dc={"key": "53"} tags=["sample_code"]
# Save the final keywords to a CSV file
# ... YOUR CODE FOR TASK 8 ...
# View a summary of our campaign work
summary = keywords_df_final.groupby(['Ad Group', 'Criterion Type'])['Keyword'].count()
print(summary)
| Generating Keywords for Google Ads/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Keras Model Conversion
# %matplotlib inline
# ### Start a CAS session
# +
import swat
sess = swat.CAS(cashost, casport)
# -
# ### Define a MNIST classification model in Keras
from keras import Sequential
from keras.layers import *
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5), strides=(1, 1),
activation='relu',padding='same',
input_shape=(28,28,1)))
model.add(MaxPooling2D(pool_size=(2, 2), padding='same', strides=(2, 2)))
model.add(Conv2D(64, (5, 5), padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2), padding='same'))
model.add(Flatten())
model.add(Dense(1000, activation='relu'))
model.add(Dense(10, activation='softmax'))
# **Note:** DLPy now only support for padding='same'.
model.summary()
# ### Convert Keras model to DLPy model
from dlpy import Model
model1 = Model.from_keras_model(conn=sess, keras_model=model, output_model_table='converted_keras_model')
# ### Now model1 is a DLPy model, fully supporting all the function in DLPy.
model1.print_summary()
model1.plot_network()
sess.terminate()
| examples/Example 3 - Keras Model Conversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import some libraries (pre made functions)
import numpy as np
import pandas as pd
import os
import seaborn as sns
import datetime as dt
import matplotlib.pyplot as plt
from datetime import date
from datetime import time
from datetime import datetime
sns.set()
# +
# Set to the folder you are working
folder = r"C:\Users\ellar\Documents\MADE\Living Lab\Python\GPS Data\Binder"
os.chdir(folder)
#Import GPS Data
df = pd.read_csv('GPSData.csv', delimiter=",")
# -
#Show a plot, that in some way looks good.
plt.figure(figsize=(8,8))
sns.jointplot(x="lat",y="lon",data=df,height=16, color = "darkblue")
#plt.color = (df_17dec["imei"])
plt.ylabel('lat')
plt.xlabel('lon')
plt.show()
# # Print Dependences
# +
# %load_ext watermark
#python, ipython, packages and machine characteristics
# %watermark -v -m -p wget,pandas,numpy,datetime,seaborn,matplotlib.pyplot,watermark
#date
print(" ")
# %watermark-u -n -t -z
# -
# %watermark
| Binder Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# from datascience import *
df_powerplant = pd.read_csv("powerplants2.csv") #.column("Population")
df_powerplant.head()
Coal_power = pd.read_csv("powerplants2.csv")["Coal_MW"]
Coal_power.head()
Coal_power[0]
Coal_power_in_kW = Coal_power*1000
Coal_power_in_kW.head()
Coal_power_in_GW = Coal_power/1000
Coal_power_in_GW.head()
powerplants2 = pd.read_csv("powerplants2.csv")
Natural_Gas_power = powerplants2["NG_MW"]
Crude_power = powerplants2["Crude_MW"]
Bio_power = powerplants2["Bio_MW"]
Hydro_power = powerplants2["Hydro_MW"]
HydroPS_power = powerplants2["HydroPS_MW"]
Nuclear_power = powerplants2["Nuclear_MW"]
Solar_power = powerplants2["Solar_MW"]
Wind_power = powerplants2["Wind_MW"]
Geo_power = powerplants2["Geo_MW"]
Other_power = powerplants2["Other_MW"]
print("The total Coal power:\t\t\t\t", Coal_power.sum(), "MW")
print("The total Natural Gas power:\t\t\t", Natural_Gas_power.sum(), "MW")
print("The total Crude Oil power:\t\t\t", Crude_power.sum(), " MW")
print("The total Bioenergy power:\t\t\t", Bio_power.sum(), " MW")
print("The total Hydroelectric power:\t\t\t", Hydro_power.sum(), " MW")
print("The total Pumped-storage Hydroelectric power:\t", HydroPS_power.sum(), " MW")
print("The total Nuclear power:\t\t\t", Nuclear_power.sum(), " MW")
print("The total Solar power:\t\t\t\t", Solar_power.sum(), " MW")
print("The total Wind power:\t\t\t\t", Wind_power.sum(), " MW")
print("The total Geothermal power:\t\t\t", Geo_power.sum(), " MW")
print("The total other power:\t\t\t\t", Other_power.sum(), " MW")
Bio_power = Bio_power.fillna(value=0)
Geo_power = Geo_power.fillna(value=0)
Other_power = Other_power.fillna(value=0)
# Array of entire power of each row, powerplant
entire_power = Coal_power + Natural_Gas_power + Crude_power + Hydro_power + HydroPS_power + Nuclear_power + Solar_power + Wind_power + Bio_power + Geo_power + Other_power
entire_power.head()
# total power of entire row
print("The total power of the entire power plants and sources :", entire_power.sum(), "MW")
print("The total power of the entire power plants and sources :", round(entire_power.sum()/1000, 2), "GW")
| Lec_2_for_Les_Powerplant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
columns = ['City_ID', 'City', 'Cloudiness', 'Country', 'Date', 'Humidity', 'Lat', 'Lng', 'Max Temp', 'Wind Speed']
df = pd.read_csv('WebVisualizations/Resources/cities.csv', names=columns)
print(df.to_html())
| csv_to_html.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
## for data
import json
import pandas as pd
import numpy as np
## for plotting
import matplotlib.pyplot as plt
import seaborn as sns
## for processing
import re
import nltk
## for bag-of-words
from sklearn import feature_extraction, model_selection, naive_bayes, pipeline, manifold, preprocessing
## for explainer
# from lime import lime_text
## for word embedding
import gensim
import gensim.downloader as gensim_api
## for deep learning
from tensorflow.keras import models, layers, preprocessing as kprocessing
from tensorflow.keras import backend as K
## for bert language model
import transformers
import pickle
import pandas as pd
## distil-bert tokenizer
tokenizer = transformers.AutoTokenizer.from_pretrained('distilbert-base-uncased', do_lower_case=True)
with open('./source/wiki_fine_dtf.pickle', 'rb') as handle:
dtf = pickle.load(handle)
# with open('./source/wiki_coarse_dtf.pickle', 'rb') as handle:
# dtf = pickle.load(handle)
'''
Preprocess a string.
:parameter
:param text: string - name of column containing text
:param lst_stopwords: list - list of stopwords to remove
:param flg_stemm: bool - whether stemming is to be applied
:param flg_lemm: bool - whether lemmitisation is to be applied
:return
cleaned text
'''
def utils_preprocess_text(text, flg_stemm=False, flg_lemm=True, lst_stopwords=None):
## clean (convert to lowercase and remove punctuations and characters and then strip)
text = re.sub(r'[^\w\s]', '', str(text).lower().strip())
## Tokenize (convert from string to list)
lst_text = text.split()
## remove Stopwords
if lst_stopwords is not None:
lst_text = [word for word in lst_text if word not in
lst_stopwords]
## Stemming (remove -ing, -ly, ...)
if flg_stemm == True:
ps = nltk.stem.porter.PorterStemmer()
lst_text = [ps.stem(word) for word in lst_text]
## Lemmatisation (convert the word into root word)
if flg_lemm == True:
lem = nltk.stem.wordnet.WordNetLemmatizer()
lst_text = [lem.lemmatize(word) for word in lst_text]
## back to string from list
text = " ".join(lst_text)
return text
# +
import timeit
start_time = timeit.default_timer()
# -
lst_stopwords = nltk.corpus.stopwords.words("english")
# lst_stopwords
dtf["text_clean"] = dtf["X"].apply(lambda x:
utils_preprocess_text(x, flg_stemm=False, flg_lemm=True,
lst_stopwords=lst_stopwords))
# dtf.head()
## split dataset
dtf_train, dtf_test = model_selection.train_test_split(dtf, test_size=0.3)
## get target
y_train = dtf_train["y"].values
y_test = dtf_test["y"].values
# +
# corpus = dtf_train["text_clean"]
# maxlen = 50
# ## add special tokens
# maxqnans = np.int((maxlen-20)/2)
# corpus_tokenized = ["[CLS] "+
# " ".join(tokenizer.tokenize(re.sub(r'[^\w\s]+|\n', '',
# str(txt).lower().strip()))[:maxqnans])+
# " [SEP] " for txt in corpus]
# ## generate masks
# masks = [[1]*len(txt.split(" ")) + [0]*(maxlen - len(
# txt.split(" "))) for txt in corpus_tokenized]
# ## padding
# txt2seq = [txt + " [PAD]"*(maxlen-len(txt.split(" "))) if len(txt.split(" ")) != maxlen else txt for txt in corpus_tokenized]
# ## generate idx
# idx = [tokenizer.encode(seq.split(" ")) for seq in txt2seq]
# ## generate segments
# segments = []
# for seq in txt2seq:
# temp, i = [], 0
# for token in seq.split(" "):
# temp.append(i)
# if token == "[SEP]":
# i += 1
# segments.append(temp)
# ## feature matrix
# X_train = [np.asarray(idx, dtype='int32'),
# np.asarray(masks, dtype='int32'),
# np.asarray(segments, dtype='int32')]
# +
def encode_data(corpus):
maxlen = 50
## add special tokens
maxqnans = np.int((maxlen-20)/2)
corpus_tokenized = ["[CLS] "+
" ".join(tokenizer.tokenize(re.sub(r'[^\w\s]+|\n', '',
str(txt).lower().strip()))[:maxqnans])+
" [SEP] " for txt in corpus]
## generate masks
masks = [[1]*len(txt.split(" ")) + [0]*(maxlen - len(
txt.split(" "))) for txt in corpus_tokenized]
## padding
txt2seq = [txt + " [PAD]"*(maxlen-len(txt.split(" "))) if len(txt.split(" ")) != maxlen else txt for txt in corpus_tokenized]
## generate idx
idx = [tokenizer.encode(seq.split(" ")) for seq in txt2seq]
## generate segments
segments = []
for seq in txt2seq:
temp, i = [], 0
for token in seq.split(" "):
temp.append(i)
if token == "[SEP]":
i += 1
segments.append(temp)
## feature matrix
result = [np.asarray(idx, dtype='int32'),
np.asarray(masks, dtype='int32'),
np.asarray(segments, dtype='int32')]
return result
# corpus = dtf_test["text_clean"]
X_test = encode_data(dtf_test["text_clean"])
X_train = encode_data(dtf_train["text_clean"])
# -
## inputs
idx = layers.Input((50), dtype="int32", name="input_idx")
masks = layers.Input((50), dtype="int32", name="input_masks")
segments = layers.Input((50), dtype="int32", name="input_segments")
## pre-trained bert
nlp = transformers.TFBertModel.from_pretrained("bert-base-uncased")
bert_out, _ = nlp([idx, masks, segments])
## fine-tuning
x = layers.GlobalAveragePooling1D()(bert_out)
x = layers.Dense(64, activation="relu")(x)
y_out = layers.Dense(len(np.unique(y_train)),
activation='softmax')(x)
## compile
model = models.Model([idx, masks, segments], y_out)
for layer in model.layers[:4]:
layer.trainable = False
model.compile(loss='sparse_categorical_crossentropy',
optimizer='adam', metrics=['accuracy'])
model.summary()
# +
# ## inputs
# idx = layers.Input((50), dtype="int32", name="input_idx")
# masks = layers.Input((50), dtype="int32", name="input_masks")
# ## pre-trained bert with config
# config = transformers.DistilBertConfig(dropout=0.2,
# attention_dropout=0.2)
# config.output_hidden_states = False
# nlp = transformers.TFDistilBertModel.from_pretrained('distilbert-base-uncased', config=config)
# bert_out = nlp(idx, attention_mask=masks)[0]
# ## fine-tuning
# x = layers.GlobalAveragePooling1D()(bert_out)
# x = layers.Dense(64, activation="relu")(x)
# y_out = layers.Dense(len(np.unique(y_train)),
# activation='softmax')(x)
# ## compile
# model = models.Model([idx, masks], y_out)
# for layer in model.layers[:3]:
# layer.trainable = False
# model.compile(loss='sparse_categorical_crossentropy',
# optimizer='adam', metrics=['accuracy'])
# model.summary()
# +
# y_train
# + jupyter={"outputs_hidden": true}
## encode y
dic_y_mapping = {n:label for n,label in
enumerate(np.unique(y_train))}
inverse_dic = {v:k for k,v in dic_y_mapping.items()}
y_train = np.array([inverse_dic[y] for y in y_train])
## train
training = model.fit(x=X_train, y=y_train, batch_size=64,
epochs=10, shuffle=True, verbose=1,
validation_split=0.3)
## test
# -
elapsed = timeit.default_timer() - start_time
print (elapsed)
predicted_prob = model.predict(X_test)
predicted = [dic_y_mapping[np.argmax(pred)] for pred in
predicted_prob]
# +
# dic_y_mapping
# +
from sklearn import metrics
classes = np.unique(y_test)
y_test_array = pd.get_dummies(y_test, drop_first=False).values
## Accuracy, Precision, Recall
accuracy = metrics.accuracy_score(list(y_test), list(predicted))
auc = metrics.roc_auc_score(y_test_array, predicted_prob, multi_class="ovr")
print("Accuracy:", round(accuracy,2))
print("Auc:", round(auc,2))
print("Detail:")
print(metrics.classification_report(y_test, predicted))
## Plot confusion matrix
cm = metrics.confusion_matrix(y_test, predicted)
fig, ax = plt.subplots()
sns.heatmap(cm, annot=True, fmt='d', ax=ax, cmap=plt.cm.Blues,
cbar=False)
ax.set(xlabel="Pred", ylabel="True", xticklabels=classes,
yticklabels=classes, title="Confusion matrix")
plt.yticks(rotation=0)
plt.savefig('bert_confusion_matrix.png')
fig, ax = plt.subplots(nrows=1, ncols=2)
## Plot roc
for i in range(len(classes)):
fpr, tpr, thresholds = metrics.roc_curve(y_test_array[:,i],
predicted_prob[:,i])
ax[0].plot(fpr, tpr, lw=3,
label='{0} (area={1:0.2f})'.format(classes[i],
metrics.auc(fpr, tpr))
)
ax[0].plot([0,1], [0,1], color='navy', lw=3, linestyle='--')
ax[0].set(xlim=[-0.05,1.0], ylim=[0.0,1.05],
xlabel='False Positive Rate',
ylabel="True Positive Rate (Recall)",
title="Receiver operating characteristic")
ax[0].legend(loc="lower right")
ax[0].grid(True)
## Plot precision-recall curve
for i in range(len(classes)):
precision, recall, thresholds = metrics.precision_recall_curve(
y_test_array[:,i], predicted_prob[:,i])
ax[1].plot(recall, precision, lw=3,
label='{0} (area={1:0.2f})'.format(classes[i],
metrics.auc(recall, precision))
)
ax[1].set(xlim=[0.0,1.05], ylim=[0.0,1.05], xlabel='Recall',
ylabel="Precision", title="Precision-Recall curve")
ax[1].legend(loc="best")
ax[1].grid(True)
# plt.show()
plt.savefig('auc.png')
| notebook/.ipynb_checkpoints/ws-16-Bert-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CS-VQE model data generation
#
# Generates all of the data required to conduct CS-VQE simulations using utils.molecule_tools.construct_molecule and the CS-VQE circuit class cs_vqe.circuit.cs_vqe_circuit.
from utils.molecule_tools import construct_molecule
from utils.cs_vqe_tools_original import greedy_dfs
from cs_vqe.circuit import cs_vqe_circuit
import json
# +
with open('data/molecule_data.json', 'r') as json_file:
molecule_data = json.load(json_file)
with open('data/model_data.json', 'r') as json_file:
model_data = json.load(json_file)
#with open('data/anz_circ_depth.json', 'r') as json_file:
# anz_data = json.load(json_file)
#speciesname_list = [mol for mol in anz_data if (anz_data[mol]['num_qubits']!=anz_data[mol]['chemaccnum'] and not anz_data[mol]['chemaccnum']==0) and mol.find('alt')==-1 and mol.find('+')==-1]
#speciesname_list
# +
#model_data = {}
for speciesname in ['B+_STO-3G_SINGLET']:#speciesname_list:
print(speciesname)
atoms, coords, multiplicity, charge, basis, sym_sector = molecule_data[speciesname].values()
molecule = construct_molecule(atoms, coords, charge, multiplicity, basis,
excitation_threshold=None, taper=True, sym_sector=sym_sector)
num_qubits = molecule['num_qubits']
ham = molecule['hamiltonian']
ansatze = molecule['ansatze']
hf_config = molecule['hf_config']
hf_energy = molecule['hf_energy']
num_tapered = molecule['num_tapered']
true_gs_nrg = molecule['true_gs_nrg']
#true_gs_vec = molecule['true_gs_vec']
terms_noncon = greedy_dfs(ham, 30, criterion='size')[-1]
print('Number of qubits:', num_qubits)
mol_circ = cs_vqe_circuit(hamiltonian = ham,
terms_noncon= terms_noncon,
num_qubits = num_qubits,
hf_config = hf_config)
ham_rotations = mol_circ.ham_rotations
noncon = mol_circ.noncon
truegs = mol_circ.truegs
G = mol_circ.G
A = mol_circ.A
X_index = mol_circ.X_index
X_qubit = mol_circ.X_qubit
cs_vqe_energy = mol_circ.cs_vqe_energy
cs_vqe_errors = mol_circ.cs_vqe_errors
chem_acc_num_q = mol_circ.chem_acc_num_q
order = mol_circ.order
ham_reduced = mol_circ.ham_reduced
reference_state= mol_circ.reference_state()
model_data[speciesname] = {"ham":ham,
"ansatze":ansatze,
"num_qubits":num_qubits,
"hf_config":hf_config,
"hf_energy":hf_energy,
"terms_noncon":terms_noncon,
"num_tapered":num_tapered,
"true_gs_nrg":true_gs_nrg,
#"true_gs_vec":true_gs_vec,
"ham_rotations":ham_rotations,
"noncon":noncon,
"truegs":truegs,
"G":G,
"A":A,
"X_index":X_index,
"X_qubit":X_qubit,
"cs_vqe_energy":cs_vqe_energy,
"cs_vqe_errors":cs_vqe_errors,
"chem_acc_num_q":chem_acc_num_q,
"order":order,
"ham_reduced":ham_reduced,
"reference_state":reference_state
}
print('Molecule constructed.\n')
# +
#data={}
#for k in model_data.keys():
# data[str(k)] = model_data[k]
with open("data/model_data.json", "w") as outfile:
json.dump(model_data, outfile)
# -
for name, a in model_data['HF_STO-3G_SINGLET']['ansatze'].items():
print(name, len(a))
model_data['B+_STO-3G_SINGLET']
| generate_model_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## 数据应用学院 Data Scientist Program
# ## Hw1
# import the necessary package at the very beginning
import numpy as np
import pandas as pd
print(str(float(100*177/891)) + '%')
# #### 1. Please rewrite following functions to lambda expressions
# Example:
#
# ```
# def AddOne(x):
# y=x+1
# return y
#
# addOneLambda = lambda x: x+1
# ```
def foolOne(x): # note: assume x is a number
y = x * 2
y -= 25
return y
## Type Your Answer Below ##
foolOne_lambda = lambda x: x*2-25
# Generate a random 3*4 matrix for test
tlist = np.random.randn(3,4)
tlist
# +
# Check if the lambda function yields same results as previous function
def test_foolOne(tlist, func1, func2):
if func1(tlist).all() == func2(tlist).all():
print("Same results!")
test_foolOne(tlist, foolOne, foolOne_lambda)
# -
def foolTwo(x): # note: assume x here is a string
if x.startswith('g'):
return True
else:
return False
## Type Your Answer Below ##
foolTwo_lambda = lambda x: x.startswith('g')
# +
# Generate a random 3*4 matrix of strings for test
# reference: https://pythontips.com/2013/07/28/generating-a-random-string/
# reference: http://www.programcreek.com/python/example/1246/string.ascii_lowercase
import random
import string
def random_string(size):
new_string = ''.join([random.choice(string.ascii_letters + string.digits) for n in range(size)])
return new_string
def test_foolTwo():
test_string = random_string(6)
if foolTwo_lambda(test_string) == foolTwo(test_string):
return True
for i in range(10):
if test_foolTwo() is False:
print('Different results!')
# -
# #### 2. What's the difference between tuple and list?
## Type Your Answer Below ##
# reference: https://docs.python.org/3/tutorial/datastructures.html
# tuple is immutable. They cannot be changed once they are made.
# tuples are easier for the python interpreter to deal with and therefore might end up being easier
# tuples might indicate that each entry has a distinct meaning and their order has some meaning (e.g., year)
# Another pragmatic reason to use tuple is when you have data which you know should not be changed (e.g., constant)
# tuples can be used as keys in dictionaries
# tuples usually contain a heterogeneous sequence of elements that are accessed via unpacking or indexing (or even by attribute in the case of namedtuples).
tuple1 = (1, 2, 3, 'a', True)
print('tuple: ', tuple1)
print('1st item of tuple: ', tuple1[0])
tuple1[0] = 4 # item assignment won't work for tuple
# tuple with just one element
tuple2 = (1) # just a number, so has no elements
print(type(tuple2))
tuple2[0]
# tuple with just one element
tuple3 = (1, )
print(type(tuple3))
tuple3[0]
# Question for TA: is tuple comprehension supported?
tuple4 = (char for char in 'abcdabcdabcd' if char not in 'ac')
print(tuple4)
# Question for TA: is the following two tuples the same?
tuple4= (1,2,'a'),(True, False)
tuple5 = ((1,2,'a'),(True, False))
print(tuple4)
print(tuple5)
# +
# lists' elements are usually homogeneous and are accessed by iterating over the list.
list1 = [1, 2, 3, 'a', True]
print('list1: ', list1)
print('1st item of list: ', list1[0])
list1[0] = 4 # item assignment works for list
# list comprehensions
list_int = [element for element in list1 if type(element)==int]
print("list_int", list2)
# +
## Type Your Answer Below ##
# A set is an unordered collection with no duplicate elements.
# set() can be used to eliminate duplicate entries
list1 = ['apple', 'orange', 'apple', 'pear', 'orange', 'banana']
set1 = set(list1)
print(set1)
# set can be used for membership testing
set2 = {1, 2, 'abc', True}
print('abc' in set2) # membership testing
set1[0] # set does not support indexing
# -
# set comprehensions
set4 = {char for char in 'abcdabcdabcd' if char not in 'ac'}
print(set4)
# #### 3. Why set is faster than list in python?
# #### Answers:
# Set and list are implemented using two different data structures - Hash tables and Dynamic arrays.
# . Python lists are implemented as dynamic arrays (which can preserve ), which must be searched one by one to compare every single member for equality, with lookup speed O(n) depending on the size of the list.
# . Python sets are implemented as hash tables, which can directly jump and locate the bucket (the position determined by the object's hash) using hash in a constant speed O(1), regardless of the size of the set.
# +
# Calculate the time cost differences between set and list
import time
import random
def compute_search_speed_difference(scope):
list1 = []
dic1 = {}
set1 = set(dic1)
for i in range(0,scope):
list1.append(i)
set1.add(i)
random_n = random.randint(0,100000) # look for this random integer in both list and set
list_search_starttime = time.time()
list_search = random_n in list1
list_search_endtime = time.time()
list_search_time = list_search_endtime - list_search_starttime # Calculate the look-up time in list
#print("The look up time for the list is:")
#print(list_search_time)
set_search_starttime = time.time()
set_search = random_n in set1
set_search_endtime = time.time()
set_search_time = set_search_endtime - set_search_starttime # Calculate the look-up time in set
#print("The look up time for the set is:")
#print(set_search_time)
speed_difference = list_search_time - set_search_time
return(speed_difference)
def test(testing_times, scope):
test_speed_difference = []
for i in range(0,testing_times):
test_speed_difference.append(compute_search_speed_difference(scope))
return(test_speed_difference)
#print(test(1000, 100000)) # test 10 times can print out the time cost differences
print("On average, the look up time for a list is more than a set in:")
print(np.mean(test(100, 1000)))
# -
# #### 4. What's the major difference between array in numpy and series in pandas?
# Pandas series (which can contain values of different data types) is much more general and flexible than the one-dimensional Numpy array(which can only contain one data type).
#
# While Numpy array has an implicitly defined integer used to access the values, the Pandas series has an explicitly defined index (which can be any data type) associated with the values (which gives the series object additonal capabilities).
#
# ### What's the relationships among Numpy, Pandas and SciPy:
# . Numpy is a libary for efficient array computations, modeled after Matlab. Arrays differ from plain Python lists in the way they are stored and handled. Array elements stay together in memory, so they can be quickly accessed. Numpy also supports quick subindexing (a[0,:,2]). Furthermore, Numpy provides vectorized mathematical functions (when you call numpy.sin(a), the sine function is applied on every element of array a), which are faster than a Python for loop.
#
# . Pandas library is good for analyzing tabular data for exploratory data analysis, statistics and visualization. It's used to understand the data you have.
#
# . Scipy provides large menu of libraries for scientific computation, such as integration, interpolation, signal processing, linear algebra, statistics. It's built upon the infrastructure of Numpy. It's good for performing scientific and engineering calculation.
#
# . Scikit-learn is a collection of advanced machine-learning algorithms for Python. It is built upon Numpy and SciPy. It's good to use the data you have to train a machine-learning algorithm.
#
## Type Your Answer Below ##
student = np.array([0, 'Alex', 3, 'M'])
print(student) # all the values' datatype is converted to str
# #### Question 5-11 are related to titanic data (train.csv) on kaggle website
# You can download the data from the following link:<br />https://www.kaggle.com/c/titanic/data
# #### 5. Read titanic data (train.csv) into pandas dataframe, and display a sample of data.
## Type Your Answer Below ##
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/pcsanwald/kaggle-titanic/master/train.csv')
df.sample(3)
df.tail(3)
df.describe()
df.info()
# #### 6. What's the percentage of null value in 'Age'?
## Type Your Answer Below ##
len(df[df.age.isnull()])/len(df)*100
# #### 7. How many unique classes in 'Embarked' ?
## Type Your Answer Below ##
df.embarked.value_counts()
print('number of classes: ', len(df.embarked.value_counts().index))
print('names of classes: ', df.embarked.value_counts().index)
# Another method
embarked_set = set(df.embarked)
print(df.embarked.unique())
# #### 8. Compare survival chance between male and female passangers.
# Please use pandas to plot a chart you think can address this question
## Type Your Answer Below ##
male_survived = df[df.survived==1][df.sex=='male']
male_survived_n = len(df.query('''sex=='male' and survived ==1'''))
female_survived = df[df.survived==1][df.sex=='female']
female_survived_n = len(df.query('''sex=='female' and survived ==1'''))
df_survived = pd.DataFrame({'male':male_survived_n, 'female': female_survived_n}, index=['Survived_number'])
df_survived
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
df_survived.plot(kind='bar', title='survived female and male', legend='True')
sns.pointplot(x='embarked', y='survived', hue='sex', data=df, palette={'male':'blue', 'female':'pink'}, markers=["*", "o"], linestyles=['-', '--'])
grid = sns.FacetGrid(df, col='embarked')
grid.map(sns.pointplot, 'pclass', 'survived', 'sex', palette={'male':'blue', 'female':'pink'}, markers=["*", "o"], linestyles=['-', '--'])
grid.add_legend()
grid = sns.FacetGrid(data_train, col='pclass')
grid.map(sns.barplot, 'embarked', 'age', 'sex')
grid.add_legend()
# #### Observations from barplot above:
# 1. In Pclass = 1 and 2, female has higher mean age than male. But in Pclass = 3, female has lower mean age than male.
# 2. Passengers in Pclass = 1 has the highest average age, followed by Pclass = 2 and Pclass = 3.
# 3. Age trend among Embarked is not abvious
#
# #### Decisions:
# Use 'Pclass'and 'Sex' in estimating missing values in 'Age'.
# #### 9. Show the table of passangers who are 23 years old.
## Type Your Answer Below ##
df_23=df.query('''age>23''')
df_23
# #### 10. Is there a Jack or Rose in our dataset?
# +
# first split name into string lists by ' '
def format_name(df):
df['split_name'] = df.name.apply(lambda x: x.split(' '))
return df
print(df.sample(3).split_name, '\n')
# for each subset string of name, check if "jack" or "rose" in it
for i in format_name(df).split_name:
for l in i:
if (("jack" in l.lower()) | ("rose" in l.lower()) ):
print("found names that contain jack or rose: ", l)
# -
# #### 11. What's the percentage of surviving when passangers' pclass is 1?
# +
## Type Your Answer Below ##
df4 = df.query('''pclass==1''')
def percent(x):
m = int(x.count())
n = m/len(df4)
return(n)
df[['survived','pclass']].query('''pclass==1''').groupby([ 'survived']).agg({'pclass':percent})
# -
# ### Refereences
# https://www.dataquest.io/blog/jupyter-notebook-tips-tricks-shortcuts/
# https://docs.python.org/3/tutorial/datastructures.html
# https://stackoverflow.com/questions/2030053/random-strings-in-python
#
| DS_HW1_Huimin Qian_052617.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <a href="https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv" target="_blank"><img align="left" src="data/cover.jpg" style="width: 76px; height: 100px; background: white; padding: 1px; border: 1px solid black; margin-right:10px;"></a>
# *This notebook contains an excerpt from the book [Machine Learning for OpenCV](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv) by <NAME>.
# The code is released under the [MIT license](https://opensource.org/licenses/MIT),
# and is available on [GitHub](https://github.com/mbeyeler/opencv-machine-learning).*
#
# *Note that this excerpt contains only the raw code - the book is rich with additional explanations and illustrations.
# If you find this content useful, please consider supporting the work by
# [buying the book](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv)!*
# <!--NAVIGATION-->
# < [Chaining Algorithms Together to Form a Pipeline](11.04-Chaining-Algorithms-Together-to-Form-a-Pipeline.ipynb) | [Contents](../README.md) |
# # Wrapping Up
#
# Congratulations! You have just made a big step toward becoming a machine learning
# practitioner. Not only are you familiar with a wide variety of fundamental machine
# learning algorithms, you also know how to apply them to both supervised and
# unsupervised learning problems.
#
# Before we part ways, I want to give you some final words of advice, point you toward some
# additional resources, and give you some suggestions on how you can further improve your
# machine learning and data science skills.
# ## Approaching a machine learning problem
#
# When you see a new machine learning problem in the wild, you might be tempted to jump
# ahead and throw your favorite algorithm at the problem—perhaps the one you understood
# best or had the most fun implementing. But knowing beforehand which algorithm will
# perform best on your specific problem is not often possible.
#
# Instead, you need to take a step back and look at the big picture.
# Here the book provides an easy-to-follow outline on how to approach machine learning problems in the wild (p.331ff.).
# ## Writing your own OpenCV based classifier in C++
#
# Since OpenCV is one of those Python libraries that does not contain a single line of Python
# code under the hood (I'm kidding, but it's close), you will have to implement your custom
# estimator in C++.
# The first step is to define a
# file `MyClass.cpp`:
#
# #include <opencv2/opencv.hpp>
# #include <opencv2/ml/ml.hpp>
# #include <stdio.h>
#
# class MyClass : public cv::ml::StatModel
# {
# public:
# MyClass()
# {
# print("MyClass constructor\n");
# }
#
# ~MyClass() {}
#
# int getVarCount() const
# {
# // returns the number of variables in the training samples
# return 0;
# }
#
# bool empty() const
# {
# return true;
# }
#
# bool isTrained() const
# {
# // returns true if the model is trained
# return false;
# }
#
# bool isClassifier() const
# {
# // returns true if the model is a classifier
# return true;
# }
#
# bool train(const cv::Ptr<cv::ml::TrainData>& trainData, int flags=0) const
# {
# // trains the model
# // trainData: training data that can be loaded from file using
# // TrainData::loadFromCSV or created with TrainData::create.
# // flags: optional flags, depending on the model. Some of the models
# // can be updated with the new training samples, not completely
# // overwritten (such as NormalBayesClassifier or ANN_MLP).
# return false;
# }
#
# bool train(cv::InputArray samples, int layout, cv::InputArray responses)
# {
# // trains the model
# // samples: training samples
# // layout: see ml::SampleTypes
# // responses: vector of responses associated with the training samples
# return false;
# }
#
# float calcError(const cv::Ptr<cv::ml::TrainData>& data, bool test, cv::OutputArray resp)
# {
# // calculates the error on the training or test set
# // data: the training data
# // test: if true, the error is computed over the test subset of the data, otherwise
# // it's computed over the training subset of the data.
# return 0.0f;
# }
#
# float predict(cv::InputArray samples, cv::OutputArray results=cv::noArray(), int flags=0) const
# {
# // predicts responses for the provided samples
# // samples: the input samples, floating-point matrix
# // results: the optional matrix of results
# // flags: the optional flags, model-dependent. see cv::ml::StatModel::Flags
# return 0.0f;
# }
# };
#
# int main()
# {
# MyClass myclass;
# return 0;
# }
# Then create a file `CMakeLists.txt`:
#
# cmake_minimum_required(VERSION 2.8)
# project(MyClass)
# find_package(OpenCV REQUIRED)
# add_executable(MyClass MyClass.cpp)
# target_link_libraries(MyClass ${OpenCV_LIBS})
# Then you can compile the file from the command line via `cmake` and `make`:
#
# $ cmake .
# $ make
# Then run the file:
#
# $ ./MyClass
#
# This should not generate any error, and print to console:
#
# MyClass constructor
# ## Writing your own Scikit-Learn based classifier in Python:
#
# Alternatively, you can write your own classifier using the scikit-learn library.
#
# You can do this by importing `BaseEstimator` and `ClassifierMixin`. The latter will
# provide a corresponding `score` method, which works for all classifiers. Optionally, you can
# overwrite the score method to provide your own.
#
# The following mixins are available:
# - `ClassifierMixin` if you are writing a classifier (will provide a basic `score` method)
# - `RegressorMixin` if you are writing a regressor (will provide a basic `score` method)
# - `ClusterMixin` if you are writing a clustering algorithm (will provide a basic `fit_predict` method)
# - `TransformerMixin` if you are writing a transformer (will provide a basic `fit_predict` method)
import numpy as np
from sklearn.base import BaseEstimator, ClassifierMixin
class MyClassifier(BaseEstimator, ClassifierMixin):
"""An example classifier"""
def __init__(self, param1=1, param2=2):
"""Called when initializing the classifier
The constructor is used to define some optional
parameters of the classifier. Store them as class
attributes for future access.
Parameters
----------
param1 : int, optional, default: 1
The first parameter
param2 : int, optional, default: 2
The second parameter
"""
self.param1 = param1
self.param2 = param2
def fit(self, X, y=None):
"""Fits the classifier to data
This should fit the classifier to the training data.
All the "work" should be done here.
Parameters
----------
X : array-like
The training data, where the first dimension is
the number of training samples, and the second
dimension is the number of features.
y : array-like, optional, default: None
Vector of class labels
Returns
-------
The fit method returns the classifier object it
belongs to.
"""
return self
def predict(self, X):
"""Predicts target labels
This should predict the target labels of some data `X`.
Parameters
----------
X : array-like
Data samples for which to predict the target labels.
Returns
-------
y_pred : array-like
Target labels for every data sample in `X`
"""
return np.zeros(X.shape[0])
# The classifier can be instantiated as follows:
myclass = MyClassifier()
# You can then fit the model to some arbitrary data:
X = np.random.rand(10, 3)
myclass.fit(X)
# And then you can proceed to predicting the target responses:
myclass.predict(X)
# ## Where to go from here?
#
# The goal of this book was to introduce you to the world of machine learning and prepare
# you to become a machine learning practitioner. Now that you know everything about the
# fundamental algorithms, you might want to investigate some topics in more depth.
#
# Although it is not necessary to understand all the details of all the algorithms we
# implemented in this book, knowing some of the theory behind them might just make you a
# better data scientist.
#
# Turn to the book to find a list of suggested reading materials, books, and machine learning software!
# ## Summary
#
# In this book, we covered a lot of theory and practice.
#
# We discussed a wide variety of fundamental machine learning algorithms, be it supervised
# or unsupervised, illustrated best practices as well as ways to avoid common pitfalls, and we
# touched upon a variety of commands and packages for data analysis, machine learning, and
# visualization.
#
# If you made it this far, you have already made a big step toward machine learning mastery.
# From here on out, I am confident you will do just fine on your own.
# All that's left to say is farewell! I hope you enjoyed the ride; I certainly did.
# <!--NAVIGATION-->
# < [Chaining Algorithms Together to Form a Pipeline](11.04-Chaining-Algorithms-Together-to-Form-a-Pipeline.ipynb) | [Contents](../README.md) |
| notebooks/12.00-Wrapping-Up.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import division
import numpy as np
import pandas as pd
# -
data_train = pd.read_csv("./data/train.csv")
data_train.head()
data_challenge = pd.read_csv("./data/challenge_data.csv")
data_challenge.head()
data_test = pd.read_csv("./data/test.csv")
data_test.head()
data_train['user_id'] = data_train['user_id'].apply(lambda x: str(x))
data_train['user_id'].describe()
data_test['user_id'] = data_test['user_id'].apply(lambda x: str(x))
data_test['user_id'].describe()
# +
data_all = pd.concat([data_train, data_test], axis=0)
data_all = data_all.sort_values('user_id')
# data_all = data_all.sample(frac=1).reset_index(drop=True)
data_all.info()
# -
data_all['user_id'].describe()
data_all.isnull().sum()
data_all['challenge_sequence'] = data_all['challenge_sequence'].apply(lambda x: 14 - x)
data_all.head(20)
data_all = data_all[['user_id', 'challenge', 'challenge_sequence']]
data_all.info()
data_all.head()
# +
import pandas as pd
from surprise import SVD
from surprise import KNNBasic
from surprise import Dataset
from surprise import Reader
# +
# A reader is still needed but only the rating_scale param is requiered.
reader = Reader(rating_scale=(1, 13))
# The columns must correspond to user id, item id and ratings (in that order).
data = Dataset.load_from_df(data_all.iloc[:], reader)
# +
# Retrieve the trainset.
trainset = data.build_full_trainset()
# Build an algorithm, and train it.
# algo = KNNBasic()
algo = SVD()
# algo.fit(trainset)
# from surprise import NormalPredictor, evaluate
# algo = NormalPredictor()
algo.fit(trainset)
# +
# uid = '4577' # raw user id (as in the ratings file).
# iid = 'CI23855' # raw item id (as in the ratings file).
# get a prediction for specific users and items, the rating is supposed to be 13.
pred_user_id_list = data_test.user_id.unique()
print(len(pred_user_id_list))
pred_challenge_id_list = data_all.challenge.unique()
print(len(pred_challenge_id_list))
# pred_challenge_id_list = data_challenge['challenge_ID'].unique()
# print(len(pred_challenge_id_list))
# +
from numpy import random
predictions = pd.DataFrame(columns=['user', 'challenge', 'rating'])
for pred_user_id in pred_user_id_list[:]:
# print("user ID is ", pred_user_id)
for pred_challenge_id in random.choice(pred_challenge_id_list, 3):
repeat_list = data_test['challenge'][data_test['user_id'] == pred_user_id].tolist()
if pred_challenge_id not in repeat_list:
pred = algo.predict(pred_user_id, pred_challenge_id, r_ui=None, verbose=False)
if float(pred.est) > 7.0:
predictions = predictions.append(pd.Series([pred_user_id, pred_challenge_id, pred.est], \
index=['user', 'challenge', 'rating']), \
ignore_index=True)
# predictions.iloc[0:9:3, 2] = 3.0
# predictions.iloc[:15, 1] = 'CI22222'
predictions.info()
predictions.head(10)
# -
predictions['rank'] = predictions.groupby('user')['rating'].rank(ascending=False)
predictions.head()
# +
predictions = predictions.sort_values(['rank'])
predictions = predictions.groupby('user').head(3)
predictions = predictions.sort_values(['user', 'rank']).reset_index(drop=True)
predictions.head(20)
# +
current = None
seq = 11
results = pd.DataFrame(columns=['user_sequence', 'challenge'])
for index, row in predictions.iterrows():
if row['user'] != current:
seq = 11
result_user_seq = row['user'] + '_' + str(seq)
result_challenge = row['challenge']
results = results.append(pd.Series([row['user'] + '_' + str(seq), row['challenge']], index=['user_sequence', 'challenge']), ignore_index=True)
seq += 1
current = row['user']
results.head(9)
# -
results.to_csv('./submission/submission_svd.csv', encoding='utf-8', index=False)
| python3_svd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction to Scikit-clean
# `scikit-clean` is a python ML library for classification in the presence of label noise. Aimed primarily at researchers, this provides implementations of several state-of-the-art algorithms, along with tools to simulate artificial noise, create complex pipelines and evaluate them.
# ### Example Usage
# Before we dive into the details, let's take a quick look to see how it works. scikit-clean, as the name implies, is built on top of scikit-learn and is fully compatible* with scikit-learn API. scikit-clean classifiers can be used as a drop in replacement for scikit-learn classifiers.
#
# In the simple example below, we corrupt a dataset using artifical label noise, and then train a model using robust logistic regression:
# +
from sklearn.datasets import make_classification, load_breast_cancer
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, cross_val_score
from skclean.simulate_noise import flip_labels_uniform, UniformNoise, CCNoise
from skclean.models import RobustLR
from skclean.pipeline import Pipeline, make_pipeline
SEED = 42
X, y = load_breast_cancer(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.30, random_state=SEED)
y_train_noisy = flip_labels_uniform(y_train, .3, random_state=SEED) # Flip labels of 30% samples
clf = RobustLR(random_state=SEED).fit(X_train,y_train_noisy)
print(clf.score(X_test, y_test))
# -
# You can use scikit-learn's built in tools with scikit-clean. For example, let's tune one hyper-parameter of `RobustLR` used above, and evaluate the resulting model using cross-validation:
# +
from sklearn.model_selection import GridSearchCV, cross_val_score
grid_clf = GridSearchCV(RobustLR(),{'PN':[.1,.2,.4]},cv=3)
cross_val_score(grid_clf, X, y, cv=5, n_jobs=5).mean() # Note: here we're training & testing here on clean data for simplicity
# -
# ### Algorithms
# Algorithms implemented in scikit-clean can be broadly categorized into two types. First we have ones that are *inherently* robust to label noise. They often modify or replace the loss functions of existing well known algorithms like SVM, Logistic Regression etc. and do not explcitly try to detect mislabeled samples in data. `RobustLR` used above is a robust variant of regular Logistic Regression. These methods can currently be found in `skclean.models` module, though this part of API is likely to change in future as no. of implementations grow.
#
# On the other hand we have *Dataset-focused* algorithms: their focus is more on identifying or cleaning the dataset, they usually rely on other existing classifiers to do the actual learning. Majority of current scikit-clean implementations fall under this category, so we describe them in a bit more detail in next section.
# ### Detectors and Handlers
# Many robust algorithms designed to handle label noise can be essentially broken down to two sequential steps: detect samples which has (probably) been mislabeled, and use that information to build robust meta classifiers on top of existing classifiers. This allows us to easily create new robust classifiers by mixing the noise detector of one paper with the noise-handler of another.
#
# In scikit-clean, the classes that implement those two tasks are called `Detector` and `Handler` respectively. During training, `Detector` will find for each sample the probability that it has been *correctly* labeled (i.e. `conf_score`). `Handler` can use that information in many ways, like removing likely noisy instances from dataset (`Filtering` class), or assigning more weight on reliable samples (`example_weighting` module) etc.
#
# Let's rewrite the above example. We'll use `KDN`: a simple neighborhood-based noise detector, and `WeightedBagging`: a variant of regular bagging that takes sample reliability into account.
# +
from skclean.detectors import KDN
from skclean.handlers import WeightedBagging
conf_score = KDN(n_neighbors=5).detect(X_train, y_train_noisy)
clf = WeightedBagging(n_estimators=50).fit(X_train, y_train_noisy, conf_score)
print(clf.score(X_test, y_test))
# -
# The above code is fine for very simple workflow. However, real world data modeling usually includes lots of sequential steps for preprocesing, feature selection etc. Moreover, hyper-paramter tuning, cross-validation further complicates the process, which, among other things, frequently leads to [Information Leakage](https://machinelearningmastery.com/data-leakage-machine-learning/). An elegant solution to this complexity management is `Pipeline`.
# ### Pipeline
# `scikit-clean` provides a customized `Pipeline` to manage modeling which involves lots of sequential steps, including noise detection and handling. Below is an example of `pipeline`. At the very first step, we introduce some label noise on training set. Some preprocessing like scaling and feature selection comes next. The last two steps are noise detection and handling respectively, these two must always be the last steps.
# +
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.svm import SVC
from sklearn.model_selection import ShuffleSplit, StratifiedKFold
from skclean.handlers import Filter
from skclean.pipeline import Pipeline # Importing from skclean, not sklearn
clf = Pipeline([
('scale', StandardScaler()), # Scale features
('feat_sel', VarianceThreshold(.2)), # Feature selection
('detector', KDN()), # Detect mislabeled samples
('handler', Filter(SVC())), # Filter out likely mislabeled samples and then train a SVM
])
inner_cv = ShuffleSplit(n_splits=5,test_size=.2,random_state=1)
outer_cv = StratifiedKFold(n_splits=5,shuffle=True,random_state=2)
clf_g = GridSearchCV(clf,{'detector__n_neighbors':[2,5,10]},cv=inner_cv)
n_clf_g = make_pipeline(UniformNoise(.3),clf_g) # Create label noise at the very first step
print(cross_val_score(n_clf_g, X, y, cv=outer_cv).mean()) # 5-fold cross validation
# -
# There are two important things to note here. First, don't use the `Pipeline` of `scikit-learn`, import from `skclean.pipeline` instead.
#
# Secondly, a group of noise hanbdlers are iterative: they call the `detect` of noise detectors multiple times (`CLNI`, `IPF` etc). Since they don't exactly follow the sequential noise detection->handling pattern, you must pass the detector in the constructor of those `Handler`s.
# +
from skclean.handlers import CLNI
clf = CLNI(classifier=SVC(), detector=KDN())
# -
# All `Handler` *can* be instantiated this way, but this is a *must* for iterative ones. (Use `iterative` attribute to check.)
# ### Noise Simulation
#
#
# Remember that as a library written primarily for researchers, you're expected to have access to "true" or "clean" labels, and then introduce noise to training data by flipping those true labels. `scikit-clean` provides several commonly used noise simulators- take a look at [this example](./Noise%20SImulators.ipynb) to understand their differences. Here we mainly focus on how to use them.
#
# Perhaps the most important thing to remember is that noise simulation should usually be the very first thing you do to your training data. In code below, `GridSearchCV` is creating a validation set *before* introducing noise and using clean labels for inner loop, leading to information leakage. This is probably NOT what you want.
clf = Pipeline([
('simulate_noise', UniformNoise(.3)), # Create label noise at first step
('scale', StandardScaler()), # Scale features
('feat_sel', VarianceThreshold(.2)), # Feature selection
('detector', KDN()), # Detect mislabeled samples
('handler', Filter(SVC())), # Filter out likely mislabeled samples and then train a SVM
])
clf_g = GridSearchCV(clf,{'detector__n_neighbors':[2,5,10]},cv=inner_cv)
print(cross_val_score(clf_g, X, y, cv=outer_cv).mean()) # 5-fold cross validation
# You can use noise simulators outside `Pipeline`, all `NoiseSimulator` classes are simple wrapper around functions. `UniformNoise` is a wrapper of `flip_labels_uniform`, as the first example of this document shows.
# ### Datasets & Performance Evaluation
# Unlike deep learning datasets which tends to be massive in size, tabular datasets are usually lot smaller. Any new algorithm is therefore compared using multiple datasets against baselines. The `skclean.utils` module provides two important functions to help researchers in these tasks:
#
# 1. `load_data`: to load several small to medium-sized preprocessed datasets on memory.
#
# 2. `compare`: These function takes several algorithms and datasets, and outputs the performances in a csv file. It supports automatic resumption of partially computed results, specially helpful for comparing long running, computationally expensive methods on big datasets.
#
# Take a look at [this notebook](./Evaluating%20Robust%20Methods.ipynb) to see how they are used.
| doc/examples/Introduction to Scikit-clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="zmRk-98sXUr4"
# #Importing the dataset from my google drive and converting it in numpy array.
# + id="RFW3zSwDgRyO"
import h5py
import numpy as np
# loading ecg data
with h5py.File('/content/drive/My Drive/data/ecg_tracings.hdf5', "r") as f:
x = np.array(f['tracings'])
# + [markdown] id="F-WSl_1vXSe3"
# #Shape of the numpy array, 827 patients, 12 lead sensors used to take readings, at 400Hz for 10 seconds.
# + id="bHBihJ_QvmYX" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636782273790, "user_tz": -330, "elapsed": 19, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="1ab4aac2-ffcc-4b02-ab66-176b4750bf2c"
s=np.shape(x)
print(s)
# + [markdown] id="VMPGTkYaYi-H"
# #Importing annotations(labels), labelled by three experiends Cardiologists and 2 other medical experts.
# + id="Pwj9OsPcvm3z"
import pandas as pd
age_gender_attr= pd.read_csv('/content/drive/My Drive/data/attributes.csv')
# annotation
cardi_1= pd.read_csv('/content/drive/My Drive/data/annotations/cardiologist1.csv')
cardi_2= pd.read_csv('/content/drive/My Drive/data/annotations/cardiologist2.csv')
cardi_student4= pd.read_csv('/content/drive/My Drive/data/annotations/cardiology_residents.csv')
cardi_student3= pd.read_csv('/content/drive/My Drive/data/annotations/emergency_residents.csv')
cardi_gold= pd.read_csv('/content/drive/My Drive/data/annotations/gold_standard.csv')
cardi_student5= pd.read_csv('/content/drive/My Drive/data/annotations/medical_students.csv')
# + [markdown] id="580bx09TZBek"
# #Printing the shape of label's file. Annotations are given for six cardiac diseases if any, otherwise patient is considered as normal.
# + id="3d7d08qlvngu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636782282379, "user_tz": -330, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="c0328db9-2244-444f-8283-67bba522d21d"
s1= age_gender_attr.shape
s2= cardi_1.shape
s3= cardi_2.shape
s4= cardi_student4.shape
s5= cardi_student3.shape
s6= cardi_gold.shape
s7= cardi_student5.shape
print("age_gender_attr- shape: ", s1)
print("annotations by cardiologist1- shape: ",s2)
print("annotations by cardiologist2- shape: ",s3)
print("annotations by cardiology student 4th year- shape: ",s4)
print("annotations by cardiology student 3rd year- shape: ",s5)
print("annotations by experienced cardiologist- shape: ",s6)
print("annotations by cardiology student 5th year- shape: ",s7)
# + [markdown] id="eaoMHgosarUg"
# #Ploting the QRS complex(ECG signal), age and gender of a random patient.
# + id="98jUalJHvn3k" colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"status": "ok", "timestamp": 1636782290506, "user_tz": -330, "elapsed": 903, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="7b249ecc-5596-4503-dc67-09c20f738ec0"
import matplotlib.pyplot as plt
import random
patient_id= random.randrange(827)
p_age_gen=age_gender_attr.loc[patient_id]
ecg_r= x[patient_id][0:1200,0]
ecg_time= np.arange(0, 3, 0.0025)
plt.plot(ecg_time, ecg_r, linewidth=0.5, linestyle='solid', color='b')
plt.title('ECG plot of patient: '+str(patient_id))
plt.xlabel('time in sec')
plt.ylabel('reading on channel0- DI at scale 1e-4V')
plt.legend([str("gen:"+str(p_age_gen.sex)+" age:"+str(p_age_gen.age)),'channel:0'], loc = 0)
plt.grid(True)
plt.show()
# + [markdown] id="RWYEO2yIcuIY"
# # age and gender of patients
# + id="GzOcf8eSvoOr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636782692887, "user_tz": -330, "elapsed": 775, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="677ae70c-2109-47a4-9cbd-ce4961389489"
age_gender_attr.head
# + [markdown] id="g2NFEVHjb0Os"
# # x contains input data and y1, y2 and y_gold are labels annotated by three experienced Cardiologist, when y1 and y2 have a conflict
# # then y_gold label will be taken
# + colab={"base_uri": "https://localhost:8080/"} id="LGSP9qA32upJ" executionInfo={"status": "ok", "timestamp": 1636783033375, "user_tz": -330, "elapsed": 757, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="91d2fb4d-b70c-4851-ac33-282434a244ba"
y1= cardi_1.to_numpy()
y2= cardi_2.to_numpy()
y_gold= cardi_gold.to_numpy()
for i in range(y1.shape[0]):
f1=0
f2=0
for j in range(6):
if y1[i][j]==1 and f1==0:
f1=1
else:
y1[i][j]=0
if y2[i][j]==1 and f2==0:
f2=1
else:
y2[i][j]=0
print(y1.shape)
print(y2.shape)
print(y_gold.shape)
# + [markdown] id="v0d0yFDPc03d"
# # Taking all three annotations by experienced Cardiologists into consideration and resolving the conflicts, if any, by giving the priority to the Gold standard as sudgested in documentation.
# + id="XNxNXlMmvokc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783225213, "user_tz": -330, "elapsed": 642, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="4aa49218-d89d-4f3e-8e5f-16f4159962e8"
y=[]
for i in range(y1.shape[0]):
a= y1[i].argmax()
b= y2[i].argmax()
if a != b:
y.append(y_gold[i][:])
else:
y.append(y1[i][:])
# adding one more column for normal patient, who does not have any abnormality
normal= []
y= np.array(y,dtype=object)
for i in range(y.shape[0]):
t=np.sum(y[i])
if t==0:
normal.append([1])
else:
normal.append([0])
normal= np.array(normal)
y= np.append(y, normal, axis=1)
print(x.shape)
print(y.shape)
# + [markdown] id="AYSqoFGQdhRm"
# # Disease wise patient's count before resampling the data
# + id="2mb1JDhbvpiT" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783244083, "user_tz": -330, "elapsed": 647, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="7e11aac2-8756-407d-cada-41da6d6bd882"
abnormalities = ['1dAVb', 'RBBB', 'LBBB', 'SB', 'AF', 'ST','normal']
print(abnormalities)
y_count= np.sum(y,axis=0)
print(y_count)
# + [markdown] id="k2KaRq2beHF8"
# # Breaking the 10 sec data in 4 sec window (It will decrease complexity and increase examples.
# + id="rXVxfIUsvqAG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783254629, "user_tz": -330, "elapsed": 1321, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="8666466b-f8b8-49fe-aed5-866b7cea92c3"
x_new=[]
y_new=[]
temp=663
for patient_id in range(827):
if y[patient_id][6]==1 and temp>=0: #skip reading for normal patients
temp-=1
continue
for channel in range(12):
s=0 #starting from 47 because 0-46 is padding bits
for j in range(7): # 5 samples from each patient's 10 sec reading
p = np.zeros(1600, dtype = float)
p = x[patient_id][s:s+1600,channel]
x_new.append(p)
y_new.append(y[patient_id][:])
s=s+400
x_new= np.array(x_new,dtype=object)
y_new= np.array(y_new, dtype=object)
print(x_new.shape)
print(y_new.shape)
# + [markdown] id="tx1084VtezE-"
# # Disease wise patient's count, after resampling the data.
# + id="5KU9-Y_DvqTi" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783265207, "user_tz": -330, "elapsed": 999, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="b450bc7c-75f3-4e1e-a0fa-6ac34fc82aee"
print("after resampling the data")
abnormalities = ['1dAVb', 'RBBB', 'LBBB', 'SB', 'AF', 'ST','normal']
'''
1dAVb= 1st degree AV block
RBBB= Right bundle branch block
LBBB= Left bundle branch block
SB= Sinus Bradycardia
AF= Atrial Fibbrilation
ST= Sinus Tachycardia
normal= no abnormality
'''
print(abnormalities)
y_count= np.sum(y_new,axis=0)
print(y_count)
# + [markdown] id="zuxk83ByfEUR"
# # changing the shape of label, by giving it's class value in range [0,6], [0:'1dAVb', 1:'RBBB', 2:'LBBB', 3:'SB', 4:'AF', 5:'ST',6:'normal']
# + id="G1c2IWzUvqiV" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783280237, "user_tz": -330, "elapsed": 651, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="60ddbc1e-a698-4487-bdf7-cf0f68cd5d03"
y= np.zeros(y_new.shape[0], dtype = float)
for i in range(y_new.shape[0]):
y[i]= np.argmax(y_new[i][:])
y= np.array(y,dtype=object)
print(x_new.shape)
print(y.shape)
# + [markdown] id="PtPD02Ecylr6"
# # spliting the data in testing(20%) and training(80%) dataset
# + id="icpzilUYvqxz" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783298247, "user_tz": -330, "elapsed": 1704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="fe0d0615-02e3-40de-ddf3-b62be2ed9b36"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x_new, y, test_size = 0.2, random_state = 1)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# + [markdown] id="_XTFUT5gzL7M"
# # Normalising the ECG readings, normalization is to change the values of numeric columns in the dataset to use a common scale, without distorting differences in the ranges of values or losing information.
# + id="J0bP6e4mvrEo"
import tensorflow as tf
# normalising the data values
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
#print(X_train.mean(axis=0))
#print(X_train.std(axis=0))
# + [markdown] id="qQ0muR9szyfN"
# # Reshaping the input to feed in CNN model.
# + id="6ULvF84swkLc" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636783315143, "user_tz": -330, "elapsed": 677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="a7d2aadf-31e4-4622-c69a-1e5ddbc4514e"
print(X_train.shape)
print(X_test.shape)
X_train= np.reshape(X_train,(10953,1600,1))
X_test= np.reshape(X_test,(2739,1600,1))
y_train= np.reshape(y_train,(10953,1))
y_test= np.reshape(y_test,(2739,1))
y_train= np.asarray(y_train).astype(np.float64)
y_test= np.asarray(y_test).astype(np.float64)
print("dimensions after reshaping")
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# + [markdown] id="WMI2wVOz0GEo"
# # Defining the CNN model
# + id="XM2AEWWKwkod"
from keras.models import Sequential
from keras.layers import Dense, Conv1D, Flatten, MaxPooling1D
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from keras.layers import Dropout
from numpy import unique
model = Sequential()
model.add(Conv1D(32, 16, activation="relu", input_shape=(1600,1)))
model.add(Conv1D(64, 16, activation="relu"))
model.add(Dropout(0.3))
model.add(MaxPooling1D())
model.add(Conv1D(64, 32, activation="relu"))
model.add(Dropout(0.5))
model.add(MaxPooling1D())
model.add(Conv1D(128, 32, activation="relu"))
model.add(Dropout(0.5))
model.add(MaxPooling1D())
model.add(Flatten())
model.add(Dense(128, activation="relu"))
model.add(Dropout(0.3))
model.add(Dense(256, activation="relu"))
model.add(Dropout(0.5))
model.add(Dense(7, activation = 'softmax'))
# + [markdown] id="i2aR0ZLA0VE6"
# # Compiling the model
# + id="tpWfrreuwlB_"
model.compile(loss = 'sparse_categorical_crossentropy',
optimizer = "adam",
metrics = ['accuracy'])
# + [markdown] id="gUyQrjZd0iID"
# #Printing the model summary
# + id="Pglc4OZrwlXJ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784052093, "user_tz": -330, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="87bdfb63-cb5b-4f43-a8ff-26d0a3104d82"
model.summary()
# + [markdown] id="dx3H43SN0xsR"
# #Defining a callback for early stopping, to avoid overfitting.
# + id="4shALEHCzuNQ"
callback = tf.keras.callbacks.EarlyStopping(monitor='accuracy', patience=4)
# + [markdown] id="6DpKG5O01MJT"
# # Training the model on ~11,000 inputs for 25 epochs.
# + id="TBU0lzUsw2Fn" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784405280, "user_tz": -330, "elapsed": 344998, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="265d2aad-b82a-40dc-8d21-42347c178d8f"
history=model.fit(X_train, y_train, batch_size=32,epochs=25, verbose=1, validation_data=(X_test, y_test), callbacks=[callback])
# + id="xcntl9NI0dAR"
# + id="k8Na-4bUw2qH" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784428813, "user_tz": -330, "elapsed": 544, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="b9ccc74a-8b50-405a-fe19-61b90d8fea6f"
print(history.history['accuracy'])
# + [markdown] id="59Wbfabg1uu8"
# # Printing test loss and test accuracy.
# + id="Uw9OoMV2w3CZ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784470263, "user_tz": -330, "elapsed": 1981, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="e9b17c78-6a86-44c0-d762-9d9bf8a28fde"
test_loss, test_acc = model.evaluate(X_test, y_test, verbose=2)
print("test accuracy is: ",test_acc*100,"%")
# + [markdown] id="M4ji6B8_13zp"
# #Predicting the diseases for test data using our trained model.
# + id="2AWt9pPlw3pO"
y_pred = model.predict(X_test)
# + [markdown] id="rTK5MyZN2IFv"
# #Printing the F1-score, Precision and Recall of our model.
# Precision: Precision quantifies the number of positive class predictions that actually belong to the positive class.
# Recall: Recall quantifies the number of positive class predictions made out of all positive examples in the dataset.
# + id="5UEF8mNrw4Nu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784497719, "user_tz": -330, "elapsed": 650, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="8acf0362-ec28-49c8-f416-3455b0e61079"
from sklearn.metrics import precision_recall_fscore_support
pre_rec_F1= precision_recall_fscore_support(y_test, y_pred.argmax(axis=1), average='macro')
print("Precision, recall, f1-score: ",pre_rec_F1)
pre_rec_F1_= precision_recall_fscore_support(y_test, y_pred.argmax(axis=1), average='micro')
print("Precision, recall, f1-score: ",pre_rec_F1_)
# + [markdown] id="Ovxzeu632k-W"
# # Defining our Confusion Matrix.
# + id="2uARVlRtxVcR"
from sklearn.metrics import confusion_matrix
#cm = confusion_matrix(y_test, y_pred)
cm= confusion_matrix(y_test, y_pred.argmax(axis=1))
cm_df = pd.DataFrame(cm,
index = ['1dAVb', 'RBBB', 'LBBB', 'SB', 'AF', 'ST','normal'],
columns = ['1dAVb', 'RBBB', 'LBBB', 'SB', 'AF', 'ST','normal'])
# + [markdown] id="pQplhtXc2zvv"
# #Plotting the confusion matrix
# + id="lF64UKxfxWch" colab={"base_uri": "https://localhost:8080/", "height": 295} executionInfo={"status": "ok", "timestamp": 1636784520908, "user_tz": -330, "elapsed": 799, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="95fc0ffe-20d3-4ac1-c927-7e32b9a5c213"
import seaborn as sns
plt.figure(figsize=(5,4))
sns.heatmap(cm_df, annot=True)
plt.title('Confusion Matrix')
plt.ylabel('Actual Values')
plt.xlabel('Predicted Values')
plt.show()
# + [markdown] id="Whm8Tb9G3E6o"
# #Saving our Model.
# + id="B70j55iixXwG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1636784548323, "user_tz": -330, "elapsed": 3506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GitmS10PcMCviLH3wiZ8LZG0ysTjmbTnBOickdz=s64", "userId": "14863231659108778653"}} outputId="0004b31b-3548-4961-989e-fa6c71327998"
model.save('/content/drive/My Drive/data/model_ecg5_gold')
# + id="40z1iWP_9hZm"
| ECG5_gold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Compressione mediante PCA
# +
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# -
from pylab import subplot,imshow,title,gray,NullLocator
from numpy import linalg
import numpy as np
# +
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
plt.style.use('fivethirtyeight')
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.serif'] = 'Ubuntu'
plt.rcParams['font.monospace'] = 'Ubuntu Mono'
plt.rcParams['font.size'] = 10
plt.rcParams['axes.labelsize'] = 10
plt.rcParams['axes.labelweight'] = 'bold'
plt.rcParams['axes.titlesize'] = 10
plt.rcParams['xtick.labelsize'] = 8
plt.rcParams['ytick.labelsize'] = 8
plt.rcParams['legend.fontsize'] = 10
plt.rcParams['figure.titlesize'] = 12
plt.rcParams['image.cmap'] = 'jet'
plt.rcParams['image.interpolation'] = 'none'
plt.rcParams['figure.figsize'] = (16, 8)
plt.rcParams['lines.linewidth'] = 2
plt.rcParams['lines.markersize'] = 8
colors = ['#008fd5', '#fc4f30', '#e5ae38', '#6d904f', '#8b8b8b', '#810f7c',
'#137e6d', '#be0119', '#3b638c', '#af6f09', '#008fd5', '#fc4f30', '#e5ae38', '#6d904f', '#8b8b8b',
'#810f7c', '#137e6d', '#be0119', '#3b638c', '#af6f09']
cmap = mcolors.LinearSegmentedColormap.from_list("", ["#82cafc", "#069af3", "#0485d1", colors[0], colors[8]])
# -
# Funzione che proietta i vettori in $M$ nello spazio generato dai primi numpc autovettori da vec
def pca(M,vec,numpc=0):
# extract eigenvectors corresponding to the first numpc eigenvalues, this is a basis in a spce of dimension numpc
basis = vec[:,range(numpc)]
# projection of the data in the new space of dimension numpc
projections = np.dot(M,basis)
#return eigenvectors and projections
return basis,projections
# Calcolo di autovalori e autovettori della matrice di covarianza associata alle righe di $A$
def get_eigen(A):
# computing eigenvalues and eigenvectors of the covariance matrix
# of the set of vectors corresponding to rows of A
# compute mean for each column (feature)
means = np.mean(A,axis=0)
# subtract the mean (along columns): each feature has now zero mean
M = (A-means)
# derive the covariance matrix of the set of vectors corresponding to rows of A
c = np.cov(M, rowvar=0)
# compute the sets of eigenvalues and eigenvectors of the covariance matrix
[evals,evects] = np.linalg.eig(c)
# compute indices of eigenvalues sorted in descending order
idx = np.argsort(evals)
# derive indices of eigenvalues sorted in ascending order
idx = idx[::-1]
# sort eigenvectors (columns of evects) according to the sorted eigenvalues
evects = evects[:,idx]
# sort eigenvalues in ascending order
evals = evals[idx]
return evals, evects, M
img =plt.imread('../dataset/austen.jpg') # load an image
bw = np.mean(img,2)
full_pc = bw.shape[0]
val,vec,bw_c=get_eigen(bw)
imshow(bw_c)
# Plot degli autovalori ordinati in modo non crescente
fig = plt.figure()
ax = fig.gca()
ax.plot(val)
ax.grid()
plt.title('Autovalori')
plt.show()
# Plot delle immagini corrispondenti alla proiezione dell'immagine originale nello spazio generato dai primi $i$ autovettori, proiettata nuovamente nello spazio originale
i = 1
dist = []
fig = plt.figure(figsize=(16,12))
fig.patch.set_facecolor('white')
for numpc in [0,1,2,3,4,5]+list(range(10,51,10))+[100]: # 0 10 20 ... full_pc
# compute projections on a space of dimension numpc
coeff, score = pca(bw_c,vec,numpc)
# compute projections back into a space of original dimension
bw_rec = np.dot(score,coeff.T)
# add back feature means
bw_rec = bw_rec+np.mean(bw,axis=0)
# difference in Frobenius norm between original and compressed images
dist.append(linalg.norm(bw-bw_rec,'fro'))
# plot image
ax = subplot(4,3,i,frame_on=False)
ax.xaxis.set_major_locator(NullLocator()) # remove ticks
ax.yaxis.set_major_locator(NullLocator())
i += 1
imshow(bw_rec)
title('Primi '+str(numpc)+' autovalori')
gray()
fig = plt.figure()
fig.patch.set_facecolor('white')
ax = fig.gca()
imshow(bw)
plt.title('Tutti gli autovalori ('+str(full_pc)+')')
plt.show()
# Differenza tra l'immagine originaria e quelle derivanti dalla compressione, misurata mediante norma di Frobenius
fig = plt.figure()
fig.patch.set_facecolor('white')
ax = fig.gca()
ax.plot(dist, c=colors[0])
ax.grid()
plt.title('Distanza in termini di norma di Frobenius')
plt.show()
| codici/PCA_austen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UE4eky2QYcXB"
# If you are interested in graident boosting, here is a good place to start: https://xgboost.readthedocs.io/en/latest/tutorials/model.html
#
# This is a supervised machine learning method.
# + id="fg_LmZjejXi_" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612719882168, "user_tz": 420, "elapsed": 30236, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="c0f9dd30-9c29-48b2-80c8-6354bd3d2457"
# !pip install xgboost --upgrade
# !pip install scikit-learn --upgrade
# + id="qC2ECegCYcXD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612719882852, "user_tz": 420, "elapsed": 30913, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="93d3d6a0-c426-44cf-e133-ffe134c460bc"
# If you have installation questions, please reach out
import pandas as pd # data storage
import xgboost as xgb # graident boosting
import numpy as np # math and stuff
import matplotlib.pyplot as plt # plotting utility
import sklearn # ML and stats
print('XGBoost ver:', xgb.__version__)
print('scikit ver:', sklearn.__version__)
from sklearn.preprocessing import MinMaxScaler, RobustScaler
from sklearn.model_selection import cross_val_score, KFold, train_test_split
from sklearn.utils.class_weight import compute_sample_weight
from sklearn.metrics import accuracy_score, max_error, mean_squared_error
from sklearn.model_selection import GridSearchCV
# + id="WNiabSVfYjTE" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612721262543, "user_tz": 420, "elapsed": 1410601, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="da01cba7-d67a-4fce-f185-088be124e758"
from google.colab import drive
drive.mount('/content/drive')
# + id="Hk1AsPnSYcXQ" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612721264026, "user_tz": 420, "elapsed": 1412080, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="82226351-c1a5-4dfe-db6e-deb3d978da4b"
df = pd.read_csv('drive/My Drive/1_lewis_research/core_to_wl_merge/Merged_dataset_inner_imputed_12_21_2020.csv')
# + [markdown] id="eMQ02l7x0qD_"
# Let's drop some columns, and show off some of the data
# + id="Ws9xTzdwYzgX" colab={"base_uri": "https://localhost:8080/", "height": 469} executionInfo={"status": "ok", "timestamp": 1612721264244, "user_tz": 420, "elapsed": 1412293, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="6906466a-a4ba-4a15-f5e2-5016d31edbcf"
df = df.drop(['Unnamed: 0', 'Unnamed: 0.1', 'LiveTime2','ScanTime2', 'LiveTime1','ScanTime1',
'ref_num', 'API', 'well_name', 'sample_num' ], axis=1)
print(df.columns.values) # printing all column names
df.describe()
# + [markdown] id="pQPTpuJJ1VY1"
# This is the dataset we want to test/train on. Should have features along with what we are trying to predict.
# + id="91nAGubNYcYo" executionInfo={"status": "ok", "timestamp": 1612721264244, "user_tz": 420, "elapsed": 1412291, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
dataset = df[[
'depth_ft', 'CAL', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'PE_smooth',
'gz_pchip_interp' # Trying to predict gz_pchip_interp
]]
# + [markdown] id="T52yBCFGYcYt"
# In the next code block, we will remove the rows without data, and change string NaN's to np.nans
# + id="tUO4fhDeYcYu" colab={"base_uri": "https://localhost:8080/", "height": 257} executionInfo={"status": "ok", "timestamp": 1612721264245, "user_tz": 420, "elapsed": 1412289, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="8cc89d69-0cbe-4a74-a248-14a2e7fce173"
dataset.replace('NaN',np.nan, regex=True, inplace=True)# Should be good already
# dataset = dataset.dropna() # not needed
dataset.head(3)
# + id="MxCYJ2GVYcZA" executionInfo={"status": "ok", "timestamp": 1612721264245, "user_tz": 420, "elapsed": 1412287, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
# Features we will use for prediction
X = dataset[['depth_ft', 'CAL', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID',
'PE_smooth']]
# What we are trying to predict
Y = dataset[['gz_pchip_interp']]
Y_array = np.array(Y.values)
# + [markdown] id="rfNwgw_MYcZJ"
# ## Starting to set up the ML model params
# + [markdown] id="HVx6Fd103Qo-"
# Setting up the test and train sets.
# + id="q_Zq4vu_YcZK" executionInfo={"status": "ok", "timestamp": 1612721264245, "user_tz": 420, "elapsed": 1412286, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}}
seed = 42 # random seed is only used if you want to compare exact answers with friends
test_size = 0.25 # how much data you want to withold, .15 - 0.3 is a good starting point
X_train, X_test, y_train, y_test = train_test_split(X.values, Y_array, test_size=test_size)
# + [markdown] id="-ySy_-2TYcZO"
# ### Let's try some hyperparameter tuning (this takes forever!)
# + [markdown] id="aU6jtQCFYcZO"
# Hyperparameter testing does a grid search to find the best parameters, out of the parameters below. This turned out to be really slow on my laptop. Please skip this!
# + id="R8i9doQmYcZP" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612721265274, "user_tz": 420, "elapsed": 1413312, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="1869d224-f320-4204-f337-d0d0a31de3f8"
xg_reg = xgb.XGBRegressor(objective ='reg:squarederror',
colsample_bytree = 0.9, learning_rate = 0.1,
max_depth = 5, alpha = 10, n_estimators = 60)
xg_reg.fit(X_train,y_train)
preds = xg_reg.predict(X_test)
rmse = mean_squared_error(y_test, preds, squared=False)
print("Root Mean Squared Error: %f" % (rmse))
max = max_error(y_test, preds)
print("Max Error: %f" % (max))
# + [markdown] id="PWEEZlhoV2Bm"
# ### Parameters to search during tuning
# + id="trJgcHlqcIF6" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612723643031, "user_tz": 420, "elapsed": 1565959, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="3a7ff49b-01e8-4fac-a9b5-f713d621d3f9"
parameters = {
'max_depth': range (4, 8, 1),
'n_estimators': range(50, 300, 25),
'colsample_bytree':[ 0.6, 0.8, 0.9, 1],
'learning_rate': [ 0.5, 0.3, 0.2, 0.1]
}
estimator = xgb.XGBRegressor(tree_method='gpu_hist',
gpu_id=0) # uses the GPU
# Grid Search
grid_search = GridSearchCV(
estimator=estimator,
param_grid=parameters,
n_jobs = 8,
cv = 5,
verbose = 2
)
grid_search.fit(X_train, y_train)
# + [markdown] id="_olH3GBuYcZf"
# Now plug in the hyperparameters into the training model.
# + id="F_AVSe-pYcZg" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1612723646893, "user_tz": 420, "elapsed": 3875, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="d5e2a9dc-6566-498d-989c-250ff479d3cc"
model1 = xgb.XGBRegressor(n_estimators=grid_search.best_estimator_.n_estimators,
max_depth = grid_search.best_estimator_.max_depth,
learning_rate=grid_search.best_estimator_.learning_rate,
colsample_bytree=grid_search.best_estimator_.colsample_bytree)
model1.fit(X_train, y_train)
preds = model1.predict(X_test)
rmse2 = mean_squared_error(y_test, preds, squared=False)
print("Mean Squared Error: %f" % (rmse2))
max1 = np.sqrt(max_error(y_test, preds))
print("Max Error: %f" % (max1))
# + [markdown] id="rlRyKrsfcKQJ"
# Let's plot the error, we are looking for low and centered around 0.
# + colab={"base_uri": "https://localhost:8080/", "height": 244} id="4sNv4HnBr80H" executionInfo={"status": "ok", "timestamp": 1612723647097, "user_tz": 420, "elapsed": 209, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="c0f314a2-9370-437d-f907-61bcf321c5f8"
error = preds - y_test.T
plt.figure(figsize=(6,3))
plt.hist(error[0], bins=25)
plt.xlabel('Prediction Error Grainsize, (phi)')
plt.xlim((-4,4))
# + id="PAX4Se0cqCsh" colab={"base_uri": "https://localhost:8080/", "height": 298} executionInfo={"status": "ok", "timestamp": 1612723647870, "user_tz": 420, "elapsed": 776, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04445590536399793096"}} outputId="559319be-e005-485c-d2ec-2bef694cd338"
sorted_idx = model1.feature_importances_.argsort()
plt.barh(X.columns[sorted_idx], model1.feature_importances_[sorted_idx])
plt.xlabel("Xgboost Feature Importance")
# + [markdown] id="QbaiGl8VXC8K"
# *fin*
| xgb/XGB_regression_grainsize_no_GR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from keras import layers
from keras.models import Model, Sequential
from keras import backend as K
from sklearn.metrics import mean_squared_error
from skimage.measure import compare_ssim as SSIM
import numpy as np
from sklearn.model_selection import train_test_split
from keras.optimizers import Adam
_optimizer = Adam(0.0002, 0.5)
from keras import losses
# from tensorflow.keras import layers
# from tensorflow.keras.models import Model
import cv2
import matplotlib.pyplot as plt
from IPython import display
from tqdm import tqdm
# -
def load_imgs(path, number, train_type):
result=np.empty((number, 48, 48, 3), dtype="float64")
for i in range(number):
I = cv2.imread(path + "{:04}_{}.jpeg".format(i+1, train_type))
result[i, :, :, :] = I
return result/result.max()
# +
'''load images, parse test/validation set'''
dataNum = 4000
dataPath = "mf_testcase/"
x1 = load_imgs(dataPath, dataNum, 1)
x2 = load_imgs(dataPath, dataNum, 2)
y = load_imgs("testcase_pro/", dataNum, 0)
y = y[:, 8:-8, 8:-8, :]
mask1 = load_imgs(dataPath, dataNum, 4)
mask1 = mask1[:, 8:-8, 8:-8, :1]
mask2 = 1-mask1
mask = np.concatenate((mask1, mask2), axis = 3)
x_train1, x_test1, x_train2, x_test2, y_train, y_test, mask_train, mask_test = train_test_split(
x1, x2, y, mask, test_size=0.25)
# -
# input size: Nonex48x48x3
# output size: Nonex36x36x16
def pre_convblock(x):
y = layers.Conv2D(filters = 16, kernel_size = (5, 5), padding = "valid", activation = "relu")(x)
y = layers.Conv2D(filters = 32, kernel_size = (5, 5), padding = "valid", activation = "relu")(y)
y = layers.Conv2D(filters = 16, kernel_size = (5, 5), padding = "valid", activation = "relu")(y)
return y
# tensor format: [batch, in_height, in_width, in_channels]
# input size: Nonex36x36x32 (cat: 16+16)
# output size: Nonex32x32x2
# softmax is applied along the channel axis.
def post_convblock(x):
y = layers.Conv2D(filters = 64, kernel_size=(3, 3), padding = "valid", activation = "relu")(x)
y = layers.Conv2D(filters = 32, kernel_size=(1, 1), padding = "valid", activation = "relu")(y)
y = layers.Conv2D(filters = 2, kernel_size=(3, 3), padding = "valid", activation = 'tanh')(y)
# y = layers.Conv2D(filters = 2, kernel_size=(3, 3), padding = "valid")(y)
y = layers.Softmax(axis = -1)(y)
return y
# input: out-of-focus image block A & block B, which denote the same area of the whole picture.
# each: 48x48x3
# output:
def fusionnet(inTensor1, inTensor2):
out1 = pre_convblock(inTensor1)
out2 = pre_convblock(inTensor2)
x = layers.Concatenate(axis = -1)([out1, out2])
y = post_convblock(x)
return y
def fusionnetPos(y):
y, inTensor1, inTensor2 = y
# crop the input images to the same size as network output.
inCrop1 = layers.Cropping2D(cropping=((8, 8), (8, 8)))(inTensor1)
inCrop2 = layers.Cropping2D(cropping=((8, 8), (8, 8)))(inTensor2)
# y1, y2 = tf.split(y, [1, 1], axis = 3)
# extend y1&y2 dimension to 3, consistant to color channels
y1 = y[:, :, :, :1]
y2 = y[:, :, :, 1:]
y1 = K.tile(y1, [1, 1, 1, 3])
y2 = K.tile(y2, [1, 1, 1, 3])
y1 = layers.Multiply()([inCrop1, y1])
y2 = layers.Multiply()([inCrop2, y2])
y = layers.Add()([y1, y2])
return y
img1 = layers.Input(shape=(48, 48, 3))
img2 = layers.Input(shape=(48, 48, 3))
intermed = fusionnet(img1, img2) # intermed: mask layer
pred = layers.Lambda(fusionnetPos)([intermed, img1, img2])
'''2 outputs: 'pred' for GAN loss and 'intermed' for mask loss'''
generator = Model(inputs = [img1, img2], outputs = [pred, intermed])
# generator.summary()
# +
'''tv_loss: designed constraint on mask. WORKS NOT WELL.'''
def tv_loss(y_true, y_pred):
#mapping = tf.cast(y_pred > 0.5, y_pred.dtype)
#loss = tf.reduce_mean(tf.image.total_variation(mapping))
# mapping = 0.25 - tf.square(y_pred-0.5) # 0.25-(x-0.5)^2
# sigma = 0.2
# mu = 0.5
# mapping = tf.exp(-0.5*tf.square((y_pred - mu)/sigma))
mapping = tf.square(y_pred)*tf.square(y_pred-1)/(tf.square(y_pred-0.5)+0.25)
loss = tf.reduce_mean(mapping)
return loss
# +
'''train on generator using MSE of pred & mask.'''
# lambda_tv = 1
# generator.compile(loss=[losses.mean_squared_error, tv_loss], loss_weights=[1, 0.001], optimizer= _optimizer)
# generator.fit([x_train1, x_train2], [y_train, mask_train], batch_size=64, epochs=5)
# prediction, a = generator.predict([x1, x2])
# get_layer_output = K.function([generator.layers[0].input, generator.layers[1].input],
# [generator.layers[-2].output])
# layer_output = np.array(get_layer_output([x1, x2])[0])
# imgIdx = 6
# fig=plt.figure(figsize=(12, 12))
# columns = 6
# rows = 1
# fig.add_subplot(rows, columns, 1)
# plt.imshow(x1[imgIdx, 8:-8, 8:-8, :])
# fig.add_subplot(rows, columns, 2)
# plt.imshow(x2[imgIdx, 8:-8, 8:-8, :])
# fig.add_subplot(rows, columns, 3)
# plt.imshow(mask[imgIdx, :, :, 0])
# fig.add_subplot(rows, columns, 4)
# plt.imshow(layer_output[imgIdx, :, :, 0])
# fig.add_subplot(rows, columns, 5)
# plt.imshow(prediction[imgIdx, :, :, :])
# fig.add_subplot(rows, columns, 6)
# plt.imshow(y[imgIdx, :, :, :])
# plt.show()
# # fig.savefig(dataPath+"results/struc_loss{:.2E}.png".format(lambda_tv))
# -
def dis_block(x, _filters, _strides, bn = True):
y = layers.Conv2D(filters = _filters, kernel_size = (5, 5), strides = _strides,
padding='same')(x)
y = layers.LeakyReLU(alpha=0.2)(y)
if bn:
y = layers.BatchNormalization(momentum=0.8)(y)
return y
def disnet(x):
y = dis_block(x, 32, (2, 2), bn = False)
y = dis_block(y, 64, (1, 1))
y = dis_block(y, 64, (2, 2))
y = dis_block(y, 128, (1, 1))
y = dis_block(y, 128, (2, 2))
# y = dis_block(y, 256, (2, 2))
# y = dis_block(y, 256, (2, 2))
y = layers.Flatten()(y)
y = layers.Dense(1, activation='sigmoid')(y)
return y
dis_input = layers.Input(shape=(32, 32, 3))
dis_output = disnet(dis_input)
discriminator = Model(inputs = dis_input, outputs = dis_output)
# discriminator.summary()
# +
# discriminator.compile(loss='mse', optimizer= _optimizer)
# y_fake = generator.predict([x_train1, x_train2])
# dis_input = np.concatenate((y_fake, y_train), axis = 0)
# label = np.append(np.zeros((y_fake.shape[0], )), np.ones((y_train.shape[0], )))
# for e in range(20):
# discriminator.fit(dis_input, label)
# label = discriminator.predict(x_test1[:, 8:-8, 8:-8, :])
# print(np.sum(label < 0.5))
# print(label)
# -
def gannet(x):
img1, img2 = x
pred, intermed = generator([img1, img2])
prob = discriminator(pred)
discriminator.trainable = False
return (prob, intermed)
prob, intermed = gannet([img1, img2])
gan = Model(inputs = [img1, img2], outputs= [prob, intermed])
# gan.summary()
# +
# prediction = gan.predict([x1, x2])
# mask = prediction[1]
# plt.imshow(mask[6, :, :, 0])
# mask_tensor = tf.convert_to_tensor(mask)
# tv = tf.image.total_variation(mask_tensor)
# sess = tf.Session()
# loss = tv.eval(session=sess)
# print(loss[:10]/32/32/2)
# -
def plot_loss(losses):
display.clear_output(wait=True)
display.display(plt.gcf())
plt.figure(figsize=(10,8))
plt.plot(losses["d"], label='discriminitive loss')
plt.plot(losses["g"], label='generative loss')
plt.legend()
plt.show()
epoch = 100
batchSize = 64
losses = {"d":[], "g":[]}
for e in tqdm(range(epoch)):
rand_idx = np.random.randint(0, x_train1.shape[0], size = batchSize)
img_batch1 = x_train1[rand_idx, :, :, :]
img_batch2 = x_train2[rand_idx, :, :, :]
mask_batch = mask_train[rand_idx, :, :, :]
y_batch = y_train[np.random.randint(0, y_train.shape[0], size = batchSize), :, :, :]
img_fake = generator.predict([img_batch1, img_batch2])[0]
img_valid = y_batch
dis_input = np.concatenate((img_fake, img_valid), axis = 0)
label = np.append(np.zeros((batchSize, )), np.ones((batchSize, )))
discriminator.trainable = True
discriminator.compile(loss='binary_crossentropy', optimizer=_optimizer)
d_loss = discriminator.train_on_batch(dis_input, label)
losses["d"].append(d_loss)
gan_label = np.ones((batchSize,))
discriminator.trainable = False
gan.compile(loss = ['binary_crossentropy', tv_loss], loss_weights=[1, 0.1], optimizer=_optimizer)
g_loss = gan.train_on_batch([img_batch1, img_batch2], [gan_label, mask_batch])
losses["g"].append(g_loss[1])
if e % 5 == 4:
plot_loss(losses)
imgIdx = 16
prediction = generator.predict([x1[imgIdx:imgIdx+1, :, :, :], x2[imgIdx:imgIdx+1, :, :, :]])[0]
#ssim = SSIM(y[imgIdx, :, :, :], prediction[0, :, :, :], data_range=prediction[0, :, :, :].max() - prediction[0, :, :, :].min(), multichannel=True)
#print(ssim)
get_layer_output = K.function([generator.layers[0].input, generator.layers[1].input],
[generator.layers[-2].output])
layer_output = np.array(get_layer_output([x1[imgIdx:imgIdx+1, :, :, :], x2[imgIdx:imgIdx+1, :, :, :]])[0])
fig=plt.figure(figsize=(12, 12))
columns = 6
rows = 1
fig.add_subplot(rows, columns, 1)
plt.imshow(x1[imgIdx, 8:-8, 8:-8, :])
fig.add_subplot(rows, columns, 2)
plt.imshow(x2[imgIdx, 8:-8, 8:-8, :])
fig.add_subplot(rows, columns, 3)
plt.imshow(mask[imgIdx, :, :, 0])
fig.add_subplot(rows, columns, 4)
plt.imshow(layer_output[0, :, :, 0])
fig.add_subplot(rows, columns, 5)
plt.imshow(prediction[0, :, :, :])
fig.add_subplot(rows, columns, 6)
plt.imshow(y[imgIdx, :, :, :])
plt.show()
#fig.savefig(dataPath+"results/struc_ep{}_{}.png".format(e,98))
# +
# prediction = generator.predict([x_test1, x_test2])
# get_layer_output = K.function([generator.layers[0].input, generator.layers[1].input],
# [generator.layers[-2].output])
# layer_output = np.array(get_layer_output([x_test1, x_test2])[0])
# +
# np.save('x_test1.npy', x_test1)
# np.save('x_test2.npy', x_test2)
# np.save('y_test.npy', y_test)
# np.save('prediction.npy', prediction[0])
# np.save('layer_output.npy', layer_output)
# np.save('mask_test.npy', mask_test)
# +
# # ssimList = []
# # ada = 999
# for imgIdx in range(1000):
# # ssim = SSIM(y_test[imgIdx, :, :, :], prediction[0][imgIdx, :, :, :], data_range=1, multichannel=True)
# # ssimList.append(ssim)
# # ssim1 = SSIM(y_test[imgIdx, :, :, :], x_test1[imgIdx, 8:-8, 8:-8, :], data_range=1, multichannel=True)
# # ssim2 = SSIM(y_test[imgIdx, :, :, :], x_test2[imgIdx, 8:-8, 8:-8, :], data_range=1, multichannel=True)
# # print(ssim1)
# # print(ssim2)
# # print(ssim)
# # imgIdx = 312
# fig=plt.figure(figsize=(12, 12))
# columns = 6
# rows = 1
# fig.add_subplot(rows, columns, 1)
# plt.imshow(x_test1[imgIdx, 8:-8, 8:-8, :])
# fig.add_subplot(rows, columns, 2)
# plt.imshow(x_test2[imgIdx, 8:-8, 8:-8, :])
# fig.add_subplot(rows, columns, 3)
# plt.imshow(layer_output[imgIdx, :, :, 0])
# fig.add_subplot(rows, columns, 4)
# plt.imshow(prediction[0][imgIdx, :, :, :])
# fig.add_subplot(rows, columns, 5)
# plt.imshow(y_test[imgIdx, :, :, :])
# fig.add_subplot(rows, columns, 6)
# plt.imshow(mask_test[imgIdx, :, :, 0])
# plt.show()
# fig.savefig(dataPath+"results/{}.png".format(imgIdx))
# +
# plt.hist(ssimList, bins = 20)
# +
# generator.save('generator.h5')
| fusionnet-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 机器学习100天——第二天:简单线性回归
# ## 第一步:数据预处理
# 这里导入我们需要的库,值得注意的是,这里比第一天多了一个matplotlib.pyplot,matplotlib是python上的一个2D绘图库,
# matplotlib下的模块pyplot是一个有命令样式的函数集合,
# matplotlib.pyplot是为我们对结果进行图像化作准备的。
import sys
sys.path.append('/home/ec2-user/.local/lib/python3.7/site-packages')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# 导入相关数据
dataset = pd.read_csv('../datasets/studentscores.csv')
print(dataset)
# 这里我们需要使用pandas的iloc(区分于loc根据index来索引,iloc利用行号来索引)方法来对数据进行处理,第一个参数为行号,:表示全部行,第二个参数 :1表示截到第1列(也就是取第0列)
X = dataset.iloc[ : , : 1 ].values
Y = dataset.iloc[ : , 1 ].values
print("X:",X)
print("Y:",Y)
# 导入sklearn库的cross_validation类来对数据进行训练集、测试集划分
# +
from sklearn.model_selection import train_test_split
#拆分数据,0.25作为测试集
X_train, X_test, Y_train, Y_test = train_test_split( X, Y, test_size = 1/4, random_state = 0)
# -
# ## 训练线性回归
# +
from sklearn.linear_model import LinearRegression
#使用训练集对模型进行训练
regressor = LinearRegression()
regressor = regressor.fit(X_train, Y_train)
# -
# ## 预测结果
Y_pred = regressor.predict(X_test)
# ## 可视化
# ### 训练集结果可视化
#散点图
plt.scatter(X_train , Y_train, color = 'red')
#线图
plt.plot(X_train , regressor.predict(X_train), 'bo-')
plt.show()
# ### 测试集结果可视化
#散点图
plt.scatter(X_test , Y_test, color = 'red')
#线图
plt.plot(X_test ,Y_pred, 'bo-')
plt.show()
| Code/Day_02_Simple_Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# ## Disclaimer
# This notebook is only working under the versions:
#
# - JuMP 0.19
#
# - MathOptInterface 0.8.4
#
# - GLPK 0.9.1
# **Description**: Shows how to solve [Sudoku](http://en.wikipedia.org/wiki/Sudoku) puzzles using integer programming and JuMP.
#
# **Author**: <NAME>
#
# **License**: <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-sa/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/">Creative Commons Attribution-ShareAlike 4.0 International License</a>.
# # Solving Sudoku with JuMP
#
# <img style="width: 200px; height: auto" src="http://upload.wikimedia.org/wikipedia/commons/f/ff/Sudoku-by-L2G-20050714.svg">
# <p style="text-align: center"><i>A partially solved Sudoku puzzle</i></p>
#
# <a href="http://en.wikipedia.org/wiki/Sudoku">Sudoku</a> is a popular number puzzle. The goal is to place the digits 1,...,9 on a nine-by-nine grid, with some of the digits already filled in. Your solution must satisfy the following rules:
#
# * The numbers 1 to 9 must appear in each 3x3 square
# * The numbers 1 to 9 must appear in each row
# * The numbers 1 to 9 must appear in each column
#
# This isn't an optimization problem, its actually a *feasibility* problem: we wish to find a feasible solution that satsifies these rules. You can think of it as an optimization problem with an objective of 0.
#
# We can model this problem using 0-1 integer programming: a problem where all the decision variables are binary. We'll use JuMP to create the model, and then we can solve it with any integer programming solver.
# +
# Import additional packages
import Pkg;
Pkg.add("RDatasets")
Pkg.add("JuMP")
Pkg.add("MathOptInterface")
Pkg.add("GLPK")
# Load packages
using MathOptInterface
using RDatasets
using JuMP
using GLPK
# shortcuts
const MOI = MathOptInterface
const MOIU = MathOptInterface.Utilities
# -
# We will define a binary variable (a variable that is either 0 or 1) for each possible number in each possible cell. The meaning of each variable is as follows:
#
# x[i,j,k] = 1 if and only if cell (i,j) has number k
#
# where `i` is the row and `j` is the column.
# +
# Create a model
sudoku = Model(with_optimizer(GLPK.Optimizer))
# Create our variables
@variable(sudoku, x[i=1:9, j=1:9, k=1:9], Bin)
# -
# Now we can begin to add our constraints. We'll actually start with something obvious to us as humans, but what we need to enforce: that there can be only one number per cell.
for i = 1:9, j = 1:9 # Each row and each column
# Sum across all the possible digits
# One and only one of the digits can be in this cell,
# so the sum must be equal to one
@constraint(sudoku, sum(x[i,j,k] for k in 1:9) == 1)
end
# Next we'll add the constraints for the rows and the columns. These constraints are all very similar, so much so that we can actually add them at the same time.
for ind = 1:9 # Each row, OR each column
for k = 1:9 # Each digit
# Sum across columns (j) - row constraint
@constraint(sudoku, sum(x[ind,j,k] for j in 1:9) == 1)
# Sum across rows (i) - column constraint
@constraint(sudoku, sum(x[i,ind,k] for i in 1:9) == 1)
end
end
# Finally, we have the to enforce the constraint that each digit appears once in each of the nine 3x3 sub-grids. Our strategy will be to index over the top-left corners of each 3x3 square with `for` loops, then sum over the squares.
for i = 1:3:7, j = 1:3:7, k = 1:9
# i is the top left row, j is the top left column
# We'll sum from i to i+2, e.g. i=4, r=4, 5, 6
@constraint(sudoku, sum(x[r,c,k] for r in i:i+2, c in j:j+2) == 1)
end
# The final step is to add the initial solution as a set of constraints. We'll solve the problem that is in the picture at the start of the notebook. We'll put a `0` if there is no digit in that location.
# The given digits
init_sol = [ 5 3 0 0 7 0 0 0 0;
6 0 0 1 9 5 0 0 0;
0 9 8 0 0 0 0 6 0;
8 0 0 0 6 0 0 0 3;
4 0 0 8 0 3 0 0 1;
7 0 0 0 2 0 0 0 6;
0 6 0 0 0 0 2 8 0;
0 0 0 4 1 9 0 0 5;
0 0 0 0 8 0 0 7 9]
for i = 1:9, j = 1:9
# If the space isn't empty
if init_sol[i,j] != 0
# Then the corresponding variable for that digit
# and location must be 1
@constraint(sudoku, x[i,j,init_sol[i,j]] == 1)
end
end
# solve problem
optimize!(sudoku)
# test if optimization worked out properly
@show has_values(sudoku)
@show termination_status(sudoku) == MOI.OPTIMAL
@show primal_status(sudoku) == MOI.FEASIBLE_POINT
# To display the solution, we need to look for the values of `x[i,j,k]` that are 1.
# Extract the values of x
x_val = value.(x)
# Create a matrix to store the solution
sol = zeros(Int,9,9) # 9x9 matrix of integers
for i in 1:9, j in 1:9, k in 1:9
# Integer programs are solved as a series of linear programs
# so the values might not be precisely 0 and 1. We can just
# round them to the nearest integer to make it easier
if round(Int,x_val[i,j,k]) == 1
sol[i,j] = k
end
end
# Display the solution
sol
# Which is the correct solution:
# <img style="width: 200px; height: auto" src="http://upload.wikimedia.org/wikipedia/commons/3/31/Sudoku-by-L2G-20050714_solution.svg">
# <p style="text-align: center"><i>A completed Sudoku puzzle</i></p>
| julia_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot all slopes scenarios
# Stupid notebook to plot all slopes in one pretty chart
# %matplotlib widget
import numpy as np
import math
import bezier
import cmocean.cm as cmo
import matplotlib.pyplot as plt
import matplotlib.patheffects as PathEffects
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter, AutoMinorLocator)
def slopeFunction(slope_radians, length):
return math.tan(slope_radians) * length
# +
# TODO improvement: Define whole channel dimensions in gridsteps!
channel = {
"width": 400, # [m] has to be at least x gridstep wide
"slope": 1.00, # [deg]
"length": 15000, # [m] has to be multiple of y gridstep
"depth": 150, # [m]
"basinslope": 0.2, # [m] the 'basin' slope
}
grid = {
"length": 36000, # y [m] has to be multiple of y gridstep
"width": 26000, # x [m] has to be multiple of x gridstep
"x_gridstep": 200, # [m]
"y_gridstep": 200, # [m]
"dims": [],
}
# in the y direction of the grid that is
ylist = np.array([i for i in range(0, grid['length'] + grid['y_gridstep'], grid['y_gridstep'])]) + 100 # + 100 is default start y in REFGRID
# -
# initial_depth = 150
basinSlope = 0.2
# +
break_point_index = int(channel['length']/grid['y_gridstep'])
smoothen_over = 10
# reduced version of that in JulesD3D
def makeCrossSection(channelSlope, basinSlope, initial_depth=150):
channel_slope_radians = math.radians(channelSlope)
channel_slope_range = range(0, channel['length'] + grid['y_gridstep'], grid['y_gridstep'])
# channel slope depths
channel_slope_list = np.array([slopeFunction(channel_slope_radians, i) for i in channel_slope_range])
channel_slope_last_depth = channel_slope_list[-1] + channel['depth']
basin_slope_radians = math.radians(basinSlope)
basin_length = grid['length'] - channel['length']
basin_list_range = range(0, basin_length, grid['y_gridstep'])
basin_list = np.array([slopeFunction(basin_slope_radians, i) for i in basin_list_range]) + channel_slope_last_depth
channel_slope_list += channel['depth']
depth_list = np.concatenate((channel_slope_list, basin_list), axis=0) + initial_depth
normal_cross_section = [ylist, depth_list] # x, y values for smoothening
# ----smoothen slope break -----
x_cross_section = ylist.copy()
depth_cross_section = depth_list.copy()
# Smooth with bezier curve between these points
start_smooth_index, end_smooth_index = [break_point_index - smoothen_over,\
break_point_index + smoothen_over]
# Prepare section to be smoothed for Bezier
nodes_x = np.array(x_cross_section[start_smooth_index:end_smooth_index])
nodes_y = np.array(depth_cross_section[start_smooth_index:end_smooth_index])
# Feed nodes into bezier instance
nodes = np.array([nodes_x, nodes_y]) # Bezier class wants it like this
curved = bezier.Curve.from_nodes(nodes)
# Get new depth (y-values in bezier funct) from bezier instance
s_vals_channel = np.linspace(0.0, 1.0, 2 * smoothen_over)
smoothened_channel_part = curved.evaluate_multi(s_vals_channel)[1]
fig_q, ax_q = plt.subplots()
fig_q.suptitle('Bathymetry cross-sections (Unsmoothened!)')
ax_q.plot(range(start_smooth_index, end_smooth_index), -smoothened_channel_part)
ax_q.set_xlabel('N (grid number)')
ax_q.set_ylabel('Depth [m]')
ax_q.grid()
smooth_cross_section = depth_cross_section.copy()
smooth_cross_section[start_smooth_index:end_smooth_index] = smoothened_channel_part
return smooth_cross_section
# -
depths085 = makeCrossSection(0.85, basinSlope)
depths100 = makeCrossSection(1, basinSlope)
depths125 = makeCrossSection(1.25, basinSlope)
depths1_25.shape
# + jupyter={"source_hidden": true}
# fig, ax = plt.subplots(figsize=(10, 5))
# fig.suptitle('Slopes of different model scenarios', fontsize=16)
# ax.set_title('Smoothened slope breaks')
# # ax.set_aspect('equal')
# ax.set_xlim((0, 36000))
# ax.set_ylim(700, 300)
# ax.plot(ylist, -depths1_00[:-1])
# arrowprops = dict(facecolor='black', shrink=0.05),
# text_100 = ax.annotate('Slope 1.00°', xy=(10000, 450), xytext=(12000, 400), fontsize=13, arrowprops=dict(facecolor='black', shrink=0.02)) #, arrowprops=arrowprops) #
# text_100.set_path_effects([PathEffects.withStroke(linewidth=1.5, foreground='bisque')])
# ax.plot(ylist, depths1_125[:-1])
# # text_100 = ax.text(10000, -560, "Slope 1.125°", fontsize=13) #, rotation=-8)
# # text_125.set_path_effects([PathEffects.withStroke(linewidth=1.5, foreground='w')])
# text_1125 = ax.annotate('Slope 1.0°', xy=(10000, 450), xytext=(12000, 400), fontsize=13, arrowprops=dict(facecolor='black', shrink=0.02)) #, arrowprops=arrowprops) #
# text_1125.set_path_effects([PathEffects.withStroke(linewidth=1.5, foreground='bisque')])
# ax.plot(ylist, depths1_25[:-1])
# text_125 = ax.text(9000, -600, "Slope 1.25°", fontsize=13) #, rotation=-8)
# text_125.set_path_effects([PathEffects.withStroke(linewidth=1.5, foreground='w')])
# ax.set_xlabel('Length $n$ [m]')
# ax.set_ylabel('Depth [m]')
# # ax.grid()
# -
plt.close('all')
# +
fig, ax = plt.subplots(figsize=(10, 5))
fig.suptitle('Slopes of Different Scenarios', fontsize=16)
# ax.set_title('Smoothened slope breaks')
# ax.set_aspect('equal')
ax.set_xlim((0, 36000))
ax.set_ylim(700, 300)
ax.xaxis.set_minor_locator(MultipleLocator(1000))
ax.yaxis.set_minor_locator(MultipleLocator(10))
ax.plot(ylist, depths100, c="orange", marker='+', markersize=2.8)
ax.plot(ylist, depths125, c='olive', marker='+', markersize=2.8)
arrow_props = dict(width=1, headwidth=8, facecolor='black', shrink=0.02)
text_085 = ax.annotate('Slope 0.85·°', xy=(14500, 510), xytext=(15500, 460), fontsize=13, arrowprops=arrow_props) #, arrowprops=arrowprops) #
text_085.set_path_effects([PathEffects.withStroke(linewidth=0.5, foreground='blueviolet')])
text_100 = ax.annotate('Slope 1.00°', xy=(10000, 470), xytext=(11000, 430), fontsize=13, arrowprops=arrow_props) #, arrowprops=arrowprops) #
text_100.set_path_effects([PathEffects.withStroke(linewidth=0.5, foreground='orange')])
ax.plot(ylist, depths085, c="blueviolet", marker='+', markersize=2.8)
text_125 = ax.annotate('Slope 1.25°', xy=(8000, 475), xytext=(1500, 540), horizontalalignment='left',
fontsize=13, arrowprops=arrow_props)
text_125.set_path_effects([PathEffects.withStroke(linewidth=0.5, foreground='olive')])
ax.set_xlabel('Length $n$ [m]', fontsize=14)
ax.set_ylabel('Depth [m]', fontsize=14)
# ax.grid()
# +
# plt.close("all")
| PlotScenarioSlopes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df = pd.DataFrame(np.random.rand(10,4),
index = 'A B C D E F G H I J'.split(),
columns = "W X Y Z".split())
df.dropna(thresh = 2)
dat = {'cust' :
'1001 1001 1002 1002 1003 1003'.split(),
'custname' :
'UIpath datrob goog chrysler ford gm'.split(),
'profit' : [2005,3245,1245,8765,5463,3547]}
dat
df = pd.DataFrame(dat)
df
dfg = df.groupby('cust')
dfg.mean()
df1 = pd.DataFrame ({'Cust':'101 102 103 104'.split(),
'Sales':
[13456, 45321, 54385,53212],
'Priority':
'CAT0 CAT1 CAT2 CAT3'.split(),
'Prime': 'yes no no yes'.split()},
index = [0,1,2,3])
df1
df2 = pd.DataFrame({'Cust':'101 102 103 104'.split(),
'Sales':
[13456, 45321, 54385,53212],
'Payback':
'CAT4 CAT5 CAT6 CAT7'.split(),
'Imp': 'yes no no yes'.split()},
index = [4,5,6,7])
df2
pd.concat([df1,df2],sort = False)
pd.merge (df1,df2, how = 'inner', on = 'Cust')
df1.join(df2, how = 'outer', on='Sales', lsuffix='_left', rsuffix='_right')
df3 = pd.DataFrame({'Cust':'101 102 103 104'.split(),
'Sales1':
[13456, 45321, 54385,53212],
'Payback1':
'CAT4 CAT5 CAT6 CAT7'.split(),
'Imp1': 'yes no no yes'.split()},
index = [4,5,6,7])
df3
df2
df2.join (df3, how = "inner", on = 'Cust')
| JoinsConcatenationsMerges.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: GeoPySpark
# language: python
# name: gps
# ---
# # Finding the Optimal Location for a New Park
#
# This example notebook will show how to find the next potential location for a new park in San Fransisco. To accomplish this, three factors will be taken into consideration when deciding on a possible spot: existing parks, schools, and Bay Area Regional Transit (BART) stops. By calculating Euclidean Distance for these three factors and then weighing them together, we will be able to produce a visual representation of where is and is not a good location for a new park.
#
# ## Importing the Libraries
# +
import geopyspark as gps
import fiona
from pyspark import SparkContext, StorageLevel
from shapely.geometry import MultiPoint, MultiPolygon, shape
import folium
# -
# ## Setup the SparkContext
conf = gps.geopyspark_conf(appName="park-siting", master="local[*]")
sc = SparkContext.getOrCreate(conf=conf)
# ## Set map display parameters
center = [37.8, -122.2]
zoom_start = 9.5
# ## Download the Geometries as GeoJsons
# !curl -o /tmp/bart.geojson https://s3.amazonaws.com/geopyspark-demo/bayarea/bart.geojson
# !curl -o /tmp/school.geojson https://s3.amazonaws.com/geopyspark-demo/bayarea/school.geojson
# !curl -o /tmp/parks.geojson https://s3.amazonaws.com/geopyspark-demo/bayarea/parks.geojson
# ## Read in the GeoJsons as Shapely Geometries
# +
with fiona.open("/tmp/bart.geojson") as source:
bart_crs = source.crs['init']
bart = MultiPoint([shape(f['geometry']) for f in source])
with fiona.open("/tmp/school.geojson") as source:
schools_crs = source.crs['init']
schools = MultiPoint([shape(f['geometry']) for f in source])
with fiona.open("/tmp/parks.geojson") as source:
parks_crs = source.crs['init']
parks = MultiPolygon([shape(f['geometry']) for f in source])
# -
# ## Calculate Euclidean Distance for Each Geometry
#
# Three new `TiledRasterLayer`s will be produced from the Euclidean Distance calculations for each geometry. All resulting layers will have a `zoom_level` of 12.
# +
bart_layer = gps.euclidean_distance(geometry=bart,
source_crs=bart_crs,
zoom=12)
schools_layer = gps.euclidean_distance(geometry=schools,
source_crs=schools_crs,
zoom=12)
parks_layer = gps.euclidean_distance(geometry=parks,
source_crs=parks_crs,
zoom=12)
# Persists each layer to memory and disk
bart_layer.persist(StorageLevel.MEMORY_AND_DISK)
schools_layer.persist(StorageLevel.MEMORY_AND_DISK)
parks_layer.persist(StorageLevel.MEMORY_AND_DISK)
# -
# ## Weighing the Layers Together
# +
weighted_layer = -1 * bart_layer - schools_layer + 3 * parks_layer
# Persists the weighted layer to memory and disk
weighted_layer.persist(StorageLevel.MEMORY_AND_DISK)
# -
# ## Reprojecting, Pyramiding, and Calculating the Histogram
# The following code may take awhile to complete
reprojected = weighted_layer.tile_to_layout(layout=gps.GlobalLayout(),
target_crs="EPSG:3857")
pyramid = reprojected.pyramid(resample_method=gps.ResampleMethod.AVERAGE)
histogram = pyramid.get_histogram()
# ## Creating the ColorMap
#
# The below code creates a `ColorMap` instance using the `Histogram` from `pyramid` for its `breaks`. For the color, the `matplotlib` color palette, `viridus` will be used.
color_map = gps.ColorMap.build(breaks=histogram,
colors='viridis')
# ## Running the Server
# +
tms = gps.TMS.build(source=pyramid,
display=color_map)
tms.bind('0.0.0.0')
# -
m = folium.Map(tiles='OpenStreetMap', location=center, zoom_start=zoom_start)
folium.TileLayer(tiles=tms.url_pattern, overlay=True, attr='GeoPySpark tiles').add_to(m)
folium.GeoJson(data='/tmp/bart.geojson', name='BART stops').add_to(m)
folium.GeoJson(data='/tmp/parks.geojson', name='Parks').add_to(m)
folium.LayerControl().add_to(m)
m
# ## Cleaning up
tms.unbind()
| notebooks/Park citing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sbn
# %matplotlib inline
df = pd.read_csv("data.csv")
# +
fig, ax = plt.subplots(nrows=3, ncols=2, figsize=(10,12))
for j, atoms in enumerate([3, 10]):
for i, phase in enumerate(["Complex Decharge", "Complex VDW", "Complex Recharge"]):
plt.setp(ax[i][j].spines['bottom'], color='#D2B9D3', lw=3, zorder=-2)
plt.setp(ax[i][j].spines['left'], color='#D2B9D3', lw=3, zorder=-2)
for dire in ['top', 'right']:
ax[i][j].spines[dire].set_color('none')
ax[i][j].xaxis.set_ticks_position('bottom')
ax[i][j].yaxis.set_ticks_position('left')
newdf = df[(df.atoms == atoms)&(df.phase == phase)]
dG = newdf.loc[(newdf.time == 1.0)&(newdf.direction == "forward"), "dG"]
std = newdf.loc[(newdf.time == 1.0)&(newdf.direction == "forward"), "std"]
line0 = ax[i][j].fill_between([0, 1], dG-std, dG+std, color='#D2B9D3', zorder=-5)
forward_newdf = newdf[newdf.direction == "forward"]
for ii, row in forward_newdf.iterrows():
line1 = ax[i][j].plot([row['time']]*2, [row['dG'] - row['std'], row['dG'] + row['std']],
color='#736AFF', ls='-', lw=3, solid_capstyle='round', zorder=1)
line11 = ax[i][j].plot(forward_newdf['time'], forward_newdf['dG'], color='#736AFF', ls='-', lw=3, marker='o', mfc='w', mew=2.5,
mec='#736AFF', ms=12, zorder=2)
reverse_newdf = newdf[newdf.direction == "reverse"]
for ii, row in reverse_newdf.iterrows():
line2 = ax[i][j].plot([row['time']]*2, [row['dG'] - row['std'], row['dG'] + row['std']], color='#C11B17', ls='-', lw=3, solid_capstyle='round', zorder=3)
line22 = ax[i][j].plot(reverse_newdf['time'], reverse_newdf['dG'], color='#C11B17', ls='-', lw=3, marker='o', mfc='w', mew=2.5, mec='#C11B17', ms=12, zorder=4)
ax[i][j].set_xlim(0, 1)
ax[i][j].set_title("%s - %s Atoms" % (phase, atoms), fontsize=20)
ax[i][j].legend((line1[0], line2[0]), (r'$Forward$', r'$Reverse$'), frameon=False, fontsize=15)
ax[i][j].set_xlabel(r'$\mathrm{Fraction\/of\/the\/simulation\/time}$', fontsize=16, color='#151B54')
ax[i][j].set_ylabel(r'$\mathrm{\Delta G(\/k_BT)}$' , fontsize=16, color='#151B54')
ax[i][j].tick_params(axis='x', color='#D2B9D3', labelsize=10)
ax[i][j].tick_params(axis='y', color='#D2B9D3', labelsize=10)
fig.tight_layout()
fig.savefig("Figure.pdf")
# -
fig, ax = plt.subplots(nrows=3, ncols=2, figsize=(10,12))
for j, atoms in enumerate([3, 10]):
for i, phase in enumerate(["Complex Decharge", "Complex VDW", "Complex Recharge"]):
plt.setp(ax[i][j].spines['bottom'], color='black', lw=2, zorder=-2)
plt.setp(ax[i][j].spines['left'], color='black', lw=2, zorder=-2)
for dire in ['top', 'right']:
ax[i][j].spines[dire].set_color('none')
ax[i][j].xaxis.set_ticks_position('bottom')
ax[i][j].yaxis.set_ticks_position('left')
newdf = df[(df.atoms == atoms)&(df.phase == phase)]
dG = newdf.loc[(newdf.time == 1.0)&(newdf.direction == "forward"), "dG"]
std = newdf.loc[(newdf.time == 1.0)&(newdf.direction == "forward"), "std"]
line0 = ax[i][j].fill_between([0, 1], dG-std, dG+std, color='#D2B9D3', zorder=-5)
forward_newdf = newdf[newdf.direction == "forward"]
for ii, row in forward_newdf.iterrows():
line1 = ax[i][j].plot([row['time']]*2, [row['dG'] - row['std'], row['dG'] + row['std']],
color='#736AFF', ls='-', lw=3, solid_capstyle='round', zorder=1)
line11 = ax[i][j].plot(forward_newdf['time'], forward_newdf['dG'], color='#736AFF', ls='-', lw=3, marker='o', mfc='w', mew=2.5,
mec='#736AFF', ms=12, zorder=2)
reverse_newdf = newdf[newdf.direction == "reverse"]
for ii, row in reverse_newdf.iterrows():
line2 = ax[i][j].plot([row['time']]*2, [row['dG'] - row['std'], row['dG'] + row['std']], color='#C11B17', ls='-', lw=3, solid_capstyle='round', zorder=3)
line22 = ax[i][j].plot(reverse_newdf['time'], reverse_newdf['dG'], color='#C11B17', ls='-', lw=3, marker='o', mfc='w', mew=2.5, mec='#C11B17', ms=12, zorder=4)
ax[i][j].set_xlim(0, 1)
#ax[i][j].set_title("%s - %s Atoms" % (phase, atoms), fontsize=20)
ax[i][j].legend((line1[0], line2[0]), (r'$Forward$', r'$Reverse$'), frameon=False, fontsize=15)
ax[i][j].set_xlabel(r'Fraction of the simulation time', fontsize=16, color='black')
ax[i][j].set_ylabel(r'$\mathrm{\Delta G\, [k_BT]}$' , fontsize=16, color='black')
ax[i][j].tick_params(axis='x', color='black', labelsize=14)
ax[i][j].tick_params(axis='y', color='black', labelsize=14)
fig.tight_layout()
| paper/figures/fig10_forward_reverse/fig9_forward_reverse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # XMM-LSS DECAM merging
#
# Both DES and DECaLS provide DECam fluxes which have overlapping coverage. We chose which to use DES preferentially. In this notebook we cross match both catalogues and take the DES fluxes where available, using DECaLS otherwise
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
import datetime
print("This notebook was executed on: \n{}".format(datetime.datetime.now()))
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
import os
import time
from astropy import units as u
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table
import numpy as np
from pymoc import MOC
from herschelhelp_internal.masterlist import merge_catalogues, nb_merge_dist_plot
from herschelhelp_internal.utils import coords_to_hpidx, ebv, gen_help_id, inMoc
# +
TMP_DIR = os.environ.get('TMP_DIR', "./data_tmp")
OUT_DIR = os.environ.get('OUT_DIR', "./data")
SUFFIX = os.environ.get('SUFFIX', time.strftime("_%Y%m%d"))
try:
os.makedirs(OUT_DIR)
except FileExistsError:
pass
# -
# ## I - Reading the prepared pristine catalogues
des = Table.read("{}/DES.fits".format(TMP_DIR))
decals = Table.read("{}/DECaLS.fits".format(TMP_DIR))
# +
for col in des.colnames:
if '_decam_' in col:
des[col].name = col.replace('_decam_', '_des-decam_')
for col in decals.colnames:
if '_decam_' in col:
decals[col].name = col.replace('_decam_', '_decals-decam_')
# -
# ## II - Merging tables
#
# We first merge the optical catalogues and then add the infrared ones: HSC, VHS, VICS82, UKIDSS-LAS, PanSTARRS, SHELA, SpIES.
#
# At every step, we look at the distribution of the distances to the nearest source in the merged catalogue to determine the best crossmatching radius.
# ### DES
master_catalogue = des
master_catalogue['des_ra'].name = 'ra'
master_catalogue['des_dec'].name = 'dec'
# ## Add DECaLS
nb_merge_dist_plot(
SkyCoord(master_catalogue['ra'], master_catalogue['dec']),
SkyCoord(decals['decals_ra'], decals['decals_dec'])
)
# Given the graph above, we use 0.8 arc-second radius
master_catalogue = merge_catalogues(master_catalogue, decals, "decals_ra", "decals_dec", radius=0.8*u.arcsec)
# ### Cleaning
#
# When we merge the catalogues, astropy masks the non-existent values (e.g. when a row comes only from a catalogue and has no counterparts in the other, the columns from the latest are masked for that row). We indicate to use NaN for masked values for floats columns, False for flag columns and -1 for ID columns.
# +
for col in master_catalogue.colnames:
if "m_" in col or "merr_" in col or "f_" in col or "ferr_" in col or "stellarity" in col:
master_catalogue[col].fill_value = np.nan
elif "flag" in col:
master_catalogue[col].fill_value = 0
elif "id" in col:
master_catalogue[col].fill_value = -1
master_catalogue = master_catalogue.filled()
# -
master_catalogue[:10].show_in_notebook()
# ## III - Merging flags and stellarity
#
# Each pristine catalogue contains a flag indicating if the source was associated to a another nearby source that was removed during the cleaning process. We merge these flags in a single one.
# +
flag_cleaned_columns = [column for column in master_catalogue.colnames
if 'flag_cleaned' in column]
flag_column = np.zeros(len(master_catalogue), dtype=bool)
for column in flag_cleaned_columns:
flag_column |= master_catalogue[column]
master_catalogue.add_column(Column(data=flag_column, name="decam_flag_cleaned"))
master_catalogue.remove_columns(flag_cleaned_columns)
# -
# Each pristine catalogue contains a flag indicating the probability of a source being a Gaia object (0: not a Gaia object, 1: possibly, 2: probably, 3: definitely). We merge these flags taking the highest value.
# +
flag_gaia_columns = [column for column in master_catalogue.colnames
if 'flag_gaia' in column]
master_catalogue.add_column(Column(
data=np.max([master_catalogue[column] for column in flag_gaia_columns], axis=0),
name="decam_flag_gaia"
))
master_catalogue.remove_columns(flag_gaia_columns)
# -
# Each prisitine catalogue may contain one or several stellarity columns indicating the probability (0 to 1) of each source being a star. We merge these columns taking the highest value.
# +
stellarity_columns = [column for column in master_catalogue.colnames
if 'stellarity' in column]
master_catalogue.add_column(Column(
data=np.nanmax([master_catalogue[column] for column in stellarity_columns], axis=0),
name="decam_stellarity"
))
master_catalogue.remove_columns(stellarity_columns)
# -
# ## VIII - Cross-identification table
#
# We are producing a table associating to each HELP identifier, the identifiers of the sources in the pristine catalogue. This can be used to easily get additional information from them.
master_catalogue.add_column(Column(data=(np.char.array(master_catalogue['des_id'].astype(str))
+ np.char.array(master_catalogue['decals_id'].astype(str) )),
name="decam_intid"))
# +
id_names = []
for col in master_catalogue.colnames:
if '_id' in col:
id_names += [col]
if '_intid' in col:
id_names += [col]
print(id_names)
# -
# ## VI - Choosing between multiple values for the same filter
#
#
decam_origin = Table()
decam_origin.add_column(master_catalogue['decam_intid'])
decam_stats = Table()
decam_stats.add_column(Column(data=['g','r','i','z','y'], name="Band"))
for col in ["DES", "DECaLS"]:
decam_stats.add_column(Column(data=np.full(5, 0), name="{}".format(col)))
decam_stats.add_column(Column(data=np.full(5, 0), name="use {}".format(col)))
decam_stats.add_column(Column(data=np.full(5, 0), name="{} ap".format(col)))
decam_stats.add_column(Column(data=np.full(5, 0), name="use {} ap".format(col)))
decam_bands = ['g','r','i','z','y'] # Lowercase naming convention (k is Ks)
for band in decam_bands:
if (band == 'i') or (band == 'y'):
master_catalogue["f_des-decam_{}".format(band)].name = "f_decam_{}".format(band)
master_catalogue["ferr_des-decam_{}".format(band)].name = "ferr_decam_{}".format(band)
master_catalogue["m_des-decam_{}".format(band)].name = "m_decam_{}".format(band)
master_catalogue["merr_des-decam_{}".format(band)].name = "merr_decam_{}".format(band)
master_catalogue["f_ap_des-decam_{}".format(band)].name = "f_ap_decam_{}".format(band)
master_catalogue["ferr_ap_des-decam_{}".format(band)].name = "ferr_ap_decam_{}".format(band)
master_catalogue["m_ap_des-decam_{}".format(band)].name = "m_ap_decam_{}".format(band)
master_catalogue["merr_ap_des-decam_{}".format(band)].name = "merr_ap_decam_{}".format(band)
master_catalogue["flag_des-decam_{}".format(band)].name = "flag_decam_{}".format(band)
continue
# DECam total flux
has_des = ~np.isnan(master_catalogue['f_des-decam_' + band])
has_decals = ~np.isnan(master_catalogue['f_decals-decam_' + band])
use_des = has_des
use_decals = has_decals & ~has_des
f_decam = np.full(len(master_catalogue), np.nan)
f_decam[use_des] = master_catalogue['f_des-decam_' + band][use_des]
f_decam[use_decals] = master_catalogue['f_decals-decam_' + band][use_decals]
ferr_decam = np.full(len(master_catalogue), np.nan)
ferr_decam[use_des] = master_catalogue['ferr_des-decam_' + band][use_des]
ferr_decam[use_decals] = master_catalogue['ferr_decals-decam_' + band][use_decals]
m_decam = np.full(len(master_catalogue), np.nan)
m_decam[use_des] = master_catalogue['m_des-decam_' + band][use_des]
m_decam[use_decals] = master_catalogue['m_decals-decam_' + band][use_decals]
merr_decam = np.full(len(master_catalogue), np.nan)
merr_decam[use_des] = master_catalogue['merr_des-decam_' + band][use_des]
merr_decam[use_decals] = master_catalogue['merr_decals-decam_' + band][use_decals]
flag_decam = np.full(len(master_catalogue), False, dtype=bool)
flag_decam[use_des] = master_catalogue['flag_des-decam_' + band][use_des]
flag_decam[use_decals] = master_catalogue['flag_decals-decam_' + band][use_decals]
master_catalogue.add_column(Column(data=f_decam, name="f_decam_" + band))
master_catalogue.add_column(Column(data=ferr_decam, name="ferr_decam_" + band))
master_catalogue.add_column(Column(data=m_decam, name="m_decam_" + band))
master_catalogue.add_column(Column(data=merr_decam, name="merr_decam_" + band))
master_catalogue.add_column(Column(data=flag_decam, name="flag_decam_" + band))
old_des_columns = ['f_des-decam_' + band,
'ferr_des-decam_' + band,
'm_des-decam_' + band,
'merr_des-decam_' + band,
'flag_des-decam_' + band]
old_decals_columns = ['f_decals-decam_' + band,
'ferr_decals-decam_' + band,
'm_decals-decam_' + band,
'merr_decals-decam_' + band,
'flag_decals-decam_' + band]
old_columns = old_des_columns + old_decals_columns
master_catalogue.remove_columns(old_columns)
origin = np.full(len(master_catalogue), ' ', dtype='<U5')
origin[use_des] = "DES"
origin[use_decals] = "DECaLS"
decam_origin.add_column(Column(data=origin, name= 'f_decam_' + band ))
# DECam aperture flux
has_ap_des = ~np.isnan(master_catalogue['f_ap_des-decam_' + band])
has_ap_decals = ~np.isnan(master_catalogue['f_ap_decals-decam_' + band])
use_ap_des = has_ap_des
use_ap_decals = has_ap_decals & ~has_ap_des
f_ap_decam = np.full(len(master_catalogue), np.nan)
f_ap_decam[use_ap_des] = master_catalogue['f_ap_des-decam_' + band][use_ap_des]
f_ap_decam[use_ap_decals] = master_catalogue['f_ap_decals-decam_' + band][use_ap_decals]
ferr_ap_decam = np.full(len(master_catalogue), np.nan)
ferr_ap_decam[use_ap_des] = master_catalogue['ferr_ap_des-decam_' + band][use_ap_des]
ferr_ap_decam[use_ap_decals] = master_catalogue['ferr_ap_decals-decam_' + band][use_ap_decals]
m_ap_decam = np.full(len(master_catalogue), np.nan)
m_ap_decam[use_ap_des] = master_catalogue['m_ap_des-decam_' + band][use_ap_des]
m_ap_decam[use_ap_decals] = master_catalogue['m_ap_decals-decam_' + band][use_ap_decals]
merr_ap_decam = np.full(len(master_catalogue), np.nan)
merr_ap_decam[use_ap_des] = master_catalogue['merr_ap_des-decam_' + band][use_ap_des]
merr_ap_decam[use_ap_decals] = master_catalogue['merr_ap_decals-decam_' + band][use_ap_decals]
master_catalogue.add_column(Column(data=f_ap_decam, name="f_ap_decam_" + band))
master_catalogue.add_column(Column(data=ferr_ap_decam, name="ferr_ap_decam_" + band))
master_catalogue.add_column(Column(data=m_ap_decam, name="m_ap_decam_" + band))
master_catalogue.add_column(Column(data=merr_ap_decam, name="merr_ap_decam_" + band))
old_ap_des_columns = ['f_ap_des-decam_' + band,
'ferr_ap_des-decam_' + band,
'm_ap_des-decam_' + band,
'merr_ap_des-decam_' + band]
old_ap_decals_columns = ['f_ap_decals-decam_' + band,
'ferr_ap_decals-decam_' + band,
'm_ap_decals-decam_' + band,
'merr_ap_decals-decam_' + band]
old_ap_columns = old_ap_des_columns + old_ap_decals_columns
master_catalogue.remove_columns(old_ap_columns)
origin_ap = np.full(len(master_catalogue), ' ', dtype='<U5')
origin_ap[use_ap_des] = "DES"
origin_ap[use_ap_decals] = "DECaLS"
decam_origin.add_column(Column(data=origin_ap, name= 'f_ap_decam_' + band ))
decam_stats['DES'][decam_stats['Band'] == band] = np.sum(has_des)
decam_stats['DECaLS'][decam_stats['Band'] == band] = np.sum(has_decals)
decam_stats['use DES'][decam_stats['Band'] == band] = np.sum(use_des)
decam_stats['use DECaLS'][decam_stats['Band'] == band] = np.sum(use_decals)
decam_stats['DES ap'][decam_stats['Band'] == band] = np.sum(has_ap_des)
decam_stats['DECaLS ap'][decam_stats['Band'] == band] = np.sum(has_ap_decals)
decam_stats['use DES ap'][decam_stats['Band'] == band] = np.sum(use_ap_des)
decam_stats['use DECaLS ap'][decam_stats['Band'] == band] = np.sum(use_ap_decals)
decam_stats.show_in_notebook()
decam_origin.write("{}/xmm-lss_decam_fluxes_origins{}.fits".format(OUT_DIR, SUFFIX), overwrite=True)
# ## IX - Saving the catalogue
master_catalogue.colnames
master_catalogue["ra"].name = "decam_ra"
master_catalogue["dec"].name = "decam_dec"
master_catalogue["flag_merged"].name = "decam_flag_merged"
# +
columns = ["decam_intid", "des_id", "decals_id",
'decam_ra', 'decam_dec', 'decam_flag_merged',
'decam_flag_cleaned', 'decam_flag_gaia', 'decam_stellarity']
bands = [column[5:] for column in master_catalogue.colnames if 'f_ap' in column]
for band in bands:
columns += ["f_ap_{}".format(band), "ferr_ap_{}".format(band),
"m_ap_{}".format(band), "merr_ap_{}".format(band),
"f_{}".format(band), "ferr_{}".format(band),
"m_{}".format(band), "merr_{}".format(band),
"flag_{}".format(band)]
# -
# We check for columns in the master catalogue that we will not save to disk.
print("Missing columns: {}".format(set(master_catalogue.colnames) - set(columns)))
master_catalogue[:10].show_in_notebook()
master_catalogue[columns].write("{}/decam_merged_catalogue_xmm-lss.fits".format(TMP_DIR), overwrite=True)
| dmu1/dmu1_ml_XMM-LSS/2.6_DECAM_merging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#-*- coding:utf-8 -*-
from commonTool import *
# +
# 1. 빠른 실행을 위하여 기초 데이터 로컬에 저장해 놓기
# 기업 데이터가 변경된 경우에 한 번씩 해 주면 됨.
def saveAnnualBCP():
jobName = 'prepare to evaluate index'
begin(jobName)
workingDataPath = outputRawPath
mkdir(workingDataPath)
scriptPathName = crawlegoScriptPath + 'EVALUATE-PREPARE.xml'
parameter = {
'OUT_PATH': workingDataPath,
'DO_SERVER': '192.168.127.12'
}
print('ret code', runDashScript(scriptPathName, parameter))
end(jobName)
saveAnnualBCP()
# -
# +
# 2. 지표를 계산하여 저장하는 스크립트 실행
cachedDataPath = outputRawPath
workingDataPath = outputRawPath + 'index' + os.path.sep
mkdir(workingDataPath)
scriptPathName = crawlegoScriptPath + 'EVALUATE-INDEX.xml'
bf = open(resourceDir + 'BASIS-DAYS.txt', 'r', encoding='utf-8')
lineText = bf.readline() # 제목
while True:
lineText = bf.readline()
if not lineText:
break
lineText = lineText.strip()
if len(lineText) <= 0:
continue
# 0.YYYYMM, 1.FIRST_DAY, 2.SECOND_DAY, 3.PREV_DAY, 4.LAST_DAY, 5.NEXT_DAY
dayData = lineText.split('\t')
if dayData[0][0:4] == '2012': continue # 데이터 부족
if dayData[0] != yyyymm(): continue
parameter = {
'IN_PATH': cachedDataPath + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': workingDataPath + (os.path.sep if runServerType == 1 else ''),
'RES_PATH': resourceDir,
'BASIS_DATE': dayData[3], # 매달 마지막 전날을 기준으로 하자
'PERIOD': 'ANNUAL', # QUARTER
'DO_SERVER': '192.168.127.12'
}
print(dayData[0], 'index ret code', runDashScript(scriptPathName, parameter))
bf.close()
# +
# 3. 지표를 계산하여 저장하는 스크립트 실행 (분기)
cachedDataPath = outputRawPath
workingDataPath = outputRawPath + 'index' + os.path.sep
mkdir(workingDataPath)
scriptPathName = crawlegoScriptPath + 'EVALUATE-INDEX.xml'
bf = open(resourceDir + 'BASIS-DAYS.txt', 'r', encoding='utf-8')
lineText = bf.readline() # 제목
while True:
lineText = bf.readline()
if not lineText:
break
lineText = lineText.strip()
if len(lineText) <= 0:
continue
# 0.YYYYMM, 1.FIRST_DAY, 2.SECOND_DAY, 3.PREV_DAY, 4.LAST_DAY, 5.NEXT_DAY
dayData = lineText.split('\t')
if dayData[0] < '202002': continue # 데이터 부족
# if dayData[0] != yyyymm(): continue
parameter = {
'IN_PATH': cachedDataPath + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': workingDataPath + (os.path.sep if runServerType == 1 else ''),
'RES_PATH': resourceDir,
'BASIS_DATE': dayData[3], # 매달 마지막 전날을 기준으로 하자
'PERIOD': 'QUARTER', # QUARTER
'DO_SERVER': '192.168.127.12'
}
print(dayData[0], 'index ret code', runDashScript(scriptPathName, parameter))
bf.close()
# -
# +
# 4. 모멘텀 계산
cachedDataPath = outputRawPath
momentumDataPath = outputRawPath + 'momentum' + os.path.sep
mkdir(momentumDataPath)
scriptPathName = crawlegoScriptPath + 'EVALUATE-MM.xml'
bf = open(resourceDir + 'BASIS-DAYS.txt', 'r', encoding='utf-8')
lineText = bf.readline() # 제목
while True:
lineText = bf.readline()
if not lineText:
break
lineText = lineText.strip()
if len(lineText) <= 0:
continue
# 0.YYYYMM, 1.FIRST_DAY, 2.SECOND_DAY, 3.PREV_DAY, 4.LAST_DAY, 5.NEXT_DAY
dayData = lineText.split('\t')
if dayData[0][0:4] == '2012': continue # 데이터 부족
# if dayData[0] != yyyymm(): continue
parameter = {
'IN_PATH': cachedDataPath + (os.path.sep if runServerType == 1 else ''),
'OUT_PATH': momentumDataPath + (os.path.sep if runServerType == 1 else ''),
'RES_PATH': resourceDir + (os.path.sep if runServerType == 1 else ''),
'BASIS_DATE': dayData[3], # 매달 마지막 전날을 기준으로 하자
'DO_SERVER': '192.168.127.12'
}
print(dayData[0], 'momentum ret code', runDashScript(scriptPathName, parameter))
bf.close()
# -
# +
momentumDataPath = outputRawPath + 'momentum' + os.path.sep
scriptPathName = crawlegoScriptPath + 'EVALUATE-MM-UPLOAD.xml'
parameter = {
'IN_PATH': momentumDataPath,
'DO_SERVER': '192.168.127.12'
}
print('ret code', runDashScript(scriptPathName, parameter))
# -
| evaluateIndex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/edeediong/DataCamp-Projects/blob/Kontrol/Fake_News_Classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9q5xorP6V2m9" colab_type="text"
#
# + id="FHM3j5cSVz9V" colab_type="code" colab={}
# + [markdown] id="NS7aQSWmWZFZ" colab_type="text"
# # **Training and testing the "fake news" model with TfidfVectorizer**
#
# ---
#
#
# + [markdown] id="To_7C-6DVy90" colab_type="text"
# # **Instructions**
#
# * Instantiate a MultinomialNB classifier called nb_classifier.
# * Fit the classifier to the training data.
# * Compute the predicted tags for the test data.
# * Calculate and print the accuracy score of the classifier.
# * Compute the confusion matrix. As in the previous exercise, specify the keyword argument labels=['FAKE', 'REAL'] so that the resulting confusion matrix is easier to read.
# + id="Sd4Er1-_VxOR" colab_type="code" colab={}
# Create a Multinomial Naive Bayes classifier: nb_classifier
nb_classifier = MultinomialNB()
# Fit the classifier to the training data
trsinfit = nb_classifier.fit(tfidf_train,y_train)
# Create the predicted tags: pred
pred = nb_classifier.predict(tfidf_test)
# Calculate the accuracy score: score
score = metrics.accuracy_score(y_test,pred)
print(score)
# Calculate the confusion matrix: cm
cm = metrics.confusion_matrix(y_test,pred,labels = ['FAKE','REAL'])
print(cm)
# + [markdown] id="l5MPz5IZVNMk" colab_type="text"
# # **Instructions**
#
# * Create a list of alphas to try using np.arange(). Values should range from 0 to 1 with steps of 0.1.
#
# * Create a function train_and_predict() that takes in one argument: alpha. The function should:
# * Instantiate a MultinomialNB classifier with alpha=alpha.
# * Fit it to the training data.
# * Compute predictions on the test data.
# * Compute and return the accuracy score.
# * Using a for loop, print the alpha, score and a newline in between. Use your train_and_predict() function to compute the score. Does the score change along with the alpha? What is the best alpha?
# + id="mSK-oREjTpLI" colab_type="code" colab={}
# Create the list of alphas: alphas
alphas = np.arange(0,1,0.1)
# Define train_and_predict()
def train_and_predict(alphas):
# Instantiate the classifier: nb_classifier
nb_classifier = MultinomialNB(alpha=alpha)
# Fit to the training data
nb_classifier.fit(tfidf_train,y_train)
# Predict the labels: pred
pred = nb_classifier.predict(tfidf_test)
# Compute accuracy: score
score = metrics.accuracy_score(y_test,pred)
return score
# Iterate over the alphas and print the corresponding score
for alpha in alphas:
print('Alpha: ', alpha)
print('Score: ', train_and_predict(0.1))
print()
# + [markdown] id="C7qsSLbSULC3" colab_type="text"
# # **Instructions**
#
# * Save the class labels as class_labels by accessing the .classes_ attribute of nb_classifier.
#
# * Extract the features using the .get_feature_names() method of tfidf_vectorizer.
#
# * Create a zipped array of the classifier coefficients with the feature names and sort them by the coefficients. To do this, first use zip() with the arguments nb_classifier.coef_[0] and feature_names. Then, use sorted() on this.
#
# * Print the top 20 weighted features for the first label of class_labels.
#
# * Print the bottom 20 weighted features for the second label of class_labels.
# + id="q1pZu-QTTrTh" colab_type="code" colab={}
# Get the class labels: class_labels
class_labels = nb_classifier.classes_
# Extract the features: feature_names
feature_names = tfidf_vectorizer.get_feature_names()
# Zip the feature names together with the coefficient array and sort by weights: feat_with_weights
feat_with_weights = sorted(zip(nb_classifier.coef_[0], feature_names))
# Print the first class label and the top 20 feat_with_weights entries
print(class_labels[0], feat_with_weights[:20])
# Print the second class label and the bottom 20 feat_with_weights entries
print(class_labels[1], feat_with_weights[-20:])
| Fake_News_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NAME : <NAME>
# # email : <EMAIL>
# # EXERCISE 1
# +
a = 93
b = 17
c = a - b
d = 2*abs(c)
if 0 < c < 16:
print(c)
else:
print(d)
# -
# # EXERCISE 2
# +
a = 14
b = 12
c = 3.5
d = a + b + c
e = a*b*c
if a == b == c:
print(e)
else:
print(d)
# -
# # EXERCISE 3
# +
def test_number5(x, y):
if x == y or abs(x-y) == 5 or (x+y) == 5:
return True
else:
return False
print(test_number5(7, 2))
print(test_number5(3, 2))
print(test_number5(2, 2))
# -
# # EXCERCISE 4
#
# +
x = int(input("Input first number: "))
y = int(input("Input second number: "))
z = int(input("Input third number: "))
a1 = min(x, y, z)
a3 = max(x, y, z)
a2 = (x + y + z) - a1 - a3
print("Numbers in sorted order: ", a1, a2, a3)
# -
#
# # EXCERCISE 5
def sum_of_cubes(n):
n -= 2
total = 0
while n > 0:
total += n * n * n
n -= 1
return total
print("Sum of cubes: ",sum_of_cubes(3))
| Fareed_Bello.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kaipak/from-eyes-to-ears/blob/master/mask_training_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="mik6aVzBEt7E" colab_type="code" outputId="378a5bfe-3505-465a-e7a2-cdd17edb7481" colab={"base_uri": "https://localhost:8080/", "height": 442}
# !pip install numpy scikit-image Shapely awscli
# + id="-Renkz992hR6" colab_type="code" colab={}
import pandas as pd
import os
import cv2
import numpy as np
from google.colab.patches import cv2_imshow
from skimage import measure
from shapely.geometry import Polygon, MultiPolygon
import boto3
from botocore import UNSIGNED
from botocore.config import Config
import pickle
# + id="nFtmQr0dul0V" colab_type="code" outputId="37ca7f20-8f03-4f34-b656-fca449939711" colab={"base_uri": "https://localhost:8080/", "height": 51}
# !rm -rf train validation test *.zip
# !mkdir tmp
# !mkdir train
# !mkdir train/images
# !mkdir validation
# !mkdir validation/images
# !mkdir test
# !mkdir test/images
# !mkdir masks/
# + [markdown] id="wNYIaiSh3s3-" colab_type="text"
# ## Start generating a phase dataset HERE
# + id="r7Y_tJLgvNqz" colab_type="code" colab={}
# Valid phases are: train, validation, test
phase = 'train'
filter_categories = ["Saxophone", "Tank", "Goose", "Guitar"]
# + [markdown] id="s0a3FQ5REoT-" colab_type="text"
# ## Run the cell that match the current phase
# + id="dNMxy-bZDMxB" colab_type="code" outputId="e5e39b7c-d00a-4ab4-aa57-cf3ff095d339" colab={"base_uri": "https://localhost:8080/", "height": 799}
# !rm *.csv
# !wget https://storage.googleapis.com/openimages/v5/class-descriptions-boxable.csv
# !wget https://storage.googleapis.com/openimages/v5/train-annotations-human-imagelabels-boxable.csv
# !rm -rf masks/
# !mkdir masks/
# %cd masks/
# !wget https://storage.googleapis.com/openimages/v5/train-masks/train-masks-0.zip
# !unzip train-masks-0.zip > /dev/null
# !wget https://storage.googleapis.com/openimages/v5/train-masks/train-masks-1.zip
# !unzip train-masks-1.zip > /dev/null
# %cd ..
# + id="5GLKyZ18DLGV" colab_type="code" outputId="898ca775-7d63-49ea-8708-c7e902a72741" colab={"base_uri": "https://localhost:8080/", "height": 799}
# !rm *.csv
# !wget https://storage.googleapis.com/openimages/v5/class-descriptions-boxable.csv
# !wget https://storage.googleapis.com/openimages/v5/validation-annotations-human-imagelabels-boxable.csv
# !rm -rf masks/
# !mkdir masks/
# %cd masks/
# !wget https://storage.googleapis.com/openimages/v5/validation-masks/validation-masks-0.zip
# !unzip validation-masks-0.zip > /dev/null
# !wget https://storage.googleapis.com/openimages/v5/validation-masks/validation-masks-1.zip
# !unzip validation-masks-1.zip > /dev/null
# %cd ..
# + id="8EhjVLU5iE0A" colab_type="code" outputId="59246809-3198-4c94-dee0-2dff4ed7c42f" colab={"base_uri": "https://localhost:8080/", "height": 799}
# !rm *.csv
# !wget https://storage.googleapis.com/openimages/v5/class-descriptions-boxable.csv
# !wget https://storage.googleapis.com/openimages/v5/test-annotations-human-imagelabels-boxable.csv
# !rm -rf masks/
# !mkdir masks/
# %cd masks/
# !wget https://storage.googleapis.com/openimages/v5/test-masks/test-masks-0.zip
# !unzip test-masks-0.zip > /dev/null
# !wget https://storage.googleapis.com/openimages/v5/test-masks/test-masks-1.zip
# !unzip test-masks-1.zip > /dev/null
# %cd ..
# + id="LiTUlJsi23vw" colab_type="code" colab={}
classes = pd.read_csv('class-descriptions-boxable.csv', header=None, names=["LabelName", "class"])
image_classes = pd.read_csv(f'{phase}-annotations-human-imagelabels-boxable.csv')
# + id="_ToAZYAI4LDm" colab_type="code" colab={}
# Find where we have data
training_imageids = classes.join(image_classes.set_index('LabelName'), on='LabelName').filter(["ImageID", "LabelName","class"])
training_imageids = training_imageids[training_imageids["ImageID"].str.startswith('0') | training_imageids["ImageID"].str.startswith('1')]
training_imageids['cutLabelName'] = training_imageids["LabelName"].str.replace('/', '')
masks = pd.DataFrame([f for f in [file.split('_') + [file] for file in os.listdir('masks')] if len(f) == 4], columns=['ImageID', 'cutLabelName', 'extension', 'mask_filename'])
training_imageids = training_imageids.merge(masks, how='inner', left_on=['ImageID', 'cutLabelName'], right_on = ['ImageID', 'cutLabelName']).filter(["ImageID", "LabelName", "class", 'mask_filename'])
# + [markdown] id="RErA9o-apRFS" colab_type="text"
# ## Get a summary to pick classes for training
# + id="V2YRkNS1I-Bf" colab_type="code" outputId="18176195-dbd4-448f-a6d5-3e0db89a16d9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
i = pd.get_option('display.max_rows', None)
pd.set_option('display.max_rows', None)
print(training_imageids.groupby("class").count()[['ImageID']])
pd.set_option('display.max_rows', i)
# + id="EaGivpmbWYqg" colab_type="code" outputId="3564acf8-0d88-47e7-c844-d4a7200dfe05" colab={"base_uri": "https://localhost:8080/", "height": 34}
filtered = training_imageids[training_imageids["class"].isin(filter_categories)].filter(["ImageID", "mask_filename", "class"])
filtered["info"] = list(zip(filtered['mask_filename'], filtered['class']))
masks = filtered.groupby("ImageID")["info"].apply(list).values.tolist()
print(len(masks))
# + id="cLrqTjQVYdes" colab_type="code" outputId="c892969d-cd93-4cb1-9aa7-e8196540fabf" colab={"base_uri": "https://localhost:8080/", "height": 1000}
s3 = boto3.client('s3', config=Config(signature_version=UNSIGNED))
categories = []
metadata = []
max_size = 500
cat_counts = {}
n = 1
for m in masks:
annotations = []
catset = set()
for f, c in m:
catset.add(c)
cont = False
for c in catset:
if c in cat_counts:
if cat_counts[c] > max_size:
cont = True
break
else:
cat_counts[c] += 1
else:
cat_counts[c] = 1
if cont:
#print(f"we have collected enough for categores {catset}")
continue
for file, category in m:
if category not in categories:
categories.append(category)
category_id = categories.index(category)
fullname = f"masks/{file}"
image_id = file.split('_')[0]
mask = (cv2.imread(fullname)[...,0]/255.0).astype(np.uint8)
contours = measure.find_contours(mask, 0.5, positive_orientation='low')
segmentations = []
polygons = []
for contour in contours:
# Flip from (row, col) representation to (x, y)
# and subtract the padding pixel
for i in range(len(contour)):
row, col = contour[i]
contour[i] = (col - 1, row - 1)
try:
# Make a polygon and simplify it
poly = Polygon(contour)
poly = poly.simplify(1.0, preserve_topology=False)
if poly.type == 'MultiPolygon':
for p in poly:
polygons.append(p)
segmentation = np.array(p.exterior.coords).ravel().tolist()
if len(segmentation) >= 6:
segmentations.append(segmentation)
else:
continue
else:
polygons.append(poly)
segmentation = np.array(poly.exterior.coords).ravel().tolist()
if len(segmentation) >= 6:
segmentations.append(segmentation)
else:
continue
except:
print("Error processing image mask, skipping")
continue
if len(segmentations) == 0:
print("Can't build polygons, skipping")
continue
# Combine the polygons to calculate the bounding box and area
multi_poly = MultiPolygon(polygons)
x, y, max_x, max_y = multi_poly.bounds
width = max_x - x
height = max_y - y
bbox = (x, y, width, height)
area = multi_poly.area
annotation = {
'bbox': multi_poly.bounds,
'bbox_mode': 0,
'category_id': category_id,
'iscrowd': 0,
'segmentation': segmentations
}
annotations.append(annotation)
if len(annotations) > 0:
h, w = mask.shape
md = {
'annotations': annotations,
'file_name': f'{phase}/images/{image_id}.jpg',
'height': h,
'width': w,
'image_id': image_id
}
metadata.append(md)
image_file = f'{image_id}.jpg'
print(f"{n} - Fetching image: {image_file} - Categories: {catset} - Current Counters: {cat_counts}")
s3.download_file('open-images-dataset', f'{phase}/{image_file}', f'tmp/{image_file}')
img = cv2.imread(f'tmp/{image_file}')
os.unlink(f'tmp/{image_file}')
img = cv2.resize(img, (w, h))
cv2.imwrite(f'{phase}/images/{image_file}', img)
n += 1
print(f"Data ready for phase: {phase}")
print("Encoding Metadata")
pickle.dump({"categories": categories, "metadata": metadata}, open(f"{phase}/metadata.pkl", "wb"))
print("Done")
# + [markdown] id="ZKLFEeOK32Dn" colab_type="text"
# ## Package and send final datasets to GDrive (ONLY RUN ONCE ALL DATASET ARE CREATED)
# + id="TB54olm10oe0" colab_type="code" outputId="e8087390-9245-48ac-b19d-67f13653d178" colab={"base_uri": "https://localhost:8080/", "height": 34}
# !rm *.zip
# !zip -r train.zip train
# !zip -r validation.zip validation
# !zip -r test.zip test
from google.colab import drive
drive.mount('/content/drive')
# !cp *.zip "drive/My Drive"
drive.flush_and_unmount()
# + id="cZlYgcSaguHr" colab_type="code" colab={}
| mask_training_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Astronomical Interactives
# [*Astro Interactives*](https://juancab.github.io/AstroInteractives/) is a series of Jupyter Notebooks meant to provide interactive demos and simulations for introductory astronomy courses (and in a few cases, upper-division astrophysics courses). These notebooks were initially developed by
# - <NAME> (Professor for Physics and Astronomy at Minnesota State University Moorhead)
# - <NAME> (Minnesota State University Moorhead Physics Major, Class of 2019)
# - <NAME> (Minnesota State University Moorhead Physics Major, Class of 2019)
#
# The goal was to replace many of the Adobe Flash-based tools that we use in labs using applications we created in-house. This allows much greater tunability to our needs AND is likely to be usable much farther into the future since Adobe Flash's end of life is set for the end of 2020.
#
# This software is provide "as-is" and may contain bugs/errors that could mis-represent astronomical reality. You use this software at your own risk! Keep in mind the authors are not professional programers!
# ## Available Notebooks
# The currently available Interactives are the following:
#
# ### General Astronomy
# 1. [Small Angle Approximation Interactive](Interactives/SmallAngleEquation.ipynb) - This interactive was designed to allow students who are uncomfortable with mathematics visualize what is happening when the small angle approximation is valid.
# 2. [Flux vs.Luminosity Interactive](Interactives/FluxVsLuminositySimulation.ipynb) - This interactive is meant to be used in discussions of the concepts of flux/brightness, luminosity, and the inverse square law.
# 3. [Doppler Shift Interactives](Interactives/DopplerShift.ipynb) - These Doppler Shift interactives are designed to allow students to first model simple Doppler Shift and then to model the more complicated time-varying Doppler Shift seen in binary star and exoplanetary systems.
#
# ### Solar System and Planetary Astronomy
# 1. [Radioactive Decay Interactives](Interactives/Radioactivity.ipynb) - This pair of interactive plots is used to illustrate the concept of radioactive decay, half-life, and how these concepts can be used to determine the age of some rocks via the geochron method.
# 2. [Exoplanet System Simulation Interactive](Interactives/Exoplanet_Sim.ipynb) - This interactive contains a full 3D simulation of a star system containing a single Jovian exoplanet, including a radial velocity or light curve for that system.
#
# ### Stellar Astronomy
# 1. [Center of Mass Interactive](Interactives/Center_of_Mass.ipynb) - This interactive model simply allows a student to adjust the mass and separation of two stars and see the shifts of their orbits, illustrating the center of mass position.
# 2. [Binary Star Simulation Interactive](Interactives/Binary_Star_Sim.ipynb) - This interactive contains a full 3D simulation of a binary star system, including a radial velocity or light curve for that system. The user can control most physical parameters of the system.
# 3. [Luminosity Calculator Interactive](Interactives/LuminosityCalculator.ipynb) - This interactive is used to illustrate to students how the radius and temperature of a star affect its luminosity. The colors of the stars accurately correspond to the temperature, but the stars produced might not have radii and temperatures that correspond to real stars.
# 4. [Blackbody Spectra Interactives](Interactives/Blackbody_Simulation.ipynb) - This set of interactive figures can be used to explore the properties of the Blackbody spectrum as applied to stars.
# 5. [HR Diagram Interactives](Interactives/HR_Diagram.ipynb) - This set of interactive plots allows a student to explore the data that goes into the HR diagram, then how main sequence fitting can be used to determine the distance to a star cluster, and finally, how detailed stellar evolution models can be used to also determine the age of a star cluster.
#
# ### Galactic and Extragalactic Astrophysics
#
# The following interactives are meant to be used in an upper-division astrophysics class as a way of introducing $\ell$-v diagrams and 21-cm spectra. These are not really meant to introduce the concepts but rather to allow students to explore how the distribution of neutral hydrogen gas both in position and velocity affects the observed $\ell$-v diagram of the Milky Way galaxy and the HI spectra of external galaxies.
# 1. [Synthetic $\ell$-v Diagram](Interactives/Synthetic_LV_Diagram.ipynb) - This interactive takes a Milky Way-like rotation curve and neutral gas profile and generates a synthetic $\ell$-v diagram. Users can then simply trace out a new rotation curve or neutral gas profile and see the corresponding $\ell$-v diagram.
# 2. [Synthetic HI Spectra](Interactives/Synthetic_Galaxy_HI_Spectra.ipynb) - This interactive model allows a student to see the single-dish (unresolve) HI spectra or the resolved HI spectra (aka velocity map) corresponding to a given model galaxy. As with the $\ell$-v diagram interactive, users can trace out a new rotation curve or neutral gas profile and see the corresponding spectra.
#
# ## Known issues
# - In some interactives, the animations are buffered by the web browser so that they lag behind the user inputs. Efforts have been made to optimize the code to avoid these problems.
# ## Documentation/Help
# - The documentation for how the Python notebooks do what they do are in the code comments for now. We hope to eventually add copies of some of the lab exercises we do so people can see these on context.
# - Send us an email at <EMAIL> if you need any help/information.
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + run_control={"marked": false}
import logging
reload(logging)
log_fmt = '%(asctime)-9s %(levelname)-8s: %(message)s'
logging.basicConfig(format=log_fmt)
# Change to info once the notebook runs ok
logging.getLogger().setLevel(logging.INFO)
# Generate plots inline
# %matplotlib inline
import os
# -
# # Target Connectivity
# ## Board specific settings
# Boards specific settings can be collected into a JSON
# platform description file:
# !ls -la $LISA_HOME/libs/utils/platforms/
# !cat $LISA_HOME/libs/utils/platforms/hikey.json
# ## Single configuration dictionary
# + run_control={"marked": false}
# Check which Android devices are available
# !adb devices
# + run_control={"marked": false}
ADB_DEVICE = '00b43d0b08a8a4b8'
# ADB_DEVICE = '607A87C400055E6E'
# -
# Unified configuration dictionary
my_conf = {
# Target platform
"platform" : 'android',
# Location of external tools (adb, fastboot, systrace, etc)
# These properties can be used to override the environment variables:
# ANDROID_HOME and CATAPULT_HOME
"ANDROID_HOME" : "/opt/android-sdk-linux",
"CATAPULT_HOME" : "/home/derkling/Code/catapult",
# Boards specific settings can be collected into a JSON
# platform description file, to be placed under:
# LISA_HOME/libs/utils/platforms
"board" : 'hikey',
# If you have multiple Android device connected, here
# we can specify which one to target
"device" : ADB_DEVICE,
# Folder where all the results will be collected
"results_dir" : "ReleaseNotes_v16.09",
}
# +
from env import TestEnv
te = TestEnv(my_conf, force_new=True)
target = te.target
# -
# # Energy Meters Support
# - Simple unified interface for multiple acquisition board
# - exposes two simple methods: **reset()** and **report()**
# - reporting **energy** consumptions
# - reports additional info supported by the specific probe,<br>
# e.g. collected samples, stats on current and voltages, etc.
# +
from time import sleep
def sample_energy(energy_meter, time_s):
# Reset the configured energy counters
energy_meter.reset()
# Run the workload you want to measure
#
# In this simple example we just wait some time while the
# energy counters accumulate power samples
sleep(time_s)
# Read and report the measured energy (since last reset)
return energy_meter.report(te.res_dir)
# -
# - Channels mapping support
# - allows to give a custom name to each channel used
# ## ARM Energy Probe (AEP)
# Requirements:
# 1. the **caimin binary tool** must be availabe in PATH<br>
# https://github.com/ARM-software/lisa/wiki/Energy-Meters-Requirements#arm-energy-probe-aep
# 2. the **ttyACMx device** created once you plug in the AEP device
# !ls -la /dev/ttyACM*
ACM_DEVICE = '/dev/ttyACM1'
# ### Direct usage
# +
# Energy Meters Configuration for ARM Energy Probe
aep_conf = {
'conf' : {
# Value of the shunt resistor [Ohm] for each channel
'resistor_values' : [0.010],
# Device entry assigned to the probe on the host
'device_entry' : ACM_DEVICE,
},
'channel_map' : {
'BAT' : 'CH0'
}
}
from energy import AEP
ape_em = AEP(target, aep_conf, '/tmp')
# -
nrg_report = sample_energy(ape_em, 2)
print nrg_report
# !cat $nrg_report.report_file
# ### Usage via TestEnv
my_conf = {
# Configure the energy meter to use
"emeter" : {
# Require usage of an AEP meter
"instrument" : "aep",
# Configuration parameters require by the AEP device
"conf" : {
# Value of the shunt resistor in Ohm
'resistor_values' : [0.099],
# Device entry assigned to the probe on the host
'device_entry' : ACM_DEVICE,
},
# Map AEP's channels to logical names (used to generate reports)
'channel_map' : {
'BAT' : 'CH0'
}
},
# Other target configurations
"platform" : 'android',
"board" : 'hikey',
"device" : ADB_DEVICE,
"results_dir" : "ReleaseNotes_v16.09",
"ANDROID_HOME" : "/opt/android-sdk-linux",
"CATAPULT_HOME" : "/home/derkling/Code/catapult",
}
# +
from env import TestEnv
te = TestEnv(my_conf, force_new=True)
# -
for i in xrange(1,11):
nrg_report = sample_energy(te.emeter, 1)
nrg_bat = float(nrg_report.channels['BAT'])
print "Sample {:2d}: {:.3f}".format(i, nrg_bat)
# ## BayLibre's ACME board (ACME)
# Requirements:
# 1. the **iio-capture tool** must be available in PATH<br>
# https://github.com/ARM-software/lisa/wiki/Energy-Meters-Requirements#iiocapture---baylibre-acme-cape
# 2. the ACME CAPE should be reacable by network
# !ping -c1 baylibre-acme.local | grep '64 bytes'
# ### Direct usage
# +
# Energy Meters Configuration for BayLibre's ACME
acme_conf = {
"conf" : {
#'iio-capture' : '/usr/bin/iio-capture',
#'ip_address' : 'baylibre-acme.local',
},
"channel_map" : {
"Device0" : 0,
"Device1" : 1,
},
}
from energy import ACME
acme_em = ACME(target, acme_conf, '/tmp')
# -
nrg_report = sample_energy(acme_em, 2)
print nrg_report
# !cat $nrg_report.report_file
# ### Usage via TestEnv
# + run_control={"marked": false}
my_conf = {
# Configure the energy meter to use
"emeter" : {
# Require usage of an AEP meter
"instrument" : "acme",
"conf" : {
#'iio-capture' : '/usr/bin/iio-capture',
#'ip_address' : 'baylibre-acme.local',
},
'channel_map' : {
'Device0' : 0,
'Device1' : 1,
},
},
# Other target configurations
"platform" : 'android',
"board" : 'hikey',
"device" : ADB_DEVICE,
"results_dir" : "ReleaseNotes_v16.09",
"ANDROID_HOME" : "/opt/android-sdk-linux",
"CATAPULT_HOME" : "/home/derkling/Code/catapult",
}
# + run_control={"marked": false}
from env import TestEnv
te = TestEnv(my_conf, force_new=True)
target = te.target
# -
for i in xrange(1,11):
nrg_report = sample_energy(te.emeter, 1)
nrg_bat = float(nrg_report.channels['Device1'])
print "Sample {:2d}: {:.3f}".format(i, nrg_bat)
# # Android Integration
# A new Android library has been added which provides APIs to:
# - simplify the interaction with a device
# - execute interesting workloads and benchmarks
# - make it easy the integration of new workloads and benchmarks
#
# Not intended to replace WA, but instead to provide a Python based<br>
# programming interface to **automate reproducible experiments** on<br>
# and Android device.
# ## System control APIs
# +
from android import System
print "Supported functions:"
for f in dir(System):
if "__" in f:
continue
print " ", f
# -
# Capturing main useful actions, for example:
# - ensure we set AIRPLAIN_MODE before measuring scheduler energy
# - provide simple support for input actions (relative swipes)
# +
# logging.getLogger().setLevel(logging.DEBUG)
# -
# Example (use tab to complete)
System.
System.menu(target)
System.back(target)
youtube_apk = System.list_packages(target, 'YouTube')
if youtube_apk:
System.start_app(target, youtube_apk[0])
logging.getLogger().setLevel(logging.INFO)
# ## Screen control APIs
# +
from android import Screen
print "Supported functions:"
for f in dir(Screen):
if "__" in f:
continue
print " ", f
# +
#logging.getLogger().setLevel(logging.DEBUG)
# -
# Example (use TAB to complete)
Screen.
Screen.set_brightness(target, auto=False, percent=100)
Screen.set_orientation(target, auto=False, portrait=False)
# +
# logging.getLogger().setLevel(logging.INFO)
# -
# ## Workloads Execution
# A simple workload class allows to easily add a wrapper for the exection
# of a specific Android application.
#
# **NOTE:** To keep things simple, LISA does not provide APKs installation support.
#
# *All the exposes APIs assume that the required packages are already installed<br>
# in the target. Whenever a package is missing, LISA reports that and it's up<br>
# to the user to install it before using it.*
#
# A wrapper class usually requires to specify:
# - a package name<br>
# which will be used to verify if the APK is available in the target
# - a run method<br>
# which usually exploits the other Android APIs to defined a **reproducible
# exection** of the specified workload
#
# A reproducible experiment should take care of:
# - setups wirelesse **connections status**
# - setup **screen orientation and backlight** level
# - properly collect **energy measurements** across the sensible part of the experiment
# - possibly collect **frames statistics** whenever available
# ### Example of YouTube integration
# Here is an example wrapper which allows to play a YouTube<br>
# video for a specified number of seconds:
#
# https://github.com/ARM-software/lisa/blob/master/libs/utils/android/workloads/youtube.py
# ### Example usage of the Workload API
# +
# logging.getLogger().setLevel(logging.DEBUG)
# +
from android import Workload
# Get the list of available workloads
wloads = Workload(te)
wloads.availables(target)
# +
yt = Workload.get(te, name='YouTube')
# Playback big bug bunny for 15s starting from 1:20s
video_id = 'XSGBVzeBUbk'
video_url = "https://youtu.be/{}?t={}s".format(video_id, 80)
# Play video and measure energy consumption
results = yt.run(te.res_dir,
video_url, video_duration_s=16,
collect='energy')
# -
results
framestats = results[0]
# !cat $framestats
# ## Benchmarks
# Android benchmarks can be integrated as standalone Notebook, like for example
# what we provide for PCMark:
# https://github.com/ARM-software/lisa/blob/master/ipynb/android/benchmarks/Android_PCMark.ipynb
#
# Alternatively we are adding other benchmarks as predefined Android workloads.
# ### UiBench support
# Here is an example of UiBench workload which can run a specified number
# of tests:
#
# + run_control={"marked": false}
from android import Workload
ui = Workload.get(te, name='UiBench')
# Play video and measure energy consumption
results = ui.run(te.res_dir,
ui.test_GlTextureView,
duration_s=5,
collect='energy')
# -
results
framestats = results[0]
# !cat $framestats
# # Improved Trace Analysis support
# The Trace module is a wrapper around the TRAPpy library which has been
# updated to:
#
# - support parsing of **systrace** file format<br>
# requires catapult locally installed<br>
# https://github.com/catapult-project/catapult
# - parsing and DataFrame generation for **custom events**
# ## Create an example trace
# **NOTE:** the cells in this sections are required just to create
# a trace file to be used by the following sections
# The following exanples uses an HiKey board
ADB_DEVICE = '607A87C400055E6E'
# +
# logging.getLogger().setLevel(logging.DEBUG)
# + run_control={"marked": false}
# Unified configuration dictionary
my_conf = {
# Tools required
"tools" : ['rt-app', 'trace-cmd'],
# RTApp calibration
#"modules" : ['cpufreq'],
"rtapp-calib" : {
"0": 254, "1": 252, "2": 252, "3": 251,
"4": 251, "5": 252, "6": 251, "7": 251
},
# FTrace configuration
"ftrace" : {
# Events to trace
"events" : [
"sched_switch",
"sched_wakeup",
"sched_wakeup_new",
"sched_wakeup_tracking",
"sched_stat_wait",
"sched_overutilized",
"sched_contrib_scale_f",
"sched_load_avg_cpu",
"sched_load_avg_task",
"sched_tune_config",
"sched_tune_filter",
"sched_tune_tasks_update",
"sched_tune_boostgroup_update",
"sched_boost_cpu",
"sched_boost_task",
"sched_energy_diff",
"cpu_capacity",
"cpu_frequency",
"cpu_idle",
"walt_update_task_ravg",
"walt_update_history",
"walt_migration_update_sum",
],
# # Kernel functions to profile
# "functions" : [
# "pick_next_task_fair",
# "select_task_rq_fair",
# "enqueue_task_fair",
# "update_curr_fair",
# "dequeue_task_fair",
# ],
# Per-CPU buffer configuration
"buffsize" : 10 * 1024,
},
# Target platform
"platform" : 'android',
"board" : 'hikey',
"device" : ADB_DEVICE,
"results_dir" : "ReleaseNotes_v16.09",
"ANDROID_HOME" : "/opt/android-sdk-linux",
"CATAPULT_HOME" : "/home/derkling/Code/catapult",
}
# + run_control={"marked": false}
from env import TestEnv
te = TestEnv(my_conf, force_new=True)
target = te.target
# + run_control={"marked": false}
from wlgen import RTA,Ramp
# Let's run a simple RAMP task
rta = RTA(target, 'ramp')
rta.conf(
kind='profile',
params = {
'ramp' : Ramp().get()
}
);
# + run_control={"marked": false}
te.ftrace.start()
target.execute("echo 'my_marker: label=START' > /sys/kernel/debug/tracing/trace_marker",
as_root=True)
rta.run(out_dir=te.res_dir)
target.execute("echo 'my_marker: label=STOP' > /sys/kernel/debug/tracing/trace_marker",
as_root=True)
te.ftrace.stop()
trace_file = os.path.join(te.res_dir, 'trace.dat')
te.ftrace.get_trace(trace_file)
# -
# ## DataFrame namespace
# + run_control={"marked": false}
from trace import Trace
events_to_parse = my_conf['ftrace']['events']
events_to_parse += ['my_marker']
trace = Trace(te.platform, trace_file, events=events_to_parse)
# + run_control={"marked": false}
trace.available_events
# -
# Use TAB to complete
trace.data_frame.
rt_tasks = trace.data_frame.rt_tasks()
rt_tasks.head()
lat_df = trace.data_frame.latency_df('ramp')
lat_df.head()
custom_df = trace.data_frame.trace_event('my_marker')
custom_df
ctxsw_df = trace.data_frame.trace_event('sched_switch')
ctxsw_df.head()
# ## Analysis namespace
# Use TAB to complete
trace.analysis.
# + run_control={"marked": false}
trace.analysis.tasks.plotTasks(tasks='ramp',
signals=['util_avg', 'boosted_util',
'sched_overutilized', 'residencies'])
# -
lat_data = trace.analysis.latency.plotLatency('ramp')
lat_data.T
trace.analysis.frequency.plotClusterFrequencies()
trace.analysis.frequency.plotClusterFrequencyResidency(pct=True, active=True)
trace.analysis.frequency.plotClusterFrequencyResidency(pct=True, active=False)
# + run_control={"marked": false}
rtapp_df = trace.data_frame.rtapp_tasks()
rtapp_df
# + run_control={"marked": false}
for task in rtapp_df.index.tolist():
trace.analysis.perf.plotRTAppPerf(task)
# + run_control={"marked": false}
ramp_df = trace.data_frame.rtapp_samples('ramp')
ramp_df.head()
# -
rt_tasks = trace.data_frame.rt_tasks()
rt_tasks.head()
# # Notebooks
# ## New collection of examples
# Each new API introduced in LISA has an associated notebook which shows a
# complete example of its usage.<br>
# Examples are usually defined to:
#
# - setup the connection to a target (usually a JUNO board)
# - configure a workload (usually using RTApp)
# - run workload and collect required measures
# - show the most common functions exposed by the new API
# - Energy meters APIs:<br>
# https://github.com/ARM-software/lisa/tree/master/ipynb/examples/energy_meter
# - Trace analysis APIs:<br>
# https://github.com/ARM-software/lisa/tree/master/ipynb/examples/trace_analysis
# ## Examples for Android specific APIs
| ipynb/releases/ReleaseNotes_v16.10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="s1fcvWw6se0T"
# %tensorflow_version 2.x
# + id="HNj9DALrxkdd" colab={"base_uri": "https://localhost:8080/"} outputId="47149c4c-d627-4d00-c02c-882ce3b59f4a"
# !nvidia-smi
# + id="tKvoSER01pNp" colab={"base_uri": "https://localhost:8080/"} outputId="fbc0db22-3cec-4293-bee9-d39c5655e064"
# !pip3 install tensorflow-addons --upgrade
# + id="QvgeZbR_yhBN"
# !rm -rf sample_data
# + id="buJ4ImYNxlH5" colab={"base_uri": "https://localhost:8080/"} outputId="36d65900-e1c1-4bdb-da50-28dc035a08b4"
# !git clone https://github.com/omerferhatt/neuro-image-reconstruction
# + id="r77sd9pIyj7v" colab={"base_uri": "https://localhost:8080/"} outputId="98c2fe1f-926a-4336-ef32-afbe642a44a4"
# %cd neuro-image-reconstruction/
# + id="m3DO4mY9yqzG" colab={"base_uri": "https://localhost:8080/"} outputId="15ccb3c0-ccaa-49b4-8910-f5c6ccb7ba88"
# !gdown --id 1dzRr8XGqrSIKJT26m7BVRn7-6s5zK_r3 -O data/mind-big-data.zip
# !gdown --id 1ycaA1npRPV0DklrQW7Mz9zmAXJDHyJw2 -O data/mind-big-data-imagenet.zip
# + id="kLrgdmRhzFjk"
# !unzip -q data/mind-big-data -d data/
# !unzip -q data/mind-big-data-imagenet.zip -d data/
# + id="HGimVvEKzw7L"
# !rm -rf data/mind-big-data-imagenet.zip && rm -rf data/mind-big-data.zip
# + id="eWki24-s4UfP" colab={"base_uri": "https://localhost:8080/"} outputId="339bed2a-0907-4cf5-f63a-22a60bd0d0b3"
# %cd /content/neuro-image-reconstruction/
# + id="KNiAROns1nYq" colab={"base_uri": "https://localhost:8080/"} outputId="232f0caa-403b-4759-afcf-68138baf0c20"
# !python3 main.py
| notebooks/neuro_image_reconsturction_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 9
# ## Problem 9-9
# + slideshow={"slide_type": "skip"}
# %pylab notebook
# -
# ### Description
# How many pulses per second must be supplied to the control unit of the motor in [Problem 9-8](Ch9-Problem_9-08.ipynb) to achieve
# a rotational speed of 600 r/min?
p = 12
n_m = 600 # [r/min]
# ### SOLUTION
# From Equation (9-20),
#
# $$n_m = \frac{1}{3p}n_\text{pulses}$$
n_pulses = 3*p*n_m
print('''
n_pulses = {:.0f} pulses/min = {:.0f} pulses/sec
============================================'''.format(n_pulses, n_pulses/60))
| Chapman/Ch9-Problem_9-09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importação da biblioteca aara acessar a página web.
import urllib.request
# Definimos a url para abertura.
with urllib.request.urlopen('https://www.python.org/') as url:
page = url.read()
# Impressão do conteúdo.
print(page)
# Importando o pacote bs4 e a biblioteca BeautifulSoup.
from bs4 import BeautifulSoup
soup = BeautifulSoup(page, 'html.parser')
soup.title
soup.title.string
soup.a
soup.find_all('a')
tables = soup.find('table')
print(tables)
| web/Web Scraping/Web scraping com site do python/Script funcional.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ml4ocn]
# language: python
# name: conda-env-ml4ocn-py
# ---
# # Full GP Example
# This is a walk-through of a full example using the GP algorithm. This algorithm is a bit expensive to train but I think it is a good starting point. We can upgrade this method to more sparse methods so that we can train
import numpy as np
from sklearn.datasets import make_regression
from sklearn.model_selection import train_test_split
from sklearn.base import clone
from sklearn.decomposition import PCA
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import WhiteKernel, ConstantKernel, RBF
from sklearn.multioutput import MultiOutputRegressor
from sklearn.compose import TransformedTargetRegressor
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
import time as time
# +
import sys
sys.path.insert(0, "/Users/eman/Documents/code_projects/ml4ocean")
from src.models.utils import MultiTaskGP, PCATargetTransform
# +
# Make Fake Dataset
X, y = make_regression(
n_samples=1000,
n_features=10, # Total Features
n_informative=3, # Informative Features
n_targets=10,
bias=10,
noise=0.8,
random_state=123
)
# Training and Testing
xtrain, xtest, ytrain, ytest = train_test_split(X, y, train_size=500, random_state=123)
# -
# ## GP - Standard
# +
# define kernel function
kernel = ConstantKernel() * RBF() + WhiteKernel()
# define GP model
gp_model = GaussianProcessRegressor(
kernel=kernel, # kernel function (very important)
normalize_y=True, # good standard practice
random_state=123, # reproducibility
n_restarts_optimizer=10, # good practice (avoids local minima)
)
# train GP Model
t0 = time.time()
gp_model.fit(xtrain, ytrain)
t1 = time.time() - t0
# Predictions
ypred, ystd = gp_model.predict(xtest, return_std=True)
# +
# Get Stats
mae = mean_absolute_error(ypred, ytest)
mse = mean_squared_error(ypred, ytest)
rmse = np.sqrt(mse)
r2 = r2_score(ypred, ytest)
print(
f"MAE: {mae:.3f}\nMSE: {mse:.3f}\nRMSE: {rmse:.3f}\nR2: {r2:.3f}"
f" \nTime: {t1:.3} seconds"
)
# -
# ## GP - MultiOutput w. PCA Transformer (Manually)
# +
# define kernel function
kernel = ConstantKernel() * RBF() + WhiteKernel()
# define GP model
gp_model = GaussianProcessRegressor(
kernel=kernel, # kernel function (very important)
normalize_y=True, # good standard practice
random_state=123, # reproducibility
n_restarts_optimizer=10, # good practice (avoids local minima)
)
# Define target transformer
pca_model = PCA(n_components=3)
# Transform Targes
ytrain_red = pca_model.fit_transform(ytrain)
# train GP Model
t0 = time.time()
gp_model.fit(xtrain, ytrain_red)
t1 = time.time() - t0
# Predictions
ypred_red, ystd = gp_model.predict(xtest, return_std=True)
# Inverse transform predictions
ypred = pca_model.inverse_transform(ypred_red)
# +
# Get Stats
mae = mean_absolute_error(ypred, ytest)
mse = mean_squared_error(ypred, ytest)
rmse = np.sqrt(mse)
r2 = r2_score(ypred, ytest)
print(
f"MAE: {mae:.3f}\nMSE: {mse:.3f}\nRMSE: {rmse:.3f}\nR2: {r2:.3f}"
f" \nTime: {t1:.3} seconds"
)
# -
# #### w. TargetTransformerClass
#
# **Note**: This does not give confidence intervals. I will have to modify the code-base later for this to work.
# +
# define kernel function
kernel = ConstantKernel() * RBF() + WhiteKernel()
# define GP model
gp_model = GaussianProcessRegressor(
kernel=kernel, # kernel function (very important)
normalize_y=True, # good standard practice
random_state=123, # reproducibility
n_restarts_optimizer=10, # good practice (avoids local minima)
)
# Define target transformer
pca_model = PCA(n_components=3)
# Define Wrapper for target transformation
full_regressor = TransformedTargetRegressor(
regressor=gp_model,
transformer=pca_model, # same number of components as informative
check_inverse=False # PCA is not a direct inverse transformation
)
# train GP Model
t0 = time.time()
full_regressor.fit(xtrain, ytrain)
t1 = time.time() - t0
# Predictions
ypred = full_regressor.predict(xtest)
# Get Stats
mae = mean_absolute_error(ypred, ytest)
mse = mean_squared_error(ypred, ytest)
rmse = np.sqrt(mse)
r2 = r2_score(ypred, ytest)
print(
f"MAE: {mae:.3f}\nMSE: {mse:.3f}\nRMSE: {rmse:.3f}\nR2: {r2:.3f}"
f" \nTime: {t1:.3} seconds"
)
# -
# ## GP - Multitask w. PCA Transformer
# **Note**: still a working progress...
# +
# define kernel function
kernel = ConstantKernel() * RBF() + WhiteKernel()
# define GP model
gp_model = GaussianProcessRegressor(
kernel=kernel, # kernel function (very important)
normalize_y=True, # good standard practice
random_state=123, # reproducibility
n_restarts_optimizer=10, # good practice (avoids local minima)
)
# Define Multioutput function
gp_model_multi = MultiOutputRegressor(
gp_model,
n_jobs=1, # Number of cores to use to parallelize the training
)
# Define target transformer
pca_model = PCA(n_components=3)
# Define Wrapper for target transformation
full_regressor = TransformedTargetRegressor(
regressor=gp_model_multi,
transformer=pca_model, # same number of components as informative
check_inverse=False # PCA is not a direct inverse transformation
)
# Fit Model
t0 = time.time()
full_regressor.fit(xtrain, ytrain)
t1 = time.time() - t0
# Predict with test set
ypred = full_regressor.predict(xtest)
# Get Stats
mae = mean_absolute_error(ypred, ytest)
mse = mean_squared_error(ypred, ytest)
rmse = np.sqrt(mse)
r2 = r2_score(ypred, ytest)
print(
f"GP Model:\n"
f"MAE: {mae:.3f}\nMSE: {mse:.3f}\nRMSE: {rmse:.3f}\nR2: {r2:.3f}"
f" \nTime: {t1:.3} seconds"
)
# -
# Interestingly enough, we got worse results for this model than the previous. Probably because of the uninformative features for each layer. It makes me skeptical to use the multi-task GP.
| notebooks/na_data/demos/full_gp_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bqplot import DateScale, LinearScale, ColorScale, Axis, ColorAxis, Figure, FlexLine
import numpy as np
# ## Get Data
dates = np.arange("2005-02", "2005-03", dtype="datetime64[D]")
size = len(dates)
spx = 100 + 5 * np.cumsum(np.random.randn(size))
vix = 10 + np.cumsum(np.random.randn(size))
# ## Displaying extra dimension with color
# +
lin_x = DateScale()
lin_y = LinearScale()
col_line = ColorScale(colors=["green", "white", "red"])
ax_x = Axis(scale=lin_x, label="Date", label_location="end")
ax_y = Axis(scale=lin_y, orientation="vertical", label="Index", label_offset="4ex")
ax_col = ColorAxis(label="Vol", scale=col_line, tick_format="0.2f")
fig_margin = dict(top=50, left=80, right=20, bottom=70)
fl = FlexLine(
x=dates, y=spx, color=vix, scales={"x": lin_x, "color": col_line, "y": lin_y}
)
Figure(marks=[fl], axes=[ax_x, ax_y, ax_col], fig_margin=fig_margin)
# -
# ## Displaying extra dimension with width
# +
lin_x = DateScale()
lin_y = LinearScale()
width_line = LinearScale()
ax_x = Axis(scale=lin_x, label="Date")
ax_y = Axis(scale=lin_y, orientation="vertical", label="SPX Index")
fl2 = FlexLine(
x=dates,
y=spx,
width=vix,
scales={"x": lin_x, "width": width_line, "y": lin_y},
stroke_width=5,
)
Figure(marks=[fl2], axes=[ax_x, ax_y])
| examples/Marks/Object Model/FlexLine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# Notebook to analyze the **False Negatives** Results for each model in the context of traceability between features and bug reports.
# + [markdown] toc-hr-collapsed=false
# # Load Libraries and Datasets
# + jupyter={"source_hidden": true}
from mod_finder_util import mod_finder_util
mod_finder_util.add_modules_origin_search_path()
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib_venn import venn3
from modules.models_runner import feat_br_strat_runner
from modules.utils import firefox_dataset_p2 as fd
from modules.utils import aux_functions
from modules.utils import tokenizers as tok
import warnings; warnings.simplefilter('ignore')
# -
# # Run All Models
# ## Volunteers Only Strategy
# + jupyter={"source_hidden": true}
vol_strat_runner = feat_br_strat_runner.Feat_BR_Vol_Strat_Runner()
vol_strat_runner.execute()
oracle = vol_strat_runner.get_oracle()
evaluator_4 = vol_strat_runner.get_evaluator()
evals_df_4 = vol_strat_runner.get_evals_df()
# -
# ### Load Datasets
# + jupyter={"outputs_hidden": true}
bugreports = fd.Datasets.read_selected_bugreports_df()
features = fd.Datasets.read_features_df()
# -
# ### Tokenization
tokenizer = tok.PorterStemmerBased_Tokenizer()
bugreports['tokens'] = bugreports.apply(lambda row : tokenizer.__call__(row['br_desc']), axis=1)
features['tokens'] = features.apply(lambda row : tokenizer.__call__(row['feat_desc']), axis=1)
# ### Results
# + jupyter={"outputs_hidden": true}
evals_df_4.head()
# -
# ### Grouping Results by Model
# + [markdown] toc-hr-collapsed=true
# #### Min Recall
# + jupyter={"source_hidden": true}
group = evals_df_4.groupby('model').perc_recall.min()
print(group)
print()
bm25_min_recall = group[group.index == 'bm25'].values[0]
lsi_min_recall = group[group.index == 'lsi'].values[0]
lda_min_recall = group[group.index == 'lda'].values[0]
wv_min_recall = group[group.index == 'wordvector'].values[0]
bm25_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='bm25', perc_recall=bm25_min_recall))
lsi_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lsi', perc_recall=lsi_min_recall))
lda_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lda', perc_recall=lda_min_recall))
wv_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='wordvector', perc_recall=wv_min_recall))
venn3([bm25_fn_set, lsi_fn_set, lda_fn_set], ['BM25','LSI','LDA'])
plt.title('Comparison False Negatives by Model (BM25, LSI, LDA) - Min Recall')
plt.show()
venn3([bm25_fn_set, wv_fn_set, lda_fn_set], ['BM25','WV','LDA'])
plt.title('Comparison False Negatives by Model (BM25, WV, LDA) - Min Recall')
plt.show()
venn3([lsi_fn_set, wv_fn_set, lda_fn_set], ['LSI','WV','LDA'])
plt.title('Comparison False Negatives by Model (LSI, WV, LDA) - Min Recall')
plt.show()
venn3([lsi_fn_set, wv_fn_set, bm25_fn_set], ['LSI','WV','BM25'])
plt.title('Comparison False Negatives by Model (LSI, WV, BM25) - Min Recall')
plt.show()
# -
# ##### Exclusive Cases
# +
print("BM25 Exclusive FN:")
bm25_exc_set = bm25_fn_set - lsi_fn_set - lda_fn_set - wv_fn_set
#display(bm25_exc_set)
print("len(bm25_exc_set): {}".format(len(bm25_exc_set)))
print("\n\nLSI Exclusive FN:")
lsi_exc_set = lsi_fn_set - bm25_fn_set - lda_fn_set - wv_fn_set
#display(lsi_exc_set)
print("len(lsi_exc_set): {}".format(len(lsi_exc_set)))
print("\n\nLDA Exclusive FN:")
lda_exc_set = lda_fn_set - lsi_fn_set - bm25_fn_set - wv_fn_set
#display(lda_exc_set)
print("len(lda_exc_set): {}".format(len(lda_exc_set)))
print("\n\nWV Exclusive FN:")
wv_exc_set = wv_fn_set - lda_fn_set - lsi_fn_set - bm25_fn_set
#display(wv_exc_set)
print("len(wv_exc_set): {}".format(len(wv_exc_set)))
# -
# ##### Word Clouds
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#aux_functions.create_wordcloud_feat_br(bm25_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - BM25",
# wc_br_title="FP - Bug Reports - BM25")
#aux_functions.create_wordcloud_feat_br(lsi_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - LSI",
# wc_br_title="FP - Bug Reports - LSI")
aux_functions.create_wordcloud_feat_br(lda_exc_set, bugreports=bugreports, features=features,
wc_feat_title="FP - Features - LDA",
wc_br_title="FP - Bug Reports - LDA")
#aux_functions.create_wordcloud_feat_br(wv_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - Word Vector",
# wc_br_title="FP - Bug Reports - Word Vector")
# -
# ##### False Negatives Amount by Model
print("LSI FN Amount: {}".format(len(lsi_fn_set)))
print("LDA FN Amount: {}".format(len(lda_fn_set)))
print("BM25 FN Amount: {}".format(len(bm25_fn_set)))
print("WV FN Amount: {}".format(len(wv_fn_set)))
# + [markdown] toc-hr-collapsed=true
# #### Max Recall
# + jupyter={"source_hidden": true}
group = evals_df_4.groupby('model').perc_recall.max()
print(group)
print()
bm25_max_recall = group[group.index == 'bm25'].values[0]
lsi_max_recall = group[group.index == 'lsi'].values[0]
lda_max_recall = group[group.index == 'lda'].values[0]
wv_max_recall = group[group.index == 'wordvector'].values[0]
bm25_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='bm25', perc_recall=bm25_max_recall))
lsi_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lsi', perc_recall=lsi_max_recall))
lda_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lda', perc_recall=lda_max_recall))
wv_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='wordvector', perc_recall=wv_max_recall))
venn3([bm25_fn_set, lsi_fn_set, lda_fn_set], ['BM25','LSI','LDA'])
plt.title('Comparison False Negatives by Model (BM25, LSI, LDA) - Max Recall')
plt.show()
venn3([bm25_fn_set, wv_fn_set, lda_fn_set], ['BM25','WV','LDA'])
plt.title('Comparison False Negatives by Model (Bm25, WV, LDA) - Max Recall')
plt.show()
venn3([lsi_fn_set, wv_fn_set, lda_fn_set], ['LSI','WV','LDA'])
plt.title('Comparison False Negatives by Model (LSI, WV, LDA) - Max Recall')
plt.show()
venn3([lsi_fn_set, wv_fn_set, bm25_fn_set], ['LSI','WV','BM25'])
plt.title('Comparison False Negatives by Model (LSI, WV, BM25) - Max Recall')
plt.show()
# -
# ##### Exclusive Cases
# +
print("BM25 Exclusive FN:")
bm25_exc_set = bm25_fn_set - lsi_fn_set - lda_fn_set - wv_fn_set
#display(bm25_exc_set)
print("len(bm25_exc_set): {}".format(len(bm25_exc_set)))
print("\n\nLSI Exclusive FN:")
lsi_exc_set = lsi_fn_set - bm25_fn_set - lda_fn_set - wv_fn_set
#display(lsi_exc_set)
print("len(lsi_exc_set): {}".format(len(lsi_exc_set)))
print("\n\nLDA Exclusive FN:")
lda_exc_set = lda_fn_set - lsi_fn_set - bm25_fn_set - wv_fn_set
#display(lda_exc_set)
print("len(lda_exc_set): {}".format(len(lda_exc_set)))
print("\n\nWV Exclusive FN:")
wv_exc_set = wv_fn_set - lda_fn_set - lsi_fn_set - bm25_fn_set
#display(wv_exc_set)
print("len(wv_exc_set): {}".format(len(wv_exc_set)))
# -
# ##### Word Clouds
# + jupyter={"outputs_hidden": true, "source_hidden": true}
#aux_functions.create_wordcloud_feat_br(bm25_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - BM25",
# wc_br_title="FP - Bug Reports - BM25")
#aux_functions.create_wordcloud_feat_br(lsi_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - LSI",
# wc_br_title="FP - Bug Reports - LSI")
aux_functions.create_wordcloud_feat_br(lda_exc_set, bugreports=bugreports, features=features,
wc_feat_title="FP - Features - LDA",
wc_br_title="FP - Bug Reports - LDA")
#aux_functions.create_wordcloud_feat_br(wv_exc_set, bugreports=bugreports, features=features,
# wc_feat_title="FP - Features - Word Vector",
# wc_br_title="FP - Bug Reports - Word Vector")
# -
# ##### False Negatives Amount by Model
print("LSI FN Amount: {}".format(len(lsi_fn_set)))
print("LDA FN Amount: {}".format(len(lda_fn_set)))
print("BM25 FN Amount: {}".format(len(bm25_fn_set)))
print("WV FN Amount: {}".format(len(wv_fn_set)))
# + [markdown] toc-hr-collapsed=true
# #### Min Precision
# + jupyter={"outputs_hidden": true, "source_hidden": true}
group = evals_df_4.groupby('model').perc_precision.min()
print(group)
print()
bm25_min_prec = group[group.index == 'bm25'].values[0]
lsi_min_prec = group[group.index == 'lsi'].values[0]
lda_min_prec = group[group.index == 'lda'].values[0]
wv_min_prec = group[group.index == 'wordvector'].values[0]
bm25_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='bm25', perc_precision=bm25_min_prec))
lsi_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lsi', perc_precision=lsi_min_prec))
lda_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lda', perc_precision=lda_min_prec))
wv_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='wordvector', perc_precision=wv_min_prec))
venn3([bm25_fn_set, lsi_fn_set, lda_fn_set], ['BM25','LSI','LDA'])
plt.title('Comparison False Negatives by Model (BM25, LSI, LDA) - Min Precision')
plt.show()
venn3([bm25_fn_set, wv_fn_set, lda_fn_set], ['BM25','WV','LDA'])
plt.title('Comparison False Negatives by Model (BM25, WV, LDA) - Min Precision')
plt.show()
venn3([lsi_fn_set, wv_fn_set, lda_fn_set], ['LSI','WV','LDA'])
plt.title('Comparison False Negatives by Model (LSI, WV, LDA) - Min Precision')
plt.show()
venn3([lsi_fn_set, wv_fn_set, bm25_fn_set], ['LSI','WV','BM25'])
plt.title('Comparison False Negatives by Model (LSI, WV, BM25) - Min Precision')
plt.show()
# + [markdown] jupyter={"source_hidden": true}
# ##### Exclusive Cases
# + jupyter={"outputs_hidden": true, "source_hidden": true}
print("BM25 Exclusive FN:")
display(bm25_fn_set - lsi_fn_set - lda_fn_set - wv_fn_set)
print("\n\nLSI Exclusive FN:")
display(lsi_fn_set - bm25_fn_set - lda_fn_set - wv_fn_set)
print("\n\nLDA Exclusive FN:")
display(lda_fn_set - lsi_fn_set - bm25_fn_set - wv_fn_set)
print("\n\nWV Exclusive FN:")
display(wv_fn_set - lda_fn_set - lsi_fn_set - bm25_fn_set)
# + jupyter={"outputs_hidden": true, "source_hidden": true}
print("LSI FN Amount: {}".format(len(lsi_fn_set)))
print("LDA FN Amount: {}".format(len(lda_fn_set)))
print("BM25 FN Amount: {}".format(len(bm25_fn_set)))
print("WV FN Amount: {}".format(len(wv_fn_set)))
# + [markdown] toc-hr-collapsed=true
# #### Max Precision
# + jupyter={"outputs_hidden": true, "source_hidden": true}
group = evals_df_4.groupby('model').perc_precision.max()
print(group)
print()
bm25_max_prec = group[group.index == 'bm25'].values[0]
lsi_max_prec = group[group.index == 'lsi'].values[0]
lda_max_prec = group[group.index == 'lda'].values[0]
wv_max_prec = group[group.index == 'wordvector'].values[0]
bm25_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='bm25', perc_precision=bm25_max_prec))
lsi_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lsi', perc_precision=lsi_max_prec))
lda_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='lda', perc_precision=lda_max_prec))
wv_fn_set = aux_functions.get_false_negatives(oracle, aux_functions.get_trace_links_df(evaluations_df=evals_df_4, model='wordvector', perc_precision=wv_max_prec))
venn3([bm25_fn_set, lsi_fn_set, lda_fn_set], ['BM25','LSI','LDA'])
plt.title('Comparison False Negatives by Model (BM25, LSI, LDA) - Max Precision')
plt.show()
venn3([bm25_fn_set, wv_fn_set, lda_fn_set], ['BM25','WV','LDA'])
plt.title('Comparison False Negatives by Model (BM25, WV, LDA) - Max Precision')
plt.show()
venn3([lsi_fn_set, wv_fn_set, lda_fn_set], ['LSI','WV','LDA'])
plt.title('Comparison False Negatives by Model (LSI,WV,LDA) - Max Precision')
plt.show()
venn3([lsi_fn_set, wv_fn_set, bm25_fn_set], ['LSI','WV','BM25'])
plt.title('Comparison False Negatives by Model (LSI, WV, BM25) - Max Precision')
plt.show()
# + [markdown] jupyter={"source_hidden": true}
# ##### Exclusive Cases
# + jupyter={"outputs_hidden": true, "source_hidden": true}
print("BM25 Exclusive FN:")
display(bm25_fn_set - lsi_fn_set - lda_fn_set - wv_fn_set)
print("\n\nLSI Exclusive FN:")
display(lsi_fn_set - bm25_fn_set - lda_fn_set - wv_fn_set)
print("\n\nLDA Exclusive FN:")
display(lda_fn_set - lsi_fn_set - bm25_fn_set - wv_fn_set)
print("\n\nWV Exclusive FN:")
display(wv_fn_set - lda_fn_set - lsi_fn_set - bm25_fn_set)
# + jupyter={"outputs_hidden": true, "source_hidden": true}
print("LSI FN Amount: {}".format(len(lsi_fn_set)))
print("LDA FN Amount: {}".format(len(lda_fn_set)))
print("BM25 FN Amount: {}".format(len(bm25_fn_set)))
print("WV FN Amount: {}".format(len(wv_fn_set)))
| notebooks/firefox_p2/feat_br_tracing/feat_br_fn_analysis.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// # The Basics
//
// This notebook illustrates the basics of the language, and how to use the Jupyter notebook with C++
// <div style="background: #efffed;
// border: 1px solid grey;
// margin: 8px 0 8px 0;
// text-align: center;
// padding: 8px; ">
// <i class="fa-play fa"
// style="font-size: 40px;
// line-height: 40px;
// margin: 8px;
// color: #444;">
// </i>
// <div>
// To run the selected code cell, hit <pre style="background: #efffed">Shift + Enter</pre>
// </div>
// </div>
// The `#include` directive brings definitions of functions and classes not defined by the language but in different libraries. Cells containing `#include` directives should not contain any definition or declaration.
#include <iostream>
#include <cstddef>
// `std::cout` is used to print text or the value of a variable. It is required to include `<iostream>` before using it, so the previous cell must have been executed.
//
std::cout << "This is a text" << std::endl;
int a = 2;
std::cout << a << std::endl;
// Omitting the `;` in the last statement of a cell results in an output being printed. Notice that this is specific to the notebook, this is NOT valid C++.
a
// - To insert a cell above (resp. below) the current cell, quit the editing mode (by hiting the ESC key) and hit `a` (resp `b`).
// - To delete the current cell, quit the editing mode and hit `d` twice.
// - Executing the last cell of the notebook always creates a new cell after.
| basics/basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.9 ('tfp')
# language: python
# name: python3
# ---
import scvelo as scv
scv.settings.verbosity = 0
dataset = '../data/Pancreas/endocrinogenesis_day15.h5ad'
label = 'clusters'
exp_metrics = {}
import unitvelo as utv
cluster_edges = [
("Pre-endocrine", "Alpha"),
("Pre-endocrine", "Beta"),
("Pre-endocrine", "Delta"),
("Pre-endocrine", "Epsilon")]
# ## scVelo stochastic
# +
title = 'scVelo stochastic mode'
adata = scv.datasets.pancreas()
adata.uns['datapath'] = dataset
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000)
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.velocity(adata, mode='stochastic')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, color=label, title=title)
# -
scv.pp.neighbors(adata)
adata_velo = adata[:, adata.var.loc[adata.var['velocity_genes'] == True].index]
exp_metrics["model_dyn"] = utv.evaluate(adata_velo, cluster_edges, label, 'velocity')
# ## scVelo dynamic
# +
title = 'scVelo dynamical mode'
adata = scv.datasets.pancreas()
adata.uns['datapath'] = dataset
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000)
scv.pp.moments(adata, n_pcs=30, n_neighbors=30)
scv.tl.recover_dynamics(adata, n_jobs=20)
scv.tl.velocity(adata, mode='dynamical')
scv.tl.velocity_graph(adata)
scv.pl.velocity_embedding_stream(adata, color=label, title=title)
# -
scv.pp.neighbors(adata)
adata_velo = adata[:, adata.var.loc[adata.var['velocity_genes'] == True].index]
exp_metrics["model_dyn"] = utv.evaluate(adata_velo, cluster_edges, label, 'velocity')
# ## UniTVelo
velo_config = utv.config.Configuration()
velo_config.R2_ADJUST = False
velo_config.IROOT = None
velo_config.FIT_OPTION = '2'
velo_config.ASSIGN_POS_U = True
adata = utv.run_model(dataset, label, config_file=velo_config)
scv.pl.velocity_embedding_stream(adata, color=adata.uns['label'], title='')
scv.pp.neighbors(adata)
adata_velo = adata[:, adata.var.loc[adata.var['velocity_genes'] == True].index]
exp_metrics["model_dyn"] = utv.evaluate(adata_velo, cluster_edges, label, 'velocity')
| notebooks/SuppFig6_Pancreas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests, json, io, urllib
from zipfile import ZipFile
#import zipfile
#from contextlib import closing
import urllib.request as ur
import pandas as pd
import subprocess
from simpledbf import Dbf5
from bis2 import gc2
import string
q='''https://www.sciencebase.gov/catalog/items?filter=tags={%22scheme%22:%22BIS%22,%22name%22:%22NHDPlusV2.1%22}&fields=files,id,tags&format=json'''
#Returns
nhdItems = requests.get(q).json()
# Set up the actions/targets for this particular instance
thisRun = {}
thisRun["instance"] = "DataDistillery"
thisRun["db"] = "BCB"
thisRun["baseURL"] = gc2.sqlAPI(thisRun["instance"],thisRun["db"])
thisRun["schema"] = "sfr"
thisRun["commitToDB"] = False
#Query comids already in the table
q_recordToSearch = "SELECT comid as lookup FROM nhd.nhdplusv2_plusflowlinevaa "
recordToSearch = requests.get(thisRun["baseURL"]+"&q="+q_recordToSearch).json()
p = recordToSearch['features']
lookup = []
#for record returned, add record to list
for f in p:
lookup.append(f['properties']['lookup'])
lenList = len(lookup)
cntExisting = 0
existingList = []
for item in nhdItems['items']:
#Identify which NHD Region via tags
for tag in item['tags']:
if 'Reg' in tag['name']:
region = tag['name']
#Look at files and find NHDPlusAttributes File
for file in item['files']:
fileName = file['name']
if 'NHDPlusAttributes' in fileName:
#Download file
print ('Retrieving region ' + region + ', file:' + fileName)
fileUrl = file['url']
ur.urlretrieve(fileUrl, fileName)
#Unzip file. Ideally this will be transformed to work in memory
subprocess.call(r'"C:\Program Files\7-Zip\7z.exe" x ' + fileName)
#Convert dbf to dataframe
dbf = Dbf5('PlusFlowLineVAA.dbf')
df = dbf.to_dataframe()
for row in df.itertuples():
if row.ComID in lookup:
continue
else:
q = "insert into nhd.nhdplusv2_plusflowlinevaa \
(comid, fdate, streamleve, streamorde, streamcalc, fromnode, tonode, hydroseq, levelpath1, \
pathlength, terminalpat, arbolatesu, divergence, startflag, terminalfl, dnlevel, thinnercod, \
uplevelpat, uphydroseq, dnlevelpat, dnminorhyd, dndraincou, dnhydroseq, frommeas, tomeas, \
reachcode, lengthkm, fcode, rtndiv, outdiv, diveeffect, vpuin, vpuout, areasqkm, totdasqkm, \
divdasqkm, tidal, totma, wbareatype) VALUES \
('" + str(row.ComID) + "' ,'" + str(row.Fdate) + "' ,'" + str(row.StreamLeve) + "' ,'" \
+ str(row.StreamOrde) + "' ,'" + str(row.StreamCalc) + "' ,'" + str(row.FromNode) + "' ,'" \
+ str(row.ToNode) + "' ,'" + str(row.Hydroseq) + "' ,'" + str(row.LevelPathI) + "' ,'" \
+ str(row.Pathlength)+ "' ,'" + str(row.TerminalPa) + "' ,'" + str(row.ArbolateSu) + "' ,'" \
+ str(row.Divergence) + "' ,'" + str(row.StartFlag) + "' ,'" + str(row.TerminalFl) + "' ,'" \
+ str(row.DnLevel) + "' ,'" + str(row.ThinnerCod) + "' ,'" + str(row.UpLevelPat) + "' ,'" \
+ str(row.UpHydroseq) + "' ,'" + str(row.DnLevelPat) + "' ,'" + str(row.DnMinorHyd) + "' ,'" \
+ str(row.DnDrainCou) + "' ,'" + str(row.DnHydroseq) + "' ,'" + str(row.FromMeas) + "' ,'" \
+ str(row.ToMeas) + "' ,'" + str(row.ReachCode) + "' ,'" + str(row.LengthKM) + "' ,'" \
+ str(row.Fcode) + "' ,'" + str(row.RtnDiv) + "' ,'" + str(row.OutDiv) + "' ,'" \
+ str(row.DivEffect) + "' ,'" + str(row.VPUIn) + "' ,'" + str(row.VPUOut) + "' ,'" \
+ str(row.AreaSqKM) + "' ,'" + str(row.TotDASqKM) + "' ,'" + str(row.DivDASqKM) + "' ,'" \
+ str(row.Tidal) + "' ,'" + str(row.TOTMA) + "' ,'" + str(row.WBAreaType) + "')"
gc2Key = gc2.gc2Keys["datadistillery_bcb"]
payload = "q=%s&key=%s"%(q,gc2Key)
url= gc2.baseURLs["sqlapi_datadistillery_bcb"]
finalUrl = url + '?q=' +q + '&key=' + gc2Key
try:
#r = requests.post(url,data=payload)
r = requests.get(finalUrl)
#r.get(finalUrl).json()
status = str(r.status_code)
if r.status_code == 200:
continue
else:
print (str(row.ComID) + ' failed with code: ' + status)
except requests.exceptions.HTTPError:
print (str(row.ComID) + ' failed with code: ' + status)
# +
import requests, json, io, urllib
from zipfile import ZipFile
import zipfile
#from contextlib import closing
import urllib.request as ur
import pandas as pd
import subprocess
from simpledbf import Dbf5
from bis2 import gc2
import string
import tempfile
import os
q='''https://www.sciencebase.gov/catalog/items?filter=tags={%22scheme%22:%22BIS%22,%22name%22:%22NHDPlusV2.1%22}&fields=files,id,tags&format=json'''
#Returns
nhdItems = requests.get(q).json()
# Set up the actions/targets for this particular instance
thisRun = {}
thisRun["instance"] = "DataDistillery"
thisRun["db"] = "BCB"
thisRun["baseURL"] = gc2.sqlAPI(thisRun["instance"],thisRun["db"])
thisRun["schema"] = "sfr"
thisRun["commitToDB"] = False
dfAll = None
for item in nhdItems['items']:
#Identify which NHD Region via tags
for tag in item['tags']:
if 'Reg' in tag['name']:
region = tag['name']
regFolder = '/' + region
if not os.path.exists(regFolder):
os.makedirs(regFolder)
os.chdir(regFolder)
#Look at files and find NHDPlusAttributes File
for file in item['files']:
fileName = file['name']
if 'NHDPlusAttributes' in fileName:
print ('Retrieving region ' + region + ', file:' + fileName)
fileUrl = file['url']
ur.urlretrieve(fileUrl, fileName)
subprocess.call(r'"C:\Program Files\7-Zip\7z.exe" x ' + fileName)
#with zipfile.ZipFile(fileName, "r") as zip_ref:
# zip_ref.extractall(tmpdirname)
# zip_ref.close()
#Convert dbf to dataframe
dbf = Dbf5('PlusFlowLineVAA.dbf')
df = dbf.to_dataframe()
if dfAll is None:
dfAll = df
else:
dfAll.append(df)
# +
import requests, json, io, urllib
from zipfile import ZipFile
import zipfile
#from contextlib import closing
import urllib.request as ur
import pandas as pd
import subprocess
from simpledbf import Dbf5
from bis2 import gc2
import string
import tempfile
import os
q='''https://www.sciencebase.gov/catalog/items?filter=tags={%22scheme%22:%22BIS%22,%22name%22:%22NHDPlusV2.1%22}&fields=files,id,tags&format=json'''
#Returns
nhdItems = requests.get(q).json()
# Set up the actions/targets for this particular instance
thisRun = {}
thisRun["instance"] = "DataDistillery"
thisRun["db"] = "BCB"
thisRun["baseURL"] = gc2.sqlAPI(thisRun["instance"],thisRun["db"])
thisRun["schema"] = "sfr"
thisRun["commitToDB"] = False
dfAll = None
for item in nhdItems['items']:
#Identify which NHD Region via tags
for tag in item['tags']:
if 'Reg' in tag['name']:
region = tag['name']
if not os.path.exists('proc'):
os.makedirs('proc')
os.chdir('proc')
#Look at files and find NHDPlusAttributes File
for file in item['files']:
fileName = file['name']
if 'NHDPlusAttributes' in fileName:
print ('Retrieving region ' + region + ', file:' + fileName)
fileUrl = file['url']
ur.urlretrieve(fileUrl, fileName)
subprocess.call(r'"C:\Program Files\7-Zip\7z.exe" x ' + fileName)
#with zipfile.ZipFile(fileName, "r") as zip_ref:
# zip_ref.extractall(tmpdirname)
# zip_ref.close()
#Convert dbf to dataframe
dbf = Dbf5('PlusFlowLineVAA.dbf')
dbf = dbf.to_dataframe()
if dfAll is None:
dfAll = df
else:
dfAll.append(df)
#df = None
#os.remove('NHDPlusAttributes.zip')
#os.remove('PlusFlow.dbf')
#os.remove('PlusFlowLineVAA.dbf')
#os.remove('elevslope.dbf')
# -
dfAll.size
import tempfile
with tempfile.TemporaryDirectory() as tmpdirname:
if 'NHDPlusAttributes' in fileName:
#Download file
print ('Retrieving region ' + region + ', file:' + fileName)
fileUrl = file['url']
ur.urlretrieve(fileUrl, fileName)
df.head()
#zFile = ZipFile(fileName, 'r')
#pd.read_table(zFile.open('PlusFlow.dbf'))
#if 'Hydrography' in fileName:
# subprocess.call(r'"C:\Program Files\7-Zip\7z.exe" x ' + fileName)
#Gave error BadZipFile: File is not a zip file
#mysock = urllib.request.urlopen(fileUrl)
#memfile = io.BytesIO(mysock.read())
#ZipFile.infolist(memfile)
#with ZipFile(memfile, 'r') as ftpl:
#ftpl.namelist()
#for file in flist:
# test = ftpl.extract(file)
# print (test)
#Gave error BadZipFile: File is not a zip file
#r = requests.get(fileUrl)
#with closing(r), ZipFile(io.BytesIO(r.content)) as archive:
# print({member.filename: archive.read(member) for member in archive.infolist()})
print (finalurl)
for item in nhdFiles['items']:
for tag in item['tags']:
if 'Reg' in tag['name']:
region = tag['name']
print (region)
print (nhdFiles)
# +
for item:
for file in files:
unzip file into memory
if fileName = Flowline*
convert correct file to dataframe
# +
# Set up the actions/targets for this particular instance
thisRun = {}
thisRun["instance"] = "DataDistillery"
thisRun["db"] = "BCB"
thisRun["baseURL"] = gc2.sqlAPI(thisRun["instance"],thisRun["db"])
thisRun["schema"] = "sfr"
thisRun["commitToDB"] = False
#Query comids already in the table
q_recordToSearch = "SELECT comid as lookup FROM nhd.nhdplusv2_plusflowlinevaa "
recordToSearch = requests.get(thisRun["baseURL"]+"&q="+q_recordToSearch).json()
p = recordToSearch['features']
lookup = []
#for record returned, add record to list
for f in p:
string = f['properties']['lookup']
if string.startswith('"') and string.endswith('"'):
string = string[1:-1] #For some reason these values are coming back with double quotes, this removes outer quote
lookup.append(string)
lenList = len(lookup)
cntExisting = 0
existingList = []
# -
print (lookup)
| GC2DD_Insert/nhdplusv2/NHDPlusV21_To_GC2DD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sysrevenv
# language: python
# name: sysrevenv
# ---
# # Barplot of README page content
# Import the libraries for README page analysis.
import matplotlib.pylab as plt
import pandas as pd
import numpy as np
import seaborn as sns
# Import the CSV file for readme pages
readme_pages = pd.read_csv('data/readme_pages.csv', header=0)
# Sum the number of elemenets across organizations
readme_pages_totals = readme_pages.sum()
print(readme_pages_totals)
# Remove the header since we will replace the header in the next step.
readme_new_labels = readme_pages_totals.drop("organization_or_project_name")
readme_new_labels.head()
# Convert the series to a dataframe, and add in new header for plotting.
readme_pages_df = pd.DataFrame({'README_elements':readme_new_labels.index, 'count':readme_new_labels.values})
readme_pages_df
# Calculate the percent of each element compared to overall number of organizations.
readme_pages_df['percent'] = (readme_pages_df['count'] / (len(readme_pages.index))) * 100
print(readme_pages_df)
# Arrange dataframe into descending order for plotting.
result = readme_pages_df.sort_values('percent', ascending=False)
print(result)
# Make plot using `seaborn` library
sns.set_style("white")
sns.set_context("paper", font_scale=1.7)
readme_barplot = sns.barplot(x="README_elements",
y = "percent",
data = readme_pages_df,
color="#482677FF",
order = result['README_elements'])
readme_barplot.set(xlabel = "README page elements",
ylabel = "Percent of READMEs")
readme_barplot.set_xticklabels(readme_barplot.get_xticklabels(),
rotation=90)
readme_barplot.set_xticklabels(['about','contribute','citation','license','version','getting started','funding','visual structure','history','resources'])
sns.despine(top=True)
plt.tight_layout(h_pad=2)
plt.ylim(0,100)
#plt.savefig('figures/README_content.jpg', dpi=300)
| readme_page_analysis.ipynb |