code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ml)
# language: python
# name: ml
# ---
# ## Exploring Activations
# I want to look at the activations generated by our network. This is mostly just out of curiosity since I wouldn't expect any big problems with ResNets.
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai.callbacks import ActivationStats, HookCallback
from fastai.vision import *
import pandas as pd
import os
# +
DATA = Path('data')
CSV_TRN_CURATED = DATA/'train_curated.csv'
CSV_TRN_NOISY = DATA/'train_noisy.csv'
CSV_TRN_MERGED = DATA/'train_merged.csv'
CSV_SUBMISSION = DATA/'sample_submission.csv'
TRN_CURATED = DATA/'train_curated'
TRN_NOISY = DATA/'train_noisy'
TEST = DATA/'test'
WORK = Path('work')
IMG_TRN_CURATED = WORK/'image/trn_curated'
IMG_TRN_NOISY = WORK/'image/trn_noisy'
IMG_TRN_MERGED = WORK/'image/trn_merged'
IMG_TEST = WORK/'image/test'
for folder in [WORK, IMG_TRN_CURATED, IMG_TRN_NOISY, IMG_TEST]:
Path(folder).mkdir(exist_ok=True, parents=True)
train_df = pd.read_csv(CSV_TRN_CURATED)
train_noisy_df = pd.read_csv(CSV_TRN_NOISY)
test_df = pd.read_csv(CSV_SUBMISSION)
# +
tfms = get_transforms(do_flip=True, max_rotate=0, max_lighting=0.1, max_zoom=0, max_warp=0.)
src = (ImageList.from_csv(WORK/'image', Path('../../')/CSV_TRN_CURATED, folder='trn_curated', suffix='.jpg')
.split_by_rand_pct(0.2)
.label_from_df(label_delim=',')
)
data = (src.transform(tfms, size=128)
.databunch(bs=64).normalize(imagenet_stats)
)
# +
# Modified from: https://forums.fast.ai/t/confused-by-output-of-hook-output/29514/4
class StoreHook(HookCallback):
def on_train_begin(self, **kwargs):
super().on_train_begin(**kwargs)
self.hists = []
def hook(self, m, i, o):
return o
def on_batch_end(self, train, **kwargs):
if (train):
self.hists.append(self.hooks.stored[0].cpu().histc(40,0,10))
# Simply pass in a learner and the module you would like to instrument
def probeModule(learn, module):
hook = StoreHook(learn, modules=flatten_model(module))
learn.callbacks += [ hook ]
return hook
# Thanks to @ste for initial version of histgram plotting code
def get_hist(h):
return torch.stack(h.hists).t().float().log1p()
# -
f_score = partial(fbeta, thresh=0.2)
learn = cnn_learner(data, models.resnet18, pretrained=False, metrics=[f_score])
learn.unfreeze()
# +
#Hook the output of the first conv and each resnet block.
hooks = [probeModule(learn, learn.model[0][0]),
probeModule(learn, learn.model[0][4]),
probeModule(learn, learn.model[0][5]),
probeModule(learn, learn.model[0][6]),
probeModule(learn, learn.model[0][7])]
names = ['conv1',
'conv2_x',
'conv3_x',
'conv4_x',
'conv5_x']
learn.fit_one_cycle(10, max_lr=1e-2)
# +
fig,axes = plt.subplots(5, figsize=(15,10))
for i, (ax,h) in enumerate(zip(axes.flatten(), hooks)):
ax.imshow(get_hist(h), origin='lower', aspect='auto')
ax.axis('off')
ax.text(0, -5, names[i], bbox={'facecolor':'red', 'alpha':0.0, 'pad':10})
plt.tight_layout()
# +
#Hook the output of the first conv and each resnet block.
hooks = [probeModule(learn, learn.model[0][0]),
probeModule(learn, learn.model[0][4]),
probeModule(learn, learn.model[0][5]),
probeModule(learn, learn.model[0][6]),
probeModule(learn, learn.model[0][7])]
names = ['conv1',
'conv2_x',
'conv3_x',
'conv4_x',
'conv5_x']
learn.fit_one_cycle(10, max_lr=slice(1e-6, 1e-2))
# +
fig,axes = plt.subplots(5, figsize=(15,10))
for i, (ax,h) in enumerate(zip(axes.flatten(), hooks)):
ax.imshow(get_hist(h), origin='lower', aspect='auto')
ax.axis('off')
ax.text(0, -5, names[i], bbox={'facecolor':'red', 'alpha':0.0, 'pad':10})
plt.tight_layout()
# -
| 01_ExploringActivations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import numpy as np
from tqdm.autonotebook import tqdm
import gym
import time
# %matplotlib inline
import matplotlib.pyplot as plt
# -
def moving_average(values, n=100) :
ret = np.cumsum(values, dtype=float)
ret[n:] = ret[n:] - ret[:-n]
return ret[n - 1:] / n
env = gym.make('FrozenLake-v0')
policy_to_action = {0:"L",1:"D",2:"R",3:"U"}
ACTION_DIM = env.action_space.n
MAX_STEPS = env.spec.max_episode_steps
STATE_DIM = env.observation_space.n
NUM_EPISODES = 1000000
START_ALPHA = 0.1
ALPHA_TAPER = 0.01
START_EPSILON = 1
EPSILON_TAPER = 0.0001
GAMMA = 0.9
# +
Q = np.zeros((STATE_DIM,ACTION_DIM),dtype=np.float64)
state_visits_count = {}
update_counts = np.zeros((STATE_DIM,ACTION_DIM),dtype=np.int64)
def updateQ( prev_state,action,reward,cur_state):
alpha = START_ALPHA / ( 1 + update_counts[prev_state][action]*ALPHA_TAPER )
update_counts[prev_state][action] += 1
Q[prev_state][action] += alpha * ( reward + GAMMA * np.max(Q[cur_state]) - Q[prev_state][action] )
def epsilon_greedy(s,eps=START_EPSILON):
if np.random.random() > 1-eps:
return np.argmax(Q[s])
else:
return env.action_space.sample()
# +
total_rewards = 0
deltas = []
verbose = True
start = time.time()
for episode in tqdm(range(NUM_EPISODES),desc = "Progress : "):
eps = START_EPSILON / ( 1.0 + EPSILON_TAPER * episode )
if verbose and episode % (NUM_EPISODES/10) == 0:
print("EPISODES : {} | AVG_REWARD : {} | EPSILON : {}".format(episode,total_rewards/(NUM_EPISODES/10),eps))
total_rewards=0
biggest_change = 0
curr_state = env.reset()
for _ in range(MAX_STEPS):
action = epsilon_greedy(curr_state,eps=eps)
state_visits_count[curr_state] = state_visits_count.get(curr_state,0)+1
prev_state = curr_state
curr_state, reward, done, _ = env.step(action)
total_rewards += reward
oldq = Q[prev_state][action]
updateQ(prev_state,action,reward,curr_state)
biggest_change = max( biggest_change , np.abs( oldq - Q[prev_state][action] ))
if done:
break
deltas.append(biggest_change)
mean_state_visit = np.mean( list(state_visits_count.values()) )
print('EACH STATE WAS VISITED {} TIMES ON AN AVERAGE'.format( mean_state_visit ))
Value_F = np.zeros(STATE_DIM)
Policy_F = np.zeros(STATE_DIM)
for s in range(STATE_DIM):
Value_F[s] = np.max(Q[s])
Policy_F[s] = np.argmax(Q[s])
print("TIME TAKEN {} ".format(time.time()-start))
gpolicy = list(map(lambda a: policy_to_action[a],Policy_F))
print("Optimal Policy :\n {} ".format(np.reshape(gpolicy,(int(np.sqrt(STATE_DIM)),int(np.sqrt(STATE_DIM))))))
print("Optimal Values :\n {}".format(np.reshape(Value_F,(int(np.sqrt(STATE_DIM)),int(np.sqrt(STATE_DIM))))))
plt.plot(moving_average(deltas,n=10000))
plt.show()
# +
"""
Lets see our success rate
"""
games = 1000
won = 0
for _ in range(games):
state = env.reset()
while True:
action = int(Policy_F[state])
(state,reward,is_done,_) = env.step(action)
if is_done:
if reward>0:
won+=1
env.close()
break
print("Success Rate : {}".format(won/games))
# -
| Homework-Assignments/Week 4 - Frozen Lake (Q Learning).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gym
import inventory
import torch
import numpy as np
import matplotlib.pyplot as plt
from stable_baselines3 import PPO
from stable_baselines3.ppo import MlpPolicy
from stable_baselines3.common.env_util import make_vec_env
from stable_baselines3.common.evaluation import evaluate_policy
from stable_baselines3.common.callbacks import EvalCallback
from inventory.envs.inventory_env import Inventory
# +
p = 1
L = 1
load_path = './results/logs2/' + str(p) + "_" + str(L)
policy_data = np.load(load_path + "/evaluations.npz") #load in training data for model
n_episodes = len(policy_data['ep_lengths'][0]) #how many episodes were used to evaluate the policy?
n_evals = len(policy_data['timesteps']) #how many times was the policy evaluated as it was trained?
policy_performance = []
for i in range(0, n_evals): #calculate mean policy performance for each time the policy was evaluated as it was trained
policy_performance.append(-float(sum(policy_data['results'][i])/n_episodes))
const_order_val = (2*p + 1)**(1/2)-1 #policy performance of best constant order policy
plt.plot(policy_data['timesteps'], policy_performance)
plt.xlabel("Number of Learning Steps")
plt.ylabel("Approximate Long Run Average Cost")
plt.axhline(y = const_order_val, color = 'red', linestyle = '--')
plt.title("Trained Policy Performance vs Learning Steps Completed (L = {}, p = {})".format(L,p))
plt.show()
# +
p = 1
L = 1 #only use for L=1!!!
load_path = './results/logs2/' + str(p) + "_" + str(L)
model = PPO.load(load_path + "/best_model.zip")
def g(x,y): #assuming you have I_t = x and x_{1,t} = y, get the deterministic action from trained policy
action, _states = model.predict(np.array([x,y]), deterministic = True)
return action
#evaluate actions on a grid for I_t, x_{1,t} in [0,2]x[0,2]
x = np.linspace(0, 2, 30)
y = np.linspace(0, 2, 30)
z = []
for j in y:
for i in x:
z.append(g(i,j))
X, Y = np.meshgrid(x, y)
Z = np.array(z).reshape(30,30) #reshape for plotting with contourf
plt.contourf(X, Y, Z, 20)
plt.xlabel("I_t (On Hand Inventory)")
plt.ylabel("x_{1,t} (Inventory Order in Pipeline)")
plt.title("Trained Policy Mean Action (L = 1, p = {})".format(p))
plt.colorbar();
# -
| .ipynb_checkpoints/PPO_policy_plotting-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pickle
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
# # Color
# read color features
color_features = pd.read_csv('../data/color_features.csv', sep=',', header=None)
color_features
# create pca
pca_color = PCA(100)
# fit pca
pca_color.fit(color_features)
# save pca model
with open('../data/pca_color.pckl', 'wb') as handle:
pickle.dump(pca_color, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
color_features_pca = pca_color.transform(color_features)
print('dimension', color_features_pca.shape)
# save new features
np.savetxt('../data/color_features_pca.csv', color_features_pca, delimiter=',')
# # Color center subregions
# read color features
color_features = pd.read_csv('../data/color_features_center_subregions.csv', sep=',', header=None)
color_features
# create pca
pca_color = PCA(100)
# fit pca
pca_color.fit(color_features)
# save pca model
with open('../data/pca_color_center_subregions.pckl', 'wb') as handle:
pickle.dump(pca_color, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
color_features_pca = pca_color.transform(color_features)
print('dimension', color_features_pca.shape)
# save new features
np.savetxt('../data/color_features_center_subregions_pca.csv', color_features_pca, delimiter=',')
# # HOG
# read hog features
hog_features = pd.read_csv('../data/hog_features.csv', sep=',', header=None)
hog_features
# create pca
pca_hog = PCA(100)
# fit pca
pca_hog.fit(hog_features)
# save pca model
with open('../data/pca_hog.pckl', 'wb') as handle:
pickle.dump(pca_hog, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
hog_features_pca = pca_hog.transform(hog_features)
print('dimension', hog_features_pca.shape)
# save new features
np.savetxt('../data/hog_features_pca.csv', hog_features_pca, delimiter=',')
# # Neural Network EfficientNet
# read nn features
nn_features = pd.read_csv('../data/nn_features.csv', sep=',', header=None)
nn_features
# create pca
pca_nn = PCA(100)
# fit pca
pca_nn.fit(nn_features)
# save pca model
with open('../data/pca_nn.pckl', 'wb') as handle:
pickle.dump(pca_nn, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
nn_features_pca = pca_nn.transform(nn_features)
print('dimension', nn_features_pca.shape)
# save new features
np.savetxt('../data/nn_features_pca.csv', nn_features_pca, delimiter=',')
# # Neural Network ResNet
# read nn features
nn_features = pd.read_csv('../data/nn_resnet_features.csv', sep=',', header=None)
nn_features
# create pca
pca_nn = PCA(100)
# fit pca
pca_nn.fit(nn_features)
# save pca model
with open('../data/pca_nn_resnet.pckl', 'wb') as handle:
pickle.dump(pca_nn, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
nn_features_pca = pca_nn.transform(nn_features)
print('dimension', nn_features_pca.shape)
# save new features
np.savetxt('../data/nn_resnet_features_pca.csv', nn_features_pca, delimiter=',')
# # HOG + Color
### read HOG features
hog_features = pd.read_csv('../data/HOG_features.csv', sep=',', header=None)
# create pca hog
pca_hc_hog = PCA(100)
# fit pca hog
pca_hc_hog.fit(hog_features)
print('New dimension:', pca_hc_hog.n_components_)
# save pca hog model
with open('../data/pca_hc_hog.pckl', 'wb') as handle:
pickle.dump(pca_hc_hog, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
hog_features_pca = pca_hc_hog.transform(hog_features)
print('dimension', hog_features_pca.shape)
### read color features
color_features = pd.read_csv('../data/color_features.csv', sep=',', header=None)
# create pca color
pca_hc_color = PCA(200)
# fit pca color
pca_hc_color.fit(color_features)
print('New dimension:', pca_hc_color.n_components_)
# save pca color model
with open('../data/pca_hc_color.pckl', 'wb') as handle:
pickle.dump(pca_hc_color, handle, protocol=pickle.HIGHEST_PROTOCOL)
# transform features
color_features_pca = pca_hc_color.transform(color_features)
print('dimension', color_features_pca.shape)
### merge features
merged_features_pca = np.hstack([hog_features_pca, color_features_pca])
print('dimension', merged_features_pca.shape)
# save merged features
np.savetxt('../data/hog_color_features_pca.csv', merged_features_pca, delimiter=',')
# +
# # standardize data
# scaler = StandardScaler()
# scaler.fit(merged)
# +
# # save scaler model
# with open('../data/scaler_std.pckl', 'wb') as handle:
# pickle.dump(scaler, handle, protocol=pickle.HIGHEST_PROTOCOL)
# +
# merged = scaler.transform(merged)
# +
# merged.shape
# +
# pca = PCA(0.95)
# +
# pca.fit(merged)
# +
# # save pca model
# with open('../data/pca_std.pckl', 'wb') as handle:
# pickle.dump(pca, handle, protocol=pickle.HIGHEST_PROTOCOL)
# +
# pca_merged = pca.transform(merged)
# +
# pca_merged.shape
# +
# np.savetxt('../data/merged_color_hog_pca_std.csv', pca_merged, delimiter=',')
| notebooks/FeaturesPCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neighborhood definitions
# In py-clEsperanto (and in [CLIJ as well](https://clij.github.io/clij2-docs/md/neighbors_of_neighbors/)), we're using adjacency graphs to investigate relationships between neighboring labeled object, in practice: cells.
#
# This notebook demonstrates the considered neighborhood definitions.
#
# We consider the "neighborhood" of a pixel always includs the pixel itself per default. That sounds intuitive in the first place and leads to some unnatural behaviour in some situations.
#
# Work is in progress. Feedback is very welcome: robert.haase at tu-dresden.de
# +
import pyclesperanto_prototype as cle
import numpy as np
import matplotlib
from numpy.random import random
cle.select_device("RTX")
# +
# Generate artificial cells as test data
tissue = cle.artificial_tissue_2d()
# fill it with random measurements
values = random([int(cle.maximum_of_all_pixels(tissue))])
for i, y in enumerate(values):
if (i != 95):
values[i] = values[i] * 10 + 45
else:
values[i] = values[i] * 10 + 90
measurements = cle.push(np.asarray([values]))
# visualize measurments in space
example_image = cle.replace_intensities(tissue, measurements)
# -
# ## Example data
# Let's take a look at an image with arbitrarily shaped pixels. Let's call them "cells". In our example image, there is one cell in the center with higher intensity:
cle.imshow(example_image, min_display_intensity=30, max_display_intensity=90, color_map='jet')
# ## Touching neighbors
# We can show all cells that belong to the "touch" neighborhood by computing the local maximum intensity in this neighborhood. Let's visualize the touching neighbor graph as mesh first.
# +
mesh = cle.draw_mesh_between_touching_labels(tissue)
# make lines a bit thicker for visualization purposes
mesh = cle.maximum_sphere(mesh, radius_x=1, radius_y=1)
cle.imshow(mesh)
# -
# From those neighbor-graph one can compute local properties, for example the maximum:
# +
local_maximum = cle.maximum_of_touching_neighbors_map(example_image, tissue)
cle.imshow(local_maximum, min_display_intensity=30, max_display_intensity=90, color_map='jet')
# -
# ## Neighbors of touching neighbors
# You can also extend the neighborhood by considering neighbors of neighbor (of neighbors (of neighbors)). How far you go, can be configured with a radius parameter. Note: Radiu==0 means, no neighbors are taken into account, radius==1 is identical with touching neighbors, radius > 1 are neighbors of neighbors:
for radius in range(0, 5):
local_maximum = cle.maximum_of_touching_neighbors_map(example_image, tissue, radius=radius)
cle.imshow(local_maximum, min_display_intensity=30, max_display_intensity=90, color_map='jet')
# ## N nearest neighbors
# You can also define a neighborhood from the distances between cells. As distance measurement, we use the Euclidean distance between label centroids. Also in this case you an configure how far the neighborhood should range by setting the number of nearest neighbors n. As mentioned above, neighborhoods include the center pixel. Thus, the neighborhood of a pixel and its nearest neighbor contains two neighbors:
for n in range(1, 10):
print("n = ", n)
mesh = cle.draw_mesh_between_n_closest_labels(tissue, n=n)
# make lines a bit thicker for visualization purposes
mesh = cle.maximum_sphere(mesh, radius_x=1, radius_y=1)
cle.imshow(mesh)
for n in range(1, 10):
print("n = ", n)
local_maximum = cle.maximum_of_n_nearest_neighbors_map(example_image, tissue, n=n)
cle.imshow(local_maximum, min_display_intensity=30, max_display_intensity=90, color_map='jet')
# ## Proximal neighbors
# We can also compute the local maximum of cells with centroid distances below a given upper threshold:
local_maximum = cle.maximum_of_proximal_neighbors_map(example_image, tissue, max_distance=20)
cle.imshow(local_maximum, min_display_intensity=30, max_display_intensity=90, color_map='jet')
local_maximum = cle.maximum_of_proximal_neighbors_map(example_image, tissue, max_distance=50)
cle.imshow(local_maximum, min_display_intensity=30, max_display_intensity=90, color_map='jet')
| demo/neighbors/neighborhood_definitions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 単変量線形回帰の予測区間
# +
import numpy as np
from scipy import stats
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
# -
# # 真の回帰直線
# * $y = x + 1$
#
# # 観測値
# * $x = 0.0, 0.5. 1.0, \dots, 10.0$
# * サンプル数: 21
# * $Y_i = x_i + 1 + \varepsilon, \varepsilon \sim N(0, 0.5^2)$
# 実験設定
SAMPLE_SIZE = 21
SIGMA = 0.5
# 実験を管理するクラス
class Experiment:
def __init__(self, random_seed, sigma, sample_size):
np.random.seed(random_seed)
# 実験設定
self.sigma = sigma
self.sample_size = sample_size
# サンプルを生成
self.x_train = np.array([0.5 * i for i in range(sample_size)])
self.y_true = self.x_train + 1
self.y_train = self.y_true + np.random.normal(0.0, sigma, sample_size)
# 回帰係数を算出
self.x_mean = np.mean(self.x_train)
self.s_xx = np.sum((self.x_train - self.x_mean) ** 2)
self.y_mean = np.mean(self.y_train)
self.s_xy = np.sum((self.x_train - self.x_mean) * (self.y_train - self.y_mean))
# 回帰係数
self.coef = self.s_xy / self.s_xx
self.intercept = self.y_mean - self.coef * self.x_mean
# 不偏標本分散
s2 = np.sum((self.y_train - self.intercept - self.coef * self.x_train) ** 2) / (sample_size - 2)
self.s = np.sqrt(s2)
# t分布(自由度N-2)の上側2.5%点
self.t = stats.t.ppf(1-0.025, df=sample_size-2)
# サンプルデータを取得する
def get_sample(self, index):
return (self.x_train[index], self.y_train[index])
# 予測
def predict(self, x):
return self.intercept + self.coef * x
# 真の値
def calc_true_value(self, x):
return x + 1
# 95%信頼区間
def calc_confidence_interval(self, x):
band = self.t * self.s * np.sqrt(1 / self.sample_size + (x - self.x_mean)**2 / self.s_xx)
upper_confidence = self.predict(x) + band
lower_confidence = self.predict(x) - band
return (lower_confidence, upper_confidence)
# 95%予測区間
def calc_prediction_interval(self, x):
band = self.t * self.s * np.sqrt(1 + 1 / self.sample_size + (x - self.x_mean)**2 / self.s_xx)
upper_confidence = self.predict(x) + band
lower_confidence = self.predict(x) - band
return (lower_confidence, upper_confidence)
# 観測値, 95%予測区間を描画する
def plot(self):
# 学習データ
plt.scatter(self.x_train, self.y_train, color='royalblue', alpha=0.2)
# 予測区間
lower_confidence, upper_confidence = self.calc_prediction_interval(self.x_train)
plt.plot(self.x_train, upper_confidence, color='green', linestyle='dashed', label='95% prediction interval')
plt.plot(self.x_train, lower_confidence, color='green', linestyle='dashed')
x_max = max(self.x_train)
plt.xlim([0, x_max])
plt.ylim([0.5, x_max + 1.5])
plt.legend();
# 観測値, 95%予測区間, 95%信頼区間を描画する
def plot_with_confidence(self):
# 学習データ
plt.scatter(self.x_train, self.y_train, color='royalblue', alpha=0.2)
# 信頼区間
lower_confidence, upper_confidence = self.calc_confidence_interval(self.x_train)
plt.plot(self.x_train, upper_confidence, color='royalblue', linestyle='dashed', label='95% confidence interval')
plt.plot(self.x_train, lower_confidence, color='royalblue', linestyle='dashed')
# 予測区間
lower_confidence, upper_confidence = self.calc_prediction_interval(self.x_train)
plt.plot(self.x_train, upper_confidence, color='green', linestyle='dashed', label='95% prediction interval')
plt.plot(self.x_train, lower_confidence, color='green', linestyle='dashed')
x_max = max(self.x_train)
plt.xlim([0, x_max])
plt.ylim([0.5, x_max + 1.5])
plt.legend();
# あるxにおける予測区間と観測値を描画する
def plot_at_x(self, x):
plt.xlim([x-0.5, x+0.5])
plot_x = np.array([x-0.5, x, x+0.5])
plot_y = self.predict(plot_x)
# 学習データ
index = int(2 * x)
plt.scatter(self.x_train[index], self.y_train[index], color='royalblue', label='sample')
# 予測区間
lb, ub = self.calc_prediction_interval(x)
error = (ub - lb) / 2
plt.errorbar(plot_x[1], plot_y[1], fmt='o', yerr=error, capsize=5, color='green', label='95% prediction interval')
plt.xlim([x-0.5, x+0.5])
plt.legend();
# # 実験
# +
# 観測した標本から95%予測区間を求める
random_seed = 12
experiment = Experiment(random_seed, SIGMA, SAMPLE_SIZE)
experiment.plot()
# +
# 観測した標本から95%予測区間, 信頼区間を求める
random_seed = 12
experiment = Experiment(random_seed, SIGMA, SAMPLE_SIZE)
experiment.plot_with_confidence()
# -
# ## 予測区間に観測値が含まれるケース
x = 2.5
experiment.plot_at_x(x)
# ## 予測区間に観測値が含まれないケース
x = 4.5
experiment.plot_at_x(x)
# # 実験を1万回繰り返す
# * 予測区間に観測値が含まれる割合を計測する
# +
experiment_count = 10000
count = 0
for i in range(experiment_count):
experiment = Experiment(i, SIGMA, SAMPLE_SIZE)
x = np.random.uniform(0, 10, 1)[0]
y = x + 1 + np.random.normal(0.0, SIGMA, 1)
lb, ub = experiment.calc_prediction_interval(x)
# 予測区間に観測値が含まれるかチェック
count += 1 if (lb <= y and y <= ub) else 0
print('予測区間に観測が含まれる割合: {:.1f}%'.format(100 * count / experiment_count))
| scikit-learn/SimpleLinearRegression/PredictionInterval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark import SparkContext, SparkConf
import os
import sys
sc.stop()
# +
dataset = "./data-1-sample.txt"
conf = (SparkConf()
.setAppName("hamroun"))
sc = SparkContext(conf=conf)
# +
#Data Loading
print("Data Loading")
original_data = sc.textFile(dataset)
original_data = original_data.map(lambda s: float(s))
#Definition of groupe Number
nb_groups = 10.0
count = original_data.count()
sum = original_data.sum()
max_data = original_data.max()
min_data = original_data.min()
quotient = round( (max_data/nb_group) - (min_data/nb_group))
print("number of groups =", nb_groups)
print("Count =", count)
print("Sum = %.8f" % sum)
print("Max = %.8f" % max_data)
print("Min = %.8f" % min_data)
print("quotient = %.2f" % quotient)
print("creation of key-value RDD")
keys_value_data = original_data.map(lambda number: ((int( number/quotient - min_data/quotient )),number))
keys_data = keys_value_data.groupByKey()
len_by_key = keys_data.mapValues(len).sortByKey()
list_lengths = len_by_key.values().collect()
list_keys = len_by_key.keys().collect()
accumulated_number = 0
print("Researching of group which contains the median.")
for current_id,current_length in zip(list_keys,list_lengths ):
accumulated_number += current_length
if(accumulated_number > count / 2):
break
median_index = int(count / 2 ) - (accumulated_number - list_lengths[current_id])
print("the id of the group which contains the median value : ")
print(current_id)
print("the id of the median in the selected group : ")
print(median_index)
selected_group = keys_data.mapValues(list).lookup(current_id)
median = sorted(selected_group[0])[median_index]
print("median = %.8f" % median)
sc.stop()
| median/median.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="z1PSyPUbbo3D"
# ### PARTE 1 - COME INTERAGIRE CON LE API
# + colab={"base_uri": "https://localhost:8080/"} id="SQA6Qao9X0lK" outputId="8d810aaf-03f2-4f78-dc8a-99f15143dcd8" language="bash"
# pip install requests
# + [markdown] id="uJSNWsLjYb9C"
# We want to access the latest currency exchange information, so we will be using GET /latest.json.
#
# Clicking on it brings us to the documentation, which tells us the URL we want to send data to is https://openexchangerates.org/api/latest.json.
#
# Let's add that to our program:
#
# - https://docs.openexchangerates.org/
# - https://openexchangerates.org/
#
# + id="6KLlvJ5xX8FC"
import requests
APP_ID = "batman"
ENDPOINT = "https://openexchangerates.org/api/latest.json"
# + [markdown] id="NZVikCVYY6Oi"
# This means that we have to use Python to send a GET request to this URL, making sure to include our App ID in there. There are a few different types of requests, such as GET and POST. This is just another piece of data the API expects. Depending on the request type, sometimes APIs will do different things in a single endpoint.
#
# Then we'll get back something like this (shown also in the official documentation):
#
# ```json
# {
# disclaimer: "https://openexchangerates.org/terms/",
# license: "https://openexchangerates.org/license/",
# timestamp: 1449877801,
# base: "USD",
# rates: {
# AED: 3.672538,
# AFN: 66.809999,
# ALL: 125.716501,
# AMD: 484.902502,
# ANG: 1.788575,
# AOA: 135.295998,
# ARS: 9.750101,
# AUD: 1.390866,
# /* ... */
# }
# }
# ```
#
# + colab={"base_uri": "https://localhost:8080/"} id="zEesJaBmYJ0B" outputId="825e9fcc-1374-4261-e79b-7091dea6fab4"
import requests
import json
#Le API sono gratutite quindi per questa volta non mi scroccate i token :D
with open("credenziali.json") as jsonFile:
jsonObject = json.load(jsonFile)
jsonFile.close()
TOKEN = jsonObject['API_TOKEN']
APP_ID = TOKEN
ENDPOINT = "https://openexchangerates.org/api/latest.json"
response = requests.get(f"{ENDPOINT}?app_id={APP_ID}")
print(response.content)
# + colab={"base_uri": "https://localhost:8080/"} id="4bMmv3SOZLAj" outputId="efdefdb5-28f9-42c4-c6f0-4b17d5ccc5d2"
response = requests.get(f"{ENDPOINT}?app_id={APP_ID}")
exchange_rates = response.json()
usd_amount = 1000
gbp_amount = usd_amount * exchange_rates['rates']['GBP']
print(f"USD{usd_amount} is GBP{gbp_amount}")
# + [markdown] id="fxB05bFCbt2D"
# ### PARTE 2 - COME SVILUPPARE DELLE API
# + id="j3RXMHcubxRq" colab={"base_uri": "https://localhost:8080/"} outputId="84b865d5-0af1-44d7-fc7d-da07dfe057c4"
# !git clone https://github.com/mongodb-developer/rewrite-it-in-rust.git
# + colab={"base_uri": "https://localhost:8080/"} id="O0tu27f2clIV" outputId="0ded894d-364e-4c2a-e605-7a50e76fa902"
# %cd /content/rewrite-it-in-rust/flask-cocktail-api
# + colab={"base_uri": "https://localhost:8080/"} id="6FALHZStc2-K" outputId="5dc579c7-ea2e-4b10-f125-f939774d32ae"
# !pip install -e .
# + id="cisy7VXHc6DX"
# !export MONGO_URI="mongodb+srv://USERNAME:<EMAIL>W<EMAIL>@<EMAIL>.azure.mongodb.net/cocktails?retryWrites=true&w=majority"
# + id="t_0ByK8gc_BF"
# !sudo apt-get install gcc python-dev libkrb5-dev
# !python -m pip install 'pymongo[srv]'
# + id="r_W6E77PdBmQ"
# !python -m pip install pymongo[snappy,gssapi,srv,tls]
# + id="Bjr85BqDecVR"
# !mongoimport --uri "$MONGO_URI" --file ./recipes.json
# + id="bMRbWeXheyuK"
# !FLASK_DEBUG=true FLASK_APP=cocktailapi flask run
# + [markdown] id="UC--qedIfVCs"
#
#
# ```json
# {
# "_links": {
# "last": {
# "href": "http://localhost:5000/cocktails/?page=5"
# },
# "next": {
# "href": "http://localhost:5000/cocktails/?page=5"
# },
# "prev": {
# "href": "http://localhost:5000/cocktails/?page=3"
# },
# "self": {
# "href": "http://localhost:5000/cocktails/?page=4"
# }
# },
# "recipes": [
# {
# "_id": "5f7daa198ec9dfb536781b0d",
# "date_added": null,
# "date_updated": null,
# "ingredients": [
# {
# "name": "Light rum",
# "quantity": {
# "unit": "oz",
# }
# },
# {
# "name": "Grapefruit juice",
# "quantity": {
# "unit": "oz",
# }
# },
# {
# "name": "Bitters",
# "quantity": {
# "unit": "dash",
# }
# }
# ],
# "instructions": [
# "Pour all of the ingredients into an old-fashioned glass almost filled with ice cubes",
# "Stir well."
# ],
# "name": "<NAME>",
# "slug": "monkey-wrench"
# },
# ]
# ```
#
#
# + id="gKHzDdM_fR_L"
# model.py
class Cocktail(BaseModel):
id: Optional[PydanticObjectId] = Field(None, alias="_id")
slug: str
name: str
ingredients: List[Ingredient]
instructions: List[str]
date_added: Optional[datetime]
date_updated: Optional[datetime]
def to_json(self):
return jsonable_encoder(self, exclude_none=True)
def to_bson(self):
data = self.dict(by_alias=True, exclude_none=True)
if data["_id"] is None:
data.pop("_id")
return data
# + id="0xKImiYFfe3b"
# objectid.py
class PydanticObjectId(ObjectId):
"""
ObjectId field. Compatible with Pydantic.
"""
@classmethod
def __get_validators__(cls):
yield cls.validate
@classmethod
def validate(cls, v):
return PydanticObjectId(v)
@classmethod
def __modify_schema__(cls, field_schema: dict):
field_schema.update(
type="string",
examples=["5eb7cf5a86d9755df3a6c593", "5eb7cfb05e32e07750a1756a"],
)
ENCODERS_BY_TYPE[PydanticObjectId] = str
# + id="CDdrqdR2fhvj"
@app.route("/cocktails/", methods=["POST"])
def new_cocktail():
raw_cocktail = request.get_json()
raw_cocktail["date_added"] = datetime.utcnow()
cocktail = Cocktail(**raw_cocktail)
insert_result = recipes.insert_one(cocktail.to_bson())
cocktail.id = PydanticObjectId(str(insert_result.inserted_id))
print(cocktail)
return cocktail.to_json()
# + id="fuiMgIXxfjJj"
@app.route("/cocktails/<string:slug>", methods=["GET"])
def get_cocktail(slug):
recipe = recipes.find_one_or_404({"slug": slug})
return Cocktail(**recipe).to_json()
# + id="6C5-7rb6flF7"
@app.route("/cocktails/")
def list_cocktails():
"""
GET a list of cocktail recipes.
The results are paginated using the `page` parameter.
"""
page = int(request.args.get("page", 1))
per_page = 10 # A const value.
# For pagination, it's necessary to sort by name,
# then skip the number of docs that earlier pages would have displayed,
# and then to limit to the fixed page size, ``per_page``.
cursor = recipes.find().sort("name").skip(per_page * (page - 1)).limit(per_page)
cocktail_count = recipes.count_documents({})
links = {
"self": {"href": url_for(".list_cocktails", page=page, _external=True)},
"last": {
"href": url_for(
".list_cocktails", page=(cocktail_count // per_page) + 1, _external=True
)
},
}
# Add a 'prev' link if it's not on the first page:
if page > 1:
links["prev"] = {
"href": url_for(".list_cocktails", page=page - 1, _external=True)
}
# Add a 'next' link if it's not on the last page:
if page - 1 < cocktail_count // per_page:
links["next"] = {
"href": url_for(".list_cocktails", page=page + 1, _external=True)
}
return {
"recipes": [Cocktail(**doc).to_json() for doc in cursor],
"_links": links,
}
# + [markdown] id="lJvsgacffue0"
# ### Gestire Errori ed eccezioni
# + id="BpNnloNFfwwp"
@app.errorhandler(404)
def resource_not_found(e):
"""
An error-handler to ensure that 404 errors are returned as JSON.
"""
return jsonify(error=str(e)), 404
@app.errorhandler(DuplicateKeyError)
def resource_not_found(e):
"""
An error-handler to ensure that MongoDB duplicate key errors are returned as JSON.
"""
return jsonify(error=f"Duplicate key error."), 400
| riassunto_per_frettolosi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Exploring precision and recall
#
# The goal of this second notebook is to understand precision-recall in the context of classifiers.
#
# * Use Amazon review data in its entirety.
# * Train a logistic regression model.
# * Explore various evaluation metrics: accuracy, confusion matrix, precision, recall.
# * Explore how various metrics can be combined to produce a cost of making an error.
# * Explore precision and recall curves.
#
# Because we are using the full Amazon review dataset (not a subset of words or reviews), in this assignment we return to using GraphLab Create for its efficiency. As usual, let's start by **firing up GraphLab Create**.
#
# Make sure you have the latest version of GraphLab Create (1.8.3 or later). If you don't find the decision tree module, then you would need to upgrade graphlab-create using
#
# ```
# pip install graphlab-create --upgrade
# ```
# See [this page](https://dato.com/download/) for detailed instructions on upgrading.
import graphlab
from __future__ import division
import numpy as np
graphlab.canvas.set_target('ipynb')
# # Load amazon review dataset
products = graphlab.SFrame('amazon_baby.gl/')
# # Extract word counts and sentiments
# As in the first assignment of this course, we compute the word counts for individual words and extract positive and negative sentiments from ratings. To summarize, we perform the following:
#
# 1. Remove punctuation.
# 2. Remove reviews with "neutral" sentiment (rating 3).
# 3. Set reviews with rating 4 or more to be positive and those with 2 or less to be negative.
# +
def remove_punctuation(text):
import string
return text.translate(None, string.punctuation)
# Remove punctuation.
review_clean = products['review'].apply(remove_punctuation)
# Count words
products['word_count'] = graphlab.text_analytics.count_words(review_clean)
# Drop neutral sentiment reviews.
products = products[products['rating'] != 3]
# Positive sentiment to +1 and negative sentiment to -1
products['sentiment'] = products['rating'].apply(lambda rating : +1 if rating > 3 else -1)
# -
# Now, let's remember what the dataset looks like by taking a quick peek:
products
# ## Split data into training and test sets
#
# We split the data into a 80-20 split where 80% is in the training set and 20% is in the test set.
train_data, test_data = products.random_split(.8, seed=1)
# ## Train a logistic regression classifier
#
# We will now train a logistic regression classifier with **sentiment** as the target and **word_count** as the features. We will set `validation_set=None` to make sure everyone gets exactly the same results.
#
# Remember, even though we now know how to implement logistic regression, we will use GraphLab Create for its efficiency at processing this Amazon dataset in its entirety. The focus of this assignment is instead on the topic of precision and recall.
model = graphlab.logistic_classifier.create(train_data, target='sentiment',
features=['word_count'],
validation_set=None)
# # Model Evaluation
# We will explore the advanced model evaluation concepts that were discussed in the lectures.
#
# ## Accuracy
#
# One performance metric we will use for our more advanced exploration is accuracy, which we have seen many times in past assignments. Recall that the accuracy is given by
#
# $$
# \mbox{accuracy} = \frac{\mbox{# correctly classified data points}}{\mbox{# total data points}}
# $$
#
# To obtain the accuracy of our trained models using GraphLab Create, simply pass the option `metric='accuracy'` to the `evaluate` function. We compute the **accuracy** of our logistic regression model on the **test_data** as follows:
accuracy= model.evaluate(test_data, metric='accuracy')['accuracy']
print "Test Accuracy: %s" % accuracy
# ## Baseline: Majority class prediction
#
# Recall from an earlier assignment that we used the **majority class classifier** as a baseline (i.e reference) model for a point of comparison with a more sophisticated classifier. The majority classifier model predicts the majority class for all data points.
#
# Typically, a good model should beat the majority class classifier. Since the majority class in this dataset is the positive class (i.e., there are more positive than negative reviews), the accuracy of the majority class classifier can be computed as follows:
baseline = len(test_data[test_data['sentiment'] == 1])/len(test_data)
print "Baseline accuracy (majority class classifier): %s" % baseline
# ** Quiz Question:** Using accuracy as the evaluation metric, was our **logistic regression model** better than the baseline (majority class classifier)?
# Yes
# ## Confusion Matrix
#
# The accuracy, while convenient, does not tell the whole story. For a fuller picture, we turn to the **confusion matrix**. In the case of binary classification, the confusion matrix is a 2-by-2 matrix laying out correct and incorrect predictions made in each label as follows:
# ```
# +---------------------------------------------+
# | Predicted label |
# +----------------------+----------------------+
# | (+1) | (-1) |
# +-------+-----+----------------------+----------------------+
# | True |(+1) | # of true positives | # of false negatives |
# | label +-----+----------------------+----------------------+
# | |(-1) | # of false positives | # of true negatives |
# +-------+-----+----------------------+----------------------+
# ```
# To print out the confusion matrix for a classifier, use `metric='confusion_matrix'`:
confusion_matrix = model.evaluate(test_data, metric='confusion_matrix')['confusion_matrix']
confusion_matrix
# **Quiz Question**: How many predicted values in the **test set** are **false positives**?
1406+1443
# ## Computing the cost of mistakes
#
#
# Put yourself in the shoes of a manufacturer that sells a baby product on Amazon.com and you want to monitor your product's reviews in order to respond to complaints. Even a few negative reviews may generate a lot of bad publicity about the product. So you don't want to miss any reviews with negative sentiments --- you'd rather put up with false alarms about potentially negative reviews instead of missing negative reviews entirely. In other words, **false positives cost more than false negatives**. (It may be the other way around for other scenarios, but let's stick with the manufacturer's scenario for now.)
#
# Suppose you know the costs involved in each kind of mistake:
# 1. \$100 for each false positive.
# 2. \$1 for each false negative.
# 3. Correctly classified reviews incur no cost.
#
# **Quiz Question**: Given the stipulation, what is the cost associated with the logistic regression classifier's performance on the **test set**?
1406 + 1443*100
# ## Precision and Recall
# You may not have exact dollar amounts for each kind of mistake. Instead, you may simply prefer to reduce the percentage of false positives to be less than, say, 3.5% of all positive predictions. This is where **precision** comes in:
#
# $$
# [\text{precision}] = \frac{[\text{# positive data points with positive predicitions}]}{\text{[# all data points with positive predictions]}} = \frac{[\text{# true positives}]}{[\text{# true positives}] + [\text{# false positives}]}
# $$
# So to keep the percentage of false positives below 3.5% of positive predictions, we must raise the precision to 96.5% or higher.
#
# **First**, let us compute the precision of the logistic regression classifier on the **test_data**.
precision = model.evaluate(test_data, metric='precision')['precision']
print "Precision on test data: %s" % precision
# **Quiz Question**: Out of all reviews in the **test set** that are predicted to be positive, what fraction of them are **false positives**? (Round to the second decimal place e.g. 0.25)
(1443)/(26689+1443)
# **Quiz Question:** Based on what we learned in lecture, if we wanted to reduce this fraction of false positives to be below 3.5%, we would: (see the quiz)
# A complementary metric is **recall**, which measures the ratio between the number of true positives and that of (ground-truth) positive reviews:
#
# $$
# [\text{recall}] = \frac{[\text{# positive data points with positive predicitions}]}{\text{[# all positive data points]}} = \frac{[\text{# true positives}]}{[\text{# true positives}] + [\text{# false negatives}]}
# $$
#
# Let us compute the recall on the **test_data**.
recall = model.evaluate(test_data, metric='recall')['recall']
print "Recall on test data: %s" % recall
# **Quiz Question**: What fraction of the positive reviews in the **test_set** were correctly predicted as positive by the classifier?
(26689)/float(26689+1443)
# **Quiz Question**: What is the recall value for a classifier that predicts **+1** for all data points in the **test_data**?
(26689)/float(26689+1406)
# # Precision-recall tradeoff
#
# In this part, we will explore the trade-off between precision and recall discussed in the lecture. We first examine what happens when we use a different threshold value for making class predictions. We then explore a range of threshold values and plot the associated precision-recall curve.
#
# ## Varying the threshold
#
# False positives are costly in our example, so we may want to be more conservative about making positive predictions. To achieve this, instead of thresholding class probabilities at 0.5, we can choose a higher threshold.
#
# Write a function called `apply_threshold` that accepts two things
# * `probabilities` (an SArray of probability values)
# * `threshold` (a float between 0 and 1).
#
# The function should return an SArray, where each element is set to +1 or -1 depending whether the corresponding probability exceeds `threshold`.
def apply_threshold(probabilities, threshold):
### YOUR CODE GOES HERE
# +1 if >= threshold and -1 otherwise.
return probabilities.apply(lambda x : +1 if x >= threshold else -1)
# Run prediction with `output_type='probability'` to get the list of probability values. Then use thresholds set at 0.5 (default) and 0.9 to make predictions from these probability values.
probabilities = model.predict(test_data, output_type='probability')
predictions_with_default_threshold = apply_threshold(probabilities, 0.5)
predictions_with_high_threshold = apply_threshold(probabilities, 0.9)
print "Number of positive predicted reviews (threshold = 0.5): %s" % (predictions_with_default_threshold == 1).sum()
print "Number of positive predicted reviews (threshold = 0.9): %s" % (predictions_with_high_threshold == 1).sum()
# **Quiz Question**: What happens to the number of positive predicted reviews as the threshold increased from 0.5 to 0.9?
# Becomes optimistic
# ## Exploring the associated precision and recall as the threshold varies
# By changing the probability threshold, it is possible to influence precision and recall. We can explore this as follows:
# +
# Threshold = 0.5
precision_with_default_threshold = graphlab.evaluation.precision(test_data['sentiment'],
predictions_with_default_threshold)
recall_with_default_threshold = graphlab.evaluation.recall(test_data['sentiment'],
predictions_with_default_threshold)
# Threshold = 0.9
precision_with_high_threshold = graphlab.evaluation.precision(test_data['sentiment'],
predictions_with_high_threshold)
recall_with_high_threshold = graphlab.evaluation.recall(test_data['sentiment'],
predictions_with_high_threshold)
# -
print "Precision (threshold = 0.5): %s" % precision_with_default_threshold
print "Recall (threshold = 0.5) : %s" % recall_with_default_threshold
print "Precision (threshold = 0.9): %s" % precision_with_high_threshold
print "Recall (threshold = 0.9) : %s" % recall_with_high_threshold
# **Quiz Question (variant 1)**: Does the **precision** increase with a higher threshold?
# Yes
# **Quiz Question (variant 2)**: Does the **recall** increase with a higher threshold?
# No
# ## Precision-recall curve
#
# Now, we will explore various different values of tresholds, compute the precision and recall scores, and then plot the precision-recall curve.
threshold_values = np.linspace(0.5, 1, num=100)
print threshold_values
# For each of the values of threshold, we compute the precision and recall scores.
# +
precision_all = []
recall_all = []
probabilities = model.predict(test_data, output_type='probability')
for threshold in threshold_values:
predictions = apply_threshold(probabilities, threshold)
precision = graphlab.evaluation.precision(test_data['sentiment'], predictions)
recall = graphlab.evaluation.recall(test_data['sentiment'], predictions)
precision_all.append(precision)
recall_all.append(recall)
# -
# Now, let's plot the precision-recall curve to visualize the precision-recall tradeoff as we vary the threshold.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_pr_curve(precision, recall, title):
plt.rcParams['figure.figsize'] = 7, 5
plt.locator_params(axis = 'x', nbins = 5)
plt.plot(precision, recall, 'b-', linewidth=4.0, color = '#B0017F')
plt.title(title)
plt.xlabel('Precision')
plt.ylabel('Recall')
plt.rcParams.update({'font.size': 16})
plot_pr_curve(precision_all, recall_all, 'Precision recall curve (all)')
# -
# **Quiz Question**: Among all the threshold values tried, what is the **smallest** threshold value that achieves a precision of 96.5% or better? Round your answer to 3 decimal places.
for threshold in threshold_values:
predictions = apply_threshold(probabilities, threshold)
precision = graphlab.evaluation.precision(test_data['sentiment'], predictions)
print threshold, precision
# **0.838383838384**
# **Quiz Question**: Using `threshold` = 0.98, how many **false negatives** do we get on the **test_data**? (**Hint**: You may use the `graphlab.evaluation.confusion_matrix` function implemented in GraphLab Create.)
predictions = apply_threshold(probabilities, 0.98)
precision = graphlab.evaluation.confusion_matrix(test_data['sentiment'], predictions)
precision
# This is the number of false negatives (i.e the number of reviews to look at when not needed) that we have to deal with using this classifier.
# 1406
# # Evaluating specific search terms
# So far, we looked at the number of false positives for the **entire test set**. In this section, let's select reviews using a specific search term and optimize the precision on these reviews only. After all, a manufacturer would be interested in tuning the false positive rate just for their products (the reviews they want to read) rather than that of the entire set of products on Amazon.
#
# ## Precision-Recall on all baby related items
#
# From the **test set**, select all the reviews for all products with the word 'baby' in them.
baby_reviews = test_data[test_data['name'].apply(lambda x: 'baby' in x.lower())]
# Now, let's predict the probability of classifying these reviews as positive:
probabilities = model.predict(baby_reviews, output_type='probability')
# Let's plot the precision-recall curve for the **baby_reviews** dataset.
#
# **First**, let's consider the following `threshold_values` ranging from 0.5 to 1:
threshold_values = np.linspace(0.5, 1, num=100)
# **Second**, as we did above, let's compute precision and recall for each value in `threshold_values` on the **baby_reviews** dataset. Complete the code block below.
# +
precision_all = []
recall_all = []
for threshold in threshold_values:
# Make predictions. Use the `apply_threshold` function
## YOUR CODE HERE
predictions = apply_threshold(probabilities, threshold)
# Calculate the precision.
# YOUR CODE HERE
precision = graphlab.evaluation.precision(baby_reviews['sentiment'], predictions)
# YOUR CODE HERE
recall = graphlab.evaluation.recall(baby_reviews['sentiment'], predictions)
# Append the precision and recall scores.
precision_all.append((precision, threshold))
recall_all.append((recall, threshold))
# -
# **Quiz Question**: Among all the threshold values tried, what is the **smallest** threshold value that achieves a precision of 96.5% or better for the reviews of data in **baby_reviews**? Round your answer to 3 decimal places.
print precision_all
# **0.86363636363636365**
# **Quiz Question:** Is this threshold value smaller or larger than the threshold used for the entire dataset to achieve the same specified precision of 96.5%?
#
# **Finally**, let's plot the precision recall curve.
# **smaller**
plot_pr_curve(precision_all, recall_all, "Precision-Recall (Baby)")
| Course3/week6/module-9-precision-recall-assignment-blank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ds]
# language: python
# name: conda-env-ds-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
#
# # MAT281
# ### Aplicaciones de la Matemática en la Ingeniería
# + [markdown] slideshow={"slide_type": "slide"}
# ## Módulo 02
# ## Clase 01: Computación Científica
# + [markdown] slideshow={"slide_type": "slide"}
# ## Objetivos
#
# * Conocer las librerías de computación científica
# * Trabajar con arreglos *matriciales*
# * Álgebra lineal con numpy
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Contenidos
# * [Scipy.org](#scipy.org)
# * [Numpy Arrays](#arrays)
# * [Operaciones Básicas](#operations)
# * [Broadcasting](#roadcasting)
# * [Álgebra Lineal](#linear_algebra)
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='scipy.org'></a>
# ## SciPy.org
# -
# **SciPy** es un ecosistema de software _open-source_ para matemática, ciencia y engeniería. Los principales son:
#
# * Numpy: Arrays N-dimensionales. Librería base, integración con C/C++ y Fortran.
# * Scipy library: Computación científica (integración, optimización, estadística, etc.)
# * Matplotlib: Visualización 2D:
# * IPython: Interactividad (Project Jupyter).
# * Simpy: Matemática Simbólica.
# * Pandas: Estructura y análisis de datos.
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='arrays'></a>
# ## Numpy Arrays
# -
# Los objetos principales de Numpy son los comúnmente conocidos como Numpy Arrays (la clase se llama `ndarray`), corresponden a una tabla de elementos, todos del mismo tipo, indexados por una tupla de enternos no-negativos. En Numpy, las dimensiones son llamadas _axes_.
import numpy as np
# +
a = np.array(
[
[ 0, 1, 2, 3, 4],
[ 5, 6, 7, 8, 9],
[10, 11, 12, 13, 14]
]
)
type(a)
# -
# Los atributos más importantes de un `ndarray` son:
a.shape # the dimensions of the array.
a.ndim # the number of axes (dimensions) of the array.
a.size # the total number of elements of the array.
a.dtype # an object describing the type of the elements in the array.
a.itemsize # the size in bytes of each element of the array.
# ### Crear Numpy Arrays
# Hay varias formas de crear arrays, el constructor básico es el que se utilizó hace unos momentos, `np.array`. El _type_ del array resultante es inferido de los datos proporcionados.
# +
a_int = np.array([2, 6, 10])
a_float = np.array([2.1, 6.1, 10.1])
print(f"a_int: {a_int.dtype.name}")
print(f"a_float: {a_float.dtype.name}")
# -
# ### Constantes
np.zeros((3, 4))
np.ones((2, 3, 4), dtype=np.int) # dtype can also be specified
np.identity(4) # Identity matrix
# ### Range
# Numpy proporciona una función análoga a `range`.
range(10)
type(range(10))
np.arange(10)
type(np.arange(10))
np.arange(3, 10)
np.arange(2, 20, 3, dtype=np.float)
np.arange(9).reshape(3, 3)
# Bonus
np.linspace(0, 100, 5)
# ### Random
np.random.uniform(size=5)
np.random.normal(size=(2, 3))
# ### Acceder a los elementos de un array
x1 = np.arange(0, 30, 4)
x2 = np.arange(0, 60, 3).reshape(4, 5)
print("x1:")
print(x1)
print("\nx2:")
print(x2)
x1[1] # Un elemento de un array 1D
x1[:3] # Los tres primeros elementos
x2[0, 2] # Un elemento de un array 2D
x2[0] # La primera fila
x2[:, 1] # Todas las filas y la segunda columna
x2[:, 1:3] # Todas las filas y de la segunda a la tercera columna
x2[:, 1:2] # What?!
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='operations'></a>
# ## Operaciones Básias
# -
# Numpy provee operaciones vectorizadas, con tal de mejorar el rendimiento de la ejecución.
# Por ejemplo, pensemos en la suma de dos arreglos 2D.
A = np.random.random((5,5))
B = np.random.random((5,5))
# Con los conocimientos de la clase pasada, podríamos pensar en iterar a través de dos `for`, con tal de llenar el arreglo resultando. algo así:
def my_sum(A, B):
n, m = A.shape
C = np.empty(shape=(n, m))
for i in range(n):
for j in range(m):
C[i, j] = A[i, j] + B[i, j]
return C
# %timeit my_sum(A, B)
# Pero la suma de `ndarray`s es simplemente con el signo de suma (`+`):
# %timeit A + B
# Para dos arrays tan pequeños la diferencia de tiempo es considerable, ¡Imagina con millones de datos!
# Los clásicos de clásicos:
x = np.arange(5)
print(f"x = {x}")
print(f"x + 5 = {x + 5}")
print(f"x - 5 = {x - 5}")
print(f"x * 2 = {x * 2}")
print(f"x / 2 = {x / 2}")
print(f"x // 2 = {x // 2}")
print(f"x ** 2 = {x ** 2}")
print(f"x % 2 = {x % 2}")
# ¡Júntalos como quieras!
-(0.5 + x + 3) ** 2
# Al final del día, estos son alias para funciones de Numpy, por ejemplo, la operación suma (`+`) es un _wrapper_ de la función `np.add`
np.add(x, 5)
# Podríamos estar todo el día hablando de operaciones, pero básicamente, si piensas en alguna operación lo suficientemente común, es que la puedes encontrar implementada en Numpy. Por ejemplo:
np.abs(-(0.5 + x + 3) ** 2)
np.log(x + 5)
np.exp(x)
np.sin(x)
# ### ¿Y para dimensiones mayores?
# La idea es la misma, pero siempre hay que tener cuidado con las dimensiones y `shape` de los arrays.
print("A + B: \n")
print(A + B)
print("\n" + "-" * 80 + "\n")
print("A - B: \n")
print(A - B)
print("\n" + "-" * 80 + "\n")
print("A * B: \n")
print(A * B) # Producto elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A / B: \n")
print(A / B) # División elemento a elemento
print("\n" + "-" * 80 + "\n")
print("A @ B: \n")
print(A @ B) # Producto matricial
# ### Operaciones Booleanas
print(f"x = {x}")
print(f"x > 2 = {x > 2}")
print(f"x == 2 = {x == 2}")
print(f"x == 2 = {x == 2}")
# +
aux1 = np.array([[1, 2, 3], [2, 3, 5], [1, 9, 6]])
aux2 = np.array([[1, 2, 3], [3, 5, 5], [0, 8, 5]])
B1 = aux1 == aux2
B2 = aux1 > aux2
print("B1: \n")
print(B1)
print("\n" + "-" * 80 + "\n")
print("B2: \n")
print(B2)
print("\n" + "-" * 80 + "\n")
print("~B1: \n")
print(~B1) # También puede ser np.logical_not(B1)
print("\n" + "-" * 80 + "\n")
print("B1 | B2 : \n")
print(B1 | B2)
print("\n" + "-" * 80 + "\n")
print("B1 & B2 : \n")
print(B1 & B2)
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='broadcasting'></a>
# ## Broadcasting
# -
# ¿Qué pasa si las dimensiones no coinciden? Observemos lo siguiente:
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
a + b
# Todo bien, dos arrays 1D de 3 elementos, la suma retorna un array de 3 elementos.
a + 3
# Sigue pareciendo normal, un array 1D de 3 elementos, se suma con un `int`, lo que retorna un array 1D de tres elementos.
M = np.ones((3, 3))
M
M + a
# Magia! Esto es _broadcasting_. Una pequeña infografía es la siguiente:
# 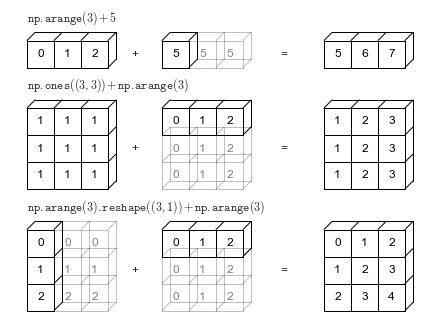
# Resumen: A lo menos los dos arrays deben coincidir en una dimensión. Luego, el array de dimensión menor se extiende con tal de ajustarse a las dimensiones del otro.
#
# La documentación oficial de estas reglas la puedes encontrar [aquí](https://numpy.org/devdocs/user/basics.broadcasting.html).
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='lineal_algebra'></a>
# ## Álgebra Lineal
# -
# Veamos algunas operaciones básicas de álgebra lineal, las que te servirán para el día a día.
a = np.array([[1.0, 2.0], [3.0, 4.0]])
print(a)
# Transpuesta
a.T # a.transpose()
# Determinante
np.linalg.det(a)
# Inversa
np.linalg.inv(a)
# Traza
np.trace(a)
# Número de condición
np.linalg.cond(a)
# Sistemas lineales
y = np.array([[5.], [7.]])
np.linalg.solve(a, y)
# Valores y vectores propios
np.linalg.eig(a)
# Descomposición QR
np.linalg.qr(a)
| m02_data_analysis/m02_c01_scientific computing/m02_c01_scientific computing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.optim.lr_scheduler import StepLR, MultiStepLR
import numpy as np
# import matplotlib.pyplot as plt
from math import *
import time
import copy
torch.cuda.set_device(2)
torch.set_default_tensor_type('torch.DoubleTensor')
# activation function
def activation(x):
return x * torch.sigmoid(x)
# build ResNet with one blocks
class Net(torch.nn.Module):
def __init__(self,input_width,layer_width):
super(Net,self).__init__()
self.layer_in = torch.nn.Linear(input_width, layer_width)
self.layer1 = torch.nn.Linear(layer_width, layer_width)
self.layer2 = torch.nn.Linear(layer_width, layer_width)
self.layer_out = torch.nn.Linear(layer_width, 1)
def forward(self,x):
y = self.layer_in(x)
y = y + activation(self.layer2(activation(self.layer1(y)))) # residual block 1
output = self.layer_out(y)
return output
dimension = 1
input_width,layer_width = dimension, 4
net = Net(input_width,layer_width).cuda() # network for u on gpu
# defination of exact solution
def u_ex(x):
temp = 1.0
for i in range(dimension):
temp = temp * torch.sin(pi*x[:, i])
u_temp = 1.0 * temp
return u_temp.reshape([x.size()[0], 1])
# defination of f(x)
def f(x):
temp = 1.0
for i in range(dimension):
temp = temp * torch.sin(pi*x[:, i])
u_temp = 1.0 * temp
f_temp = dimension * pi**2 * u_temp
return f_temp.reshape([x.size()[0],1])
# generate points by random
def generate_sample(data_size):
sample_temp = torch.rand(data_size, dimension)
return sample_temp.cuda()
def model(x):
x_temp = x.cuda()
D_x_0 = torch.prod(x_temp, axis = 1).reshape([x.size()[0], 1])
D_x_1 = torch.prod(1.0 - x_temp, axis = 1).reshape([x.size()[0], 1])
model_u_temp = D_x_0 * D_x_1 * net(x)
return model_u_temp.reshape([x.size()[0], 1])
# loss function to DRM by auto differential
def loss_function(x):
# x = generate_sample(data_size).cuda()
# x.requires_grad = True
u_hat = model(x)
grad_u_hat = torch.autograd.grad(outputs = u_hat, inputs = x, grad_outputs = torch.ones(u_hat.shape).cuda(), create_graph = True)
grad_u_sq = ((grad_u_hat[0]**2).sum(1)).reshape([len(grad_u_hat[0]), 1])
part = torch.sum(0.5 * grad_u_sq - f(x) * u_hat) / len(x)
return part
data_size = 200
x = generate_sample(data_size).cuda()
x.requires_grad = True
def get_weights(net):
""" Extract parameters from net, and return a list of tensors"""
return [p.data for p in net.parameters()]
def set_weights(net, weights, directions=None, step=None):
"""
Overwrite the network's weights with a specified list of tensors
or change weights along directions with a step size.
"""
if directions is None:
# You cannot specify a step length without a direction.
for (p, w) in zip(net.parameters(), weights):
p.data.copy_(w.type(type(p.data)))
else:
assert step is not None, 'If a direction is specified then step must be specified as well'
if len(directions) == 2:
dx = directions[0]
dy = directions[1]
changes = [d0*step[0] + d1*step[1] for (d0, d1) in zip(dx, dy)]
else:
changes = [d*step for d in directions[0]]
for (p, w, d) in zip(net.parameters(), weights, changes):
p.data = w + torch.Tensor(d).type(type(w))
def set_states(net, states, directions=None, step=None):
"""
Overwrite the network's state_dict or change it along directions with a step size.
"""
if directions is None:
net.load_state_dict(states)
else:
assert step is not None, 'If direction is provided then the step must be specified as well'
if len(directions) == 2:
dx = directions[0]
dy = directions[1]
changes = [d0*step[0] + d1*step[1] for (d0, d1) in zip(dx, dy)]
else:
changes = [d*step for d in directions[0]]
new_states = copy.deepcopy(states)
assert (len(new_states) == len(changes))
for (k, v), d in zip(new_states.items(), changes):
d = torch.tensor(d)
v.add_(d.type(v.type()))
net.load_state_dict(new_states)
def get_random_weights(weights):
"""
Produce a random direction that is a list of random Gaussian tensors
with the same shape as the network's weights, so one direction entry per weight.
"""
return [torch.randn(w.size()) for w in weights]
def get_random_states(states):
"""
Produce a random direction that is a list of random Gaussian tensors
with the same shape as the network's state_dict(), so one direction entry
per weight, including BN's running_mean/var.
"""
return [torch.randn(w.size()) for k, w in states.items()]
def get_diff_weights(weights, weights2):
""" Produce a direction from 'weights' to 'weights2'."""
return [w2 - w for (w, w2) in zip(weights, weights2)]
def get_diff_states(states, states2):
""" Produce a direction from 'states' to 'states2'."""
return [v2 - v for (k, v), (k2, v2) in zip(states.items(), states2.items())]
def normalize_direction(direction, weights, norm='filter'):
"""
Rescale the direction so that it has similar norm as their corresponding
model in different levels.
Args:
direction: a variables of the random direction for one layer
weights: a variable of the original model for one layer
norm: normalization method, 'filter' | 'layer' | 'weight'
"""
if norm == 'filter':
# Rescale the filters (weights in group) in 'direction' so that each
# filter has the same norm as its corresponding filter in 'weights'.
for d, w in zip(direction, weights):
d.mul_(w.norm()/(d.norm() + 1e-10))
elif norm == 'layer':
# Rescale the layer variables in the direction so that each layer has
# the same norm as the layer variables in weights.
direction.mul_(weights.norm()/direction.norm())
elif norm == 'weight':
# Rescale the entries in the direction so that each entry has the same
# scale as the corresponding weight.
direction.mul_(weights)
elif norm == 'dfilter':
# Rescale the entries in the direction so that each filter direction
# has the unit norm.
for d in direction:
d.div_(d.norm() + 1e-10)
elif norm == 'dlayer':
# Rescale the entries in the direction so that each layer direction has
# the unit norm.
direction.div_(direction.norm())
def normalize_directions_for_weights(direction, weights, norm='filter', ignore='biasbn'):
"""
The normalization scales the direction entries according to the entries of weights.
"""
assert(len(direction) == len(weights))
for d, w in zip(direction, weights):
if d.dim() <= 1:
if ignore == 'biasbn':
d.fill_(0) # ignore directions for weights with 1 dimension
else:
d.copy_(w) # keep directions for weights/bias that are only 1 per node
else:
normalize_direction(d, w, norm)
def normalize_directions_for_states(direction, states, norm='filter', ignore='ignore'):
assert(len(direction) == len(states))
for d, (k, w) in zip(direction, states.items()):
if d.dim() <= 1:
if ignore == 'biasbn':
d.fill_(0) # ignore directions for weights with 1 dimension
else:
d.copy_(w) # keep directions for weights/bias that are only 1 per node
else:
normalize_direction(d, w, norm)
def ignore_biasbn(directions):
""" Set bias and bn parameters in directions to zero """
for d in directions:
if d.dim() <= 1:
d.fill_(0)
def create_random_direction(net, dir_type='weights', ignore='biasbn', norm='filter'):
"""
Setup a random (normalized) direction with the same dimension as
the weights or states.
Args:
net: the given trained model
dir_type: 'weights' or 'states', type of directions.
ignore: 'biasbn', ignore biases and BN parameters.
norm: direction normalization method, including
'filter" | 'layer' | 'weight' | 'dlayer' | 'dfilter'
Returns:
direction: a random direction with the same dimension as weights or states.
"""
# random direction
if dir_type == 'weights':
weights = get_weights(net) # a list of parameters.
direction = get_random_weights(weights)
normalize_directions_for_weights(direction, weights, norm, ignore)
elif dir_type == 'states':
states = net.state_dict() # a dict of parameters, including BN's running mean/var.
direction = get_random_states(states)
normalize_directions_for_states(direction, states, norm, ignore)
return direction
def tvd(m, l_i):
# load model parameters
pretrained_dict = torch.load('net_params_DGM_ResNet_Uniform.pkl')
# get state_dict
net_state_dict = net.state_dict()
# remove keys that does not belong to net_state_dict
pretrained_dict_1 = {k: v for k, v in pretrained_dict.items() if k in net_state_dict}
# update dict
net_state_dict.update(pretrained_dict_1)
# set new dict back to net
net.load_state_dict(net_state_dict)
weights_temp = get_weights(net)
states_temp = net.state_dict()
step_size = 2 * l_i / m
grid = np.arange(-l_i, l_i + step_size, step_size)
num_direction = 1
loss_matrix = torch.zeros((num_direction, len(grid)))
for temp in range(num_direction):
weights = weights_temp
states = states_temp
direction_temp = create_random_direction(net, dir_type='weights', ignore='biasbn', norm='filter')
normalize_directions_for_states(direction_temp, states, norm='filter', ignore='ignore')
directions = [direction_temp]
for dx in grid:
itemindex_1 = np.argwhere(grid == dx)
step = dx
set_states(net, states, directions, step)
loss_temp = loss_function(x)
loss_matrix[temp, itemindex_1[0]] = loss_temp
# clear memory
torch.cuda.empty_cache()
# get state_dict
net_state_dict = net.state_dict()
# remove keys that does not belong to net_state_dict
pretrained_dict_1 = {k: v for k, v in pretrained_dict.items() if k in net_state_dict}
# update dict
net_state_dict.update(pretrained_dict_1)
# set new dict back to net
net.load_state_dict(net_state_dict)
weights_temp = get_weights(net)
states_temp = net.state_dict()
interval_length = grid[-1] - grid[0]
TVD = 0.0
for temp in range(num_direction):
for index in range(loss_matrix.size()[1] - 1):
TVD = TVD + np.abs(float(loss_matrix[temp, index] - loss_matrix[temp, index + 1]))
Max = np.max(loss_matrix.detach().numpy())
Min = np.min(loss_matrix.detach().numpy())
TVD = TVD / interval_length / num_direction / (Max - Min)
return TVD, Max, Min
# +
M = 100
m = 20
l_i = 1.0
TVD_DRM = 0.0
time_start = time.time()
Max = []
Min = []
Result = []
for count in range(M):
TVD_temp, Max_temp, Min_temp = tvd(m, l_i)
# print(Max_temp, Min_temp)
Max.append(Max_temp)
Min.append(Min_temp)
Result.append(TVD_temp)
print('Current direction TVD of DGM is: ', TVD_temp)
TVD_DRM = TVD_DRM + TVD_temp
print((count + 1) / M * 100, '% finished.')
# print('Max of all is: ', np.max(Max))
# print('Min of all is: ', np.min(Min))
TVD_DRM = TVD_DRM / M
print('All directions average TVD of DGM is: ', TVD_DRM)
print('Variance TVD of DRM is: ', np.sqrt(np.var(Result, ddof = 1)))
print("Value of roughness index is: ", np.sqrt(np.var(Result, ddof = 1)) / TVD_DRM)
time_end = time.time()
print('Total time costs: ', time_end - time_start, 'seconds')
# -
| code/Results1D/randomInitialization/DRM_index_ResNet_Uniform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import category_encoders as ce
import numpy as np
# +
df = pd.DataFrame({
'name': ['<NAME>', 'Walter', 'Donny', 'The Stranger', 'Brandt', 'Bunny'],
'haircolor': ['brown', np.nan, 'brown', 'silver', 'blonde', 'blonde'],
'gender': ['male', 'male', 'male', 'male', 'male', 'female'],
'drink': ['caucasian', 'beer', 'beer', 'sasparilla', 'unknown', 'unknown'],
'age': [48, 49, 45, 63, 40, 23]
},
columns=['name', 'haircolor', 'gender', 'drink', 'age']
)
test_df = pd.DataFrame({
'name': ['<NAME>'],
'haircolor': ['black'],
'gender': ['male'],
'drink': ['caucasian'],
'age': [48]
},
columns=['name', 'haircolor', 'gender', 'drink', 'age']
)
# -
encoder = ce.OneHotEncoder(cols=['haircolor']) #, handle_unknown='return_nan')
df_onehot = encoder.fit_transform(df)
df_onehot
df
encoder, encoder.category_mapping, encoder.mapping
test_onehot = encoder.transform(test_df)
test_onehot
| notebook/categorical_transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
from bs4 import BeautifulSoup, NavigableString, Tag
import urllib.request
import requests
import pandas as pd
import warnings; warnings.simplefilter('ignore')
response = requests.get('https://www.baseball-reference.com/leagues/MLB/2019-schedule.shtml')
soup = BeautifulSoup(response.text,'html.parser')
alldates = soup.find_all('h3')
data = pd.DataFrame(columns = ['Date','VisitingTeam','VisitingScore','HomeTeam','HomeScore'])
# +
### Scrape game results for all MLB games in 2019
datelist=[]
VisitingTeamList = []
VisitingTeamScoreList = []
HomeTeamList = []
HomeScoreList = []
flag = 0 ## to break out of loops
for header in soup.find_all('h3')[8:190]: ## Games start at 8
nextNode = header
while True:
nextNode = nextNode.nextSibling
if nextNode is None:
break
if (isinstance(nextNode, Tag) and flag == 0):
if nextNode.name == "h2":
break
if header.text == 'Today\'s Games': ## Stop scraping when getting to Today's date
flag +=1
break
datelist.append(header.text) ## Append date
try:
VisitingTeamList.append(nextNode.get_text().splitlines()[1])
except IndexError:
continue
try:
VisitingTeamScoreList.append(nextNode.get_text().splitlines()[2])
except IndexError:
continue
try:
HomeTeamList.append(nextNode.get_text().splitlines()[4])
except IndexError:
continue
try:
HomeScoreList.append(nextNode.get_text().splitlines()[5])
except IndexError:
continue
# -
## Above scraping gets one instance of the date in excess, remove it
for x in set(datelist):
datelist.remove(x)
data['Date'] = datelist
data['VisitingTeam'] = pd.Series(VisitingTeamList)
data['VisitingScore'] = pd.Series(VisitingTeamScoreList)
data['HomeTeam'] = pd.Series(HomeTeamList)
data['HomeScore'] = pd.Series(HomeScoreList)
data['VisitingScore']=data['VisitingScore'].map(lambda x: x.strip(' ()'))
data['HomeScore']=data['HomeScore'].map(lambda x: x.strip(' ()'))
data['Date'] = pd.to_datetime(data['Date'])
data.loc[data['HomeScore'] > data['VisitingScore'],'HomeWin'] = 1
data.loc[data['HomeScore'] < data['VisitingScore'],'HomeWin'] = 0
# +
data['VisitingTeam'] = data['VisitingTeam'].str.strip()
data['HomeTeam'] = data['HomeTeam'].str.strip()
intteamdict = {'Toronto Blue Jays': 'Toronto', 'New York Yankees': 'Yankees', 'Boston Red Sox': 'Redsox', 'Baltimore Orioles': 'Orioles'}
def WinColumn(team): #Figure out if each team won or lost based on whether they were home or away
data[intteamdict[team] + 'Win'] = ""
data.loc[(data['HomeTeam'] == team) & (data['HomeWin'] == 1), intteamdict[team] + 'Win'] = 1
data.loc[(data['HomeTeam'] == team) & (data['HomeWin'] == 0), intteamdict[team] + 'Win'] = -1
data.loc[(data['VisitingTeam'] == team) & (data['HomeWin'] == 0), intteamdict[team] + 'Win'] = 1
data.loc[(data['VisitingTeam'] == team) & (data['HomeWin'] == 1), intteamdict[team] + 'Win'] = -1
WinColumn('Toronto Blue Jays')
WinColumn('New York Yankees')
WinColumn('<NAME>')
WinColumn('Baltimore Orioles')
# -
# +
### Create team specific df
Toronto_df = data.loc[data['TorontoWin'] != ""]
Orioles_df = data.loc[data['OriolesWin'] != ""]
Yankees_df = data.loc[data['YankeesWin'] != ""]
Redsox_df = data.loc[data['RedsoxWin'] != ""]
Toronto_df['GameNumber'] = list(range(1,toronto_df.shape[0]+1))
Orioles_df['GameNumber'] = list(range(1,orioles_df.shape[0]+1))
Yankees_df['GameNumber'] = list(range(1,yankees_df.shape[0]+1))
Redsox_df['GameNumber'] = list(range(1,redsox_df.shape[0]+1))
# -
# +
## Create WinLoss and WinLoss10 columns
def WinLossColumn(team_df,team):
if team == 'Toronto':
team_df.loc[team_df['GameNumber'] ==1,'WinLossRatio'] = 0
team_df.loc[team_df['GameNumber'] ==1,'WinLossRatioLast10'] = 0
team_df.loc[team_df['GameNumber'].isin(list(range(2, team_df.shape[0]+1))),'WinLossRatio'] = team_df[team + 'Win'].cumsum()[:-1].values
team_df.loc[team_df['GameNumber'].isin(list(range(2, team_df.shape[0]+1))),'WinLossRatioLast10'] = team_df[team + 'Win'].rolling(min_periods=1, window=10).sum()[:-1].values
else:
team_df.loc[team_df['GameNumber'] ==1,team+'WinLossRatio'] = 0
team_df.loc[team_df['GameNumber'] ==1,team+'WinLossRatioLast10'] = 0
team_df.loc[team_df['GameNumber'].isin(list(range(2, team_df.shape[0]+1))),team+'WinLossRatio'] = team_df[team + 'Win'].cumsum()[:-1].values
team_df.loc[team_df['GameNumber'].isin(list(range(2, team_df.shape[0]+1))),team+'WinLossRatioLast10'] = team_df[team + 'Win'].rolling(min_periods=1, window=10).sum()[:-1].values
WinLossColumn(Toronto_df,'Toronto')
WinLossColumn(Orioles_df,'Orioles')
WinLossColumn(Redsox_df,'Redsox')
WinLossColumn(Yankees_df,'Yankees')
# +
## Set datetime so you are able to merge on it
def setdatetime(df):
df['Date'] = pd.to_datetime(df['Date'],format= '%Y-%m-%d')
df.set_index('Date',inplace=True,drop=True)
setdatetime(Toronto_df)
setdatetime(Yankees_df)
setdatetime(Orioles_df)
setdatetime(Redsox_df)
# -
## The only columns you need for other teams are Date and WinLossRatio
Redsox_df = Redsox_df[['RedsoxWinLossRatio','RedsoxWinLossRatioLast10']]
Orioles_df = Orioles_df[['OriolesWinLossRatio','OriolesWinLossRatioLast10']]
Yankees_df = Yankees_df[['YankeesWinLossRatio','YankeesWinLossRatioLast10']]
##Merge on date
Toronto_df = pd.merge_asof(Toronto_df, Redsox_df, left_index=True, right_index=True)
Toronto_df = pd.merge_asof(Toronto_df, Orioles_df, left_index=True, right_index=True)
Toronto_df = pd.merge_asof(Toronto_df, Yankees_df, left_index=True, right_index=True)
## Create GamesBack column
Toronto_df['GamesBack'] = Toronto_df['WinLossRatio'] - Toronto_df[['YankeesWinLossRatio','OriolesWinLossRatio','RedsoxWinLossRatio']].max(axis=1)
# +
## Use same team names as historical data
teamdict = {
"Arizona DBacks":"ARI",
"Atlanta Braves":"ATL",
"Baltimore Orioles":"BAL",
"Boston Red Sox":"BOS",
"Chicago Cubs":"CHN",
"Chicago White Sox":"CHA",
"Cincinnati Reds":"CIN",
"Cleveland Indians":"CLE",
"Colorado Rockies":"COL",
"Detroit Tigers":"DET",
"Houston Astros":"HOU",
"Kansas City Royals":"KCA",
"Los Angeles Angels":"LAA",
"Los Angeles Dodgers":"LAN",
"Miami Marlins":"FLO",
"Milwaukee Brewers":"MIL",
"Minnesota Twins":"MIN",
"New York Mets":"NYN",
"New York Yankees":"NYA",
"Oakland Athletics":"OAK",
"Philadelphia Phillies":"PHI",
"Pittsburgh Pirates":"PIT",
"San Diego Padres":"SDN",
"San Francisco Giants":"SFN",
"Seattle Mariners":"SEA",
"St. Louis Cardinals":"SLN",
"Tampa Bay Rays":"TBA",
"Texas Rangers":"TEX",
"Toronto Blue Jays":"TOR",
"Washington Nationals":"WAS",
}
Toronto_df['HomeTeam'] = Toronto_df['HomeTeam'].map(teamdict)
Toronto_df['VisitingTeam'] = Toronto_df['VisitingTeam'].map(teamdict)
# -
Toronto_df = Toronto_df[['WinLossRatio','WinLossRatioLast10','RedsoxWinLossRatio','OriolesWinLossRatio','YankeesWinLossRatio','GamesBack','VisitingTeam']]
Toronto_df.to_csv('Toronto2019WinLoss.csv')
| 2019_AllTeams_WinLoss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature importance
# In this notebook we illustrate the computation of feature importance for a gaussian process.
from fastai.tabular.all import *
from tabularGP import tabularGP_learner
# + [markdown] heading_collapsed=true
# ## Data
# + [markdown] hidden=true
# Builds a regression problem on a subset of the adult dataset:
# + hidden=true
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv').sample(1000)
procs = [FillMissing, Normalize, Categorify]
# + hidden=true
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race', 'salary']
cont_names = ['education-num', 'fnlwgt']
dep_var = 'age'
# + hidden=true
data = TabularDataLoaders.from_df(df, path, procs=procs, cat_names=cat_names, cont_names=cont_names, y_names=dep_var)
# + [markdown] heading_collapsed=true
# ## Model
# + hidden=true
learn = tabularGP_learner(data)
# + hidden=true
learn.fit_one_cycle(5, max_lr=1e-3)
# -
# ## Feature importance
# You can get direct access to the importance of each feature, stored in a dataframe, using the `feature_importance` field.
# This information is deduced from the parameters of the kernel (mostly the scales) and can be accessed instantly:
learn.feature_importance
# You can also plot them directly for a quick analysis (parameters are forwarded to the [Pandas plotting function](https://pandas.pydata.org/pandas-docs/version/0.23.4/generated/pandas.DataFrame.plot.html)):
learn.plot_feature_importance()
| examples/4_feature_importance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="NT0fNtYLFyP8"
# # Named Entity Recognition (NER)
# spaCy has an **'ner'** pipeline component that identifies token spans fitting a predetermined set of named entities. These are available as the `ents` property of a `Doc` object.
# + colab={"base_uri": "https://localhost:8080/", "height": 390} colab_type="code" id="Ec_QhKxTGCFR" outputId="3b998edf-78d5-4f41-f2e0-597b712971e4"
pip install -U spacy[cuda92]
# + colab={} colab_type="code" id="M655KUdkFyP9"
# Perform standard imports
import spacy
spacy.prefer_gpu()
nlp = spacy.load('en_core_web_sm')
# + colab={} colab_type="code" id="I_FDKDZzFyQA"
# Write a function to display basic entity info:
def show_ents(doc):
if doc.ents:
for ent in doc.ents:
print(ent.text+' - '+ent.label_+' - '+str(spacy.explain(ent.label_)))
else:
print('No named entities found.')
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="GOCYs6wKFyQC" outputId="1e2452e5-5289-47ef-8c9e-573a30c04041"
doc = nlp(u'May I go to Washington, DC next May to see the Washington Monument?')
show_ents(doc)
# + [markdown] colab_type="text" id="_yYw2u_MFyQG"
# Here we see tokens combine to form the entities `Washington, DC`, `next May` and `the Washington Monument`
# + [markdown] colab_type="text" id="a9NQ3CWWFyQH"
# ## Entity annotations
# `Doc.ents` are token spans with their own set of annotations.
# <table>
# <tr><td>`ent.text`</td><td>The original entity text</td></tr>
# <tr><td>`ent.label`</td><td>The entity type's hash value</td></tr>
# <tr><td>`ent.label_`</td><td>The entity type's string description</td></tr>
# <tr><td>`ent.start`</td><td>The token span's *start* index position in the Doc</td></tr>
# <tr><td>`ent.end`</td><td>The token span's *stop* index position in the Doc</td></tr>
# <tr><td>`ent.start_char`</td><td>The entity text's *start* index position in the Doc</td></tr>
# <tr><td>`ent.end_char`</td><td>The entity text's *stop* index position in the Doc</td></tr>
# </table>
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="_cP9qnCEFyQI" outputId="5eeaf018-d62d-4d17-fc30-7425e55ad6cc"
doc = nlp(u'Can I please borrow 500 dollars from you to buy some Microsoft stock?')
for ent in doc.ents:
print(ent.text, ent.start, ent.end, ent.start_char, ent.end_char, ent.label_)
# + [markdown] colab_type="text" id="Bqvuj6byFyQK"
# ## NER Tags
# Tags are accessible through the `.label_` property of an entity.
# <table>
# <tr><th>TYPE</th><th>DESCRIPTION</th><th>EXAMPLE</th></tr>
# <tr><td>`PERSON`</td><td>People, including fictional.</td><td>*<NAME>*</td></tr>
# <tr><td>`NORP`</td><td>Nationalities or religious or political groups.</td><td>*The Republican Party*</td></tr>
# <tr><td>`FAC`</td><td>Buildings, airports, highways, bridges, etc.</td><td>*Logan International Airport, The Golden Gate*</td></tr>
# <tr><td>`ORG`</td><td>Companies, agencies, institutions, etc.</td><td>*Microsoft, FBI, MIT*</td></tr>
# <tr><td>`GPE`</td><td>Countries, cities, states.</td><td>*France, UAR, Chicago, Idaho*</td></tr>
# <tr><td>`LOC`</td><td>Non-GPE locations, mountain ranges, bodies of water.</td><td>*Europe, Nile River, Midwest*</td></tr>
# <tr><td>`PRODUCT`</td><td>Objects, vehicles, foods, etc. (Not services.)</td><td>*Formula 1*</td></tr>
# <tr><td>`EVENT`</td><td>Named hurricanes, battles, wars, sports events, etc.</td><td>*Olympic Games*</td></tr>
# <tr><td>`WORK_OF_ART`</td><td>Titles of books, songs, etc.</td><td>*The Mona Lisa*</td></tr>
# <tr><td>`LAW`</td><td>Named documents made into laws.</td><td>*Roe v. Wade*</td></tr>
# <tr><td>`LANGUAGE`</td><td>Any named language.</td><td>*English*</td></tr>
# <tr><td>`DATE`</td><td>Absolute or relative dates or periods.</td><td>*20 July 1969*</td></tr>
# <tr><td>`TIME`</td><td>Times smaller than a day.</td><td>*Four hours*</td></tr>
# <tr><td>`PERCENT`</td><td>Percentage, including "%".</td><td>*Eighty percent*</td></tr>
# <tr><td>`MONEY`</td><td>Monetary values, including unit.</td><td>*Twenty Cents*</td></tr>
# <tr><td>`QUANTITY`</td><td>Measurements, as of weight or distance.</td><td>*Several kilometers, 55kg*</td></tr>
# <tr><td>`ORDINAL`</td><td>"first", "second", etc.</td><td>*9th, Ninth*</td></tr>
# <tr><td>`CARDINAL`</td><td>Numerals that do not fall under another type.</td><td>*2, Two, Fifty-two*</td></tr>
# </table>
# + [markdown] colab_type="text" id="Xb0zcaWpFyQL"
# ___
# ## Adding a Named Entity to a Span
# Normally we would have spaCy build a library of named entities by training it on several samples of text.<br>In this case, we only want to add one value:
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="9iNiaZL6FyQL" outputId="19d9802e-76d2-49f7-fb01-bfbee6f43336"
doc = nlp(u'Tesla to build a U.K. factory for $6 million')
show_ents(doc)
# + [markdown] colab_type="text" id="AgW5MSwFFyQN"
# <font color=green>Right now, spaCy does not recognize "Tesla" as a company.</font>
# + colab={} colab_type="code" id="Ce6dZ1kJFyQO"
from spacy.tokens import Span
# Get the hash value of the ORG entity label
ORG = doc.vocab.strings[u'ORG']
# Create a Span for the new entity
new_ent = Span(doc, 0, 1, label=ORG)
# Add the entity to the existing Doc object
doc.ents = list(doc.ents) + [new_ent]
# + [markdown] colab_type="text" id="Z1VysmzTFyQP"
# <font color=green>In the code above, the arguments passed to `Span()` are:</font>
# - `doc` - the name of the Doc object
# - `0` - the *start* index position of the span
# - `1` - the *stop* index position (exclusive)
# - `label=ORG` - the label assigned to our entity
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="Qbkdml1RFyQQ" outputId="9f69cf94-7ec5-46ea-8b57-7fb7c512e239"
show_ents(doc)
# + [markdown] colab_type="text" id="tL1UcSS3FyQS"
# ___
# ## Adding Named Entities to All Matching Spans
# What if we want to tag *all* occurrences of "Tesla"? In this section we show how to use the PhraseMatcher to identify a series of spans in the Doc:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="jMQSN9SNFyQS" outputId="693d31dc-5697-4407-d64d-0f015dcbd656"
doc = nlp(u'Our company plans to introduce a new vacuum cleaner. '
u'If successful, the vacuum cleaner will be our first product.')
show_ents(doc)
# + colab={} colab_type="code" id="UXbWFRgSFyQV"
# Import PhraseMatcher and create a matcher object:
from spacy.matcher import PhraseMatcher
matcher = PhraseMatcher(nlp.vocab)
# + colab={} colab_type="code" id="x6gDIZNCFyQW"
# Create the desired phrase patterns:
phrase_list = ['vacuum cleaner', 'vacuum-cleaner']
phrase_patterns = [nlp(text) for text in phrase_list]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="wZMuaZ7mFyQZ" outputId="d6f5c7be-a7a7-4217-8d50-119b35c2c6db"
# Apply the patterns to our matcher object:
matcher.add('newproduct', None, *phrase_patterns)
# Apply the matcher to our Doc object:
matches = matcher(doc)
# See what matches occur:
matches
# + colab={} colab_type="code" id="F_1UGfzMFyQb"
# Here we create Spans from each match, and create named entities from them:
from spacy.tokens import Span
PROD = doc.vocab.strings[u'PRODUCT']
new_ents = [Span(doc, match[1],match[2],label=PROD) for match in matches]
doc.ents = list(doc.ents) + new_ents
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="-18PVOtqFyQe" outputId="f1ccd39f-38ed-4439-89f0-70a79523591f"
show_ents(doc)
# + [markdown] colab_type="text" id="6RP3XqcZFyQg"
# ___
# ## Counting Entities
# While spaCy may not have a built-in tool for counting entities, we can pass a conditional statement into a list comprehension:
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="1RDvY1YuFyQh" outputId="2def8844-f93c-40a6-ffb4-9d856d703ca4"
doc = nlp(u'Originally priced at $29.50, the sweater was marked down to five dollars.')
show_ents(doc)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="s6wMXGbFFyQk" outputId="5df7579c-8c82-49b3-b444-8ee690b47f06"
len([ent for ent in doc.ents if ent.label_=='MONEY'])
# + [markdown] colab_type="text" id="5GufDzOeFyQn"
# ## <font color=blue>Problem with Line Breaks</font>
#
# <div class="alert alert-info" style="margin: 20px">There's a <a href='https://github.com/explosion/spaCy/issues/1717'>known issue</a> with <strong>spaCy v2.0.12</strong> where some linebreaks are interpreted as `GPE` entities:</div>
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Hnrh9ipdFyQo" outputId="607d4651-688d-4def-e116-9085691616d5"
spacy.__version__
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="902BVPS2FyQr" outputId="bb95a8c0-219d-43ff-d6e1-4f0ec01093e9"
doc = nlp(u'Originally priced at $29.50,\nthe sweater was marked down to five dollars.')
show_ents(doc)
# + [markdown] colab_type="text" id="VHYmRCxYFyQt"
# ### <font color=blue>However, there is a simple fix that can be added to the nlp pipeline:</font>
# + colab={} colab_type="code" id="PDSoRI5wFyQu"
# Quick function to remove ents formed on whitespace:
def remove_whitespace_entities(doc):
doc.ents = [e for e in doc.ents if not e.text.isspace()]
return doc
# Insert this into the pipeline AFTER the ner component:
nlp.add_pipe(remove_whitespace_entities, after='ner')
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="ruQbOCoyFyQv" outputId="b2fc4a24-b493-496a-a140-ced006e68d6c"
# Rerun nlp on the text above, and show ents:
doc = nlp(u'Originally priced at $29.50,\nthe sweater was marked down to five dollars.')
show_ents(doc)
# + [markdown] colab_type="text" id="L_0o_qo5FyQx"
# For more on **Named Entity Recognition** visit https://spacy.io/usage/linguistic-features#101
# + [markdown] colab_type="text" id="rnuXo59CFyQx"
# ___
# ## Noun Chunks
# `Doc.noun_chunks` are *base noun phrases*: token spans that include the noun and words describing the noun. Noun chunks cannot be nested, cannot overlap, and do not involve prepositional phrases or relative clauses.<br>
# Where `Doc.ents` rely on the **ner** pipeline component, `Doc.noun_chunks` are provided by the **parser**.
# + [markdown] colab_type="text" id="5EBL_yb1FyQy"
# ### `noun_chunks` components:
# <table>
# <tr><td>`.text`</td><td>The original noun chunk text.</td></tr>
# <tr><td>`.root.text`</td><td>The original text of the word connecting the noun chunk to the rest of the parse.</td></tr>
# <tr><td>`.root.dep_`</td><td>Dependency relation connecting the root to its head.</td></tr>
# <tr><td>`.root.head.text`</td><td>The text of the root token's head.</td></tr>
# </table>
# + colab={"base_uri": "https://localhost:8080/", "height": 70} colab_type="code" id="JX28D_z9FyQy" outputId="a3e6547a-4fb2-49dd-b2bb-885687ad221e"
doc = nlp(u"Autonomous cars shift insurance liability toward manufacturers.")
for chunk in doc.noun_chunks:
print(chunk.text+' - '+chunk.root.text+' - '+chunk.root.dep_+' - '+chunk.root.head.text)
# + [markdown] colab_type="text" id="imJT3cP7FyQ1"
# ### `Doc.noun_chunks` is a generator function
# Previously we mentioned that `Doc` objects do not retain a list of sentences, but they're available through the `Doc.sents` generator.<br>It's the same with `Doc.noun_chunks` - lists can be created if needed:
# + colab={"base_uri": "https://localhost:8080/", "height": 167} colab_type="code" id="4e0yg8IUFyQ1" outputId="08258aaa-4d85-452d-a012-a353f3629fa7"
len(doc.noun_chunks)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ji9sjTVgFyQ3" outputId="3f6a0053-4f0d-4726-8eeb-afcd553b512b"
len(list(doc.noun_chunks))
# + [markdown] colab_type="text" id="a_K9Cm3uFyQ6"
# For more on **noun_chunks** visit https://spacy.io/usage/linguistic-features#noun-chunks
# + colab={} colab_type="code" id="uOEPsXSPGcMC"
# + colab={} colab_type="code" id="SUfqIN7vIGJ-"
# Perform standard imports
import spacy
nlp = spacy.load('en_core_web_sm')
# Import the displaCy library
from spacy import displacy
# + colab={} colab_type="code" id="gz-j8GpkIGKA" outputId="e11a8944-2b76-4b88-a039-0ba949a86bce"
doc = nlp(u'Over the last quarter Apple sold nearly 20 thousand iPods for a profit of $6 million. '
u'By contrast, Sony sold only 7 thousand Walkman music players.')
displacy.render(doc, style='ent', jupyter=True)
# + colab={} colab_type="code" id="XebKurgEIgDq"
# + colab={} colab_type="code" id="1hFosKW7IGKD" outputId="41c61c67-414b-4403-b9f3-a93726a57760"
for sent in doc.sents:
displacy.render(nlp(sent.text), style='ent', jupyter=True)
# + colab={} colab_type="code" id="v0uEcP0cIGKG"
doc2 = nlp(u'Over the last quarter Apple sold nearly 20 thousand iPods for a profit of $6 million. '
u'By contrast, my kids sold a lot of lemonade.')
# + colab={} colab_type="code" id="NX_s73r2IGKI" outputId="3acc985f-3bc9-4c91-d328-794b73692803"
for sent in doc2.sents:
displacy.render(nlp(sent.text), style='ent', jupyter=True)
# + colab={} colab_type="code" id="qXdOLILxIGKL" outputId="9a29e302-6bcc-4081-ceef-8b4d2b5613a3"
for sent in doc2.sents:
docx = nlp(sent.text)
if docx.ents:
displacy.render(docx, style='ent', jupyter=True)
else:
print(docx.text)
# + colab={} colab_type="code" id="KyQvseRaIGKO" outputId="9912c9cc-4a70-4520-c792-940c472cf936"
options = {'ents': ['ORG', 'PRODUCT']}
displacy.render(doc, style='ent', jupyter=True, options=options)
# + colab={} colab_type="code" id="dL7fp6miIGKQ" outputId="b870e49a-1045-4689-e25c-abaf6dad8e38"
colors = {'ORG': 'linear-gradient(90deg, #aa9cfc, #fc9ce7)', 'PRODUCT': 'radial-gradient(yellow, green)'}
options = {'ents': ['ORG', 'PRODUCT'], 'colors':colors}
displacy.render(doc, style='ent', jupyter=True, options=options)
# + [markdown] colab_type="text" id="3cuZW1l1IGKT"
# ___
# # Creating Visualizations Outside of Jupyter
# If you're using another Python IDE or writing a script, you can choose to have spaCy serve up HTML separately.
#
# Instead of `displacy.render()`, use `displacy.serve()`:
# + colab={} colab_type="code" id="C-xYfpzuIGKT" outputId="c0226ed6-e700-4e77-84fe-7efa6ceabc70"
displacy.serve(doc, style='ent', options=options)
| Data-Science-Portfolio-master/Text Analytics/Named_Entity_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
df = pd.read_excel("Data/New_Demos.xlsx")
df_D = df[['Station', "White", "Latino", "Black", "Asian"]]
df_G = df[['Station', "Male", "Female"]]
## Get percentages from counts
df_D = df_D.set_index("Station")
df_D = df_D.div(df_D.sum(axis=1), axis=0).multiply(100).round(decimals=1)
df_D.loc['US Population'] = [60.0, 18.4, 12.4, 5.6]
df_D['Other'] = 0.0
df_D.at['US Population', "Other"] = 3.6
df_G = df_G.set_index("Station")
df_G = df_G.div(df_G.sum(axis=1), axis=0).multiply(100).round(decimals=1)
df_G.loc['US Population'] = [49.2, 50.8]
def mse(dataframe, size):
baseline = list(dataframe.iloc[5])
sums = []
for item in range(5):
comparison = list(dataframe.iloc[item])
## Calculate MSE
amount = [(abs(j-i)**2)/size for i,j in zip(baseline, comparison)]
sums.append(sum(amount))
sums = ['%.1f' % elem for elem in sums]
sums = [float(elem) for elem in sums]
sums.append(0)
return(sums)
df_D['MSE'] = mse(df_D, 5)
df_G['MSE'] = mse(df_G, 2)
df_D = df_D.sort_values(by="MSE")
df_G = df_G.sort_values(by="MSE")
df_D
# -
df_G
df
# #### US Demographics
# +
df_G = df_G[['Male', "Female"]]
df_D = df_D[["White", "Latino", "Black", "Asian", "Other"]]
q = df_G.transpose()
w = df_D.transpose()
fig = plt.figure(figsize=(16,8))
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['US Population'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['US Population'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral', "steelblue"])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/US_Pop.png", dpi=400)
# +
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
from plotly import tools
import numpy as np
cmap = plt.get_cmap('Set2')
colors = [cmap(i) for i in np.linspace(0, 1, 8)]
gender_pie = go.Pie(labels=q['US Population'].index, values=q['US Population'], marker=dict(colors=colors[1:3]
, line=dict(color='#FFF', width=2)),
domain={'x': [0.0, .4], 'y': [0.0, 1]}
, showlegend=False, name='Gender', textinfo='label+percent')
demo_pie = go.Pie(labels=w['US Population'].index, values=w['US Population'], marker=dict(colors=colors
, line=dict(color='#FFF', width=2)),
domain={'x': [.6, 1], 'y': [0.0, 1]}
, showlegend=False, name='Demographics', textinfo='label+percent')
layout = go.Layout(height = 600,
width = 1000,
autosize = False,
title = 'Gender and Racial Demographics of US Population')
fig = go.Figure(data = [gender_pie,demo_pie ], layout = layout)
py.iplot(fig, filename='basic_pie_chart')
# +
## Remove Other Category from Pie Charts (Didn't collect Other Category)
df_G = df_G[['Male', "Female"]]
df_D = df_D[["White", "Latino", "Black", "Asian"]]
q = df_G.transpose()
w = df_D.transpose()
# -
# #### Fox News
# +
fig = plt.figure(figsize=(16,8), dpi=200)
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['Fox'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['Fox'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral'])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/Fox.png", dpi=400)
# + [markdown] heading_collapsed=true
# #### CNN
# + hidden=true
fig = plt.figure(figsize=(16,8), dpi=200)
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['CNN'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['CNN'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral'])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/CNN.png", dpi=400)
# + [markdown] heading_collapsed=true
# #### ABC
# + hidden=true
fig = plt.figure(figsize=(16,8), dpi=400)
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['ABC'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['ABC'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral'])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/ABC.png", dpi=400)
# + [markdown] heading_collapsed=true
# #### CBS
# + hidden=true
fig = plt.figure(figsize=(16,8), dpi=200)
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['CBS'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['CBS'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral'])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/CBS.png", dpi=400)
# + [markdown] heading_collapsed=true
# #### NBC
# + hidden=true
fig = plt.figure(figsize=(16,8), dpi=200)
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
z = q['NBC'].plot(kind='pie', ax=ax1, autopct='%1.1f%%', startangle=90, fontsize=16, colors=['dodgerblue', 'pink'])
t = w['NBC'].plot(kind='pie', ax=ax2, autopct='%1.1f%%', startangle=90, fontsize=12, colors=['thistle', 'seagreen', 'gold', 'coral'])
plt.ylabel('')
ax1.yaxis.label.set_visible(False)
plt.savefig("Images/NBC.png", dpi=400)
# -
# #### End of Script
| News_Demo_Graphics .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
from astropy.io import fits
import astropy.units as u
import pickle as pickle
import os as os
import pandas as pd
from spectools_ir.utils import extract_hitran_data, spec_convol, make_rotation_diagram, get_molmass
from spectools_ir.utils import compute_thermal_velocity, sigma_to_fwhm, fwhm_to_sigma, wn_to_k, spec_convol_R
from spectools_ir.utils import get_miri_mrs_resolution, get_miri_mrs_wavelengths, make_miri_mrs_figure
from spectools_ir.flux_calculator import calc_fluxes, make_lineshape
from spectools_ir.slabspec import make_spec
from spectools_ir.slab_fitter import Config, LineData,Retrieval
from spectools_ir.slab_fitter import corner_plot, trace_plot, find_best_fit, compute_model_fluxes
from spectools_ir.slab_fitter import calc_solid_angle, calc_radius
from spectools_ir.slab_fitter import read_data_from_file, get_samples
# -
import spectools_ir
spectools_ir.__file__
# # Flux_calculator example use
# Flux calculator will loop through a set of provided wavelengths, then perform a Gaussian and numerical computation of line flux at each location. If one begins by using the helper tool "extract_hitran_data", the output will include transition-specific data in addition to the computed line fluxes.
# ### Read in HITRAN data with extract_hitran_data
# This example uses extract_hitran_data to find fundamental CO vibrational transitions in the M band. A spectrum is read in from a FITS file.
hitran_data=extract_hitran_data('CO',4.58,5.2,vup=1) #Outputs an astropy table
#Read in FITS data containing spectrum.
infile='nirspec_doar44_glue.dat' #my file name
wave,flux = np.loadtxt(infile, unpack=True, skiprows=29) #make sure to skip correct amount of rows in data set
# +
#Plot spectrum.
fig=plt.figure(figsize=(18, 6))
ax1=fig.add_subplot(211)
ax1.plot(wave,flux)
ax1.set_xlim(4.645,4.783)
for i,mywave in enumerate(hitran_data['wave']):
if( (mywave>4.645) & (mywave<4.783) ):
ax1.axvline(mywave,color='C1')
ax1.text(hitran_data['wave'][i],0.67,hitran_data['Qpp'][i].strip())
ax1.set_ylabel('Flux [Jy]',fontsize=14)
ax2=fig.add_subplot(212)
ax2.plot(wave,flux)
ax2.set_xlim(4.95,5.10)
for i,mywave in enumerate(hitran_data['wave']):
if( (mywave>4.95) & (mywave<5.10) ):
ax2.axvline(mywave,color='C1')
ax2.text(hitran_data['wave'][i],0.67,hitran_data['Qpp'][i].strip())
ax2.set_xlabel('Wavelength [$\mu$m]',fontsize=14)
ax2.set_ylabel('Flux [Jy]',fontsize=14)
# +
#How to fix where P/R labels are? Change value next to hitran_data in ax2.text
# -
# ### Calculate line fluxes for dataset, using locations specified in hitran dataset
# calc_fluxes is now used to compute the line fluxes in the spectrum. (Some additional code is added to read and write in the flux values for use in later parts of this notebook.) Users can provide input values of the continuum level (cont), FWHM (fwhm_v) and Doppler shift of lines relative to vacuum rest wavelength (v_dop). Users can also choose the size of the fit interval (sep_v), whether or not to vet the fits (vet_fits; highly recommended) and whether or not to plot (highly recommended). The routine does little to no vetting on its own, so user vetting is highly recommended.
# +
#Calculate fluxes.
#Uncomment the following two lines to run the code
lineflux_data=calc_fluxes(wave,flux,hitran_data, v_dop=-5,fwhm_v=50.,sep_v=250.,cont=0.5,vet_fits=True, plot=True, ymax=0.8, ymin=0.4)
pickle.dump(lineflux_data,open('lineflux_data.p','wb')) #save for convenience
#lineflux_data=pickle.load(open('lineflux_data.p','rb')) #restore lineflux_data if you don't want to re-run
# +
#Removed P35, P33, P29, P13, R0, R1
# -
# Note that flux_calculator calculates fluxes using both a Gaussian fit and numerical integration. The numerical integration sums from -3$\sigma$ to +3$\sigma$, where $\sigma$ is provided by the Gaussian fit. These values should be similar for Gaussian line shapes, but may be quite different for more complex line shapes, as for AA Tau.
#
# The figure below shows how the two compare for this example.
lineflux_data
fig=plt.figure()
ax1=fig.add_subplot(111)
ax1.plot(lineflux_data['wave'],lineflux_data['lineflux']*1e16, '-o', label='Numerical flux')
ax1.plot(lineflux_data['wave'],lineflux_data['lineflux_Gaussian']*1e16, '-o', label='Gaussian flux')
ax1.legend()
ax1.set_xlabel('Wavelength [microns]')
ax1.set_ylabel(r'Line flux [10$^{-16}$ W m$^{-2}$]')
# ### Create a composite lineshape using info from flux_calculator output
# Flux calculator also has a routine that can make a composite lineshape from calc_fluxes output.
lineshape_data=make_lineshape(wave,flux,lineflux_data)
# +
fig=plt.figure()
ax1=fig.add_subplot(111)
ax1.plot(lineshape_data[0],lineshape_data[1])
ax1.set_xlabel('Velocity [km/s]')
ax1.set_ylabel('Arbitrary flux')
# -
# ### Display resultant fluxes in a rotation diagram
# Spectools_ir.utils has a function to make a rotation diagram, which can use calc_fluxes output as its dataset. This example compares rotation diagrams for the Gaussian and numerical fluxes.
#
# make_rotation_diagrams has a "units" keyword with values 'mks' (SI), 'cgs', or 'mixed'. Based on some precedent in the literature, 'mixed' units keeps most values in SI units, but wavenumbers in inverse cm.
# +
rot_numerical=make_rotation_diagram(lineflux_data,fluxkey='lineflux')
rot_Gaussian=make_rotation_diagram(lineflux_data,fluxkey='lineflux_Gaussian')
fig=plt.figure()
ax1=fig.add_subplot(111)
ax1.plot(rot_numerical['x'],rot_numerical['y'],'C0o',label='Numerical fit')
ax1.plot(rot_Gaussian['x'],rot_Gaussian['y'],'C1o', label='Gaussian fit')
ax1.set_xlabel('Eup [K]')
ax1.set_ylabel(r'ln(F/(gA$\tilde{\nu}$))')
ax1.legend()
# -
# # Slabspec example use
# Slabspec makes a model spectrum from a "slab" of gas - a column of gas with defined area, column density, and (single) temperature. The example below shows the creation of a CO spectrum from 4.58 to 5.2 microns. Units are assumed to be SI (mks). If convol_fwhm is specified, the model output will be convolved with a Gaussian with the given FWHM value (in km/s). Both the convolved and unconvolved spectra are included in the output.
#
# **Important note**: The model can produce wonky output if the model resolution ('res' parameter: resolution in microns) is not sufficient. It's highly recommended that one check the output of a model against a model with higher resolution to make sure the output remains the same.
# ### Make a slab model spectrum with make_spec
au=1.5e11
#Make slab model
#This can take a few seconds to run. It is primarily limited by the convolution time.
model_output=make_spec('CO',10**(21.6),1370,np.pi*(0.158*au)**2., wmin=4.58,wmax=5.2,convol_fwhm=30.,d_pc=146.3,res=1e-5,vup=1)
#Plot slab model
fig=plt.figure(figsize=(10,3))
ax1=fig.add_subplot(111)
ax1.plot(model_output['spectrum']['wave'],model_output['spectrum']['flux']+1,label='Not convolved')
ax1.plot(model_output['spectrum']['wave'],model_output['spectrum']['convolflux']+1,'r',label='Convolved')
ax1.legend()
# ### Output model fluxes as a rotation diagram (compare to data if desired)
# Output from slabspec can also be displayed as a rotation diagram.
rd_model=make_rotation_diagram(model_output['lineparams'])
rd_data=make_rotation_diagram(lineflux_data)
fig=plt.figure()
ax1=fig.add_subplot(111)
ax1.plot(rd_model['x'],rd_model['y'],' C0o',label='Model')
ax1.plot(rd_data['x'],rd_data['y'],' C1*',label='Data')
ax1.set_xlabel('Eup [K]')
ax1.set_ylabel(r'ln(F/(gA$\tilde{\nu}$))')
ax1.set_xlim(3000,6000)
ax1.set_ylim(-60,-53)
ax1.legend()
# # Slab_fitter example use
# slab_fitter uses the MCMC fitting code "emcee" to fit line fluxes with a slab model. Priors are assumed to be flat, with boundaries provided by the user as input parameters.
#
# In contrast to the slab model, slab_fitter uses solid angle in place of area. Helper functions calc_solid_angle(radius, distance) and calc_radius(solid_angle,distance) can be used to convert between the two. Calculations assume a face-on disk shape.
# ### Set up configuration
# Configuration parameters are all stored in config.json, or another user-specified json file. User must specify the range of temperatures, column densities, and solid angles, the number of walkers, the number of burnin samples, and the number of samples per walker.
#
# slab_fitter naturally takes output of flux_calculator as input. (Other input examples are discussed below).
# +
myconfig=Config(config_file='/Users/erichegonzales/Desktop/eriche-thesis/data/doar44.json') #reads config.json by default, or can specify path to file as input parameter
myconfig.display()
data=lineflux_data
mydata=LineData(data)
myretrieval=Retrieval(myconfig,mydata)
# -
# ### Run retrieval
# Running the retrieval can take a significant amount of time, so only run when ready. (Code below currently takes 9 minutes on my personal laptop.)
# +
mychain=myretrieval.run_emcee() #Run retrieval. May take a while, depending on number of samples
pickle.dump(mychain,open('test_chain.p','wb')) #Save for convenience and later retrieval
os.system('afplay /System/Library/Sounds/Sosumi.aiff')
mychain=pickle.load(open('test_chain.p','rb'))
postsamples=get_samples(mychain,myconfig.getpar('Nburnin')) #Get list of samples, after removing burnin, from chain
# -
# ### Evaluate retrieval results
# slab_fitter includes a few simple routines for evaluating and viewing the retrieval results.
#
# **Note:** The examples here are not necessarily well-behaved samples. Users should refer to emcee documentation and other sources for more info about MCMC sample vetting.
#Trace plot - examine samples
trace_plot(postsamples)
#Corner plot - see posterior distributions
fig=corner_plot(postsamples)
#Best fit parameters - best fit parameters plus/minus 1-sigma intervals
best_fit=find_best_fit(postsamples,show=True)
# +
#Rotation diagram
modelflux=compute_model_fluxes(mydata,postsamples) #Compute model fluxes for ecah line in data
rotdiag=mydata.rot_diagram(modelfluxes=modelflux,units='mks') #Rotation diagram for data and model
fig=plt.figure()
ax1=fig.add_subplot(111)
ax1.plot(rotdiag['x'],rotdiag['modely'],'bo',label='Model')
ax1.plot(rotdiag['x'],rotdiag['modely'],'b',label='_None_')
ax1.plot(rotdiag['x'],rotdiag['y'],'ro',label='Data')
ax1.set_xlabel('Upper Level Energy [Kelvin]')
#ax1.set_ylabel(r'F[W m$^{-2}$]/($\~{\nu}$ [m$^{-1}$]g A[s$^{-1}$])',fontsize=14)
ax1.set_ylabel(r'F]/($\~{\nu}$ g A)',fontsize=14)
ax1.legend()
# -
| data/.ipynb_checkpoints/DoAr44_data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="gqwrcHFAVwgs"
# # Quick Start
# + [markdown] colab_type="text" id="oz-X6ZrGVwgu"
# In this section, we would like to show an overview to give a quick start.
# -
# ## Operation
# There are some operations supported in `tednet`, and it is convinient to use them.
import tednet as tdt
# **Create matrix whose diagonal elements are ones**
diag_matrix = tdt.eye(5, 5)
print(diag_matrix)
# **Take Pytorch tensor to Numpy narray**
print(type(diag_matrix))
diag_matrix = tdt.to_numpy(diag_matrix)
print(type(diag_matrix))
# **Take Numpy narray to Pytorch tensor**
diag_matrix = tdt.to_tensor(diag_matrix)
print(type(diag_matrix))
# ## Tensor Decomposition Networks (Tensor Ring for Sample)
# **To use tensor ring decomposition models, simply calling the tensor ring module is enough.**
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ri9QCNEAVwgw" outputId="efd8c64b-836e-449b-a385-dd3a63a5b4a2"
import tednet.tnn.tensor_ring as tr
# -
# **Here, we would like to give a case of building the TR-LeNet5.**
# + colab={} colab_type="code" id="5Awp7wdwVwg3"
# Define a TR-LeNet5
model = tr.TRLeNet5(10, [6, 6, 6, 6])
| docs/source/quick_start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # sentence segmentation in text
# # 01- Extract Text from PDF Files using NLP
# First step I will do is to bulid a model that ectract text from PDF file to txt file.
# We have to install PyPDF2 library
# Learn more about PyPDF2 library:https://pythonhosted.org/PyPDF2/
# !pip install PyPDF2
import PyPDF2 as pdf
# I will try it with an AI Lacture (L4)
file= open('Presentation.pdf' , 'rb')
file
pdf_reader = pdf.PdfFileReader(file)
pdf_reader
pdf_reader.getIsEncrypted()
pdf_reader.getNumPages()
page1= pdf_reader.getPage(0)
page1.extractText()
page3= pdf_reader.getPage(2)
page3.extractText()
pdf_writer = pdf.PdfFileWriter()
# !pip install nltk
import nltk
nltk.download('punkt')
# ##I will do the rest of the work later
| 1st task for university graduation project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Визуализация данных в Matplotlib.
# ## Задание 1
# Загрузите модуль pyplot библиотеки matplotlib с псевдонимом plt, а также библиотеку numpy с псевдонимом np.
# Примените магическую функцию %matplotlib inline для отображения графиков в Jupyter Notebook и настройки конфигурации ноутбука со значением 'svg' для более четкого отображения графиков.
# Создайте список под названием x с числами 1, 2, 3, 4, 5, 6, 7 и список y с числами 3.5, 3.8, 4.2, 4.5, 5, 5.5, 7.
# С помощью функции plot постройте график, соединяющий линиями точки с горизонтальными координатами из списка x и вертикальными - из списка y.
# Затем в следующей ячейке постройте диаграмму рассеяния (другие названия - диаграмма разброса, scatter plot).
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
x = [1, 2, 3, 4, 5, 6, 7]
y = [3.5, 3.8, 4.2, 4.5, 5, 5.5, 7]
plt.plot(x ,y)
plt.show()
plt.scatter(x, y)
plt.show()
# ## Задание 2
# С помощью функции linspace из библиотеки Numpy создайте массив t из 51 числа от 0 до 10 включительно.
# Создайте массив Numpy под названием f, содержащий косинусы элементов массива t.
# Постройте линейную диаграмму, используя массив t для координат по горизонтали, а массив f - для координат по вертикали. Линия графика должна быть зеленого цвета.
# Выведите название диаграммы - 'График f(t)'.
# Также добавьте названия для горизонтальной оси - 'Значения t' и для вертикальной - 'Значения f'.
# Ограничьте график по оси x значениями 0.5 и 9.5, а по оси y - значениями -2.5 и 2.5.
t = np.linspace(0, 10, 51)
print(t)
f = np.cos(t)
print(f)
plt.plot(t, f, color='green')
plt.title('График f(t)')
plt.xlabel('Значения t')
plt.ylabel('Значения f')
plt.axis([0.5, 9.5, -2.5, 2.5])
plt.show()
# ## Задание 3
# С помощью функции linspace библиотеки Numpy создайте массив x из 51 числа от -3 до 3 включительно.
# Создайте массивы $y_1$, $y_2$, $y_3$, $y_4$ по следующим формулам:
# $$y_1 = x^2$$
# $$y_2 = 2 * x + 0.5$$
# $$y_3 = -3 * x - 1.5$$
# $$y_4 = \sin(x)$$
# Используя функцию subplots модуля matplotlib.pyplot, создайте объект matplotlib.figure.Figure с названием fig и массив объектов Axes под названием ax, причем так, чтобы у вас было 4 отдельных графика в сетке, состоящей из двух строк и двух столбцов. В каждом графике массив x используется для координат по горизонтали.
# В левом верхнем графике для координат по вертикали используйте $y_1$, в правом верхнем - $y_2$, в левом нижнем - $y_3$, в правом нижнем - $y_4$.
# Дайте название графикам: 'График $y_1$', 'График $y_2$' и т.д.
# Для графика в левом верхнем углу установите границы по оси x от -5 до 5.
# Установите размеры фигуры 8 дюймов по горизонтали и 6 дюймов по вертикали.
# Вертикальные и горизонтальные зазоры между графиками должны составлять 0.3.
x = np.linspace(-3, 3, 51)
print(x)
y1 = x**2
print(y1)
y2 = 2 * x + 0.5
print(y2)
y3 = -3 * x - 1.5
print(y3)
y4 = np.sin(x)
print(y4)
fig, ax = plt.subplots(nrows=2, ncols=2)
ax1, ax2, ax3, ax4 = ax.flatten()
ax1.plot(x, y1)
ax2.plot(x, y2)
ax3.plot(x, y3)
ax4.plot(x, y4)
ax1.set_title('График $y_1$')
ax2.set_title('График $y_2$')
ax3.set_title('График $y_3$')
ax4.set_title('График $y_4$')
ax1.set_xlim([-5, 5])
fig.set_size_inches(8, 6)
plt.subplots_adjust(wspace=0.3, hspace=0.3)
plt.show()
#
# ## Задание 4
# В этом задании мы будем работать с датасетом, в котором приведены данные по мошенничеству с кредитными данными:
# Credit Card Fraud Detection (информация об авторах: <NAME>, <NAME>, <NAME> and <NAME>. Calibrating Probability with Undersampling for Unbalanced Classification. In Symposium on Computational Intelligence and Data Mining (CIDM), IEEE, 2015).
# Данный датасет является примером несбалансированных данных, так как мошеннические операции с картами встречаются реже обычных.
# Импортруйте библиотеку Pandas, а также используйте для графиков стиль “fivethirtyeight”.
# Посчитайте с помощью метода value_counts количество наблюдений для каждого значения целевой переменной Class и примените к полученным данным метод plot, чтобы построить столбчатую диаграмму. Затем постройте такую же диаграмму, используя логарифмический масштаб.
# На следующем графике постройте две гистограммы по значениям признака V1 - одну для мошеннических транзакций (Class равен 1) и другую - для обычных (Class равен 0). Подберите значение аргумента density так, чтобы по вертикали графика было расположено не число наблюдений, а плотность распределения. Число бинов должно равняться 20 для обеих гистограмм, а коэффициент alpha сделайте равным 0.5, чтобы гистограммы были полупрозрачными и не загораживали друг друга. Создайте легенду с двумя значениями: “Class 0” и “Class 1”. Гистограмма обычных транзакций должна быть серого цвета, а мошеннических - красного. Горизонтальной оси дайте название “Class”.
import pandas as pd
plt.style.use('fivethirtyeight')
creditcard = pd.read_csv('creditcard.csv')
class_list = creditcard['Class'].value_counts()
print(class_list)
class_list.plot(kind='barh')
plt.show()
class_list.plot(kind='barh', logx=True)
plt.show()
class0 = creditcard.loc[creditcard['Class'] == 0, ['V1']]
class1 = creditcard.loc[creditcard['Class'] == 1, ['V1']]
plt.hist(class0['V1'], bins=20, density=True, alpha=0.5, label='Class 0', color='grey')
plt.hist(class1['V1'], bins=20, density=True, alpha=0.5, label='Class 1', color='red')
plt.legend()
plt.show()
| Lesson04/Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyluos
from pyluos import Device
device = Device ('/dev/cu.usbmodem142302')
print(device.nodes)
print(device.services)
print(device.lcd.parameters.blink)
device.servo.rot_position = 100
print(device.services)
print(device.servo.rot_position)
device.servo.target_rot_position = 0
print(device.servo.rot_position)
print
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### LDA Tutorial
#
# Taken from https://www.machinelearningplus.com/nlp/topic-modeling-python-sklearn-examples/
#
# +
import numpy as np
import pandas as pd
import re, nltk, spacy, gensim
from sklearn.decomposition import LatentDirichletAllocation,TruncatedSVD
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.model_selection import GridSearchCV
from pprint import pprint
import pyLDAvis
import pyLDAvis.sklearn
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Import Dataset 20-Newsgroups Dataset
df = pd.read_json('https://raw.githubusercontent.com/selva86/datasets/master/newsgroups.json')
print(df.target_names.unique())
df.head()
# +
##remove emails and new line characters
data = df.content.values.tolist()
# Remove Emails
data = [re.sub('[\w.-]+@[\w.-]+.\w+', '', sent) for sent in data]
# Remove new line characters
data = [re.sub('\n+', ' ', sent) for sent in data]
# Remove distracting single quotes
data = [re.sub("\'", "", sent) for sent in data]
pprint(data[:1])
# -
# +
## deacc remove punctuations
def sen_to_word(sentences):
for sentence in sentences:
yield(gensim.utils.simple_preprocess(str(sentence), deacc=True))
data_words = list(sen_to_word(data))
print(data_words[:1])
# -
def lemmatization(texts, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV']):
"""https://spacy.io/api/annotation"""
texts_out = []
for sent in texts:
doc = nlp(" ".join(sent))
texts_out.append(" ".join([token.lemma_ if token.lemma_ not in ['-PRON-'] else '' for token in doc if token.pos_ in allowed_postags]))
return texts_out
# +
# Initialize spacy 'en' model, keeping only tagger component (for efficiency)
# Run in terminal: python3 -m spacy download en
nlp = spacy.load('en', disable=['parser', 'ner'])
# Do lemmatization keeping only Noun, Adj, Verb, Adverb
data_lemmatized = lemmatization(data_words, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
print(data_lemmatized[:2])
# -
# +
vectorizer = CountVectorizer(analyzer='word',
min_df=10, # minimum reqd occurences of a word
stop_words='english', # remove stop words
lowercase=True, # convert all words to lowercase
token_pattern='[a-zA-Z0-9]{3,}', # num chars > 3
# max_features=50000, # max number of uniq words
)
data_vectorized = vectorizer.fit_transform(data_lemmatized)
# -
# +
# Materialize the sparse data
data_dense = data_vectorized.todense()
# Compute Sparsicity = Percentage of Non-Zero cells
print("Sparsicity: ", ((data_dense > 0).sum()/data_dense.size)*100, "%")
# -
# Build LDA Model
lda_model = LatentDirichletAllocation(n_components=20, # Number of components
max_iter=10, # Max learning iterations
learning_method='batch',
random_state=100, # Random state
batch_size=128, # n docs in each learning iter
evaluate_every = -1, # compute perplexity every n iters, default: Don't
n_jobs = -1, # Use all available CPUs
)
lda_output = lda_model.fit_transform(data_vectorized)
print(lda_model) # Model attributes
# +
# Log Likelyhood: Higher the better
print("Log Likelihood: ", lda_model.score(data_vectorized))
# Perplexity: Lower the better. Perplexity = exp(-1. * log-likelihood per word)
print("Perplexity: ", lda_model.perplexity(data_vectorized))
# See model parameters
pprint(lda_model.get_params())
# -
# ### Grid Search to Optimize Parameters
#
#
# The most important tuning parameter for LDA models is n_components (number of topics).
#
# Besides these, other possible search params could be learning_offset (downweigh early iterations. Should be > 1) and max_iter.
# +
# Define Search Param
#search_params = {'n_components': [10, 15, 20, 25, 30], 'learning_decay': [.5, .7, .9], 'max_iter': [5, 10, 15]}
search_params = {'n_components': [10, 15, 20], 'learning_decay': [.7, .9]}
# Init the Model
lda = LatentDirichletAllocation()
# Init Grid Search Class
model = GridSearchCV(lda, param_grid=search_params)
# Do the Grid Search
model.fit(data_vectorized)
# -
# +
# Best Model
best_lda_model = model.best_estimator_
# Model Parameters
print("Best Model's Params: ", model.best_params_)
# Log Likelihood Score
print("Best Log Likelihood Score: ", model.best_score_)
# Perplexity
print("Model Perplexity: ", best_lda_model.perplexity(data_vectorized))
# -
# +
# Get Log Likelyhoods from Grid Search Output
n_topics = [10, 15, 20]
log_likelyhoods_5 = [round(gscore.mean_validation_score) for gscore in model.grid_scores_ if gscore.parameters['learning_decay']==0.5]
log_likelyhoods_7 = [round(gscore.mean_validation_score) for gscore in model.grid_scores_ if gscore.parameters['learning_decay']==0.7]
log_likelyhoods_9 = [round(gscore.mean_validation_score) for gscore in model.grid_scores_ if gscore.parameters['learning_decay']==0.9]
# Show graph
plt.figure(figsize=(12, 8))
plt.plot(n_topics, log_likelyhoods_5, label='0.5')
plt.plot(n_topics, log_likelyhoods_7, label='0.7')
plt.plot(n_topics, log_likelyhoods_9, label='0.9')
plt.title("Choosing Optimal LDA Model")
plt.xlabel("Num Topics")
plt.ylabel("Log Likelyhood Scores")
plt.legend(title='Learning decay', loc='best')
plt.show()
# -
# +
# Create Document - Topic Matrix
lda_output = best_lda_model.transform(data_vectorized)
# column names
topicnames = ["Topic" + str(i) for i in range(best_lda_model.n_topics)]
# index names
docnames = ["Doc" + str(i) for i in range(len(data))]
# Make the pandas dataframe
df_document_topic = pd.DataFrame(np.round(lda_output, 2), columns=topicnames, index=docnames)
# Get dominant topic for each document
dominant_topic = np.argmax(df_document_topic.values, axis=1)
df_document_topic['dominant_topic'] = dominant_topic
# Styling
def color_green(val):
color = 'green' if val > .1 else 'black'
return 'color: {col}'.format(col=color)
def make_bold(val):
weight = 700 if val > .1 else 400
return 'font-weight: {weight}'.format(weight=weight)
# Apply Style
df_document_topics = df_document_topic.head(15).style.applymap(color_green).applymap(make_bold)
df_document_topics
# -
df_topic_distribution = df_document_topic['dominant_topic'].value_counts().reset_index(name="Num Documents")
df_topic_distribution.columns = ['Topic Num', 'Num Documents']
df_topic_distribution
pyLDAvis.enable_notebook()
panel = pyLDAvis.sklearn.prepare(best_lda_model, data_vectorized, vectorizer, mds='tsne')
panel
# +
# Topic-Keyword Matrix
df_topic_keywords = pd.DataFrame(best_lda_model.components_)
# Assign Column and Index
df_topic_keywords.columns = vectorizer.get_feature_names()
df_topic_keywords.index = topicnames
# View
df_topic_keywords.head()
# -
# +
# Show top n keywords for each topic
def show_topics(vectorizer=vectorizer, lda_model=lda_model, n_words=20):
keywords = np.array(vectorizer.get_feature_names())
topic_keywords = []
for topic_weights in lda_model.components_:
top_keyword_locs = (-topic_weights).argsort()[:n_words]
topic_keywords.append(keywords.take(top_keyword_locs))
return topic_keywords
topic_keywords = show_topics(vectorizer=vectorizer, lda_model=best_lda_model, n_words=15)
# Topic - Keywords Dataframe
df_topic_keywords = pd.DataFrame(topic_keywords)
df_topic_keywords.columns = ['Word '+str(i) for i in range(df_topic_keywords.shape[1])]
df_topic_keywords.index = ['Topic '+str(i) for i in range(df_topic_keywords.shape[0])]
df_topic_keywords
# -
# +
# Define function to predict topic for a given text document.
nlp = spacy.load('en', disable=['parser', 'ner'])
def predict_topic(text, nlp=nlp):
global sent_to_words
global lemmatization
# Step 1: Clean with simple_preprocess
mytext_2 = list(sent_to_words(text))
# Step 2: Lemmatize
mytext_3 = lemmatization(mytext_2, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV'])
# Step 3: Vectorize transform
mytext_4 = vectorizer.transform(mytext_3)
# Step 4: LDA Transform
topic_probability_scores = best_lda_model.transform(mytext_4)
topic = df_topic_keywords.iloc[np.argmax(topic_probability_scores), :].values.tolist()
return topic, topic_probability_scores
# Predict the topic
mytext = ["Some text about christianity and bible"]
topic, prob_scores = predict_topic(text = mytext)
print(topic)
# -
# +
# Construct the k-means clusters
from sklearn.cluster import KMeans
clusters = KMeans(n_clusters=15, random_state=100).fit_predict(lda_output)
# Build the Singular Value Decomposition(SVD) model
svd_model = TruncatedSVD(n_components=2) # 2 components
lda_output_svd = svd_model.fit_transform(lda_output)
# X and Y axes of the plot using SVD decomposition
x = lda_output_svd[:, 0]
y = lda_output_svd[:, 1]
# Weights for the 15 columns of lda_output, for each component
print("Component's weights: \n", np.round(svd_model.components_, 2))
# Percentage of total information in 'lda_output' explained by the two components
print("Perc of Variance Explained: \n", np.round(svd_model.explained_variance_ratio_, 2))
# -
# Plot
plt.figure(figsize=(12, 12))
plt.scatter(x, y, c=clusters)
plt.xlabel('Component 2')
plt.xlabel('Component 1')
plt.title("Segregation of Topic Clusters", )
# +
from sklearn.metrics.pairwise import euclidean_distances
nlp = spacy.load('en', disable=['parser', 'ner'])
def similar_documents(text, doc_topic_probs, documents = data, nlp=nlp, top_n=5, verbose=False):
topic, x = predict_topic(text)
dists = euclidean_distances(x.reshape(1, -1), doc_topic_probs)[0]
doc_ids = np.argsort(dists)[:top_n]
if verbose:
print("Topic KeyWords: ", topic)
print("Topic Prob Scores of text: ", np.round(x, 1))
print("Most Similar Doc's Probs: ", np.round(doc_topic_probs[doc_ids], 1))
return doc_ids, np.take(documents, doc_ids)
# -
# Get similar documents
mytext = ["Some text about christianity and bible"]
doc_ids, docs = similar_documents(text=mytext, doc_topic_probs=lda_output, documents = data, top_n=1, verbose=True)
print('\n', docs[0][:500])
| Machine_Learning/LDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="pXMdEU-qHQHh"
# This study guide should reinforce and provide practice for all of the concepts you have seen in the past week. There are a mix of written questions and coding exercises, both are equally important to prepare you for the sprint challenge as well as to be able to speak on these topics comfortably in interviews and on the job.
#
# If you get stuck or are unsure of something remember the 20 minute rule. If that doesn't help, then research a solution with google and stackoverflow. Only once you have exausted these methods should you turn to your Team Lead - they won't be there on your SC or during an interview. That being said, don't hesitate to ask for help if you truly are stuck.
#
# Have fun studying!
# + [markdown] colab_type="text" id="V-vPPhtthaNU"
# # Resources
# + [markdown] colab_type="text" id="PCvm7I44H3Oe"
# [Category Encoders](https://contrib.scikit-learn.org/categorical-encoding/)
#
# [Logistic Regression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html)
#
# [Decision Tree Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html)
#
# [Hyperparameter Tuning](https://scikit-learn.org/stable/modules/grid_search.html)
#
# [Confusion Matrix](https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html)
#
# [Scoring Metrics](https://scikit-learn.org/stable/modules/model_evaluation.html)
# + colab={} colab_type="code" id="b_7n8s1mak9k"
import pandas as pd
# + [markdown] colab_type="text" id="xL8fg3mIhd_L"
# Use the dataframe below for all questions unless otherwise stated
# + colab={"base_uri": "https://localhost:8080/", "height": 206} colab_type="code" id="3WWarojafXt5" outputId="be403850-88a6-465a-981d-43f0ebbaa2e1"
# https://www.kaggle.com/ronitf/heart-disease-uci
df = pd.read_csv('https://raw.githubusercontent.com/bundickm/Study-Guides/master/data/hearts.csv', index_col=0)
df.head()
# + [markdown] colab_type="text" id="eDMTLmTzhufS"
# # Basics and Data Preparation
# + [markdown] colab_type="text" id="x7LH-U2KiMfO"
# Define the following terms in your own words, do not simply copy and paste a definition found elsewhere but reword it to be understandable and memorable to you. *Double click the markdown to add your definitions.*
# <br/><br/>
#
# **Logistic Regression:** `A statistical model using a logistic function to classify a binary dependent variable`
#
# **Imbalanced Classes:** ` A disproportionate ratio of observations in each class in a dataset`
#
# **Leakage:** `Unrealistic improvements in model performance by poor data management between training and test subsets`
#
# **Categorical Encoding:** `Transforming categorical data into numerical format through techniques such as One Hot and Ordinal Encoding`
#
# **Skew:** `A measure of asymmetry from the shape of a normal distribution with respect to data of interest`
#
# **Log Transformation:** `A mathematical transformation applied to skewed dataset to conform to a normal distribution`
#
# **Outliers:** `Observations in a dataset which significantly deviate from most other observations`
# + [markdown] colab_type="text" id="fCIXoJtDnXDZ"
# Answer the following questions in plain english as much as possible.
# <br/><br/>
#
# What are some ways to deal with imbalanced classes?
# ```
# Your Answer Here
# ```
#
# What are some possible sources of data leakage?
# ```
# Your Answer Here
# ```
#
# What are some indicators or methods for detecting data leakage?
# ```
# Your Answer Here
# ```
#
# What is the relationship between skew and log transformation?
# ```
# Your Answer Here
# ```
# + [markdown] colab_type="text" id="q2M6tTW02sJe"
# Using the dataset above, complete the following:
# - Train/Test/Validation Split
# - Get a baseline
# - Perform EDA with visuals
# - Clean up any nulls, duplicate columns, or outliers you might find
# - Engineer at least 2 features
# - Use One Hot or Ordinal Encoding on one feature
# -
# ### Train/Test/Validation Split
# + colab={} colab_type="code" id="XZlA2iue6LFN"
from sklearn.model_selection import train_test_split
# +
train, test = train_test_split(df, train_size=0.8, test_size=0.2, random_state=8)
train, validate = train_test_split(train, train_size=0.8, test_size=0.2, random_state=8)
train.shape, validate.shape, test.shape
# -
# ### Get a baseline
train.target.value_counts(normalize=True)*100
# ### Perform EDA with visuals
train.head()
import seaborn as sns
import matplotlib.pyplot as plt
sns.jointplot(x="age",y="trestbps", data=train)
plt.show()
sns.jointplot(x="trestbps",y="thalach", data=train)
plt.show()
sns.jointplot(x="trestbps",y="oldpeak", data=train)
plt.show()
sns.jointplot(x="chol",y="trestbps", data=train)
plt.show()
train.groupby("target")["chol"].mean().plot.bar()
train.groupby("target")["trestbps"].mean().plot.bar()
train.groupby("target")["thalach"].mean().plot.bar()
# ### Clean up any nulls, duplicate columns, or outliers you might find
train.isna().sum()
train["age"].value_counts(dropna=False).sort_index().plot.bar(width=1)
train["sex"].value_counts(dropna=False, normalize=True).plot(kind="bar")
train.describe(include="all")
# ### Engineer at least 2 features
# +
train["cholesterol years"] = (train["chol"] / train["chol"].mean()) + (train["age"] / train["age"].mean())
train.head()
# -
train.groupby("target")["cholesterol years"].mean()
train["max heart rate years"] = (train["thalach"] / train["thalach"].mean()) + (train["age"] / train["age"].mean())
train
train.groupby("target")["max heart rate years"].mean()
train["number of major vessel years"] = train["ca"] * (train["age"] / train["age"].mean())
train.groupby("target")["number of major vessel years"].mean()
# +
validate["cholesterol years"] = ((validate["chol"].copy() / validate["chol"].mean()) + (validate["age"].copy() / validate["age"].mean()))
validate["max heart rate years"] = ((validate["thalach"].copy() / validate["thalach"].mean()) + (validate["age"] .copy()/ validate["age"].mean()))
validate["number of major vessel years"] = validate["ca"] * (validate["age"] / validate["age"].mean())
test["cholesterol years"] = ((test["chol"].copy() / test["chol"].mean()) + (test["age"].copy() / test["age"].mean()))
test["max heart rate years"] = ((test["thalach"].copy() / test["thalach"].mean()) + (test["age"].copy() / test["age"].mean())).copy()
test["number of major vessel years"] = test["ca"] * (test["age"] / test["age"].mean())
# -
train.dtypes
# ### Use One Hot or Ordinal Encoding on one feature
# +
# The status_group column is the target
target = 'target'
# Get a dataframe with all train columns except the target
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Get a series with the cardinality of the nonnumeric features
cardinality = train_features.select_dtypes(exclude='number').nunique()
# Get a list of all categorical features with cardinality <= 50
categorical_features = cardinality[cardinality<=50].index.tolist()
# Combine the lists
features = numeric_features + categorical_features
# -
X_train = train[features]
y_train = train[target]
X_validate = validate[features]
y_validate = validate[target]
X_test = test[features]
y_test = test[target]
# +
# I will include Ordinal Encoding in my prediction pipeline
# + [markdown] colab_type="text" id="14JotyiJ8RjD"
# # Model Building
# + [markdown] colab_type="text" id="dhEv3JQ78TdH"
# Define the following terms in your own words, do not simply copy and paste a definition found elsewhere but reword it to be understandable and memorable to you. *Double click the markdown to add your definitions.*
# <br/><br/>
#
# **Decision Tree:** `Your Answer Here`
#
# **Ensemble Methods (Ensemble Models):** `Your Answer Here`
#
# **Gradient Descent:** `Your Answer Here`
#
# **Bagging:** `Your Answer Here`
#
# **Boosting:** `Your Answer Here`
#
# **Hyperparameters:** `Your Answer Here`
# + [markdown] colab_type="text" id="Bl8D-WwF8W8l"
# Build a random forest classifier using the dataset you cleaned and prepped above.
# -
import category_encoders as ce
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
# + colab={} colab_type="code" id="T1dTbiC78g7r"
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
StandardScaler(),
RandomForestClassifier(n_jobs=-1, min_samples_leaf=2, random_state=8)
)
pipeline.fit(X_train, y_train)
print ('Train Accuracy', pipeline.score(X_train, y_train))
print ('Validation Accuracy', pipeline.score(X_validate, y_validate))
# + [markdown] colab_type="text" id="oE5S3glR-wNp"
# Graph your model's feature importances
# + colab={} colab_type="code" id="Jptjbm16-3SY"
model = pipeline.named_steps["randomforestclassifier"]
encoder = pipeline.named_steps["ordinalencoder"]
encoded_columns = encoder.transform(X_validate).columns
importances = pd.Series(model.feature_importances_, encoded_columns)
plt.figure(figsize=(10,10))
importances.sort_values().plot(kind="barh", color="red")
plt.title("Random Forest Classifier Feature Importances")
plt.show()
# + [markdown] colab_type="text" id="nu9wLKlg-3q3"
# In 2-3 sentences, explain how to interpret and use the feature importances to further refine or help explain your model.
#
# ```
# Your Answer Here
# ```
#
# How does feature importance differ from drop-column importances and permutation importances?
#
# ```
# Your Answer Here
# ```
# + [markdown] colab_type="text" id="df3-NCv18hUC"
# Build a logisitic regression model using the dataset you cleaned and prepped above.
# + colab={} colab_type="code" id="N2Kzqo358qbc"
from sklearn.linear_model import LogisticRegression
# +
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(strategy='mean'),
StandardScaler(),
LogisticRegression(solver='lbfgs', n_jobs=-1, random_state=8)
)
pipeline.fit(X_train, y_train)
print ('Train Accuracy', pipeline.score(X_train, y_train))
print ('Validation Accuracy', pipeline.score(X_validate, y_validate))
# + [markdown] colab_type="text" id="inxDK4qx_WM7"
# Plot the coefficients of your model.
# +
model = pipeline.named_steps["logisticregression"]
coefficients = pd.Series(model.coef_[0], X_train.columns)
coefficients.sort_values().plot.barh(color="#A0522D");
plt.title("Logistic Regression Coefficients", size="x-large")
plt.show()
# + [markdown] colab_type="text" id="Kz_obfF9_kaF"
# In 2-3 sentences, explain how to interpret and use the coefficients to further refine or help explain your model.
#
# ```
# Your Answer Here
# ```
# + [markdown] colab_type="text" id="lZXahsuH_yUl"
# What is an example of an ensemble method?
#
# ```
# Your Answer Here
# ```
#
# What do we mean by hyperparameter tuning and how can we automate the tuning process?
#
# ```
# Your Answer Here
# ```
# + [markdown] colab_type="text" id="UfkSVMIS_7dg"
# # Metrics and Model Evaluation
# + [markdown] colab_type="text" id="Vb659NzBAQAY"
# Define the following terms in your own words, do not simply copy and paste a definition found elsewhere but reword it to be understandable and memorable to you. *Double click the markdown to add your definitions.*
# <br/><br/>
#
# **ROC:** `Your Answer Here`
#
# **ROC-AUC:** `Your Answer Here`
#
# **Discrimination Threshold:** `Your Answer Here`
#
# **Precision:** `Your Answer Here`
#
# **Recall:** `Your Answer Here`
#
# **F1 Score:** `Your Answer Here`
#
# **Confusion Matrix:** `Your Answer Here`
# + [markdown] colab_type="text" id="0F8klqRlCj1k"
# Choose one of your models above to complete the following:
# - Get your model's validation accuracy (This may be done multiple times if you are refining your model)
# - Get your model's test accuracy
# - Create a confusion matrix for your model
# - Calculate the Accuracy, F1 Score, Precision, and Recall by hand
# - Use SKLearn to calculate accuracy, F1 score, precision, and recall to confirm your work.
# -
# ### Get your model's validation accuracy (This may be done multiple times if you are refining your model)
# + colab={} colab_type="code" id="MLFsf9me_49e"
print ('Validation Accuracy', pipeline.score(X_validate, y_validate))
# -
# ### Get your model's test accuracy
print ('Testing Accuracy', pipeline.score(X_test, y_test))
# ### Create a confusion matrix for your model
# +
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(pipeline, X_test, y_test, values_format='.0f', xticks_rotation="vertical", cmap="cividis")
plt.show()
# -
# ### Calculate the Accuracy, F1 Score, Precision, and Recall by hand
correct_predictions = 19 + 29
correct_predictions
total_predictions = 19 + 3 + 10 + 29
total_predictions
classification_accuracy = correct_predictions / total_predictions
classification_accuracy
# +
correct_predictions_0 = 19
correct_predictions_1 = 29
total_predictions_0 = 19 + 10
total_preditions_1 = 3 + 29
# -
precision_0 = correct_predictions_0 / total_predictions_0
precision_0
# ### Use SKLearn to calculate accuracy, F1 score, precision, and recall to confirm your work.
from sklearn.metrics import classification_report
y_pred = pipeline.predict(X_validate)
print(classification_report(y_validate, y_pred))
# + [markdown] colab_type="text" id="elfpNZo-Fxe-"
# Give an example of when we would use precision to score our model and explain why precision is the best metric for that situation.
#
# ```
# Your Answer Here
# ```
#
# Give an example of when we would use recall to score our model and explain why recall is the best metric for that situation.
#
# ```
# Your Answer Here
# ```
# + [markdown] colab_type="text" id="Bdhtv_4KJrgQ"
# Find your model's ROC-AUC Score
# + colab={} colab_type="code" id="uFR91njwJv8b"
y_pred_proba = pipeline.predict_proba(X_validate)[:,1]
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_validate, y_pred_proba)
# -
from sklearn.metrics import roc_auc_score
roc_auc_score(y_validate, y_pred_proba)
# + [markdown] colab_type="text" id="DYOLWhoUJfQU"
# Plot your model's ROC Curve
# + colab={} colab_type="code" id="VI45S71RJkRy"
plt.scatter(fpr, tpr)
plt.title('ROC curve')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate');
# -
| module4-classification-metrics/Unit_2_Sprint_2_Tree_Ensembles_Study_Guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''pytorch'': conda)'
# name: python37664bitpytorchconda0cdad03962454fdfb22b6d3ea1ad8fae
# ---
import torch
# require a place to store the gradient of y with respect to x.
x = torch.arange(4.0, requires_grad=True) # it tells the framework we need allocate gradient space for x in the future.
x
x.grad
y = 2 * torch.dot(x, x)
y
# 1 + 2**2 + 3**2 = 14,then 14*2 = 28
y.backward()
x.grad
# +
x.grad.zero_()
y = x * x
u = y.detach()
z = u * x
z.sum().backward()
x.grad == u
# -
x.grad.zero_()
y.sum().backward()
x.grad == 2 * x
def f(a):
b = a * 2
while b.norm().item() < 1000:#return the number that is in tensor()'s brackets.
b = b * 2
if b.sum().item() > 0:
c = b
else:
c = 100 * b
return c
a.grad == (d / a)
a = torch.randn(size=(1,), requires_grad=True)
# record the calculation
d = f(a)
d.backward()
a.grad == (d / a)
b = torch.randn(size=(1,), requires_grad=True)
b = b + 1000
# record the calculation
d = f(b)
d.backward()
b.grad == (d / b)
#
b = torch.randn(size=(1,), requires_grad=True)
b = b + 1000
# record the calculation
d = f(b)
d.backward()
d
b
t = b.grad
t
b.grad == (d / b)
| Ch02_Preliminaries/2-5-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1D Kalman Filter
#
# Now, you're ready to implement a 1D Kalman Filter by putting all these steps together. Let's take the case of a robot that moves through the world. As a robot moves through the world it locates itself by performing a cycle of:
# 1. sensing and performing a measurement update and
# 2. moving and performing a motion update
#
# You've programmed each of these steps individually, so now let's combine them in a cycle!
#
# After implementing this filter, you should see that you can go from a very uncertain location Gaussian to a more and more certain Gaussian, as pictured below. The code in this notebooks is really just a simplified version of the Kalman filter that runs in the Google self-driving car that is used to track surrounding vehicles and other objects.
#
# <img src='images/gaussian_updates.png' height=70% width=70% />
#
# +
# import math functions
from math import *
import matplotlib.pyplot as plt
import numpy as np
# gaussian function
def f(mu, sigma2, x):
''' f takes in a mean and squared variance, and an input x
and returns the gaussian value.'''
coefficient = 1.0 / sqrt(2.0 * pi *sigma2)
exponential = exp(-0.5 * (x-mu) ** 2 / sigma2)
return coefficient * exponential
# +
# the update function
def update(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters.'''
# Calculate the new parameters
new_mean = (var2*mean1 + var1*mean2)/(var2+var1)
new_var = 1/(1/var2 + 1/var1)
return [new_mean, new_var]
# the motion update/predict function
def predict(mean1, var1, mean2, var2):
''' This function takes in two means and two squared variance terms,
and returns updated gaussian parameters, after motion.'''
# Calculate the new parameters
new_mean = mean1 + mean2
new_var = var1 + var2
return [new_mean, new_var]
# -
# ### QUIZ: For the given measurements and motions, write complete 1D Kalman filter code that loops through all of these in order.
#
# Your complete code should look at sensor measurements then motions in that sequence until all updates are done!
#
# ### Initial Uncertainty
#
# You'll see that you are given initial parameters below, and this includes and nitial location estimation, `mu` and squared variance, `sig`. Note that the initial estimate is set to the location 0, and the variance is extremely large; this is a state of high confusion much like the *uniform* distribution we used in the histogram filter. There are also values given for the squared variance associated with the sensor measurements and the motion, since neither of those readings are perfect, either.
#
# You should see that even though the initial estimate for location (the initial `mu`) is far from the first measurement, it should catch up fairly quickly as you cycle through measurements and motions.
# +
# measurements for mu and motions, U
measurements = [5., 6., 7., 9., 10.]
motions = [1., 1., 2., 1., 1.]
# initial parameters
measurement_sig = 4.
motion_sig = 2.
mu = 0.
sig = 10000.
## TODO: Loop through all measurements/motions
# this code assumes measurements and motions have the same length
# so their updates can be performed in pairs
for n in range(len(measurements)):
# measurement update, with uncertainty
mu, sig = update(mu, sig, measurements[n], measurement_sig)
print('Update: [{}, {}]'.format(mu, sig))
# motion update, with uncertainty
mu, sig = predict(mu, sig, motions[n], motion_sig)
print('Predict: [{}, {}]'.format(mu, sig))
# print the final, resultant mu, sig
print('\n')
print('Final result: [{}, {}]'.format(mu, sig))
# -
| 3_Motion_SLAM/1D Kalman Filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="u3Zq5VrfiDqB"
# ##### Copyright 2021 The TensorFlow Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="form" id="3jTEqPzFiHQ0"
#@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="x97n3SaNmNpB"
# # Variational Inference on Probabilistic Graphical Models with Joint Distributions
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/probability/examples/Variational_Inference_and_Joint_Distributions"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/probability/blob/main/tensorflow_probability/examples/jupyter_notebooks/Variational_Inference_and_Joint_Distributions.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/probability/blob/main/tensorflow_probability/examples/jupyter_notebooks/Variational_Inference_and_Joint_Distributions.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/probability/examples/jupyter_notebooks/Variational_Inference_and_Joint_Distributions.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="SVcOch4u2bVS"
# Variational Inference (VI) casts approximate Bayesian inference as an optimization problem and seeks a 'surrogate' posterior distribution that minimizes the KL divergence with the true posterior. Gradient-based VI is often faster than MCMC methods, composes naturally with optimization of model parameters, and provides a lower bound on model evidence that can be used directly for model comparison, convergence diagnosis, and composable inference.
#
# TensorFlow Probability offers tools for fast, flexible, and scalable VI that fit naturally into the TFP stack. These tools enable the construction of surrogate posteriors with covariance structures induced by linear transformations or normalizing flows.
#
# VI can be used to estimate Bayesian [credible intervals](https://en.wikipedia.org/wiki/Credible_interval) for parameters of a regression model to estimate the effects of various treatments or observed features on an outcome of interest. Credible intervals bound the values of an unobserved parameter with a certain probability, according to the posterior distribution of the parameter conditioned on observed data and given an assumption on the parameter's prior distribution.
#
# In this Colab, we demonstrate how to use VI to obtain credible intervals for parameters of a Bayesian linear regression model for radon levels measured in homes (using [Gelman et al.'s (2007) Radon dataset](http://www.stat.columbia.edu/~gelman/arm/); see [similar examples](https://mc-stan.org/users/documentation/case-studies/radon.html#Correlations-among-levels) in Stan). We demonstrate how TFP `JointDistribution`s combine with `bijectors` to build and fit two types of expressive surrogate posteriors:
#
# - a standard Normal distribution transformed by a block matrix. The matrix may reflect independence among some components of the posterior and dependence among others, relaxing the assumption of a mean-field or full-covariance posterior.
# - a more complex, higher-capacity [inverse autoregressive flow](https://arxiv.org/abs/1606.04934).
#
# The surrogate posteriors are trained and compared with results from a mean-field surrogate posterior baseline, as well as ground-truth samples from Hamiltonian Monte Carlo.
# + [markdown] id="pt5Lzw4hjd6A"
# ## Overview of Bayesian Variational Inference
#
# Suppose we have the following generative process, where $\theta$ represents random parameters, $\omega$ represents deterministic parameters, and the $x_i$ are features and the $y_i$ are target values for $i=1,\ldots,n$ observed data points:
# \begin{align*}
# &\theta \sim r(\Theta) && \text{(Prior)}\\
# &\text{for } i = 1 \ldots n: \nonumber \\
# &\quad y_i \sim p(Y_i|x_i, \theta, \omega) && \text{(Likelihood)}
# \end{align*}
#
# VI is then characterized by:
# $\newcommand{\E}{\operatorname{\mathbb{E}}}
# \newcommand{\K}{\operatorname{\mathbb{K}}}
# \newcommand{\defeq}{\overset{\tiny\text{def}}{=}}
# \DeclareMathOperator*{\argmin}{arg\,min}$
#
# \begin{align*}
# -\log p(\{y_i\}_i^n|\{x_i\}_i^n, \omega)
# &\defeq -\log \int \textrm{d}\theta\, r(\theta) \prod_i^n p(y_i|x_i,\theta, \omega) && \text{(Really hard integral)} \\
# &= -\log \int \textrm{d}\theta\, q(\theta) \frac{1}{q(\theta)} r(\theta) \prod_i^n p(y_i|x_i,\theta, \omega) && \text{(Multiply by 1)}\\
# &\le - \int \textrm{d}\theta\, q(\theta) \log \frac{r(\theta) \prod_i^n p(y_i|x_i,\theta, \omega)}{q(\theta)} && \text{(Jensen's inequality)}\\
# &\defeq \E_{q(\Theta)}[ -\log p(y_i|x_i,\Theta, \omega) ] + \K[q(\Theta), r(\Theta)]\\
# &\defeq ``\text{expected negative log likelihood"} + ``\text{kl regularizer"}
# \end{align*}
#
# (Technically we're assuming $q$ is [absolutely continuous](https://en.wikipedia.org/wiki/Absolute_continuity#Absolute_continuity_of_measures) with respect to $r$. See also, [Jensen's inequality](https://en.wikipedia.org/wiki/Jensen%27s_inequality).)
#
# Since the bound holds for all q, it is obviously tightest for:
#
# $$q^*,w^* = \argmin_{q \in \mathcal{Q},\omega\in\mathbb{R}^d} \left\{ \sum_i^n\E_{q(\Theta)}\left[ -\log p(y_i|x_i,\Theta, \omega) \right] + \K[q(\Theta), r(\Theta)] \right\}$$
#
#
#
#
# Regarding terminology, we call
#
# - $q^*$ the "surrogate posterior," and,
# - $\mathcal{Q}$ the "surrogate family."
#
# $\omega^*$ represents the maximum-likelihood values of the deterministic parameters on the VI loss. See [this survey](https://arxiv.org/abs/1601.00670) for more information on variational inference.
# + [markdown] id="pt532xMzBJiR"
# ## Example: Bayesian hierarchical linear regression on Radon measurements
#
# Radon is a radioactive gas that enters homes through contact points with the
# ground. It is a carcinogen that is the primary cause of lung cancer in
# non-smokers. Radon levels vary greatly from household to household.
#
# The EPA did a study of radon levels in 80,000 houses. Two important predictors
# are:
# - Floor on which the measurement was taken (radon higher in basements)
# - County uranium level (positive correlation with radon levels)
#
# Predicting radon levels in houses grouped by county is a classic problem in Bayesian hierarchical modeling, introduced by [Gelman and Hill (2006)](http://www.stat.columbia.edu/~gelman/arm/). We will build a hierarchical linear model to predict radon measurements in houses, in which the hierarchy is the grouping of houses by county. We are interested in credible intervals for the effect of location (county) on the radon level of houses in Minnesota. In order to isolate this effect, the effects of floor and uranium level are also included in the model. Additionaly, we will incorporate a contextual effect corresponding to the mean floor on which the measurement was taken, by county, so that if there is variation among counties of the floor on which the measurements were taken, this is not attributed to the county effect.
# + id="i00BTGk5tiwe"
# !pip3 install -q tf-nightly tfp-nightly
# + id="H9omoz32_Y9F"
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_probability as tfp
import warnings
tfd = tfp.distributions
tfb = tfp.bijectors
plt.rcParams['figure.facecolor'] = '1.'
# + id="BFKYEEfY1FhB"
# Load the Radon dataset from `tensorflow_datasets` and filter to data from
# Minnesota.
dataset = tfds.as_numpy(
tfds.load('radon', split='train').filter(
lambda x: x['features']['state'] == 'MN').batch(10**9))
# Dependent variable: Radon measurements by house.
dataset = next(iter(dataset))
radon_measurement = dataset['activity'].astype(np.float32)
radon_measurement[radon_measurement <= 0.] = 0.1
log_radon = np.log(radon_measurement)
# Measured uranium concentrations in surrounding soil.
uranium_measurement = dataset['features']['Uppm'].astype(np.float32)
log_uranium = np.log(uranium_measurement)
# County indicator.
county_strings = dataset['features']['county'].astype('U13')
unique_counties, county = np.unique(county_strings, return_inverse=True)
county = county.astype(np.int32)
num_counties = unique_counties.size
# Floor on which the measurement was taken.
floor_of_house = dataset['features']['floor'].astype(np.int32)
# Average floor by county (contextual effect).
county_mean_floor = []
for i in range(num_counties):
county_mean_floor.append(floor_of_house[county == i].mean())
county_mean_floor = np.array(county_mean_floor, dtype=log_radon.dtype)
floor_by_county = county_mean_floor[county]
# + [markdown] id="EU9ieWyOjddQ"
# The regression model is specified as follows:
#
# $\newcommand{\Normal}{\operatorname{\sf Normal}}$
# \begin{align*}
# &\text{uranium_weight} \sim \Normal(0, 1) \\
# &\text{county_floor_weight} \sim \Normal(0, 1) \\
# &\text{for } j = 1\ldots \text{num_counties}:\\
# &\quad \text{county_effect}_j \sim \Normal (0, \sigma_c)\\
# &\text{for } i = 1\ldots n:\\
# &\quad \mu_i = ( \\
# &\quad\quad \text{bias} \\
# &\quad\quad + \text{county_effect}_{\text{county}_i} \\
# &\quad\quad +\text{log_uranium}_i \times \text{uranium_weight} \\
# &\quad\quad +\text{floor_of_house}_i \times \text{floor_weight} \\
# &\quad\quad +\text{floor_by_county}_{\text{county}_i} \times \text{county_floor_weight} ) \\
# &\quad \text{log_radon}_i \sim \Normal(\mu_i, \sigma_y)
# \end{align*}
# in which $i$ indexes the observations and $\text{county}_i$ is the county in which the $i$th observation was taken.
#
# We use a county-level random effect to capture geographical variation. The parameters `uranium_weight` and `county_floor_weight` are modeled probabilistically, and `floor_weight` and the constant `bias` are deterministic. These modeling choices are largely arbitrary, and are made for the purpose of demonstrating VI on a probabilistic model of reasonable complexity. For a more thorough discussion of multilevel modeling with fixed and random effects in TFP, using the radon dataset, see [Multilevel Modeling Primer](https://github.com/tensorflow/probability/blob/main/tensorflow_probability/examples/jupyter_notebooks/Multilevel_Modeling_Primer.ipynb) and [Fitting Generalized Linear Mixed-effects Models Using Variational Inference](https://github.com/tensorflow/probability/blob/main/tensorflow_probability/examples/jupyter_notebooks/Linear_Mixed_Effects_Model_Variational_Inference.ipynb).
# + id="awL6fCUh6OCF"
# Create variables for fixed effects.
floor_weight = tf.Variable(0.)
bias = tf.Variable(0.)
# Variables for scale parameters.
log_radon_scale = tfp.util.TransformedVariable(1., tfb.Exp())
county_effect_scale = tfp.util.TransformedVariable(1., tfb.Exp())
# Define the probabilistic graphical model as a JointDistribution.
@tfd.JointDistributionCoroutineAutoBatched
def model():
uranium_weight = yield tfd.Normal(0., scale=1., name='uranium_weight')
county_floor_weight = yield tfd.Normal(
0., scale=1., name='county_floor_weight')
county_effect = yield tfd.Sample(
tfd.Normal(0., scale=county_effect_scale),
sample_shape=[num_counties], name='county_effect')
yield tfd.Normal(
loc=(log_uranium * uranium_weight + floor_of_house* floor_weight
+ floor_by_county * county_floor_weight
+ tf.gather(county_effect, county, axis=-1)
+ bias),
scale=log_radon_scale[..., tf.newaxis],
name='log_radon')
# Pin the observed `log_radon` values to model the un-normalized posterior.
target_model = model.experimental_pin(log_radon=log_radon)
# + [markdown] id="UlkQTJSlkjJ1"
# ## Expressive surrogate posteriors
#
# Next we estimate the posterior distributions of the random effects using VI with two different types of surrogate posteriors:
# - A constrained multivariate Normal distribution, with covariance structure induced by a blockwise matrix transformation.
# - A multivariate Standard Normal distribution transformed by an [Inverse Autoregressive Flow](https://arxiv.org/abs/1606.04934), which is then split and restructured to match the support of the posterior.
# + [markdown] id="3QG0scmDcdTw"
# ### Multivariate Normal surrogate posterior
# + [markdown] id="K8soBr2oBHSV"
# To build this surrogate posterior, a trainable linear operator is used to induce correlation among the components of the posterior.
# + id="sJuvC5ykBAiK"
# Determine the `event_shape` of the posterior, and calculate the size of each
# `event_shape` component. These determine the sizes of the components of the
# underlying standard Normal distribution, and the dimensions of the blocks in
# the blockwise matrix transformation.
event_shape = target_model.event_shape_tensor()
flat_event_shape = tf.nest.flatten(event_shape)
flat_event_size = tf.nest.map_structure(tf.reduce_prod, flat_event_shape)
# The `event_space_bijector` maps unconstrained values (in R^n) to the support
# of the prior -- we'll need this at the end to constrain Multivariate Normal
# samples to the prior's support.
event_space_bijector = target_model.experimental_default_event_space_bijector()
# + [markdown] id="LxLqBKBgsQPg"
# Construct a `JointDistribution` with vector-valued standard Normal components, with sizes determined by the corresponding prior components. The components should be vector-valued so they can be transformed by the linear operator.
# + id="0ceaCfU8sPjg"
base_standard_dist = tfd.JointDistributionSequential(
[tfd.Sample(tfd.Normal(0., 1.), s) for s in flat_event_size])
# + [markdown] id="uu0d8uWS4luv"
# Build a trainable blockwise lower-triangular linear operator. We'll apply it to the standard Normal distribution to implement a (trainable) blockwise matrix transformation and induce the correlation structure of the posterior.
#
# Within the blockwise linear operator, a trainable full-matrix block represents full covariance between two components of the posterior, while a block of zeros (or `None`) expresses independence. Blocks on the diagonal are either lower-triangular or diagonal matrices, so that the entire block structure represents a lower-triangular matrix.
#
# Applying this bijector to the base distribution results in a multivariate Normal distribution with mean 0 and (Cholesky-factored) covariance equal to the lower-triangular block matrix.
# + id="dUCks9qg6nU2"
operators = (
(tf.linalg.LinearOperatorDiag,), # Variance of uranium weight (scalar).
(tf.linalg.LinearOperatorFullMatrix, # Covariance between uranium and floor-by-county weights.
tf.linalg.LinearOperatorDiag), # Variance of floor-by-county weight (scalar).
(None, # Independence between uranium weight and county effects.
None, # Independence between floor-by-county and county effects.
tf.linalg.LinearOperatorDiag) # Independence among the 85 county effects.
)
block_tril_linop = (
tfp.experimental.vi.util.build_trainable_linear_operator_block(
operators, flat_event_size))
scale_bijector = tfb.ScaleMatvecLinearOperatorBlock(block_tril_linop)
# + [markdown] id="dHI0bziq44od"
# After applying the linear operator to the standard Normal distribution, apply a multipart `Shift` bijector to allow the mean to take nonzero values.
# + id="ceS386lN448r"
loc_bijector = tfb.JointMap(
tf.nest.map_structure(
lambda s: tfb.Shift(
tf.Variable(tf.random.uniform(
(s,), minval=-2., maxval=2., dtype=tf.float32))),
flat_event_size))
# + [markdown] id="gLO_8C0_Hd7f"
# The resulting multivariate Normal distribution, obtained by transforming the standard Normal distribution with the scale and location bijectors, must be reshaped and restructured to match the prior, and finally constrained to the support of the prior.
# + id="PnnU3lJ7H-pj"
# Reshape each component to match the prior, using a nested structure of
# `Reshape` bijectors wrapped in `JointMap` to form a multipart bijector.
reshape_bijector = tfb.JointMap(
tf.nest.map_structure(tfb.Reshape, flat_event_shape))
# Restructure the flat list of components to match the prior's structure
unflatten_bijector = tfb.Restructure(
tf.nest.pack_sequence_as(
event_shape, range(len(flat_event_shape))))
# + [markdown] id="HK3n0iqc5Ei3"
# Now, put it all together -- chain the trainable bijectors together and apply them to the base standard Normal distribution to construct the surrogate posterior.
# + id="xlrIbELO5EWR"
surrogate_posterior = tfd.TransformedDistribution(
base_standard_dist,
bijector = tfb.Chain( # Note that the chained bijectors are applied in reverse order
[
event_space_bijector, # constrain the surrogate to the support of the prior
unflatten_bijector, # pack the reshaped components into the `event_shape` structure of the posterior
reshape_bijector, # reshape the vector-valued components to match the shapes of the posterior components
loc_bijector, # allow for nonzero mean
scale_bijector # apply the block matrix transformation to the standard Normal distribution
]))
# + [markdown] id="bVmf3qld5oPP"
# Train the multivariate Normal surrogate posterior.
# + id="J5c5mhh-F9l-"
optimizer = tf.optimizers.Adam(learning_rate=1e-2)
@tf.function(jit_compile=True)
def fit_vi():
return tfp.vi.fit_surrogate_posterior(
target_model.unnormalized_log_prob,
surrogate_posterior,
optimizer=optimizer,
num_steps=10**4,
sample_size=16,
)
mvn_loss = fit_vi()
mvn_samples = surrogate_posterior.sample(1000)
mvn_final_elbo = tf.reduce_mean(
target_model.unnormalized_log_prob(*mvn_samples)
- surrogate_posterior.log_prob(mvn_samples))
print('Multivariate Normal surrogate posterior ELBO: {}'.format(mvn_final_elbo))
plt.plot(mvn_loss)
plt.xlabel('Training step')
_ = plt.ylabel('Loss value')
# + [markdown] id="_Wh2eps0fQCZ"
# Since the trained surrogate posterior is a TFP distribution, we can take samples from it and process them to produce posterior credible intervals for the parameters.
#
# The box-and-whiskers plots below show 50% and 95% [credible intervals](https://en.wikipedia.org/wiki/Credible_interval) for the county effect of the two largest counties and the regression weights on soil uranium measurements and mean floor by county. The posterior credible intervals for county effects indicate that location in St. Louis county is associated with lower radon levels, after accounting for other variables, and that the effect of location in Hennepin county is near neutral.
#
# Posterior credible intervals on the regression weights show that higher levels of soil uranium are associated with higher radon levels, and counties where measurements were taken on higher floors (likely because the house didn't have a basement) tend to have higher levels of radon, which could relate to soil properties and their effect on the type of structures built.
#
# The (deterministic) coefficient of floor is negative, indicating that lower floors have higher radon levels, as expected.
# + id="600DiJ8xfQf-"
st_louis_co = 69 # Index of St. Louis, the county with the most observations.
hennepin_co = 25 # Index of Hennepin, with the second-most observations.
def pack_samples(samples):
return {'County effect (St. Louis)': samples.county_effect[..., st_louis_co],
'County effect (Hennepin)': samples.county_effect[..., hennepin_co],
'Uranium weight': samples.uranium_weight,
'Floor-by-county weight': samples.county_floor_weight}
def plot_boxplot(posterior_samples):
fig, axes = plt.subplots(1, 4, figsize=(16, 4))
# Invert the results dict for easier plotting.
k = list(posterior_samples.values())[0].keys()
plot_results = {
v: {p: posterior_samples[p][v] for p in posterior_samples} for v in k}
for i, (var, var_results) in enumerate(plot_results.items()):
sns.boxplot(data=list(var_results.values()), ax=axes[i],
width=0.18*len(var_results), whis=(2.5, 97.5))
# axes[i].boxplot(list(var_results.values()), whis=(2.5, 97.5))
axes[i].title.set_text(var)
fs = 10 if len(var_results) < 4 else 8
axes[i].set_xticklabels(list(var_results.keys()), fontsize=fs)
results = {'Multivariate Normal': pack_samples(mvn_samples)}
print('Bias is: {:.2f}'.format(bias.numpy()))
print('Floor fixed effect is: {:.2f}'.format(floor_weight.numpy()))
plot_boxplot(results)
# + [markdown] id="WnWb8WSDcjEK"
# ### Inverse Autoregressive Flow surrogate posterior
# + [markdown] id="SUHcK4WzJ27o"
# Inverse Autoregressive Flows (IAFs) are normalizing flows that use neural networks to capture complex, nonlinear dependencies among components of the distribution. Next we build an IAF surrogate posterior to see whether this higher-capacity, more fiexible model outperforms the constrained multivariate Normal.
# + id="R0FFLYnaGRrc"
# Build a standard Normal with a vector `event_shape`, with length equal to the
# total number of degrees of freedom in the posterior.
base_distribution = tfd.Sample(
tfd.Normal(0., 1.), sample_shape=[tf.reduce_sum(flat_event_size)])
# Apply an IAF to the base distribution.
num_iafs = 2
iaf_bijectors = [
tfb.Invert(tfb.MaskedAutoregressiveFlow(
shift_and_log_scale_fn=tfb.AutoregressiveNetwork(
params=2, hidden_units=[256, 256], activation='relu')))
for _ in range(num_iafs)
]
# Split the base distribution's `event_shape` into components that are equal
# in size to the prior's components.
split = tfb.Split(flat_event_size)
# Chain these bijectors and apply them to the standard Normal base distribution
# to build the surrogate posterior. `event_space_bijector`,
# `unflatten_bijector`, and `reshape_bijector` are the same as in the
# multivariate Normal surrogate posterior.
iaf_surrogate_posterior = tfd.TransformedDistribution(
base_distribution,
bijector=tfb.Chain([
event_space_bijector, # constrain the surrogate to the support of the prior
unflatten_bijector, # pack the reshaped components into the `event_shape` structure of the prior
reshape_bijector, # reshape the vector-valued components to match the shapes of the prior components
split] + # Split the samples into components of the same size as the prior components
iaf_bijectors # Apply a flow model to the Tensor-valued standard Normal distribution
))
# + [markdown] id="j4pzY9dPrBny"
# Train the IAF surrogate posterior.
# + id="WyQayFhIz1Bq"
optimizer=tf.optimizers.Adam(learning_rate=1e-2)
@tf.function(jit_compile=True)
def fit_vi():
return tfp.vi.fit_surrogate_posterior(
target_model.unnormalized_log_prob,
iaf_surrogate_posterior,
optimizer=optimizer,
num_steps=10**4,
sample_size=4
)
iaf_loss = fit_vi()
iaf_samples = iaf_surrogate_posterior.sample(1000)
iaf_final_elbo = tf.reduce_mean(
target_model.unnormalized_log_prob(*iaf_samples)
- iaf_surrogate_posterior.log_prob(iaf_samples))
print('IAF surrogate posterior ELBO: {}'.format(iaf_final_elbo))
plt.plot(iaf_loss)
plt.xlabel('Training step')
_ = plt.ylabel('Loss value')
# + [markdown] id="tzrbAezxPLeB"
# The credible intervals for the IAF surrogate posterior appear similar to those of the constrained multivariate Normal.
# + id="QmKl4G1BGIIl"
results['IAF'] = pack_samples(iaf_samples)
plot_boxplot(results)
# + [markdown] id="IWKqLYPOZOO_"
# ### Baseline: Mean-field surrogate posterior
#
# VI surrogate posteriors are often assumed to be mean-field (independent) Normal distributions, with trainable means and variances, that are constrained to the support of the prior with a bijective transformation. We define a mean-field surrogate posterior in addition to the two more expressive surrogate posteriors, using the same general formula as the multivariate Normal surrogate posterior.
# + id="GoPeLGAjZLbS"
# A block-diagonal linear operator, in which each block is a diagonal operator,
# transforms the standard Normal base distribution to produce a mean-field
# surrogate posterior.
operators = (tf.linalg.LinearOperatorDiag,
tf.linalg.LinearOperatorDiag,
tf.linalg.LinearOperatorDiag)
block_diag_linop = (
tfp.experimental.vi.util.build_trainable_linear_operator_block(
operators, flat_event_size))
mean_field_scale = tfb.ScaleMatvecLinearOperatorBlock(block_diag_linop)
mean_field_loc = tfb.JointMap(
tf.nest.map_structure(
lambda s: tfb.Shift(
tf.Variable(tf.random.uniform(
(s,), minval=-2., maxval=2., dtype=tf.float32))),
flat_event_size))
mean_field_surrogate_posterior = tfd.TransformedDistribution(
base_standard_dist,
bijector = tfb.Chain( # Note that the chained bijectors are applied in reverse order
[
event_space_bijector, # constrain the surrogate to the support of the prior
unflatten_bijector, # pack the reshaped components into the `event_shape` structure of the posterior
reshape_bijector, # reshape the vector-valued components to match the shapes of the posterior components
mean_field_loc, # allow for nonzero mean
mean_field_scale # apply the block matrix transformation to the standard Normal distribution
]))
optimizer=tf.optimizers.Adam(learning_rate=1e-2)
@tf.function(jit_compile=True)
def fit_vi():
return tfp.vi.fit_surrogate_posterior(
target_model.unnormalized_log_prob,
mean_field_surrogate_posterior,
optimizer=optimizer,
num_steps=10**4,
sample_size=16)
mean_field_loss = fit_vi()
mean_field_samples = mean_field_surrogate_posterior.sample(1000)
mean_field_final_elbo = tf.reduce_mean(
target_model.unnormalized_log_prob(*mean_field_samples)
- mean_field_surrogate_posterior.log_prob(mean_field_samples))
print('Mean-field surrogate posterior ELBO: {}'.format(mean_field_final_elbo))
plt.plot(mean_field_loss)
plt.xlabel('Training step')
_ = plt.ylabel('Loss value')
# + [markdown] id="qv3VzGvMX83Q"
# In this case, the mean field surrogate posterior gives similar results to the more expressive surrogate posteriors, indicating that this simpler model may be adequate for the inference task.
# + id="3_P2nrNSGiG5"
results['Mean Field'] = pack_samples(mean_field_samples)
plot_boxplot(results)
# + [markdown] id="6FtKcJUyTToh"
# ### Ground truth: Hamiltonian Monte Carlo (HMC)
#
# We use HMC to generate "ground truth" samples from the true posterior, for comparison with results of the surrogate posteriors.
# + id="bwTmpfxuC_A4"
num_chains = 8
num_leapfrog_steps = 3
step_size = 0.4
num_steps=20000
flat_event_shape = tf.nest.flatten(target_model.event_shape)
enum_components = list(range(len(flat_event_shape)))
bijector = tfb.Restructure(
enum_components,
tf.nest.pack_sequence_as(target_model.event_shape, enum_components))(
target_model.experimental_default_event_space_bijector())
current_state = bijector(
tf.nest.map_structure(
lambda e: tf.zeros([num_chains] + list(e), dtype=tf.float32),
target_model.event_shape))
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=target_model.unnormalized_log_prob,
num_leapfrog_steps=num_leapfrog_steps,
step_size=[tf.fill(s.shape, step_size) for s in current_state])
hmc = tfp.mcmc.TransformedTransitionKernel(
hmc, bijector)
hmc = tfp.mcmc.DualAveragingStepSizeAdaptation(
hmc,
num_adaptation_steps=int(num_steps // 2 * 0.8),
target_accept_prob=0.9)
chain, is_accepted = tf.function(
lambda current_state: tfp.mcmc.sample_chain(
current_state=current_state,
kernel=hmc,
num_results=num_steps // 2,
num_burnin_steps=num_steps // 2,
trace_fn=lambda _, pkr:
(pkr.inner_results.inner_results.is_accepted),
),
autograph=False,
jit_compile=True)(current_state)
accept_rate = tf.reduce_mean(tf.cast(is_accepted, tf.float32))
ess = tf.nest.map_structure(
lambda c: tfp.mcmc.effective_sample_size(
c,
cross_chain_dims=1,
filter_beyond_positive_pairs=True),
chain)
r_hat = tf.nest.map_structure(tfp.mcmc.potential_scale_reduction, chain)
hmc_samples = pack_samples(
tf.nest.pack_sequence_as(target_model.event_shape, chain))
print('Acceptance rate is {}'.format(accept_rate))
# + [markdown] id="GSRQTbT-T07X"
# Plot sample traces to sanity-check HMC results.
# + id="z34B7sa05KX1"
def plot_traces(var_name, samples):
fig, axes = plt.subplots(1, 2, figsize=(14, 1.5), sharex='col', sharey='col')
for chain in range(num_chains):
s = samples.numpy()[:, chain]
axes[0].plot(s, alpha=0.7)
sns.kdeplot(s, ax=axes[1], shade=False)
axes[0].title.set_text("'{}' trace".format(var_name))
axes[1].title.set_text("'{}' distribution".format(var_name))
axes[0].set_xlabel('Iteration')
warnings.filterwarnings('ignore')
for var, var_samples in hmc_samples.items():
plot_traces(var, var_samples)
# + [markdown] id="ioys0R7QYzH1"
# All three surrogate posteriors produced credible intervals that are visually similar to the HMC samples, though sometimes under-dispersed due to the effect of the ELBO loss, as is common in VI.
# + id="hZ1GUl1dJtpl"
results['HMC'] = hmc_samples
plot_boxplot(results)
# + [markdown] id="V8Y-O_CsT7vH"
# ## Additional results
#
# + cellView="form" id="OUnECXkG42uZ"
#@title Plotting functions
plt.rcParams.update({'axes.titlesize': 'medium', 'xtick.labelsize': 'medium'})
def plot_loss_and_elbo():
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
axes[0].scatter([0, 1, 2],
[mvn_final_elbo.numpy(),
iaf_final_elbo.numpy(),
mean_field_final_elbo.numpy()])
axes[0].set_xticks(ticks=[0, 1, 2])
axes[0].set_xticklabels(labels=[
'Multivariate Normal', 'IAF', 'Mean Field'])
axes[0].title.set_text('Evidence Lower Bound (ELBO)')
axes[1].plot(mvn_loss, label='Multivariate Normal')
axes[1].plot(iaf_loss, label='IAF')
axes[1].plot(mean_field_loss, label='Mean Field')
axes[1].set_ylim([1000, 4000])
axes[1].set_xlabel('Training step')
axes[1].set_ylabel('Loss (negative ELBO)')
axes[1].title.set_text('Loss')
plt.legend()
plt.show()
plt.rcParams.update({'axes.titlesize': 'medium', 'xtick.labelsize': 'small'})
def plot_kdes(num_chains=8):
fig, axes = plt.subplots(2, 2, figsize=(12, 8))
k = list(results.values())[0].keys()
plot_results = {
v: {p: results[p][v] for p in results} for v in k}
for i, (var, var_results) in enumerate(plot_results.items()):
ax = axes[i % 2, i // 2]
for posterior, posterior_results in var_results.items():
if posterior == 'HMC':
label = posterior
for chain in range(num_chains):
sns.kdeplot(
posterior_results[:, chain],
ax=ax, shade=False, color='k', linestyle=':', label=label)
label=None
else:
sns.kdeplot(
posterior_results, ax=ax, shade=False, label=posterior)
ax.title.set_text('{}'.format(var))
ax.legend()
# + [markdown] id="WXzsxJcG1kPH"
# ### Evidence Lower Bound (ELBO)
#
# IAF, by far the largest and most flexible surrogate posterior, converges to the highest Evidence Lower Bound (ELBO).
# + id="cKf_nCvpxohJ"
plot_loss_and_elbo()
# + [markdown] id="5ag72K8X3tpJ"
# ### Posterior samples
#
# Samples from each surrogate posterior, compared with HMC ground truth samples (a different visualization of the samples shown in the box plots).
# + id="_yjwsHIoftLX"
plot_kdes()
# + [markdown] id="45bp131ngAxT"
# ## Conclusion
# + [markdown] id="0xXnCEkKgDa5"
# In this Colab, we built VI surrogate posteriors using joint distributions and multipart bijectors, and fit them to estimate credible intervals for weights in a regression model on the radon dataset. For this simple model, more expressive surrogate posteriors performed similarly to a mean-field surrogate posterior. The tools we demonstrated, however, can be used to build a wide range of flexible surrogate posteriors suitable for more complex models.
| site/en-snapshot/probability/examples/Variational_Inference_and_Joint_Distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# # ACOPF with [PowerSimulations.jl](https://github.com/NREL-SIIP/PowerSimulations.jl) using [PowerModels.jl](https://github.com/lanl-ansi/PowerModels.jl)
# **Originally Contributed by**: <NAME>
# ## Introduction
# PowerSimulations.jl supports non-linear AC optimal power flow through a deep integration
# with [PowerModels.jl](https://github.com/lanl-ansi/PowerModels.jl). This example shows a
# single multi-period optimization of economic dispatch with a full representation of
# AC optimal power flow.
# ## Dependencies
# We can use the a TAMU synthetic ERCOT dataset that is included in the PowerSystemsTestData.
# +
using SIIPExamples
using PowerSystems
using PowerSimulations
using Dates
pkgpath = pkgdir(SIIPExamples)
PowerSystems.download(PowerSystems.TestData; branch = "master") # *note* add `force=true` to get a fresh copy
base_dir = pkgdir(PowerSystems);
# -
# The TAMU data format relies on a folder containing `.m` or `.raw` files and `.csv`
# files for the time series data. We have provided a parser for the TAMU data format with
# the `TamuSystem()` function.
TAMU_DIR = joinpath(base_dir, "data", "ACTIVSg2000");
sys = TamuSystem(TAMU_DIR)
transform_single_time_series!(sys, 2, Hour(1))
# Since we'll be doing non-linear optimization, we need a solver that supports non-linear
# problems. Ipopt is quite good.
using Ipopt
solver = optimizer_with_attributes(Ipopt.Optimizer)
# In the [OperationsProblem example](../../notebook/3_PowerSimulations_examples/1_operations_problems.ipynb)
# we defined a unit-commitment problem with a copper plate representation of the network.
# Here, we want do define an economic dispatch (linear generation decisions) with an ACOPF
# network representation.
# So, starting with the network, we can select from _almost_ any of the endpoints on this
# tree:
TypeTree(PSI.PM.AbstractPowerModel, init_expand = 10, scopesep="\n")
# For now, let's just choose a standard ACOPF formulation.
devices = Dict(
:Generators => DeviceModel(ThermalStandard, ThermalDispatch),
:Loads => DeviceModel(PowerLoad, StaticPowerLoad),
:QLoads => DeviceModel(FixedAdmittance, StaticPowerLoad)
)
ed_template = template_economic_dispatch(network = ACPPowerModel, devices = devices)
# Now we can build a 4-hour economic dispatch / ACOPF problem with the TAMU data.
problem = OperationsProblem(
EconomicDispatchProblem,
ed_template,
sys,
horizon = 1,
optimizer = solver,
balance_slack_variables = true,
)
# And solve it ...
solve!(problem)
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
| notebook/3_PowerSimulations_examples/06_ACOPF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Stelath/pytorch-learning/blob/main/Pytorch_Learning_6_GPU.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="2wt1-dxsr8mp"
# Get Dataset
# + id="oYT7c-WToLV3"
# !wget -q https://download.microsoft.com/download/3/E/1/3E1C3F21-ECDB-4869-8368-6DEBA77B919F/kagglecatsanddogs_3367a.zip
# + id="A6ODwaTWrzbE"
# !unzip -q kagglecatsanddogs_3367a.zip
# + [markdown] id="wsqXtNMpsEDR"
# Import Librarys
# + id="ItjpuQlZsCMx"
import os
import cv2
import numpy as np
from tqdm import tqdm
# + [markdown] id="exSxcm9Cs_ET"
# Import Dataset
# + id="xjW6ySjYs-YI" colab={"base_uri": "https://localhost:8080/"} outputId="57135950-3192-45ee-eccc-82caf19cb40c"
REBUILD_DATA = True
class DogsVSCats():
IMG_SIZE = 50
CATS = 'PetImages/Cat'
DOGS = 'PetImages/Dog'
LABELS = {CATS: 0, DOGS: 1}
training_data = []
cat_count = 0
dog_count = 0
def make_training_data(self):
for label in self.LABELS:
print(label)
for f in tqdm(os.listdir(label)):
try:
path = os.path.join(label, f)
img = cv2.imread(path, cv2.IMREAD_GRAYSCALE)
img = cv2.resize(img, (self.IMG_SIZE, self.IMG_SIZE))
self.training_data.append([np.array(img), np.eye(2)[self.LABELS[label]]])
if label == self.CATS:
self.cat_count += 1
elif label == self.DOGS:
self.dog_count += 1
except Exception as e:
pass
# print(str(e))
np.random.shuffle(self.training_data)
np.save('training_data.npy', self.training_data)
print()
print('Cats:', self.cat_count)
print('Dogs:', self.dog_count)
if REBUILD_DATA:
dogvcats = DogsVSCats()
dogvcats.make_training_data()
# + id="TRljgL0k5-Bj"
training_data = np.load('training_data.npy', allow_pickle=True)
print(len(training_data))
print(training_data[0])
# + id="96zgN4Tu9sLE"
import matplotlib.pyplot as plt
plt.imshow(training_data[2][0], cmap='gray')
plt.show()
# + id="G_fHBmlk9z27"
training_data[2][1]
# + [markdown] id="HqE6ePQ5_bkr"
# Build Model
# + id="rzAGDyfO_ar1"
import torch
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 32, 5)
self.conv2 = nn.Conv2d(32, 64, 5)
self.conv3 = nn.Conv2d(64, 128, 5)
x = torch.randn(50, 50).view(-1, 1, 50, 50)
self._to_linear = None
self.convs(x)
self.fc1 = nn.Linear(self._to_linear, 512)
self.fc2 = nn.Linear(512, 2)
def convs(self, x):
x = F.max_pool2d(F.relu(self.conv1(x)), (2, 2))
x = F.max_pool2d(F.relu(self.conv2(x)), (2, 2))
x = F.max_pool2d(F.relu(self.conv3(x)), (2, 2))
if self._to_linear is None:
self._to_linear = x[0].shape[0] * x[0].shape[1] * x[0].shape[2]
return x
def forward(self, x):
x = self.convs(x)
x = x.view(-1, self._to_linear)
x = F.relu(self.fc1(x))
x = self.fc2(x)
# Use activation function cause its probably better
return F.softmax(x, dim = 1)
# + [markdown] id="osr147kmRj0t"
# CUDA
# + id="94FwKHM7RT1y"
print('CUDA Available:', torch.cuda.is_available())
if torch.cuda.is_available():
device = torch.device('cuda:0')
print('Running on the GPU')
else:
device = torch.device('cpu')
print('Running on the CPU')
net = Net().to(device)
# + [markdown] id="85xuSL_jRl4r"
# Optimization
# + id="CARjWQ2PGYHn"
import torch.optim as optim
optimizer = optim.Adam(net.parameters(), lr=0.001)
loss_function = nn.MSELoss()
# + [markdown] id="YYOv0nYARYUw"
# Training & Testing
# + id="0CHV9jO2Kk-G"
X = torch.Tensor([i[0] for i in training_data]).view(-1, 50, 50)
X = X/255.0 # Scale imagery so pixle values are between 0 and 1 not 0 and 255
y = torch.Tensor([i[1] for i in training_data])
VAL_PCT = 0.1
val_size = int(len(X) * VAL_PCT)
print(val_size)
train_X = X[:-val_size]
train_y = y[:-val_size]
test_X = X[-val_size:]
test_y = y[-val_size:]
print(len(train_X))
print(len(test_X))
# + id="ZwaV-3xgK1dH"
def train(net, EPOCHS=3, BATCH_SIZE=100):
for epoch in range(EPOCHS):
for i in tqdm(range(0, len(train_X), BATCH_SIZE)):
batch_X = train_X[i:i + BATCH_SIZE].view(-1, 1, 50, 50)
batch_y = train_y[i:i + BATCH_SIZE]
batch_X, batch_y = batch_X.to(device), batch_y.to(device) # NEEDED FOR CUDA
net.zero_grad()
outputs = net(batch_X)
loss = loss_function(outputs, batch_y)
loss.backward()
optimizer.step()
print(f'Epoch: {epoch}\nLoss: {loss}')
# + id="TiYt7RyJLckl"
def test(net):
correct = 0
total = 0
with torch.no_grad():
for i in tqdm(range(len(test_X))):
real_class = torch.argmax(test_y[i]).to(device) # NEEDED FOR CUDA
net_out = net(test_X[i].view(-1, 1, 50, 50).to(device))[0]
predicted_class = torch.argmax(net_out)
if predicted_class == real_class:
correct += 1
total += 1
print()
print('Accuracy:', round(correct / total, 3))
# + id="tx1XbZdyVROD"
train(net, EPOCHS=12)
# + id="KFegjH39Xy5b"
test(net)
| Pytorch_Learning_6_GPU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment1
#
# Name: <NAME>
#
# ID: 11712121
# ## 1 Part I: the perceptron
# ### Code
# +
import numpy as np
import matplotlib.pyplot as plt
def gen_gaussian_distribution(size, mean=None, cov=None):
if not mean:
mean = np.random.randn(2)
if not cov:
cov = np.eye(2)
data = np.random.multivariate_normal(mean, cov, size)
return data
class Perceptron(object):
def __init__(self, n_inputs, max_epochs=1e2, learning_rate=1e-2):
"""
Initializes perceptron object.
Args:
n_inputs: number of inputs.
max_epochs: maximum number of training cycles.
learning_rate: magnitude of weight changes at each training cycle
"""
self.n_inputs = n_inputs
self.max_epochs = max_epochs
self.learning_rate = learning_rate
self.weights = np.zeros(self.n_inputs)
self.bias = 0
def forward(self, input):
"""
Predict label from input
Args:
input: array of dimension equal to n_inputs.
"""
sum = np.sign(np.dot(input, self.weights))
label = np.where(sum > 0, 1, -1)
return label
def train(self, training_inputs, labels):
"""
Train the perceptron
Args:
training_inputs: list of numpy arrays of training points.
labels: arrays of expected output value for the corresponding point in training_inputs.
"""
train_size = len(training_inputs)
epochs = 0
while epochs < self.max_epochs:
epochs += 1
for i in range(train_size):
if np.any(labels[i] * (np.dot(self.weights, training_inputs[i]) + self.bias) <= 0):
self.weights = self.weights + (self.learning_rate * labels[i] * training_inputs[i]).T
self.bias = self.bias + self.learning_rate * labels[i]
def score(self, test_inputs, test_labels):
pred_arr = np.where(self.forward(test_inputs) > 0, 1, -1)
true_size = len(np.where(pred_arr == test_labels)[0])
return true_size / len(test_labels)
def main():
p = Perceptron(2)
"""
gen dataset
"""
data_size = 100
train_size = 80
x1 = gen_gaussian_distribution(data_size, [5, 5])
x2 = gen_gaussian_distribution(data_size, [-5, -5])
y1 = a_label = np.ones(data_size, dtype=np.int16)
y2 = -y1
x_train = np.concatenate((x1[:train_size], x2[:train_size]), axis=0)
y_train = np.concatenate((y1[:train_size], y2[:train_size]), axis=0)
x_test = np.concatenate((x1[train_size:], x2[train_size:]), axis=0)
y_test = np.concatenate((y1[train_size:], y2[train_size:]), axis=0)
"""
train model
"""
p.train(x_train, y_train)
"""
test model
"""
acc = p.score(x_test, y_test)
if __name__ == "__main__":
main()
# -
# ### 1.1 Task 1
#
# > Generate a dataset of points in R2. To do this, define two Gaussian distributions and sample 100 points from each. Your dataset should then contain a total of 200 points, 100 from each distribution. Keep 80 points per distribution as the training (160 in total), 20 for the test (40 in total).
# +
"""
gen dataset
"""
data_size = 100
train_size = 80
x1 = gen_gaussian_distribution(data_size, [5, 5])
x2 = gen_gaussian_distribution(data_size, [-5, -5])
y1 = a_label = np.ones(data_size, dtype=np.int16)
y2 = -y1
x_train = np.concatenate((x1[:train_size], x2[:train_size]), axis=0)
y_train = np.concatenate((y1[:train_size], y2[:train_size]), axis=0)
x_test = np.concatenate((x1[train_size:], x2[train_size:]), axis=0)
y_test = np.concatenate((y1[train_size:], y2[train_size:]), axis=0)
# plt
plt.plot(x1[:,0], x1[:,1], 'x')
plt.plot(x2[:,0], x2[:,1], 'x')
plt.axis('equal')
plt.savefig('./img/fig1.png')
plt.show()
# -
# We set `mean1 = [5, 5]` and `mean2 = [-5, -5]`, generate `cov1 and cov2` with `np.eve(2)` which returns a 2-D array with ones on the diagonal and zeros elsewhere.
# ### 1.2 Task 2
#
# > Implement the perceptron following the specs in perceptron.py and the pseudocode in perceptronslides.pdf.
# ### 1.3 Task 3
#
# > Train the perceptron on the training data (160 points) and test in on the remaining 40 test points. Compute the classification accuracy on the test set.
# +
p = Perceptron(2)
"""
gen dataset
"""
data_size = 100
train_size = 80
x1 = gen_gaussian_distribution(data_size, [5, 5])
x2 = gen_gaussian_distribution(data_size, [-5, -5])
y1 = a_label = np.ones(data_size, dtype=np.int16)
y2 = -y1
x_train = np.concatenate((x1[:train_size], x2[:train_size]), axis=0)
y_train = np.concatenate((y1[:train_size], y2[:train_size]), axis=0)
x_test = np.concatenate((x1[train_size:], x2[train_size:]), axis=0)
y_test = np.concatenate((y1[train_size:], y2[train_size:]), axis=0)
"""
train model
"""
p.train(x_train, y_train)
"""
test model
"""
acc = p.score(x_test, y_test)
print(f'Perceptron test accuracy: {acc * 100}%')
# -
# ### 1.4 Task 4
#
# > Experiment with different sets of points (generated as described in Task 1). What happens during the training if the means of the two Gaussians are too close and/or if their variance is too high?
for _ in range(10):
p = Perceptron(2)
"""
gen dataset
"""
data_size = 100
train_size = 80
x1 = gen_gaussian_distribution(data_size, [1, 1])
x2 = gen_gaussian_distribution(data_size, [1, 1])
y1 = a_label = np.ones(data_size, dtype=np.int16)
y2 = -y1
x_train = np.concatenate((x1[:train_size], x2[:train_size]), axis=0)
y_train = np.concatenate((y1[:train_size], y2[:train_size]), axis=0)
x_test = np.concatenate((x1[train_size:], x2[train_size:]), axis=0)
y_test = np.concatenate((y1[train_size:], y2[train_size:]), axis=0)
"""
train model
"""
p.train(x_train, y_train)
"""
test model
"""
acc = p.score(x_test, y_test)
print(f'Perceptron test accuracy: {acc * 100}%')
# We run 10 times for the close Gaussians([1, 1], [1, 1]), and accuracy is lower then 50%.
for _ in range(10):
p = Perceptron(2)
"""
gen dataset
"""
data_size = 100
train_size = 80
x1 = gen_gaussian_distribution(data_size, [-5, -5])
x2 = gen_gaussian_distribution(data_size, [1, 1])
y1 = a_label = np.ones(data_size, dtype=np.int16)
y2 = -y1
x_train = np.concatenate((x1[:train_size], x2[:train_size]), axis=0)
y_train = np.concatenate((y1[:train_size], y2[:train_size]), axis=0)
x_test = np.concatenate((x1[train_size:], x2[train_size:]), axis=0)
y_test = np.concatenate((y1[train_size:], y2[train_size:]), axis=0)
"""
train model
"""
p.train(x_train, y_train)
"""
test model
"""
acc = p.score(x_test, y_test)
print(f'Perceptron test accuracy: {acc * 100}%')
# We run 10 times for the Gaussians with high variance([-5, -5], [1, 1]), and accuracy is higher then the close Gaussians, over 90%.
| CS324_Deep-Learning/Assignmnet1/Part 1/11712121_assignment1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Setup
# +
from day2 import puzzle3
from day2 import puzzle4
sample_input2 = "1-3 a: abcde\n1-3 b: cdefg\n2-9 c: ccccccccc"
input2 = open("input2.txt", "r").read()
# -
# # Puzzle Timings
# %%timeit
puzzle3(input2)
# %%timeit
puzzle4(input2)
# # Sample Timings
# %%timeit
puzzle3(sample_input2)
# %%timeit
puzzle4(sample_input2)
| AdventofCode2020/timings/Timing Day 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Quiz : `# TODO` 부분에 코드를 입력하여 아래와 같은 출력결과가 나오도록 코드를 작성하세요.
# %matplotlib inline
# %config InlineBackend.figure_formats = {'png', 'retina'}
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.font_manager as fm
# 데이터 로드
kospi_df = pd.read_csv("kospi.csv")
kospi_df.tail(1)
# #### 1. 모든 기업(name)이 포함되는 월별 상승률(cr)를 데이터 프레임으로 출력하고, 출력된 데이터 프레임을 리스트의 딕셔너리포멧으로 출력하세요.
# +
# 모든 기업(name)이 포함되는 월별 상승률(cr)를 데이터 프레임으로 출력
# -
# 사용할 column select 후, 작업을 진행할 df_1 생성
columns = ["cr", "dt"]
df_1 = kospi_df[columns]
df_1.tail(5)
# month column 추가
df_1["month"] = 0
df_1.tail(1)
# checkpoint
df_2 = df_1
df_2.tail(1)
# dt range 확인
df_2["dt"].max(), df_2["dt"].min()
# +
# month data 입력
df_2.loc[df_2["dt"] <= 20191231, "month"] = '2019-12'
df_2.loc[df_2["dt"] <= 20191130, "month"] = '2019-11'
df_2.loc[df_2["dt"] <= 20191031, "month"] = '2019-10'
df_2.loc[df_2["dt"] <= 20190930, "month"] = '2019-09'
df_2.loc[df_2["dt"] <= 20190831, "month"] = '2019-08'
df_2.loc[df_2["dt"] <= 20190731, "month"] = '2019-07'
df_2
# -
col1 = ["month", "cr"]
df_3 = df_2[col1]
df_3.tail(1)
df_3.loc[df_3['month']=='2019-07', 'cr'].mean()
month = ["2019-07", "2019-08", "2019-09", "2019-10", "2019-11", "2019-12"]
cr = [
round(df_3.loc[df_3['month']=='2019-07', 'cr'].mean(), 6),
round(df_3.loc[df_3['month']=='2019-08', 'cr'].mean(), 6),
round(df_3.loc[df_3['month']=='2019-09', 'cr'].mean(), 6),
round(df_3.loc[df_3['month']=='2019-10', 'cr'].mean(), 6),
round(df_3.loc[df_3['month']=='2019-11', 'cr'].mean(), 6),
round(df_3.loc[df_3['month']=='2019-12', 'cr'].mean(), 6),
]
month, cr
datas = [
{"month":"2019-07", "cr":df_3.loc[df_3['month']=='2019-07', 'cr'].mean()},
{"month":"2019-08", "cr":df_3.loc[df_3['month']=='2019-08', 'cr'].mean()},
{"month":"2019-09", "cr":df_3.loc[df_3['month']=='2019-09', 'cr'].mean()},
{"month":"2019-10", "cr":df_3.loc[df_3['month']=='2019-10', 'cr'].mean()},
{"month":"2019-11", "cr":df_3.loc[df_3['month']=='2019-11', 'cr'].mean()},
{"month":"2019-12", "cr":df_3.loc[df_3['month']=='2019-12', 'cr'].mean()},
]
result_df = pd.DataFrame(datas)
result_df
# +
# TODO
result_df
# +
# 결과 데이터를 소수 셋째자리까지 출력되도록 아래와 같이 리스트의 딕셔너리 포멧으로 출력
# -
datas = [
{"month":"2019-07", "cr":round(df_3.loc[df_3['month']=='2019-07', 'cr'].mean(), 3)},
{"month":"2019-08", "cr":round(df_3.loc[df_3['month']=='2019-08', 'cr'].mean(), 3)},
{"month":"2019-09", "cr":round(df_3.loc[df_3['month']=='2019-09', 'cr'].mean(), 3)},
{"month":"2019-10", "cr":round(df_3.loc[df_3['month']=='2019-10', 'cr'].mean(), 3)},
{"month":"2019-11", "cr":round(df_3.loc[df_3['month']=='2019-11', 'cr'].mean(), 3)},
{"month":"2019-12", "cr":round(df_3.loc[df_3['month']=='2019-12', 'cr'].mean(), 3)},
]
datas
# +
# TODO
result_datas
# -
# #### 2. 2019년 11월 01일에서 2019년 12월 5일까지 일별 상승률 상위 5개 회사의 평균 상승율을 소수점 셋째 자리까지 출력하세요.
df_4 = kospi_df
df_4.tail(1)
# 20191101~20191205까지의 일별 상승률
df_5 = df_4[df_4["dt"] >= 20191101]
df_5 = df_5[df_5["dt"] <= 20191205]
col2 = ["name", "cr"]
df_5[col2]
# 평균 일별 상승률
df_6 = df_5[col2]
df_7 = df_6.groupby("name").agg("mean").reset_index()[["name", "cr"]]
df_7
# 평균 일별 상승률 상위 5개
df_7.sort_values("cr", ascending=False)[:5]
# 평균 일별 상승률 상위 5개 회사에 대한 평균
mean_cr = round(df_7.sort_values("cr", ascending=False)[:5].mean(), 3)
mean_cr
# 일별 상승률 상위 5개 회사
df_5[col2].sort_values("cr", ascending=False)[:5]
# 일별 상승률 상위 5개 회사에 대한 평균
mean_cr = round(df_5[col2].sort_values("cr", ascending=False)[:5].mean(), 3)
mean_cr
# +
# TODO
mean_cr
# -
# #### 3. 전체 기간에서 평균 등락이 높은 3개의 회사와 평균 등락을 출력하세요. (cv: 등락)
kospi_df.tail(1)
df_8 = kospi_df
df_8.tail(1)
df_9 = df_8.groupby("name").agg("mean").reset_index()[["name", "cv"]]
result_dict = df_9.sort_values("cv", ascending=False)[:3]
result_dict.to_dict()
result_dict
# +
# TODO
result_dict
# -
# #### 4. 한진칼, 한국전력, 이마트 기업의 종가(ncv) 데이터를 아래와 같이 정규화하여 그래프를 출력하세요.
# - 정규화(Nomalization) 수식
# - 데이터의 스케일이 0 ~ 1 사이의 값으로 변경됩니다.
#
# $$y_i = \frac{x_i-min(x)}{max(x)-min(x)}$$
#
#
# - companies 변수에 회사이름을 변경하면 변경된 회사의 데이터로 출력되도록 코드를 작성하세요. (직접적인 회사이름은 companies 변수 설정에서만 사용이 가능합니다.)
# +
# 한글 폰트 적용 변경하는 방법
import matplotlib.font_manager as fm
font_location = "C:/Windows/fonts/Myungjo.ttf" # windows
font_location = "/Library/Fonts/AppleMyungjo.ttf" # mac 카탈리나 이전 버전
font_location = "/System/Library/Fonts/Supplemental/AppleMyungjo.ttf" # mac 카탈리나 버전
font_name = fm.FontProperties(fname=font_location).get_name()
mpl.rc("font", family=font_name)
mpl.rcParams["font.family"] # 현재 폰트 확인
# +
column = ["name", "ncv", "dt"]
df_b = df_a[column]
companies = ["한진칼", "한국전력", "이마트"]
df_b["name" == "삼성전자",]
# +
# 위에서 찾은 3개의 회사 데이터만 필터링
companies = ["한진칼", "한국전력", "이마트"]
# TODO
df
# -
# 정규화 함수 작성
def nomalization(df, company):
# TODO
# +
# 기업별 정규화한 데이터 컬럼 적용
df["nomal"] = 0
# TODO
df
# +
# 기업별 정규화한 값으로 그래프 그리기
plt.figure(figsize=(20, 5))
# TODO
plt.show()
# -
# #### 5. 아래와 같이 기업별 상관계수를 출력하세요.
# +
companies = ["한진칼", "한국전력", "이마트"]
# TODO
result_df
| python/Exam/01_python_numpy_pandas_gunhoko.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="okAZVx9nb9Es"
# # Drive mount code
# + id="Re1cOmeea_7J" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632560787290, "user_tz": -360, "elapsed": 533, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="0d939459-2c9f-49b4-d018-def0136cdc55"
from google.colab import drive
drive.mount('./drive')
# + [markdown] id="nW4ElwLgysH_"
# # Dataset Creation
# + [markdown] id="Ay2TDwZCy96_"
# ## Download dataset 2
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="wlb4s-FrcGxi" executionInfo={"status": "ok", "timestamp": 1632563411634, "user_tz": -360, "elapsed": 2516, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="da618ae0-564b-49e9-e912-dd6e3de6411c"
# !gdown --id 1L7gRzRsyWzQ6jtRg86q81aVfKbREYv2A
# + [markdown] id="9RQpoQIizIfo"
# ## Import package
# + colab={"base_uri": "https://localhost:8080/"} id="uLl0_bE2cC-C" executionInfo={"status": "ok", "timestamp": 1632563416457, "user_tz": -360, "elapsed": 4827, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="e0c9f86c-33e3-4095-f53a-902cc7270b64"
# import some importent library or packages
import matplotlib.pyplot as plt
import warnings
import time,sys
import copy
import pandas as pd
import numpy as np
import cv2
import os
import pathlib
import zipfile
import torch
import torchvision
from torchvision import models
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader, ConcatDataset
import torch.optim as optim
import pathlib
import shutil
from pathlib import Path
from collections import Counter
# !pip install torchsummary
from torchsummary import summary
from sklearn.utils import shuffle
# !pip install torchviz
from torchviz import make_dot, make_dot_from_trace
warnings.filterwarnings('ignore')
torch.manual_seed(0)
# + [markdown] id="tzq1MJWAzLy3"
# ## Read downloaded dataset
# + id="SG4wK48ZcGFy" executionInfo={"status": "ok", "timestamp": 1632563416463, "user_tz": -360, "elapsed": 59, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
df = pd.read_csv('/content/Dataset 2.csv',encoding='unicode_escape')
# + colab={"base_uri": "https://localhost:8080/"} id="yZCvTyl6keZe" executionInfo={"status": "ok", "timestamp": 1632563416466, "user_tz": -360, "elapsed": 58, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="a6b221df-b706-4ef0-8119-4a8d15b78159"
df.columns ## columns name
# + colab={"base_uri": "https://localhost:8080/"} id="l8lCKytvnQxF" executionInfo={"status": "ok", "timestamp": 1632563416468, "user_tz": -360, "elapsed": 55, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="6316a6de-ccf0-4c76-f6c0-52c18f9ca813"
df['polarity'].value_counts() ## get number of data in `polarity` class
# + colab={"base_uri": "https://localhost:8080/"} id="wU8m4CcPkiV1" executionInfo={"status": "ok", "timestamp": 1632563416469, "user_tz": -360, "elapsed": 54, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="0b95e673-df18-4cb6-930c-6810f5c20f01"
df['text'].value_counts()
# + [markdown] id="BnThLglw3HEu"
# ## Dataset splitting
# + id="NddseGZ0n-Y8" executionInfo={"status": "ok", "timestamp": 1632563416470, "user_tz": -360, "elapsed": 52, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
## split dataset based on the class
traning_split_size = 0.8
df_class_1 = df[df['polarity'] == 1]
df_class_0 = df[df['polarity'] == 0]
trainSize = int(len(df_class_0) * traning_split_size)
Traning_class_0 = df_class_0[:trainSize]
Test_class_0 = df_class_0[trainSize:]
trainSize = int(len(df_class_1) * traning_split_size)
Traning_class_1 = df_class_1[:trainSize]
Test_class_1 = df_class_1[trainSize:]
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="MJnpde_qvabR" executionInfo={"status": "ok", "timestamp": 1632563416471, "user_tz": -360, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="91b94165-8578-4c88-bac0-9c6d07d25890"
## traning dataset create
li = [Traning_class_0,Traning_class_1]
frame = pd.concat(li, axis=0, ignore_index=True)
frame = shuffle(frame)
frame.reset_index(inplace=True, drop=True)
frame.to_csv('Train-Dataset.csv',index=False)
frame
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="UC7KHE3Jxnax" executionInfo={"status": "ok", "timestamp": 1632563416472, "user_tz": -360, "elapsed": 48, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="b069021a-9b77-4768-9263-01781eedeb34"
## testing dataset create
li = [Test_class_0,Test_class_1]
frame = pd.concat(li, axis=0, ignore_index=True)
frame = shuffle(frame)
frame.reset_index(inplace=True, drop=True)
frame.to_csv('Test-Dataset.csv',index=False)
frame
# + [markdown] id="SwhD-3_x3t5V"
# # Text cleaning
# + colab={"base_uri": "https://localhost:8080/"} id="vf7x0ff83u0F" executionInfo={"status": "ok", "timestamp": 1632563495158, "user_tz": -360, "elapsed": 5514, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="84b0c92a-90a2-4cc4-fb6c-41a32ac6afaa"
# import some importent library or packages
import matplotlib.pyplot as plt
import warnings
import time,sys,re,string
import copy
import pandas as pd
import numpy as np
import cv2
import os
import pathlib
import zipfile
import torch
import torchvision
from torchvision import models
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader, ConcatDataset
import torch.optim as optim
import pathlib
import shutil
from pathlib import Path
from collections import Counter
# !pip install torchsummary
from torchsummary import summary
from sklearn.utils import shuffle
# !pip install torchviz
from torchviz import make_dot, make_dot_from_trace
try:
import contractions
except:
# !pip install contractions
import contractions
from torchtext.legacy.data import Field, TabularDataset, BucketIterator
from torchtext.data.utils import get_tokenizer
from torchtext.vocab import build_vocab_from_iterator, Vectors, GloVe
import nltk
try:
nltk.data.find('tokenizers/punkt')
nltk.data.find('averaged_perceptron_tagger')
nltk.data.find('brown')
except LookupError:
nltk.download('averaged_perceptron_tagger')
nltk.download('brown')
nltk.download('punkt')
from nltk import sent_tokenize,word_tokenize
warnings.filterwarnings('ignore')
torch.manual_seed(0)
# + id="UqxEVwxP37g2" executionInfo={"status": "ok", "timestamp": 1632563495161, "user_tz": -360, "elapsed": 213, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
traning_df = pd.read_csv('/content/Train-Dataset.csv')
testing_df = pd.read_csv('/content/Test-Dataset.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="gulKysGivn3d" executionInfo={"status": "ok", "timestamp": 1632563495164, "user_tz": -360, "elapsed": 210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="a79e7b4a-3cbb-4abb-f549-4a64463546ca"
traning_df
# + id="qdQ4ajcoOdX6" executionInfo={"status": "ok", "timestamp": 1632563496779, "user_tz": -360, "elapsed": 1677, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
def clean_text(text):
'''Make text lowercase, remove text in square brackets,remove links,remove punctuation
and remove words containing numbers. removibng miltiple full stop'''
text = str(text).lower()
text = re.sub('\[.*?\]', '', text)
text = re.sub('https?://\S+|www\.\S+', '', text)
text = re.sub('<.*?>+', '', text)
text = re.sub('[%s]' % re.escape(string.punctuation), '', text)
text = re.sub('\n', '', text)
text = re.sub('\w*\d\w*', '', text)
text = re.sub(r'\.+', ".", text)
return text
def replace_text(text):
text = str(text).lower()
text = text.encode('ascii', 'ignore').decode('utf-8')
return text
for dta in [traning_df,testing_df]:
dta['text_cleaning'] = dta.text.apply(lambda x: x.strip().lower() )
dta['text_cleaning'] = dta.text_cleaning.apply(lambda x : " ".join(x.split()) )
dta['text_cleaning'] = dta.text_cleaning.apply(lambda x: contractions.fix(x) )
dta['text_cleaning'] = dta.text_cleaning.apply(lambda x: clean_text(x) )
dta['text_cleaning'] = dta.text_cleaning.apply(lambda x: replace_text(x) )
dta['number_of_word'] = dta.text_cleaning.apply(lambda x: len(word_tokenize(x)) )
dta['number_of_letter'] = dta.text_cleaning.apply(lambda x: len(x) )
del dta['text']
dta['text'] = dta['text_cleaning']
del dta['text_cleaning']
for dta in [traning_df,testing_df]:
word_count_zero = dta[dta['number_of_word'] == 0]
dta.drop(word_count_zero.index,inplace=True)
letter_count_zero = dta[dta['number_of_letter'] == 0]
dta.drop(letter_count_zero.index,inplace=True)
# # Training DF
# traning_df['text_cleaning'] = traning_df.text.apply(lambda x: x.strip().lower() )
# traning_df['text_cleaning'] = traning_df.text_cleaning.apply(lambda x : " ".join(x.split()) )
# traning_df['text_cleaning'] = traning_df.text_cleaning.apply(lambda x: contractions.fix(x) )
# traning_df['text_cleaning'] = traning_df.text_cleaning.apply(lambda x: clean_text(x) )
# traning_df['text_cleaning'] = traning_df.text_cleaning.apply(lambda x: replace_text(x) )
# # Testing DF
# testing_df['text_cleaning'] = testing_df.text.apply(lambda x: x.strip().lower() )
# testing_df['text_cleaning'] = traning_df.text_cleaning.apply(lambda x : " ".join(x.split()) )
# testing_df['text_cleaning'] = testing_df.text_cleaning.apply(lambda x: contractions.fix(x) )
# testing_df['text_cleaning'] = testing_df.text_cleaning.apply(lambda x: clean_text(x) )
# testing_df['text_cleaning'] = testing_df.text_cleaning.apply(lambda x: replace_text(x) )
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="OptnUdYqPe_D" executionInfo={"status": "ok", "timestamp": 1632563496795, "user_tz": -360, "elapsed": 29, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="ad4795a5-07fa-42a3-ace3-6a229f8d1ba1"
traning_df
# + colab={"base_uri": "https://localhost:8080/"} id="CnDFX37x7tcc" executionInfo={"status": "ok", "timestamp": 1632563497232, "user_tz": -360, "elapsed": 460, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="504b10ec-d0ae-4905-8531-713df7cedb49"
alltext_length = []
allword_length = []
for val in traning_df.text:
word_tok = word_tokenize(val)
alltext_length.append(len(val))
allword_length.append(len(word_tok))
print(max(alltext_length))
print(max(allword_length))
# traning_df.text.apply(lambda x: len(x) )
# + colab={"base_uri": "https://localhost:8080/"} id="doo23zv2ZNj4" executionInfo={"status": "ok", "timestamp": 1632563498351, "user_tz": -360, "elapsed": 1123, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="66bb9334-1088-4b1f-ff34-c35bae164a69"
for val in traning_df.text:
word_tok = word_tokenize(val)
if len(word_tok) <= 1:
print(val)
# + id="U5lXljaX4nx8" executionInfo={"status": "ok", "timestamp": 1632563498353, "user_tz": -360, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
traning_df.to_csv('/content/Train-Dataset-prcessed.csv',index=False)
testing_df.to_csv('/content/Test-Dataset-prcessed.csv',index=False)
# + [markdown] id="pS1_ntiM5itz"
# # Training & `LSTM` modeling
# + colab={"base_uri": "https://localhost:8080/"} id="VCpz-8Tl6EX0" executionInfo={"status": "ok", "timestamp": 1632565712040, "user_tz": -360, "elapsed": 7747, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="e5bc01eb-1cc0-47d8-9ba3-afb80d408185"
# import some importent library or packages
import matplotlib.pyplot as plt
import warnings
import time,sys,re,string
import copy
import pandas as pd
import numpy as np
import cv2
import os
import pathlib
import zipfile
import torch
import torchvision
from torchvision import models
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader, ConcatDataset
import torch.optim as optim
import pathlib
import shutil
from pathlib import Path
from collections import Counter
# !pip install torchsummary
# !pip install torchinfo
from torchinfo import summary
from sklearn.utils import shuffle
# !pip install torchviz
from torchviz import make_dot, make_dot_from_trace
try:
import contractions
except:
# !pip install contractions
import contractions
from torchtext.legacy.data import Field, TabularDataset, BucketIterator
from torchtext.data.utils import get_tokenizer
from torchtext.vocab import build_vocab_from_iterator, Vectors, GloVe
from torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence
import nltk
try:
nltk.data.find('tokenizers/punkt')
nltk.data.find('averaged_perceptron_tagger')
nltk.data.find('brown')
except LookupError:
nltk.download('averaged_perceptron_tagger')
nltk.download('brown')
nltk.download('punkt')
from nltk import sent_tokenize,word_tokenize
warnings.filterwarnings('ignore')
torch.manual_seed(0)
# + id="-4O9tu5u6LC6" executionInfo={"status": "ok", "timestamp": 1632565712042, "user_tz": -360, "elapsed": 17, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
traning_df = pd.read_csv('/content/Train-Dataset-prcessed.csv')
testing_df = pd.read_csv('/content/Test-Dataset-prcessed.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="kQmdyRqE6fwK" executionInfo={"status": "ok", "timestamp": 1632565712042, "user_tz": -360, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="f170b7b2-1648-401a-c274-28127a1ec36a"
traning_df
# + id="7n0l74pAlyr_" executionInfo={"status": "ok", "timestamp": 1632565712702, "user_tz": -360, "elapsed": 670, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
label_field = Field(sequential=False, use_vocab=False, batch_first=True, dtype=torch.float)
text_field = Field(tokenize='spacy', lower=True, include_lengths=True, batch_first=True)
fields = [('polarity', label_field), ('text', text_field)]
train_data = TabularDataset(path="/content/Train-Dataset-prcessed.csv",
format="csv",
fields=fields,
skip_header=True)
valid_data = TabularDataset(path="/content/Test-Dataset-prcessed.csv",
format="csv",
fields=fields,
skip_header=True)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
train_iter = BucketIterator(train_data, batch_size=32, sort_key=lambda x: len(x.text),
device=device, sort=True, sort_within_batch=True)
valid_iter = BucketIterator(valid_data, batch_size=32, sort_key=lambda x: len(x.text),
device=device, sort=True, sort_within_batch=True)
text_field.build_vocab(train_data,) # min_freq=3,vectors = "glove.6B.100d"
label_field.build_vocab(train_data)
# + id="9GXT5L5ADSU3" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632565712702, "user_tz": -360, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="bc026445-8bbc-431e-9cf0-fe0d98fd07a1"
#No. of unique tokens in text
print("Size of TEXT vocabulary:",len(text_field.vocab))
#No. of unique tokens in label
print("Size of LABEL vocabulary:",len(label_field.vocab))
#Commonly used words
print(text_field.vocab.freqs.most_common(10))
#Word dictionary
print(text_field.vocab.stoi)
# + id="J_GgfPEM1Rri" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632565713807, "user_tz": -360, "elapsed": 1108, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="84e10388-1286-4a93-f1d9-d77a6fc323c6"
for batch in train_iter:
print(batch.polarity)
print(batch.text)
# + [markdown] id="_oF4NWmtDAEE"
# ## Code
#
# + id="hNxd9n-6JUeb" executionInfo={"status": "ok", "timestamp": 1632565713808, "user_tz": -360, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
import torch.nn as nn
class LSTMTagger(torch.nn.Module):
def __init__(self, embedding_dim, hidden_dim, vocab_size, tagset_size):
super(LSTMTagger, self).__init__()
self.hidden_dim = hidden_dim
self.word_embeddings = nn.Embedding(vocab_size, embedding_dim)
# The LSTM takes word embeddings as inputs, and outputs hidden states
# with dimensionality hidden_dim.
self.lstm = nn.LSTM(embedding_dim, hidden_dim)
# The linear layer that maps from hidden state space to tag space
self.hidden2tag = nn.Linear(hidden_dim, tagset_size)
def forward(self, sentence):
embeds = self.word_embeddings(sentence)
# print(len(sentence))
lstm_out, _ = self.lstm(embeds.view(len(sentence), 1, -1))
x = embeds.view(len(sentence), 1, -1)
# print(x.shape)
lstm_out, _ = self.lstm(x)
tag_space = self.hidden2tag(lstm_out.view(len(sentence), -1))
tag_scores = torch.nn.functional.log_softmax(tag_space, dim=1)
return tag_scores
# + id="Ve_OtXk4JcID" executionInfo={"status": "ok", "timestamp": 1632565713808, "user_tz": -360, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
EMBEDDING_DIM = 6
HIDDEN_DIM = 6
model = LSTMTagger(EMBEDDING_DIM, HIDDEN_DIM, len(text_field.vocab), 1)
loss_function = torch.nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.00001)
criterion = torch.nn.BCEWithLogitsLoss()
# + id="ubro-1vsQyAi" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632565713809, "user_tz": -360, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="fb6190f2-b214-4607-fa62-c3ef6861636a"
from torchinfo import summary
summary( model, input_size=(1,),dtypes=[torch.long],)
# + id="cVP8QA7eKHXT" executionInfo={"status": "ok", "timestamp": 1632565713809, "user_tz": -360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
def binary_accuracy(preds, y):
rounded_preds = torch.round(torch.sigmoid(preds))
correct = (rounded_preds == y).float()
acc = correct.sum() / len(correct)
return acc
# + id="-sm8DhuuKIAF" executionInfo={"status": "ok", "timestamp": 1632565713810, "user_tz": -360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
# training function
def train(model, iterator):
epoch_loss = 0
epoch_acc = 0
model.train()
for i,batch in enumerate( iterator,1):
text, text_lengths = batch.text
optimizer.zero_grad()
predictions = model(text,).squeeze(1)
loss = criterion(predictions, batch.polarity)
acc = binary_accuracy(predictions, batch.polarity)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
print_val = f"running_loss : {(loss.item()):.6f}\t"
print_val += f"running_corrects : {acc.item():.6f}\t"
sys.stdout.write('\r' + str(print_val))
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + id="sqx1YxbhKIUS" executionInfo={"status": "ok", "timestamp": 1632565713810, "user_tz": -360, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
def evaluate(model, iterator):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
text, text_lengths = batch.text
predictions = model(text,).squeeze(1)
loss = criterion(predictions, batch.polarity)
acc = binary_accuracy(predictions, batch.polarity)
epoch_acc += acc.item()
epoch_loss += loss.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# + id="RAvQiQFkKI3j" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632565901034, "user_tz": -360, "elapsed": 187233, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="1ecce775-ed4c-4f59-f44e-cf1fafd36af0"
t = time.time()
loss=[]
acc=[]
val_acc=[]
val_loss=[]
num_epochs = 100
for epoch in range(num_epochs):
train_loss, train_acc = train(model, train_iter)
valid_loss, valid_acc = evaluate(model, valid_iter)
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}% | Val. Acc: {valid_acc*100:.2f}%')
# print(f'\t')
loss.append(train_loss)
acc.append(train_acc)
val_loss.append(valid_loss)
val_acc.append(valid_acc)
print(f'time:{time.time()-t:.3f}')
# + [markdown] id="bo5j5q2H2IZZ"
# # Classification Performance Metrics
# + id="QaNZD97cLleV" executionInfo={"status": "ok", "timestamp": 1632565906892, "user_tz": -360, "elapsed": 433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
# import some importent library or packages
import glob,sys,os
import matplotlib.pyplot as plt
import warnings
import numpy as np
import seaborn as sn
import pandas as pd
import pathlib
import zipfile
import torch
import torch.nn as nn
import torchvision
import torchvision.transforms as transforms
from torchvision import models
from torch.utils.data import Dataset, DataLoader, ConcatDataset
import torch.optim as optim
import time,sys
import copy
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
warnings.filterwarnings('ignore')
# + id="dQJ35QIc2L3Z" executionInfo={"status": "ok", "timestamp": 1632565907748, "user_tz": -360, "elapsed": 36, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
_tranning_loss = loss
_tranning_acc = acc
_validation_loss = val_loss
_validation_acc = val_acc
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="4LZpToji3Jxy" executionInfo={"status": "ok", "timestamp": 1632565907750, "user_tz": -360, "elapsed": 38, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="c0cb631b-5096-4be5-8569-82d8efcd5c3b"
plt.figure(figsize=(10,5))
plt.title("Loss graph")
plt.plot(_tranning_loss,label="Tranning Loss")
plt.plot(_validation_loss,label="Validation Loss")
plt.xlabel("iterations")
plt.ylabel("Loss")
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 350} id="9I3Z4eo13K-T" executionInfo={"status": "ok", "timestamp": 1632565907752, "user_tz": -360, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="441552eb-4b60-45e4-ff99-d82a01a4066b"
plt.figure(figsize=(10,5))
plt.title("Accuracy graph")
plt.plot(_tranning_acc,label="Tranning Accuracy")
plt.plot(_validation_acc,label="Validation Accuracy")
plt.xlabel("iterations")
plt.ylabel("Accuracy")
plt.legend()
plt.show()
# + id="1xjZ8_bBAeMH" executionInfo={"status": "ok", "timestamp": 1632565908212, "user_tz": -360, "elapsed": 493, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
y_true_tensor = torch.tensor([])
y_pred_tensor = torch.tensor([])
model.eval()
with torch.no_grad():
for batch in valid_iter:
text, text_lengths = batch.text
predictions = model(text).squeeze(1)
rounded_preds = torch.round(torch.sigmoid(predictions))
correct = (rounded_preds == batch.polarity).float()
# print(rounded_preds,batch.polarity)
y_true_tensor = torch.cat((y_true_tensor,batch.polarity))
y_pred_tensor = torch.cat((y_pred_tensor,rounded_preds))
# acc = binary_accuracy(predictions, batch.polarity)
# print(acc)
# + id="Uil8uwrOLUz2" executionInfo={"status": "ok", "timestamp": 1632565908214, "user_tz": -360, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}}
y_true = y_true_tensor.type(torch.LongTensor).tolist()
y_pred = y_pred_tensor.type(torch.LongTensor).tolist()
# + colab={"base_uri": "https://localhost:8080/"} id="evonmYksLWOY" executionInfo={"status": "ok", "timestamp": 1632565908215, "user_tz": -360, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="f3139e67-beaf-486d-c6b7-06babf1ef470"
matrice = confusion_matrix(y_true,y_pred)
matrice
# + colab={"base_uri": "https://localhost:8080/", "height": 336} id="QvWx7SNHW2cs" executionInfo={"status": "ok", "timestamp": 1632565908215, "user_tz": -360, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="802f5ae7-d310-4960-dd09-9be0657fa414"
df_cm = pd.DataFrame(matrice,columns=[0,1],index=[0,1])
plt.figure(figsize=(5,5))
sn.heatmap(df_cm, annot=True,annot_kws={"size": 10},fmt='g',cmap='Blues',)
plt.title(f"Confusion matrix")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="U9nF3NHxUzH7" executionInfo={"status": "ok", "timestamp": 1632565908216, "user_tz": -360, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "07570094735222624973"}} outputId="a12b6105-f667-4890-cdd1-92f515a43e0d"
classify_report = classification_report(y_true, y_pred, )
print(classify_report)
| CSE 4238 - Soft Computing Lab - Assignment 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo of PYPIT on GMOS Longslit [v1.1]
# v1.1 -- B600
# +
# import
from importlib import reload
import os
import glob
import numpy as np
from astropy.io import fits
# A few core routines
from pypit.core import arsetup
from pypit.core import arsort
from pypit import arpixels
from pypit.core import arprocimg
from pypit.core import arwave
from pypit.core import arsave
from pypit import arutils
from pypit import arload
# Classes
from pypit import calibrations
from pypit import fluxspec
from pypit import pypitsetup
from pypit import scienceimage
# Spectrgraph and Settings
from pypit.spectrographs.util import load_spectrograph
from pypit.par import pypitpar
# -
# ## To play along, you need the Development suite and the $PYPIT_DEV environmental variable pointed at it
os.getenv('PYPIT_DEV')
# ## Spectrograph + Settings
spectro_name='gemini_gmos_south'
spectrograph = load_spectrograph(spectrograph=spectro_name)
spectrograph
# ### Settings
par = pypitpar.PypitPar()
par.keys()
par['calibrations']['biasframe']['useframe'] = 'overscan'
par['calibrations']['biasframe']['number'] = 0
par['calibrations']['pixelflatframe']['number'] = 1
par['calibrations']['traceframe']['number'] = 1
# ## Build the fitstbl
# ### Files
gemini_gmos_files = glob.glob(os.getenv('PYPIT_DEV')+'RAW_DATA/Gemini_GMOS/B600/S2018*')
gemini_gmos_files.sort()
len(gemini_gmos_files)
gemini_gmos_files
# restricting to 530nm
gemini_gmos_files_530 = gemini_gmos_files[3:7] + [gemini_gmos_files[-5]]
gemini_gmos_files_530
# ### Looking at the Headers
hdul = fits.open(gemini_gmos_files_530[2])
hdul[0].header
hdul.info()
hdul[1].header
# ### PypitSetup
# Init
reload(pypitsetup)
setupc = pypitsetup.PypitSetup(gemini_gmos_files_530, spectrograph_name='gemini_gmos_south', par=par)
fitstbl = setupc.build_fitstbl(gemini_gmos_files_530)
fitstbl
# ## Image type
# Classifies the images
# Adds image type columns to the fitstbl
filetypes = setupc.type_data(flag_unknown=True)
# ### Show
setupc.fitstbl[['filename','arc','bias','pixelflat','science','standard','trace','unknown' ]]
# # KLUDGING
# +
setupc.fitstbl['unknown'] = False
setupc.fitstbl[0:2]['science'] = True
setupc.fitstbl[2]['pixelflat'] = True
setupc.fitstbl[2]['trace'] = True
setupc.fitstbl[3]['arc'] = True
setupc.fitstbl[-1]['standard'] = True
#setupc.fitstbl[-1]['science'] = False
#setupc.fitstbl[5]['pixelflat'] = True
#setupc.fitstbl[5]['trace'] = True
#
'''
setupc.fitstbl[-3]['standard'] = True
setupc.fitstbl[-3]['science'] = False
setupc.fitstbl[-2]['pixelflat'] = True
setupc.fitstbl[-2]['trace'] = True
setupc.fitstbl[-1]['arc'] = True
'''
# -
setupc.fitstbl[['filename','arc','bias','pixelflat','science','standard','trace','unknown' ]]
# ## Match to science
fitstbl = setupc.match_to_science()
# ### Setup dict
setup_dict = setupc.build_setup_dict()
setup_dict
setupc.fitstbl[['filename','arc','bias','pixelflat','science','standard','sci_ID']]
# ----
# ## Setup + datasec
# Image IDs
sci_ID = 1 # First exposure ID
det = 1 #
dnum = 'det01'
# Index in fitstbl
scidx = np.where((fitstbl['sci_ID'] == sci_ID) & fitstbl['science'])[0][0]
scidx
# ### Setup
setup = 'A_01_aa'
# ## Calibrations
reload(calibrations)
caliBrate = calibrations.MultiSlitCalibrations(fitstbl, spectrograph=spectrograph, par=par['calibrations'], save_masters=False, write_qa=False)
caliBrate.reset(setup, 1, sci_ID)#, spectrograph.calib_par)
# ## datasec_img
datasec_img = caliBrate.get_datasec_img()
datasec_img.shape
# ## Bias
caliBrate.par['biasframe']['useframe'] = 'overscan'
bias = caliBrate.get_bias()
bias
# ----
# ## Arc Image frame
arc = caliBrate.get_arc()
caliBrate.show(arc)
# ----
# ## Bad pixel mask
bpm = caliBrate.get_bpm()
np.sum(bpm)
# ----
# ## pixlocn
pixlocn = caliBrate.get_pixlocn()
pixlocn.shape
# ----
# ## Trace slit(s)
caliBrate.msbpm[:,0:37] = 1.
caliBrate.msbpm[:,-20:] = 1.
# +
#caliBrate.spectrograph.calib_par['slits']
# -
caliBrate.par['slits']['sigdetect'] = 300.
caliBrate.par['slits']['pca']['params'] = [1,0]
tslits_dict, maskslits = caliBrate.get_slits()
caliBrate.show(caliBrate.traceSlits.mstrace)
caliBrate.traceSlits.show('siglev')
caliBrate.traceSlits.show('edges')
# ### Mask the uninteresting slits
caliBrate.maskslits[0:2]= True
caliBrate.maskslits[-1]= True
caliBrate.maskslits
# ----
# ## Wavelength Calibration
# +
#caliBrate.show(caliBrate.waveCalib.msarc)
# +
#caliBrate.waveCalib.arccen.shape
# +
#caliBrate.waveCalib.maskslits
# -
caliBrate.get_wv_calib()
caliBrate.wv_calib.keys()
# ----
# ## Wave Tilts
# Settings kludges
tilt_settings = dict(tilts=settings.argflag['trace']['slits']['tilts'].copy(),
masters=settings.argflag['reduce']['masters'])
tilt_settings['tilts']['function'] = settings.argflag['trace']['slits']['function']
# Instantiate
waveTilts = wavetilts.WaveTilts(msarc, settings=tilt_settings,
det=det, setup=setup,
tslits_dict=tslits_dict, settings_det=settings_det,
pixlocn=pixlocn)
# Run
mstilts, wt_maskslits = waveTilts.run(maskslits=maskslits, wv_calib=wv_calib)
waveTilts.show('fweight', slit=0)
waveTilts.show('tilts', slit=0)
# ----
# ## Pixel Flat Field
# Settings
flat_settings = dict(flatfield=settings.argflag['reduce']['flatfield'].copy(),
slitprofile=settings.argflag['reduce']['slitprofile'].copy(),
combine=settings.argflag['pixelflat']['combine'].copy(),
masters=settings.argflag['reduce']['masters'].copy(),
detector=settings.spect[dnum])
# Instantiate
pixflat_image_files = arsort.list_of_files(fitstbl, 'pixelflat', sci_ID)
flatField = flatfield.FlatField(file_list=pixflat_image_files, msbias=msbias,
spectrograph=spectrograph,
settings=flat_settings,
tslits_dict=tslits_dict,
tilts=mstilts, det=det, setup=setup,
datasec_img=datasec_img)
# Run
mspixflatnrm, slitprof = flatField.run(armed=False)
flatField.show('norm')
# ----
# ## Wavelength Image
# Settings
wvimg_settings = dict(masters=settings.argflag['reduce']['masters'].copy())
# Instantiate
waveImage = waveimage.WaveImage(mstilts, wv_calib, settings=wvimg_settings,
setup=setup, maskslits=maskslits,
slitpix=tslits_dict['slitpix'])
# Build
mswave = waveImage._build_wave()
waveImage.show('wave')
# ----
# ## Science Image
# ### File list
sci_image_files = arsort.list_of_files(fitstbl, 'science', sci_ID)
# Settings
sci_settings = tsettings.copy()
# ### Instantiate
# Instantiate
sciI = scienceimage.ScienceImage(file_list=sci_image_files, datasec_img=datasec_img,
bpm=msbpm, det=det, setup=setup, settings=sci_settings,
maskslits=maskslits, pixlocn=pixlocn, tslits_dict=tslits_dict,
tilts=mstilts, fitstbl=fitstbl, scidx=scidx)
# ### Name, time
# Names and time
obstime, basename = sciI.init_time_names(settings.spect['mosaic']['camera'],
timeunit=settings.spect["fits"]["timeunit"])
basename
# ### Process
# Process (includes Variance image and CRs)
dnoise = (settings_det['darkcurr'] * float(fitstbl["exptime"][scidx])/3600.0)
sciframe, rawvarframe, crmask = sciI._process(
msbias, mspixflatnrm, apply_gain=True, dnoise=dnoise)
sciI.show('sci')
# ### Global sky sub
# Global skysub
settings_skysub = {}
settings_skysub['skysub'] = settings.argflag['reduce']['skysub'].copy()
global_sky, modelvarframe = sciI.global_skysub(settings_skysub)
sciI.show('skysub')
# ### Find objects
_, nobj = sciI.find_objects()
# ### Repeat the last 2 steps
# Mask the objects
global_sky, modelvarframe = sciI.global_skysub(settings_skysub, use_tracemask=True)
# Another round of finding objects
_, nobj = sciI.find_objects()
# ### Extraction -- New algorithm in development
specobjs, finalvar, finalsky = sciI.extraction(mswave)
# ### Flexure
flex_list = arwave.flexure_obj(
specobjs, maskslits, settings.argflag['reduce']['flexure']['method'],
spectrograph,
skyspec_fil = settings.argflag['reduce']['flexure']['spectrum'],
mxshft = settings.argflag['reduce']['flexure']['maxshift'])
# QA
arwave.flexure_qa(specobjs, maskslits, basename, det, flex_list)
# ### Heliocentric (optional)
vel, vel_corr = arwave.geomotion_correct(specobjs, maskslits, fitstbl, scidx,
obstime, settings.spect,
settings.argflag['reduce']['calibrate']['refframe'])
sci_dict = {}
sci_dict['meta'] = {}
sci_dict['meta']['vel_corr'] = vel_corr
# ----
# ## Write
# ### 1D spectra
outfile = 'Science/spec1d_{:s}.fits'.format(basename)
helio_dict = dict(refframe=settings.argflag['reduce']['calibrate']['refframe'],
vel_correction=sci_dict['meta']['vel_corr'])
arsave.save_1d_spectra_fits([specobjs], fitstbl[scidx], outfile,
helio_dict=helio_dict, obs_dict=settings.spect['mosaic'])
# ### 2D images
# Write 2D images for the Science Frame
arsave.save_2d_images(
sci_dict, fitstbl, scidx,
settings.spect['fits']['headext{0:02d}'.format(1)], setup,
settings.argflag['run']['directory']['master']+'_'+spectrograph, # MFDIR
'Science/', basename)
# ----
# ## Fluxing (optional)
# ### Reduce a standard star
# +
std_dict = {}
# Reduce standard here; only legit if the mask is the same
std_idx = arsort.ftype_indices(fitstbl, 'standard', sci_ID)[0]
#
std_image_files = arsort.list_of_files(fitstbl, 'standard', sci_ID)
std_dict[std_idx] = {}
# Instantiate for the Standard
stdI = scienceimage.ScienceImage(file_list=std_image_files, datasec_img=datasec_img,
bpm=msbpm, det=det, setup=setup, settings=sci_settings,
maskslits=maskslits, pixlocn=pixlocn, tslits_dict=tslits_dict,
tilts=mstilts, fitstbl=fitstbl, scidx=std_idx,
objtype='standard')
# Names and time
_, std_basename = stdI.init_time_names(settings.spect['mosaic']['camera'],
timeunit=settings.spect["fits"]["timeunit"])
# Process (includes Variance image and CRs)
stdframe, _, _ = stdI._process(msbias, mspixflatnrm, apply_gain=True, dnoise=dnoise)
# Sky
_ = stdI.global_skysub(settings_skysub)
# Find objects
_, nobj = stdI.find_objects()
_ = stdI.global_skysub(settings_skysub, use_tracemask=True)
# Extract
stdobjs, _, _ = stdI.extraction(mswave)
# Save for fluxing and output later
std_dict[std_idx][det] = {}
std_dict[std_idx][det]['basename'] = std_basename
std_dict[std_idx][det]['specobjs'] = arutils.unravel_specobjs([stdobjs])
# -
# ### Sensitivity function
# Settings
fsettings = settings.spect.copy()
fsettings['run'] = settings.argflag['run']
fsettings['reduce'] = settings.argflag['reduce']
# Build the list of stdobjs
reload(fluxspec)
all_std_objs = []
for det in std_dict[std_idx].keys():
all_std_objs += std_dict[std_idx][det]['specobjs']
FxSpec = fluxspec.FluxSpec(settings=fsettings, std_specobjs=all_std_objs,
setup=setup) # This takes the last setup run, which is as sensible as any..
sensfunc = FxSpec.master(fitstbl[std_idx], save=False)
all_std_objs
# Show
FxSpec.show_sensfunc()
# ### Flux
# Load
sci_specobjs, sci_header = arload.load_specobj('Science/spec1d_OFF_J1044p6306_LRISr_2016Feb16T112439.fits')
#
FxSpec.sci_specobjs = sci_specobjs
FxSpec.sci_header = sci_header
# Flux
FxSpec.flux_science()
# Write
FxSpec.write_science('Science/spec1d_OFF_J1044p6306_LRISr_2016Feb16T112439.fits')
| doc/demos/GMOS_longslit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <a href="https://www.bigdatauniversity.com"><img src = "https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width = 400, align = "center"></a>
#
# <h1 align=center><font size = 5> Classification with Python</font></h1>
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In this notebook we try to practice all the classification algorithms that we learned in this course.
#
# We load a dataset using Pandas library, and apply the following algorithms, and find the best one for this specific dataset by accuracy evaluation methods.
#
# Lets first load required libraries:
# + button=false new_sheet=false run_control={"read_only": false}
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
# %matplotlib inline
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### About dataset
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# This dataset is about past loans. The __Loan_train.csv__ data set includes details of 346 customers whose loan are already paid off or defaulted. It includes following fields:
#
# | Field | Description |
# |----------------|---------------------------------------------------------------------------------------|
# | Loan_status | Whether a loan is paid off on in collection |
# | Principal | Basic principal loan amount at the |
# | Terms | Origination terms which can be weekly (7 days), biweekly, and monthly payoff schedule |
# | Effective_date | When the loan got originated and took effects |
# | Due_date | Since it’s one-time payoff schedule, each loan has one single due date |
# | Age | Age of applicant |
# | Education | Education of applicant |
# | Gender | The gender of applicant |
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets download the dataset
# + button=false new_sheet=false run_control={"read_only": false}
# !wget -O loan_train.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/loan_train.csv
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Data From CSV File
# + button=false new_sheet=false run_control={"read_only": false}
df = pd.read_csv('loan_train.csv')
df.head()
# -
df.shape
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Convert to date time object
# + button=false new_sheet=false run_control={"read_only": false}
df['due_date'] = pd.to_datetime(df['due_date'])
df['effective_date'] = pd.to_datetime(df['effective_date'])
df.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Data visualization and pre-processing
#
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Let’s see how many of each class is in our data set
# + button=false new_sheet=false run_control={"read_only": false}
df['loan_status'].value_counts()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# 260 people have paid off the loan on time while 86 have gone into collection
#
# -
# Lets plot some columns to underestand data better:
# notice: installing seaborn might takes a few minutes
# !conda install -c anaconda seaborn -y
# +
import seaborn as sns
bins = np.linspace(df.Principal.min(), df.Principal.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'Principal', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + button=false new_sheet=false run_control={"read_only": false}
bins = np.linspace(df.age.min(), df.age.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'age', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Pre-processing: Feature selection/extraction
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Lets look at the day of the week people get the loan
# + button=false new_sheet=false run_control={"read_only": false}
df['dayofweek'] = df['effective_date'].dt.dayofweek
bins = np.linspace(df.dayofweek.min(), df.dayofweek.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'dayofweek', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We see that people who get the loan at the end of the week dont pay it off, so lets use Feature binarization to set a threshold values less then day 4
# + button=false new_sheet=false run_control={"read_only": false}
df['weekend'] = df['dayofweek'].apply(lambda x: 1 if (x>3) else 0)
df.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Convert Categorical features to numerical values
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets look at gender:
# + button=false new_sheet=false run_control={"read_only": false}
df.groupby(['Gender'])['loan_status'].value_counts(normalize=True)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# 86 % of female pay there loans while only 73 % of males pay there loan
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets convert male to 0 and female to 1:
#
# + button=false new_sheet=false run_control={"read_only": false}
df['Gender'].replace(to_replace=['male','female'], value=[0,1],inplace=True)
df.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## One Hot Encoding
# #### How about education?
# + button=false new_sheet=false run_control={"read_only": false}
df.groupby(['education'])['loan_status'].value_counts(normalize=True)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Feature befor One Hot Encoding
# + button=false new_sheet=false run_control={"read_only": false}
df[['Principal','terms','age','Gender','education']].head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Use one hot encoding technique to conver categorical varables to binary variables and append them to the feature Data Frame
# + button=false new_sheet=false run_control={"read_only": false}
Feature = df[['Principal','terms','age','Gender','weekend']]
Feature = pd.concat([Feature,pd.get_dummies(df['education'])], axis=1)
Feature.drop(['Master or Above'], axis = 1,inplace=True)
Feature.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Feature selection
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets defind feature sets, X:
# + button=false new_sheet=false run_control={"read_only": false}
X = Feature
X[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# What are our lables?
# + button=false new_sheet=false run_control={"read_only": false}
y = df['loan_status'].values
y[0:5]
# -
# ### Try to understand the corelation of loan_status and the selected features
merge = pd.concat([X, df['loan_status']], axis=1, sort=False)
merge.head()
merge.corr(method='pearson')
# ### Split dataset for test and train
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=4)
print("X_train size is ", X_train.shape, "\n", "X_test size is ", X_test.shape, "\n",
"y_train size is ", y_train.shape, "\n", "y_test size is ", y_test.shape)
print(X_train[0:5])
y_train[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Normalize Data
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Data Standardization give data zero mean and unit variance (technically should be done after train test split )
# + button=false new_sheet=false run_control={"read_only": false}
X = preprocessing.StandardScaler().fit(X).transform(X.astype(float))
X[0:5]
# -
# also need to normalize the test and train dataset
X_train = preprocessing.StandardScaler().fit(X_train).transform(X_train.astype(float))
X_train[0:5]
X_test = preprocessing.StandardScaler().fit(X_test).transform(X_test.astype(float))
X_test[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Classification
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now, it is your turn, use the training set to build an accurate model. Then use the test set to report the accuracy of the model
# You should use the following algorithm:
# - K Nearest Neighbor(KNN)
# - Decision Tree
# - Support Vector Machine
# - Logistic Regression
#
#
#
# __ Notice:__
# - You can go above and change the pre-processing, feature selection, feature-extraction, and so on, to make a better model.
# - You should use either scikit-learn, Scipy or Numpy libraries for developing the classification algorithms.
# - You should include the code of the algorithm in the following cells.
# -
# # K Nearest Neighbor(KNN)
# Notice: You should find the best k to build the model with the best accuracy.
# **warning:** You should not use the __loan_test.csv__ for finding the best k, however, you can split your train_loan.csv into train and test to find the best __k__.
# ### k-Nearest Neighbors test - find the best k value
# +
# finding a suitable k value
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import jaccard_similarity_score
import matplotlib.pyplot as plt
# %matplotlib inline
k_range = range(1, 10)
accuracy_score = []
for k in k_range:
KNN = KNeighborsClassifier(n_neighbors = k).fit(X_train, y_train)
# perform the test
knn_yhat = KNN.predict(X_test)
print("Test set Accuracy at k=", k, ": ", jaccard_similarity_score(y_test, knn_yhat))
accuracy_score.append(jaccard_similarity_score(y_test, knn_yhat))
# plot the relationship between K and testing accuracy
plt.plot(k_range, accuracy_score)
plt.xlabel('Value of K for KNN')
plt.ylabel('Testing Accuracy')
# -
# #### The result shows that the best accuracy came from k = 7
# ### Perform k-Nearest Neighbors test using k = 7
# for KNN
from sklearn.neighbors import KNeighborsClassifier
# perform the test
KNN = KNeighborsClassifier(n_neighbors = 7).fit(X_train, y_train)
KNN
# # Decision Tree
# ### Decision Trees test - find the best Depth
# +
# findinng the best depth level
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import f1_score
from sklearn.metrics import jaccard_similarity_score
# Compare accuracy result for depth = 3, 4 and 5
d_range = range(3, 6)
f1 = []
ja = []
for d in d_range:
DT = DecisionTreeClassifier(criterion="entropy", max_depth=d)
DT.fit(X_train, y_train)
dt_yhat = DT.predict(X_test)
f1.append(f1_score(y_test, dt_yhat, average='weighted'))
ja.append(jaccard_similarity_score(y_test, dt_yhat))
result = pd.DataFrame(f1, index=['d=3','d=4', 'd=5'])
result.columns = ['F1-score']
result.insert(loc=1, column='Jacard', value=ja)
result.columns.name = "Depth"
result
# -
# #### The result shows that using Depth=5 will give a higer accuracy
# ### Perform Decision Trees using Depth = 5
# for Decision Trees
from sklearn.tree import DecisionTreeClassifier
# prepare DT setting
DT = DecisionTreeClassifier(criterion="entropy", max_depth=5)
# perform the test
DT.fit(X_train, y_train)
DT
# # Support Vector Machine
# ### Support Vector Machines test - find the best kernel function
# +
# for SVM
from sklearn import svm
from sklearn.metrics import jaccard_similarity_score
from sklearn.metrics import f1_score
# import Matplotlib (scientific plotting library)
import matplotlib.pyplot as plt
# %matplotlib inline
func_list = ['linear', 'poly', 'rbf', 'sigmoid']
accuracy_score = []
for func in func_list:
SVM = svm.SVC(kernel=func)
SVM.fit(X_train, y_train)
svm_yhat = SVM.predict(X_test)
accuracy_score.append(f1_score(y_test, svm_yhat, average='weighted'))
# plot the comparison among 4 kernel functions
import numpy as np
import matplotlib.pyplot as plt
y_pos = np.arange(len(func_list))
plt.bar(y_pos, accuracy_score, align='center', alpha=0.5)
plt.xticks(y_pos, func_list)
plt.ylabel('Accuracy')
plt.xlabel('Kernel Functions')
plt.title('Accuracy Comparison for 4 Kernal Functions')
plt.show()
# -
# #### The found best kernel function is rbf
# ### Perform Support Vector Machines using rbf kernel function
# for SVM
from sklearn import svm
# prepare SVM setting
SVM = svm.SVC(kernel='rbf')
# perform the test
SVM.fit(X_train, y_train)
SVM
# # Logistic Regression
# ### Logistic Regression test - find the best parameters
# +
# for Logistic Regression
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import log_loss
# import Matplotlib (scientific plotting library)
import matplotlib.pyplot as plt
# %matplotlib inline
c_list = [0.1, 0.01, 0.001]
solver_list = ['newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga']
idx = []
accuracy_score = []
for idx1, c in enumerate(c_list):
for idx2, sol in enumerate(solver_list):
idx.append(idx2 + idx1 * 5)
# perform the test
LR = LogisticRegression(C=c, solver=sol).fit(X_train, y_train)
# it can predict the outcome
lr_yhat = LR.predict(X_test)
lr_prob = LR.predict_proba(X_test)
print("Test ", (idx2 + idx1 * 5), ": Accuracy at c =", c,"solver=", sol,
"is : ", log_loss(y_test, lr_prob))
accuracy_score.append(log_loss(y_test, lr_prob))
lr_prob = LR.predict_proba(X_test)
log_loss(y_test, lr_prob)
# plot the relationship between K and testing accuracy
plt.plot(idx, accuracy_score)
plt.xlabel('Parameter value')
plt.ylabel('Testing Accuracy')
# -
# #### The result shows that using c=0.001 and solver=liblinear gives the highest accuracy
# ### Perform Logistic Regression test using c=0.001 and solver=liblinear
# for Logistic Regression
from sklearn.linear_model import LogisticRegression
# prepare LR setting
LR = LogisticRegression(C=0.001, solver='liblinear').fit(X_train, y_train)
LR
# # Model Evaluation using Test set
# ### First, download and load the test set:
# !wget -O loan_test.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/loan_test.csv
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Test set for evaluation
# + button=false new_sheet=false run_control={"read_only": false}
test_df = pd.read_csv('loan_test.csv')
# convert date time
test_df['due_date'] = pd.to_datetime(test_df['due_date'])
test_df['effective_date'] = pd.to_datetime(test_df['effective_date'])
test_df['dayofweek'] = test_df['effective_date'].dt.dayofweek
# evaulate weekend field
test_df['weekend'] = test_df['dayofweek'].apply(lambda x: 1 if (x>3) else 0)
# convert male to 0 and female to 1
test_df['Gender'].replace(to_replace=['male','female'], value=[0,1],inplace=True)
# work out education level
test_feature = test_df[['Principal','terms','age','Gender','weekend']]
test_feature = pd.concat([test_feature,pd.get_dummies(test_df['education'])], axis=1)
test_feature.drop(['Master or Above'], axis = 1,inplace=True)
# Testing feature
X_loan_test = test_feature
# normalize the test data
X_loan_test = preprocessing.StandardScaler().fit(X_loan_test).transform(X_loan_test)
# and target result
y_loan_test = test_df['loan_status'].values
y_loan_test[0:5]
print (X_loan_test[0:5])
print (X_loan_test.shape)
print (y_loan_test[0:5])
print (y_loan_test.shape)
# -
# # Evaluate Result
#
# Evaulate the result by using 3 diferent algorithms
# ### Jaccard
# +
# Jaccard setup
from sklearn.metrics import jaccard_similarity_score
# evaluate KNN
knn_yhat = KNN.predict(X_loan_test)
jc1 = round(jaccard_similarity_score(y_loan_test, knn_yhat), 2)
# evaluate Decision Trees
dt_yhat = DT.predict(X_loan_test)
jc2 = round(jaccard_similarity_score(y_loan_test, dt_yhat), 2)
#evaluate SVM
svm_yhat = SVM.predict(X_loan_test)
jc3 = round(jaccard_similarity_score(y_loan_test, svm_yhat), 2)
# evaluate Logistic Regression
lr_yhat = LR.predict(X_loan_test)
jc4 = round(jaccard_similarity_score(y_loan_test, lr_yhat), 2)
list_jc = [jc1, jc2, jc3, jc4]
list_jc
# -
# ### F1-score
# +
# F1-score setup
from sklearn.metrics import f1_score
# evaluate KNN
fs1 = round(f1_score(y_loan_test, knn_yhat, average='weighted'), 2)
# evaluate Desision Trees
fs2 = round(f1_score(y_loan_test, dt_yhat, average='weighted'), 2)
# evaluate SVM
fs3 = round(f1_score(y_loan_test, svm_yhat, average='weighted'), 2)
# evaluate Logistic Regression
fs4 = round(f1_score(y_loan_test, lr_yhat, average='weighted'),2 )
list_fs = [fs1, fs2, fs3, fs4]
list_fs
# -
# ### LogLoss
# LogLoss
from sklearn.metrics import log_loss
lr_prob = LR.predict_proba(X_loan_test)
list_ll = ['NA', 'NA', 'NA', round(log_loss(y_loan_test, lr_prob), 2)]
list_ll
# # Report
# You should be able to report the accuracy of the built model using different evaluation metrics:
# +
import pandas as pd
# fomulate the report format
df = pd.DataFrame(list_jc, index=['KNN','Decision Tree','SVM','Logistic Regression'])
df.columns = ['Jaccard']
df.insert(loc=1, column='F1-score', value=list_fs)
df.insert(loc=2, column='LogLoss', value=list_ll)
df.columns.name = 'Algorithm'
df
# -
# | Algorithm | Jaccard | F1-score | LogLoss |
# |--------------------|---------|----------|---------|
# | KNN | ? | ? | NA |
# | Decision Tree | ? | ? | NA |
# | SVM | ? | ? | NA |
# | LogisticRegression | ? | ? | ? |
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Want to learn more?
#
# IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: [SPSS Modeler](http://cocl.us/ML0101EN-SPSSModeler).
#
# Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at [Watson Studio](https://cocl.us/ML0101EN_DSX)
#
#
# <hr>
# Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Thanks for completing this lesson!
#
# Notebook created by: <a href = "https://ca.linkedin.com/in/saeedaghabozorgi"><NAME></a>
| ML -IBM/templ/Felix NB for Machine Learning Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Download GNPS data <br>
# Replace Job ID below with your GNPS job ID:
# ! curl -d "" 'https://gnps.ucsd.edu/ProteoSAFe/DownloadResult?task=b817262cb6114e7295fee4f73b22a3ad&view=download_cytoscape_data' -o GNPS_output_graphML.zip
# ! unzip -d GNPS_output_graphML/ GNPS_output_graphML.zip
import pandas as pd
import os
from pyMolNetEnhancer import *
from networkx import *
# import MS2LDA data, replace the MS2LDA job ID in the link below with your MS2LDA job ID:
motifs = pd.read_csv('http://ms2lda.org/basicviz/get_gnps_summary/907')
motifs.head()
edges = pd.read_csv('GNPS_output_graphML/networking_pairs_results_file_filtered/' + str(os.listdir('GNPS_output_graphML/networking_pairs_results_file_filtered/')[0]), sep = '\t')
edges.head()
# create network data with mapped motifs
motif_network = Mass2Motif_2_Network(edges,motifs,prob = 0.01,overlap = 0.3, top = 5)
motif_network['nodes'].head()
motif_network['edges'].head()
# write network data with mapped motifs to files:
# The edges file can be importet as network into Cytoscape, whereas the nodes file can be imported as table. Select column 'CLUSTERID1' as Source Node, column 'interact' as Interaction Type and 'CLUSTERID2' as Target Node. When importing the nodes file as table, make sure to select the semicolon delimiter under Advanced Options.
motif_network['edges'].to_csv("Mass2Motifs_Edges_FeatureBased.tsv",sep='\t',index=False)
motif_network['nodes'].to_csv("Mass2Motifs_Nodes_FeatureBased.tsv",sep='\t',index=True)
# create graphML file
MG = make_motif_graphml(motif_network['nodes'],motif_network['edges'])
# write graphML file
nx.write_graphml(MG, "Motif_Network_FeatureBased.graphml", infer_numeric_types = True)
# ## map chemical class information
# read in chemical class information (to create this file follow descriptions in ChemicalClasses_2_Network_FeatureBased.ipynb)
final = pd.read_csv('ClassyFireResults_Network.txt', sep = "\t")
final.head()
graphML_classy = make_classyfire_graphml(MG,final)
# write graphML file containing motif and chemical class information
nx.write_graphml(graphML_classy, "Motif_ChemicalClass_Network_FeatureBased.graphml", infer_numeric_types = True)
| Example_notebooks/Mass2Motifs_2_Network_FeatureBased.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os import path
from collections import deque
from ga4stpg.graph import UGraph
from ga4stpg.graph.disjointsets import DisjointSets
from ga4stpg.graph.priorityqueue import PriorityQueue
from ga4stpg.graph.reader import read_problem
from ga4stpg.graph.util import has_cycle, how_many_components
from ga4stpg.util import STEIN_B
from ga4stpg.tree.evaluation import EvaluateTreeGraph
from ga4stpg.tree.generate import GenerateBasedPrimRST
# +
INDEX_SELECT = 4
datasets = path.join('..', '..', 'ppgi-stpg-gpx', 'datasets', 'ORLibrary')
filename = path.join(datasets, STEIN_B[INDEX_SELECT][0])
print(filename)
STPG = read_problem(filename)
# +
from ga4stpg.edgeset import UEdge, EdgeSet
class Segment:
def __init__(self):
self.edges = EdgeSet()
self.cost = 0
self.portals = set()
def __len__(self):
return len(self.edges)
def __str__(self):
return f'Segment <{len(self.edges)}>'
def __iter__(self):
return iter(self.edges)
@property
def bounds(self):
return frozenset(self.portals)
def add(self, v, u):
self.edges.add(UEdge(v, u))
# +
from random import randrange
def f_weight(v, u):
if STPG.graph.has_edge(v, u):
return STPG.graph.weight(v, u)
else:
return 0
def find_segments(graph, common_nodes):
visited = set()
# start = None
# index = randrange(0, len(common_nodes))
# for i, nro in enumerate(common_nodes):
# if i == index:
# start = nro
# break
# assert start is not None
# stack_outter = [start]
stack_outter = list(common_nodes)
result = list()
def search(start, neighbor):
segment = Segment()
segment.portals.add(start)
segment.add(start, neighbor)
segment.cost += f_weight(start, neighbor)
stack_inner = [neighbor]
while stack_inner:
u = stack_inner.pop()
visited.add(u)
if u not in common_nodes:
counter = 0
for w in graph.adjacent_to(u):
if w not in visited:
stack_inner.append(w)
segment.add(u, w)
segment.cost += f_weight(u, w)
counter += 1
if counter == 0:
segment.portals.add(u)
else:
stack_outter.append(u)
segment.portals.add(u)
# end while
return segment
# end search
while stack_outter:
s = stack_outter.pop()
visited.add(s)
for v in graph.adjacent_to(s):
if v not in visited:
seg = search(s, v)
result.append(seg)
return result
# -
def check_portals(portals, disjoint):
'''Verifica se os vértices portais de um segmento
se conectam à mesma partição de vértices comuns.
Faz essa verificação em tempo O(n)
'''
f_check = set()
for p in portals:
if p not in disjoint:
return False
k = disjoint.find(p)
if k in f_check:
return False
f_check.add(k)
return True
# +
edges = [
(1, 3),
(1, 15),
(15, 17),
(3, 5),
(5, 11),
(11, 13),
(5, 7),
(7, 9),
(7, 23),
]
red = UGraph()
for v, u in edges:
red.add_edge(v, u)
# +
edges = [
(1, 17),
(1, 14),
(14, 18),
(18, 5),
(5, 23),
(23, 13),
(5, 9)
]
blue = UGraph()
for v, u in edges:
blue.add_edge(v, u)
# -
# ## Crossing two random tree generated by Prim Random Steneir Tree
# +
generator = GenerateBasedPrimRST(STPG)
red = generator()
blue = generator()
# -
red.edges == blue.edges
has_cycle(red), how_many_components(red)
has_cycle(blue), how_many_components(blue)
child = UGraph()
red_only = UGraph()
blue_only = UGraph()
for v, u in red.gen_undirect_edges():
if blue.has_edge(v, u):
child.add_edge(v, u)
else:
red_only.add_edge(v, u)
for v, u in blue.gen_undirect_edges():
if not red.has_edge(v, u):
blue_only.add_edge(v, u)
has_cycle(red_only), how_many_components(red_only)
has_cycle(blue_only), how_many_components(blue_only)
has_cycle(child), how_many_components(child)
#
red_leaves = { v for v in red.vertices if red.degree(v) == 1}
blue_leaves = { v for v in blue.vertices if blue.degree(v) == 1}
# +
common_nodes_red = set(red_only.vertices) & set(blue.vertices)
common_nodes_red
# +
common_nodes_blue = set(blue_only.vertices) & set(red.vertices)
common_nodes_blue
# -
red_segments = find_segments(red_only, common_nodes_red)
blue_segments = find_segments(blue_only, common_nodes_blue)
len(red_segments), len(blue_segments)
# +
queue = PriorityQueue()
for seg in red_segments:
# print(seg, seg.cost)
queue.push(seg.cost, seg)
for seg in blue_segments:
# print(seg, seg.cost)
queue.push(seg.cost, seg)
len(queue)
# +
common_nodes = set(red.vertices) | set(blue.vertices)
# +
dset = DisjointSets()
nodes = set(child.vertices)
nodes.update(common_nodes)
for v in nodes:
dset.make_set(v)
for v, u in child.gen_undirect_edges():
dset.union(v, u)
# -
len(dset)
dset.get_disjoint_sets()
while queue:
seg = queue.pop()
if check_portals(seg.portals, dset):
# print('adding segment', seg)
print(seg)
for edge in seg:
print(edge, end=" ")
v, u = edge
child.add_edge(v, u)
portals = iter(seg.portals)
# Update dset
print("")
p_last = next(portals)
print("Portals: ", p_last, end=" ")
for p in portals:
print(p, end=" ")
dset.union(p_last, p)
p_last = p
print("\n\n")
# else:
# print("####### >>> rejecting: ", seg)
for seg in red_segments:
parent = dict()
for p in seg.portals:
if p in dset:
parent[p] = dset.find(p)
print(parent)
for seg in blue_segments:
parent = dict()
for p in seg.portals:
if p in dset:
parent[p] = dset.find(p)
print(parent)
has_cycle(child)
how_many_components(child)
for seg in blue_segments:
print(seg.portals)
for edge in seg:
print(edge, end=' ')
print("")
for seg in red_segments:
print(seg.portals)
for edge in seg:
print(edge, end=' ')
print("")
| Studies_on_DFS_Partitioning_Segmentation/DFS_identificar_segmentos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter_23
#
# ## 1. Introduction
#
# All in this chapter is a higher level use of LIST.
#
# ## 2. Map
# +
things = [2, 5, 9]
things_1 = []
for thing in things:
things_1.append(thing*2)
print(things_1)
# -
things_2 = map(lambda x: x*2, things)
print(things_2)
print(list(things_2))
# ## 3. Filter
# +
nums = [3, 4, 6, 7, 0, 1]
new_list = []
for num in nums:
if num % 2 == 0:
new_list.append(num)
print(new_list)
# -
new_seq = filter(lambda num: num % 2 == 0, nums)
print(new_seq)
print(list(new_seq))
# ## 4. List Comprehensions
#
# ```
# [<transformer_expression> for <loop_var> in <sequence> if <filtration_expression>]
# ```
# +
things = [2, 5, 9]
yourlist = [value * 2 for value in things]
print(yourlist)
print([x*2 for x in things if x % 2 == 0])
# -
# ## 5. Zip
#
# One more common pattern with lists, besides accumulation, is to step through a pair of lists (or several lists), doing something with all of the first items, then something with all of the second items, and so on.
# +
L1 = [3, 4, 5]
L2 = [1, 2, 3]
L3 = []
for i in range(len(L1)):
L3.append(L1[i] + L2[i])
print(L3)
# -
L1 = [3, 4, 5]
L2 = [1, 2, 3]
L4 = list(zip(L1, L2))
print(L4)
# +
L1 = [3, 4, 5]
L2 = [1, 2, 3]
L3 = []
L4 = list(zip(L1, L2))
for (x1, x2) in L4:
L3.append(x1+x2)
print(L3)
# -
# ## Practices
#
# 2.1 Using map, create a list assigned to the variable `greeting_doubled` that doubles each element in the list `lst`.
# +
lst = [["hi", "bye"], "hello", "goodbye", [9, 2], 4]
greeting_doubled = list(map(lambda x: x*2,lst))
print(greeting_doubled)
# -
# 2.2 Below, we have provided a list of strings called `abbrevs`. Use map to produce a new list called `abbrevs_upper` that contains all the same strings in upper case.
# +
abbrevs = ["usa", "esp", "chn", "jpn", "mex", "can", "rus", "rsa", "jam"]
abbrevs_upper = list(map(lambda x: x.upper(),abbrevs))
print(abbrevs_upper)
# -
# 3.1 Write code to assign to the variable `filter_testing` all the elements in lst_check that have a w in them using filter.
# +
lst_check = ['plums', 'watermelon', 'kiwi', 'strawberries', 'blueberries', 'peaches', 'apples', 'mangos', 'papaya']
filter_testing = list(filter(lambda x: 'w' in x, lst_check))
print(filter_testing)
# -
# 3.2 Using filter, filter `lst` so that it only contains words containing the letter “o”. Assign to variable `lst2`.
# +
lst = ["witch", "halloween", "pumpkin", "cat", "candy", "wagon", "moon"]
lst2 = list(filter(lambda x: 'o' in x, lst))
print(lst2)
# -
# 4.2 The for loop below produces a list of numbers greater than 10. Below the given code, use list comprehension to accomplish the same thing. Assign it the the variable `lst2`.
# +
L = [12, 34, 21, 4, 6, 9, 42]
lst2 = [x for x in L if x>10 ]
print(list2)
# +
tester = {'info': [{"name": "Lauren", 'class standing': 'Junior', 'major': "Information Science"},\
{'name': 'Ayo', 'class standing': "Bachelor's", 'major': 'Information Science'},\
{'name': 'Kathryn', 'class standing': 'Senior', 'major': 'Sociology'},\
{'name': 'Nick', 'class standing': 'Junior', 'major': 'Computer Science'},\
{'name': 'Gladys', 'class standing': 'Sophomore', 'major': 'History'},\
{'name': 'Adam', 'major': 'Violin Performance', 'class standing': 'Senior'}]}
compri = [x['name'] for x in tester['info']]
print(compri)
# -
# 5.1 Below we have provided two lists of numbers, `L1` and `L2`. Using zip and list comprehension, create a new list, `L3`, that sums the two numbers if the number from L1 is greater than 10 and the number from L2 is less than 5. This can be accomplished in one line of code.
# +
L1 = [1, 5, 2, 16, 32, 3, 54, 8, 100]
L2 = [1, 3, 10, 2, 42, 2, 3, 4, 3]
L3 = [x1+x2 for x1,x2 in zip(L1,L2) if x1>10 and x2<5]
print(L3)
# -
# ## Exercise
#
# 1. Write equivalent code using map instead of the manual accumulation below and assign it to the variable test
# +
things = [3, 5, -4, 7]
accum = []
for thing in things:
accum.append(thing+1)
print(accum)
# -
test = list(map(lambda x: x+1, things))
print(test)
# 2. Use manual accumulation to define the lengths function below.
def lengths(strings):
"""lengths takes a list of strings as input and returns a list of numbers that are the lengths
of strings in the input list. Use manual accumulation!"""
# fill in this function's definition to make the test pass.
leng = []
for string in strings:
leng.append(len(string))
return leng
# 3. Now define lengths using map instead.
def lengths(strings):
"""lengths takes a list of strings as input and returns a list of numbers that are the lengths
of strings in the input list. Use map!"""
# fill in this function's definition to make the test pass.
leng = map(lambda x: len(x),strings)
return list(leng)
# 4. Now define lengths using a list comprehension instead.
def lengths(strings):
"""lengths takes a list of strings as input and returns a list of numbers that are the lengths
of strings in the input list. Use a list comprehension!"""
# fill in this function's definition to make the test pass.
return [len(x) for x in strings]
# 5. Write a function called positives_Acc that receives list of numbers as the input (like [3, -1, 5, 7]) and returns a list of only the positive numbers, [3, 5, 7], via manual accumulation.
# +
things = [3, 5, -4, 7]
def positives_Acc(things):
outList = []
for thing in things:
if thing > 0:
outList.append(thing)
return outList
# -
# 6. Write a function called positives_Fil that receives list of things as the input and returns a list of only the positive things, [3, 5, 7], using the filter function.
# +
things = [3, 5, -4, 7]
def positives_Fil(things):
return list(filter(lambda x : x>0, things))
# -
# 7. Write a function called positives_Li_Com that receives list of things as the input and returns a list of only the positive things, [3, 5, 7], using the list comprehension.
# +
things = [3, 5, -4, 7]
def positives_Li_Com(things):
return [x for x in things if x>0]
# -
# 8. Define longwords using manual accumulation.
def longwords(strings):
"""Return a shorter list of strings containing only the strings with more than four characters. Use manual accumulation."""
# write your code here
strLong = []
for string in strings:
if len(string) > 4:
strLong.append(string)
return strLong
# 9.Define longwords using filter
def longwords_Fil(strings):
"""Return a shorter list of strings containing only the strings with more than four characters. Use the filter function."""
# write your code here
return list(filter(lambda x:len(x)>4,strings))
# 10. Define longwords using a list comprehension.
def longwords_Li_Comp(strings):
"""Return a shorter list of strings containing only the strings with more than four characters. Use a list comprehension."""
# write your code here
return [x for x in strings if len(x)>4]
# 11. Write a function called longlengths that returns the lengths of those strings that have at least 4 characters. Try it with a list comprehension.
def longlengths(strings):
return [len(x) for x in strings if len(x)>3]
# 12. Write a function called longlengths that returns the lengths of those strings that have at least 4 characters. Try it using map and filter.
def longlengths(strings):
result = list(map(lambda x: len(x),strings))
result = list(filter(lambda x: x>4,result))
return result
# 13. Write a function that takes a list of numbers and returns the sum of the squares of all the numbers. Try it using an accumulator pattern.
def sumSquares(L):
sum = 0
for num in L:
sum = num**2 + sum
return sum
# 14. Write a function that takes a list of numbers and returns the sum of the squares of all the numbers. Try it using map and sum.
def sumSquares(L):
sqNum = list(map(lambda x: x**2,L))
return sum(sqNum)
# 15. Use the zip function to take the lists below and turn them into a list of tuples, with all the first items in the first tuple, etc.
# +
L1 = [1, 2, 3, 4]
L2 = [4, 3, 2, 3]
L3 = [0, 5, 0, 5]
tups = zip(L1,L2,L3)
# -
# 16. Use zip and map or a list comprehension to make a list consisting the maximum value for each position. For L1, L2, and L3, you would end up with a list [4, 5, 3, 5].
maxs = [max(x) for x in zip(L1,L2,L3)]
# 17. Write code to assign to the variable compri_sample all the values of the key name in the dictionary tester if they are Juniors. Do this using list comprehension.
# +
tester = {'info': [{"name": "Lauren", 'class standing': 'Junior', 'major': "Information Science"},{'name': 'Ayo', 'class standing': "Bachelor's", 'major': 'Information Science'}, {'name': 'Kathryn', 'class standing': 'Senior', 'major': 'Sociology'}, {'name': 'Nick', 'class standing': 'Junior', 'major': 'Computer Science'}, {'name': 'Gladys', 'class standing': 'Sophomore', 'major': 'History'}, {'name': 'Adam', 'major': 'Violin Performance', 'class standing': 'Senior'}]}
compri_sample=[x["name"] for x in tester["info"] if x["class standing"] == 'Junior']
# -
# 18. Challenge The nested for loop given takes in a list of lists and combines the elements into a single list. Do the same thing using a list comprehension for the list L. Assign it to the variable result2.
# +
def onelist(lst):
result = []
for each_list in lst:
for item in each_list:
result.append(item)
return result
L = [["hi", "bye"], ["hello", "goodbye"], ["hola", "adios", "bonjour", "au revoir"]]
result2 = [x for y in L for x in y]
# -
# 19. Challenge: Write code to assign to the variable class_sched all the values of the key important classes. Do this using list comprehension.
# +
tester = {'info': [
{"name": "Lauren", 'class standing': 'Junior', 'major': "Information Science", 'important classes': ['SI 106', 'ENGLISH 125', 'SI 110', 'AMCULT 202']},
{'name': 'Ayo', 'class standing': "Bachelor's", 'major': 'Information Science', "important classes": ['SI 106', 'SI 410', 'PSYCH 111']},
{'name': 'Kathryn', 'class standing': 'Senior', 'major': 'Sociology', 'important classes': ['WOMENSTD 220', 'SOC 101', 'ENS 384']},
{'name': 'Nick', 'class standing': 'Junior', 'major': 'Computer Science', "important classes": ['SOC 101', 'AMCULT 334', 'EECS 281']},
{'name': 'Gladys', 'class standing': 'Sophomore', 'major': 'History', 'important classes': ['ENGLISH 125', 'HIST 259', 'ENGLISH 130']},
{'name': 'Adam', 'major': 'Violin Performance', 'class standing': 'Senior', 'important classes': ['PIANO 101', 'STUDIO 300', 'THEORY 229', 'MUSC 356']}]}
class_sched = [y for x in tester['info'] for y in x['important classes']]
# -
# 20. Challenge: Below, we have provided a list of lists that contain numbers. Using list comprehension, create a new list threes that contains all the numbers from the original list that are divisible by 3. This can be accomplished in one line of code.
# +
nums = [[4, 3, 12, 10], [8, 7, 6], [5, 18, 15, 7, 11], [9, 4], [24, 20, 17], [3, 5]]
threes = [y for x in nums for y in x if y%3==0]
# -
# ## Chapter Assessment
# 1. Write code to assign to the variable map_testing all the elements in lst_check while adding the string “Fruit: ” to the beginning of each element using mapping.
# +
lst_check = ['plums', 'watermelon', 'kiwi', 'strawberries', 'blueberries', 'peaches', 'apples', 'mangos', 'papaya']
map_testing = list(map(lambda x: "Fruit: "+ x, lst_check))
# -
# 2. Below, we have provided a list of strings called countries. Use filter to produce a list called b_countries that only contains the strings from countries that begin with B.
# +
countries = ['Canada', 'Mexico', 'Brazil', 'Chile', 'Denmark', 'Botswana', 'Spain', 'Britain', 'Portugal', 'Russia', 'Thailand', 'Bangladesh', 'Nigeria', 'Argentina', 'Belarus', 'Laos', 'Australia', 'Panama', 'Egypt', 'Morocco', 'Switzerland', 'Belgium']
b_countries = list(filter(lambda x: x[0] is 'B',countries))
# -
# 3. Below, we have provided a list of tuples that contain the names of Game of Thrones characters. Using list comprehension, create a list of strings called first_names that contains only the first names of everyone in the original list.
# +
people = [('Snow', 'Jon'), ('Lannister', 'Cersei'), ('Stark', 'Arya'), ('Stark', 'Robb'), ('Lannister', 'Jamie'), ('Targaryen', 'Daenerys'), ('Stark', 'Sansa'), ('Tyrell', 'Margaery'), ('Stark', 'Eddard'), ('Lannister', 'Tyrion'), ('Baratheon', 'Joffrey'), ('Bolton', 'Ramsey'), ('Baelish', 'Peter')]
first_names =[x[1] for x in people]
# -
# 4. Use list comprehension to create a list called lst2 that doubles each element in the list, lst.
# +
lst = [["hi", "bye"], "hello", "goodbye", [9, 2], 4]
lst2 = [x*2 for x in lst]
# -
# 5. Below, we have provided a list of tuples that contain students’ names and their final grades in PYTHON 101. Using list comprehension, create a new list passed that contains the names of students who passed the class (had a final grade of 70 or greater).
# +
students = [('Tommy', 95), ('Linda', 63), ('Carl', 70), ('Bob', 100), ('Raymond', 50), ('Sue', 75)]
passed = [x[0] for x in students if x[1]>=70]
# -
# 6. Write code using zip and filter so that these lists (l1 and l2) are combined into one big list and assigned to the variable opposites if they are both longer than 3 characters each.
# +
l1 = ['left', 'up', 'front']
l2 = ['right', 'down', 'back']
opposites = filter(lambda x:len(x[0])>3 and len(x[1])>3,zip(l1,l2))
# -
# 7. Below, we have provided a species list and a population list. Use zip to combine these lists into one list of tuples called pop_info. From this list, create a new list called endangered that contains the names of species whose populations are below 2500.
# +
species = ['golden retriever', 'white tailed deer', 'black rhino', 'brown squirrel', 'field mouse', 'orangutan', 'sumatran elephant', 'rainbow trout', 'black bear', 'blue whale', 'water moccasin', 'giant panda', 'green turtle', 'blue jay', 'japanese beetle']
population = [10000, 90000, 1000, 2000000, 500000, 500, 1200, 8000, 12000, 2300, 7500, 100, 1800, 9500, 125000]
pop_info = zip(species,population)
endangered = [x[0] for x in pop_info if x[1]<2500]
| python3/Chapter_23_Map_Filter_List_Comprehensions_and_Zip(Not finished).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading data into machine learning
#
# In this notebook, we will review how to load data before any machine learning takes place.
# ## Generated datasets
# ## Regression
# +
from sklearn.datasets import make_regression
X, y = make_regression()
# -
X
y
# ## Classification
# +
from sklearn.datasets import make_classification
X, y = make_classification()
# -
X
y
# ## Sample Datasets
from sklearn.datasets import fetch_openml
iris = fetch_openml(data_id=61, as_frame=True)
print(iris.DESCR)
X = iris.data
X
y = iris.target
y
import matplotlib.pyplot as plt
plt.scatter(X['sepallength'], X['sepalwidth'], c=y.cat.codes)
iris_df = iris.frame
iris_df.head()
import seaborn as sns
sns.set_theme(font_scale=1.5)
iris_df.columns
sns.relplot(data=iris_df, x='sepallength', y='sepalwidth',
hue='class', height=6);
sns.displot(data=iris_df, x='sepallength', hue='class', kind='kde', aspect=2);
sns.jointplot(data=iris_df, x="sepallength", y="sepalwidth", height=10, hue='class');
# # Exercise 1
#
# 1. Load the wine dataset from the `sklearn.datasets` module using the `load_wine` function.
# 2. Print the description of the dataset.
# 3. What is the number of classes and features in this dataset?
# 4. Is this a classifiation of a regression problem? Hint: The target column is called `target`.
# 5. Use `sns.jointplot` to explore the relationship between the `alcohol` and `hue` features.
# +
# # %load solutions/01-ex1-solution.py
| notebooks/01-loading-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <h1> Time series prediction, end-to-end </h1>
#
# This notebook illustrates several models to find the next value of a time-series:
# <ol>
# <li> DNN
# <li> CNN
# <li> LSTM
# </ol>
#
# <p>
# <b>Note:</b>
# See [(Time series prediction with RNNs and TensorFlow)](../05_artandscience/d_customestimator.ipynb) for a very similar example, except that it works with multiple short sequences.
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
SEQ_LEN = 50
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['SEQ_LEN'] = str(SEQ_LEN)
# %%datalab project set -p $PROJECT
# <h3> Simulate some time-series data </h3>
#
# Essentially a set of sinusoids with random amplitudes and frequencies.
import tensorflow as tf
print tf.__version__
# +
import numpy as np
import seaborn as sns
def create_time_series():
freq = (np.random.random()*0.5) + 0.1 # 0.1 to 0.6
ampl = np.random.random() + 0.5 # 0.5 to 1.5
noise = [np.random.random()*0.3 for i in xrange(SEQ_LEN)] # -0.3 to +0.3 uniformly distributed
x = np.sin(np.arange(0,SEQ_LEN) * freq) * ampl + noise
return x
flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
for i in xrange(0, 5):
sns.tsplot( create_time_series(), color=flatui[i%len(flatui)] ); # 5 series
# +
def to_csv(filename, N):
with open(filename, 'w') as ofp:
for lineno in xrange(0, N):
seq = create_time_series()
line = ",".join(map(str, seq))
ofp.write(line + '\n')
import os
try:
os.makedirs('data/sines/')
except OSError:
pass
to_csv('data/sines/train-1.csv', 1000) # 1000 sequences
to_csv('data/sines/valid-1.csv', 250)
# -
# !head -5 data/sines/*-1.csv
# <h3> Train model locally </h3>
#
# Make sure the code works as intended.
# %bash
# run module as-is
DATADIR=$(pwd)/data/sines
OUTDIR=$(pwd)/trained/sines
# echo $DATADIR $OUTDIR
# rm -rf $OUTDIR
export PYTHONPATH=${PYTHONPATH}:${PWD}/sinemodel
python -m trainer.task \
--train_data_paths="${DATADIR}/train-1.csv" \
--eval_data_paths="${DATADIR}/valid-1.csv" \
--output_dir=${OUTDIR} \
--job-dir=./tmp \
--model=lstmN --train_steps=10 --sequence_length=$SEQ_LEN
# Try out online prediction. This is how the REST API will work after you train on Cloud ML Engine
# %writefile data/sines/test.json
{"rawdata": [0.0748873916376,0.447740615977,0.717589472426,0.858772978179,0.608724336456,0.302446739938,0.0280110368029,-0.531982131283,-0.468104981652,-0.494012545404,-0.126079933123,0.199709971324,0.553663376401,0.778562711683,0.767834294938,0.671534068434,0.392766129638,-0.219178436795,-0.377270233974,-0.617684784386,-0.557375138859,-0.13441895655,0.179344392905,0.694238738373,0.742613945325,0.802871888808,0.527062793736,0.315789725344,-0.25938141901,-0.439878866953,-0.715512272927,-0.484315619215,-0.0648125337349,0.334649573669,0.746666758013,0.70962911156,0.743632509136,0.567667711174,0.370128693895,-0.188000053146,-0.593534425513,-0.717821580268,-0.430561591089,-0.0302571755165,0.301924685353,0.505925169202,0.911040529317,0.881595070159,0.565253317297]}
# %bash
OUTDIR=$(pwd)/trained/sines
MODEL_DIR=$OUTDIR/export/Servo/$(ls $OUTDIR/export/Servo/)
# echo $MODEL_DIR
gcloud ml-engine local predict --model-dir=$MODEL_DIR --json-instances=data/sines/test.json
# <h3> Cloud ML Engine </h3>
#
# Now to train on Cloud ML Engine with more data.
import shutil
shutil.rmtree('data/sines', ignore_errors=True)
os.makedirs('data/sines/')
for i in xrange(0,10):
to_csv('data/sines/train-{}.csv'.format(i), 1000) # 1000 sequences
to_csv('data/sines/valid-{}.csv'.format(i), 250)
# %bash
gsutil -m rm -rf gs://${BUCKET}/sines/*
gsutil -m cp data/sines/*.csv gs://${BUCKET}/sines
# %bash
for MODEL in cnn dnn lstm lstm2 lstmN; do
OUTDIR=gs://${BUCKET}/sinewaves/$MODEL
JOBNAME=sines_${MODEL}_$(date -u +%y%m%d_%H%M%S)
REGION=us-central1
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=${PWD}/sinemodel/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=BASIC_GPU \
--runtime-version=1.2 \
-- \
--train_data_paths="gs://${BUCKET}/sines/train*.csv" \
--eval_data_paths="gs://${BUCKET}/sines/valid*.csv" \
--output_dir=$OUTDIR \
--train_steps=10000 --sequence_length=$SEQ_LEN --model=$MODEL
done
# ## Results
#
# When I ran it, these were the RMSEs that I got for different models:
#
# | Model | Sequence length | # of steps | Minutes | RMSE |
# | --- | ----| --- | --- | --- |
# | dnn | 50 | 10000 | 17 min | 0.111 |
# | cnn | 50 | 10000 | 17 min | 0.098 |
# | lstm | 50 | 10000 | 22 min | 0.134 |
# | lstm2 | 50 | 10000 | 27 min |0.103 |
# | lstmN | 50 | 10000 | 29 min | 0.091 |
#
# Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/09_sequence/sinewaves.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: growth
# language: python
# name: growth
# ---
# # Growth Tutorial Overview
#
# This barebones tutorial will walk you through the steps needed to perform a single tissue growth simulation, visualize the resultant synthetic cell culture, and generate synthetic fluorescence measurement data.
#
# **<font color='red'>Please note that documentation for this project is very sparse as this package is not intended for open-source distribution. We are simply making the code available so other researchers may reproduce the results published in our Fly-QMA manuscript.</font>**
# ### Running a growth simulation
# +
from growth import Culture
# define a target culture size
num_cells = 100
# initialize a synthetic cell culture
culture = Culture(reference_population=num_cells)
# run the growth simulation
culture.grow(min_population=num_cells, division_rate=0.1, recombination_rate=0.1)
# -
# ### Visualizing a synthetic cell culture
# visualize the synthetic cell culture
culture.plot(colorby='genotype', s=25)
# +
# %%capture
# animate the entire simulation
video = culture.animate(interval=200, colorby='genotype', s=25, repeat_delay=5000)
# +
from IPython.core.display import HTML
HTML(video)
# -
# ### Generating synthetic fluorescence measurements
# generate synthetic fluorescence measurements
measurements = culture.measure(ambiguity=0.1)
measurements.head()
# +
import matplotlib.pyplot as plt
# %matplotlib inline
fig, ax = plt.subplots(figsize=(3, 2))
_ = ax.hist(measurements.clonal_marker, bins=10)
_ = ax.set_xlabel('Clonal Marker Fluorescence')
| tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deduplication Example
# ## Boilerplate
# %load_ext autoreload
# %autoreload 2
from importlib import reload
import logging
reload(logging)
logging.basicConfig(format='%(asctime)s %(levelname)s:%(message)s', level=logging.INFO, datefmt='%H:%M:%S')
# +
import sys
sys.path.insert(0, '..')
# -
import entity_embed
# +
import torch
import numpy as np
random_seed = 42
torch.manual_seed(random_seed)
np.random.seed(random_seed)
# -
# ## Load Dataset
# We'll use the [Music Brainz 20K from Database Group Leipzig](https://dbs.uni-leipzig.de/research/projects/object_matching/benchmark_datasets_for_entity_resolution). From the site: "The Music Brainz dataset is based on real records about songs from the MusicBrainz database but uses the DAPO data generator to create duplicates with modified attribute values. The generated dataset consists of five sources and contains duplicates for 50% of the original records in two to five sources. All duplicates are generated with a high degree of corruption to stress-test the ER and clustering approaches."
#
# Here is it's [README](https://www.informatik.uni-leipzig.de/~saeedi/musicBrainz_readme.txt):
# ```
# 5 sources
# ----------
# TID: a unique record's id (in the complete dataset).
# CID: cluster id (records having the same CID are duplicate)
# CTID: a unique id within a cluster (if two records belong to the same cluster they will have the same CID but different CTIDs). These ids (CTID) start with 1 and grow until cluster size.
# SourceID: identifies to which source a record belongs (there are five sources). The sources are deduplicated.
# Id: the original id from the source. Each source has its own Id-Format. Uniqueness is not guaranteed!! (can be ignored).
# number: track or song number in the album.
# length: the length of the track.
# artist: the interpreter (artist or band) of the track.
# year: date of publication.
# language: language of the track.
# ```
# Let's download the CSV dataset to a temporary directory:
# +
import urllib
import tempfile
dataset_url = 'https://www.informatik.uni-leipzig.de/~saeedi/musicbrainz-20-A01.csv.dapo'
tf = tempfile.NamedTemporaryFile(mode='r', delete=False)
tf.close()
urllib.request.urlretrieve(dataset_url, tf.name);
# -
# Now we must read the CSV dataset into a `dict` called `record_dict`.
#
# `record_dict` will contain all records from the dataset, and each record will have the indication of the true cluster it belongs to in the field `CID`.
#
# So `CID` is our `cluster_field`. Entity Embed needs that to train, validate, and test.
#
# We'll dynamically attribute an `id` to each record using `enumerate`. Entity Embed needs that too.
# +
import csv
record_dict = {}
cluster_field = 'CID'
with open(tf.name, newline='') as f:
for current_record_id, record in enumerate(csv.DictReader(f)):
record['id'] = current_record_id
record[cluster_field] = int(record[cluster_field]) # convert cluster_field to int
record_dict[current_record_id] = record
# -
# Here's an example of a record:
record_dict[83]
# That's a great song, but it's actually called "Berimbau", not "Berimbou"! And it's a Brazilian song, in Portuguese. This a small example on how noisy is this dataset...
# How many clusters this dataset has?
cluster_total = len(set(record[cluster_field] for record in record_dict.values()))
cluster_total
# From all clusters, we'll use only 20% for training, and other 20% for validation to test how well we can generalize:
# +
from entity_embed.data_utils import utils
train_record_dict, valid_record_dict, test_record_dict = utils.split_record_dict_on_clusters(
record_dict=record_dict,
cluster_field=cluster_field,
train_proportion=0.2,
valid_proportion=0.2,
random_seed=random_seed)
# -
# Note we're splitting the data on **clusters**, not records, so the record counts vary:
len(train_record_dict), len(valid_record_dict), len(test_record_dict)
# Clean up the temporary files:
# +
import os
os.remove(tf.name)
# -
# ## Preprocess
# We'll perform a very minimal preprocessing of the dataset. We want to simply force ASCII chars, lowercase all chars, and strip leading and trailing whitespace.
#
# The fields we'll clean are the ones we'll use:
field_list = ['number', 'title', 'artist', 'album', 'year', 'language']
# +
import unidecode
def clean_str(s):
return unidecode.unidecode(s).lower().strip()
for record in record_dict.values():
for field in field_list:
record[field] = clean_str(record[field])
# -
utils.subdict(record_dict[83], field_list)
# Forcing ASCII chars in this dataset is useful to improve recall because there's little difference between accented and not-accented chars here. Also, this dataset contains mostly latin chars.
# ## Configure Entity Embed fields
# Now we will define how record fields will be numericalized and encoded by the neural network. First we set an `alphabet`, here we'll use ASCII numbers, letters, symbols and space:
# +
from entity_embed.data_utils.field_config_parser import DEFAULT_ALPHABET
alphabet = DEFAULT_ALPHABET
''.join(alphabet)
# -
# It's worth noting you can use any alphabet you need, so the accent removal we performed is optional.
# Then we set an `field_config_dict`. It defines `field_type`s that determine how fields are processed in the neural network:
field_config_dict = {
'number': {
'field_type': "STRING",
'alphabet': alphabet,
'max_str_len': None, # compute
},
'title': {
'field_type': "MULTITOKEN",
'tokenizer': "entity_embed.default_tokenizer",
'alphabet': alphabet,
'max_str_len': None, # compute
},
'title_semantic': {
'key': 'title',
'field_type': "SEMANTIC_MULTITOKEN",
'tokenizer': "entity_embed.default_tokenizer",
'vocab': "fasttext.en.300d",
},
'artist': {
'field_type': "MULTITOKEN",
'tokenizer': "entity_embed.default_tokenizer",
'alphabet': alphabet,
'max_str_len': None, # compute
},
'album': {
'field_type': "MULTITOKEN",
'tokenizer': "entity_embed.default_tokenizer",
'alphabet': alphabet,
'max_str_len': None, # compute
},
'album_semantic': {
'key': 'album',
'field_type': "SEMANTIC_MULTITOKEN",
'tokenizer': "entity_embed.default_tokenizer",
'vocab': "fasttext.en.300d",
},
'year': {
'field_type': "STRING",
'alphabet': alphabet,
'max_str_len': None, # compute
},
'language': {
'field_type': "STRING",
'alphabet': alphabet,
'max_str_len': None, # compute
},
}
# Then we use our `field_config_dict` to get a `record_numericalizer`. This object will convert the strings from our records into tensors for the neural network.
#
# The same `record_numericalizer` must be used on ALL data: train, valid, test. This ensures numericalization will be consistent. Therefore, we pass `record_list=record_dict.values()`:
# +
from entity_embed import FieldConfigDictParser
record_numericalizer = FieldConfigDictParser.from_dict(field_config_dict, record_list=record_dict.values())
# -
# ## Initialize Data Module
# under the hood, Entity Embed uses [pytorch-lightning](https://pytorch-lightning.readthedocs.io/en/latest/), so we need to create a datamodule object:
# +
from entity_embed import DeduplicationDataModule
batch_size = 32
eval_batch_size = 64
datamodule = DeduplicationDataModule(
train_record_dict=train_record_dict,
valid_record_dict=valid_record_dict,
test_record_dict=test_record_dict,
cluster_field=cluster_field,
record_numericalizer=record_numericalizer,
batch_size=batch_size,
eval_batch_size=eval_batch_size,
random_seed=random_seed
)
# -
# We've used `DeduplicationDataModule` because we're doing Deduplication of a single dataset/table (a.k.a. Entity Clustering, Entity Resolution, etc.).
#
# We're NOT doing Record Linkage of two datasets here. Check the other notebook [Record-Linkage-Example](./Record-Linkage-Example.ipynb) if you want to learn how to do it with Entity Embed.
# ## Training
# Now the training process! Thanks to pytorch-lightning, it's easy to train, validate, and test with the same datamodule.
#
# We must choose the K of the Approximate Nearest Neighbors, i.e., the top K neighbors our model will use to find duplicates in the embedding space. Below we're setting it on `ann_k` and initializing the `EntityEmbed` model object:
# +
from entity_embed import EntityEmbed
ann_k = 100
model = EntityEmbed(
record_numericalizer,
ann_k=ann_k,
)
# -
# To train, Entity Embed uses [pytorch-lightning Trainer](https://pytorch-lightning.readthedocs.io/en/latest/common/trainer.html) on it's `EntityEmbed.fit` method.
#
# Since Entity Embed is focused in recall, we'll use `valid_recall_at_0.3` for early stopping. But we'll set `min_epochs = 5` to avoid a very low precision.
#
# `0.3` here is the threshold for **cosine similarity of embedding vectors**, so possible values are between -1 and 1. We're using a validation metric, and the training process will run validation on every epoch end due to `check_val_every_n_epoch=1`.
#
# We also set `tb_name` and `tb_save_dir` to use Tensorboard. Run `tensorboard --logdir notebooks/tb_logs` to check the train and valid metrics during and after training.
trainer = model.fit(
datamodule,
min_epochs=5,
max_epochs=100,
check_val_every_n_epoch=1,
early_stop_monitor="valid_recall_at_0.3",
tb_save_dir='tb_logs',
tb_name='music',
)
# `EntityEmbed.fit` keeps only the weights of the best validation model. With them, we can check the best performance on validation set:
model.validate(datamodule)
# And we can check which fields are most important for the final embedding:
model.get_pool_weights()
# ## Testing
# Again with the best validation model, we can check the performance on the test set:
model.test(datamodule)
# Entity Embed achieves Recall of ~0.99 with Pair-Entity ratio below 100 on a variety of datasets. **Entity Embed aims for high recall at the expense of precision. Therefore, this library is suited for the Blocking/Indexing stage of an Entity Resolution pipeline.** A scalabale and noise-tolerant Blocking procedure is often the main bottleneck for performance and quality on Entity Resolution pipelines, so this library aims to solve that. Note the ANN search on embedded records returns several candidate pairs that must be filtered to find the best matching pairs, possibly with a pairwise classifier. See the [Record-Linkage-Example](./Record-Linkage-Example.ipynb) for an example of matching.
# ## t-sne visualization
# Let's visualize a small sample of the test embeddings and see if they look properly clustered. First, get the embedding vectors:
test_vector_dict = model.predict(
record_dict=test_record_dict,
batch_size=eval_batch_size
)
# Then, produce the visualization:
vis_sample_size = 10
test_cluster_dict = utils.record_dict_to_cluster_dict(test_record_dict, cluster_field)
vis_cluster_dict = dict(sorted(test_cluster_dict.items(), key=lambda x: len(x[1]), reverse=True)[:vis_sample_size])
vis_x = np.stack([test_vector_dict[id_] for cluster in vis_cluster_dict.values() for id_ in cluster])
vis_y = np.array([cluster_id for cluster_id, cluster in vis_cluster_dict.items() for __ in cluster])
# +
from sklearn.manifold import TSNE
tnse = TSNE(metric='cosine', perplexity=20, square_distances=True, random_state=random_seed)
tsne_results = tnse.fit_transform(vis_x)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import itertools
plt.figure(figsize=(16,10))
ax = sns.scatterplot(
x=tsne_results[:,0],
y=tsne_results[:,1],
hue=vis_y,
palette=sns.color_palette("hls", len(vis_cluster_dict.keys())),
legend="full",
alpha=0.8
)
for id_, (x, y) in zip(itertools.chain.from_iterable(vis_cluster_dict.values()), tsne_results):
# text = id_
text = test_record_dict[id_]['title'][:30]
ax.text(x + 2, y + 2, text)
# -
# ## Testing manually (like a production run)
# When running in production, you only have access to the trained `model` object and the production `record_dict` (without the `cluster_field` filled, of course).
#
# So let's simulate that by removing `cluster_field` from the `test_record_dict`:
# +
import copy
prod_test_record_dict = copy.deepcopy(test_record_dict)
for record in prod_test_record_dict.values():
del record[cluster_field]
# -
# Then call `predict_pairs` with some `ann_k` and `sim_threshold`:
# +
sim_threshold = 0.3
found_pair_set = model.predict_pairs(
record_dict=prod_test_record_dict,
batch_size=eval_batch_size,
ann_k=ann_k,
sim_threshold=sim_threshold
)
len(found_pair_set)
# -
# Let's check now the metrics of the found duplicate pairs:
# +
from entity_embed.evaluation import pair_entity_ratio
pair_entity_ratio(len(found_pair_set), len(prod_test_record_dict))
# +
from entity_embed.evaluation import precision_and_recall
precision_and_recall(found_pair_set, datamodule.test_pos_pair_set)
# -
# Same numbers of the `trainer.test`, so our manual testing is fine.
# Finally, we can check the false positives and negatives to see if they're really difficult:
false_positives = list(found_pair_set - datamodule.test_pos_pair_set)
len(false_positives)
false_negatives = list(datamodule.test_pos_pair_set - found_pair_set)
len(false_negatives)
cos_similarity = lambda a, b: np.dot(a, b)
for (id_left, id_right) in false_positives[:3]:
display(
(
cos_similarity(test_vector_dict[id_left], test_vector_dict[id_right]),
utils.subdict(record_dict[id_left], field_list), utils.subdict(record_dict[id_right], field_list)
)
)
for (id_left, id_right) in false_negatives[:3]:
display(
(
cos_similarity(test_vector_dict[id_left], test_vector_dict[id_right]),
utils.subdict(record_dict[id_left], field_list), utils.subdict(record_dict[id_right], field_list)
)
)
| notebooks/Deduplication-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Considering the Trending Technologies we will choose some of them to see which programming languages are mostly used for them.
import pandas as pd
df = pd.read_csv('Github_Data_2019-2021_ready_for_text_analysis.csv')
focus = ['open source', 'deep learning', 'data science', 'machine learning', 'guided project', 'web development',
'starter project']
for tech in focus:
print('\nFor ' +tech + ' : ', list(set(df[df['clean'].str.contains(tech)]['language'])))
| Programming Languages Used in top technologies/2019-2021/2019-2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import numpy as np
from plotly.offline import plot
import plotly.graph_objs as go
# # load BCG, ECG data
from fs import load_ecg,load_bcg
import ecgpeaks
import autopeaks
from glob import glob
data_dir = "/home/guo/physio/BCG_data/bcg_ecg_data/bcg_ecg_data(500HZ)/"
# ### load bcg
bcgs = glob(data_dir+"*.txt")
bcgs
bcg_file = bcgs[4]
bcg_file
bcg = load_bcg(bcg_file,filter=False,notch=False)
# ### load ecg
edfs = glob(data_dir+"*.edf")
edf = edfs[3]
edf
ecg = load_ecg(edf)
ecg_peak_indices, ecg_peak_values = ecgpeaks.findpeaks_in_ecg(ecg)
# # BCG findpeaks
from findpeaks_clinical import findpeaks_clinical
bcg = bcg[530000:820000] #0
#bcg = bcg[180000:440000] 1
peak_indices, peak_values, intervals = findpeaks_clinical(-bcg)
plot([go.Scatter(y=np.diff(peak_indices),name="Raw interval",mode="lines+markers"),go.Scatter(y=intervals,name="intervals_denosing",mode="lines+markers")])
plot([go.Scatter(y=-bcg)])
# # BCG peaks denosing
| findpeaks_clinical/clinical_bcg_findpeaks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # 三、分组密码
# ## 分组密码的定义
# 将明文切割成若干等长的模块(block),然后对每个模块分别加密。加密的过程是一个迭代过程,每一轮加密使用原始密钥生成的对于子密钥,利用前一轮加密的密文作
# 为原文。典型的分组密码包括3DES(模块64位,密钥168位,迭代48次),AES(模块128位,密钥128位或192位或256位,迭代10次)。
#
# 从加密速度来看,分组密码的速度显著慢于流密码。
#
# 伪随机函数(PRF)根据密钥和明文生成与明文同长的随机密文,也即是普通映射,允许存在多对一关系;伪随机变换(PRP)根据密钥和明文对明文加密,**但密文与明>文存在唯一对应关系**,也即是一一映射,因而存在反函数,可以有效求逆。故PRP是一种特殊的PRF。分组密钥最终是要构造一个PRP。
#
# 一个安全的伪随机函数加密的密文,攻击者无法辨别是来自真随机函数还是伪随机函数。一个安全的伪随机变换加密的密文,攻击者也无法辨别是来自伪随机变换还是伪>随机函数。
#
# 多次使用同一个安全的伪随机函数,可以作为伪随机数生成器使用。并且多次使用这个过程相互独立,因此可以实现并行计算,从而大幅提升效率的。
#
# ## 数据加密标准(Data Encryption Standard,DES)
# DES是基于IBM的Lucifer密码,参考美国国家标准改造的加密方式。模块长度64位,密钥长度56位,1997年已经被穷举搜索破解,从而被AES加密算法替代。
#
# DES算法的核心,是Feistel网络结构。给定D个伪随机函数,两组长度为n的模块。每次迭代过程,左模块变为上次迭代时的右模块,而右模块变为上次迭代时左模块与伪>随机函数加密的右模块的异或,共计迭代D次。
#
# 该网络模拟,可以在任意伪随机函数不可逆的情况下,整体可逆。并且逆算法(解密算法)依然沿用原有的伪随机函数,仅调换次序。对于硬件来说,效率很高。
#
# Feistel网络是分组密码构建可逆函数的主流办法,但AES并未采取该办法。
#
# 可以证明,只要每个伪随机函数是安全的,则只要三轮迭代的Feistel网络就是一个安全的伪随机变换。
#
# DES算法的过程,给定64位的模块,经过序列转换之后,成为两个32位的子模块,进入Feistel网络,16轮迭代之后,对两个子模块做之前序列转换的逆运算,得到密文。
#
# Feistel网络中的伪随机函数这样确定:32位子模块,通过重复抽样成为48位子模块,与密钥生成的对应48位密钥做异或运算。得到的结果分位8份,每份6位,将6位利用>查阅表格成为4位,8份重组为32位。32位经过序列变换,输出32位。
#
# 查阅表格成为S-boxes。六位数字的最左和最右位组合作为纵轴,有4种。六位数字的中间四位作为横轴,有16种。从而组合出对应4位的32种组合。
#
# 4位序列如果恰好可以由6位序列线性异或而成,则会造成整体DES加密不安全。
#
# ## 穷举搜索
# 对于DES加密,任意给定一组明文和密文,对应的密钥最多只有一份的概率大于99.5%(1-1/256)。
#
# 给定两组明文和密文,DES密钥唯一的概率1-1/271,AES密钥唯一的概率是1-1/2<sup>128</sup>。因此对于穷举搜索而言,只要获得两组明文和密文即可。
#
# DES的密钥长度是56位,因为存在256种可能。在发明的时候,这个数字不可能穷举。直到1997年,有机构花三个月时间穷举破解了DES加密。而经过特殊设计造价达到25万
# 美元的硬件EFF machine在1998年只需要三天就可以破解。06年,一台价值仅一万美元的机器,花7天时间就可以破解DES。
#
# 解决方案一:3DES。三组相互独立的密钥,3E((k<sub>1</sub>,k<sub>2</sub>,k<sub>3</sub> ),m)=E(k<sub>1</sub>,D(k<sub>2</sub>,E(k<sub>3</sub>,m)))。3DES的
# 密钥长度因此达到了3 * 56=168位,加密速度只有DES的三分之一。之所中间加上一道解密算法,是为了适配很多集成DES的硬件。把三组密钥设为相同,可以回到DES算法
# 。
#
# 为什么不用2DES?2E((k<sub>1</sub>,k<sub>2</sub> ),m)=E(k<sub>1</sub>,E(k<sub>2</sub>,m))。尽管2DES的密钥长度达到了112位,但穷举算法并不需要花费2<sup>112</sup>的时间。将k<sub>2</sub>的256种可能穷举,加密算法对明文列出所有可能的中间密文。然后反向,解密算法对密文计算所有的中间明文。寻找匹配。这样需要
# 花费的时间为2<sup>56</sup> * log2<sup>56</sup> + 2<sup>56</sup> * log2<sup>56</sup> < 2<sup>63</sup> << 2<sup>112</sup>。同理,穷举破译3DES也只需要2<sup>118</sup>,而非2<sup>168</sup>。
# 就目前的计算能力来说,2<sup>63</sup>是一种可以达到的数字,而2<sup>118</sup>暂时还不能。2<sup>90</sup>以上的量级都可以认为是安全的。
#
# 解决方案二:DESX。三组相互独立的密钥,EX((k<sub>1</sub>,k<sub>2</sub>,k<sub>3</sub> ),m)=k<sub>1</sub>⊕E(k<sub>2</sub>,m⊕k<sub>3</sub> )。密钥长度184,穷举可以在2120时间内完成。注意,去除k1或者k3将使加密失效。另外,DESX虽然可以应对穷举搜索,但无法应对其他攻击。保密性不如3DES。
#
# ## 分组密码的其它攻击方式
# 旁路攻击:对系统的物理学分析和实现方式分析,而非通过密码学的分析,来破解密码系统。如监测加密时间,电流等。
#
# 故障攻击:最后一轮加密过程的计算错误可能会暴露密钥。
#
# 线性差分攻击:某几位的加密过程存在固定模式,对明文与密文做异或运算,能够预测结果的概率为1/2+ε。那么给定1⁄ε<sup>2</sup> 对明文和密文,就可以在1⁄ε<sup>2</sup> 的时间内破解出对应位数的密钥。剩下的穷举破解,可大幅缩减整体穷举的时间。对于DES算法,其中14位密钥可以通过线性差分攻击破译,而ε为1/2<sup>21</sup>。也就是说,给定2<sup>42</sup>对密文和明文,总破解时间只需要2<sup>42</sup>+2<sup>42</sup>的时间,非常不安全。
#
# 量子攻击:基于量子计算机,每一位可表示的数字超过两位。传统基于图灵架构的计算机,穷举的时间为O(|X|),而量子计算机穷举的时间是O(|X|<sup>1/2</sup>)。目>前,量子计算机离成形依然非常遥远。
#
# ## 高级加密标准(Advanced Encryption Standard,AES)
# 核心是Subs-Perm网络(替换-变换网络),共10轮迭代。每轮迭代过程,首先与子密钥异或,然后按照查询表格替换,最后变更次序(最后一轮除外)。
#
# 模块共16字节,写成4 * 4的矩阵。子密钥也同样写成矩阵形式,异或。
#
# 替换-变换过程包括三个函数。1)字节替换函数:构造替换框(s-box)替换明文矩阵。2)行位移,第二行向左移动一个单位,第三行两个,第四行三个。3)取其中一列
# ,替换以其他列线性表示。
#
# 提前生成替换-变化函数,可以大幅提高加密速度,但也会使代码变得臃肿。浏览器通常包含了由JavaScript编写的AES模块,提前生成函数提前计算。Intel包括AMD的处>理器通常也都内置了AES计算的相应指令集。
#
# 128位密钥的AES,最佳密钥恢复攻击需要花费2<sup>126</sup>单位时间,比穷举快四倍。而AES-256可能面临相似密钥攻击,收集2<sup>99</sup>对明文密文,可以在2<sup>99</sup>单位时间内恢复密钥。
#
# ## 利用伪随机数生成器(PRG)构造分组密码
# 对一个密钥使用一次伪随机生成器,生成的序列一分为二,得到两份密文,就构造出了一个对应1位密钥的伪随机函数。对两个子序列继续使用伪随机生成器,生成的序列
# 继续一分为二,得到四份密文,就构造出了一个对应2位密钥的伪随机函数。
#
# 依次类推,可以得到对于任意长度密钥的伪随机函数。
#
# 尽管通过伪随机生成器生成的伪随机函数不可逆,但是利用Feistel网络,构造三层,就可以得到伪随机序列,从而成功构造分组密码。
#
# 在实践当中,由于这种办法速度太慢,实际上没有采用这种办法来构造分组密码。
#
#
# # 四、使用分组密码
# ## 回顾:伪随机序列和伪随机函数
# PRP和PRF的安全性也是用语义安全(统计量优势)来定义的。
#
# 当明文长度足够长的时候,一个安全的PRP,同时也是安全的PRF。二者统计量优势之差的绝对值小于q<sup>2</sup>⁄(2|x|),其中q是可获得的明文密文对,而x是明文长>度。
#
# 从这一章开始,可以不再过多考虑AES和3DES的内部工作原理,只要知道它们在现阶段都是安全的PRP,主要考虑如何使用就可以。
#
# ## 运行模式:一次性密钥
# 邮件的加密就是这种模式,每段新明文,都使用新密钥。
#
# 如果密钥永不更换,这种形式叫做电子密码本(Electronic Code Book,ECB)。那么内容相同的模块(block)会得到相同的密文,加密是不安全的。以图片为例,加密>后的图片就会依然呈现出某种模式。
#
# ECB不具备语义安全,可以非常简单地被攻破,攻击者只要发送的两条明文中,一条存在重复的情况,则就可以通过返回密文是否存在重复来辨别。
#
# 安全的分组密码使用方法,是对每个模块添加计数器,传入密钥和计数器用伪随机函数加密。
#
# ## 多次密钥的安全性
# 文件系统和IPsec都是多次使用相同的AES密钥。
#
# 对于多次密钥,攻击者可以使用选择明文攻击(Chosen Plaintext Attack,CPA)。先传入两条相同的明文,得到对应密文。在传入之前的明文和新明文,可以清楚地分>辨所得密文是否来自先前的明文。
#
# 解决方案一:在加密过程中加入随机性,使得一条明文可以映射为多条密文。这样对同一条明文加密两次很大可能得到两条不同的密文,而明文由于需要包含随机数,会>长于密文。在伪随机函数安全的前提下,只要随机数取值空间足够大很大可能不会出现重复,该加密方案对于选择明文攻击就是语义安全的。
#
# 解决方案二:引入不重复数nonce,每次加密,传入明文、密钥和nonce。密钥重复使用,但需要保证(key,nonce)对不重复。Nonce可以随机,也可以是计数器。
#
# ## 运行模式:多次密钥
# CBC和CRT,课件里结合图讲得更清楚,这里不仔细记录了。
| Block Ciphers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn')
# +
from jupyterworkflow.data import get_fremont_data
data = get_fremont_data()
pivoted = data.pivot_table('Total',index=data.index.time,columns=data.index.date)
pivoted.plot(legend=False,alpha=0.01)
# -
pivoted.index
import numpy as np
np.unique(data.index.time)
# +
# !head=24 fremonts.scv
assert len(np.unique(data.index.time) == 24)
# -
# [](http://spacemacs.org)
| fast_download.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3.9
# language: python
# name: python3.9
# ---
import matplotlib.pyplot as plt
wins = {60: 17, 100: 14, 120: 19, 150: 17, 170: 13, 180: 11, 210: 10, 220: 10, 270: 9, 280: 11, 10: 16, 80: 14, 110: 17, 130: 14, 140: 19, 160: 16, 190: 14, 240: 9, 320: 7, 330: 5, 340: 6, 350: 3, 380: 4, 400: 3, 410: 4, 420: 4, 460: 3, 480: 4, 490: 4, 510: 3, 530: 3, 540: 3, 50: 18, 90: 17, 200: 10, 230: 11, 260: 11, 300: 6, 370: 3, 390: 4, 440: 3, 450: 3, 470: 3, 500: 4, 520: 3, 600: 2, 620: 3, 630: 2, 640: 3, 650: 2, 670: 3, 690: 3, 710: 3, 720: 3, 750: 2, 760: 2, 20: 20, 30: 11, 40: 22, 70: 15, 250: 8, 290: 4, 310: 6, 360: 3, 430: 2, 560: 3, 570: 3, 580: 1, 590: 2, 610: 2, 660: 2, 700: 1, 740: 1, 770: 1, 790: 1, 800: 1, 810: 1, 820: 1, 830: 1, 840: 1, 860: 1, 550: 1}
loss = {10: 15, 20: 11, 30: 20, 40: 9, 50: 13, 70: 16, 80: 17, 90: 14, 110: 12, 130: 14, 140: 9, 160: 11, 190: 10, 200: 12, 230: 7, 240: 8, 250: 8, 260: 5, 290: 9, 300: 7, 100: 15, 150: 10, 170: 14, 210: 11, 270: 7, 280: 4, 310: 5, 360: 5, 370: 5, 390: 3, 430: 4, 440: 3, 450: 2, 470: 2, 500: 1, 520: 2, 180: 14, 220: 9, 330: 5, 350: 5, 400: 3, 460: 2, 510: 2, 550: 3, 560: 1, 570: 1, 580: 3, 590: 2, 610: 2, 660: 1, 680: 3, 700: 2, 730: 2, 740: 1, 770: 1, 780: 2, 60: 14, 120: 10, 320: 3, 340: 3, 380: 3, 410: 2, 480: 1, 490: 1, 530: 2, 540: 2, 600: 2, 630: 2, 650: 2, 850: 1, 420: 2, 640: 1, 620: 1}
for k in range(10, 861, 10):
if k not in wins:
wins[k] = 0
if k not in loss:
loss[k] = 0
max([wins[k] + loss[k] for k in range(10, 861, 10)])
plt.figure(dpi=150, facecolor='white')
plt.plot(range(10, 861, 10), [wins[i] for i in range(10, 861, 10)], 'g-', label='wins')
plt.plot(range(10, 861, 10), [loss[i] for i in range(10, 861, 10)], 'r-', label='loss')
plt.xlabel('Number of samples')
plt.ylabel('Number of wins/loss')
plt.legend()
for k in wins:
total = wins[k] + loss[k]
wins[k] = wins[k] / total
loss[k] = loss[k] / total
plt.figure(dpi=150, facecolor='white')
plt.plot(range(10, 861, 10), [wins[i] for i in range(10, 861, 10)], 'g-', label='wins')
plt.plot(range(10, 861, 10), [loss[i] for i in range(10, 861, 10)], 'r-', label='loss')
plt.xlabel('Number of samples')
plt.ylabel('Percentage of wins/loss')
plt.legend()
plt.figure(dpi=150, facecolor='white')
plt.plot(range(10, 861, 10), [loss[i] for i in range(10, 861, 10)], 'w-')
plt.plot(range(10, 861, 10), [wins[i] for i in range(10, 861, 10)], 'g-', label='wins')
plt.plot(range(10, 861, 10), [0.5 for i in range(10, 861, 10)], 'r--')
plt.xlabel('Number of samples')
plt.ylabel('Percentage of wins/loss')
plt.legend()
sum([wins[k] >= 0.5 for k in range(10, 861, 10)]) / len(wins)
| src/data-size-vs-perf/Data size vs. win rate (DODGE vs. DODGE + fuzzy sampling).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing the three algorithms by Neal
# +
import numpy as np
import scipy.stats as stats
import subprocess
import matplotlib.pyplot as plt
from google.protobuf.internal.decoder import _DecodeVarint32
import sys
sys.path.insert(0, '..')
from proto.py.algorithm_state_pb2 import AlgorithmState
import arviz as az
# import pip
# pip.main(["install", "arviz"])
# Utility to save files with Unix-like newlines
def save_np(filename, npobj):
with open(filename, 'wb') as f:
np.savetxt(f, npobj, fmt='%1.5f')
# -
# Generate data
rng = 20201124
np.random.seed(rng)
n = 200
mean1 = -3.0
mean2 = +3.0
norm1 = np.random.normal(loc=mean1, scale=1.0, size=int(n/2))
norm2 = np.random.normal(loc=mean2, scale=1.0, size=int(n/2))
uni_data = np.concatenate((norm1, norm2))
# Generate grid
uni_grid = np.arange(-10, +10, 0.1)
# Save to file
save_np("../resources/csv/in/uni_data.csv", uni_data)
save_np("../resources/csv/in/uni_grid.csv", uni_grid)
# True density of data
true_pdf = 0.5 * stats.norm.pdf(uni_grid, mean1, 1.0) + \
0.5 * stats.norm.pdf(uni_grid, mean2, 1.0)
# Define list of algorithms
algos = ["Neal2", "Neal3", "Neal8"]
# Run the executable
for algo in algos:
cmd = ["../build/run",
"../algo_marg_settings.asciipb",
"NNIG", "../resources/asciipb/nnig_ngg_prior.asciipb",
"DP", "../resources/asciipb/dp_gamma_prior.asciipb",
f"../{algo}.recordio",
"../resources/csv/in/uni_data.csv",
"../resources/csv/in/uni_grid.csv",
f"../resources/csv/out/uni_{algo}_dens.csv",
f"../resources/csv/out/uni_{algo}_mass.csv",
f"../resources/csv/out/uni_{algo}_nclu.csv",
f"../resources/csv/out/uni_{algo}_clus.csv"
]
output = subprocess.run(cmd, capture_output=True)
print(output)
# ## Clustering
# Read clusterings
clusterings = dict.fromkeys(algos)
for algo in algos:
clusterings[algo] = np.loadtxt(f"../resources/csv/out/uni_{algo}_clus.csv")
# Compare clusterings by counting misclassified points
print(np.linalg.norm(clusterings["Neal2"]-clusterings["Neal3"], 1))
print(np.linalg.norm(clusterings["Neal2"]-clusterings["Neal8"], 1))
print(np.linalg.norm(clusterings["Neal3"]-clusterings["Neal8"], 1))
# ## Density estimation
# Densities
plt.figure(figsize=(16, 8))
for algo in algos:
matr = np.genfromtxt(f"../resources/csv/out/uni_{algo}_dens.csv", delimiter=',')
plt.plot(uni_grid, np.exp(np.mean(matr, axis=0)))
plt.plot(uni_grid, true_pdf, color="red", linestyle="--")
plt.legend(algos + ["true"])
plt.title("Univariate densities")
# ## Effective Sample Size
# Utility to read file collector, courtesy of
# github.com/mberaha/utils/blob/master/proto_utils/py/recordio.py
def readManyFromFile(filename, msgType):
out = []
with open(filename, "rb") as fp:
buf = fp.read()
n = 0
while n < len(buf):
msg_len, new_pos = _DecodeVarint32(buf, n)
n = new_pos
msg_buf = buf[n:n+msg_len]
try:
msg = msgType()
msg.ParseFromString(msg_buf)
out.append(msg)
n += msg_len
except Exception as e:
break
return out
# Compute Effective Sample Sizes for each algorithm
ESS = dict.fromkeys(algos)
for algo in algos:
# Read chain
chain = readManyFromFile(f"../{algo}.recordio", MarginalState)
# Record number of clusters at each iteration
n_clusters = np.empty(len(chain))
for i in range(len(chain)):
state = chain[i]
n_clusters[i] = len(state.cluster_states)
ESS[algo] = az.ess(n_clusters)
# Times of MCMC, collected via the progressbar
filecoll_times = dict(zip(algos, [5.690, 6.824, 8.636]))
memocoll_times = dict(zip(algos, [5.617, 6.040, 7.348]))
# Display computed ESS
for key, val in ESS.items():
print(key, "ESS =", val, "-> ESS/time =", val/filecoll_times[key], sep="\t")
# # Marginal (Neal2) vs conditional (BlockedGibbs)
# Run Neal2:
cmd = ("build/run algo_marg_settings.asciipb "
"NNIG resources/asciipb/nnig_ngg_prior.asciipb "
"DP resources/asciipb/dp_gamma_prior.asciipb '' "
"resources/csv/in/uni_data.csv resources/csv/in/uni_grid.csv "
"resources/csv/out/uni_dens.csv resources/csv/out/uni_nclu.csv "
"resources/csv/out/uni_clus.csv").split()
subprocess.run(cmd, capture_output=True)
# Run Blocked Gibbs:
cmd = ("../build/run ../algo_cond_settings.asciipb "
"NNIG ../resources/asciipb/nnig_ngg_prior.asciipb "
"TruncSB ../resources/asciipb/truncsb_py_prior.asciipb '' "
"../resources/csv/in/uni_data.csv ../resources/csv/in/uni_grid.csv "
"../resources/csv/out/truncsb_dens.csv ../resources/csv/out/truncsb_nclu.csv "
"../resources/csv/out/truncsb_clus.csv").split()
subprocess.run(cmd, capture_output=True)
# Read density estimates:
matr1 = np.genfromtxt("../resources/csv/out/uni_dens.csv", delimiter=',')
matr2 = np.genfromtxt("../resources/csv/out/truncsb_dens.csv", delimiter=',')
# Plot density estimates:
plt.figure(figsize=(16, 8))
plt.plot(uni_grid, np.exp(np.mean(matr1, axis=0)))
plt.plot(uni_grid, np.exp(np.mean(matr2, axis=0)))
plt.plot(uni_grid, true_pdf, color="red", linestyle="--")
plt.legend(["Neal2", "BlockedGibbs", "true"])
plt.title("Density estimates")
| python/comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Masterthesis
# language: python
# name: myenv
# ---
# +
import numpy as np
# grAdapt
import grAdapt
from grAdapt.space.datatype import Float, Integer
from grAdapt.models import Sequential
# sklearn
# Import datasets, classifiers and performance metrics
from sklearn.metrics import log_loss
from sklearn import datasets, svm, metrics
from sklearn.model_selection import train_test_split
# The digits dataset
digits = datasets.load_digits()
# plot
import matplotlib.pyplot as plt
# -
# ## 1. Load NIST dataset
# +
# The digits dataset
digits = datasets.load_digits()
# To apply a classifier on this data, we need to flatten the image, to
# turn the data in a (samples, feature) matrix:
n_samples = len(digits.images)
data = digits.images.reshape((n_samples, -1))
# -
# ## 2. Fit SVM
# +
# Create a classifier: a support vector classifier
classifier = svm.SVC(gamma=0.001, probability=True)
# Split data into train and test subsets
X_train, X_test, y_train, y_test = train_test_split(
data, digits.target, test_size=0.5, shuffle=False)
# We learn the digits on the first half of the digits
classifier.fit(X_train, y_train)
# -
classifier.score(X_test, y_test)
# ## 3. Display a test image which has been rightfully classified
plt.imshow(X_test[500].reshape(8, 8), cmap=plt.cm.binary)
classifier.predict_proba(X_test[500].reshape(1, -1))
# ## 4. Goal
# We aim to perturbate the image above such that is will be missclassified. A One Pixel Attack is an adversarial example itself with the only difference that only one pixel of the image is perturbated. This reduces the number of dimensions to deal in the optimization problem. We optimize two values: One for the amount of perturbation and one for which pixel we attack. Both values are contrained to be integers. If the domain constraint is not satisfied by the perturbation, 1 is returned. 0 is returned if the perturbated image has been misclassified.
#
# Differently than adversarial attacks, the norm of the perturbation does not has to be minimized. One pixel is already a small perturbation.
# ### 4.1 Define Black-Box
def one_pixel_attack(per):
# perturbation has shape (2)
# X_test[500] has shape (64,)
pixel_position = int(per[0])
amount = int(per[1])
perturbation = np.zeros((64,))
perturbation[pixel_position] = amount
perturbated_image = (X_test[500] + perturbation).reshape(1, -1)
# image not valid because domain contraint not satisfied
#print(perturbated_image[0][pixel_position].shape)
#print(perturbated_image.shape)
if perturbated_image[0][pixel_position] > 16:
return 1
# let's target the label 5
# this is also the second highest probabilty of the image
target_label = np.zeros((10,))
target_label[6] = 1
pred_label_perturbated = classifier.predict_proba(perturbated_image)
# return cross entropy
#print(target_label.shape)
#print(pred_label_perturbated.shape)
loss = log_loss(target_label, pred_label_perturbated.reshape(10,))
#print(loss)
return loss
# ### 4.2 grAdapt
# Using NoGradient accelerates the training process when dealing with high dimensional optimization problems (64 dimension). Only escape functions are used to obtain the next point. The best point is used as the mean and surrounding points are then evaluated.
# +
pos_bound = Integer(0, 63)
amount_bound = Integer(0, 16)
bounds = [pos_bound, amount_bound]
# -
#sur = grAdapt.surrogate.NoGradient()
model = Sequential(random_state=1)#, surrogate=sur)
res = model.minimize(one_pixel_attack, bounds, 1000)
# #### 4.2.1 Plot Loss
plt.title('Loss')
plt.plot(res['y'], label='grAdapt: Training loss')
plt.legend(loc='upper right')
#plt.yscale('log')
plt.show()
res['y_sol']
# #### 4.2.2 Plot x values
plt.scatter(res['x'][:,0], res['x'][:,1], s=2)
# #### 4.2.3 Plot original and perturbated image
# +
pixel_position, amount = res['x_sol']
pixel_position = int(pixel_position)
amount = int(amount)
perturbation = np.zeros((64,))
perturbation[pixel_position] = amount
perturbated_image = (X_test[500] + perturbation).reshape(1, -1)
# -
fig=plt.figure(figsize=(8, 8))
columns = 2
rows = 1
img = [X_test[500], perturbated_image]
labels = ['Original', 'Perturbated', ]
for i in range(1, columns*rows +1):
fig.add_subplot(rows, columns, i, title=labels[i-1])
plt.imshow(img[i-1].reshape(8, 8), cmap=plt.cm.binary)
plt.show()
classifier.predict_proba(perturbated_image.reshape(1, -1))
target_label = np.zeros((10,))
target_label[6] = 1
classifier.predict_proba(perturbated_image).reshape(10,)
log_loss(target_label, classifier.predict_proba(perturbated_image).reshape(10,))
# ### 4.3 BFGS with scipy
import scipy
x0 = grAdapt.utils.sampling.sample_points_bounds(bounds, 1, random_state=1)
res_scipy = scipy.optimize.minimize(one_pixel_attack, x0, bounds=bounds)
res_scipy
# #### 4.3.1 Plot original and perturbated image
# +
pixel_position, amount = np.round(res_scipy.x)
pixel_position = int(pixel_position)
amount = int(amount)
perturbation = np.zeros((64,))
perturbation[pixel_position] = amount
perturbated_image_scipy = (X_test[500] + perturbation).reshape(1, -1)
# -
fig=plt.figure(figsize=(8, 8))
columns = 2
rows = 1
img = [X_test[500], perturbated_image_scipy]
labels = ['Original', 'Perturbated', ]
for i in range(1, columns*rows +1):
fig.add_subplot(rows, columns, i, title=labels[i-1])
plt.imshow(img[i-1].reshape(8, 8), cmap=plt.cm.binary)
plt.show()
classifier.predict(perturbated_image_scipy.reshape(1, -1))
# ## 5. Conclusion
# Both grAdapt and BFGS struggle to perturbate the image by one pixel to yield for misclassification.
| examples/Adversarial Examples/2. SVM NIST Dataset (One Pixel Attack).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. Thêm các thư viện cần thiết
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.utils import np_utils
from keras.datasets import mnist
import csv
import random
def load_data(path_data):
data = []
label = []
with open(path_data, 'r') as csv_file:
result = csv.reader(csv_file)
data_csv = []
# đọc từng dòng của file và thêm vào list rows, mỗi phần tử của list là một dòng
for row in result:
data_csv.append(row)
for letter in data_csv:
x = np.array([int(j) for j in letter[1:]])
x = x.reshape(28,28)
data.append(x)
label.append(int(letter[0]))
return data, label
# load data and label train
train_data = []
train_label = []
path_data_train = './Input_image/Plate_license_train.csv'
train_data, train_label = load_data(path_data_train)
# load data and label val
val_data = []
val_label = []
path_data_val = './Input_image/Plate_license_val.csv'
val_data, val_label = load_data(path_data_val)
# load data and lable test
test_data = []
test_label = []
path_data_test = './Input_image/Plate_license_test.csv'
test_data, test_label = load_data(path_data_test)
def Shuffle_data(data, label):
i = int(len(data))
shuffle_order = list(range(i))
random.shuffle(shuffle_order)
data = np.array(data)
label = np.array(label)
data = data[shuffle_order]
label = label[shuffle_order]
return data, label
train_data, train_label = Shuffle_data(train_data, train_label)
val_data, val_label = Shuffle_data(val_data, val_label)
test_data, test_label = Shuffle_data(test_data, test_label)
# +
IMG_SIZE = 28
N_CLASSES = 10
input_shape=(28,28,1)
# -
train_data = train_data.reshape(train_data.shape[0], IMG_SIZE, IMG_SIZE, 1)
val_data = val_data.reshape(val_data.shape[0], IMG_SIZE, IMG_SIZE, 1)
test_data = test_data.reshape(test_data.shape[0], IMG_SIZE, IMG_SIZE, 1)
# +
original_label = test_label
train_label = np_utils.to_categorical(train_label, N_CLASSES)
val_label = np_utils.to_categorical(val_label, N_CLASSES)
test_label = np_utils.to_categorical(test_label, N_CLASSES)
# +
# 5. Định nghĩa model
model = Sequential()
# Thêm Convolutional layer với 32 kernel, kích thước kernel 3*3
# dùng hàm sigmoid làm activation và chỉ rõ input_shape cho layer đầu tiên
model.add(Conv2D(32, (3, 3), activation='sigmoid', input_shape=(28,28,1)))
# Thêm Convolutional layer
model.add(Conv2D(32, (3, 3), activation='sigmoid'))
# Thêm Max pooling layer
model.add(MaxPooling2D(pool_size=(2,2)))
# Flatten layer chuyển từ tensor sang vector
model.add(Flatten())
# Thêm Fully Connected layer với 128 nodes và dùng hàm sigmoid
model.add(Dense(128, activation='sigmoid'))
# Output layer với 10 node và dùng softmax function để chuyển sang xác xuất.
model.add(Dense(10, activation='softmax'))
# -
# 6. Compile model, chỉ rõ hàm loss_function nào được sử dụng, phương thức
# đùng để tối ưu hàm loss function.
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
H = model.fit(train_data, train_label, validation_data=(val_data, val_label),
batch_size=32, epochs=10, verbose=1)
# 8. Vẽ đồ thị loss, accuracy của traning set và validation set
fig = plt.figure()
numOfEpoch = 10
plt.plot(np.arange(0, numOfEpoch), H.history['loss'], label='training loss')
plt.plot(np.arange(0, numOfEpoch), H.history['val_loss'], label='validation loss')
plt.plot(np.arange(0, numOfEpoch), H.history['acc'], label='accuracy')
plt.plot(np.arange(0, numOfEpoch), H.history['val_acc'], label='validation accuracy')
plt.title('Accuracy and Loss')
plt.xlabel('Epoch')
plt.ylabel('Loss|Accuracy')
plt.legend()
# 10. Dự đoán ảnh
plt.imshow(test_data[50].reshape(28,28), cmap='gray')
plt.show()
y_predict = model.predict(test_data[50].reshape(1,28,28,1))
print('Giá trị dự đoán: ', np.argmax(y_predict))
model.save('CNN_1_9_1CNN.hdf5')
del model
from keras.models import load_model
model = load_model('CNN_1_9_1CNN.hdf5')
| SEGMENT AND DECTECT CHARACTER/training/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 10. Introduction to Artificial Neural Networks with Keras
# +
from tensorflow import keras
from tensorflow.keras.datasets import boston_housing
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from scipy.stats import reciprocal
from sklearn.model_selection import train_test_split, RandomizedSearchCV
# +
# Ingestion
###########
(train_data, y_train), (test_data, y_test) = boston_housing.load_data()
# Preprocessing
###############
sc = StandardScaler()
x_train = sc.fit_transform(train_data)
x_test = sc.transform(test_data)
x_train__train, x_train__val, y_train__train, y_train__val = train_test_split(x_train, y_train, test_size=0.15,
random_state=0)
NUM_FEATURES = x_train.shape[1:]
# -
# ### Fine-Tuning Neural Network Hyperparameters
#
# Some of the things to consider when using neural networks are:
# 1. Architecture
# 2. For an MLP, the no. of layers, size of layers / no. of neurons, type of activation function, weight inisialisation logic etc.
#
# How do you know what combinations of hyperparameters is the best fo the problem?
# One way is to simply try many combinations of hyperparameters and see which ones work the best during k-fold CV. For this, we can wrap the model around a parameter search algorithm like `GridSearchCV` or `RandomizedSearchCV`.
#
# Using a function call, let's build a way to initialise models with keyword arguments.
def build_model(n_hidden_layers=1, n_neurons=64, learning_rate=3e-3, input_shape=(13,),
dropout=0.0, kernel_regularizer=None):
m = keras.models.Sequential()
for l in range(n_hidden_layers):
m.add(keras.layers.Dense(n_neurons, activation='relu', input_shape=input_shape,
kernel_regularizer=kernel_regularizer))
if 0.0 < dropout:
m.add(keras.layers.Dropout(0.5))
m.add(keras.layers.Dense(1))
optimizer = keras.optimizers.RMSprop(learning_rate=learning_rate)
m.compile(optimizer=optimizer, loss='mse', metrics=['mae'])
return m
# Here we execute a simple workflow on the baseline model.
# BASELINE MODEL
model0 = build_model()
history0 = model0.fit(x_train__train, y_train__train,
epochs=20, batch_size=32,
validation_data=(x_train__val, y_train__val), verbose=0) # Train
model0.predict(x_test[:10]) # Predict
# Let's now build a `keras_reg` that wraps the Keras model, and now can be treated like a classifier like one in `sklearn`. This allows us to implement sklearn functions like `RandomizedSearch`.
keras_reg = keras.wrappers.scikit_learn.KerasRegressor(build_fn=build_model)
# Training baseline model using keras_clf (treating it like a model from sklearn)
stop_early_checkpoint = keras.callbacks.EarlyStopping(patience=3, restore_best_weights=True)
keras_reg.fit(x_train__train, y_train__train,
epochs=20, batch_size=32, callbacks=[stop_early_checkpoint],
validation_data=(x_train__val, y_train__val), verbose=0) # Train
keras_reg.predict(x_test[:10]) # Predict
# +
r = keras.regularizers.l2(l=0.01)
# Training model with regularisation
stop_early_checkpoint = keras.callbacks.EarlyStopping(patience=3, restore_best_weights=True)
keras_reg2 = keras.wrappers.scikit_learn.KerasRegressor(build_fn=build_model,
kernel_regularizer=r)
keras_reg2.fit(x_train__train, y_train__train,
epochs=20, batch_size=32, callbacks=[stop_early_checkpoint],
validation_data=(x_train__val, y_train__val), verbose=0) # Train
keras_reg2.predict(x_test[:10]) # Predict
# +
# Impementing RandomizedSearch on a Keras model.
param_dist = {
'n_hidden_layers' : (1,2,3,4,5),
'n_neurons' : (6,7,8,9,10,11),
'learning_rate' : reciprocal(3e-4, 3e-2),
'kernel_regularizer' : (None,
keras.regularizers.l2(l=0.01),
keras.regularizers.l1(l=0.001),),
'dropout' : (0.0, 0.1, 0.2, 0.3, 0.4, 0.5),
}
rnd_search_cv = RandomizedSearchCV(keras_reg, param_dist, n_iter=10, cv=4)
rnd_search_cv.fit(x_train__train, y_train__train,
epochs=10, batch_size=512, callbacks=[stop_early_checkpoint],
validation_data=(x_train__val, y_train__val), verbose=0)
# -
# Obtaining the best model params
print(rnd_search_cv.best_params_)
print(rnd_search_cv.best_score_)
# And from here, you can save the model, evalauate on test set and if happy, deploy it to production.
| chap04/textbook-chap-4-3b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (traders_nlp)
# language: python
# name: traders_nlp
# ---
# +
from grid_search_tools import GSTools
from ptstrategy_cointegration import CointStrategy
from custom_analyzer import Metrics
from pandas_datafeed import PandasData
from pair_selector import *
import backtrader as bt
import backtrader.feeds as btfeeds
import pandas as pd
import warnings
import glob
import os
import uuid
import itertools
import json
import datetime
# %load_ext autoreload
# %autoreload 2
# +
# INPUT PARAMETERS
DIR = "../ib-data/nyse-daily-tech/"
BT_START_DT = '2018-03-19'
TEST_PERIOD = 200
PRE_PAIR_FORMATION = 252 + 252 + 252 + 52 - 60 - 52
PAIR_FORMATION_LEN = 60
# top PCT percentage of the pairs with lowest distance will be backtested
PCT = 0.9
# STRATEGY PARAMETERS
LOOKBACK_VALUE = [20, 40, 52]
ENTER_THRESHOLD_SIZE = [2, 3]
EXIT_THRESHOLD_SIZE = [0.5]
LOSS_LIMIT = [-0.005]
MAX_LOOKBACK = max(LOOKBACK_VALUE)
CONSIDER_BORROW_COST = False
CONSIDER_COMMISSION = True
# ADDITIONAL INFO
OTHER_INFO = ""
# Where to save the ouputs
DST_DIR = "../backtest-results/cointegration/experiment3-y3/"
# +
CONFIG = {
'DIR': DIR,
'BT_START_DT': BT_START_DT,
'TEST_PERIOD': TEST_PERIOD,
'PRE_PAIR_FORMATION': PRE_PAIR_FORMATION,
'PAIR_FORMATION_LEN': PAIR_FORMATION_LEN,
'PCT': PCT,
'LOOKBACK_VALUE': LOOKBACK_VALUE,
'ENTER_THRESHOLD_SIZE': ENTER_THRESHOLD_SIZE,
'EXIT_THRESHOLD_SIZE': EXIT_THRESHOLD_SIZE,
'LOSS_LIMIT': LOSS_LIMIT,
'MAX_LOOKBACK': MAX_LOOKBACK,
'CONSIDER_BORROW_COST': CONSIDER_BORROW_COST,
'CONSIDER_COMMISSION': CONSIDER_COMMISSION,
'DST_DIR': DST_DIR,
'OTHER_INFO': OTHER_INFO,
}
# create json string
CONFIG_JSON_STR = json.dumps(CONFIG)
# create directory if neccessary
if not os.path.exists(DST_DIR):
os.makedirs(DST_DIR)
# save json string to a file
with open(DST_DIR + 'config.json', 'w') as outfile:
json.dump(CONFIG_JSON_STR, outfile)
# +
print("---------------------------------------------------------------------")
###################################################################################################################
# Load data
data = GSTools.load_csv_files(DIR)
dt_idx = GSTools.get_trading_dates(data)
print("Initial number of datafeeds: " + str(len(dt_idx)) + ".")
###################################################################################################################
# get position of intended start date of backtest
bt_start_idx = dt_idx.get_loc(BT_START_DT)
size = PRE_PAIR_FORMATION + PAIR_FORMATION_LEN + MAX_LOOKBACK + (len(dt_idx) - bt_start_idx)
print("To fulfill BT_START_DT, PAIR_FORMATION_LEN and MAX_LOOKBACK, size = " + str(size) + ".")
# get datafeeds which fulfill size requirement
data = GSTools.cut_datafeeds(data, size=size)
print("After cutting datafeeds, " + str(len(data.keys())) + " datafeeds remaining.")
###################################################################################################################
# just to be safe, sync the start end dates of the dataframes
data, start_dt, end_dt = GSTools.sync_start_end(data)
dt_idx = GSTools.get_trading_dates(data)
print("Backtest start date: " + str(dt_idx[PRE_PAIR_FORMATION + PAIR_FORMATION_LEN + MAX_LOOKBACK]))
print("Backtest end date: " + str(dt_idx[PRE_PAIR_FORMATION + PAIR_FORMATION_LEN + MAX_LOOKBACK + TEST_PERIOD - 1]))
###################################################################################################################
# get aggregated close prices
close_df = GSTools.get_aggregated(data, col='close')
if close_df.isnull().values.any():
warnings.warn("There are null values in the aggregated close price df.")
else:
print("No null values detected in aggregated close price df.")
###################################################################################################################
# total number of stocks remaining
N = len(data.keys())
# number of pairs of interest
K = int(PCT * N * (N-1) / 2)
###################################################################################################################
# pair selection
good_pairs = coint(df=close_df[PRE_PAIR_FORMATION:PRE_PAIR_FORMATION + PAIR_FORMATION_LEN], intercept=True, sig_level=0.005)
good_pairs.sort(key=lambda x: x[2])
good_pairs = good_pairs[0 : K]
print("From " + str(int(N * (N-1) / 2)) + " pairs, " + str(len(good_pairs)) + " pairs passed the cointegration test.")
print("---------------------------------------------------------------------")
# +
# combinations of parameters
param_combinations = list(itertools.product(LOOKBACK_VALUE, ENTER_THRESHOLD_SIZE, EXIT_THRESHOLD_SIZE, LOSS_LIMIT))
# list to store MACRO results
macro_results = []
for i, params in enumerate(param_combinations, 1):
# set params
print("Running " + str(i) + "/" + str(len(param_combinations)))
print (str(datetime.datetime.now()))
print("Backtesting all pairs using parameters " + str(params))
# list to store MICRO results
results = []
for pair in good_pairs:
# get names of both stock
stk0, stk1, _ = pair
# get data of both stock
stk0_df, stk1_df = data[stk0], data[stk1]
stk0_df_test = stk0_df[PRE_PAIR_FORMATION + PAIR_FORMATION_LEN : PRE_PAIR_FORMATION + PAIR_FORMATION_LEN + MAX_LOOKBACK + TEST_PERIOD]
stk1_df_test = stk1_df[PRE_PAIR_FORMATION + PAIR_FORMATION_LEN : PRE_PAIR_FORMATION + PAIR_FORMATION_LEN + MAX_LOOKBACK + TEST_PERIOD]
# Create a cerebro
cerebro = bt.Cerebro()
# Create data feeds
data0 = bt.feeds.PandasData(dataname=stk0_df_test, timeframe=(bt.TimeFrame.Days), datetime=0)
data1 = bt.feeds.PandasData(dataname=stk1_df_test, timeframe=(bt.TimeFrame.Days), datetime=0)
# add data feeds to cerebro
cerebro.adddata(data0)
cerebro.adddata(data1)
# Add the strategy
cerebro.addstrategy(CointStrategy,
lookback=params[0],
max_lookback=MAX_LOOKBACK,
enter_threshold_size=params[1],
exit_threshold_size=params[2],
loss_limit=params[3],
consider_borrow_cost=CONSIDER_BORROW_COST,
consider_commission=CONSIDER_COMMISSION
)
# Add analyzers
cerebro.addanalyzer(bt.analyzers.SharpeRatio, _name='mysharpe')
cerebro.addanalyzer(Metrics,
lookback=MAX_LOOKBACK,
_name='metrics')
# Add the commission - only stocks like a for each operation
cerebro.broker.setcash(1000000)
# And run it
strat = cerebro.run()
# get MICRO metrics
sharperatio = strat[0].analyzers.mysharpe.get_analysis()['sharperatio']
returnstd = strat[0].analyzers.metrics.returns_std()
startcash = cerebro.getbroker().startingcash
endcash = cerebro.getbroker().getvalue()
profit = (endcash - startcash) / startcash
results.append((stk0 + "-" + stk1, sharperatio, profit, returnstd))
# convert to dataframe
results_df = pd.DataFrame(results)
results_df.columns = ['pair', 'sharpe_ratio', 'overall_return', 'returns_std']
# save as csv
uuid_str = str(uuid.uuid4())
path = DST_DIR + str(uuid_str) + ".csv"
results_df.to_csv(path_or_buf=path, index=False)
# calculate MACRO attributes
avg_sharpe_ratio = results_df['sharpe_ratio'].mean()
median_sharpe_ratio = results_df['sharpe_ratio'].median()
avg_overall_return = results_df['overall_return'].mean()
median_overall_return = results_df['overall_return'].median()
overall_return_std = results_df['overall_return'].std()
macro_results.append((params[0],
params[1],
params[2],
params[3],
avg_sharpe_ratio,
median_sharpe_ratio,
avg_overall_return,
median_overall_return,
overall_return_std,
uuid_str
))
# nextline
print("")
macro_results_df = pd.DataFrame(macro_results)
macro_results_df.columns = ['lookback',
'enter_threshold_size',
'exit_threshold_size',
'loss_limit',
'avg_sharpe_ratio',
'median_sharpe_ratio',
'avg_overall_return',
'median_overall_return',
'overall_return_std',
'uuid']
macro_results_df.to_csv(DST_DIR + 'summary.csv', index=False)
# -
macro_results_df = pd.read_csv(DST_DIR + "summary.csv")
macro_results_df
macro_results_df[macro_results_df['median_overall_return'] == max(macro_results_df['median_overall_return'])]
macro_results_df
| jupyter_py/5c-Cointegration-Method-Grid-Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
# # 1. Loading Data
df = pd.read_csv('../data/WA_Fn-UseC_-Marketing-Customer-Value-Analysis.csv')
df.shape
df.head()
df['Engaged'] = df['Response'].apply(lambda x: 0 if x == 'No' else 1)
df.head()
# # 2. Data Analysis
list(df.columns)
# #### - Engagement Rate
engagement_rate_df = pd.DataFrame(
df.groupby('Engaged').count()['Response'] / df.shape[0] * 100.0
)
engagement_rate_df
engagement_rate_df.T
# #### - By Renew Offer Type
# +
engagement_by_offer_type_df = pd.pivot_table(
df, values='Response', index='Renew Offer Type', columns='Engaged', aggfunc=len
).fillna(0.0)
engagement_by_offer_type_df.columns = ['Not Engaged', 'Engaged']
# -
engagement_by_offer_type_df
# +
engagement_by_offer_type_df.plot(
kind='pie',
figsize=(15, 7),
startangle=90,
subplots=True,
autopct=lambda x: '%0.1f%%' % x
)
plt.show()
# -
# #### - By Sales Channel
# +
engagement_by_sales_channel_df = pd.pivot_table(
df, values='Response', index='Sales Channel', columns='Engaged', aggfunc=len
).fillna(0.0)
engagement_by_sales_channel_df.columns = ['Not Engaged', 'Engaged']
# -
engagement_by_sales_channel_df
# +
engagement_by_sales_channel_df.plot(
kind='pie',
figsize=(15, 7),
startangle=90,
subplots=True,
autopct=lambda x: '%0.1f%%' % x
)
plt.show()
# -
# #### - Total Claim Amount Distributions
# +
ax = df[['Engaged', 'Total Claim Amount']].boxplot(
by='Engaged',
showfliers=False,
figsize=(7,5)
)
ax.set_xlabel('Engaged')
ax.set_ylabel('Total Claim Amount')
ax.set_title('Total Claim Amount Distributions by Enagements')
plt.suptitle("")
plt.show()
# +
ax = df[['Engaged', 'Total Claim Amount']].boxplot(
by='Engaged',
showfliers=True,
figsize=(7,5)
)
ax.set_xlabel('Engaged')
ax.set_ylabel('Total Claim Amount')
ax.set_title('Total Claim Amount Distributions by Enagements')
plt.suptitle("")
plt.show()
# -
# #### - Income Distributions
# +
ax = df[['Engaged', 'Income']].boxplot(
by='Engaged',
showfliers=True,
figsize=(7,5)
)
ax.set_xlabel('Engaged')
ax.set_xlabel('Income')
ax.set_title('Income Distributions by Enagements')
plt.suptitle("")
plt.show()
# -
df.groupby('Engaged').describe()['Income'].T
# # 3. Regression Analysis with Continuous Variables Only
import statsmodels.api as sm #updated
df.describe()
df['Income'].dtype
df['Customer Lifetime Value'].dtype
continuous_vars = [
'Customer Lifetime Value', 'Income', 'Monthly Premium Auto',
'Months Since Last Claim', 'Months Since Policy Inception',
'Number of Open Complaints', 'Number of Policies',
'Total Claim Amount'
]
logit = sm.Logit(
df['Engaged'],
df[continuous_vars]
)
logit_fit = logit.fit()
logit_fit.summary()
# # 4. Regression Analysis with Categorical Variables
df.describe()
# #### - Different ways to handle categorical variables
# ###### 1. factorize
labels, levels = df['Education'].factorize()
labels
levels
# ###### 2. pandas' Categorical variable series
categories = pd.Categorical(
df['Education'],
categories=['High School or Below', 'Bachelor', 'College', 'Master', 'Doctor']
)
categories.categories
categories.codes
# ###### 3. dummy variables
pd.get_dummies(df['Education']).head(10)
# #### - Adding Gender
gender_values, gender_labels = df['Gender'].factorize()
df['GenderFactorized'] = gender_values
gender_values
gender_labels
df
# #### - Adding Education Level
categories = pd.Categorical(
df['Education'],
categories=['High School or Below', 'Bachelor', 'College', 'Master', 'Doctor']
)
categories.codes
categories.categories
df['EducationFactorized'] = categories.codes
df.head()
# #### - Regression Analysis with Categorical Variables
logit = sm.Logit(
df['Engaged'],
df[[
'GenderFactorized',
'EducationFactorized'
]]
)
logit_fit = logit.fit()
logit_fit.summary()
# # 5. Regression Analysis with Both Continuous and Categorical Variables
logit = sm.Logit(
df['Engaged'],
df[['Customer Lifetime Value',
'Income',
'Monthly Premium Auto',
'Months Since Last Claim',
'Months Since Policy Inception',
'Number of Open Complaints',
'Number of Policies',
'Total Claim Amount',
'GenderFactorized',
'EducationFactorized'
]]
)
logit_fit = logit.fit()
logit_fit.summary()
| RegressionAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8 (py38)
# language: python
# name: py38
# ---
# +
import os
import time
import numpy as np
import pandas
import geopandas
from shapely.ops import unary_union
from shapely.geometry import Point, Polygon
from matplotlib import pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.patches import FancyBboxPatch
import re
from pprint import pprint
from matplotlib.path import Path
from matplotlib.patches import BoxStyle
thisyear = 2018
#this_state = 'MO'
script_dir = '/home/idies/workspace/Storage/raddick/jordanraddick.com/gerrymandering/'
data_dir = '/home/idies/workspace/Storage/raddick/jordanraddick.com/gerrymandering/districts_with_data/'
output_dir = data_dir
acs5_dir = '/home/idies/workspace/Temporary/raddick/census_scratch/acs5/{0:.0f}/estimates/'.format(thisyear)
shapefiledir = '/home/idies/workspace/Temporary/raddick/census_scratch/shapefiles/{0:.0f}/'.format(thisyear)
extras_dir = '/home/idies/workspace/Storage/raddick/census/extras/'
#separated_roads_dir = output_dir = '/home/idies/workspace/Temporary/raddick/jordanraddick.com_temp/roads/{0:}/'.format(this_state.lower())
water_area_tol = 1 * 1000 * 1000
overlap_area_tract_tol = 22000
overlap_area_bg_tol = 4000
#smallest tract in US is Cook County, Illinois Tract 307.02 (area = 22,094 m^2)
#smallest block group in US is Miami-Dade County, FL, Census Tract 2703, block group 7 (area = 4,436 m^2)
equal_area_crs = {'init': 'epsg:2163'} # An equal area projection: https://epsg.io/2163
scale = 1
#map_buffer = 0.25 # extra room on each edge of the maps, in degres
#plt.rc('axes', prop_cycle=default_cycler)
district_color_cycle = ['red', 'green', 'orange', 'cyan', 'yellow', 'pink', 'gray', 'lime', 'navajowhite', 'cornflowerblue', 'darkseagreen', 'thistle', 'tomato', 'silver', 'blueviolet', 'olive', 'peru', 'dodgerblue']
district_contrast_color_cycle = ['green', 'red', 'navy', 'black', 'purple', 'lime', 'black', 'red', 'red', 'black', 'red', 'black', 'black', 'white', 'yellow', 'yellow', 'yellow', 'yellow']
debug = 1
g = 0
def makebox(manual_xlim, manual_ylim):
points = []
points.append(Point(manual_xlim[0],manual_ylim[1]))
points.append(Point(manual_xlim[1],manual_ylim[1]))
points.append(Point(manual_xlim[1],manual_ylim[0]))
points.append(Point(manual_xlim[0],manual_ylim[0]))
coords = [(p.x, p.y) for p in points]
metrobox = Polygon(coords)
return metrobox
# we may derive from matplotlib.patches.BoxStyle._Base class.
# You need to override transmute method in this case.
class shield(BoxStyle._Base):
"""
A simple box.
"""
def __init__(self, pad=0.3):
"""
The arguments need to be floating numbers and need to have
default values.
*pad*
amount of padding
"""
self.pad = pad
super().__init__()
def transmute(self, x0, y0, width, height, mutation_size):
"""
Given the location and size of the box, return the path of
the box around it.
- *x0*, *y0*, *width*, *height* : location and size of the box
- *mutation_size* : a reference scale for the mutation.
Often, the *mutation_size* is the font size of the text.
You don't need to worry about the rotation as it is
automatically taken care of.
"""
# padding
pad = mutation_size * self.pad
# width and height with padding added.
width, height = width + 2.*pad, \
height + 2.*pad,
# boundary of the padded box
x0, y0 = x0-pad, y0-pad,
x1, y1 = x0+width, y0 + height
cp = [(0.5*(x0+x1), (y0-2.*pad)), # bottom
(x1, y0), # right lower-mid
(x1+pad, (y0+y1)/2.), # far right mid
(x1, y1+pad), # top right corner
((x0+x1)/2,y1), # dip from top
(x0, y1+pad), # top left corner
(x0-pad, (y0+y1)/2.), # far left mid
(x0, y0), # left lower-mid
((x0+x1)/2., (y0-2.*pad)), # return to bottom
((x0+x1)/2., (y0-2.*pad))]
com = [Path.MOVETO, # start
Path.CURVE4, # curve to right mid
Path.LINETO, # line to far right mid
Path.LINETO, # line to top right corner
Path.CURVE3, # line to dip
Path.LINETO, # line to top left corner
Path.CURVE4, # curve to far left mid
Path.LINETO, # line to left lower-mid
Path.LINETO,
Path.CLOSEPOLY]
path = Path(cp, com)
return path
BoxStyle._style_list["shield"] = shield
def parse_road_name(thename):
try:
annotator = thisrow['FULLNAME'][re.search('\d',thisrow['FULLNAME']).start():]
except AttributeError:
try:
annotator = thisrow['FULLNAME'][re.search('Hwy',thisrow['FULLNAME']).end():]
except AttributeError:
annotator = thisrow['FULLNAME']
return annotator
road_label_format = {
'I': { 'labelsize': 16, 'thecolor': 'yellow', 'thebbox': dict(boxstyle="shield", fc='blue', ec='orange') },
'U': { 'labelsize': 14, 'thecolor': 'black', 'thebbox': dict(boxstyle="shield", fc='white', ec='black') },
'S': { 'labelsize': 12, 'thecolor': 'black', 'thebbox': dict(boxstyle="square,pad=0.25", fc='white', ec='black')},
'C': { 'labelsize': 10, 'thecolor': 'black', 'thebbox': dict(boxstyle="sawtooth,pad=0.5", fc='white') },
'M': { 'labelsize': 16, 'thecolor': 'black'},
'O': { 'labelsize': 11, 'thecolor': 'yellow' }
}
old_map_buffer_ratio = -1
print('ok')
# -
# # Get congressional district shapefiles
#
# WITH data on metro and city areas as a percentage of total district area.
#
# This assumes that you have already run <code>districts-cities.ipynb</code>.
# +
s = time.time()
print('reading congressional districts with urban/rural area data...')
cd_gdf = geopandas.read_file(data_dir+'cd116_with_areas_and_types_435.shp')
print('Fixing at-large districts by setting district number to 1...')
cd_gdf.loc[cd_gdf['CD116FP'] == 0, 'CD116FP'] = 1 # At-large districts will be called District 1
cd_gdf = cd_gdf.rename(columns = {'pct_metro_': 'pct_metro_area', 'pct_city_a': 'pct_city_area'})
cd_gdf.loc[:, 'CD116FP'] = cd_gdf['CD116FP'].apply(lambda x: int(x))
cd_gdf = cd_gdf.set_index('GEOID')
e = time.time()
g = g + (e-s)
print('Read {0:,.0f} districts in {1:,.1f} seconds.'.format(len(cd_gdf), e-s))
#sorted(cd_gdf[cd_gdf['CD116FP'] == 0]['STUSAB'].tolist())
# -
# # Load block groups (nationwide)
# +
s = time.time()
bg_file_list = [shapefiledir+'BG/'+x for x in os.listdir(shapefiledir+'BG/') if ((x[-4:] == '.shp'))]
bg_gdf = geopandas.GeoDataFrame()
for i in range(0, len(bg_file_list)):
if (debug >= 1):
if ((np.mod(i,10) == 0) | (i == len(bg_file_list)-1)):
print('\tReading file {0:,.0f} of {1:,.0f}...'.format(i+1, len(bg_file_list)))
bg_gdf_i = geopandas.read_file(bg_file_list[i])
bg_gdf = pandas.concat((bg_gdf, bg_gdf_i), axis=0, sort=False)
print('converting block group identifiers and coords to numeric...')
bg_gdf.loc[:, 'STATEFP'] = pandas.to_numeric(bg_gdf['STATEFP'], errors='coerce')
bg_gdf.loc[:, 'COUNTYFP'] = pandas.to_numeric(bg_gdf['COUNTYFP'], errors='coerce')
bg_gdf.loc[:, 'TRACTCE'] = pandas.to_numeric(bg_gdf['TRACTCE'].apply(lambda x: x[0:4]+'.'+x[4:]), errors='coerce')
bg_gdf.loc[:, 'BLKGRPCE'] = pandas.to_numeric(bg_gdf['BLKGRPCE'], errors='coerce')
bg_gdf.loc[:, 'INTPTLAT'] = pandas.to_numeric(bg_gdf['INTPTLAT'], errors='coerce')
bg_gdf.loc[:, 'INTPTLON'] = pandas.to_numeric(bg_gdf['INTPTLON'], errors='coerce')
#bg_gdf.loc[:, 'NAME'] = pandas.to_numeric(tract_gdf['NAME'], errors='coerce')
# bg_gdf = tract_gdf.sort_values(by='NAME')
print('assigning GEOID as index...')
bg_gdf.loc[:, 'GEOID'] = bg_gdf['GEOID'].apply(lambda x: '15000US'+str(x))
bg_gdf = bg_gdf.set_index('GEOID')
e = time.time()
g = g + (e-s)
if (debug >= 1):
print('Read {0:,.0f} census block groups in {1:,.1f} seconds!'.format(len(bg_gdf), e-s))
#bg_gdf.sample(1).T
# -
# ## Look up state and county names from numbers
# +
s = time.time()
print('looking up state names from numbers...')
state_codes_df = pandas.read_csv(extras_dir+'statecodes.csv')
state_codes_df = state_codes_df.rename(columns={'STATE': 'STATEFP'})
bg_gdf = bg_gdf.reset_index().merge(state_codes_df, how='left', on='STATEFP').set_index('GEOID')
print('looking up county names from numbers...')
county_names_df = pandas.read_excel(extras_dir+'all-geocodes-v2019.xlsx', header=4)
county_names_df = county_names_df[county_names_df['Summary Level'] == 50]
county_names_df = county_names_df.rename(columns={'State Code (FIPS)': 'STATEFP', 'County Code (FIPS)': 'COUNTYFP', 'Area Name (including legal/statistical area description)': 'COUNTY_NAME' })
bg_gdf = bg_gdf.reset_index().merge(county_names_df[['STATEFP', 'COUNTYFP', 'COUNTY_NAME']], how='left', on=['STATEFP', 'COUNTYFP']).set_index('GEOID')
e = time.time()
g = g + (e-s)
print('Added state and county names in {0:,.1f} seconds!'.format(e-s))
# -
# ## Get population data, and join onto shapefiles
# +
s = time.time()
print('reading ACS5 census data for {0:.0f}...'.format(thisyear))
acs5_estimates_df = pandas.read_csv(acs5_dir+'estimates_acs{0:}_tract_bg_gerrymandering.csv'.format(thisyear), index_col='GEOID')
print('joining population data onto block group shapefiles...')
bg_gdf = bg_gdf.join(acs5_estimates_df[['B01001_001', 'Geography Name']], how='left')
bg_gdf = bg_gdf.rename(columns={'B01001_001': 'total_population'})
e = time.time()
g = g + (e-s)
print('\nadded ACS5 census data to {0:,.0f} block groups in {1:,.0f} seconds!'.format(len(bg_gdf), e-s))
# -
# # Geo-match congressional districts
#
# If a tract overlaps with only one district, match that tract to its district.
# If it overlaps multiple districts, divide into block groups and match each block group to its matching districts.
# # READ FROM FILE
# ## Get what has been matched so far
# +
# s = time.time()
# print('reading districts that have been matched so far...')
# assembler_df = pandas.read_csv(output_dir+'raw_district_files/ak_to_ca.csv', encoding='utf-8', index_col='GEOID')
# assembler_gdf = geopandas.GeoDataFrame(assembler_df.join(bg_gdf.geometry))
# assembler_gdf.crs = bg_gdf.crs
# # cd_gdf = cd_gdf[~cd_gdf['STUSAB'].isin(assembler_gdf['STUSAB'].drop_duplicates().tolist())]
# # bg_gdf = bg_gdf[~bg_gdf['STUSAB'].isin(assembler_gdf['STUSAB'].drop_duplicates().tolist())]
# #print('backing up...')
# # cd_gdf_bk = cd_gdf
# # bg_gdf_bk = bg_gdf
# #assembler_gdf_bk = assembler_gdf
# e = time.time()
# g = g + (e-s)
# print('Kept {0:,.0f} block groups in {1:,.1f} seconds!'.format(len(bg_gdf),e-s))
# assembler_gdf.groupby('STUSAB')['block_based_district'].max().sort_values(ascending=False)
# -
# ### READ AFRESH
# +
s = time.time()
print('reading districts that have been matched so far...')
assembler_df = pandas.read_csv(data_dir+'block_group_with_district_1_36.csv', encoding='utf-8', index_col='GEOID')
assembler_gdf = geopandas.GeoDataFrame(assembler_df.join(bg_gdf.geometry))
assembler_gdf.crs = bg_gdf.crs
# cd_gdf = cd_gdf[~cd_gdf['STUSAB'].isin(assembler_gdf['STUSAB'].drop_duplicates().tolist())]
# bg_gdf = bg_gdf[~bg_gdf['STUSAB'].isin(assembler_gdf['STUSAB'].drop_duplicates().tolist())]
print('backing up...')
cd_gdf_bk = cd_gdf
bg_gdf_bk = bg_gdf
assembler_gdf_bk = assembler_gdf
e = time.time()
g = g + (e-s)
print('Kept {0:,.0f} block groups in {1:,.1f} seconds!'.format(len(bg_gdf),e-s))
# -
# ## California
#
# These are harder, because first we have to know which block groups overlap multiple districts.
#
# Create a function to mark those.
# ### Create function to mark block groups with the districts they overlap
def mark_block_groups_with_districts_bitmask_values(block_group_info_gdf, congressional_district_info_gdf, debug=0):
#working_with_state = block_group_info_gdf['STATE_NAME'].head(1).values[0]
marked_s = pandas.Series()
equal_area_crs = {'init': 'epsg:2163'} # An equal area projection: https://epsg.io/2163
cnt = 0
#if (debug > 0):
# print('\tAssigning bitmask values to block groups in {0:}...'.format(working_with_state))
for ix, thisrow in block_group_info_gdf.iterrows():
bitmasker = 0
if (debug > 1):
print(ix)
print('\t\tMatching {0:}, {1:} census tract {2:}, block group {3:}...'.format(thisrow['STATE_NAME'], thisrow['COUNTY_NAME'], thisrow['TRACTCE'], thisrow['BLKGRPCE']))
print('\n')
if ((np.mod(cnt,1000) == 0) | (cnt == len(block_group_info_gdf) - 1)):
if (debug > 1):
print('\t\t\tprocessing row {0:,.0f} of {1:,.0f}...'.format(cnt+1, len(block_group_info_gdf)))
else:
print('\t\tprocessing row {0:,.0f} of {1:,.0f}...'.format(cnt+1, len(block_group_info_gdf)))
for jx, thatrow in congressional_district_info_gdf.iterrows():
if (thisrow.geometry.intersects(thatrow.geometry)):
this_district_overlap_area = block_group_info_gdf[block_group_info_gdf.index == ix].to_crs(equal_area_crs).geometry.values[0].intersection(congressional_district_info_gdf[congressional_district_info_gdf.index == jx].to_crs(equal_area_crs).geometry.values[0]).area
if (this_district_overlap_area >= overlap_area_bg_tol):
bitmasker = bitmasker + 2**(thatrow['CD116FP']-1)
if (debug > 1):
print('\t\t\t\tIntersects District {0:.0f} with overlap area {1:,.1f} km^2...'.format(thatrow['CD116FP'], this_district_overlap_area/1000000))
marked_s.loc[ix] = bitmasker
cnt = cnt + 1
return marked_s
print('defined district-marking function!')
#assembler_gdf.groupby('STUSAB')['block_based_district'].max().sort_values(ascending=False)
# ## Find block group / district overlaps, assign districts to non-overlapping block groups
#
# Uses function defined above
# +
s = time.time()
print('getting from backup...')
bg_gdf = bg_gdf_bk
assembler_gdf = assembler_gdf_bk
for this_state in ['CA']:
state_full_name = bg_gdf[bg_gdf['STUSAB'] == this_state]['STATE_NAME'].values[0]
print('\n')
print('Processing {0:}...'.format(state_full_name))
assembler_i_gdf = bg_gdf[bg_gdf['STUSAB'] == this_state]
district_bitmask_values_s = mark_block_groups_with_districts_bitmask_values(
bg_gdf[bg_gdf['STUSAB'] == this_state][['STATE_NAME', 'COUNTY_NAME', 'TRACTCE', 'BLKGRPCE', 'total_population', 'geometry']],
cd_gdf[cd_gdf['STUSAB'] == this_state][['STUSAB', 'CD116FP', 'geometry']],
debug
)
assembler_i_gdf = assembler_i_gdf.assign(congressional_districts_bitmask_values = district_bitmask_values_s)
if (debug > 0):
print('\tconverting bitmask values to human-readable bitmasks...')
nDistrictsForBitmaskeration = cd_gdf[cd_gdf['STUSAB'] == this_state]['CD116FP'].max()
assembler_i_gdf = assembler_i_gdf.assign(congressional_districts_bitmask =
assembler_i_gdf['congressional_districts_bitmask_values'].apply(lambda x: 'x'+np.binary_repr(int(x)).zfill(nDistrictsForBitmaskeration)[::-1])
)
if (debug > 0):
print('\tcounting number of districts each block group overlaps...')
assembler_i_gdf = assembler_i_gdf.assign(nDistricts =
assembler_i_gdf['congressional_districts_bitmask'].apply(lambda x: x[1:].count("1"))
)
if (debug > 0):
print('\tassigning congressional district to each block group with only one overlap...')
assembler_i_gdf = assembler_i_gdf.assign(block_based_district = np.nan)
assembler_i_gdf.loc[
assembler_i_gdf['nDistricts'] == 1,
'block_based_district'] = assembler_i_gdf[
assembler_i_gdf['nDistricts'] == 1
]['congressional_districts_bitmask'].apply(lambda x: x.find("1"))
print('\t...-1 otherwise...')
assembler_i_gdf.loc[assembler_i_gdf['nDistricts'] > 1, 'block_based_district'] = -1
#assembler_i_gdf.to_file(output_dir+'raw_district_files/nj_raw.shp')
if (debug > 0):
print('\tjoining {0:} to the rest of the assembler dataframe...'.format(state_full_name))
assembler_gdf = pandas.concat((assembler_gdf, assembler_i_gdf), axis=0, sort=True)
if (debug > 0):
print("\n")
e = time.time()
g = g + (e-s)
print('matched districts for {0:,.0f} block groups in {1:} in {2:,.0f} minutes {3:,.0f} seconds!'.format(len(assembler_i_gdf), this_state, np.floor((e-s)/60), np.floor((e-s)%60)))
print('backing up...')
assembler_gdf_justparsed = assembler_gdf
e = time.time()
g = g + (e-s)
#print('Read {0:,.0f} rows in {1:,.0f} seconds!'.format(e-s, ))
print('Got {0:,.0f} block groups in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(assembler_gdf), np.floor((e-s)/60), np.floor((e-s)%60)))
# print(assembler_gdf.groupby('STUSAB').size())
# print(assembler_gdf.groupby(['STUSAB', 'nDistricts']).size())
#print(assembler_gdf.groupby(['STUSAB', 'block_based_district']).size())
# assembler_gdf.sample(2).T
#assembler_gdf[assembler_gdf['STUSAB'] == this_state].groupby('congressional_districts_bitmask').size()
assembler_gdf.groupby('STUSAB')['block_based_district'].max().sort_values(ascending=False)
# matched districts for 23,212 block groups in CA in 51 minutes 17 seconds!
# backing up...
# Got 201,478 block groups in 51 minutes 17 seconds!
# +
# s = time.time()
# pandas.DataFrame(assembler_gdf[[x for x in assembler_gdf.columns if x != 'geometry']]
# ).to_csv(output_dir+'raw_district_files/ak_to_ca.csv', encoding='utf-8')
# e = time.time()
# g = g + (e-s)
# print('Wrote out {0:,.0f} block groups in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(dallas_gdf), np.floor((e-s)/60), np.floor((e-s)%60)))
# +
s = time.time()
print('Assigning districts based on examination to block groups that overlap multiple districts...')
print('assigning districts...')
### CALIFORNIA
##### Bay Area
assembler_gdf.loc[['15000US060971506122'], 'block_based_district'] = 5
assembler_gdf.loc[['15000US060952522024', '15000US060952522014'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US060952521021', '15000US060952522021'], 'block_based_district'] = 5
assembler_gdf.loc[['15000US060133180002', '15000US060133560022', '15000US060133180003', '15000US060133601013', '15000US060133592022'], 'block_based_district'] = 5
assembler_gdf.loc[['15000US060139900000', '15000US060133560024', '15000US060133211033', '15000US060133200041', '15000US060133211014', '15000US060133630004', '15000US060133601012', '15000US060133601012', '15000US060133601022'], 'block_based_district'] = 11
assembler_gdf.loc[['15000US060770052061', '15000US060770052062', '15000US060770051192', '15000US060770051313', '15000US060770051351'], 'block_based_district'] = 9
assembler_gdf.loc[['15000US060770055011', '15000US060770052021', '15000US060770051221'], 'block_based_district'] = 10
assembler_gdf.loc[['15000US060133551083', '15000US060133040011', '15000US060133040021', '15000US060133060032', '15000US060133080011', '15000US060133551081'], 'block_based_district'] = 9
assembler_gdf.loc[['15000US060133551071', '15000US060133551121'], 'block_based_district'] = 11
assembler_gdf.loc[['15000US060133452031', '15000US060133452023', '15000US060133451051'], 'block_based_district'] = 11
assembler_gdf.loc[['15000US060133551171', '15000US060133551161', '15000US060133451131', '15000US060133451142', '15000US060133451141', '15000US060133551143'], 'block_based_district'] = 15
assembler_gdf.loc[['15000US060750179021'], 'block_based_district'] = 12
assembler_gdf.loc[['15000US060759901000', '15000US060750255001', '15000US060750605023', '15000US060750330003', '15000US060750330004', '15000US060750303022', '15000US060750304003' , '15000US060750308005'], 'block_based_district'] = 12
assembler_gdf.loc[['15000US060750263023', '15000US060750255002', '15000US060750306002', '15000US060750308004'], 'block_based_district'] = 14
assembler_gdf.loc[['15000US060014338001', '15000US060014338003'], 'block_based_district'] = 13
assembler_gdf.loc[['15000US060019900000', '15000US060014301021', '15000US060014338002', '15000US060014338004', '15000US060014304002', '15000US060014328003'], 'block_based_district'] = 15
assembler_gdf.loc[['15000US060819901000', '15000US060816138002', '15000US060816117002', '15000US060816117004', '15000US060816096033' , '15000US060816104001', '15000US060816100001', '15000US060816102032', '15000US060816102012', '15000US060816109003'], 'block_based_district'] = 14
assembler_gdf.loc[['15000US060816134003', '15000US060816134002', '15000US060816097001', '15000US060816099001', '15000US060816105001'], 'block_based_district'] = 18
assembler_gdf.loc[['15000US060014507013', '15000US060014415242', '15000US060014419243', '15000US060014415031'], 'block_based_district'] = 15
assembler_gdf.loc[['15000US060014431031', '15000US060014418001', '15000US060014418003', '15000US060014419241'], 'block_based_district'] = 17
assembler_gdf.loc[['15000US060855077023', '15000US060855079041', '15000US060855079042', '15000US060855080043', '15000US060855059003', '15000US060855077031', '15000US060855077032', '15000US060855077034', '15000US060855077035', '15000US060855078052', '15000US060855084015'], 'block_based_district'] = 17
assembler_gdf.loc[['15000US060855117071', '15000US060855047001', '15000US060855076001', '15000US060855100011', '15000US060855058002' , '15000US060855059002'], 'block_based_district'] = 18
assembler_gdf.loc[['15000US060855043081', '15000US060855052031', '15000US060855057003'], 'block_based_district'] = 17
assembler_gdf.loc[['15000US060855135001', '15000US060855042011', '15000US060855051002'], 'block_based_district'] = 19
assembler_gdf.loc[['15000US060855119112', '15000US060855119111', '15000US060855119113', '15000US060855120271', '15000US060855120521', '15000US060855029022', '15000US060855022022' ,'15000US060855022023', '15000US060855020011', '15000US060855058001', '15000US060855020023', '15000US060855026011'], 'block_based_district'] = 18
assembler_gdf.loc[['15000US060855121001', '15000US060855120242', '15000US060855120523', '15000US060855029032', '15000US060855029033', '15000US060855029021', '15000US060855026012', '15000US060855023022', '15000US060855005003', '15000US060855005004', '15000US060855020022'], 'block_based_district'] = 19
assembler_gdf.loc[['15000US060871212002', '15000US060871212001'], 'block_based_district'] = 18
assembler_gdf.loc[['15000US060879901000', '15000US060871212003', '15000US060871004001', '15000US060871005003', '15000US060871012003', '15000US060871207003'], 'block_based_district'] = 20
assembler_gdf.loc[['15000US060855126022', '15000US060855124011', '15000US060855125092', '15000US060855125101'], 'block_based_district'] = 19
assembler_gdf.loc[['15000US060855126021', '15000US060855125051', '15000US060855125062', '15000US060855126043', '15000US060855126032'], 'block_based_district'] = 20
####### Southern California
assembler_gdf.loc[['15000US060710092021'], 'block_based_district'] = 8
assembler_gdf.loc[['15000US060290060073'], 'block_based_district'] = 23
assembler_gdf.loc[['15000US060379012054', '15000US060379012053', '15000US060379007031'], 'block_based_district'] = 23
assembler_gdf.loc[['15000US060379003001', '15000US060379012131', '15000US060379006022', '15000US060379006062', '15000US060379102021', '15000US060379103011'], 'block_based_district'] = 25
assembler_gdf.loc[['15000US061110001001'], 'block_based_district'] = 24
assembler_gdf.loc[['15000US061119901000'], 'block_based_district'] = 26
assembler_gdf.loc[['15000US061110074053'], 'block_based_district'] = 26
assembler_gdf.loc[['15000US061110075111'], 'block_based_district'] = 30
assembler_gdf.loc[['15000US061110075061', '15000US061110075071', '15000US061110075142', '15000US061110079012', '15000US061110084022', '15000US061110075143'], 'block_based_district'] = 25
assembler_gdf.loc[['15000US061110084021', '15000US061110085003', '15000US061110075091', '15000US061110079012'], 'block_based_district'] = 26
assembler_gdf.loc[['15000US060379108111'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060379302002'], 'block_based_district'] = 28
assembler_gdf.loc[['15000US060379110013'], 'block_based_district'] = 25
assembler_gdf.loc[['15000US060379303012'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060379108051', '15000US060379108101'], 'block_based_district'] = 25
assembler_gdf.loc[['15000US061110075121', '15000US060371081021', '15000US060371081011'], 'block_based_district'] = 25
assembler_gdf.loc[['15000US060371132351', '15000US060371082021', '15000US060371066032', '15000US060371066431', '15000US060371082012', '15000US060371112061', '15000US060371112062'], 'block_based_district'] = 30
assembler_gdf.loc[['15000US060378003261', '15000US060378003263'], 'block_based_district'] = 26
assembler_gdf.loc[['15000US060378003242', '15000US060378003262'], 'block_based_district'] = 33
assembler_gdf.loc[['15000US060379301011', '15000US060374604011', '15000US060374617002', '15000US060374639002', '15000US060374639002'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060374608002', '15000US060374639002', '15000US060374639001', '15000US060374637002'], 'block_based_district'] = 28
assembler_gdf.loc[['15000US060710008081', '15000US060710008172'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060374005011', '15000US060374002051', '15000US060374002043', '15000US060374009002', '15000US060374039011'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060374016012', '15000US060374004031', '15000US060374006042', '15000US060374039013', '15000US060374008001'], 'block_based_district'] = 32
assembler_gdf.loc[['15000US060374315022', '15000US060374315023', '15000US060374315021', '15000US060374315022', '15000US060374321012', '15000US060374321025', '15000US060374321011', '15000US060374321022', '15000US060374329021'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060374310022','15000US060374313001', '15000US060374313002', '15000US060374303014', '15000US060374325001', '15000US060374325004', '15000US060374315011'], 'block_based_district'] = 32
assembler_gdf.loc[['15000US060374017011', '15000US060374017012'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060374017041'], 'block_based_district'] = 35
assembler_gdf.loc[['15000US060374824022', '15000US060374825221'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060375300031'], 'block_based_district'] = 38
assembler_gdf.loc[['15000US060375304002'], 'block_based_district'] = 27
assembler_gdf.loc[['15000US060379302001', '15000US060371041243', '15000US060379800261', '15000US060371021031', '15000US060371021042'], 'block_based_district'] = 28
assembler_gdf.loc[['15000US060371041242', '15000US060371032001', '15000US060371222002', '15000US060371211021', '15000US060371211023', '15000US060371032002', '15000US060379800211'], 'block_based_district'] = 29
assembler_gdf.loc[['15000US060371897013', '15000US060373108005', '15000US060373117006', '15000US060371437001'], 'block_based_district'] = 28
assembler_gdf.loc[['15000US060373110002', '15000US060373116004'], 'block_based_district'] = 30
assembler_gdf.loc[['15000US060371923001', '15000US060371944011', '15000US060371944023', '15000US060371924104', '15000US060371926201'], 'block_based_district'] = 28
assembler_gdf.loc[['15000US060371864041', '15000US060371924103', '15000US060371925201', '15000US060371925202', '15000US060371927002', '15000US060371958021', '15000US060371957103', '15000US060371926202'], 'block_based_district'] = 34
assembler_gdf.loc[['15000US060371093003', '15000US060371114001', '15000US060379800081', '15000US060371252003', '15000US060371284004', '15000US060371285002'], 'block_based_district'] = 29
assembler_gdf.loc[['15000US060371066031', '15000US060379800221', '15000US060371311002', '15000US060371311003', '15000US060371321022', '15000US060371236013', '15000US060371251001', '15000US060371255021', '15000US060371255022', '15000US060371256001', '15000US060371433004'], 'block_based_district'] = 30
assembler_gdf.loc[['15000US060371415001', '15000US060378002032'], 'block_based_district'] = 30
assembler_gdf.loc[['15000US060378002041'], 'block_based_district'] = 33
assembler_gdf.loc[['15000US060374013031', '15000US060374013042'], 'block_based_district'] = 32
assembler_gdf.loc[['15000US060374084023', '15000US060374083031', '15000US060374339022', '15000US060374340031', '15000US060374340032', '15000US060374331012', '15000US060374331021', '15000US060374334011', '15000US060374334012', '15000US060374334022', '15000US060374335043'], 'block_based_district'] = 32
assembler_gdf.loc[['15000US060375003001', '15000US060374083021', '15000US060374338012', '15000US060374335041'], 'block_based_district'] = 38
assembler_gdf.loc[['15000US060374082022', '15000US060374082111', '15000US060374035001'], 'block_based_district'] = 32
assembler_gdf.loc[['15000US060374082112', '15000US060374033031', '15000US060374082121', '15000US060374082122'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060372756021'], 'block_based_district'] = 43
assembler_gdf.loc[['15000US060372741001', '15000US060372753022', '15000US060372737003'], 'block_based_district'] = 33
assembler_gdf.loc[['15000US060372737001', '15000US060372738001', '15000US060372712002', '15000US060372713003', '15000US060372655101', '15000US060372656022', '15000US060372656021', '15000US060372657001', '15000US060372657003', '15000US060372676002', '15000US060372676003'], 'block_based_district'] = 37
assembler_gdf.loc[['15000US060376503005', '15000US060376507012', '15000US060376511021', '15000US060376514011', '15000US060376707011'], 'block_based_district'] = 33
assembler_gdf.loc[['15000US060372766011', '15000US060379800281'], 'block_based_district'] = 43
assembler_gdf.loc[['15000US060379903000'], 'block_based_district'] = 44
assembler_gdf.loc[['15000US060372964021', '15000US060372964023'], 'block_based_district'] = 33
assembler_gdf.loc[['15000US060372214021', '15000US060372242002'], 'block_based_district'] = 34
assembler_gdf.loc[['15000US060375309011', '15000US060375309022'], 'block_based_district'] = 34
assembler_gdf.loc[['15000US060375310001', '15000US060372260012', '15000US060375311013'], 'block_based_district'] = 40
assembler_gdf.loc[['15000US060710004013'], 'block_based_district'] = 35
assembler_gdf.loc[['15000US060374033213', '15000US060710001051', '15000US060374024041', '15000US060710001132', '15000US060710001152'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060372240201', '15000US060372240202', '15000US060372246002', '15000US060372311002'], 'block_based_district'] = 37
assembler_gdf.loc[['15000US060372318001'], 'block_based_district'] = 40
assembler_gdf.loc[['15000US060372381002', '15000US060372382002', '15000US060372382003', '15000US060372756031', '15000US060372756033', '15000US060372761003'], 'block_based_district'] = 37
assembler_gdf.loc[['15000US060372384001', '15000US060372384003', '15000US060372383102', '15000US060372383201'], 'block_based_district'] = 43
assembler_gdf.loc[['15000US060375016001', '15000US060375002025', '15000US060375002012'], 'block_based_district'] = 38
assembler_gdf.loc[['15000US060591103011', '15000US060374085033', '15000US060375002011'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060375531003', '15000US060375302021', '15000US060375302023', '15000US060375531004', '15000US060375541051', '15000US060375541052', '15000US060375542043', '15000US060375543013'], 'block_based_district'] = 38
assembler_gdf.loc[['15000US060375531001', '15000US060375318003', '15000US060375319021', '15000US060375542011'], 'block_based_district'] = 40
assembler_gdf.loc[['15000US060591101023'], 'block_based_district'] = 38
assembler_gdf.loc[['15000US060375715031', '15000US060591101111', '15000US060591103041'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060590218262', '15000US060590219152', '15000US060590219151'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060590219241', '15000US060590219243', '15000US060590758131', '15000US060590758133'], 'block_based_district'] = 45
assembler_gdf.loc[['15000US060591104023'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060591102011'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060590117222', '15000US060591104011', '15000US060590117202', '15000US060590116021', '15000US060590116022', '15000US060590868011', '15000US060590868032', '15000US060591104022'], 'block_based_district'] = 39
assembler_gdf.loc[['15000US060590117141', '15000US060590762022', '15000US060590864073', '15000US060590867011', '15000US060590868021', '15000US060590868013', '15000US060590117201', '15000US060590117203'], 'block_based_district'] = 46
assembler_gdf.loc[['15000US060591103012', '15000US060591103013'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060375351012'], 'block_based_district'] = 40
assembler_gdf.loc[['15000US060375353001', '15000US060375353003'], 'block_based_district'] = 44
assembler_gdf.loc[['15000US060372410013', '15000US060372911102'], 'block_based_district'] = 43
assembler_gdf.loc[['15000US060375435011'], 'block_based_district'] = 44
assembler_gdf.loc[['15000US060375706021', '15000US060375706022', '15000US060375706024', '15000US060375717033'], 'block_based_district'] = 44
assembler_gdf.loc[['15000US060375440021', '15000US060375723012'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060590755151'], 'block_based_district'] = 45
assembler_gdf.loc[['15000US060590755041', '15000US060590755052'], 'block_based_district'] = 45
assembler_gdf.loc[['15000US060590744063', '15000US060590754033', '15000US060590757012', '15000US060590757013', '15000US060590758132'], 'block_based_district'] = 46
assembler_gdf.loc[['15000US060590626143', '15000US060590626213', '15000US060590423331', '15000US060590626493', '15000US060590423202', '15000US060590423201'], 'block_based_district'] = 45
assembler_gdf.loc[['15000US060590626042', '15000US060590626102', '15000US060590626411', '15000US060590626412', '15000US060590626343', '15000US060590423352', '15000US060590423351', '15000US060590626361'], 'block_based_district'] = 48
assembler_gdf.loc[['15000US060590320411', '15000US060590320222', '15000US060590320431', '15000US060590320432'], 'block_based_district'] = 45
assembler_gdf.loc[['15000US060590320564', '15000US060590320223', '15000US060590320534', '15000US060590320563', '15000US060590320572'], 'block_based_district'] = 49
assembler_gdf.loc[['15000US060590878063', '15000US060590883012', '15000US060591102031', '15000US060590878011', '15000US060590878021', '15000US060590878061', '15000US060590878062'], 'block_based_district'] = 46
assembler_gdf.loc[['15000US060591102021', '15000US060590878031', '15000US060590878051', '15000US060590883021', '15000US060591102022', '15000US060591102023'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060590741061', '15000US060590741062', '15000US060590741082', '15000US060590748032'], 'block_based_district'] = 46
assembler_gdf.loc[['15000US060590996031', '15000US060591100073', '15000US060591100082', '15000US060590996021', '15000US060590997011', '15000US060590997012', '15000US060590997013', '15000US060591100081'], 'block_based_district'] = 47
assembler_gdf.loc[['15000US060590890011', '15000US060590996031', '15000US060590997022'], 'block_based_district'] = 48
assembler_gdf.loc[['15000US060599901000', '15000US060590423051', '15000US060590423052', '15000US060590423151', '15000US060590423152', '15000US060590423153', '15000US060590423155'], 'block_based_district'] = 48
assembler_gdf.loc[['15000US060590423241', '15000US060590423111', '15000US060590423113'], 'block_based_district'] = 49
####### Rest of state
assembler_gdf.loc[['15000US060210103002', '15000US060210105013'], 'block_based_district'] = 1
assembler_gdf.loc[['15000US060210102003' , '15000US060210102004', '15000US060210104007'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US060570009003', '15000US060610213041', '15000US060610215012', '15000US060610218022', '15000US060610218023', '15000US060610215022'], 'block_based_district'] = 1
assembler_gdf.loc[['15000US060610220141', '15000US060610213042', '15000US060610213043', '15000US060610205011', '15000US060610203001', '15000US060610203003', '15000US060610215021'], 'block_based_district'] = 4
assembler_gdf.loc[['15000US060971534031', '15000US060971534032', '15000US060971535013', '15000US060971506072', '15000US060971506073', '15000US060971527022', '15000US060971527024'], 'block_based_district'] = 2
assembler_gdf.loc[['15000US060971530052', '15000US060971512014', '15000US060971535011', '15000US060971526001', '15000US060971524003', '15000US060971527021', '15000US060971529062', '15000US060971526005'], 'block_based_district'] = 5
assembler_gdf.loc[['15000US060330001002', '15000US060330010002', '15000US060330004005'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US060330012001', '15000US060330012002', '15000US060330012003', '15000US060330009001', '15000US060330009002', '15000US060330009003', '15000US060330003001', '15000US060330004001', '15000US060330004002', '15000US060330004003', '15000US060330004004'], 'block_based_district'] = 5
assembler_gdf.loc[['15000US060670096182'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US060670099004', '15000US060670040121', '15000US061130104012', '15000US061130101021', '15000US060670074172', '15000US060670074312'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US061130104021', '15000US060670096011', '15000US060670070011', '15000US060670070191', '15000US060670071011', '15000US060670071064', '15000US060670074271', '15000US060670074301', '15000US061130102031'], 'block_based_district'] = 6
assembler_gdf.loc[['15000US060670096381'], 'block_based_district'] = 7
assembler_gdf.loc[['15000US060670095011'], 'block_based_district'] = 9
assembler_gdf.loc[['15000US060670096382'], 'block_based_district'] = 3
assembler_gdf.loc[['15000US060190064033'], 'block_based_district'] = 4
assembler_gdf.loc[['15000US060190064031', '15000US060190064034'], 'block_based_district'] = 22
assembler_gdf.loc[['15000US060670092011', '15000US060670048013', '15000US060670048011', '15000US060670048012', '15000US060670048022', '15000US060670052043', '15000US060670054043', '15000US060670054045'], 'block_based_district'] = 6
assembler_gdf.loc[['15000US060670092012', '15000US060670050022', '15000US060670047012', '15000US060670096141', '15000US060670096142', '15000US060670059011', '15000US060670056011', '15000US060670056012', '15000US060670056013', '15000US060670056051', '15000US060670056061', '15000US060670081271', '15000US060670081272', '15000US060670081291', '15000US060670081303', '15000US060670081311'], 'block_based_district'] = 7
assembler_gdf.loc[['15000US060670094081'], 'block_based_district'] = 7
assembler_gdf.loc[['15000US060670094073'], 'block_based_district'] = 9
assembler_gdf.loc[['15000US060710108021', '15000US060710110021', '15000US060710111014', '15000US060710085005', '15000US060710086022', '15000US060710087064', '15000US060710086012', '15000US060710076043', '15000US060710079043', '15000US060710079013', '15000US060710079014'], 'block_based_district'] = 8
assembler_gdf.loc[['15000US060710020101', '15000US060710020191', '15000US060710020211', '15000US060710020222', '15000US060710045031', '15000US060710061007', '15000US060710085003', '15000US060710085005', '15000US060710086013', '15000US060710086021'], 'block_based_district'] = 31
assembler_gdf.loc[['15000US060710074092', '15000US060710074093', '15000US060710076031', '15000US060710065001', '15000US060710064021', '15000US060710064022', '15000US060710074091', '15000US060710074072', '15000US060710074082'], 'block_based_district'] = 8
assembler_gdf.loc[['15000US060710074032', '15000US060710074042', '15000US060710063023', '15000US060710074101', '15000US060710074103', '15000US060710074081', '15000US060710074102', '15000US060710076013'], 'block_based_district'] = 31
assembler_gdf.loc[['15000US060770049022', '15000US060770050031'], 'block_based_district'] = 9
assembler_gdf.loc[['15000US060770050032', '15000US060770049011', '15000US060770049013', '15000US060770049023'], 'block_based_district'] = 10
assembler_gdf.loc[['15000US060190019003', '15000US060190010002', '15000US060190041001', '15000US060190041002'], 'block_based_district'] = 16
assembler_gdf.loc[['15000US060190015002', '15000US060190076001', '15000US060190018001', '15000US060190015001'], 'block_based_district'] = 21
assembler_gdf.loc[['15000US060190046021' , '15000US060190046023'], 'block_based_district'] = 16
assembler_gdf.loc[['15000US060190031043', '15000US060190042054', '15000US060190046011', '15000US060190014143', '15000US060190014143'], 'block_based_district'] = 22
assembler_gdf.loc[['15000US060190068022', '15000US060190068023', '15000US060190060001', '15000US060190062012', '15000US060190062024'], 'block_based_district'] = 21
assembler_gdf.loc[['15000US060190062021'], 'block_based_district'] = 22
assembler_gdf.loc[['15000US060290032041', '15000US060290037002', '15000US060290032024', '15000US060290063041', '15000US060290013002', '15000US060290013006', '15000US060290014002', '15000US060290012011', '15000US060290012012', '15000US060290009061', '15000US060290014004'], 'block_based_district'] = 21
assembler_gdf.loc[['15000US060290027001', '15000US060290007004', '15000US060290014003'], 'block_based_district'] = 23
assembler_gdf.loc[['15000US061070006004', '15000US061070008001', '15000US061070033002', '15000US061070033003', '15000US061070007012', '15000US061070025004', '15000US061070028001', '15000US061070028002'], 'block_based_district'] = 22
assembler_gdf.loc[['15000US061070001003', '15000US061070014003', '15000US061070014005', '15000US061070014002', '15000US061070014004', '15000US061070025002', '15000US061070025003'], 'block_based_district'] = 23
assembler_gdf.loc[['15000US061110012061', '15000US061110024001', '15000US061110025002'], 'block_based_district'] = 24
assembler_gdf.loc[['15000US061110009033', '15000US061110012041', '15000US061110023003', '15000US061110028004', '15000US061110028005', '15000US061110025004'], 'block_based_district'] = 26
assembler_gdf.loc[['15000US060710040042', '15000US060710036122', '15000US060710040042', '15000US060710023042', '15000US060710022071'], 'block_based_district'] = 31
assembler_gdf.loc[['15000US060710034042', '15000US060710039001'], 'block_based_district'] = 35
assembler_gdf.loc[['15000US060650438221'], 'block_based_district'] = 41
assembler_gdf.loc[['15000US060650438231', '15000US060650438232'], 'block_based_district'] = 36
assembler_gdf.loc[['15000US060650444021', '15000US060650444031', '15000US060650438121', '15000US060650438201', '15000US060650435172', '15000US060650433041', '15000US060650433043', '15000US060650433044', '15000US060650433173', '15000US060650437011'], 'block_based_district'] = 36
assembler_gdf.loc[['15000US060650432393', '15000US060650427441', '15000US060650427451', '15000US060650427452', '15000US060650427232'], 'block_based_district'] = 42
assembler_gdf.loc[['15000US060650406031', '15000US060650406072', '15000US060650410043', '15000US060650420081', '15000US060650429022', '15000US060650420072', '15000US060650429031', '15000US060650420091', '15000US060650420092', '15000US060650414041', '15000US060650414122', '15000US060650420051'], 'block_based_district'] = 41
assembler_gdf.loc[['15000US060650420071', '15000US060650429021', '15000US060650414091', '15000US060650420082', '15000US060650414092', '15000US060650420091', '15000US060650420041', '15000US060650414093', '15000US060650420052', '15000US060650420031'], 'block_based_district'] = 42
assembler_gdf.loc[['15000US060650427312', '15000US060650490002'], 'block_based_district'] = 41
assembler_gdf.loc[['15000US060650427281', '15000US060650427091', '15000US060650427092', '15000US060650427321'], 'block_based_district'] = 42
assembler_gdf.loc[['15000US060650426202', '15000US060650426241', '15000US060650429011'], 'block_based_district'] = 41
assembler_gdf.loc[['15000US060650427191', '15000US060650427301', '15000US060650426231', '15000US060650427311', '15000US060650490003'], 'block_based_district'] = 42
assembler_gdf.loc[['15000US060650432183', '15000US060650512001', '15000US060650512002', '15000US060650432571', '15000US060650432572'], 'block_based_district'] = 42
assembler_gdf.loc[['15000US060650432672', '15000US060650432671', '15000US060650432462', '15000US060650432653', '15000US060650432522'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730190011', '15000US060730193033', '15000US060730186121', '15000US060730171102', '15000US060730200132', '15000US060730203061', '15000US060739901000'], 'block_based_district'] = 49
assembler_gdf.loc[['15000US060730199032', '15000US060730200191', '15000US060730200193', '15000US060730200271', '15000US060730192071', '15000US060730192082', '15000US060730199031', '15000US060730203071', '15000US060730203072'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730171101', '15000US060730171061', '15000US060730171062', '15000US060730173042', '15000US060730170292', '15000US060730083051', '15000US060730170321', '15000US060730173062', '15000US060730083121', '15000US060730083391', '15000US060730173061', '15000US060730170301'], 'block_based_district'] = 49
assembler_gdf.loc[['15000US060730083123', '15000US060730170291'], 'block_based_district'] = 52
assembler_gdf.loc[['15000US060730210001', '15000US060730212041', '15000US060730212043'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730211001', '15000US060730211002', '15000US060730212022', '15000US060730213022'], 'block_based_district'] = 51
assembler_gdf.loc[['15000US060730134191'], 'block_based_district'] = 53
assembler_gdf.loc[['15000US060730166061', '15000US060730208071', '15000US060730208012'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730207091', '15000US060730207062', '15000US060730207063', '15000US060730204011', '15000US060730204012', '15000US060730208011'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730207101', '15000US060730170101', '15000US060730170201', '15000US060730170212'], 'block_based_district'] = 52
assembler_gdf.loc[['15000US060730166052', '15000US060730166162', '15000US060730136051', '15000US060730136052', '15000US060730154041', '15000US060730154044'], 'block_based_district'] = 50
assembler_gdf.loc[['15000US060730213032', '15000US060730162021', '15000US060730163011', '15000US060730154031', '15000US060730154043'], 'block_based_district'] = 53
assembler_gdf.loc[['15000US060730051002', '15000US060730052001', '15000US060730099021', '15000US060730106012'], 'block_based_district'] = 52
assembler_gdf.loc[['15000US060730213021', '15000US060730100141', '15000US060730100142', '15000US060730032042', '15000US060730027034', '15000US060730027121', '15000US060730030031', '15000US060730031131', '15000US060730041004'], 'block_based_district'] = 51
assembler_gdf.loc[['15000US060730133141', '15000US060730032041', '15000US060730032071', '15000US060730032121', '15000US060730134121', '15000US060730016002', '15000US060730031132', '15000US060730041002', '15000US060730046002'], 'block_based_district'] = 53
assembler_gdf.loc[['15000US060730098052', '15000US060730089021', '15000US060730056001', '15000US060730065001', '15000US060730098013', '15000US060730098025', '15000US060730095112', '15000US060730095092', '15000US060730095073'], 'block_based_district'] = 52
assembler_gdf.loc[['15000US060730086001', '15000US060730090002', '15000US060730056002', '15000US060730059003', '15000US060730065002', '15000US060730065003', '15000US060730098011', '15000US060730098021', '15000US060730095091', '15000US060730097042'], 'block_based_district'] = 53
e = time.time()
g = g + (e-s)
print('Grand total time: {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor(g/60), np.floor(g%60)))
# print(assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == -1)
# ].groupby('congressional_districts_bitmask').size().sort_index(ascending=False))
#assembler_gdf.groupby('STUSAB').size()
# -
# # WRITE OUTPUT
# +
s = time.time()
pandas.DataFrame(assembler_gdf[[x for x in assembler_gdf.columns if x != 'geometry']]
).to_csv(output_dir+'block_group_with_district_all.csv', encoding='utf-8')
e = time.time()
g = g + (e-s)
print('matched districts for {0:,.0f} block groups in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(assembler_gdf), np.floor((e-s)/60), np.floor((e-s)%60)))
print('GRAND TOTAL TIME: {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor(g/60), np.floor(g%60)))
# -
# # TEST that output wrote correctly
# +
s = time.time()
z = pandas.read_csv(output_dir+'block_group_with_district_all.csv', index_col='GEOID')
bg_file_list = [shapefiledir+'BG/'+x for x in os.listdir(shapefiledir+'BG/') if ((x[-4:] == '.shp'))]
bg_gdf = geopandas.GeoDataFrame()
for i in range(0, len(bg_file_list)):
if (debug >= 1):
if ((np.mod(i,10) == 0) | (i == len(bg_file_list)-1)):
print('\tReading file {0:,.0f} of {1:,.0f}...'.format(i+1, len(bg_file_list)))
bg_gdf_i = geopandas.read_file(bg_file_list[i])
bg_gdf = pandas.concat((bg_gdf, bg_gdf_i), axis=0, sort=False)
e = time.time()
print('done in {0:,.1f} seconds'.format(e-s))
s = time.time()
print('converting block group identifiers to numeric...')
bg_gdf.loc[:, 'STATEFP'] = pandas.to_numeric(bg_gdf['STATEFP'], errors='coerce')
bg_gdf.loc[:, 'COUNTYFP'] = pandas.to_numeric(bg_gdf['COUNTYFP'], errors='coerce')
bg_gdf.loc[:, 'TRACTCE'] = pandas.to_numeric(bg_gdf['TRACTCE'].apply(lambda x: str(x)[0:4]+'.'+str(x)[4:]), errors='coerce')
bg_gdf.loc[:, 'BLKGRPCE'] = pandas.to_numeric(bg_gdf['BLKGRPCE'], errors='coerce')
print('assigning GEOID as index...')
bg_gdf.loc[:, 'GEOID'] = bg_gdf['GEOID'].apply(lambda x: '15000US'+str(x))#
bg_gdf = bg_gdf.set_index('GEOID')
newtestgdf = geopandas.GeoDataFrame(data=z.join(bg_gdf.geometry, how='left'), crs=bg_gdf.crs, geometry='geometry')
e = time.time()
print('done in {0:,.1f} seconds'.format(e-s))
# -
fig,ax = plt.subplots(1,1)
newtestgdf.plot(ax=ax)
plt.xlim(-130,-60)
plt.ylim(20,50)
plt.show()
# # CALIFORNIA|
# ## Get water areas for whole state
# +
s = time.time()
this_state = 'CA'
this_state_number = 6
#this_state_number = state_codes_df[state_codes_df['STUSAB'] == this_state.upper()].index.values[0]
if (debug >= 1):
print('reading area water shapefiles in {0:}...'.format(this_state))
water_gdf = geopandas.GeoDataFrame()
water_file_list = [shapefiledir+'AREAWATER/'+x for x in os.listdir(shapefiledir+'AREAWATER/') if ((x[-4:] == '.shp') and ('tl_2018_{0:02d}'.format(this_state_number) in x))]
for i in range(0, len(water_file_list)):
if (debug >= 1):
if ((np.mod(i,10) == 0) | (i == len(water_file_list)-1)):
print('\tReading file {0:,.0f} of {1:,.0f}...'.format(i+1, len(water_file_list)))
water_gdf_i = geopandas.read_file(water_file_list[i])
#water_gdf_i = water_gdf_i[water_gdf_i['AWATER'] >= water_area_tol]
water_gdf = pandas.concat((water_gdf, water_gdf_i), axis=0, sort=False)
water_gdf = water_gdf.set_index('HYDROID')
e = time.time()
g = g + (e-s)
if (debug >= 1):
#print('Read {0:,.0f} bodies of water with area greater than or equal to {1:,.0f} km^2 in {2:,.0f} seconds!'.format(len(water_gdf), water_area_tol/(1000*1000), e-s))
print('Read {0:,.0f} bodies of linear water in {1:,.0f} seconds!'.format(len(water_gdf), e-s))
#print(assembler_gdf.groupby(['STUSAB', 'block_based_district']).size())
s = time.time()
if (debug >= 1):
print('reading place shapefiles in {0:}...'.format(this_state))
place_gdf = geopandas.GeoDataFrame()
place_file_list = [shapefiledir+'PLACE/'+x for x in os.listdir(shapefiledir+'PLACE/') if ((x[-4:] == '.shp') and ('tl_2018_{0:02d}'.format(this_state_number) in x))]
for i in range(0, len(place_file_list)):
if (debug >= 1):
if ((np.mod(i,10) == 0) | (i == len(water_file_list)-1)):
print('\tReading file {0:,.0f} of {1:,.0f}...'.format(i+1, len(place_file_list)))
place_gdf_i = geopandas.read_file(place_file_list[i])
place_gdf = pandas.concat((place_gdf, place_gdf_i), axis=0, sort=False)
place_gdf.loc[:, 'INTPTLON'] = pandas.to_numeric(place_gdf['INTPTLON'], errors='coerce')
place_gdf.loc[:, 'INTPTLAT'] = pandas.to_numeric(place_gdf['INTPTLAT'], errors='coerce')
place_gdf = place_gdf.set_index('GEOID')
e = time.time()
g = g + (e-s)
if (debug >= 1):
#print('Read {0:,.0f} bodies of water with area greater than or equal to {1:,.0f} km^2 in {2:,.0f} seconds!'.format(len(water_gdf), water_area_tol/(1000*1000), e-s))
print('Read {0:,.0f} places in {1:,.1f} seconds!'.format(len(place_gdf), e-s))
#place_gdf.head(1)
s = time.time()
if (debug >= 1):
print('reading roads shapefiles in {0:}...'.format(this_state))
roads_gdf = geopandas.GeoDataFrame()
roads_file_list = [shapefiledir+'ROADS/'+x for x in os.listdir(shapefiledir+'ROADS/') if ((x[-4:] == '.shp') and ('tl_2018_{0:02d}'.format(this_state_number) in x))]
for i in range(0, len(roads_file_list)):
if (debug >= 1):
if ((np.mod(i,10) == 0) | (i == len(water_file_list)-1)):
print('\tReading file {0:,.0f} of {1:,.0f}...'.format(i+1, len(roads_file_list)))
roads_gdf_i = geopandas.read_file(roads_file_list[i])
roads_gdf = pandas.concat((roads_gdf, roads_gdf_i), axis=0, sort=False)
roads_gdf = roads_gdf.set_index('LINEARID')
# #roads_gdf.head(1).T
# # Road Types: C = County, I = Interstate, M = Common name, O = Other, S = State hwy, U = US hwy
old_map_buffer_ratio = -1
e = time.time()
g = g + (e-s)
if (debug >= 1):
print('Read {0:,.0f} roads in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(roads_gdf), np.floor((e-s)/60), np.floor((e-s)%60)))
# s = time.time()
# water_gdf.to_file(shapefiledir+'texas/water_texas.shp')
# place_gdf.to_file(shapefiledir+'texas/place_texas.shp')
# roads_gdf.to_file(shapefiledir+'texas/roads_texas.shp')
# e = time.time()
print('Got California water/place/road shapefiles in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# print('Wrote outfiles in {0:,.0f} seconds!'.format(e-s))
# -
# ## Define Bay Area rectangle
# +
this_state = 'CA'
nDistricts = cd_gdf[cd_gdf['STUSAB'] == this_state]['CD116FP'].max()
color_reset_point = 18
#### BAY AREA
s = time.time()
print('Defining Bay Area...')
bay_area_west_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'San Mateo County')].geometry.tolist()).bounds[0] - 0.025 # West border of San mateo County
bay_area_north_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Napa County')].geometry.tolist()).bounds[1] + 0.025 # South border of Napa County
bay_area_east_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Santa Clara County')].geometry.tolist()).bounds[2] + 0.025 # East border of Santa Clara
bay_area_south_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Santa Clara County')].geometry.tolist()).bounds[1] - 0.025 # South border of Santa Clara County
xlimits = [bay_area_west_limit, bay_area_east_limit]
ylimits = [bay_area_south_limit,bay_area_north_limit]
print('\tMaking box...')
bay_area_geo = makebox(xlimits, ylimits)
print('\tMaking dataframe...')
bay_area_gdf = geopandas.GeoDataFrame(data=[[bay_area_geo]], columns=['geometry'], crs=assembler_gdf.crs, geometry='geometry')
print('\tFinding block groups in Bay Area...')
if ('in_bay_area' not in assembler_gdf.columns):
assembler_gdf = assembler_gdf.assign(in_bay_area = False)
assembler_gdf.loc[(
(assembler_gdf['STUSAB'] == this_state)
& (assembler_gdf.geometry.intersects(bay_area_geo))
), 'in_bay_area'] = True
print('\tFinding congressional districts in Bay Area...')
bay_area_districts = cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf.geometry.intersects(bay_area_geo))]['CD116FP'].drop_duplicates().sort_values().tolist()
e = time.time()
print('Found {0:,.0f} block groups in the Bay Area in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(assembler_gdf[assembler_gdf['in_bay_area'] == True]), np.floor((e-s)/60), np.floor((e-s)%60)))
s = time.time()
print('Dividing water areas, places, and roads...')
water_bay_area_gdf = water_gdf[water_gdf.geometry.apply(lambda x: x.intersects(bay_area_geo))]
print('Plotting...')
legend_location = 'upper right'
fig, ax = plt.subplots(1,1,figsize=(12,8))
legend_list = []
for i in bay_area_districts:
cd_gdf[
(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)
].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], edgecolor='white', lw=1)
legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1], label='District {0:,.0f}'.format(i)))
print('labeling districts...')
for ix, thisrow in cd_gdf[
(cd_gdf['STUSAB'] == this_state)
& (cd_gdf['CD116FP'].isin(bay_area_districts))
& ((cd_gdf['INTPTLON'] >= xlimits[0]) & (cd_gdf['INTPTLON'] <= xlimits[1]))
& ((cd_gdf['INTPTLAT'] >= ylimits[0]) & (cd_gdf['INTPTLAT'] <= ylimits[1]))
].iterrows():
annotator = thisrow['CD116FP']
showpoint = (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT']))
plt.annotate(annotator, showpoint, showpoint,
fontsize=24, color='black', backgroundcolor='white', ha='center', va='center')
print('plotting block groups...')
assembler_gdf[(assembler_gdf['in_bay_area'] == True)].plot(ax=ax, color='none', edgecolor='black', lw=0.05)
#ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
water_bay_area_gdf.plot(ax=ax, color='blue')
bay_area_gdf.plot(ax=ax, color='none', edgecolor='yellow', lw=1)
plt.xlim(xlimits)
plt.ylim(ylimits)
plt.show()
# -
# ## Define Southern California rectangle
# +
this_state = 'CA'
nDistricts = cd_gdf[cd_gdf['STUSAB'] == this_state]['CD116FP'].max()
color_reset_point = 18
#### SOUTHERN CALIFORNIA
s = time.time()
print('Defining Southern California..')
so_cal_west_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Los Angeles County')].geometry.tolist()).bounds[0] - 0.025 # West border of Ventura County
so_cal_north_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Los Angeles County')].geometry.tolist()).bounds[3] + 0.025 # North border of Los Angeles County
so_cal_east_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Los Angeles County')].geometry.tolist()).bounds[2] + 0.025 # East border of Los Angeles County
so_cal_south_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['COUNTY_NAME'] == 'Los Angeles County')].geometry.tolist()).bounds[1] - 0.025 # South border of Los Angeles County
#so_cal_south_limit = unary_union(assembler_gdf[(assembler_gdf['STUSAB'] == this_state)].geometry.tolist()).bounds[1] - 0.025 # South border of California
xlimits = [so_cal_west_limit, so_cal_east_limit]
ylimits = [so_cal_south_limit, so_cal_north_limit]
print('\tMaking box...')
so_cal_geo = makebox(xlimits, ylimits)
print('\tMaking dataframe...')
so_cal_gdf = geopandas.GeoDataFrame(data=[[so_cal_geo]], columns=['geometry'], crs=assembler_gdf.crs, geometry='geometry')
print('\tFinding block groups in Bay Area...')
if ('in_so_cal' not in assembler_gdf.columns):
assembler_gdf = assembler_gdf.assign(in_so_cal = False)
assembler_gdf.loc[(
(assembler_gdf['STUSAB'] == this_state)
& (assembler_gdf.geometry.intersects(so_cal_geo))
), 'in_so_cal'] = True
print('\tFinding congressional districts in Southern California...')
so_cal_districts = cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf.geometry.intersects(so_cal_geo))]['CD116FP'].drop_duplicates().sort_values().tolist()
e = time.time()
print('Found {0:,.0f} block groups in Southern California in {1:,.0f} minutes {2:,.0f} seconds!'.format(len(assembler_gdf[assembler_gdf['in_so_cal'] == True]), np.floor((e-s)/60), np.floor((e-s)%60)))
s = time.time()
print('Dividing water areas, places, and roads...')
water_so_cal_gdf = water_gdf[water_gdf.geometry.apply(lambda x: x.intersects(so_cal_geo))]
place_so_cal_gdf = place_gdf[place_gdf.geometry.apply(lambda x: x.intersects(so_cal_geo))]
roads_so_cal_gdf = roads_gdf[roads_gdf.geometry.apply(lambda x: x.intersects(so_cal_geo))]
print('Plotting...')
legend_location = 'upper right'
fig, ax = plt.subplots(1,1,figsize=(12,12))
legend_list = []
for i in so_cal_districts:
cd_gdf[
(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)
].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], edgecolor='white', lw=1)
legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1], label='District {0:,.0f}'.format(i)))
print('labeling districts...')
for ix, thisrow in cd_gdf[
(cd_gdf['STUSAB'] == this_state)
& (cd_gdf['CD116FP'].isin(so_cal_districts))
& ((cd_gdf['INTPTLON'] >= xlimits[0]) & (cd_gdf['INTPTLON'] <= xlimits[1]))
& ((cd_gdf['INTPTLAT'] >= ylimits[0]) & (cd_gdf['INTPTLAT'] <= ylimits[1]))
].iterrows():
annotator = thisrow['CD116FP']
showpoint = (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT']))
plt.annotate(annotator, showpoint, showpoint,
fontsize=14, color='black', backgroundcolor='white', ha='center', va='center')
print('plotting block groups...')
assembler_gdf[(assembler_gdf['in_so_cal'] == True)].plot(ax=ax, color='none', edgecolor='black', lw=0.05)
#ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
water_so_cal_gdf.plot(ax=ax, color='blue')
so_cal_gdf.plot(ax=ax, color='none', edgecolor='pink', lw=1)
plt.xlim(xlimits)
plt.ylim(ylimits)
plt.show()
# -
# ## Define regions outside Bay Area & Southern California rectangles
# +
if ('in_neither' not in assembler_gdf.columns):
assembler_gdf = assembler_gdf.assign(in_neither = False)
assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['in_bay_area'] == False) & (assembler_gdf['in_so_cal']), 'in_neither'] = True
this_state = 'CA'
nDistricts = cd_gdf[cd_gdf['STUSAB'] == 'CA']['CD116FP'].max()
color_reset_point = 18
legend_location = 'upper right'
print('splitting water...')
water_other_gdf = water_gdf[~(water_gdf.index.isin(water_bay_area_gdf.index) & ~(water_gdf.index.isin(water_so_cal_gdf.index)))]
print('splitting places...')
place_other_gdf = place_gdf[~(place_gdf.index.isin(water_bay_area_gdf.index) & ~(place_gdf.index.isin(place_so_cal_gdf.index)))]
print('splitting roads...')
roads_other_gdf = roads_gdf[~(roads_gdf.index.isin(water_bay_area_gdf.index) & ~(roads_gdf.index.isin(roads_so_cal_gdf.index)))]
fig, ax = plt.subplots(1,1,figsize=(12,8))
#legend_list = []
for i in range(1,nDistricts+1):
cd_gdf[
(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)
].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], edgecolor='white')
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1], label='District {0:,.0f}'.format(i)))
assembler_gdf[
(assembler_gdf['STUSAB'] == this_state)
& (assembler_gdf['in_bay_area'] == True)
].plot(ax=ax, color='none', edgecolor='pink', lw=0.1)
assembler_gdf[
(assembler_gdf['STUSAB'] == this_state)
& (assembler_gdf['in_so_cal'] == True)
].plot(ax=ax, color='none', edgecolor='yellow', lw=0.1)
assembler_gdf[
(assembler_gdf['STUSAB'] == this_state)
& (assembler_gdf['in_neither'] == True)
].plot(ax=ax, color='none', edgecolor='black', lw=0.1)
bay_area_gdf.plot(ax=ax, color='none', edgecolor='pink', lw=3)
so_cal_gdf.plot(ax=ax, color='none', edgecolor='yellow', lw=3)
#ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
plt.show()
# -
# ## BAY AREA
# +
# assembler_gdf.loc[['15000US060971506122'], 'block_based_district'] = 5
# assembler_gdf.loc[['15000US060952522024', '15000US060952522014'], 'block_based_district'] = 3
# assembler_gdf.loc[['15000US060952521021', '15000US060952522021'], 'block_based_district'] = 5
# assembler_gdf.loc[['15000US060133180002', '15000US060133560022', '15000US060133180003', '15000US060133601013', '15000US060133592022'], 'block_based_district'] = 5
# assembler_gdf.loc[['15000US060139900000', '15000US060133560024', '15000US060133211033', '15000US060133200041', '15000US060133211014', '15000US060133630004', '15000US060133601012', '15000US060133601012', '15000US060133601022'], 'block_based_district'] = 11
# assembler_gdf.loc[['15000US060770052061', '15000US060770052062', '15000US060770051192', '15000US060770051313', '15000US060770051351'], 'block_based_district'] = 9
# assembler_gdf.loc[['15000US060770055011', '15000US060770052021', '15000US060770051221'], 'block_based_district'] = 10
# assembler_gdf.loc[['15000US060133551083', '15000US060133040011', '15000US060133040021', '15000US060133060032', '15000US060133080011', '15000US060133551081'], 'block_based_district'] = 9
# assembler_gdf.loc[['15000US060133551071', '15000US060133551121'], 'block_based_district'] = 11
# assembler_gdf.loc[['15000US060133452031', '15000US060133452023', '15000US060133451051'], 'block_based_district'] = 11
# assembler_gdf.loc[['15000US060133551171', '15000US060133551161', '15000US060133451131', '15000US060133451142', '15000US060133451141', '15000US060133551143'], 'block_based_district'] = 15
# assembler_gdf.loc[['15000US060750179021'], 'block_based_district'] = 12
# assembler_gdf.loc[['15000US060759901000', '15000US060750255001', '15000US060750605023', '15000US060750330003', '15000US060750330004', '15000US060750303022', '15000US060750304003' , '15000US060750308005'], 'block_based_district'] = 12
# assembler_gdf.loc[['15000US060750263023', '15000US060750255002', '15000US060750306002', '15000US060750308004'], 'block_based_district'] = 14
# assembler_gdf.loc[['15000US060014338001', '15000US060014338003'], 'block_based_district'] = 13
# assembler_gdf.loc[['15000US060019900000', '15000US060014301021', '15000US060014338002', '15000US060014338004', '15000US060014304002', '15000US060014328003'], 'block_based_district'] = 15
# assembler_gdf.loc[['15000US060819901000', '15000US060816138002', '15000US060816117002', '15000US060816117004', '15000US060816096033' , '15000US060816104001', '15000US060816100001', '15000US060816102032', '15000US060816102012', '15000US060816109003'], 'block_based_district'] = 14
# assembler_gdf.loc[['15000US060816134003', '15000US060816134002', '15000US060816097001', '15000US060816099001', '15000US060816105001'], 'block_based_district'] = 18
# assembler_gdf.loc[['15000US060014507013', '15000US060014415242', '15000US060014419243', '15000US060014415031'], 'block_based_district'] = 15
# assembler_gdf.loc[['15000US060014431031', '15000US060014418001', '15000US060014418003', '15000US060014419241'], 'block_based_district'] = 17
# assembler_gdf.loc[['15000US060855077023', '15000US060855079041', '15000US060855079042', '15000US060855080043', '15000US060855059003', '15000US060855077031', '15000US060855077032', '15000US060855077034', '15000US060855077035', '15000US060855078052', '15000US060855084015'], 'block_based_district'] = 17
# assembler_gdf.loc[['15000US060855117071', '15000US060855047001', '15000US060855076001', '15000US060855100011', '15000US060855058002' , '15000US060855059002'], 'block_based_district'] = 18
# assembler_gdf.loc[['15000US060855043081', '15000US060855052031', '15000US060855057003'], 'block_based_district'] = 17
# assembler_gdf.loc[['15000US060855135001', '15000US060855042011', '15000US060855051002'], 'block_based_district'] = 19
# assembler_gdf.loc[['15000US060855119112', '15000US060855119111', '15000US060855119113', '15000US060855120271', '15000US060855120521', '15000US060855029022', '15000US060855022022' ,'15000US060855022023', '15000US060855020011', '15000US060855058001', '15000US060855020023', '15000US060855026011'], 'block_based_district'] = 18
# assembler_gdf.loc[['15000US060855121001', '15000US060855120242', '15000US060855120523', '15000US060855029032', '15000US060855029033', '15000US060855029021', '15000US060855026012', '15000US060855023022', '15000US060855005003', '15000US060855005004', '15000US060855020022'], 'block_based_district'] = 19
# assembler_gdf.loc[['15000US060871212002', '15000US060871212001'], 'block_based_district'] = 18
# assembler_gdf.loc[['15000US060879901000', '15000US060871212003', '15000US060871004001', '15000US060871005003', '15000US060871012003', '15000US060871207003'], 'block_based_district'] = 20
# assembler_gdf.loc[['15000US060855126022', '15000US060855124011', '15000US060855125092', '15000US060855125101'], 'block_based_district'] = 19
# assembler_gdf.loc[['15000US060855126021', '15000US060855125051', '15000US060855125062', '15000US060855126043', '15000US060855126032'], 'block_based_district'] = 20
# assembler_gdf[
# (assembler_gdf['STUSAB'] == 'CA')
# & (assembler_gdf['in_bay_area'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# ].groupby('congressional_districts_bitmask').size().sort_index(ascending=False)
# +
# s = time.time()
# #assembler_gdf = assembler_gdf_bk
# this_state = 'CA'
# test_this_bitmask = 'x00000000000000000011000000000000000000000000000000000'
# bay_area_districts = []
# for i in range(0, nDistricts+1):
# if (test_this_bitmask[i] == '1'):
# bay_area_districts.append(i)
# print(i)
# manual_plot_limits = False
# show_small_roads = True
# show_places = False
# show_water = True
# # 19 vs 20
# manual_xlim = (-121.64, -121.43)
# manual_ylim = (36.90, 37.11)
# legend_location = 'upper right'
# map_buffer_ratio = .1
# fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# #fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('Plotting official districts...')
# for ax in (ax1, ax2):
# legend_list = []
# #for i in range(1, len(test_this_bitmask_philly)):
# for i in bay_area_districts:
# if (test_this_bitmask[i] == '1'):
# if (i <= 18):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[i-1], label='District {0:.0f}'.format(i))
# else:
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:.0f}'.format(i))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1], label='District {0:,.0f}'.format(i)))
# print('plotting tracts that overlap multiple districts...')
# for ax in (ax1,ax2):
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_bay_area'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].plot(ax=ax, color='none', edgecolor='white', linewidth=5, zorder=100)
# if (manual_plot_limits):
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# else:
# bounding_box = unary_union(
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_bay_area'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].geometry.tolist()
# ).bounds
# # print(bounding_box)
# xlimits = (bounding_box[0], bounding_box[2])
# ylimits = (bounding_box[1], bounding_box[3])
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# ax.set_xlim(xlimits[0] - xbuffer, xlimits[1] + xbuffer)
# ax.set_ylim(ylimits[0] - ybuffer, ylimits[1] + ybuffer)
# ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
# print('labeling block groups...')
# ##### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['in_bay_area'] == True)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# if (show_water):
# print('plotting water...')
# water_bay_area_gdf.plot(ax=ax2, color='blue')
# print('plotting roads...')
# if (show_small_roads):
# roads_bay_area_gdf[~roads_bay_area_gdf['RTTYP'].isin(['I','U','S','C'])].plot(ax=ax2, color='black', linewidth=0.25)
# roads_bay_area_gdf[roads_bay_area_gdf['RTTYP'] == 'C'].plot(ax=ax2, color='black', linewidth=0.5)
# roads_bay_area_gdf[roads_bay_area_gdf['RTTYP'] == 'S'].plot(ax=ax2, color='black', linewidth=1)
# roads_bay_area_gdf[roads_bay_area_gdf['RTTYP'] == 'U'].plot(ax=ax2, color='black', linewidth=1.5)
# roads_bay_area_gdf[roads_bay_area_gdf['RTTYP'] == 'I'].plot(ax=ax2, color='black', linewidth=2)
# if (show_places):
# print('plotting places...')
# place_bay_area_gdf.plot(ax=ax, color='none', edgecolor='yellow', linewidth=3)
# print('\tlabeling places in map area...')
# for ix, thisrow in place_bay_area_gdf[
# ((place_bay_area_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_bay_area_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_bay_area_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_bay_area_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].sort_values(by='NAME').iterrows():
# print('\t{0:}...'.format(thisrow['NAME']))
# annotator = thisrow['NAME'].upper()
# ax2.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='red', backgroundcolor='white', fontsize=9, ha='center'
# )
# print('\tlabeling places in chosen tracts...')
# for ix, thisrow in place_bay_area_gdf[
# ((place_bay_area_gdf['INTPTLON'] >= xlimits[0]) & (place_bay_area_gdf['INTPTLON'] <= xlimits[1]))
# & ((place_bay_area_gdf['INTPTLAT'] >= ylimits[0]) & (place_bay_area_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='red', backgroundcolor='white', fontsize=12, ha='center', va='center')
# plt.xticks(fontsize=14)
# plt.yticks(fontsize=14)
# plt.show()
# explore_more = b
# oldi = -1
# old_map_bufer_ratio = -1
# print('manual_xlim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_xlim()[0]*100)/100, np.ceil(ax.get_xlim()[1]*100)/100))
# print('manual_ylim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_ylim()[0]*100)/100, np.ceil(ax.get_ylim()[1]*100)/100))
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# +
# s = time.time()
# #explore_more = ['15000US060855119112']
# label_small_roads = True
# label_small_road_types = []
# show_places = False
# show_water = True
# basesize = 12
# aspect_ratio = xspan / yspan
# oldi = -1
# i = 0
# map_buffer_ratio = .1
# suffixes_to_count_dict = {'Rd': 0, 'Dr': 0, 'St': 0,'Ave': 0, 'Blvd': 0, 'Ln': 0, 'Cir': 0, 'Way': 0, 'Ct': 0, 'Pkwy': 0, 'Pl': 0 }
# other_roads = []
# if (oldi != i):
# print('Running this cell for the first time...')
# this_block_group_id = explore_more[i]
# print('Examining block group {0:}'.format(this_block_group_id))
# print('Population: {0:,.0f}'.format(assembler_gdf.loc[this_block_group_id]['total_population']))
# block_group_gdf = assembler_gdf[assembler_gdf.index == this_block_group_id]
# total_block_group_area = assembler_gdf[assembler_gdf.index == this_block_group_id].to_crs(equal_area_crs).geometry.apply(lambda x: x.area).values[0]
# print('Area: {0:,.1f} km^2'.format(total_block_group_area/1000000))
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0], x.bounds[2])).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1], x.bounds[3])).values[0]
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# aspect_ratio = xspan / yspan
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0] - xbuffer, x.bounds[2] + xbuffer)).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1] - ybuffer, x.bounds[3] + ybuffer)).values[0]
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# map_area_geo = makebox(xlimits, ylimits)
# map_area_gdf = geopandas.GeoDataFrame(data=[[map_area_geo]], columns=['geometry'], crs=block_group_gdf.crs, geometry='geometry')
# print('Aspect ratio: {0:.3f}'.format(aspect_ratio))
# print('\n')
# fig, ax = plt.subplots(1,1,figsize=(basesize*aspect_ratio,basesize))
# legend_list = []
# for j in range(1, len(test_this_bitmask)):
# if (test_this_bitmask[j] == '1'):
# print('Matching District {0:}...'.format(j))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(j,color_reset_point)-1], label='District {0:,.0f}'.format(j)))
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].plot(ax=ax, color=district_color_cycle[np.mod(j,color_reset_point)-1])
# overlap_area = geopandas.overlay(block_group_gdf.to_crs(equal_area_crs), cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].to_crs(equal_area_crs), how='intersection').area.values[0]
# print('\tOverlap area: {0:,.1f} km^2 ({1:.1%})'.format(overlap_area / 1000000, overlap_area / total_block_group_area))
# print('\n')
# block_group_gdf.plot(ax=ax, color='none', edgecolor='white', lw=8, zorder=100)
# map_area_gdf.plot(ax=ax, color='none', edgecolor='yellow', lw=16)
# print('finding water overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# water_in_map_area_gdf = water_bay_area_gdf[water_bay_area_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# #water_here_gdf = geopandas.overlay(water_gdf, block_group_gdf, how='intersection')
# water_in_map_area_gdf.plot(ax=ax, color='blue')
# print('finding road overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# print('\twith block group...')
# roads_in_block_group_gdf = roads_bay_area_gdf[roads_bay_area_gdf.geometry.apply(lambda x: x.intersects(block_group_gdf.geometry.values[0]))]
# print('\twith full map area...')
# roads_in_map_area_gdf = roads_bay_area_gdf[roads_bay_area_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isnull()].plot(ax=ax, color=road_label_format['O']['thecolor'], linewidth=0.5)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isin(['M', 'O'])].plot(ax=ax, color=road_label_format['M']['thecolor'], linewidth=1)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'C'].plot(ax=ax, color=road_label_format['C']['thecolor'], linewidth=2)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'S'].plot(ax=ax, color=road_label_format['S']['thecolor'], linewidth=3)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'U'].plot(ax=ax, color=road_label_format['U']['thecolor'], linewidth=4)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'I'].plot(ax=ax, color=road_label_format['I']['thecolor'], linewidth=5)
# print('labeling big roads in map area...')
# for ix, thisrow in roads_in_map_area_gdf[
# (roads_in_map_area_gdf['FULLNAME'].notnull())
# & (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S']))].iterrows():
# #& (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S', 'C']))].iterrows():
# annotator = parse_road_name(thisrow['FULLNAME'])
# centerpoint = Point((thisrow.geometry.centroid.x, thisrow.geometry.centroid.y))
# if (map_area_geo.contains(centerpoint)):
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', ha='center', va='center',
# fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], bbox=road_label_format[thisrow['RTTYP']]['thebbox'])
# if (label_small_roads):
# print('labeling small roads in block group...')
# for ix, thisrow in roads_in_block_group_gdf[(roads_in_block_group_gdf['FULLNAME'].notnull()) & (roads_in_block_group_gdf['RTTYP'].isin(['M', 'O']))].iterrows():
# #print('\t{0:}'.format(thisrow['FULLNAME']))
# is_other_road_type = False
# try:
# road_suffix = thisrow['FULLNAME'][::-1][:re.search("\s",thisrow['FULLNAME'][::-1]).start()][::-1].strip()
# if (road_suffix in suffixes_to_count_dict):
# suffixes_to_count_dict[road_suffix] = suffixes_to_count_dict[road_suffix] + 1
# except AttributeError:
# is_other_road_type = True
# other_roads.append(thisrow['FULLNAME'])
# if ((road_suffix in label_small_road_types) ):#| (is_other_road_type)):
# annotator = thisrow['FULLNAME']
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], ha='center', va='center'
# , zorder=200)
# plt.title("Block Group {0:}".format(this_block_group_id), fontsize=24)
# if (label_small_roads):
# print("\tAnalysis of road names:")
# for k,v in suffixes_to_count_dict.items():
# print('\t\t{0:}: {1:,.0f}'.format(k,v))
# print('\tOther road names:')
# for x in other_roads:
# print('\t\t',x)
# print('\n')
# if (show_places):
# print('finding places...')
# print('\tshowing place names...')
# for ix, thisrow in place_bay_area_gdf.iterrows():
# if (thisrow.geometry.intersects(assembler_gdf.loc[this_block_group_id].geometry)):
# print('\t\t',str(thisrow['NAME']))
# print('\tplotting places...')
# place_dallas_gdf.plot(ax=ax, color='none', edgecolor='cyan', lw=3)
# print('\tlabeling places...')
# for ix, thisrow in place_bay_area_gdf[
# ((place_bay_area_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_bay_area_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_bay_area_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_bay_area_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=11, ha='center', va='center')
# print('http://maps.google.com/maps?ll={1:.3f},{0:.3f}&spn={2:.3f},{3:.3f}&t=m'.format((xlimits[0]+xlimits[1])/2, (ylimits[0]+ylimits[1])/2, xspan, yspan))
# ax.legend(handles=legend_list, fontsize=11, loc=legend_location)
# plt.xlim(xlimits)
# plt.ylim(ylimits)
# # plt.xticks(fontsize=24)
# # plt.yticks(fontsize=24)
# plt.show()
# oldi = i
# old_map_buffer_ratio = map_buffer_ratio
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# #map_area_geo
# if (label_small_roads):
# print(other_roads)
# pprint(this_block_group_id)
# +
# s = time.time()
# this_state = 'CA'
# nDistricts = 53
# #assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['block_based_district'] < 0), 'block_based_district'] = -5
# thedistricts = [-1,19,20]
# # # 19 vs 20
# manual_xlim = (-121.64, -121.43)
# manual_ylim = (36.90, 37.11)
# legend_location = 'lower right'
# show_real_district_boundaries = True
# district_boundary_color = 'white'
# map_buffer_ratio = .05
# fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# #fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('plotting block-group-based districts...')
# for ax in (ax1,ax2):
# legend_list = []
# for i in thedistricts:
# print(i)
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == i)
# ].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], edgecolor='black', lw=0.25)
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:,.0f}'.format(i)))
# if (show_real_district_boundaries):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'].isin(thedistricts))].plot(ax=ax, color='none', edgecolor=district_boundary_color, lw=3)
# ax.legend(handles=legend_list, fontsize=14, loc=legend_location)
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# ax.set_xlim(xlimits)
# ax.set_ylim(ylimits)
# print('labeling block groups...')
# #### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] < 0)
# #& (assembler_gdf.index == '15000US060014415031')
# & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# print('plotting water...')
# water_bay_area_gdf[
# (water_bay_area_gdf.geometry.apply(lambda x: x.centroid.x >= xlimits[0])) & ((water_bay_area_gdf.geometry.apply(lambda x: x.centroid.x) <= xlimits[1]))
# & (water_bay_area_gdf.geometry.apply(lambda x: x.centroid.y >= ylimits[0])) & ((water_bay_area_gdf.geometry.apply(lambda x: x.centroid.y) <= ylimits[1]))
# ].plot(ax=ax2, color='blue')
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# plt.show()
# -
# ## Plot Bay Area
# ## Southern California
# +
# # assembler_gdf.loc[['15000US060710092021'], 'block_based_district'] = 8
# # assembler_gdf.loc[['15000US060290060073'], 'block_based_district'] = 23
# # assembler_gdf.loc[['15000US060379012054', '15000US060379012053', '15000US060379007031'], 'block_based_district'] = 23
# # assembler_gdf.loc[['15000US060379003001', '15000US060379012131', '15000US060379006022', '15000US060379006062', '15000US060379102021', '15000US060379103011'], 'block_based_district'] = 25
# # assembler_gdf.loc[['15000US061110001001'], 'block_based_district'] = 24
# # assembler_gdf.loc[['15000US061119901000'], 'block_based_district'] = 26
# # assembler_gdf.loc[['15000US061110074053'], 'block_based_district'] = 26
# # assembler_gdf.loc[['15000US061110075111'], 'block_based_district'] = 30
# # assembler_gdf.loc[['15000US061110075061', '15000US061110075071', '15000US061110075142', '15000US061110079012', '15000US061110084022', '15000US061110075143'], 'block_based_district'] = 25
# # assembler_gdf.loc[['15000US061110084021', '15000US061110085003', '15000US061110075091', '15000US061110079012'], 'block_based_district'] = 26
# # assembler_gdf.loc[['15000US060379108111'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060379302002'], 'block_based_district'] = 28
# # assembler_gdf.loc[['15000US060379110013'], 'block_based_district'] = 25
# # assembler_gdf.loc[['15000US060379303012'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060379108051', '15000US060379108101'], 'block_based_district'] = 25
# # assembler_gdf.loc[['15000US061110075121', '15000US060371081021', '15000US060371081011'], 'block_based_district'] = 25
# # assembler_gdf.loc[['15000US060371132351', '15000US060371082021', '15000US060371066032', '15000US060371066431', '15000US060371082012', '15000US060371112061', '15000US060371112062'], 'block_based_district'] = 30
# # assembler_gdf.loc[['15000US060378003261', '15000US060378003263'], 'block_based_district'] = 26
# # assembler_gdf.loc[['15000US060378003242', '15000US060378003262'], 'block_based_district'] = 33
# # assembler_gdf.loc[['15000US060379301011', '15000US060374604011', '15000US060374617002', '15000US060374639002', '15000US060374639002'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060374608002', '15000US060374639002', '15000US060374639001', '15000US060374637002'], 'block_based_district'] = 28
# # assembler_gdf.loc[['15000US060710008081', '15000US060710008172'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060374005011', '15000US060374002051', '15000US060374002043', '15000US060374009002', '15000US060374039011'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060374016012', '15000US060374004031', '15000US060374006042', '15000US060374039013', '15000US060374008001'], 'block_based_district'] = 32
# # assembler_gdf.loc[['15000US060374315022', '15000US060374315023', '15000US060374315021', '15000US060374315022', '15000US060374321012', '15000US060374321025', '15000US060374321011', '15000US060374321022', '15000US060374329021'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060374310022','15000US060374313001', '15000US060374313002', '15000US060374303014', '15000US060374325001', '15000US060374325004', '15000US060374315011'], 'block_based_district'] = 32
# # assembler_gdf.loc[['15000US060374017011', '15000US060374017012'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060374017041'], 'block_based_district'] = 35
# # assembler_gdf.loc[['15000US060374824022', '15000US060374825221'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060375300031'], 'block_based_district'] = 38
# # assembler_gdf.loc[['15000US060375304002'], 'block_based_district'] = 27
# # assembler_gdf.loc[['15000US060379302001', '15000US060371041243', '15000US060379800261', '15000US060371021031', '15000US060371021042'], 'block_based_district'] = 28
# # assembler_gdf.loc[['15000US060371041242', '15000US060371032001', '15000US060371222002', '15000US060371211021', '15000US060371211023', '15000US060371032002', '15000US060379800211'], 'block_based_district'] = 29
# # assembler_gdf.loc[['15000US060371897013', '15000US060373108005', '15000US060373117006', '15000US060371437001'], 'block_based_district'] = 28
# # assembler_gdf.loc[['15000US060373110002', '15000US060373116004'], 'block_based_district'] = 30
# # assembler_gdf.loc[['15000US060371923001', '15000US060371944011', '15000US060371944023', '15000US060371924104', '15000US060371926201'], 'block_based_district'] = 28
# # assembler_gdf.loc[['15000US060371864041', '15000US060371924103', '15000US060371925201', '15000US060371925202', '15000US060371927002', '15000US060371958021', '15000US060371957103', '15000US060371926202'], 'block_based_district'] = 34
# # assembler_gdf.loc[['15000US060371093003', '15000US060371114001', '15000US060379800081', '15000US060371252003', '15000US060371284004', '15000US060371285002'], 'block_based_district'] = 29
# # assembler_gdf.loc[['15000US060371066031', '15000US060379800221', '15000US060371311002', '15000US060371311003', '15000US060371321022', '15000US060371236013', '15000US060371251001', '15000US060371255021', '15000US060371255022', '15000US060371256001', '15000US060371433004'], 'block_based_district'] = 30
# # assembler_gdf.loc[['15000US060371415001', '15000US060378002032'], 'block_based_district'] = 30
# # assembler_gdf.loc[['15000US060378002041'], 'block_based_district'] = 33
# # assembler_gdf.loc[['15000US060374013031', '15000US060374013042'], 'block_based_district'] = 32
# # assembler_gdf.loc[['15000US060374084023', '15000US060374083031', '15000US060374339022', '15000US060374340031', '15000US060374340032', '15000US060374331012', '15000US060374331021', '15000US060374334011', '15000US060374334012', '15000US060374334022', '15000US060374335043'], 'block_based_district'] = 32
# # assembler_gdf.loc[['15000US060375003001', '15000US060374083021', '15000US060374338012', '15000US060374335041'], 'block_based_district'] = 38
# # assembler_gdf.loc[['15000US060374082022', '15000US060374082111', '15000US060374035001'], 'block_based_district'] = 32
# # assembler_gdf.loc[['15000US060374082112', '15000US060374033031', '15000US060374082121', '15000US060374082122'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060372756021'], 'block_based_district'] = 43
# # assembler_gdf.loc[['15000US060372741001', '15000US060372753022', '15000US060372737003'], 'block_based_district'] = 33
# # assembler_gdf.loc[['15000US060372737001', '15000US060372738001', '15000US060372712002', '15000US060372713003', '15000US060372655101', '15000US060372656022', '15000US060372656021', '15000US060372657001', '15000US060372657003', '15000US060372676002', '15000US060372676003'], 'block_based_district'] = 37
# # assembler_gdf.loc[['15000US060376503005', '15000US060376507012', '15000US060376511021', '15000US060376514011', '15000US060376707011'], 'block_based_district'] = 33
# # assembler_gdf.loc[['15000US060372766011', '15000US060379800281'], 'block_based_district'] = 43
# # assembler_gdf.loc[['15000US060379903000'], 'block_based_district'] = 44
# # assembler_gdf.loc[['15000US060372964021', '15000US060372964023'], 'block_based_district'] = 33
# # assembler_gdf.loc[['15000US060372214021', '15000US060372242002'], 'block_based_district'] = 34
# # assembler_gdf.loc[['15000US060375309011', '15000US060375309022'], 'block_based_district'] = 34
# # assembler_gdf.loc[['15000US060375310001', '15000US060372260012', '15000US060375311013'], 'block_based_district'] = 40
# # assembler_gdf.loc[['15000US060710004013'], 'block_based_district'] = 35
# # assembler_gdf.loc[['15000US060374033213', '15000US060710001051', '15000US060374024041', '15000US060710001132', '15000US060710001152'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060372240201', '15000US060372240202', '15000US060372246002', '15000US060372311002'], 'block_based_district'] = 37
# # assembler_gdf.loc[['15000US060372318001'], 'block_based_district'] = 40
# # assembler_gdf.loc[['15000US060372381002', '15000US060372382002', '15000US060372382003', '15000US060372756031', '15000US060372756033', '15000US060372761003'], 'block_based_district'] = 37
# # assembler_gdf.loc[['15000US060372384001', '15000US060372384003', '15000US060372383102', '15000US060372383201'], 'block_based_district'] = 43
# # assembler_gdf.loc[['15000US060375016001', '15000US060375002025', '15000US060375002012'], 'block_based_district'] = 38
# # assembler_gdf.loc[['15000US060591103011', '15000US060374085033', '15000US060375002011'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060375531003', '15000US060375302021', '15000US060375302023', '15000US060375531004', '15000US060375541051', '15000US060375541052', '15000US060375542043', '15000US060375543013'], 'block_based_district'] = 38
# # assembler_gdf.loc[['15000US060375531001', '15000US060375318003', '15000US060375319021', '15000US060375542011'], 'block_based_district'] = 40
# # assembler_gdf.loc[['15000US060591101023'], 'block_based_district'] = 38
# # assembler_gdf.loc[['15000US060375715031', '15000US060591101111', '15000US060591103041'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060590218262', '15000US060590219152', '15000US060590219151'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060590219241', '15000US060590219243', '15000US060590758131', '15000US060590758133'], 'block_based_district'] = 45
# # assembler_gdf.loc[['15000US060591104023'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060591102011'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060590117222', '15000US060591104011', '15000US060590117202', '15000US060590116021', '15000US060590116022', '15000US060590868011', '15000US060590868032', '15000US060591104022'], 'block_based_district'] = 39
# # assembler_gdf.loc[['15000US060590117141', '15000US060590762022', '15000US060590864073', '15000US060590867011', '15000US060590868021', '15000US060590868013', '15000US060590117201', '15000US060590117203'], 'block_based_district'] = 46
# # assembler_gdf.loc[['15000US060591103012', '15000US060591103013'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060375351012'], 'block_based_district'] = 40
# # assembler_gdf.loc[['15000US060375353001', '15000US060375353003'], 'block_based_district'] = 44
# # assembler_gdf.loc[['15000US060372410013', '15000US060372911102'], 'block_based_district'] = 43
# # assembler_gdf.loc[['15000US060375435011'], 'block_based_district'] = 44
# # assembler_gdf.loc[['15000US060375706021', '15000US060375706022', '15000US060375706024', '15000US060375717033'], 'block_based_district'] = 44
# # assembler_gdf.loc[['15000US060375440021', '15000US060375723012'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060590755151'], 'block_based_district'] = 45
# # assembler_gdf.loc[['15000US060590755041', '15000US060590755052'], 'block_based_district'] = 45
# # assembler_gdf.loc[['15000US060590744063', '15000US060590754033', '15000US060590757012', '15000US060590757013', '15000US060590758132'], 'block_based_district'] = 46
# # assembler_gdf.loc[['15000US060590626143', '15000US060590626213', '15000US060590423331', '15000US060590626493', '15000US060590423202', '15000US060590423201'], 'block_based_district'] = 45
# # assembler_gdf.loc[['15000US060590626042', '15000US060590626102', '15000US060590626411', '15000US060590626412', '15000US060590626343', '15000US060590423352', '15000US060590423351', '15000US060590626361'], 'block_based_district'] = 48
# # assembler_gdf.loc[['15000US060590320411', '15000US060590320222', '15000US060590320431', '15000US060590320432'], 'block_based_district'] = 45
# # assembler_gdf.loc[['15000US060590320564', '15000US060590320223', '15000US060590320534', '15000US060590320563', '15000US060590320572'], 'block_based_district'] = 49
# # assembler_gdf.loc[['15000US060590878063', '15000US060590883012', '15000US060591102031', '15000US060590878011', '15000US060590878021', '15000US060590878061', '15000US060590878062'], 'block_based_district'] = 46
# # assembler_gdf.loc[['15000US060591102021', '15000US060590878031', '15000US060590878051', '15000US060590883021', '15000US060591102022', '15000US060591102023'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060590741061', '15000US060590741062', '15000US060590741082', '15000US060590748032'], 'block_based_district'] = 46
# # assembler_gdf.loc[['15000US060590996031', '15000US060591100073', '15000US060591100082', '15000US060590996021', '15000US060590997011', '15000US060590997012', '15000US060590997013', '15000US060591100081'], 'block_based_district'] = 47
# # assembler_gdf.loc[['15000US060590890011', '15000US060590996031', '15000US060590997022'], 'block_based_district'] = 48
# # assembler_gdf.loc[['15000US060599901000', '15000US060590423051', '15000US060590423052', '15000US060590423151', '15000US060590423152', '15000US060590423153', '15000US060590423155'], 'block_based_district'] = 48
# # assembler_gdf.loc[['15000US060590423241', '15000US060590423111', '15000US060590423113'], 'block_based_district'] = 49
# assembler_gdf[
# (assembler_gdf['STUSAB'] == 'CA')
# & (assembler_gdf['in_so_cal'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# ].groupby('congressional_districts_bitmask').size().sort_index(ascending=False)
# +
# s = time.time()
# this_state = 'CA'
# test_this_bitmask = 'x00000000000000000000000000000000000000000000000110000'
# so_cal_districts = []
# for i in range(0, nDistricts+1):
# if (test_this_bitmask[i] == '1'):
# so_cal_districts.append(i)
# print(i)
# manual_plot_limits = False
# show_small_roads = True
# show_places = False
# show_water = True
# legend_location = 'upper right'
# map_buffer_ratio = .1
# manual_xlim = (-117.76,-117.72)
# manual_ylim = (33.47,33.52)
# fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# #fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('Plotting official districts...')
# for ax in (ax1, ax2):
# legend_list = []
# for i in so_cal_districts:
# if (test_this_bitmask[i] == '1'):
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1],label='District {0:,.0f}'.format(i)))
# if (i <= 18):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[i-1], label='District {0:.0f}'.format(i))
# else:
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:.0f}'.format(i))
# ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
# print('plotting tracts that overlap multiple districts...')
# for ax in (ax1,ax2):
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_so_cal'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].plot(ax=ax, color='none', edgecolor='white', linewidth=5, zorder=100)
# if (manual_plot_limits):
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# else:
# bounding_box = unary_union(
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_so_cal'] == True)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].geometry.tolist()
# ).bounds
# xlimits = (bounding_box[0], bounding_box[2])
# ylimits = (bounding_box[1], bounding_box[3])
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# ax.set_xlim(xlimits[0] - xbuffer, xlimits[1] + xbuffer)
# ax.set_ylim(ylimits[0] - ybuffer, ylimits[1] + ybuffer)
# print('labeling block groups...')
# ##### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['in_so_cal'] == True)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# # & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# # & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# if (show_water):
# print('plotting water...')
# water_so_cal_gdf.plot(ax=ax2, color='blue')
# print('plotting roads...')
# if (show_small_roads):
# roads_so_cal_gdf[~roads_so_cal_gdf['RTTYP'].isin(['I','U','S','C'])].plot(ax=ax2, color='black', linewidth=0.25)
# roads_so_cal_gdf[roads_so_cal_gdf['RTTYP'] == 'C'].plot(ax=ax2, color='black', linewidth=0.5)
# roads_so_cal_gdf[roads_so_cal_gdf['RTTYP'] == 'S'].plot(ax=ax2, color='black', linewidth=1)
# roads_so_cal_gdf[roads_so_cal_gdf['RTTYP'] == 'U'].plot(ax=ax2, color='black', linewidth=1.5)
# roads_so_cal_gdf[roads_so_cal_gdf['RTTYP'] == 'I'].plot(ax=ax2, color='black', linewidth=2)
# if (show_places):
# print('plotting places...')
# place_so_cal_gdf.plot(ax=ax, color='none', edgecolor='yellow', linewidth=3)
# print('\tlabeling places in map area...')
# for ix, thisrow in place_so_cal_gdf[
# ((place_so_cal_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_so_cal_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_so_cal_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_so_cal_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].sort_values(by='NAME').iterrows():
# print('\t{0:}...'.format(thisrow['NAME']))
# annotator = thisrow['NAME'].upper()
# ax2.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='red', backgroundcolor='white', fontsize=9, ha='center'
# )
# print('\tlabeling places in chosen tracts...')
# for ix, thisrow in place_so_cal_gdf[
# ((place_so_cal_gdf['INTPTLON'] >= xlimits[0]) & (place_so_cal_gdf['INTPTLON'] <= xlimits[1]))
# & ((place_so_cal_gdf['INTPTLAT'] >= ylimits[0]) & (place_so_cal_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='red', backgroundcolor='white', fontsize=12, ha='center', va='center')
# plt.xticks(fontsize=14)
# plt.yticks(fontsize=14)
# plt.show()
# explore_more = b
# oldi = -1
# old_map_bufer_ratio = -1
# print('manual_xlim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_xlim()[0]*100)/100, np.ceil(ax.get_xlim()[1]*100)/100))
# print('manual_ylim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_ylim()[0]*100)/100, np.ceil(ax.get_ylim()[1]*100)/100))
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# +
# ## 48 vs 49
# # manual_xlim = (-117.77, -117.65)
# # manual_ylim = (33.47, 33.57)
# s = time.time()
# legend_loc = 'upper right'
# #explore_more = ['15000US060590626102']
# label_small_roads = True
# label_small_road_types = []
# show_places = False
# show_water = True
# basesize = 12
# aspect_ratio = xspan / yspan
# oldi = -1
# i = 1
# map_buffer_ratio = .1
# suffixes_to_count_dict = {'Rd': 0, 'Dr': 0, 'St': 0,'Ave': 0, 'Blvd': 0, 'Ln': 0, 'Cir': 0, 'Way': 0, 'Ct': 0, 'Pkwy': 0, 'Pl': 0 }
# other_roads = []
# if (oldi != i):
# print('Running this cell for the first time...')
# this_block_group_id = explore_more[i]
# print('Examining block group {0:}'.format(this_block_group_id))
# print('Population: {0:,.0f}'.format(assembler_gdf.loc[this_block_group_id]['total_population']))
# block_group_gdf = assembler_gdf[assembler_gdf.index == this_block_group_id]
# total_block_group_area = assembler_gdf[assembler_gdf.index == this_block_group_id].to_crs(equal_area_crs).geometry.apply(lambda x: x.area).values[0]
# print('Area: {0:,.1f} km^2'.format(total_block_group_area/1000000))
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0], x.bounds[2])).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1], x.bounds[3])).values[0]
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# aspect_ratio = xspan / yspan
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0] - xbuffer, x.bounds[2] + xbuffer)).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1] - ybuffer, x.bounds[3] + ybuffer)).values[0]
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# map_area_geo = makebox(xlimits, ylimits)
# map_area_gdf = geopandas.GeoDataFrame(data=[[map_area_geo]], columns=['geometry'], crs=block_group_gdf.crs, geometry='geometry')
# print('Aspect ratio: {0:.3f}'.format(aspect_ratio))
# print('\n')
# fig, ax = plt.subplots(1,1,figsize=(basesize*aspect_ratio,basesize))
# legend_list = []
# for j in range(1, len(test_this_bitmask)):
# if (test_this_bitmask[j] == '1'):
# print('Matching District {0:}...'.format(j))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(j,color_reset_point)-1], label='District {0:,.0f}'.format(j)))
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].plot(ax=ax, color=district_color_cycle[np.mod(j,color_reset_point)-1])
# overlap_area = geopandas.overlay(block_group_gdf.to_crs(equal_area_crs), cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].to_crs(equal_area_crs), how='intersection').area.values[0]
# print('\tOverlap area: {0:,.1f} km^2 ({1:.1%})'.format(overlap_area / 1000000, overlap_area / total_block_group_area))
# print('\n')
# block_group_gdf.plot(ax=ax, color='none', edgecolor='white', lw=8, zorder=100)
# #map_area_gdf.plot(ax=ax, color='none', edgecolor='yellow', lw=16)
# print('finding water overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# water_in_map_area_gdf = water_so_cal_gdf[water_so_cal_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# #water_here_gdf = geopandas.overlay(water_gdf, block_group_gdf, how='intersection')
# water_in_map_area_gdf.plot(ax=ax, color='blue')
# print('finding road overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# print('\twith block group...')
# roads_in_block_group_gdf = roads_so_cal_gdf[roads_so_cal_gdf.geometry.apply(lambda x: x.intersects(block_group_gdf.geometry.values[0]))]
# print('\twith full map area...')
# roads_in_map_area_gdf = roads_so_cal_gdf[roads_so_cal_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isnull()].plot(ax=ax, color=road_label_format['O']['thecolor'], linewidth=0.5)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isin(['M', 'O'])].plot(ax=ax, color=road_label_format['M']['thecolor'], linewidth=1)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'C'].plot(ax=ax, color=road_label_format['C']['thecolor'], linewidth=2)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'S'].plot(ax=ax, color=road_label_format['S']['thecolor'], linewidth=3)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'U'].plot(ax=ax, color=road_label_format['U']['thecolor'], linewidth=4)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'I'].plot(ax=ax, color=road_label_format['I']['thecolor'], linewidth=5)
# print('labeling big roads in map area...')
# for ix, thisrow in roads_in_map_area_gdf[
# (roads_in_map_area_gdf['FULLNAME'].notnull())
# & (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S']))].iterrows():
# #& (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S', 'C']))].iterrows():
# annotator = parse_road_name(thisrow['FULLNAME'])
# centerpoint = Point((thisrow.geometry.centroid.x, thisrow.geometry.centroid.y))
# if (map_area_geo.contains(centerpoint)):
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', ha='center', va='center',
# fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], bbox=road_label_format[thisrow['RTTYP']]['thebbox'])
# if (label_small_roads):
# print('labeling small roads in block group...')
# for ix, thisrow in roads_in_block_group_gdf[(roads_in_block_group_gdf['FULLNAME'].notnull()) & (roads_in_block_group_gdf['RTTYP'].isin(['M', 'O']))].iterrows():
# #print('\t{0:}'.format(thisrow['FULLNAME']))
# is_other_road_type = False
# try:
# road_suffix = thisrow['FULLNAME'][::-1][:re.search("\s",thisrow['FULLNAME'][::-1]).start()][::-1].strip()
# if (road_suffix in suffixes_to_count_dict):
# suffixes_to_count_dict[road_suffix] = suffixes_to_count_dict[road_suffix] + 1
# except AttributeError:
# is_other_road_type = True
# other_roads.append(thisrow['FULLNAME'])
# if ((road_suffix in label_small_road_types) ):#| (is_other_road_type)):
# annotator = thisrow['FULLNAME']
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], ha='center', va='center'
# , zorder=200)
# plt.title("Block Group {0:}".format(this_block_group_id), fontsize=24)
# if (label_small_roads):
# print("\tAnalysis of road names:")
# for k,v in suffixes_to_count_dict.items():
# print('\t\t{0:}: {1:,.0f}'.format(k,v))
# print('\tOther road names:')
# for x in other_roads:
# print('\t\t',x)
# print('\n')
# if (show_places):
# print('finding places...')
# print('\tshowing place names...')
# for ix, thisrow in place_bay_area_gdf.iterrows():
# if (thisrow.geometry.intersects(assembler_gdf.loc[this_block_group_id].geometry)):
# print('\t\t',str(thisrow['NAME']))
# print('\tplotting places...')
# place_dallas_gdf.plot(ax=ax, color='none', edgecolor='cyan', lw=3)
# print('\tlabeling places...')
# for ix, thisrow in place_bay_area_gdf[
# ((place_bay_area_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_bay_area_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_bay_area_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_bay_area_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=11, ha='center', va='center')
# print('http://maps.google.com/maps?ll={1:.3f},{0:.3f}&spn={2:.3f},{3:.3f}&t=m'.format((xlimits[0]+xlimits[1])/2, (ylimits[0]+ylimits[1])/2, xspan, yspan))
# ax.legend(handles=legend_list, fontsize=11, loc=legend_location)
# plt.xlim(xlimits)
# plt.ylim(ylimits)
# # plt.xticks(fontsize=24)
# # plt.yticks(fontsize=24)
# plt.show()
# oldi = i
# old_map_buffer_ratio = map_buffer_ratio
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# #map_area_geo
# if (label_small_roads):
# print(other_roads)
# pprint(this_block_group_id)
# +
# s = time.time()
# this_state = 'CA'
# nDistricts = 53
# u = -1
# assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['block_based_district'] < 0), 'block_based_district'] = u
# thedistricts = [u,48,49]
# ## 48 vs 49
# manual_xlim = (-117.77, -117.65)
# manual_ylim = (33.47, 33.57)
# legend_location = 'upper right'
# show_real_district_boundaries = True
# district_boundary_color = 'white'
# map_buffer_ratio = .05
# fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# #fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('plotting block-group-based districts...')
# for ax in (ax1,ax2):
# legend_list = []
# for i in thedistricts:
# print(i)
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == i)
# ].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], edgecolor='black', lw=0.25)
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:,.0f}'.format(i)))
# if (show_real_district_boundaries):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'].isin(thedistricts))].plot(ax=ax, color='none', edgecolor=district_boundary_color, lw=3)
# ax.legend(handles=legend_list, fontsize=14, loc=legend_location)
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# ax.set_xlim(xlimits)
# ax.set_ylim(ylimits)
# print('labeling block groups...')
# #### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] < 0)
# #& (assembler_gdf.index == '15000US060710092021')
# & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# print('plotting water...')
# water_so_cal_gdf[
# (water_so_cal_gdf.geometry.apply(lambda x: x.centroid.x >= xlimits[0])) & ((water_so_cal_gdf.geometry.apply(lambda x: x.centroid.x) <= xlimits[1]))
# & (water_so_cal_gdf.geometry.apply(lambda x: x.centroid.y >= ylimits[0])) & ((water_so_cal_gdf.geometry.apply(lambda x: x.centroid.y) <= ylimits[1]))
# ].plot(ax=ax2, color='blue')
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# plt.show()
# assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['block_based_district'] < 0), 'block_based_district'] = -1
# -
# # REST OF STATE
# +
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_bay_area'] == False)
# & (assembler_gdf['in_so_cal'] == False)
# & (assembler_gdf['block_based_district'] == -1)
# ].groupby('congressional_districts_bitmask').size().sort_index(ascending=False)
# +
# s = time.time()
# this_state = 'CA'
# test_this_bitmask = 'x00000000000000000000000000000000000000000000000000011'
# other_ca_districts = []
# for i in range(0, nDistricts+1):
# if (test_this_bitmask[i] == '1'):
# other_ca_districts.append(i)
# print(i)
# manual_plot_limits = False
# show_roads = False
# show_places = False
# show_water = False
# legend_location = 'upper right'
# map_buffer_ratio = .05
# manual_xlim = (-117.04,-117)
# manual_ylim = (32.79,32.81)
# fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# #fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('Plotting official districts...')
# for ax in (ax1, ax2):
# legend_list = []
# for i in other_ca_districts:
# if (test_this_bitmask[i] == '1'):
# if (i <= 18):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[i-1], label='District {0:.0f}'.format(i))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1],label='District {0:,.0f}'.format(i)))
# else:
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == i)].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:.0f}'.format(i))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i,color_reset_point)-1],label='District {0:,.0f}'.format(i)))
# ax.legend(handles=legend_list, fontsize=12, loc=legend_location)
# print('plotting tracts that overlap multiple districts...')
# for ax in (ax1,ax2):
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_bay_area'] == False)
# & (assembler_gdf['in_so_cal'] == False)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].plot(ax=ax, color='none', edgecolor='white', linewidth=5, zorder=100)
# if (manual_plot_limits):
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# else:
# bounding_box = unary_union(
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['in_bay_area'] == False)
# & (assembler_gdf['in_so_cal'] == False)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# ].geometry.tolist()
# ).bounds
# xlimits = (bounding_box[0], bounding_box[2])
# ylimits = (bounding_box[1], bounding_box[3])
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# ax.set_xlim(xlimits[0] - xbuffer, xlimits[1] + xbuffer)
# ax.set_ylim(ylimits[0] - ybuffer, ylimits[1] + ybuffer)
# print('labeling block groups...')
# ##### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == -1)
# & (assembler_gdf['congressional_districts_bitmask'] == test_this_bitmask)
# & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# if (show_water):
# print('plotting water...')
# water_other_gdf.plot(ax=ax2, color='blue')
# if (show_roads):
# print('plotting roads...')
# roads_other_gdf[~roads_other_gdf['RTTYP'].isin(['I','U','S','C'])].plot(ax=ax2, color='black', linewidth=0.25)
# roads_other_gdf[roads_other_gdf['RTTYP'] == 'C'].plot(ax=ax2, color='black', linewidth=0.5)
# roads_other_gdf[roads_other_gdf['RTTYP'] == 'S'].plot(ax=ax2, color='black', linewidth=1)
# roads_other_gdf[roads_other_gdf['RTTYP'] == 'U'].plot(ax=ax2, color='black', linewidth=1.5)
# roads_other_gdf[roads_other_gdf['RTTYP'] == 'I'].plot(ax=ax2, color='black', linewidth=2)
# if (show_places):
# print('plotting places...')
# place_other_gdf.plot(ax=ax, color='none', edgecolor='yellow', linewidth=3)
# print('\tlabeling places in map area...')
# for ix, thisrow in place_other_gdf[
# ((place_other_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_so_cal_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_other_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_other_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].sort_values(by='NAME').iterrows():
# print('\t{0:}...'.format(thisrow['NAME']))
# annotator = thisrow['NAME'].upper()
# ax2.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='red', backgroundcolor='white', fontsize=9, ha='center'
# )
# print('\tlabeling places in chosen tracts...')
# for ix, thisrow in place_other_gdf[
# ((place_other_gdf['INTPTLON'] >= xlimits[0]) & (place_other_gdf['INTPTLON'] <= xlimits[1]))
# & ((place_other_gdf['INTPTLAT'] >= ylimits[0]) & (place_other_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='red', backgroundcolor='white', fontsize=12, ha='center', va='center')
# plt.xticks(fontsize=14)
# plt.yticks(fontsize=14)
# plt.show()
# explore_more = b
# oldi = -1
# old_map_bufer_ratio = -1
# print('manual_xlim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_xlim()[0]*100)/100, np.ceil(ax.get_xlim()[1]*100)/100))
# print('manual_ylim = ({0:,.2f}, {1:,.2f})'.format(np.floor(ax.get_ylim()[0]*100)/100, np.ceil(ax.get_ylim()[1]*100)/100))
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# +
# # manual_xlim = (-117.23, -117.00)
# # manual_ylim = (32.71, 32.84)
# s = time.time()
# legend_loc = 'upper right'
# #explore_more = ['15000US060730134191']
# label_small_roads = True
# label_small_road_types = ['Dr','St','Ave','Blvd']
# show_places = True
# show_water = True
# basesize = 12
# aspect_ratio = xspan / yspan
# i = 0
# map_buffer_ratio = 1
# suffixes_to_count_dict = {'Rd': 0, 'Dr': 0, 'St': 0,'Ave': 0, 'Blvd': 0, 'Ln': 0, 'Cir': 0, 'Way': 0, 'Ct': 0, 'Pkwy': 0, 'Pl': 0 }
# other_roads = []
# if (oldi != i):
# print('Running this cell for the first time...')
# this_block_group_id = explore_more[i]
# print('Examining block group {0:}'.format(this_block_group_id))
# print('Population: {0:,.0f}'.format(assembler_gdf.loc[this_block_group_id]['total_population']))
# block_group_gdf = assembler_gdf[assembler_gdf.index == this_block_group_id]
# total_block_group_area = assembler_gdf[assembler_gdf.index == this_block_group_id].to_crs(equal_area_crs).geometry.apply(lambda x: x.area).values[0]
# print('Area: {0:,.1f} km^2'.format(total_block_group_area/1000000))
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0], x.bounds[2])).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1], x.bounds[3])).values[0]
# xspan = xlimits[1] - xlimits[0]
# yspan = ylimits[1] - ylimits[0]
# aspect_ratio = xspan / yspan
# xbuffer = xspan * map_buffer_ratio
# ybuffer = yspan * map_buffer_ratio
# xlimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[0] - xbuffer, x.bounds[2] + xbuffer)).values[0]
# ylimits = block_group_gdf.geometry.apply(lambda x: (x.bounds[1] - ybuffer, x.bounds[3] + ybuffer)).values[0]
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# map_area_geo = makebox(xlimits, ylimits)
# map_area_gdf = geopandas.GeoDataFrame(data=[[map_area_geo]], columns=['geometry'], crs=block_group_gdf.crs, geometry='geometry')
# print('Aspect ratio: {0:.3f}'.format(aspect_ratio))
# print('\n')
# fig, ax = plt.subplots(1,1,figsize=(basesize*aspect_ratio,basesize))
# legend_list = []
# for j in range(1, len(test_this_bitmask)):
# if (test_this_bitmask[j] == '1'):
# print('Matching District {0:}...'.format(j))
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(j,color_reset_point)-1], label='District {0:,.0f}'.format(j)))
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].plot(ax=ax, color=district_color_cycle[np.mod(j,color_reset_point)-1])
# overlap_area = geopandas.overlay(block_group_gdf.to_crs(equal_area_crs), cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'] == j)].to_crs(equal_area_crs), how='intersection').area.values[0]
# print('\tOverlap area: {0:,.1f} km^2 ({1:.1%})'.format(overlap_area / 1000000, overlap_area / total_block_group_area))
# print('\n')
# block_group_gdf.plot(ax=ax, color='none', edgecolor='white', lw=8, zorder=100)
# #map_area_gdf.plot(ax=ax, color='none', edgecolor='yellow', lw=16)
# print('finding water overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# water_in_map_area_gdf = water_other_gdf[water_other_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# #water_here_gdf = geopandas.overlay(water_gdf, block_group_gdf, how='intersection')
# if (show_water):
# water_in_map_area_gdf.plot(ax=ax, color='blue')
# print('finding road overlap...')
# if ((oldi != i) | (old_map_buffer_ratio != map_buffer_ratio)):
# print('\twith block group...')
# roads_in_block_group_gdf = roads_other_gdf[roads_other_gdf.geometry.apply(lambda x: x.intersects(block_group_gdf.geometry.values[0]))]
# print('\twith full map area...')
# roads_in_map_area_gdf = roads_other_gdf[roads_other_gdf.geometry.apply(lambda x: x.intersects(map_area_geo))]
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isnull()].plot(ax=ax, color=road_label_format['O']['thecolor'], linewidth=0.5)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'].isin(['M', 'O'])].plot(ax=ax, color=road_label_format['M']['thecolor'], linewidth=1)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'C'].plot(ax=ax, color=road_label_format['C']['thecolor'], linewidth=2)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'S'].plot(ax=ax, color=road_label_format['S']['thecolor'], linewidth=3)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'U'].plot(ax=ax, color=road_label_format['U']['thecolor'], linewidth=4)
# roads_in_map_area_gdf[roads_in_map_area_gdf['RTTYP'] == 'I'].plot(ax=ax, color=road_label_format['I']['thecolor'], linewidth=5)
# print('labeling big roads in map area...')
# for ix, thisrow in roads_in_map_area_gdf[
# (roads_in_map_area_gdf['FULLNAME'].notnull())
# & (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S']))].iterrows():
# #& (roads_in_map_area_gdf['RTTYP'].isin(['I', 'U', 'S', 'C']))].iterrows():
# annotator = parse_road_name(thisrow['FULLNAME'])
# centerpoint = Point((thisrow.geometry.centroid.x, thisrow.geometry.centroid.y))
# if (map_area_geo.contains(centerpoint)):
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', ha='center', va='center',
# fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], bbox=road_label_format[thisrow['RTTYP']]['thebbox'])
# if (label_small_roads):
# print('labeling small roads in block group...')
# for ix, thisrow in roads_in_block_group_gdf[(roads_in_block_group_gdf['FULLNAME'].notnull()) & (roads_in_block_group_gdf['RTTYP'].isin(['M', 'O']))].iterrows():
# #print('\t{0:}'.format(thisrow['FULLNAME']))
# is_other_road_type = False
# try:
# road_suffix = thisrow['FULLNAME'][::-1][:re.search("\s",thisrow['FULLNAME'][::-1]).start()][::-1].strip()
# if (road_suffix in suffixes_to_count_dict):
# suffixes_to_count_dict[road_suffix] = suffixes_to_count_dict[road_suffix] + 1
# except AttributeError:
# is_other_road_type = True
# other_roads.append(thisrow['FULLNAME'])
# if ((road_suffix in label_small_road_types) ):#| (is_other_road_type)):
# annotator = thisrow['FULLNAME']
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=road_label_format[thisrow['RTTYP']]['labelsize'], ha='center', va='center'
# , zorder=200)
# plt.title("Block Group {0:}".format(this_block_group_id), fontsize=24)
# if (label_small_roads):
# print("\tAnalysis of road names:")
# for k,v in suffixes_to_count_dict.items():
# print('\t\t{0:}: {1:,.0f}'.format(k,v))
# print('\tOther road names:')
# for x in other_roads:
# print('\t\t',x)
# print('\n')
# if (show_places):
# print('finding places...')
# print('\tshowing place names...')
# for ix, thisrow in place_other_gdf.iterrows():
# if (thisrow.geometry.intersects(assembler_gdf.loc[this_block_group_id].geometry)):
# print('\t\t',str(thisrow['NAME']))
# print('\tplotting places...')
# place_other_gdf.plot(ax=ax, color='none', edgecolor='cyan', lw=3)
# print('\tlabeling places...')
# for ix, thisrow in place_other_gdf[
# ((place_other_gdf['INTPTLON'] >= xlimits[0] - xbuffer) & (place_other_gdf['INTPTLON'] <= xlimits[1] + xbuffer))
# & ((place_other_gdf['INTPTLAT'] >= ylimits[0] - ybuffer) & (place_other_gdf['INTPTLAT'] <= ylimits[1] + ybuffer))
# ].iterrows():
# annotator = thisrow['NAME'].upper()
# plt.annotate(annotator, (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y), (thisrow.geometry.centroid.x, thisrow.geometry.centroid.y),
# color='black', backgroundcolor='white', fontsize=11, ha='center', va='center')
# print('http://maps.google.com/maps?ll={1:.3f},{0:.3f}&spn={2:.3f},{3:.3f}&t=m'.format((xlimits[0]+xlimits[1])/2, (ylimits[0]+ylimits[1])/2, xspan, yspan))
# ax.legend(handles=legend_list, fontsize=11, loc=legend_location)
# plt.xlim(xlimits)
# plt.ylim(ylimits)
# # plt.xticks(fontsize=24)
# # plt.yticks(fontsize=24)
# plt.show()
# oldi = i
# old_map_buffer_ratio = map_buffer_ratio
# e = time.time()
# g = g + (e-s)
# print('Done in {0:,.0f} minutes {1:,.0f} seconds!'.format(np.floor((e-s)/60), np.floor((e-s)%60)))
# #map_area_geo
# if (label_small_roads):
# print(other_roads)
# pprint(this_block_group_id)
# +
# s = time.time()
# this_state = 'CA'
# nDistricts = 53
# show_water = True
# u = -10
# assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['block_based_district'] < 0), 'block_based_district'] = u
# thedistricts = [u,50,51,52,53]
# legend_location = 'lower right'
# show_real_district_boundaries = True
# district_boundary_color = 'white'
# map_buffer_ratio = .05
# #fig, (ax1, ax2) = plt.subplots(1,2,figsize=(16,8))
# fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16,8))
# print('plotting block-group-based districts...')
# for ax in (ax1,ax2):
# legend_list = []
# for i in thedistricts:
# print(i)
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == i)
# ].plot(ax=ax, color=district_color_cycle[np.mod(i, color_reset_point)-1], lw=0.25)
# assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == i)
# ].plot(ax=ax, color='none', edgecolor='black', lw=0.25, zorder=99)
# legend_list.append(mpatches.Patch(color=district_color_cycle[np.mod(i, color_reset_point)-1], label='District {0:,.0f}'.format(i)))
# if (show_real_district_boundaries):
# cd_gdf[(cd_gdf['STUSAB'] == this_state) & (cd_gdf['CD116FP'].isin(thedistricts))].plot(ax=ax, color='none', edgecolor=district_boundary_color, lw=3)
# ax.legend(handles=legend_list, fontsize=14, loc=legend_location)
# xlimits = (manual_xlim[0], manual_xlim[1])
# ylimits = (manual_ylim[0], manual_ylim[1])
# ax.set_xlim(xlimits)
# ax.set_ylim(ylimits)
# print('labeling block groups...')
# #### add labels for block groups with overlap
# b = []
# for ix, thisrow in assembler_gdf[
# (assembler_gdf['STUSAB'] == this_state)
# & (assembler_gdf['block_based_district'] == u)
# #& (assembler_gdf.index == '15000US060650426202')
# & ((assembler_gdf['INTPTLON'] >= xlimits[0]) & (assembler_gdf['INTPTLON'] <= xlimits[1]))
# & ((assembler_gdf['INTPTLAT'] >= ylimits[0]) & (assembler_gdf['INTPTLAT'] <= ylimits[1]))
# ].iterrows():
# b.append(ix)
# annotator = ix[-6:]
# ax1.annotate(annotator,
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# (float(thisrow['INTPTLON']), float(thisrow['INTPTLAT'])),
# color='black', backgroundcolor='white', fontsize=10, ha='center', zorder=1000
# )
# if (show_water):
# print('plotting water...')
# water_other_gdf[
# (water_other_gdf.geometry.apply(lambda x: x.centroid.x >= xlimits[0])) & ((water_other_gdf.geometry.apply(lambda x: x.centroid.x) <= xlimits[1]))
# & (water_other_gdf.geometry.apply(lambda x: x.centroid.y >= ylimits[0])) & ((water_other_gdf.geometry.apply(lambda x: x.centroid.y) <= ylimits[1]))
# ].plot(ax=ax2, color='blue')
# b = sorted(b)
# for i in range(0,len(b)):
# print(b[i])
# plt.show()
# assembler_gdf.loc[(assembler_gdf['STUSAB'] == this_state) & (assembler_gdf['block_based_district'] < 0), 'block_based_district'] = -1
# +
# # 52 vs 53
# manual_xlim = (-117.24, -116.99)
# manual_ylim = (32.71, 32.84)
# # 51 vs 53
# manual_xlim = (-117.16, -116.65)
# manual_ylim = (32.55, 32.77)
# # 51 vs 52
# manual_xlim = (-117.26, -117.09)
# manual_ylim = (32.57, 32.75)
## 50 vs 53
#manual_xlim = (-117.05, -116.87)
#manual_ylim = (32.64, 32.86)
## 50 vs 52
# manual_xlim = (-117.17, -116.67)
# manual_ylim = (32.80, 33.22)
# # 50 vs 51
# manual_xlim = (-116.93, -116.00)
# manual_ylim = (32.49, 33.39)
## 49 vs 52
# manual_xlim = (-117.29, -117.05)
# manual_ylim = (32.82, 33.10)
# # 49 vs 50
# manual_xlim = (-117.27, -117.04)
# manual_ylim = (33.00, 33.56)
# # 42 vs 50
# manual_xlim = (-117.31, -117.02)
# manual_ylim = (33.43, 33.55)
# 41 vs 42
# manual_xlim = (-117.32, -117.01)
# manual_ylim = (33.67, 33.97)
# # 36 vs 42
# manual_xlim = (-117.19, -116.63)
# manual_ylim = (33.37, 33.98)
# ### 31 vs 35
# manual_xlim = (-117.57, -117.30)
# manual_ylim = (34.00, 34.17)
# # ## 24 vs 26
# manual_xlim = (-119.53, -119.01)
# manual_ylim = (34.21, 34.55)
# # 22 vs 23
# manual_xlim = (-119.36, -118.96)
# manual_ylim = (36.02, 36.72)
# # 21 vs 23
# manual_xlim = (-119.74, -118.72)
# manual_ylim = (35.05, 35.55)
# # 21 vs 22
# manual_xlim = (-119.61, -119.44)
# manual_ylim = (36.53, 36.76)
# ## 16 vs 22
# manual_xlim = (-119.90, -119.66)
# manual_ylim = (36.68, 36.82)
#
## 16 vs 21
# manual_xlim = (-120.11, -119.62)
# manual_ylim = (36.58, 36.87)
# # # 8 vs 31
# manual_xlim = (-117.63, -116.98)
# manual_ylim = (33.96, 34.39)
# ## 6 vs 7
# manual_xlim = (-121.47, -121.24)
# manual_ylim = (38.40, 38.76)
# # # 3 vs 6
# manual_xlim = (-121.73, -121.34)
# manual_ylim = (38.16, 38.85)
# ## 3 vs 5
# manual_xlim = (-123.17, -122.30)
# manual_ylim = (38.68, 39.67)
# ## 2 vs 5
# manual_xlim = (-122.88, -122.51)
# manual_ylim = (38.22, 38.62)
# # # 1 vs 4
# manual_xlim = (-121.55, -119.86)
# manual_ylim = (38.81, 39.60)
# # # 1 vs 3
# manual_xlim = (-122.29, -121.91)
# manual_ylim = (39.43, 39.80)
# ## 46 vs 47
# manual_xlim = (-118.03, -117.92)
# manual_ylim = (33.78, 33.84)
# ## 45 vs 49
# manual_xlim = (-117.70, -117.38)
# manual_ylim = (33.49, 33.74)
# # 45 vs 48
# manual_xlim = (-117.91, -117.64)
# manual_ylim = (33.52, 33.71)
# # 45 vs 46
# manual_xlim = (-117.86, -117.80)
# manual_ylim = (33.73, 33.85)
# # 39 vs 46
# manual_xlim = (-118.02, -117.81)
# manual_ylim = (33.82, 33.88)
# # 39 vs 45
# manual_xlim = (-117.86, -117.61)
# manual_ylim = (33.73, 33.90)
# # 38 vs 40
# manual_xlim = (-118.16, -118.10)
# manual_ylim = (33.86, 34.05)
# # 38 vs 39
# manual_xlim = (-118.05, -117.94)
# manual_ylim = (33.83, 34.02)
# # 37 vs 43
# manual_xlim = (-118.43, -118.26)
# manual_ylim = (33.95, 34.00)
# # # 37 vs 40
# manual_xlim = (-118.29, -118.25)
# manual_ylim = (34.00, 34.05)
# # 35 vs 39
# manual_xlim = (-117.86, -117.63)
# manual_ylim = (33.91, 34.09)
# # 34 vs 40
# manual_xlim = (-118.27, -118.16)
# manual_ylim = (34.01, 34.06)
# # 33 vs 43
# manual_xlim = (-118.47, -118.28)
# manual_ylim = (33.73, 34.00)
# # 33 vs 37
# manual_xlim = (-118.47, -118.41)
# manual_ylim = (33.96, 34.08)
# 32 vs 39
# manual_xlim = (-118.02, -117.80)
# manual_ylim = (33.97, 34.09)
# 32 vs 38
# manual_xlim = (-118.09, -117.98)
# manual_ylim = (33.99, 34.08)
# # 29 vs 30
# manual_xlim = (-118.55, -118.33)
# manual_ylim = (34.13, 34.36)
# # 28 vs 30
# manual_xlim = (-118.39, -118.31)
# manual_ylim = (34.11, 34.19)
# # 28 vs 29
# manual_xlim = (-118.43, -118.22)
# manual_ylim = (34.19, 34.36)
# # 27 vs 32
# manual_xlim = (-118.11, -117.63)
# manual_ylim = (34.06, 34.20)
# # 27 vs 28
# manual_xlim = (-118.33, -118.13)
# manual_ylim = (34.10, 34.33)
# # 25 vs 26
# manual_xlim = (-118.86, -118.63)
# manual_ylim = (34.18, 34.40)
# # 23 vs 25
# manual_xlim = (-118.42, -118.00)
# manual_ylim = (34.56, 34.80)
# # # 19 vs 20
# manual_xlim = (-117.71, -117.38)
# manual_ylim = (34.14, 34.41)
# # 18 vs 20
# manual_xlim = (-122.12, -121.96)
# manual_ylim = (36.93, 37.10)
# # 18 vs 19
# manual_xlim = (-121.98, -121.63)
# manual_ylim = (37.10, 37.37)
# # 17 vs 19
# manual_xlim = (-122.03, -121.13)
# manual_ylim = (36.90, 37.54)
# manual_xlim = (-122.03, -121.7)
# manual_ylim = (37.3, 37.54)
# # 17 vs 18
# manual_xlim = (-122.23, -121.91)
# manual_ylim = (37.17, 37.46)
# # 15 vs 17
# manual_xlim = (-122.18, -121.63)
# manual_ylim = (37.43, 37.69)
# # 14 vs 18
# manual_xlim = (-122.44, -122.1)
# manual_ylim = (37.26, 37.54)
# # 13 vs 15
# manual_xlim = (-122.195, -122)
# manual_ylim = (37.63, 37.83)
# # 12 vs 14
# manual_xlim = (-122.51, -122.4)
# manual_ylim = (37.7, 37.76)
# # 11 vs 15
# manual_xlim = (-122.1,-121.83)
# manual_ylim = (37.72,37.84)
# 9 vs 11
# manual_xlim = (-122,-121.5)
# manual_ylim = (37.7,38.1)
# # 9 vs 10
# manual_xlim = (-121.6,-121.1)
# manual_ylim = (37.68,37.9)
# # 5 vs 11
# manual_xlim = (-122.33,-122.04)
# manual_ylim = (37.77,38.05)
| gerrymander53.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# Most examples work across multiple plotting backends equivalent, this example is also available for:
#
# * [Bokeh - radial_heatmap](../bokeh/radial_heatmap.ipynb)
# +
import numpy as np
import pandas as pd
import holoviews as hv
hv.extension("matplotlib")
# -
# # Declaring data
#
# ### NYC Taxi Data
#
# Let's dive into a concrete example, namely the New York - Taxi Data ([For-Hire Vehicle (“FHV”) records](http://www.nyc.gov/html/tlc/html/about/trip_record_data.shtml)). The following data contains hourly pickup counts for the entire year of 2016.
#
# **Considerations**: Thinking about taxi pickup counts, we might expect higher taxi usage during business hours. In addition, public holidays should be clearly distinguishable from regular business days. Furthermore, we might expect high taxi pickup counts during Friday and Saterday nights.
#
# **Design**: In order model the above ideas, we decide to assign days with hourly split to the *radial segments* and week of year to the *annulars*. This will allow to detect daily/hourly periodicity and weekly trends. To get you more familiar with the mapping of segemnts and annulars, take a look at the following radial heatmap:
# +
# load example data
df_nyc = pd.read_csv("../../../assets/nyc_taxi.csv.gz", parse_dates=["Pickup_date"])
# create relevant time columns
df_nyc["Day & Hour"] = df_nyc["Pickup_date"].dt.strftime("%A %H:00")
df_nyc["Week of Year"] = df_nyc["Pickup_date"].dt.strftime("Week %W")
df_nyc["Date"] = df_nyc["Pickup_date"].dt.strftime("%Y-%m-%d")
heatmap = hv.HeatMap(df_nyc, ["Day & Hour", "Week of Year"], ["Pickup_Count", "Date"])
# -
# # Plot
# **At first glance**: First, let's take a closer look at the mentioned segments and annulars. **Segments** correspond to *hours of a given day* whereas **annulars** represent entire *weeks*. If you use the hover tool, you will quickly get an idea of how segments and annulars are organized. **Color** decodes the pickup values with blue being low and red being high.
#
# **Plot improvements**: The above plot clearly shows systematic patterns however the default plot options are somewhat disadvantageous. Therefore, before we start to dive into the results, let's increase the readability of the given plot:
#
# - **Remove annular ticks**: The information about week of year is not very important. Therefore, we hide it via `yticks=None`.
# - **Custom segment ticks**: Right now, segment labels are given via day and hour. We don't need hourly information and we want every day to be labeled. We can use a tuple here which will be passed to `xticks=("Friday", ..., "Thursday")`
# - **Add segment markers**: Moreover, we want to aid the viewer in distingushing each day more clearly. Hence, we can provide marker lines via `xmarks=7`.
# - **Rotate heatmap**: The week starts with Monday and ends with Sunday. Accordingly, we want to rotate the plot to have Sunday and Monday be at the top. This can be done via `start_angle=np.pi*19/14`. The default order is defined by the global sort order which is present in the data. The default starting angle is at 12 o'clock.
#
# Let's see the result of these modifications:
# +
# %%opts HeatMap [radial=True fig_size=300 yticks=None xmarks=7 ymarks=3 start_angle=np.pi*19/14]
# %%opts HeatMap [xticks=("Friday", "Saturday", "Sunday", "Monday", "Tuesday", "Wednesday", "Thursday")]
heatmap
# -
# After tweaking the plot defaults, we're comfortable with the given visualization and can focus on the story the plot tells us.
#
# **There are many interesting findings in this visualization:**
#
# 1. Taxi pickup counts are high between 7-9am and 5-10pm during weekdays which business hours as expected. In contrast, during weekends, there is not much going on until 11am.
# 2. Friday and Saterday nights clearly stand out with the highest pickup densities as expected.
# 3. Public holidays can be easily identified. For example, taxi pickup counts are comparetively low around Christmas and Thanksgiving.
# 4. Weather phenomena also influence taxi service. There is a very dark blue stripe at the beginning of the year starting at Saterday 23rd and lasting until Sunday 24th. Interestingly, there was one of the [biggest blizzards](https://www.weather.gov/okx/Blizzard_Jan2016) in the history of NYC.
| examples/gallery/demos/matplotlib/nyc_radial_heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings("ignore")
def load_data():
reviewsFile = open('../data/reviews.txt','r')
reviews = list(map(lambda x:x[:-1],reviewsFile.readlines()))
reviewsFile.close()
labelsFile = open('../data/labels.txt','r')
labels = list(map(lambda x:x[:-1],labelsFile.readlines()))
labelsFile.close()
return reviews,labels
reviews,labels = load_data()
from nltk.tokenize import RegexpTokenizer
tokenizer = RegexpTokenizer("\w+\'?\w+|\w+")
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
from spacy.lang.en.stop_words import STOP_WORDS
exceptionStopWords = {
'again',
'against',
'ain',
'almost',
'among',
'amongst',
'amount',
'anyhow',
'anyway',
'aren',
"aren't",
'below',
'bottom',
'but',
'cannot',
'couldn',
"couldn't",
'didn',
"didn't",
'doesn',
"doesn't",
'don',
"don't",
'done',
'down',
'except',
'few',
'hadn',
"hadn't",
'hasn',
"hasn't",
'haven',
"haven't",
'however',
'isn',
"isn't",
'least',
'mightn',
"mightn't",
'move',
'much',
'must',
'mustn',
"mustn't",
'needn',
"needn't",
'neither',
'never',
'nevertheless',
'no',
'nobody',
'none',
'noone',
'nor',
'not',
'nothing',
'should',
"should've",
'shouldn',
"shouldn't",
'too',
'top',
'up',
'very'
'wasn',
"wasn't",
'well',
'weren',
"weren't",
'won',
"won't",
'wouldn',
"wouldn't",
}
stop_words = set(stop_words).union(STOP_WORDS)
final_stop_words = stop_words-exceptionStopWords
import spacy
nlp = spacy.load("en",disable=['parser', 'tagger', 'ner'])
def make_token(review):
return tokenizer.tokenize(str(review))
def remove_stopwords(review):
return [token for token in review if token not in final_stop_words]
def lemmatization(review):
lemma_result = []
for words in review:
doc = nlp(words)
for token in doc:
lemma_result.append(token.lemma_)
return lemma_result
def pipeline(review):
review = make_token(review)
review = remove_stopwords(review)
return lemmatization(review)
# %%time
reviews = list(map(lambda review: pipeline(review),reviews))
reviews[:2]
from gensim.models import Word2Vec
embedding_dimension = 100
model = Word2Vec(reviews,size=embedding_dimension, window=3, min_count=3, workers=4)
model.sg
word_vectors = model.wv
del model
len(word_vectors.vocab)
| section3/s3v3-Exploring word embeddings with Gensim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bus
#
# This bus has a passenger entry and exit control system to monitor the number of occupants it carries and thus detect when there is too high a capacity.
#
# At each stop the entry and exit of passengers is represented by a tuple consisting of two integer numbers.
# ```
# bus_stop = (in, out)
# ```
# The succession of stops is represented by a list of these tuples.
# ```
# stops = [(in1, out1), (in2, out2), (in3, out3), (in4, out4)]
# ```
#
# ## Goals:
# * lists, tuples
# * while/for loops
# * minimum, maximum, length
# * average, standard deviation
#
# ## Tasks
# 1. Calculate the number of stops.
# 2. Assign to a variable a list whose elements are the number of passengers at each stop (in-out),
# 3. Find the maximum occupation of the bus.
# 4. Calculate the average occupation. And the standard deviation.
#
# variables
pass_in = 0
pass_out = 0
pass_total = 0
bus_stop = ()
stops = []
passengers = []
# +
# 1. Calculate the number of stops.
import time
print('Welcome to the Bus Game!\nWe are going to be a bus driver.')
print("\n\nLet's start with instructions: ")
print('1) You should type the number of passegenrs In and Out.')
print('\n2) When the the passagens bus number comes to zero, the bus and game stops.!')
nstop = 0
while True:
pass_in = int(input('Passengers get IN: '))
pass_out = int(input('Passengers get OUT: '))
pass_total += (pass_in - pass_out)
print(f'The Bus has {pass_total} passegenrs')
passengers.append(pass_total)
stops.append( ((pass_in),(pass_out)) )
if pass_total == 0:
print(f'The bus has stopped. The number total os stops was {len(passengers)}')
break
for i in range(2):
print('|----|\n 0--0\n')
time.sleep(1)
# -
# 3. Find the maximum occupation of the bus.
print(passengers)
print(max(passengers))
# 4. Calculate the average occupation. And the standard deviation.
import statistics
print(f"In and out's: {stops}")
print(f"Average occupation: {sum(passengers)/len(passengers)}")
print(f'Standard Deviation: {statistics.pstdev(passengers)}')
| bus/bus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Edge detection
#
# Different edge detection is applied
#
# 1. Laplacian edge detection
# 2. Sobel method
# 3. Canny edge detection
import cv2
import numpy as np
from matplotlib import pyplot as plt
# read the image
img1 = cv2.imread('road.png',0)
# ## Laplacian edge detection
# +
# remove noise
img = cv2.GaussianBlur(img1,(3,3),0)
# convolute with proper kernels
laplacian = cv2.Laplacian(img,cv2.CV_64F)
plt.subplot(2,2,1),plt.imshow(img,cmap = 'gray')
plt.title('Original'), plt.xticks([]), plt.yticks([])
plt.subplot(2,2,2),plt.imshow(laplacian,cmap = 'gray')
plt.title('Laplacian'), plt.xticks([]), plt.yticks([])
plt.show()
# -
# ## Sobel method
# +
# Create a custom kernel
# 3x3 array for edge detection
sobel_y = np.array([[ -1, -2, -1],
[ 0, 0, 0],
[ 1, 2, 1]])
## TODO: Create and apply a Sobel x operator
sobel_x = np.array([[-1,0,1],
[ -2, 0 , 2],
[ -1,0,1]])
# +
# Filter the image using filter2D, which has inputs: (grayscale image, bit-depth, kernel)
filtered_image_y = cv2.filter2D(img1, -1, sobel_y)
filtered_image_x = cv2.filter2D(img1, -1, sobel_x)
plt.subplot(2,2,1),plt.imshow(img1,cmap = 'gray')
plt.title('Gray scale'), plt.xticks([]), plt.yticks([])
plt.subplot(2,2,2),plt.imshow(filtered_image_x,cmap = 'gray')
plt.title('Sobel X'), plt.xticks([]), plt.yticks([])
plt.subplot(2,2,3),plt.imshow(filtered_image_y,cmap = 'gray')
plt.title('Sobel Y'), plt.xticks([]), plt.yticks([])
plt.show()
# -
# ## Canny edge detection
# +
# canny edge detection
edges = cv2.Canny(img,250,250)
plt.subplot(121),plt.imshow(img,cmap = 'gray')
plt.title('Original Image'), plt.xticks([]), plt.yticks([]);
plt.subplot(122),plt.imshow(edges,cmap = 'gray')
plt.title('Edge Image'), plt.xticks([]), plt.yticks([]);
# -
# ### Out of all, canny edge detection is better
| Edge_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="SB26xiG-H5ZT"
# # Housing Market
# + [markdown] id="xdmcOlDUH5ZW"
# ### Introduction:
#
# This time we will create our own dataset with fictional numbers to describe a house market. As we are going to create random data don't try to reason of the numbers.
#
# ### Step 1. Import the necessary libraries
# + id="54WPjRowH5ZX"
import pandas as pd
import numpy as np
# + [markdown] id="_4A79p9zH5ZX"
# ### Step 2. Create 3 differents Series, each of length 100, as follows:
# 1. The first a random number from 1 to 4
# 2. The second a random number from 1 to 3
# 3. The third a random number from 10,000 to 30,000
# + id="QcGeiWxkH5ZY"
random1= np.random.randint(1,4,size=100)
random2=np.random.randint(1,3,size=100)
random3= np.random.randint(10000,30000,size=100)
sr1=pd.Series(random1)
sr2=pd.Series(random2)
sr3=pd.Series(random3)
# + [markdown] id="qxtHEBi3H5ZY"
# ### Step 3. Let's create a DataFrame by joinning the Series by column
# + id="4jYZwNnTH5ZY" outputId="120f8d36-985d-4602-9fbd-a523fd324fde" colab={"base_uri": "https://localhost:8080/", "height": 423}
df= pd.concat([sr1,sr2,sr3],axis=1)
# + [markdown] id="Io8fdd_tH5ZZ"
# ### Step 4. Change the name of the columns to bedrs, bathrs, price_sqr_meter
# + id="bEiqe7zNH5ZZ" outputId="4f677f8b-6707-44d3-8ba1-faa89060c3a5" colab={"base_uri": "https://localhost:8080/", "height": 423}
df.rename(columns={0:'bedrs',1:'bathrs',2:'price_sqr_meter'})
# + [markdown] id="nHiiO4b8H5Za"
# ### Step 5. Create a one column DataFrame with the values of the 3 Series and assign it to 'bigcolumn'
# + id="gIbxJuLSH5Za" outputId="ea8ac2c8-4792-4af6-d245-cd64aa59e828" colab={"base_uri": "https://localhost:8080/", "height": 423}
bigcolumn=pd.concat([sr1,sr2,sr3],axis=0)
bigcolumn=bigcolumn.to_frame()
bigcolumn
# + [markdown] id="Z4yabDNcH5Za"
# ### Step 6. Oops, it seems it is going only until index 99. Is it true?
# + id="rrUv6t-1H5Zb" outputId="30da38ca-42c7-42a5-c5d0-2a39f6f706b7" colab={"base_uri": "https://localhost:8080/"}
len(bigcolumn)
# + [markdown] id="JiQRGG02H5Zb"
# ### Step 7. Reindex the DataFrame so it goes from 0 to 299
# + id="m9_LZeEqH5Zc" outputId="d9c40fe8-ee36-4d6f-dc24-51464654fd27" colab={"base_uri": "https://localhost:8080/", "height": 423}
bigcolumn.reset_index(drop=True, inplace= True)
bigcolumn
# + id="2P63xZs8cbnz"
| 05_Merge/Housing Market/Exercises_solved.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import os
import numpy as np
import pandas
import matplotlib.pyplot as plt
import plot_domain
fig = plot_domain.henry_domain()
# ## Model background
# Here is an example based on the Henry saltwater intrusion problem. The synthetic model is a 2-dimensional SEAWAT model (X-Z domain) with 1 row, 120 columns and 20 layers. The left boundary is a specified flux of freshwater, the right boundary is a specified head and concentration saltwater boundary. The model has two stress periods: an initial steady state (calibration) period, then a transient period with less flux (forecast).
# The inverse problem has 603 parameters: 600 hydraulic conductivity pilot points, 1 global hydraulic conductivity, 1 specified flux multiplier for history matching and 1 specified flux multiplier for forecast conditions. The inverse problem has 36 obseravtions (21 heads and 15 concentrations) measured at the end of the steady-state calibration period. The forecasts of interest of the distance from the left model edge to the 10% seawater concentration in the basal model layer and the concentration at location 10. Both of there forecasts are "measured" at the end of the forecast stress period. The forecasts are both in the Jacobian matrix as zero-weight observations named `pd_ten` and `C_obs10_2`.I previously calculated the jacobian matrix, which is in the `henry/` folder, along with the PEST control file.
#
#
# ##Using `pyemu`
import pyemu
# First create a linear_analysis object. We will use `schur` derived type, which replicates the behavior of the `PREDUNC` suite of PEST. We pass it the name of the jacobian matrix file. Since we don't pass an explicit argument for `parcov` or `obscov`, `pyemu` attempts to build them from the parameter bounds and observation weights in a pest control file (.pst) with the same base case name as the jacobian. Since we are interested in forecast uncertainty as well as parameter uncertainty, we also pass the names of the forecast sensitivity vectors we are interested in, which are stored in the jacobian as well. Note that the `forecasts` argument can be a mixed list of observation names, other jacobian files or PEST-compatible ASCII matrix files.
la = pyemu.Schur(jco=os.path.join("henry", "pest.jcb"),verbose=False)
# The screen output can be redirected to a log file by passing a file name to the `verbose` keyword argument. Or screen output can be stopped by passing `False` to the `verbose` argument
# We can inspect the parcov and obscov attributes by saving them to files. We can save them PEST-compatible ASCII or binary matrices (`.to_ascii()` or `.to_binary()`), PEST-compatible uncertainty files (`.to_uncfile()`), or simply as numpy ASCII arrays (`numpy.savetxt()`). In fact, all matrix and covariance objects (including the forecasts) have these methods.
#
la.parcov.to_uncfile(os.path.join("henry", "parcov.unc"), covmat_file=os.path.join("henry","parcov.mat"))
# When saving an uncertainty file, if the covariance object is diagonal (`self.isdiagonal == True`), then you can force the uncertainty file to use standard deviation blocks instead of covariance matrix blocks by explicitly passing `covmat_file` as `None`:
la.obscov.to_uncfile(os.path.join("henry", "obscov.unc"), covmat_file=None)
# ## Posterior parameter uncertainty analysis
# Let's calculate and save the posterior parameter covariance matrix:
la.posterior_parameter.to_ascii(os.path.join("henry", "posterior.mat"))
# You can open this file in a text editor to examine. The diagonal of this matrix is the posterior variance of each parameter. Since we already calculated the posterior parameter covariance matrix, additional calls to the `posterior_parameter` decorated method only require access:
#
la.posterior_parameter.to_dataframe().sort_index().sort_index(axis=1).iloc[0:3,0:3] #look so nice in the notebook
# We can see the posterior variance of each parameter along the diagonal of this matrix. Now, let's make a simple plot of prior vs posterior uncertainty for the 600 pilot point parameters
par_sum = la.get_parameter_summary().sort_index()
par_sum.loc[par_sum.index[:20],"percent_reduction"].plot(kind="bar",figsize=(10,4),edgecolor="none")
par_sum.iloc[0:10,:]
# We can see that the at most, the uncertainty of any one of the 600 hydraulic conductivity parameters is only reduced by 5% and the uncertainty of many parameters has not been reduced at all, meaning these parameters are not informed by the observations.
# ## Prior forecast uncertainty
# Now let's examine the prior and posterior variance of the forecasts:
la.get_forecast_summary()
# It is interesting that the uncertainty of the forecasts is reduced substantially even though the uncertainty for any one parameter is only slightly reduced. This is because the right combinations of forecast-sensitive parameters are being informed by the observations.
# ## Data worth
# Now, let's try to identify which observations are most important to reducing the posterior uncertainty (e.g.the forecast worth of every observation). We simply recalculate Schur's complement without some observations and see how the posterior forecast uncertainty increases
#
# Let's see which observations are most important, which is measured by the increase in forecast uncertainty when that observation is left out
df = la.get_removed_obs_importance()
df = 100.0 * (df - df.loc["base",:])/df
ax = df.plot(kind="bar",figsize=(15,8))
ax.set_ylabel("percent uncertainty increase")
# ```base``` row are the results of Schur's complement calculation using all observations. The increase in posterior forecast uncertainty for the ```head``` and ```conc``` cases show how much forecast uncertainty increases when the head and concentrations observations are not used in history matching
#
# ## parameter contribution to forecast uncertainty
#
#
# Lets look at which parameters are contributing most to forecast uncertainty, which we estimate as the decrese in forecast uncertainty from "perfect" knowledge of one or more parameters. for demostration purposes, lets group the hydraulic conductivity parameters by row.
# +
par_groups = {}
for pname in la.pst.par_names:
if pname.startswith('k'):
row = "k_row_"+pname[2:4]
if row not in par_groups.keys():
par_groups[row] = []
par_groups[row].append(pname)
par_groups["global_k"] = "global_k"
par_groups["histmatch_mult"] = "mult1"
par_groups["forecast_mult"] = "mult2"
df = la.get_par_contribution(par_groups)
df.sort_index(inplace=True)
df
# -
df
df.plot(kind="bar",figsize=(10,5))
plt.show()
# We see that the largest contributions to forecast uncertainty depends on the forecast. Forecast ```pd_ten``` is most sensitive to hydraulic conductivity parameters in row 10. However, Forecast ```c_obs10_2``` is most sensitive to the ```forecast_mult``` parameter.
| examples/Schurexample_henry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# <script>
# function code_toggle() {
# if (code_shown){
# $('div.input').hide('500');
# $('#toggleButton').val('Show Code')
# } else {
# $('div.input').show('500');
# $('#toggleButton').val('Hide Code')
# }
# code_shown = !code_shown
# }
#
# $( document ).ready(function(){
# code_shown=false;
# $('div.input').hide()
# });
# </script>
# <form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>
# -
# ## Code visibility (relevant for HTML only)
# Use the Show/Hide Code button on the top left to make to make the code visible or hide it. It will be hidden in the HTML files by default.
# # How to use code cells in this notebook
# If a code cell starts with
# ```python
# # RUN
# ```
# Run the cell by CTRL+Enter, or the Run button above.
#
# If a code cell starts with
# ```python
# # USER INPUT
# ```
# User input is needed before running the cell. Usually there will be a cell preceding this which gives an example for the values to be provided.
#
# If a code cell starts with
# ```python
# # OPTIONAL USER INPUT
# ```
# User input is needed before running the cell. However, some defaults are provided, so make sure that either the settings will work for your run, or change them appropriately.
#
# If a cell starts with
# #### Example cell
# These cells are not code cells but examples of user inputs from the test data analysis for the actual code cell that follows it, informing the user about the formatting etc.
#
# **Important note on entering input:** When entering user input, please make sure you follow the formatting provided in the example cells. For example, when the parameter is text, make sure you have quotation marks around the parameters but when it is a number, do not enclose in quotes. If it is a list, then provide a list in brackets.
# RUN
import sys
sys.path.append("/opt/src")
import mip_functions as mip
import probe_summary_generator
import pickle
import json
import copy
import os
import numpy as np
import subprocess
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
from matplotlib.lines import Line2D
plt.rcParams['svg.fonttype'] = 'none'
import pandas as pd
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
import allel
wdir = "/opt/analysis/"
data_dir = "/opt/data/"
targeted_table = pd.read_csv("targeted_mutation_prevalence.csv",
header=list(range(2)), index_col=list(range(4)))
targeted_counts = pd.read_csv("targeted_mutation_counts.csv",
header=list(range(2)), index_col=list(range(4)))
k13_prev = pd.read_csv("k13_min5_prev.csv",
header=list(range(2)), index_col=list(range(4)))
k13_counts = pd.read_csv("k13_min5_counts.csv",
header=list(range(2)), index_col=list(range(4)))
targeted_table.head()
# +
os.environ['PROJ_LIB'] = "/opt/conda/share/proj/"
import geopandas as gp
from mpl_toolkits.basemap import Basemap
from matplotlib.patches import Polygon
from matplotlib import cm
# -
combined = pd.read_csv("combined_prevalences.csv", index_col=0)
combined.head()
import matplotlib as mpl
import matplotlib.gridspec as gridspec
average = combined.pivot_table(index=["Year", "District"],
columns="Mutation Name", values="Prevalence")
average.head()
import geopy
locator = geopy.Nominatim(user_agent="myGeocoder")
district_to_coordinate = {}
for d in combined["District"].unique():
gc = locator.geocode(d + ", Uganda")
district_to_coordinate[d] = [gc.longitude, gc.latitude]
district_to_coordinate
mutations = ["dhps-Ala581Gly", "dhfr-ts-Ile164Leu", "k13-Ala675Val"]
fig, axes = plt.subplots(len(mutations), 2)
years = [2018, 2019]
for i in range(len(mutations)):
mut = mutations[i]
max_value = average[mut].max()
for j in range(2):
ax = axes[i, j]
yr = years[j]
av = average.loc[yr]
m_dict = av[mut].to_dict()
norm = mpl.colors.Normalize(vmin=0, vmax=max_value)
m = Basemap(llcrnrlat=-1.55, llcrnrlon=29.5, urcrnrlat=4.3, urcrnrlon=35.1,
resolution="h", ax=ax)
shp = m.readshapefile("/opt/data/project_data/gadm36_UGA_2", 'states', drawbounds=True, linewidth=0.2)
polygons = {}
for nshape, seg in enumerate(m.states):
polygons[nshape] = Polygon(seg, aa=True, lw=0.2)
m.drawcountries()
m.fillcontinents("lightgrey", lake_color="deepskyblue")
for d in m_dict:
for p in polygons:
poly = polygons[p]
if poly.contains_point(district_to_coordinate[d]):
poly.set_facecolor(cm.get_cmap("Reds")(norm(m_dict[d])))
ax.add_patch(poly)
ax.text(*district_to_coordinate[d], d.capitalize(),
fontsize=3, ha="center", va="bottom")
break
if j == 1:
cbar = fig.colorbar(cm.ScalarMappable(cmap=cm.get_cmap("Reds"), norm=norm),
ax=axes[i,:], shrink=0.75, pad=0.01)
cbar.ax.tick_params(labelsize=4)
cbar.set_label("Prevalence", fontsize=6, x=0)
# ax.set_title(mut, fontdict={"fontsize": 8})
if i == 0:
ax.set_title(2019, fontdict={"fontsize": 10})
else:
ax.set_ylabel(mut, fontdict={"fontsize": 8})
if i == 0:
ax.set_title(2018, fontdict={"fontsize": 10})
fig.set_dpi(300)
fig.set_size_inches(4, 6)
| base_resources/Example-plotting-data-on-maps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
def safelog(x):
return np.log(x + 1e-100)
# ## Read Data
# +
images = np.genfromtxt('hw03_data_set_images.csv',delimiter=',').reshape(5, 39, 320)
labels = np.char.strip(np.genfromtxt('hw03_data_set_labels.csv',delimiter=',',dtype=str),'"').reshape(5,39)
for index, key in enumerate(['A', 'B', 'C', 'D', 'E']):
labels[labels == key] = index + 1
labels = labels.astype(int)
# +
training_images = images[:,:25,:].reshape(125, 320)
training_l = labels[:,:25].reshape(125,)
test_images = images[:,25:,:].reshape(70, 320)
test_l = labels[:,25:].reshape(70,)
K = np.max(training_l)
N_train = training_l.shape[0]
N_test = test_l.shape[0]
training_labels = np.zeros((N_train, K)).astype(int)
training_labels[range(N_train), training_l - 1] = 1
test_labels = np.zeros((N_test, K)).astype(int)
test_labels[range(N_test), test_l - 1] = 1
# -
pcd = []
for i in range(5):
pcd.append(np.sum(training_images[i*25:(i+1)*25], axis=0).flatten()/(training_images.shape[0]/K))
print("pcd[0] => ", pcd[0][:10], "...")
print("pcd[1] => ", pcd[1][:10], "...")
print("pcd[2] => ", pcd[2][:10], "...")
print("pcd[3] => ", pcd[3][:10], "...")
print("pcd[4] => ", pcd[4][:10], "...")
fig, axs = plt.subplots(1,5,figsize=(15,15), sharey=True)
for i in range(5):
axs[i].imshow(pcd[i].reshape(16,20).T, cmap="Greys",interpolation='none')
plt.show()
# +
def score_func(x, pcd):
return [np.dot(x[i].T, safelog(pcd[c])) + np.dot((1-x[i].T), safelog(1 - pcd[c])) + safelog(0.2) for c in range(5)]
training_scores = np.zeros((125,5))
for i in range(125):
training_scores[i] = score_func(training_images, pcd)
# +
y_predicted = np.argmax(training_scores, axis = 1) + 1
confusion_matrix = pd.crosstab(y_predicted, np.sum(training_labels * np.array([1,2,3,4,5]), axis=1), rownames = ['y_predicted'], colnames = ['y_train'])
print("\n",confusion_matrix)
print("\n====================")
print("\n====================")
# +
test_scores = np.zeros((70,5))
for i in range(70):
test_scores[i] = score_func(test_images, pcd)
y_predicted_test = np.argmax(test_scores, axis = 1) + 1
confusion_matrix = pd.crosstab(y_predicted_test, np.sum(test_labels * np.array([1,2,3,4,5]), axis=1), rownames = ['y_predicted'], colnames = ['y_test'])
print(confusion_matrix)
# -
| 3 - Naïve Bayes’ Classifier/hw03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import scipy
import matplotlib.pyplot as plt
from tqdm import tqdm
import sklearn
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from imblearn.over_sampling import SMOTE
import librosa
from dataset import ALCDataset
# %matplotlib inline
# -
import warnings
warnings.filterwarnings('ignore')
SR = 16000
# ### Load data
alc_dataset = ALCDataset('/Users/mazeyu/Desktop/CMU/20fall/18797/project/code/data')
data_train, label_train = alc_dataset.load_data('train', percentage=1.0, num_threads=4)
data_dev1, label_dev1 = alc_dataset.load_data('d1', percentage=1.0, num_threads=4)
data_dev2, label_dev2 = alc_dataset.load_data('d2', percentage=1.0, num_threads=4)
data_test, label_test = alc_dataset.load_data('test', percentage=1.0, num_threads=4)
# +
assert len(data_train) == len(label_train)
assert len(data_dev1) == len(label_dev1)
assert len(data_dev2) == len(label_dev2)
assert len(data_test) == len(label_test)
print('#train: {}'.format(len(data_train)))
print('#dev1: {}'.format(len(data_dev1)))
print('#dev2: {}'.format(len(data_dev2)))
print('#test: {}'.format(len(data_test)))
# -
# ### Feature extraction
class ALCFeature:
def __init__(self, sr):
self.sr = sr
def delete_silence(self, audio, top_db=20):
result = []
intervals = librosa.effects.split(audio, top_db=top_db, frame_length=2048, hop_length=512)
for interval in intervals:
result.append(audio[interval[0]: interval[1]])
result = np.concatenate(result)
return result
def get_mfcc(self, data, label, n_mfcc=20, wsize=0.1, concat=5, scale=True):
x_mfcc = []
y_mfcc = []
record_mfcc = []
wsize = int(wsize * self.sr)
for i in tqdm(range(len(data)), ncols=100, ascii=True, desc='MFCC feature'):
audio = self.delete_silence(data[i])
x = librosa.feature.mfcc(audio, sr=self.sr, n_mfcc=n_mfcc, n_fft=2048, hop_length=512, win_length=wsize, window='hann')
if scale:
x = x - np.min(x, axis=1, keepdims=True)
for j in range(x.shape[1] // concat):
slice_ = x[:, j * concat: (j + 1) * concat]
x_mfcc.append(slice_.flatten())
y_mfcc.append(label[i])
record_mfcc.append(i)
x_mfcc = np.stack(x_mfcc)
y_mfcc = np.array(y_mfcc)
record_mfcc = np.array(record_mfcc)
return x_mfcc, y_mfcc, record_mfcc
def get_pncc(self, data, label):
pass
def get_cqt(self, data, label, n_chroma=12, wsize=0.1, concat=5, scale=True):
x_cqt = []
y_cqt = []
record_cqt = []
wind = np.hamming(int(wsize * self.sr))
for i in tqdm(range(len(data)), ncols=100, ascii=True, desc='CQT feature'):
audio = self.delete_silence(data[i])
x = librosa.feature.chroma_cqt(audio, sr=self.sr, n_chroma=n_chroma, hop_length=512, window=wind)
if scale:
x = x - np.min(x, axis=1, keepdims=True)
for j in range(x.shape[1] // concat):
slice_ = x[:, j * concat: (j + 1) * concat]
x_cqt.append(slice_.flatten())
y_cqt.append(label[i])
record_cqt.append(i)
x_cqt = np.stack(x_cqt)
y_cqt = np.array(y_cqt)
record_cqt = np.array(record_cqt)
return x_cqt, y_cqt, record_cqt
# +
alc_feature = ALCFeature(SR)
smote = SMOTE(random_state=0)
# # MFCC
# x_train, y_train, record_train = alc_feature.get_mfcc(data_train, label_train)
# x_balance, y_balance = smote.fit_resample(x_train, y_train)
# x_dev1, y_dev1, record_dev1 = alc_feature.get_mfcc(data_dev1, label_dev1)
# x_dev2, y_dev2, record_dev2 = alc_feature.get_mfcc(data_dev2, label_dev2)
# x_test, y_test, record_test = alc_feature.get_mfcc(data_test, label_test)
# CQT
x_train, y_train, record_train = alc_feature.get_cqt(data_train, label_train)
x_balance, y_balance = smote.fit_resample(x_train, y_train)
x_dev1, y_dev1, record_dev1 = alc_feature.get_cqt(data_dev1, label_dev1)
x_dev2, y_dev2, record_dev2 = alc_feature.get_cqt(data_dev2, label_dev2)
x_test, y_test, record_test = alc_feature.get_cqt(data_test, label_test)
# -
pca = sklearn.decomposition.PCA(n_components=50)
pca.fit(x_train)
x_train = pca.transform(x_train)
x_balance = pca.transform(x_balance)
x_dev1 = pca.transform(x_dev1)
x_dev2 = pca.transform(x_dev2)
x_test = pca.transform(x_test)
# ### Classification model
class ALCModel:
def __init__(self, method, verbose=None):
if method == 'lr':
if verbose is None:
verbose = 0
self.clf = LogisticRegression(verbose=verbose)
elif method == 'svm':
if verbose is None:
verbose = False
self.clf = SVC(C=1.0, kernel='rbf', verbose=verbose)
elif method == 'forest':
if verbose is None:
verbose = 0
self.clf = RandomForestClassifier(n_estimators=100, verbose=verbose)
elif method == 'adaboost':
self.clf = AdaBoostClassifier(n_estimators=100)
else:
pass
def fit(self, x, y):
self.clf.fit(x, y)
def predict(self, x, record):
prediction = []
probability = []
raw_pred = self.clf.predict(x)
for i in range(record[-1] + 1):
this_pred = raw_pred[record == i]
if len(this_pred) == 0:
this_prob = np.random.uniform(low=0.0, high=1.0)
this_pred = np.random.choice([0, 1])
else:
this_prob = np.mean(this_pred)
this_pred = np.argmax(np.bincount(this_pred))
prediction.append(this_pred)
probability.append(this_prob)
prediction = np.array(prediction)
probability = np.array(probability)
return prediction, probability
def evaluate(self, x, record, label, roc=False):
pred, prob = self.predict(x, record)
acc = np.mean(pred == label)
report = sklearn.metrics.classification_report(label, pred)
if roc:
fpr, tpr, thresholds = sklearn.metrics.roc_curve(label, prob)
plt.figure()
plt.plot(fpr, tpr)
plt.title('ROC Curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
return acc, report
model = ALCModel('svm')
# model.fit(x_train, y_train)
model.fit(x_balance, y_balance)
acc, report = model.evaluate(x_test, record_test, label_test, roc=True)
print(report)
model = ALCModel('lr')
# model.fit(x_train, y_train)
model.fit(x_balance, y_balance)
acc, report = model.evaluate(x_test, record_test, label_test, roc=True)
print(report)
model = ALCModel('forest')
# model.fit(x_train, y_train)
model.fit(x_balance, y_balance)
acc, report = model.evaluate(x_test, record_test, label_test, roc=True)
print(report)
model = ALCModel('adaboost')
# model.fit(x_train, y_train)
model.fit(x_balance, y_balance)
acc, report = model.evaluate(x_test, record_test, label_test, roc=True)
print(report)
| feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="KYJOgQSzdkt6"
# # ResNet50 With Pandas Python UDFs
#
#
#
# + [markdown] colab_type="text" id="LORil9Jzdkt8"
# # Launch Spark
#
# Three configuration items have to be added to the Spark configuration to enable Arrow as it is disabled by default. This can be done without modifying SparkLauncher now, but you can just modify that if you like.
#
# ```python
# # Apache Arrow Config
# conf.set('spark.yarn.appMasterEnv.ARROW_PRE_0_15_IPC_FORMAT', '1')
# conf.set('spark.executorEnv.ARROW_PRE_0_15_IPC_FORMAT', '1')
# conf.set('spark.sql.execution.arrow.enabled', 'true')
# ```
# + colab={} colab_type="code" id="3qW5QRCEdkt8" jupyter={"outputs_hidden": true} outputId="5d695fd6-9cae-49bf-f612-1cb9d9b4b9e8"
# !pip install keras --no-cache-dir
# !pip install tensorflow --no-cache-dir
# + colab={} colab_type="code" id="yh4lzKlCdkuB" outputId="437fe97e-b2e0-4756-a30d-a21501ea9b25"
import import_ipynb
from data603 import SparkLauncher
from data603 import HDFS
#import extra libraries
import io
from io import StringIO, BytesIO
# get a configuration object
conf = SparkLauncher.get_spark_conf()
# add a file to the configuration that will get copied to all the nodes on the cluster
conf.set('spark.yarn.dist.files', 'keras_data/resnet50_weights_tf_dim_ordering_tf_kernels.h5')
# Apache Arrow Config
conf.set('spark.yarn.appMasterEnv.ARROW_PRE_0_15_IPC_FORMAT', '1')
conf.set('spark.executorEnv.ARROW_PRE_0_15_IPC_FORMAT', '1')
conf.set('spark.sql.execution.arrow.enabled', 'true')
# launch the cluster using the configuration
spark = SparkLauncher.get_spark_session(pack_venv = False, conf = conf)
hdfs = HDFS.get_hdfs()
# + [markdown] colab_type="text" id="dUiH5dswdkuF"
# # Read Dataframe
#
# This must be done _BEFORE_ the UDF is defined because the UDF needs the schemas of the dataframes it will be using.For this section I have a parquet file written out with the bounding boxes extracted of several cat types.
# + colab={} colab_type="code" id="vGmef7h-dkuG"
image_chips = spark.read.parquet("/user/has1/chips_image.parquet")
image_chips = image_chips.drop('data') # remove the full-image data.
# + colab={} colab_type="code" id="bi3tYeC1dkuJ" outputId="e1da802d-90d7-4f48-d1c9-6fefacced2ef"
image_chips.count()
# + colab={} colab_type="code" id="kioYDyTjdkuM" outputId="e6f73f8e-0839-49bc-995f-8a3a63481268"
len(image_chips.columns)
# + colab={} colab_type="code" id="zxNCzn9jdkuO" outputId="45706321-bab0-4313-fc77-feee7b610d7f"
#Checking to see its there
image_chips.show(10)
# + colab={} colab_type="code" id="j-nwFABpdkuT" outputId="4898c3dc-cb0b-4dc5-fa93-b1d4f8339a69"
image_chips.printSchema()
# + [markdown] colab_type="text" id="owO-rBwHdkuX"
# # Add In a Grouping Column
#
#
# + colab={} colab_type="code" id="Q6ggPncCdkuY"
from pyspark.sql.functions import monotonically_increasing_id
from pyspark.sql.functions import col, lit, udf
from pyspark.sql.types import IntegerType
def group_id(n):
ret = n % 10
return ret
udf_group_id = udf(group_id, IntegerType())
# create the counter 1 - # of rows
image_chips = image_chips.withColumn("n", monotonically_increasing_id())
# modulo the counter to get a repeating pattern of 0,1,2,3,4,5,6,7,8,9 for the group number
image_chips = image_chips.withColumn("grp", udf_group_id("n"))
# + [markdown] colab_type="text" id="BPccki9Jdkub"
# # Create Ouput Column(s)
#
#
# + colab={} colab_type="code" id="19KgppcYdkuc"
# create two empty columns for result of udf
image_chips = image_chips.withColumn('prediction_label', lit(""))
image_chips = image_chips.withColumn('prediction_confidence', lit(0.0))
# + [markdown] colab_type="text" id="X5HSPHMGdkue"
# # Create the Pandas UDF
#
#
# + colab={} colab_type="code" id="YMwg4TD3dkuf"
import pandas as pd
from pyspark.sql.functions import pandas_udf, PandasUDFType
@pandas_udf(image_chips.schema, PandasUDFType.GROUPED_MAP)
def evaluate_chip(pdf):
# pdf is a pandas dataframe
import io
import os
from keras.applications.resnet50 import ResNet50
from keras.applications.resnet50 import preprocess_input
from keras.applications.resnet50 import decode_predictions
from keras.preprocessing.image import load_img
from keras.preprocessing.image import img_to_array
# Load Model Data
model = ResNet50(weights = f'{os.getcwd()}/resnet50_weights_tf_dim_ordering_tf_kernels.h5',
include_top = True)
# Create arrays to hold prediction outputs.
prediction_label = []
prediction_confidence = []
for chip_data in pdf['chip_data']:
# Load the image
img = load_img(io.BytesIO(chip_data), target_size = (224,224))
# Prepare Image
image = img_to_array(img)
image = image.reshape((1, image.shape[0], image.shape[1], image.shape[2]))
image = preprocess_input(image)
# Run prediction
yhat = model.predict(image)
# Decode Predictions
label = decode_predictions(yhat)
label = label[0][0] # Get first prediction (most confident)
# Save prediction results in arrays
prediction_label.append(label[1])
prediction_confidence.append(label[2])
# Assign result array data to the correct columns in the pandas dataframe
pdf['prediction_label'] = prediction_label
pdf['prediction_confidence'] = prediction_confidence
return pdf
# + [markdown] colab_type="text" id="s5Bgl8dQdkui"
# # Group the Image Dataframe, Apply Pandas UDF
#
# Using the group column to separate the data into processing chunks, call `apply` on each chunk to apply the Pandas UDF.
# + colab={} colab_type="code" id="f0qs3zpDdkui"
image_chips = image_chips.groupby('grp').apply(evaluate_chip)
# + [markdown] colab_type="text" id="9TuDSYiFdkum"
# # View the Result!
#
# Since Spark does lazy evaluation, this next line will take some time to process, but if we've done everything right, we shouldn't blow up the cluster memory limits.
# + colab={} colab_type="code" id="5P53K6DXdkum" outputId="f1d396d2-4891-4857-d9e9-8fa00714cf88"
image_chips.select(['prediction_label', 'prediction_confidence']).show(100)
#image_chips.select(['prediction_label', 'prediction_confidence'])
# + [markdown] colab_type="text" id="lP85oizNdkup"
# # Analyze the Result!
#
# At this point the `image_chips` dataframe is any other Spark dataframe, so process it accordingly.
# + colab={} colab_type="code" id="w31IxgdTdkuq"
ic_summary = image_chips.filter('prediction_confidence > 0.90')\
.groupby('prediction_label')\
.count()\
.filter("count > 100")\
.sort(col("count").desc())
# + colab={} colab_type="code" id="-KPK9x6Hdkus" outputId="12fbd959-c29d-4c95-cf1f-a0546f0d8e70"
ic_summary.show()
# + [markdown] colab_type="text" id="a9KST4cVdkuv"
# # I've only Found One Issue
#
# So far, the issue I found was that `toPandas()` on a dataframe no longer works. We might be able to fix this, but at this point it's a small loss. You can use `.collect()` and `.show` to view data instead.
# + colab={} colab_type="code" id="gKr1sK4_dkuv" jupyter={"outputs_hidden": true} outputId="3ff654d5-9998-47bc-e713-06b09a2a3f1f"
ic_summary.toPandas()
# + colab={} colab_type="code" id="coZvlifDdkuy"
spark.stop()
| oldCode/projectPart2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vasudevgupta7/bigbird/blob/main/notebooks/evaluate_nq.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="XZmucHS1ND0g" outputId="2ebe909f-43d2-43bd-e7cd-be773f6fd1b8"
# !nvidia-smi
# + id="gKcvuGzhJVUq"
# %%capture
# !git clone https://github.com/vasudevgupta7/bigbird
# !cd bigbird/natural-questions && pip3 install -r requirements.txt
# + colab={"base_uri": "https://localhost:8080/"} id="wg9sZulYmJkk" outputId="911b260b-a390-41e0-c87e-7cfc17b6dc2d"
# cd bigbird/natural-questions
# + colab={"base_uri": "https://localhost:8080/"} id="h0i-0EsuWZPA" outputId="9f9e8442-2825-4eca-e1ba-2304b62997e6"
# !mkdir natural-questions-validation
# !wget https://huggingface.co/datasets/vasudevgupta/natural-questions-validation/resolve/main/natural_questions-validation.arrow -P natural-questions-validation
# !wget https://huggingface.co/datasets/vasudevgupta/natural-questions-validation/resolve/main/dataset_info.json -P natural-questions-validation
# !wget https://huggingface.co/datasets/vasudevgupta/natural-questions-validation/resolve/main/state.json -P natural-questions-validation
# + colab={"base_uri": "https://localhost:8080/"} id="q8BttvymQP3u" outputId="cc2b61d9-a77b-45b5-8ec2-dc247b966ec0"
from datasets import load_dataset, load_from_disk
dataset = load_from_disk("natural-questions-validation")
dataset
# + id="bDms6scxZlz6"
def format_dataset(sample):
question = sample['question']['text']
context = sample['document']['tokens']['token']
is_html = sample['document']['tokens']['is_html']
long_answers = sample['annotations']['long_answer']
short_answers = sample['annotations']['short_answers']
context_string = " ".join([context[i] for i in range(len(context)) if not is_html[i]])
# 0 - No ; 1 - Yes
for answer in sample['annotations']['yes_no_answer']:
if answer == 0 or answer == 1:
return {"question": question, "context": context_string, "short": [], "long": [], "category": "no" if answer == 0 else "yes"}
short_targets = []
for s in short_answers:
short_targets.extend(s['text'])
short_targets = list(set(short_targets))
long_targets = []
for s in long_answers:
if s['start_token'] == -1:
continue
answer = context[s['start_token']: s['end_token']]
html = is_html[s['start_token']: s['end_token']]
new_answer = " ".join([answer[i] for i in range(len(answer)) if not html[i]])
if new_answer not in long_targets:
long_targets.append(new_answer)
category = "long_short" if len(short_targets + long_targets) > 0 else "null"
return {"question": question, "context": context_string, "short": short_targets, "long": long_targets, "category": category}
# + colab={"base_uri": "https://localhost:8080/"} id="ub7MRJdBe247" outputId="3a7a77cb-3537-45c2-8a0c-614271441768"
dataset = dataset.map(format_dataset).remove_columns(["annotations", "document", "id"])
# + colab={"base_uri": "https://localhost:8080/", "height": 151, "referenced_widgets": ["a92b261c1ac74524b3eee8e8afcd7df9", "7276f1d7421245dc9ff13fccadfde9fd", "01a236fbe4ac45898654849a51bba080", "1c5e075d504746cd9769e6192e747428", "31c9d34424a44a6bb5f2d6913222090e", "32b073da7f05449da911e6a3d58fe55d", "f54d459b7507437fa258e900425151d3", "4ad75d04b4824daaaacd278edd668cd0"]} id="O5NZr2cjYLAH" outputId="2f440f2a-a8fe-4941-f57f-e587d582fc68"
short_validation_dataset = dataset.filter(lambda x: (len(x['question']) + len(x['context'])) < 4 * 4096)
short_validation_dataset = short_validation_dataset.filter(lambda x: x['category'] != "null")
short_validation_dataset
# + id="03nV8RZpYPdI"
PUNCTUATION_SET_TO_EXCLUDE = set(''.join(['‘', '’', '´', '`', '.', ',', '-', '"']))
def get_sub_answers(answers, begin=0, end=None):
return [" ".join(x.split(" ")[begin:end]) for x in answers if len(x.split(" ")) > 1]
def expand_to_aliases(given_answers, make_sub_answers=False):
if make_sub_answers:
# if answers are longer than one word, make sure a predictions is correct if it coresponds to the complete 1: or :-1 sub word
# *e.g.* if the correct answer contains a prefix such as "the", or "a"
given_answers = given_answers + get_sub_answers(given_answers, begin=1) + get_sub_answers(given_answers, end=-1)
answers = []
for answer in given_answers:
alias = answer.replace('_', ' ').lower()
alias = ''.join(c if c not in PUNCTUATION_SET_TO_EXCLUDE else ' ' for c in alias)
answers.append(' '.join(alias.split()).strip())
return set(answers)
# + id="l5KD7eKaYXdH"
def get_best_valid_start_end_idx(start_scores, end_scores, top_k=1, max_size=100):
best_start_scores, best_start_idx = torch.topk(start_scores, top_k)
best_end_scores, best_end_idx = torch.topk(end_scores, top_k)
widths = best_end_idx[:, None] - best_start_idx[None, :]
mask = torch.logical_or(widths < 0, widths > max_size)
scores = (best_end_scores[:, None] + best_start_scores[None, :]) - (1e8 * mask)
best_score = torch.argmax(scores).item()
return best_start_idx[best_score % top_k], best_end_idx[best_score // top_k]
# + id="-rSFgoWeKXyj"
def evaluate(example):
# encode question and context so that they are seperated by a tokenizer.sep_token and cut at max_length
inputs = tokenizer(example["question"], example["context"], return_tensors="pt", max_length=4096, padding="max_length", truncation=True)
inputs = {k: inputs[k].to(DEVICE) for k in inputs}
with torch.no_grad():
outputs = model(**inputs)
start_scores = outputs['start_logits']
end_scores = outputs['end_logits']
_, category = outputs["cls_out"].max(dim=-1)
predicted_category = CATEGORY_MAPPING[category.item()]
example['targets'] = example['long'] + example['short']
if example['category'] in ['yes', 'no', 'null']:
example['targets'] = [example['category']]
example['has_tgt'] = example['category'] != 'null'
# Now target can be: "yes", "no", "null", "list of long & short answers"
if predicted_category in ['yes', 'no', 'null']:
example['output'] = [predicted_category]
example['match'] = example['output'] == example['targets']
example['has_pred'] = predicted_category != 'null'
return example
max_size = 38 if predicted_category == "short" else 1024
start_score, end_score = get_best_valid_start_end_idx(start_scores[0], end_scores[0], top_k=8, max_size=max_size)
input_ids = inputs["input_ids"][0].cpu().tolist()
example["output"] = [tokenizer.decode(input_ids[start_score: end_score+1])]
answers = expand_to_aliases(example["targets"], make_sub_answers=True)
predictions = expand_to_aliases(example["output"])
# if there is a common element, it's a exact match
example["match"] = len(list(answers & predictions)) > 0
example["has_pred"] = predicted_category != 'null' and len(predictions) > 0
return example
# + id="vG8-p763FjaN" colab={"base_uri": "https://localhost:8080/"} outputId="b92f79ab-a1a2-48f1-8e23-7a8e8f4aa476"
import torch
import numpy as np
from train_nq import BigBirdForNaturalQuestions
from params import CATEGORY_MAPPING
from transformers import BigBirdTokenizer
CATEGORY_MAPPING = {v: k for k, v in CATEGORY_MAPPING.items()}
CATEGORY_MAPPING
# + colab={"base_uri": "https://localhost:8080/", "height": 262, "referenced_widgets": ["2d4e55e101d94bf18a576912ad573bb0", "dbea65321bc8474d8c5da6450a51051d", "<KEY>", "96e5c6868d8f4d858edb908745776e75", "ef6f3127b7fe465ea0851b1f6b382e3b", "befc7b5edb3e4205bb4cb3e68a7acacc", "7e06f985d4b24a9bb926e724eccfda0f", "b6433fa8b6ce41a6b5096010995c23c1", "65212dfdc4cb46f09fde38d5350a3e9c", "aa055c2b94ba44fe8c65da597703a6ec", "e93b195d28bc4ae99970d82343cf2f2c", "<KEY>", "<KEY>", "4a4377a20083444d911d462f587dda42", "<KEY>", "<KEY>", "d530be617e2b4e9da605b5add82b5c58", "<KEY>", "e73364b8a37648738ee9be1461ed5a0c", "98453f4b5e724121abe9f395560e39cf", "dda68efd7a25451cac24e197a74559c3", "eaa51b636261404d88c4394a4067d09c", "<KEY>", "f99529ff22424d609e857422dfa876c3", "<KEY>", "fee46ad24fca4f64917e580273763e0d", "488d0045392e470d9215b25907c78697", "1e2ca51d3c844e91ade56be8f2ae1085", "ad639372435e47be889ad81793d694ce", "<KEY>", "<KEY>", "279c1272072c4959a189ed41531175d0", "<KEY>", "082f62ce95af4b2a99fa339616538aa9", "23f6894f42b243d38e25224d0ed69993", "<KEY>", "9b29aee4d69844ed9e277adec3f8f787", "<KEY>", "<KEY>", "0062b210b82949eb81bd42e232650384"]} id="FgBtSb7XKfU7" outputId="f62cc2b6-f18e-4391-9534-e41b51307eef"
MODEL_ID = "vasudevgupta/bigbird-roberta-natural-questions"
DEVICE = torch.device("cuda" if torch.cuda.is_available() else "cpu")
revision = "b962e30f2367cbc5e35b2c0d64faa9bad469e2e2"
model = BigBirdForNaturalQuestions.from_pretrained(MODEL_ID, revision=revision).to(DEVICE)
tokenizer = BigBirdTokenizer.from_pretrained(MODEL_ID, revision=revision)
# + id="GWiMuSiMK85d" colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["99092795f53945e6815162ae061adead", "3a4c173e70444767b81c02efd5b43c9f", "fdd34006f89e4cf58fd243e9e888aed0", "3f095077ebb7427fa5f4e50b5f40cf13", "dff59adf90df4615ab3027629f490ac2", "67ee5da33d4c4316a968d3f34ab70029", "914729ce4e6a445b94fe0d81e868b13e", "c53d27f5843a41229131136e54a073c8"]} outputId="eb8d3332-af84-44da-c28a-1760bd9c7a46"
def evaluate_print(example, verbose=0):
example = evaluate(example)
if verbose != 0:
print("TARGET", example["short"] if len(example["short"]) > 0 else example['long'])
print("PREDICTION", example["output"], end="\n\n")
return example
short_validation_dataset = short_validation_dataset.map(lambda x: evaluate_print(x, 0))
# + id="a5hs_Ym4Lc-2" colab={"base_uri": "https://localhost:8080/", "height": 83, "referenced_widgets": ["505efaf8832247d18b38a4270e94293e", "0f60f005e2114cf6a209ac39a7160961", "769585183a244dda883141f7070ed74b", "5c236ec4ecbc4580afd44292dd833c4e", "2db678fe25514bbfaf841d8535178dd4", "97f54815cba54e70a0edaeab4999d7aa", "ab7aa876fe2f47a1a2b360ab53a1067e", "8d288c71b85947d09ac8f7ceae8ed064"]} outputId="040b3e1e-ca03-4797-c1ea-9bad289849a5"
total = len(short_validation_dataset)
matched = len(short_validation_dataset.filter(lambda x: x["match"] == 1))
print("EM score:", (matched / total)*100, "%")
# + [markdown] id="tyql0RUKVP-G"
# We are getting **Exact Match ~ 47.45** 💥💥
# <!-- # this f1 is as per official nq script from here (https://github.com/google-research-datasets/natural-questions/blob/master/nq_eval.py)
# has_pred = len(short_validation_dataset.filter(lambda x: x["has_pred"]))
# has_tgt = len(short_validation_dataset.filter(lambda x: x["has_tgt"]))
# matched = len(short_validation_dataset.filter(lambda x: x["match"]))
# precision = matched / has_pred
# recall = matched / has_tgt
# print("F1 score:", 2*precision*recall / (precision + recall)) -->
# + id="8GFcWmv-QFMB"
| notebooks/evaluate_nq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # chemdiab dataset example
#
# Data fom
# [locfit R package](https://cran.r-project.org/web/packages/locfit/locfit.pdf).
#
# Numeric variables are rw, fpg, ga, ina and sspg. Classifier cc is the Diabetic type.
#
# Originally from **<NAME>. and <NAME>. (1979). An attempt to define the nature of chemical diabetes using a multidimensional analysis. Diabetologia 16, 17-24.**
#
#
# +
# import data
import numpy as np
import inspect
from os import path
dtype = [("rw",np.float64),("fpg",np.float64),("ga",np.float64),
("ina",np.float64),('sspg', np.float64),('cc', 'S20')]
file_path = path.dirname(path.abspath(
inspect.getfile(inspect.currentframe())))
file_path = path.join(path.dirname(file_path),"chemdiab.tab")
chemdiab = np.genfromtxt("chemdiab.tab", names=True,
dtype=dtype)
# get only numeric columns as numpy array
nam_cols = list(chemdiab.dtype.names[0:5])
num_cols = chemdiab[nam_cols]
num_cols = num_cols.view(np.float64)\
.reshape(num_cols.shape + (-1,))
cat, ind = np.unique(chemdiab[chemdiab.dtype.names[5]],
return_inverse=True)
# +
# clean and visualize data
from sklearn.preprocessing import StandardScaler, FunctionTransformer
from cartographer.filterers import KernelDensityFilterer
from seaborn import plt
std_scaler = StandardScaler()
scaled = std_scaler.fit_transform(num_cols)
fig, axs = plt.subplots(2,3, figsize=(16,8))
flat_axs = axs.flatten()
for i, name in enumerate(nam_cols):
flat_axs[i].hist(scaled[:,i])
flat_axs[i].set_title(name)
flat_axs[5].hist(ind)
flat_axs[5].set_title("category")
fig
# +
from cartographer.mapper import Mapper
from cartographer.coverers import HyperRectangleCoverer
from sklearn.cluster import DBSCAN
from cartographer.visualization import html_graph
from IPython.core.display import HTML
m = Mapper(coverer=HyperRectangleCoverer(intervals=10, overlap=0.5),
filterer=KernelDensityFilterer(bandwidth=1.0),
clusterer=DBSCAN(min_samples=5,eps=2.))
m.fit(scaled)
HTML(html_graph(m, {"ind": ind}, {"kde": m.filterer.transform(scaled)}))
| examples/chemdiab_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Train POS Tagger in French by Spark NLP
# ### Based on Universal Dependency `UD_French-GSD`
#
# +
import sys
import time
#Spark ML and SQL
from pyspark.ml import Pipeline, PipelineModel
from pyspark.sql.functions import array_contains
from pyspark.sql import SparkSession
from pyspark.sql.types import StructType, StructField, IntegerType, StringType
#Spark NLP
import sparknlp
from sparknlp.annotator import *
from sparknlp.common import RegexRule
from sparknlp.base import DocumentAssembler, Finisher
# -
# ### Let's create a Spark Session for our app
# +
spark = sparknlp.start()
print("Spark NLP version: ", sparknlp.version())
print("Apache Spark version: ", spark.version)
# -
# Let's prepare our training datasets containing `token_posTag` like `de_DET`. You can download this data set from Amazon S3:
#
# ```
# wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/fr/pos/UD_French/UD_French-GSD_2.3.txt -P /tmp
# ```
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/fr/pos/UD_French/UD_French-GSD_2.3.txt -P /tmp
from sparknlp.training import POS
training_data = POS().readDataset(spark, '/tmp/UD_French-GSD_2.3.txt', '_', 'tags')
training_data.show()
# +
document_assembler = DocumentAssembler() \
.setInputCol("text")
sentence_detector = SentenceDetector() \
.setInputCols(["document"]) \
.setOutputCol("sentence")
tokenizer = Tokenizer() \
.setInputCols(["sentence"]) \
.setOutputCol("token")\
.setExceptions(["jusqu'", "aujourd'hui", "États-Unis", "lui-même", "celui-ci", "c'est-à-dire", "celle-ci", "au-dessus", "etc.", "sud-est", "Royaume-Uni", "ceux-ci", "au-delà", "elle-même", "peut-être", "sud-ouest", "nord-ouest", "nord-est", "Etats-Unis", "Grande-Bretagne", "Pays-Bas", "eux-mêmes", "porte-parole", "Notre-Dame", "puisqu'", "week-end", "quelqu'un", "celles-ci", "chef-lieu"])\
.setPrefixPattern("\\A([^\\s\\p{L}\\d\\$\\.#]*)")\
.setSuffixPattern("([^\\s\\p{L}\\d]?)([^\\s\\p{L}\\d]*)\\z")\
.setInfixPatterns([
"([\\p{L}\\w]+'{1})",
"([\\$#]?\\d+(?:[^\\s\\d]{1}\\d+)*)",
"((?:\\p{L}\\.)+)",
"((?:\\p{L}+[^\\s\\p{L}]{1})+\\p{L}+)",
"([\\p{L}\\w]+)"
])
posTagger = PerceptronApproach() \
.setNIterations(1) \
.setInputCols(["sentence", "token"]) \
.setOutputCol("pos") \
.setPosCol("tags")
pipeline = Pipeline(stages=[
document_assembler,
sentence_detector,
tokenizer,
posTagger
])
# -
# Let's train our Pipeline by using our training dataset
model = pipeline.fit(training_data)
# This is our testing DataFrame where we get some sentences in French. We are going to use our trained Pipeline to transform these sentence and predict each token's `Part Of Speech`.
dfTest = spark.createDataFrame([
"Je sens qu'entre ça et les films de médecins et scientifiques fous que nous avons déjà vus, nous pourrions emprunter un autre chemin pour l'origine.",
"On pourra toujours parler à propos d'Averroès de décentrement du Sujet."
], StringType()).toDF("text")
predict = model.transform(dfTest)
predict.select("token.result", "pos.result").show(truncate=50)
| tutorials/old_generation_notebooks/jupyter/3- Build your own French POS tagger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# > ### <font color= "1F284B" >**WEB SCRAPER**</font><font color= "3B4D90" >_Inmobiliaria Gilmar.</font>
# ------
# ##### <font color= "1F284B" >PASO 0. IMPORTAR BIBLIOTECAS.</font>
from bs4 import BeautifulSoup
from selenium import webdriver
import pandas as pd
from time import sleep, strftime
# ------
# ##### <font color= "1F284B" >PASO 1. FUNCIÓN PARA SACAR INFORMACIÓN DE LA WEB.</font>
def sacar_info(url):
chrome_driver = "C:/Users/nuria/Bootcamp Data Science/02. Febrero/chromedriver"
options = webdriver.ChromeOptions()
options.add_argument('--incognito')
driver = webdriver.Chrome(executable_path=chrome_driver,options = options)
driver.get(url)
# Sacamos las características de cada vivienda.
zona_barrio = driver.find_element_by_xpath("//p[@class = 'zona']").text
precio = driver.find_element_by_xpath("//div[@class = 'referencia']/p[3]").text.split(": ")[1]
precio_m2 = driver.find_element_by_xpath("//div[@class = 'referencia']/p[4][1]").text.split(": ")[1]
m2_construidos = driver.find_element_by_xpath("//ul[2]/li[1]/span").text
habs = driver.find_element_by_xpath("//ul[2]/li[2]/span").text.split(" ")[0]
baños = driver.find_element_by_xpath("//ul[2]/li[3]/span").text.split(" ")[0]
terraza = driver.find_element_by_xpath("//ul[2]/li[4]/span").text.split(", ")[0]
trastero = driver.find_element_by_xpath("//ul[2]/li[5]/span").text
garaje = driver.find_element_by_xpath("//ul[2]/li[6]/span").text.split(",")[0]
return zona_barrio, precio, precio_m2, m2_construidos, habs, baños, terraza, trastero, garaje
# ------
# ##### <font color= "1F284B" >PASO 2. SACAMOS INFORMACIÓN DE CADA BARRIO.</font>
# #### <font color= "1F284B" >1. Centro.</font>
urls_1centro = []
for i in range(1,103):
try:
urls_1centro.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/centro/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_1centro = []
for i in urls_1centro:
info_piso = sacar_info(i)
lista_1centro.append(info_piso)
df_1_centro = pd.DataFrame(lista_1centro, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_1_centro.to_excel("df_1_centro.xlsx",index=True)
# #### <font color= "1F284B" >2. Arganzuela.</font>
urls_2arganzuela = []
for i in range(1,47):
try:
urls_2arganzuela.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/arganzuela/gilmar(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_2arganzuela = []
for i in urls_2arganzuela:
info_piso = sacar_info(i)
lista_2arganzuela.append(info_piso)
df_2_arganzuela = pd.DataFrame(lista_2arganzuela, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_2_arganzuela.to_excel("df_2_arganzuela.xlsx",index=True)
#
# #### <font color= "1F284B" >3. Retiro.</font>
urls_3retiro = []
for i in range(1,85):
try:
urls_3retiro.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/retiro/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_3retiro = []
for i in urls_3retiro:
info_piso = sacar_info(i)
lista_3retiro.append(info_piso)
df_3_retiro = pd.DataFrame(lista_3retiro, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_3_retiro.to_excel("df_3_retiro.xlsx",index=True)
# #### <font color= "1F284B" >4. Salamanca.</font>
urls_4salamanca = []
for i in range(1,152):
try:
urls_4salamanca.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/salamanca/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_4salamanca = []
for i in urls_4salamanca:
info_piso = sacar_info(i)
lista_4salamanca.append(info_piso)
df_4_salamanca = pd.DataFrame(lista_4salamanca, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_4_salamanca.to_excel("df_4_salamanca.xlsx",index=True)
# #### <font color= "1F284B" >5. Chamartín.</font>
urls_5chamartin = []
for i in range(1,107):
try:
urls_5chamartin.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/chamartin/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_5chamartin = []
for i in urls_5chamartin:
info_piso = sacar_info(i)
lista_5chamartin.append(info_piso)
df_5_chamartin = pd.DataFrame(lista_5chamartin, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_5_chamartin.to_excel("df_5_chamartin.xlsx",index=True)
# #### <font color= "1F284B" >6. Tetuán. (No hay pisos de este barrio en gilmar)</font>
# #### <font color= "1F284B" >7. Chamberí.</font>
urls_7chamberi = []
for i in range(1,53):
try:
urls_7chamberi.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/chamberi/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_7chamberi = []
for i in urls_7chamberi:
info_piso = sacar_info(i)
lista_7chamberi.append(info_piso)
df_7_chamberi = pd.DataFrame(lista_7chamberi, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_5_chamartin.to_excel("df_5_chamartin.xlsx",index=True)
# #### <font color= "1F284B" >8. Fuencarral.</font>
urls_8fuencarral = []
for i in range(1,29):
try:
urls_8fuencarral.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/fuencarral/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_8fuencarral = []
for i in urls_8fuencarral:
info_piso = sacar_info(i)
lista_8fuencarral.append(info_piso)
df_8_fuencarral = pd.DataFrame(lista_8fuencarral, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_8_fuencarral.to_excel("df_8_fuencarral.xlsx",index=True)
# #### <font color= "1F284B" >9. Fuencarral.</font>
urls_9moncloa = []
for i in range(1,32):
try:
urls_9moncloa.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/moncloa/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_9moncloa = []
for i in urls_9moncloa:
info_piso = sacar_info(i)
lista_9moncloa.append(info_piso)
df_9_moncloa = pd.DataFrame(lista_9moncloa, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_9_moncloa.to_excel("df_9_moncloa.xlsx",index=True)
# #### <font color= "1F284B" >10. Latina.</font>
for i in range(1,12):
try:
urls_10latina.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/latina/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_10latina = []
for i in urls_10latina:
info_piso = sacar_info(i)
lista_10latina.append(info_piso)
df_10_latina = pd.DataFrame(lista_10latina, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_10_latina.to_excel("df_10_latina.xlsx",index=True)
# #### <font color= "1F284B" >11. Carabanchel.</font>
urls_11carabanchel = []
for i in range(1,27):
try:
urls_11carabanchel.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/carabanchel/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_11carabanchel = []
for i in urls_11carabanchel:
info_piso = sacar_info(i)
lista_11carabanchel.append(info_piso)
df_11_carabanchel = pd.DataFrame(lista_11carabanchel, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_11_carabanchel.to_excel("df_11_carabanchel.xlsx",index=True)
# #### <font color= "1F284B" >12. Usera. (No hay pisos de este barrio en gilmar)</font>
# #### <font color= "1F284B" >13. <NAME>.</font>
urls_13ptevallecas = []
for i in range(1,7):
try:
urls_13ptevallecas.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/puente%20de%20vallecas/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_13ptevallecas = []
for i in urls_13ptevallecas:
info_piso = sacar_info(i)
lista_13ptevallecas.append(info_piso)
df_13_ptevallecas = pd.DataFrame(lista_13ptevallecas, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_13_ptevallecas.to_excel("df_13_ptevallecas.xlsx",index=True)
# #### <font color= "1F284B" >14. Moratalaz.</font>
urls_14moratalaz = []
for i in range(1,4):
try:
urls_14moratalaz.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/moratalaz/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_14moratalaz = []
for i in urls_14moratalaz:
info_piso = sacar_info(i)
lista_14moratalaz.append(info_piso)
df_14_moratalaz = pd.DataFrame(lista_14moratalaz, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_14_moratalaz.to_excel("df_14_moratalaz.xlsx",index=True)
# #### <font color= "1F284B" >15. Ciudad Lineal.</font>
urls_15clineal = []
for i in range(1,37):
try:
urls_15clineal.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/ciudad%20lineal/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_15clineal = []
for i in urls_15clineal:
info_piso = sacar_info(i)
lista_15clineal.append(info_piso)
df_15_clineal = pd.DataFrame(lista_15clineal, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_15_clineal.to_excel("df_15_clineal.xlsx",index=True)
# #### <font color= "1F284B" >16. Hortaleza. (No hay pisos de este barrio en gilmar)</font>
# #### <font color= "1F284B" >17. Villaverde.</font>
urls_17villaverde = []
for i in range(1,7):
try:
urls_17villaverde.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/villaverde/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_17villaverde = []
for i in urls_17villaverde:
info_piso = sacar_info(i)
lista_17villaverde.append(info_piso)
df_17_villaverde = pd.DataFrame(lista_17villaverde, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_17_villaverde.to_excel("df_17_villaverde.xlsx",index=True)
# #### <font color= "1F284B" >18. Villa de vallecas.</font>
urls_18villavallecas = []
for i in range(1,3):
try:
urls_18villavallecas.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/villa%20de%20vallecas/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_18villavallecas = []
for i in urls_18villavallecas:
info_piso = sacar_info(i)
lista_18villavallecas.append(info_piso)
df_18_villavallecas = pd.DataFrame(lista_18villavallecas, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_18_villavallecas.to_excel("df_18_villavallecas.xlsx",index=True)
# #### <font color= "1F284B" >19. Vicálvaro.</font>
urls_19vicalvaro = []
for i in range(1,2):
try:
urls_19vicalvaro.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/vicalvaro/gilmar(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_19vicalvaro = []
for i in urls_19vicalvaro:
info_piso = sacar_info(i)
lista_19vicalvaro.append(info_piso)
df_19_vicalvaro = pd.DataFrame(lista_19vicalvaro, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_19_vicalvaro.to_excel("df_19_vicalvaro.xlsx",index=True)
# #### <font color= "1F284B" >20. San Blás.</font>
urls_20sanblas = []
for i in range(1,11):
try:
urls_20sanblas.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/San%20blas/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_20sanblas = []
for i in urls_20sanblas:
info_piso = sacar_info(i)
lista_20sanblas.append(info_piso)
df_20_sanblas = pd.DataFrame(lista_20sanblas, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_20_sanblas.to_excel("df_20_sanblas.xlsx",index=True)
# #### <font color= "1F284B" >21. Barajas.</font>
urls_21barajas = []
for i in range(1,3):
try:
urls_21barajas.append("C:/Users/nuria/Bootcamp%20Data%20Science/Proyecto%20Final/Html%20scraper/barajas/gilmar%20(" + str(i) + ").html")
except:
print("No hay más htmls")
lista_21barajas = []
for i in urls_21barajas:
info_piso = sacar_info(i)
lista_21barajas.append(info_piso)
df_21_barajas = pd.DataFrame(lista_21barajas, columns = ["Barrio", "Precio", "Precio_m2", "m2", "habitaciones", "baños", "terraza", "trastero", "garaje"])
df_21_barajas.to_excel("df_21_barajas.xlsx",index=True)
| PARTE 1.2. Web Scraping gilmar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Diversas fuentes de datos
#
# 
# ## Datos relacionales
#
# Los tipos de datos más simples que hemos visto consisten de una sola tabla con algunas variables (columnas) y algunos registros (filas). Este tipo de datos es fácil de analizar, y muchas veces podemos reducir nuestros datos a una única tabla antes de empezar a correr algoritmos de aprendizaje de máquinas sobre ellos.
#
# Sin embargo, los datos en el mundo real no necesariamente son tan "bonitos". La mayoría de datos reales que nos encontramos son complejos y desordenados, y no son fáciles de organizar en una sola tabla sin antes hacer un buen trabajo en su procesamiento.
#
# Adicionalmente, muchas veces podemos reducir el costo de guardar los datos en memoria distribuyendo los datos en varias tablas con relaciones definidas, en lugar de una sola tabla que concentre toda la información.
#
# El día de hoy vamos a revisar un poco como combinar datos de diferentes fuentes, y cómo podemos generar características bastante útiles.
#
# Como ejemplo tomaremos datos de las 10 compañías top en el índice [Fortune Global 500](https://en.wikipedia.org/wiki/Fortune_Global_500). Para trabajarlas, usaremos la función `read_html` de pandas, la cual nos permitirá ingerir los datos directamente desde la página.
# Importar librerías
import pandas as pd
# Cargar datos en memoria usando pd.read_html
data = pd.read_html("https://en.wikipedia.org/wiki/Fortune_Global_500")
type(data)
for i in range(len(data)):
print(type(data[i]))
fortune500 = data[0]
fortune500
# ### ¿Qué es lo que hay detrás de la función `pd.read_html`?
#
# Los pasos se pueden detallar tanto como se quiera, pero escencialmente son #:
#
# 1. Hacer un **GET request** a la página web (usando la librería [requests](https://docs.python-requests.org/en/master/)):
# Importar librería requests
import requests
help(requests.get)
# Hacer un get request a la página
response = requests.get("https://en.wikipedia.org/wiki/Fortune_Global_500")
help(response.json)
# ¿Qué obtenemos con este request?
# Atributo text
response.text
# Inspeccionar página ...
#
# Entonces, obtenemos todos los datos de la página. Lo "único" que nos hace falta es:
#
# 2. Llevar estos datos a un formato adecuado usando un **HTML parser** (usamos [Beautiful Soup](https://www.crummy.com/software/BeautifulSoup/bs4/doc/)):
# Importar bs4.BeautifulSoup
from bs4 import BeautifulSoup
help(BeautifulSoup)
# Instanciar un objeto tipo BeautifulSoup con los contenidos del request
soup = BeautifulSoup(response.text, "html")
# [Entrada de stackoverflow donde se discuten los diferentes parsers](https://stackoverflow.com/questions/25714417/beautiful-soup-and-table-scraping-lxml-vs-html-parser)
# ¿Qué contiene nuestra "sopa"?
soup
# Podemos hacer búsqueda de diferentes objetos:
# Título
soup.find("title")
# Tablas
table = soup.find_all("table")[0].text
table.split("\n1")[1].split("\n2")
# Observamos que la tabla la podríamos "parsear" usando la clase `str`. Acá podríamos hacer uso del parser que nos provee pandas:
fortune500 = pd.read_html(str(soup.find("table")))[0]
fortune500
# De este modo podemos obtener información relevante de páginas web públicas.
#
# Hay mucho más en cuanto al tema de scraping de páginas web.
#
# - Si para obtener información de una página debes navegar en ella, hacer clicks en botones o cosas por el estilo, hay otra librería que nos puede ayudar a automatizar estas tareas. Su nombre es [Selenium](https://selenium-python.readthedocs.io/).
#
# - Por otra parte, cuando una página web no quiere que sus contenidos sean obtenidos de manera masiva y repetitiva, normalmente incluyen sistemas "antibots":
#
# 
#
# - Otra práctica común, es que limiten los requests cuando identifican que se hacen con la misma dirección ip.
# Volviendo a nuestros datos:
# Datos de fortune 500
fortune500
# Una pregunta que quisieramos resolver es, ¿Cuál es el ingreso promedio por empleado?
#
# Podemos buscar estos datos en Wikipedia también. Yo ya los "scrapeé" manualmente por ustedes para que los usemos en la clase:
other_data = [
{"name": "Walmart",
"employees": 2200000,
"year founded": 1962
},
{"name": "State Grid Corporation of China",
"employees": 1566000,
"year founded": 2002
},
{"name": "China National Petroleum Corporation",
"employees": 460724,
"year founded": 1988
},
{"name": "Berkshire Hathaway Inc.",
"employees": 360000,
"year founded": 1839
},
{"name": "BP plc",
"employees": 70100,
"year founded": 1909
},
{"name": "China Petrochemical Corporation",
"employees": 582648,
"year founded": 1998
},
{"name": "<NAME>",
"employees": 86000,
"year founded": 1907
},
{"name": "Toyota Motor Corporation",
"employees": 364445,
"year founded": 1937
},
{"name": "<NAME>",
"employees": 66800,
"year founded": 1933
},
{"name": "Apple Inc.",
"employees": 147000,
"year founded": 1976
},
{"name": "<NAME>",
"employees": 307342,
"year founded": 1937
},
{"name": "Amazon.com, Inc.",
"employees":1298000,
"year founded": 1994
}
]
employees_info = pd.DataFrame(other_data)
employees_info
fortune500
# Pensaríamos que podría ser tan fácil como hacer un merge entre ambas tablas sobre el nombre de las columnas. Sin embargo, es fácil notar que no todos los nombres coinciden.
# Diccionario de mapeo entre nombres
name_map = {"Walmart": "Walmart",
"China Petrochemical Corporation": "Sinopec Group",
"State Grid Corporation of China": "State Grid",
"China National Petroleum Corporation": "China National Petroleum",
"Royal Dutch Shell": "Royal Dutch Shell",
"<NAME>": "<NAME>",
"Volkswagen AG": "Volkswagen",
"BP plc": "BP",
"Amazon.com, Inc.": "Amazon.com",
"Toyota Motor Corporation": "Toyota Motor"
}
# Hacer un map de los nombres en el dataframe inicial
employees_info["company"] = employees_info["name"].map(name_map)
employees_info
# Hacer el merge
fortune500_with_employees = fortune500.merge(right=employees_info[["company", "employees"]],
left_on="Company",
right_on="company",
how="left")
fortune500_with_employees
fortune500_with_employees.dtypes
fortune500_with_employees["Revenue in USD"] = \
fortune500_with_employees["Revenue in USD"].apply(
lambda s: int(s[1:4]) * 10**9
)
fortune500_with_employees
fortune500_with_employees.dtypes
# Responder la pregunta
fortune500_with_employees["Revenue per employee"] = \
fortune500_with_employees["Revenue in USD"] / fortune500_with_employees["employees"]
fortune500_with_employees.sort_values(by="Revenue per employee", ascending=False)
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Semana4/Clase8_DiversasFuentesDatos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# .. _nb_radar:
# -
# ## Radar Plot
#
#
# Let us produce some data first:
# + code="visualization/usage_radar.py" section="load_data"
import numpy as np
np.random.seed(3)
ideal_point = np.array([0.15, 0.1, 0.2, 0.1, 0.1])
nadir_point = np.array([0.85, 0.9, 0.95, 0.9, 0.85])
F = np.random.random((1, 5)) * (nadir_point - ideal_point) + ideal_point
print(F)
# -
# If the values should not be normalized, then we can plot the ideal and nadir point in addition.
# This keeps the absolute values of each objective. The outer shape represents the nadir point, the inner area the ideal point. All points will lie in the area spanned by those two points.
# + code="visualization/usage_radar.py" section="radar"
from pymoo.visualization.radar import Radar
plot = Radar(bounds=[ideal_point, nadir_point], normalize_each_objective=False)
plot.add(F)
plot.show()
# -
# But if the scale of the objective is too different, then normalization is recommended. Then, the ideal point is just the point in the middle and the nadir point is now symmetric.
# + code="visualization/usage_radar.py" section="radar_norm"
plot = Radar(bounds=[ideal_point, nadir_point])
plot.add(F)
plot.show()
# + code="visualization/usage_radar.py" section="radar_custom"
F = np.random.random((6, 5)) * (nadir_point - ideal_point) + ideal_point
plot = Radar(bounds=[ideal_point, nadir_point],
axis_style={"color": 'blue'},
point_style={"color": 'red', 's': 30})
plot.add(F[:3], color="red", alpha=0.8)
plot.add(F[3:], color="green", alpha=0.8)
plot.show()
# -
# ### API
# + raw_mimetype="text/restructuredtext" active=""
# .. autoclass:: pymoo.visualization.radar.Radar
# :noindex:
| doc/source/visualization/radar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: learn-env
# language: python
# name: learn-env
# ---
import pandas as pd
#from gensim.models import Word2Vec
#from gensim.models import KeyedVectors
from collections import Counter
import numpy as np
import pickle
df = pd.read_csv('data/complete_dict.csv')
df.head()
# +
##pickle df for use in webapp
#pickle.dump(df, open('webapp/cham_words_df.pkl', 'wb'))
# -
sentences = [word for word in df['word']]
sentences
# +
#model = Word2Vec(sentences=sentences, iter=5, min_count=1, size=300, workers=4)
#, size=100, window=5, min_count=5, workers=8, sg=1)
#model.save("w2c")
# +
#model_1 = KeyedVectors.load_word2vec_format(model)
# +
#model.wv.most_similar(positive=['abuno'], topn=3)
# -
def edits1(word):
"All edits that are one edit away from `word`."
letters = "åabcdefghijklmnñopqrstuvwxyz'"
splits = [(word[:i], word[i:]) for i in range(len(word) + 1)]
deletes = [L + R[1:] for L, R in splits if R]
transposes = [L + R[1] + R[0] + R[2:] for L, R in splits if len(R)>1]
replaces = [L + c + R[1:] for L, R in splits if R for c in letters]
inserts = [L + c + R for L, R in splits for c in letters]
return set(deletes + transposes + replaces + inserts)
len(edits1("a'abang"))
def known(words):
return set(w for w in words if w in WORDS)
WORDS = Counter(sentences)
# +
#pickle WORDS for use in webapp
#pickle.dump(WORDS, open('webapp/cham_words_dict.pkl', 'wb'))
# -
WORDS['ababang']
known(edits1("a'abang"))
def edits2(word):
return (e2 for e1 in edits1(word) for e2 in edits1(e1))
len(set(edits2("a'abang")))
known(edits2("a'abang"))
known(edits2("aabang"))
def P(word, N=sum(WORDS.values())):
return WORDS[word] / N
# +
def correction(word):
return max(candidates(word), key=P)
def candidates(word):
return known([word]) or known(edits1(word)) or known(edits2(word)) or [word]
# -
correction("aabg")
def correction_2(word):
words = candidates(word)
return sorted(words, key=lambda x: P(x), reverse=True)[:5]
correction_2('hayi')
def correction_3(word):
recs = {}
words = candidates(word)
word_recs = sorted(words, key=lambda x: P(x), reverse=True)[:5]
for i in word_recs:
#recs[i] = df.loc[df['word'] == i, ['definition']]
recs[i] = df.loc[df['word'] == i].definition.values[0]
return recs
correction_3("hayi")
def check_spelling():
print( "What word would you like to spell-check?")
word = input()
if [word] == correction_2(word):
return print("No Spelling Suggestions Available")
else:
return correction_3(word)
check_spelling()
# +
#rename, edit, and clean functions
# -
| model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plot value for a partially observable system
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
#parameters
x = 1
T = 1
v_1 = lambda sg: np.power(x,2)/(1+T) + np.power(sg,2)*np.log(1+T) + (1 - np.power(sg,2))*T
v_2 = lambda s: np.power(x,2)/(1+T) + 0*s
s = np.linspace(0,1,11)
plt.plot(s, v_1(s), label='linear terminal')
'''
plt.plot(s, v_2(s), linestyle='dashed', label='quadratic terminal')
plt.ylim(0.2, 1.8)
'''
plt.legend()
plt.show()
| src/plot2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overlay Tutorial
#
# This notebook gives an overview of how the Overlay class should be used efficiently.
#
# The redesigned Overlay class has three main design goals
# * Allow overlay users to find out what is inside an overlay in a consistent manner
# * Provide a simple way for developers of new hardware designs to test new IP
# * Facilitate reuse of IP between Overlays
#
# This tutorial is primarily designed to demonstrate the final two points, walking through the process of interacting with a new IP, developing a driver, and finally building a more complex system from multiple IP blocks. All of the code and block diagrams can be found at [https://github.com/PeterOgden/overlay_tutorial]. For these examples to work copy the contents of the overlays directory into the home directory on the PYNQ-Z1 board.
# ## Developing a Single IP
#
# For this first example we are going to use a simple design with a single IP contained in it. This IP was developed using HLS and adds two 32-bit integers together. The full code for the accelerator is:
#
# ```C++
# void add(int a, int b, int& c) {
# #pragma HLS INTERFACE ap_ctrl_none port=return
# #pragma HLS INTERFACE s_axilite port=a
# #pragma HLS INTERFACE s_axilite port=b
# #pragma HLS INTERFACE s_axilite port=c
#
# c = a + b;
# }
# ```
#
# With a block diagram consisting solely of the HLS IP and required glue logic to connect it to the ZYNQ7 IP
#
# 
#
# To interact with the IP first we need to load the overlay containing the IP.
# +
from pynq import Overlay
overlay = Overlay('/home/xilinx/tutorial_1.bit')
# -
# Creating the overlay will automatically download it. We can now use a question mark to find out what is in the overlay.
# +
# overlay?
# -
# All of the entries are accessible via attributes on the overlay class with the specified driver. Accessing the `scalar_add` attribute of the will create a driver for the IP - as there is no driver currently known for the `Add` IP core `DefaultIP` driver will be used so we can interact with IP core.
# +
add_ip = overlay.scalar_add
# add_ip?
# -
# By providing the HWH file along with overlay we can also expose the register map associated with IP.
add_ip.register_map
# We can interact with the IP using the register map directly
add_ip.register_map.a = 3
add_ip.register_map.b = 4
add_ip.register_map.c
# Alternatively by reading the driver source code generated by HLS we can determine that offsets we need to write the two arguments are at offsets `0x10` and `0x18` and the result can be read back from `0x20`.
add_ip.write(0x10, 4)
add_ip.write(0x18, 5)
add_ip.read(0x20)
# ## Creating a Driver
#
# While the `UnknownIP` driver is useful for determining that the IP is working it is not the most user-friendly API to expose to the eventual end-users of the overlay. Ideally we want to create an IP-specific driver exposing a single `add` function to call the accelerator. Custom drivers are created by inheriting from `UnknownIP` and adding a `bindto` class attribute consisting of the IP types the driver should bind to. The constructor of the class should take a single `description` parameter and pass it through to the super class `__init__`. The description is a dictionary containing the address map and any interrupts and GPIO pins connected to the IP.
# +
from pynq import DefaultIP
class AddDriver(DefaultIP):
def __init__(self, description):
super().__init__(description=description)
bindto = ['xilinx.com:hls:add:1.0']
def add(self, a, b):
self.write(0x10, a)
self.write(0x18, b)
return self.read(0x20)
# -
# Now if we reload the overlay and query the help again we can see that our new driver is bound to the IP.
# +
overlay = Overlay('/home/xilinx/tutorial_1.bit')
# overlay?
# -
# And we can access the same way as before except now our custom driver with an `add` function is created instead of `DefaultIP`
overlay.scalar_add.add(15,20)
# ## Reusing IP
#
# Suppose we or someone else develops a new overlay and wants to reuse the existing IP. As long as they import the python file containing the driver class the drivers will be automatically created. As an example consider the next design which, among other things includes a renamed version of the `scalar_add` IP.
#
# 
#
# Using the question mark on the new overlay shows that the driver is still bound.
# +
overlay = Overlay('/home/xilinx/tutorial_2.bit')
# overlay?
# -
# ## IP Hierarchies
#
# The block diagram above also contains a hierarchy looking like this:
#
# 
#
# Containing a custom IP for multiple a stream of numbers by a constant and a DMA engine for transferring the data. As streams are involved and we need correctly handle `TLAST` for the DMA engine the HLS code is a little more complex with additional pragmas and types but the complete code is still relatively short.
#
# ```C
# typedef ap_axiu<32,1,1,1> stream_type;
#
# void mult_constant(stream_type* in_data, stream_type* out_data, ap_int<32> constant) {
# #pragma HLS INTERFACE s_axilite register port=constant
# #pragma HLS INTERFACE ap_ctrl_none port=return
# #pragma HLS INTERFACE axis port=in_data
# #pragma HLS INTERFACE axis port=out_data
# out_data->data = in_data->data * constant;
# out_data->dest = in_data->dest;
# out_data->id = in_data->id;
# out_data->keep = in_data->keep;
# out_data->last = in_data->last;
# out_data->strb = in_data->strb;
# out_data->user = in_data->user;
#
# }
# ```
#
# Looking at the HLS generated documentation we again discover that to set the constant we need to set the register at offset `0x10` so we can write a simple driver for this purpose
class ConstantMultiplyDriver(DefaultIP):
def __init__(self, description):
super().__init__(description=description)
bindto = ['Xilinx:hls:mult_constant:1.0']
@property
def constant(self):
return self.read(0x10)
@constant.setter
def constant(self, value):
self.write(0x10, value)
# The DMA engine driver is already included inside the PYNQ driver so nothing special is needed for that other than ensuring the module is imported. Reloading the overlay will make sure that our newly written driver is available for use.
# +
import pynq.lib.dma
overlay = Overlay('/home/xilinx/tutorial_2.bit')
dma = overlay.const_multiply.multiply_dma
multiply = overlay.const_multiply.multiply
# -
# The DMA driver transfers numpy arrays allocated using `pynq.allocate`. Lets test the system by multiplying 5 numbers by 3.
# +
from pynq import allocate
import numpy as np
in_buffer = allocate(shape=(5,), dtype=np.uint32)
out_buffer = allocate(shape=(5,), dtype=np.uint32)
for i in range(5):
in_buffer[i] = i
multiply.constant = 3
dma.sendchannel.transfer(in_buffer)
dma.recvchannel.transfer(out_buffer)
dma.sendchannel.wait()
dma.recvchannel.wait()
out_buffer
# -
# While this is one way to use the IP, it still isn't exactly user-friendly. It would be preferable to treat the entire hierarchy as a single entity and write a driver that hides the implementation details. The overlay class allows for drivers to be written against hierarchies as well as IP but the details are slightly different.
#
# Hierarchy drivers are subclasses of `pynq.DefaultHierarchy` and, similar to `DefaultIP` have a constructor that takes a description of hierarchy. To determine whether the driver should bind to a particular hierarchy the class should also contain a static `checkhierarchy` method which takes the description of a hierarchy and returns `True` if the driver should be bound or `False` if not. Similar to `DefaultIP`, any classes that meet the requirements of subclasses `DefaultHierarchy` and have a `checkhierarchy` method will automatically be registered.
#
# For our constant multiply hierarchy this would look something like:
# +
from pynq import DefaultHierarchy
class StreamMultiplyDriver(DefaultHierarchy):
def __init__(self, description):
super().__init__(description)
def stream_multiply(self, stream, constant):
self.multiply.constant = constant
with allocate(shape=(len(stream),), \
dtype=np.uint32) as in_buffer,\
allocate(shape=(len(stream),), \
dtype=np.uint32) as out_buffer:
for i, v, in enumerate(stream):
in_buffer[i] = v
self.multiply_dma.sendchannel.transfer(in_buffer)
self.multiply_dma.recvchannel.transfer(out_buffer)
self.multiply_dma.sendchannel.wait()
self.multiply_dma.recvchannel.wait()
result = out_buffer.copy()
return result
@staticmethod
def checkhierarchy(description):
if 'multiply_dma' in description['ip'] \
and 'multiply' in description['ip']:
return True
return False
# -
# We can now reload the overlay and ensure the higher-level driver is loaded
# +
overlay = Overlay('/home/xilinx/tutorial_2.bit')
# overlay?
# -
# and use it
overlay.const_multiply.stream_multiply([1,2,3,4,5], 5)
# ## Overlay Customisation
#
# While the default overlay is sufficient for many use cases, some overlays will require more customisation to provide a user-friendly API. As an example the default AXI GPIO drivers expose channels 1 and 2 as separate attributes meaning that accessing the LEDs in the base overlay requires the following contortion
base = Overlay('base.bit')
base.leds_gpio.channel1[0].on()
# To mitigate this the overlay developer can provide a custom class for their overlay to expose the subsystems in a more user-friendly way. The base overlay includes custom overlay class which performs the following functions:
# * Make the AXI GPIO devices better named and range/direction restricted
# * Make the IOPs accessible through the `pmoda`, `pmodb` and `ardiuno` names
# * Create a special class to interact with RGB LEDs
#
# The result is that the LEDs can be accessed like:
# +
from pynq.overlays.base import BaseOverlay
base = BaseOverlay('base.bit')
base.leds[0].on()
# -
# Using a well defined class also allows for custom docstrings to be provided also helping end users.
# +
# base?
# -
# ### Creating a custom overlay
#
# Custom overlay classes should inherit from `pynq.UnknownOverlay` taking a the full path of the bitstream file and possible additional keyword arguments. These parameters should be passed to `super().__init__()` at the start of `__init__` to initialise the attributes of the Overlay. This example is designed to go with our tutorial_2 overlay and adds a function to more easily call the multiplication function
class TestOverlay(Overlay):
def __init__(self, bitfile, **kwargs):
super().__init__(bitfile, **kwargs)
def multiply(self, stream, constant):
return self.const_multiply.stream_multiply(stream, constant)
# To test our new overlay class we can construct it as before.
overlay = TestOverlay('/home/xilinx/tutorial_2.bit')
overlay.multiply([2,3,4,5,6], 4)
# ## Included Drivers
#
# The pynq library includes a number of drivers as part of the `pynq.lib` package. These include
#
# * AXI GPIO
# * AXI DMA (simple mode only)
# * AXI VDMA
# * AXI Interrupt Controller (internal use)
# * Pynq-Z1 Audio IP
# * Pynq-Z1 HDMI IP
# * Color convert IP
# * Pixel format conversion
# * HDMI input and output frontends
# * Pynq Microblaze program loading
| docs/source/overlay_design_methodology/overlay_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="9TMFWyRksLsB"
# #**Ctrl4AI**
#
# A helper package for Machine Learning and Deep Learning solutions
# + [markdown] colab_type="text" id="beg1hNYg431k"
# **Developers:** <NAME>
# + [markdown] colab_type="text" id="zqHLRp9LDB2K"
# 
# + [markdown] colab_type="text" id="xX5djzDxs9LZ"
# **Highlights**
#
# - Open Source Package with emphasis on data preprocessing so far.
# - Self intelligent methods that can be employed at the levels of abstraction or customization.
# - The flow of auto-preprocessing is orchestrated compatible to the learning type.
# - Parameter tuning allows users to transform the data precisely to their specifications.
# - Developed computations for inspecting the data to discover its type, distribution, correlation etc. which are handled in the background.
#
#
#
#
#
#
#
#
# + [markdown] colab_type="text" id="JJHV68yyup90"
# # **Install & Import**
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="ytOOXvTR6oHc" outputId="e0a392ba-ae5c-4c37-ca87-fb8f98b5e1a9"
pip install ctrl4ai --upgrade
# + colab={} colab_type="code" id="OhKL1Vda6wID"
from ctrl4ai import preprocessing
from ctrl4ai import automl
# + [markdown] colab_type="text" id="G-qtyPJ1u6Ql"
# # **Usage**
# + [markdown] colab_type="text" id="gMLSu6k6IbN7"
# For documentation, please read [HELP.md](https://github.com/vkreat-tech/ctrl4ai/blob/master/HELP.md)
# + colab={"base_uri": "https://localhost:8080/", "height": 428} colab_type="code" id="-YhCgOUBLk6B" outputId="29c35578-b396-4b94-9a30-6d638456cd4b"
help(automl.preprocess)
# + [markdown] colab_type="text" id="2MhiLi-j33UO"
# # **Inbuilt datasets**
#
#
# + colab={} colab_type="code" id="W3NwMs_vuHlP"
from ctrl4ai import datasets
# + colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="RNrmJ14Opu23" outputId="c96e6158-7597-4293-f4b3-dff0cfb826b4"
dataset1=datasets.trip_fare()
dataset1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="qXI-T-r0Ydwl" outputId="0499e6ab-cb1b-4db5-8177-bd577e1b10ed"
dataset2=datasets.titanic()
dataset2.head()
# + [markdown] colab_type="text" id="z2U7WPIXE0-g"
# # **AutoML**
# + [markdown] colab_type="text" id="B_4JaOQaL86f"
# ## **Preprocessing**
# + colab={"base_uri": "https://localhost:8080/", "height": 530} colab_type="code" id="5F-wxMNeE8U1" outputId="bc3e701e-4658-4d61-f159-dce5176144b8"
dataset1_labels,dataset1_processed=automl.preprocess(dataset1,'supervised',target_variable='fare_amount',target_type='continuous')
dataset1_processed.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="VioTyisZJjkw" outputId="ad7ef4bf-b42a-49df-b391-a04616567f61"
dataset2_labels,dataset2_processed=automl.preprocess(dataset2,'supervised',target_variable='Survived',target_type='categorical',impute_null_method='KNN',tranform_categorical='one_hot_encoding',define_continuous_cols=['Fare'])
dataset2_processed.head()
# + [markdown] colab_type="text" id="E4lO52gxBfL8"
# ## **Collinearity Check**
# + [markdown] colab_type="text" id="eUm-t71fIoMh"
# - Calculates the association between variables in a dataset.
# - Auto detects the type of data and checks Pearson correlation between two continuous variables, CramersV correlation between two categorical variables, Kendalls Tau correlation between a categorical and a continuos variable to find the correlation
# + colab={"base_uri": "https://localhost:8080/", "height": 565} colab_type="code" id="f1dnNSeC63U4" outputId="c3333e4e-ace4-4c12-834e-8f90d53e5b88"
automl.master_correlation(dataset1_processed)
# + [markdown] colab_type="text" id="FDMSoBJ6yn8M"
# # **Preprocessing - Custom Methods**
# + [markdown] colab_type="text" id="DGHcFbXGKpv6"
# ## **Derived Features**
# + [markdown] colab_type="text" id="6c01D-QMDNLA"
# Having the timestamp fields or geographical coordinates as it is doesn't serve any purpose for classification / regression algorithms. So, the goal should be to derive maximum information out of the them.
# + colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="HomgeLFh72Ik" outputId="f0ca747d-4c8c-475c-f343-01e41d439cf5"
dataset1= preprocessing.get_timediff(dataset1,'pickup_datetime','dropoff_datetime')
dataset1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="LWBCBtl679U-" outputId="5fec14d5-ff48-4d2e-93be-f64d701db84f"
dataset1=preprocessing.get_distance(dataset1,'pickup_latitude','pickup_longitude','dropoff_latitude','dropoff_longitude')
dataset1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="GeDKE6eO9Bhf" outputId="a7663dac-01d7-4f18-8b83-83fce7e18ba8"
dataset1=preprocessing.derive_from_datetime(dataset1)
dataset1.head()
# + [markdown] colab_type="text" id="TZB3_HuJ4Wn_"
# ## **Feature Elimination**
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="zW3bs2mMR0gz" outputId="01ca7006-2016-4dc1-f239-c307a4238905"
dataset1=preprocessing.drop_null_fields(dataset1,dropna_threshold=0.5)
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="OI6fHuT94-tu" outputId="a5783abd-2288-4a4b-fcc2-9e339f3cfb74"
dataset1=preprocessing.drop_single_valued_cols(dataset1)
# + [markdown] colab_type="text" id="lATGBVXm85K7"
# ## *Dealing with Categorical Data*
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="37PH7cRO7Z9-" outputId="2841a0ba-ea22-429c-9611-5b2851fee01d"
dataset2=preprocessing.get_ohe_df(dataset2,ignore_cols=['Age'])
dataset2.head()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="YyTg-Hyt-dtd" outputId="901b3426-a446-4c84-b885-6a6bdd6c092d"
col_labels,dataset1=preprocessing.get_label_encoded_df(dataset1)
dataset1.head()
# + [markdown] colab_type="text" id="WEd0Gsvl_TXy"
# ## **Data Cleansing**
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="su986mY2_ahf" outputId="fd0cba97-f788-47e6-d7d0-a36255cbfeab"
dataset2=preprocessing.drop_non_numeric(dataset2)
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="q_9ccZPZARRX" outputId="f04d44a9-5706-425c-d0fa-703ec1739153"
dataset2.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="n4KFFe_8_73B" outputId="9be5d241-c711-44ab-8c0a-4bb25ded221a"
dataset2=preprocessing.impute_nulls(dataset2,method='KNN')
dataset2.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="rxRZss9RAXfX" outputId="769e3e17-f47d-4e08-8f4a-55bc2a3d060c"
dataset1.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="MVUARFWrAaPR" outputId="ea58b0f1-45a8-45c0-e033-453273b8cc04"
dataset1=preprocessing.impute_nulls(dataset1)
dataset1.isnull().sum()
# + [markdown] colab_type="text" id="FqMh-8sBMjZ9"
# ## **Feature Selection**
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="fNAlR9F4Mpsc" outputId="75af0e35-7b98-4b28-8d31-036514f19e4d"
col_corr,correlated_features=preprocessing.get_correlated_features(dataset1,'fare_amount','continuous')
dataset1=dataset1[correlated_features]
dataset1.head()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="qXAqUBhgljvH" outputId="9be50f4d-ef38-4fbf-ec9a-fbfd222a88ed"
col_corr
# + [markdown] colab_type="text" id="WEEDoxQxBxna"
# # **Standardization/Normalization**
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="BeerE0LUB_LO" outputId="125db197-c6d4-494a-ce16-21236572f531"
dataset2=preprocessing.log_transform(dataset1)
dataset2.head()
# + colab={"base_uri": "https://localhost:8080/"} colab_type="code" id="U9XRe2ntElEx" outputId="682de1dd-4a7b-4e4f-e8e7-98c4f68e9efd"
automl.scale_transform(dataset1_processed,method='robust')
| README.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ## SICP 习题 (1.24) 解题总结
;; SICP 习题 1.24 要求使用费马检测检测素数,可以说我的噩梦是从这道题开始的,从这道题开始的好几个星期内完全处于怀疑自己智商的状态中,因为我发现我要面对的不是会不会解题这个问题,而是我能不能理解题目的问题。
;;
;; 后来的努力证明,普通人也是可以理解复杂的数学问题的,所以各位可以继续努力!!
;;
;; 说到费马检测,首先是要去看看最朴素的素数检测方法,就是使用我们之前的smallest-divisor找最小因数的过程,如果一个数的最小因数就是它自己,那么这个数就是素数。
;;
;; 如果你对我上面说到的还是不太明白的话,就需要回去看看数论中有关素数,合数的基本讨论了。没事,我也是特意在网上找了一些资料重新看了有关素数的讨论才开始继续下面的解题过程的。
;;
;; 因为上面提到的朴素的素数检测方法比较耗时,所以大家就开始找方法更快地检测一个数是不是素数。费马检测就是其中的著名方法,SICP书中也比较详细地讲解了费马检测。
;;
;; 首先要明确的就是费马检测这个方法是一个“概率方法”,就是通过这个方法可以发现一个数是素数的可能性大不大,并不能准确地判断一个数是不是素数。
;;
;; 有关“概率方法”这个想法一定要理解清楚,后面好几道题都和这个概念有关。
;;
;; 然后就是理解费马检测的具体操作,如果要判断一个数n是不是素数,最基本的就是找一个比n小,比1大的数a,如果((a的n次方)对n求模)= a 的话,这个数n是素数的可能性就很大。
;;
;; 现在的问题是如何求((a的n次方)对n求模)),其实我最早想到的就是用我们前面的题目中做的快速求n次方的过程fast-expt,再加上remainder过程就可以了。没想到后来这个方法还在习题1.25中作为反例出现!伤自尊呀!
;;
;; 后来就去看别人实现的((a的n次方)对n求模))的过程,出乎我意料地长成这个样子:
;; +
(define (expmod base exp m)
(cond ((= exp 0) 1)
((even? exp)
(remainder (square (expmod base (/ exp 2) m))
m))
(else
(remainder (* base (expmod base (- exp 1) m))
m))))
(define (square x)
(* x x))
;; -
;; 看了几遍以后也算看明白了,其实和fast-expt同一个原理,通过这个我自己叫做“折半”的方法可以再对数步数完成计算。
;;
;; 简单测试一下:
;;
;; 10 的 4 次方 对 7 求模
(expmod 10 4 7)
;; 接着看费马测试的过程就很简单了,实现如下:
(define (fermat-test n)
(define (try-it a)
(= (expmod a n n ) a))
(try-it (+ 1 (random (- n 1)))))
;; 其实就是通过random过程随机找一个比n小比1大的数,然后通过expmod过程进行检测。
;;
;; 先跑着测试一下:
(fermat-test 101)
;; 不过,以上方法只是对数n做了一次费马检测,如果数n通过检测的话只能说n这个数是素数的可能性大。如何让这种方法更厉害一点呢?简单的方法就是多做几次费马检测,如果都通过的话那n这个数是素数的可能性就更大了。
;;
;; 过程如下:
(define (fast-prime? n times)
(cond ((= times 0) #t)
((fermat-test n) (fast-prime? n (- times 1)))
(else #f)))
(fast-prime? 101 3)
;; 上面的过程可以指定一个数n进行费马检测,同时指定检测次数,检测次数越大,出来的结果就越准确。
;;
;; 事实上,悄悄告诉你,不管你检测多少次,有些数就是可以骗过费马检测的,那些数不是素数,不过它们可以百分百通过费马检测,后面的习题还会讨论这一点。
;;
;; 最后,结合之前的习题,可以通过以下过程对一个数n进行素数检测,同时报告检测所需要的时间,可以发现,下面的过程中调用fast-prime?时指定检测次数为100次。
(define (start-prime-test n start-time)
(if (fast-prime? n 100)
(begin
(report-prime n (- (real-time-clock) start-time))
#t)
#f))
(import "time")
(define (real-time-clock) (python-eval "time.time()"))
(define (report-prime number elapsed-time)
(display number)
(display " *** ")
(display elapsed-time)
(newline))
(define (timed-prime-test n)
(start-prime-test n (real-time-clock)))
(timed-prime-test 101)
(start-prime-test 101 (real-time-clock))
;; +
(define (find-prime start end number)
(if (even? start)
(find-prime (+ start 1) end number)
(find-prime-iter start end 0 number)))
(define (find-prime-iter start end cur-number max-number)
(if (and (< start end) (< cur-number max-number))
(if (timed-prime-test start)
(find-prime-iter (+ start 2) end (+ 1 cur-number) max-number)
(find-prime-iter (+ start 2) end cur-number max-number))
cur-number))
;; -
;; 通过以上方法就可以回答题中的有关计算时间的问题了。
;;
;; 我测试了比100,10000,100000000,10000000000000000大的三个素数,测试100, 10000的时候不明显,测试到100000000,10000000000000000的时候就比较明显了,1000000000比10000多用了一倍时间,而10000000000000000比1000000000又多用了一倍时间。
;;
;; 部分测试代码运行如下:
(find-prime 1000 1200 3)
(find-prime 10000 12000 3)
(find-prime 10000000000000000 10000000000200000 3)
;; 一切符合理论上的对数步数的预期。
;;
;; 同时惊叹一下,能找到对数步数的算法真的很牛X,计算10000000000000000左右的数值只比计算1000000000左右的数值花多了一倍的时间!
| cn/.ipynb_checkpoints/sicp-1-24-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural network forward propagation
import numpy as np
from math import exp
def tangent(x):
return 2 / (1 + np.exp(-2 * x)) -1
def sigmoid(x):
return 1/(1 + np.exp(-x))
def forward_propagation(x, Wji, Wj0, Wkj, Wk0):
""" NN forward propagation step
x: np.array
Sample
Wji: np.array
First layer weights vector
Wj0: np.array
First layer bias
Wkj: np.array
Output layer weights vector
Wk0: np.array
Output layer bias
"""
print('x:', x)
y = tangent(np.matmul(Wji, x) + Wj0)
print('y:', y)
z = sigmoid(np.matmul(Wkj, y) + Wk0)
print('z:', z)
return z
# ## Examples
Wji = np.array([[-0.7057, 1.9061, 2.6605, -1.1359],
[0.4900, 1.9324, -0.4269, -5.1570],
[0.9438, -5.4160, -0.3431, -0.2931]
])
Wj0 = np.array([4.8432, 0.3973, 2.1761])
Wkj = np.array([[-1.1444, 0.3115, -9.9812],
[0.0106, 11.5477, 2.6479]
])
Wk0 = np.array([2.5230, 2.6463])
X = np.array([[1, 0, 1, 0],
[0, 1, 0, 1],
[1, 1, 0, 0],
])
outputs = []
for x in X:
z = forward_propagation(x, Wji, Wj0, Wkj, Wk0)
outputs.append(z)
#remove the e-n formatting for decimal numbers
np.set_printoptions(suppress=True)
print('outputs:', outputs)
| NN-forward_propagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <font size="+5">#02. Machine Learning & Linear Regression</font>
# <ul>
# <li>Doubts? → Ask me in <img src="https://emoji.gg/assets/emoji/3970-discord.png" style="height: 1em; vertical-align: middle;"> <a href="https://discord.gg/cmB3KGsqMy">Discord</a></li>
# <li>Tutorials → <img src="https://openmoji.org/php/download_asset.php?type=emoji&emoji_hexcode=E044&emoji_variant=color" style="height: 1em; vertical-align: middle;"> <a href="https://www.youtube.com/channel/UCovCte2I3loteQE_kRsfQcw">YouTube</a></li>
# <li>Book Private Lessons → <span style="color: orange">@</span> <a href="https://sotastica.com/reservar">sotastica</a></li>
# </ul>
# # Load the Data
# > By running the following commands:
#
# ```python
# import seaborn as sns
# sns.get_dataset_names()
# ```
#
# > You'll observe a list of posible `tables` that we could work with.
# > You may use whichever you'd like.
# > Just substitue `?` by the name of the dataset you want.
#
# ```python
# sns.load_dataset(name=?)
# ```
# # `LinearRegression()` Model in Python
# ## Build the Model
# > 1. **Necesity**: Build Model
# > 2. **Google**: How do you search for the solution?
# > 3. **Solution**: Find the `function()` that makes it happen
# ### Code Thinking
# > - By the time you get to the actual `function()` to compute the model
# > - You'll notice they're asking you for two parameters:
# > 1. `X`: **explanatory variable**
# > 2. `y`: **target varaible**
# > It's recommended that you make a `Data Visualization`
# >
# > - To analyze if the two variables are related?
# > - i.e., do you think `X` variable is good to predict `y`?
# > - If so, the Linear Regression would be great!
# ### Scatterplot with Variables from DataFrame
# > You may observe in the plot that the points could be related through a line.
# >
# > Otherwise, you may select another pair of variables.
# ### Finally `fit()` the Model
# > - Pass the `objects` that **contains the sequence of numbers** of the two variables
# > - To the `parameters` of the function that computes the Linear Regression Model
# # Model Interpretation
# > 1. Especify the **mathematical equation** of the model.
# > 2. **Interpret the coefficients** from the equation.
# # Calculate Predictions
# > - `model.predict()`
# # Visualize the Model
# > 1. **Real Data**.
# > 2. **Model: predictions** calculated with the mathematical equation.
# # Real vs. Predicted Data
# > How good is our model?
# >
# > 1. Create a new `DataFrame` column to assign the predictions.
# >
# > - `df['pred'] = predicciones`
# > - `df.sample(10)` to check if predictions are equal to reality:
# > 2. How to measure the **model's error**?
# > - How good is our model to predict reality?
# > - `model.score()`
# # Achieved Goals
# _Double click on **this cell** and place an `X` inside the square brackets (i.e., [X]) if you think you understand the goal:_
#
# - [ ] Understand **how the Machine Learns/Optimizes the model**
# - [ ] No more than to find the best numbers in a mathematical equation.
# - [ ] The `function()` as the indivisible part of programming.
# - As the **atom**: the key element in the Universe.
# - [ ] Understand **what's going on inside the computer** as you execute a function.
# - Is the code downloaded online, or could we use it withoug internet conection.
# - [ ] Use **programming as a tool**.
# - It's a **means to an end. NOT an end itself** (as hard as it may be to get started with).
# - [ ] Understand that statistics is not a perfect science.
# - It tries to **approximate to the reality** the best way it can.
# - [ ] **Measuring the error** is nothing more than measuring the difference between reality and predictions.
# - [ ] We may use more than one mathematical formula to calculate the same variable.
# - Different models/equations/algorightms to calculate the same variable.
# - Some models might be better than other ones.
| #03. Machine Learning & Linear Regression/03practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">TensorFlow Neural Network Lab</h1>
# <img src="image/notmnist.png">
# In this lab, you'll use all the tools you learned from *Introduction to TensorFlow* to label images of English letters! The data you are using, <a href="http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html">notMNIST</a>, consists of images of a letter from A to J in differents font.
#
# The above images are a few examples of the data you'll be training on. After training the network, you will compare your prediction model against test data. Your goal, by the end of this lab, is to make predictions against that test set with at least an 80% accuracy. Let's jump in!
# To start this lab, you first need to import all the necessary modules. Run the code below. If it runs successfully, it will print "`All modules imported`".
# +
import hashlib
import os
import pickle
from urllib.request import urlretrieve
import numpy as np
from PIL import Image
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelBinarizer
from sklearn.utils import resample
from tqdm import tqdm
from zipfile import ZipFile
print('All modules imported.')
# -
# The notMNIST dataset is too large for many computers to handle. It contains 500,000 images for just training. You'll be using a subset of this data, 15,000 images for each label (A-J).
# +
def download(url, file):
"""
Download file from <url>
:param url: URL to file
:param file: Local file path
"""
if not os.path.isfile(file):
print('Downloading ' + file + '...')
urlretrieve(url, file)
print('Download Finished')
# Download the training and test dataset.
download('https://s3.amazonaws.com/udacity-sdc/notMNIST_train.zip', 'notMNIST_train.zip')
download('https://s3.amazonaws.com/udacity-sdc/notMNIST_test.zip', 'notMNIST_test.zip')
# Make sure the files aren't corrupted
assert hashlib.md5(open('notMNIST_train.zip', 'rb').read()).hexdigest() == 'c8673b3f28f489e9cdf3a3d74e2ac8fa',\
'notMNIST_train.zip file is corrupted. Remove the file and try again.'
assert hashlib.md5(open('notMNIST_test.zip', 'rb').read()).hexdigest() == '5d3c7e653e63471c88df796156a9dfa9',\
'notMNIST_test.zip file is corrupted. Remove the file and try again.'
# Wait until you see that all files have been downloaded.
print('All files downloaded.')
# +
def uncompress_features_labels(file):
"""
Uncompress features and labels from a zip file
:param file: The zip file to extract the data from
"""
features = []
labels = []
with ZipFile(file) as zipf:
# Progress Bar
filenames_pbar = tqdm(zipf.namelist(), unit='files')
# Get features and labels from all files
for filename in filenames_pbar:
# Check if the file is a directory
if not filename.endswith('/'):
with zipf.open(filename) as image_file:
image = Image.open(image_file)
image.load()
# Load image data as 1 dimensional array
# We're using float32 to save on memory space
feature = np.array(image, dtype=np.float32).flatten()
# Get the the letter from the filename. This is the letter of the image.
label = os.path.split(filename)[1][0]
features.append(feature)
labels.append(label)
return np.array(features), np.array(labels)
# Get the features and labels from the zip files
train_features, train_labels = uncompress_features_labels('notMNIST_train.zip')
test_features, test_labels = uncompress_features_labels('notMNIST_test.zip')
# Limit the amount of data to work with a docker container
docker_size_limit = 150000
train_features, train_labels = resample(train_features, train_labels, n_samples=docker_size_limit)
# Set flags for feature engineering. This will prevent you from skipping an important step.
is_features_normal = False
is_labels_encod = False
# Wait until you see that all features and labels have been uncompressed.
print('All features and labels uncompressed.')
# -
# <img src="image/mean_variance.png" style="height: 75%;width: 75%; position: relative; right: 5%">
#
# ## Problem 1
# The first problem involves normalizing the features for your training and test data.
#
# Implement Min-Max scaling in the `normalize()` function to a range of `a=0.1` and `b=0.9`. After scaling, the values of the pixels in the input data should range from 0.1 to 0.9.
#
# Since the raw notMNIST image data is in [grayscale](https://en.wikipedia.org/wiki/Grayscale), the current values range from a min of 0 to a max of 255.
#
# Min-Max Scaling:
# $
# X'=a+{\frac {\left(X-X_{\min }\right)\left(b-a\right)}{X_{\max }-X_{\min }}}
# $
#
# *If you're having trouble solving problem 1, you can view the solution [here](https://github.com/udacity/CarND-TensorFlow-Lab/blob/master/solutions.ipynb).*
# +
# Problem 1 - Implement Min-Max scaling for grayscale image data
def normalize_grayscale(image_data):
"""
Normalize the image data with Min-Max scaling to a range of [0.1, 0.9]
:param image_data: The image data to be normalized
:return: Normalized image data
"""
a = 0.1
b = 0.9
X_min = 0
X_max = 255
return a + (((image_data - X_min)*(b - a)) / (X_max - X_min))
### DON'T MODIFY ANYTHING BELOW ###
# Test Cases
np.testing.assert_array_almost_equal(
normalize_grayscale(np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 255])),
[0.1, 0.103137254902, 0.106274509804, 0.109411764706, 0.112549019608, 0.11568627451, 0.118823529412, 0.121960784314,
0.125098039216, 0.128235294118, 0.13137254902, 0.9],
decimal=3)
np.testing.assert_array_almost_equal(
normalize_grayscale(np.array([0, 1, 10, 20, 30, 40, 233, 244, 254,255])),
[0.1, 0.103137254902, 0.13137254902, 0.162745098039, 0.194117647059, 0.225490196078, 0.830980392157, 0.865490196078,
0.896862745098, 0.9])
if not is_features_normal:
train_features = normalize_grayscale(train_features)
test_features = normalize_grayscale(test_features)
is_features_normal = True
print('Tests Passed!')
# +
if not is_labels_encod:
# Turn labels into numbers and apply One-Hot Encoding
encoder = LabelBinarizer()
encoder.fit(train_labels)
train_labels = encoder.transform(train_labels)
test_labels = encoder.transform(test_labels)
# Change to float32, so it can be multiplied against the features in TensorFlow, which are float32
train_labels = train_labels.astype(np.float32)
test_labels = test_labels.astype(np.float32)
is_labels_encod = True
print('Labels One-Hot Encoded')
# +
assert is_features_normal, 'You skipped the step to normalize the features'
assert is_labels_encod, 'You skipped the step to One-Hot Encode the labels'
# Get randomized datasets for training and validation
train_features, valid_features, train_labels, valid_labels = train_test_split(
train_features,
train_labels,
test_size=0.05,
random_state=832289)
print('Training features and labels randomized and split.')
# +
# Save the data for easy access
pickle_file = 'notMNIST.pickle'
if not os.path.isfile(pickle_file):
print('Saving data to pickle file...')
try:
with open('notMNIST.pickle', 'wb') as pfile:
pickle.dump(
{
'train_dataset': train_features,
'train_labels': train_labels,
'valid_dataset': valid_features,
'valid_labels': valid_labels,
'test_dataset': test_features,
'test_labels': test_labels,
},
pfile, pickle.HIGHEST_PROTOCOL)
except Exception as e:
print('Unable to save data to', pickle_file, ':', e)
raise
print('Data cached in pickle file.')
# -
# # Checkpoint
# All your progress is now saved to the pickle file. If you need to leave and comeback to this lab, you no longer have to start from the beginning. Just run the code block below and it will load all the data and modules required to proceed.
# +
# %matplotlib inline
# Load the modules
import pickle
import math
import numpy as np
import tensorflow as tf
from tqdm import tqdm
import matplotlib.pyplot as plt
# Reload the data
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
pickle_data = pickle.load(f)
train_features = pickle_data['train_dataset']
train_labels = pickle_data['train_labels']
valid_features = pickle_data['valid_dataset']
valid_labels = pickle_data['valid_labels']
test_features = pickle_data['test_dataset']
test_labels = pickle_data['test_labels']
del pickle_data # Free up memory
print('Data and modules loaded.')
# -
# <img src="image/weight_biases.png" style="height: 60%;width: 60%; position: relative; right: 10%">
#
# ## Problem 2
# For the neural network to train on your data, you need the following <a href="https://www.tensorflow.org/api_docs/python/tf/dtypes/DType">float32</a> tensors:
#
# - `features`
# - Placeholder tensor for feature data (`train_features`/`valid_features`/`test_features`)
# - `labels`
# - Placeholder tensor for label data (`train_labels`/`valid_labels`/`test_labels`)
# - `weights`
# - Variable Tensor with random numbers from a truncated normal distribution.
# - See <a href="https://www.tensorflow.org/api_docs/python/tf/random/truncated_normal">`tf.truncated_normal()` documentation</a> for help.
# - `biases`
# - Variable Tensor with all zeros.
# - See <a href="https://www.tensorflow.org/api_docs/python/tf/zeros"> `tf.zeros()` documentation</a> for help.
#
# *If you're having trouble solving problem 2, review "TensorFlow Linear Function" section of the class. If that doesn't help, the solution for this problem is available [here](https://github.com/udacity/CarND-TensorFlow-Lab/blob/master/solutions.ipynb).*
# +
features_count = 784
labels_count = 10
# TODO: Set the features and labels tensors
features = tf.placeholder(tf.float32)
labels = tf.placeholder(tf.float32)
# TODO: Set the weights and biases tensors
weights = tf.Variable(tf.truncated_normal((features_count, labels_count)))
biases = tf.Variable(tf.zeros(labels_count))
### DON'T MODIFY ANYTHING BELOW ###
#Test Cases
from tensorflow.python.ops.variables import Variable
assert features._op.name.startswith('Placeholder'), 'features must be a placeholder'
assert labels._op.name.startswith('Placeholder'), 'labels must be a placeholder'
assert isinstance(weights, Variable), 'weights must be a TensorFlow variable'
assert isinstance(biases, Variable), 'biases must be a TensorFlow variable'
assert features._shape == None or (\
features._shape.dims[0].value is None and\
features._shape.dims[1].value in [None, 784]), 'The shape of features is incorrect'
assert labels._shape == None or (\
labels._shape.dims[0].value is None and\
labels._shape.dims[1].value in [None, 10]), 'The shape of labels is incorrect'
assert weights._variable._shape == (784, 10), 'The shape of weights is incorrect'
assert biases._variable._shape == (10), 'The shape of biases is incorrect'
assert features._dtype == tf.float32, 'features must be type float32'
assert labels._dtype == tf.float32, 'labels must be type float32'
# Feed dicts for training, validation, and test session
train_feed_dict = {features: train_features, labels: train_labels}
valid_feed_dict = {features: valid_features, labels: valid_labels}
test_feed_dict = {features: test_features, labels: test_labels}
# Linear Function WX + b
logits = tf.matmul(features, weights) + biases
prediction = tf.nn.softmax(logits)
# Cross entropy
cross_entropy = -tf.reduce_sum(labels * tf.log(prediction), axis=1)
# some students have encountered challenges using this function, and have resolved issues
# using https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits
# please see this thread for more detail https://discussions.udacity.com/t/accuracy-0-10-in-the-intro-to-tensorflow-lab/272469/9
# Training loss
loss = tf.reduce_mean(cross_entropy)
# Create an operation that initializes all variables
init = tf.global_variables_initializer()
# Test Cases
with tf.Session() as session:
session.run(init)
session.run(loss, feed_dict=train_feed_dict)
session.run(loss, feed_dict=valid_feed_dict)
session.run(loss, feed_dict=test_feed_dict)
biases_data = session.run(biases)
assert not np.count_nonzero(biases_data), 'biases must be zeros'
print('Tests Passed!')
# +
# Determine if the predictions are correct
is_correct_prediction = tf.equal(tf.argmax(prediction, 1), tf.argmax(labels, 1))
# Calculate the accuracy of the predictions
accuracy = tf.reduce_mean(tf.cast(is_correct_prediction, tf.float32))
print('Accuracy function created.')
# -
# <img src="image/learn_rate_tune.png" style="height: 60%;width: 60%">
#
# ## Problem 3
# Below are 3 parameter configurations for training the neural network. In each configuration, one of the parameters has multiple options. For each configuration, choose the option that gives the best acccuracy.
#
# Parameter configurations:
#
# Configuration 1
# * **Epochs:** 1
# * **Batch Size:**
# * 2000
# * 1000
# * 500
# * 300
# * 50
# * **Learning Rate:** 0.01
#
# Configuration 2
# * **Epochs:** 1
# * **Batch Size:** 100
# * **Learning Rate:**
# * 0.8
# * 0.5
# * 0.1
# * 0.05
# * 0.01
#
# Configuration 3
# * **Epochs:**
# * 1
# * 2
# * 3
# * 4
# * 5
# * **Batch Size:** 100
# * **Learning Rate:** 0.2
#
# The code will print out a Loss and Accuracy graph, so you can see how well the neural network performed.
#
# *If you're having trouble solving problem 3, you can view the solution [here](https://github.com/udacity/CarND-TensorFlow-Lab/blob/master/solutions.ipynb).*
# +
# TODO: Find the best parameters for each configuration
epochs = 5
batch_size = 128
learning_rate = 0.2
### DON'T MODIFY ANYTHING BELOW ###
# Gradient Descent
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
# The accuracy measured against the validation set
validation_accuracy = 0.0
# Measurements use for graphing loss and accuracy
log_batch_step = 50
batches = []
loss_batch = []
train_acc_batch = []
valid_acc_batch = []
with tf.Session() as session:
session.run(init)
batch_count = int(math.ceil(len(train_features)/batch_size))
for epoch_i in range(epochs):
# Progress bar
batches_pbar = tqdm(range(batch_count), desc='Epoch {:>2}/{}'.format(epoch_i+1, epochs), unit='batches')
# The training cycle
for batch_i in batches_pbar:
# Get a batch of training features and labels
batch_start = batch_i*batch_size
batch_features = train_features[batch_start:batch_start + batch_size]
batch_labels = train_labels[batch_start:batch_start + batch_size]
# Run optimizer and get loss
_, l = session.run(
[optimizer, loss],
feed_dict={features: batch_features, labels: batch_labels})
# Log every 50 batches
if not batch_i % log_batch_step:
# Calculate Training and Validation accuracy
training_accuracy = session.run(accuracy, feed_dict=train_feed_dict)
validation_accuracy = session.run(accuracy, feed_dict=valid_feed_dict)
# Log batches
previous_batch = batches[-1] if batches else 0
batches.append(log_batch_step + previous_batch)
loss_batch.append(l)
train_acc_batch.append(training_accuracy)
valid_acc_batch.append(validation_accuracy)
# Check accuracy against Validation data
validation_accuracy = session.run(accuracy, feed_dict=valid_feed_dict)
loss_plot = plt.subplot(211)
loss_plot.set_title('Loss')
loss_plot.plot(batches, loss_batch, 'g')
loss_plot.set_xlim([batches[0], batches[-1]])
acc_plot = plt.subplot(212)
acc_plot.set_title('Accuracy')
acc_plot.plot(batches, train_acc_batch, 'r', label='Training Accuracy')
acc_plot.plot(batches, valid_acc_batch, 'x', label='Validation Accuracy')
acc_plot.set_ylim([0, 1.0])
acc_plot.set_xlim([batches[0], batches[-1]])
acc_plot.legend(loc=4)
plt.tight_layout()
plt.show()
print('Validation accuracy at {}'.format(validation_accuracy))
# -
# ## Test
# Set the epochs, batch_size, and learning_rate with the best learning parameters you discovered in problem 3. You're going to test your model against your hold out dataset/testing data. This will give you a good indicator of how well the model will do in the real world. You should have a test accuracy of at least 80%.
# +
# TODO: Set the epochs, batch_size, and learning_rate with the best parameters from problem 3
epochs = 5
batch_size = 128
learning_rate = 0.2
### DON'T MODIFY ANYTHING BELOW ###
# The accuracy measured against the test set
test_accuracy = 0.0
with tf.Session() as session:
session.run(init)
batch_count = int(math.ceil(len(train_features)/batch_size))
for epoch_i in range(epochs):
# Progress bar
batches_pbar = tqdm(range(batch_count), desc='Epoch {:>2}/{}'.format(epoch_i+1, epochs), unit='batches')
# The training cycle
for batch_i in batches_pbar:
# Get a batch of training features and labels
batch_start = batch_i*batch_size
batch_features = train_features[batch_start:batch_start + batch_size]
batch_labels = train_labels[batch_start:batch_start + batch_size]
# Run optimizer
_ = session.run(optimizer, feed_dict={features: batch_features, labels: batch_labels})
# Check accuracy against Test data
test_accuracy = session.run(accuracy, feed_dict=test_feed_dict)
assert test_accuracy >= 0.80, 'Test accuracy at {}, should be equal to or greater than 0.80'.format(test_accuracy)
print('Nice Job! Test Accuracy is {}'.format(test_accuracy))
# -
# # Multiple layers
# Good job! You built a one layer TensorFlow network! However, you want to build more than one layer. This is deep learning after all! In the next section, you will start to satisfy your need for more layers.
| lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Advertising Prediction Notebook
# In this notebook, the Advertising dataset of Kaggle is analyzed (https://www.kaggle.com/fayomi/advertising). It consists of 10 columns with a total of 1000 rows. The use case consists of predicting whether a website visitor will click on an advertisement or not based on his demographics and internet usage data.
# The notebook's approach is based on the CRISP-DM model, which clearly divides the phases in a data science project.
# <img src="https://statistik-dresden.de/wp-content/uploads/2012/04/CRISP-DM_Process_Diagram1.png" alt="CRISP-DM Modell" width="300" height="300">
#
# ## 1. Business Understanding
#
# From an economic perspective, it is necessary for Facebook to keep customers on their streaming platform for as long as possible. However, the question now arises as to how this can be achieved. The problem here is that Netflix has a good data basis, but this must first be processed and then developed into a digital service. Furthermore, it must be specified which data is available at all, which services can be implemented based on this data, and what added value the customer and Netflix itself derive from this. As a service, this paper deals with a recommendation model that suggests a selection of similar offers to the user for each film or series offered.
# ## 2. Data and Data Understanding
#
# In this notebook, the advertising dataset of Kaggle is analyzed. It consists of 10 columns with a total of 1000 rows. The use case consists of predicting whether a website visitor will click on an advertisement or not based on his demographics and internet usage data. The Clicked on Ad target value is perfectly balanced between the two categories (0.1), as the mean value is exactly 0.5. This means that there are the same number of values for both categories (500 each). In addition, we can see that the Ad Topic Line and City features have very many unique values (1000 and 969 "unique" values, respectively), which means that. It can be seen that there are significant differences between the user profiles. Users who click on an ad (Clicked on Ad=1) spend less time on the website on average, are older (approx. 40), have a lower income and use the Internet significantly less. From this information, a rough user profile can already be derived, which could also be relevant for a company's marketing and sales to optimize their measures based on the user profiles.
# ## 2.1. Import of Relevant Modules
# +
# Import required libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# Set style for the visualization libra
# %matplotlib inline
sns.set_style('whitegrid')
plt.style.use("fivethirtyeight")
# -
# ## 2. 2. Read Data.csv
# Load the CSV-file in a DataFrame
data = pd.read_csv('https://storage.googleapis.com/ml-service-repository-datastorage/Predicting_clicks_on_online_advertising_by_Facebook_data.csv')
data.head()
# # 3. Data Analysis
#
# The purpose of this chapter is to review, analyze, and prepare the data.
# Info of the DataFrame
data.info()
# Compute different metrics for each column
data.describe(include="all")
# ### Missing values
# Number of missing values in each column
data.isnull().sum()
# ### Duplicates
# Displays duplicate records
data.duplicated().sum()
# ## 3.1 Explorative data analysis
# In this chapter, initial analyses and visualizations will be made.
# Create Pairplots
sns.pairplot(data, hue='Clicked on Ad')
# For each label, count the occurence
data['Clicked on Ad'].value_counts()
# It can be seen that the data set is perfectly balanced, i.e. there are exactly 500 records for both classes.
# User profile analysis
data.groupby('Clicked on Ad')['Daily Time Spent on Site', 'Age', 'Area Income',
'Daily Internet Usage'].mean()
# Scatterplot: Daily Time Spent on Site vs. Age in context of Clicked on Ad
sns.scatterplot(x="Daily Time Spent on Site", y="Age", data=data, hue="Clicked on Ad")
# Scatterplot: Daily Time Spent on Site vs. Area Income in context of Clicked on Ad
sns.scatterplot(x="Daily Time Spent on Site", y="Area Income", data=data, hue="Clicked on Ad")
# Scatterplot: Daily Time Spent on Site vs. Daily Internet Usage in context of Clicked on Ad
sns.scatterplot(x="Daily Time Spent on Site", y="Daily Internet Usage", data=data, hue="Clicked on Ad")
# Scatterplot: Age vs. Daily Internet Usage in context of Clicked on Ad
sns.scatterplot(x="Age", y="Daily Internet Usage", data=data, hue="Clicked on Ad")
# ## 3.2 Distribution plots for all features with numerical values
#
# Distribution plots are created to identify outliers in the data and better understand the data
# Distribution plot of Age
sns.distplot(data["Age"])
plt.title("Age Distribution")
# Cut the left 1% and right 99% quantile to avoid outliers
q_small = data["Age"].quantile(0.01)
q_big = data["Age"].quantile(0.99)
data = data[(data["Age"]>q_small) & (data["Age"]<q_big)]
# Distribution plot of Daily Time Spent on Site
sns.distplot(data["Daily Time Spent on Site"])
plt.title("Daily Time Spent on Site Distribution")
# Cut the left 1% and right 99% quantile to avoid outliers
q_small = data["Daily Time Spent on Site"].quantile(0.01)
q_big = data["Daily Time Spent on Site"].quantile(0.99)
data = data[(data["Daily Time Spent on Site"]>q_small) & (data["Daily Time Spent on Site"]<q_big)]
# Distribution plot of Area Income
sns.distplot(data["Area Income"])
plt.title("Area Income Distribution")
# Cut the left 1% and right 99% quantile to avoid outliers
q_small = data["Area Income"].quantile(0.01)
q_big = data["Area Income"].quantile(0.99)
data = data[(data["Area Income"]>q_small) & (data["Area Income"]<q_big)]
# Distribution plot of Area Income with method Boxcox and lambda = 1.5
# The other functions have also been tried out, but the boxcox method fits the best
from scipy.stats import boxcox
#function = lambda x: 1/x or np.log(x) or np.sqrt(x)
#function = lambda x: np.log(x)
#log_data = data["Area Income"].apply(function)
data['Area Income'] = boxcox(data['Area Income'], lmbda=1.5)
sns.distplot(data['Area Income'])
plt.title("Area Income: Boxcox")
# Distribution plot of Daily Internet Usage
sns.distplot(data["Daily Internet Usage"])
plt.title("Daily Internet Usage Distribution")
# Cut the left 1% and right 99% quantile to avoid outliers
q_small = data["Daily Internet Usage"].quantile(0.01)
q_big = data["Daily Internet Usage"].quantile(0.99)
data = data[(data["Daily Internet Usage"]>q_small) & (data["Daily Internet Usage"]<q_big)]
# Distribution plot of Clicked on Ad
sns.distplot(data["Clicked on Ad"])
plt.title("Clicked on Ad Distribution")
# <a id="cell1.3"></a>
# ## 4. Correlations
# Now the correlations of all numerical features are calculated and reflected in a correlation matrix.
# Create heatmap
sns.heatmap(data.corr(), annot=True)
# t can be seen that Daily Time Spent on Site and Daily Internet Usage correlate. There is also a strong negative correlation between Daily Internet Usage / Daily Time Spent on Site and Clicked on Ad.
# However, significant correlations that lead to the removal of a feature do not exist (assumption: if correlation is greater than 0.9).
# <a id="cell2"></a>
# # 5. Data preparation
# In this section, the dataset will be prepared for machine learning.
#
# <a id="cell2.1"></a>
# ## 5.1 Feature Engineering
# In this section, feature engineering is performed. Here, important information is extracted from the raw data.
#
# <a id="cell2.1.1"></a>
# ### 5.1.1 Timestamp
# There is a Timestamp feature in the dataset. This could be important for prediction, as there can be a correlation between the user click and the time.
# Extract datetime variables using timestamp column
data['Timestamp'] = pd.to_datetime(data['Timestamp'])
# Converting timestamp column into datatime object in order to extract new features
data['Month'] = data['Timestamp'].dt.month
# Creates a new column called Month
data['Day'] = data['Timestamp'].dt.day
# Creates a new column called Day
data['Hour'] = data['Timestamp'].dt.hour
# Creates a new column called Hour
data["Weekday"] = data['Timestamp'].dt.dayofweek
# Creates a new column called Weekday with sunday as 6 and monday as 0
data = data.drop(['Timestamp'], axis=1) # deleting timestamp
# In this section, feature engineering is performed. Here, important information is extracted from the raw data.
#
# Look at first 5 rows of the newly created DataFrame
data.head()
# Create heatmap
sns.set(rc={'figure.figsize':(14,14)})
sns.heatmap(data.corr(), annot=True)
# Barplots for the Weekday feature in context of the Clicked on Ad
ax = sns.barplot(x="Weekday", y="Clicked on Ad", data=data, estimator=sum)
# Creating pairplot to check effect of datetime variables on target variable (variables which were created)
pp = sns.pairplot(data, hue= 'Clicked on Ad', vars = ['Month', 'Day', 'Hour', 'Weekday'], palette= 'husl')
# There is probably no sifnificant effect through time.
# Info of the dataframe
data.info()
# Reset the index
data.reset_index(drop=True, inplace=True)
# <a id="cell2.1.2"></a>
# ### 5.1.2 Age
# We can make intervals for age.
# Creating Bins on Age column
data['Age_bins'] = pd.cut(data['Age'], bins=[0, 18, 30, 45, 70], labels=['Young', 'Adult','Mid', 'Elder'])
# Count for each category of Age_bins
data['Age_bins'].value_counts()
# Dummy encoding on Age_bins column
data = pd.concat([data, pd.get_dummies(data['Age_bins'], prefix='Age', drop_first=True)], axis=1)
# <a id="cell2.2"></a>
# ## 5.2 Final dataset creation
# Remove redundant and no predictive power features
data.drop(['Country', 'Ad Topic Line', 'City', 'Day', 'Month', 'Weekday',
'Hour', 'Age', 'Age_bins'], axis = 1, inplace = True)
# <a id="cell2.3"></a>
# ## 5.3 Record splitting and standardization
# Split the data set into features (X) and target variable (y).
# First 5 rows of the dataset
data.head()
# Prepare and split data for prediction
from sklearn.model_selection import train_test_split
X = data.drop(['Clicked on Ad'],1)
y = data['Clicked on Ad']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25)
# Standardization of the Features
from sklearn.preprocessing import StandardScaler
stdsc = StandardScaler()
X_train_std = stdsc.fit_transform(X_train)
X_test_std = stdsc.transform(X_test)
# Dimensions of the different splits (rows -> number of samples, columns -> number of features)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# <a id="cell3"></a>
# # 6 Model building
# Now the individual algorithms can be trained and evaluated.
# Import required libraries for the model creation
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix, plot_confusion_matrix
from sklearn.metrics import roc_auc_score, accuracy_score, confusion_matrix, classification_report
# <a id="cell3.1"></a>
# ## 6.1 Logistic regression
# Sample sigmoid curve
x = np.linspace(-6, 6, num=1000)
plt.figure(figsize=(10, 6))
plt.plot(x, (1 / (1 + np.exp(-x))))
plt.title("Sigmoid Function")
# +
from sklearn.linear_model import LogisticRegression
# Create a Logistic Regression Classifier
lr = LogisticRegression(penalty="l2", C= 0.1, random_state=42)
lr.fit(X_train_std, y_train)
# Predict and evaluate using model
lr_training_pred = lr.predict(X_train_std)
lr_test_pred = lr.predict(X_test_std)
lr_training_prediction = accuracy_score(y_train, lr_training_pred)
lr_test_prediction = accuracy_score(y_test, lr_test_pred)
print( "Accuracy of Logistic regression training set:", round(lr_training_prediction,3))
print( "Accuracy of Logistic regression test set:", round(lr_test_prediction,3))
print(classification_report(y_test, lr.predict(X_test_std)))
tn, fp, fn, tp = confusion_matrix(y_test, lr_test_pred).ravel()
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1_score = 2*((precision*recall)/(precision+recall))
print("True Positive: %i" %tp)
print("False Positive: %i" %fp)
print("True Negative: %i" %tn)
print("False Negative: %i" %fn)
print(f"Precision: {precision:.2%}")
print(f"Recall: {recall:.2%}")
print(f"F1-Score: {f1_score:.2%}")
# -
print('Intercept:', lr.intercept_)
weights = pd.Series(lr.coef_[0],
index=X.columns.values)
weights.sort_values(ascending = False)
# In particular, Daily Time Spent on Site, Daily Internet Usage, and Area Income have a greater impact.
# <a id="cell3.2"></a>
# ## 6.2 Decision Tree
# +
from sklearn.tree import DecisionTreeClassifier
# Create a Decision Tree Classifier
estimator = DecisionTreeClassifier(max_leaf_nodes=4, random_state=0)
# Predict and evaluate using model
estimator.fit(X_train_std,y_train)
# Predict and evaluate using model
rf_training_pred = estimator.predict(X_train_std)
rf_test_pred = estimator.predict(X_test_std)
rf_training_prediction = accuracy_score(y_train, rf_training_pred)
rf_test_prediction = accuracy_score(y_test, rf_test_pred)
print("Accuracy of Decision Tree training set:", round(rf_training_prediction,3))
print("Accuracy of Decision Tree test set:", round(rf_test_prediction,3))
print(classification_report(y_test, lr.predict(X_test_std)))
tn, fp, fn, tp = confusion_matrix(y_test, rf_test_pred).ravel()
precision = tp/(tp+fp)
recall = tp/(tp+fn)
f1_score = 2*((precision*recall)/(precision+recall))
print("True Positive: %i" %tp)
print("False Positive: %i" %fp)
print("True Negative: %i" %tn)
print("False Negative: %i" %fn)
print(f"Precision: {precision:.2%}")
print(f"Recall: {recall:.2%}")
print(f"F1-Score: {f1_score:.2%}")
# -
| Marketing/Predicting clicks on online advertising by Facebook/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp ipytyping.annotations
# -
# hide
# %load_ext autoreload
# %autoreload 2
#exporti
from pathlib import Path
from collections.abc import MutableMapping
from typing import Dict, Optional, Iterable, Any, Union
from ipywidgets import Layout
from ipyannotator.mltypes import OutputImageLabel, OutputLabel
from ipyannotator.custom_input.buttons import ImageButton, ImageButtonSetting, ActionButton
# hide
import ipytest
import pytest
ipytest.autoconfig(raise_on_error=True)
# ## Annotation Types
#
# The current notebook store the annotation data typing. Every annotator stores its data in a particular way, this notebook designs the store and it's casting types.
#exporti
class AnnotationStore(MutableMapping):
def __init__(self, annotations: Optional[Dict] = None):
self._annotations = annotations or {}
def __getitem__(self, key: str):
return self._annotations[key]
def __delitem__(self, key: str):
if key in self:
del self._annotations[key]
def __setitem__(self, key: str, value: Any):
self._annotations[key] = value
def __iter__(self):
return iter(self._annotations)
def __len__(self):
return len(self._annotations)
def __repr__(self):
return "{}({!r})".format(self.__class__.__name__, self._annotations)
# ### LabelStore Data Type
#
# The `LabelStore` stores a path as a key it's answer in the format: `{'<path>': {'answer': <bool>}`.
#exporti
class LabelStore(AnnotationStore):
def __getitem__(self, key: str):
assert isinstance(key, str)
return self._annotations[key]
def __delitem__(self, key: str):
assert isinstance(key, str)
if key in self:
del self._annotations[key]
def __setitem__(self, key: str, value: Optional[Dict[str, bool]]):
assert isinstance(key, str)
if value:
assert isinstance(value, dict)
self._annotations[key] = value
# The following cell will define a cast from the annotation to a custom widget called `ImageButton`.
#exporti
def _label_store_to_image_button(
annotation: LabelStore,
width: int = 150,
height: int = 150,
disabled: bool = False
) -> Iterable[ImageButton]:
button_setting = ImageButtonSetting(
display_label=False,
image_width=f'{width}px',
image_height=f'{height}px'
)
buttons = []
for path, value in annotation.items():
image_button = ImageButton(button_setting)
image_button.image_path = str(path)
image_button.label_value = Path(path).stem
image_button.active = value.get('answer', False)
image_button.disabled = disabled
buttons.append(image_button)
return buttons
#exporti
def _label_store_to_button(
annotation: LabelStore,
disabled: bool
) -> Iterable[ActionButton]:
layout = {
'width': 'auto',
'height': 'auto'
}
buttons = []
for label, value in annotation.items():
button = ActionButton(layout=Layout(**layout))
button.description = label
button.value = label
button.tooltip = label
button.disabled = disabled
if value.get('answer', True):
button.layout.border = 'solid 2px #f7f01e'
buttons.append(button)
return buttons
#exporti
class LabelStoreCaster: # pylint: disable=too-few-public-methods
"""Factory that casts the correctly widget
accordingly with the input"""
def __init__(
self,
output: Union[OutputImageLabel, OutputLabel],
width: int = 150,
height: int = 150,
widgets_disabled: bool = False
):
self.width = width
self.height = height
self.output = output
self.widgets_disabled = widgets_disabled
def __call__(self, annotation: LabelStore) -> Iterable:
if isinstance(self.output, OutputImageLabel):
return _label_store_to_image_button(
annotation,
self.width,
self.height,
self.widgets_disabled
)
if isinstance(self.output, OutputLabel):
return _label_store_to_button(
annotation,
disabled=self.widgets_disabled
)
raise ValueError(
f"output should have type OutputImageLabel or OutputLabel. {type(self.output)} given"
)
@pytest.fixture
def str_label_fixture():
return {
'A': {'answer': False},
'B': {'answer': True}
}
@pytest.fixture
def img_label_fixture():
return {
'../data/projects/capture1/pics/pink25x25.png': {'answer': False},
}
# %%ipytest
def test_it_cast_label_store_to_image_button(img_label_fixture):
label_store = LabelStore()
label_store.update(img_label_fixture)
output = OutputImageLabel()
caster = LabelStoreCaster(output)
image_buttons = caster(label_store)
for image_button in image_buttons:
assert isinstance(image_button, ImageButton)
assert len(image_buttons) == 1
# %%ipytest
def test_it_cast_label_store_to_button(str_label_fixture):
label_store = LabelStore()
label_store.update(str_label_fixture)
output = OutputLabel(class_labels=list(str_label_fixture.keys()))
caster = LabelStoreCaster(output)
buttons = caster(label_store)
assert len(buttons) == 2
for button in buttons:
assert isinstance(button, ActionButton)
assert buttons[0].description == 'A'
assert buttons[1].description == 'B'
assert buttons[0].value == 'A'
assert buttons[1].value == 'B'
# %%ipytest
def test_it_can_disable_widgets(str_label_fixture):
label_store = LabelStore()
label_store.update(str_label_fixture)
output = OutputLabel(class_labels=list(str_label_fixture.keys()))
caster = LabelStoreCaster(output, widgets_disabled=True)
buttons = caster(label_store)
for button in buttons:
assert button.disabled is True
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/00c_annotation_types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Particle filtering
#
# Particle filtering is not working yet - **WORK IN PROGRESS!**
# +
import os, sys
sys.path.append(os.path.abspath('../../main/python'))
import datetime as dt
import numpy as np
import numpy.testing as npt
import matplotlib.pyplot as plt
from thalesians.tsa.distrs import NormalDistr as N
import thalesians.tsa.filtering as filtering
import thalesians.tsa.filtering.kalman as kalman
import thalesians.tsa.filtering.particle as particle
import thalesians.tsa.numpyutils as npu
import thalesians.tsa.processes as proc
import importlib
importlib.reload(particle)
importlib.reload(proc)
# -
# ## A single-process, univariate example
#
# First we need a **process model**. In this case it will be a single stochastic process,
process = proc.WienerProcess.create_from_cov(mean=3., cov=0.0001)
# This we pass to a newly created Kalman filter, along with the initial time and initial state. The latter takes the form of a normal distribution. We have chosen to use Python `datetime`s as our data type for time, but we could have chosen `int`s or something else.
t0 = dt.datetime(2017, 5, 12, 16, 18, 25, 204000)
pf = particle.ParticleFilter(t0, state_distr=N(mean=100., cov=0.0000000000001), process=process)
# Next we create an **observable**, which incorporates a particular **observation model**. In this case, the observation model is particularly simple, since we are observing the entire state of the Kalman filter. Our observation model is a 1x1 identity:
observable = pf.create_observable(kalman.KalmanFilterObsModel.create(1.), process)
# Let's roll forward the time by one hour:
t1 = t0 + dt.timedelta(hours=1)
# What is our predicted observation at this time? Since we haven't observed any actual information, this is our **prior** observation estimate:
prior_predicted_obs1 = observable.predict(t1)
prior_predicted_obs1
# We confirm that this is consistent with how our (linear-Gaussian) process model scales over time:
np.mean(pf._prior_particles), 100. + 3./24.
prior_predicted_obs1
prior_predicted_obs1 = observable.predict(t1)
npt.assert_almost_equal(prior_predicted_obs1.distr.mean, 100. + 3./24.)
npt.assert_almost_equal(prior_predicted_obs1.distr.cov, 250. + 25./24.)
npt.assert_almost_equal(prior_predicted_obs1.cross_cov, prior_predicted_obs1.distr.cov)
# Let us now actually *observe* our observation. Say, the observation is 100.35 and the observation noise covariance is 100.0:
observable.observe(time=t1, obs=N(mean=100.35, cov=100.0))
# Having seen an actual observation, let us obtain the **posterior** observation estimate:
posterior_predicted_obs1 = observable.predict(t1); posterior_predicted_obs1
# We can now fast-forward the time, by two hours, say, and repeat the process:
# +
t2 = t1 + dt.timedelta(hours=2)
prior_predicted_obs2 = observable.predict(t2)
npt.assert_almost_equal(prior_predicted_obs2.distr.mean, 100.28590504 + 2.*3./24.)
npt.assert_almost_equal(prior_predicted_obs2.distr.cov, 71.513353115 + 2.*25./24.)
npt.assert_almost_equal(prior_predicted_obs2.cross_cov, prior_predicted_obs2.distr.cov)
observable.observe(time=t2, obs=N(mean=100.35, cov=100.0))
posterior_predicted_obs2 = observable.predict(t2)
npt.assert_almost_equal(posterior_predicted_obs2.distr.mean, 100.45709020)
npt.assert_almost_equal(posterior_predicted_obs2.distr.cov, 42.395213845)
npt.assert_almost_equal(posterior_predicted_obs2.cross_cov, posterior_predicted_obs2.distr.cov)
# -
# ## A multi-process, multivariate example
# The real power of our Kalman filter interface is demonstrated for process models consisting of several (independent) stochastic processes:
process1 = proc.WienerProcess.create_from_cov(mean=3., cov=25.)
process2 = proc.WienerProcess.create_from_cov(mean=[1., 4.], cov=[[36.0, -9.0], [-9.0, 25.0]])
# Such models are common in finance, where, for example, the dynamics of a yield curve may be represented by a (multivariate) stochastic process, whereas the idiosyncratic spread for each bond may be an independent stochastic process.
#
# Let us pass `process1` and `process2` as a (compound) process model to our Kalman filter, along with the initial time and state:
t0 = dt.datetime(2017, 5, 12, 16, 18, 25, 204000)
kf = kalman.KalmanFilter(
t0,
state_distr=N(
mean=[100.0, 120.0, 130.0],
cov=[[250.0, 0.0, 0.0],
[0.0, 360.0, 0.0],
[0.0, 0.0, 250.0]]),
process=(process1, process2))
# We shall now create several **observables**, each corresponding to a distinct **observation model**. The first one will observe the entire state:
state_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(1.0, np.eye(2)),
process1, process2)
# The second observable will observe the first coordinate of the first process:
coord0_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(1.),
process1)
# The third, the first coordinate of the second process:
coord1_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(npu.row(1., 0.)),
process2)
# The fourth, the second coordinate of the second process:
coord2_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(npu.row(0., 1.)),
process2)
# The fifth will observe the sum of the entire state (across the two processes):
sum_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(npu.row(1., 1., 1.)),
process1, process2)
# And the sixth a certain linear combination thereof:
lin_comb_observable = kf.create_observable(
kalman.KalmanFilterObsModel.create(npu.row(2., 0., -3.)),
process1, process2)
# Fast-forward the time by one hour:
t1 = t0 + dt.timedelta(hours=1)
# Let's predict the state at this time...
predicted_obs1_prior = state_observable.predict(t1)
predicted_obs1_prior
# And check that it is consistent with the scaling of the (multivariate) Wiener process with time:
npt.assert_almost_equal(predicted_obs1_prior.distr.mean,
npu.col(100.0 + 3.0/24.0, 120.0 + 1.0/24.0, 130.0 + 4.0/24.0))
npt.assert_almost_equal(predicted_obs1_prior.distr.cov,
[[250.0 + 25.0/24.0, 0.0, 0.0],
[0.0, 360.0 + 36.0/24.0, -9.0/24.0],
[0.0, -9.0/24.0, 250 + 25.0/24.0]])
npt.assert_almost_equal(predicted_obs1_prior.cross_cov, predicted_obs1_prior.distr.cov)
# Suppose that a new observation arrives, and we observe each of the three coordinates individually:
state_observable.observe(time=t1, obs=N(mean=[100.35, 121.0, 135.0],
cov=[[100.0, 0.0, 0.0],
[0.0, 400.0, 0.0],
[0.0, 0.0, 100.0]]));
# Let's look at our (posterior) predicted state:
state_observable.predict(t1)
# Let's also look at the predictions for the individual coordinates:
coord0_observable.predict(t1)
coord1_observable.predict(t1)
coord2_observable.predict(t1)
# The predicted sum:
sum_observable.predict(t1)
# And the predicted linear combination:
lin_comb_observable.predict(t1)
# Let's now go 30 minutes into the future:
t2 = t1 + dt.timedelta(minutes=30)
# And observe only the first coordinate of the second process, with a pretty high confidence:
coord1_observable.observe(time=t2, obs=N(mean=125.25, cov=4.))
# How does our predicted state change?
state_observable.predict(t2)
# Thirty minutes later...
t3 = t2 + dt.timedelta(minutes=30)
# We observe the *sum* of the three coordinates, rather than the individual coordinates:
sum_observable.observe(time=t3, obs=N(mean=365.00, cov=9.))
# How has our prediction of the state changed?
state_observable.predict(t3)
# And what is its predicted sum?
sum_observable.predict(t3)
| tsa/src/jupyter/python/particle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Shahid-coder/python-colab/blob/main/03_strings.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="XRQ08AZ5UznB"
# # Strings
# Strings in python are surrounded by either single quotation marks, or double quotation marks.
#
# 'hello' is the same as "hello".
#
# You can display a string literal with the print() function:
# + id="hDdLFfxAT1q1" colab={"base_uri": "https://localhost:8080/"} outputId="bd4435bb-ca1e-46f2-acee-c2269cfebc77"
print("Hello")
print('Hello')
# + [markdown] id="CE5o_nl_VP7H"
# # Slicing
# You can return a range of characters by using the slice syntax.
#
# Specify the start index and the end index, separated by a colon, to return a part of the string.
# + colab={"base_uri": "https://localhost:8080/"} id="aulXMwmcU8U2" outputId="1eb22e41-6d5c-40f1-b255-786112e0aa4d"
b = "Hello, World!"
print(b[2:5])
# + [markdown] id="IOaH80X5VrH8"
# # Slice To the End
# By leaving out the end index, the range will go to the end:
# + colab={"base_uri": "https://localhost:8080/"} id="iqNTcwGfVN4i" outputId="5f7c6c4c-1ff9-42a1-bebf-71d5a7cf7e6d"
b = "Hello, World!"
print(b[2:])
# + [markdown] id="HTPQ6hLQV1fZ"
# # Negative Indexing
# Use negative indexes to start the slice from the end of the string:
# + colab={"base_uri": "https://localhost:8080/"} id="KvNcqgvzVxMu" outputId="a77e3f8d-d380-4e48-d22a-40d2a698422e"
b = "Hello, World!"
print(b[-5:-2])
# + [markdown] id="QnyS7HZeWWvw"
# # Python - Modify Strings
# Python has a set of built-in methods that you can use on strings.
# + [markdown] id="wjledfvZWgjz"
# Upper Case:
# + colab={"base_uri": "https://localhost:8080/"} id="OPsGyqGKWfOM" outputId="f2b965a8-4ae8-41e7-c0f8-efc9a16e532b"
a = "Hello, World!"
print(a.upper())
# + [markdown] id="OZItRE9cWnaq"
# Lower Case:
# + colab={"base_uri": "https://localhost:8080/"} id="qw932ACxWmLi" outputId="4d4debd0-872a-4ee6-f7a3-17f12d16b2c4"
a = "Hello, World!"
print(a.lower())
# + [markdown] id="mONksT5DWu6I"
# #### Remove Whitespace:
# Whitespace is the space before and/or after the actual text, and very often you want to remove this space.
# + [markdown] id="GaGmBXZwW3sA"
# The strip() method removes any whitespace from the beginning or the end:
# + colab={"base_uri": "https://localhost:8080/"} id="h9hr3bNhWr4G" outputId="7885ed76-07c5-4b80-9bbd-0f1a9dd5a227"
a = " Hello, World! "
print(a.strip()) # returns "Hello, World!"
# + [markdown] id="QsYCJqLUXDXF"
# # Replace String
# The replace() method replaces a string with another string:
# + colab={"base_uri": "https://localhost:8080/"} id="rMYChue8W9y4" outputId="554d1699-80f7-424b-c57c-2e57b6b21376"
a = "Hello, World!"
print(a.replace("H", "J"))
# + [markdown] id="BQbY2IvYXOqT"
# # Split String
# The split() method returns a list where the text between the specified separator becomes the list items.
# + colab={"base_uri": "https://localhost:8080/"} id="UssdzTZIXI1M" outputId="2078cae8-a1e5-4add-b45b-5847f83ccc66"
a = "Hello, World!"
print(a.split(",")) # returns ['Hello', ' World!']
# + [markdown] id="kY-3QD7NXab0"
# # String Concatenation
# To concatenate, or combine, two strings you can use the + operator.
# + [markdown] id="q6WDCgjZXmSZ"
# Merge variable a with variable b into variable c:
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="3cwqWVmXXUxn" outputId="563ce50f-0b21-47c2-f51f-cb2ce0f67525"
a = "Hello"
b = "World"
c = a + b
c
# + [markdown] id="kzzr6By-Xn5J"
# To add a space between them, add a " ":
#
# + colab={"base_uri": "https://localhost:8080/"} id="23UTlKK9Xf-H" outputId="65532b0b-3fb4-462c-fb98-72923939a3fc"
a = "Hello"
b = "World"
c = a + " " + b
print(c)
# + [markdown] id="-ymbQlhtXzFj"
# String Format
# As we learned in the Python Variables chapter, we cannot combine strings and numbers.But we can combine strings and numbers by using the format() method!
#
#
# + [markdown] id="Di0VlS7hX-yV"
# The format() method takes the passed arguments, formats them, and places them in the string where the placeholders {} are:
# + colab={"base_uri": "https://localhost:8080/"} id="L91YQso9Xta7" outputId="09ac599e-024d-4e4b-da5f-e0ca6d7c19a3"
age = 36
txt = "My name is John, and I am {}"
print(txt.format(age))
# + [markdown] id="IL7pBUrkYLwB"
# The format() method takes unlimited number of arguments, and are placed into the respective placeholders:
# + colab={"base_uri": "https://localhost:8080/"} id="lgyNi5l_YDyQ" outputId="5ed574be-c975-46f4-9232-f56c37e95967"
quantity = 3
itemno = 567
price = 49.95
myorder = "I want {} pieces of item {} for {} dollars."
print(myorder.format(quantity, itemno, price))
# + [markdown] id="baIezYHQYRFd"
# You can use index numbers {0} to be sure the arguments are placed in the correct placeholders:
# + colab={"base_uri": "https://localhost:8080/"} id="SgmLJW0QYN0e" outputId="89eddff4-3060-4ad3-9811-0dce7e163af4"
quantity = 3
itemno = 567
price = 49.95
myorder = "I want to pay {2} dollars for {0} pieces of item {1}."
print(myorder.format(quantity, itemno, price))
# + [markdown] id="qmwWi05eYfO4"
# Escape Character
# To insert characters that are illegal in a string, use an escape character.
#
# An escape character is a backslash \ followed by the character you want to insert.
#
# An example of an illegal character is a double quote inside a string that is surrounded by double quotes:
# + id="cefLvdTtYr7j"
txt = "We are the so-called "Vikings" from the north."
# + [markdown] id="jlTk7OkcYnU6"
# The escape character allows you to use double quotes when you normally would not be allowed:
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="O-Sj4t55YVpO" outputId="40e77b74-f968-4be6-b5d4-3e693b23723e"
txt = "We are the so-called \"Vikings\" from the north."
txt
# + [markdown] id="AWjQHtS4ZIWN"
# # String Methods
# Python has a set of built-in methods that you can use on strings.
# + colab={"base_uri": "https://localhost:8080/"} id="4dh-2aYOZM4K" outputId="c89250d3-eae0-44ba-d9e8-0dc45ed7dc2a"
# capitalize
# The capitalize() method returns a string where the first character is upper case.
txt = "hello, and welcome to my world."
x = txt.capitalize()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="mzOhlrs7ZemL" outputId="5e4c11fd-06dc-4438-b9d9-24b1cdb0356d"
#Case fold
#Converts string into lower case
txt = "Hello, And Welcome To My World!"
x = txt.casefold()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="VJd7CoHVZ5dQ" outputId="4389bde9-5663-4731-c83d-514c7b25c30a"
# center()
#Returns a centered string
txt = "banana"
x = txt.center(20)
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="aqOys0a4Z_3F" outputId="ef7d950d-0539-441d-ba78-511baf58ca96"
# count
# Returns the number of times a specified value occurs in a string
txt = "I love apples, apple are my favorite fruit"
x = txt.count("apple")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="CCh413vxaQ6K" outputId="cfd4938b-1d88-46cf-c05e-f87be6435d24"
# encode()
# Returns an encoded version of the string
txt = "My name is Ståle"
x = txt.encode()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="yH0UgGaxaY2T" outputId="60e929f2-b18b-4254-d5cd-dc1525c6ccf1"
# endswith
# Returns true if the string ends with the specified value
txt = "Hello, welcome to my world."
x = txt.endswith(".")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="HvLb9qBaavMb" outputId="6467a52d-7e02-45b5-bbce-10629db08248"
# expandtabs()
#Sets the tab size of the string
txt = "H\te\tl\tl\to"
x = txt.expandtabs(2)
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="RD0ZiEqMa6QE" outputId="b477ff72-909e-461d-f37b-aeb7d0167228"
# find()
# Searches the string for a specified value and returns the position of where it was found
txt = "Hello, welcome to my world."
x = txt.find("welcome")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="2FpUHazqa_01" outputId="b988e763-0559-4f18-bbf9-a489123a29ff"
# format()
# Formats specified values in a string
txt = "For only {price:.2f} dollars!"
print(txt.format(price = 49))
# + id="7HRVwF8dbaxs"
# format_map
# Formats specified values in a string
# + colab={"base_uri": "https://localhost:8080/"} id="Tgn-zHb5bjGY" outputId="254f6af7-13c2-435b-b30f-f42e86c7fe18"
# index()
#Searches the string for a specified value and returns the position of where it was found
txt = "Hello, welcome to my world."
x = txt.index("welcome")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="SgWnfAo3brKM" outputId="4434714f-0e81-4ef2-9510-384658c2ff19"
#isalnum()
#Returns True if all characters in the string are alphanumeric
txt = "Company12"
x = txt.isalnum()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="_b_VQd8Ub3qE" outputId="34ec4962-0991-4ea0-ff3c-b560701fefbe"
#isalpha
#Returns True if all characters in the string are in the alphabet
txt = "CompanyX"
x = txt.isalpha()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="7L90sIdwcILr" outputId="c01fa073-b1be-4953-cdc3-0b969b704711"
#isdecimal()
#Returns True if all characters in the string are decimals
txt = "\u0033" #unicode for 3
x = txt.isdecimal()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="vAwjaGbLca4d" outputId="60db2d0c-d216-4c57-fefa-420c6fcbaf14"
#isdigits()
#Returns True if all characters in the string are digits
txt = "50800"
x = txt.isdigit()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="88Pw37Z8cj0c" outputId="fcc258b0-cdc8-4fde-b4bf-0a738fb4e547"
#isidentifier()
#Returns True if the string is an identifier
txt = "Demo"
x = txt.isidentifier()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="udUFLFzwc0e9" outputId="3c53a9b7-8096-4149-8c60-70361289f1c0"
#islower()
#Returns True if all characters in the string are lower case
txt = "hello world!"
x = txt.islower()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="ysAncXfHeZR1" outputId="50bbb50f-7ced-4ea3-f0b5-10804e1d6e12"
#isnumeric()
#Returns True if all characters in the string are numeric
txt = "565543"
x = txt.isnumeric()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="-rD9mFiJejTm" outputId="bb4b0963-d3c2-409c-8430-ae6cc21ec64b"
#isprintable()
#Returns True if all characters in the string are printable
txt = "Hello! Are you #1?"
x = txt.isprintable()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="9oByXvyqe0F9" outputId="8815ffe7-581e-479e-ba6f-9fcada90f225"
#isspace()
#Returns True if all characters in the string are whitespaces
txt = " "
x = txt.isspace()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="mFMkqTxde-gW" outputId="af064fdf-1406-4491-9a15-3907fb8f5871"
#istitle()
#Returns True if the string follows the rules of a title
txt = "Hello, And Welcome To My World!"
x = txt.istitle()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="30Re3fLafPGZ" outputId="e20f01f2-c5b8-41d4-909e-de29d0108cac"
#isupper()
#Returns True if all characters in the string are upper case
txt = "THIS IS NOW!"
x = txt.isupper()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="AEO1PdSCfYdN" outputId="4d764e72-ab6d-47de-bd29-e53796e471c7"
#join()
#Joins the elements of an iterable to the end of the string
myTuple = ("John", "Peter", "Vicky")
x = "#".join(myTuple)
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="1c1VGIH5fr9i" outputId="7d384a24-aadc-4aa5-dcad-a4de6104ddaf"
#ljust()
#Returns a left justified version of the string
txt = "banana"
x = txt.ljust(20)
print(x, "is my favorite fruit.")
# + colab={"base_uri": "https://localhost:8080/"} id="X9FgGjoif5Y6" outputId="b09fa396-1e1a-499b-ab47-655b6bd5ce88"
#lower()
#Converts a string into lower case
txt = "<NAME>"
x = txt.lower()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="eS8u6eRggDT8" outputId="c19d6f4d-03ee-4970-b3c4-32e382349d86"
#lstrip()
#Returns a left trim version of the string
txt = " banana "
x = txt.lstrip()
print("of all fruits", x, "is my favorite")
# + colab={"base_uri": "https://localhost:8080/"} id="Q40ViMaOgPIW" outputId="8428d768-eec0-4a3c-f530-7e3da0be5f38"
#maketrans()
#Returns a translation table to be used in translations
txt = "Hello Sam!"
mytable = txt.maketrans("S", "P")
print(txt.translate(mytable))
# + colab={"base_uri": "https://localhost:8080/"} id="nv27XC_LgYxp" outputId="124d9b9f-95b1-4b60-c3bc-32d8657e594e"
#partition()
#Returns a tuple where the string is parted into three parts
'''
Search for the word "bananas", and return a tuple with three elements:
1 - everything before the "match"
2 - the "match"
3 - everything after the "match"
'''
txt = "I could eat bananas all day"
x = txt.partition("bananas")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="mUCcC0z3glvd" outputId="26b05799-d00d-472b-d3d6-a7cccc579556"
#replace()
#Returns a string where a specified value is replaced with a specified value
txt = "I like bananas"
x = txt.replace("bananas", "apples")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="SirxJ5DMg41Q" outputId="cf4b0c67-34a5-408c-9324-389c4066dc17"
#rfind()
#Searches the string for a specified value and returns the last position of where it was found
txt = "Mi casa, su casa."
x = txt.rfind("casa")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="Q-T0almNhFcP" outputId="30995966-d41f-4b9f-b4f0-dd09f0a8992e"
#rindex()
#Searches the string for a specified value and returns the last position of where it was found
txt = "Mi casa, su casa."
x = txt.rindex("casa")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="j8_3rGbGhMTL" outputId="3a8028d0-3093-42e1-be73-7245cf917c80"
#rjust()
#Returns a right justified version of the string
txt = "banana"
x = txt.rjust(20)
print(x, "is my favorite fruit.")
# + colab={"base_uri": "https://localhost:8080/"} id="Q_G4uvC5hj7F" outputId="ff470dfb-7854-4f72-9168-a456cf44106b"
#rpartition()
#Returns a tuple where the string is parted into three parts
'''
Search for the last occurrence of the word "bananas", and return a tuple with three elements:
1 - everything before the "match"
2 - the "match"
3 - everything after the "match"
'''
txt = "I could eat bananas all day, bananas are my favorite fruit"
x = txt.rpartition("bananas")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="ySATgMpHh4eY" outputId="c1f5656e-9657-4911-998e-dc18668263ce"
#rsplit()
#Splits the string at the specified separator, and returns a list
txt = "apple, banana, cherry"
x = txt.rsplit(", ")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="O63_obLKiFEr" outputId="72c112f1-7fb8-410b-c7f5-2839aad557d2"
#rstrip()
#Returns a right trim version of the string
txt = " banana "
x = txt.rstrip()
print("of all fruits", x, "is my favorite")
# + colab={"base_uri": "https://localhost:8080/"} id="Mo94a6TTibUg" outputId="ce43f334-7c9f-4531-f3ad-edbc1c9d14d2"
#rsplit()
#Splits the string at the specified separator, and returns a list
txt = "welcome to the jungle"
x = txt.split()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="VXAM0pyRimdw" outputId="f326467a-41df-48e9-8fb1-913f67da83cf"
#splitlines()
#Splits the string at line breaks and returns a list
txt = "Thank you for the music\nWelcome to the jungle"
x = txt.splitlines()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="1kRSCKI6iv90" outputId="e87eea23-2dbe-4d6d-ebab-ae86a3d5e530"
#startswith()
#Returns true if the string starts with the specified value
txt = "Hello, welcome to my world."
x = txt.startswith("Hello")
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="gy1B98aci-hB" outputId="a7545ac4-b760-448a-d8f1-01f5218cfca3"
#strip()
#Returns a trimmed version of the string
txt = " banana "
x = txt.strip()
print("of all fruits", x, "is my favorite")
# + colab={"base_uri": "https://localhost:8080/"} id="nrSk6H-FjJ2z" outputId="6dcc4f24-950a-4229-b808-d2f2b1170036"
#swapcase
#Swaps cases, lower case becomes upper case and vice versa
txt = "Hello My Name Is PETER"
x = txt.swapcase()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="1PrnZ0TsjUbC" outputId="765fb1bb-2a79-4ea6-ec58-e59cf4b23d12"
#title()
#Converts the first character of each word to upper case
txt = "Welcome to my world"
x = txt.title()
print(x)
# + colab={"base_uri": "https://localhost:8080/"} id="UOITMO5ajmAD" outputId="5a12ed2c-ae1a-4517-a7aa-691b4a51e208"
#translate()
#Returns a translated string
#use a dictionary with ascii codes to replace 83 (S) with 80 (P):
mydict = {83: 80}
txt = "Hello Sam!"
print(txt.translate(mydict))
# + colab={"base_uri": "https://localhost:8080/"} id="Cx-NKCLDjx2J" outputId="4620095a-2d88-4575-ce2f-d78ede569c8a"
#upper()
#Converts a string into upper case
txt = "Hello my friends"
x = txt.upper()
print(x)
# + id="6zK2Fqe7j-f3"
#zfill()
#Fills the string with a specified number of 0 values at the beginning
txt = "50"
x = txt.zfill(10)
print(x)
| 03_strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
## comment out if running in colab
import os
os.chdir('../')
# ## if useing Google colab, uncomment the following code
# # !git clone https://github.com/nicktfranklin/SEM.git
# import os
# os.chdir('./SEM/')
# # !pip install tensorflow==1.9
# # !pip install keras==2.2
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from models import *
from opt import encode
from tqdm import tnrange
sns.set_context('paper', font_scale=1.5)
# +
np.random.seed(0)
# figure out how many dimensions we need using plates formula
n = 18 # number of atomic values in the language
k = 6; # maximum number of terms to be combined
err = 0.02; # error probability
from opt import plate_formula, embed, encode, decode, embed_onehot
d = plate_formula(n, k, err)
print d
def embed(d):
return np.random.randn(1, d) / np.sqrt(d)
verb_property = embed(d) * 1.00
noun_property = embed(d) * 1.00
words = {
'Ask': embed(d),
'Answer': embed(d),
'Chase': embed(d),
'Tom': (embed(d) + noun_property),
'Charan':(embed(d) + noun_property),
'Jill': (embed(d) + noun_property),
'Talia': (embed(d) + noun_property),
'Tony': (embed(d) + noun_property),
'Clarisa': (embed(d) + noun_property),
'Bruce': (embed(d) + noun_property),
'Dick': (embed(d) + noun_property),
'Dog': (embed(d) + noun_property),
'Cat': (embed(d) + noun_property),
}
roles = {
'Agent': embed(d),
'Patient': embed(d),
'Verb': embed(d)
}
# +
from sklearn.metrics.pairwise import cosine_distances, euclidean_distances
keys = words.keys()
keys.sort()
print keys
sns.heatmap(cosine_distances([words[w].reshape(-1) for w in keys]))
# -
keys = words.keys()
keys.sort()
print keys
sns.heatmap(euclidean_distances([words[w].reshape(-1) for w in keys]))
# +
sentences = [
['Ask', 'Tom', 'Charan'], ['Answer', 'Charan', 'Tom'],
# ['Ask', 'Tom', 'Jill'], ['Answer', 'Jill', 'Tom'],
# ['Ask', 'Tom', 'Talia'], ['Answer', 'Talia', 'Tom'],
# ['Ask', 'Tom', 'Tony'], ['Answer', 'Tony', 'Tom'],
# ['Ask', 'Tom', 'Clarisa'], ['Answer', 'Clarisa', 'Tom'],
# ['Ask', 'Charan', 'Tom'], ['Answer', 'Tom', 'Charan'],
# ['Ask', 'Charan', 'Jill'], ['Answer', 'Jill', 'Charan'],
# ['Ask', 'Charan', 'Talia'], ['Answer', 'Talia', 'Charan'],
# ['Ask', 'Charan', 'Tony'], ['Answer', 'Tony', 'Charan'],
# ['Ask', 'Charan', 'Clarisa'], ['Answer', 'Clarisa', 'Charan'],
['Ask', 'Jill', 'Talia'], ['Answer', 'Talia', 'Jill'],
# ['Ask', 'Jill', 'Tom'], ['Answer', 'Tom', 'Jill'],
# ['Ask', 'Jill', 'Charan'], ['Answer', 'Charan', 'Jill'],
# ['Ask', 'Jill', 'Tony'], ['Answer', 'Tony', 'Jill'],
# ['Ask', 'Jill', 'Clarisa'], ['Answer', 'Charan', 'Jill'],
# ['Ask', 'Talia', 'Tom'], ['Answer', 'Tom', 'Talia'],
# ['Ask', 'Talia', 'Jill'], ['Answer', 'Jill', 'Talia'],
# ['Ask', 'Talia', 'Charan'], ['Answer', 'Charan', 'Talia'],
# ['Ask', 'Talia', 'Tony'], ['Answer', 'Tony', 'Talia'],
# ['Ask', 'Talia', 'Clarisa'], ['Answer', 'Clarisa', 'Talia'],
# ['Ask', 'Tony', 'Tom'], ['Answer', 'Tom', 'Tony'],
# ['Ask', 'Tony', 'Jill'], ['Answer', 'Jill', 'Tony'],
# ['Ask', 'Tony', 'Charan'], ['Answer', 'Charan', 'Tony'],
# ['Ask', 'Tony', 'Talia'], ['Answer', 'Talia', 'Tony'],
['Ask', 'Tony', 'Clarisa'], ['Answer', 'Clarisa', 'Tony'],
# ['Ask', 'Clarisa', 'Tom'], ['Answer', 'Tom', 'Clarisa'],
# ['Ask', 'Clarisa', 'Jill'], ['Answer', 'Jill', 'Clarisa'],
# ['Ask', 'Clarisa', 'Charan'], ['Answer', 'Charan', 'Clarisa'],
# ['Ask', 'Clarisa', 'Talia'], ['Answer', 'Talia', 'Clarisa'],
# ['Ask', 'Clarisa', 'Tony'], ['Answer', 'Tony', 'Clarisa'],
]
x_train = []
for s in sentences:
x_train.append(
encode(words[s[0]], roles['Verb'])
+ encode(words[s[1]], roles['Agent'])
+ encode(words[s[2]], roles['Patient'])
)
test_a = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Answer'], roles['Verb'])
+ encode(words['Charan'], roles['Agent'])
+ encode(words['Tom'], roles['Patient']),
]
test_b = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Bruce'], roles['Agent'])
+ encode(words['Dick'], roles['Patient']),
encode(words['Answer'], roles['Verb'])
+ encode(words['Dick'], roles['Agent'])
+ encode(words['Bruce'], roles['Patient'])
]
test_c = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Chase'], roles['Verb'])
+ encode(words['Dog'], roles['Agent'])
+ encode(words['Cat'], roles['Patient']),
]
test_d = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
]
test_e = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Ask'], roles['Verb'])
+ encode(words['Bruce'], roles['Agent'])
+ encode(words['Dick'], roles['Patient']),
]
test_a2x = [
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Answer'], roles['Verb'])
+ encode(words['Charan'], roles['Agent'])
+ encode(words['Tom'], roles['Patient']),
encode(words['Ask'], roles['Verb'])
+ encode(words['Bruce'], roles['Agent'])
+ encode(words['Dick'], roles['Patient']),
encode(words['Answer'], roles['Verb'])
+ encode(words['Dick'], roles['Agent'])
+ encode(words['Bruce'], roles['Patient'])
]
# +
def angle_cos(x, y):
return np.dot(x, y.T) / (np.linalg.norm(x) * np.linalg.norm(y))
print angle_cos(test_a[0], test_a[1])
print angle_cos(test_a[0], test_b[1])
print angle_cos(test_a[0], test_c[1])
print angle_cos(test_a[0], test_d[1])
print angle_cos(test_a[0], test_e[1])
# -
np.linalg.norm(np.dot(test_a[0], test_b[1].T), axis=1)
np.linalg.norm(np.dot(test_a[0], test_c[1].T), axis=1)
# +
# SEM parameters
# df0 = 1.0
# mode = 0.15
df0 = 100.0
mode = 0.305
scale0 = (mode * (df0 + 2)) / df0
print scale0
lmda = 1.0 # stickyness parameter
alfa = 2. # concentration parameter
f_class = NonLinearEvent
# f_class = LinearDynamicSystem
f_opts=dict(var_scale0=scale0, var_df0=df0, dropout=0.5, n_hidden = d, n_epochs=100,
prior_log_prob=2.0)
sem_kwargs = dict(lmda=lmda, alfa=alfa, f_class=f_class, f_opts=f_opts)
_x = np.concatenate(x_train, axis=0)
event_types = np.array([0] * len(x_train))
event_boundaries = np.array([1, 0] * len(x_train))
sem_model = SEM(**sem_kwargs)
# +
def print_evalute(x, event_model, metric=euclidean_distances):
print "t=0, f0={}, origin={}".format(
metric(event_model.predict_f0(), x[0])[0][0], metric(np.zeros((1, d)), x[0])[0][0])
print "t=1, f(x)={}, f0={}, origin={}".format(
metric(event_model.predict_next_generative(x[0]), x[1])[0][0],
metric(event_model.predict_f0(), x[1])[0][0],
metric(np.zeros((1, d)), x[1])[0][0])
def print_models_likelihood(x, event_model_0, event_model_1):
print "t=0, p(f0_a)={}, p(f0_a)={}".format(
event_model_0.log_likelihood_f0(x[0]),
event_model_1.log_likelihood_f0(x[0])
)
print "t=1, p(f(x)_a)={}, p(f0_a)={}, p(f0_b)={}".format(
event_model_0.log_likelihood_sequence(x[0], x[1]),
event_model_0.log_likelihood_f0(x[1]),
event_model_1.log_likelihood_f0(x[1])
)
# -
# # Test Case A
# +
# Ask(Tom, Charan) -> Answer(Charan, Tom)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
print sem_model.event_models[0].Sigma.mean()
sem_model._update_state(np.concatenate(test_a), 3)
sem_model.run(np.concatenate(test_a), compile_model=False)
results = [
{
'Test Case': 'Answer(Charan,Tom)',
'Boundary': np.exp(sem_model.results.log_boundary_probability)[1],
'New Event': 1-sem_model.results.post[1, 0]
}
]
print np.exp(sem_model.results.log_boundary_probability)
print sem_model.results.e_hat
# +
sns.set_context('paper', font_scale=1.0)
def plot_results(results, x_labels):
fig, ax = plt.subplots(1, 1, figsize=(3.0, 2.0), sharey=True)
w = 0.4
y = np.exp(sem_model.results.log_boundary_probability)
ax.bar([0-w/2., 1.-w/2.], y, facecolor=[0.65, 0.65, 0.65], width=w, edgecolor='k', lw=1, label='Boundary')
ax.set_ylabel('Probability')
ax.set_title('Boundary')
ax.set_title('New Event')
ax.set_xticks([0, 1])
ax.set_xticklabels(x_labels, rotation=0)
y = 1-results.post[:, 0]
print y
ax.bar([0+w/2., 1.+w/2.], y, facecolor=[0.35, 0.35, 0.35], width=w, edgecolor='k', lw=1, label='New Event')
plt.legend(loc='center right', bbox_to_anchor=(1.5, 0.5))
sns.despine()
plot_results(sem_model.results, 'Ask(Tom,Charan) Answer(Charan,Tom)'.split(' '))
# plt.savefig('generalization_test_case_a.png', dpi=300, bbox_inches='tight')
# -
sem_model.results.post[:, 0]
print sem_model.results.restart_prob, sem_model.results.repeat_prob
print sem_model.results.log_like
print sem_model.results.log_prior
print_evalute(test_a, sem_model.event_models[0])
print_models_likelihood(test_a, sem_model.event_models[0], sem_model.event_models[1])
# +
# check the decoding here
from opt import decode
from scipy.linalg import norm
from scipy.special import logsumexp
gamma = 50.0
agent = decode(sem_model.results.x_hat[1], roles['Agent'])
patient = decode(sem_model.results.x_hat[1], roles['Patient'])
verb = decode(sem_model.results.x_hat[1], roles['Verb'])
_df = []
for w in words.keys():
_df.append({
'Word': w,
'Agent Prob': gamma * -cosine_distances(agent, words[w])[0][0],
'Patient Prob': gamma * -cosine_distances(patient, words[w])[0][0],
'Verb Prob': gamma * -cosine_distances(verb, words[w])[0][0],
})
_df = pd.DataFrame(_df)
_df['Agent Prob'] =np.exp( _df['Agent Prob'] - logsumexp(_df['Agent Prob']))
_df['Patient Prob'] =np.exp( _df['Patient Prob'] - logsumexp(_df['Patient Prob']))
_df['Verb Prob'] =np.exp( _df['Verb Prob'] - logsumexp(_df['Verb Prob']))
# plot
fig, axes = plt.subplots(1, 3, figsize=(6.5, 2.0), sharey=True)
_df.sort_values('Agent Prob', inplace=True)
_df.plot(x='Word', y='Agent Prob', kind='barh', color='grey', ax=axes[0], legend=None, title='Agent')
_df.plot(x='Word', y='Patient Prob', kind='barh', color='grey', ax=axes[1], legend=None, title='Patient')
_df.plot(x='Word', y='Verb Prob', kind='barh', color='grey', ax=axes[2], legend=None, title='Verb')
for ax in axes:
ax.plot([1. / len(_df), 1. / len(_df)], plt.gca().get_ylim(), c='r', ls='--')
ax.set_xlabel('Decoding Probability')
plt.show()
plt.savefig('DecodingA.png', dpi=300, bbox_iunches='tight')
# +
from scipy.stats import multivariate_normal
print multivariate_normal.logpdf(test_a[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
print multivariate_normal.logpdf(test_b[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
# -
a = multivariate_normal.logpdf(test_a[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
b = multivariate_normal.logpdf(test_b[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
z = logsumexp([a, b])
print np.exp(a - z), np.exp(b - z)
print np.exp(a - b)
# # Test Case B
# +
# Ask(Bruce, Dick) -> Answer(Dick, Bruce)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
sem_model._update_state(np.concatenate(test_b), 3)
sem_model.run(np.concatenate(test_b), compile_model=False)
results.append(
{
'Test Case': 'Answer(Dick,Bruce)',
'Boundary': np.exp(sem_model.results.log_boundary_probability)[1],
'New Event': 1-sem_model.results.post[1, 0]
}
)
print np.exp(sem_model.results.log_boundary_probability)
print sem_model.results.e_hat
# -
plot_results(sem_model.results, 'Ask(Bruce,Dick) Answer(Dick,Bruce)'.split(' '))
# plt.savefig('generalization_test_case_b.png', dpi=300, bbox_inches='tight')
sem_model.results.log_like
print sem_model.results.restart_prob, sem_model.results.repeat_prob
print_evalute(test_b, sem_model.event_models[0])
print_models_likelihood(test_b, sem_model.event_models[0], sem_model.event_models[1])
# +
# check the decoding here
from opt import decode
from scipy.linalg import norm
from scipy.special import logsumexp
agent = decode(sem_model.results.x_hat[1], roles['Agent'])
patient = decode(sem_model.results.x_hat[1], roles['Patient'])
verb = decode(sem_model.results.x_hat[1], roles['Verb'])
_df = []
for w in words.keys():
_df.append({
'Word': w,
'Agent Prob': -gamma * cosine_distances(agent, words[w])[0][0],
'Patient Prob': -gamma * cosine_distances(patient, words[w])[0][0],
'Verb Prob': -gamma * cosine_distances(verb, words[w])[0][0],
})
_df = pd.DataFrame(_df)
_df['Agent Prob'] =np.exp( _df['Agent Prob'] - logsumexp(_df['Agent Prob']))
_df['Patient Prob'] =np.exp( _df['Patient Prob'] - logsumexp(_df['Patient Prob']))
_df['Verb Prob'] =np.exp( _df['Verb Prob'] - logsumexp(_df['Verb Prob']))
# plot
fig, axes = plt.subplots(1, 3, figsize=(6.5, 2.0), sharey=True)
_df.sort_values('Agent Prob', inplace=True)
_df.plot(x='Word', y='Agent Prob', kind='barh', color='grey', ax=axes[0], legend=None, title='Agent')
_df.plot(x='Word', y='Patient Prob', kind='barh', color='grey', ax=axes[1], legend=None, title='Patient')
_df.plot(x='Word', y='Verb Prob', kind='barh', color='grey', ax=axes[2], legend=None, title='Verb')
for ax in axes:
ax.plot([1. / len(_df), 1. / len(_df)], plt.gca().get_ylim(), c='r', ls='--')
ax.set_xlabel('Decoding Probability')
plt.show()
plt.savefig('DecodingB.png', dpi=300, bbox_iunches='tight')
# +
b_foil = encode(words['Answer'], roles['Verb']) \
+ encode(words['Bruce'], roles['Agent']) \
+ encode(words['Dick'], roles['Patient'])
a = multivariate_normal.logpdf(test_a[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
b = multivariate_normal.logpdf(test_b[1].reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
b0 = multivariate_normal.logpdf(b_foil.reshape(-1), mean=sem_model.results.x_hat[1].reshape(-1), cov=np.diag(sem_model.event_models[0].Sigma))
z = logsumexp([a, b, b0])
print np.exp(a - z), np.exp(b - z), np.exp(b0 - z)
print np.exp(a - b), np.exp(b - b0)
# -
sem_model.results.log_boundary_probability
# # Test Case C
# +
# Ask(Tom, Charan) -> Chase(Dog, Cat)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
sem_model._update_state(np.concatenate(test_c), 3)
sem_model.run(np.concatenate(test_c), compile_model=False)
results.append(
{
'Test Case': 'Chase(Dog,Cat)',
'Boundary': np.exp(sem_model.results.log_boundary_probability)[1],
'New Event': 1-sem_model.results.post[1, 0]
}
)
print np.exp(sem_model.results.log_boundary_probability)
print sem_model.results.e_hat
# -
plot_results(sem_model.results, 'Ask(Tom,Charan) Chase(Dog,Cat)'.split(' '))
# plt.savefig('generalization_test_case_c.png', dpi=300, bbox_inches='tight')
sem_model.results.log_like
print_evalute(test_c, sem_model.event_models[0])
print_models_likelihood(test_c, sem_model.event_models[0], sem_model.event_models[1])
# # Test Case D
# +
# Ask(Tom, Charan) -> Ask(Tom, Charan)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
sem_model._update_state(np.concatenate(test_d), 3)
sem_model.run(np.concatenate(test_d), compile_model=False)
results.append(
{
'Test Case': 'Ask(Tom,Charan)',
'Boundary': np.exp(sem_model.results.log_boundary_probability)[1],
'New Event': 1-sem_model.results.post[1, 0]
}
)
print np.exp(sem_model.results.log_boundary_probability)
print sem_model.results.e_hat
# -
plot_results(sem_model.results, 'Ask(Tom,Charan) Ask(Tom,Charan)'.split(' '))
# plt.savefig('generalization_test_case_d.png', dpi=300, bbox_inches='tight')
# ## Test Case E
# +
# Ask(Tom, Charan) -> Ask(Tom, Charan)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
sem_model._update_state(np.concatenate(test_e), 3)
sem_model.run(np.concatenate(test_e), compile_model=False)
results.append(
{
'Test Case': 'Ask(Bruce, Dick)',
'Boundary': np.exp(sem_model.results.log_boundary_probability)[1],
'New Event': 1-sem_model.results.post[1, 0]
}
)
print np.exp(sem_model.results.log_boundary_probability)
print sem_model.results.e_hat
# -
plot_results(sem_model.results, 'Ask(Tom,Charan) Ask(Tom,Charan)'.split(' '))
# plt.savefig('generalization_test_case_d.png', dpi=300, bbox_inches='tight')
# # All test cases
results = pd.DataFrame(results)
results = results.loc[[0, 2, 1, 3, 4], ]
results.index = range(5)
results
x_labels = results['Test Case'].copy()
with sns.plotting_context('paper', font_scale=0.8):
fig, ax = plt.subplots(1, 1, figsize=(6.875, 2.0), sharey=True)
w = 0.35
y = results.Boundary
x = [x0 - w/2. for x0 in results.index]
ax.bar(x, y, facecolor=[0.65, 0.65, 0.65], width=w, edgecolor='k', lw=1, label='Boundary')
ax.set_ylabel('Probability')
ax.set_xticks(results.index)
ax.set_xticklabels(x_labels, rotation=0)
y = results['New Event']
x = [x0 + w/2. for x0 in results.index]
ax.bar(x, y, facecolor=[0.35, 0.35, 0.35], width=w, edgecolor='k', lw=1, label='New Event')
ax.axhline(y=0.5, ls=':', c='k')
ax.set_ylim(0, 1)
plt.legend(loc='upper left')
sns.despine()
plt.savefig('generalization_demo_results.png', dpi=300)
# +
cases = (results['Test Case'] == 'Answer(Dick,Bruce)') | (results['Test Case'] == 'Chase(Dog,Cat)') | (results['Test Case'] == 'Answer(Charan,Tom)')
# results.loc[cases]
res_ = results.loc[cases].copy()
res_ = res_.loc[[0, 2, 1], :]
y = res_.Boundary
x = range(3)
with sns.plotting_context('talk'):
# fig, ax = plt.subplots(1, 1, figsize=(7.5, 2.0), sharey=True)
fig, ax = plt.subplots(1, 1)
ax.bar(x, y, facecolor=[0.65, 0.65, 0.65], edgecolor='k', lw=1, label='Boundary')
ax.set_ylabel('Probability')
ax.set_xticks(x)
ax.set_xticklabels(['Previously\nExperienced', 'Repeat\nStructure', 'Violate\nStructure'], rotation=0)
sns.despine()
plt.savefig('StructureTalk.png', dpi=300, bbox_inches='tight')
# -
res_
# # Reduced model: No clustering
# +
words['See'] = embed(d)
test_a2x = [
encode(words['See'], roles['Verb'])
+ encode(words['Dog'], roles['Agent'])
+ encode(words['Cat'], roles['Patient']),
encode(words['Chase'], roles['Verb'])
+ encode(words['Dog'], roles['Agent'])
+ encode(words['Cat'], roles['Patient']),
encode(words['Ask'], roles['Verb'])
+ encode(words['Tom'], roles['Agent'])
+ encode(words['Charan'], roles['Patient']),
encode(words['Answer'], roles['Verb'])
+ encode(words['Charan'], roles['Agent'])
+ encode(words['Tom'], roles['Patient']),
# encode(words['Ask'], roles['Verb'])
# + encode(words['Bruce'], roles['Agent'])
# + encode(words['Dick'], roles['Patient']),
# encode(words['Answer'], roles['Verb'])
# + encode(words['Dick'], roles['Agent'])
# + encode(words['Bruce'], roles['Patient'])
]
# Ask(Tom, Charan) -> Answer(Charan, Tom)
sem_model = SEM(**sem_kwargs)
sem_model.pretrain(_x, event_types, event_boundaries)
print sem_model.event_models[0].Sigma.mean()
sem_model._update_state(np.concatenate(test_a2x), 3)
sem_model.run(np.concatenate(test_a2x), compile_model=False)
new_results = [
{
'Model': 'SEM',
'PE': sem_model.results.pe[-1] / sem_model.results.pe[-2]
}
]
print sem_model.results.surprise
# -
sem_model.results.e_hat
sem_model.results.pe / np.linalg.norm(np.concatenate(test_a2x), axis=1).mean()
sem_model.results.pe / sem_model.results.pe[-2]
# +
## these are the original parameters
# # SEM parameters
# df0 = 100.0
# mode = 0.305
# scale0 = (mode * (df0 + 2)) / df0
# print scale0
# lmda = 1.0 # stickyness parameter
# alfa = 2. # concentration parameter
# prevent boundaries from being simulated by changing the CRP parameters
sem_kwargs_reduced_model = dict(lmda=10^6, alfa=10^-6, f_class=f_class, f_opts=f_opts)
# Ask(Tom, Charan) -> Answer(Charan, Tom)
sem_model = SEM(**sem_kwargs_reduced_model)
sem_model.pretrain(_x, event_types, event_boundaries)
sem_model._update_state(np.concatenate(test_a2x), 5)
sem_model.run(np.concatenate(test_a2x), compile_model=False)
new_results += [
{
'Model': 'Reduced',
'PE': sem_model.results.pe[-1] / sem_model.results.pe[-2]
}
]
# -
with sns.plotting_context('paper', font_scale=1.0):
sns.catplot(data=pd.DataFrame(new_results), x='Model', y='PE', color='grey',
height=2.0, kind='bar')
plt.ylim([0, 1])
plt.savefig('generalization_demo_reduced_model2.png', dpi=300)
sem_model.results.pe / sem_model.results.pe[-2]
| PaperSimulations/Segmentation - Generalizing Structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Robocupido
# language: python
# name: robocupido
# ---
# # Prepocessing and data cleansing
# This Jupyter Notebook is used, as its name indicates, to preproccess and to clean the data for the model of the project "Robocupido".
import pandas as pd
import numpy as np
df = pd.read_excel ('EnamóraTec_chido.xlsx')
df
cols = df.columns
cols
df1 = df[cols[5:41]]
df1
cols = df1.columns
def binarize(dataframe, column_name):
series = list(dataframe[column_name])
new_columns = []
# column = list()
for i in series:
x = list(i.split(', '))
for y in x:
if y not in new_columns:
new_columns.append(y)
new_dataframe = pd.DataFrame(columns = new_columns)
new_row = []
for i in series:
x = list(i.split(', '))
for z in new_columns:
if z in x:
new_row.append(True)
else:
new_row.append(False)
new_dataframe.loc[len(new_dataframe)] = new_row
new_row = []
column_names = list(dataframe.columns)
index = column_names.index(column_name)
dataframe = dataframe.drop(column_name, axis = 1)
column_names_newdf = list(new_dataframe.columns)
for x in column_names_newdf:
dataframe.insert(loc=index, column=x, value=new_dataframe[x])
index += 1
return dataframe
new_df = binarize(df1, cols[11])
new_df
binarized_df = binarize(new_df, cols[-21])
binarized_df
contact = df[cols[41:45]]
contact
binarized_df_final = pd.concat([binarized_df, contact], axis=1)
binarized_df_final
from pandas import ExcelWriter
writer = ExcelWriter('binarized_final.xlsx')
binarized_df_final.to_excel(writer,'Sheet1')
writer.save()
| Preprocessing and data cleansing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # ROS Tutorial Chapter 1 Evaluation
# This notebook starts the evaluation for the Chapter one Challenge
# +
import rospy
rospy.init_node('our_first_node')
from nav_msgs.msg import Odometry
from std_msgs.msg import String
data_list = []
publisher = rospy.Publisher('/eval', String, queue_size=10)
def odom_handler(data):
result = "goal not reached"
if(data.pose.pose.position.x<= -1.57 and data.pose.pose.position.y <= -1.63):
result = "goal reached"
publisher.publish(result)
rospy.Subscriber("/odom",Odometry, odom_handler)
# -
| src/ch1/ch1_eval.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
// + [markdown] dotnet_interactive={"language": "fsharp"}
// # Prototyping Trace Log API
// + dotnet_interactive={"language": "fsharp"}
#r "nuget:Microsoft.Diagnostics.Tracing.TraceEvent"
#r "nuget:XPlot.Plotly"
#r "nuget:XPlot.GoogleCharts"
// + dotnet_interactive={"language": "fsharp"}
// Open all the libraries.
open Microsoft.Diagnostics.Tracing;
open Microsoft.Diagnostics.Tracing.Etlx;
open Microsoft.Diagnostics.Tracing.Session;
open Microsoft.Diagnostics.Tracing.Parsers.Clr;
open XPlot.Plotly;
open XPlot.GoogleCharts;
// + dotnet_interactive={"language": "fsharp"}
let ETL_FILEPATH = @"C:\Users\mukun\OneDrive\Documents\CallstackShmuff.etl\CallstackShmuff.etl"
// -
// ## Charting Allocation Events Using Trace Log
// + dotnet_interactive={"language": "fsharp"}
let session = new TraceEventSession("TestSession", ETL_FILEPATH)
let traceLog = TraceLog.OpenOrConvert(ETL_FILEPATH)
let allocationAmountForDevenv =
traceLog.Events
|> Seq.filter(fun e -> e.ProcessName = "devenv" && e.EventName.Contains("GC/AllocationTick"))
|> Seq.take 100
|> Seq.map(fun e -> (e.TimeStampRelativeMSec, float(e.PayloadByName("AllocationAmount").ToString())))
let chart =
let options = Options( title = "Allocation Amount Over Time",
vAxes = [| Axis(title = "Allocation Amount"); Axis(title = "Timestamp"); |] )
allocationAmountForDevenv
|> Chart.Line
|> Chart.WithOptions options
|> Chart.Show
// -
// ## Call Stack Collection Using Trace Log
// + dotnet_interactive={"language": "fsharp"}
open Microsoft.Diagnostics.Symbols;
open Microsoft.Diagnostics.Tracing;
open Microsoft.Diagnostics.Tracing.Etlx;
open Microsoft.Diagnostics.Tracing.Parsers;
// + dotnet_interactive={"language": "fsharp"}
let session = new TraceEventSession("TestSession", ETL_FILEPATH)
let traceLog = TraceLog.OpenOrConvert(ETL_FILEPATH)
let loadSymbols : unit =
use symbolReader = new SymbolReader(TextWriter.Null, SymbolPath.SymbolPathFromEnvironment)
traceLog.Processes
|> Seq.filter(fun p -> p.Name = "GCRealTimeMon")
|> Seq.iter(fun proc -> (
printfn $"{proc} is being"
proc.LoadedModules |> Seq.where (fun m -> not (isNull m.ModuleFile))
|> Seq.iter (fun m -> traceLog.CodeAddresses.LookupSymbolsForModule(symbolReader, m.ModuleFile))
))
// + dotnet_interactive={"language": "fsharp"}
let processCallStack (callStack : TraceCallStack) : unit =
use symbolReader = new SymbolReader(TextWriter.Null, SymbolPath.SymbolPathFromEnvironment)
let printStackFrame (callStack : TraceCallStack) : unit =
traceLog.CodeAddresses.LookupSymbolsForModule(symbolReader, callStack.CodeAddress.ModuleFile)
printfn "%s!%s" callStack.CodeAddress.ModuleName callStack.CodeAddress.FullMethodName
let rec processFrame (callStack : TraceCallStack) : unit =
if isNull callStack then
()
else
printStackFrame callStack
processFrame callStack.Caller
processFrame callStack
let printGCAllocStacksForGCRealTimeMon : unit =
traceLog.Events
|> Seq.filter(fun e -> e.ProcessName = "GCRealTimeMon" && e.EventName = "GC/AllocationTick")
|> Seq.take 1
|> Seq.iter(fun e ->(
printfn "\n"; processCallStack (e.CallStack())))
| src/Prototypes/PrototypingTraceLog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Variational Auto Encoders
#
# - Reference: Adapted from the Keras example
# - Auto-Encoding Variational Bayes
# https://arxiv.org/abs/1312.6114
import tensorflow as tf
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
from tensorflow.keras.layers import Input, Dense, Lambda, Flatten, Reshape, Conv2D, Conv2DTranspose
from tensorflow.keras.models import Model
from tensorflow.keras import metrics
from tensorflow.keras.datasets import fashion_mnist
# -
# ## Fashion MNIST
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
plt.figure(figsize=(16, 8))
for i in range(0, 18):
plt.subplot(3, 6, i + 1)
plt.imshow(x_train[i], cmap="gray")
plt.axis("off")
plt.show()
y_train[0:10]
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
# ## Standard full-connected VAE model
#
# Let's define a VAE model with fully connected MLPs for the encoder and decoder networks.
x_train_standard = x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test_standard = x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
x_train_standard.shape, x_test_standard.shape
# ### Encoder
# +
original_dim = 784
latent_dim = 2
intermediate_dim = 256
def make_encoder(original_dim, intermediate_dim, latent_dim):
x = Input(shape=(original_dim,))
hidden = Dense(intermediate_dim, activation='relu')(x)
z_mean = Dense(latent_dim)(hidden)
z_log_var = Dense(latent_dim)(hidden)
return Model(inputs=x, outputs=[z_mean, z_log_var],
name="mlp_encoder")
encoder = make_encoder(original_dim, intermediate_dim, latent_dim)
# -
# ### The VAE stochastic latent variable
#
# <img src="./images/vae_3.png" width="600px" />
#
# We use the reparametrization trick to define a random variable z that is conditioned on the input image x as follows:
#
# $$ z \sim \mathcal{N}(\mu_z(x), \sigma_z(x)) $$
#
# The reparametrization tricks defines $z$ has follows:
#
# $$ z = \mu_z(x) + \sigma_z(x) \cdot \epsilon$$
#
# with:
#
# $$ \epsilon \sim \mathcal{N}(0, 1) $$
#
# This way the dependency to between $z$ and $x$ is deterministic and differentiable. The randomness of $z$ only stems from $\epsilon$ only for a given $x$.
#
# Note that in practice the output of the encoder network parameterizes $log(\sigma^2_z(x)$ instead of $\sigma_z(x)$. Taking the exponential of $log(\sigma^2_z(x)$ ensures the positivity of the standard deviation from the raw output of the network:
# +
def sampling_func(inputs):
z_mean, z_log_var = inputs
batch_size = tf.shape(z_mean)[0]
epsilon = tf.random.normal(shape=(batch_size, latent_dim),
mean=0., stddev=1.)
return z_mean + tf.exp(z_log_var / 2) * epsilon
sampling_layer = Lambda(sampling_func, output_shape=(latent_dim,),
name="latent_sampler")
# -
# ### Decoder
# +
def make_decoder(latent_dim, intermediate_dim, original_dim):
decoder_input = Input(shape=(latent_dim,))
x = Dense(intermediate_dim, activation='relu')(decoder_input)
x = Dense(original_dim, activation='sigmoid')(x)
return Model(decoder_input, x, name="mlp_decoder")
decoder = make_decoder(latent_dim, intermediate_dim, original_dim)
# -
# By default the decoder outputs has random weights and output noise:
random_z_from_prior = np.random.normal(loc=0, scale=1, size=(1, latent_dim))
generated = decoder.predict(random_z_from_prior)
plt.imshow(generated.reshape(28, 28), cmap=plt.cm.gray)
plt.axis('off');
# The generated image is completely univariate noise: there is no apparent spatial depenedencies between the pixel values. This reflects the lack of prior structure in the randomly initialized fully-connected decoder network.
#
#
# Let's now the plug the encoder and decoder via the stochastic latent variable $z$ to get the full VAE architecture. The loss function is the negative ELBO of the variational inference problem:
# +
def make_vae(input_shape, encoder, decoder, sampling_layer):
# Build de model architecture by assembling the encoder,
# stochastic latent variable and decoder:
x = Input(shape=input_shape, name="input")
z_mean, z_log_var = encoder(x)
z = sampling_layer([z_mean, z_log_var])
x_decoded_mean = decoder(z)
vae = Model(x, x_decoded_mean)
# Define the VAE loss
xent_loss = original_dim * metrics.binary_crossentropy(
Flatten()(x), Flatten()(x_decoded_mean))
kl_loss = - 0.5 * tf.reduce_sum(1 + z_log_var - tf.square(z_mean) - tf.exp(z_log_var), axis=-1)
vae_loss = tf.reduce_mean(xent_loss + kl_loss)
vae.add_loss(vae_loss)
vae.compile(optimizer='adam')
return vae
vae = make_vae((original_dim,), encoder, decoder,
sampling_layer=sampling_layer)
vae.summary()
# -
vae.fit(x_train_standard, epochs=50, batch_size=100,
validation_data=(x_test_standard, None))
# +
# vae.save_weights("standard_weights.h5")
# -
vae.load_weights("standard_weights.h5")
# Note that the model has not yet converged even after 50 epochs. Furthermore it's is not overfitting significantly either. We chose a very low value for the latent dimension. It is likely that using the higher dimensional space could lead to a model either to optimize that would better fit the training set.
#
# By sampling a random latent vector from the prior distribution and feeding it to the decoder we can effectively sample from the image model trained by the VAE:
random_z_from_prior = np.random.normal(size=(1, latent_dim)).astype("float32")
generated = decoder(random_z_from_prior).numpy()
plt.imshow(generated.reshape(28, 28), cmap=plt.cm.gray)
plt.axis('off');
# Use `Ctrl-Enter` several times to sample from various random locations in the 2D latent space.
#
# The generated pictures are blurry but capture of the global organization of pixels required to represent samples from the 10 fashion item categories. The spatial structure has been learned and is only present in the decoder weights.
# ### 2D plot of the image classes in the latent space
#
# We can also use the encoder to set the visualize the distribution of the test set in the 2D latent space of the VAE model. In the following the colors show the true class labels from the test samples.
#
# Note that the VAE is an unsupervised model: it did not use any label information during training. However we can observe that the 2D latent space is largely structured around the categories of images used in the training set.
id_to_labels = {0: "T-shirt/top", 1: "Trouser", 2: "Pullover", 3: "Dress", 4: "Coat",
5: "Sandal", 6: "Shirt", 7: "Sneaker", 8: "Bag", 9: "Ankle boot"}
x_test_encoded, x_test_encoded_log_var = encoder(x_test_standard)
plt.figure(figsize=(7, 6))
plt.scatter(x_test_encoded[:, 0], x_test_encoded[:, 1], c=y_test,
cmap=plt.cm.tab10)
cb = plt.colorbar()
cb.set_ticks(list(id_to_labels.keys()))
cb.set_ticklabels(list(id_to_labels.values()))
cb.update_ticks()
plt.show()
# **Exercises**
#
# - One can see that the class labels 5, 7 and 9 are grouped in a cluster of the latent space. Use matplotlib to display some samples from each of those 3 classes and discover why they have been grouped together by the VAE model.
#
# - Similarly: can you qualitatively explain with matplotlib why class 0, 4 and 6 seem to be hard to disentangle in this 2D latent space discovered by the VAE model?
#
# - One can observe that the global 2D shape of the encoded dataset is approximately spherical with values with a maximum radius of size 3. Where can you explain where the shape of this marginal latent distribution come from?
# +
# # %load solutions/class_5_7_9.py
# +
# # %load solutions/class_0_4_6.py
# +
# # %load solutions/shape_marginal_latent_distribution.py
# -
# ### 2D panel view of samples from the VAE manifold
#
# The following linearly spaced coordinates on the unit square were transformed through the inverse CDF (ppf) of the Gaussian to produce values of the latent variables z. This makes it possible to use a square arangement of panels that spans the gaussian prior of the latent space.
# +
n = 15 # figure with 15x15 panels
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
grid_x = norm.ppf(np.linspace(0.05, 0.95, n)).astype(np.float32)
grid_y = norm.ppf(np.linspace(0.05, 0.95, n)).astype(np.float32)
for i, yi in enumerate(grid_x):
for j, xi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
x_decoded = decoder(z_sample).numpy()
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(10, 10))
plt.imshow(figure, cmap='Greys_r')
plt.show()
# -
# ## Anomaly detection
#
# Let's rebuild a new VAE which encodes 9 of the 10 classes, and see if we can build a measure that shows wether the data is an anomaly
# We'll call standard classes the first 9 classes, and anomalies the last class (class n°9, which is "ankle boots")
valid_indexes_train = y_train != 9
valid_indexes_test = y_test != 9
x_train_9 = x_train[valid_indexes_train]
x_test_9 = x_test[valid_indexes_test]
x_train_standard_9 = x_train_9.reshape((len(x_train_9), np.prod(x_train_9.shape[1:])))
x_test_standard_9 = x_test_9.reshape((len(x_test_9), np.prod(x_test_9.shape[1:])))
print(x_train_standard_9.shape, x_test_standard_9.shape)
anomalies_indexes = y_test == 9
anomalies = x_test_standard[anomalies_indexes]
# +
# rebuild a new encoder, decoder, and train them on the limited dataset
encoder = make_encoder(original_dim, intermediate_dim, latent_dim)
decoder = make_decoder(latent_dim, intermediate_dim, original_dim)
vae_9 = make_vae((original_dim,), encoder, decoder,
sampling_layer=sampling_layer)
vae_9.fit(x_train_standard_9, epochs=50, batch_size=100,
validation_data=(x_test_standard_9, None))
# +
# vae_9.save_weights("standard_weights_9.h5")
# -
vae_9.load_weights("standard_weights_9.h5")
# +
# For simplicity, we will do our sampling with numpy not with Keras or tensorflow
def sampling_func_numpy(inputs):
z_mean, z_log_var = inputs
batch_size = np.shape(z_mean)[0]
epsilon = np.random.normal(size=(batch_size, latent_dim),
loc=0., scale=1.).astype("float32")
return z_mean + np.exp(z_log_var / 2) * epsilon
# Compute the reconstruction error: encode, sample, then decode.
# To ensure we get a stable result, we'll run the sampling nb_sampling times
def compute_reconstruction_error(img, nb_sampling=10):
if len(img.shape) == 1:
img = np.expand_dims(img, 0)
batch_size = np.shape(img)[0]
img_encoded_mean_and_var = encoder(img)
img_encoded_samples = [sampling_func_numpy(img_encoded_mean_and_var)
for x in range(nb_sampling)]
# stack all samples
img_encoded_samples = np.vstack(img_encoded_samples)
reconstructed_samples = decoder(img_encoded_samples).numpy()
# unstack all samples
split_samples = reconstructed_samples.reshape(nb_sampling, batch_size, img.shape[-1])
errors = np.linalg.norm(split_samples - img, axis=-1)
return np.mean(errors, axis=0)
# -
errors_test = compute_reconstruction_error(x_test_standard_9)
errors_anomalies = compute_reconstruction_error(anomalies)
noise = np.random.uniform(size=(1000, 784), low=0.0, high=1.0)
errors_random = compute_reconstruction_error(noise.astype(np.float32))
# +
# most anomalous in test set
indexes = np.argsort(errors_test)[-18:]
plt.figure(figsize=(16, 8))
for i in range(0, 18):
plt.subplot(3, 6, i + 1)
plt.imshow(x_test_9[indexes][i], cmap="gray")
plt.axis("off")
plt.show()
# It shows weird shaped tops, or very complex shoes which are difficult to reconstruct
# +
# most normal in anomalies test set
indexes = np.argsort(errors_anomalies)[0:18]
plt.figure(figsize=(16, 8))
for i in range(0, 18):
plt.subplot(3, 6, i + 1)
plt.imshow(x_test[anomalies_indexes][indexes][i], cmap="gray")
plt.axis("off")
plt.show()
# Indeed most of them do not look like ankle boot (they could belong to other shoes categories)!
# +
# most anomalous in anomalies test set
indexes = np.argsort(errors_anomalies)[-18:]
plt.figure(figsize=(16, 8))
for i in range(0, 18):
plt.subplot(3, 6, i + 1)
plt.imshow(x_test[anomalies_indexes][indexes][i], cmap="gray")
plt.axis("off")
plt.show()
# -
# ### Is this method a good anomaly detection method?
#
# Let's compare the distribution of reconstruction errors from
# - standard test set images
# - class 9 images
# - random noise
#
# What can you interpret from this graph?
fig = plt.figure()
ax = fig.add_subplot(111)
a1 = ax.hist(np.random.choice(errors_test, 1000, replace=False).tolist(), color="blue", alpha=0.5,)
a2 = ax.hist(errors_anomalies.tolist(), color="red", alpha=0.5)
a3 = ax.hist(errors_random.tolist(), color="green", alpha=0.5)
plt.legend(('standard (classes 0 to 8)', 'ankle boots (class 9)', 'random pixels (white noise)'))
plt.show()
# ## Convolutional Variational Auto Encoder
x_train_conv = np.expand_dims(x_train, -1)
x_test_conv = np.expand_dims(x_test, -1)
x_train_conv.shape, x_test_conv.shape
# **Exercise**: write an encoder that uses a series of convolutional layers, with maxpooling or strided convolutions and Batch norm to encode the 2D, gray-level images into 2D latent vectors:
# +
from tensorflow.keras.layers import BatchNormalization
img_rows, img_cols, img_chns = 28, 28, 1
filters = 32
kernel_size = 3
intermediate_dim = 128
latent_dim = 2
def make_conv_encoder(img_rows, img_cols, img_chns,
latent_dim, intermediate_dim):
inp = x = Input(shape=(img_rows, img_cols, img_chns))
# TODO: write me!
return Model(inputs=inp, outputs=[z_mean, z_log_var],
name='convolutional_encoder')
conv_encoder = make_conv_encoder(img_rows, img_cols, img_chns,
latent_dim, intermediate_dim)
print(conv_encoder.summary())
conv_encoder.predict(x_train_conv[:1])
# +
# # %load solutions/conv_encoder.py
# -
# The stochastic latent variable is the same as for the fully-connected model.
sampling_layer = Lambda(sampling_func, output_shape=(latent_dim,),
name="latent_sampler")
# ## Decoder
#
# The decoder is also convolutional but instead of downsampling the spatial dimensions from (28, 28) to 2 latent dimensions, it starts from the latent space to upsample a (28, 28) dimensions using strided `Conv2DTranspose` layers.
#
# Here again BatchNormalization layers are inserted after the convolution to make optimization converge faster.
# +
def make_conv_decoder(latent_dim, intermediate_dim, original_dim,
spatial_size=7, filters=16):
decoder_input = Input(shape=(latent_dim,))
x = Dense(intermediate_dim, activation='relu')(decoder_input)
x = Dense(filters * spatial_size * spatial_size, activation='relu')(x)
x = Reshape((spatial_size, spatial_size, filters))(x)
# First up-sampling:
x = Conv2DTranspose(filters,
kernel_size=3,
padding='same',
strides=(2, 2),
activation='relu')(x)
x = BatchNormalization()(x)
x = Conv2DTranspose(filters,
kernel_size=3,
padding='same',
strides=1,
activation='relu')(x)
x = BatchNormalization()(x)
# Second up-sampling:
x = Conv2DTranspose(filters,
kernel_size=3,
strides=(2, 2),
padding='valid',
activation='relu')(x)
x = BatchNormalization()(x)
# Ouput 1 channel of gray pixels values between 0 and 1:
x = Conv2D(1, kernel_size=2, padding='valid',
activation='sigmoid')(x)
return Model(decoder_input, x, name='convolutional_decoder')
conv_decoder = make_conv_decoder(latent_dim, intermediate_dim, original_dim,
spatial_size=7, filters=filters)
print(conv_decoder.summary())
# -
generated = conv_decoder.predict(np.random.normal(size=(1, latent_dim)))
plt.imshow(generated.reshape(28, 28), cmap=plt.cm.gray)
plt.axis('off');
# This new decoder encodes some a priori knowledge on the local dependencies between pixel values in the "deconv" architectures. Depending on the randomly initialized weights, the generated images can show some local spatial structure.
#
# Try to re-execute the above two cells several times to try to see the kind of local structure that stem from the "deconv" architecture it-self for different random initializations of the weights.
#
#
# Again, let's now plug everything to together to get convolutional version of a full VAE model:
input_shape = (img_rows, img_cols, img_chns)
vae = make_vae(input_shape, conv_encoder, conv_decoder,
sampling_layer)
vae.summary()
vae.fit(x_train_conv, epochs=15, batch_size=100,
validation_data=(x_test_conv, None))
# +
# vae.save_weights("convolutional_weights.h5")
# -
vae.load_weights("convolutional_weights.h5")
generated = conv_decoder.predict(np.random.normal(size=(1, latent_dim)))
plt.imshow(generated.reshape(28, 28), cmap=plt.cm.gray)
plt.axis('off');
# ### 2D plot of the image classes in the latent space
#
# We find again a similar organization of the latent space. Compared to the fully-connected VAE space, the different class labels seem slightly better separated. This could be a consequence of the slightly better fit we obtain from the convolutional models.
x_test_encoded, _ = conv_encoder(x_test_conv)
plt.figure(figsize=(7, 6))
plt.scatter(x_test_encoded[:, 0], x_test_encoded[:, 1], c=y_test,
cmap=plt.cm.tab10)
cb = plt.colorbar()
cb.set_ticks(list(id_to_labels.keys()))
cb.set_ticklabels(list(id_to_labels.values()))
cb.update_ticks()
plt.show()
# ### 2D panel view of samples from the VAE manifold
#
# The following linearly spaced coordinates on the unit square were transformed through the inverse CDF (ppf) of the Gaussian to produce values of the latent variables z. This makes it possible to use a square arangement of panels that spans the gaussian prior of the latent space.
# +
n = 15 # figure with 15x15 panels
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
grid_x = norm.ppf(np.linspace(0.05, 0.95, n))
grid_y = norm.ppf(np.linspace(0.05, 0.95, n))
for i, yi in enumerate(grid_x):
for j, xi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
x_decoded = conv_decoder.predict(z_sample)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(10, 10))
plt.imshow(figure, cmap='Greys_r')
plt.show()
# -
# ### Semi-supervised learning
#
# Let's reuse our encoder trained on many unlabeled samples to design a supervised model that can only use supervision from a small subset of samples with labels.
#
# To keep things simple we will just build a small supervised model on top of the latent representation defined by our encoder.
#
# We assume that we only have access to a small labeled subset with 50 examples per class (instead of 5000 examples per class in the full Fashion MNIST training set):
# +
rng = np.random.RandomState(42)
small_x_train = []
small_y_train = []
num_per_class = 50
for c in range(10):
class_mask = np.where(y_train==c)[0]
idx = rng.choice(class_mask, size=num_per_class, replace=False)
small_x_train += [x_train_conv[idx]]
small_y_train += [c] * num_per_class
small_x_train = np.vstack(small_x_train)
small_y_train = np.array(small_y_train)
# reshuffle our small dataset
perm = rng.permutation(range(small_y_train.shape[0]))
small_x_train = small_x_train[perm]
small_y_train = small_y_train[perm]
small_x_train.shape, small_y_train.shape
# -
# **Exercise:**
#
# - Use `conv_encoder` to project `small_x_train` into the latent space;
# - Define a small supervised 10-class classification network and use `small_y_train` to train it;
# - What test accuracy can you reach? What is the chance level?
# - Suggest what could be changed to improve the quality of our classification on this small labeled dataset.
# +
# TODO: implement me!
# define `small_x_train_encoded` for in the input training data
# define a model named `mdl` with its layers and its loss function.
# +
# # %load solutions/small_classifier.py
# -
print(mdl.summary())
mdl.fit(small_x_train_encoded, small_y_train,
epochs=30, validation_data=[x_test_encoded, y_test])
# +
from sklearn.metrics import confusion_matrix
y_pred = mdl.predict(x_test_encoded).argmax(axis=-1)
cnf_matrix = confusion_matrix(y_test, y_pred)
print(cnf_matrix)
# +
import itertools
def plot_confusion_matrix(cm, classes, title='Confusion matrix',
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], 'd'),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
class_names = [name for id, name in sorted(id_to_labels.items())]
plot_confusion_matrix(cnf_matrix, classes=class_names)
# -
# ### Going further
#
# - Retrain the conv/deconv VAE model with a latent dim of 30 instead of 2. Generating the 2D manifold panels plots is no longer possible. However this richer latent space should make it possible to reach a much better test likelihood bound and generate higher quality images.
#
# - Adapt the convolutional architecture to retrain the model on the labeled faces in the wild (LFW) dataset instead (GPU needed).
| labs/10_unsupervised_generative_models/Variational_AutoEncoders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("datascience.csv", encoding='gb18030')
df.head()
df.shape
import jieba
def chinese_word_cut(mytext):
return " ".join(jieba.cut(mytext))
df["content_cutted"] = df.content.apply(chinese_word_cut)
df.content_cutted.head()
#执行时间比较长,需等待
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
n_features = 1000
tf_vectorizer = CountVectorizer(strip_accents = 'unicode',
max_features=n_features,
stop_words=("这个","就是","的","www","com","http","可能","一些","什么","他们","没有","自己","如果","很多","不是","但是"
,"现在","这样","因为","已经","时候","还是","可是",),
max_df = 0.5,
min_df = 10)
tf = tf_vectorizer.fit_transform(df.content_cutted)
from sklearn.decomposition import LatentDirichletAllocation
n_topics = 5
lda = LatentDirichletAllocation(n_topics=n_topics, max_iter=50,
learning_method='online',
learning_offset=50.,
random_state=0)
lda.fit(tf)
#执行时间比较长,需等待
def print_top_words(model, feature_names, n_top_words):
for topic_idx, topic in enumerate(model.components_):
print("Topic #%d:" % topic_idx)
print(" ".join([feature_names[i]
for i in topic.argsort()[:-n_top_words - 1:-1]]))
print()
n_top_words = 20
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, n_top_words)
import pyLDAvis
import pyLDAvis.sklearn
pyLDAvis.enable_notebook()
pyLDAvis.sklearn.prepare(lda, tf, tf_vectorizer)
# +
n_topics = 10
lda = LatentDirichletAllocation(n_topics=n_topics, max_iter=50,
learning_method='online',
learning_offset=50.,
random_state=0)
#运算时间长,需等待
lda.fit(tf)
print_top_words(lda, tf_feature_names, n_top_words)
pyLDAvis.sklearn.prepare(lda, tf, tf_vectorizer)
# -
| topic-lda/topic-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import pandas as pd
# +
#Got Open Weather API and key to use with the URL. Download and made request
#pull out the data that will be used such as temperature in imperial. The
#original temperature came in standard and was converted to imperial units.
#Using for loops, temperature was gathered and the data were appended.
temps = []
url = "http://api.openweathermap.org/data/2.5/forecast?zip=20850,us&units=imperial&appid=d7bb4169bffbbdccbe072168d0db38e7"
page = requests.get(url)
weather_data = page.json()
a = weather_data['list']
for i in range(0, weather_data['cnt']):
b = a[i]
c = b['main']
d = c['temp']
temps.append(d)
# +
#Continued getting information using the for lopps and appending information
#on the date and time
dt_txts = []
for i in range(0, weather_data['cnt']):
b = a[i]
f = b['dt_txt']
dt_txts.append(f)
# -
# +
#For loops was used to gather the description and
#was appended to descriptions.
descriptions = []
for i in range(0, weather_data['cnt']):
b = a[i]
h = b['weather'][0]
description = h['description']
descriptions.append(description)
# -
#Created weather_data2 a pandas data frame containing date, temperature
#and description.
weather_data2 = pd.DataFrame({'Date':dt_txts, "Temperature": temps,
"Description": descriptions})
weather_data2
#Information about the weather_data2 needed to be gathered to know the data
#type. Date and Time are objects that needed to be separated
print(weather_data2.info())
weather_data2["Date"] = pd.to_datetime(weather_data2['Date'])
weather_data2.info()
#After converting the Date and Time to data type datetime. Date and Time
#is separated from one another.
weather_data2["Time"]=weather_data2["Date"].dt.time
weather_data2['Date']=weather_data2['Date'].dt.date
weather_data2
# +
#This descibes the temperature during the 5 day period whereby temperature
#was gathered every 3 hours. It is gathered that there are 40 index, the
#average temperature is 59.684250 while the spread of dispersion is 7.309277.
weather_data2['Temperature'].describe()
# -
#The temperature was sorted from coldest to warmest
sorted_weather_data2 = weather_data2.sort_values(by='Temperature')
sorted_weather_data2
#These are the coldest temperature for the week
sorted_weather_data2.head()
#These are the warmest temperature for the week
sorted_weather_data2.tail()
#Using matplotlib, a box plot was made. The temperature is stable. Since
#there are no outliers. The smallest data value is within the lower fence
#while the largest data value stayed within the upper fence. This box plot
#is slightly right skewed.
import matplotlib
weather_data2.Temperature.plot(kind = 'box')
#The temperature was stored in a csv filed called weather every 3 hours
weather_data2.to_csv('data.csv', encoding='utf-8', index=False)
| Assignment 2/API Open Weather Map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Appendix B: Contribution/Bugs/Features
# Please have a look at the [Contribution Guidelines](https://github.com/stammler/simframe/blob/master/.github/CONTRIBUTING.md).
# ## Contributing to `simframe`
# To contribute code please open a new pull request and describe the changes to the software your pull request introduces.
#
# Please note, that we want to achieve a **code coverage of 100%**. Any addition to the software must therefore also come with unit tests. Additional features must also be described in the documentation.
# ## Bug Report
# If you encountered a bug in `simframe`, please open a new [bug report issue](https://github.com/stammler/simframe/issues/new?template=bug_report.md&title=%5BBUG%5D+Descriptive+title+of+the+bug+report) and describe the bug, the expected behavior, and steps how to reproduce the bug.
# ## Feature Request
# If you have an idea of a new feature, that is missing in `simframe`, or if you want to suggest an improvement, please open a new [feature request issue](https://github.com/stammler/simframe/issues/new?template=feature_request.md&title=%5BFEATURE%5D+Descriptive+title+of+the+feature+request).
| examples/B_contrib_bug_feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <a id="top"></a>Intersecting grids with shapes
#
# _Note: This feature requires the shapely module (which is not a dependency of flopy) so must be installed by the user._
#
# This notebook shows the grid intersection functionality in flopy. The intersection methods are available through the GridIntersect object. A flopy model grid is passed to instantiate the object. Then the modelgrid can be intersected with Points, LineStrings and Polygons through the different intersect methods. There are two intersection modes:
# - the first (default mode) is accessed by passing `method='strtree'` to `GridIntersect` and converts the modelgrid to a list of shapes that are sorted into an STR-tree to allow fast spatial queries. This works on structured and vertex grids.
# - the second only works on structured grids and is accessed by passing `method='structured'` to `GridIntersect`. These methods use information from the structured grid to limit the search space for intersections and are generally faster.
#
# This notebook showcases the functionality of the GridIntersect class.
#
#
# ### Table of Contents
# - [GridIntersect Class](#gridclass)
# - [Rectangular regular grid](#rectgrid)
# - [Polygon with regular grid](#rectgrid.1)
# - [Polyline with regular grid](#rectgrid.2)
# - [MultiPoint with regular grid](#rectgrid.3)
# - [Triangular grid](#trigrid)
# - [Polygon with triangular grid](#trigrid.1)
# - [Polyline with triangular grid](#trigrid.2)
# - [MultiPoint with triangular grid](#trigrid.3)
# - [Tests](#tests)
# - [Timings](#timings)
# Import some stuff
# +
import sys
import os
import platform
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# run installed version of flopy or add local path
try:
import flopy
import flopy.discretization as fgrid
import flopy.plot as fplot
from flopy.utils.triangle import Triangle as Triangle
from flopy.utils.gridintersect import GridIntersect
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
import flopy.discretization as fgrid
import flopy.plot as fplot
from flopy.utils.triangle import Triangle as Triangle
from flopy.utils.gridintersect import GridIntersect
import shapely
from shapely.geometry import Polygon, Point, LineString, MultiLineString, MultiPoint, MultiPolygon
from shapely.strtree import STRtree
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('matplotlib version: {}'.format(mpl.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
triangle_exe = None
# ## <a id="gridclass"></a>[GridIntersect Class](#top)
#
# This GridIntersect class takes a flopy.mfgrid and by default converts it to a list of Shapely geometries and builds a STRTree which can be used to efficiently query the grid to perform intersections. If the method is set to 'structured', the STR-tree is not built and different intersection methods are applied (written by <NAME>). The following methods are available:
# - ` _rect_grid_to_shape_list`: convert rectangular (structured) modflow grid to list of shapely geometries
# - `_sort_strtree_result`: sort STRTree by cellid (to ensure lowest cellid is returned when shapes intersect with multiple grid cells)
# - `_usg_grid_to_shape_list`: not yet implemented, convert unstructured grid to list of shapely geometries
# - `_vtx_grid_to_shape_list`: convert vertex modflow grid to list of shapely geometries
# - `_intersect_point_shapely`: intersect Shapely point with grid
# - `_intersect_polygon_shapely`: intersect Shapely Polygon with grid
# - `_intersect_linestring_shapely`: intersect Shapely LineString with grid
# - `_intersect_point_structured`: intersect Shapely point with grid, using optimized search for structured grids
# - `_intersect_polygon_structured`: intersect Shapely Polygon with grid, using optimized search for structured grids
# - `_intersect_rectangle_structured`: intersect rectangle with grid to get intersecting node ids
# - `_intersect_linestring_structured`: intersect Shapely LineString with structured grid, using optimized search for structured grids
# - `_check_adjacent_cells_intersecting_line`: helper function to check adjacent cells in a structured grid for line intersections
# - `_get_nodes_intersecting_linestring`: helper function to follow linestring through structured grid
# - `intersect_point`: intersect point with grid, method depends on whether 'structured' or 'strtree' is passed at intialization.
# - `intersect_linestring`: intersect linestring with grid, method depends on whether 'structured' or 'strtree' is passed at intialization.
# - `intersect_polygon`: intersect polygon with grid, method depends on whether 'structured' or 'strtree' is passed at intialization.
# - `plot_point`: plot intersect result for point
# - `plot_polygon`: plot intersect result for polygons
# - `plot_polyline`: plot intersect result for linestrings
# ## <a id="rectgrid"></a>[Rectangular regular grid](#top)
delc = 10*np.ones(10, dtype=np.float)
delr = 10*np.ones(10, dtype=np.float)
xoff = 0.
yoff = 0.
angrot = 0.
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None, xoff=xoff, yoff=yoff, angrot=angrot)
sgr.plot()
# ### <a id="rectgrid.1"></a>[Polygon with regular grid](#top)
# Polygon to intersect with:
p = Polygon(shell=[(15, 15), (20, 50), (35, 80.), (80, 50), (80, 40), (40, 5), (15, 12)],
holes=[[(25, 25), (25, 45), (45, 45), (45, 25)]])
# Create GridIntersect class
ix = GridIntersect(sgr)
# Do the intersect operation for a polygon
result = ix.intersect_polygon(p)
# %timeit ix.intersect_polygon(p)
# The results are returned as a numpy.recarray containing several fields based on the intersection performed. An explanation of the data in each of the possible fields is given below:
# - cellids: contains the cell ids of the intersected grid cells
# - vertices: contains the vertices of the intersected shape
# - areas: contains the area of the polygon in that grid cell (only for polygons)
# - lenghts: contains the length of the linestring in that grid cell (only for linestrings)
# - ixshapes: contains the shapely object representing the intersected shape (useful for plotting the result)
#
# Looking at the data for the polygon intersection (convert to pandas.DataFrame for prettier formatting)
# pd.DataFrame(result)
result
# Visualizing the results
# +
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
sgr.plot(ax=ax)
ix.plot_polygon(result, ax=ax)
# only cells that intersect with shape
for irow, icol in result.cellids:
h2, = ax.plot(sgr.xcellcenters[0, icol], sgr.ycellcenters[irow, 0], "kx", label="centroids of intersected gridcells")
ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# Alternatively, the intersection can be calculated using special methods optimized for structured grids. Access these methods by instantiating the GridIntersect class with the `method="structured"` keyword argument.
ixs = GridIntersect(sgr, method="structured")
# pd.DataFrame(ixs.intersect_polygon(p)).head()
ixs.intersect_polygon(p)
# ### <a id="rectgrid.2"></a>[Polyline with regular grid](#top)
# MultiLineString to intersect with:
ls1 = LineString([(95, 105), (30, 50)])
ls2 = LineString([(30, 50), (90, 22)])
ls3 = LineString([(90, 22), (0, 0)])
mls = MultiLineString(lines=[ls1, ls2, ls3])
# %timeit ix.intersect_linestring(mls)
result = ix.intersect_linestring(mls)
# +
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
sgr.plot(ax=ax)
ix.plot_linestring(result, ax=ax)
for irow, icol in result.cellids:
h2, = ax.plot(sgr.xcellcenters[0, icol], sgr.ycellcenters[irow, 0], "kx", label="centroids of intersected gridcells")
ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# Same as before, the intersect for structured grids can also be performed with a different method optimized for structured grids
ixs = GridIntersect(sgr, method="structured")
# pd.DataFrame(ixs.intersect_linestring(mls)).head()
ixs.intersect_linestring(mls)
# ### [MultiPoint with regular grid](#top)<a id="rectgrid.3"></a>
#
# MultiPoint to intersect with
mp = MultiPoint(points=[Point(50.0, 0.0), Point(45., 45.),
Point(10., 10.), Point(150., 100.)])
result = ix.intersect_point(mp)
# +
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
sgr.plot(ax=ax)
ix.plot_point(result, ax=ax, s=50)
for irow, icol in result.cellids:
h2, = ax.plot(sgr.xcellcenters[0, icol], sgr.ycellcenters[irow, 0], "kx", label="centroids of intersected cells")
ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# Same as before, the intersect for structured grids can also be performed with a different method optimized for structured grids
ixs = GridIntersect(sgr, method="structured")
# pd.DataFrame(ixs.intersect_point(mp))
ixs.intersect_point(mp)
# ## <a id="trigrid"></a>[Triangular Grid](#top)
# + active=""
# maximum_area = 50.
# x0, x1, y0, y1 = sgr.extent
# domainpoly = [(x0, y0), (x0, y1), (x1, y1), (x1, y0)]
# tri = Triangle(maximum_area=maximum_area, angle=30, model_ws=".",
# exe_name=triangle_exe)
# tri.add_polygon(domainpoly)
# tri.build(verbose=False)
# + active=""
# cell2d = tri.get_cell2d()
# vertices = tri.get_vertices()
# tgr = fgrid.VertexGrid(vertices, cell2d)
# + active=""
# fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# pmv = fplot.PlotMapView(modelgrid=tgr)
# pmv.plot_grid(ax=ax)
# -
# ### <a id="trigrid.1"></a>[Polygon with triangular grid](#top)
# + active=""
# ix2 = GridIntersect(tgr)
# + active=""
# result = ix2.intersect_polygon(p)
# + active=""
# fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# pmv = fplot.PlotMapView(ax=ax, modelgrid=tgr)
# pmv.plot_grid()
# ix.plot_polygon(result, ax=ax)
#
# # only cells that intersect with shape
# for cellid in result.cellids:
# h2, = ax.plot(tgr.xcellcenters[cellid], tgr.ycellcenters[cellid], "kx", label="centroids of intersected gridcells")
#
# ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# ### <a id="trigrid.2"></a>[LineString with triangular grid](#top)
# + active=""
# result = ix2.intersect_linestring(mls)
# + active=""
# fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# pmv = fplot.PlotMapView(ax=ax, modelgrid=tgr)
# pmv.plot_grid()
# ix2.plot_linestring(result, ax=ax, lw=3)
#
# for cellid in result.cellids:
# h2, = ax.plot(tgr.xcellcenters[cellid], tgr.ycellcenters[cellid], "kx", label="centroids of intersected gridcells")
#
# ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# ### <a id="trigrid.3"></a>[MultiPoint with triangular grid](#top)
# + active=""
# result = ix2.intersect_point(mp)
# + active=""
# fig, ax = plt.subplots(1, 1, figsize=(8, 8))
# pmv = fplot.PlotMapView(ax=ax, modelgrid=tgr)
# pmv.plot_grid()
# ix2.plot_point(result, ax=ax)
#
# for cellid in result.cellids:
# h2, = ax.plot(tgr.xcellcenters[cellid], tgr.ycellcenters[cellid], "kx", label="return one intersecting grid cell per point")
#
# ax.legend([h2], [i.get_label() for i in [h2]], loc="best")
# -
# ## <a id="tests"></a>[Tests](#top)
# Tests are written for Points, LineStrings and Polygons for both rectangular (regular) grids, triangular grids, and rotated and offset regular grids.
# !pytest --cov-report term --cov gridintersect ../../autotest/t065_test_gridintersect.py
# ## <a id="timings"></a>[Timings](#top)
# Comparing performance for the different methods in a large grid. Some helper functions are defined below
# +
def ix_shapely_point(nrnc, npoints=100):
results = []
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr)
points = np.random.random((npoints, 2)) * nrnc
for p in [Point(x, y) for x, y in points]:
results.append(ix.intersect_point(p))
return np.concatenate(results, axis=0)
def ix_structured_point(nrnc, npoints=100):
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr, method="structured")
points = np.random.random((npoints, 2)) * nrnc
mp = MultiPoint(points=[Point(x, y) for x, y in points])
return ix.intersect_point(mp)
def ix_shapely_linestring(nrnc, ls=None):
if ls is None:
ls = LineString([(0, 0), (nrnc/3, nrnc)])
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr)
return ix.intersect_linestring(ls)
def ix_structured_linestring(nrnc, ls=None):
if ls is None:
ls = LineString([(0, 0), (nrnc/3, nrnc)])
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr, method="structured")
return ix.intersect_linestring(ls)
def ix_shapely_polygon(nrnc, p=Polygon([(10, 10), (540, 430), (730, 80), (250, 0)])):
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr)
return ix.intersect_polygon(p)
def ix_structured_polygon(nrnc, p=Polygon([(10, 10), (540, 430), (730, 80), (250, 0)])):
delc = np.ones(nrnc, dtype=np.float)
delr = np.ones(nrnc, dtype=np.float)
sgr = fgrid.StructuredGrid(delc, delr, top=None, botm=None)
ix = GridIntersect(sgr, method="structured")
return ix.intersect_polygon(p)
# -
# Below are some results of `%timeit` runs of some intersections on a 1000 x 1000 structured grid. For obvious reasons not having to build the STR-tree saves a significant amount of time for large grids (~ 15 seconds on my laptop).
# nrnc = 1000 # no rows and columns
nrnc = 10 # save time when testing notebook
# For point intersections, most of the time required by the shapely approach is needed to build the STR-tree (~15 s). Obviously, the pure numpy approach used in structured mode is unbeatable.
# %timeit -n 1 -r 1 ix_shapely_point(nrnc, npoints=100)
# %timeit ix_structured_point(nrnc, npoints=2)
# For linestrings, following the linestring through the grid (in structured mode) reduces the amount of intersection calls by a significant amount. This is where the downside of the STR-tree query is obvious. The bounding box of the linestring covers about one third of the grid. The query only reduces the search-space by 2/3 leaving ~333k cells to try to intersect with. On top of the building of the STR-tree the intersection calls take another ~15 seconds.
#
# (Cutting the linestring into pieces would probably improve performance.)
# %timeit -n 1 -r 1 ix_shapely_linestring(nrnc)
# %timeit ix_structured_linestring(nrnc)
# For Polygons the difference between structured mode and shapely mode is less obvious. Building the STR-tree (~15s) and doing the intersect (~20s) takes a little bit longer than performing the intersection in structured mode. However, note that intersecting with a second similarly sized polygon in shapely mode will only require ~20s, whereas in structured mode the required time will remain ~30 seconds.
#
# For repeated intersections with Polygons, the shapely method might be preferred over the structured method.
# %timeit -n 1 -r 1 ix_shapely_polygon(nrnc)
# %timeit -n 1 -r 1 ix_structured_polygon(nrnc)
| examples/Notebooks/flopy3_grid_intersection_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Just Use Linear Regression
import numpy as np
import matplotlib.pyplot as plt
# +
np.random.seed(500)
x = np.random.uniform(-3, 3, size=100)
X = x.reshape(-1, 1)
y = 0.5 * x + 3 + np.random.normal(0, 1, size=100)
plt.scatter(x, y)
# +
from sklearn.model_selection import train_test_split
np.random.seed(500)
X_train, X_test, y_train, y_test = train_test_split(X, y)
# +
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
def PolynomialRegression(degree):
return Pipeline([
("poly", PolynomialFeatures(degree=degree)),
("std_scaler", StandardScaler()),
("lin_reg", LinearRegression())
])
# +
from sklearn.metrics import mean_squared_error
poly_reg = PolynomialRegression(degree=20)
poly_reg.fit(X_train, y_train)
y_poly_predict = poly_reg.predict(X_test)
mean_squared_error(y_test, y_poly_predict)
# +
def plot_model(model):
X_plot = np.linspace(-3, 3, 100).reshape(100, 1)
y_plot = model.predict(X_plot)
plt.scatter(x, y)
plt.plot(X_plot[:,0], y_plot, color='r')
plt.axis([-3, 3, 0, 6])
plot_model(poly_reg)
# -
# # LASSO
# +
from sklearn.linear_model import Lasso
def LassoRegression(degree, alpha):
return Pipeline([
("poly", PolynomialFeatures(degree=degree)),
("std_scaler", StandardScaler()),
("lasso_reg", Lasso(alpha=alpha))
])
# +
lasso1_reg = LassoRegression(20, 0.01)
lasso1_reg.fit(X_train, y_train)
y1_predict = lasso1_reg.predict(X_test)
mean_squared_error(y_test, y1_predict)
# -
plot_model(lasso1_reg)
# +
lasso2_reg = LassoRegression(20, 0.1)
lasso2_reg.fit(X_train, y_train)
y2_predict = lasso2_reg.predict(X_test)
mean_squared_error(y_test, y2_predict)
# -
plot_model(lasso2_reg)
# +
lasso3_reg = LassoRegression(20, 1)
lasso3_reg.fit(X_train, y_train)
y3_predict = lasso3_reg.predict(X_test)
mean_squared_error(y_test, y3_predict)
# -
plot_model(lasso3_reg)
| 06Polynomial-Regression-and-Model-Generalization/07LASSO-Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
# # Semana 11
# 21/10/2021
# O exercício a seguir pretende servir como reforço e prática no tratamento básico de dataframes com Pandas.
#
# É importantte dominar bem o básico de leitura e análise dos dados de cada coluna, assim como entender como selecionar e manipular os dados.
# ## Base de dados do Tráfico de Africanos Escravizados
#
# > Exercícios inspirados no material de Melanie Walsh, que pode ser [acessado aqui](https://melaniewalsh.github.io/Intro-Cultural-Analytics/03-Data-Analysis/02-Pandas-Basics-Part2.html)
# Esse dataset foi baixado da [Base de dados Comércio Transatlântico de Escravos](https://www.slavevoyages.org/voyage/database), parte do projeto [Slave Voyages](https://www.slavevoyages.org).
#
# Possuiu dados de mais de trinta e seis mil viagens de navios negreiros entre 1514 e 1866.
#
# Para esse dataset, selecionei apenas algumas colunas, pois a base de dados original possui mais de uma dezena de colunas.
#
# Como vcs vão perceber, esse dataset possuiu variados tipos de dados. Em muitas dessas colunas, o valor é `NaN`, portanto, é importante tratar as colunas que serão analisadas.
# ### Objetivos:
#
# 1. Importar o Pandas
# 2. Ler o CSV (`CSVs/slave_trade_database.csv`) e criar um dataframe
# 3. Analisar a estrutura do dataframe (quantidades de linhas, colunas, nomes das colunas, tipos de dados
# 4. Renomear as colunas: substitua os espaços por `_`
# 5. Calcular as bandeiras mais frequentes
# 6. Crie um novo dataframe apenas com as viagens cujo navio possui 'Brazil' na bandeira
# 7. Contar os navios com a bandeira 'Brazil' que mais repetiram viagens
# 8. Criar um dataframe apenas com navios com porcentagem de mortalidade maior que 10%
| sem10-11_Python/exercicios_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Within repo
# +
import pandas as pd
train_file = ["mesos", "usergrid", "appceleratorstudio", "appceleratorstudio", "titanium", "aptanastudio", "mule", "mulestudio"]
test_file = ["usergrid", "mesos", "aptanastudio", "titanium", "appceleratorstudio", "titanium", "mulestudio", "mule"]
mae = [1.07, 1.14, 2.75, 1.99, 2.85, 3.41, 3.14, 2.31]
df_gpt = pd.DataFrame(data={"group": ["Within Repository" for i in range(8)],
"approach": ["Deep-SE" for i in range(8)],
"train_file": train_file,
"test_file": test_file,
"mae": mae})
df = pd.read_csv("./within_repo_abe0.csv")
df = df.append(df_gpt)
df.to_csv("./within_repo_abe0.csv", index=False)
# -
# ### Cross repo
# +
import pandas as pd
train_file = ["clover", "talendesb", "talenddataquality", "mule", "talenddataquality", "mulestudio", "appceleratorstudio", "appceleratorstudio"]
test_file = ["usergrid", "mesos", "aptanastudio", "titanium", "appceleratorstudio", "titanium", "mulestudio", "mule"]
mae = [1.57, 2.08, 5.37, 6.36, 5.55, 2.67, 4.24, 2.7]
df = pd.read_csv("./cross_repo_abe0.csv")
df_gpt = pd.DataFrame(data={"group": ["Cross Repository" for i in range(8)],
"approach": ["Deep-SE" for i in range(8)],
"train_file": train_file,
"test_file": test_file,
"mae": mae})
df = df.append(df_gpt)
df.to_csv("./cross_repo_abe0.csv", index=False)
# -
| abe0/ignore_process_csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# +
import kfp
from typing import NamedTuple
from google.cloud import aiplatform
from kfp import dsl
from kfp.v2 import compiler
from kfp.v2.dsl import component
from kfp.v2.google.client import AIPlatformClient
from kfp.v2.dsl import (Artifact, Dataset, Input, InputPath, Model, Output,
OutputPath, component)
# -
# +
PROJECT_ID = 'jk-mlops-dev'
REGION = 'us-central1'
USER = 'test'
STAGING_BUCKET = 'gs://jk-vertex-us-central1'
PIPELINE_ROOT = "{}/pipeline_root/{}".format(STAGING_BUCKET, 'pipeline_runs')
VERTEX_SA = f'<EMAIL>'
PIPELINE_ROOT
# -
aiplatform.init(
project=PROJECT_ID,
location=REGION,
staging_bucket=STAGING_BUCKET
)
@component
def preprocess(
# An input parameter of type string.
message: str,
# Use Output to get a metadata-rich handle to the output artifact
# of type `Dataset`.
output_dataset_one: Output[Dataset],
# A locally accessible filepath for another output artifact of type
# `Dataset`.
output_dataset_two_path: OutputPath("Dataset"),
# A locally accessible filepath for an output parameter of type string.
output_parameter_path: OutputPath(str),
):
"""'Mock' preprocessing step.
Writes out the passed in message to the output "Dataset"s and the output message.
"""
output_dataset_one.metadata["hello"] = "there"
# Use OutputArtifact.path to access a local file path for writing.
# One can also use OutputArtifact.uri to access the actual URI file path.
with open(output_dataset_one.path, "w") as f:
f.write(message)
# OutputPath is used to just pass the local file path of the output artifact
# to the function.
with open(output_dataset_two_path, "w") as f:
f.write(message)
with open(output_parameter_path, "w") as f:
f.write(message)
@component(
base_image="python:3.9", # Use a different base image.
)
def train(
# An input parameter of type string.
message: str,
# Use InputPath to get a locally accessible path for the input artifact
# of type `Dataset`.
dataset_one_path: InputPath("Dataset"),
# Use InputArtifact to get a metadata-rich handle to the input artifact
# of type `Dataset`.
dataset_two: Input[Dataset],
# Output artifact of type Model.
imported_dataset: Input[Dataset],
model: Output[Model],
# An input parameter of type int with a default value.
num_steps: int = 3,
# Use NamedTuple to return either artifacts or parameters.
# When returning artifacts like this, return the contents of
# the artifact. The assumption here is that this return value
# fits in memory.
) -> NamedTuple(
"Outputs",
[
("output_message", str), # Return parameter.
("generic_artifact", Artifact), # Return generic Artifact.
],
):
"""'Mock' Training step.
Combines the contents of dataset_one and dataset_two into the
output Model.
Constructs a new output_message consisting of message repeated num_steps times.
"""
# Directly access the passed in GCS URI as a local file (uses GCSFuse).
with open(dataset_one_path, "r") as input_file:
dataset_one_contents = input_file.read()
# dataset_two is an Artifact handle. Use dataset_two.path to get a
# local file path (uses GCSFuse).
# Alternately, use dataset_two.uri to access the GCS URI directly.
with open(dataset_two.path, "r") as input_file:
dataset_two_contents = input_file.read()
with open(model.path, "w") as f:
f.write("My Model")
with open(imported_dataset.path, "r") as f:
data = f.read()
print("Imported Dataset:", data)
# Use model.get() to get a Model artifact, which has a .metadata dictionary
# to store arbitrary metadata for the output artifact. This metadata will be
# recorded in Managed Metadata and can be queried later. It will also show up
# in the UI.
model.metadata["accuracy"] = 0.9
model.metadata["framework"] = "Tensorflow"
model.metadata["time_to_train_in_seconds"] = 257
artifact_contents = "{}\n{}".format(dataset_one_contents, dataset_two_contents)
output_message = " ".join([message for _ in range(num_steps)])
return (output_message, artifact_contents)
@component
def read_artifact_input(
generic: Input[Artifact],
):
with open(generic.path, "r") as input_file:
generic_contents = input_file.read()
print(f"generic contents: {generic_contents}")
@dsl.pipeline(
# Default pipeline root. You can override it when submitting the pipeline.
pipeline_root=PIPELINE_ROOT,
# A name for the pipeline. Use to determine the pipeline Context.
name="metadata-pipeline-v2",
)
def pipeline(message: str):
importer = kfp.dsl.importer(
artifact_uri="gs://ml-pipeline-playground/shakespeare1.txt",
artifact_class=Dataset,
reimport=False,
)
preprocess_task = preprocess(message=message)
train_task = train(
dataset_one=preprocess_task.outputs["output_dataset_one"],
dataset_two=preprocess_task.outputs["output_dataset_two"],
imported_dataset=importer.output,
message=preprocess_task.outputs["output_parameter"],
num_steps=5,
)
read_task = read_artifact_input( # noqa: F841
train_task.outputs["generic_artifact"]
)
# +
package_path = "hw_pipeline_job.json"
compiler.Compiler().compile(
pipeline_func=pipeline, package_path=package_path)
# +
job_name = 'test_pipeline_run'
pipeline_job = aiplatform.PipelineJob(
display_name=job_name,
template_path=package_path,
enable_caching=False,
)
pipeline_job.run(
service_account=VERTEX_SA
)
# -
| kfp-nvt/kfp-nvt-sandbox.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dsref
# language: python
# name: dsref
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Circle
from matplotlib.figure import Figure
# %matplotlib inline
# #%matplotlib nbagg
import ipywidgets as widgets
from IPython.display import display
# -
# # Matplotlib
# ## Controlling the appearance
# +
# Initialize the figure and subplot
fig = plt.figure(figsize=(10,5))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2)
# Titles
fig.suptitle('Sample Figure', fontsize=14)
ax1.set_title("Scatter Plot", fontsize='large')
ax2.set_title("Curves", fontsize='large')
# Set the limits
ax1.set_xlim(0, 5)
ax1.set_ylim(0, 7)
# ticks and labels
ax1.set_xticks([1,2,3,4])
ax1.set_yticks([1,2,3,4,5,6])
ax1.set_xticklabels(['a', 'b', 'c'])
ax1.set_yticklabels(['','','','first', 'second', 'third'])
# plot
ax1.scatter([1,2,3], [4,5,6])
x = np.linspace(1., 8., 30)
ax2.plot(x, x ** 1.5, 'ro', label='density')
ax2.plot(x, 20/x, 'bx', label='temperature')
ax2.legend()
# Retrieve an element of a plot and set properties
for tick in ax1.xaxis.get_ticklabels():
tick.set_weight('bold')
tick.set_color('blue')
# You can also change the global setting
plt.rc('xtick', labelsize='medium', direction='out', color='r')
plt.rc('xtick.major', size=4, pad=4)
plt.show()
fig.savefig('datavs.png', bbox_inches='tight')
# -
# closing activities
plt.close(fig)
plt.rcdefaults()
# ## Twin Axis
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax2 = ax1.twinx()
t = np.linspace(0., 10., 100)
ax1.plot(t, t ** 2, 'b-')
ax2.plot(t, 1000 / (t + 1), 'r-')
ax1.set_ylabel('Density (cgs)', color='red')
ax2.set_ylabel('Temperature (K)', color='blue')
ax1.set_xlabel('Time (s)')
# ## Inserting subplots
fig = plt.figure(figsize=(4,3)) # create the figure with size (width,height)
ax1 = fig.add_axes([0.1,0.1,0.8,0.8])
ax1.plot([4,3,2,1])
ax2 = fig.add_axes([0.62,0.62,0.20,0.20])
ax2.plot([1,2])
plt.show()
# ## Color Bars
fig = plt.figure()
ax = fig.add_axes([0.1,0.1,0.6,0.8])
image = np.random.poisson(10., (100, 80))
i = ax.imshow(image, aspect='auto', interpolation='nearest')
colorbar_ax = fig.add_axes([0.7, 0.1, 0.05, 0.8])
fig.colorbar(i, cax=colorbar_ax)
# ## Artists, Patches, and Lines
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
c = Circle((0.5, 0.5), radius=0.2,
edgecolor='red', facecolor='blue', alpha=0.3)
ax.add_patch(c)
plt.show()
# # Interactive Graphs
# ## ipyWidgets
# + [markdown] cell_style="center"
# ### Example : Sine Wave
# + [markdown] cell_style="center"
# $A = amplitude$
#
# $\phi = phase$
#
# $\nu = frequency$
#
# $A \sin (2\pi(\nu x + \phi))$
# +
def update_plot(amp=1, phase=1, freq=1):
'''
This function is linked to the sliders, and
it replots the sine wave when sliders are changed
'''
plt.suptitle('Sine Wave')
x = np.linspace(0, 2, 1000)
y = amp + np.sin(2 * np.pi * (freq * x + phase))
plt.plot(x,y)
plt.show()
amp = widgets.FloatSlider(min=1, max=10, value=1, description='Amp:')
phase= widgets.FloatSlider(min=0, max=5, value=0, description='Phase:')
freq = widgets.FloatSlider(min=1, max=10, value=1, description='Freq:')
widgets.interact(update_plot, amp=amp, phase=phase, freq=freq)
# -
# ### Example : Polynomial
# +
# This example show an additional axis
fig, ax = plt.subplots(1, figsize=(10,4))
def g(c=10):
x = np.linspace(-5, 5, 20)
y = c * x**2
ax.clear()
ax.plot(x,y,'r--')
ax.set_xlim([-5,5])
ax.set_ylim([0,80])
fig.canvas.draw()
#plt.show()
display(fig)
c_slide = widgets.FloatSlider(value=1.0, min=0, max=3, step=0.1)
widgets.interact(g, c=c_slide)
# -
| dsref/notebooks/.ipynb_checkpoints/Data Visualization-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Cheating coin toss quantum circuit
# In this exercise we create a quantum circuit quantum circuit that tweaks the probabilistic nature of a single qubit in superposition. Like before we put the qubit in superposition $|\psi\rangle = \left(|0\rangle+|1\rangle\right)/\sqrt{2}.$
# We then add an additional Ry (rotation around the Y axis) gate, which pushes the Bloch vector $\pi/8$ closer to $|1\rangle$. we tweak the odds in favor of $|1\rangle$. The expected outcome is now $|0\rangle$ ~ 33% and $|1\rangle$ with a 66% probability.
# The result is displayed as a numeric readout, as a bar diagram.
#
# In this exercise we introduce the Ry gate, which rotates a qubit around the y-axis.
# ```
# ┌────────────┐
# q_0: |0>┤ Ry(0.3927) ├
# └────────────┘
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# Import the required libraries.
# + slideshow={"slide_type": "fragment"}
from qiskit import QuantumCircuit, execute, Aer
# Import Blochsphere visualization
from qiskit.visualization import plot_bloch_multivector, plot_histogram
# We also need to use pi in our gate input
from math import pi
# + [markdown] slideshow={"slide_type": "slide"}
# As we will be using the Bloch sphere visualization (`plot_bloch_multivector`) a bit, here's a quick function that calculates the state vector ($|\psi\rangle$) for the circuit to let you display the Bloch vector for any given state.
# + slideshow={"slide_type": "fragment"}
def get_psi(circuit):
global psi
backend = Aer.get_backend('statevector_simulator')
psi = execute(circuit, backend).result().get_statevector(circuit)
# + [markdown] slideshow={"slide_type": "slide"}
# Create an empty quantum circuit.
# + slideshow={"slide_type": "fragment"}
qc = QuantumCircuit(1,1)
print(qc)
# Display the Bloch sphere
get_psi(qc)
plot_bloch_multivector(psi)
# + [markdown] slideshow={"slide_type": "slide"}
# Add a Hadamard (super position) gate to the quantum circuit. This puts the qubit in a superposition: $|\psi\rangle = \left(|0\rangle+|1\rangle\right)/\sqrt{2}.$
# + slideshow={"slide_type": "fragment"}
qc.h(0)
print(qc)
# Display the Bloch sphere
get_psi(qc)
plot_bloch_multivector(psi)
# + [markdown] slideshow={"slide_type": "slide"}
# Now, let's rotate the Bloch vector vector $\pi$/8 radians closer to $|1\rangle$ by adding a Y rotation gate.
# + slideshow={"slide_type": "fragment"}
qc.ry(pi/8,0)
#Move the Bloch vector pi/8 radians closer to |1>.
print(qc)
# Display the Bloch sphere
get_psi(qc)
plot_bloch_multivector(psi)
# + [markdown] slideshow={"slide_type": "slide"}
# And finally add the measurement:
# + slideshow={"slide_type": "fragment"}
qc.measure(0,0)
print(qc)
# + [markdown] slideshow={"slide_type": "slide"}
# Set the backend to a local simulator.
# + slideshow={"slide_type": "fragment"}
backend = Aer.get_backend('qasm_simulator')
# + [markdown] slideshow={"slide_type": "slide"}
# Create a quantum job for the circuit, the selected backend, that runs a number of shots to simulate a sequence of coin tosses. Run the job and display the results. Display the result as a histogram. The slight tweak of the Bloch vector towards |1> turns out to have a massive result in that ~70% of the coin tosses are now |1>.
# + slideshow={"slide_type": "fragment"}
job = execute(qc, backend, shots=1000)
result = job.result()
counts = result.get_counts(qc)
print(counts)
plot_histogram(counts)
# -
| qiskit_advocates/meetups/09_23_19-Hassi_Norlen-Morgan_State_Uni/1_2_coin_toss_T.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import ipywidgets as widgets
from traitlets import Unicode, validate, Bool, Any
from ipywidgets import Color
class ImageLabelerWidget(widgets.DOMWidget):
_view_name = Unicode('ImageLabelerView').tag(sync=True)
_view_module = Unicode('ImageLabeler').tag(sync=True)
# + language="javascript"
# var isClicked = false;
# var rect, startX, startY, endX, endY;
#
# //Cargamos CSS
# var cssId = 'Style';
# if (!document.getElementById(cssId))
# {
# var head = document.getElementsByTagName('head')[0];
# var link = document.createElement('link');
# link.id = cssId;
# link.rel = 'stylesheet';
# link.type = 'text/css';
# link.href = 'Style.css';
# head.appendChild(link);
# }
#
# function updateRect(rect) {
# // rect.attr("x",endX - startX > 0 ? startX : endX)
# // .attr("y",y: endY - startY > 0 ? startY : endY)
# // .attr("width",Math.abs(endX - startX))
# // .attr("height",Math.abs(endY - startY));
# rect.attr({
# x: endX - startX > 0 ? startX : endX,
# y: endY - startY > 0 ? startY : endY,
# width: Math.abs(endX - startX),
# height: Math.abs(endY - startY)
# });
# }
#
# require.undef('ImageLabeler');
# define('ImageLabeler', ["jupyter-js-widgets"], function(widgets) {
#
# var ImageLabelerView = widgets.DOMWidgetView.extend({
# // Renderizar vista
# render: function() {
# require.config({
# paths: {
# d3: '//cdnjs.cloudflare.com/ajax/libs/d3/3.4.8/d3.min'
# }
# });
# require(['d3'], function(d3){
# //Añadimos un contenedor
# var canvas = element.append("<div id='canvas'></div>");
# // var canvas = createElement('div');
# // d3.select(canvas)
# // .attr("id","canvas")
# // .attr("width","960px")
# // .attr("height","600px");
#
# $("#canvas").width("960px");
# $("#canvas").height("600px");
#
# var margin = {top: 20, right: 20, bottom: 30, left: 40};
# var width = 880 - margin.left - margin.right;
# var height = 500 - margin.top - margin.bottom;
#
# //Creamos SVG
# var w = 600, h = 500;
# var svg = d3.select('#canvas').append('svg').attr({width: w, height: h});
#
# // Añadimos un rectángulo
# //var svg = svg.append("rect").attr("x",10).attr("y",10).attr("width",10).attr("height",10)
#
# //Añadimos la imagen
#
# svg.append("image")
# .attr("xlink:href", "http://www.google.com/intl/en_ALL/images/logo.gif")//"@Url.Content("~/Content/images/icons/refresh.png")")
# .attr("x", "0")
# .attr("y", "0")
# .attr("width", "200")
# .attr("height", "200");
#
# //Añadimos listeners
# d3.select("#canvas").on("click", function(eve){
# console.log(d3.mouse(this));
#
# if(isClicked){
# //Finalizamos rectangulo
# isClicked = false;
# //Inicializamos coords.
# endX = d3.mouse(this)[0];
# endY = d3.mouse(this)[1];
#
# //Cambiamos ref al rectangulo
# rect = null;
#
# //logger
# console.log("Finalizamos rectangulo");
# console.log(endX);
# console.log(endY);
#
# //Inicializamos coords.
# endX = d3.mouse(this)[0];
# endY = d3.mouse(this)[1];
#
# }else{
# //Inicializamos rectangulo
# isClicked = true;
# //Inicializamos coords.
# startX = d3.mouse(this)[0];
# startY = d3.mouse(this)[1];
#
# //Creamos rectangulo
# rect = svg.append("rect")
# .attr("x",startX)
# .attr("y",startY)
# .attr("width",0)
# .attr("height",0)
# .attr("class","rectangle");
# // .attr("fill","transparent")
# // .attr("stroke","green")
# // .attr("stroke-width","5px");
#
# console.log("Inicializamos rectangulo");
# console.log(startX);
# console.log(startY);
#
# rect = svg.append("rect");
# //updateRect(rect);
# }
# });
#
# d3.select("#canvas").on("mousemove", function(eve){
# if(isClicked){
# endX = d3.mouse(this)[0];
# endY = d3.mouse(this)[1];
# // updateRect(rect);
#
# //Actualizamos valores
# if (endX - startX > 0){
# var x = startX
# }else{
# var x = endX;
# }
#
# if (endY - startY > 0){
# var y = startY
# }else{
# var y = endY;
# }
# var width = Math.abs(endX - startX);
# var height = Math.abs(endY - startY);
#
# rect.attr("x", x)
# .attr("y", y)
# .attr("width", width)
# .attr("height", height)
# .attr("class","rectangle");
# }
# });
#
# });
# },
# });
#
# return {
# ImageLabelerView: ImageLabelerView
# };
# });
# -
ImageLabelerWidget()
| code/notebooks/Prototypes/Image_Labeler_Prototypes/Image_Labeler_prototype2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Continuous Control
#
# Project #2 for the Udacity Reinforcement Learning Nanodegree.
# <NAME>
# __________
#
# <img src="images/arm.jpg" width="250" />
#
# ## Introduction
#
# For this project, we will work with the [Reacher](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Learning-Environment-Examples.md#reacher) environment.
#
# In this environment, a double-jointed arm can move to target locations. A reward of +0.1 is provided for each step that the agent's hand is in the goal location. Thus, the goal of the agent is to maintain its position at the target location for as many time steps as possible.
#
# The observation space consists of 33 variables corresponding to position, rotation, velocity, and angular velocities of the arm. Each action is a vector with four numbers, corresponding to torque applicable to two joints. Every entry in the action vector should be a number between -1 and 1.
#
# The task is episodic, and in order to solve the environment, the agent must get an average score of +30 over 100 consecutive episodes.
#
# There are two versions of the environment, one with a single agent, an one with 20 agents. A solution for both versions is provided.
#
#
# ## Model
#
# The problem is solved by using a Deep Deterministic Policy Gradient (DDPG) architecture, which is based on the paper [Continuous control with deep reinforcement learning](https://arxiv.org/abs/1509.02971)
#
# The code for the agents can be found in [drltools/agent.py](https://github.com/lalopey/drl/blob/master/drltools/agent/agent.py), under the DDPGAgent class.
# The code for the PyTorch models for the Critic and the Actor can be found in [drltools/model.py](https://github.com/lalopey/drl/blob/master/drltools/model/model.py)
#
#
# ### Actor-Critic Method
#
# DDPG is an Actor-Critic method. It attempts to estimate both the policy directly, the Actor, and the value function, the Critic.
#
# Policy based approaches, which attempt to estimate the optimal policy directly, are low bias/unbiased methods. They require many samples to train, and have high variance. For example, a sampled trajetcory might have taken a good action in general, but received a low reward by chance. This will lower the odds we will pick this action in the future, and only figure out that it was a good action after many samples.
#
# Value based approaches attempt to estimate the Value function. They have lower variance than Policy based methods, as they use an estimate of an estimate for each step, but can be biased, as your estimate might be incorrect, particularly early on in training.
#
#
# ### Q-learning in continuous action space
#
# DDPG tries to bring Q-Learning to continuous action domain. In Q-learning, we update the Q-function by looking at the action that maximizes the Q function for the received future state:
#
# $$Q^{new}_{\pi}(s,a) = Q^{old}_{\pi}(s,a) + \alpha (r_t + \gamma \max_a Q_{\pi}(s_{t+1}, a) - Q^{old}_{\pi}(s,a))$$
#
# The maximum of the Q function can't be easily estimated with a continuous action space. To work through this issue, DDPG tries to estimate the maximum by estimating a deterministic policy.
#
# The Actor network approximates a deterministic policy. This will give us the action that the policy considers the best for a given state. We can use this estimate the maximum in the Q-function update. The Q-function approximation will be the Critic.
#
# ### Experience replay
#
# Like in Q-learning, we can profit from using experience replay. When the agent interacts with the environment, the sequence of observations can be highly correlated. The naive Q-learning algorithm that learns from each of these experience tuples in sequential order runs the risk of getting swayed by the effects of this correlation. By instead keeping track of a replay buffer and later sampling from this buffer at random, a method known as experience replay, we can prevent action values from oscillating or diverging catastrophically. Experience replay also allows us to learn more from individual observations multiple times, recall rare events, and in general make better use of our experience.
#
# ### Fixed Q-targets
#
# Like in Q-learning, we will have a local and target network, both for the Actor and the Critic. The local network is the network we are training, while the target network is the one we use for prediction to stabilize training. Its weights are updated slowly.
#
# ### Exploration and Noise
#
# When we have discrete action spaces, we can explore different paths by sampling from the action space, as we did through an epsilon greedy policy in the last project.
#
# For continuous action spaces, exploration is done via adding noise to the action itself. In the DDPG paper, the authors use Ornstein-Uhlenbeck process (OU) to add noise to the action output:
#
# The OU process is a continuous mean-reverting stochastic process. It generates noise that is correlated with the previous noise, as to prevent the noise from canceling out or freezing the overall dynamics.
#
# ### Batch Normalization
#
# Batch normalization is a technique for training deep neural networks that standardizes the inputs to a layer for each mini-batch. This has the effect of stabilizing the learning process and reducing the number of training episodes required to train the network.
#
# ### Training with multiple agents
#
# Since the enviornment for the 20 agents provides the same dynamics for each individual agent, and they don't interact with each other, we can use the same neural network architecture to train all the agents. We just need to add the samples generated by each agent to the replay buffer and sample for training for there.
#
#
#
# ## Results
#
# ### Agent Training and Model Architecture
#
# The model architecture used to solve the problem is as follows:
#
# - A target and local neural network for the Critic, both with the same architecture:
# - One layer with states as input, with 256 nodes. The layer is activated with ReLu and batch normalized
# - A second layer with 256 nodes, ReLu activated.
# - A final layer with a single output, as we are estimating a deterministic policy
#
#
# - A target and local neural network for the Actor, both with the same architecture:
# - One layer with states as input, with 256 nodes. The layer is activated with ReLu and batch normalized
# - A second layer, with 256 nodes, ReLu activated.
# - A final layer that maps into action space, activated with tanh so that the output is between -1 and 1 as its possible for the actions.
#
# - A batch size of 256 is used.
# - The local networks are optimized with Adam, with a learning rate of 1e-4 and 4e-4 for the actor and critic, respectively.
# - The $\gamma$ in the TD update is 0.99
# - The weights of the target network are updated through a soft update θ_target = τ*θ_local + (1 - τ)*θ_target with τ = 1e-3
# - A buffer size of 1e6 for experience replay.
# - An Ornstein-Uhlenbeck noise process with parameters $\mu=0$, $\theta=0.15$, $\sigma=0.1$ is used
# - For the single agent environment, the buffer is sampled 10 times every 20 steps, while for the 20 agent environment, its samples twice every four steps.
#
#
# ### DDPG
#
# The environment solved with DDPG is solved in 185 episodes.
#
# Episode 100 Average Score: 8.22
# Episode 185 Average Score: 30.04 Episode score (max over agents): 38.36
# Environment solved in 185 episodes with an Average Score of 30.04
#
# <img src="images/ddpg_reacher.png" width="450" />
#
# ### Multi Agent DDPG
#
# The environment for 20 agents is solved in 100 episodes. It converges very fast as all the arms learn at the same time.
#
# Episode 100 Average Score: 31.69
#
# Environment solved in 100 episodes with an Average Score of 31.69
#
# <img src="images/ddpg_reacher20.png" width="450" />
#
# # Future improvements
#
# On this project we levered what we had learned about Q-learning in the last project, and adapted it to solve the continuous action problem, through an actor-critic framework.
#
# There are other architectures that more directly use the actor-critic approach to estimating the value and policy functions at the same time. Some of them are:
#
# - [PPO](https://arxiv.org/pdf/1707.06347.pdf)
# - [A3C](https://arxiv.org/pdf/1602.01783.pdf)
# - [D4PG](https://openreview.net/pdf?id=SyZipzbCb)
#
# These architectures can train multiple non-interacting agents in parallel, which would be very efficient for our problem.
#
# Other types of agents that have recently been introduced could also be useful, including:
#
# - [TD3](https://arxiv.org/abs/1802.09477)
# - [SAC](https://arxiv.org/abs/1812.05905)
# - [PlanNet](https://ai.googleblog.com/2019/02/introducing-planet-deep-planning.html)
# - [Dreamer](https://ai.googleblog.com/2020/03/introducing-dreamer-scalable.html)
#
# We could also improve on the way the replay buffer samples through
# [Prioritized Experience Replay](https://arxiv.org/abs/1511.05952) and [Hindsight Experience Replay](https://arxiv.org/abs/1707.01495)
#
#
#
| 2 - Continuous Control -DDPG/Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: skorch
# language: python
# name: skorch
# ---
import matplotlib.pyplot as plt
import numpy as np
import torch.nn as nn
from skorch import NeuralNetClassifier
import umap
# !pip install /pstore/home/shetabs1/code/iflai/
import warnings
warnings.filterwarnings("ignore")
import os
import torch
from torchvision import transforms
import pandas as pd
import numpy as np
from torch.utils.data import DataLoader, Dataset
from sklearn.metrics import classification_report
from imblearn.over_sampling import RandomOverSampler
from iflai.dl.util import read_data, get_statistics_h5, calculate_weights
from iflai.dl.dataset import train_validation_test_split_wth_augmentation, Dataset_Generator_Preprocessed_h5
#from iflai.ml.feature_extractor import AmnisData
from iflai.dl.models import PretrainedModel
from skorch.callbacks import LRScheduler, Checkpoint
import torch.optim as optim
from skorch.helper import predefined_split
# +
seed_value = 42
os.environ['PYTHONHASHSEED']=str(seed_value)
import random
random.seed(seed_value)
np.random.seed(seed_value)
torch.manual_seed(seed_value)
# -
dataset_name = "wbc"
only_channels = [0,1,2,3,4,5,6,7,8,9,10,11]
path_to_data ="/pstore/data/DS4/White_blood_cell_dataset/Experiment_1/Donor_1/condition_1/"
scaling_factor = 255.
reshape_size = 64
num_channels = len(only_channels)
train_transform = transforms.Compose(
[transforms.RandomVerticalFlip(),
transforms.RandomHorizontalFlip(),
transforms.RandomRotation(45)])
test_transform = transforms.Compose([])
batch_size = 256
num_workers = 2
dev="cuda"
# +
# amnis_data = AmnisData(path_to_data, None)
# -
X, y, CLASS_NAMES, data_map = read_data(path_to_data)
num_classes = len(data_map.keys())
train_indx, validation_indx, test_indx = train_validation_test_split_wth_augmentation(X, y, only_classes=None)
# +
train_dataset = Dataset_Generator_Preprocessed_h5(path_to_data=path_to_data,
set_indx=train_indx,
scaling_factor=scaling_factor,
reshape_size=reshape_size,
transform=train_transform,
data_map=data_map,
only_channels=only_channels,
num_channels=num_channels)
trainloader = DataLoader(train_dataset,
batch_size=batch_size,
shuffle=False,
num_workers=num_workers)
# -
statistics = get_statistics_h5(trainloader, only_channels, None, num_channels)
y_train = [data_map.get(y[i]) for i in train_indx]
weights = calculate_weights(y_train)
class_weights = torch.FloatTensor(weights).to(dev)
oversample = RandomOverSampler(random_state=seed_value, sampling_strategy='all')
train_indx, y_train = oversample.fit_resample(np.asarray(train_indx).reshape(-1, 1), np.asarray(y_train))
train_indx = train_indx.T[0]
y_train = [data_map.get(y[i]) for i in train_indx]
y_test = [data_map.get(y[i]) for i in test_indx]
# +
train_dataset = Dataset_Generator_Preprocessed_h5(path_to_data=path_to_data,
set_indx=train_indx,
scaling_factor=scaling_factor,
reshape_size=reshape_size,
transform=train_transform,
data_map=data_map,
only_channels=only_channels,
num_channels=num_channels,
means=statistics["mean"],
stds=statistics["std"],
return_only_image=True,
)
validation_dataset = Dataset_Generator_Preprocessed_h5(path_to_data=path_to_data,
set_indx=validation_indx,
scaling_factor=scaling_factor,
reshape_size=reshape_size,
transform=test_transform,
data_map=data_map,
only_channels=only_channels,
num_channels=num_channels,
means=statistics["mean"],
stds=statistics["std"],
return_only_image=True,
)
test_dataset = Dataset_Generator_Preprocessed_h5(path_to_data=path_to_data,
set_indx=test_indx,
scaling_factor=scaling_factor,
reshape_size=reshape_size,
transform=test_transform,
data_map=data_map,
only_channels=only_channels,
num_channels=num_channels,
means=statistics["mean"],
stds=statistics["std"],
return_only_image=True,
)
# -
lrscheduler = LRScheduler(policy='StepLR', step_size=7, gamma=0.5)
checkpoint = Checkpoint(f_params='wbs_net_all.pth', monitor='valid_loss_best')
net = NeuralNetClassifier(
PretrainedModel,
criterion=nn.CrossEntropyLoss,
criterion__weight=class_weights,
lr=1e-5,
batch_size=256,
max_epochs=10,
module__output_features=num_classes,
module__num_channels=num_channels,
optimizer=optim.Adam,
iterator_train__shuffle=False,
iterator_train__num_workers=4,
iterator_valid__shuffle=False,
iterator_valid__num_workers=2,
callbacks=[lrscheduler, checkpoint],
train_split=predefined_split(validation_dataset),
device=dev
)
net.fit(train_dataset, y=None)
y_pred_net = net.predict(test_dataset)
print(classification_report(y_test, y_pred_net, target_names=CLASS_NAMES, digits=4))
plt.imshow(test_dataset[3][0][0,:,:])
| docs/notebooks/dl_classification_wbc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Function Decorators for Accelerated Code
#
# The idea is to provide a simple API for end users to interact with custom IP in the fabric, and provide a simple mechanism for overlay writers to expose that functionality to end users. The idea would be to have a decorator that marks a function as being potentially offloaded `@hardware_function(vlnv)` that handles all of the communication. The return type and argument type are then expressed using python type annotations. If the VLNV appears in the loaded bitstream then a wrapper will be returned that, upon accessing the data, will act like a numpy array of the specified type. If the VLNV is not in the block design, the function will be executed as per normal.
# ## Representation of call chains
# The first task is to provide wrappers for the call chains which are being offloaded. This is taken wholesale from the test notebook. At the moment, it is assumed that all functions take one or more streams and input and return a single stream.
# +
import numpy as np
class Wrapper:
def __init__(self, wrapped, dtype = np.int32):
self.wrapped = wrapped
self.dtype = dtype
def value(self):
return self.wrapped
class Call:
def __init__(self, func, stream_args, scalar_args, return_type = np.uint32):
self.func = func
self.args = stream_args
self.scalar_args = scalar_args
self.dtype = return_type
self.cached = None
def value(self):
return self.func(*[a.value() for a in self.args])
def hw_value(self):
return execute_hardware(self)
def __str__(self):
if self.cached is None:
self.cached = self.hw_value()
return str(self.cached)
def __getitem__(self, index):
if self.cached is None:
self.cached = self.hw_value()
return self.cached[index]
def __len__(self):
if self.cached is None:
self.cached = self.hw_value()
return len(self.cached)
# -
# ## Determining whats in the bitstream
# In order to correctly wire up the switches in the bitstream, we need to extract from the TCL file what IP is in the diagram and how it is wired. This is future work so, for now, it is hard-coded to the example bitstream but this will be changed post proof-of-concept.
# +
from collections import namedtuple
Function = namedtuple('Function', 'in_ports out_ports name')
class FunctionMetadata:
def __init__(self):
self.DMA = [([0],[0]),([5],[4])]
self.DMA_names = ['axi_dma_0', 'axi_dma_1']
self.functions = {}
self.functions['Xilinx:hls:stream_double:1.0'] = Function(in_ports=[2],out_ports=[2],name=None)
#self.functions['Xilinx:hls:stream_mult:1.0'] = Function(in_ports=[3,4],out_ports=[3],name=None)
self.functions['xilinx.com:hls:wrapped_conv_im2col_hw:1.0'] = Function(in_ports=[3,4],out_ports=[3],name=None)
self.functions['Xilinx:hls:simple_sum:1.0'] = Function(in_ports=[1],out_ports=[1],name=None)
self.functions['Xilinx:hls:mult_constant:1.0'] = Function(in_ports=[6],out_ports=[5],name='mult_constant_0')
metadata = FunctionMetadata()
# -
# ## Controlling the switch
# The next helper class controls the switch by setting routes. It is a thin wrapper around the control interface of the Xilinx AXI Stream Switch.
# +
from pynq import PL
from pynq import MMIO
class StreamingSwitch:
def __init__(self, name):
base_addr = int(PL.ip_dict["SEG_{0}_Reg".format(name)][0],16)
self.mmio = MMIO(base_addr, 256)
self.reset()
def set_route(self, in_port, out_port):
print('SWITCH: setting route {0} to {1}'.format(in_port, out_port))
self.mmio.write(0x40 + out_port * 4, in_port)
def reset(self):
for i in range(16):
# Disable the output on every port
self.mmio.write(0x40 + i * 4, 0x80000000)
def commit(self):
# Causes the switch to update atomically to the new routing
self.mmio.write(0, 2)
# -
# ## The Decorator
# Take a function and wrap it in a call object
# +
import inspect
def wrap_arg(a, dtype=np.int32):
if type(a) is Call or type(a) is Wrapper:
return a
else:
# TODO: sort out element type
return Wrapper(a, dtype);
def hardware_function(vlnv):
def decorator(func):
sig = inspect.signature(func)
ret_type = sig.return_annotation[0]
def wrapped_function(*args, **kwargs):
ba = sig.bind(*args, **kwargs)
if vlnv in metadata.functions:
stream_args = []
scalar_args = []
for param in sig.parameters.values():
if type(param.annotation) is list:
stream_args.append(wrap_arg(ba.arguments[param.name], param.annotation[0]))
else:
scalar_args.append(ba.arguments[param.name])
return Call(vlnv, stream_args, scalar_args, return_type=ret_type)
else:
# We don't have the function available so we might
# as well just call the function and return
return func(*args, **kwargs)
return wrapped_function
return decorator
# -
# ## Configuring the Switch and DMA
# The final step is to take a Call object and configure the switch accordingly. This process should also prime the DMA with the correct to be sent. We need a mechanism to set the correct size of the receiving buffer, thoughts welcome.
# Horrible hack to load the DMA driver
from pynq import Overlay
Overlay('base.bit').download()
from pynq.drivers import DMA
import pynq.drivers.dma
#Overlay('/home/xilinx/decorator_test.bit').download()
Overlay('/home/xilinx/decorator_conv_im2col.bit').download()
# ## Wrap the DMA
# Provide a simple API to the DMA. The DMA engine out to be separated out into a separate buffer as proposed separately then the DMA engine instances can be static and buffers could be returned without being copied.
class DMAWrapper:
def __init__(self,index):
print('Send DMA: create index {0} name {1}'.format(index, metadata.DMA_names[index]))
base_addr = int(PL.ip_dict["SEG_{0}_Reg".format(metadata.DMA_names[index])][0],16)
print('Send DMA: base_address {0:x}'.format(base_addr))
self.dma = DMA(base_addr, 0)
self.ports = metadata.DMA[index]
def set_data(self, data, dtype):
self.length = len(data) * dtype.itemsize
print('Send DMA: sending {0} bytes'.format(self.length))
self.dma.create_buf(self.length)
ffi = pynq.drivers.dma.ffi
buf = ffi.buffer(self.dma.buf, self.length)
view = np.frombuffer(buf, dtype, -1)
np.copyto(view, data, casting='same_kind')
def transfer(self):
print('Send DMA: transfer started')
self.dma.transfer(self.length, 0)
def wait(self):
self.dma.wait()
print('Send DMA: transfer finished')
# ## Parse the execution plan
# Next a recursive function is used to walk the execution plan. At the moment, there is no protection against using a function multiple times in a plan. That will follow later.
def prepare_execution(plan, dma, return_port):
if type(plan) is Wrapper:
d = DMAWrapper(len(dma))
d.set_data(plan.wrapped, plan.dtype())
dma.append(d)
hw_switch.set_route(d.ports[1][0], return_port)
elif type(plan) is Call:
in_ports = metadata.functions[plan.func].in_ports
out_ports = metadata.functions[plan.func].out_ports
name = metadata.functions[plan.func].name
mmio = None
if name:
mmio = MMIO(int(PL.ip_dict['SEG_{0}_Reg'.format(name)][0],16),256)
for i, a in enumerate(plan.args):
prepare_execution(a, dma, in_ports[i])
for i, a in enumerate(plan.scalar_args):
mmio.write(0x10 + 4*i, a)
hw_switch.set_route(out_ports[0], return_port)
else:
print("Unknown plan type: " + repr(plan))
# ## Execute the plan
# This is the main function that executes the plan. It first calls the parsing functions, then configures the input DMA engineswith suitable buffers and then waits for the return DMA to complete. Because the return buffer belongs to the DMA engine, a copy has to be taken. This can be changed with a modified DMA API
# +
hw_switch = StreamingSwitch('axis_switch_0')
def execute_hardware(plan):
dma = []
hw_switch.reset()
ret_dma_base = int(PL.ip_dict["SEG_{0}_Reg".format(metadata.DMA_names[0])][0],16)
ret_dma_mmio = MMIO(ret_dma_base, 256)
ret_dma = DMA(ret_dma_base, 1)
# TODO: Metadata for how big the buffer should be?
ret_dma.create_buf(8388607)
prepare_execution(plan, dma, metadata.DMA[0][0][0])
hw_switch.commit()
for d in dma:
d.transfer()
for d in dma:
d.wait()
ret_dma.transfer(8388607, 1)
ret_dma.wait()
bytes_read = ret_dma_mmio.read(0x58)
ffi = pynq.drivers.dma.ffi
buf = ffi.buffer(ret_dma.buf, bytes_read)
view = np.frombuffer(buf, plan.dtype, -1).copy()
return view
# -
# # Testing the Decorator
# Create some simple functions which map to the hardware functions and see if the decorator maps accordingly. We'll add some print statements to the python versions of the functions so we can make sure they're not called
# +
@hardware_function('Xilinx:hls:simple_sum:1.0')
def total(vs:[np.int32]) -> [np.int32]:
print("In total")
return sum(vs)
@hardware_function('Xilinx:hls:stream_double:1.0')
def double(vs:[np.int32]) -> [np.int32]:
print("In double")
return [v * 2 for v in vs]
#@hardware_function('Xilinx:hls:stream_mult:1.0')
@hardware_function('xilinx.com:hls:wrapped_conv_im2col_hw:1.0')
def mult(a:[np.int32], b:[np.int32]) -> [np.int32]:
return [a1 * b1 for (a1,b1) in zip(a,b)]
# -
# First we chain two hardware functions together. Note that no computation happens at this point as we don't know if the user wants this value or plans to use it as an intermediate value
# +
#vals = [1,2,3,4,5,6]
#vals2 = [6,5,4,3,2,1]
#inter = double(mult(vals, vals))
#t = total(inter)
#val1 = [5,5,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1]
#val1 = [6,6,
# 1,1,1,1,1,1,
# 1,1,1,1,1,1,
# 1,1,1,1,1,1,
# 1,1,1,1,1,1,
# 1,1,1,1,1,1,
# 1,1,1,1,1,1]
#val2 = [5,5,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,2]
#inter = mult(val1, val2)
#t = total(inter)
A = [8, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8,
1, 2, 3, 4, 5, 6, 7, 8
]
#B = [3, 3,
# 1, 3, 5,
# 3, 5, 1,
# 5, 3, 1
# ]
B = [8,8,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1,
1,1,1,1,1,1,1,1
]
t = mult(A, B)
print(t)
# -
# By calling print, we trigger the execution and the value is return
#print(t)
#tmp = t.hw_value() + 3
print(total(mult(A,B)))
# Because we never stored the intermediate value, if the user later requests it, we would need to redo the computation
print(inter)
# Our hardware also contains a block that multiplies by a constant. The constant is passed in using the AXI-lite interface.
# +
@hardware_function('Xilinx:hls:mult_constant:1.0')
def constant_multiply(in_data:[np.int32], constant:np.int32) -> [np.int32]:
return [v * constant for v in in_data]
print(constant_multiply([1,2,3,4,5,6,7], 5))
# -
# As `constant_multiple` is a python function like any other, we can also do function-y things to it. For example, we can use the `functools` library to partially apply the constant, giving us a new implementation of `double` in terms of `constant_multiply`
# +
import functools
new_double = functools.partial(constant_multiply, constant=2)
print(new_double(mult(vals,vals2)))
# -
# ## Open Problems
# * Allocation of receive buffer
# * Data bigger than buffer size - SG may be able to help here
# * 0-length arrays - AXI4-Stream has no concept of a 0-length stream. Maybe a word with no strb bits?
# * Current wrapper logic is patchy at best but completely proxying a python object is non-trivial
#
# ## Possible features
# * Plan partitioning for plans with more Calls than execution units/DMA engines
# * Re-use of intermediate values
# * I/O functions which configure the switch to route I/O directly
# * AXI-Master HLS support
#
# ## Performance considerations
# * Need a way for users to CMA alloc a numpy array
# * Buffers not bound to DMA so that any CMA allocated buffer can be passed
| python_notebooks/.ipynb_checkpoints/Decorator-checkpoint.ipynb |