code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
import matplotlib.pyplot as plt
#import seaborn as sns
import pandas as pd
import numpy as np
plt.rcParams['figure.dpi'] = 100
# # Using the adult dataset
adult = pd.read_csv("data/adult.csv", index_col=0)
adult.head()
adult.income.value_counts()
adult.income.value_counts().plot(kind="barh")
adult.education.value_counts()
adult.groupby("income")
adult.groupby("income")['education'].value_counts()
education_counts = adult.groupby("income")['education'].value_counts()
education_counts.unstack("income")
unstacked_education = education_counts.unstack("income")
unstacked_education.plot(kind="barh")
(unstacked_education / unstacked_education.sum(axis=0)).plot(kind="barh")
unstacked_education.columns
plt.figure()
(unstacked_education[" >50K"] / unstacked_education.sum(axis=1)).plot(kind="barh")
# # Exercise
# Group the data by gender, and compare the income distributions over genders.
# Do a similar plot for some of the other variables.
# +
# solution
# -
# # Exercise
# Apply the basic machine learning workflow to the dataset.
# For simplicity you might want to drop the "native-country" column.
# Proceed as follows:
# - separate features and target
# - do dummy encoding of the categorical features
# - split data in training and test set
# - scale the data
# - apply a machine learning model. Start with ``sklearn.linear_model.LogisticRegression``, a linear classifier.
# - visualize the coefficients in a bar-plot (if there are too many, only show the ones of larges magnitude)
| 05 - More data - the adult dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
from matplotlib import pyplot as plt
from datetime import datetime
from pymongo import MongoClient
client = MongoClient("mongodb://zui:F0reverqwerty@localhost:27017/")
db = client['hkns3']
col = db['items']
import pandas
df = pandas.DataFrame(list(db['raw'].find({"type":"max_id"})))
df['time'].astype(int)
df
datetime.fromtimestamp(db['raw'].find_one({"type":"max_id"})['time'])
df.plot(x="time", y="id")
| hkns/data/MoreDataAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 3.1 推論ベースの手法とニューラルネットワーク
import sys
sys.path.append('..')
import numpy as np
from common.trainer import Trainer
from common.optimizer import Adam
from common.layers import MatMul, SoftmaxWithLoss
from common.util import preprocess
# 単語IDが0の単語を全結合層で変換する例
c = np.array([[1, 0, 0, 0, 0, 0, 0]])
print(c.shape) # ミニバッチ処理を考えて2次元にしている
W = np.random.randn(7, 3)
h = np.dot(c, W)
print(h)
# MatMulレイヤを使う
layer = MatMul(W)
h = layer.forward(c)
print(h)
# # 3.2 シンプルなword2vec (CBOWモデル)
# ## CBOWモデルの推論処理
# **入力層 - 中間層 - 出力層** という単純なモデルを考える
# - 入力層と出力層のニューロンの数は単語ベクトルの要素数
# - 入力層の数はコンテキストとして与える単語数
# - 全ての入力層から中間層への変換は重み$\mathrm{W_{in}}$の全結合層によって,中間層から出力層への変換は重み$\mathrm{W_{out}}$の全結合層によって行われる
# - 中間層は各入力層の全結合による変換後の値を平均したもの
# - 出力層の値は各単語のスコア.Softmax関数を適用すると各単語の確率になる.
# - 学習後の重み$\mathrm{W_{in}}, \mathrm{W_{out}}$が単語の分散表現になる.中間層のニューロンの数を入力層よりも少なくすることで,密なベクトルが得られる.
# +
# サンプルデータ
c0 = np.array([[1, 0, 0, 0, 0, 0, 0]])
c1 = np.array([[0, 0, 1, 0, 0, 0, 0]])
# 重み初期化
W_in = np.random.randn(7, 3)
W_out = np.random.randn(3, 7)
# レイヤ生成
in_layer0 = MatMul(W_in)
in_layer1 = MatMul(W_in)
out_layer = MatMul(W_out)
# forward
h0 = in_layer0.forward(c0)
h1 = in_layer1.forward(c1)
h = 0.5 * (h0 + h1)
s = out_layer.forward(h)
# スコア出力
print(s)
# -
# ## CBOWモデルの学習
# ネットワークから出力されるスコアにSoftmax関数を適用することで確率を得ることができる.上の例では,単語IDが0と2の単語と共起する単語を予測するようなタスクを考えている.正解が単語IDが1の単語であれば,s[1]が最も高くなるように重みを調整することになる.
# 追加する層は
# - Softmaxレイヤ
# - Cross Entropy Errorレイヤ
#
# の2つ.すでに実装しているSoftmax with Lossレイヤで実装できる.
#
# <br/>
#
# 最終的に利用する単語の分散表現の選択肢は次の3つ
# - 入力側の重み: 行ベクトルが各単語の分散表現に対応.こちらだけを使うのが最もポピュラーな方法.
# - 出力側の重み: 列ベクトルが各単語の分散表現に対応.
# - 入出力の重み: 両方を足し合わせるなどの方法
# ***
# # 3.3 学習データの準備
# ## コンテキストとターゲット
# **コーパス → (コンテキスト & ターゲット)** という処理を行う
# まずはコーパスを作る
text = 'You say goodbye and I say hello.'
corpus, word_to_id, id_to_word = preprocess(text)
print(f'corpus: {corpus}')
print(f'id_to_word: {id_to_word}')
# コンテキストとターゲットを作る関数を実装
def create_contexts_target(corpus, window_size=1):
'''コンテキストとターゲットの作成
:param corpus: コーパス(単語IDのリスト)
:param window_size: ウィンドウサイズ(ウィンドウサイズが1のときは、単語の左右1単語がコンテキスト)
:return: tuple (contexts, target)
'''
target = corpus[window_size:-window_size] # 端の単語を除く
contexts = []
# コーパスの両端からwindow_size分を除いてループ
# ターゲットのidxになる
for idx in range(window_size, len(corpus)-window_size):
cs = []
# 今のidxから前後window_size分を見てコンテキストとして追加する
for t in range(-window_size, window_size + 1):
if t == 0:
continue
cs.append(corpus[idx + t])
contexts.append(cs)
return np.array(contexts), np.array(target)
contexts, target = create_contexts_target(corpus, window_size=1)
print(f'contexts:\n{contexts}')
print(f'target: {target}')
# ## one-hot表現への変換
# NNに入力するためone-hotベクトルに変換する
def convert_one_hot(corpus, vocab_size):
'''one-hot表現への変換
:param corpus: 単語IDのリスト(1次元もしくは2次元のNumPy配列)
:param vocab_size: 語彙数
:return: one-hot表現(2次元もしくは3次元のNumPy配列)
'''
N = corpus.shape[0]
if corpus.ndim == 1:
one_hot = np.zeros((N, vocab_size), dtype=np.int32)
for idx, word_id in enumerate(corpus):
one_hot[idx, word_id] = 1
elif corpus.ndim == 2:
C = corpus.shape[1]
one_hot = np.zeros((N, C, vocab_size), dtype=np.int32)
for idx_0, word_ids in enumerate(corpus):
for idx_1, word_id in enumerate(word_ids):
one_hot[idx_0, idx_1, word_id] = 1
return one_hot
vocab_size = len(word_to_id)
target = convert_one_hot(target, vocab_size)
contexts = convert_one_hot(contexts, vocab_size)
# コンテキストの形状は (batch_size, window_size, vocab_size) になる
# 今はコーパスが小さいので batch_size = data_size になっている
print(f'contexts.shape: {contexts.shape}')
print(f'contexts:\n{contexts}')
print()
print(f'target.shape: {target.shape}')
print(f'target:\n{target}')
# # CBOWモデルの実装
class SimpleCBOW:
def __init__(self, vocab_size, hidden_size):
# vocab_size: 語彙数
# hidden_size: 中間層のニューロン数
V, H = vocab_size, hidden_size
# 重みの初期化
W_in = 0.01 * np.random.randn(V, H).astype('f')
W_out = 0.01 * np.random.randn(H, V).astype('f')
# レイヤの生成
# in_layer はコンテキストの window_size 分必要
self.in_layer0 = MatMul(W_in)
self.in_layer1 = MatMul(W_in)
self.out_layer = MatMul(W_out)
self.loss_layer = SoftmaxWithLoss()
# すべての重みと勾配をリストにまとめる
layers = [self.in_layer0, self.in_layer1, self.out_layer]
self.params, self.grads = [], []
for layer in layers:
self.params += layer.params
self.grads += layer.grads
# メンバ変数に単語の分散表現を設定
self.word_vecs = W_in
def forward(self, contexts, target):
h0 = self.in_layer0.forward(contexts[:, 0]) # 0番目のwindowの単語のone-hotベクトルが入る
h1 = self.in_layer1.forward(contexts[:, 1]) # 1番目のwindowの単語のone-hotベクトルが入る
h = (h0 + h1) * 0.5 # 2つの入力層の値を平均する
score = self.out_layer.forward(h) # 出力層への変換
loss = self.loss_layer.forward(score, target) # 損失を計算
return loss
def backward(self, dout=1):
# 各レイヤのbackword()メソッドを呼び出し,レイヤのインスタンス変数gradを更新しつつ,下流の勾配計算に必要な偏微分を返す
# 各レイヤのインスタンス変数gradが更新されると,self.gradsも更新される (参照渡し)
ds = self.loss_layer.backward(dout)
da = self.out_layer.backward(ds)
da *= 0.5
self.in_layer1.backward(da)
self.in_layer0.backward(da)
return None
# ## 学習コードの実装
# +
window_size = 1
hidden_size = 5
batch_size = 3
max_epoch = 1000
model = SimpleCBOW(vocab_size, hidden_size)
optimizer = Adam()
trainer = Trainer(model, optimizer)
trainer.fit(contexts, target, max_epoch, batch_size)
trainer.plot()
# -
word_vecs = model.word_vecs
for word_id, word in id_to_word.items():
print(word, word_vecs[word_id])
# # 3.5 word2vecの補足
# ## CBOWモデルと確率
# 事象Aが起こる確率を $P(A)$ ,事象Bが起こる確率を $P(B)$ ,事象Aと事象Bが同時に起こる確率 (同時確率) を $P(A, B)$ ,事象Bが起きた後で事象Aが起こる確率 (事後確率) を $P(A|B)$ とする.
# <br/>
# $w_1, \cdots, w_{t-1}, w_t, w_{t+1}, \cdots, w_T$ というコーパス,ウインドウサイズ1のコンテキストを考える.
# つまりコンテキストは $w_{t-1}, w_{t+1}$ ,ターゲットは $w_t$ となる.
# これは$w_{t-1}, w_{t+1}$ が与えられたもとでの $w_t$ の確率であり,事後確率を使って次のように書ける.
#
# $$
# P(w_t | w_{t-1}, w_{t+1})
# $$
#
# CBOWはこの事後確率をモデル化したものである.
# 上式から損失関数を導く.
# 一つの単語の交差エントロピー誤差は次の通り.
#
# $$
# L = - \Sigma_k t_k \log y_k
# $$
#
# <br/>
#
# ターゲットはone-hotベクトルであるため正解ラベルに対応する $\log y_k$ のみを計算すればよい.損失関数は次の通り.
#
# $$
# L = - \log y_t = - \log P(w_t | w_{t-1}, w_{t+1})
# $$
#
# <br/>
#
# コーパス全体に拡張
#
# $$
# L = - \frac{1}{T} \Sigma_{t=1}^T \log P(w_t | w_{t-1}, w_{t+1})
# $$
# ***
# ## skip-gramモデル
# CBOWモデルとは反対に,中央の単語からその周囲のコンテキストを予測する
# - 入力層は一つで,ニューロンの数は語彙数
# - 出力層はコンテキストの数で,ニューロンの数は語彙数
#
# 確率の定義で表現する.
# $w_t$ が与えられたもとでの $w_{t-1}, w_{t+1}$ の同時確率であり,条件付き独立を仮定して変換する.
#
# $$
# P(w_{t-1}, w_{t+1} | w_t) = P(w_{t-1} | w_t) P(w_{t+1} | w_t)
# $$
#
# <br/>
#
# 交差エントロピー誤差に適用すると,skip-gramモデルの損失関数は次の通り.
#
# $$
# \begin{align}
# L &= - \log P(w_{t-1}, w_{t+1} | w_t) \\\\
# &= - \log P(w_{t-1} | w_t) P(w_{t+1} | w_t) \\\\
# &= - \{ \log P(w_{t-1} | w_t) + \log P(w_{t+1} | w_t) \}
# \end{align}
# $$
#
# <br/>
#
# コーパス全体に拡張
#
# $$
# L = - \frac{1}{T} \Sigma_{t=1}^T \{ \log P(w_{t-1} | w_t) + \log P(w_{t+1} | w_t) \}
# $$
#
| ch03/.ipynb_checkpoints/ch03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Scraping dynamic web sites with Selenium
#
# [Last week's lesson](../pt1/scraping_lecture.ipynb) involved scraping a static site, or a site that is rendered up front in HTML. Today, we'll look at how to scrape sites that change when you load or interact with the page, sometimes without the URL changing.
#
# [Selenium](https://www.selenium.dev/documentation/) was created to "automate browsers." The major use case for software like Selenium is to automate testing browser-based apps. But journalists can use software like Selenium to scrape dynamic websites.
#
# For today's lesson, we're going to scrape all the public hearings in Alameda County courts on a given day.
# +
import pandas as pd
from bs4 import BeautifulSoup
from tqdm.notebook import tqdm
from selenium import webdriver
from selenium.webdriver.support.select import Select
from selenium.webdriver.common.by import By
import time
import math
# -
# ## Open your automated browser
#
# Earlier we installed `chromedriver` using `brew`. Below, we tell Selenium to use Chrome as our automated browser.
# +
# initiate webdriver
driver = webdriver.Chrome()
# some people like to call this variable `browser` — call it whatever you like!
# -
# ## Open the website
# Navigate to a URL
driver.get('https://publicrecords.alameda.courts.ca.gov/CalendarSearch/')
# ## Find the inputs you want to interact with
#
# In last week's lecture, we used Beautiful Soup to find elements on a page. Because we want to interact with elements within Selenium's automated browser, we need to use Selenium to find elements.
#
# Tips:
# - If you want to interact with the page, use selenium
# - If you want to read or parse complex HTML, use bs4
#
# You'll use `By` to indicate how the browser will pinpoint your element. These are the [different options for `By`](https://www.selenium.dev/selenium/docs/api/py/webdriver/selenium.webdriver.common.by.html):
#
# - `CLASS_NAME`
# - `CSS_SELECTOR` (e.g. a pseudo-element)
# - `ID`
# - `LINK_TEXT` (the text inside <a> tags)
# - `NAME`
# - `PARTIAL_LINK_TEXT` (the text inside <a> tags)
# - `TAG_NAME`
# - `XPATH` (when the element doesn't have a unique identifer, you can still pinpoint with this method; Chrome has a cool way to grab the xpath of an item in Developer Tools)
#
# Luckily, the date fields have IDs, so we can select them this way:
hearing_date_from = driver.find_element(By.ID, 'FeaturedContent_txtFromdt')
hearing_date_to = driver.find_element(By.ID, 'FeaturedContent_txtTodt')
# You can use `type()` to find out whether a variable is a selenium object or a bs4 object.
type(hearing_date_from)
# ## Input dates into the dropdowns
#
# Use selenium's `send_keys()` method to input text into the date dropdowns.
hearing_date_from.send_keys('12/06/2021')
hearing_date_to.send_keys('12/06/2021')
# ## "Click" on the submit button
#
# First, you'll have to find the element by its `id` value, then `click()` on it.
submit_button = driver.find_element(By.ID, 'FeaturedContent_btFind')
submit_button.click()
# Below, I'm telling the computer to wait 5 seconds before executing the next line of code. That way the browser can finish loading the page before continuing with the code. That's crucial if I end up restarting this notebook kernel and running all cells at once. We want the browser to finish loading the page because some elements might not exist until the element exists.
time.sleep(5)
# There are better ways to wait for elements on a page. Check out the documentation to read more about [WebDriverWait()](https://selenium-python.readthedocs.io/waits.html).
# ## "Select" more rows to view
#
# When you get your search results, the courts show only 10 rows at a time. It'll be faster to scrape all the results if you can show the max amount of rows at a time (which is 50).
displayed_rows_dropdown = Select(driver.find_element(By.NAME, 'ctl00$MainContent$gvResult$ctl13$ctl13'))
displayed_rows_dropdown.select_by_visible_text('50')
# ## Get the count of results so you know how many pages you have to scrape
#
# Even though I'm parsing HTML below, I'm using Selenium instead of Beautiful Soup. I'm doing this because I haven't called Beautiful Soup yet and Selenium is capable of parsing.
records_count_container = driver.find_element(By.ID, 'MainContent_lbCnt')
records_count = records_count_container.text.split()
records_count = records_count[len(records_count) - 1]
records_count = int(records_count)
records_count
pages_to_check = math.ceil(records_count/50)
pages_to_check
# ## Figure out how to loop through the pages
# find the "Next" link — it looks like ">"
next_button = driver.find_element(By.LINK_TEXT, '>')
next_button.click()
# The below code is commented out because I don't want you to run it yet. But, you can see how one could flip through all the pages of this site.
# +
# for n in range(pages_to_check):
# next_button = driver.find_element(By.LINK_TEXT, '>')
# next_button.click()
# # wait 2 seconds
# time.sleep(2)
# -
# You can manually get back to the first page by going to the "automated" browser and clicking "1".
# ## Parse the first page of results with Beautiful Soup
#
# Now I'm going to switch to using Beautiful Soup because it's the best program to parse through a lot of HTML.
# ### Get the table by its `id`
soup = BeautifulSoup(driver.page_source, 'html.parser')
table = soup.find(id='MainContent_gvResult')
# +
# table
# + [markdown] tags=[]
# Each row of this table is a unique something. I'm not sure what that something is. It might not be a unique case. It might be something else. I'm not going to assume. Anyway, I'd like to transfer this table into a pandas dataframe.
#
# ### Create your blank dataframe
# -
hearings = pd.DataFrame(
columns=[
'Serial No.',
'Name',
'Case #',
'PFN',
'CEN',
'Dept#',
'Hearing Date',
'Hearing Time',
'Hearing Type',
'Case Type',
'Defense Atty',
'DA'
])
# ### Parse the table and put the data into a dataframe
#
# Let's go over each section below manually before running.
# +
# create a simple `page_data` list to store the page data before we make a pandas dataframe
page_data = []
rows = table.find_all('tr')
# we haven't used enumerate() yet but basically that just allows you to index an iterable
for i, row in enumerate(rows):
# we can skip the first row because that's the header row
# we can also skip any row greater than index 50 because that has the page numbers
if (i > 0) and (i <= 50):
# `cells` will get and index all the cells within a row
cells = row.find_all('td')
page_data.append({
'Serial No.' : cells[0].text.strip(),
'Name' : cells[1].text.strip(),
'Case #' : cells[2].text.strip(),
'PFN' : cells[3].text.strip(),
'CEN' : cells[4].text.strip(),
'Dept#' : cells[5].text.strip(),
'Hearing Date' : cells[6].text.strip(),
'Hearing Time' : cells[7].text.strip(),
'Hearing Type' : cells[8].text.strip(),
'Case Type' : cells[9].text.strip(),
'Defense Atty' : cells[10].text.strip(),
'DA' : cells[11].text.strip()
})
# create a dataframe with `page_data`
page_hearing = pd.DataFrame(page_data)
# -
# ## Append `page_hearing` dataframe to main `hearings` dataframe
hearings = hearings.append(page_hearing).reset_index(drop=True)
# ## View dataframe
hearings
for n in range(pages_to_check):
next_button = driver.find_element(By.LINK_TEXT, '>')
next_button.click()
time.sleep(2)
# ## Addenda
#
# If I want to search for another date, I can stay on the same page and "clear" the date fields. Then I send send new dates.
# +
hearing_date_from.clear()
hearing_date_to.clear()
hearing_date_from.send_keys('12/07/2021')
hearing_date_to.send_keys('12/07/2021')
# -
# Once you're done using the automated browser, you can close it manually or run the following:
driver.close()
# ## Classwork
#
# I'd like you to figure out how to loop through all the pages and collect all the information.
| pt2/selenium_lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.pyplot as plt
import networkx as nx
import pandas as pd
import sys
sys.path.append('/home/wrwt/Programming/pygraphmodels')
import graphmodels as gm
from itertools import permutations
from graphmodels import MatrixGraph, DGM
# %matplotlib inline
# %load_ext line_profiler
from graphmodels import AddEdge, RemoveEdge, ReverseEdge, InvalidOperation, GreedySearch, ScoreBIC
import heapq
class HeapGreedySearch:
def __init__(self, data, cls_score):
graph = nx.DiGraph()
graph.add_nodes_from(data.columns)
graph = MatrixGraph.from_networkx_DiGraph(graph, order=data.columns)
self.graph = graph
self.fscore = cls_score(graph, data)
self.ops = []
self.ops += [AddEdge(graph, self.fscore, u, v) for u, v in permutations(graph.nodes(), 2)]
self.ops += [RemoveEdge(graph, self.fscore, u, v) for u, v in permutations(graph.nodes(), 2)]
self.ops += [ReverseEdge(graph, self.fscore, u, v) for u, v in permutations(graph.nodes(), 2)]
self.op_heap = [(-op.score(), op) for op in self.ops]
heapq.heapify(self.op_heap)
def iteration(self):
while len(self.op_heap) > 0:
op = heapq.heappop(self.op_heap)[1]
if op.score() <= 1e-5:
return True
try:
op.do()
self.op_heap = [(-oper.score(), oper) for oper in self.ops]
heapq.heapify(self.op_heap)
op.score()
return False
except InvalidOperation:
pass
return True
def __call__(self, max_iter=40, verbose=True):
counter = 0
while not self.iteration() and counter < max_iter:
if verbose:
print(self.fscore.total())
counter += 1
return DGM(self.graph.to_networkx_DiGraph())
from os import listdir
import os.path
NETWORKS_PATH = '/home/wrwt/Programming/pygraphmodels/networks/'
network_filenames = listdir(NETWORKS_PATH)
true_dgm = gm.DGM.read(os.path.join(NETWORKS_PATH, 'alarm.bif'))
true_dgm.draw()
data = true_dgm.rvs(size=100000)
# %%time
gs = GreedySearch(data, ScoreBIC)
# %lprun -f ScoreBIC.__call__ res = gs(max_iter=100)
gs = GreedySearch(data, ScoreBIC)
# %%time
gs(max_iter=100).draw()
| notebooks/GSSL_heap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EXAMPLE - 7
#
# **Tasks :- Query similarity**
#
# **Tasks Description**
#
# ``Query similarity`` :- This is a sentence pair classification task which determines whether the second sentence in a sample can be inferred from the first.
#
# **Conversational Utility** :- In conversational AI context, this task can be seen as determining whether the second sentence is similar to first or not. Additionally, the probability score can also be used as a similarity score between the sentences.
#
# **Data** :- In this example, we are using the <a href="https://nlp.stanford.edu/projects/snli">SNLI</a> data which is having sentence pairs and labels.
#
# The data can be downloaded using the following ``wget`` command and unzipped using ``unzip`` command.
# !wget qim.fs.quoracdn.net/quora_duplicate_questions.tsv -P qqp_data/
# # Step -1 Data Transformations
#
# Defining transform file
#
# ```
# sample_transform:
# transform_func: qqp_query_similarity_to_tsv
# read_file_names:
# - quora_duplicate_questions.tsv
# read_dir : qqp_data
# save_dir: ../../data
# ```
# !python ../../data_transformations.py \
# --transform_file 'transform_file_qqp.yml'
# # Step -2 Data Preparation
#
# Defining task file for query similarity detection with QQP data
#
# ```
# querysimilarity:
# model_type: BERT
# config_name: bert-base-uncased
# dropout_prob: 0.2
# metrics:
# - classification_accuracy
# loss_type: CrossEntropyLoss
# class_num: 2
# task_type: SentencePairClassification
# file_names:
# - qqp_query_similarity_train.tsv
# - qqp_query_similarity_dev.tsv
# - qqp_query_similarity_test.tsv
# ```
# !python ../../data_preparation.py \
# --task_file 'tasks_file_qqp.yml' \
# --data_dir '../../data' \
# --max_seq_len 200
# # Step - 3 Running train
#
# Following command will start the training for the tasks. The log file reporting the loss, metrics and the tensorboard logs will be present in a time-stamped directory.
#
# For knowing more details about the train process, refer to <a href= "https://multi-task-nlp.readthedocs.io/en/latest/training.html#running-train">running training</a> in documentation.
# !python ../../train.py \
# --data_dir '../../data/bert-base-uncased_prepared_data' \
# --task_file 'tasks_file_qqp.yml' \
# --out_dir 'qqp_query_similarity_bert_base' \
# --epochs 3 \
# --train_batch_size 32 \
# --eval_batch_size 32 \
# --grad_accumulation_steps 2 \
# --log_per_updates 100 \
# --save_per_updates 3000 \
# --limit_save 6 \
# --max_seq_len 200 \
# --eval_while_train \
# --test_while_train \
# --silent
# # Step - 4 Infering
#
# You can import and use the ``inferPipeline`` to get predictions for the required tasks.
# The trained model and maximum sequence length to be used needs to be specified.
#
# For knowing more details about infering, refer to <a href="https://multi-task-nlp.readthedocs.io/en/latest/infering.html">infer pipeline</a> in documentation.
import sys
sys.path.insert(1, '../../')
from infer_pipeline import inferPipeline
| examples/query_pair_similarity/query_similarity_qqp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vptauffer/Ola-Mundo/blob/master/Curso_Libania_06.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="E4m0_1G0CnE8" outputId="0509a975-e95c-4f7e-8376-2ab8cbdfa7e3"
import pandas as pd
dados = pd.read_excel('http://profalibania.com.br/ds/secao05video01.xlsx')
#agrupar dados por categorias
analise = dados.groupby('Sexo').count()
print(analise)
analise = dados.groupby('Sexo').mean()
print(analise)
# Posso limitar as colunas a serem exibidas:
analise = dados[['Sexo','Idade']].groupby('Sexo').count()
print(analise)
# + id="qz4NPFhgHxPe"
| Curso_Libania_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# -*- coding: utf-8 -*-
import dataiku
import pandas as pd, numpy as np
from dataiku import pandasutils as pdu
# Read recipe inputs
titanic_prepared = dataiku.Dataset("Titanic_prepared")
titanic_prepared_df = titanic_prepared.get_dataframe()
# Compute recipe outputs from inputs
# TODO: Replace this part by your actual code that computes the output, as a Pandas dataframe
# NB: DSS also supports other kinds of APIs for reading and writing data. Please see doc.
# titanic_python_df| = titanic_prepared_df # For this sample code, simply copy input to output
# -
titanic_prepared_df
for dataset in titanic_prepared_df:
dataset['Cabin'] = dataset['Cabin'].str[:1]
# Write recipe outputs
titanic_python = dataiku.Dataset("titanic_python")
titanic_python.write_with_schema(titanic_python_df)
| ipython_notebooks/.ipynb_checkpoints/notebook editor for compute_titanic_python-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Fitting a model not (yet) in FRApy
#
# Imagine you have a very nice lensed galaxy, where you measure the metallicity, and you suspect that your metallicity might flatten at higher radius. You'd like to try that hypothesis, fitting this model and comparing it with a 'normal' gradient, but, alas!, FRApy doesn't have a *Flatten_Gradient* model!
#
# Good news is, if you are thinking of an analytical model that depends only on the distance to the galactic centre (radius) and/or the angular position, something like:
#
# $$model_{new}(r,\theta) = cte1 * arbritatly\_weird\_function(r) + cte2 * arbritatly\_weird\_function(\theta) + cte3 ... $$
#
# you might be able to get FRApy to fit it.
#
# All models in FRApy are based in on 'mother of models' called BaseModel. This class takes care of the lensing part and outputs a projected distance map and an azimuthal map that you can use in your new model.
#
# However, you have to add a bit extra magic to make it work.
#
# Here is an empty Model class that we will fill in:
class Awesome_New_Model(BaseModel):
""" Awesome New Model.
Documentation is nice :)
Parameters
----------
cx: int
x position of the centre (in pixels)
cy: int
y position of the centre (in pixels)
q: float
axis ratio (a/b)
pa: float
Position angle (0 North, +90 East )
ADD YOUR EXTRA PARAMETERS HERE
"""
def __init__(self,zlens,dplx_path,dply_path,cx=0,cy=0,q=1,pa=0,v_t =100,r_t=10):
BaseModel.__init__(self,zlens,dplx_path,dply_path,cx=0,cy=0,q=1,pa=0)
#self.extra_par = extra_par
def model_name():
"""Returns the model's name"""
return 'no_name_yet'
def model_parameters(self,verbose=True):
"""Returns the model's parameters"""
if verbose:
print('cx: x position of the centre (in pixels)')
print('cy: y position of the centre (in pixels)')
print('q: axis ratio (a/b)')
print('pa: position angle (in degrees)')
# ADD PARAMETERS HERE
return ['cx','cy','q','pa']
def print_parameter_values(self):
"""Returns the model's parameters values"""
print('cx: %d'%self.cx)
print('cy: %d'%self.cy)
print('q: %0.2f'%self.q)
print('pa: %0.2f'%self.pa)
## ADD PARAMETERS HERE
def update_model_parameters(self,par):
"""Updates the parameters of the model.
Parameters
----------
par: dictionary
dictionary in the shape {'name':parameter_name, 'value':parameter value}
"""
for name in par.keys():
if name == 'cx':
self.cx = par[name]['value']
if name == 'cy':
self.cy = par[name]['value']
if name == 'q':
self.q= par[name]['value']
if name == 'pa':
self.pa = par[name]['value']
## ADD PARAMETERS HERE
def make_model(self):
""" Makes a model using the current parameters' values and stores it
in the 'data' attribute"""
# WHERE THE MAGIC HAPPEN
return model
| examples/fitting_with_a_new_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/galeone/italian-machine-learning-course/blob/master/Introduzione_a_TensorFlow_2_0_e_TensorFlow_Datasets.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="R5qcyzY9Bbcl" colab_type="code" colab={}
# %tensorflow_version 2.x
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="odivAp-OBtuF" colab_type="text"
# Google Colab ci fornisce un ambiente pronto all'uso per usare TensorFlow 2.0.
# Nella cella precedente abbiamo usato il magic-command (comando non Python, ma specifico dei Jupyter Notebook) `%tensorflow_version` per impostare nel runtime corrente l'uso di TensorFlow 2.x.
#
# TensorFlow 2.0 è eager by default: ogni riga di codice è eseguita in ordine sequenziale, esattamente così com'è scritta.
#
# TensorFlow 1.x, invece, utilzzava un diverso paradigma di programazione. Infatti, TensorFlow era un framework che funzionava in modo *descrittivo*: prima veniva definita la computazione, e solo in seguito eseguita "all-at-once".
#
# Usare TensorFlow 1.x all'interno di Jupyter Notebook era molto scomodo, ma fortunatamente la nuova versione permette di utilizzare ogni tool creato per Python (come i notebook), essendo TensorFlow 2.0 molto più "pythonico".
#
# ## Obiettivo
#
# L'obietivo di questo notebook è risolvere un problema di classificazione su immagini, usando una rete completamente connesssa.
#
# Gli step da seguire, propri di ogni buona pipeline di sviluppo di progetti Machine Learning sono:
#
# - Ottenere ed Analizzare i dati
# - Definire la pipeline di input
# - Definire il modello
# - Definire le metriche
# - Definire il training loop
# - Allenare il modello e misurare le metriche durante ed alla fine di ogni epoca
# - Selezionare il modello migliore (basandosi sulla metrica di validation)
# - Misurare le performance sullo split di test
#
# Il tutto utilizzando come strumento di data visualization principale **TensorBoard**.
#
# ## Ottenere ed Analizzare i dati
#
# In un caso d'uso reale, avere un dataset di dati etichettati è un processo lungo e noiso (che qualcuno però deve fare); fortunatamente, per sperimentare diversi algoritmi di machine learning, esistono dataset (solitamente prodotti da universià o industrie) che sono diventati lo standard.
#
# TensorFlow, ha deciso di standardizzare e semplificare il processo di ottenimento dei dataset mediante la libreria [**TensorFlow Datasets** (tfds)](https://www.tensorflow.org/datasets/).
#
# Anziché dover manualmente scaricare e processare i dati, dai siti delle università/industrie, possiamo usare tfds, per (automaticamente):
#
# - scaricare il dataset di dati grezzi
# - applicare trasformazioni ai dati grezzi in modo tale da renderli usabili
# - trasformare questi dati in `TFRecord` (formato ottimizzato per i dati usato da TensorFlow)
# - ottenere un oggetto `tf.data.Dataset` (oggetti per pipeline di input altemete efficienti) pronto all'uso.
#
# Essendo una libreria separata, è necessario scaricarla ed installarla nel sistema usando pip:
#
# + id="KlDb8_4UDcsl" colab_type="code" colab={}
# ! pip install --upgrade tensorflow_datasets
# + [markdown] id="wokDCe4YK7xH" colab_type="text"
# Siamo ora pronti per conoscere TensorFlow datasets.
#
# La libreria è molto semplice e pratica da usare: tutto si basa sul concetto di Dataset Builder. Un dataset builder è una classe (implementata all'interno di tfds) che contiene tutto il processo logico per scaricare, trasfromare, ed ottenere il dataset sotto forma di oggetto `tf.data.Dataset`.
#
# Vedere la lista dei builders (e quindi dei dataset) disponibili è semplice:
# + id="4GUJ7cjPKjYN" colab_type="code" colab={}
import tensorflow_datasets as tfds
print(tfds.list_builders())
print(len(tfds.list_builders()))
# + [markdown] id="f7tunfZ-RbVB" colab_type="text"
# I dataset disponibili sono ~100. Per ognuno di questi è disponibile una descrizione completa sul [catalogo online](https://www.tensorflow.org/datasets/catalog/overview).
#
# Per sperimentare i nostri modelli di classificazione, scegliamo di utilizzare il dataset `"cifar10"`.
#
# Questo dataset è un dataset tipicamente utilizzato per fare benchmark di algoritmi di computer vision ed è composto da immagini a colori 32x32.
#
# TensorFlow datasets ci offre, mediante il metodo load, si ottenere sia il dataset che le **informazioni** relative al tipo di dati che questo contiene.
# + id="0xz7NBrHLnRt" colab_type="code" colab={}
data, info = tfds.load("cifar10", with_info=True, split=tfds.Split.ALL)
# + id="BEU7C5vVSkKW" colab_type="code" colab={}
print(info)
# + [markdown] id="FqsDDNP5T8em" colab_type="text"
# Grazie all'oggetto `DatasetInfo` abbiamo già una prima analisi del dataset:
#
# Il dataset viene fornito direttamente con degli split:
#
# - ci sono 50000 immagini di train
# - ci sono 10000 immagini di test
# - **non c'è un validation set** (dovremmo crearlo noi)
# - le immagini sono `32 x 32 x 3` ed il loro tipo è `tf.uint8` (il ché implica valori in [0,255])
# - le label sono 10 ed il tipo è `tf.int64` (uno scalare, non codificato one-hot)
#
# Leggendo l'[API reference di TensorFlow Datasets](https://www.tensorflow.org/datasets/api_docs/python/tfds) è possibile trovare diverse funzioni messe a nostra disposizione per poter visualizzare ed analizzare il dataset.
#
# Una delle più interessanti [`tfds.show_examples`](https://www.tensorflow.org/datasets/api_docs/python/tfds/show_examples) che dato un dataset e le sue informazioni, ci permette di visualizzare direttamente in un notebook (in quanto ritorna un oggetto matplotlib) alcuni samples dal dataset:
#
# + id="mjcMGbIHTI_T" colab_type="code" colab={}
fig = tfds.show_examples(info, data, rows=4, cols=4)
# + [markdown] id="gKdhsIN6XTVa" colab_type="text"
# Come visibile dall'immagine, all'interno del datataset è presente la coppia immagine label, ed all'interno dell'oggetto info, invece, è presente la relazione che lega la label testuale alla label numerica.
#
# Dall'API documentation è possibile trovare i metodi [str2int](https://www.tensorflow.org/datasets/api_docs/python/tfds/features/ClassLabel#str2int) ed [int2str](https://www.tensorflow.org/datasets/api_docs/python/tfds/features/ClassLabel#int2str) che permettono di passare da stringa a label e viceversa.
#
# Essendo TensorFlow 2.0 eager by default, possiamo iniziare ad utilizzarlo per creare un loop su oggetti di tipo `tf.Tensor`, prodotti all'operazione `tf.range` (equivalente alla `range` di Python). Nel loop visualizziamo la relazione tra label numerica e stringa:
# + id="9359Pb-bVZQk" colab_type="code" colab={}
for label in tf.range(10):
print(label.numpy(), " -> ", info.features["label"].int2str(label))
# + [markdown] id="B--amRprdkay" colab_type="text"
# ## Dataset API: definire la pipeline di input
#
# `data` è un oggetto di tipo `tf.data.Dataset`: la dataset API è **ottimizzata** per creare pipeline di input per il train di modelli.
#
# L'API è basata sul **method chaining**: i metodi dell'oggetto dataset, applicano trasformazioni al dataset corrente, e ritornano un dataset con la trasofmrazione applicata.
#
# La dataset API rappresenta correttamente il processo di ETL (Extract-Transform-Load) tipici di una pipeline di data science.
#
# - TensorFlow Datasets è incaricato dell'estrazione dei dati e della prima trasformazione
# - tf.data.Dataset con i suoi metodi applica la serie di trasformazioni atte a rendere utili i dati
# - L'iterazione sull'oggetto dataset è il load dei dati in memoria
#
# 
#
# La pipeline di trasformazioni che vogliamo applicare è questa:
#
# - trasformare i dati da uint a float
# - codificare one-hot le label
# - scalare le immagini nel range [-1,1]
# - "appiattire" (flatten) le immagini, per renderle anziché tensori `32 x 32 x 3`, dei tensori `32*32*3`
# - Creare gli split di train, validation, test (tre oggetti `tf.data.Dataset`)
# - Per ognuno di questi: creare dei batch di dimensione 32 per poter fare, successivamente, mini-batch gradient descent / valutazione in batch
# - ottimizzare le performance della pipeline
# + id="ltmyq97VZc3V" colab_type="code" colab={}
def transform(row):
# trasformare i dati da uint a float
row["image"] = tf.image.convert_image_dtype(row["image"], dtype=tf.float32)
# 1-hot
row["label"] = tf.one_hot(row["label"], depth=10, on_value=1, off_value=0)
# [-1,1] range
row["image"] = (row["image"] - 0.5) * 2.
# flatten
row["image"] = tf.reshape(row["image"], (-1,))
return row
dataset = data.map(transform)
# split, batch, prefetch
train = dataset.take(50000).batch(32).prefetch(1)
validation = dataset.skip(50000).take(5000).batch(32).prefetch(1)
test = dataset.skip(50000 + 5000).take(5000).batch(32).prefetch(1)
# + [markdown] id="D0W9cY6QsRCu" colab_type="text"
# ## Definizione del modello: Keras API
#
# Keras è un API specification per la definizione ed il training di modelli di machine learning che TensorFlow ha deciso di adottare.
#
# L'API è molto intuitiva da usare e si trova all'interno del modulo [`tf.keras`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/).
#
# Keras offre tre differenti modi per creare un modello:
#
# - Sequential API
# - Functional API
# - Subclassing
#
# I layer offerti da Keras sono i più disparati: dai layer dense, ai layer di attivazione, ai layer di batch normalization, a quelli di convoluzioni per lavorare su immagini o pointcloud e molti altri.
#
# Ogni layer è una classe da poter istanziare e configurare tramite i parametri del costruttore.
#
# Per esempio, il costruttore del layer `tf.keras.layers.Dense` ha la seguente firma:
#
# ```python
# __init__(
# units,
# activation=None,
# use_bias=True,
# kernel_initializer='glorot_uniform',
# bias_initializer='zeros',
# kernel_regularizer=None,
# bias_regularizer=None,
# activity_regularizer=None,
# kernel_constraint=None,
# bias_constraint=None,
# **kwargs
# )
# ```
#
# Nella quale possiamo specificare *ogni cosa* relativa all funzionamento del layer: dal numero di unità (neuroni), all'uso o meno del bias, al tipo di inizializzazione dei parametri.
#
# Dato che la definizione di un modello è completamente arbitraria, possiamo provare a partire con un semplice modello (pochi neuroni per layer, per evitare l'overfitting), che riduce la dimensionalità dell'input layer per layer, fino ad arrivare a 10 neuroni di output (le classi).
#
# Un semplice modello fully connected può essere visto come uno stack di layer `Dense`, ed è il caso d'uso canonico della Sequential API.
#
# ## Sequential API
# + id="6R_ekD8BdyyM" colab_type="code" colab={}
model = tf.keras.Sequential([
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10) # Note the linear activation
])
# + [markdown] id="_qsV69zES_uS" colab_type="text"
# Keras permette opzionalmente, di specificare la dimensione dell'input in fase di definizione del modello, oppure di lasciare che sia il Keras model alla prima esecuzione a determinarla in maniera automatica.
#
# Questo può sembrare un particolare da poco, ma in realtà è qualcosa di fondamentale.
#
# Conoscere a priori la dimensione dell'input permette di definire il **grafo computazionale** del modello completamente, e quindi poter visualizzare il "riassunto" completo del modello.
#
# Ogni keras model offre il metodo `.summary()` per ottenere una visualizzazione tablellare della struttura del modello, ma per ottenre il numero dei parametri del primo layer, è **sempre** necessario conoscere la dimensione di input.
#
# Infatti, per poter completamente definire la *matrice* dei pesi del primo layer, è necessario non solo il numero di neuroni, ma anche il numero di dimensioni dell'input.
#
# Difatti, se proviamo a invocare il metodo `.summary()` sul modello appena creato, otteniamo il seguente errore:
# + id="_OE7LXwBUvKJ" colab_type="code" colab={}
model.summary()
# + [markdown] id="ATdo0HPtUxWe" colab_type="text"
# Per poter visualizzare il summary completo, non avendo definito come attributo `input_shape` del primo layer, dobbiamo effettuare un **forward pass** del modello con un tensore di input (della dimensione coretta), in modo tale che Keras possa costrure il grafo computazionale.
# + id="scUH4OK1Uwfh" colab_type="code" colab={}
fake_input = tf.zeros((1, 32*32*3))
out = model(fake_input)
model.summary()
# + [markdown] id="53JaDxgR1hBi" colab_type="text"
# Il numero di parametri è stato correttamente calcolato (e come è possibile vedere, per un modello così semplice siamo già oltre il milione di parametri), sebbene l'`output shape` risulti "multiple", anziché essere del valore corretto.
#
# La **raccomadazione** è di specificare **sempre** in fase di creazione del modello la dimensione dell'input, un modo tale da poter ottenere summary rappresentativi ed aiutare Keras nella definizione del modello stesso.
#
# Possiamo quindi sovrascrivere il modelo precedente, creandolo ex-novo, ma specificando l'input shape. Per specificarla abbiamo due modi:
#
# - O usare un `tf.keras.layers.Input` layer
# - O usara il parametro del costruttore del primo layer dense `input_shape`
# + id="sBE-MrAKVHU7" colab_type="code" colab={}
model = tf.keras.Sequential([
tf.keras.layers.Input(shape=(32*32*3)),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10) # Note the linear activation
])
# + id="YEnIcVX-2hJG" colab_type="code" colab={}
model.summary()
# + id="xt3qN2Sa439n" colab_type="code" colab={}
tf.keras.utils.plot_model(model)
# + [markdown] id="H9eJCt3q5bX_" colab_type="text"
# ## Functional API
#
# Un modo differente per definire i modelli, è quello di usare la functional API.
#
# Ogni layer Keras è un oggetto **callable**: questo significa che è possibile utilizzare un oggetto istanziato come se fosse una funzione, che accetta un input e produce un output.
#
# Per un modello con un singolo input ed un singolo output, totalmente sequenziale (come il nostro) non è necessario utlizzarla, i quanto Sequential soddisfa pienamente ogni requisito.
#
# In ogni modo, essendo l'API più flessibile offerta da Keras per definire modelli, è bene conoscerla ed utlizzarla il più possibile, in modo tale da essere familiari con questa API quando si definiranno modelli con più input, più output e con relazioni tra i layer del modello.
# + id="LKOJ-qJC5HYH" colab_type="code" colab={}
inputs = tf.keras.layers.Input(shape=(32*32*3))
net = tf.keras.layers.Dense(512, activation=tf.nn.relu)(inputs)
net = tf.keras.layers.Dense(256, activation=tf.nn.relu)(net)
net = tf.keras.layers.Dense(128, activation=tf.nn.relu)(net)
out = tf.keras.layers.Dense(10)(net)
model = tf.keras.Model(inputs=inputs, outputs=out)
model.summary()
# + [markdown] id="zcsKI8VL8AN-" colab_type="text"
# ## Loss function
#
# All'interno del modulo [`tf.keras.losses`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/losses) troviamo una lunga lista di loss function pronte all'uso.
#
# Le loss disponibili sono tra le più disparate e scegliere quella corretta dipende da:
#
# - il problema che stiamo risolvendo (classificazione, regressione, ...)
# - il formato delle nostre label
#
# Dato che:
#
# - abbiamo codificato in one-hot le label
# - **non** abbiamo applicato una non-linearità al layer di outut del modello (volutamente)
#
# La nostra scelta deve ricadere sulle funzioni: **non** sparse e che accettano (unscaled) **logits** come input.
#
# TensorFlow, per le loss, utilizza le keywords **sparse** e **logits** per indicare se la loss function accetta label scalari (non one-hot, ed applica la loss function stessa la conversione all'interno) e output di modelli **lineari**.
#
# Quando la loss function accetta label scalari, allora è la loss function stessa che al suo interno applica la conversione a rappresentazione one-hot.
#
# Quando la loss function accetta (o permette di specificare) `from_logits=True` significa che sarà la loss function stessa a applicare la non linearità corretta all'output del modello per il calcolo della loss.
#
# Ad esempio, per un problema di classificazione multi-classe, con label rappresentate in one-hot, la loss function che viene utilzzata è la **categorical cross-entropy loss**.
#
# Quello che vogliamo, è allenare la rete neurale per **produrre una probabilità su C classi** (10 in questo caso) data un immagine di input.
#
# La loss calcola la **softmax activation** sull'output della rete (per riscalare i valori di output nel range probabilistico [0,1]) e dopo calcola la cross-entropy-loss.
#
# **Softmax**
#
# Softmax è una funzione di attivazione che riscala i valori di output di un classificatore nel range [0,1], in modo tale che la somma di tutti i valori predetti sia 1.
#
# La softmax activation viene applicata agli **score** predetti dalla rete *s*; dato che gli elementi predetti rappresentano delle classi, questi score possono essere interpretati come probabilità (predizione aereoplano con probabilità 0.8, macchina con probabilità 0.1, ecc).
#
# Per una data classe $s_i$, la funzione softmax viene calcolata come
#
# $$ f(s)_i = \frac{e^{s_i}}{\sum_{j}^{C}{e^{s_j} }} $$
#
# **Cross entropy loss**
#
# La formula della categorical cross-entropy loss è data dall'applicazione della cross-entropy tra le label $t_i$ (one-hot) e le predizioni dopo il softmax.
#
# $$ CE = - \sum_{i}^{C}{ t_i log(f(s)_i) } $$
#
# Dato che la label sono codificate in one hot, solo il componente del vettore dove il valore è 1 concorre al calcolo della loss, mentre tutti gli altri valgono zero.
#
# Dato il target vector (label reale), codificato in one hot $t$ e la sua componente ad uno nella posizione $p$ (quindi $t_p$), abbiamo che la formulazione della cross-entropy diventa:
#
# $$ CE = -log\left(\frac{e^{s_p}}{\sum_{j}^{C}{e^{s_j}}}\right) $$
#
# ### Implementazione
#
# Keras si occupa di realizzare **tutto** il calcolo della loss nella maniera più ottimizzata possibile e numericalmente stabile.
#
# Come è facile notare, dato che è la loss stessa ad applicare la funzione di attivazione (**softmax**) all'output della rete, quando abbiamo definito il modello abbiamo evitato di aggiungerla all'output layer.
# + id="QdIsvd6C7eH9" colab_type="code" colab={}
# Loss is a callable object
loss = tf.keras.losses.CategoricalCrossentropy(from_logits=True)
# + [markdown] id="iJy7dleJCnVY" colab_type="text"
# Abbiamo il modello `model`, abbiamo il dataset di train `train`, abbiamo la loss function da usare `loss`, ciò che rimane è la definizione del training loop, con annesse scelta dell'ottimizzatore e delle metriche da misurare.
#
# ## Training loop
#
# TensorFlow 2.0 offre una maniera "avanzata" per definire ed implementare training loop.
#
# Keras, d'altro canto, offre la sua maniera. In questo corso **non tratteremo** il modo Keras di definire ed eseguire i training loop, in quanto **nascondono troppi dettagli** e sono poco flessibili.
#
# L'implementazione di un "custom training loop" è considerata avanzata, ma in realtà non è nulla di complesso. Anzi, avere controllo sulla fase di forward pass e di calcolo ed applicazione dei gradienti è utile per apprendere in maniera migliore il processo di training ed è senza dubbio l'opzione più flessibile.
#
# ### Metriche
#
# Prima di definire il training loop, è bene scegliere che metriche misurare per tenere monitorate le performance ed identificare eventuali condizioni patologiche.
#
# Dato che il dataset è bilanciato, possiamo misurare **l'accuracy**
# Inoltre, possiamo misurare il valore della loss medio (sul batch) durante il training loop.
#
# TensorFlow offre metriche per misurare il valore medio di un qualsiasi scalare che varia nel tempo (`tf.keras.metrics.Mean`), oppure per il calcolo del valore medio di una specifica metrica (e.g. `tf.keras.metrics.Accuracy`).
#
# Ogni metrica implementa l'interfaccia standard di Keras relative alle metriche: questo ci garantisce che ogni oggeto appartenente al modulo `tf.keras.metrics` abbia i metodi:
#
# - `update_state` (identico al metodo `__call__`): per computare la metrica
# - `result` per ottenere il valore della metrica computato fin'ora
# - `reset_state` per resettare lo stato della metrica al valore iniziale
# + id="7FmbvgWLChd3" colab_type="code" colab={}
accuracy = tf.keras.metrics.Accuracy()
mean_loss = tf.keras.metrics.Mean(name="loss")
# + [markdown] id="iGJwZTHwcceS" colab_type="text"
# ## Loggare le metriche
#
# Per loggare le metriche abbiamo due opzioni, da implementare **sempre** assieme quando si definisce una pipeline di ML (ben definita):
#
# - loggare su stanard output/error
# - loggare su **TensorBoard**
#
# TensorBoard è un programma che viene installato assieme al modulo tensorflow e permette di visualizzare su grafici curve, istogrammi, dataset embedding, immagini, tracce audio, e molto altro.
#
# È perfettamente integrato con i Jupyter notebook e tramite il magic command `%tensorboard` è possibile lansciare un'istanza del tensorboard server.
#
# TensorBoard necessita di una cartella da "monitorare": all'interno di questa cartella vanno inseriti tutti i dati da loggare (summary).
#
# TensorFlow, tramite il modulo `tf.summary` da la possibilità di salvare su file i valori delle metriche, delle immagin utilizzate e di ogni altro tipo di dato che il modulo `tf.summary` supporta.
#
# Il concetto fondamentale per poter correttamente utilizzare i summary è quello di `FileWriter`.
#
# Questo oggetto permette di creare un **contesto** e tramite questo, scrivere i dati all'interno della cartella monitorata da tensorboard.
# Il **contesto** è fondamentale per una **buona organizzazione dei log**; infatti, è possibile creare un contesto di train, uno di validation ed uno di test, e visualizzare sullo stesso plot curve (in colori differenti) relative a contesti differenti.
#
# Ad esempio, dato un `FileWriter` `writer`, possiamo definire un contesto tramite a keyword python `with` e scrivere all'interno del contesto creato dal writer, in questo modo:
#
# ```python
# with writer.as_default():
# for step in range(100):
# # other model code would go here
# tf.summary.scalar("my_metric", 0.5, step=step)
# writer.flush()
# ```
#
# Per il nostro caso, possiamo creare tre writer differenti
# + id="Du81dKeCcbkZ" colab_type="code" colab={}
train_writer = tf.summary.create_file_writer("logs/train")
validation_writer = tf.summary.create_file_writer("logs/validation")
test_writer = tf.summary.create_file_writer("logs/test")
# + [markdown] id="-uY_E3csxvh6" colab_type="text"
# Avendo definito le metriche ed i file writer, siamo pronti a definire il training loop.
#
# ## Training loop
#
# Il loop di training consiste in due parti. Durante ogni epoca di training, dobbiamo calcolare il valore della loss sul batch, **tenere traccia** delle operazioni effettuate durante il calcolo.
#
# Tenere traccia delle operazioni fatte è di fondamentale importanza, in quanto possiamo utilizzare queste informazioni per calcolare **il gradiente**, quindi stimare la direzione dell'aggioranamento, ed usare un **ottimizzatore** per applicare l'update dei parametri nella direzione stimata.
#
# TensorFlow 2.0 ci aiuta nella modularizzazione del codice: essendo eager by default, possiamo scrivere funzioni Python che effettuano determinate operazioni e mediante **tf.function** è possibile anche accelerare il calcolo di alcune di queste, convertendo il codice in una rappresentazione a grafo altamente ottimizzata.
# + id="zV4zud5gqBCc" colab_type="code" colab={}
# Define the optimizer
optimizer = tf.keras.optimizers.SGD(learning_rate=1e-3)
def compute_loss(input_samples):
predictions = model(input_samples["image"])
loss_value = loss(input_samples["label"], predictions)
return loss_value
@tf.function
def train_step(input_samples):
with tf.GradientTape() as tape:
loss_value = compute_loss(input_samples)
gradient = tape.gradient(loss_value, model.trainable_variables)
optimizer.apply_gradients(zip(gradient, model.trainable_variables))
return loss_value
def measure_metrics(input_samples):
predicted_labels = tf.argmax(model(input_samples["image"]), axis=1)
accuracy.update_state(tf.argmax(input_samples["label"], axis=1), predicted_labels)
mean_loss.update_state(compute_loss(input_samples))
# + [markdown] id="EksCULOa-__E" colab_type="text"
# Dopo aver definito i "macroblocchi" del nostro train, possiamo definire direttamente il training loop.
# + id="7Nf07TQy-_Ct" colab_type="code" colab={}
global_step = tf.Variable(0, dtype=tf.int64, trainable=False)
epoch_counter = tf.Variable(0, dtype=tf.int64, trainable=False)
def train_loop(num_epochs):
for epoch in tf.range(epoch_counter, num_epochs):
for input_samples in train:
loss_value = train_step(input_samples)
measure_metrics(input_samples)
global_step.assign_add(1)
if tf.equal(tf.math.mod(global_step, 100), 0):
mean_loss_value = mean_loss.result()
accuracy_value = accuracy.result()
mean_loss.reset_states()
accuracy.reset_states()
tf.print(f"[{global_step.numpy()}] loss value: ", mean_loss_value," - train acc: ", accuracy_value)
with train_writer.as_default():
tf.summary.scalar("loss", mean_loss_value, step=global_step)
tf.summary.scalar("accuracy", accuracy_value, step=global_step)
tf.summary.image("images", tf.reshape(input_samples["image"], (-1, 32,32,3)), step=global_step, max_outputs=5)
# end of epoch: measure performance on validation set and log the values on tensorboard
tf.print(f"Epoch {epoch.numpy() + 1 } completed")
epoch_counter.assign(epoch + 1)
# TODO: insert validation code here
# + [markdown] id="svUBzxG7B7Io" colab_type="text"
# Dopo aver definito la funzione di train, con annessa misura delle performance di train e validation, possiamo lanciare tensorboard e subito dopo invocare la funzione di train.
# + id="efDU90l0B5NQ" colab_type="code" colab={}
# %load_ext tensorboard
# %tensorboard --logdir logs
# + id="ry0wTMidCGem" colab_type="code" colab={}
train_loop(num_epochs=2)
# + [markdown] id="s_BqJD_px14K" colab_type="text"
# ## Quali sono i problemi di questo training loop?
#
# - Non stiamo misurando le performance di validation durante il training [**esercizio 1**]
# - Al termine del train non stiamo misurando le performance sul test set [**esercizio 2**]
# - Non c'è persistenza del modello: se il train deovesse interrompersi per qualsiasi motivo (fallimento hardware e simili) dovremmo re-iniziare il training dall'inizio, in quanto **non abbiamo salvato lo stato del modello**. [sezione successiva]
# - C'è un problema con la visualizzazione delle immagini in TensorBoard: la funzione `tf.summary.image` si aspetta immagini scalate i [0,1] mentre le nostre immagini sono state riscalate in [-1,1] durante la `map_fn`. [**esercizio 3**]
# - Non effettuiamo alcuna model selection: non gestendo la persistenza del modello e non misurando le performance sul validation set, non abbiamo tenuto monitorato l'overfitting (quando le performance di train sono troppo migliori delle performance di validation) [**esercizio 4**]
#
# Alcuni di questi punti sono facilmente risolvibili con le conoscenze acquisite fin'ora, ma per gestire la persistenza è necessario introdurre il concetto di **training checkpoints**.
#
# ## Training Checkpoints
#
# Salvare lo stato di un modello in TensorFlow 2.0 è davvero facile: tutto ciò che è necessario è craere un oggetto checkpoint ed assegnargli (direttamente nel costruttore) gli oggetti che vogliamo salvare.
#
# Molti oggetti TensorFLow 2.0 sono "checkpointable", il ché significa che sono salvabili su disco dall'oggetto checkpoint (aka sono serializzabili).
#
# Per usare un checkpoint è necessario un `CheckpointManager` che permette di gestirli.
#
# esempio:
#
# ```python
# ckpt = tf.train.Checkpoint(step=tf.Variable(1), optimizer=opt, net=net)
# manager = tf.train.CheckpointManager(ckpt, './tf_ckpts', max_to_keep=3)
# ckpt.restore(manager.latest_checkpoint)
# if manager.latest_checkpoint:
# print("Restored from {}".format(manager.latest_checkpoint))
# else:
# print("Initializing from scratch.")
# ```
#
# Nel nostro esempio, vogliamo salvare lo stato del modello, dell'ottimizatore (anche se non stiamo usando un ottimizzatore che definisce variabili, ma è lo stesso una buona pratica) e il global step, così da poter riprendere il train dallo step esatto in cui è stato interrotto.
# + id="ZXnvkivWCOfB" colab_type="code" colab={}
ckpt = tf.train.Checkpoint(step=global_step, optimizer=optimizer, model=model)
manager = tf.train.CheckpointManager(ckpt, 'ckpts', max_to_keep=3)
ckpt.restore(manager.latest_checkpoint)
if manager.latest_checkpoint:
tf.print(f"Restored from {manager.latest_checkpoint}")
else:
tf.print("Initializing from scratch.")
# + [markdown] id="YmcKV28feimP" colab_type="text"
# Avendo associato un CheckpointManager ad un oggetto `Checkpoint`, possiamo usarlo per salvare/ripristinare lo stato del modello.
#
# Il manager altro non fa' che creare un'associazione tra l'oggetto `Checkpoint` **ed una cartella** (`ckpts`) dove verranno salvati gli oggetti "attaccati" al checkpoint.
#
# Il metodo da utilizzare per salvare lo stato corrente è il metodo `.save()` del manager:
# + id="koZKhm5f0JWV" colab_type="code" colab={}
manager.save()
ckpt.restore(manager.latest_checkpoint)
if manager.latest_checkpoint:
tf.print(f"Restored from {manager.latest_checkpoint}")
# + [markdown] id="rFdzxH5qfsSF" colab_type="text"
# ## Esercizio 1
#
# Crea una nuova funzione `training_loop_v2` che aggiunga le misure delle performance sul validation set alla fine di ogni epoca. Usare il `FileWriter` corretto e gestire correttamente lo stato delle metriche.
# Scrivere i risultati su tensorboard e verificare se una nuova curva appare sullo stesso grafico del training.
#
# + id="KotsNqqxe3rd" colab_type="code" colab={}
global_step = tf.Variable(0, dtype=tf.int64)
def train_loop(num_epochs):
for epoch in tf.range(num_epochs):
for input_samples in train:
loss_value = train_step(input_samples)
measure_metrics(input_samples)
global_step.assign_add(1)
if tf.equal(tf.math.mod(global_step, 100), 0):
mean_loss_value = mean_loss.result()
accuracy_value = accuracy.result()
mean_loss.reset_states()
accuracy.reset_states()
tf.print(f"[{global_step.numpy()}] loss value: ", mean_loss_value," - train acc: ", accuracy_value)
with train_writer.as_default():
tf.summary.scalar("loss", mean_loss_value, step=global_step)
tf.summary.scalar("accuracy", accuracy_value, step=global_step)
tf.summary.image("images", tf.reshape(input_samples["image"], (-1, 32,32,3)), step=global_step, max_outputs=5)
# end of epoch: measure performance on validation set and log the values on tensorboard
tf.print(f"Epoch {epoch.numpy() + 1 } completed")
# TODO: insert validation code here
# + [markdown] id="lMJCRpgpgK7j" colab_type="text"
# ## Esercizio 2
#
# Definire una funzione `test()` che misuri le performance sul test set ed invocarla al seguito dell'esecuzione della funzione `training_loop_v2` eseguita per 10 epoche.
# + id="cBges1cLgJnJ" colab_type="code" colab={}
def test():
#TODO
training_loop_v2(num_epochs=10)
test()
# + [markdown] id="ZWhJh7nbgbHi" colab_type="text"
# ## Esercizio 3
#
# Creare una funzione che scali un tensore a valori in [0,1] in un tensore a valori in [-1,1] (codice già presente nel notebook).
# Creare una seconda funzione che scali un tensore a valori in [-1,1] in [0,1]: usare `tf.summary.image` con immagini scalate nel range corretto ([0,1]): aggiornare ed eseguire le funzioni di training loop.
# + id="IkP5OCR0gKRY" colab_type="code" colab={}
def rescale(image):
return (image + 1.) / 2.
# + [markdown] id="9aoM4fI8hAb8" colab_type="text"
# ## Esercizio 4
#
# La funzione `training_loop_v2` misura correttamente l'accuracy sul training set e sul validation set.
#
# Modificare il codice della funzione per:
#
# - ripristinare lo stato del modello dall'ultimo training step raggiungo prima di iniziare il training loop sulle nuove epoche richieste (usare un checkpoint ed un checkpoint manager per salvare il modello al termine di ogni epoca)
# - dopo aver misurato l'accuracy di validation e l'accuracy di train (al termine di ogni epoca), usare un **diverso** checkpoint e checkpoint managert (su una diversa folder) per salvare il modello che ha raggiunto la miglior validation accuracy.
# - Se per 2 epoche consecutive, le performance di train sono migliori delle performance di validation interrompere il training (early stopping basato sul confronto delle metriche)
# + id="Fl48dFjCg3aJ" colab_type="code" colab={}
# + [markdown] id="dUzbNq2ZiCTE" colab_type="text"
# ## Esercizio 5
#
# Creare un nuovo notebook per risolvere lo stesso problema, ma:
#
# - non usare la funzione di one-hot encoding per le label, ma delegare l'encoding alla Keras loss adeguata: modificare ogni parte del codice necessaria ad usare le label scalari, anziché la rappresentazione one-hot.
#
# ## Esercizio 6
#
# Sperimentare!
#
# - Provare come variano le performance al variare del learning rate
# - Cambiare dataset, scegliendo tra altri presenti in TensorFlow datasets (adeguare quindi il modello)
# - Sperimentare come variano le performance al variare dell'ottimizzatore (vedesi lista degli ottimizzatori [nella documentazione](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/optimizers))
# - Variare l'architettura del modello, cambiare numero di neuroni, numero di layer, funzioni di attivazione, struttura, ...
#
# ## Conclusione
#
# La soluzione proposta per questo problema di classificazione di immagini è subottimale: stiamo usando una archietttura fully connected, con milioni di parametri, quando esiste una soluzione ben più efficiente, con un numero minore di parametri e con performance migliori: le reti neurali convoluzionali.
#
# Questi tipo di rete ha rivoluzionato il campo della computer vision e del machine learning in generale: tutt'oggi sono i blocchi fondamentali per la stragrande maggioranza delle architetture che lavorano su immagini, audio (e anche per i modelli generativi!).
#
# Nel prossimo notebook introdurremo l'operazione di convoluzione, le reti neurali convoluzionali, definiremo un'architettura deep usando stack di layer convoluzionali e risolveremo non solo il problema della classificazione, ma apprenderemo anche come **localizzare** un oggetto all'interno di una immagine, regredendo le coordinate della bounding box.
| Introduzione_a_TensorFlow_2_0_e_TensorFlow_Datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/girish342/flaskSaaS/blob/master/Ex1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="6tFUAhJCGzp4" colab_type="code" colab={}
import numpy as np
import tensorflow as tf
# + id="tGxuiqRtG_8W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cf99c5f9-a29c-4743-ee01-afd8539a4309"
a = tf.constant(4)
b = tf.constant(5)
c = a+b
print(c)
# + id="WkWXkoGaHTsT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="548ee0c2-cdde-4c5f-9e9f-251bc732cf83"
with tf.Session() as sess:
v1 = sess.run(c)
print(v1)
# + id="iCcY7XJqH5hq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="677551db-6e8d-4127-badc-8a606bad55b1"
a = tf.Variable(1, name = 'input')
b = tf.constant (2)
mid_value = tf.add(a,b)
update_value = tf.assign(a, mid_value)
tg = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(tg)
print(sess.run(a))
for i in range(3):
sess.run(update_value)
print(sess.run(a))
# + id="5vJBvlQRKm5i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="5d5a7396-50bb-480f-cc67-722015aeb2e3"
x = tf.placeholder('float32', [None,3])
y = x**2
with tf.Session() as sess:
result = sess.run(y, feed_dict = {x:
[[1.5,2,3],[3,6,4]]})
print(result)
# + id="ujPI1Z0lOx8-" colab_type="code" colab={}
| Ex1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Types in Python
# By default Python have these data types:
# + active=""
# strings - used to represent text data, the text is given under quote marks. eg. "ABCD"
# + active=""
# integer - used to represent integer numbers. eg. -1, -2, -3
# + active=""
# float - used to represent real numbers. eg. 1.2, 42.42
# + active=""
# boolean - used to represent True or False.
# + active=""
# complex - used to represent a number in complex plain. eg. 1.0 + 2.0j, 1.5 + 2.5j
# -
# # Data Types in NumPy
# + active=""
# NumPy has some extra data types, and refer to data types with one character, like i for integers, u for unsigned integers etc.
# + active=""
# Below is a list of all data types in NumPy and the characters used to represent them.
# + active=""
# i - integer
#
# b - boolean
#
# u - unsigned integer
#
# f - float
#
# c - complex float
#
# m - timedelta
#
# M - datetime
#
# O - object
#
# S - string
#
# U - unicode string
#
# v - fixed chunk of memory for other type(void)
#
# -
# # Checking the Data Type of an Array
# + active=""
# The NumPy array object has a property called dtype that returns the data type of the array:
# -
# Get the data type of an array object:
# +
import numpy as np
arr = np.array([1, 2, 3, 4])
print(arr.dtype)
# -
# Get the data type of an array containing strings:
# +
import numpy as np
arr = np.array(['apple', 'banana', 'cherry'])
print(arr.dtype)
# -
# # Creating Arrays With a Defined Data Type
# + active=""
# We use the array() function to create arrays, this function can take an optional argument: dtype that allows us to define the expected data type of the array elements:
# -
# Create an array with data type string:
# +
import numpy as np
arr = np.array([1, 2, 3, 4], dtype='S')
print(arr)
print(arr.dtype)
# + active=""
# For i, u, f, S and U we can define size as well.
# -
# Create an array with data type 4 bytes integer:
# +
import numpy as np
arr = np.array([1, 2, 3, 4], dtype='i4')
print(arr)
print(arr.dtype)
# -
# # What if a Value Can Not Be Converted?
# + active=""
# If a type is given in which elements can't be casted then NumPy will raise a ValueError.
# + active=""
# NOTE -ValueError: In Python ValueError is raised when the type of passed argument to a function is unexpected/incorrect.
# -
# A non integer string like 'a' can not be converted to integer (will raise an error):
# +
import numpy as np
arr = np.array(['a', '2', '3'], dtype='i')
# -
# # Converting Data Type on Existing Arrays
# + active=""
# The best way to change the data type of an existing array, is to make a copy of the array with the astype() method.
# + active=""
# The astype() function creates a copy of the array, and allows you to specify the data type as a parameter.
# + active=""
# The data type can be specified using a string, like 'f' for float, 'i' for integer etc. or you can use the data type directly like float for float and int for integer.
# -
# Change data type from float to integer by using 'i' as parameter value:
# +
import numpy as np
arr = np.array([1.1, 2.1, 3.1])
newarr = arr.astype('i')
print(newarr)
print(newarr.dtype)
# -
# Change data type from float to integer by using int as parameter value:
# +
import numpy as np
arr = np.array([1.1, 2.1, 3.1])
newarr = arr.astype(int)
print(newarr)
print(newarr.dtype)
# -
# Change data type from integer to boolean:
# +
import numpy as np
arr = np.array([1, 0, 3])
newarr = arr.astype(bool)
print(newarr)
print(newarr.dtype)
| 5. NumPy Data Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series Forecasting
# In this tutorial, we will demonstrate how to build a model for time series forecasting in NumPyro. Specifically, we will replicate the **Seasonal, Global Trend (SGT)** model from the [Rlgt: Bayesian Exponential Smoothing Models with Trend Modifications](https://cran.r-project.org/web/packages/Rlgt/index.html) package. The time series data that we will use for this tutorial is the **lynx** dataset, which contains annual numbers of lynx trappings from 1821 to 1934 in Canada.
# !pip install -q numpyro@git+https://github.com/pyro-ppl/numpyro
# +
import os
import matplotlib.pyplot as plt
import pandas as pd
from IPython.display import set_matplotlib_formats
import jax.numpy as jnp
from jax import random
import numpyro
import numpyro.distributions as dist
from numpyro.contrib.control_flow import scan
from numpyro.diagnostics import autocorrelation, hpdi
from numpyro.infer import MCMC, NUTS, Predictive
if "NUMPYRO_SPHINXBUILD" in os.environ:
set_matplotlib_formats("svg")
numpyro.set_host_device_count(4)
assert numpyro.__version__.startswith("0.9.2")
# -
# ## Data
# First, lets import and take a look at the dataset.
URL = "https://raw.githubusercontent.com/vincentarelbundock/Rdatasets/master/csv/datasets/lynx.csv"
lynx = pd.read_csv(URL, index_col=0)
data = lynx["value"].values
print("Length of time series:", data.shape[0])
plt.figure(figsize=(8, 4))
plt.plot(lynx["time"], data)
plt.show()
# The time series has a length of 114 (a data point for each year), and by looking at the plot, we can observe [seasonality](https://en.wikipedia.org/wiki/Seasonality) in this dataset, which is the recurrence of similar patterns at specific time periods. e.g. in this dataset, we observe a cyclical pattern every 10 years, but there is also a less obvious but clear spike in the number of trappings every 40 years. Let us see if we can model this effect in NumPyro.
#
# In this tutorial, we will use the first 80 values for training and the last 34 values for testing.
y_train, y_test = jnp.array(data[:80], dtype=jnp.float32), data[80:]
# ## Model
# The model we are going to use is called **Seasonal, Global Trend**, which when tested on 3003 time series of the [M-3 competition](https://forecasters.org/resources/time-series-data/m3-competition/), has been known to outperform other models originally participating in the competition:
#
# \begin{align}
# \text{exp-val}_{t} &= \text{level}_{t-1} + \text{coef-trend} \times \text{level}_{t-1}^{\text{pow-trend}} + \text{s}_t \times \text{level}_{t-1}^{\text{pow-season}}, \\
# \sigma_{t} &= \sigma \times \text{exp-val}_{t}^{\text{powx}} + \text{offset}, \\
# y_{t} &\sim \text{StudentT}(\nu, \text{exp-val}_{t}, \sigma_{t})
# \end{align}
#
# , where `level` and `s` follows the following recursion rules:
#
# \begin{align}
# \text{level-p} &=
# \begin{cases}
# y_t - \text{s}_t \times \text{level}_{t-1}^{\text{pow-season}} & \text{if } t \le \text{seasonality}, \\
# \text{Average} \left[y(t - \text{seasonality} + 1), \ldots, y(t)\right] & \text{otherwise},
# \end{cases} \\
# \text{level}_{t} &= \text{level-sm} \times \text{level-p} + (1 - \text{level-sm}) \times \text{level}_{t-1}, \\
# \text{s}_{t + \text{seasonality}} &= \text{s-sm} \times \frac{y_{t} - \text{level}_{t}}{\text{level}_{t-1}^{\text{pow-trend}}}
# + (1 - \text{s-sm}) \times \text{s}_{t}.
# \end{align}
# A more detailed explanation for SGT model can be found in [this vignette](https://cran.r-project.org/web/packages/Rlgt/vignettes/GT_models.html) from the authors of the Rlgt package. Here we summarize the core ideas of this model:
#
# + [Student's t-distribution](https://en.wikipedia.org/wiki/Student%27s_t-distribution), which has heavier tails than normal distribution, is used for the likelihood.
# + The expected value `exp_val` consists of a trending component and a seasonal component:
# - The trend is governed by the map $x \mapsto x + ax^b$, where $x$ is `level`, $a$ is `coef_trend`, and $b$ is `pow_trend`. Note that when $b \sim 0$, the trend is linear with $a$ is the slope, and when $b \sim 1$, the trend is exponential with $a$ is the rate. So that function can cover a large family of trend.
# - When time changes, `level` and `s` are updated to new values. Coefficients `level_sm` and `s_sm` are used to make the transition smoothly.
# + When `powx` is near $0$, the error $\sigma_t$ will be nearly constant while when `powx` is near $1$, the error will be propotional to the expected value.
# + There are several varieties of SGT. In this tutorial, we use generalized seasonality and seasonal average method.
# We are ready to specify the model using *NumPyro* primitives. In NumPyro, we use the primitive `sample(name, prior)` to declare a latent random variable with a corresponding `prior`. These primitives can have custom interpretations depending on the effect handlers that are used by NumPyro inference algorithms in the backend. e.g. we can condition on specific values using the `condition` handler, or record values at these sample sites in the execution trace using the `trace` handler. Note that these details are not important for specifying the model, or running inference, but curious readers are encouraged to read the [tutorial on effect handlers](http://pyro.ai/examples/effect_handlers.html) in Pyro.
def sgt(y, seasonality, future=0):
# heuristically, standard derivation of Cauchy prior depends on
# the max value of data
cauchy_sd = jnp.max(y) / 150
# NB: priors' parameters are taken from
# https://github.com/cbergmeir/Rlgt/blob/master/Rlgt/R/rlgtcontrol.R
nu = numpyro.sample("nu", dist.Uniform(2, 20))
powx = numpyro.sample("powx", dist.Uniform(0, 1))
sigma = numpyro.sample("sigma", dist.HalfCauchy(cauchy_sd))
offset_sigma = numpyro.sample(
"offset_sigma", dist.TruncatedCauchy(low=1e-10, loc=1e-10, scale=cauchy_sd)
)
coef_trend = numpyro.sample("coef_trend", dist.Cauchy(0, cauchy_sd))
pow_trend_beta = numpyro.sample("pow_trend_beta", dist.Beta(1, 1))
# pow_trend takes values from -0.5 to 1
pow_trend = 1.5 * pow_trend_beta - 0.5
pow_season = numpyro.sample("pow_season", dist.Beta(1, 1))
level_sm = numpyro.sample("level_sm", dist.Beta(1, 2))
s_sm = numpyro.sample("s_sm", dist.Uniform(0, 1))
init_s = numpyro.sample("init_s", dist.Cauchy(0, y[:seasonality] * 0.3))
def transition_fn(carry, t):
level, s, moving_sum = carry
season = s[0] * level**pow_season
exp_val = level + coef_trend * level**pow_trend + season
exp_val = jnp.clip(exp_val, a_min=0)
# use expected vale when forecasting
y_t = jnp.where(t >= N, exp_val, y[t])
moving_sum = (
moving_sum + y[t] - jnp.where(t >= seasonality, y[t - seasonality], 0.0)
)
level_p = jnp.where(t >= seasonality, moving_sum / seasonality, y_t - season)
level = level_sm * level_p + (1 - level_sm) * level
level = jnp.clip(level, a_min=0)
new_s = (s_sm * (y_t - level) / season + (1 - s_sm)) * s[0]
# repeat s when forecasting
new_s = jnp.where(t >= N, s[0], new_s)
s = jnp.concatenate([s[1:], new_s[None]], axis=0)
omega = sigma * exp_val**powx + offset_sigma
y_ = numpyro.sample("y", dist.StudentT(nu, exp_val, omega))
return (level, s, moving_sum), y_
N = y.shape[0]
level_init = y[0]
s_init = jnp.concatenate([init_s[1:], init_s[:1]], axis=0)
moving_sum = level_init
with numpyro.handlers.condition(data={"y": y[1:]}):
_, ys = scan(
transition_fn, (level_init, s_init, moving_sum), jnp.arange(1, N + future)
)
if future > 0:
numpyro.deterministic("y_forecast", ys[-future:])
# Note that `level` and `s` are updated recursively while we collect the expected value at each time step. NumPyro uses [JAX](https://github.com/google/jax) in the backend to JIT compile many critical parts of the NUTS algorithm, including the verlet integrator and the tree building process. However, doing so using Python's `for` loop in the model will result in a long compilation time for the model, so we use `scan` - which is a wrapper of [lax.scan](https://jax.readthedocs.io/en/latest/_autosummary/jax.lax.scan.html#jax.lax.scan) with supports for NumPyro primitives and handlers. A detailed explanation for using this utility can be found in [NumPyro documentation](http://num.pyro.ai/en/latest/primitives.html#scan). Here we use it to collect `y` values while the triple `(level, s, moving_sum)` plays the role of carrying state.
# Another note is that instead of declaring the observation site `y` in `transition_fn`
#
# ```python
# numpyro.sample("y", dist.StudentT(nu, exp_val, omega), obs=y[t])
# ```
#
# , we have used [condition](http://num.pyro.ai/en/stable/handlers.html#numpyro.handlers.condition) handler here. The reason is we also want to use this model for forecasting. In forecasting, future values of `y` are non-observable, so `obs=y[t]` does not make sense when `t >= len(y)` (caution: index out-of-bound errors do not get raised in JAX, e.g. `jnp.arange(3)[10] == 2`). Using `condition`, when the length of `scan` is larger than the length of the conditioned/observed site, unobserved values will be sampled from the distribution of that site.
# ## Inference
# First, we want to choose a good value for `seasonality`. Following [the demo in Rlgt](https://github.com/cbergmeir/Rlgt/blob/master/Rlgt/demo/lynx.R), we will set `seasonality=38`. Indeed, this value can be guessed by looking at the plot of the training data, where the second order seasonality effect has a periodicity around $40$ years. Note that $38$ is also one of the highest-autocorrelation lags.
print("Lag values sorted according to their autocorrelation values:\n")
print(jnp.argsort(autocorrelation(y_train))[::-1])
# Now, let us run $4$ MCMC chains (using the No-U-Turn Sampler algorithm) with $5000$ warmup steps and $5000$ sampling steps per each chain. The returned value will be a collection of $20000$ samples.
# %%time
kernel = NUTS(sgt)
mcmc = MCMC(kernel, num_warmup=5000, num_samples=5000, num_chains=4)
mcmc.run(random.PRNGKey(0), y_train, seasonality=38)
mcmc.print_summary()
samples = mcmc.get_samples()
# ## Forecasting
# Given `samples` from `mcmc`, we want to do forecasting for the testing dataset `y_test`. NumPyro provides a convenient utility [Predictive](http://num.pyro.ai/en/stable/utilities.html#numpyro.infer.util.Predictive) to get predictive distribution. Let's see how to use it to get forecasting values.
#
# Notice that in the `sgt` model defined above, there is a keyword `future` which controls the execution of the model - depending on whether `future > 0` or `future == 0`. The following code predicts the last 34 values from the original time-series.
predictive = Predictive(sgt, samples, return_sites=["y_forecast"])
forecast_marginal = predictive(random.PRNGKey(1), y_train, seasonality=38, future=34)[
"y_forecast"
]
# Let's get sMAPE, root mean square error of the prediction, and visualize the result with the mean prediction and the 90% highest posterior density interval (HPDI).
y_pred = jnp.mean(forecast_marginal, axis=0)
sMAPE = jnp.mean(jnp.abs(y_pred - y_test) / (y_pred + y_test)) * 200
msqrt = jnp.sqrt(jnp.mean((y_pred - y_test) ** 2))
print("sMAPE: {:.2f}, rmse: {:.2f}".format(sMAPE, msqrt))
# Finally, let's plot the result to verify that we get the expected one.
plt.figure(figsize=(8, 4))
plt.plot(lynx["time"], data)
t_future = lynx["time"][80:]
hpd_low, hpd_high = hpdi(forecast_marginal)
plt.plot(t_future, y_pred, lw=2)
plt.fill_between(t_future, hpd_low, hpd_high, alpha=0.3)
plt.title("Forecasting lynx dataset with SGT model (90% HPDI)")
plt.show()
# As we can observe, the model has been able to learn both the first and second order seasonality effects, i.e. a cyclical pattern with a periodicity of around 10, as well as spikes that can be seen once every 40 or so years. Moreover, we not only have point estimates for the forecast but can also use the uncertainty estimates from the model to bound our forecasts.
# ## Acknowledgements
#
# We would like to thank <NAME> for many helpful resources and suggestions. Fast inference would not have been possible without the support of JAX and the XLA teams, so we would like to thank them for providing such a great open-source platform for us to build on, and for their responsiveness in dealing with our feature requests and bug reports.
# ## References
#
# [1] `Rlgt: Bayesian Exponential Smoothing Models with Trend Modifications`,<br>
# <NAME>, <NAME>, <NAME>, <NAME>, Trustees of Columbia University
| notebooks/source/time_series_forecasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/ZacCranko/robustlearningexperiments/blob/master/regulariser_unit_tests.ipynb)
# + id="xdqW-kE8Srje" colab_type="code" colab={}
# Copyright 2018 Google LLC and <NAME>
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# https://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import numpy as np
import tensorflow as tf
# + id="RjNQMY67tiSF" colab_type="code" colab={}
# Copyright 2018 Google LLC.
# SPDX-License-Identifier: Apache-2.0
def power_iterate_conv(layer, num_iter):
"""Perform power iteration for a convolutional layer."""
assert isinstance(layer, tf.keras.layers.Conv2D)
weights = layer.kernel
strides = (1,) + layer.strides + (1,)
padding = layer.padding.upper()
with tf.variable_scope(None, default_name='power_iteration'):
u_var = tf.get_variable(
'u_conv', [1] + map(int, layer.output_shape[1:]),
initializer=tf.random_normal_initializer(),
trainable=False)
u = u_var
for _ in xrange(num_iter):
v = tf.nn.conv2d_transpose(
u, weights, [1] + map(int, layer.input_shape[1:]), strides, padding)
v /= tf.sqrt(tf.maximum(2 * tf.nn.l2_loss(v), 1e-12))
u = tf.nn.conv2d(v, weights, strides, padding)
u /= tf.sqrt(tf.maximum(2 * tf.nn.l2_loss(u), 1e-12))
tf.add_to_collection(tf.GraphKeys.UPDATE_OPS, tf.assign(u_var, u))
u = tf.stop_gradient(u)
v = tf.stop_gradient(v)
return tf.reduce_sum(u * tf.nn.conv2d(v, weights, strides, padding))
def power_iterate_dense(layer, num_iter):
"""Perform power iteration for a fully connected layer."""
assert isinstance(layer, tf.keras.layers.Dense)
weights = layer.kernel
output_shape, input_shape = weights.get_shape().as_list()
with tf.variable_scope(None, default_name='power_iteration'):
u_var = tf.get_variable(
'u', map(int, [output_shape]) + [1],
initializer=tf.random_normal_initializer(),
trainable=False)
u = u_var
for _ in xrange(num_iter):
v = tf.matmul(weights, u, transpose_a=True)
v /= tf.sqrt(tf.maximum(2 * tf.nn.l2_loss(v), 1e-12))
u = tf.matmul(weights, v)
u /= tf.sqrt(tf.maximum(2 * tf.nn.l2_loss(u), 1e-12))
tf.add_to_collection(tf.GraphKeys.UPDATE_OPS, tf.assign(u_var, u))
u = tf.stop_gradient(u)
v = tf.stop_gradient(v)
return tf.reduce_sum(u * tf.matmul(weights, v))
# + id="4l9ZdjxfG4tY" colab_type="code" colab={}
def operator_norm(layer, ord = 2, **kwargs):
"""Compute operator norm for a Keras layer."""
with tf.variable_scope(None, default_name='operator_norm'):
if ord == 1:
w = layer.kernel
if isinstance(layer, tf.keras.layers.Conv2D):
sum_w = tf.reduce_sum(tf.abs(w), [0, 1, 3])
else:
sum_w = tf.reduce_sum(tf.abs(w), 1)
return tf.reduce_max(sum_w)
elif ord == 2:
num_iter = kwargs.get('num_iter', 5)
if isinstance(layer, tf.keras.layers.Conv2D):
return power_iterate_conv(layer, num_iter)
else:
return power_iterate_dense(layer, num_iter)
elif ord == np.inf:
w = layer.kernel
if isinstance(layer, tf.keras.layers.Conv2D):
sum_w = tf.reduce_sum(tf.abs(w), [0, 1, 2])
else:
sum_w = tf.reduce_sum(tf.abs(w), 0)
return tf.reduce_max(sum_w)
# + id="24NAhjHvS5fs" colab_type="code" colab={}
def tf_assert_almost_equal(actual, desired, **kwargs):
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
_actual = actual.eval()
_desired = desired.eval()
return np.testing.assert_almost_equal(_actual, _desired, **kwargs)
def conv_matrix(layer):
"""Build the matrix associated with the convolution operation."""
assert isinstance(layer, tf.keras.layers.Conv2D)
with tf.variable_scope(None, default_name='build_conv_matrix'):
weights = layer.kernel
strides = (1,) + layer.strides + (1,)
padding = layer.padding.upper()
in_h, in_w, in_ch = layer.input_shape[1:4]
out_h, out_w, out_ch = layer.output_shape[1:4]
id_mx = tf.reshape(tf.eye(in_h*in_w*in_ch),
(in_h*in_w*in_ch, in_h, in_w, in_ch))
conv_mx_t = tf.reshape(tf.nn.conv2d(id_mx, weights, strides, padding),
(in_h*in_w*in_ch, out_h*out_w*out_ch))
return tf.transpose(conv_mx_t)
# + id="N7sA-821-rV4" colab_type="code" colab={}
model = tf.keras.Sequential()
conv1 = tf.keras.layers.Conv2D(32, 5, 1, padding='SAME',
input_shape=(28, 28, 1))
model.add(conv1)
model.add(tf.keras.layers.Activation('relu'))
model.add(tf.keras.layers.MaxPooling2D(2, 2, padding='SAME'))
conv2 = tf.keras.layers.Conv2D(64, 5, 1, padding='SAME')
model.add(conv2)
dense1 = tf.keras.layers.Dense(1024)
model.add(dense1)
model.add(tf.keras.layers.Activation('relu'))
dense2 = tf.keras.layers.Dense(10)
model.add(dense2)
# + id="OyXDyXml4MI5" colab_type="code" colab={}
num_iter = 200
# test dense layers
for layer in ['dense1', 'dense2']:
_layer = eval(layer)
op = "1_norm"
# axis = 1 here since _layer.kernel is stored transposed for dense layers
inf_opn_mx = tf.reduce_max(tf.reduce_sum(tf.abs(_layer.kernel), axis = 1))
tf_assert_almost_equal(operator_norm(_layer, 1), inf_opn_mx, err_msg = "%s(%s)"%(op,layer), decimal = 1)
op = "inf_norm"
# axis = 0 here since _layer.kernel is stored transposed for dense layers
inf_opn_mx = tf.reduce_max(tf.reduce_sum(tf.abs(_layer.kernel), axis = 0))
tf_assert_almost_equal(operator_norm(_layer, np.inf), inf_opn_mx, err_msg = "%s(%s)"%(op,layer), decimal = 1)
op = "spectral_norm"
spec_pow = operator_norm(_layer, 2, num_iter = num_iter)
spec_svd = tf.svd(_layer.kernel, compute_uv=False)
tf_assert_almost_equal(spec_pow, spec_svd[0], decimal = 2, err_msg = "%s(%s)"%(op,layer))
# test conv layers
for layer in ['conv1', 'conv2']:
_layer = eval(layer)
op = "1_norm"
conv_mx = conv_matrix(_layer)
desired = tf.reduce_max(tf.reduce_sum(tf.abs(conv_mx), axis = 0))
tf_assert_almost_equal(operator_norm(_layer, 1), desired, err_msg = "%s(%s)"%(op,layer), decimal = 1)
op = "inf_norm"
conv_mx = conv_matrix(_layer)
desired = tf.reduce_max(tf.reduce_sum(tf.abs(conv_mx), axis = 1))
tf_assert_almost_equal(operator_norm(_layer, np.inf), desired, err_msg = "%s(%s)"%(op,layer), decimal = 1)
op = "spectral_norm"
spec_pow = operator_norm(_layer, 2, num_iter = num_iter)
spec_svd = tf.svd(conv_matrix(_layer), compute_uv=False)
tf_assert_almost_equal(spec_pow, spec_svd[0], err_msg = "%s(%s)"%(op,layer), decimal = 2)
| regulariser_unit_tests.ipynb |
# ---
# jupyter:
# celltoolbar: Create Assignment
# jupytext:
# formats: ipynb,md:myst
# notebook_metadata_filter: all,-language_info,-toc,-latex_envs
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Answers: Lab Week 2 - Jupyterhub Tutorial
#
# ## EOSC 211
#
# ### Learning Objectives
#
# - Connect to Jupyterhub via the course website
# - Create, edit, and delete *code cells* and *markdown cells* in a Jupyter notebook
# - Assign data to *variables*
# - Execute code cells to do some calculations
# - Use comments and/or *markdown* cells to add annotations to your code
# - Save your work and submit an assignment via Canvas
# ### Intro
#
# Welcome to your first lab in **EOSC 211**! Here we will write our first bits of *Python* code, explore some of the features of *Jupyter Notebooks*, and learn how to save and submit your work for grading.
#
# The URL for this course is -url-. Here you can view all of the notes, worksheets, labs, and assignments for each week of the course. If you are on the website and would like to open an assignment to work on your own copy of it, click the rocketship icon in the top right hand corner of the screen to launch ***JuptyerHub***. After you sign in, Jupyterhub will open an interactive copy of the notebook which you can edit, download, and submit for grading.
#
# <img src=open_jhub.png width=150>
#
# #### Why Python/Jupyterhub?
#
# <img src=xkcd_python.png>
# <figcaption>Credit: <NAME>, xkcd.com</figcaption>
#
# **Python** is a *high level programming language,* meaning it is comparatively easy for humans to read. Python programs are executed by a *python interpreter,* which takes python code, reads it into a processor (the CPU on your computer, or in our case, a Jupyterhub server) and returns a result. This process is repeated over and over for each line of code, often taking the result of one line and feeding it into the next.
#
# Python is also *open source*, meaning there are no fees to download/use python, and anyone is free to add to the Python code base. This is a major reason for Python's growing popularity in the scientific community, there are many add ons (called *packages*) developed by other scientists specifically for the type of programming we want to do in this course, with more being added every day!
#
# **Jupyter Notebooks** are an IDE, or *integrated development environment* which have gained popularity amoung the science community for processing data, creating scientific figures, solving numerical equations, and sharing code. IDE's are essentially advanced text editors containing features helpful for writing code (if you really wanted to, you could write all your code in Notepad or Microsoft Word then copy/paste it into a python interpreter somewhere, but we strongly recommend *not* doing this).
#
# **JupyterHub** is a *computational environment* for running jupyter notebooks in the cloud, designed specifically for applications like teaching or collaborative research. The EOSC211 course hub is available online and requires no installation on your own machine. Later in the course, we will talk about installing python yourself, creating your own *environments*, and developing your own individual workflow.
# ### Code Cells and Markdown Cells
#
# Jupyter notebooks are divided into *cells,* which can be individually edited and run. To create a new cell, click `Insert` $\rightarrow$ `Insert Cell Below`, or press `[Alt]` + `[Enter]` to run the current cell and create a new one below. After you create a cell, you can choose either *Code* (keyboard shortcut `[Esc]` + `[y]`) or *Markdown* (`[Esc]` + `[m]`) as the cell type. You can delete cells with `Edit` $\rightarrow$ `Delete Cells`
#
# <img src=insert_new_cell.png width=180><img src=cell_type.png width=150><img src=delete_cells.png width=180>
#
# ### Code Cells
#
# Code cells are where you can write, edit, and run *Python Code.* Text entered into code cells will be shown with *syntax highlighting,* with python *reserved words* shown in green, *strings* shown in red, and *comments* in blue. You can execute code cells by pressing [$\blacktriangleright$ Run], or `[Ctrl]` + `[Enter]` (execute current cell), `[Shift]` + `[Enter]` (run current cell and select the next one), or `[Alt]` + `[Enter]` (execute current cell and create a new one below).
#
# Try it yourself! Edit the cell below, replacing the comment (lines of code beginning with "#" are ignored by the python interpreter and not executed) with:
#
# ```python
# print("Hello World")
# ```
#
# and press `[Ctrl]` + `[Enter]` to execute the code.
# your code here
print("Hello World")
# Congratulations, you have just written and executed your first Python code! The "Hello World" program is a long-standing tradition in computer programming as a first program in a new language.
#
# Python has much more capability than just printing "Hello World". You can write multiple lines of code in one cell, the *python interpreter* will execute each line of code in the order it appears, i.e.
print("Hello Earth")
print("Hello Ocean")
print("Hello Atmosphere")
# Text written in code cells needs to be *syntactically correct* in order to execute, i.e. it needs to contain a valid combination of reserved words, operations and variables that can be parsed by the python interpreter. Invalid code will raise an error:
print("Hello <NAME>")
# If there is something wrong with a code cell, the python interpreter stops execution and produces a *traceback message.* Traceback messages appear below a code cell and usually give you a helpful hint as to what is wrong and how to fix it. Some messages can be a bit cryptic; if this is the case, try searching [stack overflow](https://stackoverflow.com/) or another helpful coding site (many exist). After searching 3 different sites without success, ask the nearest classmate, professor, or TA for help.
# #### Variables
#
# Instead of just outputting things to the screen, we can also save data as a *variable*, and reference it throughout your code, like so:
planet = "Earth"
# This tells the python interpreter: *Find some space in computer memory, store the letters which form the word "Earth" there, and reference that word with a variable called **planet** (Note: this happens without outputting anything to the screen).* Throughout the rest of your code, referencing the variable "planet" will result in the word "Earth". If you change the variable to "Betelgeuse", every reference to that variable will now produce "Betelgeuse". Clever use of variables can save you time hunting through your code and replacing every instance of a word/number/value.
print(f"Welcome to {planet}")
print(f"Nice weather today here on {planet}")
print(f"{planet} will shortly be destroyed to make space for the new interstellar Vogon highway.")
# #### Cell Execution Order and Restarting the Kernel
#
# Within a cell, lines of code are executed in the order they are written. Code cells will execute in any order you decide. The order in which cells are executed appears to the left e.g. `In [7]:` indicates that a particular cell is the seventh cell to be run in the notebook. Try executing the cells below in order:
#
# * cell 1, cell 2, cell 3
# * cell 3, cell 2, cell 1
#
# Which value for `my_field` gets printed to the screen in either case?
# cell 1
my_field = "oceanography"
# cell 2
print(my_field + " is awesome!")
# cell 3
my_field = "atmospheric science"
my_field = "geology"
# Finally, press the \[$\circlearrowright$\] button to *restart the kernel* and try executing cell 2 again. What happens? Restarting the kernel tells the python interpreter to *forget all variables and start over from scratch*. The \[$\blacktriangleright\blacktriangleright$\] button restarts the kernel then immediately executes each cell in order (top to bottom, or until it encounters an error and cannot continue).
#
# <div class="alert alert-danger" role="alert">
# <strong>Important: </strong> When your instructors mark a submitted notebook, all cells will be executed in order. Before you submit, run the whole notebook from top to bottom and make sure it produces the result you want
# </div>
#
#
# Try it out! Edit the cells above (create new cells or copy/paste lines of code) so that when you press \[$\blacktriangleright\blacktriangleright$\], the resulting output to the screen is:
#
# ```
# atmospheric science is awesome!
# ```
# #### Math in Python
#
# The Python core contains functionality for doing basic math. Later in the course, we will learn how to *import* further functionality for doing not-so-basic math. The syntax for math in Python is summarized below:
#
# | Operator | Description | Example | Result |
# | --- | --- | --- | --- |
# | ``` + ``` | addition | ```2 + 3 ``` | ```5``` |
# | ``` - ``` | subtraction | ```8 - 6 ``` | ```2``` |
# | ``` - ``` | negative number | ```-4``` | ```-4``` |
# | ``` * ``` | multiplication | ``` 5 * 2 ``` | ``` 10 ``` |
# | ``` / ``` | division | ``` 6 / 3 ``` | ``` 2 ``` |
# | ``` ** ```| raises a number to a power | ``` 10**2 ``` | ``` 100 ``` |
#
# For more complex calculations involving several operators, the python interpreter will perform calculations in the order BEDMAS (brackets, exponents, division/multiplication, addiition/subtraction), just like you would do by hand.
#
# Try it out! In the code cell below (or you can create multiple code cells, up to you!), solve the following equations:
#
# **A:** $2^2 + 9$
#
# **B:** $2^2 + 9 \times 2$
#
# **C:** $(2 + 9)^2 \times 2$
# your code here
a = (2 + 9)**2.
# #### Math With Variables
#
# Just like we could print *variables* to the screen, we can assign and reference variables within calculations. For example, the circumference $C$ of a circle is:
#
# $$
# C = 2\pi r
# $$
#
# where $r$ is the radius of the circle and $\pi = 3.14$. We can code this equation in python like so:
# +
radius_of_planet = 6371 # km
circumference = 2 * 3.14 * radius_of_planet
print(circumference)
# -
# ### Markdown Cells
#
# One of the big learning goals in this course is writing code that is *well documented*. It should be easy for another programmer (or your future self) to look at your code and understand what is happening in each code cell and why. A major reason we have elected to teach this course with Jupyter Notebooks is their ability to show nicely formatted commentary between snippets of executable computer code.
#
# Unlike code cells, *markdown cells* don't execute or produce outputs, they simply *render* plain text into nicely formatted paragraphs. The cell you are reading right now is written in markdown; double click on it to see the plain text before rendering. Press `[Shift]` + `[Enter]` to re-render the cell.
#
# The main focus of this course is writing *Python* code, so you won't be expected to learn every part of the markdown syntax (but feel free to make your notebooks/labs as fancy as you like, we encourage it!). You will need to be able to edit markdown cells in order to complete long answer questions, like so:
#
# **What is your favorite color?**
# your answer here
#
# red
# ### Saving and Submitting Assignments
#
# As you create or edit a Jupyter Notebook, you can save your work by clicking `File` $\rightarrow$ `Save and Checkpoint` (or `[Ctrl]` + `[s]`). This will save your notebook on the Jupyterhub server in the cloud. To download a notebook to your own computer, select `File` $\rightarrow$ `Download as` $\rightarrow$ `Notebook (.ipynb)`.
#
# It is ***very good programming practice*** to keep at least 2 copies of each notebook in at least 2 locations (e.g. on the jupyterhub server and on your laptop) and we highly recommend doing this to prevent losing work.
#
# To hand in an assignment, download your final copy as a .ipynb and upload it via the link to that assignment on canvas
# ## To hand in by the end of this lab
#
# Write your answers in the cells below and submit your lab via Canvas
# ### Question 1
#
# **What motivated you to sign up for this course? Are you most interested in Earth, Ocean, or Atmospheric Science? (or something else?)**
#
# **(Double click to edit the markdown cell below, `[Ctrl]` + `[Enter]` to render)**
# + [markdown] nbgrader={"grade": true, "grade_id": "texttest", "locked": false, "points": 1, "schema_version": 3, "solution": true, "task": false}
# your answer here
# -
# ### Question 2 (3 points)
#
# **We can calculate travel time $t$ given distance $d$ and speed $v$ with**
#
# $$
# t = \frac{d}{v}
# $$
#
# **With $t$ reported in hours, $d$ in kilometers and $v$ in kilometers per hour. Write code in the cell below to calculate the time in days required to bike along the equator of a planet of radius `radius`, using the *variables* assigned assigned earlier in this notebook. Assume you can maintain
# an average speed of `biking_speed`. Make sure you produce a variable with the name `time_in_days` that contains the answer.**
#
# **Press the \[$\blacktriangleright\blacktriangleright$\] button to run the whole notebook again, and make sure that the code still produces the correct output. (Note: the *values* for `planet` (ie the name), `biking_speed` (numerical value in kph) and `radius` (numerical value in km) and aren't important important. Full marks are awarded for code that produces a sensible result with no errors)**
# + nbgrader={"grade": false, "grade_id": "vartest", "locked": false, "schema_version": 3, "solution": true, "task": false}
# biking_speed = xx # km/h
# reference variables assigned in earlier cells to calculate travel time around "planet"
# and assign the result to a new variable "t"
biking_speed = 20 # km/hour
hours_per_day = 24.
biking_speed = biking_speed*hours_per_day # convert hours to days
time_in_days = circumference/biking_speed
print(f"It will take {time_in_days} days to ride a bicycle around {planet}")
# + nbgrader={"grade": true, "grade_id": "cell-a31c35d4a996071e", "locked": true, "points": 3, "schema_version": 3, "solution": false, "task": false}
assert(time_in_days > 0)
# -
# ## Helpful Terminology
#
# **Python:** a high level programming language popular among scientists. Python code is saved with the extension `.py` and can be run on any computer with a python interpreter installed on it. We are using version the *miniconda distribution, version 3.7* for this course.
#
# **Jupyter Notebooks:** an *Integrated Development Environment* for writing python code that runs in a web browser (i.e. Chrome, Firefox). Notebooks are comprised of cells, which are either executable python code or rendered markdown text. Notebooks contain and execute python code, but are saved with the extension `.ipynb`
#
# **Jupyterhub:** a *computational environment* for running jupyter notebooks in the cloud, designed specifically for applications like teaching or collaborative research. The EOSC211 course hub is available online and requires no installation on your own machine. Later in the course, we will talk about installing python yourself and developing your own individual workflow.
| lab_keys/week2_lab/lab_wk2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3D analysis
#
# This tutorial shows how to run a 3D map-based analysis using three example observations of the Galactic center region with CTA.
# ## Setup
# %matplotlib inline
import matplotlib.pyplot as plt
import os
from pathlib import Path
import numpy as np
import astropy.units as u
from astropy.coordinates import SkyCoord
from gammapy.data import DataStore
from gammapy.irf import EnergyDispersion, make_mean_psf, make_mean_edisp
from gammapy.maps import WcsGeom, MapAxis, Map, WcsNDMap
from gammapy.cube import MapMaker, MapEvaluator, PSFKernel, MapDataset
from gammapy.cube.models import SkyModel, SkyDiffuseCube, BackgroundModel
from gammapy.spectrum.models import PowerLaw, ExponentialCutoffPowerLaw
from gammapy.image.models import SkyGaussian, SkyPointSource
from gammapy.utils.fitting import Fit
from regions import CircleSkyRegion
# !gammapy info --no-system
# ## Prepare modeling input data
#
# ### Prepare input maps
#
# We first use the `DataStore` object to access the CTA observations and retrieve a list of observations by passing the observations IDs to the `.get_observations()` method:
# Define which data to use and print some information
data_store = DataStore.from_dir("$GAMMAPY_DATA/cta-1dc/index/gps/")
data_store.info()
print(
"Total observation time (hours): ",
data_store.obs_table["ONTIME"].sum() / 3600,
)
print("Observation table: ", data_store.obs_table.colnames)
print("HDU table: ", data_store.hdu_table.colnames)
# Select some observations from these dataset by hand
obs_ids = [110380, 111140, 111159]
observations = data_store.get_observations(obs_ids)
# Now we define a reference geometry for our analysis, We choose a WCS based gemoetry with a binsize of 0.02 deg and also define an energy axis:
energy_axis = MapAxis.from_edges(
np.logspace(-1.0, 1.0, 10), unit="TeV", name="energy", interp="log"
)
geom = WcsGeom.create(
skydir=(0, 0),
binsz=0.02,
width=(10, 8),
coordsys="GAL",
proj="CAR",
axes=[energy_axis],
)
# The `MapMaker` object is initialized with this reference geometry and a field of view cut of 4 deg:
# %%time
maker = MapMaker(geom, offset_max=4.0 * u.deg)
maps = maker.run(observations)
# The maps are prepared by calling the `.run()` method and passing the `observations`. The `.run()` method returns a Python `dict` containing a `counts`, `background` and `exposure` map:
print(maps)
# This is what the summed counts image looks like:
counts = maps["counts"].sum_over_axes()
counts.smooth(width=0.1 * u.deg).plot(stretch="sqrt", add_cbar=True, vmax=6);
# This is the background image:
background = maps["background"].sum_over_axes()
background.smooth(width=0.1 * u.deg).plot(
stretch="sqrt", add_cbar=True, vmax=6
);
# And this one the exposure image:
exposure = maps["exposure"].sum_over_axes()
exposure.smooth(width=0.1 * u.deg).plot(stretch="sqrt", add_cbar=True);
# We can also compute an excess image just with a few lines of code:
excess = counts - background
excess.smooth(5).plot(stretch="sqrt", add_cbar=True);
# For a more realistic excess plot we can also take into account the diffuse galactic emission. For this tutorial we will load a Fermi diffuse model map that represents a small cutout for the Galactic center region:
diffuse_gal = Map.read("$GAMMAPY_DATA/fermi-3fhl-gc/gll_iem_v06_gc.fits.gz")
print("Diffuse image: ", diffuse_gal.geom)
print("counts: ", maps["counts"].geom)
# We see that the geometry of the images is completely different, so we need to apply our geometric configuration to the diffuse emission file:
# +
coord = maps["counts"].geom.get_coord()
data = diffuse_gal.interp_by_coord(
{
"skycoord": coord.skycoord,
"energy": coord["energy"]
* maps["counts"].geom.get_axis_by_name("energy").unit,
},
interp=3,
)
diffuse_galactic = WcsNDMap(maps["counts"].geom, data)
print("Before: \n", diffuse_gal.geom)
print("Now (same as maps): \n", diffuse_galactic.geom)
# -
# diffuse_galactic.slice_by_idx({"energy": 0}).plot(add_cbar=True); # this can be used to check image at different energy bins
diffuse = diffuse_galactic.sum_over_axes()
diffuse.smooth(5).plot(stretch="sqrt", add_cbar=True)
print(diffuse)
# We now multiply the exposure for this diffuse emission to subtract the result from the counts along with the background.
combination = diffuse * exposure
combination.unit = ""
combination.smooth(5).plot(stretch="sqrt", add_cbar=True);
# We can plot then the excess image subtracting now the effect of the diffuse galactic emission.
# +
excess2 = counts - background - combination
fig, axs = plt.subplots(1, 2, figsize=(15, 5))
axs[0].set_title("With diffuse emission subtraction")
axs[1].set_title("Without diffuse emission subtraction")
excess2.smooth(5).plot(
cmap="coolwarm", vmin=-1, vmax=1, add_cbar=True, ax=axs[0]
)
excess.smooth(5).plot(
cmap="coolwarm", vmin=-1, vmax=1, add_cbar=True, ax=axs[1]
);
# -
# ### Prepare IRFs
#
# To estimate the mean PSF across all observations at a given source position `src_pos`, we use `make_mean_psf()`:
# +
# mean PSF
src_pos = SkyCoord(0, 0, unit="deg", frame="galactic")
table_psf = make_mean_psf(observations, src_pos)
# PSF kernel used for the model convolution
psf_kernel = PSFKernel.from_table_psf(table_psf, geom, max_radius="0.3 deg")
# -
# To estimate the mean energy dispersion across all observations at a given source position `src_pos`, we use `make_mean_edisp()`:
# +
# define energy grid
energy = energy_axis.edges * energy_axis.unit
# mean edisp
edisp = make_mean_edisp(
observations, position=src_pos, e_true=energy, e_reco=energy
)
# -
# ### Save maps and IRFs to disk
#
# It is common to run the preparation step independent of the likelihood fit, because often the preparation of maps, PSF and energy dispersion is slow if you have a lot of data. We first create a folder:
path = Path("analysis_3d")
path.mkdir(exist_ok=True)
# And then write the maps and IRFs to disk by calling the dedicated `.write()` methods:
# +
# write maps
maps["counts"].write(str(path / "counts.fits"), overwrite=True)
maps["background"].write(str(path / "background.fits"), overwrite=True)
maps["exposure"].write(str(path / "exposure.fits"), overwrite=True)
# write IRFs
psf_kernel.write(str(path / "psf.fits"), overwrite=True)
edisp.write(str(path / "edisp.fits"), overwrite=True)
# -
# ## Likelihood fit
#
# ### Reading maps and IRFs
# As first step we read in the maps and IRFs that we have saved to disk again:
# +
# read maps
maps = {
"counts": Map.read(str(path / "counts.fits")),
"background": Map.read(str(path / "background.fits")),
"exposure": Map.read(str(path / "exposure.fits")),
}
# read IRFs
psf_kernel = PSFKernel.read(str(path / "psf.fits"))
edisp = EnergyDispersion.read(str(path / "edisp.fits"))
# -
# Let's cut out only part of the maps, so that we the fitting step does not take so long (we go from left to right one):
cmaps = {
name: m.cutout(SkyCoord(0, 0, unit="deg", frame="galactic"), 2 * u.deg)
for name, m in maps.items()
}
cmaps["counts"].sum_over_axes().plot(stretch="sqrt");
# Insted of the complete one, which was:
counts.plot(stretch="sqrt");
# ### Fit mask
#
# To select a certain spatial region and/or energy range for the fit we can create a fit mask:
# +
mask = Map.from_geom(cmaps["counts"].geom)
region = CircleSkyRegion(center=src_pos, radius=0.6 * u.deg)
mask.data = mask.geom.region_mask([region])
mask.get_image_by_idx((0,)).plot();
# -
# In addition we also exclude the range below 0.3 TeV for the fit:
coords = mask.geom.get_coord()
mask.data &= coords["energy"] > 0.3
# ### Model fit
#
# No we are ready for the actual likelihood fit. We first define the model as a combination of a point source with a powerlaw:
spatial_model = SkyPointSource(lon_0="0.01 deg", lat_0="0.01 deg")
spectral_model = PowerLaw(
index=2.2, amplitude="3e-12 cm-2 s-1 TeV-1", reference="1 TeV"
)
model = SkyModel(spatial_model=spatial_model, spectral_model=spectral_model)
# Defining a background model
#
# Often, it is useful to fit the normalisation (and also the tilt) of the background. To do so, we have to define the background as a model. In this example, we will keep the tilt fixed and the norm free.
background_model = BackgroundModel(cmaps["background"], norm=1.1, tilt=0.0)
background_model.parameters['norm'].frozen = False
background_model.parameters['tilt'].frozen = True
# Now we set up the `MapDataset` object by passing the prepared maps, IRFs as well as the model:
dataset = MapDataset(
model=model,
counts=cmaps["counts"],
exposure=cmaps["exposure"],
background_model=background_model,
mask=mask,
psf=psf_kernel,
edisp=edisp,
)
# No we run the model fit:
# %%time
fit = Fit(dataset)
result = fit.run(optimize_opts={"print_level": 1})
result.parameters.to_table()
# We can see that the background normalisation is very large (2.15). This is because of the large number of sources, and the presence of diffuse emission in the galactic center.
# ### Check model fit
#
# We check the model fit by computing a residual image. For this we first get the number of predicted counts:
npred = dataset.npred()
# And compute a residual image:
residual = cmaps["counts"] - npred
residual.sum_over_axes().smooth(width=0.05 * u.deg).plot(
cmap="coolwarm", vmin=-3, vmax=3, add_cbar=True
);
# We can also plot the best fit spectrum. For that need to extract the covariance of the spectral parameters.
# +
spec = model.spectral_model
# set covariance on the spectral model
covariance = result.parameters.covariance
spec.parameters.covariance = covariance[2:5, 2:5]
energy_range = [0.3, 10] * u.TeV
spec.plot(energy_range=energy_range, energy_power=2)
ax = spec.plot_error(energy_range=energy_range, energy_power=2)
# -
# Apparently our model should be improved by adding a component for diffuse Galactic emission and at least one second point
# source. But before we do that in the next section, we will fit the background as a model.
# ### Add Galactic diffuse emission to model
# We use both models at the same time, our diffuse model (the same from the Fermi file used before) and our model for the central source. This time, in order to make it more realistic, we will consider an exponential cut off power law spectral model for the source (note that we are not constraining the fit with any mask this time). We will again fot the normalisation of the background.
diffuse_model = SkyDiffuseCube.read(
"$GAMMAPY_DATA/fermi-3fhl-gc/gll_iem_v06_gc.fits.gz"
)
# +
spatial_model = SkyPointSource(lon_0="0.01 deg", lat_0="0.01 deg")
spectral_model = ExponentialCutoffPowerLaw(
index=2 * u.Unit(""),
amplitude=1e-12 * u.Unit("cm-2 s-1 TeV-1"),
reference=1.0 * u.TeV,
lambda_=1 / u.TeV,
)
model_ecpl = SkyModel(
spatial_model=spatial_model, spectral_model=spectral_model
)
background_model = BackgroundModel(cmaps["background"], norm=1.0, tilt=0.0)
background_model.parameters['norm'].frozen = False
background_model.parameters['tilt'].frozen = True
model_combined = diffuse_model + model_ecpl
# -
dataset_combined = MapDataset(
model=model_combined,
counts=cmaps["counts"],
exposure=cmaps["exposure"],
background_model=background_model,
psf=psf_kernel,
)
# %%time
fit_combined = Fit(dataset_combined)
result_combined = fit_combined.run()
print(model_ecpl)
# As we can see we have now two components in our model, and we can access them separately.
# Checking normalization value (the closer to 1 the better)
print("Model 1: {}\n".format(model_combined.model1))
print("Model 2: {}\n".format(model_combined.model2))
print("Background model: {}\n".format(background_model))
# You can see that the normalisation of the background has vastly improved
# We can now plot the residual image considering this improved model.
residual2 = cmaps["counts"] - dataset_combined.npred()
# Just as a comparison, we can plot our previous residual map (left) and the new one (right) with the same scale:
# +
plt.figure(figsize=(15, 5))
ax_1 = plt.subplot(121, projection=residual.geom.wcs)
ax_2 = plt.subplot(122, projection=residual.geom.wcs)
ax_1.set_title("Without diffuse emission subtraction")
ax_2.set_title("With diffuse emission subtraction")
residual.sum_over_axes().smooth(width=0.05 * u.deg).plot(
cmap="coolwarm", vmin=-2, vmax=2, add_cbar=True, ax=ax_1
)
residual2.sum_over_axes().smooth(width=0.05 * u.deg).plot(
cmap="coolwarm", vmin=-2, vmax=2, add_cbar=True, ax=ax_2
);
# -
# Finally we can check again our model (including now the diffuse emission):
spec_ecpl = model_ecpl.spectral_model
ax = spec_ecpl.plot(energy_range=energy_range, energy_power=2)
# Results seems to be better (but not perfect yet). Next step to improve our model even more would be getting rid of the other bright source (G0.9+0.1).
# Note that this notebook aims to show you the procedure of a 3D analysis using just a few observations and a cutted Fermi model. Results get much better for a more complete analysis considering the GPS dataset from the CTA First Data Challenge (DC-1) and also the CTA model for the Galactic diffuse emission, as shown in the next image:
# 
# The complete tutorial notebook of this analysis is available to be downloaded in [GAMMAPY-EXTRA](https://github.com/gammapy/gammapy-extra) repository at https://github.com/gammapy/gammapy-extra/blob/master/analyses/cta_1dc_gc_3d.ipynb).
# ## Exercises
#
# * Analyse the second source in the field of view: G0.9+0.1 and add it to the combined model.
| tutorials/analysis_3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''afml'': venv)'
# name: python37464bitafmlvenvc7329fc4ec4c463eb5286d30ad3c7824
# ---
import pandas as pd
df = pd.DataFrame({
'x1': ['a', 'a', 'a', 'b', 'b', 'b'],
'x2': [1 ,2, 3, 4, 5, 6]
})
df
df['x1'].drop_duplicates()
df['x1'].drop_duplicates(keep='first')
df['x1'].drop_duplicates(keep='last')
| notebooks/drop-duplicates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: 2db524e06e9f5f4ffedc911c917cb75e12dbc923643829bf417064a77eb14d37
# name: python3
# ---
# ## Carga dos dados
import pandas as pd
temperaturas = pd.read_csv('temps.csv')
temperaturas.head()
# ## Análise Exploratória
print('Dimensões dos dataset temps: ', temperaturas.shape)
temperaturas.describe()
print('Quantos dias do ano foram utilizados: ', 348/365 * 100, '%')
temperaturas.info()
# +
from datetime import datetime
years = temperaturas['year']
months = temperaturas['month']
days = temperaturas['day']
# Formato americano para a data: yyyy-mm-dd
dates=[str(year) + '-' + str(month) + '-' + str(day) for year, month, day in zip(years, months, days)]
dates=[datetime.strptime(date, '%Y-%m-%d') for date in dates]
# +
import matplotlib.pyplot as plt
fig, ((ax1, ax2), (ax3, ax4)) = plt.subplots(nrows=2, ncols=2, figsize=(10, 10))
fig.autofmt_xdate(rotation=45)
_ = ax1.plot(dates, temperaturas['actual'])
ax1.set_xlabel(''); ax1.set_ylabel('Temperatura'); ax1.set_title('Atual')
_ = ax2.plot(dates, temperaturas['temp_1'])
ax2.set_xlabel(''); ax2.set_ylabel('Temperatura'); ax2.set_title('Dia anterior')
_ = ax3.plot(dates, temperaturas['temp_2'])
ax3.set_xlabel(''); ax3.set_ylabel('Temperatura'); ax3.set_title('Dois dias atrás')
_ = ax4.plot(dates, temperaturas['friend'])
ax4.set_xlabel(''); ax4.set_ylabel('Temperatura'); ax4.set_title('Estimada pelo amigo')
# -
# ## Preparação dos dados
# Transforma o dado categorico em númerico (dias da semana)
temperaturas = pd.get_dummies(temperaturas)
temperaturas.head()
print('Dimensões do dataset temps: ', temperaturas.shape)
# ## Treinamento e Teste do Modelo
X = temperaturas.drop('actual', axis=1).values
y = temperaturas['actual'].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, random_state=42)
print('X_train shape', X_train.shape)
print('X_test shape', X_train.shape)
print('y_train shape', y_train.shape)
print('y_test shape', y_test.shape)
# ## Base de comparação
# +
import numpy as np
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
colunas = list(temperaturas.columns)
y_hist = X_test[:, colunas.index('average')]
print('MAE :', mean_absolute_error(y_test, y_hist))
print('MSE :', mean_squared_error(y_test, y_hist))
print('RMSE:', np.sqrt(mean_squared_error(y_test, y_hist)))
# -
# ## Treinamento do modelo
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(n_estimators=100, random_state=42)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_test)
print('MAE :', mean_absolute_error(y_test, y_pred))
print('MSE :', mean_squared_error(y_test, y_pred))
print('RMSE:', np.sqrt(mean_squared_error(y_test, y_pred)))
from sklearn.tree import plot_tree
fig = plt.figure(figsize=(25, 20))
# Escolhe uma arvore em toda a floresta, nesse caso a de número 5
_ = plot_tree(rf.estimators_[5], filled=True, feature_names=colunas, max_depth=2)
# ## Importância das variáveis independentes
importancias = list(rf.feature_importances_)
variaveis_importancias = [(coluna, importancia) for coluna, importancia in zip(colunas, importancias)]
variaveis_importancias = sorted(variaveis_importancias, key=lambda x:x[1], reverse=True)
for v, i in variaveis_importancias:
print(f'Variavel: {v:10} - Importancia: {i}')
X_valores = list(range(len(importancias)))
_=plt.bar(X_valores, importancias)
colunas = colunas[:-1]
_=plt.xticks(X_valores, colunas, rotation='vertical')
# ## Variaveis mais importantes
rf_imp = RandomForestRegressor(n_estimators=100, random_state=42)
indices = [colunas.index('temp_1'), colunas.index('average')]
X_train_imp = X_train[:, indices]
X_test_imp = X_test[:, indices]
rf_imp.fit(X_train_imp, y_train)
y_pred = rf_imp.predict(X_test_imp)
print('MAE :', mean_absolute_error(y_test, y_pred))
print('MSE :', mean_squared_error(y_test, y_pred))
print('RMSE:', np.sqrt(mean_squared_error(y_test, y_pred)))
# ## Avaliação de outros modelos
# +
from sklearn.linear_model import LinearRegression
from sklearn.svm import SVR
RANDOM_STATE = 42
def build_regressors():
regressors = []
regressors.append(
('lr',
LinearRegression(),
{'fit_intercept': [True, False]})
)
regressors.append(
('svr',
SVR(),
{'C': [0.1, 1, 10],
'gamma': [0.00001, 0.0001, 0.001, 0.01, 1],
'epsilon': [0.0001, 0.005, 0.001, 1, 5, 10]})
)
return regressors
# +
from sklearn.model_selection import KFold
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_score
train_results = []
test_results = []
names = []
scoring = 'neg_root_mean_squared_error'
regressors = build_regressors()
kfold = KFold(n_splits=3, shuffle=True, random_state=RANDOM_STATE)
for name, regressor, params in regressors:
print(f">>> pregressor : {name}")
# Treinamento
gs = GridSearchCV(regressor, param_grid=params, scoring=scoring, verbose=1)
gs.fit(X_train, y_train)
cv_results = cross_val_score(gs, X_train, y_train, cv=kfold, scoring=scoring) # análise
names.append(name)
train_results.append(abs(cv_results))
# Teste
y_pred = gs.predict(X_test)
rmse = mean_squared_error(y_test, y_pred) ** 0.5
test_results.append(rmse)
# -
fig, ax = plt.subplots()
fig.suptitle('Desempenho em treinamento')
_ = plt.boxplot(train_results)
ax.set_xticklabels(names)
ax.set_ylabel('RMSE')
ax.set_xlabel('Regressores')
fig, ax = plt.subplots()
fig.suptitle('Desempenho em test')
_ = plt.bar(names, test_results)
ax.set_ylabel('RMSE')
ax.set_xlabel('Regressores')
| aprendizado-de-maquina-i/regressao/temps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PCA Tutorial
#
# In an applied math class, you may be told that the principal components of a matrix are the eigenvalues of the matrix's covariance matrix. Depending on your background in statistics and linear algebra, this may or may not have meaning for you. Either way, it's worth unpacking this a bit.
#
# **Variance** refers to how much data points vary around their mean.
#
# **Covariance** is for two or more variables. It refers to how much one variable changes when another one is changed. For example, *height* and *hand size* tend to increase together (taller people have longer hands). If you think that covariance sounds a lot like correlation, you're right! Correlation is just covariance that is scaled to be between -1 and +1.
#
# The **covariance matrix** stores the variance and covariance values between two or more variables. It is always a square and symmetrical matrix. Let's explore one....
# +
# import packages to create data and plot it
import numpy as np
import matplotlib.pyplot as plt
# create two sets of 1000 points sampled from a normal distribution
# re-write the second set to be a noisy version of the first
# Visualize the data
plt.scatter()
plt.axis('equal');
# -
# Compute the covariance matrix of this data using the **_np.cov()_** function.
# +
# Your code here
# -
# The resulting matrix is 2x2 because we have two variables. The items on the *diagonal* (top left and bottom right) refer to the variance of each variable. (Think: why are these close to 1?) The items on the *off-diagonal* are copies of the covariance between these items. Because this is a positive value, we can see that an increase in one variable means an increase in the other.
#
# Now, what about the eigen-stuffs? Consider a matrix A and a vector x. Let's say Let's say that we want to find the vector x that will satisfy the following equation:
#
# \begin{equation*}
# Ax = w
# \end{equation*}
#
# We can think about matrix-vector multiplication as the matrix applying some transformation to x. Therefore, we are trying to find a vector that, when transformed, becomes the w vector. Imagine that vector w is a special case that is simply a scaled (larger or smaller) version of x. In this case, the matrix A is behaving like a scalar. We'll call that scalar.We'll call that a scalar $\lambda$.
#
# \begin{equation*}
# Ax = q = \lambda x
# \end{equation*}
#
# In this case, the vector x is called the eigenvector of matrix A, and $\lambda$ is called its eigenvalue.
#
# The visual analogy is this: An unordered Rubik's cube represents your original matrix A. The process of rotating represents the eigenvectors, and the solved cube represents the matrix when diagonalized, or projected into the new eigenbasis.
#
# <img src="rubiks.png" alt="drawing" width="500"/>
#
# Either way, you can think of the process of PCA this way: by doing an eigen decomposition of the covariance matrix of a set of data, we are finding new axes to explain the data that correspond to the dimensions of the highest correlation.
#
# We'll start out by doing PCA by hand to build our intuition. The first step is to compute eigenvectors and eigenvalues.
# +
# Compute the eigenvectors and eigenvalues of covMat
eigenValue, eigenVector = np.linalg.eig(covMat)
print(eigenValue)
# -
# You can see that the second eigenvalue is much larger than the first. These values are proportional to the amount of variance that the principal components explain. We will order our eigenvectors (the actual components) by their eigenvalues.
# +
# Sort the eigenvectors
eigenVector = np.sort(, axis=1) # axis=1 sorts by column
# Plot the data and eigenvectors
plt.figure()
plt.scatter(,alpha=0.5) # alpha controls the transparency of the points
plt.plot(,c='black') # 1st PC
plt.plot(,c='red') # 2nd PC
# -
# Now that you can do this by hand, we can use one of Python's built-in libraries to do this for us:
# import a machine learning library
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca.fit(X)
print(pca.components_)
# We can check to make sure that these are the same as the ones we previously calculated:
print(eigenVector)
# (Note that the difference is, for example, between the black vector pointing down and to the left versus up and to the right. It's still the same vector).
# A nice thing about using the sklearn PCA is that it will tell us how much variability each of our two PCs explain:
length = pca.explained_variance_
vector = pca.components_
# Let's now plot the data and our PCs again, but this time scale the PCs by the amount of variability they explain.
# +
# create a function that will draw pretty vectors
def drawVector(v0, v1, ax=None):
ax = ax or plt.gca()
arrowprops=dict(arrowstyle='->',
linewidth=2,
shrinkA=0, shrinkB=0)
ax.annotate('', v1, v0, arrowprops=arrowprops)
# plot data
plt.scatter(, alpha=0.2)
for i in range(2):
# Scaling eigen vectors
v = vector[i] * 3 * np.sqrt(length[i])
print(length[i])
drawVector(pca.mean_, pca.mean_ + v[::-1])
plt.axis('equal');
# -
# Now, we can rotate the axis from a data basis to a PC basis:
# +
# set up plotting to have two side-by-side figures
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
fig.subplots_adjust(left=0.0625, right=0.95, wspace=0.1)
# plot data
ax[0].scatter(X[:, 0], X[:, 1], alpha=0.2)
for i in range(2):
v = vector[i] * 3 * np.sqrt(length[i])
drawVector(pca.mean_, pca.mean_ + v, ax=ax[0])
ax[0].axis('equal');
ax[0].set(xlabel='x', ylabel='y', title='input')
# plot principal components
X_pca = pca.transform(X)
ax[1].scatter(X_pca[:, 0], X_pca[:, 1], alpha=0.2)
drawVector([0, 0], [0, -length[1]*10], ax=ax[1])
drawVector([0, 0], [length[0]*2, 0], ax=ax[1])
ax[1].axis('equal')
ax[1].set(xlabel='component 1', ylabel='component 2',
title='principal components',
xlim=(-5, 5), ylim=(-3, 3.1));
# -
# Most of what we will use PCA for is dimensionality reduction. Let's see an example of that.
pca = PCA(n_components=1)
pca.fit(X)
X_pca = pca.transform(X)
print("original shape: ", X.shape)
print("transformed shape:", X_pca.shape)
# Originally, we had two variables. By specifying 1 component, we are asking how well we can reconstruct our data with just the first principal component. To understand the effect of this dimensionality reduction, we can perform the inverse transform of this reduced data and plot it along with the original data:
X_new = pca.inverse_transform(X_pca)
plt.scatter(X[:, 0], X[:, 1], alpha=0.2)
plt.scatter(X_new[:, 0], X_new[:, 1], alpha=0.8)
plt.axis('equal');
# The blue points are the original data, while the orange points are the projected version. This makes clear what a PCA dimensionality reduction means: the information along the least important principal axis or axes is removed, leaving only the component(s) of the data with the highest variance. The fraction of variance that is cut out (proportional to the spread of points about the line formed in this figure) is roughly a measure of how much "information" is discarded in this reduction of dimensionality.
#
# This reduced-dimension dataset is in some senses "good enough" to encode the most important relationships between the points: despite reducing the dimension of the data by 50%, the overall relationship between the data points are mostly preserved.
# ### PCA on images
# The usefulness of the dimensionality reduction may not be entirely apparent in only two dimensions, but becomes much more clear when looking at high-dimensional data. For this example, we'll use a set of hand-written digits that is part of our sklearn library. We'll first load in the data:
from sklearn.datasets import load_digits
digits = load_digits()
np.shape(digits.data)
# This means that there are 1,797 images in our database, each with 64 pixels.
#
# Reshape the first image into 8x8 pictures. They are very small, but we can still recognize the numbers.
firstImage = digits.data[]
firstImage = np.reshape()
plt.imshow(, cmap='Greys')
# compute PCA using first two components
pca = PCA(n_components = ) # project from 64 to 2 dimensions
# Fit data to PCA model
projected = pca.fit_transform()
# Print the shape of the original and projected data
print()
print()
# Now we can project our data onto the first two components to see if the numbers cluster:
plt.scatter( ,c=digits.target, edgecolor='none', alpha=0.5, cmap=plt.cm.get_cmap('Spectral', 10))
plt.xlabel('component 1')
plt.ylabel('component 2')
plt.colorbar();
# ## Reconstructing data with limited components
# Let's see what our first ten digits look like with only the first two components:
# +
pca = PCA(n_components = )
train = pca.fit()
components = train.transform(digits.data)
projected = train.inverse_transform(components)
# Plot the results
fig, ax = plt.subplots(2, 10, figsize=(10, 2.5),
subplot_kw={'xticks':[], 'yticks':[]},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
# Loop through data to plot images from the database and the projected images
for i in range():
# Plotting the digits from the data base
ax[0, i].imshow(digits.data[i].reshape(8, 8), cmap='Greys')
# Plot the image projections
ax[1, i].imshow(, cmap='Greys')
ax[0, 0].set_ylabel('full-dim\ninput')
ax[1, 0].set_ylabel('2-dim\nreconstruction');
# -
# ## Choosing the right number of components
# A vital part of using PCA in practice is the ability to estimate how many components are needed to describe the data. This can be determined by looking at the cumulative explained variance ratio as a function of the number of components:
pca = PCA().fit(digits.data)
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
# This curve quantifies how much of the total, 64-dimensional variance is contained within the first $N$ components. For example, we see that with the digits the first 10 components contain approximately 75% of the variance, while you need around 50 components to describe close to 100% of the variance.
#
# Here we see that our two-dimensional projection loses a lot of information (as measured by the explained variance) and that we'd need about 20 components to retain 90% of the variance. Looking at this plot for a high-dimensional dataset can help you understand the level of redundancy present in multiple observations.
# Using the digits dataset, see what images look like using the first 5, 10, and 20 PCs
# +
# Setting up plotting structure
fig, ax = plt.subplots(4, 10, figsize=(10, 8),
subplot_kw={'xticks':[], 'yticks':[]},
gridspec_kw=dict(hspace=0.1, wspace=0.1))
# Create list to calculate PCA with different number of components
# Initializing counter for plotting
count = 0
# Outer-loop to compute PCA with different number of components
for j in range():
# Index principal component list
# Initialize PCA to compute with given number of components
# Fit data to PCA model
# Define the principal components
# Project components into vector
count = count + 1
# Inner-loop to plot projected images
for i in range():
# Plotting the images from the database
ax[0, i].imshow(digits.data[i].reshape(8, 8), cmap='Greys')
# Plot the projected images of each PCA
ax[count, i].imshow(, cmap='Greys')
ax[0, 0].set_ylabel('full-dim\ninput')
ax[1, 0].set_ylabel('5-dim\nreconstruction');
ax[2, 0].set_ylabel('10-dim\nreconstruction');
ax[3, 0].set_ylabel('20-dim\nreconstruction');
# -
# ## Representational Similarity Analysis
#
# Representational similarity analysis (RSA) is used to analyze the response similarity between various stimuli in selected regions of the brain. This is often visualized by computing and graphically displaying a **Representational Dissimilarity Matrix (RDM)**. RDM's are calculated by measuring the distance between activity patterns in the brain responding to various stimuli. The distance is measured by using multi-dimensional scaling in which each point is transformed into in a 2-dimensional space, optimizing each of their relative distances to all the other conditions.
#
# Ths *RSA_labMaterials.mat* file included in this lab is from an article published in the journal *Neuron* and is titled "Matching categorical object representaions in inferior temperoral cortex of man and monkey". In this experiment, the researchers compared the response of a monkey's inferior temperoral cotrex (IT) and a human's IT to 92 images. These images were split up into two main categories (animate and inanimate) and two subcategories each (human body vs human face and natural vs artificial). They recorded the monkey's with electrodes and the human's with an fMRI, and created two RDM's, one for each species.
#
# Their results are in the following figure:
#
# <img src="IT_RDM.jpg" alt="drawing" width="500"/>
#
# #### The dark-blue represents areas where there is a lot of correlation between various stimuli.
# #### The dark-red represents areas where there is little correlation between various stimuli.
#
#
# ***
# You will attempt to recreate the results of the above paper by using the data stored in the *RSA_labMaterials.mat* file.
#
# A look inside the file:
# - *simTruePatterns* is an array containing the multi-dimensional scaling of all the activity patterns by the human participants of the study.
# - *categoryLabels* is an array containing the name of the various categories
# - *categoryVectors* is an array which...?
#
# Load in the file using the imported **_loadmat()_** function
# Run the following:
from mat2array import loadmat
RSA_labMaterials = loadmat('RSA_labMaterials.mat')
simTruePatterns = RSA_labMaterials['simTruePatterns']
categoryLabels = RSA_labMaterials['categoryLabels']
categoryVectors = RSA_labMaterials['categoryVectors']
# Visualize the *simTruePatterns* and *categoryVectors* matrices in horizontal subplots by using **_plt.imshow_**.
import matplotlib.pyplot as plt
# Your code here
plt.figure(figsize=(7,3))
plt.subplot()
plt.imshow()
plt.subplot()
plt.imshow(, aspect='auto')
# Now, calculate the RDM for *simTruePatterns* by using the imported **_pdist_** function.
# +
from scipy.spatial.distance import pdist
distMat = pdist(, metric='correlation')
# -
# Note that your calculated RDM is a 1-dimensional array. To visualize it, use the imported function **_squareform()_** to make it a square matrix and then use **_plt.imshow()_** to see it.
from scipy.spatial.distance import squareform
distMat = squareform()
plt.figure()
plt.imshow()
# Great! You have created your first RDM based off of real data.
#
# Next, compare the RDM you created to the RDM's in the figure below.
# +
models = loadmat('92_modelRDMs.mat')
Models = models['Models']
plt.figure(figsize=(10,6))
rdmCube = np.zeros((92,92,7))
count = 0
for i in range(len(Models)):
model = vars(Models[i])
RDM = model['RDM']
name = model['name']
if name != 'monkeyIT':
rdmCube[:,:,count] = RDM
count = count + 1
plt.subplot(2,4,count)
plt.title(name)
plt.imshow(RDM)
# -
# You can actually find which models contribute most to the RDM that you made from the human IT. Using the same linear regression method as you learned in the Linear Algebra lab, visualize the predicted RDM and the two models containing the most information with respect to your calculated RDM.
# +
from numpy.linalg import inv
# Create y vector by transforming distMat back to a vector
y = squareform(, force='tovector')
# Initialize data space for each RDM model
dataCube = np.zeros()
for i in range(7):
# Loop through rdmCube to get each RDM
# Flatten each RDM to a vector
# Store each vector in dataCube
# Create vector of ones to help with accuracy
A = np.ones()
# Use vstack to concatenate the vector of ones with dataCube... REMEMBER DIMENSIONALITY
dataCube = np.vstack()
# Calculate the inverse of the dataCube multiplied by the transpose of itself
X = inv(np.matmul())
# Mutliply dataCube by the y vector
D = np.matmul(dataCube,y)
# Multiply your X and D matrices to get your beta matrix
B = np.matmul()
# Calculate predicted RDM
yHat = np.matmul(dataCube.T,B)
# Use squareform to create RDM's of the human RDM, predicted RDM,
# and the two models which influence the human RDM the most.
yHatRDM = squareform(yHat)
model1 = squareform(dataCube[1,:])
model2 = squareform(dataCube[3,:])
# Plotting
plt.figure(figsize=(6,8))
plt.subplot(2,2,1)
plt.imshow(distMat)
plt.title('Human IT')
plt.subplot(2,2,2)
plt.imshow(yHatRDM)
plt.title('Predicted RDM')
plt.subplot(2,2,3)
plt.imshow(model1)
plt.title('The first model: Animacy')
plt.subplot(2,2,4)
plt.imshow(model2)
plt.title('The second model: EVA')
# -
# Do your results makes sense? Why or why not?
#
# Answer in a comment below:
# +
# Answer:
# -
# Now that you have a better idea of what information is primarily found in the human IT, it will be helpful to understand how much information is lost in higher dimensions.
#
# Do a multi-dimensional scaling of the result by using the imported function **_cmdscale_** from **MDS** and plot the eigenvalues.
# +
from MDS import cmdscale
'''
Note: cmdscale returns 2 variables, a Y array and an e array
Y array: Configuration matrix. Each column represents a dimension.
e array: Eigenvalues of B.
'''
Y,e = cmdscale()
plt.figure()
plt.plot()
# -
# How do we determine the right number of dimensions?
# +
# Answer:
# -
# Look at result represented by the two dimensions (columns of Y).
# +
# Your code here
# -
# As you can see, the multi-dimensional scaling represents a cluster of data points, however we have to determine which each group data point corresponds to. The first two columns of the categoryVectors matrix represents the animate vs inanimate categories, with the remaining dimensions corresponding to face, body, natural, and artificial image categories respectively.
#
# In the cell below, plot the animate category in green and the inanimate category in black.
animate, = np.where()
inanimate, = np.where()
plt.figure()
plt.scatter(, c='g')
plt.scatter(, c='k')
# Next, face, body, natural, and artificial dimensions to see where each specific category is plotted.
# +
face, = np.where()
body, = np.where()
nat, = np.where()
art, = np.where()
# Plotting
plt.figure()
plt.scatter(Y2[face,0], Y2[face,1], c='g')
plt.scatter(Y2[body,0], Y2[body,1], c='y')
plt.scatter(Y2[nat,0], Y2[nat,1], c='purple')
plt.scatter(Y2[art,0], Y2[art,1], c='k')
# -
# What do these points correspond to?
# +
# Answer:
# -
# How do you interpret the two dimensions?
# +
# Answer:
# -
# How bad is the 2D approximation?
distMat2 = pdist(Y2, 'correlation')
distMat2 = squareform()
# Plotting
plt.figure()
plt.subplot(1,2,1)
plt.imshow(distMat)
plt.subplot(1,2,2)
plt.imshow(distMat2)
# How much better can we do with more dimensions?
#
# Run the following code and answer in a comment below what happens when you calculate the RDM with an increasing amount of dimensions.
plt.subplots(figsize=(10,10))
count = 1
for dim in range(2,83,10):
thisY = Y[:,:dim]
thisMat = pdist(thisY, 'correlation')
thisMat = squareform(thisMat)
plt.subplot(3,3,count)
plt.imshow(thisMat)
count = count + 1
# +
# Answer:
# -
# Now look at the help file on pdist, and see what happens when you use Euclidean distance.
# +
# Answer:
# -
# That's all for today, once again, great job!
| Lab11/PCA_RSA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JuanPablo20001404/ms-learn-ml-crash-course-python/blob/master/05.%20Logistic%20Regression%20-%20Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zjdkpcypM9Rj"
# Exercise 5 - Logistic Regression
# =====
#
# Logistic regression predicts binary (yes/no) events. For example, we may want to predict if someone will arrive at work on time, or if a person shopping will buy a product.
#
# This exercise will demonstrate simple logistic regression: predicting an outcome from only one feature.
#
# Step 1
# -----
#
# We want to place a bet on the outcome of the next football (soccer) match. It is the final of a competition, so there will not be a draw. We have historical data about our favourite team playing in matches such as this. Complete the exercise below to preview our data.
#
# ### In the cell below replace:
# #### 1. `<addFilePath>` with `'Data/football data.txt' ` (including the quotation marks)
# #### 2. `<printDataHere>` with `print(dataset.head())`
#
# #### and then __run the code__.
# + id="yLuZDhgsQryF"
# + id="yczuHnuaQtXA" outputId="617b2525-6515-455d-a75c-1dbbfaebed6b" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/drive')
# + id="tv6SFA9HM9Rm" outputId="99cc660c-ea89-4682-a409-2ff0287872a9" colab={"base_uri": "https://localhost:8080/", "height": 119}
# This part sets up the graphing configuration
import warnings
warnings.filterwarnings("ignore")
import matplotlib.pyplot as graph
# %matplotlib inline
graph.rcParams['figure.figsize'] = (15,5)
graph.rcParams["font.family"] = 'DejaVu Sans'
graph.rcParams["font.size"] = '12'
graph.rcParams['image.cmap'] = 'rainbow'
import pandas as pd
###
# REPLACE <addFilePath> BELOW WITH '/content/drive/My Drive/Colab Notebooks/Regresión/Data/football data.txt' (INCLUDING THE QUOTES) TO LOAD THE DATA FROM THAT FILE
###
dataset = pd.read_csv('/content/drive/My Drive/Colab Notebooks/Regresión/Data/football data.txt', index_col = False, sep = '\t', header = 0)
###
###
# REPLACE <printDataHere> BELOW WITH print(dataset.head()) TO PREVIEW OUR DATASET
###
print(dataset.head())
###
# + [markdown] id="uxe5DbzkM9Rv"
# This data shows the average goals per match of our team for that season in the left column. In the right column it lists a 1 if our team won the competition or a 0 if they did not.
#
# Step 2
# ----
#
# Let's graph the data so we have a better idea of what's going on here. Complete the exercise below to make an x-y scatter plot.
#
# ### In the cell below replace:
# #### 1. `<addWonCompetition>` with `'won_competition'`
# #### 2. `<addAverageGoals>` with `'average_goals_per_match'`
# #### then __run the code__.
# + id="Yk_5NEwJM9Rx" outputId="296fb1e0-1282-4fa7-c758-8fc4e7bdc896" colab={"base_uri": "https://localhost:8080/", "height": 340}
###
# REPLACE <addWonCompetition> BELOW WITH 'won_competition' (INCLUDING THE QUOTES)
###
train_Y = dataset['won_competition']
###
###
# REPLACE <addAverageGoals> BELOW WITH 'average_goals_per_match' (INCLUDING THE QUOTES)
###
train_X = dataset['average_goals_per_match']
###
# The 'won_competition' will be displayed on the vertical axis (y axis)
# The 'average_goals_per_match' will be displayed on the horizontal axis (x axis)
graph.scatter(train_X, train_Y, c = train_Y, marker = 'D')
graph.yticks([0, 1], ['No', 'Yes'])
graph.ylabel("Competition Win")
graph.ylim([-0.5, 1.5])
graph.xlabel("Average number of goals scored per match")
graph.show()
# + [markdown] id="1ZdiDnMBM9R_"
# We can see from this graph that generally, when our team has a good score average, they tend to win the competition.
#
# Step 3
# ----
#
# How can we predict whether the team will win this season? Let's apply AI to this problem, by making a logisitic regression model using this data and then graph it. This will tell us whether we will likely win this season.
#
# #### Below replace `<buildLinearRegression>` with `linear_model.LogisticRegression()` and then __run the code__.
# + id="9FjUjOIlM9SB"
import numpy as np
from sklearn import linear_model
# Here we build a logistic regression model
###
# REPLACE <buildLinearRegression> BELOW WITH linear_model.LogisticRegression() TO BUILD A LOGISTIC REGRESSION MODEL
###
clf = linear_model.LogisticRegression()
###
# This step fits (calculates) the model
# We are using our feature (x - number of goals scored) and our outcome/label (y - won/lost)
clf.fit(train_X[:, np.newaxis], train_Y)
# This works out the loss
def sigmoid(train_X):
return 1 / (1 + np.exp(-train_X))
X_test = np.linspace(0, 3, 300)
loss = sigmoid(X_test * clf.coef_ + clf.intercept_).ravel()
# + [markdown] id="TpeBwldXM9SJ"
# Alright, that's the model done. Now __run the code__ below to graph it.
# + id="T2LL5f5dM9SM" outputId="b4831640-9226-4ece-e57a-5867995621d6" colab={"base_uri": "https://localhost:8080/", "height": 340}
# This makes the graph
# The data points
graph.scatter(train_X, train_Y, c = train_Y, marker = 'D')
# The curve
graph.plot(X_test, loss, color = 'gold', linewidth = 3)
# Define the y-axis
graph.yticks([0, 1], ['No = 0.0', 'Yes = 1.0'])
graph.ylabel("Competition Win Likelihood")
graph.xlabel("Average number of goals per match")
graph.show()
# + [markdown] id="dvU0uhqBM9SV"
# We now have a line fit to our data. This yellow line is our logistic regression model.
#
# Step 4
# ------
#
# We can read the model above like so:
# * Take the average number of goals per match for the current year. Let's say it is 2.5.
# * Find 2.5 on the x-axis.
# * What value (on the y axis) does the line have at x=2.5?
# * If this value is above 0.5, then the model thinks our team will win this year. If it is less than 0.5, it thinks our team will lose.
#
# Because this line is just a mathematical function (equation) we don't have to do this visually.
#
# In the exercise below, __choose the number of goals you want to evaluate__.
#
# The code will calculate the probability that our team will win with your chosen number of goals in the match.
#
# ### In the cell below replace:
# #### 1. `<numberOfGoals>` with the number of goals in a year (any number from 0 to 3)
# #### 2. `<replaceWithP>` with `p`
# #### then __run the code__.
# + id="MM4ZpsaqM9SX" outputId="6d55aec8-b6f9-430a-d6da-0a453b44598c" colab={"base_uri": "https://localhost:8080/", "height": 374}
###
# REPLACE <numberOfGoals> BELOW WITH THE NUMBER OF GOALS IN A MATCH THIS YEAR. USE ANY NUMBER FROM 0 TO 3
###
p = 2.3
###
# Next we're going to use our model again - clf is the name of our model.
# We'll use a method to predict the probability of a positive result
# Use the variable p which we just made in this method.
###
# REPLACE <replaceWithP> BELOW WITH p TO PREDICT USING THIS VALUE
###
probOfWinning = clf.predict_proba([[ p ]])[0][1]
###
# This prints out the result
print("Probability of winning this year")
print(str(probOfWinning * 100) + "%")
# This plots the result
graph.scatter(train_X, train_Y, c = train_Y, marker = 'D')
graph.yticks([0, probOfWinning, 1], ['No = 0.0', round(probOfWinning,3), 'Yes = 1.0'])
graph.plot(X_test, loss, color = 'gold', linewidth = 3)
graph.plot(p, probOfWinning, 'ko') # result point
graph.plot(np.linspace(0, p, 2), np.full([2],probOfWinning), dashes = [6, 3], color = 'black') # dashed lines (to y-axis)
graph.plot(np.full([2],p), np.linspace(0, probOfWinning, 2), dashes = [6, 3], color = 'black') # dashed lines (to x-axis)
graph.ylabel("Competition Win Likelihood")
graph.xlabel("Average number of goals per match")
graph.show()
# + [markdown] id="EbWkZiNjM9Sf"
# Conclusion
# -----
#
# Well done! We have calculated the likelihood that our team will win this year's competition.
#
# You can go back to the course now and click __'Next Step'__
# + [markdown] id="tSycAUAHM9Sh"
# Optional: Step 5
# -----
#
# Of course, these predictions are only one model.
#
# Let's return to what we did in step 3, but we'll replace `linear_model.LogisticRegression()` with `linear_model.LogisticRegression(C=200)`. This will tell the model to make a steeper decision boundary. Then repeat Step 4 with this boundary. Did your results change?
#
# There are methods we can use to choose sensible parameters for many models. This is currently outside the scope of this course, but it is important to remember that a model is only as good as the data we give it, the parameters we choose, and the assumptions we make.
#
# #### Follow the instructions in the cell below to replace `<numberOfGoals>` and `<buildLinearRegression>` and __run the code__.
# + id="CGNXeyNHM9Si" outputId="ea982224-e51c-4486-a0ad-2c47cc432a80" colab={"base_uri": "https://localhost:8080/", "height": 374}
# Let's do that again.
# We will repeat what we did in step 3, but change the decision boundary.
import numpy as np
from sklearn import linear_model
###
# REPLACE THE <numberOfGoals> WITH THE NUMBER OF GOALS YOU WANT TO EVALUATE
###
p = 2.3
###
# Here we build the new logistic regression model.
# The C=200 is where we change the decision boundary.
###
# REPLACE <buildLinearRegression> BELOW WITH linear_model.LogisticRegression(C=200) TO BUILD A LOGISTIC REGRESSION MODEL
###
clf = linear_model.LogisticRegression(C=200)
###
# This step fits (calculates) the model
# We are using our feature (x - number of goals scored) and our outcome/label (y - won/lost)
clf.fit(train_X[:, np.newaxis], train_Y)
# This works out the loss
def sigmoid(train_X):
return 1 / (1 + np.exp(-train_X))
X_test = np.linspace(0, 3, 300)
loss = sigmoid(X_test * clf.coef_ + clf.intercept_).ravel()
# This makes the prediction for your chosen number of goals.
probOfWinning = clf.predict_proba([[p]])[0][1]
# This prints out the result.
print("Probability of winning this year")
print(str(probOfWinning * 100) + "%")
# This plots the result.
graph.scatter(train_X, train_Y, c = train_Y, marker = 'D')
graph.yticks([0, probOfWinning, 1], ['No = 0.0', round(probOfWinning,3), 'Yes = 1.0'])
graph.plot(X_test, loss, color = 'gold', linewidth = 3)
graph.plot(p, probOfWinning, 'ko') # result point
graph.plot(np.linspace(0, p, 2), np.full([2],probOfWinning), dashes = [6, 3], color = 'black') # dashed lines (to y-axis)
graph.plot(np.full([2],p), np.linspace(0, probOfWinning, 2), dashes = [6, 3], color = 'black') # dashed lines (to x-axis)
graph.ylabel("Competition Win Likelihood")
graph.xlabel("Average number of goals per match")
graph.show()
| 05. Logistic Regression - Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import geopandas as gpd
huyen = gpd.read_file("data/Bai2_Chuyen doi he TD/HUYEN_region.shp")
huyen.crs
huyen.plot()
huyen_mercator = huyen.to_crs(epsg=3395)
huyen_mercator.plot()
huyen_mercator.crs
| Bai 2 - Chuyen doi he TD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 1: Data Cleaning
#
# Let's start with getting the datafiles rounds.csv and companies.txt.
#
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# reading data files
# using encoding = "ISO-8859-1" to avoid pandas encoding error
rounds = pd.read_csv("rounds2.csv", encoding = "ISO-8859-1")
companies = pd.read_csv("companies.txt", sep="\t", encoding = "ISO-8859-1")
# -
# Look at rounds head
print(rounds.head())
# inspect the structure etc.
print(rounds.info(), "\n")
print(rounds.shape)
# The variables funding_round_code and raised_amount_usd contain some missing values, as shown above. We'll deal with them after we're done with understanding the data - column names, primary keys of tables etc.
# look at companies head
companies.head()
# companies structure
companies.info()
# Ideally, the ```permalink``` column in the companies dataframe should be the unique_key of the table, having 66368 unique company names (links, or permalinks). Also, these 66368 companies should be present in the rounds file.
#
# Let's first confirm that these 66368 permalinks (which are the URL paths of companies' websites) are not repeating in the column, i.e. they are unique.
# identify the unique number of permalinks in companies
len(companies.permalink.unique())
# Also, let's convert all the entries to lowercase (or uppercase) for uniformity.
# converting all permalinks to lowercase
companies['permalink'] = companies['permalink'].str.lower()
companies.head()
# look at unique values again
len(companies.permalink.unique())
# Thus, there are 66368 unique companies in the table and ```permalink``` is the unique primary key. Each row represents a unique company.
#
# Let's now check whether all of these 66368 companies are present in the rounds file, and if some extra ones are present.
# look at unique company names in rounds df
# note that the column name in rounds file is different (company_permalink)
len(rounds.company_permalink.unique())
# There seem to be 90247 unique values of ```company_permalink```, whereas we expected only 66368. May be this is because of uppercase/lowercase issues.
#
# Let's convert the column to lowercase and look at unique values again.
# converting column to lowercase
rounds['company_permalink'] = rounds['company_permalink'].str.lower()
rounds.head()
# Look at unique values again
len(rounds.company_permalink.unique())
# There seem to be 2 extra permalinks in the rounds file which are not present in the companies file. Let's hope that this is a data quality issue, since if this were genuine, we have two companies whose investment round details are available but their metadata (company name, sector etc.) is not available in the companies table.
# Let's have a look at the company permalinks which are in the 'rounds' file but not in 'companies'.
# companies present in rounds file but not in (~) companies file
rounds.loc[~rounds['company_permalink'].isin(companies['permalink']), :]
# All the permalinks have weird non-English characters. Let's see whether these characters are present in the original df as well.
# looking at the indices with weird characters
rounds_original = pd.read_csv("rounds2.csv", encoding = "ISO-8859-1")
rounds_original.iloc[[29597, 31863, 45176, 58473], :]
# The company weird characters appear when you import the data file. To confirm whether these characters are actually present in the given data or whether python has introduced them while importing into pandas, let's have a look at the original CSV file in Excel.
#
# The figure below shows the filtered rows - they have the usual English characters.
#
# <img src="./weird_names.PNG">
# Thus, this is most likely a data quality issue we have introduced while reading the data file into python. Specifically, this is most likely caused because of encoding.
#
# First, let's try to figure out the encoding type of this file. Then we can try specifying the encoding type at the time of reading the file. The ```chardet``` library shows the encoding type of a file.
# +
# import chardet
# rawdata = open('rounds2.csv', 'rb').read()
# result = chardet.detect(rawdata)
# charenc = result['encoding']
# print(charenc)
# +
# print(result)
# -
# Now let's try telling pandas (at the time of importing) the encoding type. Here's a list of various encoding types python can handle: https://docs.python.org/2/library/codecs.html#standard-encodings.
# +
# trying different encodings
# encoding="cp1254" throws an error
# rounds_original = pd.read_csv("rounds2.csv", encoding="cp1254")
# rounds_original.iloc[[29597, 31863, 45176], :]
# -
# Apparently, pandas cannot decode "cp1254" in this case. After searching a lot on stackoverflow and Google, the best conclusion that can be drawn is that this file is encoded using multiple encoding types (may be because the ```company_permalink``` column contains names of companies in various countries, and hence various languages).
#
# After trying various other encoding types (in vain), this answer suggested an alternate (and a more intelligent) way: https://stackoverflow.com/questions/45871731/removing-special-characters-in-a-pandas-dataframe.
#
#
rounds['company_permalink'] = rounds.company_permalink.str.encode('utf-8').str.decode('ascii', 'ignore')
rounds.loc[~rounds['company_permalink'].isin(companies['permalink']), :]
# This seems to work fine.
#
# Let's now look at the number of unique values in rounds dataframe again.
# Look at unique values again
len(rounds.company_permalink.unique())
# Now it makes sense - there are 66368 unique companies in both the ```rounds``` and ```companies``` dataframes.
#
# It is possible that a similar encoding problems are present in the companies file as well. Let's look at the companies which are present in the companies file but not in the rounds file - if these have special characters, then it is most likely because the ```companies``` file is encoded (while rounds is not).
# companies present in companies df but not in rounds df
companies.loc[~companies['permalink'].isin(rounds['company_permalink']), :]
# Thus, the ```companies``` df also contains special characters. Let's treat those as well.
# remove encoding from companies df
companies['permalink'] = companies.permalink.str.encode('utf-8').str.decode('ascii', 'ignore')
# Let's now look at the companies present in the companies df but not in rounds df - ideally there should be none.
# companies present in companies df but not in rounds df
companies.loc[~companies['permalink'].isin(rounds['company_permalink']), :]
# Thus, the encoding issue seems resolved now. Let's write these (clean) dataframes into separate files so we don't have to worry about encoding problems again.
# +
# write rounds file
rounds.to_csv("rounds_clean.csv", sep=',', index=False)
# write companies file
companies.to_csv("companies_clean.csv", sep='\t', index=False)
| 3. Investment Analysis Assignment/Solution/1_Data_Cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup Antennal Lobe Simulation Libraries
import networkx as nx
from neurokernel.LPU.InputProcessors.StepInputProcessor import StepInputProcessor
import numpy as np
name = 'DM4_step1_'
work_path = ''
out_path = ''
G_path = work_path+'{}.gexf'.format(name)
visual_components = np.load(work_path+'visual_components_{}.npy'.format(name), allow_pickle=True)
visual_neurons = np.load(work_path+'visual_neurons_{}.npy'.format(name), allow_pickle=True).item()
def large_scale_sim(name, G_path, conc = 1., DM4_b = 2.17 * 1e-2, DM4_d = 2.94, DL5_b = 0.1 * 2.94, exp_name = '', LN_3=False, LN_3_I = 10.):
G = nx.read_gexf(G_path)
G = nx.MultiDiGraph(G)
sim_name = name + '_conc_' + str(conc) + exp_name
def set_br(G):
for node in G.nodes():
if '_OTP' in node:
if 'DC1' in node:
G.nodes(data=True)[node]['br'] = 0.
elif 'DM4' in node:
G.nodes(data=True)[node]['br'] = DM4_b
G.nodes(data=True)[node]['dr'] = DM4_d
else:
G.nodes(data=True)[node]['br'] = DL5_b
G.nodes(data=True)[node]['dr'] = DM4_d
return G
G = set_br(G)
inputs = [i for i in G.nodes() if '_OTP' in i]
fi = StepInputProcessor('conc', inputs, conc, start=1., stop=3.)
print('Concentration Level:', conc)
print('Inputs:', inputs)
def simulate(G, t, inputs,
record_var_list = None,
sample_interval: int = 100,
):
from neurokernel.LPU.LPU import LPU
from neurokernel.LPU.InputProcessors.BaseInputProcessor import (
BaseInputProcessor,
)
from neurokernel.LPU.InputProcessors.ArrayInputProcessor import (
ArrayInputProcessor,
)
from neurokernel.LPU.OutputProcessors.OutputRecorder import OutputRecorder
from neurokernel.LPU.OutputProcessors.FileOutputProcessor import FileOutputProcessor
dt = t[1] - t[0]
if isinstance(inputs, BaseInputProcessor):
fi = [inputs]
elif isinstance(inputs, (list, tuple, np.ndarray)) and isinstance(
inputs[0], BaseInputProcessor
):
fi = inputs
elif isinstance(inputs, dict):
for data in inputs.values():
assert "uids" in data
assert "data" in data
assert isinstance(data["data"], np.ndarray)
fi = [ArrayInputProcessor(inputs)]
else:
raise ValueError("Input not understood")
if LN_3:
fi += [StepInputProcessor('I', ['LN_3'], LN_3_I, start=1., stop=3.)]
print('Adding current to LN3.')
fo = OutputRecorder([('I', None), ('g', None)], sample_interval=sample_interval)
fo2 = OutputRecorder([('spike_state', None)], sample_interval=1)
fo3 = OutputRecorder([('V', None)], sample_interval=1)
lpu = LPU(
dt,
"obj",
G,
device=0,
id="EOS",
input_processors=fi,
output_processors=[fo, fo2, fo3],
debug=False,
manager=False
)
lpu.run(steps=len(t))
fo.to_file(out_path+'fo_{}'.format(sim_name))
fo2.to_file(out_path+'fo2_{}'.format(sim_name))
fo3.to_file(out_path+'fo3_{}'.format(sim_name))
return fi, fo, fo2, fo3, lpu
return simulate, G, fi
# ## Simple Simulation Setup
import time
import gc
# names = ['DM4_DL5_step4_a','DM4_DL5_step4_b'] # a: keep synapses to DL5; b: keep synapses to DM4
names = ['DM4_nov_general']
simulation_traces = {}
for name in names:
for DM4_b_coeff in [0.0,0.2,0.4,0.6,0.8,1.0]:
for DL5_b_coeff in [0.0]:
time.sleep(1)
conc = 10.
G_path = '/mnt/server-home/mehmet/alclustering/{}.gexf'.format(name)
visual_components = np.load('/mnt/server-home/mehmet/alclustering/visual_components_{}.npy'.format(name), allow_pickle=True)
visual_neurons = np.load('/mnt/server-home/mehmet/alclustering/visual_neurons_{}.npy'.format(name), allow_pickle=True).item()
simulate, G, fi = large_scale_sim(name, G_path, conc=conc, DM4_b = DM4_b_coeff * 0.3, DM4_d = 2.94, DL5_b = DL5_b_coeff * 0.3,
exp_name = '_changing_balanced_default_' + str(DM4_b_coeff) + '_' + str(DL5_b_coeff), LN_3=True, LN_3_I = 20.)
dt = 1e-5
dur = 4.
steps = int((dur+dt/2)//dt)
t = np.arange(steps)*dt
fi, fo, fo2, fo3, lpu = simulate(G, t, fi)
| feedbackcircuits/examples/ALRun.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Methods and Tools
# + [markdown] slideshow={"slide_type": "slide"}
# ## Interactive Lectures
#
# **All lectures in the course will be _interactive_**
#
# They contain running code, as well as theory!
#
# * Presented and discussed in frontal lectures...
# * ...You can download PDFs
# * ...But you will also be able to _make changes and experiment_
#
# **From a software perspective, the workshorses of this approach are:**
#
# * [Jupyter](https://jupyter.org) notebooks for the presentation & interaction
# * [Docker](https://www.docker.com) containers for the setup and distribution
#
# **Both are widely used systems:**
#
# * Jupyter is a user favorite when it comes to data science
# * Docker is a state-of-the-art system for manageing services
# + [markdown] slideshow={"slide_type": "slide"}
# # A Few Words about Docker
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Docker
#
# **Docker is a system for running software in _"containers"_**
#
# <center><img src="assets/vertical-logo-monochromatic.png" width="400"/></center>
#
#
# Think of a container as a _lightweight virtual machine_:
#
# * (Essentially) the same level of isolation
# * ...But smaller disk footprint, faster setup and operation, etc.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Docker
#
# **Docker is a system for running software in _"containers"_**
#
# <center><img src="assets/vertical-logo-monochromatic.png" width="400"/></center>
#
# Using containers has _many advantages_:
#
# * Multiple environments on the same machine
# * Improved isolation, robustness, and reproducibility
# * Easier replication (scalability of cloud services)
# * ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Docker
#
# **During this course we will see many problems**
#
# ...And tackle them with _many techniques_:
#
# * Classical Machine Learning
# * Deep Learning
# * Statistics
# * Signal processing
# * Declarative optimization
# * Differential Equations
# * Agent based simulation
# * ...
#
# Managing dependences would quickly become hellish
# + [markdown] slideshow={"slide_type": "subslide"}
#
# ## Docker
#
# **With docker, we can simply use _a different container per case study_**
#
# _Inside each container_ we will have:
#
# * All the needed libraries & tools
# * A running instance of a Jupyter server
#
# _In the host_ machine (your PC):
#
# * We will just open a browser...
# * ...And connect to the Jupyter server
#
# **Two key concepts in Docker**
#
# * A _container_ is a (sort of) running, lightweight, Virtual Machine
# * An _image_ is (sort of) the content of the hard disk of the VM
#
# The image can be used to instantiate multiple containers
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Building an Image
#
# **Images in docker are built by:**
#
# * Starting from a base image on [Docker Hub](https://hub.docker.com)
# * Copying content between the host and the container
# * Running commands in the container
#
# **The process is controlled via a Dockerfile**
#
# * Just a text file with a specific syntax
# * There is an [extensive reference](https://docs.docker.com/engine/reference/builder/), but we only care about a few commands
#
# **To build an image, we can use:**
#
# ```sh
# docker build .
# ```
#
# ...From the directory with the Dockerfile
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **This is a _minimal Dockerfile_ for this lecture:**
#
# ```dockerfile
# FROM python:3.8
# RUN pip install jupyter pandas sklearn matplotlib ipympl RISE
# COPY . /app
# WORKDIR /app/notebooks
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", "--ip=0.0.0.0", "--allow-root"]
# ```
#
# * The `FROM` keyword specifies the base image
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **This is a _minimal Dockerfile_ for this lecture:**
#
# ```dockerfile
# FROM python:3.8
# RUN pip install jupyter pandas sklearn matplotlib ipympl RISE
# COPY . /app
# WORKDIR /app/notebooks
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", "--ip=0.0.0.0", "--allow-root"]
# ```
#
# * The `RUN` keyword runs a command
# * In our case, we install a number of python packages
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **This is a _minimal Dockerfile_ for this lecture:**
#
# ```dockerfile
# FROM python:3.8
# RUN pip install jupyter pandas sklearn matplotlib ipympl RISE
# COPY . /app
# WORKDIR /app/notebooks
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", "--ip=0.0.0.0", "--allow-root"]
# ```
#
# * The `COPY` keyword transfers data from the host to the container
# * The first path refers to the host
# * The second path to the container
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **This is a _minimal Dockerfile_ for this lecture:**
#
# ```dockerfile
# FROM python:3.8
# RUN pip install jupyter pandas sklearn matplotlib ipympl RISE
# COPY . /app
# WORKDIR /app/notebooks
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", "--ip=0.0.0.0", "--allow-root"]
# ```
#
# * The `WORKDIR` changes the current directory in the container
# * It's like running `cd` in the container
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **This is a _minimal Dockerfile_ for this lecture:**
#
# ```dockerfile
# FROM python:3.8
# RUN pip install jupyter pandas sklearn matplotlib ipympl RISE
# COPY . /app
# WORKDIR /app/notebooks
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", "--ip=0.0.0.0", "--allow-root"]
# ```
#
# * The `CMD` keyword is triggered only when we _run_ a container
# * It's the first command that the container should execute
# * It does nothing when building an image
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **When we run `docker build .` for our file:**
#
# * The docker daemon downloads the base image, if not already available
# * A container is started
# * All operations in the Dockerfile are executed
# * The resulting container is dumped, to create an _image_
#
# **You can check that a new image has been built using:**
#
# ```sh
# docker image ls
# ```
#
# You will see an entry with no name:
#
# ```sh
# REPOSITORY TAG IMAGE ID CREATED SIZE
# <none> <none> 96b910c1514f 3 seconds ago 1.36GB
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ## An Example Dockerfile
#
# **You can assign a name to an image using:**
#
# ```sh
# docker build . -t <name of the image>
# ```
#
# **You can remove an image with:**
#
# ```sh
# docker image rm <image name or id>
# ```
#
# * Useful to free space, however...
# * ...Images are incremental! Docker stores only the differences
# * ...So, don't worry too much about space usage
#
# **You can remove all images with no running container with:**
#
# ```sh
# docker image prune
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Running a Container
#
# **You can instantiate and run a container with:**
#
# ```sh
# docker run <image name or id>
# ```
#
# * The container `stdout` will be piped (i.e. connected) to your terminal
# * By default, this is not the case for `stdin`
# * You can make the container interactive with the `-it` options
# * You can autoremove the container at the end with `--rm`
# * You can sync folders in the host and on the container using [volumes](https://docs.docker.com/storage/volumes/)
#
# The [documentation](https://docs.docker.com/engine/reference/run/) is extensive
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Running a Container
#
# **You can obtain the list of all containers with:**
#
# ```sh
# docker ps
# ```
#
# * The option `-a` shows all containers (incl. those that are stopped)
#
# **You can remove a container with**
#
# ```sh
# docker rm <container id>
# ```
#
# * As you see, it's a very flexible system
# * ...But also a bit complex
#
# **That's why we will automate most operations using _Docker Compose_**
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Docker Compose
#
# **_Docker Compose_ is a tool to help the management of containers**
#
# In a second `docker-compose.yml` file, you specify:
#
# * Which "services" (i.e. container) should be built and run
# * How to build them
# * Which options to use when running them
# * ...
#
# All in a [human-readable, declarative format](https://yaml.org)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## A Docker Compose Example
#
# **Let's see the `docker-compose.yml` for this lecture:**
#
# ```yaml
# version: '2.0'
# services:
# jupyter:
# build: .
# ports:
# - "8888:8888"
# volumes:
# - .:/app
# ```
#
# * `version` refers to the Docker Compose syntax
# * `services` is followed by a list of the containers
# * `jupyter` is our service
# * `build` specifies where the `Dockerfile` can be found
# + [markdown] slideshow={"slide_type": "subslide"}
# ## A Docker Compose Example
#
# **Let's see the `docker-compose.yml` for this lecture:**
#
# ```yaml
# version: '2.0'
# services:
# jupyter:
# build: .
# ports:
# - "8888:8888"
# volumes:
# - .:/app
# ```
#
# * `ports` tells which ports to expose to `docker run`
# * `volumes` specifies which folders to sync
# * In our case "/app/notebooks" on the container
# * ...Will actually be "./notebooks" on the host
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Benefits of Using Docker Compose
#
# **We need to use one more tool, but now we can:**
#
# _Build and run_ a container with:
#
# ```sh
# docker-compose up
# ```
#
# * The command can also restart a stopped container
#
# _Stop_ the container with CTRL+C, or with:
#
# ```sh
# docker-compose stop
# ```
#
# Stop _and remove_ the container with:
#
# ```sh
# docker-compose down
# ```
#
# ...Which is considerably simpler than before!
# + [markdown] slideshow={"slide_type": "slide"}
# # Our Jupyter Setup
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our Setup
#
# **We will often work with this development setup**
#
# The folder with the notebooks is structured as follows:
#
# ```
# notebook1.pynb
# notebook2.pynb
# ...
# util <-- module
# assets <-- images and such
# rise.css <-- for the "slide" mode
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our Setup
#
# **We will often work with this development setup**
#
# The folder with the notebooks is structured as follows:
#
#
# ```
# notebook1.pynb
# notebook2.pynb
# ...
# util +-- __init__.py
# +-- XYZ.py <-- submodule
# +-- YZX.py <-- submodule
# +-- ...
# font
# rise.css
# ```
#
# **Most important part:** the use of _a module_ besides notebooks
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our Setup
#
# **Working with modules provides some advantages:**
#
# We do not need to keep all our code in the notebooks. We can:
#
# * _Share_ functions _between cells_
# * _Share_ functions _between notebooks_
# * IDEs can offer _more functionality_ if they recognize a module
#
# **...But also a significant disadvantage:**
#
# * Python modules are compiled first when loaded...
# * ...The loaded version is _not updated_ when the source changes
#
# This is very inconvenient at development time
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our Setup
#
# **We can circumvent this thanks to Jupyter "magic" extensions**
#
# The first one is [the "autoreload" extension](https://ipython.org/ipython-doc/3/config/extensions/autoreload.html)
# + slideshow={"slide_type": "-"}
# %load_ext autoreload
# %autoreload 2
# -
#
# * `load_ext` will enable the extension
# * `autoreload 2` will reload all modules before code execution
#
# **This is _inefficient, but convenient_ during development**
#
# * Together with the use of volumes (in docker-compose)...
# * ...This allows us to update the code without re-building the docker image
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Starting a Notebook
#
# Let's look back to the `CMD` keyword in our `Dockerfile`:
#
# ```dockerfile
# CMD ["jupyter", "notebook", "--port=8888", "--no-browser", \
# "--ip=0.0.0.0", "--allow-root"]
# ```
#
# This is translated to:
#
# ```sh
# jupyter notebook --port=8888 --no-browser --ip=0.0.0.0 --allow-root
# ```
#
# * `--port 8888`: the server listen on port 8888
# * `--no-browser`: do not open the browser (there's no browser in the container)
# * `--ip=0.0.0.0`: listen on all network interfaces
# * `--allow-root`: we operate as `root` (admin) on the container
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Starting a Notebook
#
# When we run:
#
# ```sh
# docker-compose up
# ```
#
# The output will look like:
# ```sh
# Starting ad_stat_jupyter_1 ... done
# ...
# ...Use Control-C to stop this server and shut down all kernels...
# ...
# ...To access the notebook...
# ...copy and paste one of these URLs:
# ... http://34b908cf2362:8888/?token=<PASSWORD>...
# ... or http://127.0.0.1:8888/?token=<PASSWORD>...
# ```
#
# * The last URL can be _copy-pasted in your favorite browser_
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Starting a Notebook
#
# When we run:
#
# ```sh
# docker-compose up
# ```
#
# The output will look like:
# ```sh
# Starting ad_stat_jupyter_1 ... done
# ...
# ...Use Control-C to stop this server and shut down all kernels...
# ...
# ...To access the notebook...
# ...copy and paste one of these URLs:
# ... http://34b908cf2362:8888/?token=<PASSWORD>...
# ... or http://127.0.0.1:8888/?token=<PASSWORD>...
# ```
#
# * The `token` is cached by the browser and grants access to the notebooks
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our Setup
#
# * We will also use the `ipympl` package and the `widget` jupyter magic
# * This will display basic tools to rescale and zoom images
# -
# %matplotlib widget
from matplotlib import pyplot as plt
import numpy as np
x = np.linspace(0, 2*np.pi, 100)
plt.figure(figsize=(9, 3))
plt.plot(x, np.sin(x))
plt.tight_layout()
| notebooks/1. Our Setup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Battle sequence simulator
#
# Simulate a single large horde trying to move through a chain of defenders.
#
# To run the simulation, type the size of your attacking horde and the number of defenders at each point in the chain below. The list of defending armies should be inside [square brackets] and be separated by commas. Press `Shift+Enter` or click Run in the menu to run the simulation and see the results!
#
# If the simulation is taking too long (e.g. because the armies are very large), you can redice the number of trials to speed things up.
# +
# --- Choose your settings
NUM_ATTACKING_ARMIES = 10
DEFENDING_ARMY_CHAIN = [3,1,4]
NUM_TRIALS = 100000
# --- You don't need to touch this part
import risk
# %matplotlib inline
outcomes = risk.battle_sequence_report(
NUM_ATTACKING_ARMIES,
DEFENDING_ARMY_CHAIN,
NUM_TRIALS
)
| attack-probabilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xarray as xr
import datetime
# load data
dataset = xr.open_dataset('/home/joao/Downloads/ERA5_20160823.nc')
# print variables of file
print(dataset)
# gets fields
uwind = dataset['u']
vwind = dataset['v']
# define the region of interest
Lon0, Lon1 = 310.0, 340.0 # longitud
Lat0, Lat1 = -50.0, -10.0 # latitud
Lev0, Lev1 = 1000, 200 # pressure level
Time0, Time1 = datetime.datetime(2016,8,23,0,0), datetime.datetime(2016,8,23,18,0) # DateTime
# +
# select region of interest
# here the uwind and vwind are slice but keeping their 4D array
uwind = uwind.sel(time=slice(Time0, Time1), level=slice(Lev1, Lev0),
latitude=slice(Lat1, Lat0), longitude=slice(Lon0, Lon1))
vwind = vwind.sel(time=slice(Time0, Time1), level=slice(Lev1, Lev0),
latitude=slice(Lat1, Lat0), longitude=slice(Lon0, Lon1))
# check the shape of the uwind and vwind
print(uwind.shape, vwind.shape)
# +
# calculate vorticity and divergence
import metlib
vor = metlib.relative_vorticity(uwind, vwind)*1.0e5
# +
# creates a custom color palette with custom_color_palette package
# see https://github.com/joaohenry23/custom_color_palette
import numpy as np
import matplotlib.pyplot as plt
import custom_color_palette as ccpl
mypalette, colorslabels, norm = ccpl.creates_palette([plt.cm.bwr],[-5.5,5.5])
# creating colorbar labels
tickslabels = np.arange(-5.5, 6.5, 1.0)
# +
# PLOTTING LATITUDINAL PROFILE OF VORTICITY [OPTION 01: USING MATPLOTLIB AND PCOLORMESH]
# define the profile
LonPro0 = 330.0 # longitud
LatPro0, LatPro1 = -50.0, -10.0 # latitud
LevPro0, LevPro1 = 1000, 200 # pressure level
TimePro0 = datetime.datetime(2016,8,23,18,0) # DateTime
# here vor is slice, passing from 4D array to 2D array !
vorProf = vor.sel(time=TimePro0, level=slice(LevPro1, LevPro0),
latitude=slice(LatPro1, LatPro0), longitude=LonPro0)
# gets lats and levs
latsProf = vorProf['latitude'].values
levsProf = vorProf['level'].values
# calculate corners of latsProf and levsProf
latsProfCor = np.concatenate((latsProf+(latsProf[0]-latsProf[1])/2.0,
[latsProf[-1]-(latsProf[0]-latsProf[1])/2.0]))
levsProfCor = np.concatenate((levsProf+(levsProf[0]-levsProf[1])/2.0,
[levsProf[-1]-(levsProf[0]-levsProf[1])/2.0]))
# creates 2D array of latsProfCor and levsProfCor
latsProfCor2D, levsProfCor2D = np.meshgrid(latsProfCor, levsProfCor)
# creates figure
fig = plt.figure('Latitudinal Profile of Vorticity Matplotlib Pcolormesh',figsize=(4,4), dpi=200)
ax = fig.add_axes([0.1, 0.15, 0.80, 0.75])
# plot image
img = ax.pcolormesh(latsProfCor2D, levsProfCor2D, vorProf, cmap=mypalette, norm=norm)
# customizing the plot border
plt.rcParams['axes.linewidth'] = 1.0
# plot colorbar
cbar = plt.colorbar(img, ticks=tickslabels, extend='neither',
spacing='proportional', orientation='horizontal',
cax=fig.add_axes([0.20, 0.03, 0.60, 0.025]))
cbar.ax.tick_params(labelsize=4, labelcolor='black',
width=0.5, length=1.2, direction='out', pad=1.5)
cbar.set_label(r'Vorticity [$10^{5}$/s]', size=5, color='black',
weight='normal', labelpad=2.0)
cbar.outline.set_linewidth(0.5)
# sets labels
xticks = np.arange(-50.0,0.0,10.0)
ax.set_xticks(xticks)
ax.set_xticklabels(xticks, size=6)
ax.set_xlim(xticks.min(),xticks.max())
ax.set_xlabel('Latitude', fontdict={'fontsize':7}, labelpad=2.0)
yticks = np.arange(1000,150,-50)
ax.set_yscale("log")
ax.set_yticks(yticks)
ax.set_yticklabels(yticks, size=6)
ax.set_ylim(yticks.max(),yticks.min())
ax.set_ylabel('Pressure level (hpa)', fontdict={'fontsize':7}, labelpad=2.0)
ax.set_title('{}\n{}\nLon: {} DateTime: {}'
.format(r'Example using $\bf{Matplotlib}$ and $\bf{Pcolormesh}$ [$\bf{option}$ $\bf{01}$]',
r'$\bf{Latitudinal}$ $\bf{Profile}$ $\bf{of}$ $\bf{Relative}$ $\bf{Vorticity}$',
LonPro0, TimePro0.strftime('%Y/%m/%d %H:%M GMT')),
fontdict={'fontsize':6}, pad=6.0)
plt.show()
# +
# PLOTTING LATITUDINAL PROFILE OF VORTICITY [OPTION 02: USING MATPLOTLIB AND CONTOURF]
# define the profile
LonPro0 = 330.0 # longitud
LatPro0, LatPro1 = -50.0, -10.0 # latitud
LevPro0, LevPro1 = 1000, 200 # pressure level
TimePro0 = datetime.datetime(2016,8,23,18,0) # DateTime
# here vor is slice, passing from 4D array to 2D array !
vorProf = vor.sel(time=TimePro0, level=slice(LevPro1, LevPro0),
latitude=slice(LatPro1, LatPro0), longitude=LonPro0)
# gets lats and levs
latsProf = vorProf['latitude'].values
levsProf = vorProf['level'].values
# creates 2D array of latsProf and levsProf
latsProfCen2D, levsProfCen2D = np.meshgrid(latsProf, levsProf)
# creates figure
fig = plt.figure('Latitudinal Profile of Vorticity Matplotlib Contourf',figsize=(4,4), dpi=200)
ax = fig.add_axes([0.1, 0.15, 0.80, 0.75])
# plot image
img = ax.contourf(latsProfCen2D, levsProfCen2D, vorProf, cmap=mypalette,
levels=tickslabels, extend='both')
# customizing the plot border
plt.rcParams['axes.linewidth'] = 1.0
# plot colorbar
cbar = plt.colorbar(img, ticks=tickslabels, extend='neither',
spacing='proportional', orientation='horizontal',
cax=fig.add_axes([0.20, 0.03, 0.60, 0.025]))
cbar.ax.tick_params(labelsize=4, labelcolor='black',
width=0.5, length=1.2, direction='out', pad=1.5)
cbar.set_label(r'Vorticity [$10^{5}$/s]', size=5, color='black',
weight='normal', labelpad=2.0)
cbar.outline.set_linewidth(0.5)
# sets labels
xticks = np.arange(-50.0,0.0,10.0)
ax.set_xticks(xticks)
ax.set_xticklabels(xticks, size=6)
ax.set_xlim(xticks.min(),xticks.max())
ax.set_xlabel('Latitude', fontdict={'fontsize':7}, labelpad=2.0)
yticks = np.arange(1000,150,-50)
ax.set_yscale("log")
ax.set_yticks(yticks)
ax.set_yticklabels(yticks, size=6)
ax.set_ylim(yticks.max(),yticks.min())
ax.set_ylabel('Pressure level (hpa)', fontdict={'fontsize':7}, labelpad=2.0)
ax.set_title('{}\n{}\nLon: {} DateTime: {}'
.format(r'Example using $\bf{Matplotlib}$ and $\bf{Contourf}$ [$\bf{option}$ $\bf{02}$]',
r'$\bf{Latitudinal}$ $\bf{Profile}$ $\bf{of}$ $\bf{Relative}$ $\bf{Vorticity}$',
LonPro0, TimePro0.strftime('%Y/%m/%d %H:%M GMT')),
fontdict={'fontsize':6}, pad=6.0)
plt.show()
# +
# PLOTTING TEMPORAL PROFILE OF VORTICITY IN ONE POINT [OPTION 01: USING MATPLOTLIB AND PCOLORMESH]
# define the profile
LonPro0 = 330.0 # longitud
LatPro0 = -30.0 # latitud
LevPro0, LevPro1 = 1000, 200 # pressure level
TimePro0, TimePro1 = datetime.datetime(2016,8,23,0,0), datetime.datetime(2016,8,23,18,0) # DateTime
# here vor is slice, passing from 4D array to 2D array !
vorProf = vor.sel(time=slice(TimePro0, TimePro1), level=slice(LevPro1, LevPro0),
latitude=LatPro0, longitude=LonPro0)
# here transpose vorProf array
vorProf = (vorProf.T)
# gets time and levs
timesProf = vorProf['time'].values.astype('datetime64[s]')
levsProf = vorProf['level'].values
# calculate corners of timesProf and levsProf
timesProfCor = np.concatenate((timesProf+(timesProf[0]-timesProf[1])/2.0,
[timesProf[-1]-(timesProf[0]-timesProf[1])/2.0]))
levsProfCor = np.concatenate((levsProf+(levsProf[0]-levsProf[1])/2.0,
[levsProf[-1]-(levsProf[0]-levsProf[1])/2.0]))
# creates 2D array of timesProfCor and levsProfCor
timesProfCor2D, levsProfCor2D = np.meshgrid(timesProfCor, levsProfCor)
# creates figure
fig = plt.figure('Temporal Profile of Vorticity Matplotlib Pcolormesh',figsize=(4,4), dpi=200)
ax = fig.add_axes([0.1, 0.15, 0.80, 0.75])
# plot image
img = ax.pcolormesh(timesProfCor2D, levsProfCor2D, vorProf, cmap=mypalette, norm=norm)
# customizing the plot border
plt.rcParams['axes.linewidth'] = 1.0
# plot colorbar
cbar = plt.colorbar(img, ticks=tickslabels, extend='neither',
spacing='proportional', orientation='horizontal',
cax=fig.add_axes([0.20, 0.03, 0.60, 0.025]))
cbar.ax.tick_params(labelsize=4, labelcolor='black',
width=0.5, length=1.2, direction='out', pad=1.5)
cbar.set_label(r'Vorticity [$10^{5}$/s]', size=5, color='black',
weight='normal', labelpad=2.0)
cbar.outline.set_linewidth(0.5)
# sets labels
xticks = [time.strftime('%Y/%m/%d %Hh') for time in timesProf.tolist()]
ax.set_xticks(timesProf)
ax.set_xticklabels(xticks, size=5.5)
ax.set_xlim(timesProf.min(),timesProf.max())
ax.set_xlabel('DateTime', fontdict={'fontsize':7}, labelpad=2.0)
yticks = np.arange(1000,150,-50)
ax.set_yscale("log")
ax.set_yticks(yticks)
ax.set_yticklabels(yticks, size=6)
ax.set_ylim(yticks.max(),yticks.min())
ax.set_ylabel('Pressure level (hpa)', fontdict={'fontsize':7}, labelpad=2.0)
ax.set_title('{}\n{}\nLon: {} Lat: {}'
.format(r'Example using $\bf{Matplotlib}$ and $\bf{Pcolormesh}$ [$\bf{option}$ $\bf{01}$]',
r'$\bf{Temporal}$ $\bf{Profile}$ $\bf{of}$ $\bf{Relative}$ $\bf{Vorticity}$ '+
r'$\bf{in}$ $\bf{one}$ $\bf{point}$',
LonPro0, LatPro0),
fontdict={'fontsize':6}, pad=6.0)
plt.show()
# +
# PLOTTING TEMPORAL PROFILE OF VORTICITY IN ONE POINT [OPTION 02: USING MATPLOTLIB AND CONTOURF]
# define the profile
LonPro0 = 330.0 # longitud
LatPro0 = -30.0 # latitud
LevPro0, LevPro1 = 1000, 200 # pressure level
TimePro0, TimePro1 = datetime.datetime(2016,8,23,0,0), datetime.datetime(2016,8,23,18,0) # DateTime
# here vor is slice, passing from 4D array to 2D array !
vorProf = vor.sel(time=slice(TimePro0,TimePro1), level=slice(LevPro1, LevPro0),
latitude=LatPro0, longitude=LonPro0)
# here transpose array
vorProf = (vorProf.T)
# gets time and levs
timesProf = vorProf['time'].values.astype('datetime64[s]')
levsProf = vorProf['level'].values
# creates 2D array of timesProf and levsProf
timesProfCen2D, levsProfCen2D = np.meshgrid(timesProf, levsProf)
# creates figure
fig = plt.figure('Temporal Profile of Vorticity Matplotlib Contourf',figsize=(4,4), dpi=200)
ax = fig.add_axes([0.1, 0.15, 0.80, 0.75])
# plot image
img = ax.contourf(timesProfCen2D, levsProfCen2D, vorProf, cmap=mypalette,
levels=tickslabels, extend='both')
# customizing the plot border
plt.rcParams['axes.linewidth'] = 1.0
# plot colorbar
cbar = plt.colorbar(img, ticks=tickslabels, extend='neither',
spacing='proportional', orientation='horizontal',
cax=fig.add_axes([0.20, 0.03, 0.60, 0.025]))
cbar.ax.tick_params(labelsize=4, labelcolor='black',
width=0.5, length=1.2, direction='out', pad=1.5)
cbar.set_label(r'Vorticity [$10^{5}$/s]', size=5, color='black',
weight='normal', labelpad=2.0)
cbar.outline.set_linewidth(0.5)
# sets labels
xticks = [time.strftime('%Y/%m/%d %Hh') for time in timesProf.tolist()]
ax.set_xticks(timesProf)
ax.set_xticklabels(xticks, size=5.5, rotation=0.0, ha='center', va='center')
ax.set_xlim(timesProf.min(),timesProf.max())
ax.set_xlabel('DateTime', fontdict={'fontsize':7}, labelpad=2.0)
ax.tick_params(axis='x', which='major', pad=6.5)
yticks = vorProf['level'].values
ax.set_yscale("log")
ax.set_yticks(yticks)
ax.set_yticklabels(yticks, size=6)
ax.set_ylim(yticks.max(),yticks.min())
ax.set_ylabel('Pressure level (hpa)', fontdict={'fontsize':7}, labelpad=2.0)
ax.set_title('{}\n{}\nLon: {} Lat: {}'
.format(r'Example using $\bf{Matplotlib}$ and $\bf{Contourf}$ [$\bf{option}$ $\bf{02}$]',
r'$\bf{Temporal}$ $\bf{Profile}$ $\bf{of}$ $\bf{Relative}$ $\bf{Vorticity}$ '+
r'$\bf{in}$ $\bf{one}$ $\bf{point}$',
LonPro0, LatPro0),
fontdict={'fontsize':6}, pad=6.0)
plt.show()
# -
| examples/ex05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## English Wikipedia Political Figures Articles Coverage and Quality Analysis
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import pandas as pd
import numpy as np
# ### Step 1: Set up page and population data
# **Wikipedia dataset** from Figshare.com: https://figshare.com/articles/Untitled_Item/5513449
# This dataset is downloaded from Figshare.com. <br/>
# The dataset is titled "Politicians by Country from the English-language Wikipedia", of which are data extracted from Wikimedia thru API calls. <br/>
# Both the dataset and the code used to extract the data are under CC-BY-SA 4.0 license. <br/>
# It is downloadable as a csv file titled "page_data.csv", and there are three columns and 47,197 rows in the csv file. <br/>
#
# page: article title of the page for political figures, not cleaned yet
# country: cleaned version of country name from which the category the political figure is under
# rev_id: unique identifier for revision tracking
## load page_data.csv into pandas DataFrame and examine first 5 rows
page_data = pd.read_csv('page_data.csv', sep=',')
page_data.head()
# **Population dataset** from Dropbox: https://www.dropbox.com/s/5u7sy1xt7g0oi2c/WPDS_2018_data.csv?dl=0
# This dataset is downloaded from Dropbox. <br/>
# The dataset is originally from Population Reference Bureau under International Indicators <br/>
# and it is population data for all countries from mid-2018 in millions of population. <br/>
# It is downloadable as a csv file titled "WPDS_2018_data.csv", and there are two columns and 207 rows in the csv file. <br/>
#
# Geography: country and continent names
# Population mid-2018 (millions): population data from mid-2018 in millions
## load WPDS_2018_data.csv into pandas DataFrame and examine first 5 rows
population_data = pd.read_csv('WPDS_2018_data.csv', sep=',', thousands=',')
population_data.head()
# ### Step 2: Set up article quality predictions
# For the article quality predictions, we will be using ORES API calls by passing in each articles' rev_id and getting their 'prediction' values from the json file.
# For ORES documentation, please refer to this website: https://ores.wikimedia.org/v3/#!/scoring/get_v3_scores_context
# For prediction values in ORES, there are 6 quality categories, in later analysis, we will mainly focus on the first two categories for high quality article percentage calculation.
#
# FA - Featured article
# GA - Good article
# B - B-class article
# C - C-class article
# Start - Start-class article
# Stub - Stub-class article
## import packages for making API calls to ORES
import requests
import json
## Define hearders and endpoint for API call
headers = {'User-Agent' : 'https://github.com/yd4wh', 'From' : '<EMAIL>'}
endpoint = 'https://ores.wikimedia.org/v3/scores/{project}/?models={model}&revids={revids}'
## Define a function that will recurse over all rev_ids and output quality predictions
def get_ores_quality_prediction(revids, headers, endpoint):
# define parameters for endpoints
params = {'project' : 'enwiki',
'model' : 'wp10',
'revids' : '|'.join(str(x) for x in revids)
}
# use above defined parameters to make API requests
api_call = requests.get(endpoint.format(**params))
response = api_call.json()
# loop thru each revids in the 100 group to get their quality predictions
quality_prediction = []
revid_list = []
# After testing, there are errors with revids that don't have a score associated
# therefore, when looping, also included except to pass the revids that don't have scores associated with them
for revid in revids:
# to iterate thru every revids to pull out the prediction value
try:
quality_prediction.append(response['enwiki']['scores'][str(revid)]['wp10']['score']['prediction'])
revid_list.append(revid)
# use except to pass thru the revids without scores
except:
pass
# this function will return revids and its associated prediction values
return revid_list, quality_prediction
# +
## set up revids in every 100 to be passed thru the API call function
# change revids into list
revids = list(page_data['rev_id'])
# define starting and ending points for first iteration
start = 0
end = 100
# create empty Dataframe for collecting article quality output
article_quality = pd.DataFrame()
# loop over all revids in groups of 100s
while start < len(revids):
# pull out the revids in groups of 100 for each iteration
iter_revids = revids[start:end]
# call the function to get article quality predictions
iter_result = get_ores_quality_prediction(iter_revids,headers,endpoint)
article_quality = article_quality.append(pd.DataFrame(list(iter_result)).T)
# update starting and ending points for next iteration
start += 100
end = min(start+100, len(revids))
# print out the final Dataframe with the revids that don't have scores
article_quality.head()
# -
# The ORES article quality prediction dataframe is now saved as **article_quality** with 2 columns and 47,092 rows after removing all articles that doesn't have a article score.
#
# revision_id: the revision_id that can be linked back to page_data
# article_quality: the ORES quality prediction for associated revision_id
# rename article_quality columns before merging in next step
article_quality.rename(columns={0:'revision_id',1:'article_quality'}, inplace=True)
article_quality.head()
# ### Step 3: Combine page_data, population_data and article_quality
# This step will use the common columns in page_data(rev_id, country), population_data(Geography), article_quality(revision_id) to merge all three dataframes together, and in the end build a combined dataframe together with 5 columns and 44,973 rows after removing all data point that don't match.
#
# country: country column from page_data
# article_name: page column from page_data
# revision_id: revision_id column from article_quality
# article_quality: article_quality column from article_quality
# population: Population mid-2018 (millions) column from population_data which will be in millions
# make deep copies of the three dataframes as base df for merging
df_page_data = page_data.copy(deep=True)
df_population_data = population_data.copy(deep=True)
df_article_quality = article_quality.copy(deep=True)
# +
# combine page_data and article_quality on rev_id and revision_id columns
combined_data = df_page_data.merge(df_article_quality, how='right', left_on='rev_id', right_on='revision_id')
# combine combined_data with population_data on country and geography columns
combined_data = combined_data.merge(df_population_data, how='inner', left_on='country', right_on='Geography')
combined_data.rename(columns={'page':'article_name','Population mid-2018 (millions)':'population'}, inplace=True)
combined_data.head()
# -
# clean dataframe for combined_data to just keep five colunms documented above
df_combined_data = combined_data[['country',
'article_name',
'revision_id',
'article_quality',
'population']]
df_combined_data.head()
# output final_data.csv for reproducibiilty
df_combined_data.to_csv('final_data.csv', index=False)
# ### Step 4: Analysis on articles quality by country and population
# make a deep copy of the final DataFrame for analysis
final_data = df_combined_data.copy(deep=True)
# The **percentage of articles-per-population** for each country: this measure will be calculated by taking the total number of articles in a particular country and divide it by the total population of the corresponding country. This requires us to sum the total number of articles by country and to represent population number normally.
# count total number of articles in each country using group by
article_by_country = final_data.groupby('country').count()['article_name']
# pass the series into a dataframe for merging with population data
df_article_by_country = article_by_country.to_frame(name='article_count')
# change country into a column instead of index for merging
df_article_by_country['country'] = df_article_by_country.index
df_article_by_country.head()
# merge with population_data df to calculate percentage
articles_per_population = df_article_by_country.merge(df_population_data, how='inner',
left_on='country', right_on='Geography')
# change population number into normal presentation
articles_per_population['population'] = articles_per_population['Population mid-2018 (millions)']*1000000
# calculate the percentage of articles per population by country
articles_per_population['pcnt_articles_per_population'] = 100*(articles_per_population['article_count']/articles_per_population['population'])
# The **percentage of high-quality-articles** for each country: this measure will be calculated by taking the total number of articles in a particular country that qualifies as being either "FA" or "GA" and divide it by the total number of articles about politicians of the corresponding country.
# limit articles to only "FA" and "GA" qualities
high_quality_articles = final_data.loc[final_data['article_quality'].isin(['FA','GA'])]
# count total number of high quality articles in each country using group by
quality_article_by_country = high_quality_articles.groupby('country').count()['article_name']
# pass the series into a dataframe for merging with population data
df_quality_article_by_country = quality_article_by_country.to_frame(name='high_quality_article_count')
# change country into a column instead of index for later merging
df_quality_article_by_country['country'] = df_quality_article_by_country.index
df_quality_article_by_country.head()
# merge with articles_per_population df to calculate article percentage
analysis_df = df_quality_article_by_country.merge(articles_per_population, how='right',
left_on='country', right_on='country')
# divide total number of high quality articles by total article count
analysis_df['pcnt_high_quality_articles'] = 100*(analysis_df['high_quality_article_count']/analysis_df['article_count'])
analysis_df.head()
# The combined analysis DataFrame will include all countries that have population and wikipedia articles regardless of the count of high quality articles.
# keep neccessary and non-duplicate columns
analysis_df = analysis_df[['country',
'article_count',
'high_quality_article_count',
'population',
'pcnt_articles_per_population',
'pcnt_high_quality_articles']]
analysis_df.head()
# ### Step 5: Tables of highest and lowest ranked countries by *articles_per_population* and *high_quality_articles*
# This section will display four tables that summarize the 10 highest and 10 lowest ranked countries in terms of their pcnt_articles_per_population and pcnt_high_quality_articles in the order below:
#
# 1. 10 highest-ranked countries in terms of pcnt_articles_per_population
# 2. 10 lowest-ranked countries in terms of pcnt_articles_per_population
# 3. 10 highest-ranked countries in terms of pcnt_high_quality_articles
# 4. 10 lowest-ranked countries in terms of pcnt_high_quality_articles
# 10 highest-ranked countries sorting by 'pcnt_articles_per_population'
analysis_df.sort_values(by='pcnt_articles_per_population', ascending=False).head(10)[['country',
'article_count',
'population',
'pcnt_articles_per_population']]
# 10 lowest-ranked countries sorting by 'pcnt_articles_per_population'
analysis_df.sort_values(by='pcnt_articles_per_population').head(10)[['country',
'article_count',
'population',
'pcnt_articles_per_population']]
# 10 highest-ranked countries sorting by 'pcnt_high_quality_articles'
analysis_df.sort_values(by='pcnt_high_quality_articles', ascending=False).head(10)[['country',
'high_quality_article_count',
'article_count',
'pcnt_high_quality_articles']]
# 10 lowest-ranked countries sorting by 'pcnt_high_quality_articles'
analysis_df.sort_values(by='pcnt_high_quality_articles').head(10)[['country',
'high_quality_article_count',
'article_count',
'pcnt_high_quality_articles']]
# One caveat on the lowest-ranked countries in terms of pcnt_high_quality_articles, we only included countries that have at least 1 article qualified as "GA" or "FA" and didn't include countries that don't have any high quality articles about politicians. Therefore, as a separate group of countries that don't have any high quality articles written about politicians, we've listed below in alphabetical order. There are 37 countries that dont have any articles qualified as "GA" or "FA".
countries_without_high_qulaity_articles = analysis_df.loc[pd.isnull(analysis_df['pcnt_high_quality_articles'])]
print("There are "+ str(countries_without_high_qulaity_articles.count()[1]) + " countries that don't have any high quality articles.")
countries_without_high_qulaity_articles[['country']]
| hcds-a2-bias.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="AW0RxUlxYIda" colab={"base_uri": "https://localhost:8080/"} outputId="a4d55390-13f4-48bf-f8e8-4d1892103b43"
import pandas as pd
import io
import re
from os import listdir
import io
import random
import json
import datetime
import argparse
import sys
# #!wget https://raw.githubusercontent.com/danielinux7/Multilingual-Parallel-Corpus/master/ab-ru/quran.tsv
# + colab={"base_uri": "https://localhost:8080/"} id="0YObtr_fI8lP" outputId="10137209-acd2-4ba2-dcce-f44fc1cfd65e"
alphabet_ab = re.compile('[ҟцукенгшәзхҿфывапролджҽџчсмитьбҩҵқӷӡҳԥҷҭ\.\:,;\ 0-9-\(\)"!?]+',re.I)
alphabet_ru = re.compile('[ёйцукенгшщзхъфывапролджэячсмитьбю\.\:,;\ 0-9-\(\)"!?]+',re.I)
alphabe_ab = re.compile('ҟцукенгшәзхҿфывапролджҽџчсмитьбҩҵқӷӡҳԥҷҭ',re.I)
alphabe_ru = re.compile('ёйцукенгшщзхъфывапролджэячсмитьбю',re.I)
sentence_signs = re.compile('[\.\:!?]+',re.I)
#corp=pd.read_csv("/content/quran.tsv", sep='\t')
def mis_punctuation(parallel_corpus):
filtered_punctuations = 0
ru_result_list = []
ab_result_list = []
for translation in parallel_corpus:
ru_signs = re.sub(alphabe_ru, "", translation[0]) #replacing alphabe_ru with "" in string translation[0]
ab_signs = re.sub(alphabe_ab, "", translation[1]) #replacing alphabe_ab with "" in string translation[1]
if ru_signs == ab_signs: #if both strings have same punctuattion
ru_result_list.append(translation[0])
ab_result_list.append(translation[1])
else:
filtered_punctuations += 1
print("filtered punctuations: "+str(filtered_punctuations))
return list(zip(ru_result_list,ab_result_list))
#corpa=filter_punctuation(corp)
# + id="kK_GSoZEyGFd"
| utils/Mispronunciation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clasificación II
# Ya hemos hablado que los problemas de clasificación son de los más solicitados en el _mundo real_, por lo que en esta clase veremos otro algoritmo de clasificación, pero en esta ocasión, uno no paramaétrico.
# ## K Nearest Neighbours
# El algoritmo **k Nearest Neighbors (kNN)** es un método no paramétrico: una vez que el parámetro $k$ se ha fijado, no se busca obtener ningún parámetro adicional.
#
# Sean los puntos $x^{(i)} = (x^{(i)}_1, ..., x^{(i)}_n)$ de etiqueta $y^{(i)}$ conocida, para $i=1, ..., m$.
#
# El problema de clasificación consiste en encontrar la etiqueta de un nuevo punto $x=(x_1, ..., x_m)$ para el cual no conocemos la etiqueta.
#
# 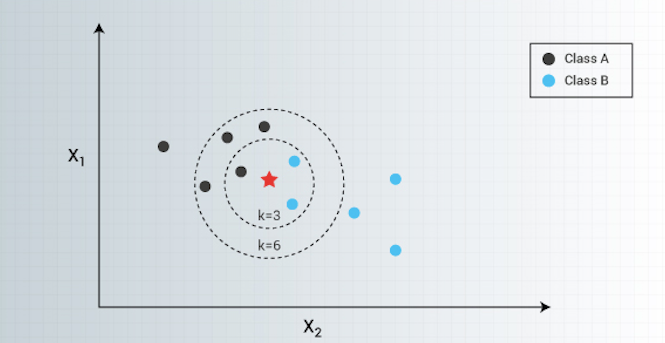
#
# [source](https://www.kdnuggets.com/2019/07/classifying-heart-disease-using-k-nearest-neighbors.html)
# La etiqueta de un punto se obtiene de la siguiente forma:
# * Para $k=1$, **1NN** asigna a $x$ la etiqueta de su vecino más cercano.
# * Para $k$ genérico, **kNN** asigna a $x$ la etiqueta más popular de los k vecinos más cercanos.
#
# La idea es la siguiente:
#
# 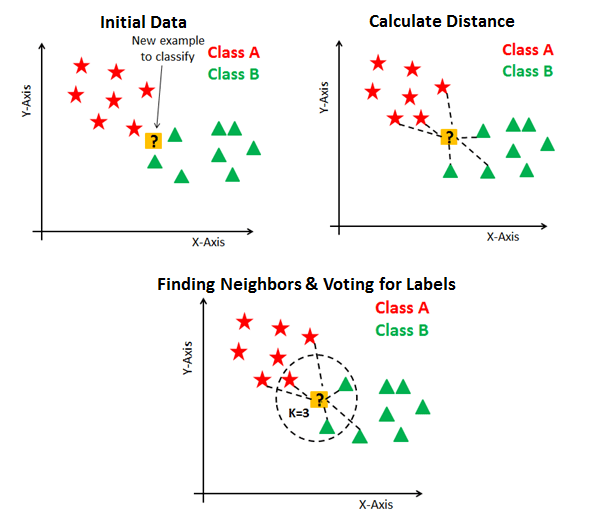
# El modelo subyacente a kNN es el conjunto de entrenamiento completo. A diferencia de otros métodos que efectivamente generalizan y resumen la información (como regresión logística, por ejemplo), cuando se necesita realizar una predicción, el algoritmo kNN mira **todos** los datos y selecciona los k datos más cercanos, para regresar la etiqueta más popular/más común. Los datos no se resumen en parámetros, sino que siempre deben mantenerse en memoria. Es un método por tanto que no escala bien con un gran número de datos.
# En caso de empate, existen diversas maneras de desempatar:
# * Elegir la etiqueta del vecino más cercano (problema: no garantiza solución).
# * Elegir la etiqueta de menor valor (problema: arbitrario).
# * Elegir la etiqueta que se obtendría con $k+1$ o $k-1$ (problema: no garantiza solución, aumenta tiempo de cálculo).
# La cercanía o similaridad entre los datos se mide de diversas maneras, pero en general depende del tipo de datos y del contexto.
#
# * Para datos reales, puede utilizarse cualquier distancia, siendo la **distancia euclidiana** la más utilizada. También es posible ponderar unas componentes más que otras. Resulta conveniente normalizar para poder utilizar la noción de distancia más naturalmente.
#
# * Para **datos categóricos o binarios**, suele utilizarse la [distancia de Hamming](https://docs.scipy.org/doc/scipy/reference/generated/scipy.spatial.distance.hamming.html).
# A continuación, una implementación de _from scratch_ en numpy:
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def knn_search(X, k, x):
""" find K nearest neighbours of data among D """
# Distancia euclidiana
d = np.linalg.norm(X - x, axis=1)
# Ordenar por cercania
idx = np.argsort(d)
# Regresar los k mas cercanos
id_closest = idx[:k]
return id_closest, d[id_closest].max()
def knn(X,Y,k,x):
# Obtener los k mas cercanos
k_closest, dmax = knn_search(X, k, x)
# Obtener las etiquetas
Y_closest = Y[k_closest]
# Obtener la mas popular
counts = np.bincount(Y_closest.flatten())
# Regresar la mas popular (cualquiera, si hay empate)
return np.argmax(counts), k_closest, dmax
def plot_knn(X, Y, k, x):
y_pred, neig_idx, dmax = knn(X, Y, k, x)
# plotting the data and the input point
fig = plt.figure(figsize=(8, 8))
plt.plot(x[0, 0], x[0, 1], 'ok', ms=16)
m_ob = Y[:, 0] == 0
plt.plot(X[m_ob, 0], X[m_ob, 1], 'ob', ms=8)
m_sr = Y[:,0] == 1
plt.plot(X[m_sr, 0], X[m_sr, 1], 'sr', ms=8)
# highlighting the neighbours
plt.plot(X[neig_idx, 0], X[neig_idx, 1], 'o', markerfacecolor='None', markersize=24, markeredgewidth=1)
# Plot a circle
x_circle = dmax * np.cos(np.linspace(0, 2*np.pi, 360)) + x[0, 0]
y_circle = dmax * np.sin(np.linspace(0, 2*np.pi, 360)) + x[0, 1]
plt.plot(x_circle, y_circle, 'k', alpha=0.25)
plt.show();
# Print result
if y_pred==0:
print("Prediccion realizada para etiqueta del punto = {} (circulo azul)".format(y_pred))
else:
print("Prediccion realizada para etiqueta del punto = {} (cuadrado rojo)".format(y_pred))
# -
# Puedes ejecutar varias veces el código anterior, variando el número de vecinos `k` para ver cómo afecta el algoritmo.
k = 5 # hyper-parameter
N = 50
np.random.seed(42)
# +
X = np.random.rand(N, 2) # random dataset
Y = np.array(np.random.rand(N) < 0.4, dtype=int).reshape(N, 1) # random dataset
x = np.random.rand(1, 2) # query point
# performing the search
plot_knn(X, Y, k, x)
# -
# ## Aplicación
import pandas as pd
from sklearn import datasets
# Utilizaremos un dataset de dígitos escritos a mano para clasificar según el valor de los píxeles
# +
# print(digits_dict["DESCR"]) # Descomenta la línea si quieren más info del dataset
# -
digits_X, digits_y = datasets.load_digits(return_X_y=True, as_frame=True)
digits = pd.concat([digits_X, digits_y], axis=1)
digits.head()
# Para hacerlo un poco más gráfico, veamos la imagen del primer número
# +
digit_images = datasets.load_digits().images
plt.imshow(digit_images[0], cmap=plt.cm.gray_r, interpolation='nearest');
# -
# Como (casi) siempre, Scikit-learn nos ofrece los algoritmos en la misma sintaxis a la que ya nos hemos ido acostumbrando.
# +
from sklearn.neighbors import KNeighborsClassifier
k = 5
neigh = KNeighborsClassifier(n_neighbors=k)
neigh.fit(digits_X, digits_y)
# -
# ¿Qué pasó? Veamos algún registro en particular
i = 0
digits_X.iloc[[i], :]
# Y su respectiva predicción
neigh.predict(digits_X.iloc[[i], :])
neigh.predict_proba(digits_X.iloc[[i], :])
# ¡No tiene ninguna gracia! Porque el elemento más cercano es él mismo! Adelantemos un poco de las clases siguientes
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(digits_X, digits_y, test_size=0.25, random_state=42)
# -
# Hagamos el ajuste solo con datos de entrenamiento
neigh_tt = KNeighborsClassifier(n_neighbors=k)
neigh_tt.fit(X_train, y_train)
i_test = 22
neigh_tt.predict(X_test.iloc[[i_test], :])
neigh_tt.predict_proba(X_test.iloc[[i_test], :])
plt.imshow(X_test.iloc[[i_test], :].to_numpy().reshape(8, 8), cmap=plt.cm.gray_r, interpolation='nearest');
| lessons/M4L04_classification_nonparametric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 참고연구 목록
# : 연구 주제 관련 직, 간접적 관련 선행연구 및 참고자료 (총 22건)
# ```
# 1) 이종우(2014), 실업팀 선수의 연봉 결정요인 분석 : 개인종목을 중심으로. 서울대학교 대학원
# 2) 권병진(2005). 미국 프로야구 Major League 선수들의 노동생산성과 연봉에 관한 연구. 서강대학교 대학원
# 3) 송종우(2008). 한국 프로스포츠 선수들의 연봉에 대한 다변량적분석. 응용통계연구, 21(3), 441-45
# 4) 승희배, 강기훈(2012). 한국 프로야구 선수들의 경기력과 연봉의 관계 분석. 한국데이터정보과학회지, 23(2), 285-298.
# 5) 신문선(2003). 한국 프로축구 선수의 연봉산정 모델 개발. 국내박사학위논문, 세종대학교 대학원, 서울.
# 6) 오광모, 이장택(2003). 데이터마이닝을 이용한 한국프로야구 선수들이 연봉에 관한 모형연구. 한국스포츠사회학회지,16(2), 295-309.
# 7) 한동섭, 김정기, 김종(2007). 스포츠 스타의 이미지와 소속 팀, 보증 제품 및 사회 이미지의 관계에 관한 연구. 한국스포츠산업.경영학회지, 12(3), 155-168.
# 8) 최명일(2002). 스포츠 스타 이미지 구성요인에 관한 연구. 국내석사학위논문, 한양대학교 대학원, 서울.
# 9) 손동만(2018). NBA 선수 연봉구조와 선수 스타파워가 팀 퍼포먼스에 미치는 영향. 서울대학교 대학원
# 10) 황선욱(2019). 주성분 기법을 활용한 NBA선수 포지션별 선수평가지수 및 연봉예측모델 개발. 동국대학교 대학원
# 11) <NAME>(2018). A Multiple Linear Regression Approach For Estimating the Market Value of Football Players in Forward Position|
# 12) <NAME>(2018), Player valuation in European football
# 13) <NAME>(2019). Predicting 2018–19 NBA’s Most Valuable Player using Machine Learning
# 14) <NAME>(2018). Can Linear Models predict a Footballer’s Value?
# 15) Yaldo, L.(2017). Computational Estimation of Football Player Wages
# 16) <NAME>(2017). BEYOND CROWD JUDGMENTS: DATA-DRIVEN ESTIMATION OF MARKET VALUE IN ASSOCIATION FOOTBALL
# 17) 김설애(2016). 원산지국가이미지가 소비자인식 및 구매의도에 미치는 영향
# 18) 문효진(2016). 국가브랜드 평판 구성요인 도출을 위한 탐색적 연구
# 19) <NAME>. & <NAME>. (1993). Measuring a Multi-Dimensional Construct: Country Image
# 20) <NAME>(2007). 국가역량 종합지수(Composite Index of National Capability, CNIC)
# 21) US News(2020). US News 최고의 나라 지수
# 22) 강지원(2020). Why do Instagram Users Tag Friends in Comments
# ```
# # 참고연구 분류 및 시사점
#
# **1. 데이터 측면**
#
# - 해외 축구에 대한 국내 연구는 거의 전무한 상태. 더군다나 ML알고리즘을 활용한 연구는 현재까지 발견하지 못함
# - 해외 축구에 대한 해외 연구는 회귀, ML알고리즘 활용한 연구가 다수 있어, 비교 연구하기에 적절.
# - (대상 데이터셋은 유사하나, 방법론 측면(선형회귀, ML알고리즘 각각 1건씩)에서 다른 논문 2건 발견)
#
# ```
#
#
# ```
# ```
# 1) 국내축구
# - 이종우(2014), 실업팀 선수의 연봉 결정요인 분석 : 개인종목을 중심으로. 서울대학교 대학원
# - 신문선(2003). 한국 프로축구 선수의 연봉산정 모델 개발. 국내박사학위논문, 세종대학교 대학원, 서울.
# - 한동섭, 김정기, 김종(2007). 스포츠 스타의 이미지와 소속 팀, 보증 제품 및 사회 이미지의 관계에 관한 연구. 한국스포츠산업.경영학회지, 12(3), 155-168.
#
# 2) 해외축구
# - 최명일(2002). 스포츠 스타 이미지 구성요인에 관한 연구. 국내석사학위논문, 한양대학교 대학원, 서울.
# - <NAME>(2018). A Multiple Linear Regression Approach For Estimating the Market Value of Football Players in Forward Position
# - <NAME>(2018), Player valuation in European football
# - Yaldo, L.(2017). Computational Estimation of Football Player Wages
# - <NAME>(2018). Can Linear Models predict a Footballer’s Value?
# - <NAME>(2017). BEYOND CROWD JUDGMENTS: DATA-DRIVEN ESTIMATION OF MARKET VALUE IN ASSOCIATION FOOTBALL
#
# 3) 타 스포츠
# - 권병진(2005). 미국 프로야구 Major League 선수들의 노동생산성과 연봉에 관한 연구. 서강대학교 대학원
# - 송종우(2008). 한국 프로스포츠 선수들의 연봉에 대한 다변량적분석. 응용통계연구, 21(3), 441-45
# - 승희배, 강기훈(2012). 한국 프로야구 선수들의 경기력과 연봉의 관계 분석. 한국데이터정보과학회지, 23(2), 285-298.
# - 오광모, 이장택(2003). 데이터마이닝을 이용한 한국프로야구 선수들이 연봉에 관한 모형연구. 한국스포츠사회학회지,16(2), 295-309.
# - 손동만(2018). NBA 선수 연봉구조와 선수 스타파워가 팀 퍼포먼스에 미치는 영향. 서울대학교 대학원
# - 황선욱(2019). 주성분 기법을 활용한 NBA선수 포지션별 선수평가지수 및 연봉예측모델 개발. 동국대학교 대학원
# - <NAME>(2019). Predicting 2018–19 NBA’s Most Valuable Player using Machine Learning
#
# ```
#
# ```
#
#
# ```
# **2. 방법 측면 : 최근 연구에서 ML 방법론의 연구가 두드러지나, 동시에 크롤링 데이터를 토대로 한 회귀 연구도 함께 활발히 진행 중**
#
# ```
# 1) 회귀 (6개)
# - 이종우(2014), 실업팀 선수의 연봉 결정요인 분석 : 개인종목을 중심으로. 서울대학교 대학원
# - 신문선(2003). 한국 프로축구 선수의 연봉산정 모델 개발. 국내박사학위논문, 세종대학교 대학원, 서울.
# - 동섭, 김정기, 김종(2007). 스포츠 스타의 이미지와 소속 팀, 보증 제품 및 사회 이미지의 관계에 관한 연구. 한국스포츠산업.경영학회지,
# - 손동만(2018). NBA 선수 연봉구조와 선수 스타파워가 팀 퍼포먼스에 미치는 영향. 서울대학교 대학원
# - <NAME>(2018). A Multiple Linear Regression Approach For Estimating the Market Value of Football Players in Forward Position
# - <NAME>(2018). Can Linear Models predict a Footballer’s Value?
# - <NAME>(2017). BEYOND CROWD JUDGMENTS: DATA-DRIVEN ESTIMATION OF MARKET VALUE IN ASSOCIATION FOOTBALL
#
# 2) 주성분 분석 (2개)
# - 승희배, 강기훈(2012). 한국 프로야구 선수들의 경기력과 연봉의 관계 분석. 한국데이터정보과학회지
# - 황선욱(2019). 주성분 기법을 활용한 NBA선수 포지션별 선수평가지수 및 연봉예측모델 개발. 동국대학교 대학원
#
#
# 2) 머신러닝 (4개)
# - 오광모, 이장택(2003). 데이터마이닝을 이용한 한국프로야구 선수들이 연봉에 관한 모형연구. 한국스포츠사회학회지,16(2)
# - <NAME>(2018), Player valuation in European football
# - <NAME>(2019). Predicting 2018–19 NBA’s Most Valuable Player using Machine Learning
# - <NAME>.(2017). Computational Estimation of Football Player Wages
#
# ```
#
#
# # 선행연구 검토에서 얻은 시사점
# ### 1. 선형회귀 관련
#
# 1) 독립변수 간 다중공선성 확인 필요. 승희배, 강기훈(2012)
#
# 2) 독립변수 selection 시, 방법론 고민 필요. Yaldo, L.(2017).
# : Relief Attribute Evaluation
#
# 3) 공격수, 수비수 간 경기력 측정 지표가 상이할 가능성 확인. 신문선(2003)
#
# 4) 독립변수 분리 네이밍 참고 : 경기력 요인 / 비경기력 요인. 이종우(2014)
#
# 5) 부분회귀 활용 : 주요 독립변수 영향력 확인. 신문선(2003)
#
# 6) 데이터의 정규성 검정 실시 필요 (선형회귀의 정규성 가정)
#
# 7) 성능 평가 방법 고민
# : 성능 평가 시, k-fold cross validation 대신, RMSE, MAE 활용. <NAME>(2017)
# : MAE, MSE, correlation coefficient, MRE
# ### 2. ML 관련
#
# 1) 독립변수 selection 시, 방법론 고민 필요. <NAME>(2018)
# : wrapper method, filter method
#
# 2) 성능 평가 방법 고민
# : (회귀) MAE, MSE, correlation coefficient, MRE / (ML) k-fold cross validation (accuracy, precision, recall, F1 score (or f-measure), and AUC-ROC)
#
# 3) ML 측면 데이터 분석 연구 흐름 : EDA -> 전처리 -> 모형 적용 -> 성능 평가 -> 결과 산출. 오광모, 이장택(2003)
#
# 4) ML 모델 구성을 CART, CART 후 NN, NN 후 CART, NN 등으로 다양하게 구성할 수 있음. 오광모, 이장택(2003)
# ### 3. 주성분 분석을 통한 예측모델 가능
#
# 1) 주성분 분석 -> 주요 설명 변수 추출 -> 선수 평가 모델 독립 변수에 활용. 황선욱(2019)
#
#
#
# ### 4. 기타
#
# 1) 농구, 야구와 달리 축구는 공신력있는 선수평가지수 및 통계량 자료(ABPR 메트릭스, 세이버 메트릭스 등)가 없음
#
# 2) 연구 방향 : 연봉 예측 / 기존 선수 평가 지수 모델 개발 가능할 듯
# ```
#
#
#
#
#
#
# ```
# # 세부 선행연구 목록
# # 1. topic관련 : 프로스포츠 연봉 관련 연구
#
#
# # 국내연구
# **1) 이종우(2014), 실업팀 선수의 연봉 결정요인 분석 : 개인종목을 중심으로. 서울대학교 대학원**
# - 사용 모델 : 정성적분석, 기술통계분석, 다중회귀분석(다변수 회귀분석)
# - 종속변수 : 연봉
# - 독립변수 : 경기력 요인 8개, 비경기력 요인 9개
#
# * 경기력요인 : 대회수준, 개인 경기력, 신기록 보유 여부, 선수 경기력 고려기간, 과거 경기력, 국가대표 여부, 국가대표 후보 여부, 신체 조건
#
# * 비경기력요인 : 종목 특성(다관왕, 종목 순위 배점), 타선수 연봉, 출신 지역, 지역내 스포츠 이벤트 개최 여부, 실업팀 연차, 실업팀 기여도, 선수 기간, 군필 여부, 감독 유형
#
# - 연구결과
# 1) 연봉과 비례(경기력요인) : 전국대회, 올림픽, 세계 선수권, 기타 국제대회, 한국 신기록, 전국가대표, 신체조건
# 2) 연봉과 비례(비경기력요인) : 실업선수 경력, 전국체전 순위 배점, 관리형 감독
#
# 2) 권병진(2005). 미국 프로야구 Major League 선수들의 노동생산성과 연봉에 관한 연구. 서강대학교 대학원
# 3) 송종우(2008). 한국 프로스포츠 선수들의 연봉에 대한 다변량적분석. 응용통계연구, 21(3), 441-45
# **4) 승희배, 강기훈(2012). 한국 프로야구 선수들의 경기력과 연봉의 관계 분석. 한국데이터정보과학회지, 23(2), 285-298.**
#
# - 사용 모델 : 사이버메트릭스 지수, 주성분 분석
#
# * 특이사항 : 독립변수 간 다중공선성 확인을 위해 "VIF분석, 고유벡터/값 분석, PCA"
#
# - 연구 결과 :
# 1) 사이버메트릭스 지수와 연봉 간 상관관계는 구단 간 차이가 있음
# 2) 사이버메트릭스 지수의 한계(수비력 반영 부족)를 미국의 방법론을 활용해 보완할 필요
# 3) 정량적 관리 방법의 우리나라의 도입 필요성 역설
# **5) 신문선(2003). 한국 프로축구 선수의 연봉산정 모델 개발. 국내박사학위논문, 세종대학교 대학원, 서울.**
#
# - 사용 모델 : 정성분석, 기초통계량분석, 상관관계 분석, 다중회귀 분석
#
# * 특이사항 : 다중공선성 처리(상관관계 분석 확인으로 마무리), 부분회귀 활용, 다중회귀모델링
#
# - 연구 결과 :
# 1) 통합 연봉 요인(9개) : 전년도 연봉, 팀성적, 구단규모, 경기출전수, 감독평가, 국가대표, 프로경력, 득점, 도움, 결정적 방어, 연령, 교체빈도, 장래성
# 2) 포지션 별 연봉 요인 차이를 둬 분석함(공격수 및 미드필더 / 수비수 / 골키퍼)
# **6) 오광모, 이장택(2003). 데이터마이닝을 이용한 한국프로야구 선수들이 연봉에 관한 모형연구. 한국스포츠사회학회지,16(2), 295-309.**
#
# - 사용 모델 : 데이터 마이닝(클레멘타인, answer tree), 예측 모형(비선형 : 신경망 분석, 의사결정 나무, 선형 : 회귀분석)
#
# * 특이사항 : 비선형 모형을 활용해 선형적으로 언급되지 않는 스타급 선수들의 연봉산정을 합리적으로 이해하고자 함
# * 특이사항 : 투수, 타자의 독립변수 항목 다르게 설정
#
# - 연구 결과 :
# 1) 타자연봉 예측 : 신경망 분석, 투수연봉 예측 : 의사결정나무 가 가장 예측 성능이 좋은 모델로 확인
# 2) 관중 동원 능력, 미래 잠재 능력 등 비경기력 요인들 확인 못한 한계점 존재
#
# **7) 한동섭, 김정기, 김종(2007). 스포츠 스타의 이미지와 소속 팀, 보증 제품 및 사회 이미지의 관계에 관한 연구. 한국스포츠산업.경영학회지, 12(3), 155-168.**
#
# - 사용 모델 : 다중회귀분석(유의미한 변수 추출)
#
# *특이사항 : 이미지 분석을 위해 100명의 학생 대상 이미지를 나타내는 "형용사" 단어 수집(총 4명의 선수(국내2명, 해외2명) 대상 조사)
#
# - 연구 결과 :
# 1) 스포츠 선수의 이미지가 사회, 국가 이미지에 미치는 영향력 유의미
# 8) 최명일(2002). 스포츠 스타 이미지 구성요인에 관한 연구. 국내석사학위논문, 한양대학교 대학원, 서울.
# **9) 손동만(2018). NBA 선수 연봉구조와 선수 스타파워가 팀 퍼포먼스에 미치는 영향. 서울대학교 대학원**
#
# * 특이사항 : 선수 스타성 지표 활용(올스타전 투표수)
# - 사용 모델 : 시계열 분석(GLS Regression with AR(1)
#
# - 연구 결과 :
# 1) 연봉 평균 비율 높을 수록 팀 성적 좋음
# 2) 연봉 격차는 팀 성적에 양적 영향
# 3) 스타파워 높을 수록 팀 성적 좋음
# **10) 황선욱(2019). 주성분 기법을 활용한 NBA선수 포지션별 선수평가지수 및 연봉예측모델 개발. 동국대학교 대학원**
#
# * 특이사항 : 주성분 분석을 통해 주요 요인 발굴 -> 선수 평가 지수 모델 개발
# - 사용 모델 : 주성분 분석, 회귀분석
# * 기존의 선수평가지수(NBA대상, PER)를 대체할 새로운 선수평가지수 개발 목적
# - 연구 결과 : 기존 PER 지수의 한계(출전시간에 반비례) 확인. 포지션별 선수 평가 지수 모델 제안.
# # 해외연구
# **1) <NAME>(2018). A Multiple Linear Regression Approach For Estimating the Market Value of Football Players in Forward Position**
#
# - 연구 내용 : 17-18시즌 데이터, 공격수 대상, 시장가치 추정
#
# - 사용 모델 : 다중 선형회귀 (step-wise regression)
# *step-wise regression 참고 : https://talkingaboutme.tistory.com/entry/ML-Stepwise-Regression
#
# - 연구 결과 :
# 1) 리그가 시장가치에 미치는 영향 적음(그런데, 아마 축구팀- 리그 사이 다중공선성이 있어 확인해봐야 함)
# 2) 신장, 주 발(오른쪽, 왼쪽), 국적, 득점 등이 시장 가치에 유효
# **[선택] 2) <NAME>(2018), Player valuation in European football**
#
# - 연구 내용 : 성과 좋은 선수들의 스킬과 특징 확인, 성과 예측
#
# * 특이사항 : 성과 예측 모델의 feature selection을 위해 'filter method', 'wrapper method'를 활용
# * ML의 feature selection
# * wrapper method : forward selection, backward selection, step-wise selection
# https://m.blog.naver.com/PostView.nhn?blogId=euleekwon&logNo=221465108279&proxyReferer=https:%2F%2Fwww.google.com%2F
#
# - 사용 모델 : 10개의 ML 알고리즘
# (RandomForest, BayesNet, Logistic, DecisionTable (a decision table majority classifier), IBk, KStar (nearest neighbor with generalized distance), NaiveBayes, J48, Part, and ZeroR (predicts the majority class for nominals or the average value for numerics)
#
# 1) 성능 측정 performace : 각 알고리즘을 10번 시행 후 평균 performance 활용
# 2) 성능 측정 지표 : accuracy, precision, recall, F1 score (or f-measure), and AUC-ROC.
# 3) feature selection : filter method, wrapper method를 통해 각각의 데이터셋 1개씩 생성
#
# - 연구 결과 :
# 1) 개별 선수들의 성공에 영향을 미치는 특징, 스킬셋 확인
# 2) 다른 연구와 달리, top player(10%, 25%, 50%) 집중 연구
# 3) 10개 ML 알고리즘의 예측 성능 비교 분석
# 4) 예측 성능이 포지션 별, 탑 티어별 차등화(f1 score)
# * 공격수, 탑 티어일수록 성공 요인이 더 두드러지게 관측됨
# 5) basic ML 기법 : EPL, 분데스리가 선수 예측에 더 높은 성능(basic ML : Zero R classifier)
# **3) <NAME>(2019). Predicting 2018–19 NBA’s Most Valuable Player using Machine Learning**
# - 컬럼 기고문
# - url : https://towardsdatascience.com/predicting-2018-19-nbas-most-valuable-player-using-machine-learning-512e577032e3
#
# - 연구 내용
# 1) 18-19 시즌, "predicting most valualbe player" (MVP award 예측, "누가 받을까?")
# 2) 문제 정의 : 랭킹화 문제. 회귀 문제로 접근해 개별 선수의 target-value(share) 예측
#
# - 사용 모델 : Linear Regression, Ridge Regression, Gradient Boosting Regressor, Random Forest Regressor, SVR
#
# - 연구 결과 :
# 1) 평균적인 score 를 측정 -> MVP 선수 예측
# **4) <NAME>(2018). Can Linear Models predict a Footballer’s Value?**
#
# - 컬럼 기고문
# - url : https://towardsdatascience.com/can-linear-models-predict-a-footballers-value-33d772211e5d
#
# - 연구 내용
# 1) relationship between a player’s popularity and his market value in the English Premier League
#
# - 사용 모델 : Linear Regression
# *주의사항 : 위키피디아 페이지 뷰 수 -> ability 지표로 활용(상세히 기술)
# **[선택] 5) Yaldo, L.(2017). Computational Estimation of Football Player Wages**
#
# - 연구 내용 : "an objective quantitative method to determine football players’ wages based on their skills"
# * 대상 : 6,082명, 2016시즌, 20개 이상의 리그(세리에A, 세리에B, ...)
# * 리그별 분석, 저평가 선수 분석 추가로 진행
#
#
# - 사용 모델 : 7개 머신러닝 알고리즘
# Additive Regression (Friedman, 2002),
# Decision Table (Kohavi, 1995),
# Nearest Neighbor with a weighted condition (Aha et al., 1991),
# K* (Cleary et al., 1995),
# Locally Weighted Learning with Naive Bayes and Linear regression classifiers (Frank et al., 2002)
# Random Committee (Seung et al., 1992),
# Random Trees (Aldous, 1993)
#
# *주의사항 : 변수 선택에 "Relief Attribute Evaluation" 방법 사용됨
# *MAE, MSE, correlation coefficient, MRE 로 성능평가('leave-one-out test strategy' 활용)
# *참고 : https://partrita.github.io/posts/regression-error/ (회귀 오류지표 종합 설명 블로그 글)
#
# - 연구 결과 :
# 1) 비경기적 요인이 wage에 포함될 것으로 예상됨
# 2) 성능 평가 방법에 따라 각 모델 별 성능 평가 순위가 달라짐
# **[선택] 6) <NAME>(2017). BEYOND CROWD JUDGMENTS: DATA-DRIVEN ESTIMATION OF MARKET VALUE IN ASSOCIATION FOOTBALL**
#
# - 연구 내용 : "estimate players’ market values using multilevel regression analysis"
# "data-driven estimation of market value"
# * 대상 : 4,217명, 6 시즌 간(09-14), 상위 5개 리그
#
# - 사용 모델 : 회귀분석
#
# *주의사항 : market value의 test 시, 즉, actual data로 "actual transfer fees"를 market value의 대체자료로 보고 활용
#
# * (actual transfer fee - model) vs (actual transfer fee - crowd source[트랜스퍼마켓])
# *model의 train은 crowd source데이터 활용.
#
# *주의사항 : 성능 평가 시, k-fold cross validation 대신, RMSE, MAE 활용
# (time-series- based evaluation approach 고민해야 할 듯)
#
# *주의사항 : 회귀 모델의 fitness 보기 위해, AIC, BIC, log-likelihood 값 등 확인
#
# *dataset : wage자료는 트랜스퍼 마켓 데이터(크라우드 소싱 데이터)로 train
#
# *주의사항 : 비 경기적 요소 데이터 포함 (위키 페이지 뷰 수, 구글 이름 검색 수, 레딧 포럼 페이지 언급 수, 유튜브 동영상 수)
# "We used four Internet metrics to measure player popularity: the number of times a player’s Wikipedia page was viewed, how often a player’s name was searched on Google, the number of times a player’s name appeared in the ―soccer forum on Reddit, and how many videos about a player were shared on YouTube. The average player had more than 100,000 Wikipe- dia page views and more than 35,000 YouTube videos."
#
# - 연구 결과 :
# 1) crowd 예측이 어느정도 유의미함. 선수의 90%(하위 이적료)는 model의 예측이 더 fit하지만, 상위 이적료 10%는 crowd-estimate가 더 fit함을 보임
# 2) 비 경기적 요소 중 위키, 유튜브, 레딧 이 유의미한 상관관계 보임
#
#
# ```
#
#
#
#
#
# ```
# # 2. topic 외 : 사회 이미지 및 국가 이미지 척도 인용
#
# # 국내연구
# 1) 김설애(2016). 원산지국가이미지가 소비자인식 및 구매의도에 미치는 영향
#
# - 원산지 국가 이미지과 구매의도 간 상관관계에 대한 연구
# - 연구 방법 : 설문 조사
# 2)문효진(2016). 국가브랜드 평판 구성요인 도출을 위한 탐색적 연구
#
# - 국가 브랜드 평판에 영향을 미치는 요인 탐구
# - 연구 방법 : 설문 조사
# # 해외연구
# 1) <NAME>. & <NAME>. (1993). Measuring a Multi-Dimensional Construct: Country Image
# - 국가 이미지에 영향을 주는 요소들의 항목 연구
# **2) <NAME>(2007). 국가역량 종합지수(Composite Index of National Capability, CNIC)**
# - 국가의 Hard Power 측정 지표
#
# - 결과 url :
# https://ko.wikipedia.org/wiki/%EA%B5%AD%EA%B0%80_%EC%97%AD%EB%9F%89_%EC%A2%85%ED%95%A9_%EC%A7%80%EC%88%98
#
# 3) monocle(2018). 국가별 소프트 파워 지수
#
# - 국가의 soft power 측정 지표
#
# - 결과 url :
# https://ko.wikipedia.org/wiki/%EA%B5%AD%EA%B0%80_%EB%B8%8C%EB%9E%9C%EB%93%9C#cite_note-5
#
# - 구체적인 결과를 보려면, monocle 잡지를 유료 구독해야한다는 한계점
# - 공신력 측면에서 한계 예상
# **4) US News(2020). US News 최고의 나라 지수**
#
# - 80개국 대상 순위 (2016 - 2020, 5개년 자료)
# - 결과 url : https://www.usnews.com/news/best-countries/overall-rankings
# - metric url : https://www.usnews.com/news/best-countries/articles/methodology
#
# ```
#
#
#
#
#
# ```
# # 3. topic 외 : SNS 데이터 분석 관련
#
# 1) 강지원(2020). Why do Instagram Users Tag Friends in Comments
#
# - instalooter : 비디오, 사진 긁어올 수 있는 루트
# 참고 url : https://blog.naver.com/moongs2chi/221612087877
# 참고 url : https://instalooter.readthedocs.io/en/latest/
#
# - selenium - webdriver 등
| 0.Project/2. Regression Anlaysis/2. Related precedent study/1. Related precedent study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
bba=[]
bcom=[]
psycho=[]
name=str(input("Enter your Name"))
batch=int(input("Enter Batch Code: BBA-1,BCom-2,Psychology-3"))
if batch==1:
a=bba.append(name)
print("BBA Students",bba)
elif batch==2:
a=bcom.append(name)
print("BCom Students",bcom)
elif batch==3:
a=psycho.append(name)
print("Psychology",psycho)
else:
print("Invalid Batch Code")
# -
.
| For Loop for Form-Database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Linear Regression
#
# It is one of the most basic regression techniques in which we want to model the relationship between a dependent variable y, and one or more independent variables X
#
# The equation can be shown as below - <br>
# #### y = w.X + b,
# where w is the weight of the independent parameter X, if x is a d-dimensional vector, then w will also be a d-dimensional vector. Weight is responsible for the steep-ness of the regression line.<br>
# b represents bias, which can thought of to be responsible for the shift in the modelled regression line. It is important to have a bias because weight cannot accomodate the shift. In simple terms, it is the intercept of the regression line.
#
# We will be training the linear regression through gradient descent. One can solve for the parameters using closed-form solution (normal solution) of linear regressions too, which is given by:
# #### w = (X<sup>T</sup>X)<sup>−1</sup>X<sup>T</sup>y
# but, closed-form solutions have some disadvantages - <br>
# 1. Non-linear functions do not usually have closed-form solutions
# 2. Closed-form solutions requires computing the inverse. It gets [too slow](https://stats.stackexchange.com/questions/23128/solving-for-regression-parameters-in-closed-form-vs-gradient-descent), when there are a lot of features
#
# ### Training Steps
# 1. Initialize weight vector and bias with zero values (or very small numbers)
# 2. Calculate $\boldsymbol{\hat{y}} = \boldsymbol{X} \cdot \boldsymbol{w} + b $
# 3. Compute the cost $J(\boldsymbol{w},b) = \frac{1}{m} \sum_{i=1}^m \Big(\hat{y}^{(i)} - y^{(i)} \Big)^2$
# 4. Compute partial derivatives of all the parameters <br>
# $ \frac{\partial J}{\partial w_j} = \frac{2}{m}\sum_{i=1}^m \Big( \hat{y}^{(i)} - y^{(i)} \Big) x^{(i)}_j$ <br>
# $ \frac{\partial J}{\partial b} = \frac{2}{m}\sum_{i=1}^m \Big( \hat{y}^{(i)} - y^{(i)} \Big)$ <br>
# The gradient can then be computed as follows: <br>
# $\nabla_{\boldsymbol{w}} J = \frac{2}{m} \boldsymbol{X}^T \cdot \big(\boldsymbol{\hat{y}} - \boldsymbol{y} \big)$ <br>
# $\nabla_{\boldsymbol{b}} J = \frac{2}{m} \big(\boldsymbol{\hat{y}} - \boldsymbol{y} \big)$
#
# 5. Update the parameters using <br>
# $w = w - \alpha \, \nabla_{\boldsymbol{w}} J$ <br>
# $b = b - \alpha \, \nabla_{\boldsymbol{b}} J$
#
# where, $\alpha$ is the learning rate
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
np.random.seed(2018)
# -
# ### Data
# +
X = 2*np.random.rand(700, 1)
y = 5 + 3*X + np.random.rand(700, 1)
fig = plt.figure(figsize=(8,6))
plt.scatter(X, y)
plt.title("Dataset")
plt.xlabel("First parameter")
plt.ylabel("Second parameter")
plt.show()
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
print("Shape of X_train is {}".format(X_train.shape))
print("Shape of X_test is {}".format(X_test.shape))
print("Shape of y_train is {}".format(y_train.shape))
print("Shape of y_test is {}".format(y_test.shape))
# -
# ### Model
# +
class LinearRegression:
def __init__(self):
pass
def train_gradient_descent(self, X, y, alpha=0.001, iterations=100):
# Step 1: Initialize the parameters
n_samples, n_features = X.shape
self.w = np.zeros(shape=(n_features, 1))
self.b = 0
J = []
for i in range(iterations):
# Step 2: Calculate y_predicted
y_hat = np.dot(X, self.w) + self.b
# Step 3: Compute the cost
cost = (1/n_samples)*np.sum((y_hat-y)**2)
J.append(cost)
if i % 100 == 0:
print("Cost at iteration {} is: {}".format(i, cost))
# Step 4: Compute partial derivatives
dJ_dw = (2/n_samples)*np.dot(X.T, (y_hat-y))
dJ_db = (2/n_samples)*np.sum((y_hat-y))
# Step 5: Update the parameters
self.w = self.w - alpha*dJ_dw
self.b = self.b - alpha*dJ_db
return self.w, self.b, J
def normal_solution(self, X, y):
self.w = np.dot(np.dot(np.linalg.inv(np.dot(X.T, X)), X.T), y)
self.b = 0
return self.w, self.b
def predict(self, X):
return (np.dot(X, self.w)) + self.b
# +
linear_regressor = LinearRegression()
w_trained, b_trained, J = linear_regressor.train_gradient_descent(X_train, y_train, alpha=0.005, iterations=600)
fig = plt.figure(figsize=(8,6))
plt.plot(np.arange(600), J)
plt.title("Gradient Descent")
plt.xlabel("Iterations")
plt.ylabel("Cost (J)")
plt.show()
# +
y_hat_train = linear_regressor.predict(X_train)
y_hat_test = linear_regressor.predict(X_test)
error_train = (1/X_train.shape[0])*np.sum((y_hat_train-y_train)**2)
error_test = (1/X_test.shape[0])*np.sum((y_hat_test-y_test)**2)
print("Error on training set: {}".format(np.round(error_train)))
print("Error on testing set: {}".format(np.round(error_test)))
# -
fig = plt.figure(figsize=(8,6))
plt.title("Dataset in blue, predictions for test set in orange")
plt.scatter(X_train, y_train)
plt.scatter(X_test, y_hat_test)
plt.xlabel("First feature")
plt.ylabel("Second feature")
plt.show()
| linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# hide
# %load_ext autoreload
# %autoreload 2
# %load_ext nb_black
# %load_ext lab_black
# +
# default_exp model_pipeline
# -
# # ModelPipeline
# > Putting it all together.
# ## Overview
#
# The functionality below uses the `NumerFrame`, `PreProcessor`, `Model` and `PostProcessor` objects to easily propagate
# data, generate predictions and postprocess them in one go.
#
# Specifically, this section introduces two objects:
# 1. `ModelPipeline`: Run all preprocessing, models and postprocessing that you define and return a `NumerFrame`.
# 2. `ModelPipelineCollection`: Manage and run multiple `ModelPipeline` objects.
# hide
from nbdev.showdoc import *
# +
#export
import uuid
import pandas as pd
from tqdm.auto import tqdm
from typeguard import typechecked
from typing import List, Union, Dict
from rich import print as rich_print
from numerblox.numerframe import NumerFrame, create_numerframe
from numerblox.preprocessing import BaseProcessor, CopyPreProcessor, GroupStatsPreProcessor, FeatureSelectionPreProcessor
from numerblox.model import BaseModel, ConstantModel, RandomModel
from numerblox.postprocessing import Standardizer, MeanEnsembler, FeatureNeutralizer
# -
# ## 1. ModelPipeline
# `ModelPipeline` handles all preprocessing, model prediction and postprocessing. It returns a `NumerFrame` with the preprocessed data, metadata and postprocessed prediction columns.
#export
@typechecked
class ModelPipeline:
"""
Execute all preprocessing, prediction and postprocessing for a given setup.
:param models: Initiliazed numerai-blocks Models (Objects inheriting from BaseModel) \n
:param preprocessors: List of initialized Preprocessors. \n
:param postprocessors: List of initialized Postprocessors. \n
:param copy_first: Whether to copy the NumerFrame as a first preprocessing step. \n
Highly recommended in order to avoid surprise behaviour by manipulating the original dataset. \n
:param pipeline_name: Unique name for pipeline. Only used for display purposes.
"""
def __init__(self,
models: List[BaseModel],
preprocessors: List[BaseProcessor] = [],
postprocessors: List[BaseProcessor] = [],
copy_first = True,
standardize = True,
pipeline_name: str = None):
self.pipeline_name = pipeline_name if pipeline_name else uuid.uuid4().hex
self.models = models
self.copy_first = copy_first
self.standardize = standardize
self.preprocessors = preprocessors
self.postprocessors = postprocessors
def preprocess(self, dataf: Union[pd.DataFrame, NumerFrame]) -> NumerFrame:
""" Run all preprocessing steps. Copies input by default. """
if self.copy_first:
dataf = CopyPreProcessor()(dataf)
for preprocessor in tqdm(self.preprocessors,
desc=f"{self.pipeline_name} Preprocessing:",
position=0):
rich_print(f":construction: Applying preprocessing: '[bold]{preprocessor.__class__.__name__}[/bold]' :construction:")
dataf = preprocessor(dataf)
return NumerFrame(dataf)
def postprocess(self, dataf: Union[pd.DataFrame, NumerFrame]) -> NumerFrame:
""" Run all postprocessing steps. Standardizes model prediction by default. """
if self.standardize:
dataf = Standardizer()(dataf)
for postprocessor in tqdm(self.postprocessors,
desc=f"{self.pipeline_name} Postprocessing: ",
position=0):
rich_print(f":construction: Applying postprocessing: '[bold]{postprocessor.__class__.__name__}[/bold]' :construction:")
dataf = postprocessor(dataf)
return NumerFrame(dataf)
def process_models(self, dataf: Union[pd.DataFrame, NumerFrame]) -> NumerFrame:
""" Run all models. """
for model in tqdm(self.models,
desc=f"{self.pipeline_name} Model prediction: ",
position=0):
rich_print(f":robot: Generating model predictions with '[bold]{model.__class__.__name__}[/bold]'. :robot:")
dataf = model(dataf)
return NumerFrame(dataf)
def pipeline(self, dataf: Union[pd.DataFrame, NumerFrame]) -> NumerFrame:
""" Process full pipeline and return resulting NumerFrame. """
preprocessed_dataf = self.preprocess(dataf)
prediction_dataf = self.process_models(preprocessed_dataf)
processed_prediction_dataf = self.postprocess(prediction_dataf)
rich_print(f":checkered_flag: [green]Finished pipeline:[green] [bold blue]'{self.pipeline_name}'[bold blue]! :checkered_flag:")
return processed_prediction_dataf
def __call__(self, dataf: Union[pd.DataFrame, NumerFrame]) -> NumerFrame:
return self.pipeline(dataf)
# Example using several preprocessor, dummy models and postprocessors
# +
model_names = ["test_0.5", "test_0.8"]
dataf = create_numerframe("test_assets/mini_numerai_version_1_data.csv", metadata={'version': 1})
preprocessors = [GroupStatsPreProcessor(), FeatureSelectionPreProcessor(feature_cols=['feature_intelligence_mean', 'feature_intelligence_std'])]
models = [ConstantModel(constant=0.5, model_name=model_names[0]), ConstantModel(constant=0.8, model_name=model_names[1])]
postprocessors = [MeanEnsembler(cols=[f"prediction_{name}" for name in model_names], final_col_name='prediction_ensembled'),
FeatureNeutralizer(feature_names=['feature_intelligence_mean', 'feature_intelligence_std'],
pred_name='prediction_ensembled', proportion=0.8)]
# -
test_pipeline = ModelPipeline(preprocessors=preprocessors, models=models,
postprocessors=postprocessors, pipeline_name="test_pipeline",
standardize=False)
processed_dataf = test_pipeline(dataf)
assert processed_dataf.meta == dataf.meta
assert isinstance(processed_dataf, NumerFrame)
processed_dataf.head(2)
# ## 2. ModelPipelineCollection
# `ModelPipelineCollection` can be used to manage and run multiple `ModelPipeline` objects.
#
# `ModelPipelineCollection` simply takes a list of `ModelPipeline` objects as input.
#export
@typechecked
class ModelPipelineCollection:
"""
Execute multiple initialized ModelPipelines in a sequence.
:param pipelines: List of initialized ModelPipelines.
"""
def __init__(self, pipelines: List[ModelPipeline]):
self.pipelines = {pipe.pipeline_name: pipe for pipe in pipelines}
self.pipeline_names = list(self.pipelines.keys())
def process_all_pipelines(self, dataf: Union[pd.DataFrame, NumerFrame]) -> Dict[str, NumerFrame]:
""" Process all pipelines and return Dictionary mapping pipeline names to resulting NumerFrames. """
result_datafs = dict()
for name, pipeline in tqdm(self.pipelines.items(),
desc="Processing Pipeline Collection"):
result_datafs[name] = self.process_single_pipeline(dataf, name)
return result_datafs
def process_single_pipeline(self, dataf: Union[pd.DataFrame, NumerFrame], pipeline_name: str) -> NumerFrame:
""" Run full model pipeline for given name in collection. """
rich_print(f":construction_worker: [bold green]Processing model pipeline:[/bold green] '{pipeline_name}' :construction_worker:")
pipeline = self.get_pipeline(pipeline_name)
dataf = pipeline(dataf)
return NumerFrame(dataf)
def get_pipeline(self, pipeline_name: str) -> ModelPipeline:
""" Retrieve model pipeline for given name. """
available_pipelines = self.pipeline_names
assert pipeline_name in available_pipelines, f"Requested pipeline '{pipeline_name}', but only the following models are in the collection: '{available_pipelines}'."
return self.pipelines[pipeline_name]
def __call__(self, dataf: Union[pd.DataFrame, NumerFrame]) -> Dict[str, NumerFrame]:
return self.process_all_pipelines(dataf=dataf)
# We introduce a different pipeline with no preprocessing or postprocessing. Only a `RandomModel`.
test_pipeline2 = ModelPipeline(models=[RandomModel()], pipeline_name="test_pipeline2")
# + [markdown] pycharm={"name": "#%% md\n"}
# We process two `ModelPipeline`s with different characteristics on the same data.
# -
collection = ModelPipelineCollection([test_pipeline, test_pipeline2])
assert collection.get_pipeline("test_pipeline2").pipeline_name == 'test_pipeline2'
result_datasets = collection(dataf=dataf)
# The `ModelPipelineCollection` returns a dictionary mapping pipeline names to `NumerFrame` objects, retaining all metadata and added prediction columns for each. Note that in this example, the 1st `NumerFrame` had a feature selection step, so it did not retain all columns. However, the second dataset retained all feature columns, because no preprocessing was done.
# + pycharm={"name": "#%%\n"}
result_datasets.keys()
# -
result_datasets['test_pipeline'].head(2)
# + pycharm={"name": "#%%\n"}
result_datasets['test_pipeline2'].head(2)
# -
# Since metadata is not manipulated in these pipelines, metadata should be the same as the original `NumerFrame` for all resulting `NumerFrame` objects.
# + pycharm={"name": "#%%\n"}
for _, result in result_datasets.items():
assert dataf.meta == result.meta
# + pycharm={"name": "#%%\n"}
result_datasets['test_pipeline'].meta
# -
# -----------------------------------------------------------------------------
# +
# hide
# Run this cell to sync all changes with library
from nbdev.export import notebook2script
notebook2script()
# + pycharm={"name": "#%%\n"}
| nbs/06_modelpipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dlnd
# language: python
# name: dlnd
# ---
# # TensorFlow 2.0
# +
import os
from glob import glob
from datetime import datetime
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras import datasets
import matplotlib.pyplot as plt
# %load_ext tensorboard
# %matplotlib inline
# -
# ## Hyperparameter Tunning
# +
num_epochs = 5
batch_size = 32
learning_rate = 0.001
dropout_rate = 0.5
input_shape = (32, 32, 3)
num_classes = 10
# -
# ## Build Model
# +
inputs = layers.Input(input_shape)
net = layers.Conv2D(32, (3, 3), padding='SAME')(inputs)
net = layers.Activation('relu')(net)
net = layers.Conv2D(32, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.MaxPooling2D(pool_size=(2, 2))(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Conv2D(64, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.Conv2D(64, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.MaxPooling2D(pool_size=(2, 2))(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Flatten()(net)
net = layers.Dense(512)(net)
net = layers.Activation('relu')(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Dense(num_classes)(net)
net = layers.Activation('softmax')(net)
model = tf.keras.Model(inputs=inputs, outputs=net, name='Basic_CNN')
# -
# Model is the full model w/o custom layers
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate), # Optimization
loss='sparse_categorical_crossentropy', # Loss Function
metrics=['accuracy']) # Metrics / Accuracy
# # Data Preprocess
train_paths = glob('../dataset/cifar/train/*.png')[:100]
test_paths = glob('../dataset/cifar/test/*.png')[:100]
def get_class_name(path):
return path.split('_')[-1].replace('.png', '')
train_labels = [get_class_name(path) for path in train_paths]
class_names = np.unique(train_labels)
def get_label(path):
fname = tf.strings.split(path, '_')[-1]
lbl_name = tf.strings.regex_replace(fname, '.png', '')
onehot = tf.cast(lbl_name == class_names, tf.uint8)
return tf.argmax(onehot) # 이번에는 onehot이 아닌 label 번호로
def load_image_label(path):
gfile = tf.io.read_file(path)
image = tf.io.decode_image(gfile)
image = tf.cast(image, tf.float32) / 255. # rescale
label = get_label(path)
return image, label
def image_preprocess(image, label):
image = tf.image.random_flip_up_down(image)
image = tf.image.random_flip_left_right(image)
return image, label
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_dataset = tf.data.Dataset.from_tensor_slices(train_paths)
train_dataset = train_dataset.map(load_image_label, num_parallel_calls=AUTOTUNE)
train_dataset = train_dataset.map(image_preprocess, num_parallel_calls=AUTOTUNE)
train_dataset = train_dataset.batch(batch_size)
train_dataset = train_dataset.shuffle(buffer_size=len(train_paths))
train_dataset = train_dataset.repeat()
test_dataset = tf.data.Dataset.from_tensor_slices(test_paths)
test_dataset = test_dataset.map(load_image_label, num_parallel_calls=AUTOTUNE)
test_dataset = test_dataset.batch(batch_size)
test_dataset = test_dataset.repeat()
# # Checkpoint
save_path = 'ckpt'
checkpoint = tf.keras.callbacks.ModelCheckpoint(save_path, monitor='val_accuracy', verbose=1, save_best_only=True, mode='max')
# ## Training
# http://localhost:6006
# +
steps_per_epoch = len(train_paths) // batch_size
validation_steps = len(test_paths) // batch_size
history = model.fit_generator(
train_dataset,
steps_per_epoch=steps_per_epoch,
validation_data=test_dataset,
validation_steps=validation_steps,
epochs=num_epochs,
callbacks=[checkpoint]
)
# -
# # Saving Model
save_path = 'my_model.h5'
model.save(save_path, include_optimizer=True)
model = tf.keras.models.load_model('my_model.h5')
# # Saving Model - 2
# +
# Save the weights
model.save_weights('model_weights.h5')
# Save the model architecture
with open('model_architecture.json', 'w') as f:
f.write(model.to_json())
# +
from tensorflow.keras.models import model_from_json
# Model reconstruction from JSON file
with open('model_architecture.json', 'r') as f:
model = model_from_json(f.read())
# Load weights into the new model
model.load_weights('model_weights.h5')
# -
# # model.h5 들여다보기
# +
import h5py
model_file = h5py.File('my_model.h5','r+')
# -
model_file.keys()
model_file['model_weights'].keys()
model_file['model_weights']['conv2d']['conv2d'].keys()
model_file['model_weights']['conv2d']['conv2d']['kernel:0']
np.array(model_file['model_weights']['conv2d']['conv2d']['kernel:0'])
weight = np.array(model_file['model_weights']['conv2d']['conv2d']['kernel:0'])
| DL_TF20/Part 21 - save and load model - h5-Antonio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## `ipydatagrid`
# ### HyperlinkRenderer
#
# A renderer which allows for rendering text-based clickable links as cells in your DataGrid. It takes two parameters:
#
# 1. `url`: A Vega Expression function which points to a full URL (with the http(s) prefix)
# 2. `urlName`: A Vega Expression function which points to a friendly URL display name
#
# **To preserve default cell selections behaviour, a link can only be opened by clicking whilst holding the `Ctrl` or `Command` key pressed.**
#
# The `HyperlinkRenderer` can be styled using all properties supported by the `TextRenderer`!
# +
import pandas as pd
import numpy as np
from ipydatagrid import VegaExpr, DataGrid, TextRenderer, HyperlinkRenderer
df = pd.DataFrame(
data={
"Name": ["Tech at Bloomberg"],
"Link": [["https://www.techatbloomberg.com/", "Tech at BBG Website"]],
}
)
link_renderer = HyperlinkRenderer(
url=VegaExpr("cell.value[0]"),
url_name=VegaExpr("cell.value[1]"),
background_color="moccasin",
text_color="blue",
font="bold 14px Arial, sans-serif",
)
grid = DataGrid(
df,
layout={"height": "120px"},
base_column_size=200,
renderers={"Link": link_renderer},
)
grid
| examples/HyperlinkRenderer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install st_annotated_text
# +
import spacy
from spacy import displacy
nlp = spacy.load("en_core_web_sm")
# -
import pandas as pd
import re
t = "Google News is a news aggregator service developed by Google. It presents a continuous flow of articles organized from thousands of publishers and magazines. Google News is available as an app on Android, iOS, and the Web. Google released a beta version in September 2002 and the official app in January 2006."
nlp_doc = nlp(t)
# +
nlp = spacy.load("en_core_web_sm")
privacy_type_mapping_filename = 'privacy_type_mapping.csv'
privacy_type_mapping = pd.read_csv(
privacy_type_mapping_filename,
index_col=0,
keep_default_na=False,
converters={"Requirements": lambda x: x.split("\n") if x else None},
).to_dict('index')
def extract_email(text):
return re.findall('[A-Za-z0-9]+[A-Za-z0-9._%+-]*@\w+.\w{2,4}', text)
def extract_phone(text):
return re.findall('(\d{3}[-\.\s]??\d{3}[-\.\s]??\d{4}|\(\d{3}\)\s*\d{3}[-\.\s]??\d{4}|\d{3}[-\.\s]??\d{4})', text)
def extract_chd(text):
return re.findall('[0-9]{16}', text)
# +
data_matchings = []
for entity in nlp_doc.ents:
if entity.label_ in privacy_type_mapping and privacy_type_mapping[entity.label_]['Requirements']:
data_matching_object = {
'type': entity.label_,
'value': entity.text,
'requirements': privacy_type_mapping[entity.label_]['Requirements'],
}
data_matchings.append(data_matching_object)
print(data_matching_object)
for extracted in extract_email(nlp_doc.text) + extract_phone(nlp_doc.text):
data_matching_object = {
'type': 'CONTACT',
'value': extracted,
'requirements': ['GLBA', 'CCPA', 'PIPEDA'],
}
data_matchings.append(data_matching_object)
print(data_matching_object)
for extracted in extract_chd(nlp_doc.text):
data_matching_object = {
'type': 'CHD',
'value': extracted,
'requirements': ['PCI'],
}
data_matchings.append(data_matching_object)
print(data_matching_object)
data_result = {
'match': bool(data_matchings),
'matchings': data_matchings,
}
# -
def allindices(string, sub, offset=0):
listindex=[]
i = string.find(sub, offset)
while i >= 0:
listindex.append(i)
i = string.find(sub, i + 1)
res = [(l, l+len(sub)) for l in listindex]
return res
allindices(t, "Android")
for m in data_matchings:
m['offsets'] = allindices(t, m['value'])
data_matchings
def include_all(l):
return "(" + ", ".join(l) + ")"
offset2data = {}
all_offsets = []
for m in data_matchings:
all_offsets += m['offsets']
for o in m['offsets']:
offset2data[o] = m
all_offsets.sort()
all_offsets
starting_index = 0
seg = []
for idx, off in enumerate(all_offsets):
if idx == 0:
seg.append(t[0:off[0]])
data = offset2data[off]
seg.append((data['value'], data['type'] + "|" + include_all(data['requirements']), "#8ef"))
seg.append(t[off[1]:])
seg
# +
import streamlit as st
from st_annotated_text import annotated_text
"""
# Annotated text example
Below is an example of how to use the annotated_text function:
"""
annotated_text(
"This ",
("is", "verb", "#8ef"),
" some ",
("annotated", "adj", "#faa"),
("text", "noun", "#afa"),
" for those of ",
("you", "pronoun", "#fea"),
" who ",
("like", "verb", "#8ef"),
" this sort of ",
("thing", "noun", "#afa"),
)
# -
pip install st_annotated_text
import matplotlib
| experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom Architecture Research
#
# > Testing variants of an idea to have multiple bodies in a neural net
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Computer Vision, Model Architecture]
# This post will cover an architecture modification for training models to give several model bodys feeding into a single custom head. We will be exploring this in the pets dataset and we see there are some improvements to be had over resnets in this instance. The goal will be to **predict pet breed**.
#
# >Credit: [<NAME>](https://github.com/jdavidberger/) read a previous blog post and had some great ideas for better baselines and things to try. This post and several of the experiments were largely inspired by our conversation on the fastai discord.
#
# Lets get started!
# +
from fastai.vision.all import *
import pickle
seed = 42
path = untar_data(URLs.PETS)
img = (path/'images').ls()[0]
# -
# # The Data
#
# We will start with creating a model for pet breeds and a model for pet species, so we will need dataloaders for each of those models. I will not cover how this works, but if you would like more details I recommend looking at the fastai datablock tutorial on their documentation page.
#
# We are going to be using **really small images (64x64)** to make this problem hard enough and fast enough to experiment with.
#collapse-hide
def label_func(fname):
return "cat" if str(fname.name[0]).isupper() else "dog"
def get_dls(get_y):
pets = DataBlock(
blocks = (ImageBlock, CategoryBlock),
get_items = get_image_files,
splitter= RandomSplitter(valid_pct = 0.2, seed=seed),
get_y= get_y,
item_tfms=Resize(128),
batch_tfms=aug_transforms(min_scale = 0.9,size=64)
)
return pets.dataloaders(path/"images",bs=64)
dls_breed = get_dls(using_attr(RegexLabeller(r'(.+)_\d+.jpg$'),'name'))
dls_breed.show_batch(max_n=3)
dls_species = get_dls(label_func)
dls_species.show_batch(max_n=3)
# It's crucial that these dataloaders show the same images in their training and test sets. If they don't, we would likely have a data leakage issue where each model has some of the others validation set in them. This would be really problematic as we will be combining these models together at the end.
#
# Lets add some quick tests to make sure that the valid imgs and the train images are the same.
breed_val = pd.DataFrame(dls_breed.valid.items,columns=['breed_imgs']).sort_values('breed_imgs')
species_val = pd.DataFrame(dls_species.valid.items,columns=['species_imgs']).sort_values('species_imgs')
assert (breed_val.breed_imgs==species_val.species_imgs).all()
breed_train = pd.DataFrame(dls_breed.train.items,columns=['breed_imgs']).sort_values('breed_imgs')
species_train = pd.DataFrame(dls_species.train.items,columns=['species_imgs']).sort_values('species_imgs')
assert (breed_train.breed_imgs==species_train.species_imgs).all()
# # The Architecture
#
# Now that we have our data let's talk about what we are going to be testing.
#
# Put simply: We are creating **resnet architectures** with **more than 1 body**. We are going to do several variations so we want a flexible wayt to create this. Let's look at our model class.
#
# I will explain what is going on in it below, as this is central to the whole expirament.
# + **body_list:** Body list takes a list of encoders (encoder_list) and cuts the heads off. So now we have a list of bodys using whatever encoders and wieghts were passed, and can do an arbitrary number of bodys using the same class.
# + **self.head:** This cretes a head that can take all the bodies and combine them. You will wee that the inputs to the head is dependent on the number of resnet bodies we have.
# + **self.split:** This just breaks out each body + head into their own parameter groups. This is important so it is easy to freeze the head that is random weights to train that first.
# + **forward:** In the forward you see how it is used. We start by passing the x (image) into each of the bodys and contatenating it together. That then is passed to the head. Exactly what we want!
#
# >Note: If any of this doesn't make sense to you, I recommend reading [Chapter 15 of the Deep Learning with Fastai and Pytorch book](https://github.com/fastai/fastbook/blob/master/15_arch_details.ipynb). That chapter has all the concepts needed to understand what's going on here.
class PetsModel(Module):
def __init__(self, encoder_list,vocab):
self.body_list = [encoder[:-1] for encoder in encoder_list]
self.head = create_head(512*len(self.body_list)*2,len(vocab))
self.split = [params(body) for body in self.body_list] + [params(self.head)]
def layer_splitter(self,model):
return self.split
def forward(self, x):
# concatenate the outputs of the cut species and cut breed models together
ftrs = torch.cat([body(x) for body in self.body_list], dim=1)
# Feed the concatenaded outputs to the model head
return self.head(ftrs)
# # The Experiment
# Now that we understand this custom architecture, let's go over what all the variations we want to compare.
#
# 1. **resnet18, resnet34, resnet50:** If this custom architecture isn't better than one of these in some way (speed to train, validation accuracy, etc), then we may as well just use a resnet. So we need to include those for comparison.
# 1. **2UntrainedBodies, 3UntrainedBodies:**
# + Uses 2 or 3 bodies respectively
# + Bodys are each resnet18
# + Uses pretrained weights with no seperate training
# 1. **2TrainedBodies, 3TrainedBodies:**
# + Uses 2 or 3 bodies respectively
# + Bodys are each resnet18
# + Takes pretrained resnet18's and seperately trains them on the task first
# 1. **2TrainedDupeBodies,3TrainedDupeBodies:**
# + Uses 2 or 3 bodies respectively
# + Bodys are each resnet18
# + Takes pretrained resnet18, train it first, then have bodys duplicates of the trained model
# 1. **2HeirarchyBodies:**
# + Uses 2 bodys
# + Bodys are each resnet18
# + Body1 is a resnet18 trained to predict species (higher level in heirarchy)
# + Body2 is a resnet18 trained to predict breed (lower level in heirarchy)
#
# # The Results
#
# I will be graphing some key findings to look at, and only graphing the best models.
#hide
def get_dfs(fname):
with open(fname, 'rb') as f:
results = pickle.load(f)
dfs = []
for exp_name in results.keys():
tmp = pd.DataFrame(results[exp_name])
for row in range(len(tmp)):
if type(tmp.iloc[row,3]) != float:
tmp.iloc[row,3] = int(tmp.iloc[row,3][:2])*60 + int(tmp.iloc[row,3][-2:])
tmp.iloc[:,3] = tmp.iloc[:,3].astype(int).cumsum()
tmp[1] = [min(10,o) for o in tmp[1]]
tmp.loc[tmp[1]==10,1] = np.nan
df = tmp
df.reset_index(drop=True,inplace=True)
dfs.append(df)
return dfs
#hide
dfs = get_dfs('ts2021-01-18 17:28:22.315808-cp148.pkl')
# Let's start with with a review of the 3 best models:
# + `2UnTrainedBodies`: Had 2 `resnet18` bodies, both using pytorch pretrained model weights.
# + `3UnTrainedBodies`: Had 3 `resnet18` bodies, each using pytorch pretrained model weights.
# + `2TrainedDupeBodies`: Fine tuned 1 `resnet18`, then used the weights from that model for both bodies
#
# Let's start by looking at validation accuracy. Key notes:
# + This looks at the best 3 models only
# + We can see that our best model models are between a resnet 34 and resnet 50 in terms of accuracy consistently in the last 20 epochs
# + Based on these results, there isn't really any value in training the bodies seperately before combining them. It's more beneficial to train the system as a whole immediately.
#hide
def plot_results(metric,models,ylim=None,xlim=None):
fig, ax = plt.subplots(1,1,figsize=(16,8))
metrics = ['Train Loss','Validation Loss','Validation Accuracy','Cumulative Train Time']
col_idx = int([i for i, s in enumerate(metrics) if metric in s][0])
for j in models:
col = dfs[j].columns[col_idx]
ax.plot(dfs[j][col], label=L(results.keys())[j])
ax.legend()
ax.set_xlabel('epoch')
ax.set_ylabel(metric)
ax.set_title(metric)
if ylim: ax.set_ylim(*ylim)
if xlim: ax.set_xlim(*xlim)
#hide_input
plot_results('Validation Accuracy',[1,6,5,8,0],(0.65,0.8),(140,160));
# Now the graph above does a great job of showing the consistency in the final epochs, but it does hide some of the early training instability. Here's a fuller view
#hide_input
plot_results('Validation Accuracy',[1,6,5,8,0],ylim=(0.5,0.8));
# Great! So this seems really promising. But the real question now is why use this approach over a `resnet50`? To do that let's look at the time to process (below).
#
# Below we see a few things:
# + The `resnet50` (which was the most accurate) takes the longest to train
# + The `resnet34` takes **slightly** more time to train as the `2TrainedDupeBodies` and the `2UntrainedBodies` model, even though the `resnet34` is less accurate.
# + The `3UntrainedBodies` model is not quite as accurate as the `resnet50` but also doesn't take as long to train.
#
# >Note: This is really important because 2 of the custom architectures have a better accuracy and trains in the slightly less time than a `resnet34`. That's great news!
#
# We will look at a zoomed in view so the differentiations are clearer. Please be aware the Y axis does not start at 0.
#hide_input
models=[1,6,5,8,0]
fig, ax = plt.subplots(1,1,figsize=(16,8))
labels = L(results.keys())[models]
values = [max(df[3]) for df in L(dfs)[models]]
ax.bar(labels,values)
ax.set_xlabel('Model')
ax.set_ylabel('Seconds')
ax.set_title('Train Time')
ax.set_ylim(2200,2400)
plt.show()
| _notebooks/2021-02-01-CustomArchitectureResearch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.model_selection import cross_val_score, train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score,f1_score,confusion_matrix
from sklearn.model_selection import GridSearchCV
dataset = pd.read_csv('../../Data/clean_profile_data_all.csv')
dataset.head()
dataset.university_name.value_counts()
target_universities=dataset.university_name.unique().tolist()
from sklearn.utils import resample
# +
resampled_dfs=[]
resampled_df = pd.DataFrame()
for each in target_universities:
if dataset[(dataset.university_name==each )].shape[0]> 600:
resampled_dfs.append(resample(dataset[(dataset.university_name==each )&(dataset.status=='accept')],replace=True,n_samples=300,random_state=123))
resampled_dfs.append(resample(dataset[(dataset.university_name==each) &(dataset.status=='reject')],replace=True,n_samples=300,random_state=123))
elif dataset[(dataset.university_name==each )].shape[0] < 200:
resampled_dfs.append(resample(dataset[(dataset.university_name==each )&(dataset.status=='accept')],replace=True,n_samples=125,random_state=123))
resampled_dfs.append(resample(dataset[(dataset.university_name==each) &(dataset.status=='reject')],replace=True,n_samples=125,random_state=123))
else:
resampled_dfs.append(dataset[(dataset.university_name==each )&(dataset.status=='accept')])
resampled_dfs.append(dataset[(dataset.university_name==each )&(dataset.status=='reject')])
resampled_df = pd.concat( [ f for f in resampled_dfs ] )
# -
resampled_df.groupby(by='university_name')['status'].value_counts()
dataset =resampled_df.copy()
dataset
#train test split for modelling
training, testing = train_test_split(dataset, test_size=0.25, random_state=5, stratify=dataset[['university_name', 'status']])
testing.groupby(by=['university_name'])['status'].value_counts()
training.shape,testing.shape
model_name=[]
model_train_acc=[]
model_test_accuracy=[]
model_train_f1=[]
model_test_f1=[]
from sklearn.model_selection import StratifiedKFold
import numpy as np
def get_result(model, X_train, X_test, Y_train, Y_test):
sc = StandardScaler()
sc.fit(X_train)
X_train = sc.transform(X_train)
X_test = sc.transform(X_test)
model.fit(X_train, Y_train)
y_pred = model.predict(X_test)
y_train_pred = model.predict(X_train)
prob_test=pd.DataFrame(model.predict_proba(X_test))
prob_train=pd.DataFrame(model.predict_proba(X_train))
test_f1_score = f1_score(Y_test, y_pred,pos_label='accept')
train_f1_score = f1_score(Y_train, y_train_pred,pos_label='accept')
train_accuracy=accuracy_score(Y_train, y_train_pred)
test_accuracy=accuracy_score(Y_test, y_pred)
test_cm = confusion_matrix(Y_test, y_pred,labels=['accept','reject'])
train_cm = confusion_matrix(Y_train, y_train_pred,labels=['accept','reject'])
model_name.append(model)
model_train_acc.append(train_accuracy)
model_test_accuracy.append(test_accuracy)
model_test_f1.append(test_f1_score)
model_train_f1.append(train_f1_score)
return [train_cm,test_cm,train_accuracy,test_accuracy,train_f1_score, test_f1_score, prob_train,prob_test, y_pred,y_train_pred, model,sc]
# +
#test and train roc
import numpy as np
from sklearn import metrics
from sklearn.metrics import roc_curve,auc
import matplotlib.pyplot as plt
import seaborn as sns
def generate_cm_roc(model_results):
test_fpr,test_tpr,test_thresholds = metrics.roc_curve(testing['status'], model_results[7][0],pos_label='accept')
test_roc_auc = auc(test_fpr, test_tpr)
train_fpr,train_tpr,train_thresholds = metrics.roc_curve(training['status'], model_results[6][0],pos_label='accept')
train_roc_auc = auc(train_fpr, train_tpr)
plt.plot(train_fpr, train_tpr, lw=2, alpha=0.5,
label='Train ROC (auc= %0.2f)' % (train_roc_auc))
plt.plot(test_fpr, test_tpr, lw=2, alpha=0.5,
label='Test ROC (auc= %0.2f)' % (test_roc_auc))
plt.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
label='Chance', alpha=.8)
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.show()
f,ax = plt.subplots(figsize=(2, 2))
sns.heatmap(model_results[0], annot=True, linewidths=1, fmt= '.1f',ax=ax)
plt.title('Train Confusion Matrix')
plt.show()
f,ax = plt.subplots(figsize=(2, 2))
sns.heatmap(model_results[1], annot=True, linewidths=1, fmt= '.1f',ax=ax)
plt.title('Test confusion matrix')
plt.show()
# -
numerical_data = training.select_dtypes(include = ['int64','float','uint8'])
categorical_data = training.select_dtypes(include = ['object'])
categorical_features = categorical_data.columns.values
numerical_features = numerical_data.columns.values
numerical_features
# +
from sklearn.svm import SVC
svc_model=SVC(gamma='auto',probability=True)
svc_model_results=get_result(svc_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
# -
print('test_accuracy:',svc_model_results[3])
print('train_accuracy:',svc_model_results[2])
print('test_f1_score:',svc_model_results[5])
print('train_f1_score:',svc_model_results[4])
generate_cm_roc(svc_model_results)
# +
from sklearn.tree import DecisionTreeClassifier
decision_tree_model=DecisionTreeClassifier()
decision_tree_model_results=get_result(decision_tree_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
# -
print('test_accuracy:',decision_tree_model_results[3])
print('train_accuracy:',decision_tree_model_results[2])
print('test_f1_score:',decision_tree_model_results[5])
print('train_f1_score:',decision_tree_model_results[4])
decision_tree_model_results[10].get_params
plt.figure(figsize=(20,5))
plt.bar(numerical_features.tolist(),decision_tree_model_results[10].feature_importances_)
# +
from sklearn.ensemble import RandomForestClassifier
random_forest_model=RandomForestClassifier(n_estimators=10)
random_forest_model_results=get_result(random_forest_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',random_forest_model_results[3])
print('train_accuracy:',random_forest_model_results[2])
print('test_f1_score:',random_forest_model_results[5])
print('train_f1_score:',random_forest_model_results[4])
# -
random_forest_model_results[10].get_params
# +
from sklearn.ensemble import RandomForestClassifier
random_forest_model=random_forest_model_results[10]
param_grid = {"n_estimators": [10,15,20,25,30],
"max_depth": [10,15,20,25],
"bootstrap": [True, False]}
random_forest_model_clf = GridSearchCV(random_forest_model,param_grid,cv=5,return_train_score=True)
random_forest_model_results=get_result(random_forest_model_clf,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',random_forest_model_results[3])
print('train_accuracy:',random_forest_model_results[2])
print('test_f1_score:',random_forest_model_results[5])
print('train_f1_score:',random_forest_model_results[4])
# -
generate_cm_roc(random_forest_model_results)
random_forest_model_results[10].best_score_
random_forest_model_results[10].cv_results_
# +
import pickle
# Dump the trained decision tree classifier with Pickle
rf_classifier_pkl_filename = 'student_university_random_forest_predict.pickel'
standard_scaler_filename = 'standard_scaler_rf_model.pickel'
random_forest_classifier_model_pkl = open(rf_classifier_pkl_filename, 'wb')
pickle.dump(random_forest_model_results[10], random_forest_classifier_model_pkl)
random_forest_classifier_model_pkl.close()
sc_rf_classifier_scaler_pkl = open(standard_scaler_filename, 'wb')
pickle.dump(random_forest_model_results[11], sc_rf_classifier_scaler_pkl)
sc_rf_classifier_scaler_pkl.close()
random_forest_classifier_model_pkl = open(rf_classifier_pkl_filename, 'rb')
random_forest_classifier_model= pickle.load(random_forest_classifier_model_pkl)
random_forest_classifier_model_pkl.close()
sc_rf_classifier_scaler_pkl = open(standard_scaler_filename, 'rb')
standard_scaler_rf_classifier= pickle.load(sc_rf_classifier_scaler_pkl)
sc_rf_classifier_scaler_pkl.close()
# -
standard_scaler_rf_classifier
predictions=random_forest_classifier_model.predict(standard_scaler_rf_classifier.transform(testing[numerical_features]))
accuracy_score(testing['status'],predictions)
# +
from sklearn.naive_bayes import GaussianNB
gnb_model=GaussianNB()
gnb_model_results=get_result(gnb_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',gnb_model_results[3])
print('train_accuracy:',gnb_model_results[2])
print('test_f1_score:',gnb_model_results[5])
print('train_f1_score:',gnb_model_results[4])
# +
from sklearn.neural_network import MLPClassifier
mlp_model=MLPClassifier(max_iter=500,solver='adam')
mlp_model_results=get_result(mlp_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',mlp_model_results[3])
print('train_accuracy:',mlp_model_results[2])
print('test_f1_score:',mlp_model_results[5])
print('train_f1_score:',mlp_model_results[4])
# +
from xgboost import XGBClassifier
xgb_model=XGBClassifier()
xgb_model_results=get_result(xgb_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',xgb_model_results[3])
print('train_accuracy:',xgb_model_results[2])
print('test_f1_score:',xgb_model_results[5])
print('train_f1_score:',xgb_model_results[4])
# +
from sklearn.ensemble import BaggingClassifier
from sklearn import tree
bagging_model=BaggingClassifier(tree.DecisionTreeClassifier(random_state=1),n_estimators = 100, max_features = 0.8)
bagging_model_results=get_result(bagging_model,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',bagging_model_results[3])
print('train_accuracy:',bagging_model_results[2])
print('test_f1_score:',bagging_model_results[5])
print('train_f1_score:',bagging_model_results[4])
# -
bagging_model_results[10].score
generate_cm_roc(bagging_model_results)
# +
param_grid = {"base_estimator__max_depth" : [30,40,50],
"max_samples" : [0.05, 0.1, 0.2,0.5,0.75,0.8,1]
}
clf = GridSearchCV(bagging_model_results[10],param_grid,cv=5,return_train_score=True)
#tree.DecisionTreeClassifier(),n_estimators = 100, max_features = 0.8
bagging_model_results=get_result(clf,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',bagging_model_results[3])
print('train_accuracy:',bagging_model_results[2])
print('test_f1_score:',bagging_model_results[5])
print('train_f1_score:',bagging_model_results[4])
generate_cm_roc(bagging_model_results)
# -
bagging_model_results[10].cv_results_
bagging_model_results[10].best_score_
predictions=bagging_model_results[10].best_estimator_.predict(bagging_model_results[11].transform(testing[numerical_features]))
accuracy_score(testing['status'],predictions)
bagging_model_results[10].best_params_
# +
import pickle
# Dump the trained decision tree classifier with Pickle
bagging_classifier_pkl_filename = 'student_university_bagging_classifier_predict.pickel'
standard_scaler_filename = 'standard_scaler_bagging_model.pickel'
bagging_classifier_model_pkl = open(bagging_classifier_pkl_filename, 'wb')
pickle.dump(bagging_model_results[10], bagging_classifier_model_pkl)
bagging_classifier_model_pkl.close()
sc_bagging_classifier_scaler_pkl = open(standard_scaler_filename, 'wb')
pickle.dump(bagging_model_results[11], sc_bagging_classifier_scaler_pkl)
sc_bagging_classifier_scaler_pkl.close()
bagging_classifier_model_pkl = open(bagging_classifier_pkl_filename, 'rb')
bagging_classifier_model= pickle.load(bagging_classifier_model_pkl)
bagging_classifier_model_pkl.close()
sc_bagging_classifier_scaler_pkl = open(standard_scaler_filename, 'rb')
standard_scaler_bagging_classifier= pickle.load(sc_bagging_classifier_scaler_pkl)
sc_bagging_classifier_scaler_pkl.close()
# -
standard_scaler_bagging_classifier
predictions=bagging_classifier_model.predict(standard_scaler_bagging_classifier.transform(testing[numerical_features]))
accuracy_score(testing['status'],predictions)
bagging_classifier_model.classes_
# +
from sklearn.neighbors import KNeighborsClassifier
knn_clf =BaggingClassifier(KNeighborsClassifier(),n_estimators = 100, max_features = 0.8)
bagging_model_results1=get_result(knn_clf,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
# -
print('test_accuracy:',bagging_model_results1[3])
print('train_accuracy:',bagging_model_results1[2])
print('test_f1_score:',bagging_model_results1[5])
print('train_f1_score:',bagging_model_results1[4])
generate_cm_roc(bagging_model_results1)
# +
from sklearn.model_selection import GridSearchCV
param_grid = {'base_estimator__n_neighbors' : [3,5,7]}
knn_clf = GridSearchCV(BaggingClassifier(KNeighborsClassifier(),n_estimators = 100, max_features = 0.8),param_grid,cv=5)
bagging_model_results1=get_result(knn_clf,training[numerical_features],testing[numerical_features],training['status'],testing['status'])
print('test_accuracy:',bagging_model_results1[3])
print('train_accuracy:',bagging_model_results1[2])
print('test_f1_score:',bagging_model_results1[5])
print('train_f1_score:',bagging_model_results1[4])
generate_cm_roc(bagging_model_results1)
# -
bagging_model_results1[10].best_score_
bagging_model_results1[10].best_params_
# +
import pickle
# Dump the trained decision tree classifier with Pickle
bagging_classifier_kNN_pkl_filename = 'student_university_kNN_bagging_classifier_predict.pickel'
standard_scaler_kNN_filename = 'standard_scaler_kNN_bagging_model.pickel'
bagging_classifier_model_pkl = open(bagging_classifier_kNN_pkl_filename, 'wb')
pickle.dump(bagging_model_results1[10], bagging_classifier_model_pkl)
bagging_classifier_model_pkl.close()
sc_bagging_classifier_scaler_pkl = open(standard_scaler_kNN_filename, 'wb')
pickle.dump(bagging_model_results1[11], sc_bagging_classifier_scaler_pkl)
sc_bagging_classifier_scaler_pkl.close()
bagging_classifier_model_pkl = open(bagging_classifier_kNN_pkl_filename, 'rb')
bagging_classifier_model= pickle.load(bagging_classifier_model_pkl)
bagging_classifier_model_pkl.close()
sc_bagging_classifier_scaler_pkl = open(standard_scaler_kNN_filename, 'rb')
standard_scaler_bagging_classifier= pickle.load(sc_bagging_classifier_scaler_pkl)
sc_bagging_classifier_scaler_pkl.close()
# -
standard_scaler_bagging_classifier
predictions=bagging_classifier_model.predict(standard_scaler_bagging_classifier.transform(testing[numerical_features]))
accuracy_score(testing['status'],predictions)
All_Model_results=pd.DataFrame()
All_Model_results["Model"]=model_name
All_Model_results['Train Accuracy']=model_train_acc
All_Model_results['Test Accuracy']=model_test_accuracy
All_Model_results['Train F1 score']=model_train_f1
All_Model_results['Test F1 score']=model_test_f1
All_Model_results
# ## model performances
All_Model_results.to_csv('All_Model_results.csv',index=False)
# # summarizing best model results
# <table style="width:100%">
# <tr>
# <th>Model</th>
# <th>Test Accuracy</th>
# <th>Train Accuracy</th>
# <th>Grid Search - Test Accuracy</th>
# <th>Grid Search - Train Accuracy</th>
# <th>Test F1 Score</th>
# <th>Train F1 Score</th>
# <th>Test AUC ROC Curve</th>
# <th>Train AUC ROC Curve</th>
# </tr>
# <tr>
# <th>Bagging - Decision Tree</th>
# <th>0.79</th>
# <th>0.99</th>
# <th>0.80</th>
# <th>0.98</th>
# <th>0.78</th>
# <th>0.99</th>
# <th>0.89</th>
# <th>0.99</th>
# </tr>
# <tr>
# <th>Random Forest Classifer</th>
# <th>0.77</th>
# <th>0.98</th>
# <th>0.78</th>
# <th>0.97</th>
# <th>0.77</th>
# <th>0.97</th>
# <th>0.87</th>
# <th>1.0</th>
# </tr>
# <tr>
# <th>Bagging - kNN(k=3(Best))</th>
# <th>0.72</th>
# <th>0.87</th>
# <th>0.74</th>
# <th>0.94</th>
# <th>0.73</th>
# <th>0.94</th>
# <th>0.82</th>
# <th>0.99</th>
# </tr>
#
# </table>
#
# - Bagging model with Decision Tree is giving us best after GridSearchCV(CV=5) results with ~80% test accuracy and 0.78 f1 score
# - Random Forest Classifier is giving us 2nd best results best after GridSearchCV(CV=5) with ~77% test accuracy and 0.77 f1 score
# - Bagging model with kNN(k=3) is giving us 3rd best results best after GridSearchCV(CV=5) with ~72% test accuracy and 0.73 f1 score
| Final Project _ Graduate Admission Predictor/Code/Modelling/all_profiles_manual_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Linear regression is used to predict the value of an outcome variable Y based on one or more input predictor variables X. The aim is to establish a linear relationship (a mathematical formula) between the predictor variable(s) and the response variable
import numpy as np
import math
import matplotlib
import matplotlib.pyplot as plt
from matplotlib import style
import pandas as pd
import sklearn #for linear and other models
import warnings
from sklearn import linear_model
from sklearn.cross_validation import train_test_split
# %matplotlib inline
#two lists xs and ys
xs=[1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24]
ys=[23,24,25,26,27,28,29,30,34,45,46,51,56,57,58,62,64,67,72,75,77,81,84,83]
len(xs),len(ys)
plt.scatter(xs,ys)
plt.ylabel("independent variable")
plt.xlabel("dependent variable")
plt.show()
#function for intercept and slope
def slope_intercept(x_val,y_val):
x=np.array(x_val)
y=np.array(y_val)
m=(((np.mean(x)*np.mean(y))-np.mean(x*y))/(np.mean(x)*np.mean(x))-np.mean(x*x))
m=round(m,2)
b=(np.mean(y)-np.mean(x)*m)
b=round(b,2)
return m,b
slope_intercept(xs,ys)
m,b=slope_intercept(xs,ys)
reg_line=[(m*x)+b for x in xs]
plt.scatter(xs,ys,color="red")
plt.plot(xs,reg_line)
plt.title("making a regression line")
plt.xlabel("independent variable")
plt.ylabel("dependent variable")
plt.show()
#rmse
import math
def rmsm(y1,y_h):
y_actual=np.array(y1)
y_pred=np.array(y_h)
error=(y_actual-y_pred)**2
error_mean=round(np.mean(error))
err_sq=math.sqrt(error_mean)
return err_sq
rmsm(ys,reg_line)
# +
import scipy.stats as stats
import matplotlib.pyplot as plt
import sklearn
import statsmodels.api as sm
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")
#special matplotlib argument for improved plots
from matplotlib import rcParams
from sklearn.datasets import load_boston
boston=load_boston()
df_x=pd.DataFrame(boston.data,columns=boston.feature_names)#data frame for independent variables
df_y=pd.DataFrame(boston.target)#dependent variable
# -
print(boston.keys())
df_x.head(13)
df_y.head(10)
df_x.shape #tells number of rows and columns
print(boston.data.shape)
names=[i for i in list(df_x)]
names
print(boston.feature_names)
print(boston.DESCR)
#In a dataset a training set is implemented to build up a model, while a test (or validation) set
#is to validate the model built.
regr=linear_model.LinearRegression()
x_train,x_test,y_train,y_test=train_test_split(df_x,df_y,test_size=0.2,random_state=4)
#use 20% of total data for data test
x_train.head()
#fit linear regression model to training data set
regr.fit(x_train,y_train)
regr.intercept_
#the coefficients
print("coefficients are:",regr.coef_)
#mean squared error
print("mean squared error: ",np.mean((regr.predict(x_test)-y_test)**2))
#variance score:1 is perfect prediction
print("variance score:",regr.score(x_test,y_test))
#coefficients of Independent variables (slope (m) of the regression line)
regr.coef_[0].tolist()
#attach slopes to these variables
pd.DataFrame(zip(names,regr.coef_[0].tolist()),columns=['names','coefficients'])
#plotting predicted x_test,y_test values
style.use("bmh")
plt.scatter(regr.predict(x_test),y_test)
plt.show()
#calculate p value
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
modedl1=sm.OLS(y_train,x_train)
result=modedl1.fit()
print(result.summary())
#select variables with p-values <0.5
model2=sm.OLS(y_train,x_train[['CRIM','ZN','CHAS','RM','DIS','RAD','TAX','PTRATIO','B','LSTAT']])
result2=model2.fit()
print(result2.summary())
#deal with multicollinearity
import seaborn
corr_df=x_train.corr(method='pearson')
print("-------------------create a correlation plot-------------------")
#create a mask to display only lower triangle
mask=np.zeros_like(corr_df)
mask[np.triu_indices_from(mask)]=True
#create heatmap using seaborn lib
#list if colormaps (parameter'cmap'is available)
seaborn.heatmap(corr_df,cmap='Accent',vmax=1.0,vmin=1.0,mask=mask,linewidths=2.5)
#show the plot
plt.yticks(rotation=0)
plt.xticks(rotation=90)
plt.show()
print("----------------------------------------end----------------------------------------!!")
| sslinearregression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rhgpM92PMCd-"
# # Flower Quickstart (Simulation with TensorFlow/Keras)
#
# Welcome to Flower, a friendly federated learning framework!
#
# In this notebook, we'll simulate a federated learning system with 100 clients. The clients will use TensorFlow/Keras to define model training and evaluation. Let's start by installing Flower Nightly, published as `flwr-nightly` on PyPI:
# + colab={"base_uri": "https://localhost:8080/"} id="BXG21C3dLj6i" outputId="f60586a0-3fc5-4801-919d-01977224a39b"
# !pip install git+https://github.com/adap/flower.git@release/0.17#egg=flwr["simulation"] # For a specific branch (release/0.17) w/ extra ("simulation")
# # !pip install -U flwr["simulation"] # Once 0.17.1 is released
# + [markdown] id="tQk9ZzCBMf9r"
# Next, we import the required dependencies. The most important imports are Flower (`flwr`) and TensorFlow:
# + id="oKvjox6uMkhj"
import os
import math
# Make TensorFlow logs less verbose
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
import flwr as fl
import tensorflow as tf
# + [markdown] id="30pJWfaTM_MC"
# With that out of the way, let's move on to the interesting bits. Federated learning systems consist of a server and multiple clients. In Flower, we create clients by implementing subclasses of `flwr.client.Client` or `flwr.client.NumPyClient`. We use `NumPyClient` in this tutorial because it is easier to implement and requires us to write less boilerplate.
#
# To implement the Flower client, we create a subclass of `flwr.client.NumPyClient` and implement the three methods `get_parameters`, `fit`, and `evaluate`:
#
# - `get_parameters`: Return the current local model parameters
# - `fit`: Receive model parameters from the server, train the model parameters on the local data, and return the (updated) model parameters to the server
# - `evaluate`: Received model parameters from the server, evaluate the model parameters on the local data, and return the evaluation result to the server
#
# We mentioned that our clients will use TensorFlow/Keras for the model training and evaluation. Keras models provide methods that make the implementation staightforward: we can update the local model with server-provides parameters through `model.set_weights`, we can train/evaluate the model through `fit/evaluate`, and we can get the updated model parameters through `model.get_weights`.
#
# Let's see a simple implementation:
# + id="vE3mqBs0NHZi"
class FlowerClient(fl.client.NumPyClient):
def __init__(self, model, x_train, y_train, x_val, y_val) -> None:
self.model = model
self.x_train, self.y_train = x_train, y_train
self.x_val, self.y_val = x_train, y_train
def get_parameters(self):
return self.model.get_weights()
def fit(self, parameters, config):
self.model.set_weights(parameters)
self.model.fit(self.x_train, self.y_train, epochs=1, verbose=2)
return self.model.get_weights(), len(self.x_train), {}
def evaluate(self, parameters, config):
self.model.set_weights(parameters)
loss, acc = self.model.evaluate(self.x_val, self.y_val, verbose=2)
return loss, len(self.x_val), {"accuracy": acc}
# -
# Our class `FlowerClient` defines how local training/evaluation will be performed and allows Flower to call the local training/evaluation through `fit` and `evaluate`. Each instance of `FlowerClient` represents a *single client* in our federated learning system. Federated learning systems have multiple clients (otherwise there's not much to federate, is there?), so each client will be represented by its own instance of `FlowerClient`. If we have, for example, three clients in our workload, we'd have three instances of `FlowerClient`. Flower calls `FlowerClient.fit` on the respective instance when the server selects a particular client for training (and `FlowerClient.evaluate` for evaluation).
#
# In this notebook, we want to simulate a federated learning system with 100 clients on a single machine. This means that the server and all 100 clients will live on a single machine and share resources such as CPU, GPU, and memory. Having 100 clients would mean having 100 instances of `FlowerClient` im memory. Doing this on a single machine can quickly exhaust the available memory resources, even if only a subset of these clients participates in a single round of federated learning.
#
# In addition to the regular capabilities where server and clients run on multiple machines, Flower therefore provides special simulation capabilities that create `FlowerClient` instances only when they are actually necessary for training or evaluation. To enable the Flower framework to create clients when necessary, we need to implement a function called `client_fn` that creates a `FlowerClient` instance on demand. Flower calls `client_fn` whenever it needs an instance of one particular client to call `fit` or `evaluate` (those instances are usually discarded after use). Clients are identified by a client ID, or short `cid`. The `cid` can be used, for example, to load different local data partitions for each client:
# + id="OtlBA2OFO0tf"
def client_fn(cid: str) -> fl.client.Client:
# Create model
model = tf.keras.models.Sequential(
[
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation="softmax"),
]
)
model.compile("adam", "sparse_categorical_crossentropy", metrics=["accuracy"])
# Load data partition (divide MNIST into NUM_CLIENTS distinct partitions)
(x_train, y_train), _ = tf.keras.datasets.mnist.load_data()
partition_size = math.floor(len(x_train) / NUM_CLIENTS)
idx_from, idx_to = int(cid) * partition_size, (int(cid) + 1) * partition_size
x_train_cid = x_train[idx_from:idx_to] / 255.0
y_train_cid = y_train[idx_from:idx_to]
# Use 10% of the client's training data for validation
split_idx = math.floor(len(x_train) * 0.9)
x_train_cid, y_train_cid = x_train_cid[:split_idx], y_train_cid[:split_idx]
x_val_cid, y_val_cid = x_train_cid[split_idx:], y_train_cid[split_idx:]
# Create and return client
return FlowerClient(model, x_train_cid, y_train_cid, x_val_cid, y_val_cid)
# + [markdown] id="6SVawWSgO48Q"
# We now have `FlowerClient` which defines client-side training and evaluation and `client_fn` which allows Flower to create `FlowerClient` instances whenever it needs to call `fit` or `evaluate` on one particular client. The last step is to start the actual simulation using `flwr.simulation.start_simulation`.
#
# The function `start_simulation` accepts a number of arguments, amongst them the `client_fn` used to create `FlowerClient` instances, the number of clients to simulate `num_clients`, the number of rounds `num_rounds`, and the strategy. The strategy encapsulates the federated learning approach/algorithm, for example, *Federated Averaging* (FedAvg).
#
# Flower comes with a number of built-in strategies, but we can also use our own strategy implementations to customize nearly all aspects of the federated learning approach. For this example, we use the built-in `FedAvg` implementation and customize it using a few basic parameters. The last step is the actual call to `start_simulation` which - you guessed it - actually starts the simulation.
# + colab={"base_uri": "https://localhost:8080/"} id="0Yxjysu5PM-A" outputId="cf5f1bab-0d94-4876-bd6a-cdb0652826ff"
NUM_CLIENTS = 100
# Create FedAvg strategy
strategy=fl.server.strategy.FedAvg(
fraction_fit=0.1, # Sample 10% of available clients for training
fraction_eval=0.05, # Sample 5% of available clients for evaluation
min_fit_clients=10, # Never sample less than 10 clients for training
min_eval_clients=10, # Never sample less than 5 clients for evaluation
min_available_clients=int(NUM_CLIENTS * 0.75), # Wait until at least 75 clients are available
)
# Start simulation
fl.simulation.start_simulation(
client_fn=client_fn,
num_clients=NUM_CLIENTS,
num_rounds=5,
strategy=strategy,
)
# -
# Congratulations! With that, you built a Flower client, customized it's instantiation through the `client_fn`, customized the server-side execution through a `FedAvg` strategy configured for this workload, and started a simulation with 100 clients (each holding their own individual partition of the MNIST dataset).
#
# Next, you can continue to explore more advanced Flower topics:
#
# - Deploy server and clients on different machines using `start_server` and `start_client`
# - Customize the server-side execution through custom strategies
# - Customize the client-side exectution through `config` dictionaries
| examples/quickstart_simulation/sim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Keithleene/CPEN-21A-ECE-2-1/blob/main/Lab_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UvPLSQLaugEc"
# #Laboratory 1
# + colab={"base_uri": "https://localhost:8080/"} id="JxH48Cw_wfwf" outputId="e6931fb5-9259-4982-cec2-d51203bf0103"
W = "Welcome to Python Programming"
print(W)
# + colab={"base_uri": "https://localhost:8080/"} id="O4TeiuF1urjZ" outputId="cd91eadf-ca01-4909-9324-974e4a4556b5"
W = "Welcome to Python Programming"
print(W)
# + colab={"base_uri": "https://localhost:8080/"} id="njJzY9h0vfk7" outputId="f039b3a6-f669-46e6-9322-e9803ee632b7"
P = "Name: <NAME>"
B = "Address: 514 Arvisu St. Biwas Tanza Cavite"
G = "Age: 19"
print(P)
print(B)
print(G)
| Lab_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
df = pd.read_csv('../Dados/provided/siscad-alunos-2019-1.csv', sep=',')
df.columns
# ### **TIPOS DE SITUAÇÃO - ALUNO**
df['situacao'].value_counts()
df.groupby(by=['curso','unidade']).count().rename(columns={'nome':'quantidade'})[['quantidade']]
df.groupby(by=['curso','unidade'])[['nome']].count()
df.groupby(by=['curso'])['curso'].count()
df['curso'].value_counts().sort_values()
# ### **ALUNOS COM MATRÍCULA REGULAR**
matriculados = df[df['situacao']=='REGULARMENTE MATRICULADO NO PERÍODO']
matriculados.count()
matriculados.to_excel('../Dados/generated/Alunos-Matriculados.xlsx',index=False)
cppp = matriculados[matriculados['unidade'] == 'CPPP']
rga_2017 = cppp[cppp['rga_descaracterizado'].str.contains('2017')]
rga_2017[rga_2017['curso'].str.contains('SISTEMAS')]
rga_2018 = cppp[cppp['rga_descaracterizado'].str.contains('2018')]
rga_2018[rga_2018['curso'].str.contains('SISTEMAS')].count()
rga_2019 = cppp[cppp['rga_descaracterizado'].str.contains('2019')]
rga_2019[rga_2019['curso'].str.contains('CIÊNCIA')].count()
rga_2019[rga_2019['curso'].str.contains('SISTEMAS')].count()
rga_2019[rga_2019['curso'].str.contains('MATEMÁTICA')].count()
rga_2019[rga_2019['curso'].str.contains('PEDAGOGIA')].count()
| Notebooks/UFMS_DATASET.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: GT
# language: python
# name: gt
# ---
# # Normal Form Games
#
# [Video](https://youtu.be/VDZ4I4IoFss?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# Game theory is the study of interactive decision making. Consider the following situation:
#
# > Two friends must decide what movie to watch at the cinema. Alice would like to watch a sport movie and Bob would like to watch a comedy. Importantly they would both rather spend their evening together then apart.
#
# To represent this mathematically we will associate **utilities** to the 4 possible outcomes:
#
# 1. Alice watches a sport movie, Bob watches a comedy: Alice receives a utility of $1$ and Bob a utility of $1$.
# 2. Alice watches a comedy, Bob watches a sport movice: Alice receives a utility of $0$ and Bob a utility of $0$.
# 3. Alice and Bob both watch a sport movie: Alice receives a utility of $3$ and Bob a utility of $2$.
# 4. Alice and Bob both watch a comedy: Alice receives a utility of $2$ and Bob a utility of $3$.
#
# This is referred to as the "battle of the sexes" and we will represent it using two matrices, $A$ will represent the utilities of Alice:
#
# $$
# A =
# \begin{pmatrix}
# 3 & 1\\
# 0 & 2
# \end{pmatrix}
# $$
#
# and matrix $B$ will represent the utilities of Bob:
#
# $$
# B =
# \begin{pmatrix}
# 2 & 1\\
# 0 & 3
# \end{pmatrix}
# $$
#
# We refer to **Alice as the row player** and **Bob as the column player**:
#
# - The row player chooses which row of the matrices the players will gain their utilities;
# - The column player chooses which column of the matrices the player will gain their utilities.
#
# Thus if the row player (Alice) chooses the first row (this corresponds to a sport movie) and the column player (Bob) chooses the second column (this corresponds to a comedy):
#
# - The row player receives a utility of $A_{12}=1$
# - The column player receives a utility of $B_{12}=1$
#
# This representation of the stategic interaction between Alice and Bob is called a **Normal Form Game**.
#
# ---
#
# ## Definition of Normal Form Game
#
# [Video](https://youtu.be/tP2WE0FdI0w?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# An \\(N\\) player normal form game consists of:
#
# - A finite set of $N$ players
# - Strategy spaces for the players: $\{S_1,S_2,S_3,\dots,S_N\}$;
# - Payoff functions for the players: $u_i:S_1\times S_2\dots\times S_N\to\mathbb{R}$
#
# ---
#
# **In this course we will only consider the case of $N=2$.**
#
# For the battle of the sexes:
#
# - We have \\(N=2\\) players (Alice and Bob)
# - The strategy spaces: $S_1=S_2=\{\text{comedy}, \text{sport movie}\}$ or equivalently $S_1=S_2=\{1, 2\}$
# - The payoff functions mapping an element of $\tilde s \in S_1\times S_2=\{(1, 1), (1, 2), (2, 1), (2, 2)\}$ to $\mathbb{R}$:
#
# $$u_1(\tilde s)=A_{\tilde s},$$
#
# $$u_2(\tilde s)=B_{\tilde s}.$$
#
# ---
#
# We can use Python to represent these games, we will use the `nashpy` library to do so and we start by building our two matrices:
import nashpy as nash
A = [[3, 1], [0, 2]]
B = [[2, 1], [0, 3]]
# We then create a `nash.Game` instance:
battle_of_the_sexes = nash.Game(A, B)
battle_of_the_sexes
# In the next chapter we will start to see how to use that for further calculations.
#
# # Examples of other common games
#
# ## Prisoners Dilemma
#
# [Video](https://youtu.be/qcQMeiUnfVQ?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# > Assume two thieves have been caught by the police and separated for questioning. If both thieves cooperate and don’t divulge any information they will each get a short sentence. If one defects he/she is offered a deal while the other thief will get a long sentence. If they both defect they both get a medium length sentence.
#
# This corresponds to:
#
# $$
# A =
# \begin{pmatrix}
# 3 & 0\\
# 5 & 1
# \end{pmatrix}\qquad
# B =
# \begin{pmatrix}
# 3 & 5\\
# 0 & 1
# \end{pmatrix}
# $$
A = [[3, 0], [5, 1]]
B = [[3, 5], [0, 1]]
prisoners_dilemma = nash.Game(A, B)
prisoners_dilemma
# ## Hawk Dove game
#
# [Video](https://youtu.be/_7HtcsVB2uU?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# > Suppose two birds of prey must share a limited resource. The birds can act like a hawk or a dove. Hawks always fight over the resource to the point of exterminating a fellow hawk and/or take a majority of the resource from a dove. Two doves can share the resource.
#
#
#
# This corresponds to:
#
# $$
# A =
# \begin{pmatrix}
# 0 & 3\\
# 1 & 2
# \end{pmatrix}\qquad
# B =
# \begin{pmatrix}
# 0 & 1\\
# 3 & 2
# \end{pmatrix}
# $$
A = [[0, 3], [1, 2]]
B = [[0, 1], [3, 2]]
hawk_dove = nash.Game(A, B)
hawk_dove
# ## Pigs
#
# [Video](https://youtu.be/ORGYJdqZkX0?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# > Consider two pigs. One dominant pig and one subservient pig. These pigs share a pen. There is a lever in the pen that delivers food but if either pig pushes the lever it will take them a little while to get to the food. If the dominant pig pushes the lever, the subservient pig has some time to eat most of the food before being pushed out of the way. If the subservient pig push the lever, the dominant pig will eat all the food. Finally if both pigs go to push the lever the subservient pig will be able to eat a third of the food.
#
# This corresponds to:
#
# $$
# A =
# \begin{pmatrix}
# 4 & 2\\
# 6 & 0
# \end{pmatrix}\qquad
# B =
# \begin{pmatrix}
# 2 & 3\\
# -1 & 0
# \end{pmatrix}
# $$
A = [[4, 2], [6, 0]]
B = [[2, 3], [-1, 0]]
pigs = nash.Game(A, B)
pigs
# ## Matching pennies
#
# [Video](https://youtu.be/80ImlktaeeY?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# >Consider two players who can choose to display a coin either Heads facing up or Tails facing up. If both players show the same face then player 1 wins, if not then player 2 wins.
#
# This corresponds to:
#
# $$
# A =
# \begin{pmatrix}
# 1 & -1\\
# -1 & 1
# \end{pmatrix}\qquad
# B =
# \begin{pmatrix}
# -1 & 1\\
# 1 & -1
# \end{pmatrix}
# $$
A = [[1, -1], [-1, 1]]
B = [[-1, 1], [1, -1]]
matching_pennies = nash.Game(A, B)
matching_pennies
# As indicated by `nashpy`, this is a `Zero sum game`:
#
# $$
# A + B = 0
# $$
#
# ---
#
# ## Definition of a zero sum game
#
# [Video](https://youtu.be/wUh1KFupLFI?list=PLnC5h3PY-znxMsG0TRYGOyrnEO-QhVwLb)
#
# A two player normal form game with payoff matrices $A, B$ is called **zero sum** iff:
#
# $$
# A = -B
# $$
#
# ---
#
# To define a zero sum game using `nashpy` we can pass a single payoff matrix (it infers what the other will be):
A = [[1, -1], [-1, 1]]
matching_pennies = nash.Game(A)
matching_pennies
| nbs/chapters/01-Normal-Form-Games.ipynb |
# ---
# title: "Cluster Usage"
# author: <NAME>
# date: December 18, 2020
# output:
# ioslides_presentation:
# keep_md: yes
# widescreen: yes
# df_print: paged
# smaller: true
# subtitle: "Just the Basics...mostly"
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <!--
# - ioslides manual:
# https://bookdown.org/yihui/rmarkdown/ioslides-presentation.html
#
# - Compile from command-line
# jupyter nbconvert Cluster_Usage.ipynb --to markdown && Rscript -e "rmarkdown::render('Cluster_Usage.md'); knitr::knit('Cluster_Usage.md', tangle=TRUE)"
#
# - Jupyter formats
# #jupyter nbextension enable splitcell/splitcell
# #jupyter nbconvert HPCC_Intro.ipynb --to slides --reveal-prefix reveal.js
# #jupyter nbconvert HPCC_Intro.ipynb --to slides --reveal-prefix "https://cdnjs.cloudflare.com/ajax/libs/reveal.js/3.3.0"
# -->
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# ## Summary
#
# * Filesystem
# * Paths
# * Quotas
# * Usage
#
# * Software
# * Module System
# * Installs
# * Management
#
# * Job Scheduling
# * Node
# * Partition
# * Limits
# * Jobs
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Paths
#
# __RoadMap__
#
# <img src="https://docs.google.com/drawings/d/e/2PACX-1vRjaVs9P2GF9oXUem-NNRH6gUD-VQ_N03wKYYHlJ373Qrqb9KPd_oZuFkTzHVFUawNX9ShIHW4u-u2l/pub?w=936&h=380">
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Paths
#
# * Symlink (<span style='color:blue;'>dotted lines</span>) - A shortcut to another directory or file
#
# * Mount (<span style='color:green;'>Local</span>/<span style='color:red;'>Shared</span>) - An entry point to a disk or storage device (ie. `'C:/'` or `Google Drive`)
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Paths
#
# __Case sensitive__
#
# All paths and commands are case sensitive, an uppercase letter is not the same as a lowercase letter.
#
# __Path Types__
#
# * Absolute path - Full path from root to current working directory
#
# ```
# /rhome/username/workshop_dir/
# ```
#
# * Relative path - Partial path or non-absolute path (current directory implied)
#
# ```
# workshop_dir/
# ```
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Quotas
#
# All storage has limits.
#
# * <span style='font-weight:bold;color:green;'>Local Storage</span> (ie. laptop hard drive)
#
# Only exists on a single machine (node) and is limited by disk size.
#
#
# * <span style='font-weight:bold;color:red;'>Shared Storage</span> (ie. Google Drive)
#
# Exists accross all machines (nodes) and is limited by a quota.
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Usage
# + [markdown] slideshow={"slide_type": "fragment"}
# Make workshop directory, if it does not already exist:
# + slideshow={"slide_type": "-"}
mkdir -p ~/workshop_dir
# + [markdown] slideshow={"slide_type": "fragment"}
# Check <span style='font-weight:bold;color:black;'>directory</span> size:
# + slideshow={"slide_type": "-"}
du -hs ~/workshop_dir
# + [markdown] slideshow={"slide_type": "fragment"}
# Check <span style='font-weight:bold;color:green;'>local</span> node storage:
# + slideshow={"slide_type": "-"}
df -h /tmp
# + slideshow={"slide_type": "-"}
df -h /scratch
# -
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filesystem: Usage
# + [markdown] slideshow={"slide_type": "-"}
# Check <span style='font-weight:bold;color:red;'>GPFS</span> storage, _"blocks"_ is used space and available space is _"quota"_:
# + slideshow={"slide_type": "-"}
check_quota home
# + slideshow={"slide_type": "-"}
check_quota bigdata
# + [markdown] slideshow={"slide_type": "-"}
# [https://hpcc.ucr.edu/manuals_linux-cluster_storage](https://hpcc.ucr.edu/manuals_linux-cluster_storage)
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Module System
#
# This system allows multiple versions of software to be loaded and unloaded.
# + [markdown] slideshow={"slide_type": "fragment"}
# To view software that is available:
# + slideshow={"slide_type": "-"}
module avail
# + [markdown] slideshow={"slide_type": "fragment"}
# To search for a specific software:
# + slideshow={"slide_type": "-"}
module avail samtools
# OR
hpcc-software samtools
# + [markdown] slideshow={"slide_type": "-"}
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Module System
# + [markdown] slideshow={"slide_type": "fragment"}
# Load software into current environment:
# + slideshow={"slide_type": "-"}
module load samtools
# + [markdown] slideshow={"slide_type": "fragment"}
# List currently loaded software modules:
# + slideshow={"slide_type": "-"}
module list
# + [markdown] slideshow={"slide_type": "fragment"}
# Remove software from current environment:
# -
module unload samtools
# + [markdown] slideshow={"slide_type": "-"}
# [https://hpcc.ucr.edu/manuals_linux-cluster_start#modules](https://hpcc.ucr.edu/manuals_linux-cluster_start#modules)
#
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Installs
# + [markdown] slideshow={"slide_type": "fragment"}
# __Python__
#
# For a basic `Python` package ([pypi](https://pypi.org/)) you can use `pip` to install it:
#
# ```bash
# pip install PKGNAME --user
# ```
# -
# For example, here is how you would install the `camelcase` package:
#
# ```bash
# pip install camelcase --user
# ```
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Installs
#
# __R__
#
# For an `R` package you can use the install fuction ([CRAN](https://cran.r-project.org/)):
# -
# ```bash
# R
# ```
#
# ```r
# install.packages('PKGNAME')
# ```
# Or you can use the install function from [BiocManager](https://www.bioconductor.org/):
# ```bash
# R
# ```
#
# ```r
# BiocManager::install('PKGNAME')
# ```
# [https://hpcc.ucr.edu/manuals_linux-cluster_package-manage.html#r-1](https://hpcc.ucr.edu/manuals_linux-cluster_package-manage.html#r-1)
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Management
#
# * <span style='font-weight:bold;color:green;'>Conda</span> - A software management system that allows you to install thousands of software packages and tools, including `R` and `Python` languages.
#
# Full instructions regarding conda setup can be found [here](https://hpcc.ucr.edu/manuals_linux-cluster_package-manage.html).
#
#
# * <span style='font-weight:bold;color:red;'>Singularity</span> - A Linux container system (similar to Docker) which allows users to prepare a Linux environment from scratch.
#
# Some singularity examples can be found [here](https://github.com/ucr-hpcc/hpcc_slurm_examples/tree/master/singularity).
#
# A previous workshop regarding custom software installs utilizing the above technologies can be found [here](https://bit.ly/2PXGWEq).
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Management
#
# __Conda__
# + [markdown] slideshow={"slide_type": "fragment"}
# List current conda virtual environments:
# -
conda env list
# + [markdown] slideshow={"slide_type": "fragment"}
# Create a `Python` 3 environment named `python3`:
# -
conda create -n python3 python=3
# + [markdown] slideshow={"slide_type": "fragment"}
# Install Python package with conda:
# -
conda install -n python3 numpy
# > __Note:__ If package fails to be found, search on the [Anaconda Website](https://anaconda.org/). After searching click on one of the results and the command for installing will be provided. Remember to add your `-n python3` environment name.
#
# <hr style='clear:both;'>
# ## Software: Management
#
# __Conda__
# After the conda environment is setup and `numpy` is installed, we can test it with the following:
conda activate python3
python -c 'import numpy as np; a = np.arange(15).reshape(3, 5); print(a)'
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Management
#
# __Singularity__
#
# > __Warning:__ This is a demo, should be used for advanced projects
#
# You may need a singularity image if...
#
# * You may want to build/control your own Linux environment
# * Your software requires older, or newer, libraries
# * Installation instructions are for `Ubuntu`
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software: Management
#
# __Singularity__
#
# First you must get your own Linux machine, and install Singularity.
# Perhaps the easiest way to do this is mentioned [here](https://sylabs.io/guides/3.7/admin-guide/installation.html#installation-on-windows-or-mac).
# -
# After this you can use __pre-built__ images or try to build a __custom__ singularity image:
#
# __Pre-Built__
#
# ```
# singularity exec docker://ubuntu:latest echo "Hello Dinosaur!"
# ```
#
# __Custom__
#
# 1. Create a Singularity definition file
# 2. Build container image based on definition file
# 3. Run shell inside image to test
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software Management
#
# __Definition File__
#
# Make file `myLinuxEnv.def` with the following content:
#
# ```
# bootstrap: docker
# From: ubuntu:latest
#
# # # %post
# apt update
# apt install httpd
# ```
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software Management
#
# __Build Container Image__
#
# Run the following command using defenition file:
# -
singularity build myLinuxEnv.sing myLinuxEnv.def
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Software Management
#
# __Test__
#
# Test the image buy going inside it:
# -
singularity shell myLinuxEnv.sing
# Once the `Singularity` image is tested, transfer it to the cluster (SCP/SFTP), and execute it within a job like so:
module load singularity
singularity exec myLinuxEnv.sing 'cat /etc/lsb-release'
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Slurm
#
# <img src="https://docs.google.com/drawings/d/e/2PACX-1vQWU7EGfVNGIhebu953CqTx3y-jufY-0ja6zcV65LN3KWLX5hBY7R2mEavvy34Gbq9fnDQeT80jEqfT/pub?w=933&h=401">
#
# [https://slurm.schedmd.com/archive/slurm-19.05.0/](https://slurm.schedmd.com/archive/slurm-19.05.0/)
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Node
#
# __What is a Compute Node?__
#
# <img src="https://docs.google.com/drawings/d/e/2PACX-1vQuTFQYXJmcIXx4873q1TlH0-44-yf1GwZicu6t7l5UrVnww08cMxD_ubYc0mpkfZ0Gsku43TT90DY0/pub?w=941&h=250">
#
# [https://hpcc.ucr.edu/hardware](https://hpcc.ucr.edu/hardware)
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Partitions
#
# <img style="float:right;" width='350px' src="https://docs.google.com/drawings/d/e/2PACX-1vQcl8tr-Tsi6TlUrUMREbrEk5ygkhllfoq82ZzrItDF13uqY-FmPwLpUqcpRGBTE7VajnpgDBwgox-v/pub?w=417&h=551">
#
# __What is a Partition?__
#
# Logical groups of nodes, to allow more efficient allocation and managment of resources.
#
# __Intel Partition__
#
# * CPU - 2 cores Default, 256 Cores Max
# * RAM - 1GB Default, 1TB Max
# * Time - 7 days Default, 30 Days Max
#
# <div style='clear:both'><a href='https://hpcc.ucr.edu/manuals_linux-cluster_jobs.html#partitions'>https://hpcc.ucr.edu/manuals_linux-cluster_jobs.html#partitions</a></div>
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Partitions
#
#
# * __Default?__
#
# Fallback to this value if not explicitly provided.
#
#
# * __Maximum?__
#
# Upper limit of what can be requested.
#
#
# For more details regarding our partitions, please review our [Cluster Jobs: Partitions](https://hpcc.ucr.edu/manuals_linux-cluster_jobs#partitions) manual page.
#
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Status
# + [markdown] slideshow={"slide_type": "fragment"}
# List all jobs owned by you and status:
# + slideshow={"slide_type": "-"}
squeue -u $USER
# + [markdown] slideshow={"slide_type": "fragment"}
# List all group jobs and status:
# + slideshow={"slide_type": "-"}
squeue -A $GROUP
# -
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Limits
# + [markdown] slideshow={"slide_type": "fragment"}
# List current Slurm limits:
# + slideshow={"slide_type": "-"}
slurm_limits
# + [markdown] slideshow={"slide_type": "fragment"}
# List CPUs currently used by you:
# + slideshow={"slide_type": "-"}
user_cpus
# + [markdown] slideshow={"slide_type": "fragment"}
# List CPUs currently used by entire group (primary):
# + slideshow={"slide_type": "-"}
group_cpus
# + [markdown] slideshow={"slide_type": "-"}
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Jobs
#
# __Submission__
# + [markdown] slideshow={"slide_type": "fragment"}
# Move into workshop directory:
# + slideshow={"slide_type": "-"}
cd ~/workshop_dir
# + [markdown] slideshow={"slide_type": "fragment"}
# Download example job submission script:
# + slideshow={"slide_type": "-"}
# Non-Stats
wget -O basic_job.sh https://bit.ly/33rozLX
# Stats Department
wget -O basic_job.sh https://bit.ly/2KBaIOs
# + [markdown] slideshow={"slide_type": "fragment"}
# Check job submission script contents (use arrow keys to navigate and `ctrl+x` to quit):
# + slideshow={"slide_type": "-"}
nano basic_job.sh
# -
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Jobs
#
# __Submission__
# + [markdown] slideshow={"slide_type": "fragment"}
# Submit as non-interactive job:
# + slideshow={"slide_type": "-"}
sbatch basic_job.sh
# + [markdown] slideshow={"slide_type": "fragment"}
# Submit interactive job:
# + [markdown] slideshow={"slide_type": "-"}
# ```bash
# srun -p short --pty bash -l
#
# # OR
#
# srun -p statsdept --pty bash -l
# ```
# -
# <hr style='clear:both;'>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Job Scheduling: Jobs
#
# __Status__
# + [markdown] slideshow={"slide_type": "fragment"}
# Check job status:
# -
squeue -u $USER
# + [markdown] slideshow={"slide_type": "fragment"}
# Check results:
# + slideshow={"slide_type": "-"}
cat slurm-2909103.out
# + [markdown] slideshow={"slide_type": "-"}
# [https://hpcc.ucr.edu/manuals_linux-cluster_jobs.html#submitting-jobs](https://hpcc.ucr.edu/manuals_linux-cluster_jobs.html#submitting-jobs)
#
# <hr style='clear:both;'>
| static/presentations/2020-12-18_Workshop/hpcc_usage/Cluster_Usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/ksachdeva/rethinking-tensorflow-probability/blob/master/notebooks/14_adventures_in_covariance.ipynb)
# # Chapter 14 - Adventures in Covariance
#
# ## Imports and utility functions
#
# +
# Install packages that are not installed in colab
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
# %tensorflow_version 2.X
# !pip install watermark
# !pip install arviz
USE_NIGHTLY_TFP = True # @param
if IN_COLAB and USE_NIGHTLY_TFP:
# !pip install --upgrade tf-nightly
# !pip install --upgrade tfp-nightly
# -
# %load_ext watermark
# +
# Core
import numpy as np
import arviz as az
import pandas as pd
import xarray as xr
import tensorflow as tf
import tensorflow_probability as tfp
# visualization
import matplotlib.pyplot as plt
from matplotlib.patches import Ellipse, transforms
# aliases
tfd = tfp.distributions
tfb = tfp.bijectors
Root = tfd.JointDistributionCoroutine.Root
# -
# %watermark -p numpy,tensorflow,tensorflow_probability,arviz,scipy,pandas
# config of various plotting libraries
# %config InlineBackend.figure_format = 'retina'
az.style.use('arviz-darkgrid')
if not USE_NIGHTLY_TFP:
assert tf.__version__ >= '2.1.0', "Tensorflow version should be at minimum 2.1.0"
assert tfp.__version__ >= '0.9.0', "TFP version should be at minimum 0.9.0"
# ## Tensorflow MCMC Sampling helpers
#
USE_XLA = False
# +
NUMBER_OF_CHAINS = 2
NUMBER_OF_BURNIN = 500
NUMBER_OF_SAMPLES = 500
NUMBER_OF_LEAPFROG_STEPS = 4
def _trace_to_arviz(trace=None,
sample_stats=None,
observed_data=None,
prior_predictive=None,
posterior_predictive=None,
inplace=True):
if trace is not None and isinstance(trace, dict):
trace = {k: np.swapaxes(v.numpy(), 1, 0)
for k, v in trace.items()}
if sample_stats is not None and isinstance(sample_stats, dict):
sample_stats = {k: v.numpy().T for k, v in sample_stats.items()}
if prior_predictive is not None and isinstance(prior_predictive, dict):
prior_predictive = {k: v[np.newaxis]
for k, v in prior_predictive.items()}
if posterior_predictive is not None and isinstance(posterior_predictive, dict):
if isinstance(trace, az.InferenceData) and inplace == True:
return trace + az.from_dict(posterior_predictive=posterior_predictive)
else:
trace = None
return az.from_dict(
posterior=trace,
sample_stats=sample_stats,
prior_predictive=prior_predictive,
posterior_predictive=posterior_predictive,
observed_data=observed_data,
)
@tf.function(autograph=False, experimental_compile=USE_XLA)
def run_chain(init_state,
bijectors,
step_size,
target_log_prob_fn,
num_leapfrog_steps=NUMBER_OF_LEAPFROG_STEPS,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN,
):
def _trace_fn_transitioned(_, pkr):
return (
pkr.inner_results.inner_results.log_accept_ratio
)
hmc_kernel = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn,
num_leapfrog_steps=num_leapfrog_steps,
step_size=step_size)
inner_kernel = tfp.mcmc.TransformedTransitionKernel(
inner_kernel=hmc_kernel,
bijector=bijectors)
kernel = tfp.mcmc.SimpleStepSizeAdaptation(
inner_kernel=inner_kernel,
target_accept_prob=.8,
num_adaptation_steps=int(0.8*burnin),
log_accept_prob_getter_fn=lambda pkr: pkr.inner_results.log_accept_ratio
)
results, sampler_stat = tfp.mcmc.sample_chain(
num_results=num_samples,
num_burnin_steps=burnin,
current_state=init_state,
kernel=kernel,
trace_fn=_trace_fn_transitioned)
return results, sampler_stat
def sample_posterior(jdc,
observed_data,
params,
num_chains=NUMBER_OF_CHAINS,
init_state=None,
bijectors=None,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN):
if init_state is None:
init_state = list(jdc.sample(NUMBER_OF_CHAINS)[:-1])
if bijectors is None:
bijectors = [tfb.Identity() for i in init_state]
target_log_prob_fn = lambda *x: jdc.log_prob(x + observed_data)
step_size = 0.1
results, sample_stats = run_chain(init_state,
bijectors,
step_size=step_size,
target_log_prob_fn=target_log_prob_fn,
num_samples=num_samples,
burnin=burnin)
stat_names = ['mean_tree_accept']
sampler_stats = dict(zip(stat_names, [sample_stats]))
posterior = dict(zip(params, results))
return _trace_to_arviz(trace=posterior, sample_stats=sampler_stats)
# -
# ## Dataset URLs
#
# +
# You could change base url to local dir or a remoate raw github content
_BASE_URL = "https://raw.githubusercontent.com/ksachdeva/rethinking-tensorflow-probability/master/data"
CHIMPANZEE_DATASET_PATH = f"{_BASE_URL}/chimpanzees.csv"
KOSTER_LECKIE_DATASET_PATH = f"{_BASE_URL}/KosterLeckie.csv"
ISLANDS_DISTMATRIX_DATASET_PATH = f"{_BASE_URL}/islandsDistMatrix.csv"
KLINE_DATASET_PATH = f"{_BASE_URL}/Kline2.csv"
PRIMATES301_DATASET_PATH = f"{_BASE_URL}/Primates301.csv"
PRIMATES301_VCOV_DATASET_PATH = f"{_BASE_URL}/Primates301_vcov_matrix.csv"
PRIMATES301_DISTANCE_DATASET_PATH = f"{_BASE_URL}/Primates301_distance_matrix.csv"
# -
# ## Code 14.1
#
# Simulate the population
a = 3.5 # average morning wait time
b = -1 # average difference afternoon wait time
sigma_a = 1 # std dev in intercepts
sigma_b = 0.5 # std dev in slopes
rho = -0.7 # correlation between intercepts and slopes
# ## Code 14.2
Mu = tf.constant([a, b])
# The value in a is the mean intercept, the wait in the morning. And the value in b is the mean slope, the difference in wait between afternoon and morning
# ## Code 14.3
cov_ab = sigma_a * sigma_b * rho
Sigma = tf.constant([[sigma_a ** 2, cov_ab], [cov_ab, sigma_b ** 2]])
# ## Code 14.4
tf.transpose(tf.reshape(tf.constant([1, 2, 3, 4]), (2, 2)))
# ## Code 14.5
# +
sigmas = tf.constant([sigma_a, sigma_b]) # standard deviations
Rho = tf.constant([[1, rho], [rho, 1]]) # correlation matrix
# now matrix multiply to get covariance matrix
Sigma = tf.linalg.tensor_diag(sigmas) @ Rho @ tf.linalg.tensor_diag(sigmas)
Sigma
# -
# ## Code 14.6
N_cafes = 20
# ## Code 14.7
# +
def build_vary_effects():
_seed = 5
tf.random.set_seed(_seed)
seed = tfp.util.SeedStream(_seed, salt="vary_effects")
Mu = tf.constant([a, b])
vary_effects = tfd.MultivariateNormalFullCovariance(
loc=Mu,
covariance_matrix=Sigma).sample((N_cafes,), seed=seed())
return vary_effects
vary_effects = build_vary_effects()
# -
# ## Code 14.8
a_cafe = vary_effects[:, 0]
b_cafe = vary_effects[:, 1]
# ## Code 14.9
# +
plt.plot(a_cafe, b_cafe, "o", mfc="none")
plt.gca().set(xlabel="intercepts (a_cafe)", ylabel="slopes (b_cafe)")
plt.xlim((1.0,5.5))
plt.ylim((-1.8,-0.25))
# overlay population distribution
# Ref: https://matplotlib.org/gallery/statistics/confidence_ellipse.html
for l in [0.1, 0.3, 0.5, 0.8, 0.99]:
pearson = Sigma[0, 1] / np.sqrt(Sigma[0, 0] * Sigma[1, 1])
ellipse = Ellipse((0, 0), np.sqrt(1 + pearson), np.sqrt(1 - pearson),
edgecolor="k", alpha=0.2, facecolor="none")
std_dev = tfd.Normal(loc=0.,scale=1.).quantile((1 + np.sqrt(l)) / 2)
scale_x = 2 * std_dev * np.sqrt(Sigma[0, 0])
scale_y = 2 * std_dev * np.sqrt(Sigma[1, 1])
scale = transforms.Affine2D().rotate_deg(45).scale(scale_x, scale_y)
ellipse.set_transform(scale.translate(Mu[0], Mu[1]) + plt.gca().transData)
plt.gca().add_patch(ellipse)
# -
# ## Code 14.10
#
# The data we’re generating describes the waiting times in 20 different cafés. Each café has a different average waiting times in the morning and in the afternoon. The average morning waiting time is the intercept, and the difference between afternoon and morning average waiting times is the slope.
# +
N_visits = 10
afternoon = np.tile(np.arange(2), N_visits * N_cafes // 2)
cafe_id = np.repeat(np.arange(N_cafes), N_visits)
def generate_data_frame():
sigma = 0.5 # std dev within cafes
_seed = 22
tf.random.set_seed(_seed)
seed = tfp.util.SeedStream(_seed, salt="generate_data_frame")
mu = tf.gather(a_cafe, cafe_id) + tf.gather(b_cafe, cafe_id) * afternoon
wait = tfd.Normal(loc=mu, scale=sigma).sample(seed=seed())
d = pd.DataFrame(dict(cafe=cafe_id, afternoon=afternoon, wait=wait))
return d
d = generate_data_frame()
d.describe()
# -
# ## Code 14.11
R = tfp.distributions.LKJ(dimension=2, concentration=2).sample((int(1e4),))
az.plot_kde(R[:, 0, 1], bw=2, label="correlation");
R.shape
# ## Code 14.12
# +
def model_14_1(cafe, afternoon):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=5.,scale=2.), sample_shape=1))
beta = yield Root(tfd.Sample(tfd.Normal(loc=-1., scale=0.5), sample_shape=1))
# sigma = yield Root(tfd.Sample(tfd.HalfCauchy(loc=0., scale=1.), sample_shape=1))
# sigma_alpha_beta = yield Root(tfd.Sample(tfd.HalfCauchy(loc=0., scale=1.), sample_shape=2))
sigma = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=1))
sigma_alpha_beta = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=2))
# Rho = yield Root(tfd.Sample(tfd.LKJ(dimension=2, concentration=2.), sample_shape=1))
Rho = yield Root(tfd.LKJ(dimension=2, concentration=2.))
Mu = tf.concat([alpha, beta], axis=-1)
scale = tf.linalg.LinearOperatorDiag(sigma_alpha_beta).matmul(tf.squeeze(Rho))
a_cafe_b_cafe = yield tfd.Sample(
tfd.MultivariateNormalTriL(
loc = Mu,
scale_tril = scale
),sample_shape=20)
# extract the alphas and betas
a_cafe = tf.gather(a_cafe_b_cafe, 0, axis=-1)
b_cafe = tf.gather(a_cafe_b_cafe, 1, axis=-1)
# extract alphas and beta using the cafe id as the index
term1 = tf.gather(a_cafe, cafe, axis=-1)
term2 = tf.gather(b_cafe, cafe, axis=-1)
# linear model
mu = term1 + term2 * afternoon
wait = yield tfd.Independent(tfd.Normal(loc=mu, scale=sigma), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_14_1 = model_14_1(tf.cast(d.cafe.values, dtype=tf.int32), tf.cast(d.afternoon.values, dtype=tf.float32))
# +
alpha_init, beta_init, sigma_init, sigma_alpha_beta_init, Rho_init, a_cafe_b_cafe_init, _ = jdc_14_1.sample(2)
# let's see Rho and make sure that what we have
# looks like a correlation matrix as a sanity check
#
# Now this will a prior sample so most likely
# we will get the correct sample however when we will do HMC
# we maybe extra care
# alpha_init, beta_init, sigma_init, sigma_alpha_beta_init, Rho_init, a_cafe_b_cafe_init
Rho_init
# +
init_rho = tf.stack([tf.eye(2) for _ in range(2)])
init_rho
# +
init_state = [
alpha_init,
beta_init,
sigma_init,
sigma_alpha_beta_init,
init_rho,
a_cafe_b_cafe_init
]
bijectors = [
tfb.Identity(), # alpha
tfb.Identity(), # beta
tfb.Identity(), # sigma
tfb.Identity(), # sigma_alpha_beta
tfb.CorrelationCholesky(), # Rho
tfb.Identity(), # a_cafe_b_cafe
]
observed_data = (tf.cast(d.wait.values, dtype=tf.float32),)
trace_14_1 = sample_posterior(
jdc_14_1,
observed_data=observed_data,
params=['alpha', 'beta', 'sigma', 'sigma_alpha_beta', 'Rho', 'a_cafe_b_cafe'],
num_samples=4000,
burnin=2000,
init_state=init_state,
bijectors=bijectors
)
# -
# ## Code 14.13
# +
post = trace_14_1.posterior
# Here comes the important part !
#
# The posterior that we get is not really Rho because of the bijector
# that we used. Hence we need to get the correlation matrix back (i.e. Rho)
states = post["Rho"].values
rhos = states @ tf.transpose(states,[0,1,3,2])
# let's look at the rhos
# we should see that it will be a collection of 2x2 matrices such that on diagnoal you would
# see the ones
rhos
# -
rhos[0]
rhos[1][:,0,1]
az.plot_kde(rhos[1][:,0,1], bw=2);
# ## Code 14.14
# +
a1 = np.array([np.mean(d.wait[(cafe_id == i) & (afternoon == 0)])
for i in range(N_cafes)])
b1 = np.array([np.mean(d.wait[(cafe_id == i) & (afternoon == 1)])
for i in range(N_cafes)]) - a1
# extract posterior means of partially pooled estimates
a2 = np.mean(post["a_cafe_b_cafe"].values[0][..., 0], 0)
b2 = np.mean(post["a_cafe_b_cafe"].values[0][..., 1], 0)
# plot both and connect with lines
plt.plot(a1, b1, "o")
plt.gca().set(xlabel="intercept", ylabel="slope",
ylim=(np.min(b1) - 0.1, np.max(b1) + 0.1),
xlim=(np.min(a1) - 0.1, np.max(a1) + 0.1))
plt.plot(a2, b2, "ko", mfc="none")
for i in range(N_cafes):
plt.plot([a1[i], a2[i]], [b1[i], b2[i]], "k", lw=0.5)
fig, ax = plt.gcf(), plt.gca()
# -
# ## Code 14.15
# +
# compute posterior mean bivariate Gaussian
Mu_est = np.array([np.mean(post["alpha"].values[0]), np.mean(post["beta"].values[0])])
rho_est = np.mean(post["Rho"][0, 1])
sa_est = np.mean(post["sigma"][:, 0])
sb_est = np.mean(post["sigma"][:, 1])
cov_ab = sa_est * sb_est * rho_est
Sigma_est = np.array([[sa_est ** 2, cov_ab], [cov_ab, sb_est ** 2]])
# draw contours
for l in [0.1, 0.3, 0.5, 0.8, 0.99]:
pearson = Sigma_est[0, 1] / np.sqrt(Sigma_est[0, 0] * Sigma_est[1, 1])
ellipse = Ellipse((0, 0), np.sqrt(1 + pearson), np.sqrt(1 - pearson),
edgecolor="k", alpha=0.2, facecolor="none")
std_dev = tfd.Normal(loc=0.,scale=1.).quantile((1 + np.sqrt(l)) / 2)
scale_x = 2 * std_dev * np.sqrt(Sigma_est[0, 0])
scale_y = 2 * std_dev * np.sqrt(Sigma_est[1, 1])
scale = transforms.Affine2D().rotate_deg(45).scale(scale_x, scale_y)
ellipse.set_transform(scale.translate(Mu_est[0], Mu_est[1]) + ax.transData)
ax.add_patch(ellipse)
fig
# -
# ## Code 14.16
# +
# convert varying effects to waiting times
wait_morning_1 = a1
wait_afternoon_1 = a1 + b1
wait_morning_2 = a2
wait_afternoon_2 = a2 + b2
# plot both and connect with lines
plt.plot(wait_morning_1, wait_afternoon_1, "o")
plt.gca().set(
xlabel="morning wait", ylabel="afternoon wait",
ylim=(np.min(wait_afternoon_1) - 0.1, np.max(wait_afternoon_1) + 0.1),
xlim=(np.min(wait_morning_1) - 0.1, np.max(wait_morning_1) + 0.1))
plt.plot(wait_morning_2, wait_afternoon_2, "ko", mfc="none")
for i in range(N_cafes):
plt.plot([wait_morning_1[i], wait_morning_2[i]],
[wait_afternoon_1[i], wait_afternoon_2[i]], "k", lw=0.5)
x = np.linspace(np.min(wait_morning_1), np.max(wait_morning_1), 101)
plt.plot(x, x, "k--", lw=1)
fig, ax = plt.gcf(), plt.gca()
# -
# ## Code 14.17
# +
# now shrinkage distribution by simulation
v = tfd.MultivariateNormalFullCovariance(loc=Mu_est, covariance_matrix=Sigma_est).sample((10000,)).numpy()
v[:,1] = v[:,0] + v[:,1] # calculate afternoon wait
Sigma_est2 = np.cov(v, rowvar=False)
Mu_est2 = Mu_est
Mu_est2[1] = Mu_est2[0] + Mu_est2[1]
# draw contours
for l in [0.1, 0.3, 0.5, 0.8, 0.99]:
pearson = Sigma_est2[0, 1] / np.sqrt(Sigma_est2[0, 0] * Sigma_est2[1, 1])
ellipse = Ellipse((0, 0), np.sqrt(1 + pearson), np.sqrt(1 - pearson),
edgecolor="k", alpha=0.2, facecolor="none")
std_dev = tfd.Normal(loc=0.,scale=1.).quantile((1 + np.sqrt(l)) / 2)
scale_x = 2 * std_dev * np.sqrt(Sigma_est2[0, 0])
scale_y = 2 * std_dev * np.sqrt(Sigma_est2[1, 1])
scale = transforms.Affine2D().rotate_deg(45).scale(scale_x, scale_y)
ellipse.set_transform(
scale.translate(Mu_est2[0], Mu_est2[1]) + ax.transData)
ax.add_patch(ellipse)
fig
# -
# ## Code 14.18
# +
d = pd.read_csv(CHIMPANZEE_DATASET_PATH, sep=";")
d["block_id"] = d.block
d["treatment"] = 1 + d.prosoc_left + 2 * d.condition
dat = dict(L=d.pulled_left.values, tid=d.treatment.values - 1,
actor=d.actor.values - 1, block_id=d.block_id.values - 1)
# actor 0 to 6 => 7
# tid 0 to 3 => 4
# +
def model_14_2(tid, actor, block_id, num_chains):
# we need to create the indexes that make
# gather_nd happy
actor_tid = tf.stack([actor, tid], axis=1)
block_tid = tf.stack([block_id, tid], axis=1)
batch_dims = 0
if num_chains > 1:
actor_tid_tiled = tf.tile(actor_tid, (num_chains, 1))
actor_tid_reshaped = tf.reshape(actor_tid_tiled, (num_chains, 504, 2))
actor_tid = actor_tid_reshaped
block_tid_tiled = tf.tile(block_tid, (num_chains, 1))
block_tid_reshaped = tf.reshape(block_tid_tiled, (num_chains, 504, 2))
block_tid = block_tid_reshaped
batch_dims = 1
def _generator():
# fixed priors
g = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=1.), sample_shape=4))
sigma_actor = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=4))
# note - not using Sample as it does not work with it
rho_actor = yield Root(tfp.distributions.LKJ(dimension=4, concentration=4.))
sigma_block = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=4))
# note - not using Sample as it does not work with it
rho_block = yield Root(tfp.distributions.LKJ(dimension=4, concentration=4.))
# adaptive priors
alpha = yield tfd.Sample(
tfd.MultivariateNormalTriL(
loc = 0.,
scale_tril = tf.linalg.LinearOperatorDiag(sigma_actor).matmul(rho_actor)
),sample_shape=7)
beta = yield tfd.Sample(
tfd.MultivariateNormalTriL(
loc = 0.,
scale_tril = tf.linalg.LinearOperatorDiag(sigma_block).matmul(rho_block)
),sample_shape=6)
print(alpha.shape)
term1 = tf.gather(g, tid, axis=-1)
term2 = tf.gather_nd(alpha, actor_tid, batch_dims=batch_dims)
term3 = tf.gather_nd(alpha, block_tid, batch_dims=batch_dims)
logit_p = term1 + term2 + term3
L = yield tfd.Independent(tfd.Binomial(total_count=1, logits=logit_p), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=False)
jdc_14_2 = model_14_2(dat["tid"], dat["actor"], dat["block_id"], num_chains=1)
jdc_14_2.sample()
# -
g_init, sigma_actor_init, rho_actor_init, sigma_block_init, rho_block_init, alpha_init, beta_init, _ = jdc_14_2.sample()
# +
init_state = [
g_init,
sigma_actor_init,
rho_actor_init,
sigma_block_init,
rho_block_init,
alpha_init,
beta_init,
]
bijectors = [
tfb.Identity(),
tfb.Exp(),
tfb.CorrelationCholesky(),
tfb.Exp(),
tfb.CorrelationCholesky(),
tfb.Identity(),
tfb.Identity()
]
observed_data = (tf.cast(dat["L"], dtype=tf.float32),)
trace_14_2 = sample_posterior(
jdc_14_2,
observed_data=observed_data,
params=['g', 'sigma_actor', 'rho_actor', 'sigma_block', 'rho_block', 'alpha', 'beta'],
num_samples=2000,
burnin=500,
init_state=init_state,
bijectors=bijectors)
# -
# What is happening is that arviz assigns 7 to the chain dimension and this screws up the
# entire structure in xarray. Really arviz does not work if there is 1 chain.
#
# Now I have problem using 2 chains so that is the real problem indeed !
trace_14_2.posterior
az.summary(trace_14_2)
| notebooks/14_adventures_in_covariance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Import scikit-learn dataset library
from sklearn import datasets
import pandas as pd
#Load dataset
iris = datasets.load_iris()
# +
# print the label species(setosa, versicolor,virginica)
print(iris.target_names)
# print the names of the four features
print(iris.feature_names)
# +
# print the iris data (top 5 records)
print(iris.data[0:5])
# print the iris labels (0:setosa, 1:versicolor, 2:virginica)
print(iris.target)
# -
print(iris.data[0:5])
#[x axis, y axis]
print(iris.data[0:5,0])
print(iris.data[0:5,1])
print(iris.data[0:5,2])
print(iris.data[0:5,3])
print(iris.data[0,0:3])
print(iris.data[0:5,0:5])
# Creating a DataFrame of given iris dataset.
data=pd.DataFrame({
'sepal length':iris.data[:,0],
'sepal width':iris.data[:,1],
'petal length':iris.data[:,2],
'petal width':iris.data[:,3],
'species':iris.target
})
data.head()
# +
# Import train_test_split function
from sklearn.model_selection import train_test_split
print(data.head())
X = data[['sepal length', 'sepal width', 'petal length', 'petal width']] # Features
y = data['species'] # Labels
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3) # 70% training and 30% test
# +
#Import Random Forest Model
from sklearn.ensemble import RandomForestClassifier
#Create a Gaussian Classifier, n_estimators is the amount of decision trees created
clf = RandomForestClassifier(n_estimators=1000, n_jobs=-1)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
# -
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
clf.predict([[3, 5, 4, 2]])
| AIC/random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Premier league: How has VAR impacted the rankings?
#
# There has been much debate about the video assistant referee (VAR) when it was introduced last year (in 2019).
# The goal is to lead to fairer refereeing, but concerns are high on whether this will really be the case and the fact that it could break the rythm of the game.
#
# We will let football analysts – or soccer analysts depending on where you are reading this notebook from – answer this question. But one thing we can look at is how has VAR impacted the league so far.
#
# This is what we will do in this notebook, alongside some other simulations we found interesting.
#
# <div style="text-align:center"><a href="https://www.atoti.io/?utm_source=gallery&utm_content=premier-league" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover.png" alt="atoti" /></a></div>
# ## Importing the data
#
# The data we will use is composed of events. An event can be anything that happens in a game: kick-off, goal, foul, etc.
# In this dataset, we only kept kick-off and goal events to build our analysis.
# Note that in the goal events we also have all the goals that were later cancelled by VAR during a game.
#
# We will first start by importing atoti and creating a session.
# +
import atoti as tt
session = tt.create_session()
# -
# Then load the events in a store
events = session.read_csv(
"s3://data.atoti.io/notebooks/premier-league/events.csv",
separator=";",
table_name="events",
)
events.head()
# ### Creating a cube
#
# We create a cube on the event store so that some matches or teams that ended with no goal will still be reflected in the pivot tables.
#
# When creating a cube in the default auto mode, a hierarchy will be created for each non float column, and average and sum measures for each float column. This setup can later be edited, or you could also define all hierarchies/measures by yourself switching to manual mode.
cube = session.create_cube(events)
cube.schema
# Let's assign measures/levels/hierarchies to shorter variables
m = cube.measures
lvl = cube.levels
h = cube.hierarchies
h["Day"] = [events["Day"]]
# ## Computing the rankings from the goals
#
# Computing the first measure below to count the total goals scored for each event. At this point the total still includes the potential own goals and VAR-refused goals.
m["Team Goals (incl Own Goals)"] = tt.agg.sum(
tt.where(lvl["EventType"] == "Goal", tt.agg.count_distinct(events["EventId"]), 0.0),
scope=tt.scope.origin(lvl["EventType"]),
)
# In this data format, own goals are scored by players from a Team, but those points should be attributed to the opponent. Therefore we will isolate the own goals in a separate measure.
m["Team Own Goals"] = tt.agg.sum(
tt.where(lvl["IsOwnGoal"] == True, m["Team Goals (incl Own Goals)"], 0.0),
scope=tt.scope.origin(lvl["IsOwnGoal"]),
)
# And deduce the actual goals scored for the team
m["Team Goals"] = m["Team Goals (incl Own Goals)"] - m["Team Own Goals"]
# At this point we can already have a look at the goals per team. By right clicking on the chart we have sorted it descending by team goals.
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["ALL_MEASURES"], "values": ["[Measures].[Team Goals]", "[Measures].[Team Own Goals]"], "verticalSubplots": [], "xAxis": ["[events].[Team].[Team]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[Team Goals], [Measures].[Team Own Goals]} ON COLUMNS, NON EMPTY Order(Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), [Measures].[Team Goals], BDESC) ON ROWS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-clustered-column-chart"}} tags=[]
session.visualize()
# -
# For a particular match, the `Opponent Goals` are equal to the `Team Goals` if we switch to the data facts where Team is replaced by Opponent and Opponent by Team
m["Opponent Goals"] = tt.agg.sum(
tt.at(
m["Team Goals"],
{lvl["Team"]: lvl["Opponent"], lvl["Opponent"]: lvl["Team"]},
),
scope=tt.scope.origin(lvl["Team"], lvl["Opponent"]),
)
m["Opponent Own Goals"] = tt.agg.sum(
tt.at(
m["Team Own Goals"],
{lvl["Team"]: lvl["Opponent"], lvl["Opponent"]: lvl["Team"]},
),
scope=tt.scope.origin(lvl["Team"], lvl["Opponent"]),
)
# We are now going to add two measures `Team Score` and `Opponent Score` to compute the result of a particular match.
m["Team Score"] = m["Team Goals"] + m["Opponent Own Goals"]
m["Opponent Score"] = m["Opponent Goals"] + m["Team Own Goals"]
# We can now visualize the result of each match of the season
# + atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[Team Score]", "[Measures].[Opponent Score]"], "rows": ["[events].[Day].[Day] => [events].[Team].[Team] => [events].[Opponent].[Opponent]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[Team Score], [Measures].[Opponent Score]} ON COLUMNS, NON EMPTY Hierarchize(Union(Crossjoin(Descendants({[events].[Day].[AllMember]}, 1, SELF_AND_BEFORE), [events].[Team].DefaultMember, [events].[Opponent].DefaultMember), Crossjoin([events].[Day].[AllMember].[1], Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), [events].[Opponent].DefaultMember), Crossjoin([events].[Day].[AllMember].[1], [events].[Team].[AllMember].[Arsenal], Hierarchize(Descendants({[events].[Opponent].[AllMember]}, 1, SELF_AND_BEFORE))), Crossjoin([events].[Day].[AllMember].[1], [events].[Team].[AllMember].[Aston Villa], Hierarchize(Descendants({[events].[Opponent].[AllMember]}, 1, SELF_AND_BEFORE))), Crossjoin([events].[Day].[AllMember].[1], [events].[Team].[AllMember].[Bournemouth], Hierarchize(Descendants({[events].[Opponent].[AllMember]}, 1, SELF_AND_BEFORE))))) ON ROWS FROM [events]", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# We now have the team goals/score and those of the opponent for each match. However, these measures include VAR cancelled goals. Let's create new measures that takes into account VAR.
m["VAR team goals impact"] = m["Team Goals"] - tt.filter(
m["Team Goals"], lvl["IsCancelledAfterVAR"] == False
)
m["VAR opponent goals impact"] = m["Opponent Goals"] - tt.filter(
m["Opponent Goals"], lvl["IsCancelledAfterVAR"] == False
)
# We can visualize that in details, there are already 4 goals cancelled by VAR on the first day of the season !
# + atoti={"widget": {"columnWidths": {"[Measures].[VAR team goals impact]": 217.36248779296875}, "mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[VAR team goals impact]", "[Measures].[Team Goals]"], "rows": ["[events].[Day].[Day]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[VAR team goals impact], [Measures].[Team Goals]} ON COLUMNS, NON EMPTY Hierarchize(Descendants({[events].[Day].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# Now that for any game we have the number of goals of each team, we can compute how many points teams have earned.
# Following the FIFA World Cup points system, three points are awarded for a win, one for a draw and none for a loss (before, winners received two points).
# We create a measure for each of this condition.
m["Points for victory"] = 3.0
m["Points for tie"] = 1.0
m["Points for loss"] = 0.0
m["Points"] = tt.agg.sum(
tt.where(
m["Team Score"] > m["Opponent Score"],
m["Points for victory"],
tt.where(
m["Team Score"] == m["Opponent Score"],
m["Points for tie"],
m["Points for loss"],
),
),
scope=tt.scope.origin(lvl["League"], lvl["Day"], lvl["Team"]),
)
# The previous points were computed including VAR-refused goals.
# Filtering out these goals gives the actual rankings of the teams, as you would find on any sports websites.
m["Actual Points"] = tt.filter(m["Points"], lvl["IsCancelledAfterVAR"] == False)
# And here we have our ranking. We will dive into it in the next section.
#
# ## Rankings and VAR impact
#
# Color rules were added to show teams that benefited from the VAR in green and those who lost championship points because of it in red.
m["Difference in points"] = m["Actual Points"] - m["Points"]
# + atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[Points]", "[Measures].[Actual Points]", "[Measures].[Difference in points]"], "rows": ["[events].[Team].[Team]"]}, "query": {"mdx": "SELECT NON EMPTY Order(Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), [Measures].[Actual Points], DESC) ON ROWS, NON EMPTY {[Measures].[Points], [Measures].[Actual Points], [Measures].[Difference in points]} ON COLUMNS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# More than half of the teams have had their points total impacted by VAR.
# Though it does not impact the top teams, it definitely has an impact in the ranking of many teams, Manchester United would have lost 2 ranks and Tottenham 4 for example!
#
# We could also visualize the difference of points in a more graphical way:
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["ALL_MEASURES"], "values": ["[Measures].[Points]", "[Measures].[Actual Points]"], "verticalSubplots": [], "xAxis": ["[events].[Team].[Team]"]}, "query": {"mdx": "SELECT NON EMPTY {[Measures].[Points], [Measures].[Actual Points]} ON COLUMNS, NON EMPTY Order(Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), [Measures].[Actual Points], BDESC) ON ROWS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}} tags=[]
session.visualize()
# -
# Since the rankings are computed from the goal level, we can perform any kind of simulation we want using simple UI filters.
# You can filter the pivot table above to see what would happen if we only keep the first half of the games? If we only keep matches played home? What if we filter out Vardy, would Leicester lose some places?
# Note that if you filter out VAR-refused goals, the `Points` measures takes the same value as the `Actual Points`.
#
# ## Evolution of the rankings over time
#
# Atoti also enables you to define cumulative sums over a hierarchy, we will use that to see how the team rankings evolved during the season.
m["Points cumulative sum"] = tt.agg.sum(
m["Actual Points"], scope=tt.scope.cumulative(lvl["Day"])
)
# + atoti={"widget": {"mapping": {"horizontalSubplots": [], "splitBy": ["[events].[Team].[Team]", "ALL_MEASURES"], "values": ["[Measures].[Points cumulative sum]"], "verticalSubplots": [], "xAxis": ["[events].[Day].[Day]"]}, "query": {"mdx": "SELECT NON EMPTY Hierarchize(Descendants({[events].[Day].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS, NON EMPTY Crossjoin(Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), {[Measures].[Points cumulative sum]}) ON COLUMNS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}} tags=[]
session.visualize()
# -
# We can notice that data is missing for the 28th match of Manchester City. This is because the game was delayed due to weather, and then never played because of the COVID-19 pandemic.
# ## Players most impacted by the VAR
#
# Until now we looked at most results at team level, but since the data exists at goal level, we could have a look at which players are most impacted by the VAR.
m["Valid player goals"] = tt.filter(
m["Team Goals"], lvl["IsCancelledAfterVAR"] == False
)
# + atoti={"widget": {"mapping": {"columns": ["ALL_MEASURES"], "measures": ["[Measures].[Team Goals]", "[Measures].[Valid player goals]", "[Measures].[Team Goals - Valid player goals]"], "rows": ["[events].[Scorer].[Scorer]"]}, "query": {"mdx": "WITH Member [Measures].[Team Goals - Valid player goals] AS [Measures].[Team Goals] - [Measures].[Valid player goals], CAPTION = \"Team Goals - Valid player goals\" SELECT NON EMPTY Hierarchize(Descendants({[events].[Scorer].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS, NON EMPTY {[Measures].[Team Goals], [Measures].[Valid player goals], [Measures].[Team Goals - Valid player goals] } ON COLUMNS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# -
# Unsurprisingly Mané is the most impacted player. He is also one of the top scorers with only Vardy scoring more goals (you can sort on the Team Goals column to verify).
# More surprisingly, Boly has had all the goals of his season cancelled by VAR and Antonio half of them..
#
# ## Simulation of a different scoring system
#
# Although we are all used to a scoring system giving 3 points for a victory, 1 for a tie and 0 per lost match this was not always the case. Before the 1990's many european leagues only gave 2 points per victory, reason for the change being to encourage teams to score more goals during the games.
# The premier league gifts us well with plenty of goals scored (take it from someone watching the French ligue 1), but how different would the results be with the old scoring system?
#
# atoti enables us to simulate this very easily. We simply have to create a new scenario where we replace the number of points given for a victory.
# We first setup a simulation on that measure.
scoring_system_simulation = cube.create_parameter_simulation(
name="Scoring system simulations",
measures={"Points for victory": 3.0},
base_scenario_name="Current System",
)
# And create a new scenario where we give it another value
scoring_system_simulation += ("Old system", 2.0)
# And that's it, no need to define anything else, all the measures will be re-computed on demand with the new value in the new scenario.
# Let's compare the rankings between the two scoring systems.
# + atoti={"widget": {"columnWidths": {"[Measures].[Actual Points],[Scoring system simulations].[Scoring system simulations].[Current System - Old system]": 202, "[Measures].[Actual Points],[Scoring system simulations].[Scoring system simulations].[Current System]": 217.59375, "[Measures].[Actual Points],[Scoring system simulations].[Scoring system simulations].[Old system]": 178.59375}, "mapping": {"columns": ["[Scoring system simulations].[Scoring system simulations].[Scoring system simulations]", "ALL_MEASURES"], "measures": ["[Measures].[Actual Points]"], "rows": ["[events].[Scorer].[Scorer]"]}, "query": {"mdx": "WITH Member [Scoring system simulations].[Scoring system simulations].[Current System - Old system] AS [Scoring system simulations].[Scoring system simulations].[Current System] - [Scoring system simulations].[Scoring system simulations].[Old system], CAPTION = \"Current System - Old system\" SELECT NON EMPTY Crossjoin(Hierarchize(Union([Scoring system simulations].[Scoring system simulations].[Scoring system simulations].Members, [Scoring system simulations].[Scoring system simulations].[Scoring system simulations].Members, [Scoring system simulations].[Scoring system simulations].[Current System - Old system])), [Measures].[Actual Points]) ON COLUMNS, NON EMPTY Hierarchize(Descendants({[events].[Scorer].[AllMember]}, 1, SELF_AND_BEFORE)) ON ROWS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "pivot-table"}} tags=[]
session.visualize()
# + atoti={"widget": {"filters": [], "mapping": {"horizontalSubplots": [], "splitBy": ["[Scoring system simulations].[Scoring system simulations].[Scoring system simulations]", "ALL_MEASURES"], "values": ["[Measures].[Actual Points]"], "verticalSubplots": [], "xAxis": ["[events].[Team].[Team]"]}, "query": {"mdx": "SELECT NON EMPTY Order(Hierarchize(Descendants({[events].[Team].[AllMember]}, 1, SELF_AND_BEFORE)), [Measures].[Actual Points], BDESC) ON ROWS, NON EMPTY Crossjoin([Scoring system simulations].[Scoring system simulations].[Scoring system simulations].Members, {[Measures].[Actual Points]}) ON COLUMNS FROM [events] CELL PROPERTIES VALUE, FORMATTED_VALUE, BACK_COLOR, FORE_COLOR, FONT_FLAGS", "updateMode": "once"}, "serverKey": "default", "widgetKey": "plotly-line-chart"}} tags=[]
session.visualize()
# -
# Surprisingly, having only 2 points for a win would only have made Burnley and West Ham lose 2 ranks, but no other real impact on the standings.
#
# <div style="text-align:center"><a href="https://www.atoti.io/?utm_source=gallery&utm_content=premier-league" target="_blank" rel="noopener noreferrer"><img src="https://data.atoti.io/notebooks/banners/discover-try.png" alt="atoti" /></a></div>
| notebooks/var-impact-in-premier-league/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from os import path
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# +
path_name = path.join(path.abspath('..'), 'data')
file_name = 'corrective_maint_job_types.csv'
data_file = path.join(path_name, file_name)
df = pd.read_csv(data_file)
# -
df.head()
pivot = df.pivot_table('notif', index='job_type', columns='network', aggfunc='count')
#pivot
import seaborn
seaborn.countplot(df['job_type'],
palette="Blues_d",
order=df['job_type'].value_counts().iloc[:10].index
)
# +
import pandas as pd
import seaborn as sns
from datetime import date
from datetime import time
from datetime import datetime
df['notif_date'] = pd.to_datetime(df['notif_date'])
new_df = df[['notif_date','job_type']]
new_df.set_index('notif_date')
monthly_data=[]
labels = ['code','year', 'month', 'count']
for cat, subcategories in new_df.groupby('job_type'):
if cat!='SPF':
job_type = cat
for yr, year_df in subcategories.groupby(df['notif_date'].dt.year):
for mnth, month_df in year_df.groupby(df['notif_date'].dt.month):
new_data=[cat, yr, mnth, len(month_df)]
#print(new_data)
monthly_data.append(new_data)
output = pd.DataFrame.from_records(monthly_data, columns=labels)
g = sns.FacetGrid(output, col='code', aspect=1, col_wrap=4)
g.map(sns.boxplot, 'month', 'count')
# +
# add lineplot
job_type = 'SF5'
data = output.loc[output['code'] == job_type]
title = 'Seasonal Analysis - ' + job_type
plot = sns.boxplot(x='month', y='count', data=data)
#plot = sns.lineplot(x='month', y='count', data=data, color='blue').set_title(title)
# format plot
plot.axes.set_title("Seasonal Analysis - " + job_type, fontsize=16)
plot.set_xlabel("Month", fontsize=14)
plot.set_ylabel("Count", fontsize=14)
plot.tick_params(labelsize=10)
# -
data.head()
# +
monthly_output = new_df[new_df['job_type'] == 'SF5']
monthly_output['notif_date']=pd.to_datetime(monthly_output['notif_date'])
monthly_output['notif_date']=monthly_output['notif_date'].dt.strftime('%Y-%m')
monthly_output = monthly_output.pivot_table('job_type', index='notif_date', aggfunc='count')
monthly_output = monthly_output.reset_index()
# +
#monthly_output.head()
# +
#_ = monthly_output.plot(x='notif_date', y='job_type', figsize=(12,4))
# -
def AR(data, maxlag = 1):
"""
Implements Autoregression (AR)
parameters:
data: times series data
maxlag: degree of the model (1 by default)
return:
X, y and b (A hat)
"""
#linear model of order max lag 8)
#data = df['sunspots'].values
maxz = data.size #to separete test and fit data, this should be reduced
# Computing matrix X
X = np.array([data[i:(maxz-maxlag-1)+i] for i in range(0, maxlag)])
# Performing the transponse of X
X = X.T
# Computing matrix y
y = np.array(data[maxlag+1:maxz])
# Computing the Pseudo-Inverse of X (as X is a rectangular matrix)
Xinv = np.linalg.pinv(X.copy())
# Solving the linear system b = Xinv * y
b = np.dot(Xinv, y)
return (X, y, b)
def pred_local_constant_model(data, X, y, maxlag = 1, nstep = 100):
"""
Implements yp = xi * b to predict nsteps
parameters:
data: times series data
X: data representation
y: value of the function for each row in X
maxlag: degree of the model (1 by default)
nstep: number of steps (100 by default)
return:
predictions
"""
#now free-rn from a local constant model
xinit = data[-maxlag-1:-1]
xi = xinit
step = 0
yp = data[-1]
#distance function
dist = lambda x1, x2: math.sqrt(((x1-x2)**2).sum())
while step < nstep:
# 1) Compute the distance to the points xi with respect X
xdm = [dist(xi,xj) for xj in X]
# 2) Sort the distances, rather neat way of finding the second
# smallest without doing a full sort
ind = np.argpartition(xdm,2)[2]
# 3) Find the second smallest value to predict the true future
xp = y[ind]
# update xi and yp
xi = np.delete(xi,0)
xi = np.append(xi,xp)
yp = np.append(yp,xp)
# increment and repeat steps 1, 2 and 3 until step == nstep
step += 1
return yp
# play with hyperparameters
maxlag = 3
nsteps = 365 # predict the next nsteps
# +
X, y, b = AR(monthly_output['job_type'].values, maxlag)
yp = np.dot(X, b)
plt.figure(figsize=(14,8))
#plt.plot(monthly_output['notif_date'][maxlag+1:], monthly_output['job_type'][maxlag+1:], label='raw')
plt.plot(monthly_output['notif_date'][maxlag+1:], y, label='original')
plt.plot(monthly_output['notif_date'][maxlag+1:], yp, label='estimated')
plt.xlabel('date')
plt.ylabel('count')
plt.legend()
plt.show()
# +
# Visualising the Predictions
from datetime import timedelta
# get daily time series data spefic to required job type
daily_output = new_df[new_df['job_type'] == 'SF5']
daily_output = daily_output.pivot_table('job_type', index='notif_date', aggfunc='count')
daily_output = daily_output.reset_index()
#Calculate the linear autoregressive model
X, y, b = AR(daily_output['job_type'].values, maxlag)
yp2 = pred_local_constant_model(daily_output['job_type'].values, X, y, maxlag, nsteps)
# Set up dataframe to capture output data in suitable format
columns = ['notif_date', 'job_type']
days = pd.DataFrame(index=daily_output.index, columns=columns)
days['notif_date'] = daily_output['notif_date']
pd.to_datetime(days['notif_date'])
# set up temporary dataframe to capture estimated values
estimates = pd.DataFrame()
# initiate start values for loop variables
next_date = daily_output['notif_date'].values[-1]
next_index = daily_output.index[-1]+1
# determine estimates for range
for i in range(nsteps+1):
next_date = next_date + np.timedelta64(1,'D')
next_count = yp2[i]
estimate = [(next_date, next_count)]
estimates = estimates.append(estimate)
# format dataframe to be in consistent format with days dataframe
estimates.columns = columns
# Append estimated values to actual values
days = days.append(estimates, ignore_index=True)
# Aggregate daily estimates to monthly estimates
#days['notif_date'] = days['notif_date'].dt.strftime('%Y-%m')
#days = days.pivot_table('job_type', index='notif_date', aggfunc='sum')
#days = days.reset_index()
# Aggregate daily actuals to monthly actuals
#daily_output['notif_date'] = pd.to_datetime(daily_output['notif_date'])
#daily_output['notif_date'] = daily_output['notif_date'].dt.strftime('%Y-%m')
#daily_output = daily_output.pivot_table('job_type', index='notif_date', aggfunc='sum')
#daily_output = daily_output.reset_index()
# Plot actual and forecast values
plt.figure(figsize=(14,8))
plt.plot(daily_output['notif_date'], daily_output['job_type'], c='b', label='original')
plt.plot(days['notif_date'], days['job_type'], c='r', label='estimated')
plt.xlabel('date')
plt.ylabel('count')
plt.legend()
plt.show()
| notebooks/.ipynb_checkpoints/Analysis_JobTypes-AR-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="2NnuRIZedJmK"
# ## Content Based Filtering by hand
#
# This lab illustrates how to implement a content based filter using low level Tensorflow operations.
# The code here follows the technique explained in Module 2 of Recommendation Engines: Content Based Filtering.
#
#
# -
# !pip install tensorflow==2.5
# Make sure to restart your kernel to ensure this change has taken place.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="IzbZLmz1dJmL" outputId="f4f882d9-6752-4b8d-8d7d-83eb61690d89"
import numpy as np
import tensorflow as tf
print(tf.__version__)
# + [markdown] colab_type="text" id="36uCjFhldJmR"
# To start, we'll create our list of users, movies and features. While the users and movies represent elements in our database, for a content-based filtering method the features of the movies are likely hand-engineered and rely on domain knowledge to provide the best embedding space. Here we use the categories of Action, Sci-Fi, Comedy, Cartoon, and Drama to describe our movies (and thus our users).
#
# In this example, we will assume our database consists of four users and six movies, listed below.
# + colab={} colab_type="code" id="ElQV43fxdJmS"
users = ['Ryan', 'Danielle', 'Vijay', 'Chris']
movies = ['Star Wars', 'The Dark Knight', 'Shrek', 'The Incredibles', 'Bleu', 'Memento']
features = ['Action', 'Sci-Fi', 'Comedy', 'Cartoon', 'Drama']
num_users = len(users)
num_movies = len(movies)
num_feats = len(features)
num_recommendations = 2
# + [markdown] colab_type="text" id="s6iJCViqdJmU"
# ### Initialize our users, movie ratings and features
#
# We'll need to enter the user's movie ratings and the k-hot encoded movie features matrix. Each row of the users_movies matrix represents a single user's rating (from 1 to 10) for each movie. A zero indicates that the user has not seen/rated that movie. The movies_feats matrix contains the features for each of the given movies. Each row represents one of the six movies, the columns represent the five categories. A one indicates that a movie fits within a given genre/category.
# + colab={} colab_type="code" id="_0asiLTwdJmV"
# each row represents a user's rating for the different movies
users_movies = tf.constant([
[4, 6, 8, 0, 0, 0],
[0, 0, 10, 0, 8, 3],
[0, 6, 0, 0, 3, 7],
[10, 9, 0, 5, 0, 2]],dtype=tf.float32)
# features of the movies one-hot encoded
# e.g. columns could represent ['Action', 'Sci-Fi', 'Comedy', 'Cartoon', 'Drama']
movies_feats = tf.constant([
[1, 1, 0, 0, 1],
[1, 1, 0, 0, 0],
[0, 0, 1, 1, 0],
[1, 0, 1, 1, 0],
[0, 0, 0, 0, 1],
[1, 0, 0, 0, 1]],dtype=tf.float32)
# + [markdown] colab_type="text" id="aCW5BtGudJmX"
# ### Computing the user feature matrix
#
# We will compute the user feature matrix; that is, a matrix containing each user's embedding in the five-dimensional feature space.
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="isMCBMOFdJmY" outputId="cf7eaa50-95ab-4e8f-916b-27c26d6421dd"
users_feats = tf.matmul(users_movies,movies_feats)
users_feats
# + [markdown] colab_type="text" id="Ps7XXoYwdJmc"
# Next we normalize each user feature vector to sum to 1. Normalizing isn't strictly neccesary, but it makes it so that rating magnitudes will be comparable between users.
# + colab={"base_uri": "https://localhost:8080/", "height": 118} colab_type="code" id="y81EeooodJmc" outputId="904beb39-0a6f-49e0-971f-5198003e7adb"
users_feats = users_feats/tf.reduce_sum(users_feats,axis=1,keepdims=True)
users_feats
# + [markdown] colab_type="text" id="kqOPr51tdJmf"
# #### Ranking feature relevance for each user
#
# We can use the users_feats computed above to represent the relative importance of each movie category for each user.
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="PKLqAD3adJmg" outputId="d535513e-72cd-4120-ef6d-82424efb20d4"
top_users_features = tf.nn.top_k(users_feats, num_feats)[1]
top_users_features
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="pvUmu7MUdJmj" outputId="a9e89bb0-330b-4687-866e-0f209910d8c0"
for i in range(num_users):
feature_names = [features[int(index)] for index in top_users_features[i]]
print('{}: {}'.format(users[i],feature_names))
# + [markdown] colab_type="text" id="Yne0CyZMdJmn"
# ### Determining movie recommendations.
#
# We'll now use the `users_feats` tensor we computed above to determine the movie ratings and recommendations for each user.
#
# To compute the projected ratings for each movie, we compute the similarity measure between the user's feature vector and the corresponding movie feature vector.
#
# We will use the dot product as our similarity measure. In essence, this is a weighted movie average for each user.
# -
users_ratings = tf.matmul(users_feats,tf.transpose(movies_feats))
users_ratings
# + [markdown] colab_type="text" id="o07wODzddJmq"
# The computation above finds the similarity measure between each user and each movie in our database. To focus only on the ratings for new movies, we apply a mask to the all_users_ratings matrix.
#
# If a user has already rated a movie, we ignore that rating. This way, we only focus on ratings for previously unseen/unrated movies.
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="xUgOnV3AdJmr" outputId="2672899f-d626-4e33-e730-7d8b051a3954"
users_ratings_new = tf.where(tf.equal(users_movies, tf.zeros_like(users_movies)),
users_ratings,
tf.zeros_like(tf.cast(users_movies, tf.float32)))
users_ratings_new
# + [markdown] colab_type="text" id="YyNvH46zdJmu"
# Finally let's grab and print out the top 2 rated movies for each user
# + colab={"base_uri": "https://localhost:8080/", "height": 101} colab_type="code" id="PdDGgmSpdJmv" outputId="a921b943-383b-4984-cffd-e0eb5c7ab41e"
top_movies = tf.nn.top_k(users_ratings_new, num_recommendations)[1]
top_movies
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="dCB7Dv9_dJmx" outputId="0d00e5c6-f7bc-4fae-a359-283f2fdb1c4c"
for i in range(num_users):
movie_names = [movies[index] for index in top_movies[i]]
print('{}: {}'.format(users[i],movie_names))
| courses/machine_learning/deepdive/10_recommend/content_based_by_hand.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="FsSzU5I8_d9G" outputId="35773363-2264-4264-ed4c-4c2db2a110a3" colab={"base_uri": "https://localhost:8080/", "height": 890}
import os
assert os.environ['COLAB_TPU_ADDR'], 'Make sure to select TPU from Edit > Notebook settings > Hardware accelerator'
DIST_BUCKET="gs://tpu-pytorch/wheels"
TORCH_WHEEL="torch-1.15-cp36-cp36m-linux_x86_64.whl"
TORCH_XLA_WHEEL="torch_xla-1.15-cp36-cp36m-linux_x86_64.whl"
TORCHVISION_WHEEL="torchvision-0.3.0-cp36-cp36m-linux_x86_64.whl"
# Install Colab TPU compat PyTorch/TPU wheels and dependencies
# !pip uninstall -y torch torchvision
os.chdir('/content/')
# !gsutil cp "$DIST_BUCKET/$TORCH_WHEEL" .
# !gsutil cp "$DIST_BUCKET/$TORCH_XLA_WHEEL" .
# !gsutil cp "$DIST_BUCKET/$TORCHVISION_WHEEL" .
# !pip install "$TORCH_WHEEL"
# !pip install "$TORCH_XLA_WHEEL"
# !pip install "$TORCHVISION_WHEEL"
# !sudo apt-get install libomp5
# + id="ZplZJTMUVo5p" outputId="2815d89d-f589-4030-fa23-b41ea1fa88a1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
VERSION = "20200516" # @param ["1.5" , "20200516", "nightly"]
# !curl https://raw.githubusercontent.com/pytorch/xla/master/contrib/scripts/env-setup.py -o pytorch-xla-env-setup.py
# !python pytorch-xla-env-setup.py --version $VERSION
# + id="BFY7G2vStrRR" outputId="1e906838-1eef-4a87-ffc0-ebcfbfe5772f" colab={"base_uri": "https://localhost:8080/", "height": 894}
# !pip3 install -r "/content/drive/My Drive/htfl/Freeze.txt"
# + id="7jdrjXSfCUuH" outputId="0994a4a2-be45-47d1-ec53-b8a696f6e8ac" colab={"base_uri": "https://localhost:8080/", "height": 50}
import os
print(os.environ["COLAB_TPU_ADDR"])
import torch
# imports the torch_xla package
import torch_xla
import torch_xla.core.xla_model as xm
import torch_xla.distributed.parallel_loader as pl
import torch_xla.distributed.data_parallel as dp
import torch_xla.distributed.xla_multiprocessing as xmp
num_cores = 8
devices = (
xm.get_xla_supported_devices(
max_devices=num_cores) if num_cores != 0 else [])
print("Devices: {}".format(devices))
# + id="4XBeULqTaeYI" outputId="9ef2c527-3d12-4967-92df-735494d28cd0" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta3_bert_config.json
# + id="6t_oe_vAp7yF" outputId="dc5e1d76-56dd-4ef5-b569-2c8ade7520d6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta3_large_bert_config.json
# + id="pjJu5pyBvymR" outputId="8b703710-7847-4ee6-b28d-99cdcb4a4bab" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta_large_bert_config.json
# + id="g6txCS_6jCz6" outputId="27b3ed69-740e-4944-9063-53edd79d5555" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta_large_bert_100_config.json
# + id="m39eQNUwrJdp" outputId="2b6cbdb9-49e6-4330-c2e1-af61129e5228" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta3_large_bert_100_config.json
# + id="qMVVVDYNu2Qi" outputId="ec824bf9-6985-4fcf-908a-9eb755c457b3" colab={"base_uri": "https://localhost:8080/", "height": 1000} pycharm={"name": "#%%\n"}
import os
os.chdir('/content/drive/My Drive/rlfl')
# !python3 train.py -c config/roberta3_bert_100_config.json
| CAIL2020/rlfl_dump/imagecls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_default
# language: python
# name: conda_default
# ---
# %cd ../src
from util import *
from rnn_util import load_embeddings_by_index
glove_50_embeds, glove_50_word_to_idx = load_embeddings_by_index(GLOVE_FILES[50])
len(glove_50_embeds), len(glove_50_word_to_idx)
amazon_glove_embeds, amazon_glove_word_to_idx = load_embeddings_by_index(GLOVE_AMAZON_FILE, 2)
len(amazon_glove_embeds), len(amazon_glove_word_to_idx)
amazon_glove_word_to_idx
i=0
for x in pol_reader():
i += 1
print(i)
import nltk
c = "I just *love* when you say it isn't $5.33 nice to do that!!! /s"
nltk.word_tokenize(c)
| notebooks/rnn2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="TMAuBkINOJRn"
import os
import re
import json
import pickle
import random
from template_config import *
from collections import defaultdict
from nltk.stem.porter import PorterStemmer
from nltk.stem.wordnet import WordNetLemmatizer
# + id="ikglVOlgOJRp"
import nltk
nltk.download('wordnet')
# + id="ummA7CRTOJRq"
ps = PorterStemmer()
lmtzr = WordNetLemmatizer()
# + id="nfovr_-UOJRr"
def read_in_all_data(data_path=DATA_PATH):
training_data = json.load(open(os.path.join(data_path, "train_spider.json")))
tables_org = json.load(open(os.path.join(data_path, "tables.json")))
tables = {tab['db_id']: tab for tab in tables_org}
return training_data, tables
# + id="8C2NDusXOJRr"
def get_all_question_query_pairs(data):
question_query_pairs = []
for item in data:
question_query_pairs.append((item['question_toks'], item['query'], item['db_id']))
return question_query_pairs
# + id="tsSh3SnnOJRs"
training_data, tables = read_in_all_data("data")
train_qq_pairs = get_all_question_query_pairs(training_data)
# + id="JmIDsISeOJRt" outputId="73bfcb8a-d503-4b0b-a399-262065eab489"
print("Training question-query pair count: {}".format(len(train_qq_pairs)))
# + id="s1mhvelTOJRu"
def is_value(token):
"""
as values can either be a numerical digit or a string literal, then we can
detect if a token is a value by matching with regex
"""
is_number = True
try:
float(token)
except ValueError:
is_number = False
is_string = token.startswith("\"") or token.startswith("\'") or token.endswith("\"") or token.endswith("\'")
return is_number or is_string
def remove_all_from_clauses(query_keywords):
"""
remove all keywords from from clauses, until there is no more from clauses
e.g. select {} from {} as {} where {} = {} --> select {} where {} = {}
"""
# remove from clause by deleting the range from "FROM" to "WHERE" or "GROUP"
start_location = 0
count = 0
while "FROM" in query_keywords:
count += 1
if count > 5:
break
print("error query_keywords: ", query_keywords)
from_location = query_keywords.index("FROM")
end_token_locations = [len(query_keywords)] # defaulting to the end of the list
for end_token in ["WHERE", "GROUP", "ORDER"]:
try:
end_token_locations.append(query_keywords.index(end_token, start_location))
except ValueError:
pass
query_keywords = query_keywords[:from_location] + [FROM_SYMBOL] + query_keywords[min(end_token_locations):]
start_location = min(end_token_locations)
return query_keywords
def strip_query(query, table):
"""
returns (stripped query, non keywords)
"""
#get table column names info
column_types = table['column_types']
table_names_original = [cn.lower() for cn in table['table_names_original']]
table_names = [cn.lower() for cn in table['table_names']]
column_names = [cn.lower() for i, cn in table['column_names']]
column_names_original = [cn.lower() for i, cn in table['column_names_original']]
#clean query: replace values, numbers, column names with SYMBOL
query_keywords = []
columns = table_names_original + table_names
query = query.replace(";","")
query = query.replace("\t","")
query = query.replace("(", " ( ").replace(")", " ) ")
# then replace all stuff enclosed by "" with a numerical value to get it marked as {VALUE}
str_1 = re.findall("\"[^\"]*\"", query)
str_2 = re.findall("\'[^\']*\'", query)
values = str_1 + str_2
for val in values:
query = query.replace(val.strip(), VALUE_STR_SYMBOL)
query_tokenized = query.split(' ')
float_nums = re.findall("[-+]?\d*\.\d+", query)
query_tokenized = [VALUE_NUM_SYMBOL if qt in float_nums else qt for qt in query_tokenized]
query = " ".join(query_tokenized)
int_nums = [i.strip() for i in re.findall("[^tT]\d+", query)]
query_tokenized = [VALUE_NUM_SYMBOL if qt in int_nums else qt for qt in query_tokenized]
nums = float_nums + int_nums
#query_tokenized = query.split(' ')
cols_dict = {}
for token in query_tokenized:
if len(token.strip()) == 0: # in case there are more than one space used
continue
if IGNORE_COMMAS_AND_ROUND_BRACKETS:
keywords_dict = SQL_KEYWORDS_AND_OPERATORS_WITHOUT_COMMAS_AND_BRACES
else:
keywords_dict = SQL_KEYWORDS_AND_OPERATORS
if token.upper() not in keywords_dict and token != VALUE_STR_SYMBOL and token != VALUE_NUM_SYMBOL:
token = token.upper()
if USE_COLUMN_AND_VALUE_REPLACEMENT_TOKEN:
token = re.sub("[T]\d+\.", '', token)
token = re.sub(r"\"|\'", '', token)
token = re.sub("[T]\d+", '', token).lower()
# if token in table_names_original:
# query_keywords.append(TABLE_SYMBOL)
# continue
if token != '' and token in column_names_original:
try:
tok_ind = column_names_original.index(token)
except:
print("\ntable: {}".format(table['db_id']))
print("\ntoken: {}".format(token))
print("column_names_original: {}".format(column_names_original))
print("query: {}".format(query))
print("query_tokenized: {}".format(query_tokenized))
exit(1)
col_type = column_types[tok_ind]
col_name = column_names[tok_ind]
columns.append(col_name)
columns.append(token)
if token not in cols_dict:
cols_dict[token] = COLUMN_SYMBOL.replace("}", str(len(cols_dict)))
query_keywords.append(cols_dict[token])
elif token in table_names_original:
query_keywords.append(TABLE_SYMBOL)
continue
else:
query_keywords.append(token.upper())
if "FROM" in query_keywords:
query_keywords = remove_all_from_clauses(query_keywords)
if USE_LIMITED_KEYWORD_SET:
query_keywords = [kw for kw in query_keywords if kw in LIMITED_KEYWORD_SET]
columns_lemed = [lmtzr.lemmatize(w) for w in " ".join(columns).split(" ") if w not in LOW_CHAR]
columns_lemed_stemed = [ps.stem(w) for w in columns_lemed]
return " ".join(query_keywords), values, nums, columns_lemed_stemed
def filter_string(cs):
return "".join([c.upper() for c in cs if c.isalpha() or c == ' '])
def process_question(question, values, nums, columns):
question = " ".join(question).lower()
values = [re.sub(r"\"|\'", '', val) for val in values]
for val in values:
val = val.lower()
try:
question = re.sub(r'\b'+val+r'\b', VALUE_STR_SYMBOL, question)
except:
continue
for num in nums:
num = num.strip()
question = re.sub(r'\b'+num+r'\b', VALUE_NUM_SYMBOL, question)
question_toks = question.split(" ")
question_lemed = [lmtzr.lemmatize(w) for w in question_toks]
question_lemed_stemed = [ps.stem(w) for w in question_lemed]
replace_inds = [i for i, qt in enumerate(question_lemed_stemed) if qt in columns]
#print("question_stemed: {}".format(question_stemed))
#print("replace_inds: {}".format(replace_inds))
for ind in replace_inds:
question_toks[ind] = COLUMN_SYMBOL
question_template = ' '.join(question_toks)
return question_template
# + id="ZCE1SVHuOJRx"
KEY_KEYWORD_SET = {"SELECT", "WHERE", "GROUP", "HAVING", "ORDER", "BY", "LIMIT", "EXCEPT", "UNION", "INTERSECT"}
ALL_KEYWORD_SET = {"SELECT", "WHERE", "GROUP", "HAVING", "DESC", "ORDER", "BY", "LIMIT", "EXCEPT", "UNION",
"INTERSECT", "NOT", "IN", "OR", "LIKE", "(", ">", ")", "COUNT"}
WHERE_OPS = ['=', '>', '<', '>=', '<=', '!=', 'LIKE', 'IS', 'EXISTS']
AGG_OPS = ['MAX', 'MIN', 'SUM', 'AVG']
DASC = ['ASC', 'DESC']
def general_pattern(pattern):
general_pattern_list = []
for x in pattern.split(" "):
if x in KEY_KEYWORD_SET:
general_pattern_list.append(x)
return " ".join(general_pattern_list)
def sub_pattern(pattern):
general_pattern_list = []
for x in pattern.split(" "):
if x in ALL_KEYWORD_SET:
general_pattern_list.append(x)
return " ".join(general_pattern_list)
def tune_pattern(pattern):
general_pattern_list = []
cols_dict = {}
for x in pattern.split(" "):
if "{COLUMN" in x:
if x not in cols_dict:
cols_dict[x] = COLUMN_SYMBOL.replace("}", str(len(cols_dict))+"}")
general_pattern_list.append(cols_dict[x])
continue
if "{VALUE" in x:
general_pattern_list.append("{VALUE}")
continue
if x == 'DISTINCT':
continue
elif x in DASC:
general_pattern_list.append("{DASC}")
elif x in WHERE_OPS:
general_pattern_list.append("{OP}")
elif x in AGG_OPS:
general_pattern_list.append("{AGG}")
else:
general_pattern_list.append(x)
return " ".join(general_pattern_list)
# + id="HG_oUG7eOJRy"
training_question_pattern_pairs = []
training_patterns = set()
pattern_question_dict = defaultdict(list)
# train_qq_pairs
for eid, (question, query, bd_id) in enumerate(train_qq_pairs):
table = tables[bd_id]
if eid % 500 == 0:
print("processing eid: ", eid)
pattern, values, nums, columns = strip_query(query, table)
question_template = process_question(question, values, nums, columns)
gen_pattern = general_pattern(pattern)
more_pattern = sub_pattern(pattern)
tu_pattern = tune_pattern(pattern)
pattern_question_dict[tu_pattern].append(' '.join(question) + " ||| " +
question_template + " ||| " + more_pattern
+ " ||| " + query)
# print("\n--------------------------------------")
# print("original question: {}".format(' '.join(question).encode('utf-8')))
# print("question: {}".format(question_template.encode('utf-8')))
# print("query: {}".format(query))
# print("pattern: {}".format(pattern))
# print("values: {}".format(values))
# print("nums: {}".format(nums))
# print("columns: {}".format(columns))
# + id="7cz7OBJqOJRz" outputId="835ce2ed-9a58-4377-b395-15a96173a11a"
print("total pattern number: {}".format(len(pattern_question_dict)))
pattern_question_dict = sorted(pattern_question_dict.items(), key=lambda kv: len(kv[1]), reverse=True)
# + id="joTm3AK4OJRz"
# filter_nums = [762, 275, 241, 204, 202, 164, 98, 59, 55, 48]
# + id="jNfqLpCsOJR0"
for sql, qts in pattern_question_dict:
# if len(qts) not in filter_nums:
# continue
print("\n--------------------------------------------")
print("SQL Pattern: {}".format(sql))
print("count: {}".format(len(qts)))
for qt in qts:
q, q_template, sql, sql_more = qt.split("|||")
print("question: ", q.replace("""'""", "").replace("""``""", ''))
# print("question: ", q_template.replace("""'""", "").replace("""``""", ''))
print("SQL: {} \n".format(sql_more))
# for qt in qts:
# q, q_template, sql_temp, sql_more = qt.split("|||")
# # print("question: ", q_template)
# # print("sql_temp: ", sql_temp)
# # print("sql_more: ", sql_more)
# if sql == 'SELECT {COLUMN0} {FROM} WHERE {COLUMN4} {OP} {VALUE_STR} AND {COLUMN5} {OP} {VALUE_STR}':
# print(sql_more)
# + id="c71M2EUVOJR2"
for sql, qts in pattern_question_dict:
print("\n")
print("SQL Pattern: {}".format(sql))
print("count: ", len(qts))
# + id="6SkVDZm8OJR2"
for sql_template, qts in pattern_question_dict:
print("\n--------------------------------------")
print("SQL Pattern: {}".format(sql_template))
print("count: ", len(qts))
sql_dict = defaultdict(int)
for qt in qts:
q, q_template, sql, sql_more = qt.split("|||")
sql_dict[sql] += 1
sql_count = sorted(sql_dict.items(), key=lambda kv: kv[1])
for sql, count in sql_count:
print("SQL: {}, count: {}".format(sql, count))
# + id="PSt2Ca_TOJR3"
for sql_template, qts in pattern_question_dict:
print("\n--------------------------------------")
print("SQL Pattern: {}".format(sql_template))
print("count: ", len(qts))
for qt in qts:
q, q_template, sql, sql_more = qt.split("|||")
print("question: ", q)
print("SQL: {} \n".format(sql_more))
| data_synthesis/generate_cfg_patterns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import math
import torch.nn as nn
import torch.optim as optim
import torch.utils
import PIL
from matplotlib import pyplot as plt
from PIL import Image
from torchvision import transforms
from torchvision import datasets
#Downloading CIFAR-10
data_path = '../data-unversioned/p1ch7/'
cifar10 = datasets.CIFAR10(data_path, train=True, download=True)
cifar10_val = datasets.CIFAR10(data_path, train=False, download=True) #下载太慢请开代理
# +
# 引入normalize的数据初始化
tensor_cifar10_normalize_train = datasets.CIFAR10(data_path, train=True, download=False,
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.4915, 0.4823, 0.4468),
(0.2470, 0.2435, 0.2616))
]))
tensor_cifar10_normalize_val = datasets.CIFAR10(data_path, train=True, download=False,
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.4915, 0.4823, 0.4468),
(0.2470, 0.2435, 0.2616))
]))
# +
# Build the dataset and DataLoader
label_map = {0: 0, 2: 1} # 占位符
class_names = ['airplane', 'bird']
# 训练集
cifar2 = [(img, label_map[label])
for img, label in tensor_cifar10_normalize_train
if label in [0, 2]]
# 验证集
cifar2_val = [(img, label_map[label])
for img, label in tensor_cifar10_normalize_val
if label in [0, 2]]
train_loader = torch.utils.data.DataLoader(cifar2, batch_size=64, shuffle=True)
# -
class Animator: #@save
"""在动画中绘制数据。"""
def __init__(self, xlabel=None, ylabel=None, legend=None, xlim=None,
ylim=None, xscale='linear', yscale='linear',
fmts=('-', 'm--', 'g-.', 'r:'), nrows=1, ncols=1,
figsize=(3.5, 2.5)):
# 增量地绘制多条线
if legend is None:
legend = []
d2l.use_svg_display()
self.fig, self.axes = d2l.plt.subplots(nrows, ncols, figsize=figsize)
if nrows * ncols == 1:
self.axes = [self.axes, ]
# 使用lambda函数捕获参数
self.config_axes = lambda: d2l.set_axes(
self.axes[0], xlabel, ylabel, xlim, ylim, xscale, yscale, legend)
self.X, self.Y, self.fmts = None, None, fmts
def add(self, x, y):
# 向图表中添加多个数据点
if not hasattr(y, "__len__"):
y = [y]
n = len(y)
if not hasattr(x, "__len__"):
x = [x] * n
if not self.X:
self.X = [[] for _ in range(n)]
if not self.Y:
self.Y = [[] for _ in range(n)]
for i, (a, b) in enumerate(zip(x, y)):
if a is not None and b is not None:
self.X[i].append(a)
self.Y[i].append(b)
self.axes[0].cla()
for x, y, fmt in zip(self.X, self.Y, self.fmts):
self.axes[0].plot(x, y, fmt)
self.config_axes()
display.display(self.fig)
display.clear_output(wait=True)
# +
device = torch.device('cuda:0')
model_F3 = nn.Sequential(
nn.Linear(3072, 512),
nn.Tanh(),
nn.Linear(512, 2),
nn.LogSoftmax(dim=1))
model_F3.to(device)
lr = 1e-2
optimizer = optim.SGD(model_F3.parameters(),lr =lr)
loss_fn = nn.NLLLoss()
n_epochs = 100
for epoch in range(n_epochs):
for imgs, labels in train_loader:
imgs, labels = imgs.to(device), labels.to(device)
batch_size = imgs.shape[0]
outputs = model_F3(imgs.view(batch_size, -1))
loss = loss_fn(outputs, labels)
#out = model_F3(img.view(-1).unsqueeze(0)).to(device)
#loss = loss_fn(out,torch.tensor([label]))
optimizer.zero_grad()
loss.backward()
optimizer.step()
print("Epoch: %d, Loss: %f" % (epoch, float(loss)))
# +
device = (torch.device('cuda')
if torch.cuda.is_available()
else torch.device('cpu'))
print(f"Training on device {device}.")
# -
class model_chap7(nn.Module):
def __init__(self, config):
super(model_chap7, self).__init__()
self.conv1 = nn.Conv2d(3, 32, kernel_size=3, padding=1)
self.conv2 = nn.Conv2d(32, 16, kernel_size=3, padding=1)
self.fc1 = nn.Linear(16 * 8 * 8, 32)
self.fc2 = nn.Linear(32, 2)
def forward(self,torch_input):
encoder = self.encoder(torch_input)
decoder = self.decoder(encoder)
return decoder
def loss(self,decoder,input,optimizer,mask_input):
loss_fn = nn.NLLLoss()
return cost,rmse
class NetWidth(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3, 32, kernel_size=3, padding=1)
self.conv2 = nn.Conv2d(32, 16, kernel_size=3, padding=1)
self.fc1 = nn.Linear(16 * 8 * 8, 32)
self.fc2 = nn.Linear(32, 2)
def forward(self, x):
out = F.max_pool2d(torch.tanh(self.conv1(x)), 2)
out = F.max_pool2d(torch.tanh(self.conv2(out)), 2)
out = out.view(-1, 16 * 8 * 8)
out = torch.tanh(self.fc1(out))
out = self.fc2(out)
return out
autorec_config = \
{
'train_ratio': 0.9,
'num_epoch': 100,
'batch_size': 100,
'optimizer': 'SGD',
'adam_lr': 1e-2,
'lambda': 1,
'device_id': 2,
'use_cuda': True,
'model_name': 'model_chap7'
}
# 实例化AutoRec对象
model = model_chap7(autorec_config)
'''Train'''
def train(epoch):
loss = 0
for step, (batch_x, batch_y) in enumerate(train_loader):
batch_x = batch_x.type(torch.FloatTensor)
decoder = rec(batch_x) # 第一步,数据的前向传播,计算预测值
loss, rmse = rec.loss(decoder=decoder, input=batch_x, optimizer=optimer, mask_input=batch_mask_x) # 第二步,计算误差
optimer.zero_grad() # 反向传播前,梯度归零
loss.backward() # 第三步,反向传播
optimer.step() # 一步更新所有参数
cost_all += loss
RMSE += rmse
RMSE = np.sqrt(RMSE.detach().cpu().numpy() / (train_mask_r == 1).sum())
animator.add(epoch + 1, RMSE)
| chap7/chap7_v3_gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyse class attendance via Zoom report
#
# Here's how to use this notebook:
#
# 
#
# For instance in the class I built this for, there's a subdirectory under this notebook which has the following files
#
# 
#
# -----
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
import seaborn as sns
sns.set_style('white')
import pandas as pd
import glob
import sys
def load_data(path):
''' loads all csv data in path,
assumes files naming format is 'zoomus_meeting_report_section_year_month_day.csv' '''
if path[-1]!='/': # make sure path has a trailing slash
path = path + '/'
files = glob.glob(path+'*.csv') # gets all .csv filenames in directory
if not files:
print('Oops! No csv files in ' + path)
raise OSError
else:
dfs = []
for afile in files:
section = afile.split('_')[3]
date = pd.Timestamp( '-'.join( afile.split('_')[4:] ).split('.')[0] )
# print(section,date)
data = pd.read_csv(afile)
data['Section'] = section
data['Date'] = date
data = data.set_index(['Date','Section','Name'])
dfs.append( data )
results = pd.concat(dfs).sort_index()
print('Found {} csv files covering {} sections over {} dates in {}'.format(
len(files), len(results.reset_index().Section.unique()),
len(results.reset_index().Date.unique()), path ))
return results
def ind_att(df):
''' when students join-then-leave-then-rejoin they get multiple entries in the attendance report
this fct is meant to be called on data grouped by Name (plus other vars if desired) and it
combines the different lines into a single description of the attendance for the student '''
datadict={}
datadict['Email'] = df['Email'].iloc[0] # doesn't matter which, all should be same
datadict['Join Time'] = df['Join Time'].min() # first time they arrive
datadict['Leave Time'] = df['Leave Time'].max() # last time they left
datadict['Duration(Minutes)'] = df['Duration(Minutes)'].sum() # total of all visits
# this is how many times the student left and reentered.
# maybe use this to figure out which students have bad connections that may hurt their work?
datadict['In-n-outs'] = df['Duration(Minutes)'].count()
return pd.DataFrame(datadict,index=df.reset_index().Name).drop_duplicates()
def individual_attendance(df):
''' calls ind_att to get the work done, handles dataframe cleanup
the droplevel is because att_report ends up duplicating Name into the index a 2nd time '''
return ( df
.groupby(['Date','Section','Name'])
.apply(ind_att)
.droplevel(-1) )
def section_attendance_report(df):
'''reports how many people were in each section on each date'''
return ( individual_attendance(df)
.groupby([ 'Date','Section'])
.count()['Join Time'] # join time is never NA, use it for counting
.rename('Attendees')
.to_frame() )
def individual_date_attendance_report(df):
'''reports how many dates someone attended;
UPDATED: if they do more than 1 section per day (like a TA) they will now only show up once
since we dont want to give attendance extra credit for someone who goes to every section 1 week ;) '''
return ( individual_attendance(df)
.groupby(['Name','Date'])
.count()['Join Time'] # join time is never NA, use it for counting
.rename('Attendances')
# we don't care how many sections each day someone went to
# we just want how many days they attended at least one section; collapse all non-zero to 1
.apply( lambda x: 1)
.groupby(['Name']) # now lets count up all the days they were there
.count()
.rename('Dates attended')
.to_frame() )
# +
# if you organized your directory differently, you should change 'Attendance/' to the correct directory location
attendance = load_data('Attendance/')
attendance
# NB: I've obfuscated the student information so that I can distribute this notebook to others
# without compromising student privacy
# when you do this you will see the full name and email as the students entered it into Zoom
# -
sections = section_attendance_report(attendance)
sections
sections.hist();
plt.xlabel('Attendees in a section')
plt.ylabel('# of sections')
plt.title('Histogram of sections with a given level of attendance');
sns.lineplot( data=sections.reset_index(),
x='Date',y='Attendees', hue='Section')
plt.title('Attendance over time in each section');
# +
# this generates a report on how many dates somone attended discussion sections
individuals = individual_date_attendance_report(attendance)
individuals
# -
# if you wanted to write out the attendance report to look at it inside Excel or something you can
individuals.to_excel('dates_attended.xlsx')
# this is how you could check who was there just for section A03 on Jan 13th
results = individual_attendance(attendance.loc['2020-01-13','A03',slice(None)])
results
| Attendance report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bfCN2QLKlrHs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="979b765c-d104-46fd-9710-826756f9bb4b"
# !git clone https://github.com/lessw2020/mish.git
# + id="FweRsNG-mXgg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="6388f54f-cab8-4d0c-bfb5-e95d011a09ce"
# !git clone https://github.com/mgrankin/over9000.git
# + id="L_p13UODlz_Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ec872522-5835-45d9-a45e-8455a75ddd3a"
# %cd mish
# + id="xdBjArECn0Yy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="aa9ae37e-65b4-4bfb-a731-2281e1e651d0"
from mxresnet import *
# + id="PEBecFdOoHpX" colab_type="code" colab={}
import torch, math
from torch.optim.optimizer import Optimizer
import itertools as it
class Ralamb(Optimizer):
def __init__(self, params, lr=1e-3, betas=(0.9, 0.999), eps=1e-8, weight_decay=0):
defaults = dict(lr=lr, betas=betas, eps=eps, weight_decay=weight_decay)
self.buffer = [[None, None, None] for ind in range(10)]
super(Ralamb, self).__init__(params, defaults)
def __setstate__(self, state):
super(Ralamb, self).__setstate__(state)
def step(self, closure=None):
loss = None
if closure is not None:
loss = closure()
for group in self.param_groups:
for p in group['params']:
if p.grad is None:
continue
grad = p.grad.data.float()
if grad.is_sparse:
raise RuntimeError('Ralamb does not support sparse gradients')
p_data_fp32 = p.data.float()
state = self.state[p]
if len(state) == 0:
state['step'] = 0
state['exp_avg'] = torch.zeros_like(p_data_fp32)
state['exp_avg_sq'] = torch.zeros_like(p_data_fp32)
else:
state['exp_avg'] = state['exp_avg'].type_as(p_data_fp32)
state['exp_avg_sq'] = state['exp_avg_sq'].type_as(p_data_fp32)
exp_avg, exp_avg_sq = state['exp_avg'], state['exp_avg_sq']
beta1, beta2 = group['betas']
# Decay the first and second moment running average coefficient
# m_t
exp_avg.mul_(beta1).add_(1 - beta1, grad)
# v_t
exp_avg_sq.mul_(beta2).addcmul_(1 - beta2, grad, grad)
state['step'] += 1
buffered = self.buffer[int(state['step'] % 10)]
if state['step'] == buffered[0]:
N_sma, radam_step_size = buffered[1], buffered[2]
else:
buffered[0] = state['step']
beta2_t = beta2 ** state['step']
N_sma_max = 2 / (1 - beta2) - 1
N_sma = N_sma_max - 2 * state['step'] * beta2_t / (1 - beta2_t)
buffered[1] = N_sma
# more conservative since it's an approximated value
if N_sma >= 5:
radam_step_size = math.sqrt((1 - beta2_t) * (N_sma - 4) / (N_sma_max - 4) * (N_sma - 2) / N_sma * N_sma_max / (N_sma_max - 2)) / (1 - beta1 ** state['step'])
else:
radam_step_size = 1 / (1 - beta1 ** state['step'])
buffered[2] = radam_step_size
update = torch.zeros_like(p_data_fp32)
if N_sma >= 5:
denom = exp_avg_sq.sqrt().add_(group['eps'])
update.addcdiv_(radam_step_size, exp_avg, denom)
else:
update.add_(radam_step_size, exp_avg)
if group['weight_decay'] != 0:
update.add_(group['weight_decay'], p_data_fp32)
radam_norm = update.pow(2).sum().sqrt()
weight_norm = p.data.pow(2).sum().sqrt()
if weight_norm == 0 or radam_norm == 0:
trust_ratio = 1
else:
trust_ratio = weight_norm / radam_norm
trust_ratio = max(0, min(10, trust_ratio))
state['weight_norm'] = weight_norm
state['adam_norm'] = radam_norm
state['trust_ratio'] = trust_ratio
p_data_fp32.add_(-update * trust_ratio * group['lr'])
p.data.copy_(p_data_fp32)
return loss
# + id="C7gbBokWmBkM" colab_type="code" colab={}
from fastai.vision import *
# + id="jHeXN8wUmQDG" colab_type="code" colab={}
path = untar_data(URLs.IMAGEWOOF)
# + id="X_eXff-kmRZR" colab_type="code" colab={}
data = (ImageList.from_folder(path).split_by_folder(valid='val')
.label_from_folder().transform(([flip_lr(p=0.5)], []), size=128)
.databunch(bs=64, num_workers=2)
.presize(128, scale=(0.35,1))
.normalize(imagenet_stats))
# + id="miE82HL5mUO2" colab_type="code" colab={}
opt_func=partial(Ralamb, betas = (0.9,0.99), eps=1e-6)
# + id="h1-pWlf7mc2a" colab_type="code" colab={}
from fastai.script import *
from fastai.vision import *
from fastai.callbacks import *
from fastai.distributed import *
# + id="NuoWTk-wmdfJ" colab_type="code" colab={}
lr = 1e-2
# + id="ku7zASqwmebx" colab_type="code" colab={}
res = []
num_epoch=5
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
metrics=[accuracy, top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func=LabelSmoothingCrossEntropy())
n = len(learn.data.train_dl)
anneal_start = int(n*5*0.7)
phase0 = TrainingPhase(anneal_start).schedule_hp('lr', lr)
phase1 = TrainingPhase(n*5 - anneal_start).schedule_hp('lr', lr, anneal=annealing_cos)
phases = [phase0, phase1]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(num_epoch)
loss, acc, topk = learn.validate()
res.append(acc.numpy())
# + id="8eLS4NYRq2vW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f10df1a0-5b5b-4527-9dcf-664470c775eb"
print(res)
# + id="JDenE8xjmh_l" colab_type="code" colab={}
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
metrics=[accuracy, top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func=LabelSmoothingCrossEntropy())
n = len(learn.data.train_dl)
anneal_start = int(n*5*0.7)
phase0 = TrainingPhase(anneal_start).schedule_hp('lr', lr)
phase1 = TrainingPhase(n*5 - anneal_start).schedule_hp('lr', lr, anneal=annealing_cos)
phases = [phase0, phase1]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(num_epoch)
loss, acc, topk = learn.validate()
res.append(acc.numpy())
# + id="Zq7pGbJvnUI_" colab_type="code" colab={}
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
metrics=[accuracy, top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func=LabelSmoothingCrossEntropy())
n = len(learn.data.train_dl)
anneal_start = int(n*5*0.7)
phase0 = TrainingPhase(anneal_start).schedule_hp('lr', lr)
phase1 = TrainingPhase(n*5 - anneal_start).schedule_hp('lr', lr, anneal=annealing_cos)
phases = [phase0, phase1]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(num_epoch)
loss, acc, topk = learn.validate()
res.append(acc.numpy())
# + id="wQKa41nWnUe9" colab_type="code" colab={}
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
metrics=[accuracy, top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func=LabelSmoothingCrossEntropy())
n = len(learn.data.train_dl)
anneal_start = int(n*5*0.7)
phase0 = TrainingPhase(anneal_start).schedule_hp('lr', lr)
phase1 = TrainingPhase(n*5 - anneal_start).schedule_hp('lr', lr, anneal=annealing_cos)
phases = [phase0, phase1]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(num_epoch)
loss, acc, topk = learn.validate()
res.append(acc.numpy())
# + id="sQUjfI0TnVF1" colab_type="code" colab={}
learn = Learner(data, mxresnet50(c_out=10), wd=1e-2, opt_func=opt_func,
metrics=[accuracy, top_k_accuracy],
bn_wd=False, true_wd=True,
loss_func=LabelSmoothingCrossEntropy())
n = len(learn.data.train_dl)
anneal_start = int(n*5*0.7)
phase0 = TrainingPhase(anneal_start).schedule_hp('lr', lr)
phase1 = TrainingPhase(n*5 - anneal_start).schedule_hp('lr', lr, anneal=annealing_cos)
phases = [phase0, phase1]
sched = GeneralScheduler(learn, phases)
learn.callbacks.append(sched)
learn.fit(num_epoch)
loss, acc, topk = learn.validate()
res.append(acc.numpy())
# + id="Jk6QoC3onVcB" colab_type="code" colab={}
| ImageWoofTests/with_fix_did_not_work.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://www.kaggle.com/kamaljp/feature-engineering-nn-modeling-wids?scriptVersionId=86681868" target="_blank"><img align="left" alt="Kaggle" title="Open in Kaggle" src="https://kaggle.com/static/images/open-in-kaggle.svg"></a>
# ### Purpose of the Notebook:
#
# EDA was done, which gave good understanding of the building, environment conditions
# https://www.kaggle.com/kamaljp/building-energy-usage-edanmodeling?kernelSessionId=85754348
#
# Modeling was tried with multiple Machine learning and introductory Neural Nets
# https://www.kaggle.com/kamaljp/modeling-site-eui-wids/notebook?kernelSessionId=85945920
#
# The results were still not better than first Lasso Regresor model score along with the EDA. The data is holding its secrets so strong. This notebook explores the Feature Engineering using dimensionality reduction methods, and use that reduced dataset to model using lasso regressor. Here goes nothing...
#
# ### What to Expect
#
# Truncated SVD is used to reduce the dimensions from 127 to 60. You can see the impact it has had [here](#tsnevis). Following in the same path, tSNE further reduces the entire components to just 2 parameters.
# The visuals are as usual insightful,but this time they are having depth to it.
#
# The model training, and results generation has been converted into functions. The entire model, instantiate, cross_validate, train and test has been wrapped into a [function](#peek2). This is done after using the process multiple times in earlier notebooks. Similarly, function for svd calculation, svd variance explanation are also made into functions.
#
#
# ### Sneek Peek
#
# tSNE is about inter-dimensional data transformation. The tSNE visual of the [training and testing](#peek1) data shows the power of this algorithm. Here were [errors](#error) that caused challenges in executing functions. The final [results](#finres) is sufficiently strange, but the values seems to make sense.
#
# PS: Use the blue links to go the exact location of the code and related activity
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _kg_hide-output=true
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import os
import numpy as np
import pandas as pd
import seaborn as sns
import plotly.express as px
from plotly.offline import init_notebook_mode
from plotly.subplots import make_subplots
import plotly.graph_objects as go
init_notebook_mode(connected=True)
pd.set_option('display.max_columns', 5000)
import warnings
warnings.filterwarnings("ignore")
#os.mkdir('/kaggle/working/individual_charts/')
import matplotlib.pyplot as plt
# Load the data
#Will come in handy to wrap the lengthy texts
import textwrap
#useful libraries and functions
from itertools import repeat
#Libraries that give a different visual possibilities
from pandas import option_context
from plotly.subplots import make_subplots
def long_sentences_seperate(sentence, width=30):
try:
splittext = textwrap.wrap(sentence,width)
text = '<br>'.join(splittext)#whitespace is removed, and the sentence is joined
return text
except:
return sentence
def load_csv(base_dir,file_name):
"""Loads a CSV file into a Pandas DataFrame"""
file_path = os.path.join(base_dir,file_name)
df = pd.read_csv(file_path,low_memory=False)
return df
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
# ## <a id='contents'> Contents </a>
#
# ### [tSNE with Truncated SVD](#tsne)
#
# #### [Transformation](#tsnemod)
#
# #### [Visualisating features](#tsnevis)
#
# #### [Machine Learning on Features](#tsnelea)
#
# #### [Dimension reduction results](#finres)
#
# #### [Lessons of the Exercise ](#FinCon)
#
# ### [DNN with Encoded Dataset ](#dnn)
#
# #### [Model Building](#dmod)
#
# #### [Fitting model ](#dfit)
#
# #### [Result & Understanding](#dres)
#
# + _kg_hide-input=true
base_dir = '../input/widsdatathon2022/'
file_name = 'train.csv'
dataset_main = load_csv(base_dir,file_name)
# + _kg_hide-input=true
#Single Factor Bar Graphs
def uni_factor(factor):
grp_factor = dataset_main.groupby(factor)['floor_area'].count().reset_index()
grp_factor.sort_values(by='floor_area',inplace=True,ascending = False)
grp_factor[factor] = grp_factor[factor].astype('category')
vis = px.bar(data_frame=grp_factor,y = factor,x ='floor_area',color= factor)
vis.update_layout(yaxis = {'categoryorder' : 'total ascending'},
title = 'Number of Buildings based on ' + factor,
height = 800)
vis.show()
#Single Factor Histogram Graphs
def uni_hist_plot(independent,dependent):
vis = px.histogram(data_frame=dataset_main,x=dependent,color=independent)
vis.update_layout(title='Distribution of '+ dependent + ' based on '+ independent)
vis.show()
#Single Factor Box Plot Graphs
def uni_box_plot(independent,dependent):
vis = px.box(data_frame=dataset_main,x=dependent,color=independent)
vis.update_layout(title='Box plot of '+ dependent + ' based on '+ independent)
vis.show()
#Two Factor Scatter Plot Graphs
def two_factor(factor1, factor2,independent):
vis = px.scatter(data_frame=dataset_main,y = factor1,x =factor2,color=independent,
facet_col=independent,facet_col_wrap=3)
vis.update_layout(title = 'Relation between ' + factor1 + ' and ' +factor2 + ' in ' + independent + 'condition',
height = 1000)
vis.show()
#Single factor target average Bar Graph
def avg_on_factor(factor,target):
grp_factor = dataset_main.groupby(factor)[target].mean().reset_index()
grp_factor.sort_values(by=target,inplace=True,ascending = False)
grp_factor[factor] = grp_factor[factor].astype('category')
vis = px.bar(data_frame=grp_factor,y = factor,x =target,color= factor)
vis.update_layout(yaxis = {'categoryorder' : 'total ascending'},
title = 'Average of '+target +' on basis of ' + factor,
height = 800)
vis.show()
def corr_heat_map(df,title):
#Building the dataset with column that are numerical
df = df[df.columns[df.dtypes != 'object']]
df_corr_mat = df.corr() #building the correlation matrix
#Building the lower triagle of the correlation matrix
df_corr_mat_lt = df_corr_mat.where(np.tril(np.ones(df_corr_mat.shape)).astype(np.bool))
vis = px.imshow(df_corr_mat_lt,aspect="auto",
height=1000,color_continuous_scale='spectral',width=900)
vis.update_layout(title=title)
vis.show()
# + _kg_hide-input=true _kg_hide-output=true
#Collecting garbage memory and deleting unwanted Dataframes, that have served their purpose earlier
import gc
gc.collect()
# + _kg_hide-input=true
#https://stackoverflow.com/a/46581125/16388185
def clean_dataset(df):
assert isinstance(df, pd.DataFrame), "df needs to be a pd.DataFrame"
df.dropna(inplace=True)
indices_to_keep = ~df.isin([np.nan, np.inf, -np.inf]).any(1)
return df[indices_to_keep].astype(np.float64)
# + _kg_hide-input=true _kg_hide-output=true
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
from sklearn.linear_model import ElasticNet
from sklearn.tree import DecisionTreeRegressor
#Import Model Packages
from sklearn.decomposition import PCA
from sklearn.decomposition import TruncatedSVD
from numpy.linalg import inv, eig, svd
from sklearn.manifold import TSNE
from sklearn.decomposition import KernelPCA
# Error Metrics
from sklearn.metrics import mean_squared_error
# -
# #### [Back to Contents](#contents)
# #### <a id='tsne'> tSNE with Truncated SVD </a>
#
# Dataset prepared by seperating the categorical and the numerical values, followed by one hot encoding the categorical columns. Intention is to understand how tSNE creates the features from so many parameters that make the dataset. Then use that features to model.
#
# [1) Transformation](#tsnemod)
#
# [2) Visualisating Features](#tsnevis)
#
# [3) Machine learning on Features](#tsnelea)
# ##### <a id='tsnemod'> Transformation</a>
# + _kg_hide-output=true _kg_hide-input=true
dataset_main.direction_max_wind_speed = dataset_main.direction_max_wind_speed.fillna(1)
dataset_main.direction_peak_wind_speed = dataset_main.direction_peak_wind_speed.fillna(1)
dataset_main.max_wind_speed = dataset_main.max_wind_speed.fillna(1)
dataset_main.days_with_fog = dataset_main.days_with_fog.fillna(dataset_main.days_with_fog.median())
dataset_main.loc[dataset_main.energy_star_rating.isna(),'energy_star_rating'] = np.median(dataset_main.loc[~dataset_main.energy_star_rating.isna(),'energy_star_rating'])
dataset_main.loc[dataset_main.year_built.isna(),'year_built'] = np.median(dataset_main.loc[~dataset_main.year_built.isna(),'year_built'])
#That leaves the Energy star rating. We need to check the test set provided for deciding
# Segregating the columns with categorical value
data_cat = dataset_main[dataset_main.columns[dataset_main.dtypes == 'object']]
data_num = dataset_main[dataset_main.columns[dataset_main.dtypes != 'object']]
data_num.drop('Year_Factor',axis=1,inplace=True)
#A simple and straight_forward one-hot encoding using Pandas' Get_Dummies
data_cat = pd.get_dummies(data_cat,columns=data_cat.columns)
data_model = pd.merge(left=data_cat,right=data_num,left_index=True,right_index=True)
# + _kg_hide-input=true
#Creating the X the Independent variables and Y the Target or Dependent variables
data_model = clean_dataset(data_model)
#We lost another 2500 entries to the big number and infinity issues
X = data_model.iloc[:,:-2]
Y = data_model.site_eui
# + _kg_hide-input=true
# split out validation dataset for the end
validation_size = 0.2
#In case the data is not dependent on the time series, then train and test split randomly
seed = 7
# X_train, X_validation, Y_train, Y_validation = train_test_split(X, Y, test_size=validation_size, random_state=seed)
#In case the data is not dependent on the time series, then train and test split should be done based on sequential sample
#This can be done by selecting an arbitrary split point in the ordered list of observations and creating two new datasets.
train_size = int(len(X) * (1-validation_size))
X_train, X_test = X[0:train_size], X[train_size:len(X)]
Y_train, Y_test = Y[0:train_size], Y[train_size:len(X)]
# -
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X_train)
rescaledDataset = pd.DataFrame(scaler.fit_transform(X_train),
columns = X_train.columns, index = X_train.index)
# summarize transformed data
X_train.dropna(how='any', inplace=True)
rescaledDataset.dropna(how='any', inplace=True)
scaler = StandardScaler().fit(X_test)
rescaledtestset = pd.DataFrame(scaler.fit_transform(X_test),
columns = X_test.columns, index = X_test.index)
# summarize transformed data
X_test.dropna(how='any', inplace=True)
rescaledtestset.dropna(how='any', inplace=True)
#Function to calculate the explained variance
def svd_cal(data, components):
ncomps = components
#Instantiating SVD
svd = TruncatedSVD(n_components=ncomps)
svd_fit = svd.fit(data)
#predicting the target values using the rescaled data
Y_pred = svd.fit_transform(data)
#calculating the cumulative variance preserved
return round(svd_fit.explained_variance_ratio_.cumsum()[-1]*100,2)
# +
# plotting the retained variance based on the parameters
exp_var = []
for para in range(5, 70):
exp_var.append(svd_cal(data= rescaledDataset, components= para))
vis1 = go.Figure()
vis1.add_trace(go.Scatter(x=np.arange(1, len(exp_var) + 1),y=exp_var,mode='lines',name='explained_variance'))
vis1.update_xaxes(title = "Cumulative Sum")
vis1.update_yaxes(title = "Number of Parameters")
vis1.show()
# -
#Function to reduce the dimensions
def svd_red(dataset,comps):
svd = TruncatedSVD(n_components=comps)
svd_fit = svd.fit(dataset)
Y_pred = svd.fit_transform(dataset)
dfsvd = pd.DataFrame(Y_pred, columns=['c{}'.format(c) for c in range(comps)], index=dataset.index)
return dfsvd
# Based on the number of components, the final X_train svd is executed
dfsvd = svd_red(dataset=rescaledDataset,comps=60)
dfsvd_test = svd_red(dataset=rescaledtestset,comps=60)
print(dfsvd_test.shape)
print(dfsvd.shape)
# #### [Back to Contents](#contents)
# ##### <a id='tsnevis'> Visualising the Parameters </a>
#
# The parameters themselves don't seem to have even slightest correlation. However they all have some negative or positive correlation with the target value
# + _kg_hide-input=true
svdcol = dfsvd.columns
dfsvd['signal'] = Y_train
corr_heat_map(dfsvd,'Visualising the Truncated Parameters')
# + _kg_hide-input=true
#Two Factor tsne Scatter Plot Graphs
def tsne_scatter(data):
vis = px.scatter(data_frame=data,y = 'y',x = 'x',color='signal')
vis.update_layout(title = 'Scatterplot of a Multiple dimension dataset reduced to 2D using t-SNE',
height = 1000)
vis.show()
# + _kg_hide-input=true _kg_hide-output=true
#Reducing the SVD dataframe further down to 2 dimensions using tSNE
tsne = TSNE(n_components=2, random_state=0)
Z = tsne.fit_transform(dfsvd[svdcol])
dftsne = pd.DataFrame(Z, columns=['x','y'], index=dfsvd.index)
dftsne['signal'] = Y_train
#Reducing the SVD test dataframe further down to 2 dimensions using tSNE
tsne = TSNE(n_components=2, random_state=0)
Z = tsne.fit_transform(dfsvd_test)
dftsne_test = pd.DataFrame(Z, columns=['x','y'], index=dfsvd_test.index)
dftsne_test['signal'] = Y_test
# -
# ##### <a id='peek1'> tSNE Visualisation linking SVD Train & Test variables to Target </a>
tsne_scatter(dftsne)
tsne_scatter(dftsne_test)
# #### [Back to Contents](#contents)
# ##### <a id='tsnelea'> Machine learning on Features </a>
# + _kg_hide-input=true
num_folds = 10
scoring = 'neg_mean_squared_error'
#scoring ='neg_mean_absolute_error'
#scoring = 'r2'
models = []
models.append(('LASSO', Lasso()))
models.append(('EN', ElasticNet()))
# -
# <a id='peek2'> Machine learning function </a>
# + _kg_hide-input=true
def machine_learn(X_train,Y_train,X_test,Y_test,models):
names = []
kfold_results = []
test_results = []
train_results = []
for name, model in models:
names.append(name)
## K Fold analysis:
kfold = KFold(n_splits=num_folds, random_state=seed)
#converted mean square error to positive. The lower the beter
cv_results = -1* cross_val_score(model, X_train, Y_train, cv=kfold, scoring=scoring)
kfold_results.append(cv_results)
# Full Training period
print('{} model fit Started'.format(model))
res = model.fit(X_train, Y_train)
print('{} model fit Completed'.format(model))
#The error function is root of mean_squared_error
train_result = np.sqrt(mean_squared_error(res.predict(X_train), Y_train))
train_results.append(train_result)
# Test results
#The error function is root of mean_squared_error
test_result = np.sqrt(mean_squared_error(res.predict(X_test), Y_test))
test_results.append(test_result)
print('{} test result'.format(model))
return kfold_results, train_result, test_result
# + _kg_hide-input=true _kg_hide-output=true
#Learning using the SVD dataframe with 60 parameters
dfsvd = dfsvd[svdcol] #the trainin data has to be removed
kfold_results, train_result, test_result = machine_learn(X_train=dfsvd,Y_train=Y_train,
X_test=dfsvd_test,Y_test = Y_test,
models=models)
# + _kg_hide-input=true
kfold = pd.DataFrame(kfold_results,columns=range(1,11)).T
kfold.columns = [x[0] for x in models]
# + _kg_hide-input=true
visB = go.Figure()
for kf in kfold.columns:
visB.add_trace(go.Box(x=kfold[kf],name=kf))
visB.update_xaxes(type='log')
visB.update_layout(title='Kfold Error Results')
visB.show()
# + _kg_hide-input=true
# compare algorithms
fig = plt.figure()
names = [x[0] for x in models]
ind = np.arange(len(names)) # the x locations for the groups
width = 0.35 # the width of the bars
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
plt.bar(ind - width/2, train_result, width=width, label='Train Error')
plt.bar(ind + width/2, test_result, width=width, label='Test Error')
fig.set_size_inches(15,8)
plt.legend()
ax.set_xticks(ind)
ax.set_xticklabels(names)
plt.show()
# -
# The SVD dataset has not provided any good result, the test error is still above 56, the value that was achieved in the earlier notebooks. Next step is to use the tSNE reduced dataset
# + _kg_hide-input=true _kg_hide-output=true
#Learning using the SVD dataframe with 60 parameters
kfold_results, train_result, test_result = machine_learn(X_train=dftsne,Y_train=Y_train,
X_test=dftsne_test,Y_test = Y_test,
models=models)
# + _kg_hide-input=true
kfold = pd.DataFrame(kfold_results,columns=range(1,11)).T
kfold.columns = [x[0] for x in models]
visT = go.Figure()
for kf in kfold.columns:
visT.add_trace(go.Box(x=kfold[kf],name=kf))
visT.update_xaxes(type='log')
visT.update_layout(title='tSNE Kfold Error Results')
visT.show()
# -
# <a id='peek3'> tSNE ML test and train results </a>
# + _kg_hide-input=true
# compare algorithms
fig = plt.figure()
names = [x[0] for x in models]
ind = np.arange(len(names)) # the x locations for the groups
width = 0.35 # the width of the bars
fig.suptitle('Algorithm Comparison')
ax = fig.add_subplot(111)
plt.bar(ind - width/2, train_result, width=width, label='Train Error')
plt.bar(ind + width/2, test_result, width=width, label='Test Error')
fig.set_size_inches(15,8)
plt.legend()
ax.set_xticks(ind)
ax.set_xticklabels(names)
plt.show()
# -
# #### [Back to Contents](#contents)
# <a id='error'> error in filling null values </a>
#
# In the below cell, the commands to fill null values for the Energy_star_rating and Year_built created considerable head ache, still I realised the mistake. Always use the proper iloc or loc indexing, is the lesson
# + _kg_hide-input=true
#loading test_set
test_main = load_csv(base_dir='../input/widsdatathon2022',file_name='test.csv')
#Need to find an effective way to deal with all the columns, so lets try describe
test_main.describe()
test_main.direction_max_wind_speed = test_main.direction_max_wind_speed.fillna(1)
test_main.direction_peak_wind_speed = test_main.direction_peak_wind_speed.fillna(1)
test_main.max_wind_speed = test_main.max_wind_speed.fillna(1)
#direction_max_wind_speed, direction_peak_wind_speed, max_wind_speed all can be safely filled with 1
test_main.days_with_fog = test_main.days_with_fog.fillna(0)
test_main.loc[test_main.energy_star_rating.isna(),'energy_star_rating'] = np.median(test_main.loc[~test_main.energy_star_rating.isna(),'energy_star_rating'])
test_main.loc[test_main.year_built.isna(),'year_built'] = np.median(test_main.loc[~test_main.year_built.isna(),'year_built'])
# Segregating the columns with categorical value
test_cat = test_main[test_main.columns[test_main.dtypes == 'object']]
test_num = test_main[test_main.columns[test_main.dtypes != 'object']]
test_num.drop('Year_Factor',axis=1,inplace=True)
#A simple and straight_forward one-hot encoding using Pandas' Get_Dummies
test_cat = pd.get_dummies(test_cat,columns=test_cat.columns)
test_model = pd.merge(left=test_cat,right=test_num,left_index=True,right_index=True)
# + _kg_hide-input=true
scaler = StandardScaler().fit(test_model)
scaledtest_act = pd.DataFrame(scaler.fit_transform(test_model),
columns = test_model.columns, index = test_model.index)
# summarize transformed data
scaledtest_act.dropna(how='any', inplace=True)
# + _kg_hide-input=true
#Reducing the original re-scaled dataframe to 2 dimensions using tSNE
# Based on the number of components, the final X_train svd is executed
test_svd = svd_red(dataset=test_model,comps=60)
tsne = TSNE(n_components=2, random_state=0)
Z = tsne.fit_transform(test_svd)
tsne_test = pd.DataFrame(Z, columns=['x','y'], index=test_svd.index)
# -
# #### [Back to Contents](#contents)
# #### <a id='finres'> Dimension reduction results </a>
# + _kg_hide-input=true
#Use the entire train and test set for training the lasso model for final result.
total_data_tsne = pd.concat([dftsne,dftsne_test])
#Seperate the Signal from the Independent variables.
Y = total_data_tsne.signal
X = total_data_tsne[['x','y']]
lasso = Lasso()
lasso.fit(X, Y)
#After 3 notebooks, the results arrive by this seemingly naive command
test_result = lasso.predict(tsne_test)
# + _kg_hide-input=true
my_submission_1 = pd.DataFrame({'id': test_main.id, 'site_eui': test_result})
# you could use any filename. We choose submission here
my_submission_1.to_csv('submission.csv', index=False)
# -
# ##### <a id='FinCon'> Lessons of the Exercise </a>
#
# The [results](#peek3) are downright strange. The test and training error are almost 0. Is this even possible? Public score was 62!!!!. Not any better. But there are lessons learnt
#
#
# The [simple errors](#error) in filling null values can take away precious time. In the below cells there is place where I fill the null values of the energy_star_rating with median value. Initially, I had it all wrong. Due to which, I was unable to execute the tSNE algorithms.
#
# 1) The important point learnt is using the tSNE and SVD for transforming the data in order to improve the final results
#
# 2) Visualising tSNE of the data gives a different perspective about the kind of data, and the space in which the data is present.
#
# 3) Challenge of working with the correct data, and creating functions and loops to automate the process.
#
# I came across the Ubiquant Competition. The dataset in that competition has 300 features, like the WIDS. https://www.kaggle.com/lonnieqin/ubiquant-market-prediction-with-dnn?kernelSessionId=85994078
#
# Next I am planning to try the DNN from this notebook. There are some modifications done to architecture. But the dense layers working on the main features have been kept same
# #### [Back to Contents](#contents)
# #### <a id='#dnn'> DNN with Encoded Dataset </a>
#
# #### [Model Building](#dmod)
#
# #### [Fitting model ](#dfit)
#
# #### [Result & Understanding](#dres)
# + _kg_hide-input=true
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow import keras
# -
# #### <a id='dmod'> DNN Model Building </a>
def preprocess(X, y):
print(X)
print(y)
return X, y
def make_dataset(feature, y, batch_size=1024, mode="train"):
ds = tf.data.Dataset.from_tensor_slices((feature, y))
ds = ds.map(preprocess)
if mode == "train":
ds = ds.shuffle(256)
ds = ds.batch(batch_size).cache().prefetch(tf.data.experimental.AUTOTUNE)
return ds
#In the model, the integerlook up model tree is not required
def get_model():
features_inputs = tf.keras.Input((127, ), dtype=tf.float16) # feature input of the WIDS
feature_x = layers.Dense(64, activation='relu')(features_inputs)
feature_x = layers.Dense(64, activation='relu')(feature_x)
feature_x = layers.Dense(64, activation='relu')(feature_x)
feature_x = layers.Dense(32, activation='relu')(feature_x)
feature_x = layers.Dense(16, activation='relu')(feature_x)
output = layers.Dense(1)(feature_x)
rmse = keras.metrics.RootMeanSquaredError(name="rmse")
model = tf.keras.Model(inputs=features_inputs, outputs=output)
model.compile(optimizer=tf.optimizers.Adam(0.0001), loss='mse', metrics=['mse', "mae", "mape", rmse])
return model
# + _kg_hide-input=true
model = get_model()
model.summary()
# + _kg_hide-input=true
keras.utils.plot_model(model, show_shapes=True)
# -
# #### <a id='dfit'> DNN Model Fitting </a>
# +
X = data_model.iloc[:,:-2]
Y = data_model.site_eui
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X)
rescaledData = pd.DataFrame(scaler.fit_transform(X),
columns = X.columns, index = X.index)
# summarize transformed data
X.dropna(how='any', inplace=True)
rescaledData.dropna(how='any', inplace=True)
# -
# %%time
from sklearn.model_selection import KFold
kfold = KFold(5, shuffle=True, random_state=42)
models = []
for index, (train_indices, valid_indices) in enumerate(kfold.split(rescaledData)):
X_train, X_val = rescaledData.iloc[train_indices], rescaledData.iloc[valid_indices]
y_train, y_val = Y.iloc[train_indices], Y.iloc[valid_indices]
train_ds = make_dataset(X_train, y_train)
valid_ds = make_dataset(X_val, y_val, mode="valid")
model = get_model()
checkpoint = keras.callbacks.ModelCheckpoint(f"model_{index}.tf", save_best_only=True, save_weights_only=True)
early_stop = keras.callbacks.EarlyStopping(patience=5)
history = model.fit(train_ds, epochs=100, validation_data=valid_ds, callbacks=[checkpoint, early_stop])
models.append(model)
pd.DataFrame(history.history, columns=["mse", "val_mse"]).plot()
plt.title("MSE")
plt.show()
pd.DataFrame(history.history, columns=["mae", "val_mae"]).plot()
plt.title("MAE")
plt.show()
pd.DataFrame(history.history, columns=["rmse", "val_rmse"]).plot()
plt.title("RMSE")
plt.show()
del X_train
del X_val
del y_train
del y_val
del train_ds
del valid_ds
gc.collect()
break
# #### <a id='dres'> Results and checking </a>
def preprocess_test(feature):
return (feature), 0
def make_test_dataset(feature, batch_size=1024):
ds = tf.data.Dataset.from_tensor_slices((feature))
ds = ds.map(preprocess_test)
ds = ds.batch(batch_size).cache().prefetch(tf.data.experimental.AUTOTUNE)
return ds
def inference(models, ds):
y_preds = []
for model in models:
y_pred = model.predict(ds)
y_preds.append(y_pred)
return np.mean(y_preds, axis=0)
test_ds = make_test_dataset(scaledtest_act)
y_pred = inference(models,test_ds)
my_submission= pd.DataFrame({'id': test_main.id, 'site_eui': y_pred.flatten()})
# you could use any filename. We choose submission here
my_submission.to_csv('submission.csv', index=False)
my_submission.head(2)
# #### [Back to Contents](#contents)
| feature-engineering-nn-modeling-wids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn as sk
from sklearn.neighbors import KNeighborsRegressor
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
# -
# Набор данных взят с https://www.kaggle.com/aungpyaeap/fish-market
# Параметры нескольких популярных промысловых рыб
# length 1 = Body height
# length 2 = Total Length
# length 3 = Diagonal Length
fish_data = pd.read_csv("datasets/Fish.csv", delimiter=',')
print(fish_data)
# +
# Выделим входные параметры и целевое значение
x_labels = ['Height', 'Width']
y_label = 'Weight'
data = fish_data[x_labels + [y_label]]
print(data)
# -
# Определим размер валидационной и тестовой выборок
val_test_size = round(0.2*len(data))
print(val_test_size)
# Генерируем уникальный seed
my_code = "Соловьёв"
seed_limit = 2 ** 32
my_seed = int.from_bytes(my_code.encode(), "little") % seed_limit
# Создадим обучающую, валидационную и тестовую выборки
random_state = my_seed
train_val, test = train_test_split(data, test_size=val_test_size, random_state=random_state)
train, val = train_test_split(train_val, test_size=val_test_size, random_state=random_state)
print(len(train), len(val), len(test))
# +
# Выделим обучающую, валидационную и тестовую выборки
train_x = train[x_labels]
train_y = np.array(train[y_label]).reshape(-1,1)
val_x = val[x_labels]
val_y = np.array(val[y_label]).reshape(-1,1)
test_x = test[x_labels]
test_y = np.array(test[y_label]).reshape(-1,1)
# +
# Нормируем значения параметров
scaler_x = MinMaxScaler()
scaler_x.fit(train_x)
scaled_train_x = scaler_x.transform(train_x)
scaler_y = MinMaxScaler()
scaler_y.fit(train_y)
scaled_train_y = scaler_y.transform(train_y)
# -
# Создадим модель метода k-ближайших соседей и обучим ее на нормированных данных. По умолчанию k = 5.
minmse=10
mink=0
a=[]
scaled_val_x = scaler_x.transform(val_x)
scaled_val_y = scaler_y.transform(val_y)
for k in range(1,51):
model1 = KNeighborsRegressor(n_neighbors = k)
model1.fit(scaled_train_x, scaled_train_y)
val_predicted = model1.predict(scaled_val_x)
mse1 = mean_squared_error(scaled_val_y, val_predicted)
a.append(mse1)
if mse1<minmse:
minmse=mse1
mink=k
print("Минимальная среднеквадратичная ошибка",minmse)
print("Значение k, соответствующее минимальной среднеквадратичной ошибке",mink)
print()
print(a)
model1 = KNeighborsRegressor(n_neighbors = 4)
model1.fit(scaled_train_x, scaled_train_y)
val_predicted = model1.predict(scaled_val_x)
mse1 = mean_squared_error(scaled_val_y, val_predicted)
print(mse1)
# +
# Проверим результат на тестевойой выборке.
scaled_test_x = scaler_x.transform(test_x)
scaled_test_y = scaler_y.transform(test_y)
test_predicted = model1.predict(scaled_test_x)
mse2 = mean_squared_error(scaled_test_y,test_predicted)
print(mse2)
# -
| 2020 Осенний семестр/Практическое задание 7/Соловьёв -задание 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="0wMBwdaUE4e7"
from google.colab import drive
drive.mount('/content/MyDrive/')
# + id="OgIsFjHNcq7L"
# !pip install turicreate
# + id="AIW5s8uURguL"
import turicreate
# + id="OUSfNPyYdMFj"
#!7z x '/content/MyDrive/MyDrive/Colab_Books/amazon_baby.sframe.zip'
# + id="XlTfyhP2fEeN"
sf = turicreate.SFrame('/content/MyDrive/MyDrive/Amazon/amazon_baby.sframe')
# + id="7wmFgAJIfT9i"
sf.head(3)
# + id="oGCZZ2aYfU0h"
sf.shape
# + id="DB2QpaJEfaIb"
#sf.export_csv('/content/MyDrive/MyDrive/Amazon/amazon_baby.csv')
# + id="orfiq463STIT"
selected_words = ['awesome', 'great', 'fantastic', 'amazing', 'love', 'horrible', 'bad', 'terrible', 'awful', 'wow', 'hate']
# + id="ih21oYr2J2yn"
#Make the word_count column to count each word in each row of reviews.
sf['word_count'] = turicreate.text_analytics.count_words(sf['review'])
# + colab={"base_uri": "https://localhost:8080/", "height": 166} id="cY8_cZgpK_VZ" outputId="05f682c2-501f-40d6-d970-9440e8902e96"
sf.head(3)
# + id="Y3YPh5usUar3"
for word in selected_words:
sf[word] = sf['word_count'].apply(lambda x: x[word] if word in x else 0)
# + id="GwUF3iOrTkVf" colab={"base_uri": "https://localhost:8080/", "height": 245} outputId="d233c8a9-b783-4603-9623-1c5dcbc162e2"
sf.head(3)
# + id="DiQ45pD_LDOU" colab={"base_uri": "https://localhost:8080/", "height": 245} outputId="947048d9-04b7-411a-b17b-6f2944b2f289"
sf.tail(3)
# + colab={"base_uri": "https://localhost:8080/"} id="dZqYZuMyWcgk" outputId="1a015cb0-34cc-4f89-8058-2051e41a3030"
sf.column_names
# + colab={"base_uri": "https://localhost:8080/"} id="l-RfhVQnWy8H" outputId="a4038bfb-40fb-44db-cb64-108aa01f82a6"
for col in selected_words:
print(col, sf[col].sum())
# + id="4VFgS3J2Tlfr"
#Ignore the products with rating of 3 (neutral type)
sf = sf[sf['rating'] != 3]
# + colab={"base_uri": "https://localhost:8080/", "height": 245} id="Gfk1ox1fUrnC" outputId="cff197f4-b7d6-420e-e22b-61d015bfdbbb"
sf.head(3)
# + id="oNzU56lLU5d3"
#Define the sentiment score (integer)
sf['sentiment'] = sf['rating'] >= 4
# + colab={"base_uri": "https://localhost:8080/", "height": 245} id="BlxpuTvIVEh-" outputId="98a86ac7-8328-45bc-fe5b-ce1298cc99f6"
sf.head(3)
# + id="a7xZN7H2gL3r"
train_data, test_data = sf.random_split(.8, seed=0)
# + id="kCPM1MilgRJg" colab={"base_uri": "https://localhost:8080/", "height": 364} outputId="1958a870-04c9-40da-ec66-942df2b56a16"
#Create the Logistic classifier model
selected_words_model = turicreate.logistic_classifier.create(train_data, target = 'sentiment', features = selected_words,
validation_set = test_data)
# + colab={"base_uri": "https://localhost:8080/"} id="C5CPR_N2Vnz1" outputId="61cb3ecb-8509-4e88-bcb3-c0fd0a8e47ec"
selected_words_model.coefficients
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="ArDSHufvWLIR" outputId="9ab952e5-992d-48c0-fc52-6b77ab7b21a2"
selected_words_model.coefficients
# + colab={"base_uri": "https://localhost:8080/"} id="ySmhCgDnWitB" outputId="8fc94d30-1026-4f93-e42e-eb5e3ffd746f"
selected_words_model.evaluate(test_data)
# + id="sOtEZaHtXCci"
diaper_champ_reviews = sf[sf['name'] == 'Baby Trend Diaper Champ']
# + colab={"base_uri": "https://localhost:8080/", "height": 245} id="wH2v0r9mXOwk" outputId="f40c6b6e-9057-4284-ca61-fcff72934e72"
diaper_champ_reviews.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 245} id="Zw0VIh_4XUfz" outputId="e2e94e49-52cf-4fdd-df82-52870b8db7c7"
#To see the most positive sentiment
diaper_champ_reviews = diaper_champ_reviews.sort('sentiment', ascending=False)
diaper_champ_reviews.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 122} id="sMHM2sXFYvY7" outputId="aebccd30-d4fe-483c-c31b-db2e992ebd66"
diaper_champ_reviews['review'][0]
# + colab={"base_uri": "https://localhost:8080/"} id="ozADfjhxZN_g" outputId="71ef0c93-54db-45f5-84f7-d4800e9fc8b8"
selected_words_model.predict(diaper_champ_reviews[0:1], output_type='probability')
# + id="-EmUHQKRjLa4"
turicreate.visualization.histogram(sf['predicted_sentiment'], xlabel='Sentiment Value',
ylabel='Counts', title='Sentiment Graph')
| 2_ML_Combined_Courses/Course/Week-2/Colab_Amazon_Quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="1asyEVYMOq6c" colab_type="code" colab={}
from http import HTTPStatus
from http.client import HTTPException
import requests
import http
import hashlib
import json
# + id="TpIoaJ3bTsTl" colab_type="code" colab={}
def get_http_connection(host):
try:
connection = http.client.HTTPSConnection(host)
return connection
except HTTPException as e:
raise(e)
# + id="db16aDUPV_6W" colab_type="code" colab={}
def fetch(connection, url, body=None, headers={}, encode_chunked=False ):
try:
connection.request('GET', url=url, body=body, headers=headers, encode_chunked=encode_chunked)
response = connection.getresponse()
if response.status == 200 :
return response
except HTTPException as e:
print(e)
return -1
print(response.status)
return -1
# + id="kWz_QRWIQNSF" colab_type="code" colab={}
def fetchJSON(host, url):
connection = get_http_connection(host)
response = fetch(connection, url)
if response.status == 200 :
response_data = response.readlines()
response_data_json = json.loads(response_data[0])
return response_data_json
return -1
# + id="wcQ5AQMSFRov" colab_type="code" colab={}
def stdioFile(path, name, values, operation):
full_path = str(path+name)
with open(full_path, operation) as fileObject:
if operation == 'r' :
result = fileObject.read(values)
if operation in ('w', 'a') :
result = fileObject.write(values)
if not fileObject.closed :
fileObject.close()
return result
# + id="LlLz_YkhYDHE" colab_type="code" colab={}
def decrypt( word, shift, dic ):
new_word = str('')
for letter in word:
position = dic.find(letter)
if position == -1:
new_word += letter
else :
new_position = position + shift
if new_position > len(dic) :
new_position = len(dic) - (new_position % len(dic))
new_word += dic[new_position]
return new_word
# + id="9aWuyxyaFgvV" colab_type="code" colab={}
def upload(path, filename, full_url):
try:
response = requests.post(url = full_url, files = {'answer': open(path+filename, 'rb') })
print(response.status_code)
print(response.json)
print(response.text)
print(response.headers)
except Exception as e:
print(e)
# + id="m_EBAe3pcVol" colab_type="code" colab={}
def main():
#variables
LOCAL_CONSTS = {
"data_output" : "/content/drive/My Drive/tk/",
"api_key" : "tk.txt",
'host' : 'api.codenation.dev',
'url_get' : '/v1/challenge/dev-ps/generate-data?token={SEU_TOKEN}',
'url_post' : '/v1/challenge/dev-ps/submit-solution?token={SEU_TOKEN}',
'dic' : 'abcdefghijklmnopqrstuvwxyz',
'file_name' : 'answer.json'
}
token = stdioFile(LOCAL_CONSTS['data_output'], LOCAL_CONSTS['api_key'] , None , 'r')
url_get = LOCAL_CONSTS['url_get'].format(SEU_TOKEN=token)
full_url_post = 'https://'+LOCAL_CONSTS['host']+LOCAL_CONSTS['url_post'].format(SEU_TOKEN=token)
#in
response_data_json = fetchJSON(LOCAL_CONSTS['host'], url_get)
#transform
decrypited = decrypt(response_data_json['cifrado'], -12, LOCAL_CONSTS['dic'])
decrypited_utf8 = str(decrypited).encode('utf-8')
resume = hashlib.sha1(decrypited_utf8).hexdigest()
#out
response_data_json['decifrado'] = decrypited
response_data_json['resumo_criptografico'] = resume
str_response_data_json = str(response_data_json)
str_response_data_json = str_response_data_json.replace("\'", '\"')
stdioFile(LOCAL_CONSTS['data_output'], LOCAL_CONSTS['file_name'] , str_response_data_json, 'w')
stdioFile(LOCAL_CONSTS['data_output'], LOCAL_CONSTS['file_name'] , None , 'r')
#upload
file_path =LOCAL_CONSTS['data_output']+LOCAL_CONSTS['file_name']
upload(LOCAL_CONSTS['data_output'], LOCAL_CONSTS['file_name'] ,full_url_post)
if __name__=="__main__":
main()
| use_sha1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.metrics import classification_report, accuracy_score, f1_score
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyser = SentimentIntensityAnalyzer()
df = pd.read_csv('Data/movie_reviews.csv',encoding = "ISO-8859-1")
df.head()
sents = np.zeros((df.shape[0]))
for j in range(df.shape[0]):
score = analyser.polarity_scores(df['SentimentText'][j])
sents[j] = score['compound']
y_hat = sents
y_true = df['Sentiment'].values
# +
# Error Analysis
# -
y_hat3 = (y_hat > 0).astype('int')
print(classification_report(y_true, y_hat3))
print("Accuracy Score :", accuracy_score(y_true, y_hat3))
print("F-1 Score :", f1_score(y_true, y_hat3))
print("Confusion Matrix :", classification_report(y_true, y_hat3))
| Sentiment_Analysis_using_VaderSentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from splinter import Browser
from bs4 import BeautifulSoup as bs
import os
import pandas as pd
import requests
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
# +
# NASA Mars News
# -
news_url = 'https://mars.nasa.gov/news/'
browser.visit(news_url)
html = browser.html
soup = bs(html,"html.parser")
news_title = soup.find("div",class_="content_title").text
news_p = soup.find("div", class_="article_teaser_body").text
print(f"Title: {news_title}")
print(f"Para: {news_p}")
# +
# JPL Mars Space Images - Featured Image
# +
image_url = ('https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars')
response = requests.get(image_url)
soup = bs(response.text, 'html.parser')
images = soup.find_all('a', class_="fancybox")
for image in images:
featured_image_url = image['data-fancybox-href']
featured_image_url = 'https://www.jpl.nasa.gov' + featured_image_url
featured_image_url
# +
# Mars Weather
# -
weather_url = "https://twitter.com/marswxreport?lang=en"
browser.visit(weather_url)
html_weather = browser.html
soup = bs(html_weather, "html.parser")
mars_weather = soup.find(class_="css-1dbjc4n r-1iusvr4 r-16y2uox r-1777fci r-5f2r5o r-1mi0q7o").text
print(mars_weather)
# +
# Mars Facts
# -
facts_url = "https://space-facts.com/mars/"
table = pd.read_html(facts_url)
table[0]
mars_facts = table[0]
mars_facts.columns = ["Parameter", "Values"]
mars_facts.set_index(["Parameter"])
mars_facts_html = mars_facts.to_html()
mars_facts_html = mars_facts_html.replace("\n", "")
mars_facts_html
# +
# <NAME>
# -
hemisphere_image_urls = []
# +
cerberus_url = ('https://astrogeology.usgs.gov/search/map/Mars/Viking/cerberus_enhanced')
response = requests.get(cerberus_url)
soup = bs(response.text, 'html.parser')
cerberus_img = soup.find_all('div', class_="wide-image-wrapper")
for img in cerberus_img:
pic = img.find('li')
img_url = pic.find('a')['href']
title = soup.find('h2', class_='title').text
cerberus = {"title": title, "img_url": img_url}
print(cerberus)
hemisphere_image_urls.append(cerberus)
# +
schiaparelli_url = ('https://astrogeology.usgs.gov/search/map/Mars/Viking/schiaparelli_enhanced')
response = requests.get(schiaparelli_url)
soup = bs(response.text, 'html.parser')
shiaparelli_img = soup.find_all('div', class_="wide-image-wrapper")
for img in shiaparelli_img:
pic = img.find('li')
img_url = pic.find('a')['href']
title = soup.find('h2', class_='title').text
shiaparelli = {"title": title, "img_url": img_url}
print(shiaparelli)
hemisphere_image_urls.append(shiaparelli)
# +
syrtis_major_url = ('https://astrogeology.usgs.gov/search/map/Mars/Viking/syrtis_major_enhanced')
response = requests.get(syrtis_major_url)
soup = bs(response.text, 'html.parser')
syrtris_img = soup.find_all('div', class_="wide-image-wrapper")
for img in syrtris_img:
pic = img.find('li')
img_url = pic.find('a')['href']
title = soup.find('h2', class_='title').text
syrtris = {"title": title, "img_url": img_url}
print(syrtris)
hemisphere_image_urls.append(syrtris)
# +
valles_marineris_url = ('https://astrogeology.usgs.gov/search/map/Mars/Viking/valles_marineris_enhanced')
response = requests.get(valles_marineris_url)
soup = bs(response.text, 'html.parser')
valles_marineris_img = soup.find_all('div', class_="wide-image-wrapper")
for img in valles_marineris_img:
pic = img.find('li')
img_url = pic.find('a')['href']
title = soup.find('h2', class_='title').text
valles_marineris = {"title": title, "img_url": img_url}
print(valles_marineris)
hemisphere_image_urls.append(valles_marineris)
# -
print(hemisphere_image_urls)
| mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import psycopg2
from sqlalchemy import create_engine
url = 'postgresql+psycopg2://postgres:spark123@localhost:5432/amazon'
engine = create_engine(url)
reviews_df = pd.read_sql_table('reviews', con=engine)
products_df = pd.read_sql_table('products', con=engine)
also_buy_df = pd.read_sql_table('also_buy', con=engine)
also_view_df = pd.read_sql_table('also_view', con=engine)
categories_df = pd.read_sql_table('categories', con=engine)
products_description_df = pd.read_sql_table('products_description', con=engine)
products_feature_df = pd.read_sql_table('products_feature', con=engine)
products_images_df = pd.read_sql_table('products_images', con=engine)
reviews_df.to_csv('./web_app/data/reviews.csv', index=False)
products_df.to_csv('./web_app/data/products.csv', index=False)
also_buy_df.to_csv('./web_app/data/also_buy.csv', index=False)
also_view_df.to_csv('./web_app/data/also_view.csv', index=False)
categories_df.to_csv('./web_app/data/categories.csv', index=False)
products_description_df.to_csv('./web_app/data/products_description.csv', index=False)
products_feature_df.to_csv('./web_app/data/products_feature.csv', index=False)
products_images_df.to_csv('./web_app/data/products_images.csv', index=False)
| spark_workspace/save_as_csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# author = Alexandros
import os
result_files = []
#/home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/results
for result_file in os.listdir("/home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/results"):
#print(result_file)
result_files.append(result_file)
#print(result_files)
for i, result_file in enumerate(result_files):
#print("trec_eval -q tar.qrels /results/"+result_file+" > "+result_file+"_trec_eval_q_measurements")
print("trec_eval tar.qrels /results/"+result_file+" > "+result_file+"_trec_eval_measurements")
print('\n')
#os.system("trec_eval -q tar.qrels results/"+result_file+" > "+result_file+"_trec_eval_q_measurements")
os.system("trec_eval tar.qrels results/"+result_file+" > "+result_file+"_trec_eval_measurements")
#print("python /home/alexandros/netbeans-8.2/NetBeansProjects/tar-master/scripts/tar_eval.py /home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/tar.qrels /home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/results/"+result_file+" > "+result_file+"_tar_eval_measurements")
#print('\n')
#os.system("python /home/alexandros/netbeans-8.2/NetBeansProjects/tar-master/scripts/tar_eval.py /home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/tar.qrels /home/alexandros/netbeans-8.2/NetBeansProjects/lucene4ir-master/data/pubmed/results/"+result_file+" > "+result_file+"_tar_eval_measurements")
| trec_and_tar_eval_automation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="fN_C6ZXasPot"
# # Variational AutoEncoder
#
# **Author:** [fchollet](https://twitter.com/fchollet)<br>
# **Date created:** 2020/05/03<br>
# **Last modified:** 2020/05/03<br>
# **Description:** Convolutional Variational AutoEncoder (VAE) trained on MNIST digits.
# + [markdown] id="FtTC_x9DsPoz"
# ## Setup
# + id="lXxXdO2ZsPoz"
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# + [markdown] id="DLdOl32QsPo0"
# ## Create a sampling layer
# + id="i-D6ucsKsPo0"
class Sampling(layers.Layer):
"""Uses (z_mean, z_log_var) to sample z, the vector encoding a digit."""
def call(self, inputs):
z_mean, z_log_var = inputs
batch = tf.shape(z_mean)[0]
dim = tf.shape(z_mean)[1]
epsilon = tf.keras.backend.random_normal(shape=(batch, dim))
return z_mean + tf.exp(0.5 * z_log_var) * epsilon
# + [markdown] id="HYjerB4-sPo1"
# ## Build the encoder
# + id="X1vW1b5KsPo1" outputId="35343bcb-c314-491b-e259-65816ee394d5" colab={"base_uri": "https://localhost:8080/"}
latent_dim = 2
encoder_inputs = keras.Input(shape=(28, 28, 1))
x = layers.Conv2D(32, 3, activation="relu", strides=2, padding="same")(encoder_inputs)
x = layers.Conv2D(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Flatten()(x)
x = layers.Dense(16, activation="relu")(x)
z_mean = layers.Dense(latent_dim, name="z_mean")(x)
z_log_var = layers.Dense(latent_dim, name="z_log_var")(x)
z = Sampling()([z_mean, z_log_var])
encoder = keras.Model(encoder_inputs, [z_mean, z_log_var, z], name="encoder")
encoder.summary()
# + [markdown] id="1SgIldVVsPo2"
# ## Build the decoder
# + id="63RDQcyosPo2" outputId="f64a67e1-8edd-4af6-98a8-ea0c7cb52f41" colab={"base_uri": "https://localhost:8080/"}
latent_inputs = keras.Input(shape=(latent_dim,))
x = layers.Dense(7 * 7 * 64, activation="relu")(latent_inputs)
x = layers.Reshape((7, 7, 64))(x)
x = layers.Conv2DTranspose(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Conv2DTranspose(32, 3, activation="relu", strides=2, padding="same")(x)
decoder_outputs = layers.Conv2DTranspose(1, 3, activation="sigmoid", padding="same")(x)
decoder = keras.Model(latent_inputs, decoder_outputs, name="decoder")
decoder.summary()
# + [markdown] id="HY4QD-N_sPo2"
# ## Define the VAE as a `Model` with a custom `train_step`
# + id="EGoOnnhysPo2"
class VAE(keras.Model):
def __init__(self, encoder, decoder, **kwargs):
super(VAE, self).__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
self.total_loss_tracker = keras.metrics.Mean(name="total_loss")
self.reconstruction_loss_tracker = keras.metrics.Mean(
name="reconstruction_loss"
)
self.kl_loss_tracker = keras.metrics.Mean(name="kl_loss")
@property
def metrics(self):
return [
self.total_loss_tracker,
self.reconstruction_loss_tracker,
self.kl_loss_tracker,
]
def train_step(self, data):
with tf.GradientTape() as tape:
z_mean, z_log_var, z = self.encoder(data)
reconstruction = self.decoder(z)
reconstruction_loss = tf.reduce_mean(
tf.reduce_sum(
keras.losses.binary_crossentropy(data, reconstruction), axis=(1, 2)
)
)
kl_loss = -0.5 * (1 + z_log_var - tf.square(z_mean) - tf.exp(z_log_var))
kl_loss = tf.reduce_mean(tf.reduce_sum(kl_loss, axis=1))
total_loss = reconstruction_loss + kl_loss
grads = tape.gradient(total_loss, self.trainable_weights)
self.optimizer.apply_gradients(zip(grads, self.trainable_weights))
self.total_loss_tracker.update_state(total_loss)
self.reconstruction_loss_tracker.update_state(reconstruction_loss)
self.kl_loss_tracker.update_state(kl_loss)
return {
"loss": self.total_loss_tracker.result(),
"reconstruction_loss": self.reconstruction_loss_tracker.result(),
"kl_loss": self.kl_loss_tracker.result(),
}
# + [markdown] id="x8HDICqRsPo3"
# ## Train the VAE
# + id="8LX1QiRjsPo4" outputId="1cf59db2-76c5-4d0c-e6c1-bdba770e48dc" colab={"base_uri": "https://localhost:8080/"}
(x_train, _), (x_test, _) = keras.datasets.mnist.load_data()
mnist_digits = np.concatenate([x_train, x_test], axis=0)
mnist_digits = np.expand_dims(mnist_digits, -1).astype("float32") / 255
vae = VAE(encoder, decoder)
vae.compile(optimizer=keras.optimizers.Adam())
vae.fit(mnist_digits, epochs=30, batch_size=128)
# + [markdown] id="51nB6F0rsPo4"
# ## Display a grid of sampled digits
# + id="S2UAIiP_sPo4" outputId="cbe2cd7c-d6a6-4a6a-ab2c-87cfc5e7b1b7" colab={"base_uri": "https://localhost:8080/", "height": 877}
import matplotlib.pyplot as plt
def plot_latent_space(vae, n=30, figsize=15):
# display a n*n 2D manifold of digits
digit_size = 28
scale = 1.0
figure = np.zeros((digit_size * n, digit_size * n))
# linearly spaced coordinates corresponding to the 2D plot
# of digit classes in the latent space
grid_x = np.linspace(-scale, scale, n)
grid_y = np.linspace(-scale, scale, n)[::-1]
for i, yi in enumerate(grid_y):
for j, xi in enumerate(grid_x):
z_sample = np.array([[xi, yi]])
x_decoded = vae.decoder.predict(z_sample)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[
i * digit_size : (i + 1) * digit_size,
j * digit_size : (j + 1) * digit_size,
] = digit
plt.figure(figsize=(figsize, figsize))
start_range = digit_size // 2
end_range = n * digit_size + start_range
pixel_range = np.arange(start_range, end_range, digit_size)
sample_range_x = np.round(grid_x, 1)
sample_range_y = np.round(grid_y, 1)
plt.xticks(pixel_range, sample_range_x)
plt.yticks(pixel_range, sample_range_y)
plt.xlabel("z[0]")
plt.ylabel("z[1]")
plt.imshow(figure, cmap="Greys_r")
plt.show()
plot_latent_space(vae)
# + [markdown] id="XXYDed8ZsPo6"
# ## Display how the latent space clusters different digit classes
# + id="XVX_I6susPo6"
def plot_label_clusters(vae, data, labels):
# display a 2D plot of the digit classes in the latent space
z_mean, _, _ = vae.encoder.predict(data)
plt.figure(figsize=(12, 10))
plt.scatter(z_mean[:, 0], z_mean[:, 1], c=labels)
plt.colorbar()
plt.xlabel("z[0]")
plt.ylabel("z[1]")
plt.show()
(x_train, y_train), _ = keras.datasets.mnist.load_data()
x_train = np.expand_dims(x_train, -1).astype("float32") / 255
plot_label_clusters(vae, x_train, y_train)
| Q3/batch1/DeepLearning Part -2/Chapter8/autoencoder/vae.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3
# ---
pip install pandas dash
pip install wget
# !wget "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/datasets/spacex_launch_dash.csv"
# !wget "https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/labs/module_3/spacex_dash_app.py"
# !python3 spacex_dash_app.py
# +
import pandas as pd
import dash
import dash_html_components as html
import dash_core_components as dcc
from dash.dependencies import Input, Output
import plotly.express as px
spacex_df = pd.read_csv("spacex_launch_dash.csv")
max_payload = spacex_df['Payload Mass (kg)'].max()
min_payload = spacex_df['Payload Mass (kg)'].min()
app = dash.Dash(__name__)
uniquelaunchsites = spacex_df['Launch Site'].unique().tolist()
lsites = []
lsites.append({'label': 'All Sites', 'value': 'All Sites'})
for site in uniquelaunchsites:
lsites.append({'label': site, 'value': site})
app.layout = html.Div(children=[html.H1('SpaceX Launch Records Dashboard',
style={'textAlign': 'center', 'color': '#503D36',
'font-size': 40}),
dcc.Dropdown(id='site_dropdown',options=lsites,placeholder='Select a Launch Site here', searchable = True , value = 'All Sites'),
html.Br(),
html.Div(dcc.Graph(id='success-pie-chart')),
html.Br(),
html.P("Payload range (Kg):"),
dcc.RangeSlider(
id='payload_slider',
min=0,
max=10000,
step=1000,
marks = {
0: '0 kg',
1000: '1000 kg',
2000: '2000 kg',
3000: '3000 kg',
4000: '4000 kg',
5000: '5000 kg',
6000: '6000 kg',
7000: '7000 kg',
8000: '8000 kg',
9000: '9000 kg',
10000: '10000 kg'
},
value=[min_payload,max_payload]
),
html.Div(dcc.Graph(id='success-payload-scatter-chart')),
])
@app.callback(
Output(component_id='success-pie-chart',component_property='figure'),
[Input(component_id='site_dropdown',component_property='value')]
)
def update_graph(site_dropdown):
if (site_dropdown == 'All Sites'):
df = spacex_df[spacex_df['class'] == 1]
fig = px.pie(df, names = 'Launch Site',hole=.3,title = 'Total Success Launches By all sites')
else:
df = spacex_df.loc[spacex_df['Launch Site'] == site_dropdown]
fig = px.pie(df, names = 'class',hole=.3,title = 'Total Success Launches for site '+site_dropdown)
return fig
@app.callback(
Output(component_id='success-payload-scatter-chart',component_property='figure'),
[Input(component_id='site_dropdown',component_property='value'),Input(component_id="payload_slider", component_property="value")]
)
def update_scattergraph(site_dropdown,payload_slider):
if site_dropdown == 'All Sites':
low, high = payload_slider
df = spacex_df
mask = (df['Payload Mass (kg)'] > low) & (df['Payload Mass (kg)'] < high)
fig = px.scatter(
df[mask], x="Payload Mass (kg)", y="class",
color="Booster Version",
size='Payload Mass (kg)',
hover_data=['Payload Mass (kg)'])
else:
low, high = payload_slider
df = spacex_df.loc[spacex_df['Launch Site'] == site_dropdown]
mask = (df['Payload Mass (kg)'] > low) & (df['Payload Mass (kg)'] < high)
fig = px.scatter(
df[mask], x="Payload Mass (kg)", y="class",
color="Booster Version",
size='Payload Mass (kg)',
hover_data=['Payload Mass (kg)'])
return fig
if __name__ == '__main__':
app.run_server()
| Plotly Dash lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import qcodes
import matplotlib.pyplot as plt
from functools import partial
from qcodes.plots.qcmatplotlib import MatPlot
from qcodes.plots.pyqtgraph import QtPlot
from scipy.optimize import curve_fit
import scipy.integrate as integrate
import pandas as pd
class AlazarTech():
def __init__(self,param):
self.name = param.get('name')
self.fs = param.get('sampling_rate')
self.ts = 1/self.fs
self.record_length = param.get('record_length')
self.channel_range = param.get('channel_range')
def alazar_bit2volt(self, signal):
return ()
adc_param_CryoRX = {
"name" : 'CryoRX', # For setting ...
"sampling_rate" : 1e9, # samples/s
"record_length" : 0.002, # in (s)
"channel_range" : 0.1 # in (v)
}
pandas.pd
alazar = AlazarTech(adc_param_CryoRX)
alazar.name
| Qubit Measurement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# name: python38164bited0cfbe7c3504f05a4d155e3328c3124
# ---
# # Combination Sum
#
# Given a set of candidate numbers (candidates) (without duplicates) and a target number (target), find all unique combinations in candidates where the candidate numbers sums to target.
#
# The same repeated number may be chosen from candidates unlimited number of times.
#
# ## 解析
#
# 题目来源:[LeetCode - Combination Sum - 39](https://leetcode.com/problems/combination-sum/)
#
# 典型的求子集+过滤子集的套路,使用递归求子集并且使用回溯法去重即可。
def combinationSum(candidates, target):
result = []
seen = []
def find(target, select):
for option in candidates:
if (target - sum(select) - option) > 0:
find(target, select + [option])
elif (target - sum(select) - option) == 0:
seq = sorted(select + [option])
if seq not in seen:
result.append(seq)
seen.append(seq)
find(target, [])
return result
# + tags=[]
print(combinationSum([2,3,6,7],7))
| Combination-Sum.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(qvalue)
library(dplyr)
library(ggplot2)
library(cowplot)
# common genes, i.e. genes tested for all aggregations
df_common_genes = read.csv("/hps/nobackup/stegle/users/acuomo/all_scripts/sc_eqtl/common_genes_across_all_aggregation_methods_and_bulk.csv")
common_genes = as.character(df_common_genes$gene)
length(common_genes)
# highly variable genes (top 50% CV)
Giordanos_selection = "/hps/nobackup/stegle/users/galvari/data/iPSCs/singleCell/metadata/ensembl_gene/Ensembl_75_Gene_CV_quant5.txt"
G_file = read.csv(Giordanos_selection, sep = "\t")
nrow(G_file)
head(G_file,2)
#### load bulk results
##### a-bulk
bulk_folder = "/hps/nobackup/hipsci/scratch/ComparingQtlMapping/Bulk/BulkTotal_PCA20/"
# a-bulk lead SNP only
leads_all = read.csv(paste0(bulk_folder,"top_qtl_results_all.txt"), sep = "\t")
leads_all = leads_all[leads_all$feature_id %in% common_genes,]
leads_all$q_value = qvalue(leads_all$empirical_feature_p_value)$qvalues
leads_all_sign = leads_all[leads_all$q_value < 0.05,]
a_eqtls = unique(paste0(leads_all_sign$feature_id,"-",leads_all_sign$snp_id))
# +
#### starting with 'dr' aggregations (all results)
# -
### dr-mean
dir0 = "/hps/nobackup/hipsci/scratch/ComparingQtlMapping/SingleCell/PseudoBulk/mean/Run_Output_PCA20_88_log_TPM_scater_libsize_206/"
df0 = read.csv(paste0(dir0,"qtl_results_all.txt"), sep="\t")
df0 = df0[df0$feature_id %in% common_genes,]
head(df0,2)
### dr-median
dir1 = "/hps/nobackup/hipsci/scratch/ComparingQtlMapping/SingleCell/PseudoBulk/median/Run_Output_PCA20_88_log_TPM_scater_libsize_206/"
df1 = read.csv(paste0(dir1,"qtl_results_all.txt"), sep="\t")
df1 = df1[df0$feature_id %in% common_genes,]
head(df1,2)
### dr-sum
dir2 = "/hps/nobackup/hipsci/scratch/ComparingQtlMapping/SingleCell/PseudoBulk/sum/Run_Output_PCA20_88_log_TPM_edgeR_libsize_206/"
df2 = read.csv(paste0(dir2,"qtl_results_all.txt"), sep="\t")
df2 = df2[df2$feature_id %in% common_genes,]
head(df2,2)
# +
# then assess replication of bulk eQTL in single-cell aggregation results
# -
# significant a-bulk replicated in sc-eqtl (dr-mean)
df0_bulk = inner_join(df0, leads_all_sign, by = c("snp_id","feature_id"), suffix = c(".sc",".bulk"))
df0_bulk_repl = df0_bulk[(df0_bulk$p_value.sc < 0.05 &
(df0_bulk$beta.sc*df0_bulk$beta.bulk)>0),]
a_eqtls_mean = unique(paste0(df0_bulk_repl$feature_id,"-",df0_bulk_repl$snp_id))
length(a_eqtls_mean)
# significant a-bulk replicated in sc-eqtl (dr-median)
df1_bulk = inner_join(df1, leads_all_sign, by = c("snp_id","feature_id"), suffix = c(".sc",".bulk"))
df1_bulk_repl = df1_bulk[(df1_bulk$p_value.sc < 0.05 &
(df1_bulk$beta.sc*df1_bulk$beta.bulk)>0),]
a_eqtls_median = unique(paste0(df1_bulk_repl$feature_id,"-",df1_bulk_repl$snp_id))
length(a_eqtls_median)
# significant a-bulk replicated in sc-eqtl (dr-sum)
df2_bulk = inner_join(df2, leads_all_sign, by = c("snp_id","feature_id"), suffix = c(".sc",".bulk"))
df2_bulk_repl = df2_bulk[(df2_bulk$p_value.sc < 0.05 &
(df2_bulk$beta.sc*df2_bulk$beta.bulk)>0),]
a_eqtls_sum = unique(paste0(df2_bulk_repl$feature_id,"-",df2_bulk_repl$snp_id))
length(a_eqtls_sum)
# +
# next, define the different catagories:
# 1. bulk eQTL replicated in all three (dr-mean, dr-median and dr-sum)
# -
eqtls1 = a_eqtls_mean[(a_eqtls_mean %in% a_eqtls_median) & (a_eqtls_mean %in% a_eqtls_sum)]
length(eqtls1)
# +
# 2. bulk eQTL replicated in dr-mean, dr-median but NOT dr-sum
# 3. bulk eQTL replicated in dr-mean, dr-sum but NOT dr-median
# 4. bulk eQTL replicated in dr-median, dr-sum but NOT dr-mean
# -
eqtls2 = a_eqtls_mean[(a_eqtls_mean %in% a_eqtls_median) & !(a_eqtls_mean %in% a_eqtls_sum)]
eqtls3 = a_eqtls_mean[!(a_eqtls_mean %in% a_eqtls_median) & (a_eqtls_mean %in% a_eqtls_sum)]
eqtls4 = a_eqtls_median[!(a_eqtls_median %in% a_eqtls_mean) & (a_eqtls_median %in% a_eqtls_sum)]
length(eqtls2)
length(eqtls3)
length(eqtls4)
# +
# 5. bulk eQTL replicated ONLY in dr-mean
# 6. bulk eQTL replicated ONLY in dr-median
# 7. bulk eQTL replicated ONLY in dr-sum
# -
eqtls5 = a_eqtls_mean[!(a_eqtls_mean %in% a_eqtls_median) & !(a_eqtls_mean %in% a_eqtls_sum)]
eqtls6 = a_eqtls_median[!(a_eqtls_median %in% a_eqtls_mean) & !(a_eqtls_median %in% a_eqtls_sum)]
eqtls7 = a_eqtls_sum[!(a_eqtls_sum %in% a_eqtls_mean) & !(a_eqtls_sum %in% a_eqtls_median)]
length(eqtls5)
length(eqtls6)
length(eqtls7)
# +
# 8. bulk eQTL replicated in none of the three
# -
eqtls8 = a_eqtls[!(a_eqtls %in% a_eqtls_mean) & !(a_eqtls %in% a_eqtls_median) & !(a_eqtls %in% a_eqtls_sum)]
length(eqtls8)
# +
### Collect stats
# -
df = leads_all_sign
df$eqtl = paste0(df$feature_id,"-",df$snp_id)
df_to_plot = df[df$eqtl %in% a_eqtls, c("feature_id","snp_id","eqtl","beta","q_value")]
nrow(df_to_plot)
head(df_to_plot,2)
# define categories
df_to_plot[df_to_plot$eqtl %in% eqtls1,"category"] = "all"
df_to_plot[df_to_plot$eqtl %in% eqtls2,"category"] = "no_sum"
df_to_plot[df_to_plot$eqtl %in% eqtls3,"category"] = "no_median"
df_to_plot[df_to_plot$eqtl %in% eqtls4,"category"] = "no_mean"
df_to_plot[df_to_plot$eqtl %in% eqtls5,"category"] = "only_mean"
df_to_plot[df_to_plot$eqtl %in% eqtls6,"category"] = "only_median"
df_to_plot[df_to_plot$eqtl %in% eqtls7,"category"] = "only_sum"
df_to_plot[df_to_plot$eqtl %in% eqtls8,"category"] = "none"
tail(df_to_plot)
### bulk stats
ab_stats_filename = "/hps/nobackup/stegle/users/acuomo/all_scripts/sc_eqtl/stats_HipSci_passQc.txt"
ab_stats = read.csv(ab_stats_filename, sep = "\t", row.names = 1)
head(ab_stats,2)
ab_stats$feature_id = rownames(ab_stats)
nrow(df_to_plot)
df_to_plot0 = inner_join(df_to_plot, ab_stats, by = "feature_id")
nrow(df_to_plot0)
df_to_plot0$category = factor(df_to_plot0$category, levels = c("all", "no_sum", "no_median", "no_mean",
"only_mean", "only_median","only_sum","none"))
gene_anno_filename = "/hps/nobackup/stegle/users/acuomo/all_scripts/Ensemble_75_Gene_Structures_Exon_Info.txt"
gene_anno = read.csv(gene_anno_filename, sep = "\t")
head(gene_anno,2)
df_genes = gene_anno[,c("Ensembl.Gene.ID","Gene.Start..bp.","Gene.End..bp.")]
colnames(df_genes) = c("feature_id","feature_start","feature_end")
df_genes = df_genes[-which(duplicated(df_genes$feature_id)),]
df_genes$feature_midpoint = df_genes$feature_start + 0.5*(df_genes$feature_end - df_genes$feature_start)
head(df_genes,2)
nrow(df_to_plot0)
df_to_plot1 = inner_join(df_to_plot0, df_genes, by = "feature_id")
nrow(df_to_plot1)
df_to_plot1$snp_pos = matrix(unlist(strsplit(as.character(df_to_plot1$snp_id),"_")), ncol = 4, byrow = T)[,2]
df_to_plot1$dist = as.numeric(df_to_plot1$feature_midpoint) - as.numeric(df_to_plot1$snp_pos)
head(df_to_plot1,2)
df_to_plot0 %>% group_by(category) %>% summarise(eGenes = n()) %>%
ggplot(aes(x = category, y = eGenes, fill = category)) + geom_bar(stat="identity")
# +
options(repr.plot.width = 7, repr.plot.height = 9)
p1 = df_to_plot0 %>% group_by(category) %>% summarise(eGenes = n()) %>%
ggplot(aes(x = category, y = eGenes, fill = category)) + geom_bar(stat="identity") + theme_classic() + xlab("") + theme(legend.position = "none")
+geom_errorbar(aes(ymin=eGenes-stdv, ymax=m+stdv), width=.2,
position=position_dodge(.9)) +
p2 = ggplot(df_to_plot0, aes(x = category, y = log2(MeanTPM_All+1), fill= category)) + geom_violin(colour=NA) + geom_boxplot(width=0.1) + theme_classic() + xlab("") + theme(legend.position = "none")
p3 = ggplot(df_to_plot0, aes(x = category, y = log2(VarTPM_All+1),fill= category)) + geom_violin(colour=NA) + geom_boxplot(width=0.1) + theme_classic() + xlab("") + theme(legend.position = "none")
p4 = ggplot(df_to_plot0, aes(x = category, y = log2(abs(beta)+1), fill= category)) + geom_violin(colour=NA) + geom_boxplot(width=0.1) + theme_classic() + xlab("") + theme(legend.position = "none")
p5 = ggplot(df_to_plot1, aes(x = category, y = log2(abs(dist)+1), fill= category)) + geom_violin(colour=NA) + geom_boxplot(width=0.1) + theme_classic() + theme(legend.position = "none")
plot_grid(p1,p2,p3,p4,p5, ncol=1)
# plot_grid(p1,p4,ncol=1)
# -
type.col <- c(only_mean = "#375e97", no_mean = "#ee7733", only_median = "#fb6542", no_median="#228833",
only_sum = "#ffbb00", no_sum="#AA3377", all="gray", none="gray80")
plot_grid(p1+scale_fill_manual(values = type.col),
p2+scale_fill_manual(values = type.col),
p3+scale_fill_manual(values = type.col),
p4+scale_fill_manual(values = type.col),
p5+scale_fill_manual(values = type.col), ncol=1)
fig_dir = "/hps/nobackup/stegle/users/acuomo/all_scripts/sc_eqtl/figures/"
pdf(paste0(fig_dir,"eqtl_properties_all_aggregations.pdf"), width=7, height=9)
plot_grid(p1+scale_fill_manual(values = type.col),
p2+scale_fill_manual(values = type.col),
p3+scale_fill_manual(values = type.col),
p4+scale_fill_manual(values = type.col),
p5+scale_fill_manual(values = type.col), ncol=1)
dev.off()
# + active=""
#
| ipsc/Figure_S3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Removing Columns
# import pandas
import pandas as pd
# read a dataset of UFO reports into a DataFrame
ufo = pd.read_csv('http://bit.ly/uforeports')
ufo.head()
# ## Remove Single Columns
# remove single columns(axis=1 refers to columns, axis=0 refers to row)
ufo.drop('City', axis=1, inplace=True)
ufo.head()
# ## Remove Multiple Columns
# remove multiple columns
ufo.drop(['Colors Reported', 'Time'], axis=1, inplace=True)
ufo.head()
# read a dataset of top-rated IMDb movies into a DataFrame
movies = pd.read_csv('http://bit.ly/imdbratings')
movies.head()
# remove single column(axis=1 refers to colums)
movies.drop('genre', axis=1, inplace=True)
movies.head()
# remove multiple columns from DataFrame
movies.drop(['actors_list', 'content_rating', 'duration'], axis=1, inplace=True)
movies.head()
# <h3>About the Author</h3>
# This repo was created by <a href="https://www.linkedin.com/in/jubayer28/" target="_blank"><NAME></a> <br>
# <a href="https://www.linkedin.com/in/jubayer28/" target="_blank"><NAME></a> is a student of Microbiology at Jagannath University and the founder of <a href="https://github.com/hdro" target="_blank">Health Data Research Organization</a>. He is also a team member of a bioinformatics research group known as Bio-Bio-1.
#
# <a rel="license" href="http://creativecommons.org/licenses/by-nc-sa/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-nc-sa/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-nc-sa/4.0/">Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License</a>.mm
| book/_build/jupyter_execute/pandas/07-Removing Columns in Pandas .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook calculates gridded maps with mass fractional contribution by sector to total NMVOC for MOZART VOCs species.
import xarray as xr
import numpy as np
import os
import pandas as pd
#data dir paths.
voc_dir='/geos/d21/s1878599/edgarv5_process/monthly_nmvocs4.3.2_mass_MOZART/'
save_dir='/geos/d21/s1878599/edgarv5_process/monthly_nmvocs4.3.2_fractional_MOZART/'
#create save directory if missing.
if not os.path.isdir(save_dir):
# !mkdir -p $save_dir
# ## Read VOCs
#get all vocs files as ordered dictionary of xarray datasets.
def get_vocs_arr(voc_dir):
'''
put all vocs files as ordered dictionary of xarray datasets.
voc_dir: path to where vocs files are.
output: lsit of dataset with individual vocs.
'''
vocs={}
for f in os.listdir(voc_dir):
vname=f.split('_')[3]
vocs.update({vname:xr.open_dataset(voc_dir+f)})
return vocs
vocs=get_vocs_arr(voc_dir)
# get total nmvoc summing all vocs contributions.
tot_nmvoc=sum(vocs.values())
tot_nmvoc
# ## Create fractional maps
for k in list(vocs.keys()):
print(k)
ds=(vocs[k]/tot_nmvoc).fillna(0.0) # divide eac voc mass by total NMVOC mass.
ds.attrs['title']='Monthly fractional contribution of ' + k
ds.to_netcdf(save_dir+'monthly_v432_2010_fraction_'+ k + '_.0.1x0.1.nc',format='NETCDF3_64BIT') #save new file.
# # Check total vocs
#get all vocs files as ordered dictionary of xarray datasets.
def get_vocs_arr_frac(voc_dir):
vocs={}
for f in os.listdir(voc_dir):
vname=f.split('_')[4]
vocs.update({vname:xr.open_dataset(voc_dir+f)})
return vocs
vocs_frac=get_vocs_arr_frac(save_dir)
# get total nmvoc summing all vocs contributions.
for k,v in vocs_frac.items():
vocs_frac[k]=v*tot_nmvoc
tot_nmvoc_calc=sum(vocs_frac.values())
tot_nmvoc_calc
#test
xr.testing.assert_allclose(tot_nmvoc_calc, tot_nmvoc)
| code/nmvoc4.3.2_map_mozart_fractions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="LCziYWI6-MOL" colab_type="text"
# # Assignment 3 - Practical Deep Learning Workshop
#
# + [markdown] id="kof0gpyU-MOj" colab_type="text"
# #### In this task we will work with the dataset of the Home depot product search relevance competition.
# #### Some background:
# In this competition, Home Depot is asking to help them improve their customers' shopping experience by developing a model that can accurately predict the relevance of search results.
#
# Search relevancy is an implicit measure Home Depot uses to gauge how quickly they can get customers to the right products.
#
# This data set contains a number of products and real customer search terms from Home Depot's website. The challenge is to predict a relevance score for the provided combinations of search terms and products. To create the ground truth labels, Home Depot has crowdsourced the search/product pairs to multiple human raters.
# The relevance is a number between 1 (not relevant) to 3 (highly relevant). For example, a search for "AA battery" would be considered highly relevant to a pack of size AA batteries (relevance = 3), mildly relevant to a cordless drill battery (relevance = 2), and not relevant to a snow shovel (relevance = 1).
#
# Each pair was evaluated by at least three human raters. The provided relevance scores are the average value of the ratings. There are three additional things to know about the ratings:
# • The specific instructions given to the raters is provided in relevance_instructions.docx.
# • Raters did not have access to the attributes.
# • Raters had access to product images, while the competition does not include images.
#
#
# #### Out task here is to predict the relevance for each pair listed in the test set. The test set contains both seen and unseen search terms.
#
# + id="2MX3jC58-MOv" colab_type="code" outputId="334a8066-17ad-4ccc-b00d-169c71a30341" executionInfo={"status": "ok", "timestamp": 1548071169423, "user_tz": -120, "elapsed": 2223, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.feature_extraction.text import CountVectorizer
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model, Sequential
from keras.layers import * # Dense, Embedding, LSTM
from sklearn.model_selection import train_test_split
from keras.utils.np_utils import to_categorical
from keras.regularizers import l2
import re
import pandas as pd
import numpy as np
import datetime
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# %matplotlib inline
# + id="XZkB-Gjx-7Qf" colab_type="code" outputId="63686915-2d88-4804-b0e6-8b9136cfc432" executionInfo={"status": "ok", "timestamp": 1548071197833, "user_tz": -120, "elapsed": 29788, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 128}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="yiT5fXTC-MP9" colab_type="text"
# #### First of all, we'll take a look at the data in each dataset of the input:
# + [markdown] id="T1eqOAru-MQJ" colab_type="text"
# train.csv is the training set, contains products, searches, and relevance scores.
# + id="571V6TUw-MQQ" colab_type="code" outputId="d89b884e-2266-4050-f784-dcc508350ae1" executionInfo={"status": "ok", "timestamp": 1548071209279, "user_tz": -120, "elapsed": 1939, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
train = pd.read_csv('/content/gdrive/My Drive/Colab Notebooks/input/train.csv',encoding='latin1')
train.head()
# + [markdown] id="6Fs9YZB5-MRD" colab_type="text"
# test.csv is the test set, contains products and searches. We will need to predict the relevance for these pairs.
# + id="CCp-j0Fo-MRP" colab_type="code" outputId="d853e30a-76d4-4784-be11-4c4f51d591d2" executionInfo={"status": "ok", "timestamp": 1548071214625, "user_tz": -120, "elapsed": 1537, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
test = pd.read_csv('/content/gdrive/My Drive/Colab Notebooks/input/test.csv',encoding='latin1')
test.head()
# + [markdown] id="o5QyHDbp-MR0" colab_type="text"
# product_descriptions.csv contains a text description of each product. We may join this table to the training or test set via the product_uid.
# + id="kly-_Lrw-MSC" colab_type="code" outputId="c3ccd7cd-dbbb-4d03-a727-87f92feebc0d" executionInfo={"status": "ok", "timestamp": 1548071219639, "user_tz": -120, "elapsed": 3634, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
product_descriptions = pd.read_csv('/content/gdrive/My Drive/Colab Notebooks/input/product_descriptions.csv',encoding='latin1')
product_descriptions.head()
# + [markdown] id="2DlmxNyV-MS-" colab_type="text"
# attributes.csv provides extended information about a subset of the products (typically representing detailed technical specifications). Not every product will have attributes.
# + id="_9ryYcng-MTE" colab_type="code" outputId="7f0bdac0-34e4-48dd-8d25-bfb4162de9b8" executionInfo={"status": "ok", "timestamp": 1548071225103, "user_tz": -120, "elapsed": 4242, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
attributes = pd.read_csv('/content/gdrive/My Drive/Colab Notebooks/input/attributes.csv',encoding='latin1')
attributes.head()
# + [markdown] id="XWx4D8OI-MTf" colab_type="text"
# Data fields:
# - id - a unique Id field which represents a (search_term, product_uid) pair
# - product_uid - an id for the products
# - product_title - the product title
# - product_description - the text description of the product (may contain HTML content)
# - search_term - the search query
# - relevance - the average of the relevance ratings for a given id
# - name - an attribute name
# - value - the attribute's value
#
# + [markdown] id="OH3P79hG-MTn" colab_type="text"
# ## Preprocessing the data
# + [markdown] id="O8MOqmXE-MTu" colab_type="text"
# We would like to have the products' corresponding product description, so we will merge the train and test datasets with the product_description table.
#
# Note: in order to decrease the dimensionality of the text, we will lower the characters.
# + id="Boqw3h6t-MTy" colab_type="code" outputId="795767fe-3a6b-424b-fe57-31457333c386" executionInfo={"status": "ok", "timestamp": 1548071228220, "user_tz": -120, "elapsed": 864, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain = pd.merge(train, product_descriptions, how='inner', on='product_uid')
mergedTrain.search_term = mergedTrain.search_term.apply(lambda x: x.lower())
mergedTrain.product_description = mergedTrain.product_description.apply(lambda x: x.lower())
mergedTrain.head()
# + id="AG8ijy_k-MT9" colab_type="code" outputId="f75edf70-2581-4c5c-d99c-5e0ad96aa2bf" executionInfo={"status": "ok", "timestamp": 1548071231312, "user_tz": -120, "elapsed": 856, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTest= pd.merge(test, product_descriptions, how='inner', on='product_uid')
mergedTest.search_term = mergedTest.search_term.apply(lambda x: x.lower())
mergedTest.product_description = mergedTest.product_description.apply(lambda x: x.lower())
mergedTest.head()
# + [markdown] id="nT0BmvrG-MUE" colab_type="text"
# We convert the product_description and search_term attributes' values to lists of characters.
# + id="MVcxj3yH-MUF" colab_type="code" colab={}
search_term_chars = []
product_description_chars = []
search_term_chars = mergedTrain.search_term.apply(lambda x: search_term_chars + list(x))
product_description_chars = mergedTrain.product_description.apply(lambda x: product_description_chars + list(x))
search_term_chars = [item for sublist in search_term_chars for item in sublist]
product_description_chars = [item for sublist in product_description_chars for item in sublist]
# + [markdown] id="6I0gPihE-MUQ" colab_type="text"
# And then, translate the characters to a unique integer values.
# + id="p2pdGCwJ-MUS" colab_type="code" colab={}
search_term_char_set = sorted(set(search_term_chars))
product_description_char_set = sorted(set(product_description_chars))
# translate from character to number, it's enumerator
search_term_char_to_int = dict((c, i) for i, c in enumerate(search_term_char_set))
search_term_int_to_char = dict((i, c) for i, c in enumerate(search_term_char_set))
product_description_char_to_int = dict((c, i) for i, c in enumerate(product_description_char_set))
product_description_int_to_char = dict((i, c) for i, c in enumerate(product_description_char_set))
# + id="xRlcIGMS-MUa" colab_type="code" outputId="414a6f46-e16c-490d-a987-377a6a5e5ee8" executionInfo={"status": "ok", "timestamp": 1548071242256, "user_tz": -120, "elapsed": 464, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 90}
# summarize the loaded data
n_chars = len(search_term_chars)
n_vocab = len(search_term_char_set)
print("search_term Total Characters: ", n_chars)
print("search_term Total Vocab: ", n_vocab)
n_chars2 = len(product_description_chars)
n_vocab2 = len(product_description_char_set)
print("product_description Total Characters: ", n_chars2)
print("product_description Total Vocab: ", n_vocab2)
# + id="qNnrXX4O-MUk" colab_type="code" outputId="f188dd9f-4f4a-4c5e-8621-6c850b04bc96" executionInfo={"status": "ok", "timestamp": 1548071246920, "user_tz": -120, "elapsed": 4304, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain.search_term = mergedTrain.search_term.apply(lambda x: list(x))
mergedTrain.product_description = mergedTrain.product_description.apply(lambda x: list(x))
mergedTrain.head()
# + id="FG0e7bN_-MUv" colab_type="code" colab={}
def createData(char_to_int, char_arr):
#seq_length = 100
dataX = []
for i in range(0,len(char_arr)):
dataX.append(char_to_int[char_arr[i]])
return np.asarray(dataX)
# + id="xwnz8Nmb-MU-" colab_type="code" outputId="394dcb11-847b-4b9f-d411-bb2fc078768d" executionInfo={"status": "ok", "timestamp": 1548071265259, "user_tz": -120, "elapsed": 16639, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain.search_term = mergedTrain.search_term.apply(lambda x: createData(search_term_char_to_int, x))
mergedTrain.product_description = mergedTrain.product_description.apply(lambda x: createData(product_description_char_to_int, x))
mergedTrain.head()
# + id="ovjlzjBI96a1" colab_type="code" outputId="5a2a8fa7-6a6a-43fc-c58d-e3d130299e03" executionInfo={"status": "ok", "timestamp": 1548071265270, "user_tz": -120, "elapsed": 13117, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 267}
plt.hist(np.unique(mergedTrain.relevance.values),density=True, histtype='bar')
plt.show()
# + id="Jonu-0k7_zkJ" colab_type="code" outputId="10ad5702-07e5-4cda-bbff-792099046b16" executionInfo={"status": "ok", "timestamp": 1548071265279, "user_tz": -120, "elapsed": 10704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
np.unique(mergedTrain.relevance.values).size
# + id="1XvHxr8U-MVI" colab_type="code" outputId="57a3dd73-b72a-4288-f04c-19ed4fd0297f" executionInfo={"status": "ok", "timestamp": 1548071268079, "user_tz": -120, "elapsed": 611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 108}
from sklearn import preprocessing
target = mergedTrain['relevance'].values
min_max_scaler = preprocessing.MinMaxScaler()
Y = min_max_scaler.fit_transform(target.reshape(-1, 1))
Y[:5]
# + id="1ZXo6K1w-MVV" colab_type="code" colab={}
X1 = mergedTrain['search_term'].values
X2 = mergedTrain['product_description'].values
# + id="QHeLb30d-MVg" colab_type="code" colab={}
search_terms_lens = []
for element in mergedTrain['search_term'].values:
search_terms_lens.append(len(element))
product_description_lens = []
for element in mergedTrain['product_description'].values:
product_description_lens.append(len(element))
max_length1 = max(search_terms_lens)
max_length2 = max(product_description_lens)
# + id="q10L5vrGYBZa" colab_type="code" outputId="ee219f38-9307-48c0-fc30-53144f6ed3fb" executionInfo={"status": "ok", "timestamp": 1548071271122, "user_tz": -120, "elapsed": 402, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
max_length2
# + id="SRq6AFLA-MVk" colab_type="code" colab={}
max_length = 75
def padding(seq, length):
ans = []
for i in range(0,min(len(seq),length)):
ans.append(seq[i])
if len(seq) <= length:
for i in range(0,length-len(seq)):
ans.append(0)
return ans
X1 = np.asarray([padding(x,max_length) for x in X1])
X2 = np.asarray([padding(x,max_length) for x in X2])
X1 = X1.reshape(X1.shape[0],X1.shape[1],1)
X2 = X2.reshape(X2.shape[0],X2.shape[1],1)
# + id="DXbFfj_BM1FI" colab_type="code" colab={}
X1 = X1.astype(np.float32)
X2 = X2.astype(np.float32)
# + id="UwnIr1XLDdUv" colab_type="code" outputId="7e3abd33-529a-4753-8154-30e1c7dd1252" executionInfo={"status": "ok", "timestamp": 1548071275871, "user_tz": -120, "elapsed": 3430, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
print(X1.shape)
print(X2.shape)
# + id="KWrg7kIp-MVs" colab_type="code" colab={}
st_input = Input(shape=(max_length,1), name='st_input',dtype='float32')
pd_input = Input(shape=(max_length,1), name='pd_input',dtype='float32')
def createModel():
model = Sequential()
model.add(LSTM(40))
model.add(Dense(64, activation='relu'))
return model
# + id="1P5CErP3-MVy" colab_type="code" outputId="2bd206aa-1f90-4c6e-9491-d6422013c157" executionInfo={"status": "ok", "timestamp": 1548071277642, "user_tz": -120, "elapsed": 3668, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 73}
from keras.optimizers import Adadelta
st_model = createModel()
pd_model = createModel()
def createSiameseModel(model1,model2,customLoss):
out = Lambda(function=lambda x: K.exp(-K.sum(K.abs(x[0]-x[1]), axis=1, keepdims=True)),
output_shape=lambda x: (x[0][0], 1),
name='prediction')([model1(st_input), model2(pd_input)])
siamese_net = Model(input=[st_input,pd_input],output=[out])
siamese_net.compile(loss=customLoss,optimizer=Adadelta(lr=1.0, rho=0.95,clipnorm=1.20))
return siamese_net
siamese_net1 = createSiameseModel(st_model,pd_model,'mse')
siamese_net2 = createSiameseModel(st_model,pd_model,'mae')
# + id="gGKY0Jg83jqr" colab_type="code" outputId="5d7c1509-ae4c-49f9-daa5-c411067a89ae" executionInfo={"status": "ok", "timestamp": 1548071296241, "user_tz": -120, "elapsed": 557, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 217}
st_model.summary()
# + id="YACG1GiJ-MWC" colab_type="code" outputId="594e5bab-2661-4d98-9da8-ad8a0131db43" executionInfo={"status": "ok", "timestamp": 1548071297547, "user_tz": -120, "elapsed": 564, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 344}
siamese_net1.summary()
# + id="D5nCWY89-MWP" colab_type="code" colab={}
X1_train,X1_val,X2_train,X2_val,Y_train, Y_val = train_test_split(X1,X2,Y,test_size = 0.2)
# + id="uN8i5wFE-MWU" colab_type="code" colab={}
from keras.callbacks import *
path = 'gdrive/My Drive/Colab Notebooks'
def set_callbacks(description='run1',patience=15,tb_base_logdir='./logs/'):
cp = ModelCheckpoint(path + '/best_model_weights_{}.h5'.format(description),save_best_only=True)
rlop = ReduceLROnPlateau(patience=5)
cb = [cp,rlop]
return cb
# + id="w5adk4qt-MWY" colab_type="code" outputId="899c7891-e09f-44c2-da9d-cc7000b00736" executionInfo={"status": "ok", "timestamp": 1548071480901, "user_tz": -120, "elapsed": 165646, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 217}
start = time.time()
history = siamese_net1.fit([X1_train,X2_train],Y_train,batch_size=1024, epochs=5, verbose=1, validation_data=([X1_val,X2_val],Y_val), callbacks=set_callbacks())
end = time.time()
total_time = end - start
# + id="V-48pZPv-MWe" colab_type="code" outputId="13a88a35-60dd-4fd3-a7d2-9c858c744818" executionInfo={"status": "ok", "timestamp": 1548071484411, "user_tz": -120, "elapsed": 652, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
# + id="ficxWZ0V-MWj" colab_type="code" colab={}
val_preds = siamese_net1.predict([X1_val,X2_val])
train_preds = siamese_net1.predict([X1_train,X2_train])
# + id="PI6SmNNXXqx6" colab_type="code" outputId="e7feaeb1-1f98-4dac-e9f5-7593a5510f90" executionInfo={"status": "ok", "timestamp": 1548071537319, "user_tz": -120, "elapsed": 1285, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
val_preds.shape
# + id="WC_AEkXSqTu7" colab_type="code" outputId="a9ae2dca-158c-49b7-99e0-9b3058ebaaea" executionInfo={"status": "ok", "timestamp": 1548071537325, "user_tz": -120, "elapsed": 630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 108}
val_preds[:5]
# + id="y8D6DtGa-MWn" colab_type="code" outputId="74cfc78a-d0d3-4531-8422-7645c15dc7f5" executionInfo={"status": "ok", "timestamp": 1548071545831, "user_tz": -120, "elapsed": 289, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 513}
plt.hist(val_preds,density=True, histtype='bar')
plt.show()
plt.hist(Y_val,density=True, histtype='bar')
plt.show()
# + id="KxFU9_Mbgdo3" colab_type="code" colab={}
resultsTable = pd.DataFrame(columns=['model','runtime','TrainRMSE','ValRMSE','TestRMSE','TrainMAE','ValMAE','TestMAE'])
def addToTable(modelName,runtime,train_rmse,val_rmse,test_rmse,train_mae,val_mae,test_mae):
return resultsTable.append({'model': modelName,'runtime': runtime,'TrainRMSE': train_rmse,'ValRMSE': val_rmse,
'TrainMAE': test_rmse,'TrainMAE': train_mae,'ValMAE' :val_mae,'TestMAE': test_mae},ignore_index=True)
# + id="ND_G1M_S-MW_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="0928cddd-506e-447a-b669-c43f7004768a" executionInfo={"status": "ok", "timestamp": 1548071615565, "user_tz": -120, "elapsed": 68107, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
search_term_chars2 = []
product_description_chars2 = []
search_term_chars2 = mergedTest.search_term.apply(lambda x: search_term_chars2 + list(x))
product_description_chars2 = mergedTest.product_description.apply(lambda x: product_description_chars2 + list(x))
search_term_chars2 = [item for sublist in search_term_chars2 for item in sublist]
product_description_chars2 = [item for sublist in product_description_chars2 for item in sublist]
search_term_char_set2 = sorted(set(search_term_chars2))
product_description_char_set2 = sorted(set(product_description_chars2))
# translate from character to number, it's enumerator
search_term_char_to_int2 = dict((c, i) for i, c in enumerate(search_term_char_set2))
search_term_int_to_char2 = dict((i, c) for i, c in enumerate(search_term_char_set2))
product_description_char_to_int2 = dict((c, i) for i, c in enumerate(product_description_char_set2))
product_description_int_to_char2 = dict((i, c) for i, c in enumerate(product_description_char_set2))
mergedTest.search_term = mergedTest.search_term.apply(lambda x: list(x))
mergedTest.product_description = mergedTest.product_description.apply(lambda x: list(x))
mergedTest.search_term = mergedTest.search_term.apply(lambda x: createData(search_term_char_to_int2, x))
mergedTest.product_description = mergedTest.product_description.apply(lambda x: createData(product_description_char_to_int2, x))
mergedTest.head()
# + id="yyaG_bNVi7ME" colab_type="code" colab={}
X1_test = mergedTest.search_term.values
X2_test = mergedTest.product_description.values
X1_test = np.asarray([padding(x,max_length) for x in X1_test])
X2_test = np.asarray([padding(x,max_length) for x in X2_test])
X1_test = X1_test.reshape(X1_test.shape[0],X1_test.shape[1],1)
X2_test = X2_test.reshape(X2_test.shape[0],X2_test.shape[1],1)
# + id="jlh2Ks6FlUEC" colab_type="code" colab={}
test_preds = siamese_net1.predict([X1_test,X2_test])
# + id="ayBm0bzElu22" colab_type="code" colab={}
from sklearn.metrics import mean_absolute_error as mae
from sklearn.metrics import mean_squared_error as mse
resultsTable = addToTable('CHAR_SiameseNetwork',total_time,mse(train_preds,Y_train),mse(val_preds,Y_val),'-',mae(train_preds,Y_train),mae(val_preds,Y_val),'-')
resultsTable.head()
# + [markdown] id="S_l3rmQY-MXQ" colab_type="text"
# ## ML Benchmark
# + id="jCX_kRgi-MXU" colab_type="code" outputId="46438e12-2a55-4c84-d9e9-4c80bc7e8586" executionInfo={"status": "ok", "timestamp": 1548071843217, "user_tz": -120, "elapsed": 579, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain2 = pd.merge(train, product_descriptions, how='inner', on='product_uid')
mergedTrain2.head()
# + id="ptKpWyix-MXa" colab_type="code" colab={}
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer(encoding='latin-1', analyzer='char')
vectorizer.fit(mergedTrain2['search_term'])
mltrain_x, mlval_x, mltrain_y, mlval_y = train_test_split(mergedTrain2['search_term'].values,mergedTrain2['relevance'].values, test_size = 0.2)
train_x_count = vectorizer.transform(mltrain_x)
val_x_count = vectorizer.transform(mlval_x)
# + id="QzNz1fOM-MXi" colab_type="code" outputId="c5ce362b-4530-42b1-ad84-8f7a24726ce4" executionInfo={"status": "ok", "timestamp": 1548071890475, "user_tz": -120, "elapsed": 45563, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 92}
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm,ensemble
ml = ensemble.RandomForestRegressor()
start_time = time.time()
ml.fit(train_x_count, mltrain_y)
end_time = time.time()
total_time = end_time - start_time
ml_train_preds = ml.predict(train_x_count)
ml_val_preds = ml.predict(val_x_count)
print(ml_val_preds.shape)
resultsTable = addToTable('CHAR_RandomForestBenchmark',total_time,mse(ml_train_preds,mltrain_y),mse(ml_val_preds,mlval_y),'-',mae(ml_train_preds,mltrain_y),mae(ml_val_preds,mlval_y),'-')
# + id="UBwEQ_Ge1527" colab_type="code" outputId="4d1eb868-ae65-4a48-ed2d-74315fc80cd3" executionInfo={"status": "ok", "timestamp": 1548071890483, "user_tz": -120, "elapsed": 44922, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 112}
resultsTable.head()
# + id="Kr7dK6bI-MXq" colab_type="code" outputId="6f293036-6f2c-4105-a111-d41c74e0c996" executionInfo={"status": "ok", "timestamp": 1548071919559, "user_tz": -120, "elapsed": 876, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 515}
plt.hist(ml_val_preds,density=True, histtype='bar')
plt.show()
plt.hist(mlval_y,density=True, histtype='bar')
plt.show()
# + [markdown] id="BH7kinB_-MXv" colab_type="text"
# ## Feature Extraction
# + id="9wMmeMJ5djW4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="2f5b629a-09cc-4bcb-80d2-49e54d8cb2bd" executionInfo={"status": "ok", "timestamp": 1548071930366, "user_tz": -120, "elapsed": 568, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
path
# + id="9tN-8dzt-MXw" colab_type="code" outputId="4e171963-231c-43ac-ae6f-7099cd489bf9" executionInfo={"status": "ok", "timestamp": 1548072045188, "user_tz": -120, "elapsed": 1912, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 55}
from keras import backend as K
fe_st_input = Input(shape=(max_length,1), name='st_input',dtype='float32')
fe_pd_input = Input(shape=(max_length,1), name='pd_input',dtype='float32')
input_layer1 = siamese_net1.layers[0].input[0]
input_layer2 = siamese_net1.layers[1].input[0]
fe_st_model = createModel()
fe_pd_model = createModel()
output_layer1 = siamese_net1.layers[3].get_output_at(0)
output_layer2 = siamese_net1.layers[3].get_output_at(1)
output_fn = K.function([st_input, pd_input], [output_layer1, output_layer2])
def extractFeatures(model1,model2,customLoss):
out = concatenate([model1(fe_st_input), model2(fe_pd_input)])
siamese_net = Model(input=[fe_st_input,fe_pd_input],output=[out])
siamese_net.load_weights(path + '/best_model_weights_run1.h5')
siamese_net.compile(loss=customLoss,optimizer=Adadelta(lr=1.0, rho=0.95,clipnorm=1.20))
return siamese_net
fe_model = extractFeatures(fe_st_model,fe_pd_model,'mse')
# + id="ApFV_C1wFt9I" colab_type="code" colab={}
fe_train_features = fe_model.predict([X1_train,X2_train])
fe_val_features = fe_model.predict([X1_val,X2_val])
fe_test_features = fe_model.predict([X1_test,X2_test])
# + id="dbFYrdYpFpe1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 110} outputId="5fea89de-9b55-4a11-8b51-1babc716768f" executionInfo={"status": "ok", "timestamp": 1548072553830, "user_tz": -120, "elapsed": 26340, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
randomForest = ensemble.RandomForestRegressor()
start_time = time.time()
randomForest.fit(fe_train_features, Y_train)
end_time = time.time()
total_time = end_time - start_time
fe_train_preds = randomForest.predict(fe_train_features)
fe_val_preds = randomForest.predict(fe_val_features)
print(ml_val_preds.shape)
resultsTable = addToTable('FE_RandomForest_CHAR',total_time,mse(fe_train_preds,Y_train),mse(fe_val_preds,Y_val),'-',mae(fe_train_preds,Y_train),mae(fe_val_preds,Y_val),'-')
# + id="E7Rk5t_iUD3t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e5da7662-7041-46bd-fdde-2e5b3fa5cc49" executionInfo={"status": "ok", "timestamp": 1548072905258, "user_tz": -120, "elapsed": 861, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
linear = linear_model.LinearRegression()
start_time = time.time()
linear.fit(fe_train_features, Y_train)
end_time = time.time()
total_time = end_time - start_time
fe_train_preds2= linear.predict(fe_train_features)
fe_val_preds2 = linear.predict(fe_val_features)
print(ml_val_preds.shape)
resultsTable = addToTable('FE_LinearRegression_CHAR',total_time,mse(fe_train_preds2,Y_train),mse(fe_val_preds2,Y_val),'-',mae(fe_train_preds2,Y_train),mae(fe_val_preds2,Y_val),'-')
# + id="S4eAd617UeZa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 175} outputId="dd6273af-8c0f-4ac3-fe3a-7e1c27e0a7a9" executionInfo={"status": "ok", "timestamp": 1548072909691, "user_tz": -120, "elapsed": 595, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
resultsTable.head()
# + [markdown] id="1kQLJokSc-NC" colab_type="text"
# # Word Level Embedding
# + id="yMRi6xlKdpEC" colab_type="code" outputId="c14611f8-0af9-40c6-d8a8-d6d89b738439" executionInfo={"status": "ok", "timestamp": 1548073032882, "user_tz": -120, "elapsed": 899, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain = pd.merge(train, product_descriptions, how='inner', on='product_uid')
mergedTrain.search_term = mergedTrain.search_term.apply(lambda x: x.lower())
mergedTrain.product_description = mergedTrain.product_description.apply(lambda x: x.lower())
mergedTrain.head()
# + id="xBf9HVhLdeSD" colab_type="code" outputId="efe6118f-a162-4643-db52-add3af63b4c9" executionInfo={"status": "ok", "timestamp": 1548073033641, "user_tz": -120, "elapsed": 1206, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTest= pd.merge(test, product_descriptions, how='inner', on='product_uid')
mergedTest.search_term = mergedTest.search_term.apply(lambda x: x.lower())
mergedTest.product_description = mergedTest.product_description.apply(lambda x: x.lower())
mergedTest.head()
# + id="cpOwUQVT6nTg" colab_type="code" outputId="3561bac9-9d82-4794-ac95-b57974d90002" executionInfo={"status": "ok", "timestamp": 1548073066944, "user_tz": -120, "elapsed": 1982, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 53}
import nltk
nltk.download('punkt')
from nltk.tokenize import word_tokenize
# + id="WWHsKOot6KYZ" colab_type="code" colab={}
st_words = []
for term in mergedTrain.search_term.values:
for word in word_tokenize(term):
st_words.append(word)
st_word_set = sorted(set(st_words))
st_dict = dict((c, i) for i, c in enumerate(st_word_set))
# + id="A_moSd-172rP" colab_type="code" colab={}
pd_words = []
for term in mergedTrain.product_description.values:
for word in word_tokenize(term):
pd_words.append(word)
pd_word_set = sorted(set(pd_words))
pd_dict = dict((c, i) for i, c in enumerate(pd_word_set))
# + id="VGMyeZdhYpsR" colab_type="code" colab={}
st_words2 = []
for term in mergedTest.search_term.values:
for word in word_tokenize(term):
st_words.append(word)
st_word_set2 = sorted(set(st_words2))
st_dict2 = dict((c, i) for i, c in enumerate(st_word_set2))
pd_words2 = []
for term in mergedTest.product_description.values:
for word in word_tokenize(term):
pd_words.append(word)
pd_word_set2 = sorted(set(pd_words2))
pd_dict2 = dict((c, i) for i, c in enumerate(pd_word_set2))
# + id="6oEB5jxm9L2t" colab_type="code" outputId="76328aff-c08d-4786-c8ce-65600471414a" executionInfo={"status": "ok", "timestamp": 1548073325727, "user_tz": -120, "elapsed": 195989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 206}
mergedTrain.search_term = mergedTrain.search_term.apply(lambda x: createData(st_dict, word_tokenize(x)))
mergedTrain.product_description = mergedTrain.product_description.apply(lambda x: createData(pd_dict, word_tokenize(x)))
mergedTrain.head()
# + id="mxasCojTYzZA" colab_type="code" colab={}
mergedTest.search_term = mergedTest.search_term.apply(lambda x: createData(st_dict2, word_tokenize(x)))
mergedTest.product_description = mergedTest.product_description.apply(lambda x: createData(pd_dict2, word_tokenize(x)))
mergedTest.head()
# + id="pI78eYMv5q-_" colab_type="code" outputId="1b18554f-b999-4fc9-93a2-daaf9e65b8a0" executionInfo={"status": "ok", "timestamp": 1548073325736, "user_tz": -120, "elapsed": 195506, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 108}
target = mergedTrain['relevance'].values
min_max_scaler = preprocessing.MinMaxScaler()
Y = min_max_scaler.fit_transform(target.reshape(-1, 1))
Y[:5]
# + id="dwzQQJBGCpGM" colab_type="code" colab={}
X1 = mergedTrain['search_term'].values
X2 = mergedTrain['product_description'].values
search_terms_lens = []
for element in mergedTrain['search_term'].values:
search_terms_lens.append(len(element))
product_description_lens = []
for element in mergedTrain['product_description'].values:
product_description_lens.append(len(element))
max_length1 = max(search_terms_lens)
max_length2 = max(product_description_lens)
# + id="DngJQER1C3p8" colab_type="code" colab={}
max_length = 50
def padding(seq, length):
ans = []
for i in range(0,min(len(seq),length)):
ans.append(seq[i])
if len(seq) <= length:
for i in range(0,length-len(seq)):
ans.append(0)
return ans
X1 = np.asarray([padding(x,max_length) for x in X1])
X2 = np.asarray([padding(x,max_length) for x in X2])
X1 = X1.reshape(X1.shape[0],X1.shape[1],1)
X2 = X2.reshape(X2.shape[0],X2.shape[1],1)
# + id="XUIDq0n0C5Gs" colab_type="code" outputId="18ec0663-e264-4bb3-d728-46a73bfd45c6" executionInfo={"status": "ok", "timestamp": 1548073335531, "user_tz": -120, "elapsed": 1394, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 55}
st_input = Input(shape=(max_length,1), name='st_input')
pd_input = Input(shape=(max_length,1), name='pd_input')
def createModel():
model = Sequential()
model.add(LSTM(60))
model.add(Dense(140, activation='relu'))
return model
st_model3 = createModel()
pd_model3 = createModel()
def createSiameseModel(model1,model2,customLoss):
out = Lambda(function=lambda x: K.exp(-K.sum(K.abs(x[0]-x[1]), axis=1, keepdims=True)),
output_shape=lambda x: (x[0][0], 1),
name='prediction')([model1(st_input), model2(pd_input)])
siamese_net = Model(input=[st_input,pd_input],output=[out])
siamese_net.compile(loss=customLoss,optimizer=Adadelta(lr=1.0, rho=0.95,clipnorm=1.20))
return siamese_net
siamese_net3 = createSiameseModel(st_model3,pd_model3,'mse')
siamese_net4 = createSiameseModel(st_model3,pd_model3,'mae')
# + id="Rfi_TWWCDV0_" colab_type="code" outputId="2672e34d-7e7b-4f02-d7c4-902d3497b9fc" executionInfo={"status": "ok", "timestamp": 1548073342740, "user_tz": -120, "elapsed": 672, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 344}
siamese_net3.summary()
# + id="iNJhKxS6DT-g" colab_type="code" outputId="c53dfe80-509e-4ee9-b10d-36976901bf22" executionInfo={"status": "ok", "timestamp": 1548073578398, "user_tz": -120, "elapsed": 230452, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}} colab={"base_uri": "https://localhost:8080/", "height": 217}
X1_train,X1_val,X2_train,X2_val,Y_train, Y_val = train_test_split(X1,X2,Y,test_size = 0.2)
start = time.time()
history3 = siamese_net3.fit([X1_train,X2_train],Y_train,batch_size=1024, epochs=5, verbose=1, validation_data=([X1_val,X2_val],Y_val), callbacks=set_callbacks())
end = time.time()
total_time = end - start
# + id="1sXdzs0mYbwd" colab_type="code" colab={}
# + id="ipR7KCVXXbW4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="bb9b6b5c-3539-4da5-90aa-21c5441b882f" executionInfo={"status": "error", "timestamp": 1548073630762, "user_tz": -120, "elapsed": 23372, "user": {"displayName": "<NAME>", "photoUrl": "https://lh4.googleusercontent.com/-MduOAIa-9SA/AAAAAAAAAAI/AAAAAAAABMc/zWvM0_m3ZqQ/s64/photo.jpg", "userId": "03994682708229346579"}}
val_preds = siamese_net3.predict([X1_val,X2_val])
train_preds = siamese_net3.predict([X1_train,X2_train])
test_preds = siamese_net3.predict([X1_test,X2_test])
# + id="HTrAevDDDcCQ" colab_type="code" colab={}
plt.plot(history3.history['loss'], label='train')
plt.plot(history3.history['val_loss'], label='test')
plt.legend()
plt.show()
# + id="mj15od96DqTP" colab_type="code" colab={}
predsDF = pd.DataFrame(data = preds)
testDF = pd.DataFrame(data = Y_val)
plt.hist(preds3,density=True, histtype='bar')
plt.show()
plt.hist(Y_val,density=True, histtype='bar')
plt.show()
# + id="n-Vzx8yfEe7n" colab_type="code" colab={}
resultsTable = addToTable('WORD_SiameseNetwork',total_time,mse(train_preds,Y_train),mse(val_preds,Y_val),'-',mae(train_preds,Y_train),mae(val_preds,Y_val),'-')
resultsTable.head()
| oldver/hw3dl-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find the Best Model
#
# ---
# +
import joblib
from xgboost import XGBClassifier
from pprint import pprint
from glob import glob
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer, ENGLISH_STOP_WORDS
from sklearn.pipeline import Pipeline
from sklearn.model_selection import train_test_split, GridSearchCV, cross_validate, validation_curve, cross_val_score
from sklearn.metrics import confusion_matrix, multilabel_confusion_matrix
from sklearn.metrics import roc_auc_score, roc_curve, classification_report
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import LabelEncoder
from sklearn.naive_bayes import MultinomialNB
from sklearn.svm import SVC
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import StratifiedKFold
from sklearn.base import BaseEstimator, TransformerMixin
from imblearn.over_sampling import SMOTE, ADASYN
import sklearn
import time
import datetime
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
from helpers import load_sqlite, custom_stop_words, get_random_class_labels, resample_data
from visualizer import Visualizer
def build_model(preprocessor, classifier, cv=3, scoring='roc_auc_ovr', verbose=1):
'''
Takes a dictionary with params and outputs a gridsearch model
'''
pipe = Pipeline(
[('prep', preprocessor.get('preprocessor')),
('clf', classifier.get('estimator'))])
pipe_params = dict()
pipe_params.update(preprocessor.get('params'))
pipe_params.update(classifier.get('params'))
skf = StratifiedKFold(n_splits=cv, shuffle=True, random_state=random_state)
model = GridSearchCV(pipe, param_grid=pipe_params, cv=skf, verbose=verbose, scoring=scoring, n_jobs=-1)
return model
tfidf = {
'preprocessor': TfidfVectorizer(stop_words=custom_stop_words),
'name': 'TF-IDF Vectorizer',
'params': {
"prep__ngram_range": [(1, 2)],
"prep__max_df": [.9],
"prep__use_idf": [True],
"prep__norm": ["l2"],
# "prep__strip_accents": [None, 'ascii', 'unicode'],
# "prep__ngram_range": [(1, 1), (1, 2)],
# "prep__max_features": [5000, 6000, 7000],
# "prep__min_df": np.arange(2, 20, 4),
# "prep__max_df": np.linspace(.8, .99, 5),
# "prep__norm": ("l1", "l2"),
# "prep__use_idf": [True, False]
}
}
estimators = {
"logisticregression": {
"name": "Logistic Regression",
"estimator": LogisticRegression(max_iter=1000, fit_intercept=False, C=.99),
"params": {
"clf__solver": ["lbfgs", "saga"]
}
},
"randomforestclassifier": {
"name": "Random Forest",
"estimator": RandomForestClassifier(min_samples_leaf=2, min_samples_split=.01),
"params": {
"clf__n_estimators": [300, 500, 1000],
"clf__max_depth": np.linspace(400, 1000, 5, dtype=int)
}
},
"multinomialnb": {
"name": "Multinomial Bayes Classifier",
"estimator": MultinomialNB(alpha=.1189),
"params": {
"clf__fit_prior": [True, False]
}
},
"svc": {
"name": "Support Vector Classifier",
"estimator": SVC(kernel="sigmoid", probability=True),
"params": {
"clf__C": [.99, 1]
}
},
"sgdclassifier": {
"name": "Stochastic Gradient Descent Classifier",
"estimator": SGDClassifier(alpha=.0001, fit_intercept=True, penalty="l2", loss="modified_huber"),
"params":
{
}
},
'xgbclassifier': {
'name': 'XGBoost Classifier',
'estimator': XGBClassifier(n_estimators=200),
'params': {
"clf__max_depth": [3, 5, 10],
# "clf__learning_rate": np.linspace(.001, .1, 3),
# "clf__n_estimators": [50, 100, 200],
# "clf__objective": ['binary:logistic', 'multi:softprob'],
# "clf__booster": ['gbtree', 'gblinear', 'dart'],
# "clf__gamma": np.linspace(0, 1, 3),
# "clf__subsample": [.5, 1],
# "clf__reg_lambda": np.linspace(0, 1, 3),
}
}
}
# +
random_state = 77
labels = ['python','javascript','html']
df = load_sqlite(database='reddit.sqlite', class_labels=labels)
# +
X = df['title']
y = df['subreddit']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=7)
# +
fitted_models = dict()
best_auc_score = 0.0
best_model = None
for name, estimator in estimators.items():
print("*"*50)
print(f'Model: {estimator.get("name")}')
print()
model = build_model(preprocessor=tfidf, classifier=estimator, cv=5, verbose=0)
model.fit(X_train, y_train)
print()
train_score = model.score(X_train, y_train)
test_score = model.score(X_test, y_test)
print(f'Train Score: {train_score}')
print(f'Test Score: {test_score}')
if hasattr(model, 'predict_proba'):
y_proba = model.predict_proba(X_test)
auc = roc_auc_score(y_test, y_proba, multi_class="ovr")
print(f'AUC Score: {auc}')
print()
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred, digits=3))
viz = Visualizer(X=X_train, y=y_train,
transformer=model.best_estimator_.named_steps.prep,
classifier=model.best_estimator_.named_steps.clf)
viz.plot_confusion_matrix(y_test, y_pred)
plt.show() # so it doesn't put them all at the end
print()
print()
fitted_models[name] = {
'auc_score': auc,
'train_score': train_score,
'test_score': test_score
}
if auc > best_auc_score:
best_auc_score = auc
best_model = model.best_estimator_
print("*"*50)
print('BEST MODEL SO FAR:', best_model)
print()
print()
# -
model_output = pd.DataFrame(fitted_models).T
model_output
best_model
joblib.dump(best_model, '01_best_model')
| 01_choose_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="-8eFW_wl1n39" outputId="3bc3e61e-3f4f-4fec-cc9c-3975f175b162"
import os
# Find the latest version of spark 3.0 from http://www-us.apache.org/dist/spark/ and enter as the spark version
# For example:
spark_version = 'spark-3.0.1'
# spark_version = 'spark-3.<spark version>'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-11-jdk-headless -qq > /dev/null
# !wget -q http://www-us.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + colab={"base_uri": "https://localhost:8080/"} id="BzCrgs0Z1rnw" outputId="c7b173a9-a4c3-429a-d7d2-fcb0e2e52253"
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.9.jar
# + id="0DuBth0V2PR8"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("BigData-HW-1").config("spark.driver.extraClassPath","/content/postgresql-42.2.9.jar").getOrCreate()
# + [markdown] id="D3W2XJVi2CU-"
# # Load Amazon Data into Spark DataFrame
# + colab={"base_uri": "https://localhost:8080/"} id="Na_stw7b1wfU" outputId="7a7fcda0-3970-45fe-931f-f3741ed38609"
from pyspark import SparkFiles
url = "https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Video_Games_v1_00.tsv.gz"
spark.sparkContext.addFile(url)
video_games_df = spark.read.csv(SparkFiles.get("amazon_reviews_us_Video_Games_v1_00.tsv.gz"), sep="\t", header=True, inferSchema=True)
video_games_df.show()
# + [markdown] id="K95dqQ6U2YRv"
# # Size of Data
# + colab={"base_uri": "https://localhost:8080/"} id="Cayz-3Q52IM3" outputId="ed29e5b6-f061-4045-c4b6-4cbd59ed671c"
video_games_df.count()
# + [markdown] id="C9U0rkGZ2eu7"
# # Cleaned up DataFrames to match tables
# + colab={"base_uri": "https://localhost:8080/"} id="2tMYkSIk2d-m" outputId="29509a79-02b4-405d-94ae-ecd1c9d5ab40"
from pyspark.sql.functions import to_date
# Review DataFrame
review_id_df = video_games_df.select(["review_id", "customer_id", "product_id", "product_parent", to_date("review_date", 'yyyy-MM-dd').alias("review_date")])
review_id_df.show()
# + id="g9gTNhT62je4"
products_df = video_games_df.select(["product_id", "product_title"]).drop_duplicates()
# + colab={"base_uri": "https://localhost:8080/"} id="GEsm2myd2mA9" outputId="3d4a3eaf-801a-45a8-f155-681472d005ef"
reviews_df = video_games_df.select(["review_id", "review_headline", "review_body"])
reviews_df.show(10)
# + colab={"base_uri": "https://localhost:8080/"} id="_pF2Vf3c2n2O" outputId="46fbe13e-452f-4696-964b-9e12db21a55b"
customers_df = video_games_df.groupby("customer_id").agg({"customer_id": "count"}).withColumnRenamed("count(customer_id)", "customer_count")
customers_df.show()
# + colab={"base_uri": "https://localhost:8080/"} id="WHQKbmCE2p3Q" outputId="6bafecb0-d2bb-4723-e818-a2fbf2dacaff"
vine_df = video_games_df.select(["review_id", "star_rating", "helpful_votes", "total_votes", "vine"])
vine_df.show(10)
# + [markdown] id="I8aTsEjZ2s6L"
# # Push to AWS RDS instance
# + id="W4dzUKfI2vXM"
mode = "append"
jdbc_url="jdbc:postgresql://<endpoint>:5432/my_data_class_db"
config = {"user":"postgres", "password": "<password>", "driver":"org.postgresql.Driver"}
# + id="iOxKqMsD2yVs"
# Write review_id_df to table in RDS
review_id_df.write.jdbc(url=jdbc_url, table='review_id_table', mode=mode, properties=config)
# + id="pPXyGVE-2yPJ"
# Write products_df to table in RDS
products_df.write.jdbc(url=jdbc_url, table='products', mode=mode, properties=config)
# + id="aHbca4zN2yIa"
# Write customers_df to table in RDS
customers_df.write.jdbc(url=jdbc_url, table='customers', mode=mode, properties=config)
# + id="2HfOFneW2x_F"
# Write vine_df to table in RDS
vine_df.write.jdbc(url=jdbc_url, table='vines', mode=mode, properties=config)
| level_one_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="nJ1ni9KrAAwt" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="KRbNPD7IOCK3" colab_type="text"
# #**Part 1 - Data gathering and feature engineering**
#
# + [markdown] id="C67XfGT7J9B7" colab_type="text"
# **Libraries**
# + id="ei9gSULOJ16y" colab_type="code" colab={}
import numpy as np #Linear_Algebra
import matplotlib.pyplot as plt
import pandas as pd #Data_Processing
import pandas_datareader as pdr
from scipy import stats
# %matplotlib inline
# + id="uYRZH0-jJGBs" colab_type="code" colab={}
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# + id="mB75nmgF_GGZ" colab_type="code" outputId="648f6068-54f0-4a78-b39e-e2bbf9205cbd" colab={"base_uri": "https://localhost:8080/", "height": 35}
pip install -q yfinance --upgrade
# + id="x4Vd8OdoK0Kw" colab_type="code" colab={}
#Import Yahoo Finance
import yfinance as yf
# + id="dWus539UK_tg" colab_type="code" colab={}
yf.pdr_override()
# + id="JKzsKLhxHkQd" colab_type="code" colab={}
#CISCO data
SELECTED_STOCK = 'CSCO'
start = '2010-12-17'
end = '2018-12-17'
# + id="R4TAtqVVLDaR" colab_type="code" colab={}
#Download NVIDIA stock price data for the past 10 yrs to date
stock_data = pdr.get_data_yahoo(SELECTED_STOCK, start, end)
# + id="ehOZlTd4LF18" colab_type="code" outputId="b412ffd3-934d-4726-d32b-cc7d6fcb2a53" colab={"base_uri": "https://localhost:8080/", "height": 378}
stock_data.head(10)
# + [markdown] id="soiy7GjONyhq" colab_type="text"
# **Feature Engineering**
# + id="rGL_GCq7M6_o" colab_type="code" outputId="5c0ac1de-ba9e-4f8e-ae0a-81b430029bf0" colab={"base_uri": "https://localhost:8080/", "height": 145}
#Getting the Open price
stock_data_open = stock_data.Open.values
reshaped_stock_data_open = np.reshape(stock_data_open, (-1, 1))
reshaped_stock_data_open
# + id="bxbMt6PRGxbQ" colab_type="code" outputId="e6afb1b5-9e77-40c5-d8f9-2f73c351d4bc" colab={"base_uri": "https://localhost:8080/", "height": 35}
#validity check
np.mean(reshaped_stock_data_open)==np.mean(stock_data_open)
# + [markdown] id="9yy2vIb3WuIE" colab_type="text"
# ###**Analysis**
# + id="1qLd3FmKRtIM" colab_type="code" outputId="aaa45f9a-dfed-409b-cecc-6ce67dbe5c57" colab={"base_uri": "https://localhost:8080/", "height": 145}
#Finding log returns by changing the close-close price change
stock_close = stock_data["Adj Close"]
stock_percent_change = np.log(stock_close / stock_close.shift(1)) *100
stock_percent_change.head()
# + id="G92DV1OwWEWk" colab_type="code" outputId="c29c92d1-bd27-4d24-8a75-edb8fa6a8c44" colab={"base_uri": "https://localhost:8080/", "height": 392}
#Check for normality in the log returns
plt.hist(stock_percent_change[1:], density= True)
# + id="dCQouDIqWMUq" colab_type="code" outputId="e4b2ec68-9b57-4f14-d1d2-32edb5e2a905" colab={"base_uri": "https://localhost:8080/", "height": 55}
#Using Scipy to get more info like skweness, Kurtosis
stats.describe(stock_percent_change[1:])
# + [markdown] id="mCI0NR8wXFho" colab_type="text"
# >--For investors, the high kurtosis of the return distribution(16.64) implies that the investor will experience occasional extreme returns (either positive or negative), more extreme than the usual + or - three standard deviations from the mean that is predicted by the normal distribution of returns. This phenomenon is known as kurtosis risk.
#
# >--The kurtosis isn't close to 0, so a normal distribution for the returns is not assumed
# + id="iKh9GpVIWa_P" colab_type="code" outputId="19ba8c42-b737-4a14-c9ff-069485852447" colab={"base_uri": "https://localhost:8080/", "height": 35}
print('CISCO : ', stats.kurtosistest(stock_percent_change[1:]))
# + [markdown] id="KfQkIWQrX3U1" colab_type="text"
# >CISCO : KurtosistestResult(statistic=21.6296870467075, pvalue=9.442157604570577e-104)
#
# >--Since the Z value is 21.63 which is higher than 1.96, it leads us to conclude that we're not seeing Kurtosis from a normal distribution
#
# >--since the pvalue is <0.05, we reject the null hypothesis, that is, the kurtosis is not from a normal distribution --There is a very low probability (<0.05) that we're seeing these results from a random chance.
# + [markdown] id="dGFKh2nf0u_4" colab_type="text"
# ####**Stocks Fundamental Data**
# + id="kzQU2BWo0wWL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="4fb6d84e-794f-45f0-a7eb-0a70ba977723"
# !pip install yfinance
# + id="qGu5cXT_3SfA" colab_type="code" colab={}
CISCO = yf.Ticker("CSCO")
# + [markdown] id="9l6KNRz63k_1" colab_type="text"
# #####**Key Ratios**
# + id="Knqw5XBW3XAV" colab_type="code" outputId="26e5fe13-25c7-4942-f630-78bce421ad8d" colab={"base_uri": "https://localhost:8080/", "height": 35}
# get price to book
pb = CISCO.info['priceToBook']
print('Price to Book Ratio is: %.2f' % pb)
# + [markdown] id="Uwnbbz596lSQ" colab_type="text"
# #####**Options Data**
# + id="1oBGaV297Gma" colab_type="code" colab={}
pip install nsepy
# + id="AbnvdnDW6kkX" colab_type="code" colab={}
from datetime import date
from nsepy import get_history
stock_opt = get_history(symbol="CSCO",
start=date(2019, 1, 15),
end=date(2019, 2, 1),
option_type="CE",
strike_price=2000,
expiry_date=date(2019, 2, 28))
stock_opt.head()
# + [markdown] id="qhAi4Rnr0Tr5" colab_type="text"
# **Analyze performance**
# + [markdown] id="u_xqrhhn1qiF" colab_type="text"
# ####Visualization and Analysis
# + id="FjBL3Ae6zzn3" colab_type="code" colab={}
# Install pyfolio if not already installed
# !pip install pyfolio
# + id="hq-mY3C_z3oD" colab_type="code" colab={}
import pyfolio as pf
# + id="2LbZkkm_z6nN" colab_type="code" outputId="c23d69ef-07ac-45b8-f4e4-66c06d6e32d1" colab={"base_uri": "https://localhost:8080/", "height": 163}
# Define the ticker list
tickers_list = ['CSCO']
# Import pandas and create a placeholder for the data
import pandas as pd
data = pd.DataFrame(columns=tickers_list)
# Feth the data
import yfinance as yf
for ticker in tickers_list:
data[ticker] = yf.download(ticker, period='5y',)['Adj Close']
# Compute the returns of individula stocks and then compute the daily mean returns.
# The mean return is the daily portfolio returns with the above four stocks.
data = data.pct_change().dropna().mean(axis=1)
# Print first 5 rows of the data
data.head()
# + id="UK76P0Pf1BoJ" colab_type="code" outputId="50f241e0-2e6b-4628-e926-717067a05425" colab={"base_uri": "https://localhost:8080/", "height": 1000}
pf.create_full_tear_sheet(data)
# + [markdown] id="YVP11YejzR1B" colab_type="text"
# ###**Portfolio Value at Risk(VaR)**
# + [markdown] id="eRZu9bFEFNvY" colab_type="text"
# ####**1. Value at Risk(VaR) using Variance-Covariance approach**
# + [markdown] id="YEpX7-dh0CMQ" colab_type="text"
# calculating the daily returns
# + id="rP4-UiJOpMkM" colab_type="code" colab={}
pd.options.mode.chained_assignment = None
df = stock_data[['Adj Close']]
df['Return'] = df['Adj Close'].pct_change()
# + [markdown] id="ugGiDaSdsUv_" colab_type="text"
# Determine the mean and standard deviation of the daily returns. Plot the normal curve against the daily returns
# + id="LrT8FT4NsQGj" colab_type="code" outputId="6453aa11-4f72-4342-86e7-08fad6dbcaae" colab={"base_uri": "https://localhost:8080/", "height": 301}
import matplotlib.mlab as mlab
from scipy.stats import norm
mean = np.mean(df['Return'])
std_dev = np.std(df['Return'])
df['Return'].hist(bins=70, density=True, histtype='stepfilled', alpha=0.6)
x = np.linspace(mean-3*std_dev, mean+3*std_dev, 100)
plt.plot(x, norm.pdf(x, mean, std_dev))
plt.show()
# + [markdown] id="aJspyfbpw30Q" colab_type="text"
# Calculate the VaR using point percentile function
# + id="KJY3o5r-w4rM" colab_type="code" outputId="140ba63e-572b-468c-8c73-5674c5eca972" colab={"base_uri": "https://localhost:8080/", "height": 108}
from tabulate import tabulate
VaR_90 = norm.ppf(1-0.9, mean,std_dev)
VaR_95 = norm.ppf(1-0.95, mean,std_dev)
VaR_99 = norm.ppf(1-0.99, mean,std_dev)
print (tabulate([['90%', VaR_90], ['95%', VaR_95], ['99%', VaR_99]], headers = ['Confidence Level', 'Value at Risk']))
# + [markdown] id="nw21sh0Gzb2a" colab_type="text"
# ####**2. Value at Risk(VaR) using Historical Simulation approach**
# + [markdown] id="qYCJlgXZz7dR" colab_type="text"
# Calculate the daily returns
# + id="DME0xdnKzvYL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="139a3f05-a08d-4e16-80b4-bf776125cffd"
df = df.dropna()
plt.hist(df['Return'], bins = 70)
plt.xlabel('Return')
plt.ylabel('Frequency')
plt.grid(True)
plt.show()
# + [markdown] id="uugOj6mK1m7W" colab_type="text"
# Sort the returns
# + id="QMxlaRmW1irg" colab_type="code" colab={}
df.sort_values('Return', inplace = True, ascending=True)
# + [markdown] id="fKj0rPZx2L53" colab_type="text"
# Calculate the VaR for 90%, 95%, and 99% confidence levels using quantile function
# + id="xsEqli-z2RLF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 108} outputId="bfad8d25-e1e6-4d16-892a-662203154d6a"
VaR_90 = df['Return'].quantile(0.1)
VaR_95 = df['Return'].quantile(0.05)
VaR_99 = df['Return'].quantile(0.01)
print (tabulate([['90%', VaR_90], ['95%', VaR_95], ['99%', VaR_99]], headers = ['Confidence Level', 'Value at Risk']))
# + [markdown] id="06075a3b2f1D" colab_type="text"
# >As you can see there is a substantial difference in the value-at-risk calculated from historical simulation and variance-covariance approach. This tells us that the return distribution is not normal.
# + [markdown] id="VT8mA0IYIFYg" colab_type="text"
# ##**Volatility**
# + id="Hefi5xYqIJEW" colab_type="code" outputId="dde696b7-cd7b-4520-a93d-173e86ca6a5f" colab={"base_uri": "https://localhost:8080/", "height": 768}
## Computing Volatility
# Compute the logarithmic returns using the Closing price
stock_data['Log_Ret'] = np.log(stock_data['Adj Close'] / stock_data['Adj Close'].shift(1))
# Compute Volatility using the pandas rolling standard deviation function
stock_data['Volatility'] = pd.Series(stock_data['Log_Ret']).rolling(window=252).std() * np.sqrt(252)
print(stock_data.tail(15))
# Plot the CISCO Price series and the Volatility
stock_data[['Adj Close', 'Volatility']].plot(subplots=True, color='blue',figsize=(8, 6))
# + [markdown] id="xFVQdH61RCuk" colab_type="text"
# ###**Measures of risk adjusted return based on volatility**
# + [markdown] id="TBVOLf7SRKi3" colab_type="text"
# ####Sharpe ratio
# + [markdown] id="z_ZMRpLtRQN7" colab_type="text"
# >Sharpe ratio = (Mean return − Risk-free rate) / Standard deviation of return
# + id="MkVl0hqpQ4M4" colab_type="code" colab={}
# Sharpe Ratio
def sharpe(returns, rf, days=252):
volatility = returns.std() * np.sqrt(days)
sharpe_ratio = (returns.mean() - rf) / volatility
return sharpe_ratio
# + [markdown] id="As05dR7iKPuo" colab_type="text"
# ##**Indicators**
# + [markdown] id="0RWcfXDJ3li5" colab_type="text"
# ####**1. Moving Average**
# + id="TmGgfI3C3zUI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 935} outputId="5afc5ea2-37ba-4ef5-8bdf-4fe0badbebc0"
# Moving Averages Code
# Load the necessary packages and modules
from pandas_datareader import data as pdr
import matplotlib.pyplot as plt
import fix_yahoo_finance
import pandas as pd
# Simple Moving Average
def SMA(data, ndays):
SMA = pd.Series(data['Adj Close'], name = 'SMA').rolling(window=ndays).mean()
data = data.join(SMA)
return data
# Exponentially-weighted Moving Average
def EWMA(data, ndays):
EMA = pd.Series((data['Adj Close'].ewm(span=ndays).mean()),
name = 'EWMA_' + str(ndays))
data = data.join(EMA)
return data
# Retrieve the CISCO data from Yahoo finance:
data = pdr.get_data_yahoo("CSCO", start="2010-01-01", end="2019-12-16")
data = pd.DataFrame(data)
close = data['Adj Close']
# Compute the 50-day SMA for CISCO
n = 50
SMA_CISCO = SMA(data,n)
SMA_CISCO = SMA_CISCO.dropna()
SMA = SMA_CISCO['SMA']
# Compute the 200-day EWMA for CISCO
ew = 200
EWMA_CISCO = EWMA(data,ew)
EWMA_CISCO = EWMA_CISCO.dropna()
EWMA = EWMA_CISCO['EWMA_200']
# Plotting the CISCO Price Series chart and Moving Averages below
plt.figure(figsize=(9,5))
plt.plot(data['Close'],lw=1, label='NSE Prices')
plt.plot(SMA,'g',lw=1, label='50-day SMA (green)')
plt.plot(EWMA,'r', lw=1, label='200-day EWMA (red)')
plt.legend(loc=2,prop={'size':11})
plt.grid(True)
plt.setp(plt.gca().get_xticklabels(), rotation=30)
# + [markdown] id="doUWIAIB3AQv" colab_type="text"
# ###**2. Commodity Channel Index (CCI)**
# + id="OT5fnFt33Dm4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 876} outputId="16dee8ca-14d1-404b-fc1e-b49409d75fda"
from pandas_datareader import data as pdr
import matplotlib.pyplot as plt
import fix_yahoo_finance
import pandas as pd
# Commodity Channel Index
def CCI(data, ndays):
TP = (data['High'] + data['Low'] + data['Adj Close']) / 3
CCI = pd.Series((TP - pd.Series(TP).rolling(window=ndays).mean()) / (0.015 * pd.Series(TP).rolling(window=ndays).std()),
name = 'CCI')
data = data.join(CCI)
return data
# Retrieve the CISCO data from Yahoo finance:
data = pdr.get_data_yahoo("CSCO", start="2010-01-01", end="2019-12-16")
data = pd.DataFrame(data)
# Compute the Commodity Channel Index(CCI) for CISCO based on the 20-day Moving average
n = 20
CISCO_CCI = CCI(data, n)
CCI = CISCO_CCI['CCI']
# Plotting the Price Series chart and the Commodity Channel index below
fig = plt.figure(figsize=(7,5))
ax = fig.add_subplot(2, 1, 1)
ax.set_xticklabels([])
plt.plot(data['Close'],lw=1)
plt.title('NSE Price Chart')
plt.ylabel('Adj Close Price')
plt.grid(True)
bx = fig.add_subplot(2, 1, 2)
plt.plot(CCI,'k',lw=0.75,linestyle='-',label='CCI')
plt.legend(loc=2,prop={'size':9.5})
plt.ylabel('CCI values')
plt.grid(True)
plt.setp(plt.gca().get_xticklabels(), rotation=30)
# + [markdown] id="6ELhVbRtIQ8U" colab_type="text"
# ###**3. RSI**
# + id="lHaRMsetIYEC" colab_type="code" colab={}
from datetime import datetime
import matplotlib.pyplot as plt
import pandas_datareader as pdd
import pyEX as p
ticker = 'CSCO'
timeframe = '1y'
df = p.chartDF(ticker, timeframe)
df = df[['Open']]
df.reset_index(level=0, inplace=True)
df.columns=['ds','y']
delta = df.y.diff().dropna()
u = delta * 0
d = u.copy()
u[delta > 0] = delta[delta > 0]
d[delta < 0] = -delta[delta < 0]
u[u.index[14-1]] = np.mean( u[:14])
u = u.drop(u.index[:(14-1)])
d[d.index[14-1]] = np.mean( d[:14])
d = d.drop(d.index[:(14-1)])
rs = pdd.stats.moments.ewma(u, com=14-1, adjust=False) / \
pdd.stats.moments.ewma(d, com=14-1, adjust=False)
rsi = 100 - 100 / (1 + rs)
plt.plot(df.ds, rsi, label='CISCO RSI', color='orange')
plt.legend(loc='upper left')
plt.show()
# + [markdown] id="rzNRL16N8eFB" colab_type="text"
# **Feature Scaling**
# + id="J9LGh9b_8dfI" colab_type="code" colab={}
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0,1))
scaled_data = sc.fit_transform(reshaped_stock_data_open)
# + id="0H2VDom68mgO" colab_type="code" colab={}
def timestamp(n_period, scaled_data):
x_train = []
y_train = [] #1 output to predict
for i in range(n_period,len(scaled_data)):
x_train.append(scaled_data[i-n_period:i,0])
y_train.append(scaled_data[i,0])
x_train, y_train = np.array(x_train), np.array(y_train)
#reshaping
x_train_ = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
return x_train_, x_train, y_train
# + id="IBLKryX-9B8s" colab_type="code" colab={}
x_train_, x_train, y_train = timestamp(60, scaled_data)
# + [markdown] id="4qt8AJEQBDeR" colab_type="text"
# #**Part 2 - Model Identification**
# + [markdown] id="UbHyO51ID7Zb" colab_type="text"
# ##**Decision Tree (Regression)**
# + id="nhi5NOxVBBx5" colab_type="code" outputId="5c590dcd-20f1-40ec-818b-3798e05c4bbd" colab={"base_uri": "https://localhost:8080/", "height": 290}
from sklearn.ensemble import BaggingRegressor
from sklearn.tree import DecisionTreeRegressor
dt = DecisionTreeRegressor()
decision_tree_regr = BaggingRegressor(dt, n_estimators=10, random_state=0)
decision_tree_regr.fit(x_train, y_train)
# + [markdown] id="9rPpLMR-7wWh" colab_type="text"
# ##**Recurrent Neural Network (RNN)**
# + id="0MKeO-QhKQVP" colab_type="code" colab={}
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# + id="fldj8Fov7zcN" colab_type="code" colab={}
#Importing the keras libraries and packages
from tensorflow.python.keras.layers import Dense, LSTM, Dropout
from tensorflow.python.keras import Sequential
# + id="ErAvXgsS78v-" colab_type="code" outputId="df308273-1f37-44c1-bda5-fe4d58ca30e2" colab={"base_uri": "https://localhost:8080/", "height": 92}
regressor = Sequential()
#Adding the first LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True, input_shape = (x_train_.shape[1], 1)))
regressor.add(Dropout(rate = 0.2))
# + id="bwFOfhEG7_oT" colab_type="code" outputId="1ba9c309-3dee-4d66-bb5a-a210e95fe25e" colab={"base_uri": "https://localhost:8080/", "height": 35}
x_train.shape[1]
# + id="0WM2v9AD8BxI" colab_type="code" colab={}
#Adding the second LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True))
regressor.add(Dropout(rate = 0.2))
# + id="-7iLbqxH8Chi" colab_type="code" colab={}
#Adding the third LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True))
regressor.add(Dropout(rate = 0.2))
# + id="N6EMu3oe8Evx" colab_type="code" colab={}
#Adding the fourth LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50))
regressor.add(Dropout(rate = 0.2))
# + id="_80lEuHF8JCV" colab_type="code" colab={}
#Adding the output layer
regressor.add(Dense(units=1))
# + id="-pOaXmeo8LMd" colab_type="code" colab={}
#compiling the RNN
regressor.compile(optimizer='adam', loss='mean_squared_error')
# + id="3K8DHTIi8P9P" colab_type="code" outputId="491e7f1f-6922-4b26-fe0f-b30e6fe204bc" colab={"base_uri": "https://localhost:8080/", "height": 1000}
#fitting the RNN to the training set
regressor.fit(x_train_, y_train, epochs=50, batch_size = 32)
# + [markdown] id="2JhqXHA0qTP7" colab_type="text"
# **Save the model**
# + id="w-n7T2kLo_Kn" colab_type="code" colab={}
regressor = regressor.save("regressor.h5")
# + [markdown] id="-Cg2xPnOqYqt" colab_type="text"
# **Load the model**
# + id="foORAh2ep18l" colab_type="code" colab={}
from tensorflow.python.keras.models import load_model
regressor = load_model("regressor.h5")
# + [markdown] id="uXyp7Km_qgFw" colab_type="text"
# ##**Making the predictions and visualising the results**
# + id="Ad_-p9kb-qMq" colab_type="code" colab={}
# Getting the real/test stock price of 2019
test_stock_data = pdr.get_data_yahoo(SELECTED_STOCK, start = '2018-12-18', end = '2019-12-17')
real_stock_price = test_stock_data.iloc[:, 1:2].values
# + id="PFr-P0u4_HHQ" colab_type="code" colab={}
dataset_total = pd.concat((stock_data['Open'], test_stock_data['Open']), axis = 0)
inputs = dataset_total[len(dataset_total) - len(test_stock_data) - 60:].values
inputs = inputs.reshape(-1,1)
inputs = sc.transform(inputs)
# + id="XEqukC49_Hy6" colab_type="code" colab={}
X_test = []
for i in range(60, 310): #80 because we're predicting 20 records
X_test.append(inputs[i-60:i, 0])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[1], 1))
# + id="Y8U81iIQ_MAt" colab_type="code" colab={}
predicted_stock_price = regressor.predict(X_test)
predicted_stock_price = sc.inverse_transform(predicted_stock_price) #retranform the output because our input data was scaled between 0 and 1.
# + id="hL4qM5SO_PF1" colab_type="code" outputId="12d1e3e1-1001-4cfd-d1a0-3955be3781d8" colab={"base_uri": "https://localhost:8080/", "height": 369}
# Visualising the results
plt.plot(real_stock_price, color = 'red', label = 'Real CISCO Stock Price')
plt.plot(predicted_stock_price, color = 'blue', label = 'Predicted CISCO Stock Price')
plt.title('CISCO Stock Price Prediction')
plt.xlabel('Time')
plt.ylabel('CISCO Stock Price')
plt.legend()
plt.show()
| Finance Algo with Deep Learning/v6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome To Fugue Tutorials
#
# [Join Fugue-Project on Slack](https://join.slack.com/t/fugue-project/shared_invite/<KEY>)
#
# This environment has everything setup for you, you can run Fugue on native python, Spark and Dask, with Fugue SQL support. In order to setup your own environment, you can pip install the package:
#
# ```bash
# pip install fugue[all]
# ```
#
# The simplest way to run the tutorial is to use [mybinder](https://mybinder.org/v2/gh/fugue-project/tutorials/master)
#
# [](https://mybinder.org/v2/gh/fugue-project/tutorials/master)
#
# **But it runs slow on binder**, the machine on binder isn't powerful enough for
# a distributed framework such as Spark. Parallel executions can become sequential, so some of the
# performance comparison examples will not give you the correct numbers.
#
# Alternatively, you should get decent performance if running its docker image on your own machine:
#
# ```
# docker run -p 8888:8888 fugueproject/tutorials:latest
# ```
#
# + [markdown] nbsphinx-toctree={"hidden": true}
# # Tutorials
#
# ## [For Beginners](tutorials/beginner.ipynb)
#
# ## [For Advanced Users](tutorials/advanced.ipynb)
#
# ## [Fugue-sql](tutorials/fugue_sql/index.ipynb)
# -
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
# Preprocessing
from sklearn.preprocessing import StandardScaler, OneHotEncoder, LabelEncoder, PolynomialFeatures
from sklearn.compose import ColumnTransformer
# Models
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression, LinearRegression
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
# Metrics
from sklearn.metrics import mean_squared_error, confusion_matrix, classification_report, roc_curve
# Model selection
from sklearn.model_selection import GridSearchCV, train_test_split, cross_val_score
from sklearn.pipeline import Pipeline
# Feature selection
from sklearn.feature_selection import RFE
# Plotting
import altair as alt
# +
X_train = pd.read_csv("../data/clean-data/Xtrain-clean-autism-screening.csv", index_col=0)
y_train = pd.read_csv("../data/clean-data/ytrain-clean-autism-screening.csv", index_col=0)
X_test = pd.read_csv("../data/clean-data/Xtest-clean-autism-screening.csv", index_col=0)
y_test = pd.read_csv("../data/clean-data/ytest-clean-autism-screening.csv", index_col=0)
# Make validation set
X_train, X_valid, y_train, y_valid = train_test_split(X_train, y_train, test_size=0.2, random_state=414)
numeric_features = ["age",
"result"]
one_hot_features = ["gender",
"ethnicity",
"jaundice",
"country_of_res",
"used_app_before",
"age_desc",
"relation",
"Class/ASD"]
other_columns = list(X_train.columns[0:10])
# +
preprocessor = ColumnTransformer(sparse_threshold=0,
transformers=[
("scale",
StandardScaler(),
numeric_features),
("one_hot",
OneHotEncoder(drop=None,
handle_unknown="ignore"),
one_hot_features)
])
X_train_temp = pd.DataFrame(preprocessor.fit_transform(X_train),
index = X_train.index,
columns = (numeric_features +
list(preprocessor
.named_transformers_["one_hot"]
.get_feature_names(one_hot_features)))
)
X_test_temp = pd.DataFrame(preprocessor.transform(X_test),
index = X_test.index,
columns = X_train_temp.columns)
X_valid_temp = pd.DataFrame(preprocessor.transform(X_valid),
index = X_valid.index,
columns = X_train_temp.columns)
X_train = X_train_temp.join(X_train[other_columns])
X_test = X_test_temp.join(X_test[other_columns])
X_valid = X_valid_temp.join(X_valid[other_columns])
le = LabelEncoder()
y_train = le.fit_transform(y_train.to_numpy().ravel())
y_test = le.transform(y_test.to_numpy().ravel())
y_valid = le.transform(y_valid.to_numpy().ravel())
# +
## Trying Gridsearch on different models to find best
## Initialize models
lr = LogisticRegression()
dt = DecisionTreeClassifier(random_state=414)
rf = RandomForestClassifier(random_state=414)
svm = SVC(random_state=414)
knn = KNeighborsClassifier()
# Make list for models and a list to store their values
estimators = [lr, dt, rf, svm, knn]
best_parameters = []
best_precision_scores = []
# Make list of dictionaries for parameters
params = [{'C':[0.000001, 0.00001, 0.0001, 0.001, 0.01, 0.1, 1, 10, 100, 1000],
'penalty': ['l1', 'l2']},
{'max_depth': [1, 5, 10, 15, 20, 25, None],
'max_features': [3, 5, 10, 20, 25, 50, 100, None]},
{'min_impurity_decrease': [0, 0.25, 0.5],
'max_features': [3, 5, 10, 20, 50, 100, 'auto']},
{'C':[0.000001, 0.00001, 0.0001, 0.001, 0.01, 0.1, 1, 10, 100, 1000],
'gamma':[0.000001, 0.00001, 0.0001, 0.001, 0.01, 0.1, 1, 10, 100]},
{'n_neighbors': [2, 5, 10, 15, 20, 50, 100],
'algorithm': ['auto', 'brute']}]
# Run for loop to best parameters for each model
# Scoring = recall to reduce false positives
for i in range(len(estimators)):
search = GridSearchCV(estimator=estimators[i],
param_grid=params[i],
cv = 10,
n_jobs=-1,
scoring='recall')
search_object = search.fit(X_train, y_train)
# Store the output on each iteration
best_parameters.append(search_object.best_params_)
best_precision_scores.append(search_object.best_score_)
best_parameters[np.argmax(best_precision_scores)]
# -
# +
# the best precision score comes from a decision tree classifier with max_depth=20 and max_features=50
# and recall = 0.46
dt = DecisionTreeClassifier(max_depth=20, max_features=50, random_state=414)
dt.fit(X_train, y_train).score(X_train, y_train)
# It gets almost perfect on the train set
dt.score(X_valid, y_valid)
# and ~82.6% on the validation set
prelim_matrix = pd.DataFrame(confusion_matrix(y_valid, dt.predict(X_valid)))
preliminary_matrix = prelim_matrix.rename(columns={0:"predicted negative", 1:'predicted positive'},
index={0:"predicted negative", 1:'predicted positive'}).swapaxes(0,1)
preliminary_matrix.to_csv('../data/preliminary_matrix.csv')
preliminary_matrix
# -
dt.score(X_valid, y_valid)
# +
## Subset just the questions:
questions = ['A1_Score',
'A2_Score',
'A3_Score',
'A4_Score',
'A5_Score',
'A6_Score',
'A7_Score',
'A8_Score',
'A9_Score',
'A10_Score']
questions_train_df = X_train[questions]
questions_valid_df = X_valid[questions]
questions_test_df = X_test[questions]
# +
# Attribution: <NAME>
class ForwardSelection:
def __init__(self,
model,
min_features=None,
max_features=None,
scoring=None,
cv=None):
"""
Initializes a model object that can then fit and transform train and test data
"""
self.max_features = max_features
if min_features is None:
self.min_features = 1
else:
self.min_features = min_features
self.model = model
self.scoring = scoring
self.cv = cv
self.ftr_ = []
return
def fit(self, X, y):
"""
Finds the features from X that best predict y
"""
error = np.inf
best = None
feature_index = list(range(0, (X.shape[1])))
errors = []
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=514)
X_temp = X_train
while error > 0.0:
if best is not None:
if best not in feature_index:
del feature_index[-2]
break
feature_index.remove(best)
for i in feature_index:
self.model.fit(X_temp[:, self.ftr_ + [i]], y_train)
temp_error = 1-np.mean(cross_val_score(self.model, X[:, self.ftr_ + [i]], y, scoring='f1'))
if temp_error < error:
error = temp_error
best = i
errors.append(round(error, 3))
if len(errors) > 2:
if errors[-1] >= errors[-2]:
break
if self.max_features is not None:
if len(errors) > self.max_features:
break
self.ftr_.append(best)
def transform(self, X, y=None):
"""
Transforms the test set to have the best features selected from self.fit()
"""
return X[:, self.ftr_]
# +
fs = ForwardSelection(DecisionTreeClassifier(), max_features=None)
# np.mean(cross_val_score(dt, questions_train_df, y_train, scoring='precision'))
fs.fit(questions_train_df.to_numpy(), y_train)
fs.ftr_
# No single one question is better than any other one question so forward selection won't work
# Or it just won't work with a decision tree
# +
# Find the best features using RFE
rfe=RFE(DecisionTreeClassifier(), n_features_to_select=5)
rfe.fit(questions_train_df, y_train)
# The top 5 questions:
top_five = np.where(rfe.ranking_ == 1)[0]
X_train_best_5 = questions_train_df.to_numpy()[:,top_five]
X_test_best_5 = questions_test_df.to_numpy()[:,top_five]
X_valid_best_5 = questions_valid_df.to_numpy()[:,top_five]
# -
np.where(rfe.ranking_ == 1)[0]
# +
### Best questions
# The top five questions are A4, A5, A6, A8, and A10
# +
# Can't use max_features=50 as there are only 5 features
dt2 = DecisionTreeClassifier(max_depth=20, max_features=5, random_state=414)
dt2.fit(X_train_best_5, y_train)
pd.DataFrame(confusion_matrix(y_valid, dt2.predict(X_valid_best_5)))
# Using just the top 5 questions gets a much worse result than using all the features
# -
# Recall of 0 with top five questions
print(classification_report(y_valid, dt2.predict(X_valid_best_5)))
# +
# Try all questions:
dt3 = DecisionTreeClassifier(max_depth=20, max_features=10, random_state=414)
dt3.fit(questions_train_df, y_train)
pd.DataFrame(confusion_matrix(y_valid, dt3.predict(questions_valid_df)))
# -
# Recall of 0.19 with all questions on validation set
print(classification_report(y_valid, dt3.predict(questions_valid_df)))
# Recall with 0.38 with all features on validation set
print(classification_report(y_valid, dt.predict(X_valid)))
# Recall of 0.07 with all features on test set
print(classification_report(y_test, dt.predict(X_test)))
# +
conf_matrix = pd.DataFrame(confusion_matrix(y_test, dt.predict(X_test)))
final_matrix = conf_matrix.rename(columns={0:"predicted negative", 1:'predicted positive'},
index={0:"actual negative", 1:'actual positive'}).swapaxes(0,1)
final_matrix.to_csv('../data/confusion_matrix.csv')
# +
# ROC curve
fpr, tpr, _ = roc_curve(y_valid, dt.predict_proba(X_valid)[:,1])
roc_df = pd.DataFrame({"fpr":fpr, "tpr":tpr})
line_df = pd.DataFrame({"start":[0,1], "end":[0,1]})
roc = alt.Chart(roc_df).mark_line().encode(
x = alt.X("fpr:Q"),
y = alt.Y("tpr:Q")
)
line = alt.Chart(line_df).mark_line(strokeDash=[5,5], color="orange").encode(
x = alt.X("start:Q", axis=alt.Axis(title="False Positive Rate")),
y = alt.Y("end:Q", axis=alt.Axis(title="True Positive Rate"))
)
chart = (roc + line).configure_axis(titleFontSize=20).properties(title="ROC Curve").configure_title(fontSize=20)
chart
chart.save('../img/ROC.png', webdriver='firefox')
# +
## Extra stuff
# Importance of each feature according to the model
dt.feature_importances_
# -
chart
| src/ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code"
# !pip install -r https://raw.githubusercontent.com/datamllab/automl-in-action-notebooks/master/requirements.txt
# + [markdown] colab_type="text"
# ### Load dataset
#
# + colab_type="code"
from sklearn.datasets import fetch_california_housing
house_dataset = fetch_california_housing()
# Import pandas package to format the data
import pandas as pd
# Extract features with their names into the a dataframe format
data = pd.DataFrame(house_dataset.data, columns=house_dataset.feature_names)
# Extract target with their names into a pd.Series object with name MEDV
target = pd.Series(house_dataset.target, name="MEDV")
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
data, target, test_size=0.2, random_state=42
)
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=0.2, shuffle=False
)
# + colab_type="code"
X_train.shape, X_test.shape
# + [markdown] colab_type="text"
# ### Use LightGBM GBDT model to do regression without tuning
#
# + colab_type="code"
def build_model(hp):
model = lgb.LGBMRegressor(
boosting_type="gbdt",
# you can also search model type such as:
# boosting_type=hp.Choice("model_type", ['gbdt', 'goss'], default='gbdt'),
num_leaves=hp.Int("num_leaves", 5, 50, step=1),
learning_rate=hp.Float("learning_rate", 1e-3, 1, sampling="log", default=0.01),
n_estimators=hp.Int("n_estimators", 5, 50, step=1),
)
return model
# + [markdown] colab_type="text"
# ### Customize tuner
#
# + colab_type="code"
import os
import pickle
import tensorflow as tf
import keras_tuner as kt
import lightgbm as lgb
from sklearn.metrics import mean_squared_error
class LightGBMTuner(kt.engine.base_tuner.BaseTuner):
def run_trial(self, trial, X, y, validation_data):
model = self.hypermodel.build(trial.hyperparameters) # build the model
model.fit(
X_train,
y_train,
eval_set=[validation_data],
eval_metric="mse",
early_stopping_rounds=5,
) # fit the model
X_val, y_val = validation_data
y_pred = model.predict(
X_val, num_iteration=model.best_iteration_
) # evaluate the model
eval_mse = mean_squared_error(y_val, y_pred)
self.save_model(trial.trial_id, model) # save the model to disk
# inform the oracle of the eval result, the result is a dictionary with the metric names as the keys.
return {"mse": eval_mse}
def save_model(self, trial_id, model, step=0):
fname = os.path.join(self.get_trial_dir(trial_id), "model.txt")
model.booster_.save_model(fname, num_iteration=model.best_iteration_)
def load_model(self, trial):
fname = os.path.join(self.get_trial_dir(trial.trial_id), "model.txt")
model = lgb.Booster(model_file=fname)
return model
# + [markdown] colab_type="text"
# ### Customize Bayesian Optimization search algorithm
#
# + colab_type="code"
import random
import numpy as np
from scipy import optimize as scipy_optimize
from scipy.stats import norm
from sklearn import exceptions
from sklearn import gaussian_process
from keras_tuner.engine import hyperparameters as hp_module
from keras_tuner.engine import multi_execution_tuner
from keras_tuner.engine import oracle as oracle_module
from keras_tuner.engine import trial as trial_lib
class BayesianOptimizationOracle(oracle_module.Oracle):
"""Bayesian optimization oracle.
It uses Bayesian optimization with a underlying Gaussian process model.
The acquisition function used is upper confidence bound (UCB), which can
be found in the following link:
https://www.cse.wustl.edu/~garnett/cse515t/spring_2015/files/lecture_notes/12.pdf
# Arguments
objective: String or `kerastuner.Objective`. If a string,
the direction of the optimization (min or max) will be
inferred.
max_trials: Int. Total number of trials
(model configurations) to test at most.
Note that the oracle may interrupt the search
before `max_trial` models have been tested if the search space has been
exhausted.
num_initial_points: (Optional) Int. The number of randomly generated samples
as initial training data for Bayesian optimization. (If not specified,
a trick is to use the square root of the dimensionality of the
hyperparameter space.)
beta: Float. The balancing factor of exploration and exploitation.
The larger it is, the more explorative it is.
seed: Int. Random seed.
hyperparameters: HyperParameters class instance.
Can be used to override (or register in advance)
hyperparamters in the search space.
"""
def __init__(
self,
objective,
max_trials,
beta=2.6,
acq_type="ucb",
num_initial_points=None,
seed=None,
hyperparameters=None,
*args,
**kwargs
):
super(BayesianOptimizationOracle, self).__init__(
objective=objective,
max_trials=max_trials,
hyperparameters=hyperparameters,
seed=seed,
*args,
**kwargs
)
# Use 2 as the initial number of random points if not presented.
self.num_initial_points = num_initial_points or 2
self.beta = beta
self.seed = seed or random.randint(1, 1e4)
self._random_state = np.random.RandomState(self.seed)
self.gpr = self._make_gpr()
self.acq_type = acq_type
def _make_gpr(self):
return gaussian_process.GaussianProcessRegressor(
kernel=gaussian_process.kernels.Matern(nu=2.5),
alpha=1e-4,
normalize_y=True,
random_state=self.seed,
)
def _vectorize_trials(self):
x, y = [], []
for trial in self.trials.values():
# Create a vector representation of each Trial's hyperparameters.
trial_hps = trial.hyperparameters
vector = []
nonfixed_hp_space = [
hp
for hp in self.hyperparameters.space
if not isinstance(hp, hp_module.Fixed)
]
for hp in nonfixed_hp_space:
# For hyperparameters not present in the trial (either added after
# the trial or inactive in the trial), set to default value.
if trial_hps.is_active(hp):
trial_value = trial_hps.values[hp.name]
else:
trial_value = hp.default
# Embed an HP value into the continuous space [0, 1].
prob = hp_module.value_to_cumulative_prob(trial_value, hp)
vector.append(prob)
if trial.status == "COMPLETED":
score = trial.score
if self.objective.direction == "min":
score = -1 * score
else:
continue
x.append(vector)
y.append(score)
x = np.array(x)
y = np.array(y)
return x, y
def _vector_to_values(self, vector):
hps = hp_module.HyperParameters()
vector_index = 0
for hp in self.hyperparameters.space:
hps.merge([hp])
if isinstance(hp, hp_module.Fixed):
value = hp.value
else:
prob = vector[vector_index]
vector_index += 1
value = hp_module.cumulative_prob_to_value(prob, hp)
if hps.is_active(hp):
hps.values[hp.name] = value
return hps.values
def _random_populate_space(self):
values = self._random_values()
if values is None:
return {"status": trial_lib.TrialStatus.STOPPED, "values": None}
return {"status": trial_lib.TrialStatus.RUNNING, "values": values}
def _num_completed_trials(self):
return len([t for t in self.trials.values() if t.status == "COMPLETED"])
def populate_space(self, trial_id):
if self._num_completed_trials() < self.num_initial_points:
return self._random_populate_space()
# Update Gaussian process regressor
x, y = self._vectorize_trials()
try:
self.gpr.fit(x, y)
except exceptions.ConvergenceWarning as e:
raise e
# Three acquisition functions
def _upper_confidence_bound(x):
x = x.reshape(1, -1)
mu, sigma = self.gpr.predict(x, return_std=True)
return -1 * (mu + self.beta * sigma)
def _probability_of_improvement(x):
# calculate the best surrogate score found so far
x_history, _ = self._vectorize_trials()
y_pred = self.gpr.predict(x_history, return_std=False)
y_best = max(y_pred)
# calculate mean and stdev via surrogate function
x = x.reshape(1, -1)
mu, sigma = self.gpr.predict(x, return_std=True)
# calculate the probability of improvement
z = (mu - y_best) / (sigma + 1e-9)
prob = norm.cdf(z)
return -1 * prob
def _expected_improvement(x):
# calculate the best surrogate score found so far
x_history, _ = self._vectorize_trials()
y_pred = self.gpr.predict(x_history, return_std=False)
y_best = max(y_pred)
# calculate mean and stdev via surrogate function
x = x.reshape(1, -1)
mu, sigma = self.gpr.predict(x, return_std=True)
# calculate the probability of improvement
z = (mu - y_best) / (sigma + 1e-9)
ei = (mu - y_best) * norm.cdf(z) + sigma * norm.pdf(z)
return -1 * ei
acq_funcs = {
"ucb": _upper_confidence_bound,
"pi": _probability_of_improvement,
"ei": _expected_improvement,
}
# Sampling based on acquisition functions
optimal_val = float("inf")
optimal_x = None
num_restarts = 50
bounds = self._get_hp_bounds()
x_seeds = self._random_state.uniform(
bounds[:, 0], bounds[:, 1], size=(num_restarts, bounds.shape[0])
)
for x_try in x_seeds:
# Sign of score is flipped when maximizing.
result = scipy_optimize.minimize(
acq_funcs[self.acq_type], x0=x_try, bounds=bounds, method="L-BFGS-B"
)
if result.fun[0] < optimal_val:
optimal_val = result.fun[0]
optimal_x = result.x
values = self._vector_to_values(optimal_x)
return {"status": trial_lib.TrialStatus.RUNNING, "values": values}
def _get_hp_bounds(self):
nonfixed_hp_space = [
hp
for hp in self.hyperparameters.space
if not isinstance(hp, hp_module.Fixed)
]
bounds = []
for hp in nonfixed_hp_space:
bounds.append([0, 1])
return np.array(bounds)
def get_state(self):
state = super(BayesianOptimizationOracle, self).get_state()
state.update(
{
"num_initial_points": self.num_initial_points,
"acq_type": self.acq_type,
"beta": self.beta,
"seed": self.seed,
}
)
return state
def set_state(self, state):
super(BayesianOptimizationOracle, self).set_state(state)
self.num_initial_points = state["num_initial_points"]
self.acq_type = state["acq_type"]
self.beta = state["beta"]
self.seed = state["seed"]
self._random_state = np.random.RandomState(self.seed)
self.gpr = self._make_gpr()
# + [markdown] colab_type="text"
# ### Use customized Bayesian Optimization search algorithm to tune models
#
# + colab_type="code"
bo_tuner = LightGBMTuner(
oracle=BayesianOptimizationOracle(
objective=kt.Objective("mse", "min"),
max_trials=100,
acq_type="ucb", # you can switch between different acquisition functions
seed=42,
),
hypermodel=build_model,
overwrite=True,
project_name="bo_tuner",
)
bo_tuner.search(X_train, y_train, validation_data=(X_val, y_val))
# + colab_type="code"
from sklearn.metrics import mean_squared_error
best_model = bo_tuner.get_best_models(1)[0]
y_pred_test = best_model.predict(X_test)
test_mse = mean_squared_error(y_test, y_pred_test)
print("The prediction MSE on test set: {}".format(test_mse))
# + colab_type="code"
bo_tuner.results_summary(1)
# + [markdown] colab_type="text"
# ### Plot search curves
#
# + colab_type="code"
import matplotlib.pyplot as plt
def plot_curve(x, y, xlabel, ylabel, title):
plt.plot(x, y)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
plt.title(title)
plt.show()
def plot_curves(
x, ys, xlabel, ylabel, title, ymin, ymax, legend, markers, linestyles, markevery=1
):
for i, y in enumerate(ys):
plt.plot(x, y, marker=markers[i], linestyle=linestyles[i], markevery=markevery)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
plt.title(title)
plt.ylim(ymin, ymax)
plt.legend(legend)
plt.show()
# + colab_type="code"
mse_bo = [
bo_tuner.oracle.get_trial(trial_id).score for trial_id in bo_tuner.oracle.end_order
]
ids = list(range(len(mse_bo)))
plot_curve(
ids, mse_bo, "Trials in finishing order", "Validation MSE", "Searched results"
)
high_value = float("inf")
high_mse_bo = []
for value in mse_bo:
high_value = min(high_value, value)
high_mse_bo.append(high_value)
plot_curve(
ids,
high_mse_bo,
"Trials in finishing order",
"Highest validation MSE so far",
"Searched results",
)
# + colab_type="code"
random_tuner = LightGBMTuner(
oracle=kt.oracles.RandomSearch(
objective=kt.Objective("mse", "min"), max_trials=100, seed=42
),
hypermodel=build_model,
overwrite=True,
project_name="random_tuner",
)
random_tuner.search(X_train, y_train, validation_data=(X_val, y_val))
# + colab_type="code"
from sklearn.metrics import mean_squared_error
best_model = random_tuner.get_best_models(1)[0]
y_pred_test = best_model.predict(X_test)
test_mse = mean_squared_error(y_test, y_pred_test)
print("The prediction MSE on test set: {}".format(test_mse))
# + colab_type="code"
random_tuner.results_summary(1)
# + colab_type="code"
mse_random = [
random_tuner.oracle.get_trial(trial_id).score
for trial_id in random_tuner.oracle.end_order
]
mse_bo = [
bo_tuner.oracle.get_trial(trial_id).score for trial_id in bo_tuner.oracle.end_order
]
print(len(mse_random))
print(len(mse_bo))
high_value = float("inf")
high_mse_random = []
for value in mse_random:
high_value = min(high_value, value)
high_mse_random.append(high_value)
high_value = float("inf")
high_mse_bo = []
for value in mse_bo:
high_value = min(high_value, value)
high_mse_bo.append(high_value)
plot_curves(
ids,
[mse_random, mse_bo],
"Trials in finishing order",
"Validation MSE",
"Searched results",
0,
1.5,
markers=["o", "+"],
linestyles=["-", "-."],
legend=["Random search", "Bayesian optimization"],
)
plot_curves(
ids,
[high_mse_random, high_mse_bo],
"Trials in finishing order",
"Highest validation MSE so far",
"Searched results",
0.2,
0.4,
markers=["o", "+"],
linestyles=["-", "-."],
legend=["Random search", "Bayesian optimization"],
markevery=5,
)
| 7.3-Bayesian-Optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys, os
sys.path.append(os.path.abspath(os.path.join('..', 'Scripts')))
# Now do your import
from run_model import *
solution=run_simulation(mode='test')
#20 experiments are conducted
assert len(solution)==20
real_numbers=[num.real for num in list(solution.keys())]
imag_numbers=[num.imag for num in list(solution.keys())]
assert min(real_numbers)==params.R_0_list[0]
assert max(real_numbers)==params.R_0_list[-1]
sum(np.array(imag_numbers)!=0)==0
| Tests/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 그래프, 수학 기능 추가
# Add graph and math features
import pylab as py
import numpy as np
# -
# # 1차 적분<br>First Order Numerical Integration
#
#
# [](https://www.youtube.com/watch?v=1p0NHR5w0Lc)
#
#
# 다시 면적이 1인 반원을 생각해 보자.<br>Again, let's think about the half circle with area of 1.
#
#
# $$
# \begin{align}
# \pi r^2 &= 2 \\
# r^2 &= \frac{2}{\pi} \\
# r &= \sqrt{\frac{2}{\pi}}
# \end{align}
# $$
#
#
# +
r = py.sqrt(2.0 / py.pi)
# +
def half_circle(x):
return py.sqrt(np.abs(r**2 - x**2))
# -
# $$
# y = \sqrt{r^2 - x^2}
# $$
#
#
# +
x_array = py.linspace(-r, r)
y_plus = half_circle(x_array)
py.fill_between(x_array, y_plus)
py.axis('equal')
py.grid(True)
# -
# 이번에는 사다리꼴 규칙을 이용해서 구해 보기로 하자.<br>
# This time, let's use the trapezoid rule to find its area.
#
#
# ## 사다리꼴 규칙<br>Trapezoid Rule
#
#
# 다음과 같은 사다리꼴을 생각해 보자.<br>Let's think about a trapezoid as follows.
#
#
# +
x_array = (0, 1)
y_array = (1, 2)
py.fill_between(x_array, y_array)
py.axis('equal')
py.axis('off')
py.text(-0.25, 0.5, '$y_i$')
py.text(1.15, 1, '$y_{i+1}$')
py.text(0.5, -0.3, '$\Delta x$')
# -
# 사다리꼴의 면적은 다음과 같다.<br>
# Area of a trapezoid is as follows.
#
#
# $$
# a_i=\frac{1}{2} \left( y_i + y_{i+1} \right) \Delta x
# $$
#
#
# ## 1차 적분<br>First order numerical integration
#
#
# 마찬가지로 일정 간격으로 $x$ 좌표를 나누어 보자.<br>
# Same as before, let's divide $x$ coordinates in a constant interval.
#
#
# +
d = r * 2.0
n = 10
x_interval = d / n
x_array = py.linspace(-r, r)
y_plus = half_circle(x_array)
x_array_bar = py.arange(-r, r+x_interval*0.1, x_interval)
y_array_bar = half_circle(x_array_bar)
x_interval = x_array_bar[1]-x_array_bar[0]
py.fill_between(x_array, y_plus)
xp, yp = x_array_bar[0], y_array_bar[0]
for x, y in zip(x_array_bar.tolist()[1:], y_array_bar.tolist()[1:]):
py.fill_between((xp, x), (yp, y), alpha=0.5, color=py.random((1, 3)))
xp, yp = x, y
py.axis('equal')
py.grid(True)
# -
# 사다리꼴의 면적을 하나씩 구해서 더해보자.<br>Let's accumulate the area of trapezoids.
#
#
# $$
# Area = \sum_{k=0}^{n-1} F_k
# $$
#
#
# $$
# F_k = \frac{\Delta x}{2}\left[f(x_k)+f(x_{k+1})\right]
# $$
#
#
# +
def num_int_1(f, xi, xe, delta_x):
x_array = py.arange(xi, xe+delta_x*0.1, delta_x)
integration_result = 0.0
xp = x_array[0]
yp = f(xp)
for x_i in x_array[1:]:
y_i = f(x_i)
area_i = 0.5 * (yp + y_i) * (x_i - xp)
xp, yp = x_i, y_i
integration_result += area_i
return integration_result
# +
n = 10
result = num_int_1(half_circle, -r, r, 2*r/n)
print('result =', result)
# -
# 예상한 값 1에 더 비슷한 값을 얻기 위해 더 잘게 나누어 보자<br>
# To obtain the result closer to the expected value of 1, let's divide with a narrower interval.
#
#
# +
n = 100
result = num_int_1(half_circle, -r, r, 2*r/n)
print('result =', result)
# -
# 도전 과제 1 : 다른 조건이 같을 때 0차 적분과 사다리꼴 적분의 오차를 비교해 보시오. 필요하면 해당 파이썬 함수를 복사하시오.<br>Try this 1 : Compare the errors of the zeroth and first order integrations of the half circle example above using the same conditions. Duplicate the python function if necessary.
#
#
# 도전 과제 2 : 길이 $L=3[m]$ 인 외팔보가 분포 하중 $\omega=50sin\left(\frac{1}{2L}\pi x\right)[N/m]$을 받고 있을 때 전단력과 굽힘모멘트를 구하시오.<br>Try this 2 : Calculate shear force and bending moment of a cantilever with length $L=3m$ under distributed load $\omega=50sin\left(\frac{1}{2L}\pi x\right)[N/m]$. <br>
# (ref : C 4.4, Pytel, Kiusalaas & Sharma, Mechanics of Materials, 2nd Ed, SI, Cengage Learning, 2011.)
#
#
# ## Final Bell<br>마지막 종
#
#
# +
# stackoverfow.com/a/24634221
import os
os.system("printf '\a'");
# +
| 30_num_int/01_first_order.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity 3.1: The effect of ensemble size
# ## 1. Introduction
#
# In this activity, we will perform data assimilation using Stochastic Ensemble Kalman filter, where the ensemble is generated by adding different noise to the output of the observation operator, that is, to the model variables mapped to the observations. We are interested in how the ensemble size affects the analysis.
#
# We will perform identical twin experiments as in the previous activities, but we will now use a Lorenz 96 dynamical model with 12 variables. In this model, the variables represent some atmospheric quantities at specific locations. These locations are placed on a circle and are equally spaced.
import numpy as np
from tools.common_misc import gen_obs, rmse_spread, createH
from tools.common_plots import plotRMSP
from tools.L96_model import lorenz96
from tools.L96_kfs import kfs_lor96
from tools.L96_plots import plotL96, plotL96obs, plotL96DA_kf
# ## 2. Compute the Nature Run
#
# As in the previous activities, we generate trajectories for every variable and assume these to be the truth.
# +
# The initial conditions
model = 'L96'
x0 = None # let it spin from rest (x_n(t=0) = F, for all n )
tmax = 5
Nx = 12
t, xt = lorenz96(tmax, x0, Nx) # Nx>=12
plotL96(t, xt, Nx)
# imperfect initial guess for our DA experiments
forc = 8.0
aux1 = forc * np.ones(Nx)
aux2 = range(Nx)
x0guess = aux1 + ((-1)*np.ones(Nx))**aux2
# -
# ## 3. Generate observations
#
# We now generate the observations by adding noise to selected variables at selected times. We don’t have observations for every variable, as in real life; in our example, we will have observations for every second variable. The variables are observed at every 10 time steps. The observations are shown with red dots.
# +
### 2. The observations
# Decide what variables to observe
obsgrid = '1010'
H, observed_vars = createH(obsgrid, model, Nx)
period_obs = 10
var_obs = 2
# Generating the observations
seed = 1
tobs, y, R = gen_obs(t, xt, period_obs, H, var_obs, seed)
plotL96obs(t, xt, Nx, tobs, y, observed_vars)
# -
# ## 4. Ensemble size
#
# In the experiments, we show the analysis for each ensemble member and their mean. The root mean square error of the ensemble mean is compared to the ensemble spread: a good match means that we can trust the ensemble and the uncertainty is estimated well, whereas a spread smaller than the root mean square error signals that the uncertainty is underestimated, and a spread larger than the root mean square error signals the uncertainty is overestimated.
#
# Recall that an assimilation is performed at every observation time and background is generated for every assimilation.
#
# Change the size of the ensemble (M), for example set M=2, M=10, M=100. How do the analysis mean and the root mean squared error change? What happens to the ensemble spread? Note the different scales for the root mean squared error.
# +
### 3. Data assimilation using KFa (SEnKF, LSEnKF and ETKF)
# No LETKF since R-localisation is extremely slow without parallel implementation
rho = 0.1
M = 2
lam = 2
loctype = 'GC'
met = 'SEnKF'
Xb,xb,Xa,xa,locmatrix = kfs_lor96(x0guess,t,tobs,y,H,R,rho,M,met,lam,loctype)
plotL96DA_kf(t,xt,tobs,y,Nx,observed_vars,Xb,xb,Xa,xa)
rmse_step=1
rmseb,spreadb = rmse_spread(xt,xb,Xb,rmse_step)
rmsea,spreada = rmse_spread(xt,xa,Xa,rmse_step)
plotRMSP(t,rmseb,rmsea,spreadb,spreada)
# +
M = 10
Xb,xb,Xa,xa,locmatrix = kfs_lor96(x0guess,t,tobs,y,H,R,rho,M,met,lam,loctype)
plotL96DA_kf(t,xt,tobs,y,Nx,observed_vars,Xb,xb,Xa,xa)
rmse_step=1
rmseb,spreadb = rmse_spread(xt,xb,Xb,rmse_step)
rmsea,spreada = rmse_spread(xt,xa,Xa,rmse_step)
plotRMSP(t,rmseb,rmsea,spreadb,spreada)
# +
M = 100
Xb,xb,Xa,xa,locmatrix = kfs_lor96(x0guess,t,tobs,y,H,R,rho,M,met,lam,loctype)
plotL96DA_kf(t,xt,tobs,y,Nx,observed_vars,Xb,xb,Xa,xa)
rmse_step=1
rmseb,spreadb = rmse_spread(xt,xb,Xb,rmse_step)
rmsea,spreada = rmse_spread(xt,xa,Xa,rmse_step)
plotRMSP(t,rmseb,rmsea,spreadb,spreada)
# -
# ## 5. Conclusions
#
# We performed an analysis using the Stochastic Ensemble Kalman filter. Different ensemble sizes were used and we observed its effect on the analysis: a small ensemble may lead to an analysis of poor quality and incorrect uncertainty estimates.
| 3-1_ensemble_activity_ensemble_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#A simple analysis of Lending Club Loan Data to identify some of the factors leading to different interest rates.
#For portions of this (as noted) I worked with <NAME>
import pandas as pd
import pandas.io.data as iod
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mp
import datetime as dt
# %matplotlib inline
# -
#The data can be pulled from Lending Club as a CSV
#When I completed this analysis it was published every couple of weeks
data_frame = pd.read_csv('treefo.csv', dtype=object)
data_frame.head()
# +
#hat tip to John for this part
loan_data = list()
weights_dictionary={}
for loan_id in data_frame.iloc[0:235628]['id']:
sub_df = data_frame.ix[data_frame.id== loan_id]
series_data = sub_df.iloc()[0]
int_rate = series_data['int_rate']
loan_status = series_data['loan_status']
dti = series_data['dti']
delinq = series_data['delinq_2yrs']
earliest = series_data['earliest_cr_line']
pub_records = series_data['pub_rec']
try:
earliest_v1= 2000 + int(earliest[0])
try:
int(earliest[1])
earliest_v1= 2000+int(earliest[0:2])
except:
pass
except:
earliest_v1 = 1900 + int(earliest[4:6])
if earliest_v1 == 1900:
earliest_v1 = 2000
#print(earliest_v1)
earliest_v2=2015-earliest_v1
loan_info = loan_id,int_rate,loan_status,dti,delinq,pub_records,earliest_v2
loan_data.append(loan_info)
#print(loan_data)
loan_list = list()
if dti < 10:
dti_value=1
elif 10 < dti < 20:
dti_value=2
elif dti > 20:
dti_value=3
loan_list.append(dti_value)
if delinq ==0:
delinq_value=1
elif delinq ==1:
delinq_value=2
elif delinq>1:
delinq_value=3
loan_list.append(delinq_value)
if pub_records ==0:
pub_records_value=1
elif pub_records ==1:
pub_records_value=2
elif pub_records>1:
pub_records_value=3
loan_list.append(pub_records_value)
if earliest_v2 >20:
earliest_value=1
elif 8 < earliest_v2 < 21:
earliest_value=2
elif earliest_v2 <9:
earliest_value=3
loan_list.append(earliest_value)
weight_sum= (sum(loan_list))
#print(loan_list)
#print(weight_sum)
weights_dictionary[loan_id]= weight_sum
#print(weights_dictionary)
low_risk_list=list()
med_low_risk_list=list()
med_risk_list=list()
med_hi_risk_list=list()
hi_risk_list=list()
for loan_id in weights_dictionary.keys():
if weights_dictionary[loan_id] < 6:
low_risk_list.append(loan_id)
elif 5 < weights_dictionary[loan_id] < 7:
med_low_risk_list.append(loan_id)
elif 6 < weights_dictionary[loan_id] < 9:
med_risk_list.append(loan_id)
elif 8 < weights_dictionary[loan_id] < 11:
med_hi_risk_list.append(loan_id)
elif weights_dictionary[loan_id] > 10:
hi_risk_list.append(loan_id)
# -
low_frame = data_frame.ix[data_frame['id'].isin(low_risk_list)]
low_frame = low_frame[low_frame.out_prncp != 0] #make sure it has a positive balance to invest
low_frame.to_csv('low_frame.csv')
med_low_frame = data_frame.ix[data_frame['id'].isin(med_low_risk_list)]
med_low_frame = med_low_frame[med_low_frame.out_prncp != 0]
med_low_frame.to_csv('med_low_frame.csv')
med_risk_frame = data_frame.ix[data_frame['id'].isin(med_risk_list)]
med_risk_frame = med_risk_frame[med_risk_frame.out_prncp != 0]
med_risk_frame.to_csv('med_frame.csv')
med_hi_risk_frame = data_frame.ix[data_frame['id'].isin(med_hi_risk_list)]
med_hi_risk_frame = med_hi_risk_frame[med_hi_risk_frame.out_prncp != 0]
med_hi_risk_frame.to_csv('med_hi_frame.csv')
hi_risk_frame = data_frame.ix[data_frame['id'].isin(hi_risk_list)]
hi_risk_frame = hi_risk_frame[hi_risk_frame.out_prncp != 0]
hi_risk_frame.to_csv('hi_frame.csv')
# +
#Get the 5 year rate from yahoo and compare it to the average of Lending Club's Portfolio
int_mean = np.mean(data_frame['int_rate'])
int_mean
five_year_rate = iod.get_data_yahoo('^FVX', start=dt.datetime(2015,4,29), end=dt.date.today())
latest_day = len(five_year_rate) - 1
latest_day
five_year_plot=five_year_rate.iloc[latest_day]['Close']
five_year_plot
N = 1
plots=[five_year_plot,int_mean]
ind = np.arange(N)
width = 1
fig, ax = plt.subplots()
rects1 = ax.bar(ind, five_year_plot, width, color='r')
rects2 = ax.bar(ind+width, int_mean, width, color='y')
ax.set_ylabel('Interest Rate')
ax.set_xlabel('Which one do you like?')
ax.set_title('Avg Lending Tree vs 5 year US Bond')
ax.set_xticks(ind+width)
ax.legend( (rects1[0], rects2[0]), ('5 Year Bond', 'Lending Club'))
# -
low_int_std = np.std(low_frame['int_rate'])
low_int_mean = np.mean(low_frame['int_rate'])
low_prin_mean = np.mean(low_frame['out_prncp'])
low_prin_sum = np.sum(low_frame['out_prncp'])
med_low_int_std = np.std(med_low_frame['int_rate'])
med_low_int_mean = np.mean(med_low_frame['int_rate'])
med_low_prin_mean = np.mean(med_low_frame['out_prncp'])
med_low_prin_sum = np.sum(med_low_frame['out_prncp'])
med_risk_int_std = np.std(med_risk_frame['int_rate'])
med_risk_int_mean = np.mean(med_risk_frame['int_rate'])
med_risk_prin_mean = np.mean(med_risk_frame['out_prncp'])
med_risk_prin_sum = np.sum(med_risk_frame['out_prncp'])
med_hi_int_std = np.std(med_hi_risk_frame['int_rate'])
med_hi_int_mean = np.mean(med_hi_risk_frame['int_rate'])
med_hi_prin_mean = np.mean(med_hi_risk_frame['out_prncp'])
med_hi_prin_sum = np.sum(med_hi_risk_frame['out_prncp'])
hi_int_std = np.std(hi_risk_frame['int_rate'])
hi_int_mean = np.mean(hi_risk_frame['int_rate'])
hi_prin_mean = np.mean(hi_risk_frame['out_prncp'])
hi_prin_sum = np.sum(hi_risk_frame['out_prncp'])
# +
#Check to see if we are in the right ballpark with our analysis ...
N = 5
intmeans = (low_prin_mean,med_low_prin_mean,med_risk_prin_mean,med_hi_prin_mean,hi_prin_mean)
ind = np.arange(N)
width = 0.35
fig, ax = plt.subplots()
rects1 = ax.bar(ind, intmeans, width)
rects1[0].set_color('green')
rects1[1].set_color('blue')
rects1[2].set_color('yellow')
rects1[3].set_color('orange')
rects1[4].set_color('red')
ax.set_ylabel('Average Loan Size, in $')
ax.set_title('Average Loan Size by risk level')
ax.set_xticks(ind+width)
ax.set_xticklabels( ('low','med-low','med','med-hi','hi') )
plt.show()
# +
#Mean and Standard Deviation ...
N = 5
intmeans = (low_int_mean,med_low_int_mean,med_risk_int_mean,med_hi_int_mean,hi_int_mean)
intstds = (low_int_std,med_low_int_std,med_risk_int_std,med_hi_int_std,hi_int_std)
ind = np.arange(N)
width = 0.35
fig, ax = plt.subplots()
rects1 = ax.bar(ind, intmeans, width, color='r', yerr=intstds)
rects1[0].set_color('green')
rects1[1].set_color('blue')
rects1[2].set_color('yellow')
rects1[3].set_color('orange')
rects1[4].set_color('red')
ax.set_ylabel('Interest Rate')
ax.set_title('Mean and stan dev of interest rate by risk level')
ax.set_xticks(ind+width)
ax.set_xticklabels( ('low','med-low','med','med-hi','hi') )
plt.show()
# +
top_5_low = low_frame.sort(['int_rate'], ascending=False)
top_5_low = top_5_low[0:5]
top_5_low.to_csv('top_5_low_frame.csv')
top_5_med_low = med_low_frame.sort(['int_rate'], ascending=False)
top_5_med_low = top_5_med_low[0:5]
top_5_med_low.to_csv('top_5_med_low.csv')
top_5_med = med_risk_frame.sort(['int_rate'], ascending=False)
top_5_med = top_5_med[0:5]
top_5_med.to_csv('top_5_med.csv')
top_5_med_hi = med_hi_risk_frame.sort(['int_rate'], ascending=False)
top_5_med_hi = top_5_med_hi[0:5]
top_5_med_hi.to_csv('top_5_med_hi.csv')
top_5_hi = hi_risk_frame.sort(['int_rate'], ascending=False)
top_5_hi = top_5_hi[0:5]
top_5_hi.to_csv('top_5_hi.csv')
# -
low_frame = pd.read_csv('low_frame.csv')
med_low_frame = pd.read_csv('med_low_frame.csv')
med_risk_frame = pd.read_csv('med_frame.csv')
med_hi_risk_frame = pd.read_csv('med_hi_frame.csv')
hi_risk_frame = pd.read_csv('med_hi_frame.csv')
# +
#Checking the size of the loan and graphing it.
N = 5
amtsums = (low_prin_sum,med_low_prin_sum,med_risk_prin_sum,med_hi_prin_sum,hi_prin_sum)
ind = np.arange(N)
width = 0.35
fig, ax = plt.subplots()
rects1 = ax.bar(ind, amtsums, width)
rects1[0].set_color('green')
rects1[1].set_color('blue')
rects1[2].set_color('yellow')
rects1[3].set_color('orange')
rects1[4].set_color('red')
ax.set_ylabel('Total Loan Size, in $')
ax.set_title('Total Loan Size by risk level')
ax.set_xticks(ind+width)
ax.set_xticklabels( ('low','med-low','med','med-hi','hi') )
plt.show()
# -
#get the top loans from any risk category (low the example below)
top_5_low = low_frame.sort(['int_rate'], ascending=False)
| LC_risk_analysis_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mergesort
# https://www.youtube.com/watch?v=3aTfQvs-_hA&t=17s
#
# Conceptually, a merge sort works as follows:
# 1. Divide the unsorted list into n sublists, each containing one element (a list of one element is considered sorted).
# 2. Repeatedly merge sublists to produce new sorted sublists until there is only one sublist remaining. This will be the sorted list.
# %run ../create_array.py
# a and b must be sorted arrays
def _merge(a, b):
merged = []
a_idx, b_idx = 0, 0
while a_idx < len(a) and b_idx < len(b): # Run through both lists
if a[a_idx] < b[b_idx]: # If its less than
merged.append(a[a_idx]) # Add it to the merged list
a_idx += 1
else:
merged.append(b[b_idx]) # If add b is it's less
b_idx += 1
# We extend the merged array with the last element of the list we didn't exhaust in the while loop
if a_idx == len(a):
merged.extend(b[b_idx:])
else:
merged.extend(a[a_idx:])
return merged
_merge(sorted(create_array(25, 10)), sorted(create_array(25, 10)))
def mergesort(a):
if len(a) <= 1: # Our base case
return a
left, right = mergesort(a[:len(a)//2]), mergesort(a[len(a)//2:]) # left is the 1st half of the array, vice versa
return _merge(left, right) # Now we sort
mergesort(create_array(25, 10))
| sorting/mergesort.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Table of Contents
# * [Multigroup aspatial indexes of segregation](#Multigroup-aspatial-indexes-of-segregation)
# * [Multigroup Dissimilarity Index](#Multigroup-Dissimilarity-Index)
# * [Multigroup Gini Index](#Multigroup-Gini-Index)
# * [Multigroup Normalized Exposure Index](#Multigroup-Normalized-Exposure-Index)
# * [Multigroup Information Theory Index](#Multigroup-Information-Theory-Index)
# * [Multigroup Relative Diversity Index](#Multigroup-Relative-Diversity-Index)
# * [Multigroup Squared Coefficient of Variation Index](#Multigroup-Squared-Coefficient-of-Variation-Index)
# * [Multigroup Diversity Index](#Multigroup-Diversity-Index)
# * [Simpson's Concentration Index (lambda)](#Simpson's-Concentration-Index-%28lambda%29)
# * [Simpson's Interaction Index (I)](#Simpson's-Interaction-Index-%28I%29)
# * [Multigroup Divergence Index](#Multigroup-Divergence-Index)
#
# # Multigroup aspatial indexes of segregation
# This is an example notebook of functionalities for multigroup aspatial indexes of the *segregation* module. Firstly, we need to import the packages we need:
# +
# %%capture
import libpysal
import segregation
import geopandas as gpd
# -
# Then it's time to load some data to estimate segregation. We use the data of 2000 Census Tract Data for the metropolitan area of Sacramento, CA, USA.
#
# We use a geopandas dataframe available in PySAL examples repository.
#
# For more information about the data: https://github.com/pysal/libpysal/tree/master/libpysal/examples/sacramento2
input_df = gpd.read_file(libpysal.examples.get_path("sacramentot2.shp"))
input_df.columns
# The groups of interest are White, Black, Asian and Hispanic population. Therefore, we create an auxiliary list with only the necessary columns for fitting the index.
groups_list = ['WHITE', 'BLACK', 'ASIAN','HISP']
# We also can plot the spatial distribution of the composition of each of these groups over the tracts of Sacramento:
# +
import matplotlib.pyplot as plt
for i in range(len(groups_list)):
input_df['comp_' + groups_list[i]] = input_df[groups_list[i]] / input_df['TOT_POP']
fig, axes = plt.subplots(ncols = 2, nrows = 2, figsize = (17, 10))
input_df.plot(column = 'comp_' + groups_list[0],
cmap = 'OrRd',
legend = True, ax = axes[0,0])
axes[0,0].set_title('Composition of ' + groups_list[0])
axes[0,0].set_xticks([])
axes[0,0].set_yticks([])
axes[0,0].set_facecolor('white')
input_df.plot(column = 'comp_' + groups_list[1],
cmap = 'OrRd',
legend = True, ax = axes[0,1])
axes[0,1].set_title('Composition of ' + groups_list[1])
axes[0,1].set_xticks([])
axes[0,1].set_yticks([])
axes[0,1].set_facecolor('white')
input_df.plot(column = 'comp_' + groups_list[2],
cmap = 'OrRd',
legend = True, ax = axes[1,0])
axes[1,0].set_title('Composition of ' + groups_list[2])
axes[1,0].set_xticks([])
axes[1,0].set_yticks([])
axes[1,0].set_facecolor('white')
input_df.plot(column = 'comp_' + groups_list[3],
cmap = 'OrRd',
legend = True, ax = axes[1,1])
axes[1,1].set_title('Composition of ' + groups_list[3])
axes[1,1].set_xticks([])
axes[1,1].set_yticks([])
axes[1,1].set_facecolor('white')
# -
# ## Multigroup Dissimilarity Index
# %%capture
from segregation.aspatial import MultiDissim
index = MultiDissim(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Gini Index
# %%capture
from segregation.aspatial import MultiGiniSeg
index = MultiGiniSeg(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Normalized Exposure Index
# %%capture
from segregation.aspatial import MultiNormalizedExposure
index = MultiNormalizedExposure(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Information Theory Index
# %%capture
from segregation.aspatial import MultiInformationTheory
index = MultiInformationTheory(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Relative Diversity Index
# %%capture
from segregation.aspatial import MultiRelativeDiversity
index = MultiRelativeDiversity(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Squared Coefficient of Variation Index
# %%capture
from segregation.aspatial import MultiSquaredCoefficientVariation
index = MultiSquaredCoefficientVariation(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Diversity Index
# %%capture
from segregation.aspatial import MultiDiversity
index = MultiDiversity(input_df, groups_list)
type(index)
index.statistic
# Normalized version of the multigroup diversity index
normalized_index = MultiDiversity(input_df, groups_list, normalized = True)
normalized_index.statistic
# ## Simpson's Concentration Index (lambda)
# %%capture
from segregation.aspatial import SimpsonsConcentration
index = SimpsonsConcentration(input_df, groups_list)
type(index)
index.statistic
# ## Simpson's Interaction Index (I)
# %%capture
from segregation.aspatial import SimpsonsInteraction
index = SimpsonsInteraction(input_df, groups_list)
type(index)
index.statistic
# ## Multigroup Divergence Index
# %%capture
from segregation.aspatial import MultiDivergence
index = MultiDivergence(input_df, groups_list)
type(index)
index.statistic
| notebooks/multigroup_aspatial_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BreakoutMentors/Data-Science-and-Machine-Learning/blob/main/machine_learning/lesson%203%20-%20Neural%20Networks/Intro_to_CNNs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JOquPv8N7VkH"
# # Introduction to Convolutional Neural Networks
# 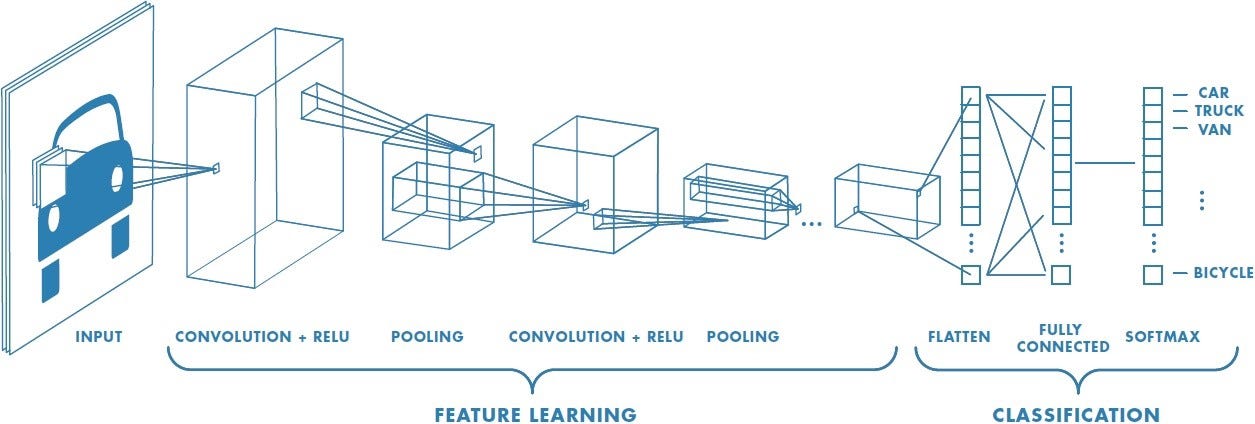
#
#
# ## What are Convolutional Neural Networks (CNNs)?
# A Convolutional Neural Network is a type of neural network and Deep Learning algorithm that has been very popular in the last 10 years for its great involvement in improving Computer Vision (CV). They are excellent at classifying objects in an images that are being implemented in many areas, especially Medicine!
#
# ## How are CNNs contributing to Medicine?
# Convolutional Neural Networks (CNNs) are frequently used to help doctors identify cancer and other diseases in medical images of patients, as they have proven to be nearly as accurate as a team of medical experts at diagnosing patients. This tech enables doctors to provide faster and better treatment, cutting a lot of time out of the diagnosis process, and saving lives as a result.
#
# CNNs are being studied and improved at the best medical universities around the world, for the hope of having CNNs being applied for the diagnosis process especially in areas where medical specialists are rare.
#
# ## What is different between a CNN and a normal Neural Network?
# The main difference between these two algorithms is in how they learn features. Specifically, the difference occurs in the *Feature Learning* section from the CNN image above. In that section, we see that an image is being inserted at the input and it goes through a process where information is being picked from the image then summerized through **Convolutional Layers**. The information that is being picked are features in the image, such as the lights, tire, and shape of the car in the image. The reason it is called *Feature Learning* is because the way it looks for features in the images is automized as well, therefore the weights used in the convolutional layers are being trained and improved overtime.
#
# You can also see that the input image gets smaller overtime, which is good since we are wanting to gather the most important features. This action of reducing the image down to its most significant features is called *encoding*. So when we use CNNs in this way, we are *encoding* the data in the images to be used for classifying whether an image contains a disease or not.
#
# The *Classification* section of the CNN is just a normal (linear) Neural Network where the encoded features are learned to then give predictions at the end. The encoded features are flattened because the flattened features will be the input of the input layer. So you can see that a CNN is just a special type of Neural Network where it learns to get the most important information from the images.
# + [markdown] id="5P1aUCNObbA9"
# ## Loading the data from Kaggle
#
# [Click here to access the dataset](https://www.kaggle.com/preetviradiya/covid19-radiography-dataset)
#
# If you do not have an account with Kaggle, please make an account with them.
#
# After making an account, you need to download a *kaggle.json* file on your local machine that provides the api to download the dataset.
#
# To download this file follow the instructions below:
# 1. Click on your account in the top right
# 2. Click account settings
# 3. Scroll down to the 'API' section
# 4. Click 'Create New API Token' then it should download
#
# The *kaggle.json* file is needed when it asks you to upload a file below before it downloads the dataset to this Colab.
# + id="m7e4fSwavrak" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 123} outputId="06fec7e3-f597-41d1-cf26-602de99d0533"
from google.colab import files
from IPython.utils import io
import os
files.upload()
os.system("mkdir -p ~/.kaggle")
os.system("cp kaggle.json ~/.kaggle/")
os.system("chmod 600 ~/.kaggle/kaggle.json")
# !kaggle datasets download -d preetviradiya/covid19-radiography-dataset
with io.capture_output() as captured:
# !unzip covid19-radiography-dataset.zip && rm covid19-radiography-dataset.zip
# + [markdown] id="HtTlliSwdySB"
# ## Looking at the distrubition of our classes
#
# The reason to look at the distribution is to understand how the number of samples per class can skew the predictions of the model. We see that their is an overwelming amount of normal X-Ray images that is greater than the other categories.
# + id="AhyttnS5we01" colab={"base_uri": "https://localhost:8080/", "height": 336} outputId="e26da0af-74d1-4abd-f671-ab05b0d0902b"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import PIL
# %matplotlib inline
dataframe = pd.read_csv("/content/metadata.csv")
dataframe['label'].value_counts().plot.bar(color=['b', 'r', 'orange', 'g'])
plt.xticks(rotation=45)
plt.title('Class Distribution')
plt.show()
# + [markdown] id="7bvHZjjmdi1o"
# ## Data Preparation
# Just as our other projects/lessons, image data needs to be prepared to be used with PyTorch. One thing you might see different is the inclusion of PyTorch's [*Dataset*](https://pytorch.org/docs/stable/data.html?highlight=dataset#torch.utils.data.Dataset) and [*Dataloader*](https://pytorch.org/docs/stable/data.html?highlight=dataloader#torch.utils.data.DataLoader) objects. These two classes allow us to easily pair images with their labels that is iterable through a for-loop to get a single batch of data, which we customize. Our data is organized by the name of the folders having the name of the class that each subdirectory image belongs to. Because of this we use [*ImageFolder*](https://pytorch.org/vision/stable/datasets.html#imagefolder) that returns a PyTorch Dataframe ready to be loaded into dataloaders.
#
# ## Image Processing
# If you remember from the past lessons where we used the *MNIST* dataset, each image was flattened as input into the neural network, which is an example of image processing but we didn't get into other transformations. PyTorch has many built-in image transformations that are very helpful for the performance of Convolutional Neural Networks. These are my favorites below:
#
# 1. [Resize](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.Resize): Resizes the images to a desired size while keeping its aspect ratio.
# 2. [Center Crop](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.CenterCrop): Crops an image in the center to the desired size (Height x Width)
# 3. [Random Resized Crop](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.RandomResizedCrop): This takes a random crop of your image with a random aspect ratio, then resizes the random crop to your desired size. This is helpful for getting different varation of objects in your images that it has to predict for.
# 4. [Random Horizontal Crop](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.RandomHorizontalFlip): Flips images horizontally with a given probability that you provide. This is really helpful to make sure the CNN learns features in the images that are influenced by the location inside the images. **This project does not use it because it is concievable that disease can effect certain areas of the lungs, therefore we want the CNN to learn features that are related to location.**
# 6. [Random Rotation](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.RandomRotation): This randomly rotates an image given your desired degrees in the range (-degrees, +degrees).
# 5. [Grayscale](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.Grayscale): This converts colored images to black-and-white images, this is helpful if you want to do make your data smaller that will make the training process faster and will hold less memory in your machine.
# 6. [ColorJitter](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.ColorJitter): This randomly changes the brightness, contrast, saturation and hue of each image. This brings some variation to the color of the images which can be helpful to have the CNN generalize the learning process.
# 7. [Padding](https://pytorch.org/vision/stable/transforms.html#torchvision.transforms.Pad): This adds a border layer around the image with a number of your choice (default is 0) which can be helpful when constructing your Convolutional Layers.
#
# You will see we only used the transformations Resize and ColorJitter below, we only used these because this dataset contains X-Ray images of infected lungs and it would be very important to keep the images as they are based on the assumption that diseases may effect different areas of the lungs. Therefore any transformations that transform the images based on location may inhibit the learning process of the model. The images were resized for the purpose of constructing the Convolutional Layers of the CNN, and changing the contrast randomly can perhaps highlight some infected areas in the images that the CNN can learn.
# + id="Td3ISkXx2zzN" colab={"base_uri": "https://localhost:8080/"} outputId="ea9444da-cfeb-4d08-ba09-0adbec1bce5f"
import torch
from torchvision.datasets import ImageFolder
import torchvision.transforms as transforms
root = "/content/COVID-19_Radiography_Dataset/COVID-19_Radiography_Dataset"
# ImageNet means and stds to normalize images in entire dataset
means = (0.485, 0.456, 0.406)
stds = (0.229, 0.224, 0.225)
# Add more transforms tomorrow
transforms = transforms.Compose([
# Image size is being resized to 3x224x224
transforms.Resize(224),
# Contrast Factor is being selected randomly between [0.5, 1.5]
transforms.ColorJitter(contrast=0.5),
# Converting images to PyTorch tensors
transforms.ToTensor(),
# Normalizing images with Means and STDs of ImageNet
transforms.Normalize(means, stds)
])
# Using ImageFolder to read in the images with their labeled folders
dataset = ImageFolder(root, transform=transforms)
# Calculating the amount of images to split the dataset
trainToTestRatio = 0.8
trainSize = int(trainToTestRatio * len(dataset))
testSize = len(dataset) - trainSize
# Calculating the amount to split the training set into validation set
valSize = int(trainSize * 0.1) # Taking 10% of training data
trainSize = trainSize - valSize # Updating the amount of training data
# Splitting Data
training_data, val_data, testing_data = torch.utils.data.random_split(dataset, lengths=[trainSize, valSize, testSize])
# Loading Data
batch_size = 16
train_dataloader = torch.utils.data.DataLoader(training_data, batch_size=batch_size, shuffle=True)
test_dataloader = torch.utils.data.DataLoader(testing_data, batch_size=batch_size)
val_dataloader = torch.utils.data.DataLoader(val_data, batch_size=batch_size)
# Using dictionaries to easily access all the dataloaders and datasets
loaders = {'train':train_dataloader, 'valid':val_dataloader, 'test':test_dataloader}
datasets = {'train':training_data, 'valid':val_data, 'test':testing_data}
print("These are the classes:", dataset.class_to_idx)
# + id="7SfHawJh3nnn" colab={"base_uri": "https://localhost:8080/"} outputId="dcd8af7a-d815-43f0-dbe8-397f2cc3d87c"
import torchvision.transforms.functional as F
# Function to reverse normalization of the images
def unNormalizeTensor(tensor, means, stds):
tensor_duplicate = tensor.detach().clone()
for channel, mean, std in zip(tensor_duplicate, means, stds):
channel.mul_(std).add_(mean)
return tensor_duplicate
# Printing the dimensions of each batch
images, labels = next(iter(train_dataloader))
print("Batch Image Dimensions:", images.size())
print("Batch Label Dimensions:", labels.size())
# + [markdown] id="6cHH8p5J0fAq"
# ### Images without Normalization
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="JmtzV8l0R3oe" outputId="790dbc28-b5d3-4af9-c507-76ad02625a02"
# Printing images and their labels
print("\nImages without Normalization:")
idx_to_class = {value:key for key,value in dataset.class_to_idx.items()}
plt.figure(figsize=(10, 9))
for i in range(5):
plt.subplot(1, 5, i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
image = F.to_pil_image(unNormalizeTensor(images[i], means, stds))
plt.imshow(image)
plt.xlabel(idx_to_class[labels[i].item()])
# + [markdown] id="3cSFhDZS0lfT"
# ### Images with Normalization
#
# You can see that normalizing the data can enhance some features in the lungs. And it looks pretty cool!
# + colab={"base_uri": "https://localhost:8080/", "height": 176} id="gc6J1a-eR-vH" outputId="1fcd195b-52db-4b38-84b8-14785f060259"
print("\nImages with Normalization:")
plt.figure(figsize=(10, 9))
for i in range(5):
plt.subplot(1, 5, i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
image = F.to_pil_image(images[i])
plt.imshow(image)
plt.xlabel(idx_to_class[labels[i].item()])
# + [markdown] id="0BcfaJ4K02m1"
# ## Constucting the Convolutional Neural Network
#
# ### Convolutional Layers
# 
#
# The GIF above shows the Convolutional Process when learning features from the images. The input is an image with 3 Color Channels for RGB, and the filters or kernels are scanning across it gathering information. There is a lot of customizability through hyperparameters.
#
# The Three Hyperparameters:
# 1. Kernel Size: The Kernels above is a 2-D matrix or 2-D array which often have a squared dimension (i.e., Height = Width) with weights inside them that are trainable (i.e., our algorithm can learn the best weights for the problem we want to solve). Kernels are responsible of finding features inside an image and condensing that information in the output, that is why they scan over the image. If you use a small kernel size, you retain a lot of information and reduce the memory cost of storing the kernal weights. On the other hand, if you have a larger kernel size, you generalize a lot of information in the image and reduce the output size of the learned features but require more memory to store the kernal weights. In this GIF, the kernel size is 3, since it is 3x3. So this would be considered a small kernal size.
#
# 2. Stride: The number of pixels to hop between to scan the image. For example, when the stride is 1, the kernel only moves to the next right pixel each scan, as shown in the GIF. But if stride is two, the filter moves two pixels to the right. The greater stride you have, the few smaller the output feature size will be. Sometimes, people increase the stride to reduce the memory required by the CNN model.
#
#
# 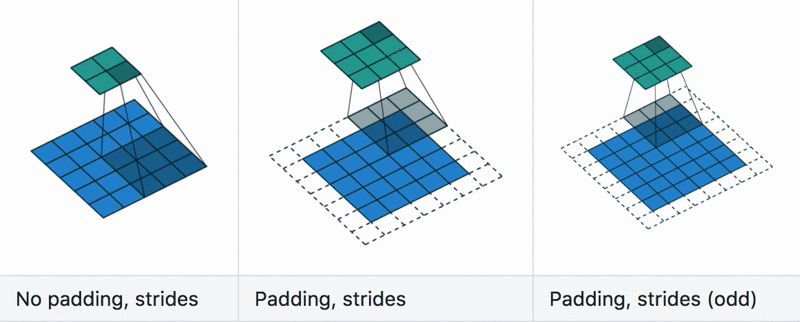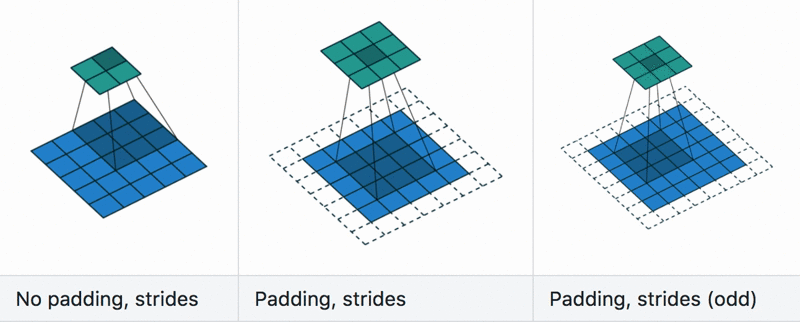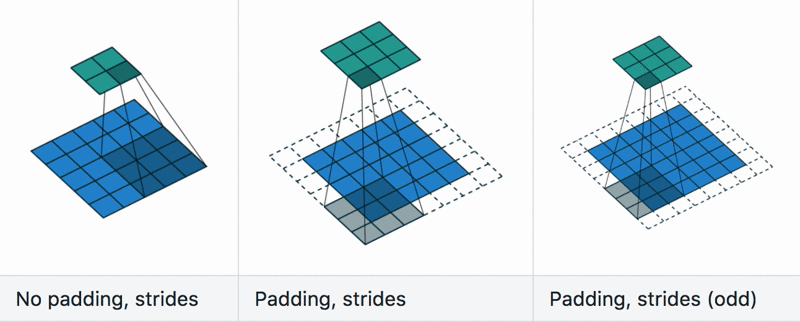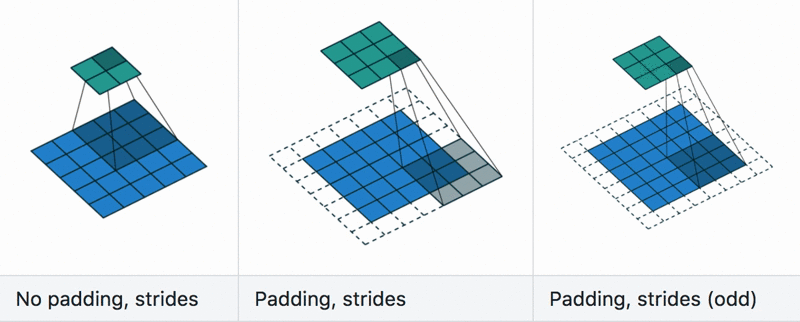
# 3. Padding: Padding as described above in the image processing section is used to add a border around the image (think blank pixels around the border of the image). This can be used to ensure that the shape of the input is compatible with the CNN model.
#
# The image above demonstrates how the output features are effected by different padding and stride settings, keeping a constant kernal size of 3x3. On the left, no padding is used and the stride is 2(`padding = 0, stride=2`). The middle example shows that the output feature size increase to 3x3 when padding is 1 (i.e., adding a border of blank pixels to the input) stride is 2. The benefit of having padding, is that you can retain information of the borders while also retaining the dimensions of the input.
#
# The outputs of the convolutional layers are called *Feature Maps*, and you can control the number of features maps that a convolutional layer produces by adjusting the `out_channels` parameter in PyTorch. This will tell PyTorch how many individual kernal to make for a given layer. For example, for the first layer I said that I want to return 16 feature maps, so there are 16 kernels used. The *in_channels* parameter details the depth of the input of the layer and the depth of the kernels, since the input is an RGB image, the input has a depth of 3 and the kernels will also have a depth of 3 such as the GIF details.
# > Note: In CNNs, *depth* is a synonym for the number of feature maps.
#
# 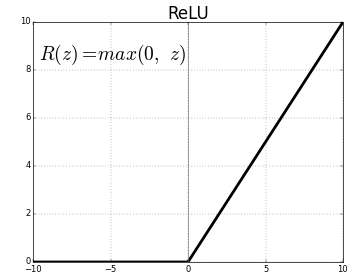
#
# After the feature maps are outputted, we pass each feature through an activation function. Generally, we use this function to find non-linear patterns in our data. The most common activation function used in CNN's is *ReLU* showed above, it is known to produce accurate results while training faster which are both great benefits! After using ReLU, the feature maps are conduced using pooling which is decribed in the next section below.
#
# ### Pooling Layers
#
# 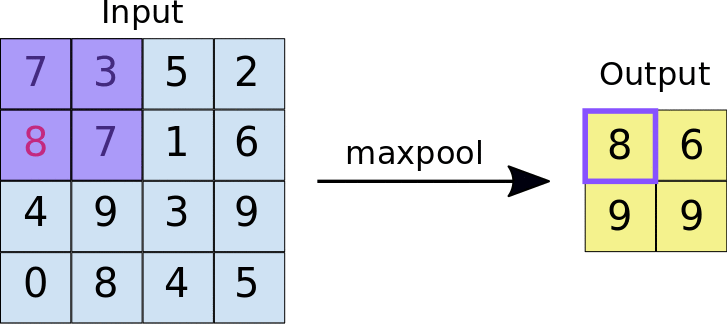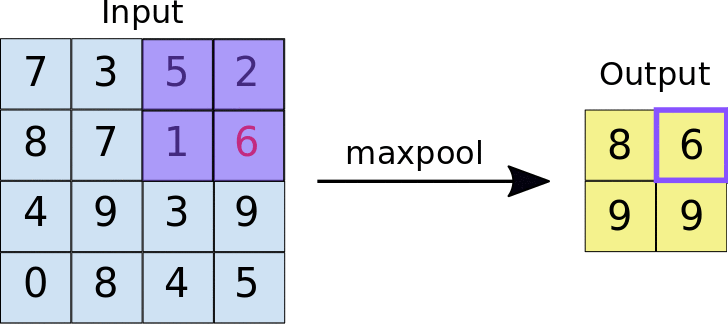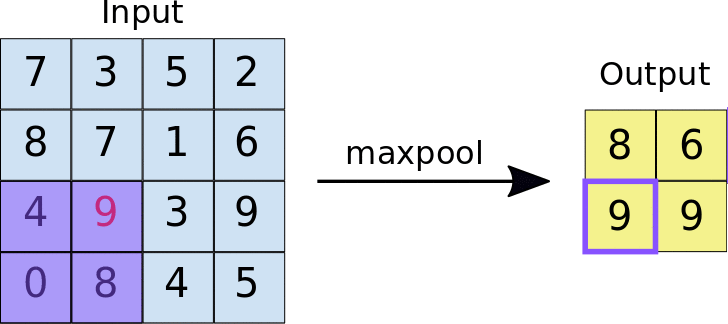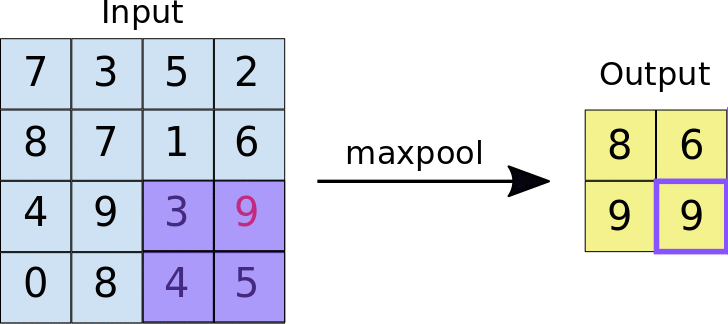
#
# Pooling is another crucial part of the *Feature Learning* process, and you can see from the GIF above that it doesn't look so different from the Convolutional Process above. That's because they are very similar. Pooling layers use kernels and strides to scan the image for the purpose of reducing the dimensions of their input. Usually the input into a pooling layer are the feature maps of a convolutional layer.
#
# The one pooling technique used here is the *MaxPool* technique, where the kernel selects the maximum number of the scanned area it is currently looking at, as shown in the GIF above. Another popular pooling technique is called *AveragePooling* where the kernels computes the average of its current scanned area and outputs that average in its output.
#
# ### Summary
# There are three steps that are usually taken with CNNs:
# 1. Use a convolutional layer to compute feature maps from the input.
# 2. Use ReLU to find non-linear patterns in the features maps.
# 3. Use a pooling technique to reduce dimensionality and save memory.
# 4. Repeat steps 1-3 for each convolutional layer.
# 5. Flatten the last output feature maps so they can be used as input into the fully-connected neural network to be used for classification.
#
#
#
# If you want to understand how the Covolutional Layers work more, here are two great resources:
#
# 1. [CNN Explainer (High-Level)](https://poloclub.github.io/cnn-explainer/): I recommend for everyone to go over this to understand the convolutional process better. It also includes lots of interactive visualizations for those of you who are visual learners!
# 2. [Stanford's CS231n (Very deep and concise)](https://cs231n.github.io/convolutional-networks/#conv): This contains all the information you can need for CNNs, and I would highly recommend you to go over somethings you are confused about if you are brave enough (lot's of math in this resource)! There is also really great formulas to figure out what your hyperparameters should be when creating your own CNNs, and I refer back to this resource a lot.
# + id="mhKHiV2XO5ED"
import torch.nn as nn
class CNN_Model(nn.Module):
# Constructing model
def __init__(self):
super(CNN_Model, self).__init__()
# Defining Batch Normalization
self.norm = nn.BatchNorm2d(num_features=3)
# Defining Convolutional Layers
self.conv1 = nn.Conv2d(in_channels=3, out_channels=12, kernel_size=3, padding=1, stride=1) # Output Dimensions = (12, 224, 224)
self.conv2 = nn.Conv2d(in_channels=12, out_channels=16, kernel_size=3, padding=1, stride=1)
self.conv3 = nn.Conv2d(in_channels=16, out_channels=20, kernel_size=3, padding=1, stride=1)
# Defining MaxPool Layer
self.maxpool = nn.MaxPool2d(kernel_size=2, stride=2)
# Defining Fully Connected Layers
self.fc1 = nn.Linear(self.conv3.out_channels * 28 * 28, 3920) # Input Dimensions before Flattening = (20, 28, 28)
self.fc2 = nn.Linear(3920, 980)
self.fc3 = nn.Linear(980, 245)
self.fc4 = nn.Linear(245, 62)
self.fc5 = nn.Linear(62, 4) # Output Layer with 4 ending nodes for the four classes
# Activation Function to use throughout
self.relu = nn.ReLU()
# Dropout to prevent overfitting
self.dropout = nn.Dropout(p=0.5)
def forward(self, input):
# Batch Normalization for faster training
x = self.norm(input)
# First Convolutional Layer
x = self.relu(self.conv1(x)) # Output Shape = (12, 224, 224)
x = self.maxpool(x) # Output Shape = (12, 112, 112)
# Second Convolutional Layer
x = self.relu(self.conv2(x)) # Output Shape = (16, 112, 112)
x = self.maxpool(x) # Output Shape = (16, 56, 56)
# Third Convolutional Lay
x = self.relu(self.conv3(x)) # Output Shape = (20, 56, 56)
x = self.maxpool(x) # Output Shape = (20, 28, 28)
# Flattening Tensor
x = x.view(-1, 20*28*28)
# Fully Connected Layers
x = self.relu(self.fc1(x))
x = self.dropout(x)
x = self.relu(self.fc2(x))
x = self.dropout(x)
x = self.relu(self.fc3(x))
x = self.dropout(x)
x = self.relu(self.fc4(x))
x = self.fc5(x)
return x
# + [markdown] id="sI6uV2SQJk5t"
# ## Define Optimizer and Loss Function
#
# This code also checks if a GPU is available to train the network
# + id="-JlbZ_H2GJVE"
device = "cuda" if torch.cuda.is_available() else "cpu"
# Intializing model and having it use a GPU if available
model = CNN_Model()
model.to(device)
# Defining Optimizer
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
# Defining Loss Function; CE Loss because we have multiple categories
criterion = nn.CrossEntropyLoss()
# + [markdown] id="3p3_dAq7J0RM"
# ## Train the model
# + colab={"base_uri": "https://localhost:8080/"} id="Nn1WLqSdM_OA" outputId="008f724a-f2e0-4847-ac59-f8d0db1384fe"
def train(n_epochs, loaders, datasets, model, optimizer, criterion, device, save_path):
"""returns trained model"""
print("Starting Training...")
# Initialize tracker for minimum validation loss
valid_loss_min = np.Inf
losses = []
valid_losses = []
for epoch in range(1, n_epochs+1):
# Initialize variables to monitor training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for images, labels in loaders['train']:
# Move to GPU if available
images, labels = images.to(device), labels.to(device)
# Zero the parameter gradients
optimizer.zero_grad()
# Calculating the output
output = model(images)
# Caluculating the Loss
loss = criterion(output, labels)
# Calculating the gradients
loss.backward()
# Performing Gradient Descent Step
optimizer.step()
# Saving the training loss
train_loss += loss.data
######################
# validate the model #
######################
model.eval()
for images, labels in loaders['valid']:
# Move to GPU if available
images, labels = images.to(device), labels.to(device)
# Getting the output
output = model(images)
# Calculating the Loss
loss = criterion(output, labels)
# Saving the validation loss
valid_loss += loss.data
# Averaging the losses
train_loss /= len(datasets['train'])
valid_loss /= len(datasets['valid'])
# Appending the losses to plot afterwards
losses.append(train_loss.item())
valid_losses.append(valid_loss.item())
# Print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch,
train_loss,
valid_loss))
# Save the model if validation loss has decreased
if valid_loss < valid_loss_min:
print('Saving Model')
torch.save(model.state_dict(), save_path)
valid_loss_min = valid_loss
# return trained model, and saved losses
return model, np.array(losses), np.array(valid_losses)
num_epochs=10
model, losses, valid_losses = train(10, loaders, datasets, model, optimizer, criterion, device, 'saved_model.pt')
# + id="nVFWdOCI21em" colab={"base_uri": "https://localhost:8080/", "height": 573} outputId="544959c6-f4e1-4f1d-ca9e-048ff608987c"
def plot_losses(losses, title):
plt.plot(losses)
plt.xlabel("Epochs")
plt.ylabel("Cross Entropy Loss")
plt.title(title)
plt.show()
plot_losses(losses, title='Training Loss')
plot_losses(valid_losses, title='Validation Loss')
# + [markdown] id="P--ORTZyJ5Y2"
# ## Test the Model
# + colab={"base_uri": "https://localhost:8080/"} id="G7ky0btzovR5" outputId="c6a3e858-4464-43c6-af7a-7aa02d97090e"
def getPredsFromLogits(logits):
# Using softmax to get an array that sums to 1, and then getting the index with the highest value
return torch.nn.functional.softmax(logits, dim=1).argmax(dim=1)
def test(loaders, model, criterion, device):
# monitor test loss and accuracy
test_loss = 0.0
correct = 0
total = 0
model.eval()
for images, labels in loaders['test']:
# move to GPU if available
images, labels = images.to(device), labels.to(device)
# forward pass: compute predicted outputs by passing inputs to the model
output = model(images)
# calculate the loss
loss = criterion(output, labels)
# update average test loss
test_loss += loss.data
# convert output probabilities to predicted class
pred = getPredsFromLogits(output)
# compare predictions to true label
correct += pred.eq(labels).sum().item()
total += pred.shape[0]
test_loss /= total
print('Test Loss: {:.6f}\n'.format(test_loss))
print('\nTest Accuracy: %2d%% (%2d/%2d)' % (100. * correct / total, correct, total))
test(loaders, model, criterion, device)
# + [markdown] id="g4p3unN93-Qx"
# ## How to load a saved model
#
# It is useful to save our trained model (i.e., like the weights it learned during training) and then reuse it later to prevent having to retrain it, which can take a very long time. This is really important when you want to work on your own projects!
# + id="hzveqgMbtp2q" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="a697a054-53f1-40c0-c1db-7a073d5b4708"
# First create a new instance of the model class
saved_model = CNN_Model()
# Second, load state dict in the file that was saved then it should work as normal!
saved_model.load_state_dict(torch.load('saved_model.pt'))
# It will be downloaded to your computer as an example for you to save yours later
files.download('saved_model.pt')
# + [markdown] id="Ke9WI6ZvJ-d4"
# ## Summary
#
# Our model here was able to get a test accuracy of 79%, but it probably can be improved on. What do you think can be changed to improve the test results? You can perhaps add more layers, change the learning rate or optimizer. Something I noticed is that may have perhaps overfitted on the images of healthy lungs that it didn't perform well on lungs that were infected. This provides an area of open creativity for you to explore to better this model.
#
# I invite you to use this model as a starting point for your own CNN medical diagnosis model. See if you can updated an get even better results! Feel free to use it on another medical imaging dataset that interest you.
#
# Here are some links to more datasets to consider:
#
# 1. [Stanford's Center for Artificial Intelligence in Medicine and Imaging](https://aimi.stanford.edu/research/public-datasets) (Highly-Recommend this one)
# 2. [Open-Access Medical Image Repositories](https://www.aylward.org/notes/open-access-medical-image-repositories)
# + id="LXN-IIC5MOs8"
| machine_learning/lesson 3 - Neural Networks/Intro_to_CNNs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Import libraries
# +
#----------------------------Reproducible----------------------------------------------------------------------------------------
import numpy as np
import tensorflow as tf
import random as rn
import os
seed=0
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
rn.seed(seed)
#session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
session_conf =tf.compat.v1.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
from keras import backend as K
#tf.set_random_seed(seed)
tf.compat.v1.set_random_seed(seed)
#sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
sess = tf.compat.v1.Session(graph=tf.compat.v1.get_default_graph(), config=session_conf)
K.set_session(sess)
#----------------------------Reproducible----------------------------------------------------------------------------------------
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
#--------------------------------------------------------------------------------------------------------------------------------
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# %matplotlib inline
matplotlib.style.use('ggplot')
import random
import scipy.sparse as sparse
import scipy.io
from keras.utils import to_categorical
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from skfeature.function.similarity_based import lap_score
from skfeature.utility import construct_W
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LinearRegression
import time
import pandas as pd
# +
def mse_check(train, val):
LR = LinearRegression(n_jobs = -1)
LR.fit(train[0], train[1])
MSELR = ((LR.predict(val[0]) - val[1]) ** 2).mean()
return MSELR
def next_batch(samples, labels, num):
# Return a total of `num` random samples and labels.
idx = np.random.choice(len(samples), num)
return samples[idx], labels[idx]
def standard_single_hidden_layer_autoencoder(X, units, O):
reg_alpha = 1e-3
D = X.shape[1]
weights = tf.get_variable("weights", [D, units])
biases = tf.get_variable("biases", [units])
X = tf.matmul(X, weights) + biases
X = tf.layers.dense(X, O, kernel_regularizer = tf.contrib.layers.l2_regularizer(reg_alpha))
return X, weights
def aefs_subset_selector(train, K, epoch_num=1000, alpha=0.1):
D = train[0].shape[1]
O = train[1].shape[1]
learning_rate = 0.001
tf.reset_default_graph()
X = tf.placeholder(tf.float32, (None, D))
TY = tf.placeholder(tf.float32, (None, O))
Y, weights = standard_single_hidden_layer_autoencoder(X, K, O)
loss = tf.reduce_mean(tf.square(TY - Y)) + alpha * tf.reduce_sum(tf.sqrt(tf.reduce_sum(tf.square(weights), axis=1)), axis=0) + tf.losses.get_total_loss()
train_op = tf.train.AdamOptimizer(learning_rate).minimize(loss)
init = tf.global_variables_initializer()
batch_size = 8
batch_per_epoch = train[0].shape[0] // batch_size
costs = []
session_config = tf.ConfigProto()
session_config.gpu_options.allow_growth = False
with tf.Session(config = session_config) as sess:
sess.run(init)
for ep in range(epoch_num):
cost = 0
for batch_n in range(batch_per_epoch):
imgs, yimgs = next_batch(train[0], train[1], batch_size)
_, c, p = sess.run([train_op, loss, weights], feed_dict = {X: imgs, TY: yimgs})
cost += c / batch_per_epoch
costs.append(cost)
return list(np.argmax(np.abs(p), axis=0)), costs
def AEFS(train, test, K, debug = True):
x_train, x_val, y_train, y_val = train_test_split(train[0], train[1], test_size = 0.1)
print("y_train.shape",y_train.shape)
bindices = []
bmse = 1e100
for alpha in [1e-3, 1e-1, 1e1, 1e3]:
print("alpha",alpha)
indices, _ = aefs_subset_selector(train, K)
mse = mse_check((train[0][:, indices], train[1]), (x_val[:, indices], y_val))
if bmse > mse:
bmse = mse
bindices = indices
if debug:
print(bindices, bmse)
return train[0][:, bindices], test[0][:, bindices]
# -
#--------------------------------------------------------------------------------------------------------------------------------
def ETree(p_train_feature,p_train_label,p_test_feature,p_test_label,p_seed):
clf = ExtraTreesClassifier(n_estimators=50, random_state=p_seed)
# Training
clf.fit(p_train_feature, p_train_label)
# Training accuracy
print('Training accuracy:',clf.score(p_train_feature, np.array(p_train_label)))
print('Training accuracy:',accuracy_score(np.array(p_train_label),clf.predict(p_train_feature)))
#print('Training accuracy:',np.sum(clf.predict(p_train_feature)==np.array(p_train_label))/p_train_label.shape[0])
# Testing accuracy
print('Testing accuracy:',clf.score(p_test_feature, np.array(p_test_label)))
print('Testing accuracy:',accuracy_score(np.array(p_test_label),clf.predict(p_test_feature)))
#print('Testing accuracy:',np.sum(clf.predict(p_test_feature)==np.array(p_test_label))/p_test_label.shape[0])
#--------------------------------------------------------------------------------------------------------------------------------
def write_to_csv(p_data,p_path):
dataframe = pd.DataFrame(p_data)
dataframe.to_csv(p_path, mode='a',header=False,index=False,sep=',')
# # 2. Loading data
# +
data_path="./Dataset/GLIOMA.mat"
Data = scipy.io.loadmat(data_path)
data_arr=Data['X']
label_arr=Data['Y'][:, 0]-1
Data=MinMaxScaler(feature_range=(0,1)).fit_transform(data_arr)
C_train_x,C_test_x,C_train_y,C_test_y= train_test_split(Data,label_arr,test_size=0.2,random_state=seed)
print('Shape of C_train_x: ' + str(C_train_x.shape))
print('Shape of C_train_y: ' + str(C_train_y.shape))
print('Shape of C_test_x: ' + str(C_test_x.shape))
print('Shape of C_test_y: ' + str(C_test_y.shape))
# -
key_feture_number=64
# # 3. Model
# +
train=(C_train_x,C_train_x)
test=(C_test_x,C_test_x)
start = time.clock()
C_train_selected_x, C_test_selected_x = AEFS((train[0], train[0]), (test[0], test[0]), key_feture_number)
time_cost=time.clock() - start
write_to_csv(np.array([time_cost]),"./log/AEFS_time"+str(key_feture_number)+".csv")
# -
# # 4. Classifying
# ### Extra Trees
# +
train_feature=C_train_x
train_label=C_train_y
test_feature=C_test_x
test_label=C_test_y
print('Shape of train_feature: ' + str(train_feature.shape))
print('Shape of train_label: ' + str(train_label.shape))
print('Shape of test_feature: ' + str(test_feature.shape))
print('Shape of test_label: ' + str(test_label.shape))
p_seed=seed
ETree(train_feature,train_label,test_feature,test_label,p_seed)
# +
train_feature=C_train_selected_x
train_label=C_train_y
test_feature=C_test_selected_x
test_label=C_test_y
print('Shape of train_feature: ' + str(train_feature.shape))
print('Shape of train_label: ' + str(train_label.shape))
print('Shape of test_feature: ' + str(test_feature.shape))
print('Shape of test_label: ' + str(test_label.shape))
p_seed=seed
ETree(train_feature,train_label,test_feature,test_label,p_seed)
# -
# # 6. Reconstruction loss
# +
from sklearn.linear_model import LinearRegression
def mse_check(train, test):
LR = LinearRegression(n_jobs = -1)
LR.fit(train[0], train[1])
MSELR = ((LR.predict(test[0]) - test[1]) ** 2).mean()
return MSELR
# +
train_feature_tuple=(C_train_selected_x,C_train_x)
test_feature_tuple=(C_test_selected_x,C_test_x)
reconstruction_loss=mse_check(train_feature_tuple, test_feature_tuple)
print(reconstruction_loss)
# -
| Python/AbsoluteAndOtherAlgorithms/7GLIOMA/AEFS_64.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import gym
import matplotlib
import numpy as np
import sys
from collections import defaultdict
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.blackjack import BlackjackEnv
from lib import plotting
matplotlib.style.use('ggplot')
# -
env = BlackjackEnv()
a= np.ones(3)
np.random.choice(np.flatnonzero(a == a.max()))
def make_epsilon_greedy_policy(epsilon, nA):
"""
Creates an epsilon-greedy policy based on a given Q-function and epsilon.
Args:
Q: A dictionary that maps from state -> action-values.
Each value is a numpy array of length nA (see below)
epsilon: The probability to select a random action . float between 0 and 1.
nA: Number of actions in the environment.
Returns:
A function that takes the observation as an argument and returns
the probabilities for each action in the form of a numpy array of length nA.
"""
def policy_fn(Q,state):
# Implement this!
actions_at_St = Q[state]
probs = np.ones(actions_at_St.size, dtype=float) * epsilon / actions_at_St.size
best_action = np.random.choice(np.flatnonzero(actions_at_St == actions_at_St.max()))#break tie randomly
probs[best_action] += (1.0 - epsilon)
action = np.random.choice(np.arange(len(probs)), p=probs)#choose an action base on their probs
return action
return policy_fn
def mc_control_epsilon_greedy(env, num_episodes, discount_factor=1.0, epsilon=0.1):
"""
Monte Carlo Control using Epsilon-Greedy policies.
Finds an optimal epsilon-greedy policy.
Args:
env: OpenAI gym environment.
num_episodes: Number of episodes to sample.
discount_factor: Gamma discount factor.
epsilon: Chance the sample a random action. Float betwen 0 and 1.
Returns:
A tuple (Q, policy).
Q is a dictionary mapping state -> action values.
policy is a function that takes an observation as an argument and returns
action probabilities
"""
# Keeps track of sum and count of returns for each state
# to calculate an average. We could use an array to save all
# returns (like in the book) but that's memory inefficient.
returns_sum = defaultdict(float)
returns_count = defaultdict(float)
# The final action-value function.
# A nested dictionary that maps state -> (action -> action-value).
Q = defaultdict(lambda: np.zeros(env.action_space.n))
# The policy we're following
policy = make_epsilon_greedy_policy(epsilon, env.action_space.n)
# Implement this!
for n in range(num_episodes):
if (n+1) % 1000 == 0:
print("\rEpisode {}/{}.".format(n+1, num_episodes), end="")
sys.stdout.flush()
obs = env.reset()
G = 0.0
episode_rewards = []
episode_states = []
episode_actions = []
while True:#sample from env untill this episode finish
episode_states.insert(0,obs)
action = policy(Q,obs)
episode_actions.insert(0,action)
obs,reward,done,_ = env.step(action)
episode_rewards.insert(0,reward)
if done:
break
#see page 123 in RL book for more details
for idx,(state,action,reward) in enumerate(zip(episode_states,episode_actions,episode_rewards)):
G = discount_factor*G +reward
past_states = episode_states[idx+1:]
past_actions = episode_actions[idx+1:]
if state not in past_states or action not in past_actions:
returns_sum[state] += G
returns_count[state] += 1.0
Q[state][action] = returns_sum[state]/returns_count[state]
return Q, policy
Q, policy = mc_control_epsilon_greedy(env, num_episodes=500000, epsilon=0.1)
# For plotting: Create value function from action-value function
# by picking the best action at each state
V = defaultdict(float)
for state, actions in Q.items():
action_value = np.max(actions)
V[state] = action_value
plotting.plot_value_function(V, title="Optimal Value Function")
| MC/MC Control with Epsilon-Greedy Policies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
import pandas as pd
import numpy as np
import sklearn
import pickle
# run streamlit
# streamlit run d:/STUDY/AAMIIN/HACKTIV8/thunder-talk/streamlit_project/app.py
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
min(X[:,3])
max(X[:,3])
load_iris().feature_names
df = load_iris()
X = df.data
y = df.target
print(df.target_names)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42, test_size=0.3)
logreg = LogisticRegression(random_state=42, solver='liblinear')
logreg.fit(X_train, y_train)
y_train_pred = logreg.predict(X_train)
y_test_pred = logreg.predict(X_test)
from sklearn.metrics import classification_report, confusion_matrix, roc_auc_score
print(classification_report(y_train, y_train_pred))
print(classification_report(y_test, y_test_pred))
print(confusion_matrix(y_train, y_train_pred))
print(confusion_matrix(y_test, y_test_pred))
pickle.dump(logreg, open('classifier.pkl', 'wb'))
X_train[0]
X_train[0]
prediction_proba = logreg.predict_proba([[5.5, 2.4, 3.7, 1. ]])
class_name = ['setosa', 'versicolor', 'virginica']
prediction_proba_df = pd.DataFrame(prediction_proba, columns=class_name)
prediction_proba_df
| streamlit_project/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import zmq
print('python: %s' % sys.executable)
print(sys.version)
print('pyzmq-%s' % zmq.__version__)
print('libzmq-%s' % zmq.zmq_version())
print('Draft API available: %s' % zmq.DRAFT_API)
# +
import zmq
context = zmq.Context()
# Socket to talk to server
print("Connecting to hello world server...")
socket = context.socket(zmq.REQ)
socket.connect("tcp://localhost:5555")
# Do 10 requests, waiting each time for a response
for request in range(10):
print("Sending request %s ..." % request)
socket.send(b"Hello")
# Get the reply.
message = socket.recv()
print("Received reply %s [ %s ]" % (request, message))
# -
from sagas.util.serializer import write_data, read_data
# An arbitrary collection of objects supported by pickle.
data = {
'a': [1, 2.0, 3, 4+6j],
'b': ("character string", b"byte string"),
'c': {None, True, False}
}
write_data('./out/data.pickle', data)
rs=read_data('./out/data.pickle')
print(rs)
| notebook/procs-zeromq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# For this lecture, we will study iterators and generators. These are special objects Python is famous for so we must know them well. We first give an overview of what these things are from an OOP perspective, and then provide a few examples to illustrate how to use them.
# Python generators and iterators are two closely related concepts. Python generators are a simple way of creating iterators. An iterator is a special type of object in Python, and a generator is a function that returns an iterator object which we can iterate over (one value at a time).
# We study generators first. Generators are essentially a special type of function. They are very similar to normal functions, whereas generators allow us to write a function that can send back a value and then later resume to pick up where it left off. A generator will allow us to generate a sequence of values over time. The main difference in syntax will be the use of a 'yield' statement instead of a 'return' statement.
#
# Therefore, in most aspects, a generator will appear very similar to a conventional function. The main difference is when a generator is compiled, they become an object that support something called an **iteration protocol**. That means when they are called in your code they don't actually return a value and then exit, the generators will automatically suspend and resume their execution and state around the last point of value generation. Put another way, while a 'return' statement terminates a function entirely, a 'yield' statement pauses the function saving all its states and later continues from there on successive calls. If a function contains at least one 'yield' statement (it may contain other 'yield' or 'return' statements), it becomes a generator function. From a processsing efficiency perspective, generators allow us to generate as we go along, instead of holding everything in memory.
#
# Let's create a generator and see how it works.
def gencube(n):
for num in range(n):
yield num**3 ## raising to the power of 3
print(type(gencube)) # function
print(type(gencube(10))) # generator
print(gencube)
print(gencube(10))
for x in gencube(10):
print(x)
# On a cursory look at the example above, the generator gencube() seems like a normal function. However, generators are best for calculating large sets of results (particularly in calculations that involve loops themselves) in cases where we don’t want to allocate the memory for all of the results at the same time. If we use a normal function, the code will look like below:
def normalcube(n):
normal_list=[]
for num in range(n):
normal_list.append(num**3)
return normal_list
print(normalcube(10))
# Notice that now by using a normal function, we have to store everything in a list called 'normal_list' within the normalcube() function. There is nothing wrong with this type of approach, but when n is large, the memory storage becomes a problem. This is where the generators come in handy.
#
# As another example, let's write a generator that creates a Fibonacci sequence.
def fibon(n):
a=1
b=1
for j in range(n):
yield a
temp=a # temporary variable
a=b
b=temp+b
for x in fibon(10):
print(x)
# Now let's discuss **iterators**, which are objects that can be iterated upon. Iterators are actually very commonplace in Python. They are often elegantly implemented within for loops, comprehensions, generators etc. but hidden in plain sight. Technically speaking, Python iterator object must implement two special methods, '\__iter\__()' and '\__next\__()', collectively called the iterator protocol. An object is called **iterable** if we can get an iterator from it. Most of built-in containers in Python like: list, tuple, string etc. are iterables. The iter() function (which in turn calls the '\__iter\__()' method) returns an iterator from them. However, note that iterables are not necessarily iterator objects.
# The key to fully understand generators and iterators is to understand the how to use the next() and iter() functions. The next() function retrieves the next item from the iterator, and if the iterator is exhausted, it returns default value (if provided). If the default parameter is omitted and iterator is exhausted, it raises 'StopIteration' error. Below is an example:
def simple_gen():
for x in range(3):
yield x
g=simple_gen()
print(type(g))
def recur_print(iterator):
try:
print(next(iterator))
except StopIteration:
print('The iterator is exhausted, no more printing!')
recur_print(g)
recur_print(g)
recur_print(g)
recur_print(g)
# The iter() function, on the other hand, returns an iterator. Below is example. You may have remember that strings are iterable, but this does not mean that the string itself is an iterator:
s='MJZ'
for letters in s:
print(letters)
# In other words, if you type next(s) from the above example, Python will raise an error.
#
# To solve this problem, we can use the iter() function. Here it basically can change the string to an iterator.
s_iter=iter(s)
print(next(s_iter))
print(next(s_iter))
print(next(s_iter))
print(type(s_iter))
| Lecture 10 Iterators and Generators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# ### Get All the necessary packages
# +
from collections import Counter
import joblib
import numpy as np
import pandas as pd
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.naive_bayes import ComplementNB
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.metrics import (
classification_report, confusion_matrix,
f1_score as calculate_f1_score, accuracy_score as calculate_accuracy_score
)
# -
# ### Get All the necessary utilities
## utilities
from utils import CleanTextTransformer, load_imdb_sentiment_analysis_dataset
# ### Load Data
(X_train, y_train), (X_test, y_test) = load_imdb_sentiment_analysis_dataset(imdb_data_path='aclImdb')
# ### Visualize dataset size
# +
keys, values, labels = [], [], []
count = Counter(y_train)
for key, value in count.items():
keys.append(key)
values.append(value)
labels.append("positive" if value else "negative")
print(count)
print()
barlist = plt.bar(keys, values)
plt.title("Frequency of Sentiments")
plt.xticks(keys, labels)
plt.ylabel('Number of Reviews')
plt.xlabel('Sentiment expressed in Reviews')
barlist[0].set_color('red')
barlist[1].set_color('green')
plt.show()
# -
# ## Using CountVectorizer
# ### Create pipeline
pipeNB = Pipeline([
("clean_text", CleanTextTransformer()),
('count', CountVectorizer(stop_words="english")),
('classifier', ComplementNB())
])
# ### Fit the model
pipeNB.fit(X_train, y_train)
# #### Save model instance
joblib.dump(pipeNB, "models/complement_naive_bayes_with_count_vectorizer.joblib")
# ### Evaluate model
# #### get the prediction (of unseen data)
y_pred = pipeNB.predict(X_test)
# #### evaluate fitted model
print("Classification Report")
print("===================================")
print(classification_report(y_test, y_pred))
print("Confusion Matrix")
print("===================================")
print(confusion_matrix(y_test, y_pred))
# #### perform cross validation
# +
accuracy, f1_score = [], []
skf = StratifiedKFold(n_splits=10, shuffle=True, random_state=100)
for train_index, test_index in tqdm(skf.split(X_train, y_train), total=10):
X_train_fold, X_test_fold = X_train[train_index], X_train[test_index]
y_train_fold, y_test_fold = y_train[train_index], y_train[test_index]
pipeNB.fit(X_train_fold, y_train_fold)
y_pred = pipeNB.predict(X_test_fold)
accuracy.append(calculate_accuracy_score(y_test_fold, y_pred))
f1_score.append(calculate_f1_score(y_test_fold, y_pred))
# make as array
f1_score = np.array(f1_score)
accuracy = np.array(accuracy)
print('\nModel Metrics ==> ')
print("================================================")
print(f'{"descr":5s} | {"accuracy":^10s} | {"f1_score":^10s}')
print("================================================")
print(f'{"Max":5s} | {accuracy.max():^10.2f} | {f1_score.max():^10.2f}')
print(f'{"Min":5s} | {accuracy.min():^10.2f} | {f1_score.min():^10.2f}')
print(f'{"Mean":5s} | {accuracy.mean():^10.2f} | {f1_score.mean():^10.2f}')
# -
# ## Using TfidfVectorizer
# ### Create pipeline
pipeNB = Pipeline([
("clean_text", CleanTextTransformer()),
('tfidf', TfidfVectorizer(stop_words="english")),
('classifier', ComplementNB())
])
# ### Fit the model
pipeNB.fit(X_train, y_train)
# #### Save model instance
joblib.dump(pipeNB, "models/complement_naive_bayes_with_tfidf_vectorizer.joblib")
# ### Evaluate model
# #### get the prediction (of unseen data)
y_pred = pipeNB.predict(X_test)
# #### evaluate fitted model
print("Classification Report")
print("===================================")
print(classification_report(y_test, y_pred))
print("Confusion Matrix")
print("===================================")
print(confusion_matrix(y_test, y_pred))
# #### perform cross validation
# +
accuracy, f1_score = [], []
skf = StratifiedKFold(n_splits=10, shuffle=True, random_state=100)
for train_index, test_index in tqdm(skf.split(X_train, y_train), total=10):
X_train_fold, X_test_fold = X_train[train_index], X_train[test_index]
y_train_fold, y_test_fold = y_train[train_index], y_train[test_index]
pipeNB.fit(X_train_fold, y_train_fold)
y_pred = pipeNB.predict(X_test_fold)
accuracy.append(calculate_accuracy_score(y_test_fold, y_pred))
f1_score.append(calculate_f1_score(y_test_fold, y_pred))
# make as array
f1_score = np.array(f1_score)
accuracy = np.array(accuracy)
print('\nModel Metrics ==> ')
print("================================================")
print(f'{"descr":5s} | {"accuracy":^10s} | {"f1_score":^10s}')
print("================================================")
print(f'{"Max":5s} | {accuracy.max():^10.2f} | {f1_score.max():^10.2f}')
print(f'{"Min":5s} | {accuracy.min():^10.2f} | {f1_score.min():^10.2f}')
print(f'{"Mean":5s} | {accuracy.mean():^10.2f} | {f1_score.mean():^10.2f}')
# -
# ## End of Notebook
| Sentiment-Analysis-with-ComplementNB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Interroger une API pour enrichir les données des bulletins communaux
# ## Imports
import json
import requests
# ## Utiliser World Geo Data pour obtenir toutes les régions administratives en Belgique et récupérer l'identifiant de la région de Bruxelles.
# https://rapidapi.com/natkapral/api/world-geo-data/
# +
url = 'https://world-geo-data.p.rapidapi.com/countries/BE/adm-divisions'
headers = {
'x-rapidapi-host': "world-geo-data.p.rapidapi.com",
'x-rapidapi-key': "4df3692d45msh4addc1a7e6190f0p1f3544jsn142deaadd4a9"
}
response = requests.get(url, headers=headers)
response = json.loads(response.content)
for region in response['divisions']:
if 'Brussels' in region['name']:
brussels_region_id = region['geonameid']
# -
# ## A partir de l'identifiant de la région de Bruxelles, faire une requête pour obtenir toutes les communes de cette région et leurs propriétés (nom et population)
# +
url = 'https://world-geo-data.p.rapidapi.com/adm-divisions/'+str(brussels_region_id)+'/cities'
response = requests.get(url, headers=headers)
response = json.loads(response.content)
for city in response['cities']:
print(f"{city['name']} - {city['population']} inhabitants")
# -
| module1/TP1/api_request.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import mpld3
mpld3.enable_notebook()
from pylab import rcParams
rcParams['figure.figsize'] = 10, 10
# +
import sys
import numpy as np
import random
import math
import tensorflow as tf
import matplotlib.pyplot as plt
sys.path.append("./../../Utils/")
# -
from readWikiData import get_wikipedia_data
# #### Get data
sentences, word2idx, idx2word, _ = get_wikipedia_data(n_files=10, n_vocab=1000, by_paragraph=True)
def get_wiki_data_skip_gram(sentences, word2idx, window_size=5):
training_data = []
vocab_size = len(word2idx)
for sentence in sentences:
if len(sentence) < window_size * 2 + 1:
continue
for i in range(len(sentence)):
left_context = sentence[max(i-window_size, 0): i]
right_context = sentence[i+1:window_size + i + 1]
centre = sentence[i]
if len(left_context + right_context) < (2*window_size):
len_left = len(left_context)
len_right = len(right_context)
if len_left < len_right:
right_context = sentence[i+1 : window_size + i + 1 + (len_right - len_left)]
else:
left_context = sentence[max(i-window_size - (len_left - len_right), 0): i]
temp = left_context + right_context
if len(temp) < window_size * 2:
print (sentence)
print (left_context)
print (right_context)
print (centre)
break
training_data.append((centre, tuple(temp)))
print (training_data[:10])
training_data = list(set(training_data))
idx2word = {v:k for k, v in word2idx.items()}
return len(word2idx), training_data, word2idx, idx2word
vocab_size, training_data, word2idx, idx2word = get_wiki_data_skip_gram(sentences, word2idx)
len(training_data)
training_data[:10]
# ##### Get batches
# +
bucket_list = []
def getNextBatchSkipGram(bi_grams_, window_size=5, batch_size=10000):
global bucket_list
docs_ids_to_select = list(set(bi_grams_) - set(bucket_list))
if len(docs_ids_to_select) < batch_size:
bucket_list = []
docs_ids_to_select = bi_grams_
# Initialize two variables
train_X = np.ndarray(shape=(batch_size, 1), dtype=np.int32)
train_label = np.ndarray(shape=(batch_size, window_size*2), dtype=np.int32)
# Get a random set of docs
random_docs = random.sample(docs_ids_to_select, batch_size)
bucket_list += random_docs
index = 0
# Iterate threw all the docs
for item in random_docs:
train_X[index] = item[0]
train_label[index] = item[1]
index += 1
return train_X, train_label
# -
getNextBatchSkipGram(training_data)
# ##### Let's design the graph for skip gram model
def init_weight(Mi, Mo):
shape_sum = float(Mi + Mo)
return np.random.uniform(-np.sqrt(6/shape_sum),np.sqrt(6/shape_sum), [Mi, Mo])
embedding_size_w = 100
vocab_size = len(word2idx)
n_neg_samples = 20
learning_rate = 10e-5
epochs = 2
batch_size=10000
mu = 0.99
window_size = 5
# Define placeholders for training
train_X = tf.placeholder(tf.int32, shape=[batch_size, 1])
train_label = tf.placeholder(tf.int32, shape=[batch_size, None])
# Define matrix for doc_embedding and word_embedding
W1 = tf.Variable(init_weight(vocab_size, embedding_size_w), name="W1", dtype=tf.float32)
# Define weights for the output unit
W2 = tf.Variable(init_weight(vocab_size, embedding_size_w), name="W2", dtype=tf.float32)
biases = tf.Variable(tf.zeros(vocab_size))
print(train_X.get_shape(), train_label.get_shape(), W1.get_shape(), W2.get_shape())
embed = tf.nn.embedding_lookup(W1, train_X[0])
loss = tf.nn.sampled_softmax_loss(weights=W2, \
biases=biases, \
labels=train_label, \
inputs=embed, \
num_sampled=n_neg_samples, \
num_classes=vocab_size,
num_true=window_size*2)
loss = tf.reduce_mean(loss)
# +
#optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate, momentum=mu).minimize(loss)
#optimizer = tf.train.AdagradOptimizer(learning_rate).minimize(loss)
global_step = tf.Variable(0, trainable=False)
starter_learning_rate = 10e-5
learning_rate = tf.train.exponential_decay(starter_learning_rate, global_step,
1000, 0.96, staircase=True)
# Passing global_step to minimize() will increment it at each step.
optimizer = (
tf.train.MomentumOptimizer(learning_rate, momentum=mu).minimize(loss, global_step=global_step)
)
# +
saver = tf.train.Saver()
with tf.Session() as sess:
init = tf.global_variables_initializer()
sess.run(init)
average_loss = 0
for step in range(epochs):
epoch_error = 0.0
temp_X , temp_labels = getNextBatchSkipGram(window_size=5, bi_grams_=training_data)
feed_dict = {train_X : temp_X, train_label : temp_labels}
op, l = sess.run([optimizer, loss],
feed_dict=feed_dict)
epoch_error += l
if step % 100 == 0:
print ("Error at epoch : ", step, " = ", epoch_error)
save_path = saver.save(sess, "./models/model_skipgram_model.ckpt")
print("Model saved in file: %s" % save_path)
# -
# ##### Embeddings
# +
W1_embedding = None
W2_embedding = None
with tf.Session() as sess:
saver = tf.train.Saver()
# Restore variables from disk.
saver.restore(sess, "./models/model_skipgram_model.ckpt")
print("Model restored.")
# Normalize word2vec
W1_embedding = W1.eval()
# Normalize word2vec
W2_embedding = W2.eval()
# -
word2vec = np.mean([W1_embedding, W2_embedding], axis=0)
# ##### Projection of embeddings using t-SNE
idx2word = {v:k for k, v in word2idx.items()}
from sklearn.manifold import TSNE
model = TSNE()
Z = model.fit_transform(word2vec)
plt.scatter(Z[:,0], Z[:,1])
for i in range(len(idx2word)):
try:
plt.annotate(s=idx2word[i].encode("utf8"), xy=(Z[i,0], Z[i,1]))
except:
print ("bad string:", idx2word[i])
plt.show()
| module2/SkipGram/word2vec_skip_gram_model.ipynb |
# <p align="center"><img src="https://raw.githubusercontent.com/satwikkansal/wtfpython/master/images/logo.png" alt=""></p>
# <h1 align="center">What the f*ck Python! 😱</h1>
# <p align="center">Exploring and understanding Python through surprising snippets.</p>
#
# Translations: [Chinese 中文](https://github.com/leisurelicht/wtfpython-cn) | [Add translation](https://github.com/satwikkansal/wtfpython/issues/new?title=Add%20translation%20for%20[LANGUAGE]&body=Expected%20time%20to%20finish:%20[X]%20weeks.%20I%27ll%20start%20working%20on%20it%20from%20[Y].)
#
# Other modes: [Interactive](https://colab.research.google.com/github/satwikkansal/wtfpython/blob/3.0/irrelevant/wtf.ipynb) | [CLI](https://pypi.python.org/pypi/wtfpython)
#
# Python, being a beautifully designed high-level and interpreter-based programming language, provides us with many features for the programmer's comfort. But sometimes, the outcomes of a Python snippet may not seem obvious at first sight.
#
# Here's a fun project attempting to explain what exactly is happening under the hood for some counter-intuitive snippets and lesser-known features in Python.
#
# While some of the examples you see below may not be WTFs in the truest sense, but they'll reveal some of the interesting parts of Python that you might be unaware of. I find it a nice way to learn the internals of a programming language, and I believe that you'll find it interesting too!
#
# If you're an experienced Python programmer, you can take it as a challenge to get most of them right in the first attempt. You may have already experienced some of them before, and I might be able to revive sweet old memories of yours! :sweat_smile:
#
# PS: If you're a returning reader, you can learn about the new modifications [here](https://github.com/satwikkansal/wtfpython/releases/).
#
# So, here we go...
#
#
# # Structure of the Examples
#
# All the examples are structured like below:
#
# > ### ▶ Some fancy Title
# >
# > ```py
# > # Set up the code.
# > # Preparation for the magic...
# > ```
# >
# > **Output (Python version(s)):**
# >
# > ```py
# > >>> triggering_statement
# > Some unexpected output
# > ```
# > (Optional): One line describing the unexpected output.
# >
# >
# > #### 💡 Explanation:
# >
# > * Brief explanation of what's happening and why is it happening.
# > ```py
# > # Set up code
# > # More examples for further clarification (if necessary)
# > ```
# > **Output (Python version(s)):**
# >
# > ```py
# > >>> trigger # some example that makes it easy to unveil the magic
# > # some justified output
# > ```
#
# **Note:** All the examples are tested on Python 3.5.2 interactive interpreter, and they should work for all the Python versions unless explicitly specified before the output.
#
# # Usage
#
# A nice way to get the most out of these examples, in my opinion, is to read them chronologically, and for every example:
# - Carefully read the initial code for setting up the example. If you're an experienced Python programmer, you'll successfully anticipate what's going to happen next most of the time.
# - Read the output snippets and,
# + Check if the outputs are the same as you'd expect.
# + Make sure if you know the exact reason behind the output being the way it is.
# - If the answer is no (which is perfectly okay), take a deep breath, and read the explanation (and if you still don't understand, shout out! and create an issue [here](https://github.com/satwikkansal/wtfPython)).
# - If yes, give a gentle pat on your back, and you may skip to the next example.
#
# PS: You can also read WTFPython at the command line using the [pypi package](https://pypi.python.org/pypi/wtfpython),
# ```sh
# $ pip install wtfpython -U
# $ wtfpython
# ```
# ---
#
# # 👀 Examples
#
#
#
# ## Hosted notebook instructions
#
# This is just an experimental attempt of browsing wtfpython through jupyter notebooks. Some examples are read-only because,
# - they either require a version of Python that's not supported in the hosted runtime.
# - or they can't be reproduced in the notebook envrinonment.
#
# The expected outputs are already present in collapsed cells following the code cells. The Google colab provides Python2 (2.7) and Python3 (3.6, default) runtimes. You can switch among these for Python2 specific examples. For examples specific to other minor versions, you can simply refer to collapsed outputs (it's not possible to control the minor version in hosted notebooks as of now). You can check the active version using
#
# ```py
# >>> import sys
# >>> sys.version
# # Prints out Python version here.
# ```
#
# That being said, most of the examples do work as expected. If you face any trouble, feel free to consult the original content on wtfpython and create an issue in the repo. Have fun!
#
# ---
#
# ### ▶ Strings can be tricky sometimes
# 1\.
#
#
a = "some_string"
id(a)
id("some" + "_" + "string") # Notice that both the ids are same.
#
# 2\.
#
a = "wtf"
b = "wtf"
a is b
a = "wtf!"
b = "wtf!"
a is b
#
# 3\.
#
#
a, b = "wtf!", "wtf!"
a is b # All versions except 3.7.x
a = "wtf!"; b = "wtf!"
a is b # This will print True or False depending on where you're invoking it (python shell / ipython / as a script)
#
#
# +
# This time in file some_file.py
a = "wtf!"
b = "wtf!"
print(a is b)
# prints True when the module is invoked!
# -
#
# 4\.
#
# **Output (< Python3.7 )**
#
#
'a' * 20 is 'aaaaaaaaaaaaaaaaaaaa'
'a' * 21 is 'aaaaaaaaaaaaaaaaaaaaa'
#
# Makes sense, right?
#
#
# #### 💡 Explanation:
# + The behavior in first and second snippets is due to a CPython optimization (called string interning) that tries to use existing immutable objects in some cases rather than creating a new object every time.
# + After being "interned," many variables may reference the same string object in memory (saving memory thereby).
# + In the snippets above, strings are implicitly interned. The decision of when to implicitly intern a string is implementation-dependent. There are some rules that can be used to guess if a string will be interned or not:
# * All length 0 and length 1 strings are interned.
# * Strings are interned at compile time (`'wtf'` will be interned but `''.join(['w', 't', 'f']` will not be interned)
# * Strings that are not composed of ASCII letters, digits or underscores, are not interned. This explains why `'wtf!'` was not interned due to `!`. CPython implementation of this rule can be found [here](https://github.com/python/cpython/blob/3.6/Objects/codeobject.c#L19)
# 
# + When `a` and `b` are set to `"wtf!"` in the same line, the Python interpreter creates a new object, then references the second variable at the same time. If you do it on separate lines, it doesn't "know" that there's already `wtf!` as an object (because `"wtf!"` is not implicitly interned as per the facts mentioned above). It's a compile-time optimization. This optimization doesn't apply to 3.7.x versions of CPython (check this [issue](https://github.com/satwikkansal/wtfpython/issues/100) for more discussion).
# + A compile unit in an interactive environment like IPython consists of a single statement, whereas it consists of the entire module in case of modules. `a, b = "wtf!", "wtf!"` is single statement, whereas `a = "wtf!"; b = "wtf!"` are two statements in a single line. This explains why the identities are different in `a = "wtf!"; b = "wtf!"`, and also explain why they are same when invoked in `some_file.py`
# + The abrupt change in the output of the fourth snippet is due to a [peephole optimization](https://en.wikipedia.org/wiki/Peephole_optimization) technique known as Constant folding. This means the expression `'a'*20` is replaced by `'aaaaaaaaaaaaaaaaaaaa'` during compilation to save a few clock cycles during runtime. Constant folding only occurs for strings having a length of less than 20. (Why? Imagine the size of `.pyc` file generated as a result of the expression `'a'*10**10`). [Here's](https://github.com/python/cpython/blob/3.6/Python/peephole.c#L288) the implementation source for the same.
# + Note: In Python 3.7, Constant folding was moved out from peephole optimizer to the new AST optimizer with some change in logic as well, so the third snippet doesn't work for Python 3.7. You can read more about the change [here](https://bugs.python.org/issue11549).
#
#
# ### ▶ Hash brownies
# 1\.
#
some_dict = {}
some_dict[5.5] = "JavaScript"
some_dict[5.0] = "Ruby"
some_dict[5] = "Python"
#
# **Output:**
#
#
some_dict[5.5]
some_dict[5.0] # "Python" destroyed the existence of "Ruby"?
some_dict[5]
complex_five = 5 + 0j
type(complex_five)
some_dict[complex_five]
#
# So, why is Python all over the place?
#
#
#
# #### 💡 Explanation
#
# * Python dictionaries check for equality and compare the hash value to determine if two keys are the same.
# * Immutable objects with the same value always have the same hash in Python.
#
5 == 5.0 == 5 + 0j
hash(5) == hash(5.0) == hash(5 + 0j)
# **Note:** Objects with different values may also have same hash (known as [hash collision](https://en.wikipedia.org/wiki/Collision_(computer_science))).
# * When the statement `some_dict[5] = "Python"` is executed, the existing value "Ruby" is overwritten with "Python" because Python recognizes `5` and `5.0` as the same keys of the dictionary `some_dict`.
# * This StackOverflow [answer](https://stackoverflow.com/a/32211042/4354153) explains the rationale behind it.
#
#
# ### ▶ Deep down, we're all the same.
#
class WTF:
pass
#
# **Output:**
#
WTF() == WTF() # two different instances can't be equal
WTF() is WTF() # identities are also different
hash(WTF()) == hash(WTF()) # hashes _should_ be different as well
id(WTF()) == id(WTF())
#
#
# #### 💡 Explanation:
#
# * When `id` was called, Python created a `WTF` class object and passed it to the `id` function. The `id` function takes its `id` (its memory location), and throws away the object. The object is destroyed.
# * When we do this twice in succession, Python allocates the same memory location to this second object as well. Since (in CPython) `id` uses the memory location as the object id, the id of the two objects is the same.
# * So, the object's id is unique only for the lifetime of the object. After the object is destroyed, or before it is created, something else can have the same id.
# * But why did the `is` operator evaluated to `False`? Let's see with this snippet.
#
class WTF(object):
def __init__(self): print("I")
def __del__(self): print("D")
#
# **Output:**
#
WTF() is WTF()
id(WTF()) == id(WTF())
# As you may observe, the order in which the objects are destroyed is what made all the difference here.
#
#
# ### ▶ Disorder within order *
#
# +
from collections import OrderedDict
dictionary = dict()
dictionary[1] = 'a'; dictionary[2] = 'b';
ordered_dict = OrderedDict()
ordered_dict[1] = 'a'; ordered_dict[2] = 'b';
another_ordered_dict = OrderedDict()
another_ordered_dict[2] = 'b'; another_ordered_dict[1] = 'a';
class DictWithHash(dict):
"""
A dict that also implements __hash__ magic.
"""
__hash__ = lambda self: 0
class OrderedDictWithHash(OrderedDict):
"""
An OrderedDict that also implements __hash__ magic.
"""
__hash__ = lambda self: 0
# -
#
# **Output**
#
dictionary == ordered_dict # If a == b
dictionary == another_ordered_dict # and b == c
ordered_dict == another_ordered_dict # the why isn't c == a ??
len({dictionary, ordered_dict, another_ordered_dict})
dictionary = DictWithHash()
dictionary[1] = 'a'; dictionary[2] = 'b';
ordered_dict = OrderedDictWithHash()
ordered_dict[1] = 'a'; ordered_dict[2] = 'b';
another_ordered_dict = OrderedDictWithHash()
another_ordered_dict[2] = 'b'; another_ordered_dict[1] = 'a';
len({dictionary, ordered_dict, another_ordered_dict})
len({ordered_dict, another_ordered_dict, dictionary}) # changing the order
#
# What is going on here?
#
#
# #### 💡 Explanation:
#
# - The reason why intransitive equality didn't hold among `dictionary`, `ordered_dict` and `another_ordered_dict` is because of the way `__eq__` method is implemented in `OrderedDict` class. From the [docs](https://docs.python.org/3/library/collections.html#ordereddict-objects)
#
# > Equality tests between OrderedDict objects are order-sensitive and are implemented as `list(od1.items())==list(od2.items())`. Equality tests between `OrderedDict` objects and other Mapping objects are order-insensitive like regular dictionaries.
# - The reason for this equality is behavior is that it allows `OrderedDict` objects to be directly substituted anywhere a regular dictionary is used.
# - Okay, so why did changing the order affect the lenght of the generated `set` object? The answer is the lack of intransitive equality only. Since sets are "unordered" collections of unique elements, the order in which elements are inserted shouldn't matter. But in this case, it does matter. Let's break it down a bit,
#
some_set = set()
some_set.add(dictionary) # these are the mapping objects from the snippets above
ordered_dict in some_set
some_set.add(ordered_dict)
len(some_set)
another_ordered_dict in some_set
some_set.add(another_ordered_dict)
len(some_set)
another_set = set()
another_set.add(ordered_dict)
another_ordered_dict in another_set
another_set.add(another_ordered_dict)
len(another_set)
dictionary in another_set
another_set.add(another_ordered_dict)
len(another_set)
# So the inconsistency is due to `another_ordered_dict in another_set` being `False` because `ordered_dict` was already present in `another_set` and as observed before, `ordered_dict == another_ordered_dict` is `False`.
#
#
# ### ▶ Keep trying... *
#
# +
def some_func():
try:
return 'from_try'
finally:
return 'from_finally'
def another_func():
for _ in range(3):
try:
continue
finally:
print("Finally!")
def one_more_func(): # A gotcha!
try:
for i in range(3):
try:
1 / i
except ZeroDivisionError:
# Let's throw it here and handle it outside for loop
raise ZeroDivisionError("A trivial divide by zero error")
finally:
print("Iteration", i)
break
except ZeroDivisionError as e:
print("Zero division error ocurred", e)
# -
#
# **Output:**
#
#
some_func()
another_func()
1 / 0
one_more_func()
#
#
# #### 💡 Explanation:
#
# - When a `return`, `break` or `continue` statement is executed in the `try` suite of a "try…finally" statement, the `finally` clause is also executed on the way out.
# - The return value of a function is determined by the last `return` statement executed. Since the `finally` clause always executes, a `return` statement executed in the `finally` clause will always be the last one executed.
# - The caveat here is, if the finally clause executes a `return` or `break` statement, the temporarily saved exception is discarded.
#
#
# ### ▶ For what?
#
some_string = "wtf"
some_dict = {}
for i, some_dict[i] in enumerate(some_string):
i = 10
#
# **Output:**
#
some_dict # An indexed dict appears.
#
#
# #### 💡 Explanation:
#
# * A `for` statement is defined in the [Python grammar](https://docs.python.org/3/reference/grammar.html) as:
# ```
# for_stmt: 'for' exprlist 'in' testlist ':' suite ['else' ':' suite]
# ```
# Where `exprlist` is the assignment target. This means that the equivalent of `{exprlist} = {next_value}` is **executed for each item** in the iterable.
# An interesting example that illustrates this:
#
for i in range(4):
print(i)
i = 10
#
# **Output:**
# ```
# 0
# 1
# 2
# 3
# ```
#
# Did you expect the loop to run just once?
#
# **💡 Explanation:**
#
# - The assignment statement `i = 10` never affects the iterations of the loop because of the way for loops work in Python. Before the beginning of every iteration, the next item provided by the iterator (`range(4)` this case) is unpacked and assigned the target list variables (`i` in this case).
#
# * The `enumerate(some_string)` function yields a new value `i` (a counter going up) and a character from the `some_string` in each iteration. It then sets the (just assigned) `i` key of the dictionary `some_dict` to that character. The unrolling of the loop can be simplified as:
#
i, some_dict[i] = (0, 'w')
i, some_dict[i] = (1, 't')
i, some_dict[i] = (2, 'f')
some_dict
#
#
# ### ▶ Evaluation time discrepancy
# 1\.
#
array = [1, 8, 15]
# A typical generator expression
gen = (x for x in array if array.count(x) > 0)
array = [2, 8, 22]
#
# **Output:**
#
#
print(list(gen)) # Where did the other values go?
#
# 2\.
#
#
# +
array_1 = [1,2,3,4]
gen_1 = (x for x in array_1)
array_1 = [1,2,3,4,5]
array_2 = [1,2,3,4]
gen_2 = (x for x in array_2)
array_2[:] = [1,2,3,4,5]
# -
#
# **Output:**
#
print(list(gen_1))
print(list(gen_2))
#
# 3\.
#
#
# +
array_3 = [1, 2, 3]
array_4 = [10, 20, 30]
gen = (i + j for i in array_3 for j in array_4)
array_3 = [4, 5, 6]
array_4 = [400, 500, 600]
# -
#
# **Output:**
#
print(list(gen))
#
#
# #### 💡 Explanation
#
# - In a [generator](https://wiki.python.org/moin/Generators) expression, the `in` clause is evaluated at declaration time, but the conditional clause is evaluated at runtime.
# - So before runtime, `array` is re-assigned to the list `[2, 8, 22]`, and since out of `1`, `8` and `15`, only the count of `8` is greater than `0`, the generator only yields `8`.
# - The differences in the output of `g1` and `g2` in the second part is due the way variables `array_1` and `array_2` are re-assigned values.
# - In the first case, `array_1` is binded to the new object `[1,2,3,4,5]` and since the `in` clause is evaluated at the declaration time it still refers to the old object `[1,2,3,4]` (which is not destroyed).
# - In the second case, the slice assignment to `array_2` updates the same old object `[1,2,3,4]` to `[1,2,3,4,5]`. Hence both the `g2` and `array_2` still have reference to the same object (which has now been updated to `[1,2,3,4,5]`).
# - Okay, going by the logic discussed so far, shouldn't be the value of `list(g)` in the third snippet be `[11, 21, 31, 12, 22, 32, 13, 23, 33]`? (because `array_3` and `array_4` are going to behave just like `array_1`). The reason why (only) `array_4` values got updated is explained in [PEP-289](https://www.python.org/dev/peps/pep-0289/#the-details)
#
# > Only the outermost for-expression is evaluated immediately, the other expressions are deferred until the generator is run.
#
#
# ### ▶ How not to use `is` operator
# The following is a very famous example present all over the internet.
#
# 1\.
#
#
a = 256
b = 256
a is b
a = 257
b = 257
a is b
#
# 2\.
#
#
a = []
b = []
a is b
a = tuple()
b = tuple()
a is b
#
# 3\.
# **Output**
#
#
a, b = 257, 257
a is b
#
# **Output (Python 3.7.x specifically)**
#
#
a, b = 257, 257
#
#
# #### 💡 Explanation:
#
# **The difference between `is` and `==`**
#
# * `is` operator checks if both the operands refer to the same object (i.e., it checks if the identity of the operands matches or not).
# * `==` operator compares the values of both the operands and checks if they are the same.
# * So `is` is for reference equality and `==` is for value equality. An example to clear things up,
#
class A: pass
A() is A() # These are two empty objects at two different memory locations.
#
# **`256` is an existing object but `257` isn't**
#
# When you start up python the numbers from `-5` to `256` will be allocated. These numbers are used a lot, so it makes sense just to have them ready.
#
# Quoting from https://docs.python.org/3/c-api/long.html
# > The current implementation keeps an array of integer objects for all integers between -5 and 256, when you create an int in that range you just get back a reference to the existing object. So it should be possible to change the value of 1. I suspect the behavior of Python, in this case, is undefined. :-)
#
#
id(256)
a = 256
b = 256
id(a)
id(b)
id(257)
x = 257
y = 257
id(x)
id(y)
#
# Here the interpreter isn't smart enough while executing `y = 257` to recognize that we've already created an integer of the value `257,` and so it goes on to create another object in the memory.
#
# Similar optimization applies to other **immutable** objects like empty tuples as well. Since lists are mutable, that's why `[] is []` will return `False` and `() is ()` will return `True`. This explains our second snippet. Let's move on to the third one,
#
# **Both `a` and `b` refer to the same object when initialized with same value in the same line.**
#
# **Output**
#
#
a, b = 257, 257
id(a)
id(b)
a = 257
b = 257
id(a)
id(b)
#
# * When a and b are set to `257` in the same line, the Python interpreter creates a new object, then references the second variable at the same time. If you do it on separate lines, it doesn't "know" that there's already `257` as an object.
#
# * It's a compiler optimization and specifically applies to the interactive environment. When you enter two lines in a live interpreter, they're compiled separately, therefore optimized separately. If you were to try this example in a `.py` file, you would not see the same behavior, because the file is compiled all at once. This optimization is not limited to integers, it works for other immutable data types like strings (check the "Strings are tricky example") and floats as well,
#
#
a, b = 257.0, 257.0
a is b
#
# * Why didn't this work for Python 3.7? The abstract reason is because such compiler optimizations are implementation specific (i.e. may change with version, OS, etc). I'm still figuring out what exact implementation change cause the issue, you can check out this [issue](https://github.com/satwikkansal/wtfpython/issues/100) for updates.
#
#
# ### ▶ `is not ...` is not `is (not ...)`
#
'something' is not None
'something' is (not None)
#
#
# #### 💡 Explanation
#
# - `is not` is a single binary operator, and has behavior different than using `is` and `not` separated.
# - `is not` evaluates to `False` if the variables on either side of the operator point to the same object and `True` otherwise.
#
#
# ### ▶ A tic-tac-toe where X wins in the first attempt!
#
# Let's initialize a row
row = [""] * 3 #row i['', '', '']
# Let's make a board
board = [row] * 3
#
# **Output:**
#
#
board
board[0]
board[0][0]
board[0][0] = "X"
board
#
# We didn't assign three `"X"`s, did we?
#
#
# #### 💡 Explanation:
#
# When we initialize `row` variable, this visualization explains what happens in the memory
#
# 
#
# And when the `board` is initialized by multiplying the `row`, this is what happens inside the memory (each of the elements `board[0]`, `board[1]` and `board[2]` is a reference to the same list referred by `row`)
#
# 
#
# We can avoid this scenario here by not using `row` variable to generate `board`. (Asked in [this](https://github.com/satwikkansal/wtfpython/issues/68) issue).
#
#
board = [['']*3 for _ in range(3)]
board[0][0] = "X"
board
#
#
# ### ▶ The sticky output function
# 1\.
#
#
# +
funcs = []
results = []
for x in range(7):
def some_func():
return x
funcs.append(some_func)
results.append(some_func()) # note the function call here
funcs_results = [func() for func in funcs]
# -
#
# **Output:**
#
#
results
funcs_results
# Even when the values of `x` were different in every iteration prior to appending `some_func` to `funcs`, all the functions return 6.
#
# 2\.
#
#
powers_of_x = [lambda x: x**i for i in range(10)]
[f(2) for f in powers_of_x]
#
#
# #### 💡 Explanation
#
# - When defining a function inside a loop that uses the loop variable in its body, the loop function's closure is bound to the variable, not its value. So all of the functions use the latest value assigned to the variable for computation.
#
# - To get the desired behavior you can pass in the loop variable as a named variable to the function. **Why this works?** Because this will define the variable again within the function's scope.
#
#
funcs = []
for x in range(7):
def some_func(x=x):
return x
funcs.append(some_func)
#
# **Output:**
#
funcs_results = [func() for func in funcs]
funcs_results
#
#
# ### ▶ The chicken-egg problem *
# 1\.
#
isinstance(3, int)
isinstance(type, object)
isinstance(object, type)
#
# So which is the "ultimate" base class? There's more to the confusion by the way,
#
# 2\.
#
#
class A: pass
isinstance(A, A)
isinstance(type, type)
isinstance(object, object)
#
# 3\.
#
#
issubclass(int, object)
issubclass(type, object)
issubclass(object, type)
#
#
#
# #### 💡 Explanation
#
# - `type` is a [metaclass](https://realpython.com/python-metaclasses/) in Python.
# - **Everything** is an `object` in Python, which includes classes as well as their objects (instances).
# - class `type` is the metaclass of class `object`, and every class (including `type`) has inherited directly or indirectly from `object`.
# - There is no real base class among `object` and `type`. The confusion in the above snippets is arising because we're thinking about these relationships (`issubclass` and `isinstance`) in terms of Python classes. The relationship between `object` and `type` can't be reproduced in pure python. To be more precise the following relationships can't be reproduced in pure Python,
# + class A is an instance of class B, and class B is an instance of class A.
# + class A is an instance of itself.
# - These relationships between `object` and `type` (both being instances of each other as well as themselves) exist in Python because of "cheating" at the implementation level.
#
#
# ### ▶ Subclass relationships
# **Output:**
#
from collections import Hashable
issubclass(list, object)
issubclass(object, Hashable)
issubclass(list, Hashable)
#
# The Subclass relationships were expected to be transitive, right? (i.e., if `A` is a subclass of `B`, and `B` is a subclass of `C`, the `A` _should_ a subclass of `C`)
#
#
# #### 💡 Explanation:
#
# * Subclass relationships are not necessarily transitive in Python. Anyone is allowed to define their own, arbitrary `__subclasscheck__` in a metaclass.
# * When `issubclass(cls, Hashable)` is called, it simply looks for non-Falsey "`__hash__`" method in `cls` or anything it inherits from.
# * Since `object` is hashable, but `list` is non-hashable, it breaks the transitivity relation.
# * More detailed explanation can be found [here](https://www.naftaliharris.com/blog/python-subclass-intransitivity/).
#
#
# ### ▶ All-true-ation *
#
all([True, True, True])
all([True, True, False])
all([])
all([[]])
all([[[]]])
#
# Why's this True-False alteration?
#
#
# #### 💡 Explanation:
#
# - The implementation of `all` function is equivalent to
#
# - ```py
# def all(iterable):
# for element in iterable:
# if not element:
# return False
# return True
# ```
#
# - `all([])` returns `True` since the iterable is empty.
# - `all([[]])` returns `False` because `not []` is `True` is equivalent to `not False` as the list inside the iterable is empty.
# - `all([[[]]])` and higher recursive variants are always `True` since `not [[]]`, `not [[[]]]`, and so on are equivalent to `not True`.
#
#
# ### ▶ The surprising comma
# **Output (< 3.6):**
#
#
# +
def f(x, y,):
print(x, y)
def g(x=4, y=5,):
print(x, y)
def h(x, **kwargs,):
# -
def h(*args,):
#
#
# #### 💡 Explanation:
#
# - Trailing comma is not always legal in formal parameters list of a Python function.
# - In Python, the argument list is defined partially with leading commas and partially with trailing commas. This conflict causes situations where a comma is trapped in the middle, and no rule accepts it.
# - **Note:** The trailing comma problem is [fixed in Python 3.6](https://bugs.python.org/issue9232). The remarks in [this](https://bugs.python.org/issue9232#msg248399) post discuss in brief different usages of trailing commas in Python.
#
#
# ### ▶ Strings and the backslashes
# **Output:**
#
print("\"")
print(r"\"")
print(r"\")
r'\'' == "\\'"
#
#
# #### 💡 Explanation
#
# - In a usual python string, the backslash is used to escape characters that may have a special meaning (like single-quote, double-quote, and the backslash itself).
#
'wt\"f'
# - In a raw string literal (as indicated by the prefix `r`), the backslashes pass themselves as is along with the behavior of escaping the following character.
#
r'wt\"f' == 'wt\\"f'
print(repr(r'wt\"f')
print("\n")
print(r"\\n")
# - This means when a parser encounters a backslash in a raw string, it expects another character following it. And in our case (`print(r"\")`), the backslash escaped the trailing quote, leaving the parser without a terminating quote (hence the `SyntaxError`). That's why backslashes don't work at the end of a raw string.
#
#
# ### ▶ not knot!
#
x = True
y = False
#
# **Output:**
#
not x == y
x == not y
#
#
# #### 💡 Explanation:
#
# * Operator precedence affects how an expression is evaluated, and `==` operator has higher precedence than `not` operator in Python.
# * So `not x == y` is equivalent to `not (x == y)` which is equivalent to `not (True == False)` finally evaluating to `True`.
# * But `x == not y` raises a `SyntaxError` because it can be thought of being equivalent to `(x == not) y` and not `x == (not y)` which you might have expected at first sight.
# * The parser expected the `not` token to be a part of the `not in` operator (because both `==` and `not in` operators have the same precedence), but after not being able to find an `in` token following the `not` token, it raises a `SyntaxError`.
#
#
# ### ▶ Half triple-quoted strings
# **Output:**
#
print('wtfpython''')
print("wtfpython""")
# The following statements raise `SyntaxError`
# print('''wtfpython')
# print("""wtfpython")
#
#
# #### 💡 Explanation:
# + Python supports implicit [string literal concatenation](https://docs.python.org/2/reference/lexical_analysis.html#string-literal-concatenation), Example,
# ```
# >>> print("wtf" "python")
# wtfpython
# >>> print("wtf" "") # or "wtf"""
# wtf
# ```
# + `'''` and `"""` are also string delimiters in Python which causes a SyntaxError because the Python interpreter was expecting a terminating triple quote as delimiter while scanning the currently encountered triple quoted string literal.
#
#
# ### ▶ What's wrong with booleans?
# 1\.
#
#
# +
# A simple example to count the number of booleans and
# integers in an iterable of mixed data types.
mixed_list = [False, 1.0, "some_string", 3, True, [], False]
integers_found_so_far = 0
booleans_found_so_far = 0
for item in mixed_list:
if isinstance(item, int):
integers_found_so_far += 1
elif isinstance(item, bool):
booleans_found_so_far += 1
# -
#
# **Output:**
#
integers_found_so_far
booleans_found_so_far
#
#
# 2\.
#
some_bool = True
"wtf" * some_bool
some_bool = False
"wtf" * some_bool
#
# 3\.
#
#
def tell_truth():
True = False
if True == False:
print("I have lost faith in truth!")
#
# **Output (< 3.x):**
#
#
tell_truth()
#
#
#
#
# #### 💡 Explanation:
#
# * `bool` is a subclass of `int` in Python
#
#
issubclass(bool, int)
issubclass(int, bool)
#
# * And thus, `True` and `False` are instances of `int`
#
isinstance(True, int)
isinstance(False, int)
#
# * The integer value of `True` is `1` and that of `False` is `0`.
#
int(True)
int(False)
#
# * See this StackOverflow [answer](https://stackoverflow.com/a/8169049/4354153) for the rationale behind it.
#
# * Initially, Python used to have no `bool` type (people used 0 for false and non-zero value like 1 for true). `True`, `False`, and a `bool` type was added in 2.x versions, but, for backward compatibility, `True` and `False` couldn't be made constants. They just were built-in variables, and it was possible to reassign them
#
# * Python 3 was backward-incompatible, the issue was finally fixed, and thus the last snippet won't work with Python 3.x!
#
#
# ### ▶ Class attributes and instance attributes
# 1\.
#
# +
class A:
x = 1
class B(A):
pass
class C(A):
pass
# -
#
# **Output:**
#
A.x, B.x, C.x
B.x = 2
A.x, B.x, C.x
A.x = 3
A.x, B.x, C.x # C.x changed, but B.x didn't
a = A()
a.x, A.x
a.x += 1
a.x, A.x
#
# 2\.
#
class SomeClass:
some_var = 15
some_list = [5]
another_list = [5]
def __init__(self, x):
self.some_var = x + 1
self.some_list = self.some_list + [x]
self.another_list += [x]
#
# **Output:**
#
#
some_obj = SomeClass(420)
some_obj.some_list
some_obj.another_list
another_obj = SomeClass(111)
another_obj.some_list
another_obj.another_list
another_obj.another_list is SomeClass.another_list
another_obj.another_list is some_obj.another_list
#
#
# #### 💡 Explanation:
#
# * Class variables and variables in class instances are internally handled as dictionaries of a class object. If a variable name is not found in the dictionary of the current class, the parent classes are searched for it.
# * The `+=` operator modifies the mutable object in-place without creating a new object. So changing the attribute of one instance affects the other instances and the class attribute as well.
#
#
# ### ▶ Non-reflexive class method *
#
class SomeClass:
def instance_method(self):
pass
@classmethod
def class_method(cls):
pass
#
# **Output:**
#
#
SomeClass.instance_method is SomeClass.instance_method
SomeClass.class_method is SomeClass.class_method
id(SomeClass.class_method) == id(SomeClass.class_method)
#
#
# #### 💡 Explanation:
#
# - The reason `SomeClass.class_method is SomeClass.class_method` is `False` is due to the `@classmethod` decorator.
#
#
SomeClass.instance_method
SomeClass.class_method
#
# A new bound method every time `SomeClass.class_method` is accessed.
#
# - `id(SomeClass.class_method) == id(SomeClass.class_method)` returned `True` because the second allocation of memory for `class_method` happened at the same location of first deallocation (See Deep Down, we're all the same example for more detailed explanation).
#
#
# ### ▶ yielding None
#
# +
some_iterable = ('a', 'b')
def some_func(val):
return "something"
# -
#
# **Output (<= 3.7.x):**
#
#
[x for x in some_iterable]
[(yield x) for x in some_iterable]
list([(yield x) for x in some_iterable])
list((yield x) for x in some_iterable)
list(some_func((yield x)) for x in some_iterable)
#
#
# #### 💡 Explanation:
# - This is a bug in CPython's handling of `yield` in generators and comprehensions.
# - Source and explanation can be found here: https://stackoverflow.com/questions/32139885/yield-in-list-comprehensions-and-generator-expressions
# - Related bug report: http://bugs.python.org/issue10544
# - Python 3.8+ no longer allows `yield` inside list comprehension and will throw a `SyntaxError`.
#
#
# ### ▶ Yielding from... return! *
# 1\.
#
#
def some_func(x):
if x == 3:
return ["wtf"]
else:
yield from range(x)
#
# **Output (> 3.3):**
#
#
list(some_func(3))
#
# Where did the `"wtf"` go? Is it due to some special effect of `yield from`? Let's validate that,
#
# 2\.
#
#
def some_func(x):
if x == 3:
return ["wtf"]
else:
for i in range(x):
yield i
#
# **Output:**
#
#
list(some_func(3))
#
# The same result, this didn't work either.
#
#
# #### 💡 Explanation:
#
# + From Python 3.3 onwards, it became possible to use `return` statement with values inside generators (See [PEP380](https://www.python.org/dev/peps/pep-0380/)). The [official docs](https://www.python.org/dev/peps/pep-0380/#enhancements-to-stopiteration) say that,
#
# > "... `return expr` in a generator causes `StopIteration(expr)` to be raised upon exit from the generator."
#
# + In the case of `some_func(3)`, `StopIteration` is raised at the beginning because of `return` statement. The `StopIteration` exception is automatically caught inside the `list(...)` wrapper and the `for` loop. Therefore, the above two snippets result in an empty list.
#
# + To get `["wtf"]` from the generator `some_func` we need to catch the `StopIteration` exception,
#
#
try:
next(some_func(3))
except StopIteration as e:
some_string = e.value
#
#
some_string
#
#
# ### ▶ Nan-reflexivity *
# 1\.
#
#
a = float('inf')
b = float('nan')
c = float('-iNf') # These strings are case-insensitive
d = float('nan')
#
# **Output:**
#
#
a
b
c
float('some_other_string')
a == -c # inf==inf
None == None # None == None
b == d # but nan!=nan
50 / a
a / a
23 + b
#
# 2\.
#
#
x = float('nan')
y = x / x
y is y # identity holds
y == y # equality fails of y
[y] == [y] # but the equality succeeds for the list containing y
#
#
#
#
# #### 💡 Explanation:
#
# - `'inf'` and `'nan'` are special strings (case-insensitive), which, when explicitly typecast-ed to `float` type, are used to represent mathematical "infinity" and "not a number" respectively.
#
# - Since according to IEEE standards ` NaN != NaN`, obeying this rule breaks the reflexivity assumption of a collection element in Python i.e. if `x` is a part of a collection like `list`, the implementations like comparison are based on the assumption that `x == x`. Because of this assumption, the identity is compared first (since it's faster) while comparing two elements, and the values are compared only when the identities mismatch. The following snippet will make things clearer,
#
#
x = float('nan')
x == x, [x] == [x]
y = float('nan')
y == y, [y] == [y]
x == y, [x] == [y]
#
# Since the identities of `x` and `y` are different, the values are considered, which are also different; hence the comparison returns `False` this time.
#
# - Interesting read: [Reflexivity, and other pillars of civilization](https://bertrandmeyer.com/2010/02/06/reflexivity-and-other-pillars-of-civilization/)
#
#
# ### ▶ Mutating the immutable!
# This might seem trivial if you know how references work in Python.
#
#
some_tuple = ("A", "tuple", "with", "values")
another_tuple = ([1, 2], [3, 4], [5, 6])
#
# **Output:**
#
some_tuple[2] = "change this"
another_tuple[2].append(1000) #This throws no error
another_tuple
another_tuple[2] += [99, 999]
another_tuple
#
# But I thought tuples were immutable...
#
#
# #### 💡 Explanation:
#
# * Quoting from https://docs.python.org/2/reference/datamodel.html
#
# > Immutable sequences
# An object of an immutable sequence type cannot change once it is created. (If the object contains references to other objects, these other objects may be mutable and may be modified; however, the collection of objects directly referenced by an immutable object cannot change.)
#
# * `+=` operator changes the list in-place. The item assignment doesn't work, but when the exception occurs, the item has already been changed in place.
#
#
# ### ▶ The disappearing variable from outer scope
#
e = 7
try:
raise Exception()
except Exception as e:
pass
#
# **Output (Python 2.x):**
#
print(e)
#
# **Output (Python 3.x):**
#
print(e)
#
#
# #### 💡 Explanation:
#
# * Source: https://docs.python.org/3/reference/compound_stmts.html#except
#
# When an exception has been assigned using `as` target, it is cleared at the end of the `except` clause. This is as if
#
#
except E as N:
foo
#
# was translated into
#
#
except E as N:
try:
foo
finally:
del N
#
# This means the exception must be assigned to a different name to be able to refer to it after the except clause. Exceptions are cleared because, with the traceback attached to them, they form a reference cycle with the stack frame, keeping all locals in that frame alive until the next garbage collection occurs.
#
# * The clauses are not scoped in Python. Everything in the example is present in the same scope, and the variable `e` got removed due to the execution of the `except` clause. The same is not the case with functions that have their separate inner-scopes. The example below illustrates this:
#
#
def f(x):
del(x)
print(x)
x = 5
y = [5, 4, 3]
#
# **Output:**
#
>>>f(x)
UnboundLocalError: local variable 'x' referenced before assignment
>>>f(y)
UnboundLocalError: local variable 'x' referenced before assignment
x
y
#
# * In Python 2.x, the variable name `e` gets assigned to `Exception()` instance, so when you try to print, it prints nothing.
#
# **Output (Python 2.x):**
#
e
print e
#
#
# ### ▶ The mysterious key type conversion
#
# +
class SomeClass(str):
pass
some_dict = {'s': 42}
# -
#
# **Output:**
#
type(list(some_dict.keys())[0])
s = SomeClass('s')
some_dict[s] = 40
some_dict # expected: Two different keys-value pairs
type(list(some_dict.keys())[0])
#
#
# #### 💡 Explanation:
#
# * Both the object `s` and the string `"s"` hash to the same value because `SomeClass` inherits the `__hash__` method of `str` class.
# * `SomeClass("s") == "s"` evaluates to `True` because `SomeClass` also inherits `__eq__` method from `str` class.
# * Since both the objects hash to the same value and are equal, they are represented by the same key in the dictionary.
# * For the desired behavior, we can redefine the `__eq__` method in `SomeClass`
#
class SomeClass(str):
def __eq__(self, other):
return (
type(self) is SomeClass
and type(other) is SomeClass
and super().__eq__(other)
)
# When we define a custom __eq__, Python stops automatically inheriting the
# __hash__ method, so we need to define it as well
__hash__ = str.__hash__
some_dict = {'s':42}
#
# **Output:**
#
s = SomeClass('s')
some_dict[s] = 40
some_dict
keys = list(some_dict.keys())
type(keys[0]), type(keys[1])
#
#
# ### ▶ Let's see if you can guess this?
#
a, b = a[b] = {}, 5
#
# **Output:**
#
a
#
#
# #### 💡 Explanation:
#
# * According to [Python language reference](https://docs.python.org/2/reference/simple_stmts.html#assignment-statements), assignment statements have the form
# ```
# (target_list "=")+ (expression_list | yield_expression)
# ```
# and
#
# > An assignment statement evaluates the expression list (remember that this can be a single expression or a comma-separated list, the latter yielding a tuple) and assigns the single resulting object to each of the target lists, from left to right.
#
# * The `+` in `(target_list "=")+` means there can be **one or more** target lists. In this case, target lists are `a, b` and `a[b]` (note the expression list is exactly one, which in our case is `{}, 5`).
#
# * After the expression list is evaluated, its value is unpacked to the target lists from **left to right**. So, in our case, first the `{}, 5` tuple is unpacked to `a, b` and we now have `a = {}` and `b = 5`.
#
# * `a` is now assigned to `{}`, which is a mutable object.
#
# * The second target list is `a[b]` (you may expect this to throw an error because both `a` and `b` have not been defined in the statements before. But remember, we just assigned `a` to `{}` and `b` to `5`).
#
# * Now, we are setting the key `5` in the dictionary to the tuple `({}, 5)` creating a circular reference (the `{...}` in the output refers to the same object that `a` is already referencing). Another simpler example of circular reference could be
#
some_list = some_list[0] = [0]
some_list
some_list[0]
some_list is some_list[0]
some_list[0][0][0][0][0][0] == some_list
# Similar is the case in our example (`a[b][0]` is the same object as `a`)
#
# * So to sum it up, you can break the example down to
#
a, b = {}, 5
a[b] = a, b
# And the circular reference can be justified by the fact that `a[b][0]` is the same object as `a`
#
a[b][0] is a
#
#
# ### ▶ Modifying a dictionary while iterating over it
#
# +
x = {0: None}
for i in x:
del x[i]
x[i+1] = None
print(i)
# -
#
# **Output (Python 2.7- Python 3.5):**
#
# ```
# 0
# 1
# 2
# 3
# 4
# 5
# 6
# 7
# ```
#
# Yes, it runs for exactly **eight** times and stops.
#
#
# #### 💡 Explanation:
#
# * Iteration over a dictionary that you edit at the same time is not supported.
# * It runs eight times because that's the point at which the dictionary resizes to hold more keys (we have eight deletion entries, so a resize is needed). This is actually an implementation detail.
# * How deleted keys are handled and when the resize occurs might be different for different Python implementations.
# * So for Python versions other than Python 2.7 - Python 3.5, the count might be different from 8 (but whatever the count is, it's going to be the same every time you run it). You can find some discussion around this [here](https://github.com/satwikkansal/wtfpython/issues/53) or in [this](https://stackoverflow.com/questions/44763802/bug-in-python-dict) StackOverflow thread.
# * Python 3.8 onwards, you'll see `RuntimeError: dictionary keys changed during iteration` exception if you try to do this.
#
#
# ### ▶ The out of scope variable
#
# +
a = 1
def some_func():
return a
def another_func():
a += 1
return a
# -
#
# **Output:**
#
some_func()
another_func()
#
#
# #### 💡 Explanation:
# * When you make an assignment to a variable in scope, it becomes local to that scope. So `a` becomes local to the scope of `another_func`, but it has not been initialized previously in the same scope, which throws an error.
# * Read [this](http://sebastianraschka.com/Articles/2014_python_scope_and_namespaces.html) short but an awesome guide to learn more about how namespaces and scope resolution works in Python.
# * To modify the outer scope variable `a` in `another_func`, use `global` keyword.
#
def another_func()
global a
a += 1
return a
#
# **Output:**
#
another_func()
#
#
# ### ▶ Deleting a list item while iterating
#
# +
list_1 = [1, 2, 3, 4]
list_2 = [1, 2, 3, 4]
list_3 = [1, 2, 3, 4]
list_4 = [1, 2, 3, 4]
for idx, item in enumerate(list_1):
del item
for idx, item in enumerate(list_2):
list_2.remove(item)
for idx, item in enumerate(list_3[:]):
list_3.remove(item)
for idx, item in enumerate(list_4):
list_4.pop(idx)
# -
#
# **Output:**
#
list_1
list_2
list_3
list_4
#
# Can you guess why the output is `[2, 4]`?
#
#
# #### 💡 Explanation:
#
# * It's never a good idea to change the object you're iterating over. The correct way to do so is to iterate over a copy of the object instead, and `list_3[:]` does just that.
#
#
some_list = [1, 2, 3, 4]
id(some_list)
id(some_list[:]) # Notice that python creates new object for sliced list.
#
# **Difference between `del`, `remove`, and `pop`:**
# * `del var_name` just removes the binding of the `var_name` from the local or global namespace (That's why the `list_1` is unaffected).
# * `remove` removes the first matching value, not a specific index, raises `ValueError` if the value is not found.
# * `pop` removes the element at a specific index and returns it, raises `IndexError` if an invalid index is specified.
#
# **Why the output is `[2, 4]`?**
# - The list iteration is done index by index, and when we remove `1` from `list_2` or `list_4`, the contents of the lists are now `[2, 3, 4]`. The remaining elements are shifted down, i.e., `2` is at index 0, and `3` is at index 1. Since the next iteration is going to look at index 1 (which is the `3`), the `2` gets skipped entirely. A similar thing will happen with every alternate element in the list sequence.
#
# * Refer to this StackOverflow [thread](https://stackoverflow.com/questions/45946228/what-happens-when-you-try-to-delete-a-list-element-while-iterating-over-it) explaining the example
# * See also this nice StackOverflow [thread](https://stackoverflow.com/questions/45877614/how-to-change-all-the-dictionary-keys-in-a-for-loop-with-d-items) for a similar example related to dictionaries in Python.
#
#
# ### ▶ Lossy zip of iterators *
#
numbers = list(range(7))
numbers
first_three, remaining = numbers[:3], numbers[3:]
first_three, remaining
numbers_iter = iter(numbers)
list(zip(numbers_iter, first_three))
list(zip(numbers_iter, remaining))
# Where did element `3` go from the `numbers` list?
#
#
# #### 💡 Explanation:
#
# - From Python [docs](https://docs.python.org/3.3/library/functions.html#zip), here's an approximate implementation of zip function,
#
def zip(*iterables):
sentinel = object()
iterators = [iter(it) for it in iterables]
while iterators:
result = []
for it in iterators:
elem = next(it, sentinel)
if elem is sentinel: return
result.append(elem)
yield tuple(result)
# - So the function takes in arbitrary number of itreable objects, adds each of their items to the `result` list by calling the `next` function on them, and stops whenever any of the iterable is exhausted.
# - The caveat here is when any iterable is exhausted, the existing elements in the `result` list are discarded. That's what happened with `3` in the `numbers_iter`.
# - The correct way to do the above using `zip` would be,
#
numbers = list(range(7))
numbers_iter = iter(numbers)
list(zip(first_three, numbers_iter))
list(zip(remaining, numbers_iter))
# The first argument of zip should be the one with fewest elements.
#
#
# ### ▶ Loop variables leaking out!
# 1\.
#
for x in range(7):
if x == 6:
print(x, ': for x inside loop')
print(x, ': x in global')
#
# **Output:**
#
6 : for x inside loop
6 : x in global
#
# But `x` was never defined outside the scope of for loop...
#
# 2\.
#
# This time let's initialize x first
x = -1
for x in range(7):
if x == 6:
print(x, ': for x inside loop')
print(x, ': x in global')
#
# **Output:**
#
6 : for x inside loop
6 : x in global
#
# 3\.
#
# **Output (Python 2.x):**
#
x = 1
print([x for x in range(5)])
print(x)
#
# **Output (Python 3.x):**
#
x = 1
print([x for x in range(5)])
print(x)
#
#
# #### 💡 Explanation:
#
# - In Python, for-loops use the scope they exist in and leave their defined loop-variable behind. This also applies if we explicitly defined the for-loop variable in the global namespace before. In this case, it will rebind the existing variable.
#
# - The differences in the output of Python 2.x and Python 3.x interpreters for list comprehension example can be explained by following change documented in [What’s New In Python 3.0](https://docs.python.org/3/whatsnew/3.0.html) changelog:
#
# > "List comprehensions no longer support the syntactic form `[... for var in item1, item2, ...]`. Use `[... for var in (item1, item2, ...)]` instead. Also, note that list comprehensions have different semantics: they are closer to syntactic sugar for a generator expression inside a `list()` constructor, and in particular, the loop control variables are no longer leaked into the surrounding scope."
#
#
# ### ▶ Beware of default mutable arguments!
#
def some_func(default_arg=[]):
default_arg.append("some_string")
return default_arg
#
# **Output:**
#
some_func()
some_func()
some_func([])
some_func()
#
#
# #### 💡 Explanation:
#
# - The default mutable arguments of functions in Python aren't really initialized every time you call the function. Instead, the recently assigned value to them is used as the default value. When we explicitly passed `[]` to `some_func` as the argument, the default value of the `default_arg` variable was not used, so the function returned as expected.
#
#
def some_func(default_arg=[]):
default_arg.append("some_string")
return default_arg
#
# **Output:**
#
some_func.__defaults__ #This will show the default argument values for the function
some_func()
some_func.__defaults__
some_func()
some_func.__defaults__
some_func([])
some_func.__defaults__
#
# - A common practice to avoid bugs due to mutable arguments is to assign `None` as the default value and later check if any value is passed to the function corresponding to that argument. Example:
#
#
def some_func(default_arg=None):
if not default_arg:
default_arg = []
default_arg.append("some_string")
return default_arg
#
#
# ### ▶ Catching the Exceptions
#
# +
some_list = [1, 2, 3]
try:
# This should raise an ``IndexError``
print(some_list[4])
except IndexError, ValueError:
print("Caught!")
try:
# This should raise a ``ValueError``
some_list.remove(4)
except IndexError, ValueError:
print("Caught again!")
# -
#
# **Output (Python 2.x):**
#
# +
Caught!
ValueError: list.remove(x): x not in list
# -
#
# **Output (Python 3.x):**
#
File "<input>", line 3
except IndexError, ValueError:
^
SyntaxError: invalid syntax
#
#
# #### 💡 Explanation
#
# * To add multiple Exceptions to the except clause, you need to pass them as parenthesized tuple as the first argument. The second argument is an optional name, which when supplied will bind the Exception instance that has been raised. Example,
#
some_list = [1, 2, 3]
try:
# This should raise a ``ValueError``
some_list.remove(4)
except (IndexError, ValueError), e:
print("Caught again!")
print(e)
# **Output (Python 2.x):**
# ```
# Caught again!
# list.remove(x): x not in list
# ```
# **Output (Python 3.x):**
#
File "<input>", line 4
except (IndexError, ValueError), e:
^
IndentationError: unindent does not match any outer indentation level
#
# * Separating the exception from the variable with a comma is deprecated and does not work in Python 3; the correct way is to use `as`. Example,
#
some_list = [1, 2, 3]
try:
some_list.remove(4)
except (IndexError, ValueError) as e:
print("Caught again!")
print(e)
# **Output:**
# ```
# Caught again!
# list.remove(x): x not in list
# ```
#
#
# ### ▶ Same operands, different story!
# 1\.
#
a = [1, 2, 3, 4]
b = a
a = a + [5, 6, 7, 8]
#
# **Output:**
#
a
b
#
# 2\.
#
a = [1, 2, 3, 4]
b = a
a += [5, 6, 7, 8]
#
# **Output:**
#
a
b
#
#
# #### 💡 Explanation:
#
# * `a += b` doesn't always behave the same way as `a = a + b`. Classes *may* implement the *`op=`* operators differently, and lists do this.
#
# * The expression `a = a + [5,6,7,8]` generates a new list and sets `a`'s reference to that new list, leaving `b` unchanged.
#
# * The expression `a += [5,6,7,8]` is actually mapped to an "extend" function that operates on the list such that `a` and `b` still point to the same list that has been modified in-place.
#
#
# ### ▶ Be careful with chained operations
#
(False == False) in [False] # makes sense
False == (False in [False]) # makes sense
False == False in [False] # now what?
True is False == False
False is False is False
1 > 0 < 1
(1 > 0) < 1
1 > (0 < 1)
#
#
# #### 💡 Explanation:
#
# As per https://docs.python.org/2/reference/expressions.html#not-in
#
# > Formally, if a, b, c, ..., y, z are expressions and op1, op2, ..., opN are comparison operators, then a op1 b op2 c ... y opN z is equivalent to a op1 b and b op2 c and ... y opN z, except that each expression is evaluated at most once.
#
# While such behavior might seem silly to you in the above examples, it's fantastic with stuff like `a == b == c` and `0 <= x <= 100`.
#
# * `False is False is False` is equivalent to `(False is False) and (False is False)`
# * `True is False == False` is equivalent to `True is False and False == False` and since the first part of the statement (`True is False`) evaluates to `False`, the overall expression evaluates to `False`.
# * `1 > 0 < 1` is equivalent to `1 > 0 and 0 < 1` which evaluates to `True`.
# * The expression `(1 > 0) < 1` is equivalent to `True < 1` and
#
int(True)
True + 1 #not relevant for this example, but just for fun
# So, `1 < 1` evaluates to `False`
#
#
# ### ▶ Name resolution ignoring class scope
# 1\.
#
x = 5
class SomeClass:
x = 17
y = (x for i in range(10))
#
# **Output:**
#
list(SomeClass.y)[0]
#
# 2\.
#
x = 5
class SomeClass:
x = 17
y = [x for i in range(10)]
#
# **Output (Python 2.x):**
#
SomeClass.y[0]
#
# **Output (Python 3.x):**
#
SomeClass.y[0]
#
#
# #### 💡 Explanation
# - Scopes nested inside class definition ignore names bound at the class level.
# - A generator expression has its own scope.
# - Starting from Python 3.X, list comprehensions also have their own scope.
#
#
# ### ▶ Needles in a Haystack *
# I haven't met even a single experience Pythonist till date who has not come across one or more of the following scenarios,
#
# 1\.
#
#
x, y = (0, 1) if True else None, None
#
# **Output:**
#
#
x, y # expected (0, 1)
#
# 2\.
#
#
# +
t = ('one', 'two')
for i in t:
print(i)
t = ('one')
for i in t:
print(i)
t = ()
print(t)
# -
#
# **Output:**
#
#
one
two
o
n
e
tuple()
#
# 3\.
#
# ```
# ten_words_list = [
# "some",
# "very",
# "big",
# "list",
# "that"
# "consists",
# "of",
# "exactly",
# "ten",
# "words"
# ]
# ```
#
# **Output**
#
#
len(ten_words_list)
#
# 4\. Not asserting strongly enough
#
#
a = "python"
b = "javascript"
#
# **Output:**
#
#
# An assert statement with an assertion failure message.
assert(a == b, "Both languages are different")
#
# 5\.
#
#
# +
some_list = [1, 2, 3]
some_dict = {
"key_1": 1,
"key_2": 2,
"key_3": 3
}
some_list = some_list.append(4)
some_dict = some_dict.update({"key_4": 4})
# -
#
# **Output:**
#
#
print(some_list)
print(some_dict)
#
# 6\.
#
#
# +
def some_recursive_func(a):
if a[0] == 0:
return
a[0] -= 1
some_recursive_func(a)
return a
def similar_recursive_func(a):
if a == 0:
return a
a -= 1
similar_recursive_func(a)
return a
# -
#
# **Output:**
#
#
some_recursive_func([5, 0])
similar_recursive_func(5)
#
#
# #### 💡 Explanation:
#
# * For 1, the correct statement for expected behavior is `x, y = (0, 1) if True else (None, None)`.
#
# * For 2, the correct statement for expected behavior is `t = ('one',)` or `t = 'one',` (missing comma) otherwise the interpreter considers `t` to be a `str` and iterates over it character by character.
#
# * `()` is a special token and denotes empty `tuple`.
#
# * In 3, as you might have already figured out, there's a missing comma after 5th element (`"that"`) in the list. So by implicit string literal concatenation,
#
#
ten_words_list
#
# * No `AssertionError` was raised in 4th snippet because instead of asserting the individual expression `a == b`, we're asserting entire tuple. The following snippet will clear things up,
#
#
a = "python"
b = "javascript"
assert a == b
assert (a == b, "Values are not equal")
assert a == b, "Values are not equal"
#
# * As for the fifth snippet, most methods that modify the items of sequence/mapping objects like `list.append`, `dict.update`, `list.sort`, etc. modify the objects in-place and return `None`. The rationale behind this is to improve performance by avoiding making a copy of the object if the operation can be done in-place (Referred from [here](http://docs.python.org/2/faq/design.html#why-doesn-t-list-sort-return-the-sorted-list)).
#
# * Last one should be fairly obvious, passing mutable object (like `list` ) results in a call by reference, whereas an immutable object (like `int`) results in a call by value.
#
# * Being aware of these nitpicks can save you hours of debugging effort in the long run.
#
#
# ### ▶ Splitsies *
#
'a'.split()
'a'.split(' ')
len(''.split())
len(''.split(' '))
#
#
# #### 💡 Explanation:
#
# - It might appear at first that the default separator for split is a single space `' '`, but as per the [docs](https://docs.python.org/2.7/library/stdtypes.html#str.split)
# > If sep is not specified or is `None`, a different splitting algorithm is applied: runs of consecutive whitespace are regarded as a single separator, and the result will contain no empty strings at the start or end if the string has leading or trailing whitespace. Consequently, splitting an empty string or a string consisting of just whitespace with a None separator returns `[]`.
# > If sep is given, consecutive delimiters are not grouped together and are deemed to delimit empty strings (for example, `'1,,2'.split(',')` returns `['1', '', '2']`). Splitting an empty string with a specified separator returns `['']`.
# - Noticing how the leading and trailing whitespaces are handled in the following snippet will make things clear,
#
' a '.split(' ')
' a '.split()
''.split(' ')
#
#
# ### ▶ All sorted? *
#
x = 7, 8, 9
sorted(x) == x
sorted(x) == sorted(x)
y = reversed(x)
sorted(y) == sorted(y)
#
#
# #### 💡 Explanation:
#
# - The `sorted` method always returns a list, and comparing lists and tuples always returns `False` in Python.
#
# - ```py
# >>> [] == tuple()
# False
# >>> x = 7, 8, 9
# >>> type(x), type(sorted(x))
# (tuple, list)
# ```
#
# - Unlike `sorted`, the `reversed` method returns an iterator. Why? Because sorting requires the iterator to be either modified in-place or use an extra container (a list), whereas reversing can simply work by iterating from the last index to the first.
#
# - So during comparison `sorted(y) == sorted(y)`, the first call to `sorted()` will consume the iterator `y`, and the next call will just return an empty list.
#
#
x = 7, 8, 9
y = reversed(x)
sorted(y), sorted(y)
#
#
# ### ▶ Midnight time doesn't exist?
#
# +
from datetime import datetime
midnight = datetime(2018, 1, 1, 0, 0)
midnight_time = midnight.time()
noon = datetime(2018, 1, 1, 12, 0)
noon_time = noon.time()
if midnight_time:
print("Time at midnight is", midnight_time)
if noon_time:
print("Time at noon is", noon_time)
# -
#
# **Output (< 3.5):**
#
#
('Time at noon is', datetime.time(12, 0))
# The midnight time is not printed.
#
#
# #### 💡 Explanation:
#
# Before Python 3.5, the boolean value for `datetime.time` object was considered to be `False` if it represented midnight in UTC. It is error-prone when using the `if obj:` syntax to check if the `obj` is null or some equivalent of "empty."
#
#
# ### ▶ Okay Python, Can you make me fly?
# Well, here you go
#
#
import antigravity
#
# **Output:**
# Sshh... It's a super-secret.
#
#
# #### 💡 Explanation:
# + `antigravity` module is one of the few easter eggs released by Python developers.
# + `import antigravity` opens up a web browser pointing to the [classic XKCD comic](http://xkcd.com/353/) about Python.
# + Well, there's more to it. There's **another easter egg inside the easter egg**. If you look at the [code](https://github.com/python/cpython/blob/master/Lib/antigravity.py#L7-L17), there's a function defined that purports to implement the [XKCD's geohashing algorithm](https://xkcd.com/426/).
#
#
# ### ▶ `goto`, but why?
#
from goto import goto, label
for i in range(9):
for j in range(9):
for k in range(9):
print("I am trapped, please rescue!")
if k == 2:
goto .breakout # breaking out from a deeply nested loop
label .breakout
print("Freedom!")
#
# **Output (Python 2.3):**
#
I am trapped, please rescue!
I am trapped, please rescue!
Freedom!
#
#
# #### 💡 Explanation:
# - A working version of `goto` in Python was [announced](https://mail.python.org/pipermail/python-announce-list/2004-April/002982.html) as an April Fool's joke on 1st April 2004.
# - Current versions of Python do not have this module.
# - Although it works, but please don't use it. Here's the [reason](https://docs.python.org/3/faq/design.html#why-is-there-no-goto) to why `goto` is not present in Python.
#
#
# ### ▶ Brace yourself!
# If you are one of the people who doesn't like using whitespace in Python to denote scopes, you can use the C-style {} by importing,
#
#
from __future__ import braces
#
# **Output:**
#
File "some_file.py", line 1
from __future__ import braces
SyntaxError: not a chance
#
# Braces? No way! If you think that's disappointing, use Java. Okay, another surprising thing, can you find where's the `SyntaxError` raised in `__future__` module [code](https://github.com/python/cpython/blob/master/Lib/__future__.py)?
#
#
# #### 💡 Explanation:
# + The `__future__` module is normally used to provide features from future versions of Python. The "future" in this specific context is however, ironic.
# + This is an easter egg concerned with the community's feelings on this issue.
# + The code is actually present [here](https://github.com/python/cpython/blob/025eb98dc0c1dc27404df6c544fc2944e0fa9f3a/Python/future.c#L49) in `future.c` file.
# + When the CPython compiler encounters a [future statement](https://docs.python.org/3.3/reference/simple_stmts.html#future-statements), it first runs the appropriate code in `future.c` before treating it as a normal import statement.
#
#
# ### ▶ Let's meet Friendly Language Uncle For Life
# **Output (Python 3.x)**
#
from __future__ import barry_as_FLUFL
"Ruby" != "Python" # there's no doubt about it
"Ruby" <> "Python"
#
# There we go.
#
#
# #### 💡 Explanation:
# - This is relevant to [PEP-401](https://www.python.org/dev/peps/pep-0401/) released on April 1, 2009 (now you know, what it means).
# - Quoting from the PEP-401
#
# > Recognized that the != inequality operator in Python 3.0 was a horrible, finger-pain inducing mistake, the FLUFL reinstates the <> diamond operator as the sole spelling.
# - There were more things that Uncle Barry had to share in the PEP; you can read them [here](https://www.python.org/dev/peps/pep-0401/).
# - It works well in an interactive environment, but it will raise a `SyntaxError` when you run via python file (see this [issue](https://github.com/satwikkansal/wtfpython/issues/94)). However, you can wrap the statement inside an `eval` or `compile` to get it working,
#
from __future__ import barry_as_FLUFL
print(eval('"Ruby" <> "Python"'))
#
#
# ### ▶ Even Python understands that love is complicated
#
import this
#
# Wait, what's **this**? `this` is love :heart:
#
# **Output:**
# ```
# The Zen of Python, by <NAME>
#
# Beautiful is better than ugly.
# Explicit is better than implicit.
# Simple is better than complex.
# Complex is better than complicated.
# Flat is better than nested.
# Sparse is better than dense.
# Readability counts.
# Special cases aren't special enough to break the rules.
# Although practicality beats purity.
# Errors should never pass silently.
# Unless explicitly silenced.
# In the face of ambiguity, refuse the temptation to guess.
# There should be one-- and preferably only one --obvious way to do it.
# Although that way may not be obvious at first unless you're Dutch.
# Now is better than never.
# Although never is often better than *right* now.
# If the implementation is hard to explain, it's a bad idea.
# If the implementation is easy to explain, it may be a good idea.
# Namespaces are one honking great idea -- let's do more of those!
# ```
#
# It's the Zen of Python!
#
#
love = this
this is love
love is True
love is False
love is not True or False
love is not True or False; love is love # Love is complicated
#
#
# #### 💡 Explanation:
#
# * `this` module in Python is an easter egg for The Zen Of Python ([PEP 20](https://www.python.org/dev/peps/pep-0020)).
# * And if you think that's already interesting enough, check out the implementation of [this.py](https://hg.python.org/cpython/file/c3896275c0f6/Lib/this.py). Interestingly, **the code for the Zen violates itself** (and that's probably the only place where this happens).
# * Regarding the statement `love is not True or False; love is love`, ironic but it's self-explanatory (if not, please see the examples related to `is` and `is not` operators).
#
#
# ### ▶ Yes, it exists!
# **The `else` clause for loops.** One typical example might be:
#
#
def does_exists_num(l, to_find):
for num in l:
if num == to_find:
print("Exists!")
break
else:
print("Does not exist")
#
# **Output:**
#
some_list = [1, 2, 3, 4, 5]
does_exists_num(some_list, 4)
does_exists_num(some_list, -1)
#
# **The `else` clause in exception handling.** An example,
#
#
try:
pass
except:
print("Exception occurred!!!")
else:
print("Try block executed successfully...")
#
# **Output:**
#
Try block executed successfully...
#
#
# #### 💡 Explanation:
# - The `else` clause after a loop is executed only when there's no explicit `break` after all the iterations. You can think of it as a "nobreak" clause.
# - `else` clause after a try block is also called "completion clause" as reaching the `else` clause in a `try` statement means that the try block actually completed successfully.
#
#
# ### ▶ Ellipsis *
#
def some_func():
Ellipsis
#
# **Output**
#
some_func()
SomeRandomString
Ellipsis
#
#
# #### 💡 Explanation
# - In Python, `Ellipsis` is a globally available built-in object which is equivalent to `...`.
#
...
# - Eliipsis can be used for several purposes,
# + As a placeholder for code that hasn't been written yet (just like `pass` statement)
# + In slicing syntax to represent the full slices in remaining direction
#
import numpy as np
three_dimensional_array = np.arange(8).reshape(2, 2, 2)
# So our `three_dimensional_array` is an array of array of arrays. Let's say we want to print the second element (index `1`) of all the innermost arrays, we can use Ellipsis to bypass all the preceding dimensions
#
three_dimensional_array[:,:,1]
three_dimensional_array[..., 1] # using Ellipsis.
# Note: this will work for any number of dimensions. You can even select slice in first and last dimension and ignore the middle ones this way (`n_dimensional_array[firs_dim_slice, ..., last_dim_slice]`)
# + In [type hinting](https://docs.python.org/3/library/typing.html) to indicate only a part of the type (like `(Callable[..., int]` or `Tuple[str, ...]`))
# + You may also use Ellipsis as a default function argument (in the cases when you want to differentiate between the "no argument passed" and "None value passed" scenarios).
#
#
# ### ▶ Inpinity
# The spelling is intended. Please, don't submit a patch for this.
#
# **Output (Python 3.x):**
#
infinity = float('infinity')
hash(infinity)
hash(float('-inf'))
#
#
# #### 💡 Explanation:
# - Hash of infinity is 10⁵ x π.
# - Interestingly, the hash of `float('-inf')` is "-10⁵ x π" in Python 3, whereas "-10⁵ x e" in Python 2.
#
#
# ### ▶ Let's mangle
# 1\.
#
class Yo(object):
def __init__(self):
self.__honey = True
self.bro = True
#
# **Output:**
#
Yo().bro
Yo().__honey
Yo()._Yo__honey
#
# 2\.
#
class Yo(object):
def __init__(self):
# Let's try something symmetrical this time
self.__honey__ = True
self.bro = True
#
# **Output:**
#
Yo().bro
Yo()._Yo__honey__
#
# Why did `Yo()._Yo__honey` work?
#
# 3\.
#
#
# +
_A__variable = "Some value"
class A(object):
def some_func(self):
return __variable # not initialized anywhere yet
# -
#
# **Output:**
#
# +
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
AttributeError: 'A' object has no attribute '__variable'
A().some_func()
# -
#
#
#
# #### 💡 Explanation:
#
# * [Name Mangling](https://en.wikipedia.org/wiki/Name_mangling) is used to avoid naming collisions between different namespaces.
# * In Python, the interpreter modifies (mangles) the class member names starting with `__` (double underscore a.k.a "dunder") and not ending with more than one trailing underscore by adding `_NameOfTheClass` in front.
# * So, to access `__honey` attribute in the first snippet, we had to append `_Yo` to the front, which would prevent conflicts with the same name attribute defined in any other class.
# * But then why didn't it work in the second snippet? Because name mangling excludes the names ending with double underscores.
# * The third snippet was also a consequence of name mangling. The name `__variable` in the statement `return __variable` was mangled to `_A__variable`, which also happens to be the name of the variable we declared in the outer scope.
# * Also, if the mangled name is longer than 255 characters, truncation will happen.
#
#
# ### ▶ Skipping lines?
# **Output:**
#
value = 11
valuе = 32
value
#
# Wut?
#
# **Note:** The easiest way to reproduce this is to simply copy the statements from the above snippet and paste them into your file/shell.
#
#
# #### 💡 Explanation
#
# Some non-Western characters look identical to letters in the English alphabet but are considered distinct by the interpreter.
#
#
ord('е') # cyrillic 'e' (Ye)
ord('e') # latin 'e', as used in English and typed using standard keyboard
'е' == 'e'
value = 42 # latin e
valuе = 23 # cyrillic 'e', Python 2.x interpreter would raise a `SyntaxError` here
value
#
# The built-in `ord()` function returns a character's Unicode [code point](https://en.wikipedia.org/wiki/Code_point), and different code positions of Cyrillic 'e' and Latin 'e' justify the behavior of the above example.
#
#
# ### ▶ Teleportation
#
# +
# `pip install nump` first.
import numpy as np
def energy_send(x):
# Initializing a numpy array
np.array([float(x)])
def energy_receive():
# Return an empty numpy array
return np.empty((), dtype=np.float).tolist()
# -
#
# **Output:**
#
energy_send(123.456)
energy_receive()
#
# Where's the Nobel Prize?
#
#
# #### 💡 Explanation:
#
# * Notice that the numpy array created in the `energy_send` function is not returned, so that memory space is free to reallocate.
# * `numpy.empty()` returns the next free memory slot without reinitializing it. This memory spot just happens to be the same one that was just freed (usually, but not always).
#
#
# ### ▶ Well, something is fishy...
#
def square(x):
"""
A simple function to calculate the square of a number by addition.
"""
sum_so_far = 0
for counter in range(x):
sum_so_far = sum_so_far + x
return sum_so_far
#
# **Output (Python 2.x):**
#
#
square(10)
#
# Shouldn't that be 100?
#
# **Note:** If you're not able to reproduce this, try running the file [mixed_tabs_and_spaces.py](/mixed_tabs_and_spaces.py) via the shell.
#
#
# #### 💡 Explanation
#
# * **Don't mix tabs and spaces!** The character just preceding return is a "tab", and the code is indented by multiple of "4 spaces" elsewhere in the example.
# * This is how Python handles tabs:
#
# > First, tabs are replaced (from left to right) by one to eight spaces such that the total number of characters up to and including the replacement is a multiple of eight <...>
# * So the "tab" at the last line of `square` function is replaced with eight spaces, and it gets into the loop.
# * Python 3 is kind enough to throw an error for such cases automatically.
#
# **Output (Python 3.x):**
#
TabError: inconsistent use of tabs and spaces in indentation
#
#
# ### ▶ `+=` is faster
#
# using "+", three strings:
timeit.timeit("s1 = s1 + s2 + s3", setup="s1 = ' ' * 100000; s2 = ' ' * 100000; s3 = ' ' * 100000", number=100)
timeit.timeit("s1 += s2 + s3", setup="s1 = ' ' * 100000; s2 = ' ' * 100000; s3 = ' ' * 100000", number=100)
#
#
# #### 💡 Explanation:
# + `+=` is faster than `+` for concatenating more than two strings because the first string (example, `s1` for `s1 += s2 + s3`) is not destroyed while calculating the complete string.
#
#
# ### ▶ Let's make a giant string!
#
# +
def add_string_with_plus(iters):
s = ""
for i in range(iters):
s += "xyz"
assert len(s) == 3*iters
def add_bytes_with_plus(iters):
s = b""
for i in range(iters):
s += b"xyz"
assert len(s) == 3*iters
def add_string_with_format(iters):
fs = "{}"*iters
s = fs.format(*(["xyz"]*iters))
assert len(s) == 3*iters
def add_string_with_join(iters):
l = []
for i in range(iters):
l.append("xyz")
s = "".join(l)
assert len(s) == 3*iters
def convert_list_to_string(l, iters):
s = "".join(l)
assert len(s) == 3*iters
# -
#
# **Output:**
#
#
# +
# Executed in ipython shell using %timeit for better readability of results.
# You can also use the timeit module in normal python shell/scriptm=, example usage below
# timeit.timeit('add_string_with_plus(10000)', number=1000, globals=globals())
NUM_ITERS = 1000
# %timeit -n1000 add_string_with_plus(NUM_ITERS)
# -
# %timeit -n1000 add_bytes_with_plus(NUM_ITERS)
# %timeit -n1000 add_string_with_format(NUM_ITERS)
# %timeit -n1000 add_string_with_join(NUM_ITERS)
l = ["xyz"]*NUM_ITERS
# %timeit -n1000 convert_list_to_string(l, NUM_ITERS)
#
# Let's increase the number of iterations by a factor of 10.
#
#
NUM_ITERS = 10000
# %timeit -n1000 add_string_with_plus(NUM_ITERS) # Linear increase in execution time
# %timeit -n1000 add_bytes_with_plus(NUM_ITERS) # Quadratic increase
# %timeit -n1000 add_string_with_format(NUM_ITERS) # Linear increase
# %timeit -n1000 add_string_with_join(NUM_ITERS) # Linear increase
l = ["xyz"]*NUM_ITERS
# %timeit -n1000 convert_list_to_string(l, NUM_ITERS) # Linear increase
#
#
# #### 💡 Explanation
# - You can read more about [timeit](https://docs.python.org/3/library/timeit.html) or [%timeit](https://ipython.org/ipython-doc/dev/interactive/magics.html#magic-timeit) on these links. They are used to measure the execution time of code pieces.
# - Don't use `+` for generating long strings — In Python, `str` is immutable, so the left and right strings have to be copied into the new string for every pair of concatenations. If you concatenate four strings of length 10, you'll be copying (10+10) + ((10+10)+10) + (((10+10)+10)+10) = 90 characters instead of just 40 characters. Things get quadratically worse as the number and size of the string increases (justified with the execution times of `add_bytes_with_plus` function)
# - Therefore, it's advised to use `.format.` or `%` syntax (however, they are slightly slower than `+` for very short strings).
# - Or better, if already you've contents available in the form of an iterable object, then use `''.join(iterable_object)` which is much faster.
# - Unlike `add_bytes_with_plus` because of the `+=` optimizations discussed in the previous example, `add_string_with_plus` didn't show a quadratic increase in execution time. Had the statement been `s = s + "x" + "y" + "z"` instead of `s += "xyz"`, the increase would have been quadratic.
#
def add_string_with_plus(iters):
s = ""
for i in range(iters):
s = s + "x" + "y" + "z"
assert len(s) == 3*iters
# %timeit -n100 add_string_with_plus(1000)
# %timeit -n100 add_string_with_plus(10000) # Quadratic increase in execution time
# - So many ways to format and create a giant string are somewhat in contrast to the [Zen of Python](https://www.python.org/dev/peps/pep-0020/), according to which,
#
# > There should be one-- and preferably only one --obvious way to do it.
#
#
# ### ▶ Minor Ones *
# * `join()` is a string operation instead of list operation. (sort of counter-intuitive at first usage)
#
# **💡 Explanation:** If `join()` is a method on a string, then it can operate on any iterable (list, tuple, iterators). If it were a method on a list, it'd have to be implemented separately by every type. Also, it doesn't make much sense to put a string-specific method on a generic `list` object API.
#
# * Few weird looking but semantically correct statements:
# + `[] = ()` is a semantically correct statement (unpacking an empty `tuple` into an empty `list`)
# + `'a'[0][0][0][0][0]` is also a semantically correct statement as strings are [sequences](https://docs.python.org/3/glossary.html#term-sequence)(iterables supporting element access using integer indices) in Python.
# + `3 --0-- 5 == 8` and `--5 == 5` are both semantically correct statements and evaluate to `True`.
#
# * Given that `a` is a number, `++a` and `--a` are both valid Python statements but don't behave the same way as compared with similar statements in languages like C, C++, or Java.
# ```py
# >>> a = 5
# >>> a
# 5
# >>> ++a
# 5
# >>> --a
# 5
# ```
#
# **💡 Explanation:**
# + There is no `++` operator in Python grammar. It is actually two `+` operators.
# + `++a` parses as `+(+a)` which translates to `a`. Similarly, the output of the statement `--a` can be justified.
# + This StackOverflow [thread](https://stackoverflow.com/questions/3654830/why-are-there-no-and-operators-in-python) discusses the rationale behind the absence of increment and decrement operators in Python.
#
# * You must be aware of the Walrus operator in Python. But have you ever heard about *the space-invader operator*?
# ```py
# >>> a = 42
# >>> a -=- 1
# >>> a
# 43
# ```
# It is used as an alternative incrementation operator, together with another one
# ```py
# >>> a +=+ 1
# >>> a
# >>> 44
# ```
# **💡 Explanation:** This prank comes from [<NAME>'s tweet](https://twitter.com/raymondh/status/1131103570856632321?lang=en). The space invader operator is actually just a malformatted `a -= (-1)`. Which is equivalent to `a = a - (- 1)`. Similar for the `a += (+ 1)` case.
#
# * Python has an undocumented [converse implication](https://en.wikipedia.org/wiki/Converse_implication) operator.
#
# ```py
# >>> False ** False == True
# True
# >>> False ** True == False
# True
# >>> True ** False == True
# True
# >>> True ** True == True
# True
# ```
#
# **💡 Explanation:** If you replace `False` and `True` by 0 and 1 and do the maths, the truth table is equivalent to a converse implication operator. ([Source](https://github.com/cosmologicon/pywat/blob/master/explanation.md#the-undocumented-converse-implication-operator))
#
# * Since we are talking operators, there's also `@` operator for matrix multiplication (don't worry, this time it's for real).
#
# ```py
# >>> import numpy as np
# >>> np.array([2, 2, 2]) @ np.array([7, 8, 8])
# 46
# ```
#
# **💡 Explanation:** The `@` operator was added in Python 3.5 keeping the scientific community in mind. Any object can overload `__matmul__` magic method to define behavior for this operator.
#
# * From Python 3.8 onwards you can use a typical f-string syntax like `f'{some_var=}` for quick debugging. Example,
# ```py
# >>> some_string = "wtfpython"
# >>> f'{some_string=}'
# "string='wtfpython'"
# ```
#
# * Python uses 2 bytes for local variable storage in functions. In theory, this means that only 65536 variables can be defined in a function. However, python has a handy solution built in that can be used to store more than 2^16 variable names. The following code demonstrates what happens in the stack when more than 65536 local variables are defined (Warning: This code prints around 2^18 lines of text, so be prepared!):
#
# ```py
# import dis
# exec("""
# def f():
# """ + """
# """.join(["X" + str(x) + "=" + str(x) for x in range(65539)]))
#
# f()
#
# print(dis.dis(f))
# ```
#
# * Multiple Python threads won't run your *Python code* concurrently (yes, you heard it right!). It may seem intuitive to spawn several threads and let them execute your Python code concurrently, but, because of the [Global Interpreter Lock](https://wiki.python.org/moin/GlobalInterpreterLock) in Python, all you're doing is making your threads execute on the same core turn by turn. Python threads are good for IO-bound tasks, but to achieve actual parallelization in Python for CPU-bound tasks, you might want to use the Python [multiprocessing](https://docs.python.org/2/library/multiprocessing.html) module.
#
# * Sometimes, the `print` method might not print values immediately. For example,
#
# ```py
# # File some_file.py
# import time
#
# print("wtfpython", end="_")
# time.sleep(3)
# ```
#
# This will print the `wtfpython` after 10 seconds due to the `end` argument because the output buffer is flushed either after encountering `\n` or when the program finishes execution. We can force the buffer to flush by passing `flush=True` argument.
#
# * List slicing with out of the bounds indices throws no errors
# ```py
# >>> some_list = [1, 2, 3, 4, 5]
# >>> some_list[111:]
# []
# ```
#
# * Slicing an iterable not always creates a new object. For example,
# ```py
# >>> some_str = "wtfpython"
# >>> some_list = ['w', 't', 'f', 'p', 'y', 't', 'h', 'o', 'n']
# >>> some_list is some_list[:] # False expected because a new object is created.
# False
# >>> some_str is some_str[:] # True because strings are immutable, so making a new object is of not much use.
# True
# ```
#
# * `int('١٢٣٤٥٦٧٨٩')` returns `123456789` in Python 3. In Python, Decimal characters include digit characters, and all characters that can be used to form decimal-radix numbers, e.g. U+0660, ARABIC-INDIC DIGIT ZERO. Here's an [interesting story](http://chris.improbable.org/2014/8/25/adventures-in-unicode-digits/) related to this behavior of Python.
#
# * You can separate numeric literals with underscores (for better readability) from Python 3 onwards.
#
# ```py
# >>> six_million = 6_000_000
# >>> six_million
# 6000000
# >>> hex_address = 0xF00D_CAFE
# >>> hex_address
# 4027435774
# ```
#
# * `'abc'.count('') == 4`. Here's an approximate implementation of `count` method, which would make the things more clear
# ```py
# def count(s, sub):
# result = 0
# for i in range(len(s) + 1 - len(sub)):
# result += (s[i:i + len(sub)] == sub)
# return result
# ```
# The behavior is due to the matching of empty substring(`''`) with slices of length 0 in the original string.
#
# **That's all folks!**
#
#
# ### ▶ First things first! *
# For some reason, the Python 3.8's "Walrus" operator (`:=`) has become quite popular. Let's check it out,
#
# 1\.
#
#
# ```py
# # Python version 3.8+
#
# >>> a = "wtf_walrus"
# >>> a
# ```
# ```py
# 'wtf_walrus'
#
# ```
#
# ```py
# >>> a := "wtf_walrus"
# ```
# ```py
# File "<stdin>", line 1
# a := "wtf_walrus"
# ^
# SyntaxError: invalid syntax
#
# ```
#
# ```py
# >>> (a := "wtf_walrus") # This works though
# >>> a
# ```
# ```py
# 'wtf_walrus'
# ```
#
#
# 2 \.
#
#
# ```py
# # Python version 3.8+
#
# >>> a = 6, 9
# >>> a
# ```
# ```py
# (6, 9)
#
# ```
#
# ```py
# >>> (a := 6, 9)
# >>> a
# ```
# ```py
# 6
#
# ```
#
# ```py
# >>> a, b = 6, 9 # Typical unpacking
# >>> a, b
# ```
# ```py
# (6, 9)
# ```
#
# ```py
# >>> (a, b = 16, 19) # Oops
# ```
# ```py
# File "<stdin>", line 1
# (a, b = 6, 9)
# ^
# SyntaxError: invalid syntax
#
# ```
#
# ```py
# >>> (a, b := 16, 19) # This prints out a weird 3-tuple
# ```
# ```py
# (6, 16, 19)
#
# ```
#
# ```py
# >>> a # a is still unchanged?
# ```
# ```py
# 6
#
# ```
#
# ```py
# >>> b
# ```
# ```py
# 16
# ```
#
#
#
#
#
# #### 💡 Explanation
#
# **Quick walrus operator refresher**
#
# The Walrus operator (`:=`) was introduced in Python 3.8, it can be useful in situations where you'd want to assign values to variables within an expression.
#
#
# ```py
# def some_func():
# # Assume some expensive computation here
# # time.sleep(1000)
# return 5
#
# # So instead of,
# if some_func():
# print(some_func()) # Which is bad practice since computation is happening twice
#
# # or
# a = some_func()
# if a:
# print(a)
#
# # Now you can concisely write
# if a := some_func():
# print(a)
# ```
# ```py
# ```
#
#
# **Output (> 3.8):**
#
#
# ```py
# 5
# 5
# 5
# ```
# ```py
# ```
#
#
# This saved one line of code, and implicitly prevented invoking `some_func` twice.
#
# - Unparenthesized "assignment expression" (use of walrus operator), is restricted at the top level, hence the `SyntaxError` in the `a := "wtf_walrus"` statement of the first snippet. Parenthesizing it worked as expected and assigned `a`.
#
# - As usual, parenthesizing of an expression containing `=` operator is not allowed. Hence the syntax error in `(a, b = 6, 9)`.
#
# - The syntax of the Walrus operator is of the form `NAME: expr`, where `NAME` is a valid identifier, and `expr` is a valid expression. Hence, iterable packing and unpacking are not supported which means,
#
# - `(a := 6, 9)` is equivalent to `((a := 6), 9)` and ultimately `(a, 9) ` (where `a`'s value is 6')
#
#
# ```py
# >>> (a := 6, 9) == ((a := 6), 9)
# ```
# ```py
# True
# ```
#
# ```py
# >>> x = (a := 696, 9)
# >>> x
# ```
# ```py
# (696, 9)
# ```
#
# ```py
# >>> x[0] is a # Both reference same memory location
# ```
# ```py
# True
# ```
#
#
# - Similarly, `(a, b := 16, 19)` is equivalent to `(a, (b := 16), 19)` which is nothing but a 3-tuple.
#
#
# ### ▶ Stubborn `del` operation
#
# ```py
# class SomeClass:
# def __del__(self):
# print("Deleted!")
# ```
# ```py
# ```
#
#
# **Output:**
# 1\.
#
# ```py
# >>> x = SomeClass()
# >>> y = x
# >>> del x # this should print "Deleted!"
# >>> del y
# ```
# ```py
# Deleted!
# ```
#
#
# Phew, deleted at last. You might have guessed what saved from `__del__` being called in our first attempt to delete `x`. Let's add more twists to the example.
#
# 2\.
#
# ```py
# >>> x = SomeClass()
# >>> y = x
# >>> del x
# >>> y # check if y exists
# ```
# ```py
# <__main__.SomeClass instance at 0x7f98a1a67fc8>
# ```
#
# ```py
# >>> del y # Like previously, this should print "Deleted!"
# >>> globals() # oh, it didn't. Let's check all our global variables and confirm
# ```
# ```py
# Deleted!
# {'__builtins__': <module '__builtin__' (built-in)>, 'SomeClass': <class __main__.SomeClass at 0x7f98a1a5f668>, '__package__': None, '__name__': '__main__', '__doc__': None}
# ```
#
#
# Okay, now it's deleted :confused:
#
#
# #### 💡 Explanation:
# + `del x` doesn’t directly call `x.__del__()`.
# + Whenever `del x` is encountered, Python decrements the reference count for `x` by one, and `x.__del__()` when x’s reference count reaches zero.
# + In the second output snippet, `y.__del__()` was not called because the previous statement (`>>> y`) in the interactive interpreter created another reference to the same object, thus preventing the reference count from reaching zero when `del y` was encountered.
# + Calling `globals` caused the existing reference to be destroyed, and hence we can see "Deleted!" being printed (finally!).
#
#
# ### ▶ Wild imports *
#
# ```py
# # File: module.py
#
# def some_weird_name_func_():
# print("works!")
#
# def _another_weird_name_func():
# print("works!")
#
# ```
# ```py
# ```
#
#
# **Output**
#
#
# ```py
# >>> from module import *
# >>> some_weird_name_func_()
# ```
# ```py
# "works!"
# ```
#
# ```py
# >>> _another_weird_name_func()
# ```
# ```py
# Traceback (most recent call last):
# File "<stdin>", line 1, in <module>
# NameError: name '_another_weird_name_func' is not defined
# ```
#
#
#
# #### 💡 Explanation:
#
# - It is often advisable to not use wildcard imports. The first obvious reason for this is, in wildcard imports, the names with a leading underscore get imported. This may lead to errors during runtime.
# - Had we used `from ... import a, b, c` syntax, the above `NameError` wouldn't have occurred.
#
# ```py
# >>> from module import some_weird_name_func_, _another_weird_name_func
# >>> _another_weird_name_func()
# ```
# ```py
# works!
# ```
#
# - If you really want to use wildcard imports, then you'd have to define the list `__all__` in your module that will contain a list of public objects that'll be available when we do wildcard imports.
#
# ```py
# __all__ = ['_another_weird_name_func']
#
# def some_weird_name_func_():
# print("works!")
#
# def _another_weird_name_func():
# print("works!")
# ```
# ```py
# ```
#
# **Output**
#
#
# ```py
# >>> _another_weird_name_func()
# ```
# ```py
# "works!"
# ```
#
# ```py
# >>> some_weird_name_func_()
# ```
# ```py
# Traceback (most recent call last):
# File "<stdin>", line 1, in <module>
# NameError: name 'some_weird_name_func_' is not defined
# ```
#
#
#
# # Contributing
#
# A few ways in which you can contribute to wtfpython,
#
# - Suggesting new examples
# - Helping with translation (See [issues labeled translation](https://github.com/satwikkansal/wtfpython/issues?q=is%3Aissue+is%3Aopen+label%3Atranslation))
# - Minor corrections like pointing out outdated snippets, typos, formatting errors, etc.
# - Identifying gaps (things like inadequate explanation, redundant examples, etc.)
# - Any creative suggestions to make this project more fun and useful
#
# Please see [CONTRIBUTING.md](/CONTRIBUTING.md) for more details. Feel free to create a new [issue](https://github.com/satwikkansal/wtfpython/issues/new) to discuss things.
#
# PS: Please don't reach out with backlinking requests, no links will be added unless they're highly relevant to the project.
#
# # Acknowledgements
#
# The idea and design for this collection were initially inspired by <NAME>'s awesome project [wtfjs](https://github.com/denysdovhan/wtfjs). The overwhelming support by Pythonistas gave it the shape it is in right now.
#
# #### Some nice Links!
# * https://www.youtube.com/watch?v=sH4XF6pKKmk
# * https://www.reddit.com/r/Python/comments/3cu6ej/what_are_some_wtf_things_about_python
# * https://sopython.com/wiki/Common_Gotchas_In_Python
# * https://stackoverflow.com/questions/530530/python-2-x-gotchas-and-landmines
# * https://stackoverflow.com/questions/1011431/common-pitfalls-in-python
# * https://www.python.org/doc/humor/
# * https://github.com/cosmologicon/pywat#the-undocumented-converse-implication-operator
# * https://www.codementor.io/satwikkansal/python-practices-for-efficient-code-performance-memory-and-usability-aze6oiq65
# * https://github.com/wemake-services/wemake-python-styleguide/search?q=wtfpython&type=Issues
#
# # 🎓 License
#
# [![WTFPL 2.0][license-image]][license-url]
#
# © [<NAME>](https://satwikkansal.xyz)
#
# [license-url]: http://www.wtfpl.net
# [license-image]: https://img.shields.io/badge/License-WTFPL%202.0-lightgrey.svg?style=flat-square
#
#
| irrelevant/wtf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <b>Calcule a integral dada</b>
# $\int_{-1}^{1} 3t^4dt$
# $\frac{3t^5}{5}, -1 \leq X \leq 1$
# $\frac{3\cdot(1)^5}{5} - \frac{3\cdot(-1)^5}{5}$
# $\frac{3}{5} + \frac{3}{5}$
# $\frac{6}{5}$
| Problemas 5.3/05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from scipy.io import loadmat
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import collections
from gtda.time_series import TakensEmbedding
from gtda.homology import VietorisRipsPersistence
from gtda.diagrams import Amplitude, NumberOfPoints, PersistenceEntropy
import scipy.io
from sklearn.model_selection import train_test_split, StratifiedKFold
from tqdm.notebook import tqdm
# -
# ## Load Data
def load_data(name, dict_name):
matdic = scipy.io.loadmat(name) #signal_100-500.mat #signal_CAR_1-500.mat
#numpy array
#data = matdic['signal_100hz']
#data = matdic['signal_100hz_CAR']
#data = matdic['ytot']
#data = matdic['signal_1hz_CAR']
data = matdic[dict_name]
labels = data[60]
diff =np.diff(labels)
index = np.where(diff) #indici dove cambia classe
index = index[0]+1
index = np.append(index,(len(labels)))
index = np.insert(index,0,0)
labels = labels[index[:-1]]
durations = np.ediff1d(index, to_begin=index[0]) #durata classi
channels = data[:60,:]
samples = []
samples_labels = []
for i in range(len(index)-1):
samples.append(channels[:,index[i]:index[i+1]])
samples_labels.append((channels[:,index[i]:index[i+1]],labels[i]))
labels= labels+1
labels = labels.astype(int)
return samples, labels
def zero_padding_tensor(samples):
save_s = []
for i in range(0, len(samples)):
save_s.append(samples[i].shape[1])
max_shape = max(save_s)
## Zero padding, all vectors with max_shape entries (=3552)
samples_tensor = np.pad(samples[0],(0, max_shape - samples[0].shape[1]), 'constant', constant_values = (0))[:60,:]
sh_0 = samples_tensor.shape[0]
sh_1 = samples_tensor.shape[1]
samples_tensor = samples_tensor.reshape((1, sh_0, sh_1 ))
for i in range(1, len(samples)):
samples_tensor = np.concatenate((samples_tensor, np.pad(samples[i],(0, max_shape - samples[i].shape[1]), 'constant', constant_values = (0))[:60,:].reshape((1, sh_0, sh_1 ))))
return samples_tensor
def list_from_tensor(Xtr, Ytr, Xte, Yte):
samples_train = []
labels_train = []
samples_test = []
labels_test = []
for i in range(0, Xtr.shape[0]):
samples_train.append(Xtr[i, :, :])
labels_train.append(Ytr[i, : ])
for i in range(0, Xte.shape[0]):
samples_test.append(Xte[i, :, :])
labels_test.append(Yte[i, :])
return samples_train, labels_train, samples_test, labels_test
def aug_pipeline(filename, dict_name, of_type_list = True, test_size = 0.20):
samples, labels = load_data(name = filename, dict_name = dict_name)
print("Data Loading Done \n")
samples_tensor = zero_padding_tensor(samples)
print("Zero Padding Done \n")
X_ = samples_tensor
#X_ = np.swapaxes(X_, 1,2)
Y_ = labels
Y_ = Y_.reshape((Y_.shape[0],1))
# there's no need for t/s split if we employ KFold CV
#Xtr, Xte, Ytr, Yte = train_test_split(X_, Y_, test_size = test_size, random_state = 0 )
#print("Train Test Split Done \n")
#Xaug, Yaug = augment_X_y(X_, Y_) #augment_X_y(Xtr, Ytr)
#print("Data Augmentation Done \n")
Xaug, Yaug = X_, Y_
Xte, Yte = np.zeros_like(X_), np.zeros_like(Y_) # placeholder for final version
#Xte, Yte = np.zeros_like(Xaug), np.zeros_like(Yaug) # placeholder for final version to be removed
if of_type_list == True:
X_train, y_train, X_test, y_train = list_from_tensor(Xaug, Yaug, Xte, Yte)
return X_train, Yaug, X_test, Yte
return Xaug, Yaug, Xte, Yte
# +
# files to be loaded as training/validation
filenames = ['../ECoG_Data/signal_CAR_1-500.mat', '../ECoG_Data/signal_CAR_100-500.mat',
'../ECoG_Data/signal_100-500.mat','../ECoG_Data/signal_50-300.mat']
dict_names = ['signal_1hz_CAR','signal_100hz_CAR',
'signal_100hz', 'signal_50hz_CAR' ]
# -
# ## TDA Features Extraction
def extract_features(X, y, time_delay=1, dimension=1, stride=10, h_dim=(0,1), n_jobs=-1):
samples, labels = X, y
TE = TakensEmbedding(time_delay=time_delay, dimension=dimension, stride=stride)
tes = []
for sample in samples:
te = TE.fit_transform(sample)
te = np.squeeze(te)
tes.append(te.T)
VR = VietorisRipsPersistence(homology_dimensions=h_dim, n_jobs=n_jobs)
X_vr = VR.fit_transform(tes)
X_features = []
#amplitude
metrics = ['bottleneck','wasserstein', 'betti', 'landscape', 'silhouette', 'heat']
for metric in metrics:
Ampl = Amplitude(metric=metric, n_jobs=n_jobs)
X_a = Ampl.fit_transform(X_vr)
X_features.append(X_a[:,0])
X_features.append(X_a[:,1])
#entropy
PE = PersistenceEntropy(normalize=True, nan_fill_value=-1, n_jobs=n_jobs)
X_pe = PE.fit_transform(X_vr)
X_features.append(X_pe[:,0])
X_features.append(X_pe[:,1])
PE = PersistenceEntropy(normalize=False, nan_fill_value=-1, n_jobs=n_jobs)
X_pe = PE.fit_transform(X_vr)
X_features.append(X_pe[:,0])
X_features.append(X_pe[:,1])
#number of points
NOP = NumberOfPoints(n_jobs=n_jobs)
X_nop = NOP.fit_transform(X_vr)
X_features.append(X_nop[:,0])
X_features.append(X_nop[:,1])
X = np.array(X_features).T
y = labels
return X, y, X_vr
# ## Classification
# +
from time import time
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.neural_network import MLPClassifier
from gtda.diagrams import PairwiseDistance # for KNeighborsClassifier
# -
from sklearn import metrics
def train_model(model, x, y):
#model = ensemble.RandomForestClassifier(**params)
# initialize stratified k-fold
kf = StratifiedKFold(n_splits=5)
# initialize accuracy list
accuracies = []
# loop over all folds
for idx in kf.split(X=x, y=y):
train_idx, test_idx = idx[0], idx[1]
xtrain = x[train_idx]
ytrain = y[train_idx]
xtest = x[test_idx]
ytest = y[test_idx]
# fit model for current fold
model.fit(xtrain, ytrain)
#create predictions
preds = model.predict(xtest)
# calculate and append accuracy
fold_accuracy = metrics.accuracy_score(
ytest,
preds
)
accuracies.append(fold_accuracy)
# return negative accuracy
avg_accuracy = np.mean(accuracies)
std_dev = np.std(accuracies)
return avg_accuracy, std_dev
# ## Random Forest
BP_RF = pd.read_csv("Outputs/tda_best_params_rf.csv")
BP_RF
#print(pd.read_csv("Outputs/best_params_rf.csv").to_latex(index = False))
rf = RandomForestClassifier(max_depth = 8, n_estimators = 1500,
criterion = 'entropy', max_features = 1.)
rf_1 = RandomForestClassifier(max_depth = 8, n_estimators = 1361,
criterion = 'entropy', max_features =0.130745)
rf_2 = RandomForestClassifier(max_depth = 3, n_estimators = 1351,
criterion = 'entropy', max_features = 1.)
rf_3 = RandomForestClassifier(max_depth = 4, n_estimators = 217,
criterion = 'entropy', max_features = 0.172762)
# +
idx = 0
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs, stds = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(rf, X, y)
accs.append(acc)
stds.append(std)
rf_acc = sum(accs)/len(accs)
rf_std = sum(stds)/len(stds)
# +
idx = 1
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_1, stds_1 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(rf_1, X, y)
accs_1.append(acc)
stds_1.append(std)
rf_1_acc = sum(accs_1)/len(accs_1)
rf_1_std = sum(stds_1)/len(stds_1)
# +
idx = 2
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_2, stds_2 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(rf_2, X, y)
accs_2.append(acc)
stds_2.append(std)
rf_2_acc = sum(accs_2)/len(accs_2)
rf_2_std = sum(stds_2)/len(stds_2)
# +
idx = 3
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_3, stds_3 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(rf_3, X, y)
accs_3.append(acc)
stds_3.append(std)
rf_3_acc = sum(accs_3)/len(accs_3)
rf_3_std = sum(stds_3)/len(stds_3)
# -
rf_acc, rf_std
rf_1_acc, rf_1_std
rf_2_acc, rf_2_std
rf_3_acc, rf_3_std
# +
rf_accs = [rf_acc, rf_1_acc, rf_2_acc, rf_3_acc]
rf_stds = [rf_std, rf_1_std, rf_2_std, rf_3_std]
BP_RF['accuracy'] = rf_accs
BP_RF['std_dev'] = rf_stds
BP_RF = BP_RF.round({'max_features': 3, 'accuracy': 2, 'std_dev': 2})
BP_RF.to_csv("Outputs/Tables/TDA_random_forest.csv")
print(BP_RF.to_latex(index = False))
# -
# ## Gradient Boosting
BP_GB = pd.read_csv("Outputs/tda_best_params_gb.csv")
BP_GB
gb = GradientBoostingClassifier(max_depth = 6, n_estimators = 100,
criterion = 'mse', subsample = 1.000000)
gb_1 = GradientBoostingClassifier(max_depth = 10, n_estimators = 100,
criterion = 'mse', subsample = 0.853072)
gb_2 = GradientBoostingClassifier(max_depth = 10, n_estimators = 1500 ,
criterion = 'friedman_mse', subsample = 0.880275)
gb_3 = GradientBoostingClassifier(max_depth = 10, n_estimators = 116,
criterion = 'friedman_mse', subsample = 0.933637)
# +
idx = 0
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs, stds = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gb, X, y)
accs.append(acc)
stds.append(std)
gb_acc = sum(accs)/len(accs)
gb_std = sum(stds)/len(stds)
# +
idx = 1
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_1, stds_1 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gb_1, X, y)
accs_1.append(acc)
stds_1.append(std)
gb_1_acc = sum(accs_1)/len(accs_1)
gb_1_std = sum(stds_1)/len(stds_1)
# +
idx = 2
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_2, stds_2 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gb_2, X, y)
accs_2.append(acc)
stds_2.append(std)
gb_2_acc = sum(accs_2)/len(accs_2)
gb_2_std = sum(stds_2)/len(stds_2)
# +
idx = 3
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_3, stds_3 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gb_3, X, y)
accs_3.append(acc)
stds_3.append(std)
gb_3_acc = sum(accs_3)/len(accs_3)
gb_3_std = sum(stds_3)/len(stds_3)
# -
gb_acc, gb_std
gb_1_acc, gb_1_std
gb_2_acc, gb_2_std
gb_3_acc, gb_3_std
# +
gb_accs = [gb_acc, gb_1_acc, gb_2_acc, gb_3_acc]
gb_stds = [gb_std, gb_1_std, gb_2_std, gb_3_std]
BP_GB['accuracy'] = gb_accs
BP_GB['std_dev'] = gb_stds
BP_GB = BP_GB.round({'subsample': 3, 'accuracy': 2, 'std_dev': 2})
#BP_GB.to_csv("Outputs/Tables/TDA_gradient_boosting.csv")
print(BP_GB.to_latex(index = False))
# -
# ## Support Vector Machine
BP_SVM = pd.read_csv("Outputs/tda_best_params_svm.csv")
BP_SVM
# +
svm = SVC(C = 2.262615 , kernel = 'linear')
svm_1 = SVC(C = 10., kernel = 'linear')
svm_2 = SVC(C = 8.255089 , kernel = 'rbf')
svm_3 = SVC(C = 0.001 , kernel = 'linear')
idx = 0
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs, stds = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(svm, X, y)
accs.append(acc)
stds.append(std)
svm_acc = sum(accs)/len(accs)
svm_std = sum(stds)/len(stds)
idx = 1
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_1, stds_1 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(svm_1, X, y)
accs_1.append(acc)
stds_1.append(std)
svm_1_acc = sum(accs_1)/len(accs_1)
svm_1_std = sum(stds_1)/len(stds_1)
idx = 2
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_2, stds_2 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(svm_2, X, y)
accs_2.append(acc)
stds_2.append(std)
svm_2_acc = sum(accs_2)/len(accs_2)
svm_2_std = sum(stds_2)/len(stds_2)
idx = 3
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_3, stds_3 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(svm_3, X, y)
accs_3.append(acc)
stds_3.append(std)
svm_3_acc = sum(accs_3)/len(accs_3)
svm_3_std = sum(stds_3)/len(stds_3)
##################################
svm_accs = [svm_acc, svm_1_acc, svm_2_acc, svm_3_acc]
svm_stds = [svm_std, svm_1_std, svm_2_std, svm_3_std]
BP_SVM['accuracy'] = svm_accs
BP_SVM['std_dev'] = svm_stds
BP_SVM = BP_SVM.round({'C': 3, 'accuracy': 2, 'std_dev': 2})
BP_SVM.to_csv("Outputs/Tables/TDA_svm.csv")
print(BP_SVM.to_latex(index = False))
# -
# ## MLP
BP_MLP = pd.read_csv("Outputs/tda_best_params_mlp.csv")
BP_MLP
# +
#print(pd.read_csv("Outputs/best_params_mlp.csv").to_latex(index = False))
mlp = MLPClassifier(hidden_layer_sizes = (180, 300, 20), max_iter = 5000, alpha = 0.007477)
mlp_1 = MLPClassifier(hidden_layer_sizes = (180, 300, 20), max_iter = 5000, alpha = 0.020931)
mlp_2 = MLPClassifier(hidden_layer_sizes = (180, 300, 20), max_iter = 5000, alpha = 0.008003)
mlp_3 = MLPClassifier(hidden_layer_sizes = (180, 300, 20), max_iter = 5000, alpha = 0.099996)
idx = 0
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs, stds = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(mlp, X, y)
accs.append(acc)
stds.append(std)
mlp_acc = sum(accs)/len(accs)
mlp_std = sum(stds)/len(stds)
idx = 1
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_1, stds_1 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(mlp_1, X, y)
accs_1.append(acc)
stds_1.append(std)
mlp_1_acc = sum(accs_1)/len(accs_1)
mlp_1_std = sum(stds_1)/len(stds_1)
idx = 2
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_2, stds_2 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(mlp_2, X, y)
accs_2.append(acc)
stds_2.append(std)
mlp_2_acc = sum(accs_2)/len(accs_2)
mlp_2_std = sum(stds_2)/len(stds_2)
idx = 3
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_3, stds_3 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(mlp_3, X, y)
accs_3.append(acc)
stds_3.append(std)
mlp_3_acc = sum(accs_3)/len(accs_3)
mlp_3_std = sum(stds_3)/len(stds_3)
##################################
mlp_accs = [mlp_acc, mlp_1_acc, mlp_2_acc, mlp_3_acc]
mlp_stds = [mlp_std, mlp_1_std, mlp_2_std, mlp_3_std]
BP_MLP['accuracy'] = mlp_accs
BP_MLP['std_dev'] = mlp_stds
BP_MLP = BP_MLP.round({'alpha': 3, 'accuracy': 2, 'std_dev': 2})
BP_MLP.to_csv("Outputs/Tables/TDA_mlp.csv")
print(BP_MLP.to_latex(index = False))
# -
# ### Gaussian Naive Bayes
BP_GNB = pd.read_csv("Outputs/tda_best_params_gnb.csv")
BP_GNB
# +
#print(pd.read_csv("Outputs/best_params_gnb.csv").to_latex(index = False))
gnb = GaussianNB(var_smoothing= 5.055780e-08)
gnb_1 = GaussianNB(var_smoothing= 7.497147e-09)
gnb_2 = GaussianNB(var_smoothing= 1.289194e-10)
gnb_3 = GaussianNB(var_smoothing= 8.680831e-08)
idx = 0
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs, stds = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gnb, X, y)
accs.append(acc)
stds.append(std)
gnb_acc = sum(accs)/len(accs)
gnb_std = sum(stds)/len(stds)
idx = 1
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_1, stds_1 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gnb_1, X, y)
accs_1.append(acc)
stds_1.append(std)
gnb_1_acc = sum(accs_1)/len(accs_1)
gnb_1_std = sum(stds_1)/len(stds_1)
idx = 2
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_2, stds_2 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gnb_2, X, y)
accs_2.append(acc)
stds_2.append(std)
gnb_2_acc = sum(accs_2)/len(accs_2)
gnb_2_std = sum(stds_2)/len(stds_2)
idx = 3
samples, y = load_data(name = filenames[idx],dict_name = dict_names[idx])
X_train, Y_train, _, _ = aug_pipeline(filename = filenames[idx],dict_name = dict_names[idx], of_type_list = True)
Y_train = np.squeeze(Y_train)
X, y, X_vr = extract_features(X_train, Y_train, h_dim=(0,1))
accs_3, stds_3 = [], []
for i in tqdm(range(0, 50)):
acc, std = train_model(gnb_3, X, y)
accs_3.append(acc)
stds_3.append(std)
gnb_3_acc = sum(accs_3)/len(accs_3)
gnb_3_std = sum(stds_3)/len(stds_3)
##################################
gnb_accs = [gnb_acc, gnb_1_acc, gnb_2_acc, gnb_3_acc]
gnb_stds = [gnb_std, gnb_1_std, gnb_2_std, gnb_3_std]
BP_GNB['accuracy'] = gnb_accs
BP_GNB['std_dev'] = gnb_stds
BP_GNB = BP_GNB.round({'accuracy': 2, 'std_dev': 2})
BP_GNB.to_csv("Outputs/Tables/TDA_gnb.csv")
print(BP_GNB.to_latex(index = False))
| Best Models - TDA & Tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pneumonia Classification Task Using Keras
# +
import os
import numpy as np
import pandas as pd
import random
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow.keras.backend as K
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Input, Dense, Flatten, Dropout, BatchNormalization
from tensorflow.keras.layers import Conv2D, SeparableConv2D, MaxPool2D, LeakyReLU, Activation, Convolution2D, MaxPooling2D
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.callbacks import ModelCheckpoint, ReduceLROnPlateau, EarlyStopping
import tensorflow as tf
# +
input_path = './images/'
fig, ax = plt.subplots(2, 3, figsize=(15, 7))
ax = ax.ravel()
plt.tight_layout()
for i, _set in enumerate(['train', 'val', 'test']):
set_path = input_path+_set
ax[i].imshow(plt.imread(set_path+'/NORMAL/'+os.listdir(set_path+'/NORMAL')[0]), cmap='gray')
ax[i].set_title(f'Set: {_set}, Condition: Normal')
ax[i+3].imshow(plt.imread(set_path+'/PNEUMONIA/'+os.listdir(set_path+'/PNEUMONIA')[0]), cmap='gray')
ax[i+3].set_title(f'Set: {_set}, Condition: Pneumonia')
# -
for _set in ['train', 'val', 'test']:
n_normal = len(os.listdir(input_path + _set + '/NORMAL'))
n_infect = len(os.listdir(input_path + _set + '/PNEUMONIA'))
print('Set: {}, normal images: {}, pneumonia images: {}'.format(_set, n_normal, n_infect))
def process_data(img_dims, batch_size):
# Data generation objects
train_datagen = ImageDataGenerator(rescale=1./255, zoom_range=0.3, vertical_flip=True)
test_val_datagen = ImageDataGenerator(rescale=1./255)
# This is fed to the network in the specified batch sizes and image dimensions
train_gen = train_datagen.flow_from_directory(
directory=input_path+'train',
target_size=(img_dims, img_dims),
batch_size=batch_size,
class_mode='binary',
shuffle=True)
test_gen = test_val_datagen.flow_from_directory(
directory=input_path+'test',
target_size=(img_dims, img_dims),
batch_size=batch_size,
class_mode='binary',
shuffle=True)
test_data = []
test_labels = []
for cond in ['/NORMAL/', '/PNEUMONIA/']:
for img in (os.listdir(input_path + 'test' + cond)):
img = plt.imread(input_path+'test'+cond+img)
img = cv2.resize(img, (img_dims, img_dims))
img = np.dstack([img, img, img])
img = img.astype('float32') / 255
if cond=='/NORMAL/':
label = 0
elif cond=='/PNEUMONIA/':
label = 1
test_data.append(img)
test_labels.append(label)
test_data = np.array(test_data)
test_labels = np.array(test_labels)
return train_gen, test_gen, test_data, test_labels
# +
img_dims = 150
epochs = 10
batch_size = 32
train_gen, test_gen, test_data, test_labels = process_data(img_dims, batch_size)
# +
# Initialising the CNN
classifier = Sequential()
# Step 1 - Convolution
classifier.add(Convolution2D(32, 3, 3, input_shape = (150, 150, 3), activation = 'relu'))
# Step 2 - Pooling
classifier.add(MaxPooling2D(pool_size = (2, 2)))
# Step 3 - Flattening
classifier.add(Flatten())
# Step 4 - Full connection
classifier.add(Dense(128, activation = 'relu'))
classifier.add(Dense(1, activation = 'sigmoid'))
# Compiling the CNN
classifier.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# -
ihist = classifier.fit_generator(
train_gen, steps_per_epoch=train_gen.samples // batch_size,
epochs=epochs, validation_data=test_gen,
validation_steps=test_gen.samples // batch_size)
# +
fig, ax = plt.subplots(1, 2, figsize=(10, 3))
ax = ax.ravel()
for i, met in enumerate(['accuracy', 'loss']):
print(i, met)
ax[i].plot(ihist.history[met])
ax[i].plot(ihist.history['val_' + met])
ax[i].set_title('Model {}'.format(met))
ax[i].set_xlabel('epochs')
ax[i].set_ylabel(met)
ax[i].legend(['train', 'val'])
# +
from sklearn.metrics import accuracy_score, confusion_matrix
preds = classifier.predict(test_data)
acc = accuracy_score(test_labels, np.round(preds))*100
cm = confusion_matrix(test_labels, np.round(preds))
tn, fp, fn, tp = cm.ravel()
print('CONFUSION MATRIX ------------------')
print(cm)
print('')
print('TEST METRICS ----------------------')
precision = tp/(tp+fp)*100
recall = tp/(tp+fn)*100
print(f'Accuracy: {acc}')
print(f'Precision: {precision}')
print(f'Recall: {recall}')
print(f'F1-score: {2*precision*recall/(precision+recall)}')
print('\nTRAIN METRIC ----------------------')
print(f'Train acc: {ihist.history["accuracy"][-1] } ' )
# -
# This links to a way more complex solution!
# Try it out as well!
#
# https://towardsdatascience.com/deep-learning-for-detecting-pneumonia-from-x-ray-images-fc9a3d9fdba8
| W8D8/PneumoniaProject.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch import nn, optim
import torch.nn.functional as F
# +
with open('alice.txt', 'r', encoding='latin1') as f:
data = f.read()
print("Extract: ", data[:50])
print("Length: ", len(data))
# -
chars = list(set(data))
indexer = {char: index for (index, char) in enumerate(chars)}
# +
indexed_data = []
for c in data:
indexed_data.append(indexer[c])
print("Indexed extract: ", indexed_data[:50])
print("Length: ", len(indexed_data))
# -
def index2onehot(batch):
batch_flatten = batch.flatten()
onehot_flat = np.zeros((batch.shape[0] * batch.shape[1], len(indexer)))
onehot_flat[range(len(batch_flatten)), batch_flatten] = 1
onehot = onehot_flat.reshape((batch.shape[0], batch.shape[1], -1))
return onehot
class LSTM(nn.Module):
def __init__(self, char_length, hidden_size, n_layers):
super().__init__()
self.hidden_size = hidden_size
self.n_layers = n_layers
self.lstm = nn.LSTM(char_length, hidden_size, n_layers, batch_first=True)
self.output = nn.Linear(hidden_size, char_length)
def forward(self, x, states):
out, states = self.lstm(x, states)
out = out.contiguous().view(-1, self.hidden_size)
out = self.output(out)
return out, states
def init_states(self, batch_size):
hidden = next(self.parameters()).data.new(self.n_layers, batch_size, self.hidden_size).zero_()
cell = next(self.parameters()).data.new(self.n_layers, batch_size, self.hidden_size).zero_()
states = (hidden, cell)
return states
# +
n_seq = 100 ## Number of sequences per batch
seq_length = 50
n_batches = math.floor(len(indexed_data) / n_seq / seq_length)
total_length = n_seq * seq_length * n_batches
x = indexed_data[:total_length]
x = np.array(x).reshape((n_seq,-1))
# -
model = LSTM(len(chars), 256, 2)
model
loss_function = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
epochs = 20
losses = []
for e in range(1, epochs+1):
states = model.init_states(n_seq)
batch_loss = []
for b in range(0, x.shape[1], seq_length):
x_batch = x[:,b:b+seq_length]
if b == x.shape[1] - seq_length:
y_batch = x[:,b+1:b+seq_length]
y_batch = np.hstack((y_batch, indexer["."] * np.ones((y_batch.shape[0],1))))
else:
y_batch = x[:,b+1:b+seq_length+1]
x_onehot = torch.Tensor(index2onehot(x_batch))
y = torch.Tensor(y_batch).view(n_seq * seq_length)
pred, states = model(x_onehot, states)
loss = loss_function(pred, y.long())
optimizer.zero_grad()
loss.backward(retain_graph=True)
optimizer.step()
batch_loss.append(loss.item())
losses.append(np.mean(batch_loss))
if e%2 == 0:
print("epoch: ", e, "... Loss function: ", losses[-1])
x_range = range(len(losses))
plt.plot(x_range, losses)
plt.xlabel("epochs")
plt.ylabel("Loss function")
plt.show()
# +
starter = "So she was considering in her own mind "
states = None
for ch in starter:
x = np.array([[indexer[ch]]])
x = index2onehot(x)
x = torch.Tensor(x)
pred, states = model(x, states)
counter = 0
while starter[-1] != "." and counter < 100:
counter += 1
x = np.array([[indexer[starter[-1]]]])
x = index2onehot(x)
x = torch.Tensor(x)
pred, states = model(x, states)
pred = F.softmax(pred, dim=1)
p, top = pred.topk(10)
p = p.detach().numpy()[0]
top = top.numpy()[0]
index = np.random.choice(top, p=p/p.sum())
starter += chars[index]
print(starter)
# -
| Activity02/Activity02.ipynb |
try:
import openmdao.api as om
import dymos as dm
except ImportError:
# !python -m pip install openmdao[notebooks]
# !python -m pip install dymos[docs]
import openmdao.api as om
import dymos as dm
# # The Phase API
# ## options
om.show_options_table('dymos.phase.Phase')
# The transcription is an instance of one of the [transcriptions](../transcriptions) available in Dymos.
#
# ## set_time_options
#
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_time_options
# :noindex:
# ```
#
# ## add_state
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_state
# :noindex:
# ```
#
# ## set_state_options
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_state_options
# :noindex:
# ```
#
# ## add_control
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_control
# :noindex:
# ```
#
# ## set_control_options
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_control_options
# :noindex:
# ```
#
# ## add_polynomial_control
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_polynomial_control
# :noindex:
# ```
#
# ## set_polynomial_control_options
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_polynomial_control_options
# :noindex:
# ```
#
# ## add_parameter
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_parameter
# :noindex:
# ```
#
# ## set_parameter_options
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_parameter_options
# :noindex:
# ```
#
# ## add_timeseries
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_timeseries
# :noindex:
# ```
#
# ## add_timeseries_output
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_timeseries_output
# :noindex:
# ```
#
# ## add_boundary_constraint
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_boundary_constraint
# :noindex:
# ```
#
# ## add_path_constraint
# ```{eval-rst}
# .. automethod:: dymos.Phase.add_path_constraint
# :noindex:
# ```
#
# ## simulate
# ```{eval-rst}
# .. automethod:: dymos.Phase.simulate
# :noindex:
# ```
#
# ## set_refine_options
# ```{eval-rst}
# .. automethod:: dymos.Phase.set_refine_options
# :noindex:
# ```
#
# ## interpolate
# ```{eval-rst}
# .. automethod:: dymos.Phase.interpolate
# :noindex:
# ```
| docs/api/phase_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Transformação de intensidade, Realce e Equalização de histograma
#
# O objetivo destas atividades é explorar várias formas de mapeamento dos valores
# dos pixels. É a categoria de funções que apenas alteram o valor do pixel aplicando-se
# uma função ou tabela de mapeamento. Veremos que é possível implementar várias
# funções de melhoria de contraste e brilho da imagem.
#
# 1. Estude atentamente o tutorial sobre Transformada de intensidade. Ele mostra como
# podemos fazer estas operações de forma eficiente através de uma tabela de mapeamento:
#
# - [Tutorial sobre Transformação de Intensidade](../master/tutorial_ti_2.ipynb)
#
# 2. Um dos exercícios tradicionais em processamento de imagens é a equalização de histograma.
# O tutorial a seguir ilustra como fazer isto de forma simples e eficiente no NumPy:
#
# - [Tutorial sobre equalização de histograma](../master/tutorial_hist_eq_2.ipynb)
#
# 3. É possível fazer a equalização da imagem utilizando também o conceito usado para
# construir um mosaico com tons de cinza igualmente distribuídos. Confira esta forma
# intuitiva e eficiente de se equalizar a distribuição de pixels na imagem.
#
# - [Tutorial equalização de histograma por montagem de mosaico](../master/tutorial_pehist_1.ipynb)
#
# 4. Acompanhando o tutorial anterior é possível estendê-lo para transformar a imagem para
# ter um histograma especificado. Acompanhe o desenvolvimento desta forma simples de
# especificação de histograma:
#
# - [Tutorial sobre especificação de histograma](../master/tutorial_pehist_2.ipynb)
#
# + [markdown] deletable=true editable=true
# ## Normalização da imagem para média zero e variância um
#
# É comum se fazer uma normalização da imagem de modo que ela fique em ponto flutuante, com média zero e variância um.
#
#
# -
# ## Correção gama
| deliver/Atividade_3_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''.venv'': venv)'
# name: python3
# ---
# # House Prices - Advanced Regression Techniques
# ### Predict sales prices and practice feature engineering, RFs, and gradient boosting
#
# https://www.kaggle.com/c/house-prices-advanced-regression-techniques/overview
# # Imports
# +
import pandas as pd
from sklearn.model_selection import train_test_split
from category_encoders import OrdinalEncoder
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import mean_squared_error
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.wrappers.scikit_learn import KerasRegressor
from keras.metrics import RootMeanSquaredError
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
# -
# # EDA
# Load and preview
df = pd.read_csv('train.csv')
df
# +
# Drop columns with too many missing value
na = df.isna().sum()
# Drop anything over 600 missing value
df = df.drop(columns=['Alley', 'FireplaceQu', 'PoolQC', 'Fence', 'MiscFeature'])
# Then Drop na
# df = df.dropna()
for col in df.columns:
if df[col].dtypes == 'object':
df[col].fillna('None')
print(f"final data shape: {df.shape}")
# +
# Split Data
train, val = train_test_split(df, random_state=1)
target = 'SalePrice'
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
# +
# Preprocess Data
pre = make_pipeline(OrdinalEncoder(), SimpleImputer(), StandardScaler())
X_train_scaled = pre.fit_transform(X_train)
X_val_scaled = pre.transform(X_val)
# +
n_features = X_train_scaled.shape[1]
model = Sequential()
model.add(Dense(128, activation='relu', kernel_initializer='he_normal', input_shape=(n_features,)))
model.add(Dense(128, activation='relu', kernel_initializer='he_normal'))
model.add(Dense(128, activation='relu', kernel_initializer='he_normal'))
model.add(Dense(1))
model.compile(optimizer='adam', loss='mse', metrics=[RootMeanSquaredError()])
model.fit(X_train_scaled, y_train, batch_size=32, epochs=50, validation_data=(X_val_scaled, y_val))
# -
# # Predict test data and save
# +
test = pd.read_csv('test.csv')
# Drop anything over 600 missing value
test = test.drop(columns=['Alley', 'FireplaceQu', 'PoolQC', 'Fence', 'MiscFeature'])
# Then Drop na
# df = df.dropna()
for col in test.columns:
if test[col].dtypes == 'object':
test[col].fillna('None')
print(f"final data shape: {test.shape}")
y_pred = model.predict(pre.transform(test)).ravel()
print(y_pred)
submit = pd.DataFrame({'Id': pd.read_csv('sample_submission.csv')['Id'], 'SalePrice': y_pred})
submit.to_csv('submission_ver2.csv', index=False)
| getting-started/home-data-for-ml-course/2_ANN.ipynb |