code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0-rc4
# language: julia
# name: julia-1.3
# ---
# # Test JLD2 compression
using Revise
using VCFTools
using MendelImpute
using GeneticVariation
using Random
using SparseArrays
using JLD2, FileIO
using CodecZlib
using BSON
using HDF5
using JLSO
using BenchmarkTools
# using Plots
# using ProfileView
# ## Check reading speed and file size for simulated data
reffile = "haplo_ref.vcf.gz"
outfile = "haplo_ref.jld2"
@time hapset = save_jld2(reffile, outfile, column_major=true);
;gzip haplo_ref.jld2 haplo_ref.jld2.gz
;ls -al haplo_ref.vcf.gz # original size of ref file (bytes)
;ls -al haplo_ref.jld2 # size of .jld2 file (bytes)
;ls -al haplo_ref.jld2.gz # size of .jld2.gz file (bytes)
20318574 / 5449864 # .jld2 is larger file size
6754174 / 5449864 # .jld2.gz is larger file size
#difference in reading speed
@time H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
@time @load "haplo_ref.jld2" hapset;
7.085228 / 0.065357
# ## Check reading speed and file size for chr22 data in 1000 genomes
reffile = "chr22.uniqueSNPs.vcf.gz"
outfile = "chr22.uniqueSNPs.jld2"
@time hapset = save_jld2(reffile, outfile, column_major=true);
;gzip chr22.uniqueSNPs.jld2 chr22.uniqueSNPs.jld2.gz
;ls -al chr22.uniqueSNPs.vcf.gz # original size of ref file (bytes)
;ls -al chr22.uniqueSNPs.jld2 # size of .jld2 file (bytes)
;ls -al chr22.uniqueSNPs.jld2.gz # size of .jld2.gz file (bytes)
615840218 / 142889955 # .jld2 is larger file size
187644621 / 142889955 # .jld2.gz is larger file size
#difference in reading speed
@time H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
@time @load "chr22.uniqueSNPs.jld2" hapset;
242.880364 / 1.496514
# ## .gz compressed jld2
#
# Not sure how to make this work
# +
reffile = "chr22.uniqueSNPs.jld2.gz"
io = GzipDecompressorStream(open(reffile, "r"))
x = read(io)
hapset = jldopen(x, "r")
# hapset = read(io, "hapset")
# jldopen(GzipDecompressorStream(open(reffile)), "r") do file
# file["bigdata"] = randn(5)
# end
# -
typeof(io) <: IOStream
# # Try BSON format
reffile = "chr22.uniqueSNPs.vcf.gz"
H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
hapset = MendelImpute.RefHaplotypes(H, true, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt);
@time bson("chr22.uniqueSNPs.bson", hapset = hapset)
@time BSON.@load "chr22.uniqueSNPs.bson" hapset
;ls -al chr22.uniqueSNPs.vcf.gz # original size of ref file (bytes)
;ls -al chr22.uniqueSNPs.bson # BSON is also large file
468468844 / 142889955
;gzip chr22.uniqueSNPs.bson chr22.uniqueSNPs.bson.gz
;ls -al chr22.uniqueSNPs.bson.gz
468468844 / 169768322
pwd()
hapset = bsonopen(x, "r")
# ## Try HDF5
reffile = "chr22.uniqueSNPs.vcf.gz"
H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
hapset = MendelImpute.RefHaplotypes(H, true, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt)
?h5write
# ## Try JLSO
reffile = "haplo_ref.vcf.gz"
H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
hapset = MendelImpute.RefHaplotypes(H, true, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt);
@time JLSO.save("haplo_ref.jlso", :hapset => hapset, format=:bson, compression=:gzip_smallest)
;ls -al haplo_ref.vcf.gz
;ls -al haplo_ref.jlso
# filesize = 5505991, format=:julia_serialize, compression=:gzip_smallest
@btime loaded = JLSO.load("haplo_ref.jlso");
# filesize = 5758746, format=:julia_serialize, compression=:gzip_fastest
@btime loaded = JLSO.load("haplo_ref.jlso");
# filesize = 6133533, format=:bson, compression=:gzip_fastest
@btime loaded = JLSO.load("haplo_ref.jlso");
# filesize = 5851597, format=:bson, compression=:gzip_smallest
@btime loaded = JLSO.load("haplo_ref.jlso");
;ls -al chr22.uniqueSNPs.vcf.gz
reffile = "chr22.uniqueSNPs.vcf.gz"
H, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt = convert_ht(Bool, reffile, trans=true, save_snp_info=true, msg = "Importing reference haplotype files...")
hapset = MendelImpute.RefHaplotypes(H, true, H_sampleID, H_chr, H_pos, H_ids, H_ref, H_alt);
@time JLSO.save("chr22.uniqueSNPs.jlso", :hapset => hapset, format=:bson, compression=:none)
;ls -al chr22.uniqueSNPs.jlso
# filesize = 156750465, format=:julia_serialize, compression=:gzip_smallest
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 163328088, format=:julia_serialize, compression=:gzip
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 179698900, format=:julia_serialize, compression=:gzip_fastest
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 163082396, format=:bson, compression=:gzip_smallest
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 169771730, format=:bson, compression=:gzip
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 186600622, format=:bson, compression=:gzip_fastest
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
# filesize = 468472432, format=:bson, compression=:none
@btime loaded = JLSO.load("chr22.uniqueSNPs.jlso") seconds=30;
@time loaded = JLSO.load("chr22.uniqueSNPs.jlso")
hs = loaded[:hapset];
all(hs.H .== hapset.H)
| simulation/compression_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inference and Validation
#
# Now that you have a trained network, you can use it for making predictions. This is typically called **inference**, a term borrowed from statistics. However, neural networks have a tendency to perform *too well* on the training data and aren't able to generalize to data that hasn't been seen before. This is called **overfitting** and it impairs inference performance. To test for overfitting while training, we measure the performance on data not in the training set called the **validation** dataset. We avoid overfitting through regularization such as dropout while monitoring the validation performance during training. In this notebook, I'll show you how to do this in PyTorch.
#
# First off, I'll implement my own feedforward network for the exercise you worked on in part 4 using the Fashion-MNIST dataset.
#
# As usual, let's start by loading the dataset through torchvision. You'll learn more about torchvision and loading data in a later part.
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import numpy as np
import time
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms
import helper
# +
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
#transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
transforms.Normalize((0.5,), (0.5,))])
# Download and load the training data
trainset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
# Download and load the test data
testset = datasets.FashionMNIST('F_MNIST_data/', download=True, train=False, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=64, shuffle=True)
# -
# ## Building the network
#
# As with MNIST, each image in Fashion-MNIST is 28x28 which is a total of 784 pixels, and there are 10 classes. I'm going to get a bit more advanced here, I want to be able to build a network with an arbitrary number of hidden layers. That is, I want to pass in a parameter like `hidden_layers = [512, 256, 128]` and the network is contructed with three hidden layers have 512, 256, and 128 units respectively. To do this, I'll use `nn.ModuleList` to allow for an arbitrary number of hidden layers. Using `nn.ModuleList` works pretty much the same as a normal Python list, except that it registers each hidden layer `Linear` module properly so the model is aware of the layers.
#
# The issue here is I need a way to define each `nn.Linear` module with the appropriate layer sizes. Since each `nn.Linear` operation needs an input size and an output size, I need something that looks like this:
#
# ```python
# # Create ModuleList and add input layer
# hidden_layers = nn.ModuleList([nn.Linear(input_size, hidden_layers[0])])
# # Add hidden layers to the ModuleList
# hidden_layers.extend([nn.Linear(h1, h2) for h1, h2 in layer_sizes])
# ```
#
# Getting these pairs of input and output sizes can be done with a handy trick using `zip`.
#
# ```python
# hidden_layers = [512, 256, 128, 64]
# layer_sizes = zip(hidden_layers[:-1], hidden_layers[1:])
# for each in layer_sizes:
# print(each)
#
# >> (512, 256)
# >> (256, 128)
# >> (128, 64)
# ```
#
# I also have the `forward` method returning the log-softmax for the output. Since softmax is a probability distibution over the classes, the log-softmax is a log probability which comes with a [lot of benefits](https://en.wikipedia.org/wiki/Log_probability). Using the log probability, computations are often faster and more accurate. To get the class probabilities later, I'll need to take the exponential (`torch.exp`) of the output. Algebra refresher... the exponential function is the inverse of the log function:
#
# $$ \large{e^{\ln{x}} = x }$$
#
# We can include dropout in our network with [`nn.Dropout`](http://pytorch.org/docs/master/nn.html#dropout). This works similar to other modules such as `nn.Linear`. It also takes the dropout probability as an input which we can pass as an input to the network.
# Generalizable network: arbitrary number of hidden layers with their sizes passed
# Additionally, drop out included
# Output: Innstead of logits/relus, log_softmax transformation
# Log_softmax = ln(softmax): larger numbers (more stable), faster (sum of logs is product of probs)
class Network(nn.Module):
def __init__(self, input_size, output_size, hidden_layers, drop_p=0.5):
''' Builds a feedforward network with arbitrary hidden layers.
Arguments
---------
input_size: integer, size of the input
output_size: integer, size of the output layer
hidden_layers: list of integers, the sizes of the hidden layers
drop_p: float between 0 and 1, dropout probability
'''
super().__init__()
# Add the first layer, input to a hidden layer
self.hidden_layers = nn.ModuleList([nn.Linear(input_size, hidden_layers[0])])
# Add a variable number of more hidden layers
layer_sizes = zip(hidden_layers[:-1], hidden_layers[1:])
self.hidden_layers.extend([nn.Linear(h1, h2) for h1, h2 in layer_sizes])
self.output = nn.Linear(hidden_layers[-1], output_size)
self.dropout = nn.Dropout(p=drop_p)
def forward(self, x):
''' Forward pass through the network, returns the output logits '''
# Forward through each layer in `hidden_layers`, with ReLU activation and dropout
for linear in self.hidden_layers:
x = F.relu(linear(x))
# Dropout is done here, after llinear/fc pass. Why?
x = self.dropout(x)
x = self.output(x)
return F.log_softmax(x, dim=1)
# # Train the network
#
# Since the model's forward method returns the log-softmax, I used the [negative log loss](http://pytorch.org/docs/master/nn.html#nllloss) as my criterion, `nn.NLLLoss()`. I also chose to use the [Adam optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Adam). This is a variant of stochastic gradient descent which includes momentum and in general trains faster than your basic SGD.
#
# I've also included a block to measure the validation loss and accuracy. Since I'm using dropout in the network, I need to turn it off during inference. Otherwise, the network will appear to perform poorly because many of the connections are turned off. PyTorch allows you to set a model in "training" or "evaluation" modes with `model.train()` and `model.eval()`, respectively. In training mode, dropout is turned on, while in evaluation mode, dropout is turned off. This effects other modules as well that should be on during training but off during inference.
#
# The validation code consists of a forward pass through the validation set (also split into batches). With the log-softmax output, I calculate the loss on the validation set, as well as the prediction accuracy.
# Create the network, define the criterion and optimizer
# Network hidden layers can be chosen by user now
model = Network(784, 10, [516, 256], drop_p=0.5)
# Negative log likelihood loss (NLLL) = Cross-Entropy for log_softmax
criterion = nn.NLLLoss()
# Adam = SGD with momentum
optimizer = optim.Adam(model.parameters(), lr=0.001)
# Implement a function for the validation pass
def validation(model, testloader, criterion):
test_loss = 0
accuracy = 0
# Note: validation is done for the WHOLE test split!
for images, labels in testloader:
# 64 x 784
images.resize_(images.shape[0], images.shape[2]*images.shape[3])
output = model.forward(images)
test_loss += criterion(output, labels).item()
# log_softmax -> prob: exp(ln(p)) = p
ps = torch.exp(output)
# ps is 64x10: batch_size x num_classes -> select max prob (= best class)
# compare max prob class to ground truth label
equality = (labels.data == ps.max(dim=1)[1])
# tensor type needs to be changed to apply mean()w
accuracy += equality.type(torch.FloatTensor).mean()
# Watch out: test_loss & accuracy need to be divided by len(testloader)
return test_loss, accuracy
epochs = 2
steps = 0
running_loss = 0
print_every = 40
for e in range(epochs):
# train() mode does dropout, eval() mode doesn't
model.train()
for images, labels in trainloader:
steps += 1
# Flatten images into a 784 long vector
images.resize_(images.size()[0], 784)
optimizer.zero_grad()
output = model.forward(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
if steps % print_every == 0:
# Make sure network is in eval mode for inference (no dropout)
model.eval()
# Turn off gradients for validation, saves memory and computations
with torch.no_grad():
test_loss, accuracy = validation(model, testloader, criterion)
print("Epoch: {}/{}.. ".format(e+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/print_every),
"Test Loss: {:.3f}.. ".format(test_loss/len(testloader)),
"Test Accuracy: {:.3f}".format(accuracy/len(testloader)))
running_loss = 0
# Make sure training is back on (activate dropout)
model.train()
# ## Inference
#
# Now that the model is trained, we can use it for inference. We've done this before, but now we need to remember to set the model in inference mode with `model.eval()`. You'll also want to turn off autograd with the `torch.no_grad()` context.
# +
# Test out your network!
# Deactivate dropout
model.eval()
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
# Convert 2D image to 1D vector
img = img.view(1, 784)
# Calculate the class probabilities (softmax) for img
with torch.no_grad():
output = model.forward(img)
ps = torch.exp(output)
# Plot the image and probabilities
helper.view_classify(img.view(1, 28, 28), ps, version='Fashion')
# -
# ## Next Up!
#
# In the next part, I'll show you how to save your trained models. In general, you won't want to train a model everytime you need it. Instead, you'll train once, save it, then load the model when you want to train more or use if for inference.
| Part 5 - Inference and Validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#############################################
#
# Test Case 1
#
#############################################
# import your libs
import hashlib
import base64
# get the urn
specification = "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme"
schema = "BPC01"
party_id = "bpcBusid01"
urn = specification + ":" + schema + "::" + party_id
urn_dict =
# urn = "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme:BPC01::bpcBusid01"
print(urn)
# make sure it's converted to lower case
lower_case_urn = urn.lower()
print(lower_case_urn)
# has to be a byte-like object to be hashed, so encode it as utf-8
urn_encoded = lower_case_urn.encode('utf-8')
print (urn_encoded)
# now create the sha256 hash of it
sha256_urn = hashlib.sha256(urn_encoded)
# print(sha256_urn) # this will be an object
# convert to human readable formats
sha256_digest = sha256_urn.digest()
print(sha256_digest)
#encode into b32
b32_urn = base64.b32encode(sha256_digest)
print(b32_urn)
# strip off the equals sign....
b32_urn_clean = b32_urn.rstrip(b"=")
print(b32_urn_clean)
# convert it back to string in case you want to do anything with it.
b32_str = b32_urn_clean.decode('utf-8')
# make sure it's in lower case again.
final = b32_str.lower()
# This should be your final answer
print(final)
#############################################
#
# Test Case 2
#
#############################################
# import your libs
import hashlib
import base64
# get the urn
specification = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523"
party_id = "0123456789"
urn = specification + ":" + schema + "::" + party_id
# urn = "urn:oasis:names:tc:ebcore:partyid-type:iso6523::0123456789"
print(urn)
# make sure it's converted to lower case
lower_case_urn = urn.lower()
print(lower_case_urn)
# has to be a byte-like object to be hashed, so encode it as utf-8
urn_encoded = lower_case_urn.encode('utf-8')
print (urn_encoded)
# now create the sha256 hash of it
sha256_urn = hashlib.sha256(urn_encoded)
# print(sha256_urn) # this will be an object
# convert to human readable formats
sha256_digest = sha256_urn.digest()
print(sha256_digest)
#encode into b32
b32_urn = base64.b32encode(sha256_digest)
print(b32_urn)
# strip off the equals sign....
b32_urn_clean = b32_urn.rstrip(b"=")
print(b32_urn_clean)
# convert it back to string in case you want to do anything with it.
b32_str = b32_urn_clean.decode('utf-8')
# make sure it's in lower case again.
final = b32_str.lower()
# This should be your final answer
print(final)
#############################################
#
# Test Case 3
#
#############################################
# import your libs
import hashlib
import base64
# get the urn
specification = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "EAN-7638725972413"
urn = specification + ":" + schema + "::" + party_id
# urn = "urn:oasis:names:tc:ebcore:partyid-type:iso6523:0088::EAN-7638725972413"
print(urn)
# make sure it's converted to lower case
lower_case_urn = urn.lower()
print(lower_case_urn)
# has to be a byte-like object to be hashed, so encode it as utf-8
urn_encoded = lower_case_urn.encode('utf-8')
print (urn_encoded)
# now create the sha256 hash of it
sha256_urn = hashlib.sha256(urn_encoded)
# print(sha256_urn) # this will be an object
# convert to human readable formats
sha256_digest = sha256_urn.digest()
print(sha256_digest)
#encode into b32
b32_urn = base64.b32encode(sha256_digest)
print(b32_urn)
# strip off the equals sign....
b32_urn_clean = b32_urn.rstrip(b"=")
print(b32_urn_clean)
# convert it back to string in case you want to do anything with it.
b32_str = b32_urn_clean.decode('utf-8')
# make sure it's in lower case again.
final = b32_str.lower()
# This should be your final answer
print(final)
#############################################
#
# Test Case 4
#
#############################################
# import your libs
import hashlib
import base64
# get the urn
specification = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "bpc-2343030383"
urn = specification + ":" + schema + "::" + party_id
# urn = "urn:oasis:names:tc:ebcore:partyid-type:iso6523::bpc-2343030383"
print(urn)
# make sure it's converted to lower case
lower_case_urn = urn.lower()
print(lower_case_urn)
# has to be a byte-like object to be hashed, so encode it as utf-8
urn_encoded = lower_case_urn.encode('utf-8')
print (urn_encoded)
# now create the sha256 hash of it
sha256_urn = hashlib.sha256(urn_encoded)
# print(sha256_urn) # this will be an object
# convert to human readable formats
sha256_digest = sha256_urn.digest()
print(sha256_digest)
#encode into b32
b32_urn = base64.b32encode(sha256_digest)
print(b32_urn)
# strip off the equals sign....
b32_urn_clean = b32_urn.rstrip(b"=")
print(b32_urn_clean)
# convert it back to string in case you want to do anything with it.
b32_str = b32_urn_clean.decode('utf-8')
# make sure it's in lower case again.
final = b32_str.lower()
# This should be your final answer
print(final)
#############################################
#
# Test Case 5
#
#############################################
# import your libs
import hashlib
import base64
# get the urn
specification = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "4035811991021"
urn = specification + ":" + schema + "::" + party_id
# urn = "urn:oasis:names:tc:ebcore:partyid-type:iso6523:0088:4035811991021"
print(urn)
# make sure it's converted to lower case
lower_case_urn = urn.lower()
print(lower_case_urn)
# has to be a byte-like object to be hashed, so encode it as utf-8
urn_encoded = lower_case_urn.encode('utf-8')
print (urn_encoded)
# now create the sha256 hash of it
sha256_urn = hashlib.sha256(urn_encoded)
# print(sha256_urn) # this will be an object
# convert to human readable formats
sha256_digest = sha256_urn.digest()
print(sha256_digest)
#encode into b32
b32_urn = base64.b32encode(sha256_digest)
print(b32_urn)
# strip off the equals sign....
b32_urn_clean = b32_urn.rstrip(b"=")
print(b32_urn_clean)
# convert it back to string in case you want to do anything with it.
b32_str = b32_urn_clean.decode('utf-8')
# make sure it's in lower case again.
final = b32_str.lower()
# This should be your final answer
print(final)
hashlib.sha256(b'urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme:bpc01::bpcbusid01')
hashlib.sha256(b'urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme:bpc01::bpcbusid01').digest()
base64.b32encode(b'\xc3{4\xfc3"\xdb\xc1u\xdcd\xe8\xbf\xe2\xad\x86\xdfjxob\x1e\'\x17\x8f\xb0\x83!\xec\x15\xab~')
# +
# #!/usr/bin/env python3
#
# File: app_logging.py
# About: Logging provider
# Development: <NAME>, <NAME>
# Date: 2021-07-16 (July 16th, 2021)
#
"""
A class to standardize log formatting across all application artifacts.
Define common loggers and format to be used across the application.
NOTE: These logs are localized and non-persistent.
If used with a Docker container,
they cease to exist when the container does.
Usage: (not meant to be called directly)
log = create_logger("app_logging")
log.debug("This message will be logged.")
"""
import logging
def create_logger(name):
"""This function creates a logger template for the einvoice package.
This funtion creates a consistant format and location for
all application log files to write to.
"""
print("Create logger with name %s" % name)
logger = logging.getLogger(name)
# It's okay to run INFO in Dev. Turn it down to DEBUG for QA
# and WARN for Prod unless troubleshooting an issue.
logger.setLevel(logging.INFO)
# create file handler which writes to a file.
file_logger = logging.FileHandler("./einvoice_output.log")
file_logger.setLevel(logging.INFO)
# create console handler with a higher log level
console_logger = logging.StreamHandler()
console_logger.setLevel(logging.INFO)
# Create a custom formatter and add it to the handlers
_format = "%(asctime)s - %(levelname)s - %(name)s - %(message)s"
datefmt = "%m/%d/%Y %I:%M:%S %p"
formatter = logging.Formatter(_format, datefmt)
file_logger.setFormatter(formatter)
console_logger.setFormatter(formatter)
# Associate the the handlers to the loggers
logger.addHandler(file_logger)
logger.addHandler(console_logger)
return logger
# +
from dataclasses import dataclass
@dataclass
class Urn:
"""Dataclass which represents the base URN for the SML query.
The base URN to be constructed as a string.
Args:
Attributes:
specification: str
The party ID specification.
schema_id: str
The party ID schema type.
party_id: str
The party ID
urn: str
The urn constructed by default values or passed into the
dataclass when called.
Returns:
Raises:
"""
specification: str
schema_id: str
party_id: str
def urn(self) -> str:
"""Construct string for the party's URN"""
# return str(f"{self.specification}:{self.schema}::{self.party_id}")
return f"{self.specification}:{self.schema}::{self.party_id}"
# +
import hashlib
import base64
from json import dumps
from dataclasses import dataclass
@dataclass
class Urn:
specification: str
schema: str
party_id: str
def urn(self) -> str:
return f"{specification}:{schema}::{party_id}"
class Hasher:
@staticmethod
def hasher(specification, schema, party_id):
urn = Urn(specification, schema, party_id)
final_urn = urn.urn()
final_urn_lower_case = final_urn.lower()
urn_lower_encoded = final_urn_lower_case.encode("utf-8")
urn_sha256_hashed = hashlib.sha256(urn_lower_encoded)
urn_sha256_digest = urn_sha256_hashed.digest()
urn_b32_hash = base64.b32encode(urn_sha256_digest)
urn_b32_cleaned = urn_b32_hash.rstrip(b"=")
lower_case_b32 = urn_b32_cleaned.lower()
final_urn_b32 = lower_case_b32.decode("utf-8")
return {
"prty_id_spec": specification,
"prty_id_schma_type": schema,
"prty_id": party_id,
"final_urn": final_urn_lower_case,
"urn_hash": final_urn_b32,
}
def write_hashes_to_file(urn_dictionary, filename):
json_str = dumps(urn_dictionary.__dict__)
with open(filename, mode="w", encoding=str) as my_file:
my_file.write(json_str)
record1 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme", "schema": "BPC01", "party_id": "bpcBusid01"}
record2 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523", "party_id": "0123456789"}
record3 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "EAN-7638725972413"}
record4 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "bpc-2343030383"}
record5 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "4035811991021"}
my_list = [record1, record2, record3, record4, record5]
for record in my_list:
hasher = Hasher()
my_hash = hasher.hasher(record["spec"], record["schema"], record["party_id"])
print(dumps(my_hash, indent=4))
print(dumps(Hasher.hasher(record1["spec"], record1["schema"], record1["party_id"]), indent=4))
print(dumps(Hasher.hasher(record2["spec"], record2["schema"], record2["party_id"]), indent=4))
print(dumps(Hasher.hasher(record3["spec"], record3["schema"], record3["party_id"]), indent=4))
print(dumps(Hasher.hasher(record4["spec"], record4["schema"], record4["party_id"]), indent=4))
print(dumps(Hasher.hasher(record5["spec"], record5["schema"], record5["party_id"]), indent=4))
# -
# + tags=[]
record1 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme", "schema": "BPC01", "party_id": "bpcBusid01"}
record2 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523", "party_id": "0123456789"}
record3 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "EAN-7638725972413"}
record4 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "bpc-2343030383"}
record5 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "4035811991021"}
my_list = [record1, record2, record3, record4, record5]
for record in my_list:
hasher = Hasher()
my_hash = hasher.hasher(record["spec"], record["schema"], record["party_id"])
print(dumps(my_hash, indent=4))
# +
# Example #1
import hashlib
import base64
spec = "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme"
schema = "BPC01"
party_id = "bpcBusid01"
urn = spec + ":" + schema + "::" + party_id
print(urn)
urnl=urn.lower()
print(urnl)
urnle = urnl.encode('utf-8')
print(urnle)
s256 = hashlib.sha256(urnle)
print(s256)
s256d = s256.digest()
print(s256d)
b32 = base64.b32encode(s256d)
print(b32)
b32r = b32.rstrip(b"=")
print(b32r)
b32rd = b32r.decode('utf-8')
print(b32rd)
final = b32rd.lower()
print(final)
# +
# Example #2
import hashlib
import base64
spec = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523"
party_id = "0123456789"
urn = spec + ":" + schema + "::" + party_id
print(urn)
urnl=urn.lower()
print(urnl)
urnle = urnl.encode('utf-8')
print(urnle)
s256 = hashlib.sha256(urnle)
print(s256)
s256d = s256.digest()
print(s256d)
b32 = base64.b32encode(s256d)
print(b32)
b32r = b32.rstrip(b"=")
print(b32r)
b32rd = b32r.decode('utf-8')
print(b32rd)
final = b32rd.lower()
print(final)
# +
# Example #3
import hashlib
import base64
spec = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "EAN-7638725972413"
urn = spec + ":" + schema + "::" + party_id
print(urn)
urnl=urn.lower()
print(urnl)
urnle = urnl.encode('utf-8')
print(urnle)
s256 = hashlib.sha256(urnle)
print(s256)
s256d = s256.digest()
print(s256d)
b32 = base64.b32encode(s256d)
print(b32)
b32r = b32.rstrip(b"=")
print(b32r)
b32rd = b32r.decode('utf-8')
print(b32rd)
final = b32rd.lower()
print(final)
# +
# Example #4
import hashlib
import base64
spec = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "bpc-2343030383"
urn = spec + ":" + schema + "::" + party_id
print(urn)
urnl=urn.lower()
print(urnl)
urnle = urnl.encode('utf-8')
print(urnle)
s256 = hashlib.sha256(urnle)
print(s256)
s256d = s256.digest()
print(s256d)
b32 = base64.b32encode(s256d)
print(b32)
b32r = b32.rstrip(b"=")
print(b32r)
b32rd = b32r.decode('utf-8')
print(b32rd)
final = b32rd.lower()
print(final)
# +
# Example #5
import hashlib
import base64
spec = "urn:oasis:names:tc:ebcore:partyid-type"
schema = "iso6523:0088"
party_id = "4035811991021"
urn = spec + ":" + schema + "::" + party_id
print(urn)
urnl=urn.lower()
print(urnl)
urnle = urnl.encode('utf-8')
print(urnle)
s256 = hashlib.sha256(urnle)
print(s256)
s256d = s256.digest()
print(s256d)
b32 = base64.b32encode(s256d)
print(b32)
b32r = b32.rstrip(b"=")
print(b32r)
b32rd = b32r.decode('utf-8')
print(b32rd)
final = b32rd.lower()
print(final)
# +
import hashlib
import base64
from json import dumps
from dataclasses import dataclass
@dataclass
class Urn:
specification: str
schema: str
party_id: str
def urn(self) -> str:
return f"{specification}:{schema}::{party_id}"
class Hasher:
@staticmethod
def hasher(specification, schema, party_id):
urn = Urn(specification, schema, party_id)
final_urn = urn.urn()
final_urn_lower_case = final_urn.lower()
urn_lower_encoded = final_urn_lower_case.encode("utf-8")
urn_sha256_hashed = hashlib.sha256(urn_lower_encoded)
urn_sha256_digest = urn_sha256_hashed.digest()
urn_b32_hash = base64.b32encode(urn_sha256_digest)
urn_b32_cleaned = urn_b32_hash.rstrip(b"=")
lower_case_b32 = urn_b32_cleaned.lower()
final_urn_b32 = lower_case_b32.decode("utf-8")
return {
"prty_id_spec": specification,
"prty_id_schma_type": schema,
"prty_id": party_id,
"final_urn": final_urn_lower_case,
"urn_hash": final_urn_b32,
}
def write_hashes_to_file(urn_dictionary, filename):
json_str = dumps(urn_dictionary.__dict__)
with open(filename, mode="w", encoding=str) as my_file:
my_file.write(json_str)
record1 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type:unregistered:myscheme", "schema": "BPC01", "party_id": "bpcBusid01"}
# record2 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523", "party_id": "0123456789"}
# record3 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "EAN-7638725972413"}
# record4 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "bpc-2343030383"}
# record5 = {"spec": "urn:oasis:names:tc:ebcore:partyid-type", "schema": "iso6523:0088", "party_id": "4035811991021"}
print(dumps(Hasher.hasher(record1["spec"], record1["schema"], record1["party_id"]), indent=4))
# print(dumps(Hasher.hasher(record2["spec"], record2["schema"], record2["party_id"]), indent=4))
# print(dumps(Hasher.hasher(record3["spec"], record3["schema"], record3["party_id"]), indent=4))
# print(dumps(Hasher.hasher(record4["spec"], record4["schema"], record4["party_id"]), indent=4))
# print(dumps(Hasher.hasher(record5["spec"], record5["schema"], record5["party_id"]), indent=4))
# -
jc4swjyiphrll4gfhlu2edehpwlkmqmsncc2lc3so7m5jvgjkewa
| einvoice/docs/jupyterlab/urn_hash_work.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a name="loesung02"></a>Solutions exercise 02
# ===
# 1.a
sentence = ['I', 'want', 'to', 'go', 'home']
# +
# 1.b
new_string = ''
for word in sentence:
new_string = new_string + word
print(new_string)
# alternatively (nicer):
new_string = ''
for word in sentence:
new_string = new_string + word + ' '
print(new_string)
# +
# 1.c
numbers = [1,2,3,4,5,6,7,8,9,10]
# even numbers
print(numbers[1::2])
# uneven numbers
print(numbers[::2])
# the first three numbers
print(numbers[:3])
# the last three numbers
print(numbers[-3:])
# -
# 1.d
my_list = [6, 9, 1, 2, 15]
new_list = []
for number in my_list:
new_list.append(number + sum(my_list))
print(new_list)
# 1.e
new_list[1::2] = [0] * int(len(new_list)/2)
print(new_list)
| 02-solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os, sys
import pandas as pd
sys.path.append('..')
missing_values = ["?"]
data_file = "sales_train.csv"
df = pd.read_csv("../data/"+data_file, na_values = missing_values)
df.head()
import explore as ex
ex.get_type_dict(df)
import transform as tr
import utility as ut
(x, y, features) = ut.extract_feature_target(df, 'CL_Street_Type', todf=False)
x
y[:5]
# ### Value Remover
vr = tr.ValueRemover(['CL_State'], ['VIC'])
sdf = vr.fit_transform(df)
sdf.head()
ex.count_levels(sdf, 'CL_State')
# ### Date Extractor
from imp import reload
reload(tr)
dx = tr.DateExtractor(['CL_Transfer_Date'], "%d/%m/%Y")
sdf = dx.fit_transform(df)
sdf.head()
# ### Imputer
ex.count_missing(df)
t = tr.ConstantImputer(numerical_columns=['Cl_Est_Land_Area'])
sdf = t.fit_transform(df)
sdf.head()
ex.count_missing(df)['Cl_Materials_In_Roof']
df['Cl_Materials_In_Roof']
t = tr.ConstantImputer(categorical_columns=['Cl_Materials_In_Roof'])
sdf = t.fit_transform(df)
sdf['Cl_Materials_In_Roof']
ex.count_missing(sdf)['Cl_Materials_In_Roof']
ex.get_column_type(df)
reload(tr)
t = tr.ConstantImputer()
sdf = t.fit_transform(df)
ex.count_missing(sdf)
# ### DataFrame Imputer
reload(tr)
t = tr.DataFrameImputer()
sdf = t.fit_transform(df)
sdf.head()
reload(ex)
ex.any_missing(df)
ex.any_missing(sdf)
ex.get_missing_columns(df)
ex.get_missing_columns(sdf)
# ### Column Remover
t = tr.ColumnRemover( ['CL_Street_Direction'])
sdf = t.fit_transform(df)
sdf.shape
df.shape
# ### Create data frame
t =tr.CreateDataFrame(['a', 'b'])
t.fit_transform([[1,2]])
# ### Encoder
reload(tr)
t = tr.Encoder(todf=False)
t.fit_transform(df[['CL_Suburb']])
t = tr.Encoder(todf=True)
t.fit_transform(df[['CL_Suburb']])
df[['CL_Street_Type']]
t = tr.ConstantImputer(categorical_columns=['CL_Street_Type'])
sdf = t.fit_transform(df)
# ## FIXME: mixed type
# mixed type...
ex.get_distinct_value(df, cols=['CL_Street_Type'])
ex.get_distinct_value(sdf, cols=['CL_Street_Type'])
t = tr.Encoder()
t.fit_transform(sdf[['CL_Street_Type']])
ex.get_distinct_value(df, cols=['CL_Street_Type', 'Cl_Toilets'])
ex.get_missing_columns(df)
t.fit_transform(df[['Cl_Toilets']])
t.fit_transform(df[['CL_State', 'CL_Suburb']])
t.fit_dict
# ### TypeSelector
reload(tr)
t = tr.TypeSelector("categorical", todf=True)
sdf = t.fit_transform(df)
sdf.head()
ex.get_column_type(sdf)
# ### Function Transformer
reload(tr)
t = tr.FunctionTransformer(ex.get_distinct_value)
t.fit_transform(sdf)
t = tr.FunctionTransformer(ex.get_distinct_value, 'Cl_Toilets')
t.fit_transform(sdf)
t = tr.FunctionTransformer(ex.get_distinct_value, 'Cl_Toilets', 'Cl_Scenic_View')
t.fit_transform(sdf)
# ### Transform Debugger
reload(tr)
reload(ut)
imputer = tr.ConstantImputer()
imputer.fit_transform(df)
len(df)
import random
random.randrange(0, 1000)
t = tr.TransformDebugger(imputer, ut.show_row)
t.fit_transform(df)
from sklearn.pipeline import Pipeline
# ### Pipleline
pipeline = Pipeline([
('Value Remover', tr.ValueRemover(['sex'], ['F'])),
])
sdf = pipeline.fit_transform(df)
sdf.shape
from imp import reload
ex.count_levels(sdf)
ex.get_distinct_value(sdf, 'sex')
| notebook/test_transformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating RESTful APIs using Flask and Python
# ## Medium Article Link: <https://medium.com/p/655bad51b24>
# ## Minimal Flask App
# ```python
# from flask import Flask
#
# app = Flask(__name__)
#
# @app.route('/hello/', methods=['GET', 'POST'])
# def welcome():
# return "Hello World!"
#
# if __name__ == '__main__':
# app.run(host='0.0.0.0', port=105)
# ```
# ## Variable Rules
# ```python
# from flask import Flask
#
# app = Flask(__name__)
#
# @app.route('/<int:number>/')
# def incrementer(number):
# return "Incremented number is " + str(number+1)
#
# @app.route('/<string:name>/')
# def hello(name):
# return "Hello " + name
#
# app.run()
# ```
# ## Return JSON Serializable Output
# ```python
# from flask import jsonify
#
# @app.route('/person/')
# def hello():
# return jsonify({'name':'Jimit',
# 'address':'India'})
# ```
# ```python
# from flask import jsonify
#
# @app.route('/numbers/')
# def print_list():
# return jsonify(list(range(5)))
# ```
# ## Redirection Behaviour
# ```python
# @app.route('/home/')
# def home():
# return "Home page"
#
# @app.route('/contact')
# def contact():
# return "Contact page"
# ```
# ## Return Status Code
# ```python
# @app.route('/teapot/')
# def teapot():
# return "Would you like some tea?", 418
# ```
# ## Before Request
# ```python
# @app.before_request
# def before():
# print("This is executed BEFORE each request.")
#
# @app.route('/hello/')
# def hello():
# return "Hello World!"
# ```
# ## Blueprints
# ### `home.py`
# ```python
# from flask import Blueprint
#
# home_bp = Blueprint('home', __name__)
#
# @home_bp.route('/hello/')
# def hello():
# return "Hello from Home Page"
# ```
# ### `contact.py`
# ```python
# from flask import Blueprint
#
# contact_bp = Blueprint('contact', __name__)
#
# @contact_bp.route('/hello/')
# def hello():
# return "Hello from Contact Page"
# ```
# ### `app.py`
# ```python
# from flask import Flask
#
# from home import home_bp
# from contact import contact_bp
#
# app = Flask(__name__)
#
# app.register_blueprint(home_bp, url_prefix='/home')
# app.register_blueprint(contact_bp, url_prefix='/contact')
#
# app.run()
# ```
# ## Logging
# ```python
# app.logger.debug('This is a DEBUG message')
# app.logger.info('This is an INFO message')
# app.logger.warning('This is a WARNING message')
# app.logger.error('This is an ERROR message')
# ```
| Creating RESTful APIs using Flask and Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 随机森林
# --------------
# **集成学习**(ensemble learning)通过构建并结合多个学习器来完成学习任务。欲得到泛化性能强的集成,集成中的个体学习器应尽可能相互独立。给定一个训练数据集,对训练样本进行采样,产生若干个不相交的子集,再从每个子集中训练出一个基学习器。这样,由于训练数据不同,每个基学习器可获得比较大的差异;但是,如果采样出的每个子集都完全不同,则每个基学习器只用到了一小部分训练数据,每个基学习器都是有偏的。此时可使用互相有重叠的采样子数据集来训练基学习器。
#
# **Bagging**是并行化集成学习方法的典型代表。给定包含有m个样本的数据集,随机取出一个样本放入采样集中,再把该样本放回初始数据集,使得下次采样时,该样本仍有可能被选中,这样经过m次随机采样操作,就得到了含有m个样本的采样集。如此,采样集中只包含初始训练集的一部分,有的多次出现,有的未出现。然后基于每个采样集,训练出一个基学习器,再将这些基学习器结合,就是Bagging的基本流程。
#
# 
#
# **随机森林**(Random Forest)是Bagging的一个扩张变体。RF在以决策树为基学习器构建Bagging集成的基础上,进一步在决策树的训练过程中引入随机属性选择。传统决策树在选择划分属性时是在当前结点的属性集合(假定有d个属性)中选择一个最优属性;而在RF中,对基决策树的每个结点,先从该结点的属性集合中随机选择一个包含k个属性的子集,然后再从这个子集中选择一个最优属性用来划分。k控制了随机性的引入程度,若$k=d$则基决策树的构建与传统决策树相同,$k=1$,则是随机选择一个属性用于划分,一般情况下,推荐$k=log_2d$。
#
# 随机森林简单、容易实现、计算开销小,在很多现实任务中展现出强大的性能,被誉为"*代表集成学习技术水平的方法*"。随机森林对Bagging只做了小改动,但是与Bagging中,基学习器的*多样性*仅通过样本扰动(通过对初始训练集采样)来实现,而在随机森林中,基学习器的多样性不仅有来自样本扰动,还有来自属性扰动。这就使得最终集成的泛化性能可通过个体学习器之间差异度的增加而进一步提升。
#
# 随机森林的收敛性与Bagging相似,起始性能往往相对较差,特别是在集成中只包含一个基学习器时。然而,随着基学习器数目的增加,随机森林通常会收敛到更低的泛化误差。由于随机森林使用的*随机型*决策树,在选择划分属性时,只需要考察一个属性子集,不像Bagging需要对所有属性进行考察,所以,它的训练效率常常优于Bagging。
#
# 
# sklearn随机森林分类器
# ---------------------------------
# ```python
# RandomForestClassifier(n_estimators=10, # 基学习器数量
# criterion='gini', # 属性选择的标准(分裂树的标准)
# max_depth=None, # 树的最大深度
# min_samples_split=2, # 最小分裂样本数
# min_samples_leaf=1, # 最小叶子节点样本数
# # 最小叶子节点权重
# min_weight_fraction_leaf=0.0,
# # 查找最佳分裂所需考虑的特征数,
# # auto:sqrt(n_features),None:n_features
# max_features='auto',
# max_leaf_nodes=None, # 最大叶子节点数
# min_impurity_split=1e-07, # 分裂的最小不纯度
# bootstrap=True,
# oob_score=False, # 是否使用袋外样本估计准确度
# n_jobs=1, # 并行job数,-1 代表全部
# andom_state=None,
# verbose=0,
# warm_start=False,
# class_weight=None)
# ```
# 在随机森林算法中可以发现Bootstrap每次约有$\frac{1}{3}$的样本不会出现在Bootstrap所采集的样本集合中,当然也就没有参加决策树的建立,这$\frac{1}{3}$的数据称为袋外数据oob(out-of-bag)。它可以用于取代测试集误差估计方法。
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.datasets import load_iris
from sklearn.ensemble import RandomForestClassifier
iris = load_iris() # 鸢尾花卉数据集
print(iris.keys())
print("feature_names:", iris.feature_names)
df = pd.DataFrame(iris.data, columns=iris.feature_names)
print(df.shape)
df['is_train'] = np.random.uniform(0, 1, len(df)) <= .75
df['species'] = pd.Categorical.from_codes(iris.target, iris.target_names)
print(df['species'][::10])
df.head()
train, test = df[df['is_train']==True], df[df['is_train']==False]
features = df.columns[:4]
print(train[features][:1])
clf = RandomForestClassifier(n_jobs=2)
y, _ = pd.factorize(train['species']) # 返回正确分类的索引
clf.fit(train[features], y)
print(iris.target_names)
preds = iris.target_names[clf.predict(test[features])]
a = np.array(["foo", "foo", "foo", "foo",
"bar", "bar", "bar", "bar",
"foo", "foo", "foo", "bar"], dtype=object)
b = np.array(["one", "one", "one", "two",
"one", "one", "one", "two",
"two", "two", "one", "one"], dtype=object)
pd.crosstab(a, b, rownames=['a'], colnames=['b']) # 求共现矩阵
pd.crosstab(test['species'], preds, rownames=['actual'], colnames=['preds'])
# +
from sklearn.cross_validation import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import make_moons, make_circles, make_classification
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis as QDA
names = ["Nearest Neighbors",
"Linear SVM",
"RBF SVM",
"Decision Tree",
"Random Forest",
"AdaBoost",
"Naive Bayes",
"LDA",
"QDA"]
classifiers = [KNeighborsClassifier(3),
SVC(kernel="linear", C=0.025),
SVC(gamma=2, C=1),
DecisionTreeClassifier(max_depth=5),
RandomForestClassifier(max_depth=5, n_estimators=10, max_features=1),
AdaBoostClassifier(),
GaussianNB(),
LDA(),
QDA()]
# -
X, y = make_classification(n_samples=100, # 生成样本数量,默认100
n_features=2, # 特征数量
n_redundant=0, # 冗余特征数量
n_informative=2, # 多信息特征数
random_state=1, # 随机数种子
n_classes=2, # 类别数量
n_clusters_per_class=1)
# +
rng = np.random.RandomState(2)
X += 2 * rng.uniform(size=X.shape)
linearly_separable = (X, y)
datasets = [make_moons(noise=0.3, random_state=0), # 生成半环形数据
make_circles(noise=0.2, factor=0.5, random_state=1), # 生成环形数据
linearly_separable]
# -
h = .02 # step size in the mesh
figure = plt.figure(figsize=(27, 9))
i = 1
# iterate over datasets
for ds in datasets:
# preprocess dataset, split into training and test part
X, y = ds
X = StandardScaler().fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.4)
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# just plot the dataset first
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright)
# and testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
i += 1
# iterate over classifiers
for name, clf in zip(names, classifiers):
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test) # 得到测试集分数(准确率)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max] x [y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
ax.contourf(xx, yy, Z, cmap=cm, alpha=.8)
# Plot also the training points
ax.scatter(X_train[:, 0],
X_train[:, 1],
c=y_train,
cmap=cm_bright)
# and testing points
ax.scatter(X_test[:, 0],
X_test[:, 1],
c=y_test,
cmap=cm_bright,
alpha=0.6)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
ax.set_title(name)
ax.text(xx.max() - .3, yy.min() + .3,
('%.2f' % score).lstrip('0'),
size=15, horizontalalignment='right')
i += 1
figure.subplots_adjust(left=.02, right=.98)
plt.show()
| ML/RandomForest/RandomForest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# name: python3
# ---
# # Data Cleaning:
# - html.parser remover
# - stemmer
# - stopword remove
#
from cleaner import cleaner
from stopword import removing_stopwords
from porter_stemmer_lemmatization import porter_stemmer
import pandas as pd
text1 = '<h1>Hello world</h1> ,my, name is hero and he is god, they said to <br />help<br /> me. movie.<br /><br />Cut <br /><br /><br /><br />'
print(text1)
text = cleaner(text1)
print(text)
text = removing_stopwords(text)
print(text)
text = porter_stemmer(text)
print(text)
# data_train = pd.read_csv('../dataset/labeledTrainData.tsv',sep='\t')
data_train = pd.read_csv('../dataset/movie_train.csv')
test = data_train.sample(22).head(1)
def clean_data(data):
new_data = cleaner(data)
new_data = removing_stopwords(new_data)
new_data = porter_stemmer(new_data)
return new_data
print(test.review.to_list()[0])
test['review'] = test['review'].apply(clean_data)
print('-----------------')
print(test.review.to_list()[0])
# # Cleaning data and store
data_train['review'] = data_train['review'].apply(clean_data)
data_train.to_csv('../dataset/cleaned_movie_train.csv',index=False)
# # Easy Method
# +
import string
def text_process(text):
#this func remove all punctuations and stopwords and finnaly returned cleaned text as list of words
nopunc = [char for char in text if char not in string.punctuation]
nopunc = ''.join(nopunc)
print(nopunc)
nopunc = removing_stopwords(nopunc)
return nopunc
text_process(text1)
| preprocesser/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
#creating a list
marks=[1,2,3,4,5,6,7,8,9,10]
# -
marks
marks[5]
marks[0:6]
# +
#adding an element
marks.append(11)
# -
marks
marks.extend([12,13])
marks
marks.append([14,15])
marks
# +
#deleting elements
marks.remove([14,15])
# -
marks
del marks[0]
marks
# +
#accessing list elements
for mark in marks:
print(mark)
# -
for mark in marks:
print(mark+1)
sam =[45, 36, 54, 78, 44]
| 23. List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
def func(x):
a = np.e**(-2*x)*np.cos(10*x)
return a
def func_integral(x):
a = -1/52.*np.e**(-2*x)*(np.cos(10*x)-(5*np.sin(10*x)))
return a
def trapezoid_core(f,x,h):
return 0.5*h*(f(x+h)+f(x))
def trapezoid_method(f,a,b,N):
x = np.linspace(0,np.pi,N)
h = x[1]-x[0]
Fint = 0.0
for i in range (0,len(x)-1,1):
Fint += trapezoid_core(f,x[i],h)
return Fint
def simpson_core(f,x,h):
return h*( f(x) + 4*f(x+h) + f(x+2*h))/3.
def simpsons_method(f,a,b,N):
x = np.linspace(0,np.pi,N)
h = x[1]-x[0]
Fint = 0.0
for i in range(0,len(x)-2,2):
Fint += simpson_core(f,x[i],h)
if ((N%2)==0):
Fint += simpson_core(f,x[-2],0.5*h)
return Fint
def romberg_core(f,a,b,i):
h = np.pi-0
dh = h/2.**(i+1)
K = h/2.**(i)
M = 0.0
for j in range(2**i):
M+=f(0+0.5*dh + j*dh)
return K*M
def romberg_integration(f,a,b,tol):
i = 0
imax = 1000
delta = 1000.0*np.fabs(tol)
I = np.zeros(imax,dtype=float)
I[0] = 0.5*(np.pi-0)*(f(np.pi)+f(0))
i += 1
while(delta>tol):
I[i] = 0.5*I[i-1] + romberg_core(f,0,np.pi,i)
delta = np.fabs( (I[i]-I[i-1])/I[i] )
print(i,I[i],I[i-1],delta)
if (delta>tol):
i+=1
if(i>imax):
print("Max iterations reached.")
raise StopIteration('Stopping iterations after',i)
return I[i]
Answer = func_integral(np.pi)-func_integral(0)
print(Answer)
print("Trapezoid")
print(trapezoid_method(func,0,np.pi,100))
print("Simpson's Method")
print(simpsons_method(func,0,np.pi,100))
print("Romberg")
tolerance = 1.0e-6
RI = romberg_integration(func,0,np.pi,tolerance)
print(RI, (RI-Answer)/Answer, tolerance)
# Romberg took 20 iterations
# Trapezoid took 50 intervals
# Simpsons took 50 intervals
| hw-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from skimage.feature import hog
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import GaussianNB
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score, roc_curve
from sklearn.preprocessing import normalize
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.wrappers.scikit_learn import KerasClassifier
from skimage.morphology import binary_erosion, binary_dilation, binary_closing,skeletonize, thin,erosion,dilation
import pickle
from commonfunctions import *
image_size = 28 # width and length
classes_n = 10 # i.e. 0, 1, 2, 3, ..., 9
image_pixels = image_size * image_size
data_path = "data/"
train_data = np.loadtxt(data_path + "train.csv",
delimiter=",")
test_data = np.loadtxt(data_path + "test.csv",
delimiter=",")
test_data[:10]
Xtrain = train_data[:,1:]
Xtest = test_data[:,1:]
Ytrain = np.asfarray(train_data[:, :1])
Ytest = np.asfarray(test_data[:, :1])
label_encoding = np.arange(10)
Ytrain_oh = (label_encoding == Ytrain).astype(np.int)
Ytest_oh = (label_encoding == Ytest).astype(np.int)
# pickle data SAVEING CODE
with open("data/pickled_mnist.pkl", "bw") as fh:
data = (Xtrain,
Xtest,
Ytrain,
Ytest,
Ytrain_oh,
Ytest_oh)
pickle.dump(data, fh)
# +
# LOADING CODE
with open("data/pickled_mnist.pkl", "br") as fh:
data = pickle.load(fh)
Xtrain = data[0]
Xtest = data[1]
Ytrain = data[2]
Ytest = data[3]
Ytrain_oh = data[4]
Ytest_oh = data[5]
# -
imgs = []
labels = []
for i in range(3):
img = Xtrain[i].reshape((28,28))
labels.append(str(Ytrain[i]))
imgs.append(img)
show_images(imgs,labels)
model = Sequential()
model.add(Dense(100, input_dim=288, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(100, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam', metrics=['accuracy'])
def hog_features(X, imgshape=(28, 28), pixels_per_cell=(6, 6)):
features = []
for row in X:
img = row.reshape(imgshape)
img_feature = hog(img, orientations=8, pixels_per_cell=pixels_per_cell, cells_per_block=(2, 2))
features.append(img_feature)
return np.array(features)
hog_features(Xtrain[1,:].reshape(1,-1))
# +
Xtrain_hog = hog_features(Xtrain)
Xtest_hog = hog_features(Xtest)
X_train_norm = normalize(Xtrain_hog)
X_test_norm = normalize(Xtest_hog)
history = model.fit(X_train_norm, Ytrain_oh,
batch_size=128, epochs=20,
verbose=2)
# -
model.evaluate(X_test_norm,Ytest_oh)
# saving the model
model_name = 'hogmodel.h5'
model.save(model_name)
print('Saved trained model as %s ' % model_name)
# plotting the metrics
fig = plt.figure()
plt.subplot(2,1,1)
plt.plot(history.history['acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
# +
def predict_img(img):
img = rgb2gray(img)
img = (img.reshape(1,-1))
Xhog = hog_features(img)
Xhog = normalize(Xhog)
Y = (model.predict(Xhog))
return (np.argmax(Y))
#hog_features(img.)
tx = io.imread("test4.tif")
predict_img(tx)
# -
filename = 'hognn_model.sav'
pickle.dump(model, open(filename, 'wb'))
# +
hog_model = load_model("model_name")
loss_and_metrics = mnist_model.evaluate(X_test, Y_test, verbose=2)
print("Test Loss", loss_and_metrics[0])
print("Test Accuracy", loss_and_metrics[1])
| GradeAutofiller/DMv2/TRAINING/HOGNN_MODEL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 3: Smart Beta Portfolio and Portfolio Optimization
#
# ## Overview
#
#
# Smart beta has a broad meaning, but we can say in practice that when we use the universe of stocks from an index, and then apply some weighting scheme other than market cap weighting, it can be considered a type of smart beta fund. A Smart Beta portfolio generally gives investors exposure or "beta" to one or more types of market characteristics (or factors) that are believed to predict prices while giving investors a diversified broad exposure to a particular market. Smart Beta portfolios generally target momentum, earnings quality, low volatility, and dividends or some combination. Smart Beta Portfolios are generally rebalanced infrequently and follow relatively simple rules or algorithms that are passively managed. Model changes to these types of funds are also rare requiring prospectus filings with US Security and Exchange Commission in the case of US focused mutual funds or ETFs.. Smart Beta portfolios are generally long-only, they do not short stocks.
#
# In contrast, a purely alpha-focused quantitative fund may use multiple models or algorithms to create a portfolio. The portfolio manager retains discretion in upgrading or changing the types of models and how often to rebalance the portfolio in attempt to maximize performance in comparison to a stock benchmark. Managers may have discretion to short stocks in portfolios.
#
# Imagine you're a portfolio manager, and wish to try out some different portfolio weighting methods.
#
# One way to design portfolio is to look at certain accounting measures (fundamentals) that, based on past trends, indicate stocks that produce better results.
#
#
# For instance, you may start with a hypothesis that dividend-issuing stocks tend to perform better than stocks that do not. This may not always be true of all companies; for instance, Apple does not issue dividends, but has had good historical performance. The hypothesis about dividend-paying stocks may go something like this:
#
# Companies that regularly issue dividends may also be more prudent in allocating their available cash, and may indicate that they are more conscious of prioritizing shareholder interests. For example, a CEO may decide to reinvest cash into pet projects that produce low returns. Or, the CEO may do some analysis, identify that reinvesting within the company produces lower returns compared to a diversified portfolio, and so decide that shareholders would be better served if they were given the cash (in the form of dividends). So according to this hypothesis, dividends may be both a proxy for how the company is doing (in terms of earnings and cash flow), but also a signal that the company acts in the best interest of its shareholders. Of course, it's important to test whether this works in practice.
#
#
# You may also have another hypothesis, with which you wish to design a portfolio that can then be made into an ETF. You may find that investors may wish to invest in passive beta funds, but wish to have less risk exposure (less volatility) in their investments. The goal of having a low volatility fund that still produces returns similar to an index may be appealing to investors who have a shorter investment time horizon, and so are more risk averse.
#
# So the objective of your proposed portfolio is to design a portfolio that closely tracks an index, while also minimizing the portfolio variance. Also, if this portfolio can match the returns of the index with less volatility, then it has a higher risk-adjusted return (same return, lower volatility).
#
# Smart Beta ETFs can be designed with both of these two general methods (among others): alternative weighting and minimum volatility ETF.
#
#
# ## Instructions
# Each problem consists of a function to implement and instructions on how to implement the function. The parts of the function that need to be implemented are marked with a `# TODO` comment. After implementing the function, run the cell to test it against the unit tests we've provided. For each problem, we provide one or more unit tests from our `project_tests` package. These unit tests won't tell you if your answer is correct, but will warn you of any major errors. Your code will be checked for the correct solution when you submit it to Udacity.
#
# ## Packages
# When you implement the functions, you'll only need to you use the packages you've used in the classroom, like [Pandas](https://pandas.pydata.org/) and [Numpy](http://www.numpy.org/). These packages will be imported for you. We recommend you don't add any import statements, otherwise the grader might not be able to run your code.
#
# The other packages that we're importing are `helper`, `project_helper`, and `project_tests`. These are custom packages built to help you solve the problems. The `helper` and `project_helper` module contains utility functions and graph functions. The `project_tests` contains the unit tests for all the problems.
# ### Install Packages
import sys
# !{sys.executable} -m pip install -r requirements.txt
# ### Load Packages
import pandas as pd
import numpy as np
import helper
import project_helper
import project_tests
# ## Market Data
# ### Load Data
# For this universe of stocks, we'll be selecting large dollar volume stocks. We're using this universe, since it is highly __liquid__.
# +
df = pd.read_csv('../../data/project_3/eod-quotemedia.csv')
percent_top_dollar = 0.2
high_volume_symbols = project_helper.large_dollar_volume_stocks(df, 'adj_close', 'adj_volume', percent_top_dollar)
df = df[df['ticker'].isin(high_volume_symbols)]
close = df.reset_index().pivot(index='date', columns='ticker', values='adj_close')
volume = df.reset_index().pivot(index='date', columns='ticker', values='adj_volume')
dividends = df.reset_index().pivot(index='date', columns='ticker', values='dividends')
# -
# ### View Data
# To see what one of these 2-d matrices looks like, let's take a look at the closing prices matrix.
project_helper.print_dataframe(close)
# # Part 1: Smart Beta Portfolio
# In Part 1 of this project, you'll build a portfolio using dividend yield to choose the portfolio weights. A portfolio such as this could be incorporated into a smart beta ETF. You'll compare this portfolio to a market cap weighted index to see how well it performs.
#
# Note that in practice, you'll probably get the index weights from a data vendor (such as companies that create indices, like MSCI, FTSE, Standard and Poor's), but for this exercise we will simulate a market cap weighted index.
#
# ## Index Weights
# The index we'll be using is based on large dollar volume stocks. Implement `generate_dollar_volume_weights` to generate the weights for this index. For each date, generate the weights based on dollar volume traded for that date. For example, assume the following is close prices and volume data:
# ```
# Prices
# A B ...
# 2013-07-08 2 2 ...
# 2013-07-09 5 6 ...
# 2013-07-10 1 2 ...
# 2013-07-11 6 5 ...
# ... ... ... ...
#
# Volume
# A B ...
# 2013-07-08 100 340 ...
# 2013-07-09 240 220 ...
# 2013-07-10 120 500 ...
# 2013-07-11 10 100 ...
# ... ... ... ...
# ```
# The weights created from the function `generate_dollar_volume_weights` should be the following:
# ```
# A B ...
# 2013-07-08 0.126.. 0.194.. ...
# 2013-07-09 0.759.. 0.377.. ...
# 2013-07-10 0.075.. 0.285.. ...
# 2013-07-11 0.037.. 0.142.. ...
# ... ... ... ...
# ```
# +
def generate_dollar_volume_weights(close, volume):
"""
Generate dollar volume weights.
Parameters
----------
close : DataFrame
Close price for each ticker and date
volume : str
Volume for each ticker and date
Returns
-------
dollar_volume_weights : DataFrame
The dollar volume weights for each ticker and date
"""
assert close.index.equals(volume.index)
assert close.columns.equals(volume.columns)
#TODO: Implement function
dollar_vol = close * volume
return dollar_vol.div(dollar_vol.sum(axis=1), axis=0)
project_tests.test_generate_dollar_volume_weights(generate_dollar_volume_weights)
# -
# ### View Data
# Let's generate the index weights using `generate_dollar_volume_weights` and view them using a heatmap.
index_weights = generate_dollar_volume_weights(close, volume)
project_helper.plot_weights(index_weights, 'Index Weights')
# ## Portfolio Weights
# Now that we have the index weights, let's choose the portfolio weights based on dividend. You would normally calculate the weights based on __trailing dividend yield__, but we'll simplify this by just calculating the __total dividend yield__ over time.
#
# Implement `calculate_dividend_weights` to return the weights for each stock based on its total dividend yield over time. This is similar to generating the weight for the index, but it's using dividend data instead.
# For example, assume the following is `dividends` data:
# ```
# Prices
# A B
# 2013-07-08 0 0
# 2013-07-09 0 1
# 2013-07-10 0.5 0
# 2013-07-11 0 0
# 2013-07-12 2 0
# ... ... ...
# ```
# The weights created from the function `calculate_dividend_weights` should be the following:
# ```
# A B
# 2013-07-08 NaN NaN
# 2013-07-09 0 1
# 2013-07-10 0.333.. 0.666..
# 2013-07-11 0.333.. 0.666..
# 2013-07-12 0.714.. 0.285..
# ... ... ...
# ```
# +
def calculate_dividend_weights(dividends):
"""
Calculate dividend weights.
Parameters
----------
ex_dividend : DataFrame
Ex-dividend for each stock and date
Returns
-------
dividend_weights : DataFrame
Weights for each stock and date
"""
#TODO: Implement function
cumsum = dividends.cumsum()
return cumsum.div(cumsum.sum(axis=1), axis=0)
project_tests.test_calculate_dividend_weights(calculate_dividend_weights)
# -
# ### View Data
# Just like the index weights, let's generate the ETF weights and view them using a heatmap.
etf_weights = calculate_dividend_weights(dividends)
project_helper.plot_weights(etf_weights, 'ETF Weights')
# ## Returns
# Implement `generate_returns` to generate returns data for all the stocks and dates from price data. You might notice we're implementing returns and not log returns. Since we're not dealing with volatility, we don't have to use log returns.
# +
def generate_returns(prices):
"""
Generate returns for ticker and date.
Parameters
----------
prices : DataFrame
Price for each ticker and date
Returns
-------
returns : Dataframe
The returns for each ticker and date
"""
#TODO: Implement function
return (prices - prices.shift()) / prices.shift()
project_tests.test_generate_returns(generate_returns)
# -
# ### View Data
# Let's generate the closing returns using `generate_returns` and view them using a heatmap.
returns = generate_returns(close)
project_helper.plot_returns(returns, 'Close Returns')
# ## Weighted Returns
# With the returns of each stock computed, we can use it to compute the returns for an index or ETF. Implement `generate_weighted_returns` to create weighted returns using the returns and weights.
# +
def generate_weighted_returns(returns, weights):
"""
Generate weighted returns.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
weights : DataFrame
Weights for each ticker and date
Returns
-------
weighted_returns : DataFrame
Weighted returns for each ticker and date
"""
assert returns.index.equals(weights.index)
assert returns.columns.equals(weights.columns)
#TODO: Implement function
return returns * weights
project_tests.test_generate_weighted_returns(generate_weighted_returns)
# -
# ### View Data
# Let's generate the ETF and index returns using `generate_weighted_returns` and view them using a heatmap.
index_weighted_returns = generate_weighted_returns(returns, index_weights)
etf_weighted_returns = generate_weighted_returns(returns, etf_weights)
project_helper.plot_returns(index_weighted_returns, 'Index Returns')
project_helper.plot_returns(etf_weighted_returns, 'ETF Returns')
# ## Cumulative Returns
# To compare performance between the ETF and Index, we're going to calculate the tracking error. Before we do that, we first need to calculate the index and ETF comulative returns. Implement `calculate_cumulative_returns` to calculate the cumulative returns over time given the returns.
# +
def calculate_cumulative_returns(returns):
"""
Calculate cumulative returns.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
Returns
-------
cumulative_returns : Pandas Series
Cumulative returns for each date
"""
#TODO: Implement function
# print(returns.T.sum())
# print(returns.sum(axis = 1))
return (returns.T.sum() + 1).cumprod()
project_tests.test_calculate_cumulative_returns(calculate_cumulative_returns)
# -
# ### View Data
# Let's generate the ETF and index cumulative returns using `calculate_cumulative_returns` and compare the two.
index_weighted_cumulative_returns = calculate_cumulative_returns(index_weighted_returns)
etf_weighted_cumulative_returns = calculate_cumulative_returns(etf_weighted_returns)
project_helper.plot_benchmark_returns(index_weighted_cumulative_returns, etf_weighted_cumulative_returns, 'Smart Beta ETF vs Index')
# ## Tracking Error
# In order to check the performance of the smart beta portfolio, we can calculate the annualized tracking error against the index. Implement `tracking_error` to return the tracking error between the ETF and benchmark.
#
# For reference, we'll be using the following annualized tracking error function:
# $$ TE = \sqrt{252} * SampleStdev(r_p - r_b) $$
#
# Where $ r_p $ is the portfolio/ETF returns and $ r_b $ is the benchmark returns.
#
# _Note: When calculating the sample standard deviation, the delta degrees of freedom is 1, which is the also the default value._
# +
def tracking_error(benchmark_returns_by_date, etf_returns_by_date):
"""
Calculate the tracking error.
Parameters
----------
benchmark_returns_by_date : Pandas Series
The benchmark returns for each date
etf_returns_by_date : Pandas Series
The ETF returns for each date
Returns
-------
tracking_error : float
The tracking error
"""
assert benchmark_returns_by_date.index.equals(etf_returns_by_date.index)
#TODO: Implement function
diff = benchmark_returns_by_date - etf_returns_by_date
return np.sqrt(252) * diff.std()
project_tests.test_tracking_error(tracking_error)
# -
# ### View Data
# Let's generate the tracking error using `tracking_error`.
smart_beta_tracking_error = tracking_error(np.sum(index_weighted_returns, 1), np.sum(etf_weighted_returns, 1))
print('Smart Beta Tracking Error: {}'.format(smart_beta_tracking_error))
# # Part 2: Portfolio Optimization
#
# Now, let's create a second portfolio. We'll still reuse the market cap weighted index, but this will be independent of the dividend-weighted portfolio that we created in part 1.
#
# We want to both minimize the portfolio variance and also want to closely track a market cap weighted index. In other words, we're trying to minimize the distance between the weights of our portfolio and the weights of the index.
#
# $Minimize \left [ \sigma^2_p + \lambda \sqrt{\sum_{1}^{m}(weight_i - indexWeight_i)^2} \right ]$ where $m$ is the number of stocks in the portfolio, and $\lambda$ is a scaling factor that you can choose.
#
# Why are we doing this? One way that investors evaluate a fund is by how well it tracks its index. The fund is still expected to deviate from the index within a certain range in order to improve fund performance. A way for a fund to track the performance of its benchmark is by keeping its asset weights similar to the weights of the index. We’d expect that if the fund has the same stocks as the benchmark, and also the same weights for each stock as the benchmark, the fund would yield about the same returns as the benchmark. By minimizing a linear combination of both the portfolio risk and distance between portfolio and benchmark weights, we attempt to balance the desire to minimize portfolio variance with the goal of tracking the index.
#
#
# ## Covariance
# Implement `get_covariance_returns` to calculate the covariance of the `returns`. We'll use this to calculate the portfolio variance.
#
# If we have $m$ stock series, the covariance matrix is an $m \times m$ matrix containing the covariance between each pair of stocks. We can use [`Numpy.cov`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.cov.html) to get the covariance. We give it a 2D array in which each row is a stock series, and each column is an observation at the same period of time. For any `NaN` values, you can replace them with zeros using the [`DataFrame.fillna`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.fillna.html) function.
#
# The covariance matrix $\mathbf{P} =
# \begin{bmatrix}
# \sigma^2_{1,1} & ... & \sigma^2_{1,m} \\
# ... & ... & ...\\
# \sigma_{m,1} & ... & \sigma^2_{m,m} \\
# \end{bmatrix}$
# +
def get_covariance_returns(returns):
"""
Calculate covariance matrices.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
Returns
-------
returns_covariance : 2 dimensional Ndarray
The covariance of the returns
"""
#TODO: Implement function
# print("\nReturns:\n", returns)
# print("\nCovariance:\n", returns.fillna(0).cov())
return returns.fillna(0).cov().values
project_tests.test_get_covariance_returns(get_covariance_returns)
# -
# ### View Data
# Let's look at the covariance generated from `get_covariance_returns`.
# +
covariance_returns = get_covariance_returns(returns)
covariance_returns = pd.DataFrame(covariance_returns, returns.columns, returns.columns)
covariance_returns_correlation = np.linalg.inv(np.diag(np.sqrt(np.diag(covariance_returns))))
covariance_returns_correlation = pd.DataFrame(
covariance_returns_correlation.dot(covariance_returns).dot(covariance_returns_correlation),
covariance_returns.index,
covariance_returns.columns)
project_helper.plot_covariance_returns_correlation(
covariance_returns_correlation,
'Covariance Returns Correlation Matrix')
# -
# ### portfolio variance
# We can write the portfolio variance $\sigma^2_p = \mathbf{x^T} \mathbf{P} \mathbf{x}$
#
# Recall that the $\mathbf{x^T} \mathbf{P} \mathbf{x}$ is called the quadratic form.
# We can use the cvxpy function `quad_form(x,P)` to get the quadratic form.
#
# ### Distance from index weights
# We want portfolio weights that track the index closely. So we want to minimize the distance between them.
# Recall from the Pythagorean theorem that you can get the distance between two points in an x,y plane by adding the square of the x and y distances and taking the square root. Extending this to any number of dimensions is called the L2 norm. So: $\sqrt{\sum_{1}^{n}(weight_i - indexWeight_i)^2}$ Can also be written as $\left \| \mathbf{x} - \mathbf{index} \right \|_2$. There's a cvxpy function called [norm()](https://www.cvxpy.org/api_reference/cvxpy.atoms.other_atoms.html#norm)
# `norm(x, p=2, axis=None)`. The default is already set to find an L2 norm, so you would pass in one argument, which is the difference between your portfolio weights and the index weights.
#
# ### objective function
# We want to minimize both the portfolio variance and the distance of the portfolio weights from the index weights.
# We also want to choose a `scale` constant, which is $\lambda$ in the expression.
#
# $\mathbf{x^T} \mathbf{P} \mathbf{x} + \lambda \left \| \mathbf{x} - \mathbf{index} \right \|_2$
#
#
# This lets us choose how much priority we give to minimizing the difference from the index, relative to minimizing the variance of the portfolio. If you choose a higher value for `scale` ($\lambda$).
#
# We can find the objective function using cvxpy `objective = cvx.Minimize()`. Can you guess what to pass into this function?
#
#
# ### constraints
# We can also define our constraints in a list. For example, you'd want the weights to sum to one. So $\sum_{1}^{n}x = 1$. You may also need to go long only, which means no shorting, so no negative weights. So $x_i >0 $ for all $i$. you could save a variable as `[x >= 0, sum(x) == 1]`, where x was created using `cvx.Variable()`.
#
# ### optimization
# So now that we have our objective function and constraints, we can solve for the values of $\mathbf{x}$.
# cvxpy has the constructor `Problem(objective, constraints)`, which returns a `Problem` object.
#
# The `Problem` object has a function solve(), which returns the minimum of the solution. In this case, this is the minimum variance of the portfolio.
#
# It also updates the vector $\mathbf{x}$.
#
# We can check out the values of $x_A$ and $x_B$ that gave the minimum portfolio variance by using `x.value`
# +
import cvxpy as cvx
def get_optimal_weights(covariance_returns, index_weights, scale=2.0):
"""
Find the optimal weights.
Parameters
----------
covariance_returns : 2 dimensional Ndarray
The covariance of the returns
index_weights : Pandas Series
Index weights for all tickers at a period in time
scale : int
The penalty factor for weights the deviate from the index
Returns
-------
x : 1 dimensional Ndarray
The solution for x
"""
assert len(covariance_returns.shape) == 2
assert len(index_weights.shape) == 1
assert covariance_returns.shape[0] == covariance_returns.shape[1] == index_weights.shape[0]
#TODO: Implement function
m = covariance_returns.shape[0]
x = cvx.Variable(m)
var = cvx.quad_form(x, covariance_returns)
nrm = cvx.norm(x - index_weights)
obj = cvx.Minimize(var + scale * nrm)
cons = [x >= 0, sum(x) == 1]
cvx.Problem(obj, cons).solve()
return x.value
project_tests.test_get_optimal_weights(get_optimal_weights)
# -
# ## Optimized Portfolio
# Using the `get_optimal_weights` function, let's generate the optimal ETF weights without rebalanceing. We can do this by feeding in the covariance of the entire history of data. We also need to feed in a set of index weights. We'll go with the average weights of the index over time.
raw_optimal_single_rebalance_etf_weights = get_optimal_weights(covariance_returns.values, index_weights.iloc[-1])
optimal_single_rebalance_etf_weights = pd.DataFrame(
np.tile(raw_optimal_single_rebalance_etf_weights, (len(returns.index), 1)),
returns.index,
returns.columns)
# With our ETF weights built, let's compare it to the index. Run the next cell to calculate the ETF returns and compare it to the index returns.
# +
optim_etf_returns = generate_weighted_returns(returns, optimal_single_rebalance_etf_weights)
optim_etf_cumulative_returns = calculate_cumulative_returns(optim_etf_returns)
project_helper.plot_benchmark_returns(index_weighted_cumulative_returns, optim_etf_cumulative_returns, 'Optimized ETF vs Index')
optim_etf_tracking_error = tracking_error(np.sum(index_weighted_returns, 1), np.sum(optim_etf_returns, 1))
print('Optimized ETF Tracking Error: {}'.format(optim_etf_tracking_error))
# -
# ## Rebalance Portfolio Over Time
# The single optimized ETF portfolio used the same weights for the entire history. This might not be the optimal weights for the entire period. Let's rebalance the portfolio over the same period instead of using the same weights. Implement `rebalance_portfolio` to rebalance a portfolio.
#
# Reblance the portfolio every n number of days, which is given as `shift_size`. When rebalancing, you should look back a certain number of days of data in the past, denoted as `chunk_size`. Using this data, compute the optoimal weights using `get_optimal_weights` and `get_covariance_returns`.
# +
def rebalance_portfolio(returns, index_weights, shift_size, chunk_size):
"""
Get weights for each rebalancing of the portfolio.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
index_weights : DataFrame
Index weight for each ticker and date
shift_size : int
The number of days between each rebalance
chunk_size : int
The number of days to look in the past for rebalancing
Returns
-------
all_rebalance_weights : list of Ndarrays
The ETF weights for each point they are rebalanced
"""
assert returns.index.equals(index_weights.index)
assert returns.columns.equals(index_weights.columns)
assert shift_size > 0
assert chunk_size >= 0
#TODO: Implement function
weights = []
# print(returns)
for i in range(0, len(returns) - chunk_size, shift_size):
chunk = returns.iloc[i:i+chunk_size,:]
cov = get_covariance_returns(chunk)
idx_weight = index_weights.iloc[i+chunk_size-1,:]
# print("")
# print(chunk)
# print(idx_weight)
# print("")
weight = get_optimal_weights(cov, idx_weight)
weights.append(weight)
return weights
project_tests.test_rebalance_portfolio(rebalance_portfolio)
# -
# Run the following cell to rebalance the portfolio using `rebalance_portfolio`.
chunk_size = 250
shift_size = 5
all_rebalance_weights = rebalance_portfolio(returns, index_weights, shift_size, chunk_size)
# ## Portfolio Turnover
# With the portfolio rebalanced, we need to use a metric to measure the cost of rebalancing the portfolio. Implement `get_portfolio_turnover` to calculate the annual portfolio turnover. We'll be using the formulas used in the classroom:
#
# $ AnnualizedTurnover =\frac{SumTotalTurnover}{NumberOfRebalanceEvents} * NumberofRebalanceEventsPerYear $
#
# $ SumTotalTurnover =\sum_{t,n}{\left | x_{t,n} - x_{t+1,n} \right |} $ Where $ x_{t,n} $ are the weights at time $ t $ for equity $ n $.
#
# $ SumTotalTurnover $ is just a different way of writing $ \sum \left | x_{t_1,n} - x_{t_2,n} \right | $
# +
def get_portfolio_turnover(all_rebalance_weights, shift_size, rebalance_count, n_trading_days_in_year=252):
"""
Calculage portfolio turnover.
Parameters
----------
all_rebalance_weights : list of Ndarrays
The ETF weights for each point they are rebalanced
shift_size : int
The number of days between each rebalance
rebalance_count : int
Number of times the portfolio was rebalanced
n_trading_days_in_year: int
Number of trading days in a year
Returns
-------
portfolio_turnover : float
The portfolio turnover
"""
assert shift_size > 0
assert rebalance_count > 0
#TODO: Implement function
s = 0
for i in range(1, len(all_rebalance_weights)):
pre = all_rebalance_weights[i - 1]
cur = all_rebalance_weights[i]
s += sum(abs(cur - pre))
return s / (rebalance_count * shift_size) * n_trading_days_in_year
project_tests.test_get_portfolio_turnover(get_portfolio_turnover)
# -
# Run the following cell to get the portfolio turnover from `get_portfolio turnover`.
print(get_portfolio_turnover(all_rebalance_weights, shift_size, len(all_rebalance_weights) - 1))
# That's it! You've built a smart beta portfolio in part 1 and did portfolio optimization in part 2. You can now submit your project.
# ## Submission
# Now that you're done with the project, it's time to submit it. Click the submit button in the bottom right. One of our reviewers will give you feedback on your project with a pass or not passed grade. You can continue to the next section while you wait for feedback.
| project_3_smart_beta/project_3_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FloPy
#
# ### Henry Saltwater Intrusion Problem
#
# In this notebook, we will use Flopy to create, run, and post process the Henry saltwater intrusion problem using SEAWAT Version 4.
# +
import os
import sys
import numpy as np
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
workspace = os.path.join('data')
#make sure workspace directory exists
if not os.path.exists(workspace):
os.makedirs(workspace)
# Input variables for the Henry Problem
Lx = 2.
Lz = 1.
nlay = 50
nrow = 1
ncol = 100
delr = Lx / ncol
delc = 1.0
delv = Lz / nlay
henry_top = 1.
henry_botm = np.linspace(henry_top - delv, 0., nlay)
qinflow = 5.702 #m3/day
dmcoef = 0.57024 #m2/day Could also try 1.62925 as another case of the Henry problem
hk = 864. #m/day
# +
# Create the basic MODFLOW model structure
modelname = 'henry'
swt = flopy.seawat.Seawat(modelname, exe_name='swtv4', model_ws=workspace)
print(swt.namefile)
# save cell fluxes to unit 53
ipakcb = 53
# Add DIS package to the MODFLOW model
dis = flopy.modflow.ModflowDis(swt, nlay, nrow, ncol, nper=1, delr=delr,
delc=delc, laycbd=0, top=henry_top,
botm=henry_botm, perlen=1.5, nstp=15)
# Variables for the BAS package
ibound = np.ones((nlay, nrow, ncol), dtype=np.int32)
ibound[:, :, -1] = -1
bas = flopy.modflow.ModflowBas(swt, ibound, 0)
# Add LPF package to the MODFLOW model
lpf = flopy.modflow.ModflowLpf(swt, hk=hk, vka=hk, ipakcb=ipakcb)
# Add PCG Package to the MODFLOW model
pcg = flopy.modflow.ModflowPcg(swt, hclose=1.e-8)
# Add OC package to the MODFLOW model
oc = flopy.modflow.ModflowOc(swt,
stress_period_data={(0, 0): ['save head', 'save budget']},
compact=True)
# Create WEL and SSM data
itype = flopy.mt3d.Mt3dSsm.itype_dict()
wel_data = {}
ssm_data = {}
wel_sp1 = []
ssm_sp1 = []
for k in range(nlay):
wel_sp1.append([k, 0, 0, qinflow / nlay])
ssm_sp1.append([k, 0, 0, 0., itype['WEL']])
ssm_sp1.append([k, 0, ncol - 1, 35., itype['BAS6']])
wel_data[0] = wel_sp1
ssm_data[0] = ssm_sp1
wel = flopy.modflow.ModflowWel(swt, stress_period_data=wel_data, ipakcb=ipakcb)
# +
# Create the basic MT3DMS model structure
#mt = flopy.mt3d.Mt3dms(modelname, 'nam_mt3dms', mf, model_ws=workspace)
btn = flopy.mt3d.Mt3dBtn(swt, nprs=-5, prsity=0.35, sconc=35., ifmtcn=0,
chkmas=False, nprobs=10, nprmas=10, dt0=0.001)
adv = flopy.mt3d.Mt3dAdv(swt, mixelm=0)
dsp = flopy.mt3d.Mt3dDsp(swt, al=0., trpt=1., trpv=1., dmcoef=dmcoef)
gcg = flopy.mt3d.Mt3dGcg(swt, iter1=500, mxiter=1, isolve=1, cclose=1e-7)
ssm = flopy.mt3d.Mt3dSsm(swt, stress_period_data=ssm_data)
# Create the SEAWAT model structure
#mswt = flopy.seawat.Seawat(modelname, 'nam_swt', mf, mt, model_ws=workspace, exe_name='swtv4')
vdf = flopy.seawat.SeawatVdf(swt, iwtable=0, densemin=0, densemax=0,
denseref=1000., denseslp=0.7143, firstdt=1e-3)
# -
# Write the input files
swt.write_input()
# Try to delete the output files, to prevent accidental use of older files
try:
os.remove(os.path.join(workspace, 'MT3D001.UCN'))
os.remove(os.path.join(workspace, modelname + '.hds'))
os.remove(os.path.join(workspace, modelname + '.cbc'))
except:
pass
v = swt.run_model(silent=True, report=True)
for idx in range(-3, 0):
print(v[1][idx])
# +
# Post-process the results
import numpy as np
import flopy.utils.binaryfile as bf
# Load data
ucnobj = bf.UcnFile(os.path.join(workspace, 'MT3D001.UCN'), model=swt)
times = ucnobj.get_times()
concentration = ucnobj.get_data(totim=times[-1])
cbbobj = bf.CellBudgetFile(os.path.join(workspace, 'henry.cbc'))
times = cbbobj.get_times()
qx = cbbobj.get_data(text='flow right face', totim=times[-1])[0]
qz = cbbobj.get_data(text='flow lower face', totim=times[-1])[0]
# Average flows to cell centers
qx_avg = np.empty(qx.shape, dtype=qx.dtype)
qx_avg[:, :, 1:] = 0.5 * (qx[:, :, 0:ncol-1] + qx[:, :, 1:ncol])
qx_avg[:, :, 0] = 0.5 * qx[:, :, 0]
qz_avg = np.empty(qz.shape, dtype=qz.dtype)
qz_avg[1:, :, :] = 0.5 * (qz[0:nlay-1, :, :] + qz[1:nlay, :, :])
qz_avg[0, :, :] = 0.5 * qz[0, :, :]
# -
# Make the plot
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1, 1, 1, aspect='equal')
ax.imshow(concentration[:, 0, :], interpolation='nearest',
extent=(0, Lx, 0, Lz))
y, x, z = dis.get_node_coordinates()
X, Z = np.meshgrid(x, z[:, 0, 0])
iskip = 3
ax.quiver(X[::iskip, ::iskip], Z[::iskip, ::iskip],
qx_avg[::iskip, 0, ::iskip], -qz_avg[::iskip, 0, ::iskip],
color='w', scale=5, headwidth=3, headlength=2,
headaxislength=2, width=0.0025)
plt.savefig(os.path.join(workspace, 'henry.png'))
plt.show();
# Extract the heads
fname = os.path.join(workspace, 'henry.hds')
headobj = bf.HeadFile(fname)
times = headobj.get_times()
head = headobj.get_data(totim=times[-1])
# Make a simple head plot
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(1, 1, 1, aspect='equal')
im = ax.imshow(head[:, 0, :], interpolation='nearest',
extent=(0, Lx, 0, Lz))
ax.set_title('Simulated Heads');
# #### Change the format of several arrays and rerun the model
swt.btn.prsity.how = "constant"
swt.btn.prsity[0].how = "internal"
swt.btn.prsity[1].how = "external"
swt.btn.sconc[0].how = "external"
swt.btn.prsity[0].fmtin = "(100E15.6)"
swt.lpf.hk[0].fmtin = "(BINARY)"
swt.write_input()
v = swt.run_model(silent=True, report=True)
for idx in range(-3, 0):
print(v[1][idx])
| examples/Notebooks/flopy3_SEAWAT_henry_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning: Intermediate report
#
# + Your name (Your ID)
# Prepare an environment for running Python codes on Jupyter notebook. The most easiest way is to use [Google Colaboratory](https://colab.research.google.com/).
#
# Write codes for the following three (and one optional) problems, and submit the notebook (`.ipynb`) as well as its HTML conversion (`.html`). *We do not accept a report in other formats (e.g., Word, PDF)*. Write a code at the specified cell in the notebook. One can add more cells if necessary.
#
# These are the links to the sample codes used in the lecture:
#
# + [Binary classification](https://github.com/chokkan/deeplearningclass/blob/master/mlp_binary.ipynb)
# + [MNIST](https://github.com/chokkan/deeplearningclass/blob/master/mnist.ipynb)
# ## 1. Multi-class classification on MNIST
#
# Train a model on the training set of MNIST, and report the accuracy of the model on the test set. One can use the same code shown in the lecture. Write a code here and show the output.
# ## 2. Confusion matrix
#
# Show a confusion matrix of the predictions of the model on the test set. This is an example of a confusion matrix.
#
# 
#
# Write a code here and show the confusion matrix.
# ## 3. Top-3 confusing examples
#
# Show the top three images where the model misrecognized their digits with strong confidences. More specifically, let $y_n$ and $\hat{y}_n$ the true and predicted, respectively, digits of the image $x_n$. We want to find three images with high $P(\hat{y}_n | x_n)$ when $y_n \neq \hat{y}_n$.
#
# Please show $y_n$, $P(y_n | x_n)$, $\hat{y}_n$, and $P(\hat{y}_n | x_n)$. This is an example of an output for an image (you need this kind of outputs for top-three images).
#
# 
#
# Write a code here and show the output.
# ## 4. Sample codes in other DL frameworks
#
# (Advanced; optional) Implement one or more sample code(s) with a different deep learning framework (e.g., Chainer, TensorFlow, DyNet) corresponding to the slides 60-66 in binary classification. *When subitting an answer to this problem, please agree that some of the submitted codes will be distributed on the Web site to improve this lecture.*
| assignment/(YourID)_report2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numerical plots
# ## distplot
# ## joinplot
# ## pairplot
import seaborn as sns
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
df = sns.load_dataset('tips')
df.head()
df['total_bill'][df['sex']=='Male'].mean()
df['total_bill'][df['sex']=='Female'].mean()
# ## Correlation with Heatmap
# A correlation heatmap uses colored cells, typically in a monochromatic scale, to show a 2D correlation matrix (table) between two discrete dimensions or event types. it is very important in feature selection
# +
# Correlation does happen between integer or float types of features
df.corr()
# -
sns.heatmap(df.corr())
# +
# Correlation does happen between integer or float types of features
df.dtypes
# -
# # distPlot, JoinPlot and PairPlot are Numerical Features
# ## JoinPlot
# A join plot allows to study the relationship between 2 numeric variables. The central chart display their correlation. it is usually a scatterplot, a hexbin plot, a 2D histogram or a 2D density plot
# Join plot is for Bivariate Analysis
# ## Bivariate Analysis
sns.jointplot(x ='tip', y = 'total_bill',data = df,kind='hex')
sns.jointplot(x ='tip', y = 'total_bill',data = df,kind='scatter')
# ## Multivariate Analaysis
# A pairplot is also known as Scatterplot, in which one variable in the same data row is matched with another variables value
# like this: pair plots are just elaborations on this. showing all variables paired with all the other variables
# PairPlot is for Multivariate Analysis
sns.pairplot(data= df)
# +
sns.pairplot(data = df, hue='sex')
# hue : string (variable name), optional
# Variable in ``data`` to map plot aspects to different colors.
# -
df['smoker'].value_counts()
df['sex'].value_counts()
sns.pairplot(data = df, hue='smoker')
# ## Univariate Analysis
# ## DistPlot
# distplot helps us to check the distribution of the columns feature
sns.distplot(df['tip'])
sns.distplot(df['tip'],kde = False, bins = 10)
# ## Categorical plots
#
# Boxplot
#
# violinplot
#
# countplot
#
# barplot
#
#
# ## CountPlot
# CountPlot is for single categorical Variable and either accepts x-axis or y-axis
# +
# Countplot
# axis is x
sns.countplot(x = 'sex',data=df)
# -
# axis is y
sns.countplot(y = 'sex',data=df)
sns.countplot('day',data= df)
# ## BarPlot
# Barplot accepts one axis as categorial and second axis as Numerical
sns.barplot(y='total_bill', x = 'sex',data = df)
sns.barplot(y='total_bill', x = 'smoker',data = df)
# ## Boxplot
# A box and whisker plot is a graph that represents information from a five-number summary
# Barplot itself accepts one axis as categorial and second axis as Numerical
sns.boxplot(y='total_bill', x = 'smoker',data = df)
sns.boxplot(y='total_bill', x = 'sex',data = df)
sns.boxplot(x = 'day' , y = 'total_bill', data = df ,palette = 'rainbow')
sns.boxplot(data = df )
sns.boxplot(x='total_bill',y='day',hue='sex',data=df)
sns.boxplot(y='total_bill',x='day',hue='smoker',data=df)
# ## ViolinPlot
# violinPlot helps us to see both the distributon of data in terms of Kernal Density estimation and the BoxPlot
sns.violinplot(y ='total_bill', x='sex',data=df)
sns.violinplot(y ='total_bill', x='day',data=df, palette='rainbow')
# ## Assignment
#
df = pd.read_csv('iris.csv')
df.head()
# +
# Numerical plots
# distplot,jointplot, pairplot
# -
df['variety'].value_counts()
df.info()
df.describe()
df.corr()
sns.heatmap(df.corr())
# +
# Distplot [Univariate Analysis]
sns.distplot(df['sepal.length'])
# -
# JointPlot
sns.jointplot('sepal.length','petal.length',data = df)
# pairplot
sns.pairplot(df)
# +
# Categorcial Plot
#box plot , bar plot, countplot, violentplot
# -
#countplot
sns.countplot('variety',data = df)
# bar plot
sns.barplot(x = 'variety',y='sepal.length',data =df)
# boxplot
sns.boxplot(x='variety',y='sepal.length',data=df)
#violinPlot
sns.violinplot(x='variety',y='sepal.length',data=df)
| Data visualization/Seaborn by KrishNaik.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.8 64-bit
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import plotly.express as px
from functools import reduce
import warnings
from sklearn import preprocessing
import numpy as np
warnings.filterwarnings('ignore')
xl = pd.ExcelFile("Calidad Speech Analytics 2021.xlsx")
print(xl.sheet_names[:4],"\n",
xl.sheet_names[4:8]) # see all sheet names ``
# +
#file_path = 'Alerta exp cliente 13 Dic.xlsx'
#file_path2 = "Alerta Operativa Calidad Manual 15-12.xlsx"
#El orden en el que están las pestañas del documento excel.
habilidades = ["Acoger",
'Asesorar',
'Asistir',
'Agilizar',
'Acoger',
'Asesorar',
'Asistir ',
'Agilizar']
class getData:
def mergeAll(df1, df2, df3):
to_merge = [df1,df2,df3]
df_merged = reduce(lambda left,right: pd.merge(left,right,on=['Ejecutivo'], how='outer'), to_merge)
df_merged = df_merged.reset_index()
df_merged = df_merged.drop(columns = ['index'])
df_merged.loc[:,~df_merged.columns.duplicated()].dropna()
return df_merged
def collectMergeSpeech(file_path):
xl = pd.ExcelFile(file_path)
sheet_names = xl.sheet_names # see all sheet names
to_merge = []
for i in range(len(sheet_names)):
to_merge.append(pd.read_excel(file_path,sheet_name = sheet_names[i]))
to_merge[i]['Habilidad'] = habilidades[i]
to_merge[i] = to_merge[i].rename({'Bloque acoger': 'Score_Acoger',"Bloque asesorar":"Score_Asesorar", "Bloque asistir":"Score_Asistir", 'Bloque Agilizar': 'Score_Agilizar'}, axis = 1)
##KONECTA
df_merged = reduce(lambda left,right: pd.merge(left,right,on=['Ejecutivo'], how='outer'), to_merge[:4])
#df_merged = df_merged.rename({"Score_Agilizar_x":"Score_Agilizar","Score_Acoger_x":"Score_Acoger","Score_Asesorar_x":"Score_Asesorar","Score_Asistir_x":"Score_Asistir"}, axis=1)
df_merged = df_merged.reset_index()
df_merged = df_merged.drop(columns = ['index'])
##UPCOM
df_merged2 = reduce(lambda one,two: pd.merge(one,two,on=['Ejecutivo'], how='outer'), to_merge[4:8])
df_merged2 = df_merged2.rename({"Score_Agilizar_x":"Score_Agilizar","Score_Acoger_x":"Score_Acoger","Score_Asesorar_x":"Score_Asesorar","Score_Asistir_x":"Score_Asistir"}, axis=1)
df_merged2 = df_merged2.reset_index()
df_merged2 = df_merged2.drop(columns = ['index'])
columns = ["Ejecutivo","Aliado_x","Score_Acoger","Score_Asesorar","Score_Asistir","Score_Agilizar"]
return df_merged[columns].append(df_merged2[columns], ignore_index = True)
def collectMergeManual(file_path):
habilidades = ["Acoger",
'Asesorar',
'Asistir',
'Agilizar',
'Acoger',
'Asesorar',
'Asistir ',
'Agilizar']
xl = pd.ExcelFile(file_path)
sheet_names = xl.sheet_names # see all sheet names
to_merge = []
for i in range(len(sheet_names)):
to_merge.append(pd.read_excel(file_path,sheet_name = sheet_names[i]))
to_merge[i]['Habilidad'] = habilidades[i]
to_merge[i] = to_merge[i].rename({"Nombre ejecutivo":"Ejecutivo",'Bloque acoger': 'Score_Acoger',"Bloque asesorar":"Score_Asesorar", "Bloque asistir":"Score_Asistir", 'Bloque Agilizar': 'Score_Agilizar'}, axis = 1)
#KONECTA
df_merged = reduce(lambda left,right: pd.merge(left,right,on=['Ejecutivo'], how='outer'), to_merge[:4])
#df_merged = df_merged.rename({"Score_Agilizar_x":"Score_Agilizar","Score_Acoger_x":"Score_Acoger","Score_Asesorar_x":"Score_Asesorar","Score_Asistir_x":"Score_Asistir"}, axis=1)
df_merged = df_merged.reset_index()
df_merged = df_merged.drop(columns = ['index'])
##UPCOM
df_merged2 = reduce(lambda one,two: pd.merge(one,two,on=['Ejecutivo'], how='outer'), to_merge[4:8])
df_merged2 = df_merged2.rename({"Score_Agilizar_x":"Score_Agilizar","Score_Acoger_x":"Score_Acoger","Score_Asesorar_x":"Score_Asesorar","Score_Asistir_x":"Score_Asistir"}, axis=1)
df_merged2 = df_merged2.reset_index()
df_merged2 = df_merged2.drop(columns = ['index'])
columns = ["Ejecutivo","Aliado_x","Score_Acoger","Score_Asesorar","Score_Asistir","Score_Agilizar"]
return df_merged[columns].append(df_merged2[columns], ignore_index = True)
def collectEPA(file_path):
xl = pd.ExcelFile(file_path)
# see all sheet names
df = pd.read_excel(file_path,sheet_name=xl.sheet_names[0])
df = df.rename(columns = {"nombre_agente":"Ejecutivo", "unidad_call":"Aliado_x"})
df = df.replace({"Konecta Pl":"KONECTA","Konecta":"KONECTA","UpCom":"UPCOM","Upcom":"UPCOM"})
return df
# -
df = getData.collectMergeManual("Alerta Operativa Calidad Manual 15-12.xlsx")
#df[["Nombre ejecutivo",'Score_Acoger','Score_Agilizar']
df[["Ejecutivo","Score_Acoger","Score_Asistir","Score_Agilizar","Score_Asesorar"]].dropna().sort_values(by='Score_Acoger',ascending = False)
# +
df_speech = getData.collectMergeSpeech("Calidad Speech Analytics 2021.xlsx")
df_speech = df_speech.loc[:,~df_speech.columns.duplicated()].dropna()
df_speech['tipoAnalisis'] = "Speech"
df_manual = getData.collectMergeManual("Alerta Operativa Calidad Manual 15-12.xlsx")
df_manual = df_manual.loc[:,~df_manual.columns.duplicated()].dropna()
df_manual["tipoAnalisis"] = "Manual"
df_total = df_speech.append(df_manual)
df_total['4Average'] = (df_total['Score_Acoger'] + df_total["Score_Asesorar"] + df_total["Score_Agilizar"] + df_total['Score_Asistir'])/4
# -
px.histogram(df_total,x="Score_Acoger",color="Aliado_x")
df_manual
fig = px.histogram(df_total, x="Score_Asistir",color="tipoAnalisis",barmode="overlay",range_x=(0.2,1))
fig.show()
#px.scatter(df_speech,x="Score_Asesorar",y="Score_Agilizar")
variables = ["Score_Acoger","Score_Asistir","Score_Agilizar","Score_Asesorar", "Aliado_x","tipoAnalisis"]
fig = px.scatter_matrix(df_total[variables],dimensions=["Score_Acoger","Score_Asistir","Score_Agilizar","Score_Asesorar"],color="tipoAnalisis")
fig.show()
df_manual.describe()-df_speech.describe()
fig = px.scatter_3d(df_total,x="Score_Asesorar",y="Score_Agilizar",z="Score_Acoger", color="tipoAnalisis")
fig.show()
fig = px.scatter(df_total,x="Score_Asesorar",y="Score_Asesorar", color="tipoAnalisis")
fig.show()
df_total.sort_values(by="4Average", ascending=False).tail(60)
df_epa = getData.collectEPA("llamadasEPACTI_21122021.xlsx")
df_final = getData.mergeAll(df_manual,df_speech,df_epa).dropna(subset=["Aliado_x_x"])
df_final
# +
from sklearn.cluster import KMeans
# k means
kmeans = KMeans(n_clusters=3, random_state=0)
df_epa['cluster'] = kmeans.fit_predict(df_epa[['r1', 'r2','r3']].replace({"h":-1,"err":-1}))
# get centroids
centroids = kmeans.cluster_centers_
cen_x = [i[0] for i in centroids]
cen_y = [i[1] for i in centroids]
cen_z = [i[2] for i in centroids]
## add to df
df_epa['cen_x'] = df_epa.cluster.map({0:cen_x[0], 1:cen_x[1], 2:cen_x[2]})
df_epa['cen_y'] = df_epa.cluster.map({0:cen_y[0], 1:cen_y[1], 2:cen_y[2]})
df_epa['cen_z'] = df_epa.cluster.map({0:cen_z[0], 1:cen_z[1], 2:cen_z[2]})
# define and map colors
colors = ['#DF2020', '#81DF20', '#2095DF']
df_epa['c'] = df_epa.cluster.map({0:colors[0], 1:colors[1], 2:colors[2]})
#plt.figure(figsize = (10,10))
#plt.scatter(df_['r1'], df_['r2'], c=df_.c, alpha = 0.6, s=10)
# -
fig = px.scatter_3d(df_epa, x='r1', y='r2', z='r3',size_max=100, color='c')
fig.show()
color = {"blue":'#DF2020','red':"#81DF20",'green':"#2095DF"}
color['blue']
df_epa[df_epa['c']==color['blue']]
| general.ipynb |
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # furniture_moving
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/furniture_moving.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/furniture_moving.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Moving furnitures (scheduling) problem in Google CP Solver.
Marriott & Stukey: 'Programming with constraints', page 112f
The model implements an experimental decomposition of the
global constraint cumulative.
Compare with the following models:
* ECLiPSE: http://www.hakank.org/eclipse/furniture_moving.ecl
* MiniZinc: http://www.hakank.org/minizinc/furniture_moving.mzn
* Comet: http://www.hakank.org/comet/furniture_moving.co
* Choco: http://www.hakank.org/choco/FurnitureMoving.java
* Gecode: http://www.hakank.org/gecode/furniture_moving.cpp
* JaCoP: http://www.hakank.org/JaCoP/FurnitureMoving.java
* SICStus: http://hakank.org/sicstus/furniture_moving.pl
* Zinc: http://hakank.org/minizinc/furniture_moving.zinc
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
import sys
from ortools.constraint_solver import pywrapcp
#
# Decompositon of cumulative.
#
# Inspired by the MiniZinc implementation:
# http://www.g12.csse.unimelb.edu.au/wiki/doku.php?id=g12:zinc:lib:minizinc:std:cumulative.mzn&s[]=cumulative
# The MiniZinc decomposition is discussed in the paper:
# <NAME>, <NAME>, <NAME>, and <NAME>.
# 'Why cumulative decomposition is not as bad as it sounds.'
# Download:
# http://www.cs.mu.oz.au/%7Epjs/rcpsp/papers/cp09-cu.pdf
# http://www.cs.mu.oz.au/%7Epjs/rcpsp/cumu_lazyfd.pdf
#
#
# Parameters:
#
# s: start_times assumption: array of IntVar
# d: durations assumption: array of int
# r: resources assumption: array of int
# b: resource limit assumption: IntVar or int
#
def my_cumulative(solver, s, d, r, b):
# tasks = [i for i in range(len(s))]
tasks = [i for i in range(len(s)) if r[i] > 0 and d[i] > 0]
times_min = min([s[i].Min() for i in tasks])
times_max = max([s[i].Max() + max(d) for i in tasks])
for t in range(times_min, times_max + 1):
bb = []
for i in tasks:
c1 = solver.IsLessOrEqualCstVar(s[i], t) # s[i] <= t
c2 = solver.IsGreaterCstVar(s[i] + d[i], t) # t < s[i] + d[i]
bb.append(c1 * c2 * r[i])
solver.Add(solver.Sum(bb) <= b)
# Somewhat experimental:
# This constraint is needed to contrain the upper limit of b.
if not isinstance(b, int):
solver.Add(b <= sum(r))
# Create the solver.
solver = pywrapcp.Solver("Furniture moving")
#
# data
#
n = 4
duration = [30, 10, 15, 15]
demand = [3, 1, 3, 2]
upper_limit = 160
#
# declare variables
#
start_times = [
solver.IntVar(0, upper_limit, "start_times[%i]" % i) for i in range(n)
]
end_times = [
solver.IntVar(0, upper_limit * 2, "end_times[%i]" % i) for i in range(n)
]
end_time = solver.IntVar(0, upper_limit * 2, "end_time")
# number of needed resources, to be minimized
num_resources = solver.IntVar(0, 10, "num_resources")
#
# constraints
#
for i in range(n):
solver.Add(end_times[i] == start_times[i] + duration[i])
solver.Add(end_time == solver.Max(end_times))
my_cumulative(solver, start_times, duration, demand, num_resources)
#
# Some extra constraints to play with
#
# all tasks must end within an hour
# solver.Add(end_time <= 60)
# All tasks should start at time 0
# for i in range(n):
# solver.Add(start_times[i] == 0)
# limitation of the number of people
# solver.Add(num_resources <= 3)
#
# objective
#
# objective = solver.Minimize(end_time, 1)
objective = solver.Minimize(num_resources, 1)
#
# solution and search
#
solution = solver.Assignment()
solution.Add(start_times)
solution.Add(end_times)
solution.Add(end_time)
solution.Add(num_resources)
db = solver.Phase(start_times, solver.CHOOSE_FIRST_UNBOUND,
solver.ASSIGN_MIN_VALUE)
#
# result
#
solver.NewSearch(db, [objective])
num_solutions = 0
while solver.NextSolution():
num_solutions += 1
print("num_resources:", num_resources.Value())
print("start_times :", [start_times[i].Value() for i in range(n)])
print("duration :", [duration[i] for i in range(n)])
print("end_times :", [end_times[i].Value() for i in range(n)])
print("end_time :", end_time.Value())
print()
solver.EndSearch()
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
| examples/notebook/contrib/furniture_moving.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/datascience-1.0
# ---
# #### SageMaker Pipelines Lambda Step
#
# This notebook illustrates how a Lambda function can be run as a step in a SageMaker Pipeline.
#
# The steps in this pipeline include -
# * Preprocessing the abalone dataset
# * Train an XGBoost Model
# * Evaluate the model performance
# * Create a model
# * Deploy the model to a SageMaker Hosted Endpoint using a Lambda Function
#
# A step to register the model into a Model Registry can be added to the pipeline using the `RegisterModel` step.
# #### Prerequisites
#
# The notebook execution role should have policies which enable the notebook to create a Lambda function. The Amazon managed policy `AmazonSageMakerPipelinesIntegrations` can be added to the notebook execution role.
#
# The policy description is -
#
# ```
#
# {
# "Version": "2012-10-17",
# "Statement": [
# {
# "Effect": "Allow",
# "Action": [
# "lambda:CreateFunction",
# "lambda:DeleteFunction",
# "lambda:InvokeFunction",
# "lambda:UpdateFunctionCode"
# ],
# "Resource": [
# "arn:aws:lambda:*:*:function:*sagemaker*",
# "arn:aws:lambda:*:*:function:*sageMaker*",
# "arn:aws:lambda:*:*:function:*SageMaker*"
# ]
# },
# {
# "Effect": "Allow",
# "Action": [
# "sqs:CreateQueue",
# "sqs:SendMessage"
# ],
# "Resource": [
# "arn:aws:sqs:*:*:*sagemaker*",
# "arn:aws:sqs:*:*:*sageMaker*",
# "arn:aws:sqs:*:*:*SageMaker*"
# ]
# },
# {
# "Effect": "Allow",
# "Action": [
# "iam:PassRole"
# ],
# "Resource": "arn:aws:iam::*:role/*",
# "Condition": {
# "StringEquals": {
# "iam:PassedToService": [
# "lambda.amazonaws.com"
# ]
# }
# }
# }
# ]
# }
#
# ```
# +
import sys
# !{sys.executable} -m pip install "sagemaker>=2.51.0"
# +
import os
import time
import boto3
import sagemaker
from sagemaker.estimator import Estimator
from sagemaker.inputs import TrainingInput
from sagemaker.processing import (
ProcessingInput,
ProcessingOutput,
Processor,
ScriptProcessor,
)
from sagemaker import Model
from sagemaker.xgboost import XGBoostPredictor
from sagemaker.sklearn.processing import SKLearnProcessor
from sagemaker.workflow.parameters import (
ParameterInteger,
ParameterString,
)
from sagemaker.workflow.pipeline import Pipeline
from sagemaker.workflow.properties import PropertyFile
from sagemaker.workflow.steps import ProcessingStep, TrainingStep, CacheConfig
from sagemaker.workflow.lambda_step import (
LambdaStep,
LambdaOutput,
LambdaOutputTypeEnum,
)
from sagemaker.workflow.step_collections import CreateModelStep
from sagemaker.workflow.conditions import ConditionLessThanOrEqualTo
from sagemaker.workflow.condition_step import ConditionStep
from sagemaker.workflow.functions import JsonGet
from sagemaker.lambda_helper import Lambda
# +
# Create the SageMaker Session
region = sagemaker.Session().boto_region_name
sm_client = boto3.client("sagemaker")
boto_session = boto3.Session(region_name=region)
sagemaker_session = sagemaker.session.Session(boto_session=boto_session, sagemaker_client=sm_client)
prefix = "lambda-step-pipeline"
account_id = boto3.client("sts").get_caller_identity().get("Account")
# +
# Define variables and parameters needed for the Pipeline steps
role = sagemaker.get_execution_role()
default_bucket = sagemaker_session.default_bucket()
base_job_prefix = "lambda-step-example"
s3_prefix = "lambda-step-pipeline"
processing_instance_count = ParameterInteger(name="ProcessingInstanceCount", default_value=1)
processing_instance_type = ParameterString(
name="ProcessingInstanceType", default_value="ml.m5.xlarge"
)
training_instance_type = ParameterString(name="TrainingInstanceType", default_value="ml.m5.xlarge")
model_approval_status = ParameterString(
name="ModelApprovalStatus", default_value="PendingManualApproval"
)
input_data = ParameterString(
name="InputDataUrl",
default_value=f"s3://sagemaker-sample-files/datasets/tabular/uci_abalone/abalone.csv",
)
model_approval_status = ParameterString(
name="ModelApprovalStatus", default_value="PendingManualApproval"
)
# Cache Pipeline steps to reduce execution time on subsequent executions
cache_config = CacheConfig(enable_caching=True, expire_after="30d")
# -
# #### Data Preparation
#
# An SKLearn processor is used to prepare the dataset for the Hyperparameter Tuning job. Using the script `preprocess.py`, the dataset is featurized and split into train, test, and validation datasets.
#
# The output of this step is used as the input to the TrainingStep
# +
# %%writefile preprocess.py
"""Feature engineers the abalone dataset."""
import argparse
import logging
import os
import pathlib
import requests
import tempfile
import boto3
import numpy as np
import pandas as pd
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
logger = logging.getLogger()
logger.setLevel(logging.INFO)
logger.addHandler(logging.StreamHandler())
# Since we get a headerless CSV file we specify the column names here.
feature_columns_names = [
"sex",
"length",
"diameter",
"height",
"whole_weight",
"shucked_weight",
"viscera_weight",
"shell_weight",
]
label_column = "rings"
feature_columns_dtype = {
"sex": str,
"length": np.float64,
"diameter": np.float64,
"height": np.float64,
"whole_weight": np.float64,
"shucked_weight": np.float64,
"viscera_weight": np.float64,
"shell_weight": np.float64,
}
label_column_dtype = {"rings": np.float64}
def merge_two_dicts(x, y):
"""Merges two dicts, returning a new copy."""
z = x.copy()
z.update(y)
return z
if __name__ == "__main__":
logger.debug("Starting preprocessing.")
parser = argparse.ArgumentParser()
parser.add_argument("--input-data", type=str, required=True)
args = parser.parse_args()
base_dir = "/opt/ml/processing"
pathlib.Path(f"{base_dir}/data").mkdir(parents=True, exist_ok=True)
input_data = args.input_data
bucket = input_data.split("/")[2]
key = "/".join(input_data.split("/")[3:])
logger.info("Downloading data from bucket: %s, key: %s", bucket, key)
fn = f"{base_dir}/data/abalone-dataset.csv"
s3 = boto3.resource("s3")
s3.Bucket(bucket).download_file(key, fn)
logger.debug("Reading downloaded data.")
df = pd.read_csv(
fn,
header=None,
names=feature_columns_names + [label_column],
dtype=merge_two_dicts(feature_columns_dtype, label_column_dtype),
)
os.unlink(fn)
logger.debug("Defining transformers.")
numeric_features = list(feature_columns_names)
numeric_features.remove("sex")
numeric_transformer = Pipeline(
steps=[
("imputer", SimpleImputer(strategy="median")),
("scaler", StandardScaler()),
]
)
categorical_features = ["sex"]
categorical_transformer = Pipeline(
steps=[
("imputer", SimpleImputer(strategy="constant", fill_value="missing")),
("onehot", OneHotEncoder(handle_unknown="ignore")),
]
)
preprocess = ColumnTransformer(
transformers=[
("num", numeric_transformer, numeric_features),
("cat", categorical_transformer, categorical_features),
]
)
logger.info("Applying transforms.")
y = df.pop("rings")
X_pre = preprocess.fit_transform(df)
y_pre = y.to_numpy().reshape(len(y), 1)
X = np.concatenate((y_pre, X_pre), axis=1)
logger.info("Splitting %d rows of data into train, validation, test datasets.", len(X))
np.random.shuffle(X)
train, validation, test = np.split(X, [int(0.7 * len(X)), int(0.85 * len(X))])
logger.info("Writing out datasets to %s.", base_dir)
pd.DataFrame(train).to_csv(f"{base_dir}/train/train.csv", header=False, index=False)
pd.DataFrame(validation).to_csv(
f"{base_dir}/validation/validation.csv", header=False, index=False
)
pd.DataFrame(test).to_csv(f"{base_dir}/test/test.csv", header=False, index=False)
# +
# Process the training data step using a python script.
# Split the training data set into train, test, and validation datasets
sklearn_processor = SKLearnProcessor(
framework_version="0.23-1",
instance_type=processing_instance_type,
instance_count=processing_instance_count,
base_job_name=f"{base_job_prefix}/sklearn-abalone-preprocess",
sagemaker_session=sagemaker_session,
role=role,
)
step_process = ProcessingStep(
name="PreprocessAbaloneData",
processor=sklearn_processor,
outputs=[
ProcessingOutput(output_name="train", source="/opt/ml/processing/train"),
ProcessingOutput(output_name="validation", source="/opt/ml/processing/validation"),
ProcessingOutput(output_name="test", source="/opt/ml/processing/test"),
],
code="preprocess.py",
job_arguments=["--input-data", input_data],
cache_config=cache_config,
)
# -
# #### Model Training
#
# Train an XGBoost model with the output of the ProcessingStep.
# +
# Define the output path for the model artifacts from the Hyperparameter Tuning Job
model_path = f"s3://{default_bucket}/{base_job_prefix}/AbaloneTrain"
image_uri = sagemaker.image_uris.retrieve(
framework="xgboost",
region=region,
version="1.0-1",
py_version="py3",
instance_type=training_instance_type,
)
xgb_train = Estimator(
image_uri=image_uri,
instance_type=training_instance_type,
instance_count=1,
output_path=model_path,
base_job_name=f"{prefix}/{base_job_prefix}/sklearn-abalone-preprocess",
sagemaker_session=sagemaker_session,
role=role,
)
xgb_train.set_hyperparameters(
objective="reg:linear",
num_round=50,
max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.7,
silent=0,
)
step_train = TrainingStep(
name="TrainAbaloneModel",
estimator=xgb_train,
inputs={
"train": TrainingInput(
s3_data=step_process.properties.ProcessingOutputConfig.Outputs["train"].S3Output.S3Uri,
content_type="text/csv",
),
"validation": TrainingInput(
s3_data=step_process.properties.ProcessingOutputConfig.Outputs[
"validation"
].S3Output.S3Uri,
content_type="text/csv",
),
},
cache_config=cache_config,
)
# -
# #### Evaluate the model
#
# Use a processing job to evaluate the model from the TrainingStep. If the output of the evaluation is True, a model will be created and a Lambda will be invoked to deploy the model to a SageMaker Endpoint.
# +
# %%writefile evaluate.py
"""Evaluation script for measuring mean squared error."""
import json
import logging
import pathlib
import pickle
import tarfile
import numpy as np
import pandas as pd
import xgboost
from sklearn.metrics import mean_squared_error
logger = logging.getLogger()
logger.setLevel(logging.INFO)
logger.addHandler(logging.StreamHandler())
if __name__ == "__main__":
logger.debug("Starting evaluation.")
model_path = "/opt/ml/processing/model/model.tar.gz"
with tarfile.open(model_path) as tar:
tar.extractall(path=".")
logger.debug("Loading xgboost model.")
model = pickle.load(open("xgboost-model", "rb"))
logger.debug("Reading test data.")
test_path = "/opt/ml/processing/test/test.csv"
df = pd.read_csv(test_path, header=None)
logger.debug("Reading test data.")
y_test = df.iloc[:, 0].to_numpy()
df.drop(df.columns[0], axis=1, inplace=True)
X_test = xgboost.DMatrix(df.values)
logger.info("Performing predictions against test data.")
predictions = model.predict(X_test)
logger.debug("Calculating mean squared error.")
mse = mean_squared_error(y_test, predictions)
std = np.std(y_test - predictions)
report_dict = {
"regression_metrics": {
"mse": {"value": mse, "standard_deviation": std},
},
}
output_dir = "/opt/ml/processing/evaluation"
pathlib.Path(output_dir).mkdir(parents=True, exist_ok=True)
logger.info("Writing out evaluation report with mse: %f", mse)
evaluation_path = f"{output_dir}/evaluation.json"
with open(evaluation_path, "w") as f:
f.write(json.dumps(report_dict))
# +
# A ProcessingStep is used to evaluate the performance of the trained model. Based on the results of the evaluation, the model is created and deployed.
script_eval = ScriptProcessor(
image_uri=image_uri,
command=["python3"],
instance_type=processing_instance_type,
instance_count=1,
base_job_name=f"{prefix}/{base_job_prefix}/sklearn-abalone-preprocess",
sagemaker_session=sagemaker_session,
role=role,
)
evaluation_report = PropertyFile(
name="AbaloneEvaluationReport",
output_name="evaluation",
path="evaluation.json",
)
step_eval = ProcessingStep(
name="EvaluateAbaloneModel",
processor=script_eval,
inputs=[
ProcessingInput(
source=step_train.properties.ModelArtifacts.S3ModelArtifacts,
destination="/opt/ml/processing/model",
),
ProcessingInput(
source=step_process.properties.ProcessingOutputConfig.Outputs["test"].S3Output.S3Uri,
destination="/opt/ml/processing/test",
),
],
outputs=[
ProcessingOutput(
output_name="evaluation",
source="/opt/ml/processing/evaluation",
destination=f"s3://{default_bucket}/{s3_prefix}/evaluation_report",
),
],
code="evaluate.py",
property_files=[evaluation_report],
cache_config=cache_config,
)
# -
# #### Create the model
#
# The model is created and the name of the model is provided to the Lambda function for deployment. The `CreateModelStep` dynamically assigns a name to the model.
# +
# Create Model
model = Model(
image_uri=image_uri,
model_data=step_train.properties.ModelArtifacts.S3ModelArtifacts,
sagemaker_session=sagemaker_session,
role=role,
predictor_cls=XGBoostPredictor,
)
step_create_model = CreateModelStep(
name="CreateModel",
model=model,
inputs=sagemaker.inputs.CreateModelInput(instance_type="ml.m4.large"),
)
# -
# #### Create the Lambda Step
#
# When defining the LambdaStep, the SageMaker Lambda helper class provides helper functions for creating the Lambda function. Users can either use the `lambda_func` argument to provide the function ARN to an already deployed Lambda function OR use the `Lambda` class to create a Lambda function by providing a script, function name and role for the Lambda function.
#
# When passing inputs to the Lambda, the `inputs` argument can be used and within the Lambda function's handler, the `event` argument can be used to retrieve the inputs.
#
# The dictionary response from the Lambda function is parsed through the `LambdaOutput` objects provided to the `outputs` argument. The `output_name` in `LambdaOutput` corresponds to the dictionary key in the Lambda's return dictionary.
# #### Define the Lambda function
#
# Users can choose the leverage the Lambda helper class to create a Lambda function and provide that function object to the LambdaStep. Alternatively, users can use a pre-deployed Lambda function and provide the function ARN to the `Lambda` helper class in the lambda step.
# +
# %%writefile lambda_helper.py
"""
This Lambda function creates an Endpoint Configuration and deploys a model to an Endpoint.
The name of the model to deploy is provided via the `event` argument
"""
import json
import boto3
def lambda_handler(event, context):
""" """
sm_client = boto3.client("sagemaker")
# The name of the model created in the Pipeline CreateModelStep
model_name = event["model_name"]
endpoint_config_name = event["endpoint_config_name"]
endpoint_name = event["endpoint_name"]
create_endpoint_config_response = sm_client.create_endpoint_config(
EndpointConfigName=endpoint_config_name,
ProductionVariants=[
{
"InstanceType": "ml.m4.xlarge",
"InitialVariantWeight": 1,
"InitialInstanceCount": 1,
"ModelName": model_name,
"VariantName": "AllTraffic",
}
],
)
create_endpoint_response = sm_client.create_endpoint(
EndpointName=endpoint_name, EndpointConfigName=endpoint_config_name
)
return {
"statusCode": 200,
"body": json.dumps("Created Endpoint!"),
"other_key": "example_value",
}
# -
# #### IAM Role
#
# The Lambda function needs an IAM role that will allow it to deploy a SageMaker Endpoint. The role ARN must be provided in the LambdaStep.
#
# The Lambda role should at minimum have policies to allow `sagemaker:CreateModel`, `sagemaker:CreateEndpointConfig`, `sagemaker:CreateEndpoint` in addition to the based Lambda execution policies.
#
# A helper function in `iam_helper.py` is available to create the Lambda function role. Please note that the role uses the Amazon managed policy - `SageMakerFullAccess`. This should be replaced with an IAM policy with least privileges as per AWS IAM best practices.
# +
from iam_helper import create_lambda_role
lambda_role = create_lambda_role("lambda-deployment-role")
# +
# Custom Lambda Step
current_time = time.strftime("%m-%d-%H-%M-%S", time.localtime())
model_name = "demo-lambda-model" + current_time
endpoint_config_name = "demo-lambda-deploy-endpoint-config-" + current_time
endpoint_name = "demo-lambda-deploy-endpoint-" + current_time
function_name = "sagemaker-lambda-step-endpoint-deploy-" + current_time
# Lambda helper class can be used to create the Lambda function
func = Lambda(
function_name=function_name,
execution_role_arn=lambda_role,
script="lambda_helper.py",
handler="lambda_helper.lambda_handler",
)
output_param_1 = LambdaOutput(output_name="statusCode", output_type=LambdaOutputTypeEnum.String)
output_param_2 = LambdaOutput(output_name="body", output_type=LambdaOutputTypeEnum.String)
output_param_3 = LambdaOutput(output_name="other_key", output_type=LambdaOutputTypeEnum.String)
step_deploy_lambda = LambdaStep(
name="LambdaStep",
lambda_func=func,
inputs={
"model_name": step_create_model.properties.ModelName,
"endpoint_config_name": endpoint_config_name,
"endpoint_name": endpoint_name,
},
outputs=[output_param_1, output_param_2, output_param_3],
)
# +
# condition step for evaluating model quality and branching execution.
# The `json_path` value is based on the `report_dict` variable in `evaluate.py`
cond_lte = ConditionLessThanOrEqualTo(
left=JsonGet(
step_name=step_eval.name,
property_file=evaluation_report,
json_path="regression_metrics.mse.value",
),
right=6.0,
)
step_cond = ConditionStep(
name="CheckMSEAbaloneEvaluation",
conditions=[cond_lte],
if_steps=[step_create_model, step_deploy_lambda],
else_steps=[],
)
# +
# Use the same pipeline name across execution for cache usage.
pipeline_name = "lambda-step-pipeline" + current_time
pipeline = Pipeline(
name=pipeline_name,
parameters=[
processing_instance_type,
processing_instance_count,
training_instance_type,
input_data,
model_approval_status,
],
steps=[step_process, step_train, step_eval, step_cond],
sagemaker_session=sagemaker_session,
)
# -
# #### Execute the Pipeline
# +
import json
definition = json.loads(pipeline.definition())
definition
# -
pipeline.upsert(role_arn=role)
execution = pipeline.start()
execution.wait()
# #### Cleaning up resources
#
# Running the following cell will delete the following resources created in this notebook -
# * SageMaker Model
# * SageMaker Endpoint Configuration
# * SageMaker Endpoint
# * SageMaker Pipeline
# * Lambda Function
# +
# Create a SageMaker client
sm_client = boto3.client("sagemaker")
# Get the model name from the EndpointCofig. The CreateModelStep properties are not available outside the Pipeline execution context
# so `step_create_model.properties.ModelName` can not be used while deleting the model.
model_name = sm_client.describe_endpoint_config(EndpointConfigName=endpoint_config_name)[
"ProductionVariants"
][0]["ModelName"]
# Delete the Model
sm_client.delete_model(ModelName=model_name)
# Delete the EndpointConfig
sm_client.delete_endpoint_config(EndpointConfigName=endpoint_config_name)
# Delete the endpoint
sm_client.delete_endpoint(EndpointName=endpoint_name)
# Delete the Lambda function
func.delete()
# Delete the Pipeline
sm_client.delete_pipeline(PipelineName=pipeline_name)
| sagemaker-pipelines/tabular/lambda-step/sagemaker-pipelines-lambda-step.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Round Trip Tear Sheet Example
# When evaluating the performance of an investing strategy, it is helpful to quantify the frequency, duration, and profitability of its independent bets, or "round trip" trades. A round trip trade is started when a new long or short position is opened and then later completely or partially closed out.
#
# The intent of the round trip tearsheet is to help differentiate strategies that profited off a few lucky trades from strategies that profited repeatedly from genuine alpha. Breaking down round trip profitability by traded name and sector can also help inform universe selection and identify exposure risks. For example, even if your equity curve looks robust, if only two securities in your universe of fifteen names contributed to overall profitability, you may have reason to question the logic of your strategy.
#
# To identify round trips, pyfolio reconstructs the complete portfolio based on the transactions that you pass in. When you make a trade, pyfolio checks if shares are already present in your portfolio purchased at a certain price. If there are, we compute the PnL, returns and duration of that round trip trade. In calculating round trips, pyfolio will also append position closing transactions at the last timestamp in the positions data. This closing transaction will cause the PnL from any open positions to realized as completed round trips.
# +
import pyfolio as pf
# %matplotlib inline
import gzip
import os
import pandas as pd
# silence warnings
import warnings
warnings.filterwarnings('ignore')
# -
transactions = pd.read_csv(gzip.open('../tests/test_data/test_txn.csv.gz'),
index_col=0, parse_dates=True)
positions = pd.read_csv(gzip.open('../tests/test_data/test_pos.csv.gz'),
index_col=0, parse_dates=True)
returns = pd.read_csv(gzip.open('../tests/test_data/test_returns.csv.gz'),
index_col=0, parse_dates=True, header=None)[1]
# Optional: Sector mappings may be passed in as a dict or pd.Series. If a mapping is
# provided, PnL from symbols with mappings will be summed to display profitability by sector.
sect_map = {'COST': 'Consumer Goods', 'INTC':'Technology', 'CERN':'Healthcare', 'GPS':'Technology',
'MMM': 'Construction', 'DELL': 'Technology', 'AMD':'Technology'}
# The easiest way to run the analysis is to call `pyfolio.create_round_trip_tear_sheet()`. Passing in a sector map is optional. You can also pass `round_trips=True` to `pyfolio.create_full_tear_sheet()` to have this be created along all the other analyses.
pf.create_round_trip_tear_sheet(returns, positions, transactions, sector_mappings=sect_map)
# Under the hood, several functions are being called. `extract_round_trips()` does the portfolio reconstruction and creates the round-trip trades.
rts = pf.round_trips.extract_round_trips(transactions,
portfolio_value=positions.sum(axis='columns') / (returns + 1))
rts.head()
pf.round_trips.print_round_trip_stats(rts)
| pyfolio/examples/round_trip_tear_sheet_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # AE outlier detection on CIFAR10
#
# ### Method
#
# The Auto-Encoder (AE) outlier detector is first trained on a batch of unlabeled, but normal (inlier) data. Unsupervised training is desireable since labeled data is often scarce. The AE detector tries to reconstruct the input it receives. If the input data cannot be reconstructed well, the reconstruction error is high and the data can be flagged as an outlier. The reconstruction error is measured as the mean squared error (MSE) between the input and the reconstructed instance.
#
# ## Dataset
#
# [CIFAR10](https://www.cs.toronto.edu/~kriz/cifar.html) consists of 60,000 32 by 32 RGB images equally distributed over 10 classes.
# +
import logging
import matplotlib.pyplot as plt
import numpy as np
import os
import tensorflow as tf
tf.keras.backend.clear_session()
from tensorflow.keras.layers import Conv2D, Conv2DTranspose, \
Dense, Layer, Reshape, InputLayer, Flatten
from tqdm import tqdm
from alibi_detect.od import OutlierAE
from alibi_detect.utils.fetching import fetch_detector
from alibi_detect.utils.perturbation import apply_mask
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.utils.visualize import plot_instance_score, plot_feature_outlier_image
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# -
# ## Load CIFAR10 data
# +
train, test = tf.keras.datasets.cifar10.load_data()
X_train, y_train = train
X_test, y_test = test
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# -
# ## Load or define outlier detector
#
# The pretrained outlier and adversarial detectors used in the example notebooks can be found [here](https://console.cloud.google.com/storage/browser/seldon-models/alibi-detect). You can use the built-in ```fetch_detector``` function which saves the pre-trained models in a local directory ```filepath``` and loads the detector. Alternatively, you can train a detector from scratch:
load_outlier_detector = True
filepath = 'my_path' # change to (absolute) directory where model is downloaded
detector_type = 'outlier'
dataset = 'cifar10'
detector_name = 'OutlierAE'
filepath = os.path.join(filepath, detector_name)
if load_outlier_detector: # load pretrained outlier detector
od = fetch_detector(filepath, detector_type, dataset, detector_name)
else: # define model, initialize, train and save outlier detector
encoding_dim = 1024
encoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(32, 32, 3)),
Conv2D(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(128, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(512, 4, strides=2, padding='same', activation=tf.nn.relu),
Flatten(),
Dense(encoding_dim,)
])
decoder_net = tf.keras.Sequential(
[
InputLayer(input_shape=(encoding_dim,)),
Dense(4*4*128),
Reshape(target_shape=(4, 4, 128)),
Conv2DTranspose(256, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2DTranspose(64, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2DTranspose(3, 4, strides=2, padding='same', activation='sigmoid')
])
# initialize outlier detector
od = OutlierAE(threshold=.015, # threshold for outlier score
encoder_net=encoder_net, # can also pass AE model instead
decoder_net=decoder_net, # of separate encoder and decoder
)
# train
od.fit(X_train,
epochs=50,
verbose=True)
# save the trained outlier detector
save_detector(od, filepath)
# ## Check quality AE model
idx = 8
X = X_train[idx].reshape(1, 32, 32, 3)
X_recon = od.ae(X)
plt.imshow(X.reshape(32, 32, 3))
plt.axis('off')
plt.show()
plt.imshow(X_recon.numpy().reshape(32, 32, 3))
plt.axis('off')
plt.show()
# ## Check outliers on original CIFAR images
X = X_train[:500]
print(X.shape)
od_preds = od.predict(X,
outlier_type='instance', # use 'feature' or 'instance' level
return_feature_score=True, # scores used to determine outliers
return_instance_score=True)
print(list(od_preds['data'].keys()))
# ### Plot instance level outlier scores
target = np.zeros(X.shape[0],).astype(int) # all normal CIFAR10 training instances
labels = ['normal', 'outlier']
plot_instance_score(od_preds, target, labels, od.threshold)
# ### Visualize predictions
X_recon = od.ae(X).numpy()
plot_feature_outlier_image(od_preds,
X,
X_recon=X_recon,
instance_ids=[8, 60, 100, 330], # pass a list with indices of instances to display
max_instances=5, # max nb of instances to display
outliers_only=False) # only show outlier predictions
# ## Predict outliers on perturbed CIFAR images
# We perturb CIFAR images by adding random noise to patches (masks) of the image. For each mask size in `n_mask_sizes`, sample `n_masks` and apply those to each of the `n_imgs` images. Then we predict outliers on the masked instances:
# nb of predictions per image: n_masks * n_mask_sizes
n_mask_sizes = 10
n_masks = 20
n_imgs = 50
# Define masks and get images:
mask_sizes = [(2*n,2*n) for n in range(1,n_mask_sizes+1)]
print(mask_sizes)
img_ids = np.arange(n_imgs)
X_orig = X[img_ids].reshape(img_ids.shape[0], 32, 32, 3)
print(X_orig.shape)
# Calculate instance level outlier scores:
all_img_scores = []
for i in tqdm(range(X_orig.shape[0])):
img_scores = np.zeros((len(mask_sizes),))
for j, mask_size in enumerate(mask_sizes):
# create masked instances
X_mask, mask = apply_mask(X_orig[i].reshape(1, 32, 32, 3),
mask_size=mask_size,
n_masks=n_masks,
channels=[0,1,2],
mask_type='normal',
noise_distr=(0,1),
clip_rng=(0,1))
# predict outliers
od_preds_mask = od.predict(X_mask)
score = od_preds_mask['data']['instance_score']
# store average score over `n_masks` for a given mask size
img_scores[j] = np.mean(score)
all_img_scores.append(img_scores)
# ### Visualize outlier scores vs. mask sizes
x_plt = [mask[0] for mask in mask_sizes]
for ais in all_img_scores:
plt.plot(x_plt, ais)
plt.xticks(x_plt)
plt.title('Outlier Score All Images for Increasing Mask Size')
plt.xlabel('Mask size')
plt.ylabel('Outlier Score')
plt.show()
ais_np = np.zeros((len(all_img_scores), all_img_scores[0].shape[0]))
for i, ais in enumerate(all_img_scores):
ais_np[i, :] = ais
ais_mean = np.mean(ais_np, axis=0)
plt.title('Mean Outlier Score All Images for Increasing Mask Size')
plt.xlabel('Mask size')
plt.ylabel('Outlier score')
plt.plot(x_plt, ais_mean)
plt.xticks(x_plt)
plt.show()
# ### Investigate instance level outlier
i = 8 # index of instance to look at
plt.plot(x_plt, all_img_scores[i])
plt.xticks(x_plt)
plt.title('Outlier Scores Image {} for Increasing Mask Size'.format(i))
plt.xlabel('Mask size')
plt.ylabel('Outlier score')
plt.show()
# Reconstruction of masked images and outlier scores per channel:
all_X_mask = []
X_i = X_orig[i].reshape(1, 32, 32, 3)
all_X_mask.append(X_i)
# apply masks
for j, mask_size in enumerate(mask_sizes):
# create masked instances
X_mask, mask = apply_mask(X_i,
mask_size=mask_size,
n_masks=1, # just 1 for visualization purposes
channels=[0,1,2],
mask_type='normal',
noise_distr=(0,1),
clip_rng=(0,1))
all_X_mask.append(X_mask)
all_X_mask = np.concatenate(all_X_mask, axis=0)
all_X_recon = od.ae(all_X_mask).numpy()
od_preds = od.predict(all_X_mask)
# Visualize:
plot_feature_outlier_image(od_preds,
all_X_mask,
X_recon=all_X_recon,
max_instances=all_X_mask.shape[0],
n_channels=3)
# ## Predict outliers on a subset of features
#
# The sensitivity of the outlier detector can not only be controlled via the `threshold`, but also by selecting the percentage of the features used for the instance level outlier score computation. For instance, we might want to flag outliers if 40% of the features (pixels for images) have an average outlier score above the threshold. This is possible via the `outlier_perc` argument in the `predict` function. It specifies the percentage of the features that are used for outlier detection, sorted in descending outlier score order.
# +
perc_list = [20, 40, 60, 80, 100]
all_perc_scores = []
for perc in perc_list:
od_preds_perc = od.predict(all_X_mask, outlier_perc=perc)
iscore = od_preds_perc['data']['instance_score']
all_perc_scores.append(iscore)
# -
# Visualize outlier scores vs. mask sizes and percentage of features used:
x_plt = [0] + x_plt
for aps in all_perc_scores:
plt.plot(x_plt, aps)
plt.xticks(x_plt)
plt.legend(perc_list)
plt.title('Outlier Score for Increasing Mask Size and Different Feature Subsets')
plt.xlabel('Mask Size')
plt.ylabel('Outlier Score')
plt.show()
# ## Infer outlier threshold value
#
# Finding good threshold values can be tricky since they are typically not easy to interpret. The `infer_threshold` method helps finding a sensible value. We need to pass a batch of instances `X` and specify what percentage of those we consider to be normal via `threshold_perc`.
print('Current threshold: {}'.format(od.threshold))
od.infer_threshold(X, threshold_perc=99) # assume 1% of the training data are outliers
print('New threshold: {}'.format(od.threshold))
| examples/od_ae_cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import ipyvolume as ipv
import numpy as np
from compas_vol.primitives import VolBox, VolPlane, VolCylinder
from compas_vol.microstructures import TPMS
from compas.geometry import Box, Frame, Point, Plane, Cylinder, Circle
from compas_vol.combinations import Intersection, Union, Subtraction
from compas_vol.modifications import Overlay, Shell
# -
vbox = VolBox(Box(Frame.worldXY(), 250, 30, 10), 1.5)
cyl = VolCylinder(Cylinder(Circle(Plane((125,0,0),(0,0,1)), 15), 10))
cbu = Union(vbox, cyl)
rx, ry, rz = np.ogrid[-130:145:550j, -16:16:64j, -8:8:32j]
dm = vbox.get_distance_numpy(rx, ry, rz)
ipv.figure(width=800, height=450)
mesh = ipv.plot_isosurface(dm, 0.0, extent=[[-130,145], [-16,16], [-8,8]], color='white')
ipv.xyzlim(145)
ipv.style.use('minimal')
ipv.show()
gyroid = TPMS(tpmstype='Gyroid', wavelength=5.0)
shell = Shell(gyroid, 2.0, 0.5)
vplane = VolPlane(Plane((0,0,0), (1,0,0)))
overlay = Overlay(shell, vplane, 0.005)
ovo = Overlay(cbu, vplane, -0.01)
intersection = Intersection(overlay, ovo)
co = VolCylinder(Cylinder(Circle(Plane((125,0,0),(0,0,1)), 12), 13))
ci = VolCylinder(Cylinder(Circle(Plane((125,0,0),(0,0,1)), 10), 20))
add = Union(intersection, co)
hole = Subtraction(add, ci)
dm = hole.get_distance_numpy(rx, ry, rz)
ipv.figure(width=800, height=450)
mesh = ipv.plot_isosurface(dm, 0.0, extent=[[-130,145], [-16,16], [-8,8]], color='white')
ipv.xyzlim(145)
ipv.style.use('minimal')
ipv.show()
from compas_vol.utilities import export_ipv_mesh
export_ipv_mesh(mesh, 'handle2.obj')
| T1/10_volumetric_modelling/wrench_clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("..")
import torch
import torch.nn as nn
import torch.optim as optim
import torch.backends.cudnn as cudnn
import os
import argparse
from inflation import BBI
import numpy as np
from experiments.cifar import cifar
from experiments.PDE_PoissonD import PDE_PoissonD
from run_experiment_hyperopt import *
from hyperopt import hp, tpe, Trials, fmin
import json
# !mkdir -p results
experiment = "cifar"
tune_epochs = 3 #number of epochs used during the tuning
n_trials = 50 #number of evaluations for the tuning, for each optimizer
check_epochs = 150 # number of epochs for checking the performance after the tuning
# for the general Poisson experiment, choose here the problem number
problem_number = None
seed = 42
#fixed BBI parameters
threshold_BBI = 2000
threshold0_BBI = 100
consEn_BBI = True
nFixedBounces_BBI = 100
deltaEn = 0.0
scanning_pars = {
'tune_epochs': tune_epochs,
'n_trials': n_trials,
'check_epochs': check_epochs,
'seed': seed,
'sgd': {'stepsize': [0.001, 0.2], 'rho': [0.8,1.0]},
'problem': problem_number,
'BBI' : {'stepsize': [0.001, 0.2]} ,
'comments': 'test Experiment. Fixed BBI pars:\n'+'\nthreshold_BBI: '+str(threshold_BBI)+
'\nthreshold0_BBI: '+str(threshold0_BBI)+
'\nconsEN_BBI: '+str(consEn_BBI)+
'\nnFixedBounces_BBI: '+str(nFixedBounces_BBI)+
'\ndeltaEn: '+str(deltaEn)
}
with open('results/scanning-parameters-'+experiment+'.json', 'w') as json_file:
json.dump(scanning_pars, json_file)
print("I am working on ", experiment)
if torch.cuda.is_available(): print("I am running on gpu")
else: print("I am running on cpu")
# +
def run_experiment_sgd(epochs = 2, stepsize = 1e-3, rho = .99):
param_list = ["main.py", experiment, "--optimizer", "sgd",
"--lr", str(stepsize), "--rho", str(rho),
"--epochs", str(epochs), "--seed", str(seed),
"--progress", "false", "--device", "cuda"]
if experiment == "PDE_PoissonD":
param_list.append("--problem")
param_list.append(str(problem_number))
return run_experiment(param_list)
def run_experiment_BBI(epochs = 2, stepsize = 1e-3, threshold = threshold_BBI, threshold0 = threshold0_BBI, consEn = consEn_BBI, nFixedBounces = nFixedBounces_BBI, deltaEn = deltaEn):
param_list = ["main.py", experiment, "--optimizer", "BBI",
"--lr", str(stepsize),
"--epochs", str(epochs),"--seed", str(seed),
"--threshold", str(threshold),
"--threshold0", str(threshold0),
"--nFixedBounces", str(nFixedBounces),
"--deltaEn", str(deltaEn),
"--consEn", str(consEn),
"--progress", "false","--device", "cuda"]
if experiment == "PDE_PoissonD":
param_list.append("--problem")
param_list.append(str(problem_number))
return run_experiment(param_list)
# -
def hyperopt_tuning(ranges, optimizer, epochs=10, n_trials=5):
def optimizer_func(pars):
return optimizer(epochs=epochs, **pars)
fspace = {}
for par, range in ranges.items(): fspace[par] = hp.uniform(par, *range)
trials = Trials()
best = fmin(fn=optimizer_func, space=fspace, algo=tpe.suggest, trials=trials, max_evals=n_trials)
return best
print("Tuning sgd: ")
best_par_sgd = hyperopt_tuning(scanning_pars['sgd'], run_experiment_sgd, epochs = tune_epochs, n_trials=n_trials)
print("best sgd parameters:", best_par_sgd)
print("Tuning BBI: ")
best_par_BBI = hyperopt_tuning(scanning_pars['BBI'],run_experiment_BBI, epochs = tune_epochs, n_trials=n_trials)
print("best BBI parameters:", best_par_BBI)
# +
print("Best parameters")
print("sgd:", best_par_sgd)
print("BBI:", best_par_BBI)
best_pars = {
'sgd': best_par_sgd,
'BBI': best_par_BBI
}
with open('results/best-parameters-'+experiment+'.json', 'w') as json_file:
json.dump(best_pars, json_file)
# + tags=[]
print("Running experiment with the best parameters for more epochs...")
print("Running BBI: ")
final_loss_BBI = run_experiment_BBI(epochs=check_epochs, **best_par_BBI)
print(final_loss_BBI)
print("Running sgd: ")
final_loss_sgd = run_experiment_sgd(epochs=check_epochs, **best_par_sgd)
print(final_loss_sgd)
# -
| CIFAR/CIFAR-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/theroyakash/DeepDream/blob/master/DeepDream.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qbuu4c3vjNNH" colab_type="code" colab={}
# Copyright 2020 theroyakash. All Rights Reserved.
#
# See privacy policy at https://www.iamroyakash.com/privacy
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + id="NBfG1p7mc4L4" colab_type="code" colab={}
import numpy as np
import matplotlib as mpl
import tensorflow as tf
import IPython.display as display
from PIL import Image
import io
import requests
from tensorflow.keras.preprocessing import image
# + id="3kIxDz0RdR43" colab_type="code" colab={}
url = 'https://images.unsplash.com/photo-1597202992582-9ee5c6672095'
# + id="O15FQvRTeJ1Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 315} outputId="879a820d-86e7-445e-d2ba-cc20b7efd876"
def download(url, max_dim=None):
'''
Download an image and read it into a NumPy array.
Args:
- ``url``: URL of the image on the web
Returns:
- Numpy array of the image
'''
img = Image.open(io.BytesIO(requests.get(f'{url}').content))
if max_dim:
img.thumbnail((max_dim, max_dim))
return np.array(img)
def deprocess(img):
'''
Normalizes image
Args:
- ``img``: PIL Image as image
'''
img = 255*(img + 1.0)/2.0
return tf.cast(img, tf.uint8)
def show(img):
display.display(PIL.Image.fromarray(np.array(img)))
# Downsizing the image makes it easier to work with.
original_img = download(url, max_dim=500)
show(original_img)
display.display(display.HTML('<span>Photo by <a href="https://unsplash.com/@theroyakash?utm_source=unsplash&utm_medium=referral&utm_content=creditCopyText">theroyakash</a> on <a href="https://unsplash.com/@theroyakash?utm_source=unsplash&utm_medium=referral&utm_content=creditCopyText">Unsplash</a></span>'))
# + id="Ri2d6cpSfd3K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="3d9e4484-a6ce-4c51-998d-af7882c53f5c"
# Downloading the base model
base_model = tf.keras.applications.InceptionV3(include_top=False, weights='imagenet')
# + id="qAJzXO9AfrVR" colab_type="code" colab={}
# Activation Maximization for the following layers
names = ['mixed3', 'mixed5']
layers = [base_model.get_layer(name).output for name in names]
# Feature extraction model
dream_model = tf.keras.Model(inputs=base_model.input, outputs=layers)
# + id="ELE3okjvf9AL" colab_type="code" colab={}
def calc_loss(img, model):
'''
Pass forward the image through the model to retrieve the activations. Converts the image into a batch of size 1.
Args:
- img
- model: Tensorflow Model
Returns:
- ``tf.reduce_sum(losses)``
'''
img_batch = tf.expand_dims(img, axis=0)
layer_activations = model(img_batch)
if len(layer_activations) == 1:
layer_activations = [layer_activations]
losses = []
for act in layer_activations:
loss = tf.math.reduce_mean(act)
losses.append(loss)
return tf.reduce_sum(losses)
# + id="7NgCLkzSgYpI" colab_type="code" colab={}
# Gradient ascent step
class DeepDream(tf.Module):
def __init__(self, model):
self.model = model
@tf.function(
input_signature=(
tf.TensorSpec(shape=[None,None,3], dtype=tf.float32),
tf.TensorSpec(shape=[], dtype=tf.int32),
tf.TensorSpec(shape=[], dtype=tf.float32),)
)
def __call__(self, img, steps, step_size):
print("Tracing")
loss = tf.constant(0.0)
for n in tf.range(steps):
with tf.GradientTape() as tape:
# This needs gradients relative to `img`
# `GradientTape` only watches `tf.Variable`s by default
tape.watch(img)
loss = calc_loss(img, self.model)
# Calculate the gradient of the loss with respect to the pixels of the input image.
gradients = tape.gradient(loss, img)
# Normalize the gradients.
gradients /= tf.math.reduce_std(gradients) + 1e-8
# In gradient ascent, the "loss" is maximized so that the input image increasingly "excites" the layers.
# You can update the image by directly adding the gradients (because they're the same shape!)
img = img + gradients*step_size
img = tf.clip_by_value(img, -1, 1)
return loss, img
# + id="GmdzpAsug4os" colab_type="code" colab={}
deepdream = DeepDream(dream_model)
# + id="0OSAZsRYg6-S" colab_type="code" colab={}
def run_dream(img, steps=100, step_size=0.01):
# Convert from uint8 to the range expected by the model.
img = tf.keras.applications.inception_v3.preprocess_input(img)
img = tf.convert_to_tensor(img)
step_size = tf.convert_to_tensor(step_size)
steps_remaining = steps
step = 0
while steps_remaining:
if steps_remaining>100:
run_steps = tf.constant(100)
else:
run_steps = tf.constant(steps_remaining)
steps_remaining -= run_steps
step += run_steps
loss, img = deepdream(img, run_steps, tf.constant(step_size))
display.clear_output(wait=True)
show(deprocess(img))
print ("Step {}, loss {}".format(step, loss))
result = deprocess(img)
display.clear_output(wait=True)
show(result)
return result
# + id="-_62rQf4hGN1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="0e92d59b-4b14-4ee8-f0d6-3791ec6f205b"
dream_img = run_dream(img=original_img,steps=100, step_size=0.01)
| DeepDream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Loading packages
import numpy as np
import matplotlib.pylab as py
import pandas as pa
import scipy.stats as st
np.set_printoptions(precision=2)
# %matplotlib inline
# # Discrete Random Variables
# In this section we show a few example of discrete random variables using Python.
# The documentation for these routines can be found at:
#
# http://docs.scipy.org/doc/scipy-0.14.0/reference/stats.html
X=st.bernoulli(p=0.3)
X.rvs(100)
# Note that "high" is not included.
X=st.randint(low=1,high=5)
X.rvs(100)
# # Continuous Random Variables
# The documentation for these routines can be found at:
#
# http://docs.scipy.org/doc/scipy-0.14.0/reference/stats.html
XUniform=st.uniform(loc=0.7,scale=0.3);
# "bins" tells you how many bars to use
# "normed" says to turn the counts into probability densities
py.hist(XUniform.rvs(1000000),bins=20,normed=True);
x = np.linspace(-0.1,1.1,100)
py.plot(x,XUniform.pdf(x))
#py.savefig('Figures/uniformPDF.png')
py.plot(XUniform.cdf(x))
#py.savefig('Figures/uniformCDF.png')
XNormal=st.norm(loc=0,scale=1);
# "bins" tells you how many bars to use
# "normed" says to turn the counts into probability densities
py.hist(XNormal.rvs(1000),bins=100,normed=True);
x = np.linspace(-3,3,100)
py.plot(x,XNormal.pdf(x))
#py.savefig('Figures/normalPDF.png')
# http://en.wikipedia.org/wiki/Carl_Friedrich_Gauss
py.plot(XNormal.cdf(x))
#py.savefig('Figures/normalCDF.png')
# Now we can look at the histograms of some of our data from Case Study 2.
data = pa.read_hdf('data.h5','movies')
data
data['title'][100000]
X=data.pivot_table('rating',index='timestamp',aggfunc='count')
X.plot()
# Warning: Some versions of Pandas use "index" and "columns", some use "rows" and "cols"
X=data.pivot_table('rating',index='title',aggfunc='sum')
#X=data.pivot_table('rating',rows='title',aggfunc='sum')
X
X.hist()
# Warning: Some versions of Pandas use "index" and "columns", some use "rows" and "cols"
X=data.pivot_table('rating',index='occupation',aggfunc='sum')
#X=data.pivot_table('rating',rows='occupation',aggfunc='sum')
X
# ## Central limit theorem
# Here we show an example of the central limit theorem. You can play around with "numberOfDistributions" and "numberOfSamples" to see how quickly this converges to something that looks Gaussian.
numberOfDistributions = 100
numberOfSamples = 1000
XTest = st.uniform(loc=0,scale=1);
# The same thing works with many distributions.
#XTest = st.lognorm(s=1.0);
XCLT=np.zeros([numberOfSamples])
for i in range(numberOfSamples):
for j in range(numberOfDistributions):
XCLT[i] += XTest.rvs()
XCLT[i] = XCLT[i]/numberOfDistributions
py.hist(XCLT,normed=True)
# # Linear Algebra
# Some basic ideas in Linear Algebra and how you can use them in Python.
import numpy as np
a=np.array([1,2,3])
a
A=np.matrix(np.random.randint(1,10,size=[3,3]))
A
x=np.matrix([[1],[2],[3]])
print x
print x.T
a*a
np.dot(a,a)
x.T*x
A*x
b = np.matrix([[5],[6],[7]])
b
Ai = np.linalg.inv(A)
print A
print Ai
A*Ai
Ai*A
xHat = Ai*b
xHat
print A*xHat
print b
# ## But matrix inversion can be very expensive.
sizes = range(100,1000,200)
times = np.zeros(len(sizes))
for i in range(len(sizes)):
A = np.random.random(size=[sizes[i],sizes[i]])
# x = %timeit -o np.linalg.inv(A)
times[i] = x.best
py.plot(sizes,times)
# ## Something slightly more advanced: Sparse matrices.
# Sparse matrices (those with lots of 0s) can often be worked with much more efficiently than general matrices than standard methods.
from scipy.sparse.linalg import spsolve
from scipy.sparse import rand,eye
mySize = 1000
A=rand(mySize,mySize,0.001)+eye(mySize)
b=np.random.random(size=[mySize])
# The sparsity structure of A.
py.spy(A,markersize=0.1)
# dense = %timeit -o np.linalg.solve(A.todense(),b)
# sparse = %timeit -o spsolve(A,b)
dense.best/sparse.best
# # Descriptive statistics
# Pandas provides many routines for computing statistics.
XNormal=st.norm(loc=0.7,scale=2);
x = XNormal.rvs(1000)
print np.mean(x)
print np.std(x)
print np.var(x)
# But empirical measures are not always good approximations of the true properties of the distribution.
sizes = np.arange(16)+1
errors = np.zeros(16)
for i in range(16):
x = XNormal.rvs(2**i)
errors[i] = np.abs(0.7-np.mean(x))
py.plot(sizes,errors)
py.plot(sizes,2/np.sqrt(sizes))
py.plot(sizes,2*2/np.sqrt(sizes),'r')
#py.savefig('Figures/errorInMean.png')
# # Playing around with data
# +
# data.pivot_table?
# -
X=data.pivot_table('rating',index='title',aggfunc='mean')
#X=data.pivot_table('rating',rows='title',aggfunc='mean')
hist(X)
X=data.pivot_table('rating',index='title',columns='gender',aggfunc='mean')
#X=data.pivot_table('rating',rows='title',cols='gender',aggfunc='mean')
py.subplot(1,2,1)
X['M'].hist()
py.subplot(1,2,2)
X['F'].hist()
py.plot(X['M'],X['F'],'.')
X.cov()
X.corr()
X=data.pivot_table('rating',index='occupation',columns='gender',aggfunc='mean')
#X=data.pivot_table('rating',rows='occupation',cols='gender',aggfunc='mean')
X
| lectures/06 Machine Learning Part 1 and Midterm Review/1_PythonExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Nd2OOOVXxXS1" colab_type="text"
# This study guide should reinforce and provide practice for all of the concepts you have seen in the past week. There are a mix of written questions and coding exercises, both are equally important to prepare you for the sprint challenge as well as to be able to speak on these topics comfortably in interviews and on the job.
#
# If you get stuck or are unsure of something remember the 20 minute rule. If that doesn't help, then research a solution with google and stackoverflow. Only once you have exausted these methods should you turn to your Team Lead - they won't be there on your SC or during an interview. That being said, don't hesitate to ask for help if you truly are stuck.
#
# Have fun studying!
# + [markdown] id="YbwATE4qeJx7" colab_type="text"
# # Applied Modeling
# + [markdown] id="fpvInKdXekFi" colab_type="text"
# ## Questions
# + [markdown] id="Q6bS8AhBZ86H" colab_type="text"
# When completing this section, try to limit your answers to 2-3 sentences max and use plain english as much as possible. It's very easy to hide incomplete knowledge and understanding behind fancy or technical words, so imagine you are explaining these things to a non-technical interviewer.
#
# 1. What is a Catagorical Feature
# ```
# A categorical feature is a feature(column) that is made of categorical variables like age or time period
# ```
#
# 2. What are Confusion Matrix?
# ```
# Confusion matrices are tables used to analyze accuracy of classification models
# ```
#
# 3. What is Permutation Importances?
# ```
# Permutation Importances is another way to compute feature importance by calculating how much a feature impacts a model
# ```
#
# 4. What is feature isolation?
# ```
# Feature isolation is working with individual features to determine impact on the model while holding all other values constant
# ```
#
# 5. What is features interaction?
# ```
# features interaction takes place when features influence the behavior of other features
# ```
#
# 6. What is Shapley Values?
# ```
# a look into the "black box" of mystery that reveals the numerical contribution each feature adds to the final prediction of a single observation
# ```
#
# 7. How do you get ROC AUC score??
# ```
# this is calculated with a built in function of SciKit Learn's Metrics module passing in the y_true and predicted probabilities of the test or validation set.
# ```
#
# + [markdown] id="dUQaIwbceohq" colab_type="text"
# ## Practice Problems Choose a dataset of your own to fill this out.
# + [markdown] id="gzQyr1iPfJBH" colab_type="text"
# Preprocessing
# You may choose which features you want to use, and whether/how you will preprocess them.
# + id="e5hj6Jjwen3J" colab_type="code" colab={}
import pandas as pd
L = pd.read_csv('Landslides.csv')
# + id="kxgJk-v63t4Q" colab_type="code" colab={}
# + [markdown] id="rq6bizULj6YF" colab_type="text"
# Fit a model with the train set. Use your model to predict probabilities for the test set.
# + id="1jSaglmDkEZ7" colab_type="code" colab={}
# + [markdown] id="i2dHVZXjhlJS" colab_type="text"
# Get your model's validation accuracy.
# Get your model's test accuracy.
# Make visualizations for model interpretation.
# + id="bnt4a3p9ZYFz" colab_type="code" colab={}
# + [markdown] id="ijwt2MNXmONV" colab_type="text"
# ## what to study
# + [markdown] id="pcvFMpPy9cul" colab_type="text"
# # 1) Preprocessing
# # 2) Modeling
# # 3) Visualization
# # 4) Confusion Matrix
# # 5) permutation Importances
# # 6) Partial Dependence Plot
# # 7) feature isolation
# # 8) Shapley Values
# # 9) ROC AUC validation score
# # 10) categorical features
# # 11) encoding
#
# + id="xEfKdJQi_Eew" colab_type="code" colab={}
| Applied_Modeling_Study_Guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Семинар 5
#
# Исследуем зависимость положения разделяющей гиперплоскости в методе опорных векторов в зависимости от значения гиперпараметра $C$.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
class_size = 500
plt.figure(figsize=(20, 10))
mean0 = [7, 5]
cov0 = [[4, 0], [0, 1]] # diagonal covariance
mean1 = [0, 0]
cov1 = [[4, 0], [0, 2]]
data0 = np.random.multivariate_normal(mean0, cov0, class_size)
data1 = np.random.multivariate_normal(mean1, cov1, class_size)
data = np.vstack((data0, data1))
y = np.hstack((-np.ones(class_size), np.ones(class_size)))
plt.scatter(data0[:, 0], data0[:, 1], c='red', s=50)
plt.scatter(data1[:, 0], data1[:, 1], c='green', s=50)
plt.legend(['y = -1', 'y = 1'])
axes = plt.gca()
axes.set_xlim([-5, 15])
axes.set_ylim([-5, 10])
plt.show()
# +
from sklearn.svm import SVC
SVM_classifier = SVC(C=0.01, kernel='linear') # changing C here
SVM_classifier.fit(data, y)
# -
SVM_classifier.coef_[0][1]
SVM_classifier.intercept_[0]
# +
from sklearn.svm import SVC
SVM_classifier = SVC(C=100, kernel='linear') # changing C here
SVM_classifier.fit(data, y)
w_1 = SVM_classifier.coef_[0][0]
w_2 = SVM_classifier.coef_[0][1]
w_0 = SVM_classifier.intercept_[0]
plt.figure(figsize=(20,10))
plt.scatter(data0[:, 0], data0[:, 1], c='red', s=50)
plt.scatter(data1[:, 0], data1[:, 1], c='green', s=50)
plt.legend(['y = -1', 'y = 1'])
x_arr = np.linspace(-10, 15, 3000)
plt.plot(x_arr, -(w_0 + w_1 * x_arr) / w_2)
axes = plt.gca()
axes.set_xlim([-5,15])
axes.set_ylim([-5,10])
plt.show()
# +
plt.figure(figsize=(20,10))
plt.scatter(data0[:, 0], data0[:, 1], c='red', s=50, label='y = -1')
plt.scatter(data1[:, 0], data1[:, 1], c='green', s=50, label='y = +1')
#plt.legend(['y = -1', 'y = 1'])
x_arr = np.linspace(-10, 15, 3000)
colors = ['red', 'orange', 'green', 'blue', 'magenta']
for i, C in enumerate([0.0001, 0.01, 1, 100, 10000]):
SVM_classifier = SVC(C=C, kernel='linear')
SVM_classifier.fit(data, y)
w_1 = SVM_classifier.coef_[0][0]
w_2 = SVM_classifier.coef_[0][1]
w_0 = SVM_classifier.intercept_[0]
plt.plot(x_arr, -(w_0 + w_1 * x_arr) / w_2, color=colors[i], label='C='+str(C))
axes = plt.gca()
axes.set_xlim([-5,15])
axes.set_ylim([-5,10])
plt.legend(loc=0)
plt.show()
# -
# Гиперпараметр $C$ отвечает за то, что является более приоритетным для классификатора, — "подгонка" под обучающую выборку или максимизация ширины разделяющей полосы.
# - При больших значениях $C$ классификатор сильно настраивается на обучение, тем самым сужая разделяющую полосу.
# - При маленьких значениях $C$ классификатор расширяет разделяющую полосу, при этом допуская ошибки на некоторых объектах обучающей выборки.
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm
# Our dataset and targets
X = np.c_[(.4, -.7),
(-1.5, -1),
(-1.4, -.9),
(-1.3, -1.2),
(-1.1, -.2),
(-1.2, -.4),
(-.5, 1.2),
(-1.5, 2.1),
(1, 1),
# --
(1.3, .8),
(1.2, .5),
(.2, -2),
(.5, -2.4),
(.2, -2.3),
(0, -2.7),
(1.3, 2.1)].T
Y = [0] * 8 + [1] * 8
# figure number
fignum = 1
# fit the model
for kernel in ('linear', 'poly', 'rbf'):
clf = svm.SVC(kernel=kernel, gamma=2)
clf.fit(X, Y)
# plot the line, the points, and the nearest vectors to the plane
plt.figure(fignum, figsize=(4, 3))
plt.clf()
plt.scatter(clf.support_vectors_[:, 0], clf.support_vectors_[:, 1], s=80,
facecolors='none', zorder=10, edgecolors='k')
plt.scatter(X[:, 0], X[:, 1], c=Y, zorder=10, cmap=plt.cm.Paired,
edgecolors='k')
plt.axis('tight')
x_min = -3
x_max = 3
y_min = -3
y_max = 3
XX, YY = np.mgrid[x_min:x_max:200j, y_min:y_max:200j]
Z = clf.decision_function(np.c_[XX.ravel(), YY.ravel()])
# Put the result into a color plot
Z = Z.reshape(XX.shape)
plt.figure(fignum, figsize=(4, 3))
plt.pcolormesh(XX, YY, Z > 0, cmap=plt.cm.Paired)
plt.contour(XX, YY, Z, colors=['k', 'k', 'k'], linestyles=['--', '-', '--'],
levels=[-.5, 0, .5])
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
plt.title(kernel)
fignum = fignum + 1
plt.show()
# +
import numpy as np
np.random.seed(0)
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import LinearSVC
from sklearn.calibration import calibration_curve
X, y = datasets.make_classification(n_samples=100000, n_features=20,
n_informative=2, n_redundant=2)
train_samples = 100 # Samples used for training the models
X_train = X[:train_samples]
X_test = X[train_samples:]
y_train = y[:train_samples]
y_test = y[train_samples:]
# Create classifiers
lr = LogisticRegression()
gnb = GaussianNB()
svc = LinearSVC(C=1.0)
rfc = RandomForestClassifier()
# #############################################################################
# Plot calibration plots
plt.figure(figsize=(10, 10))
ax1 = plt.subplot2grid((3, 1), (0, 0), rowspan=2)
ax2 = plt.subplot2grid((3, 1), (2, 0))
ax1.plot([0, 1], [0, 1], "k:", label="Perfectly calibrated")
for clf, name in [(lr, 'Logistic'),
(gnb, 'Naive Bayes'),
(svc, 'Support Vector Classification'),
(rfc, 'Random Forest')]:
clf.fit(X_train, y_train)
if hasattr(clf, "predict_proba"):
prob_pos = clf.predict_proba(X_test)[:, 1]
else: # use decision function
prob_pos = clf.decision_function(X_test)
prob_pos = \
(prob_pos - prob_pos.min()) / (prob_pos.max() - prob_pos.min())
fraction_of_positives, mean_predicted_value = \
calibration_curve(y_test, prob_pos, n_bins=10)
ax1.plot(mean_predicted_value, fraction_of_positives, "s-",
label="%s" % (name, ))
ax2.hist(prob_pos, range=(0, 1), bins=10, label=name,
histtype="step", lw=2)
ax1.set_ylabel("Fraction of positives")
ax1.set_ylim([-0.05, 1.05])
ax1.legend(loc="lower right")
ax1.set_title('Calibration plots (reliability curve)')
ax2.set_xlabel("Mean predicted value")
ax2.set_ylabel("Count")
ax2.legend(loc="upper center", ncol=2)
plt.tight_layout()
plt.show()
| week5_lin-class/sem05-svm-viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Download and prepare the Credit Approval data set from the UCI Machine Learning Repository.
#
# **Citation:**
#
# <NAME>. and <NAME>. (2019). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.
#
# ============================================
#
# **Raw data download:**
#
# To download the Credit Approval dataset from the UCI Machine Learning Repository visit [this website](http://archive.ics.uci.edu/ml/machine-learning-databases/credit-screening/) and click on crx.data to download the data set. Save crx.data to the parent folder of this directory (../crx.data).
import random
import pandas as pd
import numpy as np
# +
# load data
data = pd.read_csv('crx.data', header=None)
# create variable names according to UCI Machine Learning
# Repo information
varnames = ['A'+str(s) for s in range(1,17)]
# add variable names to dataframe columns
data.columns = varnames
# replace ? by np.nan
data = data.replace('?', np.nan)
# display
data.head()
# +
# re-cast some variables to the correct types
data['A2'] = data['A2'].astype('float')
data['A14'] = data['A14'].astype('float')
# encode target to binary
data['A16'] = data['A16'].map({'+':1, '-':0})
# display
data.head()
# -
# find categorical variables
cat_cols = [c for c in data.columns if data[c].dtypes=='O']
data[cat_cols].head()
# find numerical variables
num_cols = [c for c in data.columns if data[c].dtypes!='O']
data[num_cols].head()
# +
# fill in missing values
data[num_cols] = data[num_cols].fillna(0)
data[cat_cols] = data[cat_cols].fillna('Missing')
data.isnull().sum()
# -
# save the data
data.to_csv('creditApprovalUCI.csv', index=False)
| Data Science and Machine Learning/Machine-Learning-In-Python-THOROUGH/MACHINE_LEARNING/PYTHON_FEATURE_ENGINEERING/Chapter03/00_CreditApprovalUCI_dataPrep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib nbagg
# %load_ext autoreload
# %autoreload 2
import os
import sys
import pickle
import nltk
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import OneHotEncoder
chatbot_path = "/home/bi0max/projects/tutorials/chatbot"
if chatbot_path not in sys.path:
sys.path.append(chatbot_path)
from chatbot.config import *
from chatbot.embed import *
from chatbot import embed
# -
# read file with all words from dataset
path = os.path.join(DATA_DIR, "all_words.pickle")
all_words = pickle.load(open(path, "rb"))
# calculate occurences of each word
freq_dist = nltk.FreqDist(all_words)
freq_dist
freq_list = list(freq_dist.values())
freq_list.sort(reverse=True)
plt.plot(freq_list)
plt.title("Number of occurences in descneding order")
n_unique_words = freq_dist.B()
print(f"Number unique words: {n_unique_words}")
# Overall words (not unique) in the dataset
# If we take certain threshold of most common words, which percentage of the overall amount they will cover
freq_array = np.array(freq_list)
overall_occurences = freq_array.sum()
print(f"Overall occurences: {overall_occurences}")
thresholds = [5000, 10000, 12000, 15000, 20000]#, 50000, 100000]
for threshold in thresholds:
top_words_occurences = freq_array[:threshold].sum()
percent_covered = top_words_occurences / overall_occurences
print(f"With threshold {threshold} percent covered: {percent_covered}")
# read GLOVE file
glove_word2index, glove_index2word, word2vec_map = embed.read_glove_vecs(GLOVE_MODEL)
# Find out which words from the dataset are present in GLOVE
in_glove = np.zeros(thresholds[-1])
glove_words = list(glove_word2index.keys())
top_words = [wf[0] for wf in freq_dist.most_common(thresholds[-1])]
for i, word in enumerate(top_words):
if i % 100 == 0:
print(f"{i} Done.")
if word in glove_words:
in_glove[i] = 1
for threshold in thresholds:
n_words_in_glove = in_glove[:threshold].sum()
print(f"For threshold {threshold} {threshold - n_words_in_glove} words are not in GLOVE.")
print("Top words, which are not in GLOVE:")
np.array(top_words)[:5000][~in_glove.astype(bool)[:5000]]
# ### We will use 12000 most common words, since they cover approx. 95% of all words.
| chatbot/choose_vocab_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Identify Unknown images
# ## Setup the environment
# nuclio: ignore
import nuclio
import os
# ### Set environment variables
# nuclio: ignore
os.environ['KV_TABLE'] = os.path.join(os.getenv('V3IO_USERNAME', 'iguazio'), 'demos/face-recognition/artifacts/encodings')
# Iguazio access
# %nuclio env V3IO_USERNAME=${V3IO_USERNAME}
# %nuclio env V3IO_ACCESS_KEY=${V3IO_ACCESS_KEY}
# %nuclio env KV_TABLE=${KV_TABLE}
# ### Set cron trigger
# +
# %nuclio config spec.triggers.secs.kind = "cron"
# %nuclio config spec.triggers.secs.attributes.interval = "1h"
# -
# ### Set iguazio mount
# %nuclio mount /User users/iguazio
# +
### Base image
# -
# %nuclio config spec.build.baseImage = "python:3.6-jessie"
# ### Installations
# When installing packages while working, Please reset the kernel to allow Jupyter to load the new packages.
# %%nuclio cmd -c
pip install v3io_frames
# ### Imports
# +
# Util
import v3io_frames as v3f
from datetime import datetime, timedelta
import os
import shutil
# DB
import v3io_frames as v3f
# -
# ## Function code
def handler(context, event):
kv_table_path = os.environ['KV_TABLE']
v3io_access_key = os.environ['V3IO_ACCESS_KEY']
client = v3f.Client("framesd:8081", token = v3io_access_key, container="users")
df = client.read(backend='kv', table=kv_table_path, reset_index=True)
context.logger.info(df.head())
df2 = df[['fileName', 'camera', 'label', 'imgUrl']]
options = ['unknown']
df3 = df2[df2.fileName.str.startswith(tuple(options))]
for idx in range(len(df3)):
img_url = df3.iloc[idx]['imgUrl']
splited = img_url.split("/")
destination = "/".join((splited[0],splited[1],splited[2],splited[3],"dataset/label_pending"))
print(img_url)
print(idx)
print(splited)
print(destination)
# Move the content of
# source to destination
dest = shutil.move(img_url, destination)
# %nuclio deploy -n unknown-labeling -p default -c
| faces/notebooks/unknown-labeling-trigger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training a ConvNet PyTorch
#
# In this notebook, you'll learn how to use the powerful PyTorch framework to specify a conv net architecture and train it on the CIFAR-10 dataset.
# +
import torch
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torch.utils.data import sampler
import torchvision.datasets as dset
import torchvision.transforms as T
import numpy as np
import timeit
# -
# ## What's this PyTorch business?
#
# You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
#
# For the last part of this assignment, though, we're going to leave behind your beautiful codebase and instead migrate to one of two popular deep learning frameworks: in this instance, PyTorch (or TensorFlow, if you switch over to that notebook).
#
# Why?
#
# * Our code will now run on GPUs! Much faster training. When using a framework like PyTorch or TensorFlow you can harness the power of the GPU for your own custom neural network architectures without having to write CUDA code directly (which is beyond the scope of this class).
# * We want you to be ready to use one of these frameworks for your project so you can experiment more efficiently than if you were writing every feature you want to use by hand.
# * We want you to stand on the shoulders of giants! TensorFlow and PyTorch are both excellent frameworks that will make your lives a lot easier, and now that you understand their guts, you are free to use them :)
# * We want you to be exposed to the sort of deep learning code you might run into in academia or industry.
# ## How will I learn PyTorch?
#
# If you've used Torch before, but are new to PyTorch, this tutorial might be of use: http://pytorch.org/tutorials/beginner/former_torchies_tutorial.html
#
# Otherwise, this notebook will walk you through much of what you need to do to train models in Torch. See the end of the notebook for some links to helpful tutorials if you want to learn more or need further clarification on topics that aren't fully explained here.
# ## Load Datasets
#
# We load the CIFAR-10 dataset. This might take a couple minutes the first time you do it, but the files should stay cached after that.
# +
class ChunkSampler(sampler.Sampler):
"""Samples elements sequentially from some offset.
Arguments:
num_samples: # of desired datapoints
start: offset where we should start selecting from
"""
def __init__(self, num_samples, start = 0):
self.num_samples = num_samples
self.start = start
def __iter__(self):
return iter(range(self.start, self.start + self.num_samples))
def __len__(self):
return self.num_samples
NUM_TRAIN = 49000
NUM_VAL = 1000
cifar10_train = dset.CIFAR10('./cs231n/datasets', train=True, download=True,
transform=T.ToTensor())
loader_train = DataLoader(cifar10_train, batch_size=64, sampler=ChunkSampler(NUM_TRAIN, 0))
cifar10_val = dset.CIFAR10('./cs231n/datasets', train=True, download=True,
transform=T.ToTensor())
loader_val = DataLoader(cifar10_val, batch_size=64, sampler=ChunkSampler(NUM_VAL, NUM_TRAIN))
cifar10_test = dset.CIFAR10('./cs231n/datasets', train=False, download=True,
transform=T.ToTensor())
loader_test = DataLoader(cifar10_test, batch_size=64)
# -
# For now, we're going to use a CPU-friendly datatype. Later, we'll switch to a datatype that will move all our computations to the GPU and measure the speedup.
# +
dtype = torch.FloatTensor # the CPU datatype
# Constant to control how frequently we print train loss
print_every = 100
# This is a little utility that we'll use to reset the model
# if we want to re-initialize all our parameters
def reset(m):
if hasattr(m, 'reset_parameters'):
m.reset_parameters()
# -
# ## Example Model
#
# ### Some assorted tidbits
#
# Let's start by looking at a simple model. First, note that PyTorch operates on Tensors, which are n-dimensional arrays functionally analogous to numpy's ndarrays, with the additional feature that they can be used for computations on GPUs.
#
# We'll provide you with a Flatten function, which we explain here. Remember that our image data (and more relevantly, our intermediate feature maps) are initially N x C x H x W, where:
# * N is the number of datapoints
# * C is the number of channels
# * H is the height of the intermediate feature map in pixels
# * W is the height of the intermediate feature map in pixels
#
# This is the right way to represent the data when we are doing something like a 2D convolution, that needs spatial understanding of where the intermediate features are relative to each other. When we input data into fully connected affine layers, however, we want each datapoint to be represented by a single vector -- it's no longer useful to segregate the different channels, rows, and columns of the data. So, we use a "Flatten" operation to collapse the C x H x W values per representation into a single long vector. The Flatten function below first reads in the N, C, H, and W values from a given batch of data, and then returns a "view" of that data. "View" is analogous to numpy's "reshape" method: it reshapes x's dimensions to be N x ??, where ?? is allowed to be anything (in this case, it will be C x H x W, but we don't need to specify that explicitly).
class Flatten(nn.Module):
def forward(self, x):
N, C, H, W = x.size() # read in N, C, H, W
return x.view(N, -1) # "flatten" the C * H * W values into a single vector per image
# ### The example model itself
#
# The first step to training your own model is defining its architecture.
#
# Here's an example of a convolutional neural network defined in PyTorch -- try to understand what each line is doing, remembering that each layer is composed upon the previous layer. We haven't trained anything yet - that'll come next - for now, we want you to understand how everything gets set up. nn.Sequential is a container which applies each layer
# one after the other.
#
# In that example, you see 2D convolutional layers (Conv2d), ReLU activations, and fully-connected layers (Linear). You also see the Cross-Entropy loss function, and the Adam optimizer being used.
#
# Make sure you understand why the parameters of the Linear layer are 5408 and 10.
#
# //Input size: 32x32, HH = WW = 1 + (32 - 7) / 2 = 13, out = 13 * 13 * 32 = 5408
#
# +
# Here's where we define the architecture of the model...
simple_model = nn.Sequential(
nn.Conv2d(3, 32, kernel_size=7, stride=2),
nn.ReLU(inplace=True),
Flatten(), # see above for explanation
nn.Linear(5408, 10), # affine layer
)
# Set the type of all data in this model to be FloatTensor
simple_model.type(dtype)
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.Adam(simple_model.parameters(), lr=1e-2) # lr sets the learning rate of the optimizer
# -
# PyTorch supports many other layer types, loss functions, and optimizers - you will experiment with these next. Here's the official API documentation for these (if any of the parameters used above were unclear, this resource will also be helpful). One note: what we call in the class "spatial batch norm" is called "BatchNorm2D" in PyTorch.
#
# * Layers: http://pytorch.org/docs/nn.html
# * Activations: http://pytorch.org/docs/nn.html#non-linear-activations
# * Loss functions: http://pytorch.org/docs/nn.html#loss-functions
# * Optimizers: http://pytorch.org/docs/optim.html#algorithms
# ## Training a specific model
#
# In this section, we're going to specify a model for you to construct. The goal here isn't to get good performance (that'll be next), but instead to get comfortable with understanding the PyTorch documentation and configuring your own model.
#
# Using the code provided above as guidance, and using the following PyTorch documentation, specify a model with the following architecture:
#
# * 7x7 Convolutional Layer with 32 filters and stride of 1
# * ReLU Activation Layer
# * Spatial Batch Normalization Layer
# * 2x2 Max Pooling layer with a stride of 2
# * Affine layer with 1024 output units
# * ReLU Activation Layer
# * Affine layer from 1024 input units to 10 outputs
#
# And finally, set up a **cross-entropy** loss function and the **RMSprop** learning rule.
# +
fixed_model_base = nn.Sequential( # You fill this in!
nn.Conv2d(3, 32, kernel_size=7, stride=1),
# (32-7)/1 + 1 = 26, 26*26*32=21632
nn.ReLU(inplace=True),
nn.BatchNorm2d(32),
nn.MaxPool2d(2, stride=2),
# 21632/4 = 5408
Flatten(),
nn.Linear(5408, 1024),
nn.ReLU(inplace=True),
nn.Linear(1024, 10)
)
fixed_model = fixed_model_base.type(dtype)
loss_fn = nn.CrossEntropyLoss().type(dtype)
optimizer = optim.RMSprop(simple_model.parameters(), lr=1e-2)
# -
# To make sure you're doing the right thing, use the following tool to check the dimensionality of your output (it should be 64 x 10, since our batches have size 64 and the output of the final affine layer should be 10, corresponding to our 10 classes):
# +
## Now we're going to feed a random batch into the model you defined and make sure the output is the right size
x = torch.randn(64, 3, 32, 32).type(dtype)
x_var = Variable(x.type(dtype)) # Construct a PyTorch Variable out of your input data
ans = fixed_model(x_var) # Feed it through the model!
# Check to make sure what comes out of your model
# is the right dimensionality... this should be True
# if you've done everything correctly
np.array_equal(np.array(ans.size()), np.array([64, 10]))
# -
# ### GPU!
#
# Now, we're going to switch the dtype of the model and our data to the GPU-friendly tensors, and see what happens... everything is the same, except we are casting our model and input tensors as this new dtype instead of the old one.
#
# If this returns false, or otherwise fails in a not-graceful way (i.e., with some error message), you may not have an NVIDIA GPU available on your machine. If you're running locally, we recommend you switch to Google Cloud and follow the instructions to set up a GPU there. If you're already on Google Cloud, something is wrong -- make sure you followed the instructions on how to request and use a GPU on your instance. If you did, post on Piazza or come to Office Hours so we can help you debug.
# +
# Verify that CUDA is properly configured and you have a GPU available
torch.cuda.is_available()
# +
import copy
gpu_dtype = torch.cuda.FloatTensor
fixed_model_gpu = copy.deepcopy(fixed_model_base).type(gpu_dtype)
x_gpu = torch.randn(64, 3, 32, 32).type(gpu_dtype)
x_var_gpu = Variable(x.type(gpu_dtype)) # Construct a PyTorch Variable out of your input data
ans = fixed_model_gpu(x_var_gpu) # Feed it through the model!
# Check to make sure what comes out of your model
# is the right dimensionality... this should be True
# if you've done everything correctly
np.array_equal(np.array(ans.size()), np.array([64, 10]))
# -
# Run the following cell to evaluate the performance of the forward pass running on the CPU:
# %%timeit
ans = fixed_model(x_var)
# ... and now the GPU:
# %%timeit
torch.cuda.synchronize() # Make sure there are no pending GPU computations
ans = fixed_model_gpu(x_var_gpu) # Feed it through the model!
torch.cuda.synchronize() # Make sure there are no pending GPU computations
# You should observe that even a simple forward pass like this is significantly faster on the GPU. So for the rest of the assignment (and when you go train your models in assignment 3 and your project!), you should use the GPU datatype for your model and your tensors: as a reminder that is *torch.cuda.FloatTensor* (in our notebook here as *gpu_dtype*)
# ### Train the model.
#
# Now that you've seen how to define a model and do a single forward pass of some data through it, let's walk through how you'd actually train one whole epoch over your training data (using the simple_model we provided above).
#
# Make sure you understand how each PyTorch function used below corresponds to what you implemented in your custom neural network implementation.
#
# Note that because we are not resetting the weights anywhere below, if you run the cell multiple times, you are effectively training multiple epochs (so your performance should improve).
#
# First, set up an RMSprop optimizer (using a 1e-3 learning rate) and a cross-entropy loss function:
loss_fn = None
optimizer = None
pass
# +
# This sets the model in "training" mode. This is relevant for some layers that may have different behavior
# in training mode vs testing mode, such as Dropout and BatchNorm.
fixed_model_gpu.train()
# Load one batch at a time.
for t, (x, y) in enumerate(loader_train):
x_var = Variable(x.type(gpu_dtype))
y_var = Variable(y.type(gpu_dtype).long())
# This is the forward pass: predict the scores for each class, for each x in the batch.
scores = fixed_model_gpu(x_var)
# Use the correct y values and the predicted y values to compute the loss.
loss = loss_fn(scores, y_var)
if (t + 1) % print_every == 0:
print('t = %d, loss = %.4f' % (t + 1, loss.data[0]))
# Zero out all of the gradients for the variables which the optimizer will update.
optimizer.zero_grad()
# This is the backwards pass: compute the gradient of the loss with respect to each
# parameter of the model.
loss.backward()
# Actually update the parameters of the model using the gradients computed by the backwards pass.
optimizer.step()
# -
# Now you've seen how the training process works in PyTorch. To save you writing boilerplate code, we're providing the following helper functions to help you train for multiple epochs and check the accuracy of your model:
# +
def train(model, loss_fn, optimizer, num_epochs = 1):
for epoch in range(num_epochs):
print('Starting epoch %d / %d' % (epoch + 1, num_epochs))
model.train()
for t, (x, y) in enumerate(loader_train):
x_var = Variable(x.type(gpu_dtype))
y_var = Variable(y.type(gpu_dtype).long())
scores = model(x_var)
loss = loss_fn(scores, y_var)
if (t + 1) % print_every == 0:
print('t = %d, loss = %.4f' % (t + 1, loss.data[0]))
optimizer.zero_grad()
loss.backward()
optimizer.step()
def check_accuracy(model, loader):
if loader.dataset.train:
print('Checking accuracy on validation set')
else:
print('Checking accuracy on test set')
num_correct = 0
num_samples = 0
model.eval() # Put the model in test mode (the opposite of model.train(), essentially)
for x, y in loader:
x_var = Variable(x.type(gpu_dtype), volatile=True)
scores = model(x_var)
_, preds = scores.data.cpu().max(1)
num_correct += (preds == y).sum()
num_samples += preds.size(0)
acc = float(num_correct) / num_samples
print('Got %d / %d correct (%.2f)' % (num_correct, num_samples, 100 * acc))
# -
# ### Check the accuracy of the model.
#
# Let's see the train and check_accuracy code in action -- feel free to use these methods when evaluating the models you develop below.
#
# You should get a training loss of around 1.2-1.4, and a validation accuracy of around 50-60%. As mentioned above, if you re-run the cells, you'll be training more epochs, so your performance will improve past these numbers.
#
# But don't worry about getting these numbers better -- this was just practice before you tackle designing your own model.
torch.cuda.random.manual_seed(12345)
fixed_model_gpu.apply(reset)
train(fixed_model_gpu, loss_fn, optimizer, num_epochs=1)
check_accuracy(fixed_model_gpu, loader_val)
# ### Don't forget the validation set!
#
# And note that you can use the check_accuracy function to evaluate on either the test set or the validation set, by passing either **loader_test** or **loader_val** as the second argument to check_accuracy. You should not touch the test set until you have finished your architecture and hyperparameter tuning, and only run the test set once at the end to report a final value.
# ## Train a _great_ model on CIFAR-10!
#
# Now it's your job to experiment with architectures, hyperparameters, loss functions, and optimizers to train a model that achieves **>=70%** accuracy on the CIFAR-10 **validation** set. You can use the check_accuracy and train functions from above.
# ### Things you should try:
# - **Filter size**: Above we used 7x7; this makes pretty pictures but smaller filters may be more efficient
# - **Number of filters**: Above we used 32 filters. Do more or fewer do better?
# - **Pooling vs Strided Convolution**: Do you use max pooling or just stride convolutions?
# - **Batch normalization**: Try adding spatial batch normalization after convolution layers and vanilla batch normalization after affine layers. Do your networks train faster?
# - **Network architecture**: The network above has two layers of trainable parameters. Can you do better with a deep network? Good architectures to try include:
# - [conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
# - [conv-relu-conv-relu-pool]xN -> [affine]xM -> [softmax or SVM]
# - [batchnorm-relu-conv]xN -> [affine]xM -> [softmax or SVM]
# - **Global Average Pooling**: Instead of flattening and then having multiple affine layers, perform convolutions until your image gets small (7x7 or so) and then perform an average pooling operation to get to a 1x1 image picture (1, 1 , Filter#), which is then reshaped into a (Filter#) vector. This is used in [Google's Inception Network](https://arxiv.org/abs/1512.00567) (See Table 1 for their architecture).
# - **Regularization**: Add l2 weight regularization, or perhaps use Dropout.
#
# ### Tips for training
# For each network architecture that you try, you should tune the learning rate and regularization strength. When doing this there are a couple important things to keep in mind:
#
# - If the parameters are working well, you should see improvement within a few hundred iterations
# - Remember the coarse-to-fine approach for hyperparameter tuning: start by testing a large range of hyperparameters for just a few training iterations to find the combinations of parameters that are working at all.
# - Once you have found some sets of parameters that seem to work, search more finely around these parameters. You may need to train for more epochs.
# - You should use the validation set for hyperparameter search, and save your test set for evaluating your architecture on the best parameters as selected by the validation set.
#
# ### Going above and beyond
# If you are feeling adventurous there are many other features you can implement to try and improve your performance. You are **not required** to implement any of these; however they would be good things to try for extra credit.
#
# - Alternative update steps: For the assignment we implemented SGD+momentum, RMSprop, and Adam; you could try alternatives like AdaGrad or AdaDelta.
# - Alternative activation functions such as leaky ReLU, parametric ReLU, ELU, or MaxOut.
# - Model ensembles
# - Data augmentation
# - New Architectures
# - [ResNets](https://arxiv.org/abs/1512.03385) where the input from the previous layer is added to the output.
# - [DenseNets](https://arxiv.org/abs/1608.06993) where inputs into previous layers are concatenated together.
# - [This blog has an in-depth overview](https://chatbotslife.com/resnets-highwaynets-and-densenets-oh-my-9bb15918ee32)
#
# If you do decide to implement something extra, clearly describe it in the "Extra Credit Description" cell below.
#
# ### What we expect
# At the very least, you should be able to train a ConvNet that gets at least 70% accuracy on the validation set. This is just a lower bound - if you are careful it should be possible to get accuracies much higher than that! Extra credit points will be awarded for particularly high-scoring models or unique approaches.
#
# You should use the space below to experiment and train your network.
#
# Have fun and happy training!
# +
# Train your model here, and make sure the output of this cell is the accuracy of your best model on the
# train, val, and test sets. Here's some code to get you started. The output of this cell should be the training
# and validation accuracy on your best model (measured by validation accuracy).
model = None
loss_fn = None
optimizer = None
train(model, loss_fn, optimizer, num_epochs=1)
check_accuracy(model, loader_val)
# -
# ### Describe what you did
#
# In the cell below you should write an explanation of what you did, any additional features that you implemented, and any visualizations or graphs that you make in the process of training and evaluating your network.
# Tell us here!
# ## Test set -- run this only once
#
# Now that we've gotten a result we're happy with, we test our final model on the test set (which you should store in best_model). This would be the score we would achieve on a competition. Think about how this compares to your validation set accuracy.
best_model = None
check_accuracy(best_model, loader_test)
# ## Going further with PyTorch
#
# The next assignment will make heavy use of PyTorch. You might also find it useful for your projects.
#
# Here's a nice tutorial by <NAME> that shows off some of PyTorch's features, like dynamic graphs and custom NN modules: http://pytorch.org/tutorials/beginner/pytorch_with_examples.html
#
# If you're interested in reinforcement learning for your final project, this is a good (more advanced) DQN tutorial in PyTorch: http://pytorch.org/tutorials/intermediate/reinforcement_q_learning.html
| assignment2/Q5_PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 高斯核函数
import numpy as np
import matplotlib.pyplot as plt
x = np.arange(-4, 5, 1)
x
y = np.array((x >= -2) & (x <= 2), dtype=int)
y
plt.scatter(x[y==0], [0]*len(x[y==0]))
plt.scatter(x[y==1], [0]*len(x[y==1]))
plt.show()
# $$K(x, y) = e ^ {-\gamma ||x-y||^2}$$
# m * n 数据经过高斯后变成为 m * m 数据
def gaussian(x, l):
gamma = 1.0
return np.exp(-gamma * (x - l) ** 2)
# +
l1, l2 = -1, 1
X_new = np.empty((len(x), 2))
for i, data in enumerate(x):
X_new[i, 0] = gaussian(data, l1)
X_new[i, 1] = gaussian(data, l2)
# -
plt.scatter(X_new[y==0, 0], X_new[y==0, 1])
plt.scatter(X_new[y==1, 0], X_new[y==1, 1])
plt.show()
| ml/svm/rbf-kernel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env
# language: python
# name: env
# ---
from pymodulo.SpatialStratification import GridStratification
from pymodulo.DataLoader import CSVDataLoader
from pymodulo.VehicleSelector import GreedyVehicleSelector, MaxPointsVehicleSelector, RandomVehicleSelector
# ## Spatial stratification process
# In this example, we do a grid stratification. At this step, you need to decide the spatial granularity. Since this example uses a grid stratification, we need to decide the length of each side of a grid. In the following example, we keep this length as 1 km.
# Here, we keep a cellSide of length 1 km (the first argument)
spatial = GridStratification(1, 77.58467674255371, 12.958180959662695, 77.60617733001709, 12.977167633046893)
spatial.stratify()
# Now, `spatial.input_geojson` returns the GeoJSON containing the strata (along with stratum ID). Below, we print the first stratum that was generated. If desired, you can store this GeoJSON using the in-built Python `json` library.
spatial.input_geojson['features'][0]
# ## Data loading process
#
# In this step, we upload the vehicle mobility data to a [MongoDB](https://docs.mongodb.com/) database. You need to take care of a few things here:
# 1. You must ensure that you have a MongoDB server (local or remote) running before you continue with this process.
# 2. The input CSV file must containing the following columns: vehicle_id, timestamp, latitude, longitude.
# 3. You will need to decide upon a `temporal_granularity` (in seconds). In this example, we use a temporal granularity of 1 hour (= 3600 seconds).
# 4. Decide the database name and a collection name (inside that database) that you want to upload your data to.
dataloader = CSVDataLoader('sample_mobility_data.csv', 3600,
anonymize_data=False,
mongo_uri='mongodb://localhost:27017/',
db_name='modulo',
collection_name='mobility_data')
# At this point, if you want, you can check your MongoDB database using a [MongoDB GUI](https://www.mongodb.com/products/compass). You should see your data uploaded in the database.
#
# Now, we need to compute the stratum ID that each vehicle mobility datum falls into. Similarly, we also need to calculate the temporal ID that each datum falls into. Think of the temporal ID as referring to a "time bucket", each of length `temporal_granularity`. Both these methods return the number of records that were updated with the `stratum_id` and the `temporal_id` respectively.
dataloader.compute_stratum_id_and_update_db(spatial)
dataloader.compute_temporal_id_and_update_db()
# You can use the following helper function to fetch the vehicle mobility data stored in the database. This function will return the stored values as a Pandas DataFrame, which you can conveniently use to do any checks, operations, analysis, etc.
df = dataloader.fetch_data()
df.head()
# ## Vehicle Selection
#
# Now, we can finally use the available algorithms to select the desired number of vehicles. In the following example, we assume that we want to choose 2 vehicles.
#
# The vehicle selection ("training") process requires the vehicle mobility data from the database. We use another helper method in `DataLoader` to fetch this data as a Pandas DataFrame.
selection_df = dataloader.fetch_data_for_vehicle_selection()
# Using greedy
greedy = GreedyVehicleSelector(2, selection_df, 1589389199)
selected_vehicles = greedy.train()
greedy.test(selected_vehicles)
# Using max-points
maxpoints = MaxPointsVehicleSelector(2, selection_df, 1589389199)
selected_vehicles = maxpoints.train()
maxpoints.test(selected_vehicles)
# Using random
random_algo = RandomVehicleSelector(2, selection_df, 1589389199)
selected_vehicles = random_algo.train()
random_algo.test(selected_vehicles)
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Funciones de utilidad y aversión al riesgo
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/6/62/Risk_down_arrow.png" width="400px" height="400px" />
#
# En el módulo anterior aprendimos
# - qué es un portafolio, cómo medir su rendimiento esperado y su volatilidad;
# - un portafolio de activos riesgosos tiene menos riesgo que la suma de los riesgos individuales,
# - y que esto se logra mediante el concepto de diversificación;
# - la diversificación elimina el riesgo idiosincrático, que es el que afecta a cada compañía en particular,
# - sin embargo, el riesgo de mercado no se puede eliminar porque afecta a todos por igual.
# - Finalmente, aprendimos conceptos importantes como frontera de mínima varianza, portafolios eficientes y el portafolio de mínima varianza, que son claves en el problema de selección óptima de portafolios.
#
# Muy bien, sin embargo, para plantear el problema de selección óptima de portafolios necesitamos definir la función que vamos a optimizar: función de utilidad.
#
# **Objetivos:**
# - ¿Cómo tomamos decisiones según los economistas?
# - ¿Cómo toman decisiones los inversionistas?
# - ¿Qué son las funciones de utilidad?
#
# *Referencia:*
# - Notas del curso "Portfolio Selection and Risk Management", Rice University, disponible en Coursera.
# ___
# ## 1. Introducción
#
# La teoría económica comienza con una suposición muy importante:
# - **cada individuo actúa para obtener el mayor beneficio posible con los recursos disponibles**.
# - En otras palabras, **maximizan su propia utilidad**
# ¿Qué es utilidad?
# - Es un concepto relacionado con la felicidad, pero más amplio.
# - Por ejemplo, yo obtengo utilidad de lavar mis dientes o comer sano. Ninguna de las dos me brindan felicidad, pero lo primero mantendrá mis dientes sanos y en el largo plazo, lo segundo probablemente contribuirá a una buena vejez.
# Los economistas no se preocupan en realidad por lo que nos da utilidad, sino simplemente que cada uno de nosotros tiene sus propias preferencias.
# - Por ejemplo, a mi me gusta el café, el fútbol, los perros, la academia, viajar, entre otros.
# - Ustedes tienen sus propias preferencias también.
# La vida es compleja y con demasiada incertidumbre. Debemos tomar decisiones a cada momento, y estas decisiones involucran ciertos "trade-off".
# - Por ejemplo, normalmente tenemos una compensación entre utilidad hoy contra utilidad en el futuro.
# - Debemos balancear nuestro consumo hoy contra nuestro consumo luego.
# - Por ejemplo, ustedes gastan cerca de cuatro horas a la semana viniendo a clases de portafolios, porque esperan que esto contribuya a mejorar su nivel de vida en el futuro.
# De manera que los economistas dicen que cada individuo se comporta como el siguiente optimizador:
#
# \begin{align}
# \max & \quad\text{Utilidad}\\
# \text{s. a.} & \quad\text{Recursos disponibles}
# \end{align}
#
# ¿Qué tiene que ver todo esto con el curso?
# - En este módulo desarrollaremos herramientas para describir las preferencias de los inversionistas cuando se encuentran con decisiones de riesgo y rendimiento.
# - Veremos como podemos medir la actitud frente al riesgo, ¿cuánto te gusta o disgusta el riesgo?
# - Finalmente, veremos como podemos formular el problema de maximizar la utilidad de un inversionista para tomar la decisión de inversión óptima.
# ___
# ## 2. Funciones de utilidad.
#
# ¿Cómo tomamos decisiones?
# Por ejemplo:
# - Ustedes tienen que decidir si venir a clase o quedarse en su casa viendo Netflix, o ir al gimnasio.
# - Tienen que decidir entre irse de fiesta cada fin, o ahorrar para salir de vacaciones.
# En el caso de un portafolio, la decisión que se debe tomar es **¿cuáto riesgo estás dispuesto a tomar por qué cantidad de rendimiento?**
#
# **¿Cómo evaluarías el "trade-off" entre tener cetes contra una estrategia muy riesgosa con un posible altísimo rendimiento?**
# De manera que veremos como tomamos decisiones cuando tenemos distintas posibilidades. Específicamente, hablaremos acerca de las **preferencias**, como los economistas usan dichas preferencias para explicar las decisiones y los "trade-offs" en dichas decisiones.
#
# Usamos las **preferencias** para describir las decisiones que tomamos. Las preferencias nos dicen cómo un individuo evalúa los "trade-offs" entre distintas elecciones.
# Por definición, las preferencias son únicas para cada individuo. En el problema de selección de portafolios:
# - las preferencias que dictan cuánto riesgo estás dispuesto a asumir por cuánto rendimiento, son específicas para cada uno de ustedes.
# - Sus respuestas a esa pregunta pueden ser muy distintas, porque tenemos distintas preferencias.
#
# Ahora, nosotros no podemos *cuantificar* dichas preferencias.
# - Por esto usamos el concepto de utilidad, para medir qué tan satisfecho está un individuo con sus elecciones.
# - Así que podemos pensar en la utilidad como un indicador numérico que describe las preferencias,
# - o un índice que nos ayuda a clasificar diferentes decisiones.
# - En términos simples, **la utilidad nos ayuda a transmitir a números la noción de cómo te sientes**;
# - mientras más utilidad, mejor te sientes.
# **Función de utilidad**: manera sistemática de asignar una medida o indicador numérico para clasificar diferentes escogencias.
#
# El número que da una función de utilidad no tiene significado alguno. Simplemente es una manera de clasificar diferentes decisiones.
# **Ejemplo.**
#
# Podemos escribir la utilidad de un inversionista como función de la riqueza,
#
# $$U(W).$$
#
# - $U(W)$ nos da una medida de qué tan satisfechos estamos con el nivel de riqueza que tenemos.
# - $U(W)$ no es la riqueza como tal, sino que la función de utilidad traduce la cantidad de riqueza en un índice numérico subjetivo.
# ¿Cómo luciría gráficamente una función de utilidad de riqueza $U(W)$?
#
# <font color=blue> Ver en el tablero </font>
# - ¿Qué caracteristicas debe tener?
# - ¿Cómo es su primera derivada?
# - ¿Cómo es su segunda derivada?
# - Tiempos buenos: riqueza alta (¿cómo es la primera derivada acá?)
# - Tiempos malos: poca riqueza (¿cómo es la primera derivada acá?)
# ## 3. Aversión al riesgo
#
# Una dimensión importante en la toma de decisiones en finanzas y economía es la **incertidumbre**. Probablemente no hay ninguna decisión en economía que no involucre riesgo.
#
# - A la mayoría de las personas no les gusta mucho el riesgo.
# - De hecho, estudios del comportamiento humano de cara al riesgo, sugieren fuertemente que los seres humanos somos aversos al riesgo.
# - Por ejemplo, la mayoría de hogares poseen seguros para sus activos.
# - Así, cuando planteamos el problema de selección óptima de portafolios, suponemos que el inversionista es averso al riesgo.
# ¿Qué significa esto en términos de preferencias? ¿Cómo lo medimos?
#
# - Como seres humanos, todos tenemos diferentes genes y preferencias, y esto aplica también a la actitud frente al riesgo.
# - Por tanto, la aversión al riesgo es clave en cómo describimos las preferencias de un inversinista.
# - Individuos con un alto grado de aversión al riesgo valorarán la seguridad a un alto precio, mientras otros no tanto.
# - De manera que alguien con alta aversión al riesgo, no querrá enfrentarse a una situación con resultado incierto y querrá pagar una gran prima de seguro para eliminar dicho riesgo.
# - O equivalentemente, una persona con alta aversión al riesgo requerirá una compensación alta si se decide a asumir ese riesgo.
#
# El **grado de aversión al riesgo** mide qué tanto un inversionista prefiere un resultado seguro a un resultado incierto.
#
# Lo opuesto a aversión al riesgo es **tolerancia al riesgo**.
#
# <font color=blue> Ver en el tablero gráficamente, cómo se explica la aversión al riesgo desde las funciones de utilidad. </font>
#
# **Conclusión:** la concavidad en la función de utilidad dicta qué tan averso al riesgo es el individuo.
# ### ¿Cómo medimos el grado de aversión al riesgo de un individuo?
#
# ¿Saben cuál es su coeficiente de aversión al riesgo? Podemos estimarlo.
#
# Suponga que se puede participar en la siguiente lotería:
# - usted puede ganar $\$1000$ con $50\%$ de probabilidad, o
# - puede ganar $\$500$ con $50\%$ de probabilidad.
#
# Es decir, de entrada usted tendrá $\$500$ seguros pero también tiene la posibilidad de ganar $\$1000$.
#
# ¿Cuánto estarías dispuesto a pagar por esta oportunidad?
# Bien, podemos relacionar tu respuesta con tu coeficiente de aversión al riesgo.
#
# | Coeficiente de aversión al riesgo | Cantidad que pagarías |
# | --------------------------------- | --------------------- |
# | 0 | 750 |
# | 0.5 | 729 |
# | 1 | 707 |
# | 2 | 667 |
# | 3 | 632 |
# | 4 | 606 |
# | 5 | 586 |
# | 10 | 540 |
# | 15 | 525 |
# | 20 | 519 |
# | 50 | 507 |
# La mayoría de la gente está dispuesta a pagar entre $\$540$ (10) y $\$707$ (1). Es muy raro encontrar coeficientes de aversión al riesgo menores a 1. Esto está soportado por una gran cantidad de encuestas.
#
# - En el mundo financiero, los consultores financieros utilizan cuestionarios para medir el coeficiente de aversión al riesgo.
# **Ejemplo.** Describir en términos de aversión al riesgo las siguientes funciones de utilidad que dibujaré en el tablero.
# ___
# # Anuncios
#
# ## 1. Quiz la siguiente clase.
# ## 2. Examen Módulos 1 y 2: Martes 19 de Marzo.
# ## 3. Recordar Tarea 5 para este viernes, 15 de Marzo.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo3/Clase11_FuncionesUtilidad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import kapre
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Lambda, Permute
from kapre.time_frequency import Melspectrogram
from kapre.utils import Normalization2D
import numpy as np
import librosa
import matplotlib.pyplot as plt
def visualise_model(model, logam=False):
n_ch, nsp_src = model.input_shape[1:]
src, _ = librosa.load('../clean/Atlantic Spotted Dolphin/6102500A_2.wav', sr=SR, mono=True)
src = src[:nsp_src]
src_batch = src[np.newaxis, np.newaxis, :]
pred = model.predict(x=src_batch)
if tf.keras.backend.image_data_format() == 'channels_first':
result = pred[0, 0]
else:
result = pred[0, :, :, 0]
if logam:
result = librosa.power_to_db(result)
fig, ax = plt.subplots(figsize=(10,8))
ax.set_title('Normalized Frequency Spectrogram', size=20)
ax.imshow(result)
ax.set_ylabel('Mel bins', size=18)
ax.set_xlabel('Time (10 ms)', size=18)
ax.xaxis.set_tick_params(labelsize=14)
ax.yaxis.set_tick_params(labelsize=14)
plt.show()
return result
SR = 16000
src = np.random.random((1, SR))
model = Sequential()
model.add(Melspectrogram(sr=SR, n_mels=128,
n_dft=512, n_hop=160, input_shape=src.shape,
return_decibel_melgram=True,
trainable_kernel=False, name='melgram'))
model.add(Normalization2D(str_axis='batch'))
#model.add(Permute((2,1,3)))
model.summary()
# +
X = visualise_model(model)
plt.title('Normalized Frequency Histogram')
plt.hist(X.flatten(), bins='auto')
plt.show()
# -
| notebooks/Kapre Mel Spectrogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LoRa Data Analysis - UCB vs. TS
#
# We first declare a fixed parameters.
#
# Those parameters are not changed during the experiments.
#
# Fixed communication parameters are listed below:
# - Code Rate: 4/5
# - Frequency: 866.1 MHz
# - Bandwidth: 125 kHz
#
# End nodes:
# - were sending different types of uplink messages
# - were sending a single message each 2 minutes
# - comparison of upper confidence bound algorithm (UCB) and Thompson sampling (TS)
#
# Access points:
# - only a single access point was used
# - capture effect was also considered
# Initial declaration
# +
# %matplotlib inline
import pandas as pd # import pandas
import numpy as np # import numpy
import matplotlib as mpl # import matplotlib
import matplotlib.pyplot as plt # import plotting module
import statistics
import math
import base64
from IPython.display import set_matplotlib_formats # module for svg export
# Set output format for png figures
output_format = 'png'
set_matplotlib_formats(output_format) # set export to svg file
ts_uplink_file = 'ts_uplink_messages.csv'
ucb_uplink_file = 'ucb_uplink_messages.csv'
# -
# ## Analysis of Uplink Messages
# We read a csv file with uplink messages
ts_uplink_messages = pd.read_csv(ts_uplink_file, delimiter=',')
ucb_uplink_messages = pd.read_csv(ucb_uplink_file, delimiter=',')
# Let us have a look at various columns that are present and can be evaluated.
ts_uplink_messages.head()
ucb_uplink_messages.head()
# Remove all columns that have fixed values or there is no point in their analysis.
try:
del ts_uplink_messages['id']
del ts_uplink_messages['msg_group_number']
del ts_uplink_messages['is_primary']
del ts_uplink_messages['coderate']
del ts_uplink_messages['bandwidth']
del ts_uplink_messages['receive_time']
except KeyError:
print('Columns have already been removed')
try:
del ucb_uplink_messages['id']
del ucb_uplink_messages['msg_group_number']
del ucb_uplink_messages['is_primary']
del ucb_uplink_messages['coderate']
del ucb_uplink_messages['bandwidth']
del ucb_uplink_messages['receive_time']
except KeyError:
print('Columns have already been removed')
# ### Payload Length
adr_uplink_messages['payload_len'] = adr_uplink_messages.app_data.apply(len)
ucb_uplink_messages['payload_len'] = ucb_uplink_messages.app_data.apply(len)
# +
adr_payload_len = round(statistics.mean(adr_uplink_messages.payload_len))
ucb_payload_len = round(statistics.mean(ucb_uplink_messages.payload_len))
print(f'Mean value of payload length for ADR is {adr_payload_len} B')
print(f'Mean value of payload length for UCB is {ucb_payload_len} B')
# -
# ### Spreading Factor
# +
sf1 = adr_uplink_messages.spf.value_counts()
sf2 = ucb_uplink_messages.spf.value_counts()
diff = abs(sf1 - sf2)
diff.fillna(0)
sf_adr = [sf1, diff]
sf_adr = pd.concat(sf_adr, axis=1, sort=False).sum(axis=1)
sf_adr.sort_index(ascending=False, inplace=True)
diff = abs(sf2 - sf1)
diff.fillna(0)
sf_ucb = [sf2, diff]
sf_ucb = pd.concat(sf_ucb, axis=1, sort=False).sum(axis=1)
sf_ucb.sort_index(ascending=False, inplace=True)
# +
# Create a grouped bar chart, with job as the x-axis
# and gender as the variable we're grouping on so there
# are two bars per job.
fig, ax = plt.subplots(figsize=(10, 4))
# Define bar width. We need this to offset the second bar.
bar_width = 0.3
index = np.arange(len(sf_adr))
ax.bar(index, sf_adr, width=bar_width, color='green', label = 'ADR')
# Same thing, but offset the x.
ax.bar(index + bar_width, sf_ucb, width=bar_width, color='blue', label = 'UCB')
# Fix the x-axes.
ax.set_xticks(index + bar_width / 2)
ax.set_xticklabels(sf_ucb.index)
# Add legend.
ax.legend()
# Axis styling.
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_color('#DDDDDD')
ax.tick_params(bottom=False, left=False)
ax.set_axisbelow(True)
ax.yaxis.grid(True, color='#EEEEEE')
ax.xaxis.grid(False)
# Add axis and chart labels.
ax.set_xlabel('Spreading Factor', labelpad=15)
ax.set_ylabel('Number of Messages', labelpad=15)
ax.set_title('Utilization of Spreading Factor', pad=15)
fig.tight_layout()
# For each bar in the chart, add a text label.
for bar in ax.patches:
# The text annotation for each bar should be its height.
bar_value = round(bar.get_height())
# Format the text with commas to separate thousands. You can do
# any type of formatting here though.
text = f'{bar_value:,}'
# This will give the middle of each bar on the x-axis.
text_x = bar.get_x() + bar.get_width() / 2
# get_y() is where the bar starts so we add the height to it.
text_y = bar.get_y() + bar_value
# If we want the text to be the same color as the bar, we can
# get the color like so:
bar_color = bar.get_facecolor()
# If you want a consistent color, you can just set it as a constant, e.g. #222222
ax.text(text_x, text_y, text, ha='center', va='bottom', color=bar_color, size=10)
fig.savefig(f'adr-ucb-sf.{output_format}', dpi=300)
# -
# All nodes used the same frequency to increase a probability of collisions.
# We have only a single Access Point.
# ## Analysis of End Nodes
# Analysis of certain aspects (active time, sleep time and collisions) of end devices.
# +
adr_unique_ens = adr_uplink_messages.node_id.nunique()
ucb_unique_ens = ucb_uplink_messages.node_id.nunique()
print(f'Number of end nodes participating for ADR is {adr_unique_ens}.')
print(f'Number of end nodes participating for UCB is {ucb_unique_ens}.')
# -
adr_end_nodes = pd.read_csv(f'adr_end_nodes.csv', delimiter=',')
ucb_end_nodes = pd.read_csv(f'ucb_end_nodes.csv', delimiter=',')
# ### Collision Ratio
# +
adr_collisions = adr_end_nodes.collisions
ucb_collisions = ucb_end_nodes.collisions
adr_max_collisions = max(adr_end_nodes.collisions)
adr_min_collisions = min(adr_end_nodes.collisions)
ucb_max_collisions = max(ucb_end_nodes.collisions)
ucb_min_collisions = min(ucb_end_nodes.collisions)
max_collisions = max(adr_max_collisions, ucb_max_collisions)
min_collisions = min(adr_min_collisions, ucb_min_collisions)
range_collisions = max_collisions - min_collisions
buckets = 8
increment = range_collisions / buckets
print(f'Max number of collisions for ADR: {adr_max_collisions}')
print(f'Min number of collisions for ADR: {adr_min_collisions}')
print(f'Max number of collisions for UCB: {ucb_max_collisions}')
print(f'Min number of collisions for UCB: {ucb_min_collisions}')
# +
fig, ax = plt.subplots(figsize=(10, 4))
bar_width = 0.4
index = np.arange(buckets)
bins = []
for i in range(buckets + 1):
bins.append(round(min_collisions + i * increment))
out_adr = pd.cut(adr_collisions, bins=bins)
adr_values = out_adr.value_counts(sort=False).iloc[::-1]
out_ucb = pd.cut(ucb_collisions, bins=bins)
ucb_values = out_ucb.value_counts(sort=False).iloc[::-1]
ax.bar(index, adr_values, width=bar_width, color='green', label='ADR')
ax.bar(index + bar_width, ucb_values, width=bar_width, color='blue', label='UCB')
ax.set_xticks(index + bar_width / 2)
ax.set_xticklabels(adr_values.index, rotation=45)
ax.legend()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_color('#DDDDDD')
ax.tick_params(bottom=False, left=False)
ax.set_axisbelow(True)
ax.yaxis.grid(True, color='#EEEEEE')
ax.xaxis.grid(False)
ax.set_xlabel('Number of Collisions', labelpad=15)
ax.set_ylabel('Number of Devices', labelpad=15)
ax.set_title('Collision Rate', pad=15)
fig.tight_layout()
for bar in ax.patches:
bar_value = round(bar.get_height())
text = f'{bar_value:,}'
text_x = bar.get_x() + bar.get_width() / 2
text_y = bar.get_y() + bar_value
bar_color = bar.get_facecolor()
ax.text(text_x, text_y, text, ha='center', va='bottom', color=bar_color, size=10)
fig.savefig(f'adr-ucb-collisions.{output_format}', dpi=300)
# -
print(f'Mean collision number for ADR is {round(statistics.mean(adr_collisions))}')
print(f'Mean collision number for UCB is {round(statistics.mean(ucb_collisions))}')
# ### Ration between active time and total nodes uptime
# +
adr_energy = (adr_end_nodes.active_time / adr_end_nodes.uptime)
adr_active_time = round(statistics.mean(adr_energy) * 100, 2)
ucb_energy = (ucb_end_nodes.active_time / ucb_end_nodes.uptime)
ucb_active_time = round(statistics.mean(ucb_energy) * 100, 2)
print(f'ADR nodes spent {adr_active_time}% of their uptime in active mode')
print(f'UCB nodes spent {ucb_active_time}% of their uptime in active mode')
# -
# ### Packet Delivery Ratio (PDR)
# Evaluation of packet delivery ratio for end nodes.
# Add message count from uplink data and collisions.
# +
adr_data = adr_uplink_messages.node_id.value_counts()
adr_nodes = pd.DataFrame({}, columns = ['dev_id', 'collisions', 'messages'])
collisions = []
messages = []
dev_id = []
for index,value in adr_data.items():
dev_id.append(index)
collision_count = adr_end_nodes.loc[adr_end_nodes.dev_id == index].collisions.values[0]
collisions.append(collision_count)
messages.append(value + collision_count)
adr_nodes['dev_id'] = dev_id
adr_nodes['collisions'] = collisions
adr_nodes['messages'] = messages
# Make the same for another algorithm
ucb_data = ucb_uplink_messages.node_id.value_counts()
ucb_nodes = pd.DataFrame({}, columns = ['dev_id', 'collisions', 'messages'])
collisions = []
messages = []
dev_id = []
for index,value in ucb_data.items():
dev_id.append(index)
collision_count = ucb_end_nodes.loc[ucb_end_nodes.dev_id == index].collisions.values[0]
collisions.append(collision_count)
messages.append(value + collision_count)
ucb_nodes['dev_id'] = dev_id
ucb_nodes['collisions'] = collisions
ucb_nodes['messages'] = messages
# +
adr_nodes['pdr'] = round((1 - (adr_nodes.collisions / adr_nodes.messages))*100, 2)
adr_mean_pdr = round(statistics.mean(adr_nodes.pdr), 2)
ucb_nodes['pdr'] = round((1 - (ucb_nodes.collisions / ucb_nodes.messages))*100, 2)
ucb_mean_pdr = round(statistics.mean(ucb_nodes.pdr), 2)
print(f'Mean value of PDR for ADR is {adr_mean_pdr}%')
print(f'Mean value of PDR for UCB is {ucb_mean_pdr}%')
# +
adr_max_pdr = max(adr_nodes.pdr)
adr_min_pdr = min(adr_nodes.pdr)
ucb_max_pdr = max(ucb_nodes.pdr)
ucb_min_pdr = min(ucb_nodes.pdr)
max_pdr = max(adr_max_pdr, ucb_max_pdr)
min_pdr = min(adr_min_pdr, ucb_max_pdr)
range_pdr = max_pdr - min_pdr
buckets = 8
increment = math.ceil(range_pdr / buckets)
print(f'Max PDR for ADR: {adr_max_pdr}%')
print(f'Min PDR for ADR: {adr_min_pdr}%')
print(f'Max PDR for UCB: {ucb_max_pdr}%')
print(f'Min PDR for UCB: {ucb_min_pdr}%')
# +
fig, ax = plt.subplots(figsize=(10, 4))
bins = []
bar_width = 0.4
index = np.arange(buckets)
for i in range(buckets + 1):
bins.append(round(min_pdr + i * increment))
out_adr = pd.cut(adr_nodes.pdr, bins=bins)
adr_values = out_adr.value_counts(sort=False).iloc[::-1]
out_ucb = pd.cut(ucb_nodes.pdr, bins=bins)
ucb_values = out_ucb.value_counts(sort=False).iloc[::-1]
ax.bar(index, adr_values, width=bar_width, color='green', label='ADR')
ax.bar(index + bar_width, ucb_values, width=bar_width, color='blue', label='UCB')
ax.set_xticks(index + bar_width / 2)
ax.set_xticklabels(adr_values.index, rotation=45)
ax.legend()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_color('#DDDDDD')
ax.tick_params(bottom=False, left=False)
ax.set_axisbelow(True)
ax.yaxis.grid(True, color='#EEEEEE')
ax.xaxis.grid(False)
ax.set_xlabel('Packet Delivery Ratio [%]', labelpad=15)
ax.set_ylabel('Number of Devices', labelpad=15)
ax.set_title('Comparison of PDR', pad=15)
fig.tight_layout()
for bar in ax.patches:
bar_value = round(bar.get_height())
text = f'{bar_value:,}'
text_x = bar.get_x() + bar.get_width() / 2
text_y = bar.get_y() + bar_value
bar_color = bar.get_facecolor()
ax.text(text_x, text_y, text, ha='center', va='bottom', color=bar_color, size=10)
fig.savefig(f'adr-ucb-pdr.{output_format}', dpi=300)
# -
# ## Path of Each End Node
# Data about position are encoded as base64.
# Decode base64, extract position and save the results to original data frame.
# +
# Extracting X and Y coordinates from payload
adr_app_data = adr_uplink_messages.app_data.apply(base64.b64decode)
adr_app_data = adr_app_data.astype(str)
adr_app_data = adr_app_data.str.split(',')
df = pd.DataFrame({}, columns = ['node_id', 'x', 'y'])
x = []
y = []
for row in adr_app_data:
x.append(round(float(row[1].split('\'')[0]), 2) / 1000)
y.append(round(float(row[0].split('\'')[1]), 2) / 1000)
adr_uplink_messages['x'] = x
adr_uplink_messages['y'] = y
# Same for the second algorithm
ucb_app_data = ucb_uplink_messages.app_data.apply(base64.b64decode)
ucb_app_data = ucb_app_data.astype(str)
ucb_app_data = ucb_app_data.str.split(',')
df = pd.DataFrame({}, columns = ['node_id', 'x', 'y'])
x = []
y = []
for row in ucb_app_data:
x.append(round(float(row[1].split('\'')[0]), 2) / 1000)
y.append(round(float(row[0].split('\'')[1]), 2) / 1000)
ucb_uplink_messages['x'] = x
ucb_uplink_messages['y'] = y
# -
# Now, we draw a path for each end node based on the received coordinates.
# +
adr_unique_ens = len(adr_uplink_messages.node_id.unique())
ucb_unique_ens = len(ucb_uplink_messages.node_id.unique())
adr_cmap = mpl.cm.summer
ucb_cmap = mpl.cm.get_cmap('PuBu')
xlim = 10
ylim = 10
fig, axis = plt.subplots(nrows=1, ncols=2, figsize=(10,5))
for i in range(0, adr_unique_ens):
adr_data = adr_uplink_messages[adr_uplink_messages.node_id == adr_uplink_messages.node_id[i]]
axis[0].plot(adr_data.x, adr_data.y, color=adr_cmap(i / unique_ens))
for i in range(0, ucb_unique_ens):
ucb_data = ucb_uplink_messages[ucb_uplink_messages.node_id == ucb_uplink_messages.node_id[i]]
axis[1].plot(ucb_data.x, ucb_data.y, color=ucb_cmap(i / unique_ens))
# Add Access Point
axis[0].plot(xlim / 2, ylim / 2, '+', mew=10, ms=2, color='black')
axis[1].plot(xlim / 2, ylim / 2, '+', mew=10, ms=2, color='black')
# ax.plot(xlim / 2 + 5, ylim / 2 - 5, 'X', mew=10, ms=2, color='black')
for i in range(2):
axis[i].set_xlim([0,xlim])
axis[i].set_ylim([0,ylim])
axis[i].spines['top'].set_visible(False)
axis[i].spines['right'].set_color('#dddddd')
axis[i].spines['left'].set_visible(False)
axis[i].spines['bottom'].set_color('#dddddd')
axis[i].tick_params(bottom=False, left=False)
axis[i].set_axisbelow(True)
axis[i].yaxis.grid(True, color='#eeeeee')
axis[i].xaxis.grid(True, color='#eeeeee')
axis[i].set_xlabel('X [km]', labelpad=15)
axis[i].set_ylabel('Y [km]', labelpad=15)
axis[0].set_title('Paths of ADR Nodes', pad=15)
axis[1].set_title('Paths of UCB Nodes', pad=15)
fig.tight_layout()
fig.savefig(f'adr-ucb-paths.{output_format}', dpi=300)
# -
# The End. //////////\\\\'
| comparison/.ipynb_checkpoints/lora-ucb-vs-ts-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# - nb044ベース(cv:0.9385, sub:0.941)
# - lgbm_clf(deep)
# - batch7処理はしない
# - n_fold=6
# # Const
# +
NB = '065'
isSmallSet = False
if isSmallSet:
LENGTH = 7000
else:
LENGTH = 500_000
MOD_BATCH7 = False
PATH_TRAIN = './../data/input/train_clean.csv'
PATH_TEST = './../data/input/test_clean.csv'
PATH_SMPLE_SUB = './../data/input/sample_submission.csv'
DIR_OUTPUT = './../data/output/'
DIR_OUTPUT_IGNORE = './../data/output_ignore/'
cp = ['#f8b195', '#f67280', '#c06c84', '#6c5b7b', '#355c7d']
sr = 10*10**3 # 10 kHz
# -
# # Import everything I need :)
import warnings
warnings.filterwarnings('ignore')
import time
import gc
import itertools
import multiprocessing
import numpy as np
from scipy import signal
# from pykalman import KalmanFilter
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from fastprogress import progress_bar
from lightgbm import LGBMRegressor, LGBMClassifier
from sklearn.model_selection import KFold, train_test_split, StratifiedKFold
from sklearn.metrics import f1_score, mean_absolute_error, confusion_matrix
from sklearn.ensemble import RandomForestRegressor, RandomForestClassifier
from sklearn.tree import DecisionTreeRegressor
from sklearn.pipeline import make_pipeline
from sklearn.base import BaseEstimator, TransformerMixin
# from sklearn.svm import SVR
from sklearn.linear_model import Lasso
# from dtreeviz.trees import dtreeviz
# # My function
# +
def f1_macro(true, pred):
return f1_score(true, pred, average='macro')
def get_df_batch(df, batch):
idxs = df['batch'] == batch
assert any(idxs), 'そのようなbatchはありません'
return df[idxs]
def add_category(train, test):
train["category"] = 0
test["category"] = 0
# train segments with more then 9 open channels classes
train.loc[2_000_000:2_500_000-1, 'category'] = 1
train.loc[4_500_000:5_000_000-1, 'category'] = 1
# test segments with more then 9 open channels classes (potentially)
test.loc[500_000:600_000-1, "category"] = 1
test.loc[700_000:800_000-1, "category"] = 1
return train, test
def get_signal_mv_mean(df, n=3001):
signal_mv = np.zeros(len(df))
for bt in df['batch'].unique():
idxs = df['batch'] == bt
_signal_mv = df['signal'][idxs].rolling(n, center=True).mean().interpolate('spline', order=5, limit_direction='both').values
signal_mv[idxs] = _signal_mv
return signal_mv
def get_signal_mv_std(df, n=3001):
signal_mv = np.zeros(len(df))
for bt in df['batch'].unique():
idxs = df['batch'] == bt
_signal_mv = df['signal'][idxs].rolling(n, center=True).std().interpolate('spline', order=5, limit_direction='both').values
signal_mv[idxs] = _signal_mv
return signal_mv
def get_signal_mv_min(df, n=3001):
signal_mv = np.zeros(len(df))
for bt in df['batch'].unique():
idxs = df['batch'] == bt
_signal_mv = df['signal'][idxs].rolling(n, center=True).min().interpolate('spline', order=5, limit_direction='both').values
signal_mv[idxs] = _signal_mv
return signal_mv
def get_signal_mv_max(df, n=3001):
signal_mv = np.zeros(len(df))
for bt in df['batch'].unique():
idxs = df['batch'] == bt
_signal_mv = df['signal'][idxs].rolling(n, center=True).max().interpolate('spline', order=5, limit_direction='both').values
signal_mv[idxs] = _signal_mv
return signal_mv
def calc_shifted(s, add_minus=False, fill_value=None, periods=range(1, 4)):
s = pd.DataFrame(s)
_periods = periods
add_minus = True
periods = np.asarray(_periods, dtype=np.int32)
if add_minus:
periods = np.append(periods, -periods)
for p in progress_bar(periods):
s[f"signal_shifted_{p}"] = s['signal'].shift(
periods=p, fill_value=fill_value
)
cols = [col for col in s.columns if 'shifted' in col]
return s[cols]
def group_feat_train(_train):
train = _train.copy()
# group init
train['group'] = int(0)
# group 1
idxs = (train['batch'] == 3) | (train['batch'] == 7)
train['group'][idxs] = int(1)
# group 2
idxs = (train['batch'] == 5) | (train['batch'] == 8)
train['group'][idxs] = int(2)
# group 3
idxs = (train['batch'] == 2) | (train['batch'] == 6)
train['group'][idxs] = int(3)
# group 4
idxs = (train['batch'] == 4) | (train['batch'] == 9)
train['group'][idxs] = int(4)
return train[['group']]
def group_feat_test(_test):
test = _test.copy()
# group init
test['group'] = int(0)
x_idx = np.arange(len(test))
# group 1
idxs = (100000<=x_idx) & (x_idx<200000)
test['group'][idxs] = int(1)
idxs = (900000<=x_idx) & (x_idx<=1000000)
test['group'][idxs] = int(1)
# group 2
idxs = (200000<=x_idx) & (x_idx<300000)
test['group'][idxs] = int(2)
idxs = (600000<=x_idx) & (x_idx<700000)
test['group'][idxs] = int(2)
# group 3
idxs = (400000<=x_idx) & (x_idx<500000)
test['group'][idxs] = int(3)
# group 4
idxs = (500000<=x_idx) & (x_idx<600000)
test['group'][idxs] = int(4)
idxs = (700000<=x_idx) & (x_idx<800000)
test['group'][idxs] = int(4)
return test[['group']]
class permutation_importance():
def __init__(self, model, metric):
self.is_computed = False
self.n_feat = 0
self.base_score = 0
self.model = model
self.metric = metric
self.df_result = []
def compute(self, X_valid, y_valid):
self.n_feat = len(X_valid.columns)
if self.metric == 'auc':
y_valid_score = self.model.predict_proba(X_valid)[:, 1]
fpr, tpr, thresholds = roc_curve(y_valid, y_valid_score)
self.base_score = auc(fpr, tpr)
else:
pred = np.round(self.model.predict(X_valid)).astype('int8')
self.base_score = self.metric(y_valid, pred)
self.df_result = pd.DataFrame({'feat': X_valid.columns,
'score': np.zeros(self.n_feat),
'score_diff': np.zeros(self.n_feat)})
# predict
for i, col in enumerate(X_valid.columns):
df_perm = X_valid.copy()
np.random.seed(1)
df_perm[col] = np.random.permutation(df_perm[col])
y_valid_pred = self.model.predict(df_perm)
if self.metric == 'auc':
y_valid_score = self.model.predict_proba(df_perm)[:, 1]
fpr, tpr, thresholds = roc_curve(y_valid, y_valid_score)
score = auc(fpr, tpr)
else:
score = self.metric(y_valid, np.round(y_valid_pred).astype('int8'))
self.df_result['score'][self.df_result['feat']==col] = score
self.df_result['score_diff'][self.df_result['feat']==col] = self.base_score - score
self.is_computed = True
def get_negative_feature(self):
assert self.is_computed!=False, 'compute メソッドが実行されていません'
idx = self.df_result['score_diff'] < 0
return self.df_result.loc[idx, 'feat'].values.tolist()
def get_positive_feature(self):
assert self.is_computed!=False, 'compute メソッドが実行されていません'
idx = self.df_result['score_diff'] > 0
return self.df_result.loc[idx, 'feat'].values.tolist()
def show_permutation_importance(self, score_type='loss'):
'''score_type = 'loss' or 'accuracy' '''
assert self.is_computed!=False, 'compute メソッドが実行されていません'
if score_type=='loss':
ascending = True
elif score_type=='accuracy':
ascending = False
else:
ascending = ''
plt.figure(figsize=(15, int(0.25*self.n_feat)))
sns.barplot(x="score_diff", y="feat", data=self.df_result.sort_values(by="score_diff", ascending=ascending))
plt.title('base_score - permutation_score')
def plot_corr(df, abs_=False, threshold=0.95):
if abs_==True:
corr = df.corr().abs()>threshold
vmin = 0
else:
corr = df.corr()
vmin = -1
# Plot
fig, ax = plt.subplots(figsize=(12, 10), dpi=100)
fig.patch.set_facecolor('white')
sns.heatmap(corr,
xticklabels=df.corr().columns,
yticklabels=df.corr().columns,
vmin=vmin,
vmax=1,
center=0,
annot=False)
# Decorations
ax.set_title('Correlation', fontsize=22)
def get_low_corr_column(df, threshold):
df_corr = df.corr()
df_corr = abs(df_corr)
columns = df_corr.columns
# 対角線の値を0にする
for i in range(0, len(columns)):
df_corr.iloc[i, i] = 0
while True:
columns = df_corr.columns
max_corr = 0.0
query_column = None
target_column = None
df_max_column_value = df_corr.max()
max_corr = df_max_column_value.max()
query_column = df_max_column_value.idxmax()
target_column = df_corr[query_column].idxmax()
if max_corr < threshold:
# しきい値を超えるものがなかったため終了
break
else:
# しきい値を超えるものがあった場合
delete_column = None
saved_column = None
# その他との相関の絶対値が大きい方を除去
if sum(df_corr[query_column]) <= sum(df_corr[target_column]):
delete_column = target_column
saved_column = query_column
else:
delete_column = query_column
saved_column = target_column
# 除去すべき特徴を相関行列から消す(行、列)
df_corr.drop([delete_column], axis=0, inplace=True)
df_corr.drop([delete_column], axis=1, inplace=True)
return df_corr.columns # 相関が高い特徴量を除いた名前リスト
def reduce_mem_usage(df, verbose=True):
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage().sum() / 1024**2
for col in df.columns:
if col!='open_channels':
col_type = df[col].dtypes
if col_type in numerics:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024**2
if verbose: print('Mem. usage decreased to {:5.2f} Mb ({:.1f}% reduction)'.format(end_mem, 100 * (start_mem - end_mem) / start_mem))
return df
def plot_confusion_matrix(truth, pred, classes, normalize=False, title=''):
cm = confusion_matrix(truth, pred)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.figure(figsize=(10, 10))
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
plt.title('Confusion matrix', size=15)
plt.colorbar(fraction=0.046, pad=0.04)
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.grid(False)
plt.tight_layout()
def create_signal_mod(train):
left = 3641000
right = 3829000
thresh_dict = {
3: [0.1, 2.0],
2: [-1.1, 0.7],
1: [-2.3, -0.6],
0: [-3.8, -2],
}
train['signal'] = train['signal'].values
for ch in train[train['batch']==7]['open_channels'].unique():
idxs_noisy = (train['open_channels']==ch) & (left<train.index) & (train.index<right)
idxs_not_noisy = (train['open_channels']==ch) & ~idxs_noisy
mean = train[idxs_not_noisy]['signal'].mean()
idxs_outlier = idxs_noisy & (thresh_dict[ch][1]<train['signal'].values)
train['signal'][idxs_outlier] = mean
idxs_outlier = idxs_noisy & (train['signal'].values<thresh_dict[ch][0])
train['signal'][idxs_outlier] = mean
return train
def create_signal_mod2(train):
left = 3641000
right = 3829000
thresh_dict = {
3: [0.1, 2.0],
2: [-1.1, 0.7],
1: [-2.3, -0.6],
0: [-3.8, -2],
}
train['signal'] = train['signal'].values
for ch in train[train['batch']==7]['open_channels'].unique():
idxs_noisy = (train['open_channels']==ch) & (left<train.index) & (train.index<right)
idxs_not_noisy = (train['open_channels']==ch) & ~idxs_noisy
mean = train[idxs_not_noisy]['signal'].mean()
std = train[idxs_not_noisy]['signal'].std()
idxs_outlier = idxs_noisy & (thresh_dict[ch][1]<train['signal'].values)
noise = np.random.normal(loc=0, scale=std, size=len(train['signal'].values[idxs_outlier]))
train['signal'][idxs_outlier] = mean + noise
idxs_outlier = idxs_noisy & (train['signal'].values<thresh_dict[ch][0])
noise = np.random.normal(loc=0, scale=std, size=len(train['signal'].values[idxs_outlier]))
train['signal'][idxs_outlier] = mean + noise
return train
# +
def train_lgbm(X, y, X_te, lgbm_params, random_state=5, n_fold=5, verbose=50, early_stopping_rounds=100, show_fig=True):
# using features
print(f'features({len(X.columns)}): \n{X.columns}') if not verbose==0 else None
# folds = KFold(n_splits=n_fold, shuffle=True, random_state=random_state)
folds = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=random_state)
scores = []
oof = np.zeros(len(X))
oof_round = np.zeros(len(X))
test_pred = np.zeros(len(X_te))
df_pi = pd.DataFrame(columns=['feat', 'score_diff'])
for fold_n, (train_idx, valid_idx) in enumerate(folds.split(X, y=y)):
if verbose==0:
pass
else:
print('\n------------------')
print(f'- Fold {fold_n + 1}/{N_FOLD} started at {time.ctime()}')
# prepare dataset
X_train, X_valid = X.iloc[train_idx], X.iloc[valid_idx]
y_train, y_valid = y[train_idx], y[valid_idx]
# train
model = LGBMRegressor(**lgbm_params)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_valid, y_valid)],
verbose=verbose,
early_stopping_rounds=early_stopping_rounds)
# pred
y_valid_pred = model.predict(X_valid, model.best_iteration_)
y_valid_pred_round = np.round(y_valid_pred).astype('int8')
_test_pred = model.predict(X_te, model.best_iteration_)
if show_fig==False:
pass
else:
# permutation importance
pi = permutation_importance(model, f1_macro) # model と metric を渡す
pi.compute(X_valid, y_valid)
pi_result = pi.df_result
df_pi = pd.concat([df_pi, pi_result[['feat', 'score_diff']]])
# result
oof[valid_idx] = y_valid_pred
oof_round[valid_idx] = y_valid_pred_round
score = f1_score(y_valid, y_valid_pred_round, average='macro')
scores.append(score)
test_pred += _test_pred
if verbose==0:
pass
else:
print(f'---> f1-score(macro) valid: {f1_score(y_valid, y_valid_pred_round, average="macro"):.4f}')
print('')
print('====== finish ======')
print('score list:', scores)
print('CV mean score(f1_macro): {0:.4f}, std: {1:.4f}'.format(np.mean(scores), np.std(scores)))
print(f'oof score(f1_macro): {f1_score(y, oof_round, average="macro"):.4f}')
print('')
if show_fig==False:
pass
else:
# visualization
plt.figure(figsize=(5, 5))
plt.plot([0, 10], [0, 10], color='gray')
plt.scatter(y, oof, alpha=0.05, color=cp[1])
plt.xlabel('true')
plt.ylabel('pred')
plt.show()
# confusion_matrix
plot_confusion_matrix(y, oof_round, classes=np.arange(11))
# permutation importance
plt.figure(figsize=(15, int(0.25*len(X.columns))))
order = df_pi.groupby(["feat"]).mean()['score_diff'].reset_index().sort_values('score_diff', ascending=False)
sns.barplot(x="score_diff", y="feat", data=df_pi, order=order['feat'])
plt.title('base_score - permutation_score')
plt.show()
# submission
test_pred = test_pred/N_FOLD
test_pred_round = np.round(test_pred).astype('int8')
return test_pred_round, test_pred, oof_round, oof, type(model).__name__
# +
def train_test_split_lgbm(X, y, X_te, lgbm_params, random_state=5, test_size=0.3, verbose=50, early_stopping_rounds=100, show_fig=True):
# using features
print(f'features({len(X.columns)}): \n{X.columns}') if not verbose==0 else None
# folds = KFold(n_splits=n_fold, shuffle=True, random_state=random_state)
# folds = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=random_state)
# prepare dataset
X_train, X_valid, y_train, y_valid = train_test_split(X, y, test_size=test_size, random_state=random_state)
# train
model = LGBMRegressor(**lgbm_params, n_estimators=N_ESTIMATORS)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_valid, y_valid)],
verbose=verbose,
early_stopping_rounds=early_stopping_rounds)
# pred
oof = model.predict(X_valid, model.best_iteration_)
oof_round = np.round(oof).astype('int8')
test_pred = model.predict(X_te, model.best_iteration_)
test_pred_round = np.round(test_pred).astype('int8')
print('====== finish ======')
print(f'oof score(f1_macro): {f1_score(y_valid, oof_round, average="macro"):.4f}')
print('')
if show_fig==False:
pass
else:
# visualization
plt.figure(figsize=(5, 5))
plt.plot([0, 10], [0, 10], color='gray')
plt.scatter(y_valid, oof, alpha=0.05, color=cp[1])
plt.xlabel('true')
plt.ylabel('pred')
plt.show()
# confusion_matrix
plot_confusion_matrix(y_valid, oof_round, classes=np.arange(11))
# permutation importance
pi = permutation_importance(model, f1_macro) # model と metric を渡す
pi.compute(X_valid, y_valid)
pi.show_permutation_importance(score_type='accuracy') # loss or accuracy
plt.show()
return test_pred_round, test_pred, oof_round, oof, type(model).__name__
# -
def train_rfc(X, y, X_te, rfc_params, random_state=5, n_fold=5, verbose=2, show_fig=True):
# using features
print(f'features({len(X.columns)}): \n{X.columns}') if not verbose==0 else None
folds = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=random_state)
scores = []
oof_proba = np.zeros([len(X), len(np.unique(y))])
test_proba = np.zeros([len(X_te), len(np.unique(y))])
df_pi = pd.DataFrame(columns=['feat', 'score_diff'])
for fold_n, (train_idx, valid_idx) in enumerate(folds.split(X, y=y)):
if verbose==0:
pass
else:
print('\n------------------')
print(f'- Fold {fold_n + 1}/{N_FOLD} started at {time.ctime()}')
# prepare dataset
X_train, X_valid = X.iloc[train_idx], X.iloc[valid_idx]
y_train, y_valid = y[train_idx], y[valid_idx]
# train
model = RandomForestClassifier(**rfc_params, verbose=verbose)
model.fit(X_train, y_train)
# pred
y_valid_pred = model.predict(X_valid)
y_valid_proba = model.predict_proba(X_valid)
# y_valid_pred_round = np.round(y_valid_pred).astype('int8')
_test_pred = model.predict(X_te)
_test_proba = model.predict_proba(X_te)
if show_fig==False:
pass
else:
# permutation importance
pi = permutation_importance(model, f1_macro) # model と metric を渡す
pi.compute(X_valid, y_valid)
pi_result = pi.df_result
df_pi = pd.concat([df_pi, pi_result[['feat', 'score_diff']]])
# result
oof_proba[valid_idx] = y_valid_proba
score = f1_score(y_valid, y_valid_pred, average='macro')
scores.append(score)
test_proba += _test_proba
if verbose==0:
pass
else:
print(f'---> f1-score(macro) valid: {f1_score(y_valid, y_valid_pred, average="macro"):.4f}')
print('')
print('====== finish ======')
oof = np.argmax(oof_proba, axis=1)
print('score list:', scores)
print('CV mean score(f1_macro): {0:.4f}, std: {1:.4f}'.format(np.mean(scores), np.std(scores)))
print(f'oof score(f1_macro): {f1_score(y, oof, average="macro"):.4f}')
print('')
if show_fig==False:
pass
else:
# visualization
plt.figure(figsize=(5, 5))
plt.plot([0, 10], [0, 10], color='gray')
plt.scatter(y, oof, alpha=0.05, color=cp[1])
plt.xlabel('true')
plt.ylabel('pred')
plt.show()
# confusion_matrix
plot_confusion_matrix(y, oof, classes=np.arange(11))
# permutation importance
plt.figure(figsize=(15, int(0.25*len(X.columns))))
order = df_pi.groupby(["feat"]).mean()['score_diff'].reset_index().sort_values('score_diff', ascending=False)
sns.barplot(x="score_diff", y="feat", data=df_pi, order=order['feat'])
plt.title('base_score - permutation_score')
plt.show()
# submission
test_proba = test_proba/N_FOLD
test_pred = np.argmax(test_proba, axis=1)
# oof_pred = np.argmax(oof_proba, axis=1)
return test_pred, test_proba, oof, oof_proba, type(model).__name__
def train_lgbm_clf(X, y, X_te, lgbm_params, random_state=5, n_fold=5, verbose=50, early_stopping_rounds=100, show_fig=True):
# using features
print(f'features({len(X.columns)}): \n{X.columns}') if not verbose==0 else None
folds = StratifiedKFold(n_splits=n_fold, shuffle=True, random_state=random_state)
scores = []
oof_proba = np.zeros([len(X), len(np.unique(y))])
test_proba = np.zeros([len(X_te), len(np.unique(y))])
df_pi = pd.DataFrame(columns=['feat', 'score_diff'])
for fold_n, (train_idx, valid_idx) in enumerate(folds.split(X, y=y)):
if verbose==0:
pass
else:
print('\n------------------')
print(f'- Fold {fold_n + 1}/{N_FOLD} started at {time.ctime()}')
# prepare dataset
X_train, X_valid = X.iloc[train_idx], X.iloc[valid_idx]
y_train, y_valid = y[train_idx], y[valid_idx]
# train
model = LGBMClassifier(**lgbm_params)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_valid, y_valid)],
verbose=verbose,
early_stopping_rounds=early_stopping_rounds)
# pred
# y_valid_pred = model.predict(X_valid, model.best_iteration_)
y_valid_proba = model.predict_proba(X_valid, num_iteration=model.best_iteration_)
y_valid_pred = np.argmax(y_valid_proba, axis=1)
# _test_pred = model.predict(X_te, model.best_iteration_)
_test_proba = model.predict_proba(X_te, num_iteration=model.best_iteration_)
_test_pred = np.argmax(_test_proba, axis=1)
if show_fig==False:
pass
else:
# permutation importance
pi = permutation_importance(model, f1_macro) # model と metric を渡す
pi.compute(X_valid, y_valid)
pi_result = pi.df_result
df_pi = pd.concat([df_pi, pi_result[['feat', 'score_diff']]])
# result
oof_proba[valid_idx] = y_valid_proba
score = f1_score(y_valid, y_valid_pred, average='macro')
scores.append(score)
test_proba += _test_proba
if verbose==0:
pass
else:
print(f'---> f1-score(macro) valid: {f1_score(y_valid, y_valid_pred, average="macro"):.4f}')
print('')
print('====== finish ======')
oof = np.argmax(oof_proba, axis=1)
print('score list:', scores)
print('CV mean score(f1_macro): {0:.4f}, std: {1:.4f}'.format(np.mean(scores), np.std(scores)))
print(f'oof score(f1_macro): {f1_score(y, oof, average="macro"):.4f}')
print('')
if show_fig==False:
pass
else:
# visualization
plt.figure(figsize=(5, 5))
plt.plot([0, 10], [0, 10], color='gray')
plt.scatter(y, oof, alpha=0.05, color=cp[1])
plt.xlabel('true')
plt.ylabel('pred')
plt.show()
# confusion_matrix
plot_confusion_matrix(y, oof, classes=np.arange(11))
# permutation importance
plt.figure(figsize=(15, int(0.25*len(X.columns))))
order = df_pi.groupby(["feat"]).mean()['score_diff'].reset_index().sort_values('score_diff', ascending=False)
sns.barplot(x="score_diff", y="feat", data=df_pi, order=order['feat'])
plt.title('base_score - permutation_score')
plt.show()
# submission
test_proba = test_proba/N_FOLD
test_pred = np.argmax(test_proba, axis=1)
# oof_pred = np.argmax(oof_proba, axis=1)
return test_pred, test_proba, oof, oof_proba, type(model).__name__
# <br>
#
# ref: https://www.kaggle.com/martxelo/fe-and-ensemble-mlp-and-lgbm
# +
def calc_gradients(s, n_grads=4):
'''
Calculate gradients for a pandas series. Returns the same number of samples
'''
grads = pd.DataFrame()
g = s.values
for i in range(n_grads):
g = np.gradient(g)
grads['grad_' + str(i+1)] = g
return grads
def calc_low_pass(s, n_filts=10):
'''
Applies low pass filters to the signal. Left delayed and no delayed
'''
wns = np.logspace(-2, -0.3, n_filts)
# wns = [0.3244]
low_pass = pd.DataFrame()
x = s.values
for wn in wns:
b, a = signal.butter(1, Wn=wn, btype='low')
zi = signal.lfilter_zi(b, a)
low_pass['lowpass_lf_' + str('%.4f' %wn)] = signal.lfilter(b, a, x, zi=zi*x[0])[0]
low_pass['lowpass_ff_' + str('%.4f' %wn)] = signal.filtfilt(b, a, x)
return low_pass
def calc_high_pass(s, n_filts=10):
'''
Applies high pass filters to the signal. Left delayed and no delayed
'''
wns = np.logspace(-2, -0.1, n_filts)
# wns = [0.0100, 0.0264, 0.0699, 0.3005, 0.4885, 0.7943]
high_pass = pd.DataFrame()
x = s.values
for wn in wns:
b, a = signal.butter(1, Wn=wn, btype='high')
zi = signal.lfilter_zi(b, a)
high_pass['highpass_lf_' + str('%.4f' %wn)] = signal.lfilter(b, a, x, zi=zi*x[0])[0]
high_pass['highpass_ff_' + str('%.4f' %wn)] = signal.filtfilt(b, a, x)
return high_pass
def calc_roll_stats(s, windows=[10, 50, 100, 500, 1000, 3000]):
'''
Calculates rolling stats like mean, std, min, max...
'''
roll_stats = pd.DataFrame()
for w in windows:
roll_stats['roll_mean_' + str(w)] = s.rolling(window=w, min_periods=1).mean().interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_std_' + str(w)] = s.rolling(window=w, min_periods=1).std().interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_min_' + str(w)] = s.rolling(window=w, min_periods=1).min().interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_max_' + str(w)] = s.rolling(window=w, min_periods=1).max().interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_range_' + str(w)] = roll_stats['roll_max_' + str(w)] - roll_stats['roll_min_' + str(w)]
roll_stats['roll_q10_' + str(w)] = s.rolling(window=w, min_periods=1).quantile(0.10).interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_q25_' + str(w)] = s.rolling(window=w, min_periods=1).quantile(0.25).interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_q50_' + str(w)] = s.rolling(window=w, min_periods=1).quantile(0.50).interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_q75_' + str(w)] = s.rolling(window=w, min_periods=1).quantile(0.75).interpolate('spline', order=5, limit_direction='both')
roll_stats['roll_q90_' + str(w)] = s.rolling(window=w, min_periods=1).quantile(0.90).interpolate('spline', order=5, limit_direction='both')
# add zeros when na values (std)
# roll_stats = roll_stats.fillna(value=0)
return roll_stats
def calc_ewm(s, windows=[10, 50, 100, 500, 1000, 3000]):
'''
Calculates exponential weighted functions
'''
ewm = pd.DataFrame()
for w in windows:
ewm['ewm_mean_' + str(w)] = s.ewm(span=w, min_periods=1).mean()
ewm['ewm_std_' + str(w)] = s.ewm(span=w, min_periods=1).std()
# add zeros when na values (std)
ewm = ewm.fillna(value=0)
return ewm
def divide_and_add_features(s, signal_size=500000):
'''
Divide the signal in bags of "signal_size".
Normalize the data dividing it by 15.0
'''
# normalize
s = s/15.0
ls = []
for i in progress_bar(range(int(s.shape[0]/signal_size))):
sig = s[i*signal_size:(i+1)*signal_size].copy().reset_index(drop=True)
sig_featured = add_features(sig)
ls.append(sig_featured)
return pd.concat(ls, axis=0)
# -
# <br>
#
# ref: https://www.kaggle.com/nxrprime/single-model-lgbm-kalman-filter-ii
def Kalman1D(observations,damping=1):
# To return the smoothed time series data
observation_covariance = damping
initial_value_guess = observations[0]
transition_matrix = 1
transition_covariance = 0.1
initial_value_guess
kf = KalmanFilter(
initial_state_mean=initial_value_guess,
initial_state_covariance=observation_covariance,
observation_covariance=observation_covariance,
transition_covariance=transition_covariance,
transition_matrices=transition_matrix
)
pred_state, state_cov = kf.smooth(observations)
return pred_state
# # Preparation
# setting
sns.set()
# <br>
#
# load dataset
df_tr = pd.read_csv(PATH_TRAIN)
df_te = pd.read_csv(PATH_TEST)
# <br>
#
# 処理のしやすさのために、バッチ番号を振る
# +
batch_list = []
for n in range(10):
batchs = np.ones(500000)*n
batch_list.append(batchs.astype(int))
batch_list = np.hstack(batch_list)
df_tr['batch'] = batch_list
batch_list = []
for n in range(4):
batchs = np.ones(500000)*n
batch_list.append(batchs.astype(int))
batch_list = np.hstack(batch_list)
df_te['batch'] = batch_list
# -
# <br>
#
# group 特徴量
# +
# group 特徴量を作成
group = group_feat_train(df_tr)
df_tr = pd.concat([df_tr, group], axis=1)
group = group_feat_test(df_te)
df_te = pd.concat([df_te, group], axis=1)
if isSmallSet:
df_te['group'][1000:2000] = 1
df_te['group'][2000:3000] = 2
df_te['group'][3000:4000] = 3
df_te['group'][4000:5000] = 4
# -
# <br>
#
# group4にオフセットをかける
# +
# --- train ---
off_set_4 = 0.952472 - (-1.766044)
off_set_9 = 0.952472 - (-1.770441)
# batch4
idxs = df_tr['batch'] == 4
df_tr['signal'][idxs] = df_tr['signal'].values + off_set_4
# batch9
idxs = df_tr['batch'] == 9
df_tr['signal'][idxs] = df_tr['signal'].values + off_set_9
# --- test ---
off_set_test = 2.750
df_te['signal'] = df_te['signal'].values
idxs = df_te['group'] == 4
df_te['signal'][idxs] = df_te['signal'][idxs].values + off_set_test
# -
# <br>
#
# batch7のスパイク処理
if MOD_BATCH7:
df_tr = create_signal_mod2(df_tr)
# <br>
#
# smallset?
if isSmallSet:
print('small set mode')
# train
batchs = df_tr['batch'].values
dfs = []
for i_bt, bt in enumerate(df_tr['batch'].unique()):
idxs = batchs == bt
_df = df_tr[idxs][:LENGTH].copy()
dfs.append(_df)
df_tr = pd.concat(dfs).reset_index(drop=True)
# test
batchs = df_te['batch'].values
dfs = []
for i_bt, bt in enumerate(df_te['batch'].unique()):
idxs = batchs == bt
_df = df_te[idxs][:LENGTH].copy()
dfs.append(_df)
df_te = pd.concat(dfs).reset_index(drop=True)
# # Train
def add_features(s):
'''
All calculations together
'''
feat_list = [s]
feat_list.append(calc_gradients(s))
feat_list.append(calc_low_pass(s))
feat_list.append(calc_high_pass(s))
feat_list.append(calc_roll_stats(s))
# feat_list.append(calc_ewm(s))
feat_list.append(calc_shifted(s, fill_value=0, periods=range(1, 4)))
return pd.concat(feat_list, axis=1)
# +
# %%time
print(f'train start {time.ctime()}')
X = divide_and_add_features(df_tr['signal'], signal_size=LENGTH).reset_index(drop=True)
# _feats = get_low_corr_column(X, threshold=0.97).to_list()
# _feats.append('signal')
# X = X[_feats]
# X = reduce_mem_usage(X)
print(f'test start {time.ctime()}')
X_te = divide_and_add_features(df_te['signal'], signal_size=LENGTH).reset_index(drop=True)
# X_te = X_te[_feats]
# X_te = reduce_mem_usage(X_te)
y = df_tr['open_channels'].values
# -
X, X_te = add_category(X, X_te)
# <br>
#
# 学習データセットの作成
# +
# Configuration
N_ESTIMATORS = 2000
# N_ESTIMATORS = 20 # 最大学習回数
VERBOSE = 100 # 300回ごとに評価する
EARLY_STOPPING_ROUNDS = 50 # 200回の学習でよくならなければ、学習をとめる
# N_JOBS = multiprocessing.cpu_count() - 2
# N_JOBS = 6
# N_FOLD = 4
# KFOLD_SEED = 0
N_JOBS = 28
N_FOLD = 6
KFOLD_SEED = 42
# lgbm_params
# lgbm_params = {
# 'objective': 'regression',
# "metric": 'rmse',
# # 'reg_alpha': 0.1,
# # 'reg_lambda': 0.1,
# "boosting_type": "gbdt",
# 'learning_rate': 0.1,
# 'n_jobs': N_JOBS,
# # "subsample_freq": 1,
# # "subsample": 1,
# "bagging_seed": 2,
# # "verbosity": -1,
# 'num_leaves': 51, 'max_depth': 158, 'min_chiled_samples': 15, 'min_chiled_weight': 1, 'learning_rate': 0.07, 'colsample_bytree': 0.8
# }
# nb015
# lgbm_params = {'boosting_type': 'gbdt',
# 'metric': 'rmse',
# 'objective': 'regression',
# 'n_jobs': N_JOBS,
# 'seed': 236,
# 'num_leaves': 280,
# 'learning_rate': 0.026623466966581126,
# 'max_depth': 73,
# 'lambda_l1': 2.959759088169741,
# 'lambda_l2': 1.331172832164913,
# 'bagging_fraction': 0.9655406551472153,
# 'bagging_freq': 9,
# 'colsample_bytree': 0.6867118652742716}
# nb019
# lgbm_params = {
# 'objective': 'regression',
# "metric": 'rmse',
# "boosting_type": "gbdt",
# 'learning_rate': 0.1,
# 'n_jobs': N_JOBS,
# 'max_depth': 85,
# 'min_chiled_samples': 62,
# 'min_chiled_weight': 10,
# 'learning_rate': 0.20158497791184515,
# 'colsample_bytree': 1.0,
# 'lambda_l1': 2.959759088169741,
# 'lambda_l2': 1.331172832164913,
# # 'bagging_fraction': 0.9655406551472153,
# # 'bagging_freq': 9,
# }
# https://www.kaggle.com/nxrprime/single-model-lgbm-kalman-filter-ii
lgbm_params = {'boosting_type': 'gbdt',
'objective': 'multiclass',
# 'metric': 'rmse',
'num_class': 11,
'n_jobs': N_JOBS,
'seed': 236,
'n_estimators': N_ESTIMATORS,
'num_leaves': 280,
# 'learning_rate': 0.026623466966581126,
'learning_rate': 0.03,
'max_depth': 73,
'lambda_l1': 2.959759088169741,
'lambda_l2': 1.331172832164913,
'bagging_fraction': 0.9655406551472153,
'bagging_freq': 9,
'colsample_bytree': 0.6867118652742716}
# -
# %%time
test_pred, test_proba, oof, oof_proba, model_name = train_lgbm_clf(X, y, X_te, lgbm_params,
n_fold=N_FOLD,
verbose=VERBOSE,
random_state=KFOLD_SEED,
early_stopping_rounds=EARLY_STOPPING_ROUNDS,
show_fig=False)
# # save
# submission
save_path = f'{DIR_OUTPUT}submission_nb{NB}_{model_name}_cv_{f1_macro(y, oof):.4f}.csv'
sub = pd.read_csv(PATH_SMPLE_SUB)
# sub['open_channels'] = test_pred
sub['open_channels'] = test_pred.astype(int)
print(f'save path: {save_path}')
sub.to_csv(save_path, index=False, float_format='%.4f')
# <br>
#
# oof proba
save_path = f'{DIR_OUTPUT_IGNORE}probas_nb{NB}_{model_name}_cv_{f1_macro(y, oof):.4f}'
print(f'save path: {save_path}')
np.savez_compressed(save_path, oof_proba, test_proba)
# # analysis
# <br>
#
# 処理のしやすさのために、バッチ番号を振る
batch_list = []
for n in range(10):
batchs = np.ones(500000)*n
batch_list.append(batchs.astype(int))
batch_list = np.hstack(batch_list)
X['batch'] = batch_list
# <br>
#
# group 特徴量
# group 特徴量を作成
group = group_feat_train(X)
X = pd.concat([X, group], axis=1)
for group in sorted(X['group'].unique()):
idxs = X['group'] == group
oof_grp = oof[idxs].astype(int)
y_grp = y[idxs]
print(f'group_score({group}): {f1_score(y_grp, oof_grp, average="micro"):4f}')
# +
x_idx = np.arange(len(X))
idxs = y != oof
failed = np.zeros(len(X))
failed[idxs] = 1
# -
n = 200
b = np.ones(n)/n
failed_move = np.convolve(failed, b, mode='same')
# +
fig, axs = plt.subplots(2, 1, figsize=(20, 6))
axs = axs.ravel()
# fig = plt.figure(figsize=(20, 3))
for i_gr, group in enumerate(sorted(X['group'].unique())):
idxs = X['group'] == group
axs[0].plot(np.arange(len(X))[idxs], X['signal'].values[idxs], color=cp[i_gr], label=f'group={group}')
for x in range(10):
axs[0].axvline(x*500000 + 500000, color='gray')
axs[0].text(x*500000 + 250000, 0.6, x)
axs[0].plot(x_idx, failed_move, '.', color='black', label='failed_mv')
axs[0].set_xlim(0, 5500000)
axs[0].legend()
axs[1].plot(x_idx, y)
axs[1].set_xlim(0, 5500000)
# fig.legend()
# -
# +
| nb/.ipynb_checkpoints/065_submission-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xXKfRxJ0MuE_"
# #Using California Housing Problem
# + id="40mZnSCdMeAa" executionInfo={"status": "ok", "timestamp": 1602714110615, "user_tz": -660, "elapsed": 5284, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgxpvArds7xLNZB29BI3aCcSZkWbMPAZWZLOm3o=s64", "userId": "03940255243131127536"}} outputId="41115491-4ef5-4b86-9d6c-cdd6f93f22ee" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.datasets import fetch_california_housing
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import tensorflow as tf
from tensorflow import keras
import numpy as np
housing = fetch_california_housing()
X_train_full, X_test, y_train_full, y_test = train_test_split(housing.data, housing.target, random_state=42)
X_train, X_valid, y_train, y_valid = train_test_split(X_train_full, y_train_full, random_state=42)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_valid = scaler.transform(X_valid)
X_test = scaler.transform(X_test)
np.random.seed(42)
tf.random.set_seed(42)
# + [markdown] id="8_fD_dhtMr8y"
# Initizlising the Code with sklearn's california housing problem
# + id="vnktZaPwM89B"
input_ = keras.layers.Input(shape=X_train[:1])
hidden1 = kears.layers.Dense(30, activation='relu')(input_)
hidden2 = keras.layers.Dense(30, activation='relu')(hidden1)
concat = keras.layers.Concatenate()([input_, hidden2])
output = keras.layers.Dense(1)(concat)
model = keras.Model(inputs=[input_], outputs=[output])
# + [markdown] id="rR9wtgqoM-T-"
# Creating layers for the tensorflow network
# - The First layer is creating the input object, including the shape and dtype
# - Next create Dense layer with 30 neurons using ReLU activation function then passing through the input
# - Create the seccond hidden layer and pass in the previous layer
# - Create Concatenate layer creates a concatenate layer and immediately calls it with the given inputs
# - Create Output layer with single neuron and no activation function and call it like a function passing in the concatenated layer
# - Create a keras model
# + id="MxjYYJJ2O3Qw" executionInfo={"status": "ok", "timestamp": 1602714980178, "user_tz": -660, "elapsed": 990, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgxpvArds7xLNZB29BI3aCcSZkWbMPAZWZLOm3o=s64", "userId": "03940255243131127536"}}
X_train_A, X_train_B = X_train[:, :5], X_train[:, 2:]
X_valid_A, X_valid_B = X_valid[:, :5], X_valid[:, 2:]
X_test_A, X_test_B = X_test[:, :5], X_test[:, 2:]
X_new_A, X_new_B = X_test_A[:3], X_test_B[:3]
input_A = keras.layers.Input(shape=[5], name="wide_input")
input_B = keras.layers.Input(shape=[6], name="deep_input")
hidden1 = keras.layers.Dense(30, activation="relu")(input_B)
hidden2 = keras.layers.Dense(30, activation="relu")(hidden1)
concat = keras.layers.concatenate([input_A, hidden2])
output = keras.layers.Dense(1, name="main_output")(concat)
aux_output = keras.layers.Dense(1, name="aux_output")(hidden2)
model = keras.models.Model(inputs=[input_A, input_B],
outputs=[output, aux_output])
# + [markdown] id="xWvKU_e3PcZj"
# Creating multiple layers with more than one input
# + id="GID8IvfXPgmN" executionInfo={"status": "ok", "timestamp": 1602714992584, "user_tz": -660, "elapsed": 11000, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgxpvArds7xLNZB29BI3aCcSZkWbMPAZWZLOm3o=s64", "userId": "03940255243131127536"}} outputId="12dc08ab-7b71-401b-b6bf-487efe11355b" colab={"base_uri": "https://localhost:8080/", "height": 785}
model.compile(loss="mse", loss_weights=[0.1], optimizer=keras.optimizers.SGD(lr=1e-3))
history = model.fit([X_train_A, X_train_B], [y_train, y_train], epochs=20,
validation_data=([X_valid_A, X_valid_B], [y_valid, y_valid]))
# + [markdown] id="0STFJPvTPptj"
# Compile and Build
| Hands-on-ML/Code/Chapter 10/Functional_API.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **4. Multi class clasisification for RockPaperScissors**
#
# ## **Abstract**
#
# Aim fo the notebook is to demonstarte the multiclass clasification for data set of Rock Paper Scissiors
# Rock Paper Scissors is a dataset containing 2,892 images of diverse hands in Rock/Paper/Scissors poses. It is licensed CC By 2.0 and available for all purposes, but it’s intent is primarily for learning and research.
#
# Rock Paper Scissors contains images from a variety of different hands, from different races, ages and genders, posed into Rock / Paper or Scissors and labelled as such. You can download the training set here, and the test set here. These images have all been generated using CGI techniques as an experiment in determining if a CGI-based dataset can be used for classification against real images. I also generated a few images that you can use for predictions. You can find them here.
#
# Note that all of this data is posed against a white background.
#
# Each image is 300×300 pixels in 24-bit color
#
# Dataset was genrated by the Laurence Moroney - http://www.laurencemoroney.com/rock-paper-scissors-dataset/
# + colab={"base_uri": "https://localhost:8080/", "height": 398} colab_type="code" id="it1c0jCiNCIM" outputId="b358b039-cb92-4464-d5a2-b6d16b52c78c"
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/rps.zip \
# -O /tmp/rps.zip
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/rps-test-set.zip \
# -O /tmp/rps-test-set.zip
# + colab={} colab_type="code" id="PnYP_HhYNVUK"
import os
import zipfile
local_zip = '/tmp/rps.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp/')
zip_ref.close()
local_zip = '/tmp/rps-test-set.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp/')
zip_ref.close()
# + colab={"base_uri": "https://localhost:8080/", "height": 141} colab_type="code" id="MrxdR83ANgjS" outputId="e7107f6b-72cb-4bc9-c81d-31d38283a78e"
rock_dir = os.path.join('/tmp/rps/rock')
paper_dir = os.path.join('/tmp/rps/paper')
scissors_dir = os.path.join('/tmp/rps/scissors')
print('total training rock images:', len(os.listdir(rock_dir)))
print('total training paper images:', len(os.listdir(paper_dir)))
print('total training scissors images:', len(os.listdir(scissors_dir)))
rock_files = os.listdir(rock_dir)
print(rock_files[:10])
paper_files = os.listdir(paper_dir)
print(paper_files[:10])
scissors_files = os.listdir(scissors_dir)
print(scissors_files[:10])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="jp9dLel9N9DS" outputId="937b1320-592f-43ce-964a-bd7652ecc035"
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
pic_index = 2
next_rock = [os.path.join(rock_dir, fname)
for fname in rock_files[pic_index-2:pic_index]]
next_paper = [os.path.join(paper_dir, fname)
for fname in paper_files[pic_index-2:pic_index]]
next_scissors = [os.path.join(scissors_dir, fname)
for fname in scissors_files[pic_index-2:pic_index]]
for i, img_path in enumerate(next_rock+next_paper+next_scissors):
#print(img_path)
img = mpimg.imread(img_path)
plt.imshow(img)
plt.axis('Off')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="LWTisYLQM1aM" outputId="a3dbb307-35dc-4afc-99a9-a17f47f80a5b"
import tensorflow as tf
import keras_preprocessing
from keras_preprocessing import image
from keras_preprocessing.image import ImageDataGenerator
TRAINING_DIR = "/tmp/rps/"
training_datagen = ImageDataGenerator(
rescale = 1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
VALIDATION_DIR = "/tmp/rps-test-set/"
validation_datagen = ImageDataGenerator(rescale = 1./255)
train_generator = training_datagen.flow_from_directory(
TRAINING_DIR,
target_size=(150,150),
class_mode='categorical',
batch_size=126
)
validation_generator = validation_datagen.flow_from_directory(
VALIDATION_DIR,
target_size=(150,150),
class_mode='categorical',
batch_size=126
)
model = tf.keras.models.Sequential([
# Note the input shape is the desired size of the image 150x150 with 3 bytes color
# This is the first convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu', input_shape=(150, 150, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
# The second convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The third convolution
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The fourth convolution
tf.keras.layers.Conv2D(128, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# Flatten the results to feed into a DNN
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.5),
# 512 neuron hidden layer
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dense(3, activation='softmax')
])
model.summary()
model.compile(loss = 'categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
history = model.fit(train_generator, epochs=25, steps_per_epoch=20, validation_data = validation_generator, verbose = 1, validation_steps=3)
model.save("rps.h5")
# + colab={"base_uri": "https://localhost:8080/", "height": 298} colab_type="code" id="aeTRVCr6aosw" outputId="d46f165d-da40-4e12-9667-c03fd14c8004"
import matplotlib.pyplot as plt
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend(loc=0)
plt.figure()
plt.show()
# + colab={} colab_type="code" id="ZABJp7T3VLCU"
import numpy as np
from google.colab import files
from keras.preprocessing import image
uploaded = files.upload()
for fn in uploaded.keys():
# predicting images
path = fn
img = image.load_img(path, target_size=(150, 150))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
images = np.vstack([x])
classes = model.predict(images, batch_size=10)
print(fn)
print(classes)
# -
# ## **Refrence**
#
# https://www.coursera.org
#
# https://www.tensorflow.org/
#
# http://www.laurencemoroney.com/rock-paper-scissors-dataset/
#
# Copyright 2020 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| 4. Multi class clasisification for RockPaperScissors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df=pd.read_csv('../DATA/kc_house_data.csv')
sns.scatterplot(x='long',y='lat',data=df,hue='price',edgecolor='None',alpha=0.1)
df.head(1).count()
# +
df.head(1).count()
df = df.drop(['zipcode','id'],axis=1)
# -
df.head(1).count()
df['date'] = pd.to_datetime(df['date'])
df['year'] = df['date'].apply(lambda date : date.year)
df['month'] = df['date'].apply(lambda date : date.month)
df = df.drop('date',axis=1)
df2=df.sort_values('price',ascending=False).iloc[300:]
#sns.scatterplot(x='long',y='lat',data=df2,edgecolor='None',alpha=0.1,hue='price',palette='RdYlGn')
df2.head(1)
x = df2.drop('price',axis=1).values
y = df2['price'].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=101)
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
X_train.shape
model = Sequential()
model.add(Dense(19,activation='relu'))
model.add(Dense(19,activation='relu'))
model.add(Dense(19,activation='relu'))
model.add(Dense(19,activation='relu'))
model.add(Dense(1))
model.compile(optimizer='adam',loss='mse')
model.fit(x=X_train,y=y_train,validation_data=(X_test,y_test),batch_size=256,epochs=800)
loss = pd.DataFrame(model.history.history)
loss.plot()
from sklearn.metrics import mean_absolute_error,mean_squared_error,explained_variance_score
predictions = model.predict(X_test)
mean_absolute_error(y_test,predictions)
np.sqrt(mean_squared_error(y_test,predictions))
explained_variance_score(y_test,predictions)
plt.figure(figsize=(12,6))
plt.scatter(y_test,predictions)
plt.plot(y_test,y_test,'r')
| supervise_learning_ANN_models/real_estate_project of kingcounty.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Calibrating the Traffic
#
# After all schedules have been added to the database, now the distribution for the vehicle type of the onward transportation of the container need to be be calibrated accordingly.
# This distribution decides which vehicle type will pick up the container on its outbound journey.
# Calibrating this distribution to your input data is good practice as it gives you the chance to re-check your assumptions regarding the traffic.
# Technically speaking the distribution will be automatically adjusted during the generation process if required.
# It will issue a warning when doing so.
# In this Jupyter Notebook, it is show how the calibration can be done.
# +
import matplotlib.pyplot as plt
import seaborn as sns
from conflowgen import DatabaseChooser
from conflowgen import ContainerFlowGenerationManager
from conflowgen import InboundAndOutboundVehicleCapacityPreviewReport
from conflowgen import ContainerFlowByVehicleTypePreviewReport
from conflowgen import VehicleCapacityExceededPreviewReport
from conflowgen import ModalSplitPreviewReport
from conflowgen import ModeOfTransport
# -
# ## Load database
# Load information from database.
database_chooser = DatabaseChooser()
demo_file_name = "demo_deham_cta.sqlite"
if demo_file_name in database_chooser.list_all_sqlite_databases():
database_chooser.load_existing_sqlite_database(demo_file_name)
else:
print("Database is missing, nothing to do here")
# Get information regarding the container flow.
container_flow_generation_manager = ContainerFlowGenerationManager()
container_flow_properties = container_flow_generation_manager.get_properties()
for key, value in container_flow_properties.items():
print(f"{key:<60}: {value}")
# ## Load text reports
#
# Generate text reports.
# These have been tested with unit tests and serve as guidance for the visualizations.
# +
inbound_and_outbound_vehicle_capacity_preview_report = InboundAndOutboundVehicleCapacityPreviewReport()
report = inbound_and_outbound_vehicle_capacity_preview_report.get_report_as_text()
print("Inbound and outbound traffic: ")
print(report)
print()
container_flow_by_vehicle_type_preview_report = ContainerFlowByVehicleTypePreviewReport()
report = container_flow_by_vehicle_type_preview_report.get_report_as_text()
print("Container flow between vehicle types as defined by schedules and distributions: ")
print(report)
print()
modal_split_preview_report = ModalSplitPreviewReport()
report = modal_split_preview_report.get_report_as_text()
print("The same container flow expressed in terms of transshipment and modal split for the hinterland: ")
print(report)
print()
vehicle_capacity_exceeded_preview_report = VehicleCapacityExceededPreviewReport()
report = vehicle_capacity_exceeded_preview_report.get_report_as_text()
print("Consequences of container flow for outgoing vehicles: ")
print(report)
# -
# ## Plot inbound and outbound capacities
inbound_and_outbound_vehicle_capacity_preview_report.get_report_as_graph()
sns.set_palette(sns.color_palette())
plt.show()
# If the outbound capacity is the same like the maximum capacity, it means all capacities are used.
# This can either happen if the transport buffer is 0%.
# ## Plot intended container flows
fig = container_flow_by_vehicle_type_preview_report.get_report_as_graph()
fig.show()
# ## Plot capacity exceeded
vehicle_capacity_exceeded_preview_report.get_report_as_graph()
sns.set_palette(sns.color_palette())
plt.show()
# If a bar of the currently planned value exceeds the maximum, this means that there is a mismatch - there are more transports planned for the outbound journey of that vehicle than there is capacity left on these vehicles.
# ## Visualize transshipment share and modal split for hinterland
modal_split_preview_report.get_report_as_graph()
sns.set_palette(sns.color_palette())
plt.show()
# ## Combine visuals for tuning
#
# Here, you can adjust the mode of transport distribution and directly get the preview of how this will affect the container flow.
# Be aware that this is just a preview.
#
# For the CTA example, the calibration was based on the information available on
# https://www.hafen-hamburg.de/de/statistiken/modal-split/ during September 2021.
# At that time, 3 mio TEU were transshipped from vessel to vessel and 5.5 mio TEU were transshipped to or from the hinterland,
# corresponding to 35% were transshipment.
# Regarding the modal split for the hinterland, in September 20217% for train, 50.2% for trucks, and 2.8% for barges were reported.
# +
hypothesized_mode_of_transport_distribution = {
ModeOfTransport.truck: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) + 0.15,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) - 0.15
},
ModeOfTransport.train: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) + 0.15,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) - 0.15
},
ModeOfTransport.barge: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6),
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6)
},
ModeOfTransport.feeder: {
ModeOfTransport.truck: 0.8 / (0.8 + 1.9) * 0.502,
ModeOfTransport.train: 0.8 / (0.8 + 1.9) * 0.47,
ModeOfTransport.barge: 0.8 / (0.8 + 1.9) * 0.0028,
ModeOfTransport.feeder: 0,
ModeOfTransport.deep_sea_vessel: 1.9 / (0.8 + 1.9)
},
ModeOfTransport.deep_sea_vessel: {
ModeOfTransport.truck: 4.6 / (4.6 + 1.9) * 0.502,
ModeOfTransport.train: 4.6 / (4.6 + 1.9) * 0.47,
ModeOfTransport.barge: 4.6 / (4.6 + 1.9) * 0.0028,
ModeOfTransport.feeder: 1.9 / (4.6 + 1.9),
ModeOfTransport.deep_sea_vessel: 0
}
}
container_flow_by_vehicle_type_preview_report.hypothesize_with_mode_of_transport_distribution(hypothesized_mode_of_transport_distribution)
fig = container_flow_by_vehicle_type_preview_report.get_report_as_graph()
fig.show()
inbound_and_outbound_vehicle_capacity_preview_report.hypothesize_with_mode_of_transport_distribution(hypothesized_mode_of_transport_distribution)
inbound_and_outbound_vehicle_capacity_preview_report.get_report_as_graph()
sns.set_palette(sns.color_palette())
plt.show()
vehicle_capacity_exceeded_preview_report.hypothesize_with_mode_of_transport_distribution(hypothesized_mode_of_transport_distribution)
vehicle_capacity_exceeded_preview_report.get_report_as_graph()
sns.set_palette(sns.color_palette())
plt.show()
modal_split_preview_report.hypothesize_with_mode_of_transport_distribution(hypothesized_mode_of_transport_distribution)
modal_split_preview_report.get_report_as_graph()
plt.show()
# -
#
| examples/Jupyter_Notebook/input_data_inspection/calibrating_the_traffic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="./logo_UTN.svg" align="right" width="150" />
# #### Procesamiento Digital de Señales
#
# # Primeras pruebas hacia el análisis espectral
# #### <NAME>
#
# Comenzamos a tomar contacto con la transformada discreta de Fourier (DFT) y su implementación eficiente la FFT. Aprovechamos la oportunidad para presentar una aplicación de los notebooks de Jupyter y su potencial para presentar resultados de forma ordenada y elegante.
# +
# Módulos para Jupyter
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import matplotlib as mpl
#%% Inicialización de librerías
# Setup inline graphics: Esto lo hacemos para que el tamaño de la salida,
# sea un poco más adecuada al tamaño del documento
mpl.rcParams['figure.figsize'] = (14,7)
# Módulos para mi script propiamente dicho
import numpy as np
import matplotlib.pyplot as plt
from scipy.fftpack import fft
from pdsmodulos import print_markdown, print_subtitle, print_latex
# -
# Podemos intercalar bloques de texto y código casi sin restricciones. En este caso el código de inicialización lo dejamos resuelto en el bloque anterior.
# +
nn = 1000
fs = 1000
tt = np.arange(0.0, nn/fs, 1/fs)
ff = np.arange(0.0, fs, nn/fs)
# ahora podemos simular que los canales están desconectados,
# o que una señal de ruido blanco, normalmente distribuido ingresa al ADC.
canales_ADC = 1
a0 = 1 # Volt
f0 = nn/4 * fs/nn
# dd = np.sin(2*np.pi*f0*tt)
dd = np.random.uniform(-np.sqrt(12)/2, +np.sqrt(12)/2, size = [nn,canales_ADC])
# dd = np.random.normal(0, 1.0, size = [N,canales_ADC])
DD = fft( dd, axis = 0 )
bfrec = ff <= fs/2
plt.figure()
plt.plot( ff[bfrec], np.abs(DD[bfrec]) )
plt.ylabel('Módulo [¿Unidades?]')
plt.xlabel('Frecuencia [Hz]')
plt.figure()
plt.plot( ff[bfrec], np.abs(DD[bfrec])**2 )
plt.ylabel('Densidad de Potencia [¿Unidades?]')
plt.xlabel('Frecuencia [Hz]')
plt.show()
# -
# ## Teorema de Parseval
#
# Para practicar te dejo las siguientes consignas:
#
# 1. Editá este notebook y agregá una breve explicación de cómo aplicarías el teorema de Parseval a las señales que te presento más arriba en este mismo notebook.
# 2. Escribí la ecuación del teorema en Latex, podés copiarla de la bibliografía.
#
# $ \sum\limits_{n=0}^{N-1} ?? = \frac{1}{N} \sum\limits_{k=0}^{N-1} ?? $
#
# 3. En un bloque de código, verificá que dicho teorema se cumple, con alguna experimentación con señales que vos generes.
# +
# Algo que podría resultarte útil alguna vez es generar Markdown en tu propio código, tal vez
# para presentar una tabla, resultado, etc. Acá te dejo unos ejemplos
print_subtitle('Teorema de Parseval (generado dinámicamente desde código)')
print_markdown('Te dejo unas funciones que te pueden servir si alguna vez quisieras generar Markdown desde tus scripts.')
# ojo que la "r" antes del string es IMPORTANTE!
print_latex(r'\sigma^2 = \frac{s+2}{p+1}')
# +
## Escribí tu respuesta a partir de aquí ...
| DFT primeras pruebas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from pyspark.sql import SparkSession, DataFrame
import os
local=False
if local:
spark=SparkSession.builder.master("local[4]") \
.config('spark.jars.packages', 'org.postgresql:postgresql:42.2.24') \
.appName("RemoveDuplicates").getOrCreate()
else:
spark=SparkSession.builder \
.master("k8s://https://kubernetes.default.svc:443") \
.appName("RemoveDuplicates") \
.config("spark.kubernetes.container.image","inseefrlab/jupyter-datascience:master") \
.config("spark.kubernetes.authenticate.driver.serviceAccountName",os.environ['KUBERNETES_SERVICE_ACCOUNT']) \
.config("spark.kubernetes.namespace", os.environ['KUBERNETES_NAMESPACE']) \
.config("spark.executor.instances", "4") \
.config("spark.executor.memory","8g") \
.config('spark.jars.packages','org.postgresql:postgresql:42.2.24') \
.getOrCreate()
# ! kubectl get pods
data=["hadoop spark","hadoop flume","spark kafka","hello spark"]
textRdd=spark.sparkContext.parallelize(data)
splitRdd=textRdd.flatMap(lambda item: item.split(" "))
tupleRdd=splitRdd.map(lambda item: (item,1))
reduceRdd=tupleRdd.reduceByKey(lambda x,y:x+y)
strRdd=reduceRdd.map(lambda item: f"{item[0]}, {item[1]}")
list=strRdd.collect()
for item in list:
print(item)
strRdd.toDebugString()
spark.sparkContext.stop()
| notebooks/.ipynb_checkpoints/word_count-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 9 Jan
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
data = pd.read_csv("train.csv")
data.head()
data.info()
data.dtypes
f1 = lambda x:0 if type (x)==float else 1
f1
data["Cabin"].apply(f1)
data["Has_cabin"]=data["Cabin"].apply(f1)
data.head()
data["familysize"] = data["SibSp"] + data["Parch"] + 1
data.head()
data[(data["Embarked"]!="S")&(data["Embarked"]!="C")&(data["Embarked"]!="Q")]
data.iloc[829]
data[data["Embarked"].isnull()]
data["Embarked"].value_counts()
data["Embarked"] = data["Embarked"].fillna("S")
data[data["Fare"].isnull()]
data.iloc[829]
# # 13 Jan
data[data["Age"].isnull()]
data["Age"].isnull().sum()
age_avg = data["Age"].mean()
age_std = data["Age"].std()
age_null_count = data["Age"].isnull().sum()
age_avg
age_std
age_null_count
age_null_random_list = np.random.randint(age_avg - age_std, age_avg + age_std, size=age_null_count)
age_null_random_list
data[np.isnan(data["Age"])]
data.loc[np.isnan(data["Age"]), "Age"] = age_null_random_list
data["Age"].isnull().sum()
data.info()
data.dtypes
data["Age"] = data["Age"].astype(int)
data["Age"]
data["Sex"] = data["Sex"].map({"female": 0, "male": 1}).astype(int)
data.head()
# # Working on title
import re
def get_title(name):
title_search = re.search("([A-Za-z]+\.),name")
if title_search:
return title_search[0]
re.search("([A-Za-z]+\.)", "Cumings, Mrs. <NAME>")
zz=re.search("([A-Za-z]+\.)", "Cumings Mrs. <NAME>")
zz[0]
data["Title"] = data["Name"].apply(get_title)
data.head()
| Titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Preregistration
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # What is preregistration?
#
# - Specifying to your plan in advance, before you gather data.
# - Preregistration separates hypothesis-generating (exploratory) from hypothesis-testing (confirmatory) research
# - Both are important.
# - The same data cannot be used to generate and test a hypothesis
# - Nevertheless in reality, this appens often
# - This often happens unintentionally
# - The consequence is that it reduces the credibility of you as a scientist and your results
# - Addressing this problem through planning:
# - Improves the quality and transparency of your research
# - Helps others who may wish to build on it
# + [markdown] slideshow={"slide_type": "slide"}
# # Why would I preregister a study?
#
# - Increase reproducibility
# - Increase trust in research by being transparent
# - Fight against Questionable Research Practices
# + [markdown] slideshow={"slide_type": "slide"}
# ## I am not doing anything wrong. Why should I waste my time?
#
# - This isn't about paying a pennance for doing something wrong
# - This is about taking collective action against Questionable Research Practices
#
# #### Leading by example
# - By voluntarily being transparant about research practices, we can:
# - Regain trust in research
# - Create a research environment that is more difficult for Questionable Research Practices to thrive in
# - This is about putting the benefit of the scientific community before the ease of our current research climate
# + [markdown] slideshow={"slide_type": "slide"}
# # An example preregistration of a study of mine
#
# <a href="https://osf.io/pdwcb" target="_blank">Automate Formant Ceiling</a>
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Challenges
#
# - Changes to procedure during study administration
# - Discovery of assumption violations during analysis
# - Data are pre-existing
# - Longitudinal studies and large, multivariate datasets
# - Many experiments
# - A program of research
# - Few a priori expectations
# - Competing predictions
# - Narritive inferences and conclusions
# + [markdown] slideshow={"slide_type": "subslide"}
# # Changes to procedure during study administration
# - What if you didn't account for:
# - the fact that half of your babies are falling asleep during testing
# - your mice decided to eat each other
# - your adult subjects are bored with your study
# - you made a mistake in your procedure
# - your algorithm parameters are unsuitible
#
# <H2> Don't Panic, Document it</H2>
# <h3>With transparent reporting, readers can assess the deviations and their rationale</h3>
# + [markdown] slideshow={"slide_type": "subslide"}
# # Discovery of assumption violations during analysis
#
# - What if your data violate normality assumptions or other similar situations?
# - Some studies can be preregistered in stages
# - Register study, plan a normality check
# - Register analysis type after normality checks
# - Register a decision tree
# - Establish standard operating procedures (SOPs) that accompany one
# or many preregistrations
#
# - Just document any changes you make
# + [markdown] slideshow={"slide_type": "subslide"}
# # Data are pre-existing
# - Meta analysis?
# - Analyze old data in new way?
# - Came to a new lab and got given data?
#
# #### A study can be preregistered at any stage before analysis
# - Analysis plan must be blind to research outcomes
# - If nobody has observed data, pre-registration is possible
# - If someone has observed the data, it may not be possible to preregister it
# - A replication could be preregistered
#
#
# - Meta analysis is allowed in pre-registration if blind to research outcomes
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Longitudinal studies and large, multivariate datasets
#
# What do you do if you cannot preregister the entire design and analysis plan for all
# future papers at project onset?
#
#
# - Preregistered in stages
# - Register a decision tree
# - Establish standard operating procedures (SOPs) that accompany one or many preregistrations
# - Document changes to procedures and deviations from original preregistration
# - Analysis plan must be blind to research outcomes
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Too Many Experiments
#
# What do you do if you have a lab that runs lots of experiments? Preregistration takes too much time.
#
# - Are you running a research paradigm?
# - Create a preregistration template
# - Defines the variables and parameters for the protocol
# - Change template for each new experiment
# - Documenting process slowing you down because data collection is so easy?
# - First do an undocumented exploratory study
# - Then preregister a confirmatory replication
# + [markdown] slideshow={"slide_type": "subslide"}
# # Program of Research
# - Analysis plan must be blind to research outcomes
# - All outcomes of analysis plan must be reported
# - No selective reporting!
# - No file drawers
# - Does not overcome multiple comparisons
# + [markdown] slideshow={"slide_type": "subslide"}
# # Few *a priori* expectations
# What if you are just trying to discover something? What if you are *just* exploring?
#
# - You probably still do have a few basic predictions.
# - Run the study as an exploratory study
# - Replicate the study as a preregistered confirmatory study
# - If dataset is large enough, split in half
# - Explore in one half, confirm on other half
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Narrative inferences and conclusions
#
# What if only 1/10 statistical tests are significant. How to avoid focusing only on the "successful" parts of the study?
#
# - Preregistration prevents people from reporting only those statistical inferences that fit a narrative
# - Preregistration does not prevent selective attention of readers and authors to focus only on results deemed "successful" or "interesting"
# + [markdown] slideshow={"slide_type": "slide"}
#
# Mostly adapted from: <NAME>., <NAME>., <NAME>., & <NAME>. (2018). The preregistration revolution. Proceedings of the National Academy of Sciences, 115(11), 2600-2606.f
# + [markdown] slideshow={"slide_type": "slide"}
# # Questions and Discussion
| 03-Preregistration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Enable importing of utilities
import sys
sys.path.append('..')
# %matplotlib inline
from pylab import rcParams
rcParams['figure.figsize'] = 10, 10
# -
# # Water Classification and Analysis of Lake Chad
#
# The [previous tutorial](igarrs_chad_02.ipynb) introduced Landsat 7 imagery. The Lake Chad dataset was split into **pre** and **post** rainy season data-sets. The datasets were then cleaned up to produce a cloud-free and SLC-gap-free composite.
#
# This tutorial will focus on analyzing bodies of water using the results of a water classification algorithm called [WOFS]()
#
# <br>
# # What to expect from this notebook:
# <br>
#
# - Loading in NETCDF files
# - Introduction to WOFS for water classification
# - Built in plotting utilities of xarrays
# - Band arithmetic using xarrays
# - Analysis of lake chad; pre and post rainy season
# <br>
# # Algorithmic Process
# <br>
#
# 
#
# <br>
#
# The algorithmic process is fairly simple. It is a chain of operations on our composite imagery. The goal here is to use water classifiers on our composite imagery to create comparabe water-products. Then to use the difference between the water products as a change classifier.
#
# <br>
#
# 1. load composites for pre and post rainy season(genereated in previous notebook)
# 2. run WOFS water classifier on both composites. (This should xarrays where where 1 is water, 0 is not water)
# 3. calculate the difference between post and pre water products to generate a water change product.
# 4. count all the positive values for **water gain** estimate
# 4. counnt all the negative values for **water loss** estimate
#
# <br>
# # Loading in composites
#
# <br>
#
# In our [previous notebook](igarrs_chad_02.ipynb) two composites were created to represent cloud and SLC-gap imagery of **pre-rainy season** and **post rainy season** Landsat7 imagery. They were saved NETCDF files to use in this tutorial.
#
# Xarrays were designed with NETCDF as it's primary storage format so loading them should be a synch. Start with the import:
# <br>
#
import xarray as xr
# <br>
# ### Load Pre Rainy Season composite
pre_rain = xr.open_dataset('../demo/pre_rain.nc')
# Lets print its contents as a high level check that data is loaded.
pre_rain
# <br>
# The `pre_rain` xarray should represents an area that looks somewhat like this:
# 
# >Note: *figure above is cached result*
# <br>
# ## Load Post Rainy Season Composite
#
post_rain = xr.open_dataset('../demo/post_rain.nc')
# Lets print this one as well
post_rain
# The post xarray represents an area that looks somewhat like this:
#
# 
#
# >Note: *figure above is cached result*
# # Water classification
# The goal of water classification is to classify each pixel as water or not water. The applications of water classification can range from identifying flood-plains or coastal boundaries, to observing trends like coastal erosion or the seasonal fluctuations of water. The purpose of this section is to classify bodies of water on pre and post rainy season composites so that we can start analyzing change in lake-chad's surface area.
# <br>
#
# 
#
# <br>
# ### WOFS Water classifier
#
# WOFS( Water Observations From Space) is a water classifier developed by the Australian government following extreme flooding in 2011. It uses a [regression tree](https://en.wikipedia.org/wiki/Logistic_model_tree) machine learning model trained on several geographically and geologically varied sections of the Australian continent on over 25 years of Landsat imagery.
#
# While details of its implementation are outside of the scope of this tutorial, you can:
#
# - access the Wofs code we're about to use on [our github](https://github.com/ceos-seo/data_cube_utilities/blob/master/dc_water_classifier.py)
# - read the original research [here](http://ac.els-cdn.com/S0034425715301929/1-s2.0-S0034425715301929-main.pdf?_tid=fb86c208-613b-11e7-92ff-00000aacb35e&acdnat=1499229771_4a94d67aaa7d03881fa5b0efc74b5c8e)
#
# <br>
# ### Running the wofs classifier
#
# Running the wofs classifier is as simple as running a function call. It is typically good practice to create simple functions that accept an Xarray Dataset and return a processed XARRAY Dataset with new data-arrays within it.
# <br>
# +
from utils.data_cube_utilities.dc_water_classifier import wofs_classify
import numpy as np
clean_mask = np.ones((pre_rain.sizes['latitude'],pre_rain.sizes['longitude'])).astype(np.bool)
pre_water = wofs_classify(pre_rain, clean_mask = clean_mask, mosaic = True)
print(pre_water)
post_water = wofs_classify(post_rain, clean_mask = clean_mask, mosaic = True)
# -
# <br>
# ### The structure of Wofs Data
# An interesting feature of Xarrays is their built-in support for plotting. Any [data-arrays](http://xarray.pydata.org/en/stable/api.html#dataarray) can plot its values using a plot function. Let's see what data-arrays come with wofs classifiers:
# <br>
#
pre_water
# <br>
# The printout shows that wofs produced a dataset with a single data-array called `wofs`. Lets see what sort of values are in `wofs` by running an np.unique command on it.
# <br>
np.unique(pre_water.wofs)
# <br>
# So wofs only ever assumes one of two values. 1 for water, 0 for not water. This should produce a highly contrasted images when plotted using Xarrays built in plotting feature.
# <br>
# <br>
# ### Pre-Rain Water Classifcations
pre_water.wofs.plot(cmap = "Blues")
# ### Post-Rain Water Classifications
post_water.wofs.plot(cmap = "Blues")
# <br>
# <br>
# # Differencing Water products to reveal Water change
# The two images rendered above aren't too revealing when it comes to observing significant trends in water change. Perhaps we should take advantage of Xarrays arithmetic capabilities to detect or highlight change in our water classes.
#
# <br>
# 
# <br>
# <br>
# Arithmetic operations like addition and subtraction can be applied to xarray datasets that share the same shape. For example, the following differencing operation ....
#
# <br>
water_change = post_water - pre_water
# <br>
# ... applies the difference operator to all values within the wofs data-array with extreme efficiency. If we were, to check unique values again...
# <br>
np.unique(water_change.wofs)
# <br>
# ... then we should encounter three values. 1, 0, -1. These values can be interpreted as values indicating change in water. The table below should serve as an incredibly clear reference:
# <br>
#
# \begin{array}{|c|c|}
# \hline post & pre & diff & interpretation \\\hline
# 1 & 0 & 1-0 = +1 & water gain \\\hline
# 0 & 1 & 0-1 = -1 & water loss \\\hline
# 1 & 1 & 1-1= 0 & no-change \\\hline
# 0 & 0 & 0-0=0 & no-change \\\hline
# \end{array}
#
# <br>
#
# Understanding the intuition and logic behind this differencing, I think we're ready to take a look at a plot of water change over the area...
# <br>
# <br>
#
water_change.wofs.plot()
# <br>
#
# ### Interpreting the plot. Relying on non-visual results
#
# The plot above shows a surprisingly different story from our expectation of water growth. Large sections of lake chad seem to have dis-appeared after the rainy season. The recommended next step would be to explore change by methods of counting.
# +
## Create a boolean xarray
water_growth = (water_change.wofs == 1)
water_loss = (water_change.wofs == -1)
## Casting a 'boolean' to an 'int' makes 'True' values '1' and 'False' Values '0'. Summing should give us our totals
total_growth = water_growth.astype(np.int8).sum()
total_loss = water_loss.astype(np.int8).sum()
# -
# <br>
# The results...
# <br>
#
print("Growth:", int(total_growth.values))
print("Loss:", int(total_loss.values))
print("Net Change:", int(total_growth - total_loss))
# <br>
# ### How to interpret these results
# Several guesses can be made here as to why water was lost after the rainy season. Since that is out scope for this lecture(and beyond the breadth of this developer's knowledge) I'll leave definitive answers to the right researchers in this field.
#
# What can be provided, however, is an addititional figure regarding trends precipitation.
#
#
# <br>
# ### Bringing back more GPM Data
#
# Lets bring back the GPM data one more time and increase the time range by one year in both directions.
#
# Instead of spanning the year of **2015** to **2016**, let's do **2014** to **2017**.
# > **Load GPM**
# > Using the same code from our first [gpm tutorial](igarrs_chad_01.ipynb), let's load in three years of rainfall data:
# <br>
#
# +
import datacube
dc = datacube.Datacube(app = "chad_rainfall")
## Define Geographic boundaries using a (min,max) tuple.
latitude = (12.75, 13.0)
longitude = (14.25, 14.5)
## Specify a date range using a (min,max) tuple
from datetime import datetime
time = (datetime(2014,1,1), datetime(2017,1,2))
## define the name you gave your data while it was being "ingested", as well as the platform it was captured on.
product = 'gpm_imerg_gis_daily_global'
platform = 'GPM'
measurements = ['total_precipitation']
gpm_data = dc.load(latitude = latitude, longitude = longitude,
product = product, platform = platform,
measurements=measurements)
# -
# <br>
# >** Display Data **
# >We'll aggregate spatial axis so that we're left with a mean value of the region for each point in time. Let's plot those points in a time series.
# <br>
# +
times = gpm_data.time.values
values = gpm_data.mean(['latitude', 'longitude']).total_precipitation.values
import matplotlib.pyplot as plt
plt.plot(times, values)
# -
# # Next Steps
#
# This concludes our series on observing the rainy season's contributions to Lake Chad's surface area. Hopefully you've taken an understanding of, or even interest to datacube and xarrays.
#
# I encourage you to check out more of our notebooks on [our github](https://github.com/ceos-seo/data_cube_notebooks) with applications ranging from [landslide detection](https://github.com/ceos-seo/data_cube_notebooks/blob/master/slip.ipynb) to [fractional coverage](https://github.com/ceos-seo/data_cube_notebooks/blob/master/fractional_coverage.ipynb) or even the [Wofs water detection algorithm](https://github.com/ceos-seo/data_cube_notebooks/blob/master/water_detection.ipynb)
| notebooks/IGARSS/igarss_chad_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 支持向量机 (support vector machines, SVM)
#
# > 支持向量机(support vector machines,SVM)是一种二分类模型,它将实例的特征向量映射为空间中的一些点,SVM 的目的就是想要画出一条线,以 “最好地” 区分这两类点,以至如果以后有了新的点,这条线也能做出很好的分类。SVM 适合中小型数据样本、非线性、高维的分类问题
#
# SVM学习的基本想法是
# 求解能够正确划分训练数据集并且几何间隔最大的分离超平面
#
# 对于线性可分的数据集来说,这样的超平面有无穷多个(即感知机),但是几何间隔最大的分离超平面却是唯一的。
#
# Advantages 优势:
# * Effective in high dimensional spaces. 在高维空间中有效。
# * Still effective in cases where number of dimensions is greater than the number of samples. 在尺寸数大于样本数的情况下仍然有效。
# * Uses a subset of training points in the decision function (called support vectors), so it is also memory efficient.
# 在决策函数中使用训练点的子集(称为支持向量),因此它也具有记忆效率。
# * Versatile: different Kernel functions can be specified for the decision function. Common kernels are provided, but it is also possible to specify custom kernels.
# 通用:可以为决策函数指定不同的核函数。提供了通用内核,但也可以指定自定义内核。
#
# disadvantages 缺点:
#
# * If the number of features is much greater than the number of samples, avoid over-fitting in choosing Kernel functions and regularization term is crucial.
# 当特征个数远大于样本个数时,在选择核函数和正则化项时应避免过拟合。
#
# * SVMs do not directly provide probability estimates, these are calculated using an expensive five-fold cross-validation (see Scores and probabilities, below).
# 支持向量机不直接提供概率估计,这些是使用昂贵的五倍交叉验证计算的(见下面的分数和概率)。
#
#
# [支持向量机](https://blog.csdn.net/qq_31347869/article/details/88071930)
# [sklearn文档-svm](https://scikit-learn.org/dev/modules/svm.html#svm)
# + [markdown] pycharm={"name": "#%% md\n"}
# The sklearn.svm module includes Support Vector Machine algorithms.
#
# |Estimators | description |
# |:---- |:---- |
# | svm.LinearSVC([penalty, loss, dual, tol, C, …]) | Linear Support Vector Classification. |
# | svm.LinearSVR(*[, epsilon, tol, C, loss, …]) | Linear Support Vector Regression. |
# | svm.NuSVC(*[, nu, kernel, degree, gamma, …]) | Nu-Support Vector Classification. |
# | svm.NuSVR(*[, nu, C, kernel, degree, gamma, …]) | Nu Support Vector Regression. |
# | svm.OneClassSVM(*[, kernel, degree, gamma, …]) | Unsupervised Outlier Detection. |
# | svm.SVC(*[, C, kernel, degree, gamma, …]) | C-Support Vector Classification. |
# | svm.SVR(*[, kernel, degree, gamma, coef0, …]) | Epsilon-Support Vector Regression. |
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Classification 使用支持向量机做分类任务
#
# SVC, NuSVC and LinearSVC are classes capable of performing binary and multi-class classification on a dataset.
# + pycharm={"name": "#%%\n"}
from sklearn.svm import SVC
import numpy as np
X = np.random.randint(0,10,(50,2))
y = (X[:,0] + X[:,1]) // 10
clf = SVC()
clf.fit(X, y)
clf.predict([[2., 7.],[3,9]]) # 结果会不一样
# + pycharm={"name": "#%%\n"}
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm, datasets
def make_meshgrid(x, y, h=.02):
"""Create a mesh of points to plot in
Parameters
----------
x: data to base x-axis meshgrid on
y: data to base y-axis meshgrid on
h: stepsize for meshgrid, optional
Returns
-------
xx, yy : ndarray
"""
x_min, x_max = x.min() - 1, x.max() + 1
y_min, y_max = y.min() - 1, y.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plot_contours(ax, clf, xx, yy, **params):
"""Plot the decision boundaries for a classifier.
Parameters
----------
ax: matplotlib axes object
clf: a classifier
xx: meshgrid ndarray
yy: meshgrid ndarray
params: dictionary of params to pass to contourf, optional
"""
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
out = ax.contourf(xx, yy, Z, **params)
return out
# import some data to play with
iris = datasets.load_iris()
# Take the first two features. We could avoid this by using a two-dim dataset
X = iris.data[:, :2]
y = iris.target
# we create an instance of SVM and fit out data. We do not scale our
# data since we want to plot the support vectors
C = 1.0 # SVM regularization parameter
models = (svm.SVC(kernel='linear', C=C),
svm.LinearSVC(C=C, max_iter=10000),
svm.SVC(kernel='rbf', gamma=0.7, C=C),
svm.SVC(kernel='poly', degree=3, gamma='auto', C=C))
models = (clf.fit(X, y) for clf in models)
# title for the plots
titles = ('SVC with linear kernel',
'LinearSVC (linear kernel)',
'SVC with RBF kernel',
'SVC with polynomial (degree 3) kernel')
# Set-up 2x2 grid for plotting.
fig, sub = plt.subplots(2, 2)
plt.subplots_adjust(wspace=0.4, hspace=0.4)
X0, X1 = X[:, 0], X[:, 1]
xx, yy = make_meshgrid(X0, X1)
for clf, title, ax in zip(models, titles, sub.flatten()):
plot_contours(ax, clf, xx, yy,
cmap=plt.cm.coolwarm, alpha=0.8)
ax.scatter(X0, X1, c=y, cmap=plt.cm.coolwarm, s=20, edgecolors='k')
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xlabel('Sepal length')
ax.set_ylabel('Sepal width')
ax.set_xticks(())
ax.set_yticks(())
ax.set_title(title)
plt.show()
# + [markdown] pycharm={"name": "#%% md\n"}
# Regression
# There are three different implementations of Support Vector Regression: SVR, NuSVR and LinearSVR. LinearSVR provides a faster implementation than SVR but only considers the linear kernel, while NuSVR implements a slightly different formulation than SVR and LinearSVR. See Implementation details for further details.
# + pycharm={"name": "#%%\n"}
from sklearn.svm import SVR
X = [[0, 0], [2, 2]]
y = [0.5, 2.5]
regr = SVR()
regr.fit(X, y)
regr.predict([[1, 1]])
# + [markdown] pycharm={"name": "#%% md\n"}
# Unsupervised Outlier Detection.
#
# Estimate the support of a high-dimensional distribution.
#
# OneClassSVM is based on libsvm.
# + pycharm={"name": "#%%\n"}
from sklearn.svm import OneClassSVM
X = [[0], [0.44], [0.45], [0.46], [1]]
clf = OneClassSVM(gamma='auto')
clf.fit(X)
result = clf.predict(X)
print(result)
scores = clf.score_samples(X)
print(scores)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 使用SVM做异常检测算法
#
# Comparing anomaly detection algorithms for outlier detection on toy datasets
#
# [refrence](https://scikit-learn.org/dev/auto_examples/miscellaneous/plot_anomaly_comparison.htm)
# + pycharm={"name": "#%%\n"}
# Author: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# License: BSD 3 clause
import time
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import sklearn
print(sklearn.__version__)
# + pycharm={"name": "#%%\n"}
from sklearn.datasets import make_moons, make_blobs
from sklearn.covariance import EllipticEnvelope
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
from sklearn.kernel_approximation import Nystroem
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import SGDOneClassSVM
print(__doc__)
matplotlib.rcParams['contour.negative_linestyle'] = 'solid'
# Example settings
n_samples = 300
outliers_fraction = 0.15
n_outliers = int(outliers_fraction * n_samples)
n_inliers = n_samples - n_outliers
# define outlier/anomaly detection methods to be compared.
# the SGDOneClassSVM must be used in a pipeline with a kernel approximation
# to give similar results to the OneClassSVM
anomaly_algorithms = [
("Robust covariance", EllipticEnvelope(contamination=outliers_fraction)),
("One-Class SVM", OneClassSVM(nu=outliers_fraction, kernel="rbf",
gamma=0.1)),
("One-Class SVM (SGD)", make_pipeline(
Nystroem(gamma=0.1, random_state=42, n_components=150),
SGDOneClassSVM(nu=outliers_fraction, shuffle=True,
fit_intercept=True, random_state=42, tol=1e-6)
)),
("Isolation Forest", IsolationForest(contamination=outliers_fraction,
random_state=42)),
("Local Outlier Factor", LocalOutlierFactor(
n_neighbors=35, contamination=outliers_fraction))]
# Define datasets
blobs_params = dict(random_state=0, n_samples=n_inliers, n_features=2)
datasets = [
make_blobs(centers=[[0, 0], [0, 0]], cluster_std=0.5,
**blobs_params)[0],
make_blobs(centers=[[2, 2], [-2, -2]], cluster_std=[0.5, 0.5],
**blobs_params)[0],
make_blobs(centers=[[2, 2], [-2, -2]], cluster_std=[1.5, .3],
**blobs_params)[0],
4. * (make_moons(n_samples=n_samples, noise=.05, random_state=0)[0] -
np.array([0.5, 0.25])),
14. * (np.random.RandomState(42).rand(n_samples, 2) - 0.5)]
# Compare given classifiers under given settings
xx, yy = np.meshgrid(np.linspace(-7, 7, 150),
np.linspace(-7, 7, 150))
plt.figure(figsize=(len(anomaly_algorithms) * 2 + 4, 12.5))
plt.subplots_adjust(left=.02, right=.98, bottom=.001, top=.96, wspace=.05,
hspace=.01)
plot_num = 1
rng = np.random.RandomState(42)
for i_dataset, X in enumerate(datasets):
# Add outliers
X = np.concatenate([X, rng.uniform(low=-6, high=6, size=(n_outliers, 2))],
axis=0)
for name, algorithm in anomaly_algorithms:
t0 = time.time()
algorithm.fit(X)
t1 = time.time()
plt.subplot(len(datasets), len(anomaly_algorithms), plot_num)
if i_dataset == 0:
plt.title(name, size=18)
# fit the data and tag outliers
if name == "Local Outlier Factor":
y_pred = algorithm.fit_predict(X)
else:
y_pred = algorithm.fit(X).predict(X)
# plot the levels lines and the points
if name != "Local Outlier Factor": # LOF does not implement predict
Z = algorithm.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contour(xx, yy, Z, levels=[0], linewidths=2, colors='black')
colors = np.array(['#377eb8', '#ff7f00'])
plt.scatter(X[:, 0], X[:, 1], s=10, color=colors[(y_pred + 1) // 2])
plt.xlim(-7, 7)
plt.ylim(-7, 7)
plt.xticks(())
plt.yticks(())
plt.text(.99, .01, ('%.2fs' % (t1 - t0)).lstrip('0'),
transform=plt.gca().transAxes, size=15,
horizontalalignment='right')
plot_num += 1
plt.show()
| part02-machine-learning/2_6_svm.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Introduction to Git and GitHub
# ## Setting up
#
# In this workshop we will learn some common ways of interacting with Git and GitHub.
#
# To follow along with this interactive workshop you will need a free [Microsoft account](https://account.microsoft.com/account) and a free [GitHub account](https://github.com/join). You can follow the links to create these accounts now, or you can create them when they are needed during the workshop.
#
# ## Introduction
#
# Git is a modern version control system. A version control system records changes to a file or set of files over time so that you can recall specific versions later ([About Version Control, Pro Git book](https://git-scm.com/book/en/v2/Getting-Started-About-Version-Control)). GitHub is an online services that helps people share projects that use Git as their version control system.
#
# The environment that you are viewing this notebook in is called Jupyter. You have access to a file browser which you can access by clicking `File > Open...`. You can also access a terminal window by clicking `File > Open...` and then on the right hand side of the screen selecting `New ▾ > Terminal` which can be used with common Linux commands. This will open up terminal in a new tab.
#
# ## Workshop notebooks
#
# This workshop containes a series of exercise that can be found in this directory `learn-github`. In each of the files (called a notebook) in this directory we will learn about a common workflow for interacting with Git and GitHub.
#
# To start learning one of the workflows simply double click on one of the notebooks or click one of the links below
#
# 1. [Introduction to Git and GitHub](./1-introduction.ipynb) *(this notebook)*
# 2. [Manage projects with Git](./2-manage-projects-with-git.ipynb)
# 3. [Making changes to projects](./3-making-changes-to-projects.ipynb)
# 4. [Project collaboration with GitHub](./4-project-collaboration-with-github.ipynb)
# 5. [(Bonus) Sharing your work on GitHub](./5-sharing-your-work-on-github.ipynb)
# In each of the notebooks are a list of blocks of code called *cells*. This are the grey boxes with a darker grey border and have `In [ ]:` on the left hand side. You can run the commands in the cells by clicking on them and then pressing the
#
# <i class="fa-step-forward fa"></i><span class="toolbar-btn-label">Run</span>
#
# button. Trying running the cell below
cat .hidden
# After running the cell you should see `hello, world 😊` appear below the cell.
#
# The cells in this workshop contain Git commands. All git commands have the same format
#
# ```bash
# git <git-command> [<value-1> <value-2> ...]
# ```
#
# When you see an expression like <value> you should replace it and the brackets `<` and `>` with an appropriate value which will depend on the situation. An expression like [optional] is optional. If you use it when running a command you should replace it and the brackets `[` and `]` with optional.
#
# ## Workshop exercises
#
# Throught the workshop notebooks there are exercises for you to test your understanding of the Git commands you will learn about. The exercises will look like the following example.
#
# > ### Exercises
# >
# > **How do you solve an exercise?**
# >
# > ```bash
# > Type your solution into the code cell underneath the question and run it like any other cell
# > ```
#
# To solve most exercises you will need to research the Git commands by clicking on the links that provide more details on the Git commands. For example in [2 Manage projects with Git](./2-manage-projects-with-git.ipynb#4-Browse-your-repositories-history) there is a link to learn more about the [git-log](https://git-scm.com/docs/git-log) command. Solutions to the workshop exercises will be provided after the workshop.
| learn-github/1-introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import scanpy as sc
import pandas as pd
import numpy as np
import scipy as sp
from statsmodels.stats.multitest import multipletests
import matplotlib.pyplot as plt
import seaborn as sns
from anndata import AnnData
import os
import time
from gprofiler import GProfiler
# scTRS tools
import scdrs.util as util
import scdrs.data_loader as dl
import scdrs.method as md
# autoreload
# %load_ext autoreload
# %autoreload 2
# -
DATA_FILE='/n/holystore01/LABS/price_lab/Users/mjzhang/scTRS_data/single_cell_data/tabula_sapiens/raw_data/TabulaSapiens.h5ad'
OUT_PATH='/n/holystore01/LABS/price_lab/Users/mjzhang/scTRS_data/single_cell_data/tabula_sapiens'
# +
adata_full = sc.read_h5ad(DATA_FILE)
adata_full.X = adata_full.raw.X
del adata_full.layers["decontXcounts"]
del adata_full.raw
adata_full.obs['tissue'] = adata_full.obs['organ_tissue']
adata_full.obs['tissue_celltype'] = ['%s.%s'%(x,y) for x,y in zip(adata_full.obs['tissue'],
adata_full.obs['cell_ontology_class'])]
for method in ['smartseq2', '10X']:
adata = adata_full[adata_full.obs['method']==method].copy()
# Before filtering
print(method)
print('# n_cell=%d, n_gene=%d'%(adata.shape[0], adata.shape[1]))
print('# n_tissue=%d'%(len(set(adata.obs['organ_tissue']))))
print('# n_celltype=%d'%(len(set(adata.obs['cell_ontology_class']))))
print('# n_tissue_celltype=%d'%(len(set(adata.obs['tissue_celltype']))))
# Remove tissue-cell types with <3 cells:
sc.pp.filter_cells(adata, min_genes=250)
sc.pp.filter_genes(adata, min_cells=50)
adata.write(OUT_PATH+'/obj_%s_raw.h5ad'%method)
# After filtering
print('After filtering')
print('# n_cell=%d, n_gene=%d'%(adata.shape[0], adata.shape[1]))
print('# n_tissue=%d'%(len(set(adata.obs['tissue']))))
print('# n_celltype=%d'%(len(set(adata.obs['cell_ontology_class']))))
print('# n_tissue_celltype=%d'%(len(set(adata.obs['tissue_celltype']))))
# -
# TS FACS
DATA_PATH = '/n/holystore01/LABS/price_lab/Users/mjzhang/scDRS_data/single_cell_data/tabula_sapiens'
adata_raw = sc.read_h5ad(DATA_PATH+'/obj_smartseq2_raw.h5ad')
# Make .cov file
df_cov = pd.DataFrame(index=adata_raw.obs.index)
df_cov['const'] = 1
df_cov['n_genes'] = (adata_raw.X>0).sum(axis=1)
for donor in sorted(set(adata_raw.obs['donor'])):
if donor!='TSP1':
df_cov['donor_%s'%donor] = (adata_raw.obs['donor']==donor)*1
df_cov.to_csv(DATA_PATH+'/ts_smartseq2.cov', sep='\t')
# TS Droplet
DATA_PATH = '/n/holystore01/LABS/price_lab/Users/mjzhang/scTRS_data/single_cell_data/tabula_sapiens'
adata_raw = sc.read_h5ad(DATA_PATH+'/obj_10X_raw.h5ad')
# Make .cov file
df_cov = pd.DataFrame(index=adata_raw.obs.index)
df_cov['const'] = 1
df_cov['n_genes'] = (adata_raw.X>0).sum(axis=1)
df_cov.to_csv(DATA_PATH+'/ts_10X.cov', sep='\t')
| experiments/job.curate_data/curate_ts_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from google.cloud import storage
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = '../t5/prepare/mesolitica-tpu.json'
client = storage.Client()
bucket = client.bucket('mesolitica-tpu-general')
# +
import tensorflow as tf
import tensorflow_datasets as tfds
from t5.data import preprocessors as prep
import functools
import t5
import gin
import sentencepiece as spm
from glob import glob
import os
from tensor2tensor.data_generators import problem
from tensor2tensor.data_generators import text_problems
from tensor2tensor.utils import registry
gin.parse_config_file('pretrained_models_base_operative_config.gin')
vocab = 'sp10m.cased.t5.model'
sp = spm.SentencePieceProcessor()
sp.Load(vocab)
# +
# import sentencepiece as spm
# vocab = 'sp10m.cased.t5.model'
# sp = spm.SentencePieceProcessor()
# sp.Load(vocab)
# +
def cnn_dataset(split, shuffle_files = False):
del shuffle_files
ds = tf.data.TextLineDataset(glob('t5-data/cnn-summarization-*.tsv'))
ds = ds.map(
functools.partial(
tf.io.decode_csv,
record_defaults = ['', ''],
field_delim = '\t',
use_quote_delim = False,
),
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
ds = ds.map(lambda *ex: dict(zip(['question', 'answer'], ex)))
return ds
def cnn_preprocessor(ds):
def to_inputs_and_targets(ex):
return {
'inputs': tf.strings.join(['ringkasan: ', ex['question']]),
'targets': ex['answer'],
}
return ds.map(
to_inputs_and_targets,
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
t5.data.TaskRegistry.remove('cnn_dataset')
t5.data.TaskRegistry.add(
'cnn_dataset',
dataset_fn = cnn_dataset,
splits = ['train'],
text_preprocessor = [cnn_preprocessor],
sentencepiece_model_path = vocab,
metric_fns = [t5.evaluation.metrics.accuracy],
)
def multinews_dataset(split, shuffle_files = False):
del shuffle_files
ds = tf.data.TextLineDataset(glob('t5-data/multinews-summarization-*.tsv'))
ds = ds.map(
functools.partial(
tf.io.decode_csv,
record_defaults = ['', ''],
field_delim = '\t',
use_quote_delim = False,
),
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
ds = ds.map(lambda *ex: dict(zip(['question', 'answer'], ex)))
return ds
def multinews_preprocessor(ds):
def to_inputs_and_targets(ex):
return {
'inputs': tf.strings.join(['ringkasan: ', ex['question']]),
'targets': ex['answer'],
}
return ds.map(
to_inputs_and_targets,
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
t5.data.TaskRegistry.remove('multinews_dataset')
t5.data.TaskRegistry.add(
'multinews_dataset',
dataset_fn = multinews_dataset,
splits = ['train'],
text_preprocessor = [multinews_preprocessor],
sentencepiece_model_path = vocab,
metric_fns = [t5.evaluation.metrics.accuracy],
)
def news_dataset(split, shuffle_files = False):
del shuffle_files
ds = tf.data.TextLineDataset(glob('t5-data/news-title-*.tsv'))
ds = ds.map(
functools.partial(
tf.io.decode_csv,
record_defaults = ['', ''],
field_delim = '\t',
use_quote_delim = False,
),
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
ds = ds.map(lambda *ex: dict(zip(['question', 'answer'], ex)))
return ds
def news_preprocessor(ds):
def to_inputs_and_targets(ex):
return {
'inputs': tf.strings.join(['tajuk: ', ex['question']]),
'targets': ex['answer'],
}
return ds.map(
to_inputs_and_targets,
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
t5.data.TaskRegistry.remove('news_dataset')
t5.data.TaskRegistry.add(
'news_dataset',
dataset_fn = news_dataset,
splits = ['train'],
text_preprocessor = [news_preprocessor],
sentencepiece_model_path = vocab,
metric_fns = [t5.evaluation.metrics.accuracy],
)
# +
from tqdm import tqdm
@registry.register_problem
class Seq2Seq(text_problems.Text2TextProblem):
@property
def approx_vocab_size(self):
return 32100
@property
def is_generate_per_split(self):
return False
@property
def dataset_splits(self):
return [{
"split": problem.DatasetSplit.TRAIN,
"shards": 100,
}]
def generate_samples(self, data_dir, tmp_dir, dataset_split):
del data_dir
del tmp_dir
del dataset_split
nq_task = t5.data.TaskRegistry.get("cnn_dataset")
ds = nq_task.get_dataset(split='qa.tsv', sequence_length={"inputs": 1024, "targets": 1024})
for ex in tqdm(tfds.as_numpy(ds)):
yield ex
nq_task = t5.data.TaskRegistry.get("multinews_dataset")
ds = nq_task.get_dataset(split='qa.tsv', sequence_length={"inputs": 1024, "targets": 1024})
for ex in tqdm(tfds.as_numpy(ds)):
yield ex
nq_task = t5.data.TaskRegistry.get("news_dataset")
ds = nq_task.get_dataset(split='qa.tsv', sequence_length={"inputs": 768, "targets": 1024})
for ex in tqdm(tfds.as_numpy(ds)):
if len(ex['targets']) > 4:
yield ex
def generate_encoded_samples(self, data_dir, tmp_dir, dataset_split):
generator = self.generate_samples(data_dir, tmp_dir, dataset_split)
for sample in generator:
sample["inputs"] = sample['inputs'].tolist()
sample["targets"] = sample['targets'].tolist()
yield sample
# -
# !rm -rf t2t-summarization/data
# +
DATA_DIR = os.path.expanduser("t2t-summarization/data")
TMP_DIR = os.path.expanduser("t2t-summarization/tmp")
tf.gfile.MakeDirs(DATA_DIR)
tf.gfile.MakeDirs(TMP_DIR)
# +
from tensor2tensor.utils import registry
from tensor2tensor import problems
PROBLEM = 'seq2_seq'
t2t_problem = problems.problem(PROBLEM)
t2t_problem.generate_data(DATA_DIR, TMP_DIR)
# +
from glob import glob
files = glob('t2t-summarization/data/*')
files
# -
for file in files:
print(file)
blob = bucket.blob(file)
blob.upload_from_filename(file)
| session/summarization/t2t/t2t-summarization-generate-sentencepiece.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="8mGJmHWihAuJ"
# lambda를 사용하면 아래와 같은 함수 정의를 간단하게 나타냄
# + colab={"base_uri": "https://localhost:8080/"} id="LP86ORr0cqf_" outputId="94ee593d-a4ba-475e-8174-f134f91664e3"
(lambda first, second : first * second + 20)(10, 3)
# + id="3Plklj0mftPf"
def plus(first, second) : # 함수 정의
result = first + 20
return result
# + colab={"base_uri": "https://localhost:8080/"} id="MV_ymruOgZQG" outputId="db639d4e-4cf2-4356-c5f8-c2a1a1a3a948"
plus(10)
# + [markdown] id="B0KOLt17hJy_"
# lambda를 변수안에 저장하면 재사용가능
# + id="KYPAPnnigk73"
plus_lambda = (lambda first: first + 20) # lambda 정의
# + colab={"base_uri": "https://localhost:8080/"} id="64uUSV3rhiHf" outputId="e23e8a76-305b-47af-dbf2-87cfe7c3d5b5"
plus_lambda(10)
# + id="4rTPjJf3iHoP"
| python_lambda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/WarmXD/Elective-1-3/blob/main/Operations_and_Expressions_in_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="24BR8e3I0M09" outputId="6dae33b2-0618-4efd-b6a8-6c1e21894bbc"
x=10
y=9
print(x>y)
print(9==10)
print(10==10)
# + [markdown] id="cXykF-QK2U_i"
#
# + colab={"base_uri": "https://localhost:8080/"} id="7vuC6o441T9h" outputId="92a4c82c-5561-4acd-89fa-094b4b73c90d"
print(bool("Wayne"))
print(bool(18))
print(bool(None))
print(bool(1))
# + [markdown] id="2Ii2rSKc2XFW"
# ## Defining Function
#
# + colab={"base_uri": "https://localhost:8080/"} id="ambIQ8a-2BBp" outputId="7f2642fb-e24c-4bb4-e5c0-49dcd2f017e0"
def Pogi():
return False
print(Pogi())
# + colab={"base_uri": "https://localhost:8080/"} id="IdekGFg02_El" outputId="8ae33286-3974-4b8c-baa3-733e679ec290"
def myFunction():
return True
if myFunction():
print("Yes")
else:
print("No")
# + [markdown] id="_ewSb1hL3geB"
# ##Application 1
#
# + colab={"base_uri": "https://localhost:8080/"} id="AtLwmMbJ3jQ1" outputId="9385914d-bffa-4d41-8d18-53c9cfb8ef1a"
print (10>9)
a=6
b=7
print(a==b)
print(a!=b)
# + [markdown] id="c3CKQfr54yDR"
# ##Arithmetic Operations
# + colab={"base_uri": "https://localhost:8080/"} id="KKhHDk7U402I" outputId="dba1d33f-b375-46e0-85a5-2be362812eb6"
print(a+b)
print(a-b)
print(a*b)
print(a**b)
# + [markdown] id="m6_eKs2S5kaw"
# ##Bitwise Operators
# + colab={"base_uri": "https://localhost:8080/"} id="h80cchHw5m81" outputId="3beb07ac-80fa-4c99-c136-9893beada89f"
c = 60
d = 13
print(c&d)
print(c|d)
print(c<<1)
print(c<<2)
# + [markdown] id="aihmYqts77kj"
# #Assignment Operators
# + colab={"base_uri": "https://localhost:8080/"} id="-8_iFiN27-ea" outputId="23b9dafe-ed93-4d3e-8cfd-a4afa759761d"
c=60
d=13
c+=3
c%=3
print(c)
print(c)
# + [markdown] id="S4TkuE7s80E_"
# ##Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="WvG0pfPr81-e" outputId="3760db1a-954d-445b-f290-7328c5e2f273"
c = True
d = False
not(c and d)
# + [markdown] id="XPSiW6jk9Oki"
# ##Identity Operators
# + colab={"base_uri": "https://localhost:8080/"} id="CNPiMDkK9SGj" outputId="f46fb5da-8a83-43f1-9a4c-ebc5b58f5195"
c is d
c is not d
# + [markdown] id="SnL8edP99f9G"
# ##Application 2
# + colab={"base_uri": "https://localhost:8080/"} id="r3KP6bw49hsr" outputId="12c0447f-3ffa-4ca0-ddd2-d36e0cac0528"
e = 10
f = 5
# Implement the operations +, //, and bit shift right >> twice
print(e+f)
print(e//f)
print(e>>2)
print(f>>2)
| Operations_and_Expressions_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cambridge
# language: python
# name: cambridge
# ---
# # GANITE(Tensorflow): Train and Evaluation
# This notebook presents the solution for training and evaluating the __GANITE__ algorithm(__Tensoflow__ version) over the [Twins](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/data/twins/) dataset..
#
# The implementation of GANITE is adapted in the local `ite` library. For the Unified API version, check [this notebook](https://github.com/bcebere/ite-api/blob/main/notebooks/unified_api_train_evaluation.ipynb).
# ## GANITE
#
# Estimating Individualized Treatment Effects(__ITE__) is the task that approximates whether a given treatment influences or determines an outcome([read more](https://www.vanderschaar-lab.com/individualized-treatment-effect-inference/)).
#
# [__GANITE__](https://openreview.net/pdf?id=ByKWUeWA-)(Generative Adversarial Nets for inference of Individualized Treatment Effects) is a framework for inferring the ITE using GANs.
#
# The implementation demonstrated in this notebook is [here](https://github.com/bcebere/ite-api/tree/main/src/ite/algs/ganite) and is adapted from [this implementation](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/alg/ganite/).
# ## Setup
#
# First, make sure that all the depends are installed in the current environment.
# ```
# pip install -r requirements.txt
# pip install .
# ```
#
# Next, we import all the dependencies necessary for the task.
# +
# Double check that we are using the correct interpreter.
import sys
print(sys.executable)
# Disable TF logging
import os
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
# Depends
import ite.algs.ganite.model as alg
import ite.datasets as ds
import ite.utils.tensorflow as utils
from matplotlib import pyplot as plt
import pandas as pd
import tensorflow.compat.v1 as tf
# -
# ## Load the Dataset
#
# The example is done using the [Twins](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/data/twins/) dataset.
#
# Next, we load the dataset, process the data, and sample a training set and a test set.
#
# The logic is implemented [here](https://github.com/bcebere/ite-api/tree/main/src/ite/datasets), and it adapted from the original [GANITE pre-processing implementation](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/alg/ganite/data_preprocessing_ganite.py).
# +
train_ratio = 0.8
dataloader = ds.load("twins", train_ratio)
[Train_X, Train_T, Train_Y, Opt_Train_Y, Test_X, Test_Y] = dataloader
pd.DataFrame(data=Train_X[:5])
# -
# ## Load the model
#
# Next, we define the model.
#
#
# The constructor supports the following parameters:
# - `dim`: The number of features in X.
# - `dim_outcome`: The number of potential outcomes.
# - `dim_hidden`: hyperparameter for tuning the size of the hidden layer.
# - `depth`: hyperparameter for the number of hidden layers in the generator and inference blocks.
# - `num_iterations`: hyperparameter for the number of training epochs.
# - `alpha`: hyperparameter used for the Generator block loss.
# - `beta`: hyperparameter used for the ITE block loss.
# - `num_discr_iterations`: number of iterations executed by the discriminator.
# - `minibatch_size`: the size of the dataset batches.
#
# The hyperparameters used in this notebook are computed using the [hyperparameter tuning notebook](https://github.com/bcebere/ite-api/blob/main/notebooks/hyperparam_tuning.ipynb).
# +
dim = len(Train_X[0])
dim_outcome = Test_Y.shape[1]
model = alg.Ganite(
dim,
dim_outcome,
dim_hidden=7,
num_iterations=10000,
alpha=5,
beta=0.1,
minibatch_size=32,
num_discr_iterations=9,
depth=2,
)
assert model is not None
# -
# ## Train the model
metrics = model.train(*dataloader)
# ## Plot train metrics
# +
metrics.plot(plt, thresholds = [0.2, 0.25, 0.3, 0.35])
metrics.print()
# -
# ## Predict
#
# You can use run inferences on the model and evaluate the output.
# +
sess = tf.InteractiveSession()
hat_y = model.predict(Test_X)
utils.sqrt_PEHE(hat_y, Test_Y).eval()
# -
# ## Test
# Will can run inferences and get metrics directly
# +
test_metrics = model.test(Test_X, Test_Y)
test_metrics.print()
# -
# ## References
#
# 1. <NAME>, <NAME>, <NAME>, "GANITE: Estimation of Individualized Treatment Effects using Generative Adversarial Nets", International Conference on Learning Representations (ICLR), 2018 ([Paper](https://openreview.net/forum?id=ByKWUeWA-)).
# 2. [GANITE Reference implementation](https://bitbucket.org/mvdschaar/mlforhealthlabpub/src/master/alg/ganite/).
| notebooks/ganite_train_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 4, "hidden": false, "row": 0, "width": 4}, "report_default": {"hidden": false}}}}
# # Example User Analysis
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
import numpy as np
import pandas as pd
import plotly
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import cufflinks as cf
import plotly.figure_factory as ff
import plotly.tools as tls
name='LA'
plotly.tools.set_credentials_file(username='xxxxxxxx', api_key='<KEY>')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
gradebook = pd.read_csv('Data\gradebook.csv', low_memory=False)
users = pd.read_csv('Data\\users.csv')
users = users.set_index('imperial_user_id')
country_codes = pd.read_csv('../../../Data/ISO_country_codes.csv')
country_codes = country_codes.set_index('alpha2')
## this sets the index of the country_codes table to the two letter code, making it easy to join with the 'country_cd' column in df
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 58, "hidden": false, "row": 0, "width": 4}, "report_default": {"hidden": false}}}}
### Join the user table to the gradebook so that we can link attainment at each stage of the module to demographics
df = gradebook.join(users, 'Anonymized Coursera ID (imperial_user_id)', how = 'left')
### Strip out unnecessary info
df.columns = df.columns.str.replace('Assessment Grade: ','')
cols = [c for c in df.columns if c.lower()[:10] != 'submission']
df = df[cols]
### Pull in additional country information from ISO data so that we have full country name and alpha3 code which is required for the worldview Plotly map
df = df.join(country_codes,'country_cd', how = 'left')
total = df['Anonymized Coursera ID (imperial_user_id)'].nunique()
print(total)
### simply for easy viewing of fields:
pretty = df.iloc[1].transpose()
pretty
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
def countries (df, column):
df1 = df.groupby('alpha3').count()
data = [dict(
type = 'choropleth',
locations = df1.index.values,
z = df1[column],
text = df1[column]/total*100,
colorscale = [[0,"rgb(5, 10, 172)"],[0.35,"rgb(40, 60, 190)"],[0.5,"rgb(70, 100, 245)"],\
[0.6,"rgb(90, 120, 245)"],[0.7,"rgb(106, 137, 247)"],[1,"rgb(220, 220, 220)"]],
autocolorscale = False,
reversescale = True,
marker = dict(
line = dict (
color = 'rgb(180,180,180)',
width = 0.5
) ),
colorbar = dict(
autotick = False),
) ]
layout = dict(
geo = dict(
showframe = False,
showcoastlines = False,
projection = dict(
type = 'Mercator'
)
)
)
fig = dict( data=data, layout=layout )
plot = py.iplot( fig, validate=False, filename=name + 'learner-world-map2' )
return plot
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
def countries_ratio (df, column1, column2):
df2 = df.groupby('alpha3').count()
data = [dict(
type = 'choropleth',
locations = df2.index.values,
z = df2[column1]/df2[column2]*100,
colorscale = [[0,"rgb(5, 10, 172)"],[0.35,"rgb(40, 60, 190)"],[0.5,"rgb(70, 100, 245)"],\
[0.6,"rgb(90, 120, 245)"],[0.7,"rgb(106, 137, 247)"],[1,"rgb(220, 220, 220)"]],
autocolorscale = False,
reversescale = True,
marker = dict(
line = dict (
color = 'rgb(180,180,180)',
width = 0.5
) ),
colorbar = dict(
autotick = False),
) ]
layout = dict(
geo = dict(
showframe = False,
showcoastlines = False,
projection = dict(
type = 'Mercator'
)
)
)
fig = dict( data=data, layout=layout )
plot = py.iplot( fig, validate=False, filename=name + 'learner-world-map2' )
return plot
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
def progress(df, column,x):
df3 = df.drop(columns = ['Course Grade', 'Course Passed', 'Completed with CC'])
df3 = df3.groupby(column).count()
df3 = df3[df3['Anonymized Coursera ID (imperial_user_id)'] > total*(x/100)]
# print (df3.iloc[0])
progress = df3.transpose()
progress = progress[:-11]
breakdown = progress.iloc[0]
print (breakdown)
# plot1 = breakdown.iplot(kind='bar', sharing='private')
progress = (progress/progress.iloc[0]) * 100
plot = progress.iplot(kind='line', sharing='private')
return plot
# -
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
def learningCurve(df, column, x):
# df3 = df.groupby(column).count()
# df3 = df3[df3['Anonymized Coursera ID (imperial_user_id)'] > total*(x/100)]
# df3 = df3.iloc[0]
df = df.drop(columns = ['Course Grade', 'Course Passed', 'Completed with CC'])
df = df.groupby(column).mean()
progress = df.transpose()
progress = progress[:-1]
breakdown = progress.iloc[1]
print (breakdown)
# plot1 = breakdown.iplot(kind='bar', sharing='private')
plot = progress.iplot(kind='line', sharing='private')
return plot
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 19, "hidden": false, "row": 0, "width": 4}, "report_default": {"hidden": false}}}}
countries (df, 'Anonymized Coursera ID (imperial_user_id)')
# -
countries_ratio (df, 'Eigenvalues and eigenvectors','Anonymized Coursera ID (imperial_user_id)')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 25, "hidden": false, "row": 4, "width": 4}, "report_default": {"hidden": false}}}}
progress(df,'browser_language_cd', 1)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 25, "hidden": false, "row": 19, "width": 4}, "report_default": {"hidden": false}}}}
learningCurve(df,'educational_attainment',1)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 22, "hidden": false, "row": 29, "width": 4}, "report_default": {"hidden": false}}}}
progress(df,'reported_or_inferred_gender')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 22, "hidden": false, "row": 44, "width": 4}, "report_default": {"hidden": false}}}}
learningCurve(df,'reported_or_inferred_gender')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 26, "hidden": false, "row": 51, "width": 4}, "report_default": {"hidden": false}}}}
progress(df,'browser_language_cd',1)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 4, "height": 26, "hidden": false, "row": 58, "width": 4}, "report_default": {"hidden": false}}}}
learningCurve(df,'browser_language_cd')
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# -
| Example_User_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import sklearn
import pandas as pd
import os
import sys
import time
import tensorflow as tf
from tensorflow import keras
print(tf.__version__)
print(sys.version_info)
for module in mpl, np, pd, sklearn, tf, keras:
print(module.__name__, module.__version__)
# +
# https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt
input_filepath = "./shakespeare.txt"
text = open(input_filepath, 'r').read()
print(len(text))
print(text[0:100])
# +
# 1. generate vocab
# 2. build mapping char->id
# 3. data -> id_data
# 4. abcd -> bcd<eos>
vocab = sorted(set(text))
print(len(vocab))
print(vocab)
# -
char2idx = {char:idx for idx, char in enumerate(vocab)}
print(char2idx)
idx2char = np.array(vocab)
print(idx2char)
text_as_int = np.array([char2idx[c] for c in text])
print(text_as_int[0:10])
print(text[0:10])
# +
def split_input_target(id_text):
"""
abcde -> abcd, bcde
"""
return id_text[0:-1], id_text[1:]
char_dataset = tf.data.Dataset.from_tensor_slices(text_as_int)
seq_length = 100
seq_dataset = char_dataset.batch(seq_length + 1,
drop_remainder = True)
for ch_id in char_dataset.take(2):
print(ch_id, idx2char[ch_id.numpy()])
for seq_id in seq_dataset.take(2):
print(seq_id)
print(repr(''.join(idx2char[seq_id.numpy()])))
# +
seq_dataset = seq_dataset.map(split_input_target)
for item_input, item_output in seq_dataset.take(2):
print(item_input.numpy())
print(item_output.numpy())
# +
batch_size = 64
buffer_size = 10000
seq_dataset = seq_dataset.shuffle(buffer_size).batch(
batch_size, drop_remainder=True)
# +
vocab_size = len(vocab)
embedding_dim = 256
rnn_units = 1024
def build_model(vocab_size, embedding_dim, rnn_units, batch_size):
model = keras.models.Sequential([
keras.layers.Embedding(vocab_size, embedding_dim,
batch_input_shape = [batch_size, None]),
keras.layers.LSTM(units = rnn_units,
stateful = True,
recurrent_initializer = 'glorot_uniform',
return_sequences = True),
keras.layers.Dense(vocab_size),
])
return model
model = build_model(
vocab_size = vocab_size,
embedding_dim = embedding_dim,
rnn_units = rnn_units,
batch_size = batch_size)
model.summary()
# -
for input_example_batch, target_example_batch in seq_dataset.take(1):
example_batch_predictions = model(input_example_batch)
print(example_batch_predictions.shape)
# random sampling.
# greedy, random.
sample_indices = tf.random.categorical(
logits = example_batch_predictions[0], num_samples = 1)
print(sample_indices)
# (100, 65) -> (100, 1)
sample_indices = tf.squeeze(sample_indices, axis = -1)
print(sample_indices)
print("Input: ", repr("".join(idx2char[input_example_batch[0]])))
print()
print("Output: ", repr("".join(idx2char[target_example_batch[0]])))
print()
print("Predictions: ", repr("".join(idx2char[sample_indices])))
# +
def loss(labels, logits):
return keras.losses.sparse_categorical_crossentropy(
labels, logits, from_logits=True)
model.compile(optimizer = 'adam', loss = loss)
example_loss = loss(target_example_batch, example_batch_predictions)
print(example_loss.shape)
print(example_loss.numpy().mean())
# +
output_dir = "./text_generation_lstm3_checkpoints"
if not os.path.exists(output_dir):
os.mkdir(output_dir)
checkpoint_prefix = os.path.join(output_dir, 'ckpt_{epoch}')
checkpoint_callback = keras.callbacks.ModelCheckpoint(
filepath = checkpoint_prefix,
save_weights_only = True)
epochs = 100
history = model.fit(seq_dataset, epochs = epochs,
callbacks = [checkpoint_callback])
# -
tf.train.latest_checkpoint(output_dir)
model2 = build_model(vocab_size,
embedding_dim,
rnn_units,
batch_size = 1)
model2.load_weights(tf.train.latest_checkpoint(output_dir))
model2.build(tf.TensorShape([1, None]))
# start ch sequence A,
# A -> model -> b
# A.append(b) -> B
# B(Ab) -> model -> c
# B.append(c) -> C
# C(Abc) -> model -> ...
model2.summary()
# +
def generate_text(model, start_string, num_generate = 1000):
input_eval = [char2idx[ch] for ch in start_string]
input_eval = tf.expand_dims(input_eval, 0)
text_generated = []
model.reset_states()
# temperature > 1, random
# temperature < 1, greedy
temperature = 2
for _ in range(num_generate):
# 1. model inference -> predictions
# 2. sample -> ch -> text_generated.
# 3. update input_eval
# predictions : [batch_size, input_eval_len, vocab_size]
predictions = model(input_eval)
# predictions: logits -> softmax -> prob
# softmax: e^xi
# eg: 4,2 e^4/(e^4 + e^2) = 0.88, e^2 / (e^4 + e^2) = 0.12
# eg: 2,1 e^2/(e^2 + e) = 0.73, e / (e^2 + e) = 0.27
predictions = predictions / temperature
# predictions : [input_eval_len, vocab_size]
predictions = tf.squeeze(predictions, 0)
# predicted_ids: [input_eval_len, 1]
# a b c -> b c d
predicted_id = tf.random.categorical(
predictions, num_samples = 1)[-1, 0].numpy()
text_generated.append(idx2char[predicted_id])
# s, x -> rnn -> s', y
input_eval = tf.expand_dims([predicted_id], 0)
return start_string + ''.join(text_generated)
new_text = generate_text(model2, "All: ")
print(new_text)
# -
| JupyterNotebookCode/text_generation_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activation Functions
import numpy as np
def sigmoid(X):
return 1/(1+np.exp(-X))
def relu(X):
return np.maximum(0,X)
def softmax(X):
expo = np.exp(X)
expo_sum = np.sum(np.exp(X))
return expo/expo_sum
def tanh(x):
return np.tanh(x)
def lrelu(X, alpha):
return np.where(X > 0, X, X * alpha)
def heaviside(X, t):
return np.where((X-t) != 0, np.maximum(0, np.sign(X-t)), 0.5)
inp = np.array([
[1, 0.5, 0.2],
[-1, -0.5, -0.2],
[0.1, -0.1, 0]
])
sigmoid(inp)
relu(inp)
softmax(inp)
tanh(inp)
lrelu(inp, 0.1)
heaviside(inp, 0.1)
| .ipynb_checkpoints/0-activation_functions-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from reflex.utils import load_file
import glob
import os
import json
# +
dataset_path = '/Users/ankur/Projects/RE-Flex/data/Google_RE'
new_directory = '/Users/ankur/Projects/RE-Flex/data/Google_RE2'
for f in glob.glob(os.path.join(dataset_path, '*')):
results = []
bname = os.path.basename(f)
data = load_file(f)
for d in data:
judgements = d['judgments']
y = 0
n = 0
for j in judgements:
if j['judgment'] == 'yes':
y += 1
else:
n += 1
# We only consider examples that have a majority vote yes judgement
if y <= n:
continue
head = d['sub_label']
tail = d['obj_label']
context = None
# We take the first masked sentence that has a mask in it
for ms in d['masked_sentences']:
if '[MASK]' in ms:
context = ms.replace('[MASK]', tail)
break
if context is None:
continue
results.append({'subject': head, 'context': context, 'object': tail})
with open(os.path.join(new_directory, bname), 'w') as wf:
for r in results:
wf.write(f'{json.dumps(r)}\n')
# -
| preprocess_notebooks/preprocess_googlere.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Importing all Required Libraries
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows',50)
from sklearn.preprocessing import LabelEncoder,StandardScaler
from sklearn.decomposition import PCA
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.style as style # for styling the graphss
# style.available (to know the available list of styles)
style.use('ggplot') # chosen style
plt.rc('xtick',labelsize=13) # to globally set the tick size
plt.rc('ytick',labelsize=13) # to globally set the tick size
# Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# To display float with 2 decimal, avoid scientific printing
pd.options.display.float_format = '{:.2f}'.format
import seaborn as sns
import warnings
import math
#Library for one hot encoding
from sklearn.preprocessing import OneHotEncoder
#train test split
from sklearn.model_selection import train_test_split
#Evaluation metrics
from sklearn.metrics import accuracy_score,mean_squared_error,r2_score
#load the data
df=pd.read_csv("./Data/cleaned_merged_all_data.csv")
df.head()
columns_to_be_removed = df.isnull().sum()[df.isnull().sum().sort_values() > 197285].index
df = df.drop(columns=columns_to_be_removed)
df.shape
df.dropna(inplace=True)
df.shape
import datetime as dt
df["Invoice Date"]=pd.to_datetime(df["Invoice Date"],dayfirst=True)
df['years']=df['Invoice Date'].dt.year
# +
#For Customer Life time value prediction we need data only of 2015 hence Subset the data
cltv_df=df[['Customer No.','Cust Type','Invoice No','Make','Model','Total Amt Wtd Tax.','divisionname','years']]
#Subsetting the data
#cltv_df=cltv_df[cltv_df['years']==2015]
cltv_df
# -
#Groupby of data
cltv_group=cltv_df.groupby(['Customer No.','Cust Type','divisionname','Make','Model']).agg({'Invoice No':'nunique','Total Amt Wtd Tax.':'mean'}).reset_index()
#cltv_group.drop('year',1,inplace=True)
cltv_group
#finding customer value for year 2015
cltv_group['customer_value']=cltv_group['Invoice No']*cltv_group['Total Amt Wtd Tax.']
cltv_group
#Removing Customers with zero value
zero_group=cltv_group[cltv_group['customer_value']<=0].index
cltv_group.drop(zero_group,0,inplace=True)
cltv_group
#Creating Dataframe for Modelling
model_df=cltv_group[['Customer No.','Cust Type','divisionname','Make','Invoice No','Total Amt Wtd Tax.','customer_value']]
model_df
# +
#Renaming Columns
model_df.columns=['Cust_no.','Cust','State','Make','Count_invoice','Avg_revenue','customer_value']
model_df
# -
#Final dummyencoded dataframe
A=pd.get_dummies(data=model_df, columns=['Cust', 'State','Make'])
final_df=pd.DataFrame(A)
final_df.drop('Cust_no.',1,inplace=True)
final_df
#Train Test Split
X=final_df.drop('customer_value',1)
y=final_df['customer_value']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=9)
# +
#Implementing Decison Tree Regressor
from sklearn.tree import DecisionTreeRegressor
# +
max_depth_range = list(range(1,12))
# List to store the average RMSE for each value of max_depth:
accuracy = []
for depth in max_depth_range:
reg = DecisionTreeRegressor(max_depth = depth, random_state = 18)
reg.fit(X_train, y_train)
score = reg.score(X_test, y_test)
accuracy.append(score)
x=max_depth_range
y=accuracy
plt.figure(figsize=(15,10))
sns.pointplot(x,y)
plt.xlabel('max_depth')
plt.ylabel('accuracy')
plt.title('accuracy vs max_depth')
# +
X=final_df.drop('customer_value',1)
y=final_df['customer_value']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=18)
dt_reg = DecisionTreeRegressor(max_depth=6, random_state = 18)
dt_reg.fit(X_train, y_train)
y_pred=dt_reg.predict(X_test)
#Evaluation metric
mse =mean_squared_error(y_test,y_pred)
print('mse score:',mse)
print('=='*100)
rmse=mean_squared_error(y_test,y_pred)
rmse=np.sqrt(rmse)
print('Rmse score:',rmse)
#accuracy score on train data
train_score=dt_reg.score(X_train,y_train)
print('train score:',train_score)
print('=='*100)
#accuracy score on test data
test_score=dt_reg.score(X_test,y_test)
print('test score:',test_score)
print('=='*100)
# -
#Evaluation metric R2 score
R2_Score=r2_score(y_test,y_pred)
print('R2 score:',R2_Score)
print('=='*100)
# +
#Repeated k fold and cross val score
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import cross_val_score
rkf=RepeatedKFold(n_splits=5, n_repeats=5, random_state=9)
#Crossvalidation process
scores = cross_val_score(dt_reg, X, y, cv=rkf, scoring='r2')
print('scores',scores)
print('=='*100)
print('Mean_score:',scores.mean())
print('=='*100)
print('std_score:',scores.std())
# -
from sklearn.metrics import mean_squared_log_error
#For test
mean_squared_log_error(dt_reg.predict(X_test),y_test)
mean_squared_log_error(dt_reg.predict(X_train),y_train)
| LTV_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf2
# language: python
# name: tf2
# ---
# # Prepare data
# +
import requests
import category_encoders as ce
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
def encode_var(var, encoder, y=None):
if y is None:
encoder.fit(var)
else:
encoder.fit(var, y)
new_var = encoder.transform(var)
if isinstance(new_var, pd.DataFrame):
new_var.insert(0, 'original', var)
return new_var
else:
return pd.DataFrame({'original': var, 'encoder': new_var})
def print_res(res, rows_per_level=2):
out = pd.DataFrame(columns=res.columns)
for lvl in res.original.unique():
out = out.append(res[res.original==lvl].head(rows_per_level))
return out
# -
download_data = False
if download_data:
url = "http://mlr.cs.umass.edu/ml/machine-learning-databases/autos/imports-85.data"
r = requests.get(url)
with open('imports-85.data', 'wb') as f:
f.write(r.content)
# +
# Define the headers since the data does not have any
headers = ["symboling", "normalized_losses", "make", "fuel_type", "aspiration",
"num_doors", "body_style", "drive_wheels", "engine_location",
"wheel_base", "length", "width", "height", "curb_weight",
"engine_type", "num_cylinders", "engine_size", "fuel_system",
"bore", "stroke", "compression_ratio", "horsepower", "peak_rpm",
"city_mpg", "highway_mpg", "price"]
# Read in the CSV file and convert "?" to NaN
df = pd.read_csv("imports-85.data",
header=None, names=headers, na_values="?" )
df = df[df.price.notnull()]
df["num_cylinders"] = df["num_cylinders"].astype('category').cat.reorder_categories(ordered=True, new_categories=['two', 'three', 'four', 'five', 'six', 'eight', 'twelve'])
# -
# # **Exercise 1**
# Calculate the regressions of 'num_cylinders' on price for each possible encoding. What is the correct interpretation for the coefficients?
from sklearn.linear_model import LinearRegression
X_raw = df[['num_cylinders']]
y = df['price']
# ## Ordinal encoder
encoder = ce.OrdinalEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression()
model.fit(X,y)
"For each increase in the level of the number of cylinders, the price changes by {:6.2f} on average".format(model.coef_[0])
# ## One-Hot Encoder
encoder = ce.OneHotEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
model.intercept_
df[['num_cylinders', 'price']].groupby('num_cylinders').mean()
"The price of a car with six cylinders is, on average, {:6.2f}.".format(model.coef_[4])
# ## Dummy encoding
X = X.iloc[:,1:]
model = LinearRegression()
model.fit(X,y)
model.coef_
model.intercept_
"The price of a car with six cylinders is, on average, {:6.2f} higher than for cars with two cylinders".format(model.coef_[3])
# ## Binary Encoder
encoder = ce.BinaryEncoder()
X = encoder.fit_transform(X_raw)
model.fit(X,y)
model.coef_
X.head()
# No meaning of coefficients. Binary encoding is just a way to express non-numeric values in numbers.
# ## Base-N Encoding
encoder = ce.BaseNEncoder(base=4)
X = encoder.fit_transform(X_raw)
model.fit(X,y)
model.coef_
X.head()
# No meaning of coefficients. Base-N encoding is just a way to express non-numeric values in numbers.
# ## Simple Encoder
from simple_coding import SimpleEncoder
encoder = SimpleEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
model.intercept_
X.head()
means = df[['num_cylinders', 'price']].groupby('num_cylinders').mean()
means.mean()
"The price of a car with three cylinders is, on average, {:6.2f} lower than the price of cars with two cylinders".format(model.coef_[1])
# ## Sum Encoder
encoder = ce.SumEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
model.intercept_
X.head()
means - means.mean()
"Each coefficient represents the difference between the group average to the grand average of the price"
"Cars with two cylinders are {:6.2f} less expensive than the average price of the groups.".format(model.coef_[1])
"The grand mean of {:6.2f} is different than the sample mean of {:6.2f}".format(means.mean()[0], y.mean())
# ## Polynomical Encoder
encoder = ce.PolynomialEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
# The coefficients represent the linear, quadradic, cubic, etc. trends in the data.
X.head()
# ## Helmert Encoding
encoder = ce.HelmertEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
# The coefficients represent the change in the grand mean for levels up to k-1 to the grand mean for levels up to k.
means.iloc[:2].mean() - means.iloc[:1].mean()
"The grand mean price for cars with cylinders up to three is {:6.2f} lower than for cars with cylinders up to two".format(model.coef_[1])
means.iloc[:3].mean() - means.iloc[:2].mean()
"The grand mean price for cars with cylinders up to four is {:6.2f} lower than for cars with cylinders up to three".format(model.coef_[2])
# ## Backward Difference Encoder
encoder = ce.BackwardDifferenceEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
means.diff()
# The coefficients are the differences in average value between adjacent levels.
"Cars with three cylinders are, on average, {:6.2f} less expensive than cars with two cylinders".format(model.coef_[1])
"Cars with four cylinders are, on average, {:6.2f} less expensive than cars with three cylinders".format(model.coef_[2])
# ## Count Encoder
encoder = ce.CountEncoder(normalize=True)
#X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
#model.fit(X,y)
# But in implementation of CountEncoder with variable of type categorical
# ## Hashing Encoder
encoder = ce.HashingEncoder()
X = encoder.fit_transform(X_raw)
model = LinearRegression(fit_intercept=False)
model.fit(X,y)
model.coef_
X.head()
# No meaning to coefficients. Hashing is just a way to represent non-numeric data as numbers.
# # Exercise 2
# Try to find the best encoding for each variable to maximize the generalization performance of a linear regression model to predict the price of a car.
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.model_selection import GridSearchCV
from category_encoders import OrdinalEncoder, OneHotEncoder, BinaryEncoder, SumEncoder, PolynomialEncoder, HelmertEncoder, BackwardDifferenceEncoder, HashingEncoder
from category_encoders import TargetEncoder, JamesSteinEncoder, MEstimateEncoder, LeaveOneOutEncoder, CatBoostEncoder
y = df['price']
X = df.drop('price', axis=1)
cols_num = X.select_dtypes(include=[int, float]).columns.values
cols_cat = X.select_dtypes(exclude=[int, float]).columns.values
num_pipeline = make_pipeline(SimpleImputer(strategy='mean'), StandardScaler())
encoders = []
for col in cols_cat:
encoders.append((col, OneHotEncoder(), [col]))
cat_pipeline = ColumnTransformer(encoders)
full_pipeline = make_pipeline(ColumnTransformer([('num', num_pipeline, cols_num), ('cat', cat_pipeline, cols_cat)]), LinearRegression())
param_grid = {'columntransformer__cat__make': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__fuel_type': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__aspiration': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__num_doors': [OrdinalEncoder(), OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__body_style': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__drive_wheels': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__engine_location': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__engine_type': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__num_cylinders': [OrdinalEncoder(), OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()],
'columntransformer__cat__fuel_system': [OneHotEncoder(), BinaryEncoder(), SumEncoder(), PolynomialEncoder(), HelmertEncoder(), BackwardDifferenceEncoder(), HashingEncoder()]}
grid = GridSearchCV(full_pipeline, param_grid, cv=3, n_jobs=11)
# +
# grid.fit(X,y)
# -
# Full search fails, due to memory restrictions. Switching to greedy.
steps = []
temp_grid = {}
for col in param_grid.keys():
param_grid[col]
steps = []
temp_grid = {}
for col in param_grid.keys():
steps.append(col)
temp_grid[col] = param_grid[col]
grid = GridSearchCV(full_pipeline, temp_grid, cv=3, n_jobs=11)
grid = grid.fit(X,y)
temp_grid[col] = [grid.best_params_[col]]
grid.best_score_
y_pred = grid.predict(X)
from sklearn.metrics import r2_score
r2_score(y, y_pred)
from sklearn.model_selection import cross_val_predict
y_pred_cv = cross_val_predict(grid, X, y)
r2_score(y, y_pred_cv)
| exercises/06_encoders_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# ## Dependencies and starter code
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
from scipy.stats import linregress
# Study data files
mouse_metadata = "data/Mouse_metadata.csv"
study_results = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata)
study_results = pd.read_csv(study_results)
# Combine the data into a single dataset
combined_df= pd.merge(mouse_metadata, study_results, how='outer', on='Mouse ID')
combined_df.head()
# -
# ## Summary statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
#create the groupby to sort by drug regimen
drug_names_gb=combined_df.groupby("Drug Regimen")
#generating the summary statics table consiting of
#consisting of the mean, median, variance, standard deviation,
#and SEM of the tumor volume for each drug regimen.
mean_drug_regimen=drug_names_gb["Tumor Volume (mm3)"].mean()
median_drug_regimen=drug_names_gb["Tumor Volume (mm3)"].median()
var_drug_regimen=drug_names_gb["Tumor Volume (mm3)"].var()
std_drug_regimen=drug_names_gb["Tumor Volume (mm3)"].std()
sem_drug_regimen=drug_names_gb["Tumor Volume (mm3)"].sem()
#creating the table
summary_statistics_table=pd.DataFrame({"Drug Regimen Names":drug_names_gb.count().index})
summary_statistics_table=summary_statistics_table.set_index(["Drug Regimen Names"])
summary_statistics_table["Mean"]=mean_drug_regimen
summary_statistics_table["Median"]=median_drug_regimen
summary_statistics_table["Variance"]=var_drug_regimen
summary_statistics_table["STD"]=std_drug_regimen
summary_statistics_table["SEM"]=sem_drug_regimen
#showing the table
summary_statistics_table
# -
# ## Bar plots
# +
# Generate a bar plot showing number of data points for each treatment regimen using pandas
df_2=pd.DataFrame({"Drug Regimen":drug_names_gb.count().index})
df_2=df_2.set_index(["Drug Regimen"])
df_2["Count"]=drug_names_gb["Drug Regimen"].count()
df_2.plot(kind="bar")
# Set a title for the chart
plt.title("Number of Drug Regimen")
plt.ylabel("Count")
plt.show()
# -
# Generate a bar plot showing number of data points for each treatment regimen using pyplot
x_axis = np.arange(len(df_2))
tick_locations = [value for value in x_axis]
plt.bar(x_axis,df_2["Count"])
plt.xticks(tick_locations,drug_names_gb.count().index,rotation=45)
plt.title("Number of Drug Regimen")
plt.ylabel("Count")
plt.show()
# ## Pie plots
# Generate a pie plot showing the distribution of female versus male mice using pandas
gender_group=combined_df.groupby("Sex")
df_3=pd.DataFrame({"Gender":gender_group.count().index})
df_3.set_index(["Gender"],inplace=True)
df_3["Count"]=gender_group["Sex"].count()
df_3.plot(kind="pie",subplots=True,)
plt.show()
# Generate a pie plot showing the distribution of female versus male mice using pyplot
labels=gender_group.count().index
sizes=df_3["Count"]
plt.pie(sizes, labels=labels,
autopct="%1.1f%%", shadow=True, startangle=140)
plt.show()
# ## Quartiles, outliers and boxplots
# +
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens. Calculate the IQR and quantitatively determine if there are any potential outliers.
df_4=combined_df.loc[((combined_df["Drug Regimen"]=="Capomulin")|(combined_df["Drug Regimen"]=="Ramicane")|(combined_df["Drug Regimen"]=="Infubinol")|(combined_df["Drug Regimen"]=="Ceftamin"))
,["Mouse ID","Drug Regimen","Timepoint","Tumor Volume (mm3)"]]
Capomulin_data=df_4.loc[df_4["Drug Regimen"]=="Capomulin",["Mouse ID","Drug Regimen","Timepoint","Tumor Volume (mm3)"]]
Capomulin_data.reset_index(drop=True,inplace=True)
Infubinol_data=df_4.loc[df_4["Drug Regimen"]=="Infubinol",["Mouse ID","Drug Regimen","Timepoint","Tumor Volume (mm3)"]]
Infubinol_data.reset_index(drop=True,inplace=True)
Ceftamin_data=df_4.loc[df_4["Drug Regimen"]=="Ceftamin",["Mouse ID","Drug Regimen","Timepoint","Tumor Volume (mm3)"]]
Ceftamin_data.reset_index(drop=True,inplace=True)
Ramicane_data=df_4.loc[df_4["Drug Regimen"]=="Ramicane",["Mouse ID","Drug Regimen","Timepoint","Tumor Volume (mm3)"]]
Ramicane_data.reset_index(drop=True,inplace=True)
Capomulin_groupby=Capomulin_data.groupby("Mouse ID")
Capomulin_data_final=Capomulin_groupby["Tumor Volume (mm3)"].min()
quartiles = Capomulin_data_final.quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
print(f"The lower quartile of Capomulin is: {lowerq}")
print(f"The upper quartile of Capomulin is: {upperq}")
print(f"The interquartile range of Capomulin is: {iqr}")
print(f"The the median of Capomulin is: {quartiles[0.5]} ")
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
print(f"Values below {lower_bound} could be outliers.")
print(f"Values above {upper_bound} could be outliers.")
# +
Infubinol_groupby=Infubinol_data.groupby("Mouse ID")
Infubinol_data_final=Infubinol_groupby["Tumor Volume (mm3)"].max()
quartiles = Infubinol_data_final.quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
print(f"The lower quartile of temperatures is: {lowerq}")
print(f"The upper quartile of temperatures is: {upperq}")
print(f"The interquartile range of temperatures is: {iqr}")
print(f"The the median of temperatures is: {quartiles[0.5]} ")
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
print(f"Values below {lower_bound} could be outliers.")
print(f"Values above {upper_bound} could be outliers.")
# +
Ceftamin_groupby=Ceftamin_data.groupby("Mouse ID")
Ceftamin_data_final=Ceftamin_groupby["Tumor Volume (mm3)"].max()
quartiles = Ceftamin_data_final.quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
print(f"The lower quartile of temperatures is: {lowerq}")
print(f"The upper quartile of temperatures is: {upperq}")
print(f"The interquartile range of temperatures is: {iqr}")
print(f"The the median of temperatures is: {quartiles[0.5]} ")
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
print(f"Values below {lower_bound} could be outliers.")
print(f"Values above {upper_bound} could be outliers.")
# +
Ramicane_groupby=Ramicane_data.groupby("Mouse ID")
Ramicane_data_final=Ramicane_groupby["Tumor Volume (mm3)"].min()
quartiles = Ramicane_data_final.quantile([.25,.5,.75])
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
iqr = upperq-lowerq
print(f"The lower quartile of temperatures is: {lowerq}")
print(f"The upper quartile of temperatures is: {upperq}")
print(f"The interquartile range of temperatures is: {iqr}")
print(f"The the median of temperatures is: {quartiles[0.5]} ")
lower_bound = lowerq - (1.5*iqr)
upper_bound = upperq + (1.5*iqr)
print(f"Values below {lower_bound} could be outliers.")
print(f"Values above {upper_bound} could be outliers.")
# -
# Generate a box plot of the final tumor volume of each mouse across four regimens of interestRamicane_data
boxplot_data=[Capomulin_data_final,Infubinol_data_final,Ceftamin_data_final,Ramicane_data_final]
names=["Capomulin","Infubinol","Ceftamin","Ramicane"]
fig1, ax1 = plt.subplots()
ax1.set_title('')
ax1.set_ylabel('')
ax1.boxplot(boxplot_data)
plt.xticks([1,2,3,4],names)
ax1.set_title('Final tumor volume for all four treatment regimens')
ax1.set_ylabel("Tumor Volume (mm3")
plt.show()
# ## Line and scatter plots
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
df_5=combined_df.loc[((combined_df["Drug Regimen"]=="Capomulin")|(combined_df["Drug Regimen"]=="Ramicane")|(combined_df["Drug Regimen"]=="Infubinol")|(combined_df["Drug Regimen"]=="Ceftamin"))
,["Drug Regimen","Weight (g)","Timepoint","Tumor Volume (mm3)"]]
df_5.reset_index(drop=True,inplace=True)
Capomulin_data2=df_5.loc[df_5["Drug Regimen"]=="Capomulin",["Drug Regimen","Weight (g)","Timepoint","Tumor Volume (mm3)"]]
Capomulin_data2.reset_index(drop=True,inplace=True)
Capomulin_data2=Capomulin_data2.iloc[0:10]
Capomulin_data2.plot('Timepoint','Tumor Volume (mm3)',kind='line')
plt.title("Time point versus tumor volume")
plt.xlabel("Time Point (seconds)")
plt.ylabel('Tumor Volume (mm3)')
plt.show()
# +
# Generate a scatter plot of mouse weight versus average tumor volume for the Capomulin regimen
Capomulin_data3=df_5.loc[df_5["Drug Regimen"]=="Capomulin",["Drug Regimen","Weight (g)","Timepoint","Tumor Volume (mm3)"]]
Capomulin_data3.reset_index(drop=True,inplace=True)
gb2=Capomulin_data3.groupby("Weight (g)")
avg_tumor=gb2['Tumor Volume (mm3)'].mean()
weights=gb2["Weight (g)"].mean()
plt.scatter(weights,avg_tumor)
plt.title("Mouse weight versus average Tumor Volume")
plt.xlabel("Weight (g)")
plt.ylabel('Average Tumor Volume')
plt.show()
# -
# Calculate the correlation coefficient
correlation = st.pearsonr(weights,avg_tumor)
print(f"The correlation between both factors is {round(correlation[0],2)}")
# +
#linear regression model for mouse weight and average tumor volume for the Capomulin regimen
#variables that will be graphed
x_values = weights
y_values = avg_tumor
#code for the linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.title("Mouse weight versus average Tumor Volume")
plt.xlabel("Weight (g)")
plt.ylabel('Average Tumor Volume')
plt.show()
print("The linear Regression equation for the scatter plot is : " +str(line_eq))
# -
| pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Graph:
def __init__(self, nVertices):
self.nVertices = nVertices
self.adjMatrix = [[0 for i in range(nVertices)] for j in range(nVertices)]
def addEdge(self, v1, v2):
self.adjMatrix[v1][v2] = 1
self.adjMatrix[v2][v1] = 1
def removeEdge(self, v1, v2):
if self.containEdge(v1, v2) is False:
return
self.adjMatrix[v1][v2] = 0
self.adjMatrix[v2][v1] = 0
def containsEdge(self, v1, v2):
return True if self.adjMatrix[v1][v2] > 0 else False
def __str__(self):
return str(self.adjMatrix)
def __dfsHelper(self, sv, visited):
#print(sv, end = " ")
visited[sv] = True # Marking vertex as visited
for i in range(self.nVertices):
if self.adjMatrix[sv][i] > 0 and visited[i] is False:
self.__dfsHelper(i, visited)
return visited
def dfs(self, sv): # USing DFS for traversing all vertices of a graph
visited = [False for i in range(self.nVertices)]
return self.__dfsHelper(sv, visited)
v, e = [int (i) for i in input().split()[:2]]
g = Graph(v)
for i in range(e):
a, b = [int(x) for x in input().split()[:2]]
g.addEdge(a, b)
visited = g.dfs(0)
#print(visited)
flag = True
for j in visited:
if j is False:
flag = False
if flag:
print("true")
else:
print("false")
# +
g = Graph(7)
g.addEdge(0, 1)
g.addEdge(0, 2)
g.addEdge(0, 3)
g.addEdge(2, 4)
g.addEdge(4, 5)
g.addEdge(3, 6)
visited = g.dfs(0)
print(visited)
flag = True
for j in visited:
if j is False:
flag = False
if flag:
print("true")
else:
print("false")
| 19 Graphs - 1/19.12 Is Graph Connected.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="NqqLNJZjfi8U"
# 0. 학습 환경 설정하기
# 1. 데이터셋 불러오기
# 2. EDA
# 3. 모델 학습을 위한 데이터 전처리
# 4. 모델 학습하기
# 5. 모델 평가하기
# 6. 모델 학습 결과 심화 분석하기
#
# 출처 : 신제용 강사 (패스트캠퍼스)
#
# + [markdown] pycharm={"name": "#%% md\n"}
# ## 0 . 학습 환경 설정하기
# + id="g49RuFGrBvt7"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import os
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor
from sklearn.metrics import mean_absolute_error, mean_squared_error
from math import sqrt
# + id="mMKFOC0OBtHO"
# os.environ을 이용하여 Kaggle API Username, Key 세팅하기
os.environ['KAGGLE_USERNAME'] = 'fastcampuskim'
os.environ['KAGGLE_KEY'] = 'c939a1e37f5ca93b6406a66fc8bb08e5'
# -
# ## 1. 데이터셋 불러오기
# * 데이터 출처: https://www.kaggle.com/dgomonov/new-york-city-airbnb-open-data
# * 컬럼
# id: 항목의 ID
# name: 항목의 이름 (타이틀)
# host_id: 호스트 ID
# host_name: 호스트의 이름
# neighbourhood_group: 방이 있는 구역 그룹
# neighbourhood: 방이 있는 구역
# latitude: 방이 위치한 위도
# longitude: 방이 위치한 경도
# room_type: 방의 종류
# price: 가격 (미 달러)
# minimum_nights: 최소 숙박 일수
# number_of_reviews: 리뷰의 개수
# last_review: 마지막 리뷰 일자
# reviews_per_month: 월별 리뷰 개수
# calculated_host_listings_count: 호스트가 올린 방 개수
# availability_365: 365일 중 가능한 일수
# + id="JSblp2NsCGbh" colab={"base_uri": "https://localhost:8080/"} outputId="153ad19e-92fa-4d97-9a82-dffbcaa55024"
# Linux 명령어로 Kaggle API를 이용하여 데이터셋 다운로드하기 (!kaggle ~)
# Linux 명령어로 압축 해제하기
# !rm *.*
# !kaggle datasets download -d dgomonov/new-york-city-airbnb-open-data
# !unzip '*.zip'
# + id="RnJv-4YwCMSx"
df = pd.read_csv('AB_NYC_2019.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 864} id="x3U_iHpNfYOF" outputId="2cee770b-3f7d-439c-888c-4e4dc62891eb"
df
# + [markdown] id="9L3BNVM7tHN5"
# ## 2. EDA
#
# + id="YcR9BX23DIFW" colab={"base_uri": "https://localhost:8080/", "height": 411} outputId="a5f8ee35-6780-4c5a-bd5f-0e6987864109"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="_x9R5pzlniRF" outputId="123aeb0b-f9a0-4b7a-af43-9567392cda44"
df['room_type'].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="89i38feMmx_e" outputId="fb376c96-10d5-4013-b761-7e2f136102f9"
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="ub117b7Do1H4" outputId="1f93b1a4-5753-4033-9e7d-d56d0252001f"
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="vii-v4FIn2BK" outputId="7af781e4-a392-4d6f-f32e-cb7abd11c734"
(df['reviews_per_month'].isna() & df['last_review'].isna()).sum()
# + colab={"base_uri": "https://localhost:8080/"} id="yEPkZwtyoLUy" outputId="039dabe5-b1b9-4af2-f8ac-fd88d712aebf"
df['reviews_per_month'].isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="_1en1a5ppZck" outputId="74e8c71a-5f26-403c-ce25-7f763f572631"
(df['number_of_reviews'] == 0).sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="w7Vpx5iModag" outputId="03eb3c62-7899-4a98-9626-f0a703763947"
df['availability_365'].hist()
# + colab={"base_uri": "https://localhost:8080/"} id="6GyC17vdojIY" outputId="84942c57-ac42-426b-f1ea-ea8c58f92bd5"
(df['availability_365'] == 0).sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="WIoofJDVmzF1" outputId="2c93a36b-33e7-478d-8255-2e6e2786a86c"
df.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="lF-wjmG2pxpb" outputId="fa91fce1-63ff-4cb4-82f9-0d0573fea01b"
df.columns
# + id="VMtkp1CYpypz"
df.drop(['id', 'name', 'host_name', 'latitude', 'longitude'], axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="YwrtZAEIp7-5" outputId="f5076543-37ae-4819-ccee-3075bd86fcfb"
df.head()
# + id="b_zcrUDF7khF" colab={"base_uri": "https://localhost:8080/"} outputId="385e5ff6-0212-4a0e-d3f9-e412a990725a"
df.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 458} id="4qMUlI0XqMV4" outputId="69b10dca-060a-4612-d892-82140b0064cf"
sns.jointplot(x='host_id', y='price', data=df, kind='hex')
# + colab={"base_uri": "https://localhost:8080/", "height": 458} id="JihglMQnqny9" outputId="def7f70f-0dd4-4d06-f0ed-9574d2a07c3a"
sns.jointplot(x='reviews_per_month', y='price', data=df, kind='hex')
# + id="GZXSBFPyDh6R" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="81d330b6-013a-4195-cb09-e3adb7451406"
sns.heatmap(df.corr(), annot=True, cmap='YlOrRd')
# + id="MPmzv61vTFiw" colab={"base_uri": "https://localhost:8080/"} outputId="915deba6-b822-4b86-97d3-8965d1579ffc"
df.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="7gcHiC_JrupY" outputId="11a5cbc5-de03-4f37-8e7b-00d9154f8adf"
sns.boxplot(x='neighbourhood_group', y='price', data=df)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="OzN-Dw5Yr7aP" outputId="8e067200-c703-460d-88bd-5baa07bbce99"
sns.boxplot(x='room_type', y='price', data=df)
# + colab={"base_uri": "https://localhost:8080/"} id="nWZj4hKAsw9c" outputId="3ed92a9d-3eef-47e4-c314-4288fff2089a"
df.columns
# + colab={"base_uri": "https://localhost:8080/"} id="gx7NTZ_zszN0" outputId="70f4b911-d0ff-4a73-bf98-fcaa5f6bb6e7"
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="IrGafHOYtBfy" outputId="56f39e35-5a79-4766-ebf3-b3ed3a235df1"
df['neighbourhood_group'].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="ZHji8nlLtJwz" outputId="fc99bbaf-abf5-4cb9-c540-b21a277e8b8f"
neigh = df['neighbourhood'].value_counts()
plt.plot(range(len(neigh)), neigh)
# + id="DFNFgntWtcFB"
df['neighbourhood'] = df['neighbourhood'].apply(lambda s: s if str(s) not in neigh[50:] else 'others')
# + colab={"base_uri": "https://localhost:8080/"} id="xAr9FZ7ttsU_" outputId="f2a15b5b-10ec-42aa-e3ee-c0099958a9f8"
df['neighbourhood'].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="MJWdyw3Kt3q3" outputId="ed69d0bf-494c-4d84-8eaa-d608fe140574"
df['room_type'].value_counts()
# + colab={"base_uri": "https://localhost:8080/"} id="cy1jXTEQt79X" outputId="ed3e991a-b429-4229-e9b4-f52f4568accc"
df.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="pCjl92shuShT" outputId="578ad294-4b5d-4c40-9430-32bcc7a63951"
sns.rugplot(x='price', data=df, height=1)
# + colab={"base_uri": "https://localhost:8080/"} id="G6PmHDEcugBE" outputId="62818e85-8eba-4d76-b044-ba59403c2745"
print(df['price'].quantile(0.95))
print(df['price'].quantile(0.005))
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="ILmJBKuavIJ4" outputId="d9a44ce9-9b57-4622-8037-811346ee00b4"
sns.rugplot(x='minimum_nights', data=df, height=1)
# + colab={"base_uri": "https://localhost:8080/"} id="-y9AtGJcvRB4" outputId="33e586d9-c2e5-453d-e1c9-bcb43e466478"
print(df['minimum_nights'].quantile(0.98))
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="9DXcLeJPvdlA" outputId="ceb12c31-96ef-4bec-b4bb-ab99fc1c5bcf"
sns.rugplot(x='availability_365', data=df, height=1)
# + colab={"base_uri": "https://localhost:8080/"} id="C0YXy8RKvl84" outputId="133a6283-b2b7-4591-cf70-4bfc6049bb59"
print(df['availability_365'].quantile(0.3))
# + id="8HbUtvqs9C-C" colab={"base_uri": "https://localhost:8080/"} outputId="8f9ebb68-8f29-4466-9acc-319fbaa64790"
# quantile(), drop() 등 메소드를 이용하여 outlier 제거하고 통계 재분석하기
p1 = df['price'].quantile(0.95)
p2 = df['price'].quantile(0.005)
print(p1, p2)
# + id="iAyiIv88wEGM"
df = df[(df['price'] < p1) & (df['price'] > p2)]
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="xHVfBeOQwNUt" outputId="85c43436-c1cb-4b63-d06c-08920ac86760"
df['price'].hist()
# + colab={"base_uri": "https://localhost:8080/"} id="EB6s7-qawSRT" outputId="b7e9f5ee-e1be-422b-f59a-7ef23edd8d4d"
mn1 = df['minimum_nights'].quantile(0.98)
print(mn1)
# + id="v2KhzoXmwddb"
df = df[df['minimum_nights'] < mn1]
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="7e9l9co0w5mt" outputId="aeb7b77c-d0be-42d7-97d2-ff110529ec52"
df['minimum_nights'].hist()
# + id="SxcndS2ExTmU"
df['is_avail_zero'] = df['availability_365'].apply(lambda x: 'Zero' if x==0 else 'Nonzero')
# + id="NW-QEym6lgtX"
# fill(), dropna() 등으로 미기입된 데이터를 처리하기
df['review_exists'] = df['reviews_per_month'].isna().apply(lambda x: 'No' if x is True else 'Yes')
# + id="m8fGPMwVyyi_"
df.fillna(0, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="JgWVgoKZy4Zm" outputId="7a405436-81b4-4f25-dcf2-8da09b86d076"
df.isna().sum()
# + [markdown] id="FRfd7ABjepBS"
# ## 3. 모델 학습을 위한 데이터 전처리
# + colab={"base_uri": "https://localhost:8080/"} id="Q8hDQjWizbCV" outputId="bddf8a90-821b-483e-f7ce-3d7def8c4a9a"
df.columns
# + id="wVmEa1ChlrTc"
X_cat = df[['neighbourhood_group', 'neighbourhood', 'room_type', 'is_avail_zero', 'review_exists']]
X_cat = pd.get_dummies(X_cat)
# + id="_k_SDCh5xMgD"
from sklearn.preprocessing import StandardScaler
# + colab={"base_uri": "https://localhost:8080/"} id="XxBgGzGYzxh8" outputId="bf4252dc-7770-4dcc-b78c-f6faf21938f9"
df.columns
# + id="W3EO22NCE3wG"
# StandardScaler를 이용해 수치형 데이터를 표준화하기
scaler = StandardScaler()
X_num = df.drop(['neighbourhood_group', 'neighbourhood', 'room_type', 'price',
'last_review', 'is_avail_zero', 'review_exists'], axis=1)
scaler.fit(X_num)
X_scaled = scaler.transform(X_num)
X_scaled = pd.DataFrame(X_scaled, index=X_num.index, columns=X_num.columns)
X = pd.concat([X_scaled, X_cat], axis=1)
y = df['price']
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="ddgwL7Q90kAh" outputId="75e2d7b3-3f47-4423-d7c8-c4463eb316d8"
X.head()
# + id="F07QjOFwFNEw"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=1)
# + [markdown] id="RrWrE8Z4exup"
# ## 4. 모델 학습하기
# + id="LSSNqFUrGM6R" colab={"base_uri": "https://localhost:8080/"} outputId="d9b22b82-6a2b-4da7-a374-30a17c63a8aa"
model_reg = XGBRegressor()
model_reg.fit(X_train, y_train)
# + [markdown] id="gUo8NmHkfIpf"
# ## 5. 모델 평가하기
# + id="KDVy7fFGfUP1" colab={"base_uri": "https://localhost:8080/"} outputId="bc729969-4d2a-4a78-abff-bc6d09553a1c"
pred = model_reg.predict(X_test)
print(mean_absolute_error(y_test, pred))
print(sqrt(mean_squared_error(y_test, pred)))
# + [markdown] id="DTqb-HqPtc4I"
# ## 6. 모델 학습 결과 심화 분석하기
#
# + id="kKEP06-OmrBs" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="dc44dbfa-ec1a-4581-b11c-7d33e7bc0659"
# y_test vs. pred Scatter 플랏으로 시각적으로 분석하기
plt.scatter(x=y_test, y=pred, alpha=0.1)
plt.plot([0,350], [0, 350], 'r-')
# + id="WLnyYNJwGRgd" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="34b35d2c-f6c1-4e3b-f25f-07077aae1b61"
# err의 히스토그램으로 에러율 히스토그램 확인하기
err = (pred - y_test) / y_test
sns.histplot(err)
plt.grid()
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="7hhxUrZ12STu" outputId="fc23057a-78e3-4206-8b2f-2075f78297dc"
# err의 히스토그램으로 에러율 히스토그램 확인하기
err = pred - y_test
sns.histplot(err)
plt.grid()
# + [markdown] id="0w8vUO602pOk" pycharm={"name": "#%% md\n"}
# ## 참고
# * 퀴즈처럼 풀면서 배우는 파이썬 머신러닝 300제
# + pycharm={"name": "#%%\n"}
| season2/02.regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/thimotio/ExpertSystem/blob/master/Froth_Flotation_Fuzzy_Control_v01.ipynb)
# + [markdown] id="PajlL9-pi_25" colab_type="text"
# # Fuzzy Control Systems: Iron Ore Froth Flotation
#
#
# # Disclaimer
#
# This code was created for the educational purpose and sharing of advanced control techniques.
#
# For application in real cases of process in the industry it is necessary to model the controls as well as to identify the limits and interrelations of the variables to guarantee operational safety.
#
# **Authors are not responsible** for the results of using this code without due precautions.
#
# ---
#
# 
#
# ##The Froth Flotation
# [wikipedia](https://en.wikipedia.org/wiki/Froth_flotation)
#
# > Froth flotation is a process for selectively separating hydrophobic materials from hydrophilic. This is used in mineral processing, paper recycling and waste-water treatment industries. Historically this was first used in the mining industry, where it was one of the great enabling technologies of the 20th century. It has been described as "the single most important operation used for the recovery and upgrading of sulfide ores".[1] The development of froth flotation has improved the recovery of valuable minerals, such as copper- and lead-bearing minerals. Along with mechanized mining, it has allowed the economic recovery of valuable metals from much lower grade ore than previously.
#
#
# Foam flotation, or simply flotation, is a separation process applied to solid particles that exploits differences in surface characteristics between the various species present, treating heterogeneous mixtures of suspended particles in the aqueous phase in which a gas phase is introduced (Peres et al., 1980)
#
# The reverse cationic flotation of iron ore has this name because it occurs in an inverse way, with the gangue the specimen floated. This gangue consists mainly of SiO2 particles with induced hydrophobic characteristics. Practically, it can be considered as a direct silica flotation.
#
#
# > [1] <NAME>, "Flotation cell development," in: The AusIMM Annual Conference, Broken Hill, New South Wales, 17–21 May 1992 (The Australasian Institute of Mining and Metallurgy: Melbourne, 1992), 25–31.
#
#
# ###Froth basic control - Iron Ore
#
# Reverse quartz cationic flotation is the most commonly used itabirite iron ore concentration method for pellet feed production. Quartz is floated with etheramines partially neutralized with acetic acid and the iron minerals are depressed by unmodified starches.
#
# Although widely used, this method has high complexity and the domain of current knowledge is consolidated in the interfacial properties that govern its dynamics and phenomenological models experienced in industrial practice.
#
# There are several mineral and process variables that interfere with the dynamic behavior of flotation, and some of this knowledge is not completely elucidated. This is mainly due to the lack of means to measure specific properties that can explain more clearly its behavior in the face of the various interferences to which it is subject. This gap makes it very difficult to define more comprehensive models and that can represent in general the various flotation processes.
#
# Flotation involves the recovery of minerals in two distinct phases. The pulp phase, composed mainly of a mixture of mineral particles and gangue in aqueous medium, can be adequately described by first order kinetics. The foam phase is mainly composed of bubbles and their lamellae, and no suitable general model exists to describe this phase. As a result, the first order kinetic model is usually imposed on the entire flotation process (Mathe et al., 1998).
#
# The mechanism involved in the separation of the ore minerals and the gangue is possible thanks to the different properties of the mineral surfaces involved when in aqueous medium, at a certain pH. The addition of reagents with physicochemical characteristics capable of selectively modifying the surface properties of the minerals in relation to the solution, as well as the introduction of the gas phase into the system, creates conditions for the separation of the minerals.
#
# In this context, the operational practice combined with the technical knowledge of the state of the art of flotation allows modeling, through knowledge-based control techniques (expert system) and the development of operating support systems that control the process variables according to pre-established rules.
#
# + [markdown] id="AwwxKUKPogD4" colab_type="text"
# # The Control Problem
#
#
# ---
#
# ## Control Description
#
# Considering the typical iron ore flotation circuits we have 3 flotation steps with the aim of maximizing the metallic recovery and obtaining the necessary quality.
#
# Below we have an industrial circuit with the 3 steps (Rougher, Cleaner and Scavenger) and the current load considering the closed circuit.
#
# Two products are generated in this circuit: the flotation concentrate and the flotation reject.
#
# The products are directed to thickeners for water recovery and suitability for later stages of the process
#
# 
#
# Reagents are added to the feed to modulate the physicochemical characteristics of the pulp in order to obtain control of the desired specifications.
#
# Flotation machines have foam layer and air flow level controls.
#
# ### The reagents are:
#
# * Soda Caustica: modulates the pH of the pulp
# * Starch as the modifier in the process, preventing the amine collecting action on the iron particle
# * Amina - acts as collector and sparklingList item
#
# ## Control Strategy
#
# In this example we will consider the expert control of a Rougher column through the following variables:
#
#
# Controlled Variable (control objective)
#
# * Silica content in the concentrate
#
#
# Manipulated Variables (the ones I modify to get the goal)
#
# * Column Level
# * Air Flow in the Column
# * Amine Dosage
# * Starch dosage
#
# It is important to emphasize that this example is a simplification, and for the correct implementation one must consider all manipulated variables of the circuit in order to implement the control.
#
# The expansion of this control can be performed from the concepts presented in this example.
#
#
#
# + [markdown] id="svc_KjHN1hLe" colab_type="text"
# # Expert System: Fuzzy Controler Design
#
# Generally, expert systems evaluate the state of the process periodically (according to the control variables) and acts on the manipulated variables to correct any deviations.
#
# For the controlled variable (PV) it is common to define a target objective (SP) and to correct the control error (SP - PV). Such a variable is called an **Error**.
#
# > This error is evaluated by the operators as Low, Null or High.
#
# In order to evaluate the dynamics of the process the operators also evaluate the tendency of the error (**Gradient**). In this case, the gradient is nothing more than the difference between two error values in a time interval "d" (error (i) - error (i-d)) divided by time interval (delta t)
#
# > This Gradient is evaluated by operators such as Falling, Stable, or Rising.
#
# From the evaluation of the two information the control actions are taken as increments in the manipulated variables. The purpose of using increments is to move the operating point incrementally, without abrupt changes, however, in order to optimize the result.
#
# * **Amine increase:** increasing the amine implies in reducing the silica error, however, it has a high cost
# * **Increasing Air:** increasing the air has the objective of reducing the silica error. It acts on the dynamics of collecting the bubbles in the ore pulp and has a very low cost.
# * **Level Increment:** raising the level aims to increase the error of the silica. It is related to the pulp balance in the circuit and its productivity (low error allows to increase the productivity in the circuit by opening the control valve and increasing the foam level)
#
# For cost reduction an evaluation of the foam level will be inserted in order to increase the amine only if the level is above a low (<50%) level
# + id="Y_qwTWDxi_29" colab_type="code" colab={}
# !pip install -U scikit-fuzzy
'''Import modules'''
import numpy as np
import skfuzzy as fuzz #fuzzy controller module
import matplotlib
import matplotlib.pyplot
from skfuzzy import control as ctrl
# + [markdown] id="dRg9L6A-5n2w" colab_type="text"
# # Definition of membership funcions to Fuzzy Variables (controled and manipulated)
# + id="yOHTjBJki_3D" colab_type="code" colab={}
def NewAntecedentFuzzy_Exponencial(Name, vmin, vmax, precision, Curvas ):
'''
Defining membership function curves with S and Z Gaussian distribuctions
Parameters:
- Name: name of fuzzy variable
- vmin: minumum value of distribuction
- vmax: maximum value of distribuction
- precision: math precision for calculation
- Curves: Names of membership curves (3 or 5 names)
'''
#define the new Antecedent Fuzzy Variabel and set
FuzzyVar = ctrl.Antecedent(np.arange(vmin, vmax, precision), Name);
#if the MF has 3 membership functions
if np.size(Curvas) == 3:
mean = np.abs(vmax - vmin)/2 + vmin
std = np.abs(vmax - vmin)/6
FuzzyVar[Curvas[0]] = fuzz.zmf(FuzzyVar.universe, vmin, mean)
FuzzyVar[Curvas[1]] = fuzz.gaussmf(FuzzyVar.universe, mean, std )
FuzzyVar[Curvas[2]] = fuzz.smf(FuzzyVar.universe, mean, vmax)
if np.size(Curvas) == 5:
mean = np.abs(vmax - vmin)/2 + vmin
quarter1 = np.abs(vmax - vmin)/4 + vmin
quarter2 = np.abs(vmax - vmin)/4 + mean
std = np.abs(vmax - vmin)/12
FuzzyVar[Curvas[0]] = fuzz.zmf(FuzzyVar.universe, vmin, quarter1)
FuzzyVar[Curvas[1]] = fuzz.gaussmf(FuzzyVar.universe, quarter1, std )
FuzzyVar[Curvas[2]] = fuzz.gaussmf(FuzzyVar.universe, mean, std )
FuzzyVar[Curvas[3]] = fuzz.gaussmf(FuzzyVar.universe, quarter2, std )
FuzzyVar[Curvas[4]] = fuzz.smf(FuzzyVar.universe, quarter2, vmax)
return FuzzyVar
# + id="ty9odE1ji_3I" colab_type="code" colab={}
def NewConsequenceFuzzy_Triangular(Name, vmin, vmax, precision, Curvas ):
'''
Defining membership function curves with triangular distribuctions
Parameters:
- Name: name of fuzzy variable
- vmin: minumum value of distribuction
- vmax: maximum value of distribuction
- precision: math precision for calculation
- Curves: Names of membership curves (3 or 5 names)
'''
#define the new Antecedent Fuzzy Variabel and set
FuzzyVar = ctrl.Consequent(np.arange(vmin, vmax, precision), Name);
#if the MF has 3 membership functions
if np.size(Curvas) == 3:
mean = np.abs(vmax - vmin)/2 + vmin
FuzzyVar[Curvas[0]] = fuzz.trimf(FuzzyVar.universe, [vmin, vmin, mean] )
FuzzyVar[Curvas[1]] = fuzz.trimf(FuzzyVar.universe, [vmin, mean, vmax] )
FuzzyVar[Curvas[2]] = fuzz.trimf(FuzzyVar.universe, [mean, vmax, vmax] )
if np.size(Curvas) == 5:
mean = np.abs(vmax - vmin)/2 + vmin
quarter1 = np.abs(vmax - vmin)/4 + vmin
quarter2 = np.abs(vmax - vmin)/4 + mean
FuzzyVar[Curvas[0]] = fuzz.trimf(FuzzyVar.universe, [vmin, vmin, quarter1])
FuzzyVar[Curvas[1]] = fuzz.trimf(FuzzyVar.universe, [vmin, quarter1, mean] )
FuzzyVar[Curvas[2]] = fuzz.trimf(FuzzyVar.universe, [quarter1, mean, quarter2] )
FuzzyVar[Curvas[3]] = fuzz.trimf(FuzzyVar.universe, [mean, quarter2, vmax] )
FuzzyVar[Curvas[4]] = fuzz.trimf(FuzzyVar.universe, [quarter2, vmax, vmax] )
return FuzzyVar
# + [markdown] id="p3iZ0x4i6aJx" colab_type="text"
# ## Creat Fuzzy Variables
#
#
# ---
#
# Define the Antecedent variables (controled) and the Consequent variables (Manipulated)
# + id="PrnzIjRei_3L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1796} outputId="fa434e19-70dc-4e78-8f5f-592f80ec8392"
Names = ['Low', 'Null', 'High']
Erro = NewAntecedentFuzzy_Exponencial("Erro", -5, 5, 0.05, Names)
Level = NewAntecedentFuzzy_Exponencial("Level", 0, 100, 0.05, Names)
Erro.view()
Names = ['Falling' , 'Stable', 'Rising']
Gradient = NewAntecedentFuzzy_Exponencial("Gradient", -1, 1, 0.05, Names)
Gradient.view()
Names = ['Negative', 'Null', 'Positive']
inc_Amine = NewConsequenceFuzzy_Triangular("inc_Amine", -5, 5, 0.05, Names)
inc_Amine.view()
inc_Level = NewConsequenceFuzzy_Triangular("inc_Level", -10, 10, 0.05, Names)
inc_Level.view()
inc_Air = NewConsequenceFuzzy_Triangular("inc_Air", -1, 1, 0.05, Names)
inc_Air.view()
# + [markdown] id="BqUmY9cC5y7X" colab_type="text"
# ## Fuzzy Rules declaration
#
#
# ---
#
#
#
# ### Fuzzy rules
#
#
# Now, to make these triangles useful, we define the *fuzzy relationship*
# between input and output variables. For the purposes of our example, consider
# three simple rules:
#
# Most people would agree on these rules, but the rules are fuzzy. Mapping the
# imprecise rules into a defined, actionable tip is a challenge. This is the
# kind of task at which fuzzy logic excels.
#
#
# + [markdown] id="6ddK2wh5829j" colab_type="text"
# <html xmlns:v="urn:schemas-microsoft-com:vml"
# xmlns:o="urn:schemas-microsoft-com:office:office"
# xmlns:w="urn:schemas-microsoft-com:office:word"
# xmlns:x="urn:schemas-microsoft-com:office:excel"
# xmlns:m="http://schemas.microsoft.com/office/2004/12/omml"
# xmlns="http://www.w3.org/TR/REC-html40">
#
# <head>
# <meta http-equiv=Content-Type content="text/html; charset=windows-1252">
# <meta name=ProgId content=Word.Document>
# <meta name=Generator content="Microsoft Word 15">
# <meta name=Originator content="Microsoft Word 15">
# <link rel=File-List href="Air%20Increment_arquivos/filelist.xml">
# <!--[if gte mso 9]><xml>
# <o:DocumentProperties>
# <o:Author><NAME></o:Author>
# <o:LastAuthor><NAME></o:LastAuthor>
# <o:Revision>1</o:Revision>
# <o:TotalTime>2</o:TotalTime>
# <o:Created>2018-10-01T14:08:00Z</o:Created>
# <o:LastSaved>2018-10-01T14:10:00Z</o:LastSaved>
# <o:Pages>1</o:Pages>
# <o:Words>14</o:Words>
# <o:Characters>78</o:Characters>
# <o:Lines>1</o:Lines>
# <o:Paragraphs>1</o:Paragraphs>
# <o:CharactersWithSpaces>91</o:CharactersWithSpaces>
# <o:Version>16.00</o:Version>
# </o:DocumentProperties>
# <o:OfficeDocumentSettings>
# <o:AllowPNG/>
# </o:OfficeDocumentSettings>
# </xml><![endif]-->
# <link rel=themeData href="Air%20Increment_arquivos/themedata.thmx">
# <link rel=colorSchemeMapping
# href="Air%20Increment_arquivos/colorschememapping.xml">
# <!--[if gte mso 9]><xml>
# <w:WordDocument>
# <w:SpellingState>Clean</w:SpellingState>
# <w:GrammarState>Clean</w:GrammarState>
# <w:TrackMoves>false</w:TrackMoves>
# <w:TrackFormatting/>
# <w:HyphenationZone>21</w:HyphenationZone>
# <w:PunctuationKerning/>
# <w:ValidateAgainstSchemas/>
# <w:SaveIfXMLInvalid>false</w:SaveIfXMLInvalid>
# <w:IgnoreMixedContent>false</w:IgnoreMixedContent>
# <w:AlwaysShowPlaceholderText>false</w:AlwaysShowPlaceholderText>
# <w:DoNotPromoteQF/>
# <w:LidThemeOther>PT-BR</w:LidThemeOther>
# <w:LidThemeAsian>X-NONE</w:LidThemeAsian>
# <w:LidThemeComplexScript>X-NONE</w:LidThemeComplexScript>
# <w:Compatibility>
# <w:BreakWrappedTables/>
# <w:SnapToGridInCell/>
# <w:WrapTextWithPunct/>
# <w:UseAsianBreakRules/>
# <w:DontGrowAutofit/>
# <w:SplitPgBreakAndParaMark/>
# <w:EnableOpenTypeKerning/>
# <w:DontFlipMirrorIndents/>
# <w:OverrideTableStyleHps/>
# </w:Compatibility>
# <m:mathPr>
# <m:mathFont m:val="Cambria Math"/>
# <m:brkBin m:val="before"/>
# <m:brkBinSub m:val="--"/>
# <m:smallFrac m:val="off"/>
# <m:dispDef/>
# <m:lMargin m:val="0"/>
# <m:rMargin m:val="0"/>
# <m:defJc m:val="centerGroup"/>
# <m:wrapIndent m:val="1440"/>
# <m:intLim m:val="subSup"/>
# <m:naryLim m:val="undOvr"/>
# </m:mathPr></w:WordDocument>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <w:LatentStyles DefLockedState="false" DefUnhideWhenUsed="false"
# DefSemiHidden="false" DefQFormat="false" DefPriority="99"
# LatentStyleCount="375">
# <w:LsdException Locked="false" Priority="0" QFormat="true" Name="Normal"/>
# <w:LsdException Locked="false" Priority="9" QFormat="true" Name="heading 1"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 2"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 3"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 4"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 5"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 6"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 7"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 8"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 9"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 1"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 2"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 3"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 4"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 5"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 6"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 7"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 8"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="header"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footer"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index heading"/>
# <w:LsdException Locked="false" Priority="35" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="caption"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of figures"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope return"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="line number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="page number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of authorities"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="macro"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="toa heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 5"/>
# <w:LsdException Locked="false" Priority="10" QFormat="true" Name="Title"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Closing"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Signature"/>
# <w:LsdException Locked="false" Priority="1" SemiHidden="true"
# UnhideWhenUsed="true" Name="Default Paragraph Font"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Message Header"/>
# <w:LsdException Locked="false" Priority="11" QFormat="true" Name="Subtitle"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Salutation"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Date"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Note Heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Block Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="FollowedHyperlink"/>
# <w:LsdException Locked="false" Priority="22" QFormat="true" Name="Strong"/>
# <w:LsdException Locked="false" Priority="20" QFormat="true" Name="Emphasis"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Document Map"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Plain Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="E-mail Signature"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Top of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Bottom of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal (Web)"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Acronym"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Cite"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Code"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Definition"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Keyboard"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Preformatted"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Sample"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Typewriter"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Variable"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Table"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation subject"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="No List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Contemporary"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Elegant"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Professional"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Balloon Text"/>
# <w:LsdException Locked="false" Priority="39" Name="Table Grid"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Theme"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Placeholder Text"/>
# <w:LsdException Locked="false" Priority="1" QFormat="true" Name="No Spacing"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 1"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 1"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Revision"/>
# <w:LsdException Locked="false" Priority="34" QFormat="true"
# Name="List Paragraph"/>
# <w:LsdException Locked="false" Priority="29" QFormat="true" Name="Quote"/>
# <w:LsdException Locked="false" Priority="30" QFormat="true"
# Name="Intense Quote"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 1"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 1"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 2"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 2"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 2"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 3"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 3"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 3"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 4"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 4"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 4"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 5"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 5"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 5"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 6"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 6"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 6"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="19" QFormat="true"
# Name="Subtle Emphasis"/>
# <w:LsdException Locked="false" Priority="21" QFormat="true"
# Name="Intense Emphasis"/>
# <w:LsdException Locked="false" Priority="31" QFormat="true"
# Name="Subtle Reference"/>
# <w:LsdException Locked="false" Priority="32" QFormat="true"
# Name="Intense Reference"/>
# <w:LsdException Locked="false" Priority="33" QFormat="true" Name="Book Title"/>
# <w:LsdException Locked="false" Priority="37" SemiHidden="true"
# UnhideWhenUsed="true" Name="Bibliography"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="TOC Heading"/>
# <w:LsdException Locked="false" Priority="41" Name="Plain Table 1"/>
# <w:LsdException Locked="false" Priority="42" Name="Plain Table 2"/>
# <w:LsdException Locked="false" Priority="43" Name="Plain Table 3"/>
# <w:LsdException Locked="false" Priority="44" Name="Plain Table 4"/>
# <w:LsdException Locked="false" Priority="45" Name="Plain Table 5"/>
# <w:LsdException Locked="false" Priority="40" Name="Grid Table Light"/>
# <w:LsdException Locked="false" Priority="46" Name="Grid Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="Grid Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="Grid Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="46" Name="List Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="List Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="List Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Mention"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Smart Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hashtag"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Unresolved Mention"/>
# </w:LatentStyles>
# </xml><![endif]-->
# <style>
# <!--
# /* Font Definitions */
# @font-face
# {font-family:"Cambria Math";
# panose-1:2 4 5 3 5 4 6 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:roman;
# mso-font-pitch:variable;
# mso-font-signature:-536869121 1107305727 33554432 0 415 0;}
# @font-face
# {font-family:Calibri;
# panose-1:2 15 5 2 2 2 4 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:swiss;
# mso-font-pitch:variable;
# mso-font-signature:-536859905 -1073732485 9 0 511 0;}
# /* Style Definitions */
# p.MsoNormal, li.MsoNormal, div.MsoNormal
# {mso-style-unhide:no;
# mso-style-qformat:yes;
# mso-style-parent:"";
# margin-top:0cm;
# margin-right:0cm;
# margin-bottom:8.0pt;
# margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# span.SpellE
# {mso-style-name:"";
# mso-spl-e:yes;}
# .MsoChpDefault
# {mso-style-type:export-only;
# mso-default-props:yes;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# .MsoPapDefault
# {mso-style-type:export-only;
# margin-bottom:8.0pt;
# line-height:107%;}
# @page WordSection1
# {size:595.3pt 841.9pt;
# margin:70.85pt 3.0cm 70.85pt 3.0cm;
# mso-header-margin:35.4pt;
# mso-footer-margin:35.4pt;
# mso-paper-source:0;}
# div.WordSection1
# {page:WordSection1;}
# -->
# </style>
# <!--[if gte mso 10]>
# <style>
# /* Style Definitions */
# table.MsoNormalTable
# {mso-style-name:"Tabela normal";
# mso-tstyle-rowband-size:0;
# mso-tstyle-colband-size:0;
# mso-style-noshow:yes;
# mso-style-priority:99;
# mso-style-parent:"";
# mso-padding-alt:0cm 5.4pt 0cm 5.4pt;
# mso-para-margin-top:0cm;
# mso-para-margin-right:0cm;
# mso-para-margin-bottom:8.0pt;
# mso-para-margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# </style>
# <![endif]--><!--[if gte mso 9]><xml>
# <o:shapedefaults v:ext="edit" spidmax="1026"/>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <o:shapelayout v:ext="edit">
# <o:idmap v:ext="edit" data="1"/>
# </o:shapelayout></xml><![endif]-->
# </head>
#
# <body lang=PT-BR style='tab-interval:35.4pt'>
#
# <div class=WordSection1>
#
# <table class=MsoNormalTable border=0 cellspacing=0 cellpadding=0 width=441
# style='width:331.0pt;border-collapse:collapse;mso-yfti-tbllook:1184;
# mso-padding-alt:0cm 3.5pt 0cm 3.5pt'>
# <tr style='mso-yfti-irow:0;mso-yfti-firstrow:yes;height:15.75pt'>
# <td width=441 nowrap colspan=4 valign=bottom style='width:331.0pt;border:
# solid windowtext 1.0pt;border-right:solid black 1.0pt;background:#44546A;
# padding:0cm 3.5pt 0cm 3.5pt;height:15.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><b><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:white;
# mso-fareast-language:PT-BR'>Air <span class=SpellE>Increment</span> </span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:1;height:22.8pt'>
# <td width=74 nowrap rowspan=2 style='width:55.3pt;border-top:none;border-left:
# solid windowtext 1.0pt;border-bottom:solid black 1.0pt;border-right:solid windowtext 1.0pt;
# background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><b><span style='font-size:18.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>Erro </span></b></p>
# </td>
# <td width=368 nowrap colspan=3 valign=bottom style='width:275.7pt;border-top:
# none;border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid black 1.0pt;
# mso-border-top-alt:solid windowtext 1.0pt;background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;
# height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Gradient</span></b></span><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:2;height:30.75pt'>
# <td width=126 nowrap valign=bottom style='width:94.2pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Falling</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.85pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Stable</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.65pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Rising</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></i></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:3;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Low</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>- </span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>- </span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0 </span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:4;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Null</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'> </span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>- </span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0 </span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+ </span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:5;mso-yfti-lastrow:yes;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><i><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>High </span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0 </span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+ </span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+ </span></p>
# </td>
# </tr>
# </table>
#
# <p class=MsoNormal></p>
#
# </div>
#
# </body>
#
# </html>
#
# + id="9r8JxSv6i_3S" colab_type="code" colab={}
Air_Rules = []
Air_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Falling'], inc_Air['Negative']))
Air_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Stable'], inc_Air['Negative']))
Air_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Rising'], inc_Air['Null']))
Air_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Falling'], inc_Air['Negative']))
Air_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Stable'], inc_Air['Null']))
Air_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Rising'], inc_Air['Positive']))
Air_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Falling'], inc_Air['Null']))
Air_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Stable'], inc_Air['Positive']))
Air_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Rising'], inc_Air['Positive']))
Air_Ctrl = ctrl.ControlSystem(Air_Rules)
# + [markdown] id="HjyLRvx2CKpv" colab_type="text"
# <html xmlns:v="urn:schemas-microsoft-com:vml"
# xmlns:o="urn:schemas-microsoft-com:office:office"
# xmlns:w="urn:schemas-microsoft-com:office:word"
# xmlns:x="urn:schemas-microsoft-com:office:excel"
# xmlns:m="http://schemas.microsoft.com/office/2004/12/omml"
# xmlns="http://www.w3.org/TR/REC-html40">
#
# <head>
# <meta http-equiv=Content-Type content="text/html; charset=windows-1252">
# <meta name=ProgId content=Word.Document>
# <meta name=Generator content="Microsoft Word 15">
# <meta name=Originator content="Microsoft Word 15">
# <link rel=File-List href="Level%20%20Increment_arquivos/filelist.xml">
# <!--[if gte mso 9]><xml>
# <o:DocumentProperties>
# <o:Author><NAME></o:Author>
# <o:LastAuthor><NAME></o:LastAuthor>
# <o:Revision>2</o:Revision>
# <o:TotalTime>20</o:TotalTime>
# <o:Created>2018-10-01T14:28:00Z</o:Created>
# <o:LastSaved>2018-10-01T14:28:00Z</o:LastSaved>
# <o:Pages>1</o:Pages>
# <o:Words>14</o:Words>
# <o:Characters>78</o:Characters>
# <o:Lines>1</o:Lines>
# <o:Paragraphs>1</o:Paragraphs>
# <o:CharactersWithSpaces>91</o:CharactersWithSpaces>
# <o:Version>16.00</o:Version>
# </o:DocumentProperties>
# <o:OfficeDocumentSettings>
# <o:AllowPNG/>
# </o:OfficeDocumentSettings>
# </xml><![endif]-->
# <link rel=themeData href="Level%20%20Increment_arquivos/themedata.thmx">
# <link rel=colorSchemeMapping
# href="Level%20%20Increment_arquivos/colorschememapping.xml">
# <!--[if gte mso 9]><xml>
# <w:WordDocument>
# <w:SpellingState>Clean</w:SpellingState>
# <w:GrammarState>Clean</w:GrammarState>
# <w:TrackMoves>false</w:TrackMoves>
# <w:TrackFormatting/>
# <w:HyphenationZone>21</w:HyphenationZone>
# <w:PunctuationKerning/>
# <w:ValidateAgainstSchemas/>
# <w:SaveIfXMLInvalid>false</w:SaveIfXMLInvalid>
# <w:IgnoreMixedContent>false</w:IgnoreMixedContent>
# <w:AlwaysShowPlaceholderText>false</w:AlwaysShowPlaceholderText>
# <w:DoNotPromoteQF/>
# <w:LidThemeOther>PT-BR</w:LidThemeOther>
# <w:LidThemeAsian>X-NONE</w:LidThemeAsian>
# <w:LidThemeComplexScript>X-NONE</w:LidThemeComplexScript>
# <w:Compatibility>
# <w:BreakWrappedTables/>
# <w:SnapToGridInCell/>
# <w:WrapTextWithPunct/>
# <w:UseAsianBreakRules/>
# <w:DontGrowAutofit/>
# <w:SplitPgBreakAndParaMark/>
# <w:EnableOpenTypeKerning/>
# <w:DontFlipMirrorIndents/>
# <w:OverrideTableStyleHps/>
# </w:Compatibility>
# <m:mathPr>
# <m:mathFont m:val="Cambria Math"/>
# <m:brkBin m:val="before"/>
# <m:brkBinSub m:val="--"/>
# <m:smallFrac m:val="off"/>
# <m:dispDef/>
# <m:lMargin m:val="0"/>
# <m:rMargin m:val="0"/>
# <m:defJc m:val="centerGroup"/>
# <m:wrapIndent m:val="1440"/>
# <m:intLim m:val="subSup"/>
# <m:naryLim m:val="undOvr"/>
# </m:mathPr></w:WordDocument>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <w:LatentStyles DefLockedState="false" DefUnhideWhenUsed="false"
# DefSemiHidden="false" DefQFormat="false" DefPriority="99"
# LatentStyleCount="375">
# <w:LsdException Locked="false" Priority="0" QFormat="true" Name="Normal"/>
# <w:LsdException Locked="false" Priority="9" QFormat="true" Name="heading 1"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 2"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 3"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 4"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 5"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 6"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 7"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 8"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 9"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 1"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 2"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 3"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 4"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 5"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 6"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 7"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 8"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="header"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footer"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index heading"/>
# <w:LsdException Locked="false" Priority="35" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="caption"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of figures"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope return"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="line number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="page number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of authorities"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="macro"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="toa heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 5"/>
# <w:LsdException Locked="false" Priority="10" QFormat="true" Name="Title"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Closing"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Signature"/>
# <w:LsdException Locked="false" Priority="1" SemiHidden="true"
# UnhideWhenUsed="true" Name="Default Paragraph Font"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Message Header"/>
# <w:LsdException Locked="false" Priority="11" QFormat="true" Name="Subtitle"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Salutation"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Date"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Note Heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Block Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="FollowedHyperlink"/>
# <w:LsdException Locked="false" Priority="22" QFormat="true" Name="Strong"/>
# <w:LsdException Locked="false" Priority="20" QFormat="true" Name="Emphasis"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Document Map"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Plain Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="E-mail Signature"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Top of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Bottom of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal (Web)"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Acronym"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Cite"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Code"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Definition"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Keyboard"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Preformatted"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Sample"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Typewriter"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Variable"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Table"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation subject"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="No List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Contemporary"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Elegant"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Professional"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Balloon Text"/>
# <w:LsdException Locked="false" Priority="39" Name="Table Grid"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Theme"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Placeholder Text"/>
# <w:LsdException Locked="false" Priority="1" QFormat="true" Name="No Spacing"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 1"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 1"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Revision"/>
# <w:LsdException Locked="false" Priority="34" QFormat="true"
# Name="List Paragraph"/>
# <w:LsdException Locked="false" Priority="29" QFormat="true" Name="Quote"/>
# <w:LsdException Locked="false" Priority="30" QFormat="true"
# Name="Intense Quote"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 1"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 1"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 2"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 2"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 2"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 3"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 3"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 3"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 4"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 4"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 4"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 5"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 5"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 5"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 6"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 6"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 6"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="19" QFormat="true"
# Name="Subtle Emphasis"/>
# <w:LsdException Locked="false" Priority="21" QFormat="true"
# Name="Intense Emphasis"/>
# <w:LsdException Locked="false" Priority="31" QFormat="true"
# Name="Subtle Reference"/>
# <w:LsdException Locked="false" Priority="32" QFormat="true"
# Name="Intense Reference"/>
# <w:LsdException Locked="false" Priority="33" QFormat="true" Name="Book Title"/>
# <w:LsdException Locked="false" Priority="37" SemiHidden="true"
# UnhideWhenUsed="true" Name="Bibliography"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="TOC Heading"/>
# <w:LsdException Locked="false" Priority="41" Name="Plain Table 1"/>
# <w:LsdException Locked="false" Priority="42" Name="Plain Table 2"/>
# <w:LsdException Locked="false" Priority="43" Name="Plain Table 3"/>
# <w:LsdException Locked="false" Priority="44" Name="Plain Table 4"/>
# <w:LsdException Locked="false" Priority="45" Name="Plain Table 5"/>
# <w:LsdException Locked="false" Priority="40" Name="Grid Table Light"/>
# <w:LsdException Locked="false" Priority="46" Name="Grid Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="Grid Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="Grid Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="46" Name="List Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="List Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="List Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Mention"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Smart Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hashtag"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Unresolved Mention"/>
# </w:LatentStyles>
# </xml><![endif]-->
# <style>
# <!--
# /* Font Definitions */
# @font-face
# {font-family:"Cambria Math";
# panose-1:2 4 5 3 5 4 6 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:roman;
# mso-font-pitch:variable;
# mso-font-signature:-536869121 1107305727 33554432 0 415 0;}
# @font-face
# {font-family:Calibri;
# panose-1:2 15 5 2 2 2 4 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:swiss;
# mso-font-pitch:variable;
# mso-font-signature:-536859905 -1073732485 9 0 511 0;}
# /* Style Definitions */
# p.MsoNormal, li.MsoNormal, div.MsoNormal
# {mso-style-unhide:no;
# mso-style-qformat:yes;
# mso-style-parent:"";
# margin-top:0cm;
# margin-right:0cm;
# margin-bottom:8.0pt;
# margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# span.SpellE
# {mso-style-name:"";
# mso-spl-e:yes;}
# .MsoChpDefault
# {mso-style-type:export-only;
# mso-default-props:yes;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# .MsoPapDefault
# {mso-style-type:export-only;
# margin-bottom:8.0pt;
# line-height:107%;}
# @page WordSection1
# {size:595.3pt 841.9pt;
# margin:70.85pt 3.0cm 70.85pt 3.0cm;
# mso-header-margin:35.4pt;
# mso-footer-margin:35.4pt;
# mso-paper-source:0;}
# div.WordSection1
# {page:WordSection1;}
# -->
# </style>
# <!--[if gte mso 10]>
# <style>
# /* Style Definitions */
# table.MsoNormalTable
# {mso-style-name:"Tabela normal";
# mso-tstyle-rowband-size:0;
# mso-tstyle-colband-size:0;
# mso-style-noshow:yes;
# mso-style-priority:99;
# mso-style-parent:"";
# mso-padding-alt:0cm 5.4pt 0cm 5.4pt;
# mso-para-margin-top:0cm;
# mso-para-margin-right:0cm;
# mso-para-margin-bottom:8.0pt;
# mso-para-margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# </style>
# <![endif]--><!--[if gte mso 9]><xml>
# <o:shapedefaults v:ext="edit" spidmax="1026"/>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <o:shapelayout v:ext="edit">
# <o:idmap v:ext="edit" data="1"/>
# </o:shapelayout></xml><![endif]-->
# </head>
#
# <body lang=PT-BR style='tab-interval:35.4pt'>
#
# <div class=WordSection1>
#
# <table class=MsoNormalTable border=0 cellspacing=0 cellpadding=0 width=441
# style='width:331.0pt;border-collapse:collapse;mso-yfti-tbllook:1184;
# mso-padding-alt:0cm 3.5pt 0cm 3.5pt'>
# <tr style='mso-yfti-irow:0;mso-yfti-firstrow:yes;height:15.75pt'>
# <td width=441 nowrap colspan=4 valign=bottom style='width:331.0pt;border:
# solid windowtext 1.0pt;border-right:solid black 1.0pt;background:#44546A;
# padding:0cm 3.5pt 0cm 3.5pt;height:15.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><b><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:white;mso-fareast-language:PT-BR'>Level</span></b></span><b><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:white;mso-fareast-language:PT-BR'> <span class=SpellE>Increment</span></span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:1;height:22.8pt'>
# <td width=74 nowrap rowspan=2 style='width:55.3pt;border-top:none;border-left:
# solid windowtext 1.0pt;border-bottom:solid black 1.0pt;border-right:solid windowtext 1.0pt;
# background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><b><span style='font-size:18.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>Erro</span></b></p>
# </td>
# <td width=368 nowrap colspan=3 valign=bottom style='width:275.7pt;border-top:
# none;border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid black 1.0pt;
# mso-border-top-alt:solid windowtext 1.0pt;background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;
# height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Gradient</span></b></span><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:2;height:30.75pt'>
# <td width=126 nowrap valign=bottom style='width:94.2pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Falling</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.85pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Stable</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.65pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Rising</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:3;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Low</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:4;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Null</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;mso-background-themecolor:background1;mso-background-themeshade:
# 217;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:5;mso-yfti-lastrow:yes;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><i><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>High</span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;mso-background-themecolor:background1;mso-background-themeshade:
# 217;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;mso-background-themecolor:background1;padding:0cm 3.5pt 0cm 3.5pt;
# height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# </tr>
# </table>
#
# <p class=MsoNormal><o:p> </o:p></p>
#
# </div>
#
# </body>
#
# </html>
#
# + id="tre4bmfN_53K" colab_type="code" colab={}
Level_Rules = []
Level_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Falling'], inc_Level['Positive']))
Level_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Stable'], inc_Level['Positive']))
Level_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Rising'], inc_Level['Null']))
Level_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Falling'], inc_Level['Positive']))
Level_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Stable'], inc_Level['Null']))
Level_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Rising'], inc_Level['Negative']))
Level_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Falling'], inc_Level['Null']))
Level_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Stable'], inc_Level['Negative']))
Level_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Rising'], inc_Level['Negative']))
Level_Ctrl = ctrl.ControlSystem(Level_Rules)
# + [markdown] id="ab6_LVawBXzP" colab_type="text"
# <html xmlns:v="urn:schemas-microsoft-com:vml"
# xmlns:o="urn:schemas-microsoft-com:office:office"
# xmlns:w="urn:schemas-microsoft-com:office:word"
# xmlns:x="urn:schemas-microsoft-com:office:excel"
# xmlns:m="http://schemas.microsoft.com/office/2004/12/omml"
# xmlns="http://www.w3.org/TR/REC-html40">
#
# <head>
# <meta http-equiv=Content-Type content="text/html; charset=windows-1252">
# <meta name=ProgId content=Word.Document>
# <meta name=Generator content="Microsoft Word 15">
# <meta name=Originator content="Microsoft Word 15">
# <link rel=File-List href="Amine%20Increment_arquivos/filelist.xml">
# <!--[if gte mso 9]><xml>
# <o:DocumentProperties>
# <o:Author><NAME></o:Author>
# <o:LastAuthor><NAME></o:LastAuthor>
# <o:Revision>2</o:Revision>
# <o:TotalTime>18</o:TotalTime>
# <o:Created>2018-10-01T14:26:00Z</o:Created>
# <o:LastSaved>2018-10-01T14:26:00Z</o:LastSaved>
# <o:Pages>1</o:Pages>
# <o:Words>20</o:Words>
# <o:Characters>113</o:Characters>
# <o:Lines>1</o:Lines>
# <o:Paragraphs>1</o:Paragraphs>
# <o:CharactersWithSpaces>132</o:CharactersWithSpaces>
# <o:Version>16.00</o:Version>
# </o:DocumentProperties>
# <o:OfficeDocumentSettings>
# <o:AllowPNG/>
# </o:OfficeDocumentSettings>
# </xml><![endif]-->
# <link rel=themeData href="Amine%20Increment_arquivos/themedata.thmx">
# <link rel=colorSchemeMapping
# href="Amine%20Increment_arquivos/colorschememapping.xml">
# <!--[if gte mso 9]><xml>
# <w:WordDocument>
# <w:SpellingState>Clean</w:SpellingState>
# <w:GrammarState>Clean</w:GrammarState>
# <w:TrackMoves>false</w:TrackMoves>
# <w:TrackFormatting/>
# <w:HyphenationZone>21</w:HyphenationZone>
# <w:PunctuationKerning/>
# <w:ValidateAgainstSchemas/>
# <w:SaveIfXMLInvalid>false</w:SaveIfXMLInvalid>
# <w:IgnoreMixedContent>false</w:IgnoreMixedContent>
# <w:AlwaysShowPlaceholderText>false</w:AlwaysShowPlaceholderText>
# <w:DoNotPromoteQF/>
# <w:LidThemeOther>PT-BR</w:LidThemeOther>
# <w:LidThemeAsian>X-NONE</w:LidThemeAsian>
# <w:LidThemeComplexScript>X-NONE</w:LidThemeComplexScript>
# <w:Compatibility>
# <w:BreakWrappedTables/>
# <w:SnapToGridInCell/>
# <w:WrapTextWithPunct/>
# <w:UseAsianBreakRules/>
# <w:DontGrowAutofit/>
# <w:SplitPgBreakAndParaMark/>
# <w:EnableOpenTypeKerning/>
# <w:DontFlipMirrorIndents/>
# <w:OverrideTableStyleHps/>
# </w:Compatibility>
# <m:mathPr>
# <m:mathFont m:val="Cambria Math"/>
# <m:brkBin m:val="before"/>
# <m:brkBinSub m:val="--"/>
# <m:smallFrac m:val="off"/>
# <m:dispDef/>
# <m:lMargin m:val="0"/>
# <m:rMargin m:val="0"/>
# <m:defJc m:val="centerGroup"/>
# <m:wrapIndent m:val="1440"/>
# <m:intLim m:val="subSup"/>
# <m:naryLim m:val="undOvr"/>
# </m:mathPr></w:WordDocument>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <w:LatentStyles DefLockedState="false" DefUnhideWhenUsed="false"
# DefSemiHidden="false" DefQFormat="false" DefPriority="99"
# LatentStyleCount="375">
# <w:LsdException Locked="false" Priority="0" QFormat="true" Name="Normal"/>
# <w:LsdException Locked="false" Priority="9" QFormat="true" Name="heading 1"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 2"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 3"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 4"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 5"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 6"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 7"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 8"/>
# <w:LsdException Locked="false" Priority="9" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="heading 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index 9"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 1"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 2"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 3"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 4"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 5"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 6"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 7"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 8"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" Name="toc 9"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="header"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footer"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="index heading"/>
# <w:LsdException Locked="false" Priority="35" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="caption"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of figures"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="envelope return"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="footnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="line number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="page number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote reference"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="endnote text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="table of authorities"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="macro"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="toa heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Bullet 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Number 5"/>
# <w:LsdException Locked="false" Priority="10" QFormat="true" Name="Title"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Closing"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Signature"/>
# <w:LsdException Locked="false" Priority="1" SemiHidden="true"
# UnhideWhenUsed="true" Name="Default Paragraph Font"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="List Continue 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Message Header"/>
# <w:LsdException Locked="false" Priority="11" QFormat="true" Name="Subtitle"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Salutation"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Date"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text First Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Note Heading"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Body Text Indent 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Block Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="FollowedHyperlink"/>
# <w:LsdException Locked="false" Priority="22" QFormat="true" Name="Strong"/>
# <w:LsdException Locked="false" Priority="20" QFormat="true" Name="Emphasis"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Document Map"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Plain Text"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="E-mail Signature"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Top of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Bottom of Form"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal (Web)"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Acronym"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Address"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Cite"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Code"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Definition"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Keyboard"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Preformatted"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Sample"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Typewriter"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="HTML Variable"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Normal Table"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="annotation subject"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="No List"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Outline List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Simple 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Classic 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Colorful 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Columns 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Grid 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 4"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 5"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 7"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table List 8"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table 3D effects 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Contemporary"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Elegant"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Professional"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Subtle 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 1"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 2"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Web 3"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Balloon Text"/>
# <w:LsdException Locked="false" Priority="39" Name="Table Grid"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Table Theme"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Placeholder Text"/>
# <w:LsdException Locked="false" Priority="1" QFormat="true" Name="No Spacing"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 1"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 1"/>
# <w:LsdException Locked="false" SemiHidden="true" Name="Revision"/>
# <w:LsdException Locked="false" Priority="34" QFormat="true"
# Name="List Paragraph"/>
# <w:LsdException Locked="false" Priority="29" QFormat="true" Name="Quote"/>
# <w:LsdException Locked="false" Priority="30" QFormat="true"
# Name="Intense Quote"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 1"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 1"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 1"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 1"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 1"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 2"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 2"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 2"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 2"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 2"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 2"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 3"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 3"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 3"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 3"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 3"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 3"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 4"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 4"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 4"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 4"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 4"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 4"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 5"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 5"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 5"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 5"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 5"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 5"/>
# <w:LsdException Locked="false" Priority="60" Name="Light Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="61" Name="Light List Accent 6"/>
# <w:LsdException Locked="false" Priority="62" Name="Light Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="63" Name="Medium Shading 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="64" Name="Medium Shading 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="65" Name="Medium List 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="66" Name="Medium List 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="67" Name="Medium Grid 1 Accent 6"/>
# <w:LsdException Locked="false" Priority="68" Name="Medium Grid 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="69" Name="Medium Grid 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="70" Name="Dark List Accent 6"/>
# <w:LsdException Locked="false" Priority="71" Name="Colorful Shading Accent 6"/>
# <w:LsdException Locked="false" Priority="72" Name="Colorful List Accent 6"/>
# <w:LsdException Locked="false" Priority="73" Name="Colorful Grid Accent 6"/>
# <w:LsdException Locked="false" Priority="19" QFormat="true"
# Name="Subtle Emphasis"/>
# <w:LsdException Locked="false" Priority="21" QFormat="true"
# Name="Intense Emphasis"/>
# <w:LsdException Locked="false" Priority="31" QFormat="true"
# Name="Subtle Reference"/>
# <w:LsdException Locked="false" Priority="32" QFormat="true"
# Name="Intense Reference"/>
# <w:LsdException Locked="false" Priority="33" QFormat="true" Name="Book Title"/>
# <w:LsdException Locked="false" Priority="37" SemiHidden="true"
# UnhideWhenUsed="true" Name="Bibliography"/>
# <w:LsdException Locked="false" Priority="39" SemiHidden="true"
# UnhideWhenUsed="true" QFormat="true" Name="TOC Heading"/>
# <w:LsdException Locked="false" Priority="41" Name="Plain Table 1"/>
# <w:LsdException Locked="false" Priority="42" Name="Plain Table 2"/>
# <w:LsdException Locked="false" Priority="43" Name="Plain Table 3"/>
# <w:LsdException Locked="false" Priority="44" Name="Plain Table 4"/>
# <w:LsdException Locked="false" Priority="45" Name="Plain Table 5"/>
# <w:LsdException Locked="false" Priority="40" Name="Grid Table Light"/>
# <w:LsdException Locked="false" Priority="46" Name="Grid Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="Grid Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="Grid Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="Grid Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="Grid Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="Grid Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="Grid Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="Grid Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="Grid Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="Grid Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="46" Name="List Table 1 Light"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark"/>
# <w:LsdException Locked="false" Priority="51" Name="List Table 6 Colorful"/>
# <w:LsdException Locked="false" Priority="52" Name="List Table 7 Colorful"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 1"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 1"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 1"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 1"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 1"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 1"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 2"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 2"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 2"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 2"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 2"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 2"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 3"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 3"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 3"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 3"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 3"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 3"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 4"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 4"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 4"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 4"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 4"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 4"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 5"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 5"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 5"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 5"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 5"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 5"/>
# <w:LsdException Locked="false" Priority="46"
# Name="List Table 1 Light Accent 6"/>
# <w:LsdException Locked="false" Priority="47" Name="List Table 2 Accent 6"/>
# <w:LsdException Locked="false" Priority="48" Name="List Table 3 Accent 6"/>
# <w:LsdException Locked="false" Priority="49" Name="List Table 4 Accent 6"/>
# <w:LsdException Locked="false" Priority="50" Name="List Table 5 Dark Accent 6"/>
# <w:LsdException Locked="false" Priority="51"
# Name="List Table 6 Colorful Accent 6"/>
# <w:LsdException Locked="false" Priority="52"
# Name="List Table 7 Colorful Accent 6"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Mention"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Smart Hyperlink"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Hashtag"/>
# <w:LsdException Locked="false" SemiHidden="true" UnhideWhenUsed="true"
# Name="Unresolved Mention"/>
# </w:LatentStyles>
# </xml><![endif]-->
# <style>
# <!--
# /* Font Definitions */
# @font-face
# {font-family:"Cambria Math";
# panose-1:2 4 5 3 5 4 6 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:roman;
# mso-font-pitch:variable;
# mso-font-signature:-536869121 1107305727 33554432 0 415 0;}
# @font-face
# {font-family:Calibri;
# panose-1:2 15 5 2 2 2 4 3 2 4;
# mso-font-charset:0;
# mso-generic-font-family:swiss;
# mso-font-pitch:variable;
# mso-font-signature:-536859905 -1073732485 9 0 511 0;}
# /* Style Definitions */
# p.MsoNormal, li.MsoNormal, div.MsoNormal
# {mso-style-unhide:no;
# mso-style-qformat:yes;
# mso-style-parent:"";
# margin-top:0cm;
# margin-right:0cm;
# margin-bottom:8.0pt;
# margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# span.SpellE
# {mso-style-name:"";
# mso-spl-e:yes;}
# span.GramE
# {mso-style-name:"";
# mso-gram-e:yes;}
# .MsoChpDefault
# {mso-style-type:export-only;
# mso-default-props:yes;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-fareast-font-family:Calibri;
# mso-fareast-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# .MsoPapDefault
# {mso-style-type:export-only;
# margin-bottom:8.0pt;
# line-height:107%;}
# @page WordSection1
# {size:595.3pt 841.9pt;
# margin:70.85pt 3.0cm 70.85pt 3.0cm;
# mso-header-margin:35.4pt;
# mso-footer-margin:35.4pt;
# mso-paper-source:0;}
# div.WordSection1
# {page:WordSection1;}
# -->
# </style>
# <!--[if gte mso 10]>
# <style>
# /* Style Definitions */
# table.MsoNormalTable
# {mso-style-name:"Tabela normal";
# mso-tstyle-rowband-size:0;
# mso-tstyle-colband-size:0;
# mso-style-noshow:yes;
# mso-style-priority:99;
# mso-style-parent:"";
# mso-padding-alt:0cm 5.4pt 0cm 5.4pt;
# mso-para-margin-top:0cm;
# mso-para-margin-right:0cm;
# mso-para-margin-bottom:8.0pt;
# mso-para-margin-left:0cm;
# line-height:107%;
# mso-pagination:widow-orphan;
# font-size:11.0pt;
# font-family:"Calibri",sans-serif;
# mso-ascii-font-family:Calibri;
# mso-ascii-theme-font:minor-latin;
# mso-hansi-font-family:Calibri;
# mso-hansi-theme-font:minor-latin;
# mso-bidi-font-family:"Times New Roman";
# mso-bidi-theme-font:minor-bidi;
# mso-fareast-language:EN-US;}
# </style>
# <![endif]--><!--[if gte mso 9]><xml>
# <o:shapedefaults v:ext="edit" spidmax="1026"/>
# </xml><![endif]--><!--[if gte mso 9]><xml>
# <o:shapelayout v:ext="edit">
# <o:idmap v:ext="edit" data="1"/>
# </o:shapelayout></xml><![endif]-->
# </head>
#
# <body lang=PT-BR style='tab-interval:35.4pt'>
#
# <div class=WordSection1>
#
# <table class=MsoNormalTable border=0 cellspacing=0 cellpadding=0 width=441
# style='width:331.0pt;border-collapse:collapse;mso-yfti-tbllook:1184;
# mso-padding-alt:0cm 3.5pt 0cm 3.5pt'>
# <tr style='mso-yfti-irow:0;mso-yfti-firstrow:yes;height:15.75pt'>
# <td width=441 nowrap colspan=4 valign=bottom style='width:331.0pt;border:
# solid windowtext 1.0pt;border-right:solid black 1.0pt;background:#44546A;
# padding:0cm 3.5pt 0cm 3.5pt;height:15.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><b><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:white;mso-fareast-language:PT-BR'>Amine</span></b></span><b><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:white;mso-fareast-language:PT-BR'> <span class=SpellE>Increment</span></span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:1;height:22.8pt'>
# <td width=74 nowrap rowspan=2 style='width:55.3pt;border-top:none;border-left:
# solid windowtext 1.0pt;border-bottom:solid black 1.0pt;border-right:solid windowtext 1.0pt;
# background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><b><span style='font-size:18.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>Erro</span></b></p>
# </td>
# <td width=368 nowrap colspan=3 valign=bottom style='width:275.7pt;border-top:
# none;border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid black 1.0pt;
# mso-border-top-alt:solid windowtext 1.0pt;background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;
# height:22.8pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Gradient</span></b></span><b><span
# style='font-size:16.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></b></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:2;height:30.75pt'>
# <td width=126 nowrap valign=bottom style='width:94.2pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Falling</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.85pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Stable</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=121 nowrap valign=bottom style='width:90.65pt;border-top:none;
# border-left:none;border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#FFF2CC;padding:0cm 3.5pt 0cm 3.5pt;height:30.75pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Rising</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:3;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Low</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:4;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span class=SpellE><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'>Null</span></i></span><i><span
# style='font-size:12.0pt;mso-ascii-font-family:Calibri;mso-fareast-font-family:
# "Times New Roman";mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;
# color:black;mso-fareast-language:PT-BR'></span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>-</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:#D9D9D9;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:yellow;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# </tr>
# <tr style='mso-yfti-irow:5;mso-yfti-lastrow:yes;height:21.6pt'>
# <td width=74 nowrap style='width:55.3pt;border:solid windowtext 1.0pt;
# border-top:none;background:#DEEAF6;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><i><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>High</span></i></p>
# </td>
# <td width=126 style='width:94.2pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:white;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>0</span></p>
# </td>
# <td width=121 style='width:90.85pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:yellow;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# <td width=121 style='width:90.65pt;border-top:none;border-left:none;
# border-bottom:solid windowtext 1.0pt;border-right:solid windowtext 1.0pt;
# background:yellow;padding:0cm 3.5pt 0cm 3.5pt;height:21.6pt'>
# <p class=MsoNormal align=center style='margin-bottom:0cm;margin-bottom:.0001pt;
# text-align:center;line-height:normal'><span style='font-size:12.0pt;
# mso-ascii-font-family:Calibri;mso-fareast-font-family:"Times New Roman";
# mso-hansi-font-family:Calibri;mso-bidi-font-family:Calibri;color:black;
# mso-fareast-language:PT-BR'>+</span></p>
# </td>
# </tr>
# </table>
#
# <p class=MsoNormal>Positive incremente <span class=SpellE>only</span> <span
# class=SpellE><span class=GramE>if</span></span><span class=GramE> Leve</span>
# lis <span class=SpellE>Low</span></p>
#
# </div>
#
# </body>
#
# </html>
#
#
#
# + id="SIVD7nX9A0Re" colab_type="code" colab={}
Amine_Rules = []
Amine_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Falling'], inc_Amine['Negative']))
Amine_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Stable'], inc_Amine['Negative']))
Amine_Rules.append(ctrl.Rule(Erro['Low'] & Gradient['Rising'], inc_Amine['Null']))
Amine_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Falling'], inc_Amine['Negative']))
Amine_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Stable'], inc_Amine['Null']))
Amine_Rules.append(ctrl.Rule(Erro['Null'] & Gradient['Rising'] & Level['Low'], inc_Amine['Positive']))
Amine_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Falling'], inc_Amine['Null']))
Amine_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Stable'] & Level['Low'], inc_Amine['Positive']))
Amine_Rules.append(ctrl.Rule(Erro['High'] & Gradient['Rising'] & Level['Low'], inc_Amine['Positive']))
Amine_Ctrl = ctrl.ControlSystem(Amine_Rules)
# + [markdown] id="bm-XE6EJ54td" colab_type="text"
# ### Fuzzy Control System Creation and Simulation
#
#
# Now that we have our rules defined, we can simply create a control system
# via: **ControlSystemSimulation**
#
# In order to simulate this control system, we will create a ``ControlSystemSimulation``. Think of this object representing our controller applied to a specific set of cirucmstances.
# + id="sXrjw3y8i_3W" colab_type="code" colab={}
Control_Air = ctrl.ControlSystemSimulation(Air_Ctrl)
Control_Level = ctrl.ControlSystemSimulation(Level_Ctrl)
Control_Amine = ctrl.ControlSystemSimulation(Amine_Ctrl)
# + [markdown] id="o5nVupsRdrXm" colab_type="text"
# # Fuzzy Control Simulation
# + id="uckFPHWFG8Rt" colab_type="code" colab={}
'Initialize Variables'
#@title #Froth Flotation Simulation
#@markdown #Enter simulation process data:
#@markdown ### Configurations:
delta_t = 5.0 #@param {type: "number"}
maximum_acceptable_delta = 0.2 #@param {type: "number"}
#@markdown ### Process Variables:
Silicium = 1.4 #@param {type: "number"}
Silicium_SP = 3 #@param {type: "number"}
Froth_Level = 20.0 #@param {type: "number"}
Silicium_Before = 2.1 #@param {type: "number"}
Silicium_Error = Silicium_SP-Silicium
Grad = ((Silicium - Silicium_Before)/delta_t)/maximum_acceptable_delta
if(Grad>1):
Grad = 1.
if (Grad < -1):
Grad = -1
# + [markdown] id="qd7JvLnRGyCh" colab_type="text"
# ## Air Control Simulation
# + id="L81WnwTZi_3Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1247} outputId="e7b99f5a-daa5-4794-9017-790d780af895"
Control_Air.input['Erro'] = Silicium_Error
Control_Air.input['Gradient'] = Grad
# Crunch the numbers
Control_Air.compute()
print("-------------------------------")
print("For a Silicium Grade = " + str(Silicium))
print("Control Error = " + str(Silicium_Error))
print("Gradient Error = " + str(Grad))
print("The control Air increment should be = " + str(Control_Air.output['inc_Air']))
print("-------------------------------\n\n")
"""
Once computed, we can view the result as well as visualize it.
"""
Erro.view(sim=Control_Air)
Gradient.view(sim=Control_Air)
inc_Air.view(sim=Control_Air)
# + id="oWDwdTElHEvM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1247} outputId="6787dc7b-fcf8-4063-be8f-1b199b171bd9"
Control_Level.input['Erro'] = Silicium_Error
Control_Level.input['Gradient'] = Grad
# Crunch the numbers
Control_Level.compute()
print("-------------------------------")
print("For a Silicium Grade = " + str(Silicium))
print("Control Error = " + str(Silicium_Error))
print("Gradient Error = " + str(Grad))
print("The control Level increment should be = " + str(Control_Level.output['inc_Level']))
print("-------------------------------\n\n")
"""
Once computed, we can view the result as well as visualize it.
"""
Erro.view(sim=Control_Level)
Gradient.view(sim=Control_Level)
inc_Level.view(sim=Control_Level)
# + id="aQjfJPxbi_3c" colab_type="code" colab={} outputId="34b342f8-9cca-4db5-a9fc-336f2e060da0"
"""# We can simulate at higher resolution with full accuracy
upsampled = np.linspace(0.1, 10, 21)
x = upsampled #np.meshgrid(upsampled, upsampled)
upsampled = np.linspace(0.1, 25, 21)
y = upsampled #np.meshgrid(upsampled, upsampled)
z = np.zeros_like(np.meshgrid(x, y))
print(x)
print(y)
print(z)
# Loop through the system 21*21 times to collect the control surface
for i in range(21):
for j in range(21):
#print(y[i])
tipping.input['Silica'] = (x)
tipping.input['Ferro'] = (y[j])
tipping.compute()
test = tipping.output['inc_Amina']
print(test)
z[i][j] = np.float(test)
# Plot the result in pretty 3D with alpha blending
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # Required for 3D plotting
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(111, projection='3d')
surf = ax.plot_surface(x, y, z, rstride=1, cstride=1, cmap='viridis',
linewidth=0.4, antialiased=True)
cset = ax.contourf(x, y, z, zdir='z', offset=-2.5, cmap='viridis', alpha=0.5)
cset = ax.contourf(x, y, z, zdir='x', offset=3, cmap='viridis', alpha=0.5)
cset = ax.contourf(x, y, z, zdir='y', offset=3, cmap='viridis', alpha=0.5)
ax.view_init(30, 200)
"""
# + id="ALjLqYdXi_3g" colab_type="code" colab={}
# + id="0yW8osPGi_3j" colab_type="code" colab={}
# + id="LRz6vq0_i_3l" colab_type="code" colab={}
| Froth_Flotation_Fuzzy_Control_v01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="./img/hsfrsl.jpg"/>
#
# # House price prediction
#
# ### Intro
# In this workshop we will see the basics of machine learning and deep learning by trying to predict real estate prices. To do so, we will use two machine learning libraries: [sklearn](https://scikit-learn.org) and [pytorch](https://pytorch.org).
#
# #### What is Machine learning
# Machine learning is the study of computer algorithms that improve automatically through experience and by the use of data\
# (TL;DR: A machine learning model is an AI learning more or less by itself)
#
# ### Import
# - `pandas`: data manipulation and analysis
# - `numpy`: support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays
# +
import pandas as pd
import numpy as np
np.random.seed(0)
# -
# We said that we want to predict real estate's prices, so before building any model let's take a look at our data.
#
# All our data is stored in a csv file, we can read it by using `panda.read_csv`, it takes in argument the path to our csv.
#
# Quick note: `.sample(frac=1)` will shuffle our data in case it's sorted. We don't like sorted data.
table = pd.read_csv("data/data.csv").sample(frac=1)
# Ok, well, we readed our csv but how to show what it contains ? That is exactly what you have to figure out.
#
# **Instruction:**
# - Show the first five rows of our csv
#
# **Help:**
# - [Dataframe.head](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.head.html)
# +
# Start of your code (1 line)
# End of your code
# -
# Take time to see what columns we have.
#
# ## Data processing 1.1
#
# We see that we have a lot of data, some may be unnecessary like `date` and others may be more useful like `bedrooms` (the number of bedrooms in the house).
#
# To simplify the workshop, we decide to drop some specific columns like `yr_renovated`, `street`, and `statezip`.
#
# **Instruction:**
# - Drop `date`, `yr_renovated`, `street`, and `statezip` columns.
#
# **Help:**
# - "Tell me [how to drop a column](https://letmegooglethat.com/?q=drop+column+pandas) please :("
# +
# Start of your code (1 line)
# End of your code
# -
# I told you that the `date` column is not useful to train our model but how to know if a column is important ?
#
# For example, let's take a look at the `country` column.
# We all agree that the country can influence the price of a house, but if all the houses are in the same country, will it still be useful to precise the country ? Of course, the answer is no.
#
# **Instruction:**
# - Try to count the number of different countries in our data.
#
# **Helps:**
# - Cast a column into a `list`: https://stackoverflow.com/questions/23748995/pandas-dataframe-column-to-list
# - `set` in python: https://www.programiz.com/python-programming/set
# +
# Start of your code (1 line)
# End of your code
# -
# What a surprise ! There is only one country (wink wink) so this column is useless, you can drop it.
#
# **Instruction:**
# - Drop the `county` column
# +
# Start of your code (1 line)
# End of your code
# -
# ## Data processing 1.2
#
# Another problem in data science is extreme values in our data.\
# For example, some houses may have extreme prices, to help our model to train and generalize, it's preferable to drop them.
#
# But how to define a min and max range for our data? Well, a good start would be to print the minimum, maximum and median values in our data.
#
# **Instructions:**
# - Print the minimum price of a house in our data
# - Print the maximum price of a house in our data
# - Print the median price of a house in our data
# - Drop all houses with a price less than $10$k or higher than $2 000$k
#
# **Helps:**
# - [pandas.DataFrame.min](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.min.html)
# - [pandas.DataFrame.max](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.max.html)
# - [pandas.DataFrame.median](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.median.html)
# - [How to drop rows on a conditional expression](https://stackoverflow.com/questions/13851535/how-to-delete-rows-from-a-pandas-dataframe-based-on-a-conditional-expression)
# +
# Show the minimum, maximum an median value of the prices in all our data.
# Start of your code (3 lines)
# End of your code
percentage = sum([1 if p >= 2_000_000 or p == 0 else 0 for p in table["price"]]) / table.shape[0]
print(f"Percentage of price higter than 2 000k: {percentage:.2f}%")
# Drop all houses with a price less than 10k or highter than 2 000k
# Start of your code (2 lines)
# End of your code
print("Number of lefting rows:", table.shape[0])
# -
# ## Data processing 1.3
#
# So, we dropped useless columns, we drop extreme value, what else?
#
# Well, another issue is columns with low information, let's take `city` for example, if a city has only a few houses to sell we do not have enough information to predict well its prices and we want to drop it.
#
# **Instruction:**
# - Drop every city that appears less than 10 times
# +
# Start of your code (~3 lines)
# End of your code
print(table.shape[0])
# -
# Let's take a look at our data after droping all thoses useless informations
table.head()
# ## Data processing 1.4
#
# The penultimate step before building our model, in machine learning and expressly in deep learning we prefer to normalize our data between $0$ and $1$ to facilitate our model training.
#
# For example, in an image, all pixels are between $0$ and $255$, we can so divide each pixel by $255$ to range all the pixels between $0$ and $1$. It's the same here.
#
# **Instruction:**
# - Store the maximum price in our data into a variable named `MAX_PRICE`
# - Normalize `price`, `sqft_living`, `sqft_lot`, `sqft_above`, `sqft_basement` and `yr_built` column between $0$ and $1$.
#
# **Help:**
# - We normalize a column by dividing it by its max value
# +
# Start of your code (~7 lines)
# End of your code
# -
# Let's take a look at our data after normalization:
table.head()
# Another issue in data science is non-numerical values. Our model only handles numerical values so how to handle values that are strings like `city` ?
#
# We encode it into one hot vector !
#
# (Don't worry, we do this step for you, but I hardly recommend you to watch [this video](https://www.youtube.com/watch?v=v_4KWmkwmsU) to understand one hot encoding)
def encode_and_bind(original_dataframe, feature_to_encode):
dummies = pd.get_dummies(original_dataframe[[feature_to_encode]])
res = pd.concat([original_dataframe, dummies], axis=1)
return(res)
table = encode_and_bind(table, "city").drop(["city"], axis=1)
# Let's take a final look at our data:
table.head()
# ## Linear Regression 1.1
#
# One fundamental notion you need to understand in machine learning is labels. A label is the target that our model tries to predict. We always remove the label from our dataset and store it in another storage.
# Giving the label to our model would be like giving the answer (it's cheating).
#
# Another notion you need to understand is the training set and testing/validation set.
#
# The training is used to train and see our model's performance evolution, but we also would like to see the performance of our model on data it has never seen. This is the role of the test set.
#
# **Instructions:**
# - Split our data into two specific set: `X_train` & `X_test` (`X_train` must have 3k rows)
# - Split the labels into an other array `y_train` and `y_test` and remove it from `X_train` a,d `X_test`
#
# **Help:**
# - `my_array[0:1000]` give you the first 1k rows of your array
# +
# Start of your code (~4 lines)
# End of your code
# -
# ### Import
#
# - `sklearn`: It features various classification, regression and clustering algorithms
from sklearn.linear_model import LinearRegression
# ## Linear Regression 1.2
#
# We now want to create our model, we will use a linear regression which is already provided in the `sklearn` library.
#
# Of course in our case, it will not be a linear regression in a 2D plan but in 42 dimensions (because with have 42 variables for each prediction.\
# Hard to imagine right ?
#
# **Instruction:**
# - Create a Linear Regression using `sklearn` and train it using `X_train` and `y_train`
#
# **Helps:**
# - What a linear regression is: https://www.youtube.com/watch?v=zPG4NjIkCjc
# - [sklearn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html) on `LinearRegression`
# +
# Start of your code (~1 line)
# End of your code
# -
# ## Linear Regression 1.3
#
# As you saw, `sklearn` does all the job for us, from creating the model to train it. We just have to provide it data.
#
# So, you created and trained your model, but now it's time to know how it performs!
#
# Display the `score` of our model.\
# Quick reminder: the more it's closer to $1$, the better it is.
#
# **Help:**
# - [sklearn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html#sklearn.linear_model.LinearRegression.score) on `score`
# +
# Start of your code (~1 line)
# End of your code
# -
# Ok, so, you printed the coefficient $R^2$. It's closed to $1$ so you understand it should be good, but let's see in a more readable way the precision of our model.\
# A great way would be to display the average difference between our predictions and our labels.
#
# Let's do that !
#
# **Instruction:**
# - Print the average difference between our prediction and our labels using `X_test` and `y_test`
#
# **Helps:**
# - You don't care if the difference is negative or positive, you want the `abs`olute value (wink, wink)
# - Don't forget to use `MAX_PRICE` to see the real difference in $.
# +
# Start of your code (~1 line)
# End of your code
# -
# ## Deep Learning 1.1
#
# We see that we have an average difference arround $105 000$$, it's good but we could better by changing our model using deep learning.
#
# Deep learning is not soo hard, but it takes time to fully understand its concept and working so we will not ask you to find answer like before, just to read, pay attention and understand basic stuff.
#
# ### Import
#
# - `torch`: open source machine learning library based on the Torch library
# - `torch.nn`: Neural network layers
# - `torch.nn.fuctional`: USefull function to train our model
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
torch.manual_seed(0)
# -
# We will not go soo much into details of how does a deep learning model work but you need to remember few things:
#
# 1. **The forward propagation**: The model takes data and make predictions with it
# 2. **The backward propagation**: The model takes the labels and try to modify itself to increase its prediction
# 3. **The learning rate**: A factor variable slowing down our model training (But why should I slow down my model training ? Well that a good question, you should take a look at the link below )
# 4. **The optimizer**: the algorithm that tries to increase our model's predictions
#
#
# **Helps:**
# - [But what is a Neural Network | Deep learning, chapter 1](https://www.youtube.com/watch?v=aircAruvnKk)
# - [Neural Networks Demystified [Part 2: Forward Propagation]
# ](https://www.youtube.com/watch?v=UJwK6jAStmg)
# - [What is backpropagation really doing? | Deep learning, chapter 3
# ](https://www.youtube.com/watch?v=Ilg3gGewQ5U&t=5s)
# - [Learning Rate in a Neural Network explained
# ](https://www.youtube.com/watch?v=jWT-AX9677k)
# - [Optimizers - EXPLAINED!](https://www.youtube.com/watch?v=mdKjMPmcWjY)
# - [Layers in a Neural Network explained](https://www.youtube.com/watch?v=FK77zZxaBoI)
#
# <br/><br/>
# Now that said let's create our deep learning model. It takes place as a class inheriting from `nn.Module`, if you're not familiar with classes in python you should take a look at [this link](https://docs.python.org/3/tutorial/classes.html).
#
# In the `__init__` method we define our layers, for this step you should use `nn.Sequential`, `nn.Linear` and `nn.Sigmoid`.\
# In the `forward` method, we define the forward pass.
#
# **Instructions:**
# - In `__init__` create a `self.main` attribute composed of two layers separated by the sigmoid function.
# - In `forward` define the forward propagation
#
#
# **Helps:**
# - We have 42 columns
# - We want to predict only one value
# - [Sequential documentation](https://pytorch.org/docs/stable/generated/torch.nn.Sequential.html)
# - [Linear documentation](https://pytorch.org/docs/stable/generated/torch.nn.Linear.html)
# - [Sigmoid documentation](https://pytorch.org/docs/stable/generated/torch.nn.Sigmoid.html)
class Model(nn.Module):
def __init__(self):
super().__init__()
# Start of your code (~5 lines)
# End of your code
def forward(self, t):
# Start of your code (~1 line)
# End of your code
# Now you created our model class, we can init our model by calling `Model()`.
# We can also create our optimizer, we choose to use `Adam`, a popular optimizer, it takes as arguments all our model's parameters and the learning rate that we set to $0.05$.
network = Model()
optimizer = optim.Adam(network.parameters(), lr=0.05)
# ## Deep Learning 1.2
#
# Now that you have a model, it's time to create our training function.
#
# Quick remind: to evaluate the accuracy our model uses a *cost function*
#
# You see that we iterate over each data in our `train_set`, for each data ask your model to make à prediction (`network(data.float())`), we calculate how wrong our prediction is by comparing predictions with labels (`F.mse_loss(predictions.squeeze(1), labels.float())`) and we modify our model to improve our predictions.
#
# Basically, that it's!
#
# **Note:** Don't pay to much attention about why is there à `.float()`, why do we do `torch.tensor(labels)` or `.squeeze(1)`. It's just to make our model able to learn from our data. Of course, if you have any questions, feel free to ask.
#
# **Help:**
# - [Part 1: An Introduction To Understanding Cost Functions](https://www.youtube.com/watch?v=euhATa4wgzo)
def train(network, optimizer, train_set, train_labels):
diverenge = 0
episode_loss = 0
correct_in_episode = 0
network.train()
for index, data in enumerate(train_set):
labels = train_labels[index]
labels = torch.tensor(labels)
predictions = network(data.float())
loss = F.mse_loss(predictions.squeeze(1), labels.float())
optimizer.zero_grad()
loss.backward()
optimizer.step()
episode_loss += loss.item()
diverenge += sum(abs(labels.unsqueeze(1) - predictions))
return episode_loss / (len(train_set) * np.shape(train_set)[1])
# The test fucntion looks like the same as the train function, the only differences are that we don't do the backpropagation and we specify to pytorch that we don't want our model to train (`network.eval()`).
def test(network, optimizer, test_set, test_labels):
diverenge = 0
episode_loss = 0
correct_in_episode = 0
network.eval()
for index, data in enumerate(test_set):
labels = test_labels[index]
labels = torch.tensor(labels)
predictions = network(data.float())
loss = F.mse_loss(predictions.squeeze(1), labels.float())
episode_loss += loss.item()
diverenge += sum(abs(labels.unsqueeze(1) - predictions))
return episode_loss / (len(test_set) * np.shape(test_set)[1])
# ## Deep Learning 1.3
#
# Another notion useful to understand in deep learning is batches. Batches help our model to generalize its prediction, for model detail I invite you to watch the link below:
#
# **Help:**
# - [Batch Size in a Neural Network explained](https://www.youtube.com/watch?v=U4WB9p6ODjM)
def create_batch(data, batch_size=8):
result = []
for i in range(batch_size, data.shape[0], batch_size):
result.append(data[i - batch_size: i])
return result
# We then call our function `create_batch` and use a batch size equal to $32$.
# +
X_train = torch.tensor(create_batch(X_train, batch_size=32))
y_train = torch.tensor(create_batch(y_train, batch_size=32))
X_test = torch.tensor(create_batch(X_test, batch_size=32))
y_test = torch.tensor(create_batch(y_test, batch_size=32))
# -
# It's time to train and test our model!
#
# Like you see, we just call `train` then `test` successively for a number of epoch ?\
# But what an epoch is ? An epoch is one iteration over all our data.
for e in range(0, 17):
train_loss = train(network, optimizer, X_train, y_train)
test_loss = test(network, optimizer, X_test, y_test)
result = network(torch.tensor(X_test).float())
diff = int(sum(sum(abs(result.squeeze(2) - y_test))) * MAX_PRICE / (len(y_test) * y_train.shape[1]))
print(f"Epoch {e}\train loss:{train_loss:.5f}\ttest loss:{test_loss:.5f}\tavg diff:{diff:.5f}")
# Congratulation, you made your first machine learning AND deep learning model !!
#
# For those how ask themself: "*That it's? am I a data scientist?*"
#
# Well, not quite yet. There is a long road and a lot of things to learning in data science/machine learning/deep learning and that exactly why it's so fascinating to work in AI, there are so many things to learn.
#
# I hope you enjoyed this workshop, and one more time: **Congratulation!**
#
# *More workshops made by PoC: [https://github.com/PoCInnovation/Workshops](github.com/PoCInnovation/Workshops)*
| ai/1.5.Real_Estate/realEstate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comprehensions
#
# Comprehensions in Python provide us with a short and concise way to construct new sequences (such as lists, set, dictionary etc.) using sequences which have been already defined. Python supports the following 4 types of comprehensions:
#
# + List Comprehensions
# + Dictionary Comprehensions
# + Set Comprehensions
# + Generator Comprehensions
#
# ## List Comprehensions
# List Comprehensions provide an elegant way to create new lists. The following is the basic structure of a list comprehension:
#
# ```
# output_list = [output_exp for var in input_list if (var satisfies this condition)]
# ```
# +
# WITHOUT Using List comprehensions
input_list = [1, 2, 3, 4, 4, 5, 6, 7, 7]
output_list = []
# Using loop for constructing output list
for var in input_list:
if var % 2 == 0:
output_list.append(var)
print ("Output List using for loop:", output_list)
# +
# Using List comprehensions
input_list = [1, 2, 3, 4, 4, 5, 6, 7, 7]
print ("Output List using for loop:", [var for var in input_list if var % 2 == 0] )
# -
# replace [] with () to get generator object
print ("Output List using for loop:", (var for var in input_list if var % 2 == 0) )
# Another example using List comprehensions
print("Output List using list comprehension:",[var**2 for var in range(1, 10)] )
# ## Dictionary Comprehensions
# Extending the idea of list comprehensions, we can also create a dictionary using dictionary comprehensions. The basic structure of a dictionary comprehension looks like below.
#
# ```
# output_dict = {key:value for (key, value) in iterable if (key, value satisfy this condition)}
# ```
# +
# WITHOUT Using Dictionary comprehensions
input_list = [1, 2, 3, 4, 5, 6, 7]
output_dict = {}
# Using loop for constructing output dictionary
for var in input_list:
if var % 2 != 0:
output_dict[var] = var**3
print("Output Dictionary using for loop:",
output_dict )
# +
# Using Dictionary comprehensions
input_list = [1, 2, 3, 4, 5, 6, 7]
print ("Output Dictionary using dictionary comprehensions:", {var:var ** 3 for var in input_list if var % 2 != 0} )
# -
# Another example using Dictionary comprehensions
state = ['Gujarat', 'Maharashtra', 'Rajasthan']
capital = ['Gandhinagar', 'Mumbai', 'Jaipur']
print("Output Dictionary using dictionary comprehensions:", {key:value for (key, value) in zip(state, capital)})
# ## Set Comprehensions
# Set comprehensions are pretty similar to list comprehensions. The only difference between them is that set comprehensions use curly brackets { }. Let’s look at the following example to understand set comprehensions.
# +
# WITHOUT Using Set comprehensions
input_list = [1, 2, 3, 4, 4, 5, 6, 6, 6, 7, 7]
output_set = set()
# Using loop for constructing output set
for var in input_list:
if var % 2 == 0:
output_set.add(var)
print("Output Set using for loop:", output_set)
# -
# Using Set comprehensions
input_list = [1, 2, 3, 4, 4, 5, 6, 6, 6, 7, 7]
print("Output Set using set comprehensions:",{var for var in input_list if var % 2 == 0})
# ## Generator Comprehensions
# Generator Comprehensions are very similar to list comprehensions. One difference between them is that generator comprehensions use circular brackets whereas list comprehensions use square brackets. The major difference between them is that generators don’t allocate memory for the whole list. Instead, they generate each value one by one which is why they are memory efficient.
#
# +
# Using Generator comprehensions
input_list = [1, 2, 3, 4, 4, 5, 6, 7, 7]
print("Output values using generator comprehensions:", end = ' ')
for var in (var for var in input_list if var % 2 == 0) :
print(var, end = ' ')
| code/10. Comprehensions.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + dotnet_interactive={"language": "fsharp"}
#r "nuget: FSharp.Stats, 0.4.0"
#r "nuget: BioFSharp, 2.0.0-beta5"
#r "nuget: BioFSharp.IO, 2.0.0-beta5"
#r "nuget: Plotly.NET, 2.0.0-beta8"
#r "nuget: BIO-BTE-06-L-7_Aux, 0.0.8"
#r "nuget: Deedle, 2.3.0"
#r "nuget: ISADotNet, 0.2.4"
#r "nuget: ISADotNet.XLSX, 0.2.4"
#r "nuget: Plotly.NET.Interactive, 2.0.0-beta8"
open System.IO
open ISADotNet
open ISADotNet.API
open Deedle
open BioFSharp
open FSharpAux
open FSharp.Stats
open Plotly.NET
open System.IO
open BIO_BTE_06_L_7_Aux.FS3_Aux
open BIO_BTE_06_L_7_Aux.Deedle_Aux
// -
// # NB06c' Label efficiency for SDS
//
// [](https://mybinder.org/v2/gh/CSBiology/BIO-BTE-06-L-7/gh-pages?filepath=NB06c_Label_efficiency_BN.ipynb)
//
// [Download Notebook](https://github.com/CSBiology/BIO-BTE-06-L-7/releases/download/NB06b_NB06b_NB06c_NB06c_NB06d_NB06d/NB06c_Label_efficiency_BN.ipynb)
//
// Stable isotopic peptide labeling is the foundation of QconCAT experiments. While an excellent tool when carried out with correctly, it also exposes
// challenges and pitfalls that have to be checked and possibly accounted for. One of these pitfalls is the efficiency with which we labeled
// our QconCAT protein (Why?). In this notebook we will have a look at some high quality peptides selected in the previous notebook and
// illustrate how the label efficiency can be calculated using simulations.
//
// ## I. Reading the data
// As promised, we start this notebook with the output of the previous analysis, this notebook assumes that the data from *NB06b Data Access and Quality Control* is stored in a .txt
// + dotnet_interactive={"language": "fsharp"}
type PeptideIon =
{|
ProteinGroup : string
Synonyms : string
StringSequence : string
PepSequenceID : int
Charge : int
|}
//This is the filepath you chose in *NB06b Data Access and Quality Control*
// let filePath = @"C:\YourPath\testOut.txt"
let filePath = System.IO.Path.Combine [|__SOURCE_DIRECTORY__; "downloads"; "qualityControlResult_BN.txt"|]
// What is different about this function from the one known from the last notebook?
let qConcatDataFiltered =
Frame.ReadCsv(path = filePath, separators = "\t")
// StringSequence is the peptide sequence
|> Frame.indexRowsUsing (fun os ->
let proteinGroup = os.GetAs<string>("ProteinGroup")
{|
ProteinGroup = os.GetAs<string>("ProteinGroup");
Synonyms = os.GetAs<string>("Synonyms")
StringSequence = os.GetAs<string>("StringSequence");
PepSequenceID = os.GetAs<int>("PepSequenceID");
Charge = os.GetAs<int>("Charge");
|}
)
|> Frame.filterRows (fun k s -> k.ProteinGroup |> String.contains "QProt_newPS")
qConcatDataFiltered.ColumnKeys
|> Array.ofSeq
// -
// First we reuse a proved pattern and define a function to manipulate our frame
// + dotnet_interactive={"language": "fsharp"}
let sliceQuantColumns quantColID frame =
frame
|> Frame.filterCols (fun ck os -> ck |> String.contains ("." + quantColID))
|> Frame.mapColKeys (fun ck -> ck.Split('.') |> Array.item 0)
// -
// Besides already familiar slices...
// + dotnet_interactive={"language": "fsharp"}
let heavy = sliceQuantColumns "Heavy" qConcatDataFiltered
// -
// ... we can also use this function for information needed to reconstruct isotopic patterns.
//
// ## II. Extraction and visualization of measured isotopic envelopes.
// + dotnet_interactive={"language": "fsharp"}
let heavyPatternMz = sliceQuantColumns "heavyPatternMz" qConcatDataFiltered
let heavyPatternI = sliceQuantColumns "heavyPatternI" qConcatDataFiltered
// -
// Now, there's a challenge: The info to reconstruct an isotopic pattern is
// separated into two columns, the x component (heavyPatternMz) and the y component (heavyPatternI).
// As always, this challenged can be solved using a function!
// Hint: Note how we define a function 'floatArrayOf' that specifies how the string is parsed.
// + dotnet_interactive={"language": "fsharp"}
let getHeavyPatternsInFile fileName =
let floatArrayOf s =
if String.isNullOrEmpty s then
[||]
else
s
|> String.split (';')
|> Array.map float
let mz, intensities =
heavyPatternMz
|> Frame.getCol fileName
|> Series.mapValues floatArrayOf,
heavyPatternI
|> Frame.getCol fileName
|> Series.mapValues floatArrayOf
let zipped = Series.zipInner mz intensities
zipped
let extractedPatterns = getHeavyPatternsInFile "20210312BN2_U1"
// -
// Additionally, we can write two functions to plot the patterns of a peptide. When it comes
// to the build the chart (plotIsotopicPattern), things get a little bit trickier, but this is not necessarily your concern. Please inspect the Chart
// created by 'plotIsotopicPatternOf' and write correct descriptions for the x and the y axis. (Fill: |> Chart.withX_AxisStyle "" and |> Chart.withY_AxisStyle "")
// + dotnet_interactive={"language": "fsharp"}
let plotIsotopicPattern color mzsAndintensities =
let min,max =
mzsAndintensities |> Seq.minBy fst |> fst,
mzsAndintensities |> Seq.maxBy fst |> fst
Seq.map (fun (x,y) ->
Chart.Line([x;x],[0.;y], Showlegend = false)
|> Chart.withLineStyle (Width = 7)
) mzsAndintensities
|> Chart.Combine
|> Chart.withMarkerStyle(Size=0,Color = FSharpAux.Colors.toWebColor color)
|> Chart.withX_AxisStyle ("", MinMax = (min - 1., max + 1.))
|> Chart.withY_AxisStyle ""
type ExtractedIsoPattern =
{|
PeptideSequence : PeptideIon
Charge : int
Pattern : seq<(float*float)>
|}
let getIsotopicPattern peptideSequence charge =
let (k,(mzs,intensities)) =
extractedPatterns
|> Series.observations
|> Seq.find (fun (k,(mzs,intensities)) ->
k.StringSequence = peptideSequence && k.Charge = charge
)
{|
PeptideSequence=k
Charge = charge
Pattern = Seq.zip mzs intensities
|}
// + dotnet_interactive={"language": "fsharp"}
let examplePep1 = getIsotopicPattern "DTDILAAFR" 2
// + dotnet_interactive={"language": "fsharp"}
plotIsotopicPattern FSharpAux.Colors.Table.Office.blue examplePep1.Pattern
// + dotnet_interactive={"language": "fsharp"}
let examplePep2 = getIsotopicPattern "LTYYTPDYVVR" 2
// + dotnet_interactive={"language": "fsharp"}
plotIsotopicPattern FSharpAux.Colors.Table.Office.blue examplePep2.Pattern
// -
// ## III. Simulation of isotopic patterns: revisited.
//
// Now that we visualized the patterns of two sample peptides, we will simulate theoretical patterns
// and compare them to the ones we measured! You will recognize a lot of the used code from *NB02c Isotopic distribution*
// Note: we copy the code so you can make yourself familiar with it, of course we could also reference functions defined beforehand.
// + dotnet_interactive={"language": "fsharp"}
// create chemical formula for amino acid and add water to reflect hydrolysed state in mass spectrometer
let toFormula bioseq =
bioseq
|> BioSeq.toFormula
// peptides are hydrolysed in the mass spectrometer, so we add H2O
|> Formula.add Formula.Table.H2O
let label n15LableEfficiency formula =
let heavyN15 = Elements.Di (Elements.createDi "N15" (Isotopes.Table.N15,n15LableEfficiency) (Isotopes.Table.N14,1.-n15LableEfficiency) )
Formula.replaceElement formula Elements.Table.N heavyN15
// Predicts an isotopic distribution of the given formula at the given charge,
// normalized by the sum of probabilities, using the MIDAs algorithm
let generateIsotopicDistribution (charge:int) (f:Formula.Formula) =
IsotopicDistribution.MIDA.ofFormula
IsotopicDistribution.MIDA.normalizeByProbSum
0.01
0.000000001
charge
f
type SimulatedIsoPattern =
{|
PeptideSequence : string
Charge : int
LableEfficiency : float
SimPattern : list<(float*float)>
|}
let simulateFrom peptideSequence charge lableEfficiency =
let simPattern =
peptideSequence
|> BioSeq.ofAminoAcidString
|> toFormula
|> label lableEfficiency
|> generateIsotopicDistribution charge
{|
PeptideSequence = peptideSequence
Charge = charge
LableEfficiency = lableEfficiency
SimPattern = simPattern
|}
// + dotnet_interactive={"language": "fsharp"}
let examplePep2_Sim1 = simulateFrom "LTYYTPDYVVR" 2 0.95
plotIsotopicPattern FSharpAux.Colors.Table.Office.orange examplePep2_Sim1.SimPattern
// + dotnet_interactive={"language": "fsharp"}
let examplePep2_Sim2 = simulateFrom "LTYYTPDYVVR" 2 0.99
plotIsotopicPattern FSharpAux.Colors.Table.Office.orange examplePep2_Sim2.SimPattern
// -
// ## IV. Comparing measured and theoretical isotopic patterns.
//
// As we see, there is a discrepancy between real and simulated patterns, both in peak height and in peak count.
// But before we compare both patterns, we have to take some things into consideration.
// While both patterns are normalized in a way that their intensities
// sum to 1., they were normalized independently from each other. Since it is often not possible to
// extract all peaks of an isotopic pattern from a MS run (e.g. due to measurement inaccuracies), we have to
// write a function which filters the simulated patterns for those peaks present in the experimentally
// measured one. Then we normalize it again and have two spectra that can be compared.
// // How are distributions called that sum up to 1?
// + dotnet_interactive={"language": "fsharp"}
let normBySum (a:seq<float*float>) =
let s = Seq.sumBy snd a
Seq.map (fun (x,y) -> x,y / s) a
let compareIsotopicDistributions (measured:ExtractedIsoPattern) (simulated:SimulatedIsoPattern)=
let patternSim' =
measured.Pattern
|> Seq.map (fun (mz,intensities) ->
mz,
simulated.SimPattern
|> Seq.filter (fun (mzSim,intensitiesSim) -> abs(mzSim-mz) < 0.05 )
|> Seq.sumBy snd
)
|> normBySum
{|
Plot =
[
plotIsotopicPattern FSharpAux.Colors.Table.Office.blue measured.Pattern
plotIsotopicPattern FSharpAux.Colors.Table.Office.orange patternSim'
]
|> Chart.Combine
|}
// + dotnet_interactive={"language": "fsharp"}
let comp1 = compareIsotopicDistributions examplePep2 examplePep2_Sim1
comp1.Plot
// + dotnet_interactive={"language": "fsharp"}
let comp2 = compareIsotopicDistributions examplePep2 examplePep2_Sim2
comp2.Plot
// -
// Comparing both simulations, we see that the simulation with a label efficiency of 0.99 fits the measured spectra better than the simulation with 0.95.
// But since we do not want to find a better fit, but the best fit to our measured pattern, this is no goal that is achievable in a feasable way
// using visual inspections. As a solution we utilize the fact that isotopic patterns can be abstracted as ___ ___ (See: How are distributions called that sum up to 1?) distributions.
// A measure to compare measured and theoretical distributions is the kullback leibler divergence. The following code block extends the function
// 'compareIsotopicDistributions' to compute the KL divergence between the precisely measured distribution p and our approximation
// of p (q) using the mida algorithm.
// + dotnet_interactive={"language": "fsharp"}
/// Calculates the Kullback-Leibler divergence Dkl(p||q) from q (theory, model, description, or approximation of p)
/// to p (the "true" distribution of data, observations, or a ___ ___ precisely measured).
let klDiv (p:seq<float>) (q:seq<float>) =
Seq.fold2 (fun acc p q -> (System.Math.Log(p/q)*p) + acc ) 0. p q
let compareIsotopicDistributions' (measured:ExtractedIsoPattern) (simulated:SimulatedIsoPattern)=
let patternSim' =
measured.Pattern
|> Seq.map (fun (mz,intensities) ->
mz,
simulated.SimPattern
|> Seq.filter (fun (mzSim,intensitiesSim) -> abs(mzSim-mz) < 0.05 )
|> Seq.sumBy snd
)
|> normBySum
let klDiv = klDiv (patternSim' |> Seq.map snd) (measured.Pattern |> Seq.map snd)
{|
KLDiv = klDiv
Plot =
[
plotIsotopicPattern FSharpAux.Colors.Table.Office.blue measured.Pattern
plotIsotopicPattern FSharpAux.Colors.Table.Office.orange patternSim'
]
|> Chart.Combine
|}
// -
// ## V. Determining the lable efficiency: an optimiziation problem.
//
// Using this function we can now visualize the kullback leibler divergence between
// different models and the two peptides we measured. Since the lower the divergence. We will
// also visualize the pattern with the best fit. Please inspect the Chart created by 'Chart.Point(lableEfficiency,comparison |> Seq.map (fun x -> x.KLDiv))'
// and write correct descriptions for the x and the y axis. (Fill: |> Chart.withX_AxisStyle "" and |> Chart.withY_AxisStyle "")
// + dotnet_interactive={"language": "fsharp"}
let lableEfficiency, comparison =
[|0.95 .. 0.001 .. 0.999|]
|> Array.map (fun lableEfficiency ->
let sim = simulateFrom "DTDILAAFR" 2 lableEfficiency
let comp = compareIsotopicDistributions' examplePep1 sim
lableEfficiency,
comp
)
|> Seq.unzip
let bestFit = comparison |> Seq.minBy (fun x -> x.KLDiv)
Chart.Point(lableEfficiency,comparison |> Seq.map (fun x -> x.KLDiv))
|> Chart.withX_AxisStyle ""
|> Chart.withY_AxisStyle ""
// + dotnet_interactive={"language": "fsharp"}
bestFit.Plot
// + dotnet_interactive={"language": "fsharp"}
let lableEfficiency2, comparison2 =
[|0.95 .. 0.001 .. 0.999|]
|> Array.map (fun lableEfficiency ->
let sim = simulateFrom "LTYYTPDYVVR" 2 lableEfficiency
let comp = compareIsotopicDistributions' examplePep2 sim
lableEfficiency,
comp
)
|> Seq.unzip
let bestFit2 = comparison2 |> Seq.minBy (fun x -> x.KLDiv)
Chart.Point(lableEfficiency2,comparison2 |> Seq.map (fun x -> x.KLDiv))
|> Chart.withX_AxisStyle ""
|> Chart.withY_AxisStyle ""
// + dotnet_interactive={"language": "fsharp"}
bestFit2.Plot
// -
// Observing the output, we can make two observations: the function x(lablefficiency) = KL(measured,sim(lableeffciency)) has in both cases a local minimum
// that is similar, yet slightly different for peptides "LTYYTPDYVVR" and "DTDILAAFR", and that the best fit resembles the measured distribution closely, but not
// perfectly, what is the reason for this?
//
// Finding this local minimum will give us the best estimator for the lable efficiency. This can be done using brute force approaches (as we just did)
// or more elaborate optimization techniques. For this we will use an algorithm called 'Brent's method'. This method is more precise and speeds up the calculation time (Why?).
// How close are the estimates?
// + dotnet_interactive={"language": "fsharp"}
let calcKL peptideSequence charge lableEfficiency =
let measured =
getIsotopicPattern peptideSequence charge
let sim = simulateFrom peptideSequence charge lableEfficiency
let comp =
compareIsotopicDistributions' measured sim
comp.KLDiv
let est1 = Optimization.Brent.minimize (calcKL "DTDILAAFR" 2) 0.98 0.999
let est2 = Optimization.Brent.minimize (calcKL "LTYYTPDYVVR" 2) 0.98 0.999
// -
// Since the estimates have a certain level of uncertainty we will repeat the estimation for
// some high intensity peptides and visualize the results. Please fill the x axis description (|> Chart.withX_AxisStyle "")
// + dotnet_interactive={"language": "fsharp"}
let highIntensityPeptides =
heavy
|> Frame.getCol "20210312BN2_U1"
|> Series.sortBy (fun (x:float) -> - x)
|> Series.filter (fun k v -> k.StringSequence |> String.exists (fun x -> x='[') |> not)
let estimates =
highIntensityPeptides
|> Series.take 20
|> Series.map (fun k v ->
FSharp.Stats.Optimization.Brent.minimize (calcKL k.StringSequence k.Charge) 0.98 0.999
)
|> Series.values
|> Seq.choose id
// + dotnet_interactive={"language": "fsharp"}
Chart.BoxPlot estimates
|> Chart.withX_AxisStyle ""
// -
// Now that we know more than an educated guess of an lable efficiency estimate we can start with our main goal:
// the absolute quantification of chlamydomonas proteins!
| Notebooks/NB06c_Label_efficiency_BN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <ins>Tutorial 2.1: Conditionals & Loops</ins>
# *ASTR 211: Observational Astronomy, Spring 2021* \
# *Written by <NAME>*
# Conditionals and loops are fundamental structures in writing code that can automate things for you and adapt to various inputs. There aren't many scripts I've written since learning the basics that haven't had at least one loop or conditional statement.
# Importing libraries *at the beginning of the file*!
import numpy as np
# # Conditionals
#
# _Conditional statements_ are structures in a program that perform different tasks depending on a set condition you give it. These conditions are just **booleans**, which we talked about in the last session. Recall that booleans are just True/False statements.
#
# The easiest way to see how these work is to watch them in action. For example, let's say I have a user input their favorite number:
num = int(input("Enter your favorite number: "))
# If we wanted to tell whether or not the number that they've entered was positive or negative, we can use _if-else statements_ (i.e. a conditional statements):
# +
if (num > 0):
print("Number is positive.")
if (num < 0):
print("Number is negative.")
# -
# Now, there are a few things to notice here.
#
# 1. **The fact that only one statement is printed.** When Python comes across a line that reads `if <condition>:`, it checks the condition first and then decides what to do next. If the condition is met, of course, it keeps going. If not, it skips the entirety of the code in the conditional statement and moves on to what comes after. So when the above code is run, the number can't be simultaneously greater-than and less-than zero, and just one print statement is executed.
#
#
# 2. **The boolean in the conditional _is a boolean_.** If you were to try to write `if 12: <do something>`, you'd get an error. You could write `if (12 == 12): <do something>`, and that would be perfectly valid, but `12` alone doesn't have an inherent truth or falsity.
#
#
# 3. **Indentation/whitespace.** When using a structure like an if-else statement, _whitespace_ (empty space in the code) is important. It tells the program where to look for it's next task. If there was no indentation, Python would assume that the next line of code was separate from the conditional statement, and run the code no matter whether the condition was met or not. Everything indented under the `if` statement is what we call _nested_ under it, and thus will only run if the boolean condition is met.
#
# However, the main thing to notice is that, if you run that code and enter a number, it works! Nice. You may be wondering where the "else" in "if-else statement" comes from, however. That's because the conventional way to write the code above is as follows:
if (num > 0):
print("Number is positive.")
else:
print("Number is negative.")
# It gives you the same result with a bit of a cleaner look, since you don't have to write the alternative condition in the case of two. It basically reads "if `<condition>` do `<something>`; if literally anything else is the case, `<do this instead>`."
#
# We've missed one key point in how we decided to design this program, however. Sure, the number could be positive or negative, but what if the number they give is _zero_? We'll have to add an extra condition into the code, which we can achieve with an "if-elif-else" statement:
if (num > 0):
print("Number is positive.")
elif (num < 0):
print("Number is negative.")
else:
print("Number is zero.")
# The `elif` line just lets us define an additional condition to check. You can add as many of these as you want, for any condition that you don't necessarily want to fall under the "literally-anything-else" category.
#
# Let's think about what we might do if we want to check another condition depending on whether the first is met. For example, if I wanted the code to tell me not only the parity of the number, but also whether it was divisible by 3, I would need to add another condition. One way to do this is just to add more ifs to my statement and include a new boolean in the condition for each:
if (num > 0) and (num % 3 == 0):
print("Number is positive and divisible by 3.")
elif (num > 0) and (num % 3 != 0):
print("Number is positive and not divisible by 3.")
elif (num < 0) and (num % 3 == 0):
print("Number is negative and divisible by 3.")
elif (num < 0) and (num % 3 != 0):
print("Number is negative and not divisible by 3.")
else:
print("Number is zero.")
# Alternatively, you could use _nested_ if-statements:
# +
if (num > 0):
print("Number is positive...")
if (num % 3 == 0):
print("...and divisible by 3.")
if (num % 3 != 0):
print("...and not divisible by 3.")
elif (num < 0):
print("Number is positive...")
if (num % 3 == 0):
print("...and divisible by 3.")
if (num % 3 != 0):
print("...and not divisible by 3.")
else:
print("Number is zero.")
# -
# (**Note:** I stressed being careful with the whitespace in structures like if statements, but notice that I added some vertical spacing to my code. This doesn't affect how the code runs at all -- Python ignores blank lines. Sometimes, this is helpful to declutter your code and separate different operations.)
#
# Both of these methods produce essentially the same result. Just to put the if statement in context for when you might use it, let's look at the same scenario, this time using variables. (In astrophysics, you're more likely to be using these statements to perform calculations or modify lists based on certain conditions, rather than printing things.)
# +
if (num > 0):
parity = 'positive'
if (num % 3 == 0):
divby3 = True
if (num % 3 != 0):
divby3 = False
elif (num < 0):
parity = 'negative'
if (num % 3 == 0):
divby3 = True
if (num % 3 != 0):
divby3 = False
else: parity = 'zero'
print("Parity:", parity)
print("Divisible by 3:", divby3)
# -
# ### Fancy formatting tricks
#
# There are a couple of tricks with these statements that you might like to use to clean up your code. Before you use these, _make sure you understand how the normal formatting works,_ as these will not always be convenient! You need the intuition to know when/when not to use these.
#
# 1. **If statements with just one line nested in them can be written in one line**. With this in mind, we can turn the above piece of code into something with 4 fewer lines:
# +
if (num > 0):
parity = 'positive'
if (num % 3 == 0): divby3 = True
if (num % 3 != 0): divby3 = False
elif (num < 0):
parity = 'negative'
if (num % 3 == 0): divby3 = True
if (num % 3 != 0): divby3 = False
else: parity = 'zero'
print("Parity:", parity)
print("Divisible by 3:", divby3)
# -
# 2. **If-else statements with just one nested line in both branches can be written in one line.** The formatting on this one is a bit different, with the syntax being `<action> if <condition> else <other action>`. Using this trick, where both actions are variable definitions, we can remove two more lines from our code:
# +
if (num > 0):
parity = 'positive'
divby3 = True if (num % 3 == 0) else False
elif (num < 0):
parity = 'negative'
divby3 = True if (num % 3 == 0) else False
else: parity = 'zero'
print("Parity:", parity)
print("Divisible by 3:", divby3)
# -
# One more thing: Beacuse sometimes a user may enter some input you don't want, or if you have a condition that ends the code when met, you can use the `quit()` function. I can't demonstrate in Jupyter because it'll throw an error, but if you use it in a normal Python script, the entire program will end if the `quit()` function appears.
# # Loops
#
# Loops are just what they sound like, in that they repeat portions of code over and over again. You'll have to specify the condition that must be met in order for the loop to stop, otherwise your code will run forever (a _runtime error,_ if you'll recall from the last tutorial).
#
# There are two kinds of loops: _while loops_ and _for loops._ We'll discuss both in detail.
#
# ## While loops
#
# In the case of a while loop, the code inside the loop repeats _while_ the condition remains true. The syntax looks like `while <condition>: <code>`. Python checks to make sure that this condition is still true each time it runs the code -- if it is, it runs the code again; if not, it exits the loop and moves on.
#
# For example, if I wanted (for some reason) to count to 10 with a while loop, I could do something like this:
# +
count = 0
while (count < 10):
count = count + 1
print(count)
# -
# The main thing to notice here is that **the loop condition is not static.** In these kinds of loops, a _counter_ is usually employed, which is just a variable that will (eventually) change the value of the condition from True to False, allowing the code to exit the loop. In this case, our counter is `count`, and the loop condition depends on it. Each time the loop goes, it adds 1 to the value of `count`, until it reaches a value of 10 and makes the loop condition false.
#
# Something else you might notice is that a variable is being added to itself. This may look odd, but it's totally valid. We're taking the previous value (on the right), adding one, and _re-assigning the variable_ (on the right; see Tutorial 1.1) with this new value so it has the same name. There's a nice way to write this kind of incrementation, with the `+=` operator:
# +
count = 0
while (count < 10):
count += 1
print(count)
# -
# This does the same thing as `count = count + 1`. If we were subtracting or multiplying or dividing, we might like to use `-=` or `*=` or `/=` as well.
#
# One thing to note about while loops is the fact that you can sometimes get stuck in them, as I've mentioned in other tutorials. If your loop condition is `1==1`, that's never going to stop being true, and your code will loop until the end of time if you let it. If your code is taking an unusually long time to run, this is often why -- in order to exit an infinite loop (or stop your code mid-run for any reason at all), hit `Ctrl+C` while in the terminal (or just hit the "stop" button in the toolbar if its a Jupyter cell).
#
# ## For loops
#
# In my experience, _for loops_ are much more useful for the astrophysical applications I've been exposed to. In contrast to while loops, _for loops_ loop through a set of values rather than relying on a truth condition. Once it's seen all the values, the loop ends. This is where lists and arrays become important; the syntax looks like `for <item> in <list/array>: <code>`.
#
# Starting simply, let's just say we want to print every element of a list:
# +
lst = np.arange(1,11)
for x in lst:
print(x)
# -
# It's worth reiterating that `x` is a variable _local to the for loop,_ a temporary variable that holds the value of the list item that the loop is currently on. For example, if my list was `[1,2,3]`, `x` would be equal to 1 for the first round of the loop, 2 for the second, 3 for the third, and then the loop would end and `x` would cease to exist. Also, "x" is just the name I chose -- just like any variable, you're free to choose how you represent it.
#
# Now, let's try subtracting 1 from every element in our list. Our original list (`lst`) should read `[1,2,3,4,5,6,7,8,9,10]`. Just to make sure, I'll print it:
print(np.arange(1,11))
# Checks out. Now, I'll edit our code to subtract 1 from each value in the list rather than printing. Once the loop runs, it should read `[0,1,2,3,4,5,6,7,8,9]`:
# +
for x in lst:
x -= 1
print(lst)
# -
# Not quite. This demonstrates the fact that we're not changing the list elements when we use them this way in a for loop. We just have access to the values, which we can use for calculations, etc. In order to edit the list, you'd have to use some of the techniques described in Tutorial 1.2. However, there is an easy way to get the index of each value in the list as you move through the loop, using the `enumerate()` function:
for n,x in enumerate(lst):
print(n,x)
# As you can see, `n` gives us the index of an element, while `x` gives us the value. (Again, "n" and "x" can be called whatever you want.) The enumerate function returns these two values _in this order_, so be sure not to mix them up!
#
# As mentioned above, this is useful for editing the list itself, using the index. Let's try again to subtract 1 from each element of `lst`, this time taking advantage of `enumerate()`:
# +
for n,x in enumerate(lst):
lst[n] = x - 1
print(lst)
# -
# Success!
#
# One more thing to note is that you don't necessarily need to have a list set aside for something like this. For example, you can use the `range()` function to use a for loop over a range of numbers from 0 to whatever:
for x in range(10):
print(x)
# Alternatively, you can just stick a list after `in`:
# +
for x in [1,2,3,4,5]:
print(x)
for x in np.linspace(1,5,5):
print(x)
# -
# For multiple lists of the same size, you can access values in each with the same index using `zip()`. In other words, `zip()` takes the first element of each list as loop variables at one time, then the second element of each, so on and so forth. For example, say I have lists of wavelengths and their corresponding frequencies, and I'd like to print both at once:
# +
wav = np.linspace(300,700,10)
freq = np.linspace(10,50,10)
for x,y in zip(wav,freq):
print(x,y)
# -
# You can also nest for loops in one another, just like we did for if statements. In fact, you can nest any if statement, while loop or for loop inside of any other. Here's a random demonstration:
# +
summation = 0
for x in range(3):
for y in range(3):
if (x == y):
summation += (x + y)
print(summation)
# -
# ### Keywords
#
# There are a few special commands that can be helpful in conditionals and loops.
#
# 1. **`break`**: If a `break` line is run inside a loop, Python will exit the loop, skipping any remaining iterations and continue with the rest of the code. For example, if I were to sum the values of the list `[1,2,3,4,5]`, I should get 15. However, if I add the condition that the loop _breaks_ once the value 3 has been reached, it should be 6:
# +
summation = 0
for x in [1,2,3,4,5]:
summation += x
if x == 3:
break
print(summation)
# -
# **Note:** This is a good time to mention that you should _always check the order of your steps in loops,_ especially in cases like this. If I had placed my if statement with the `break` before the step where `x` was added to the sum, the result would have been 3 instead of 6 because it left the loop early:
# +
summation = 0
for x in [1,2,3,4,5]:
if x == 3:
break
summation += x
print(summation)
# -
# 2. **`continue`**: If a `continue` line is reached inside a loop, the current step is skipped. Going back to the `[1,2,3,4,5]` example, if we sum this list again and include a `continue` conditional on the third element, the sum should be 1+2+4+5 = 12:
# +
summation = 0
for x in [1,2,3,4,5]:
if x == 3:
continue
summation += x
print(summation)
# -
# 2. **`pass`**: If a `pass` line is reached inside a loop or conditional, nothing happens. (I use this a lot as a placeholder for code I need to finish later.) If we run the same loop with a pass conditional, it should just give the normal sum of 15:
# +
summation = 0
for x in [1,2,3,4,5]:
if x == 3:
pass
summation += x
print(summation)
# -
# ### List comprehension
#
# _Don't use this until you understand loops (and lists) well!_
#
# There's really only one good trick that I use a lot off the top of my head, something called **list comprehension**. Essentially, you can generate a list from a little for loop inside some brackets. For example, say I have two lists with some overlap between them. If I wanted to make sure that the elements of each list are unique (i.e. one list does not have any of the same elements as the other), I could use list comprehension to filter one list:
# +
lst1 = [1,2,3,4,5,6,7]
lst2 = [1,1,4,3,5,6,9,0,8,8,9,1,2]
lst2 = [x for x in lst2 if x not in lst1]
print(lst2)
# -
# **Note:** Something I probably should've mentioned in the lists/arrays tutorial is that `in` and `not in` are logical operators like `and`/`or` that can be used on lists. You can see the demonstration for `not in` in this list comprehension example; same goes for `in`.
#
# Omitting the if-statement part of the syntax, I can just create a list:
# +
lst3 = [x for x in range(4)]
print(lst3)
# -
| files/astr211_tut2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''Create a program that asks the user to enter their name and
their age. Print out a message addressed to them that tells them
the year that they will turn 100 years old.'''
import datetime
def one_hundred():
name = input('What is your name? ')
age = input('How old are you? ')
age = int(age)
now = datetime.datetime.now()
print(f'{name}, you will turn 100 in the year {now.year - age + 100}.')
one_hundred()
# +
'''Ask the user for a number. Depending on whether the number is even
or odd, print out an appropriate message to the user. Hint: how does
an even / odd number react differently when divided by 2?
If the number is a multiple of 4, print out a different message.'''
def even_or_odd():
number = input('Give me a number: ')
number1 = str(int(number) / 4)
number2 = int(number) % 2
if number1.endswith('.0'):
print('This number is divisible by 4! Yay!')
elif number2 == 0:
print('This is an even number!')
else:
print('This is an odd number!')
even_or_odd()
# +
'''Ask the user for two numbers: one number to check (call it num)
and one number to divide by (check). If check divides evenly into
num, tell that to the user. If not, print a different appropriate
message.'''
def divisible():
num = input('What number are we dividing? ')
check = input('What are we dividing it by? ')
divide = str(int(num) / int(check))
if divide.endswith('.0'):
print(f'{check} divides evenly into {num}!')
else:
print(f'{check} does not divide evenly into {num}.')
divisible()
# +
'''Take a list, say for example this one: and write a program that
prints out all the elements of the list that are less than 5.'''
a = [1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89]
def less_than_five(list):
final = []
for item in list:
if int(item) < 5:
final.append(item)
return final
less_than_five(a)
# +
'''Create a program that asks the user for a number and then prints
out a list of all the divisors of that number. (If you don’t know what
a divisor is, it is a number that divides evenly into another number.
For example, 13 is a divisor of 26 because 26 / 13 has no remainder.)'''
def all_divisors():
number = input('Give me a number: ')
number = int(number)
divisors = []
n = 1
while n < number:
divide = str(int(number) / int(n))
n = n + 1
if divide.endswith('.0'):
divisors.append(float(divide))
return divisors
all_divisors()
# +
'''Take two lists, say for example these two: and write a program that
returns a list that contains only the elements that are common between
the lists (without duplicates). Make sure your program works on two
lists of different sizes.'''
a = [1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89]
b = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13]
def common_elements(list1, list2):
common = []
for item in list1:
if item in(list2):
common.append(item)
return set(common)
common_elements(a, b)
# -
| Daily/Practice Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch
# language: python
# name: pytorch
# ---
# %reload_ext autoreload
# %autoreload 2
import torch
from fastai.vision.all import *
# %matplotlib inline
import sys
sys.path.append('/home/Public/Desktop/wjt/disentanglement_lib/')
# %env DISENTANGLEMENT_LIB_DATA=/data/disentanglement
import wandb
api = wandb.Api()
from disentanglement_lib.visualize import visualize_model
from disentanglement_lib.evaluation.metrics import mig,factor_vae,dci,modularity_explicitness,sap_score,unified_scores
from disentanglement_lib.data.ground_truth import dsprites
from disentanglement_lib.utils.hub import *
data=dsprites.DSprites([1,2])
plt.imshow(data[17][0][0])
r = api.run("erow/dlib/13j2udpx")
r.url
model = get_model('tmp',device='cpu')
encoder,decoder = convert_model(model)
def sigmoid(x):
return 1 / (1 + np.exp(-x))
mu = torch.randn(10,10)
recon=sigmoid(decoder(mu)).squeeze(3)
3*64*64
F.binary_cross_entropy(torch.Tensor(recon[0]),torch.Tensor(recon[2]))*64*64
# from disentanglement_lib.visualize.visualize_util import *
mu = torch.randn(1,10)
plt_sample_traversal(mu,lambda x:sigmoid(decoder(x)).squeeze(3),8,range(7));
z_sampled = torch.zeros(1,10)
z_fake = torch.randn_like(z_sampled,requires_grad=True)
model.load_state_dict(torch.load("tmp/model.pt"))
pos_recons = model.decode(z_sampled)
neg_recons = model.decode(z_fake)
fig,axes=plt.subplots(1,2)
axes[0].imshow(pos_recons[0,0].sigmoid().data)
axes[1].imshow(neg_recons[0,0].sigmoid().data)
tp=(tp.data>0.5).float().clamp(1e-4,1-1e-4)
pp = pos_recons.sigmoid().clamp(1e-4,1-1e-4)
np = neg_recons.sigmoid().clamp(1e-4,1-1e-4).data
plt.imshow((pp*np)[0,0].data,vmax=1,vmin=0)
(tp+pp-pp*tp).sum()
(pp*np).sum()
from disentanglement_lib.methods.shared import losses
model.load_state_dict(torch.load("tmp/model.pt"))
opt=torch.optim.Adam(model.parameters(),1e-4)
for i in range(50):
pos_recons = model.decode(z_sampled)
# neg_recons = model.decode(z_fake)
pp = pos_recons.sigmoid().clamp(1e-4,1-1e-4)
# np = neg_recons.sigmoid().clamp(1e-4,1-1e-4).data
inter = (pp*np).float()
gap = (F.binary_cross_entropy(pp,1-np.data,reduction='none')*inter).sum()
opt.zero_grad()
gap.backward()
opt.step()
if i%10==0:
print(gap.item())
fig,axes=plt.subplots(1,2)
axes[0].imshow(pos_recons[0,0].sigmoid().data)
axes[1].imshow(neg_recons[0,0].sigmoid().data)
tp = pos_recons.clamp(1e-4,1-1e-4)
pp = pos_recons.sigmoid().clamp(1e-4,1-1e-4)
np = neg_recons.sigmoid().clamp(1e-4,1-1e-4)
plt.imshow((inter_tpn).data[0,0])
F.binary_cross_entropy(pp,np.data,reduction='none').sum(),
# +
inter_tpn = tp * pp * np
inter_pt = tp * pp
inter_pn = np * pp
# pos_recon_loss = -torch.log(inter_pt).sum([1,2,3]).mean()
neg_recon_loss = -torch.log(1 - (inter_pn-inter_tpn)).sum([1,2,3]).mean()
pos_recon_loss,neg_recon_loss
# -
intersection = pos_recons.data.sigmoid() * neg_recons.sigmoid()
plt.imshow(intersection[0,0].data)
visualize_util.grid_save_images([pics], 'beta_traversal.png')
mi=np.array(r.summary['discrete_mi'])
import seaborn as sns
sns.heatmap(mi,
annot=True, fmt='.2f',
# yticklabels=[0,0.1,0.3,0.5,1],
xticklabels=['shape','scale','orientation','posX','posY'])
for i in range(means.shape[1]):
pics = activation(
latent_traversal_1d_multi_dim(_decoder, means[i, :], None))
file_name = os.path.join(results_dir, "traversals{}.jpg".format(i))
visualize_model.visualize_reconstructions(output_dir,dataset,recon_fn)
visualize_model.visualize_samples(output_dir,num_latent,_decoder)
# + jupyter={"outputs_hidden": true}
visualize_model.visualize_traversal(output_dir,dataset,_encoder,_decoder)
# + jupyter={"outputs_hidden": true}
visualize_model.visualize_intervention(output_dir, dataset, _encoder)
# -
from disentanglement_lib.visualize import visualize_scores
# + jupyter={"outputs_hidden": true}
dataset.factor_names = ['shape','scale','orientation','posX','posY']
ans=unified_scores.compute_unified_scores(dataset,representation_fn,
np.random.RandomState(), 'visualization', 10000,
100, matrix_fns=[unified_scores.mutual_information_matrix])
ans
# -
runs = api.runs("erow/fractionVAE",{"$and":[{"config.dataset":"dsprites_full"},
{"config.base":"80,30,12"},
{"config.beta":3},
{"state":"finished"}]})
len(runs)
# + jupyter={"outputs_hidden": true}
for file in r.files():
if file.name.startswith('model-'):
print(file)
ans=defaultdict(list)
representation_fn,model = get_representation_fn(r,r.config['img_size'],device='cpu',model_file=file.name)
def _decoder(latent_vectors):
with torch.no_grad():
torch_imgs = model.decoder(torch.Tensor(latent_vectors)).numpy()
return torch_imgs.transpose((0, 2, 3, 1))
def _encoder(obs):
with torch.no_grad():
obs = torch.Tensor(obs.transpose((0, 3, 1, 2))) # convert tf format to torch's
mu, logvar = model.encoder(obs)
mu, logvar = mu.numpy(), logvar.numpy()
return mu, logvar
visualize_model.visualize_intervention(output_dir, dataset, _encoder)
name = file.name
os.system(f'cp visualization/interventional_effects/interventional_effect.png visualization/interventional_effects/{name}.png')
# -
name = "m1"
# !cp visualization/interventional_effects/interventional_effect.png visualization/interventional_effects/$name.png
# %matplotlib inline
for name in ['color_dsprites','scream_dsprites','smallnorb']:
dataset = get_named_ground_truth_data(name)
for i,n in enumerate(dataset.factors_num_values):
num = min(n,6)
factors = np.zeros([num,dataset.num_factors])
factors[:,i] = np.linspace(0,n-1,num)
images = dataset.sample_observations_from_factors(factors,np.random.RandomState())
fig,axes=plt.subplots(1,num,figsize=(num,1),dpi=300)
for img,ax in zip(images,axes):
ax.imshow(img, cmap='gray')
ax.axis('off')
plt.subplots_adjust(wspace=0.05, hspace=0.05)
fig.savefig(f'datasets/{name}_{i}.png',bbox_inches='tight',pad_inches=0)
# break
fig
dataset.factors_num_values
np.linspace(0.5,1,6)
| notebooks/visualize_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Python Object-Oriented Programming (OOP)
#
# _<NAME>_
# ## Dziedziczenie (ang. *Inheritance*)
#
# 
# ### Dziedziczenie
#
# Dziedziczenie umożliwia tworzenie nowych klas, które przejmują (dziedziczą) formę i funkcjonalność klas bazowych. I tak jak dziedzic majątku może nim rozporządzać, np. doprowadzić do ruiny, tak klasy pochodne (dziedziczące) mogą rozszerzać i ulepszać funkcjonalność klas przodków.
#
# Dziedziczenie definiowane jest za pomocą składni:
# +
class KlasaBazowa:
pass
class KlasaPochodna(KlasaBazowa):
pass
# -
# Przykład (na razie bez rozszerzania klasy bazowej `KontoBankowe`):
# +
class KontoBankowe:
def __init__(self, nazwa, stan=0):
self.nazwa = nazwa
self.stan = stan
def info(self):
print("nazwa:", self.nazwa)
print("stan:", self.stan)
def wyplac(self, ilosc):
self.stan -= ilosc
def wplac(self, ilosc):
self.stan += ilosc
class KontoDebetowe(KontoBankowe):
pass
# -
# > ##### Zadanie 1
# > Sprawdź czy cały interfejs klasy bazowej `KontoBankowe` znajduje się i działa w instancji klasy pochodnej `KontoDebetowe`.
# Rozszerzenie zachowania klasy `KontoBankowe`:
class KontoDebetowe(KontoBankowe):
def __init__(self, nazwa, stan=0, limit=0):
KontoBankowe.__init__(self, nazwa, stan)
self.limit = limit
def wyplac(self, ilosc):
"""Jeżeli stan konta po operacji przekroczyłby limit, przerwij."""
if (self.stan - ilosc) < (-self.limit):
print("Brak srodkow na koncie")
else:
KontoBankowe.wyplac(self, ilosc)
# #### Super
#
# Aby wywołać metodę klasy bazowej, zamiast wpisywać długie wyrażenie `NazwaKlasyBazowej` można użyć metody `super()` zwracającej klasę rodzica. Jest to szczególnie przydatne jeśli zmienimy nazwę klasy bazowej, nie trzeba będzie wtedy wprowadzać zmian w klasach pochodnych. Przykład z kontem bankowym:
# bylo:
def __init__(self, nazwa, stan=0, limit=0):
KontoBankowe.__init__(self, nazwa, stan)
self.limit = limit
# jest:
def __init__(self, nazwa, stan=0, limit=0):
super().__init__(nazwa, stan)
self.limit = limit
# #### Zadanie utrwalające
#
# Stwórz klasę bazową dla figur geometrycznych:
# +
import math
class Figura:
def obwod(self):
"""Obliczanie obwodu."""
raise NotImplementedError
def pole(self):
"""Obliczanie pola powierzchni."""
raise NotImplementedError
# -
# Następnie zaimplementuj klasy pochodne dla następujących figur:
#
# 1. koło
# 1. trójkąt równoboczny
# 1. prostokąt
# 1. kwadrat
# 1. równoległobok
# 1. trapez prostokątny
#
# Pamiętaj o zachowaniu odpowiedniej hierarchii (dziedziczenia) oraz inicjalizacji atrybutów (np. wysokości, promienia czy długości boku) w metodzie `__init__`.
# 1. koło
class Kolo(Figura):
def __init__(self, r):
self.r = r
def obwod(self):
return 2 * math.pi * self.r
def pole(self):
return math.pi * self.r ** 2
f1 = Kolo(5)
f1.obwod()
f1.pole()
# 2. trójkąt równoboczny
class TrojkatRownoboczny(Figura):
def __init__(self, a):
self.a = a
self.h = 0.5 * a * math.sqrt(3)
def obwod(self):
return 3 * self.a
def pole(self):
return 0.5 * self.a * self.h
f2 = TrojkatRownoboczny(5)
f2.obwod()
f2.pole()
# 3. prostokąt
class Prostokat(Figura):
def __init__(self, a, b):
self.a = a
self.b = b
def obwod(self):
return 2 * (self.a + self.b)
def pole(self):
return self.a * self.b
f3 = Prostokat(2, 5)
f3.obwod()
f3.pole()
# 4. kwadrat
class Kwadrat(Prostokat):
def __init__(self, a):
self.a = a
# trik, dzięki któremu możemy dziedziczyć wprost
# z prostokąta i nie musimy wypełniać metod `obwod` i `pole`
self.b = a
f4 = Kwadrat(5)
f4.obwod()
f4.pole()
# 5. równoległobok
class Rownoleglobok(Figura):
def __init__(self, a, b, h):
# inny sposób przypisywania zmiennych
# trochę skraca ilość linii
self.a, self.b, self.h = a, b, h
def obwod(self):
return 2 * (self.a + self.b)
def pole(self):
return 0.5 * self.a * self.h
f5 = Rownoleglobok(2, 4, 3)
f5.obwod()
f5.pole()
# 6. trapez prostokątny
class TrapezProstokatny(Figura):
def __init__(self, a, b, h):
self.a, self.b, self.h = a, b, h
# czwarty bok (self.c) obliczamy z Pitagorasa
d = b - a
self.c = (h ** 2 + d ** 2) ** 0.5
def obwod(self):
return sum([self.a, self.b, self.c, self.h])
def pole(self):
return 0.5 * (self.a + self.b) * self.h
f6 = TrapezProstokatny(2, 4, 3)
f6.obwod()
f6.pole()
# #### Wielokrotne dziedziczenie
#
# Trochę jak w prawdziwym życiu w rodzinie, klasa pochodna może mieć więcej niż jednego przodka i od każdego przodka zbierać atrybuty i metody.
#
# 
#
# Przykład klasy z dwoma rodzicami:
# +
class A:
"""<NAME>"""
def __init__(self):
super().__init__()
self.a = "A"
def fa(self):
print("a:", self.a)
class B:
"""<NAME>"""
def __init__(self):
super().__init__()
self.b = "B"
def fb(self):
print("b:", self.b)
class Pochodna(B, A):
"""Dziecko"""
def __init__(self):
super().__init__()
# -
# Działanie:
d = Pochodna()
d.a
d.b
d.fa()
d.fb()
# Jak widać klasa `Pochodna` zawiera pola i metody od każdego z rodziców.
# > ##### Zadanie 3
# > W klasach `A` i `B` zmień nazwy metod `fa()` i `fb()` na `f()`. Sprawdź jak zachowa się teraz wywołanie `d.f()`, gdzie `d` jest instancją klasy pochodnej.\
# > Jak na to zachowanie wpływa zmiana kolejności rodziców przy definicji klasy pochodnej `class Pochodna(tutaj_kolejnosc)`?
# ### Ćwiczenia
# > ##### Ćwiczenie: Utwórz podrzędną klasę `Bus`, która odziedziczy wszystkie zmienne i metody klasy `Vehicle`
# > Utwórz obiekt `Bus`, który odziedziczy wszystkie zmienne i metody klasy `Vehicle` i wyświetli je.\
# > Dane wejściowe:
class Vehicle:
def __init__(self, name, max_speed, mileage):
self.name = name
self.max_speed = max_speed
self.mileage = mileage
# > Oczekiwany wynik:\
# > `Nazwa pojazdu: Szkolne Volvo Prędkość: 180 Przebieg: 12`
# +
class Vehicle:
def __init__(self, name, max_speed, mileage):
self.name = name
self.max_speed = max_speed
self.mileage = mileage
class Bus(Vehicle):
pass
# -
School_bus = Bus("Szkolne Volvo", 180, 12)
print("Nazwa pojazdu:", School_bus.name, "Prędkość:", School_bus.max_speed, "Przebieg:", School_bus.mileage)
# > ##### Ćwiczenie: Dziedziczenie klas
# > Utwórz klasę `Bus`, która dziedziczy po klasie `Vehicle`. Podaj argument pojemności w metodzie `Bus.seating_capacity()` o domyślnej wartości `50`.\
# > Dane wejściowe:
# > Użyj poniższego kodu dla swojej nadrzędnej klasy `Vehicle`. Musisz użyć przesłaniania metody.
class Vehicle:
def __init__(self, name, max_speed, mileage):
self.name = name
self.max_speed = max_speed
self.mileage = mileage
def seating_capacity(self, capacity):
return f"Liczba miejsc siedzących w {self.name} to {capacity} pasażerów"
# > Oczekiwany wynik:\
# > `Liczba miejsc siedzących w Szkolne Volvo to 50 pasażerów`
# +
class Vehicle:
def __init__(self, name, max_speed, mileage):
self.name = name
self.max_speed = max_speed
self.mileage = mileage
def seating_capacity(self, capacity):
return f"Liczba miejsc siedzących w {self.name} to {capacity} pasażerów"
class Bus(Vehicle):
def seating_capacity(self, capacity=50):
return super().seating_capacity(capacity=50)
# -
School_bus = Bus("<NAME>", 180, 12)
print(School_bus.seating_capacity())
| Python/.ipynb_checkpoints/03 Python OOP - dziedziczenie-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# ### Range of axes
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
N = 70
trace1 = go.Mesh3d(x=(70*np.random.randn(N)),
y=(55*np.random.randn(N)),
z=(40*np.random.randn(N)),
opacity=0.5,
color='rgba(244,22,100,0.6)'
)
layout = go.Layout(
scene = dict(
xaxis = dict(
nticks=4, range = [-100,100],),
yaxis = dict(
nticks=4, range = [-50,100],),
zaxis = dict(
nticks=4, range = [-100,100],),),
width=700,
margin=dict(
r=20, l=10,
b=10, t=10)
)
fig = go.Figure(data=[trace1], layout=layout)
py.iplot(fig, filename='3d-axis-range')
# -
# ### Fixed Ratio Axes
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.tools as tls
import numpy as np
N = 50
fig = tls.make_subplots(
rows=2, cols=2,
specs=[
[{'is_3d': True}, {'is_3d': True}],
[{'is_3d': True}, {'is_3d': True}]
],
print_grid=False
)
for i in [1,2]:
for j in [1,2]:
fig.append_trace(
go.Mesh3d(
x=(60*np.random.randn(N)),
y=(25*np.random.randn(N)),
z=(40*np.random.randn(N)),
opacity=0.5,
),
row=i, col=j)
fig['layout'].update(go.Layout(
width=700,
margin=dict(
r=10, l=10,
b=10, t=10)
))
# fix the ratio in the top left subplot to be a cube
fig['layout'][].update(go.Layout(
go.layout.Scene(aspectmode='cube')),
# manually force the z-axis to appear twice as big as the other two
fig['layout']['scene2'].update(go.layout.Scene(
aspectmode='manual',
aspectratio=go.layout.scene.Aspectratio(
x=1, y=1, z=2
)
))
# draw axes in proportion to the proportion of their ranges
fig['layout']['scene3'].update(go.layout.Scene(aspectmode='data'))
# automatically produce something that is well proportioned using 'data' as the default
fig['layout']['scene4'].update(go.layout.Scene(aspectmode='auto'))
py.iplot(fig, filename='3d-axis-fixed-ratio-axes')
# -
# ### Set Axes Title
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
N = 50
trace1 = go.Mesh3d(x=(60*np.random.randn(N)),
y=(25*np.random.randn(N)),
z=(40*np.random.randn(N)),
opacity=0.5,
color='yellow'
)
trace2 = go.Mesh3d(x=(70*np.random.randn(N)),
y=(55*np.random.randn(N)),
z=(30*np.random.randn(N)),
opacity=0.5,
color='pink'
)
layout = go.Layout(
scene = dict(
xaxis = dict(
title='X AXIS TITLE'),
yaxis = dict(
title='Y AXIS TITLE'),
zaxis = dict(
title='Z AXIS TITLE'),),
width=700,
margin=dict(
r=20, b=10,
l=10, t=10)
)
fig = go.Figure(data=[trace1,trace2], layout=layout)
py.iplot(fig, filename='3d-axis-titles')
# -
# ### Ticks Formatting
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
N = 50
trace1 = go.Mesh3d(x=(60*np.random.randn(N)),
y=(25*np.random.randn(N)),
z=(40*np.random.randn(N)),
opacity=0.5,
color='rgba(100,22,200,0.5)'
)
layout = go.Layout(
scene = dict(
xaxis = dict(
ticktext= ['TICKS','MESH','PLOTLY','PYTHON'],
tickvals= [0,50,75,-50]),
yaxis = dict(
nticks=5, tickfont=dict(
color='green',
size=12,
family='Old Standard TT, serif',),
ticksuffix='#'),
zaxis = dict(
nticks=4, ticks='outside',
tick0=0, tickwidth=4),),
width=700,
margin=dict(
r=10, l=10,
b=10, t=10)
)
fig = go.Figure(data=[trace1], layout=layout)
py.iplot(fig, filename='3d-axis-tick-formatting')
# -
# ### Background and Grid Color
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
N = 50
trace1 = go.Mesh3d(x=(30*np.random.randn(N)),
y=(25*np.random.randn(N)),
z=(30*np.random.randn(N)),
opacity=0.5,)
layout = go.Layout(
scene = dict(
xaxis = dict(
backgroundcolor="rgb(200, 200, 230)",
gridcolor="rgb(255, 255, 255)",
showbackground=True,
zerolinecolor="rgb(255, 255, 255)",),
yaxis = dict(
backgroundcolor="rgb(230, 200,230)",
gridcolor="rgb(255, 255, 255)",
showbackground=True,
zerolinecolor="rgb(255, 255, 255)"),
zaxis = dict(
backgroundcolor="rgb(230, 230,200)",
gridcolor="rgb(255, 255, 255)",
showbackground=True,
zerolinecolor="rgb(255, 255, 255)",),),
width=700,
margin=dict(
r=10, l=10,
b=10, t=10)
)
fig = go.Figure(data=[trace1], layout=layout)
py.iplot(fig, filename='3d-axis-background-and-grid-color')
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'3d-axes.ipynb', 'python/3d-axes/', 'Axes Formatting in 3d Plots | plotly',
'How to format axes of 3d plots in Python with Plotly.',
title = 'Format 3d Axes | plotly',
name = '3D Axes',
has_thumbnail='true', thumbnail='thumbnail/3d-axes.png',
language='python', page_type='example_index',
display_as='3d_charts', order=0.101,
ipynb= '~notebook_demo/96')
# -
| _posts/python-v3/3d/3d-axes/3d-axes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="KhLptlrdVwee" colab_type="code" outputId="83d0129c-1e1f-43cb-9dfb-68514d648399" executionInfo={"status": "ok", "timestamp": 1581766996792, "user_tz": -330, "elapsed": 3787, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd drive/My Drive/google_colab_gpu/SOP3-2/DOP
# + id="P5SRC6AxTxD4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="0745173c-356d-4dc7-b685-0fa9349faab8" executionInfo={"status": "ok", "timestamp": 1581767002191, "user_tz": -330, "elapsed": 4081, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}}
import numpy as np
from keras import layers
from keras.layers import Input, Add, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D, AveragePooling2D, MaxPooling2D, GlobalMaxPooling2D
from keras.models import Model, load_model
from keras.preprocessing import image
from keras.utils import layer_utils
from keras.utils.data_utils import get_file
from keras.applications.imagenet_utils import preprocess_input
import pydot
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
#from resnets_utils import *
from keras.initializers import glorot_uniform
import scipy.misc
from matplotlib.pyplot import imshow
# %matplotlib inline
import keras.backend as K
K.set_image_data_format('channels_last')
K.set_learning_phase(1)
# + id="8Q5mULzmUfIw" colab_type="code" colab={}
import os
import numpy as np
import tensorflow as tf
import h5py
import math
def load_dataset():
train_dataset = h5py.File('datasets/train_signs.h5', "r")
train_set_x_orig = np.array(train_dataset["train_set_x"][:]) # your train set features
train_set_y_orig = np.array(train_dataset["train_set_y"][:]) # your train set labels
test_dataset = h5py.File('datasets/test_signs.h5', "r")
test_set_x_orig = np.array(test_dataset["test_set_x"][:]) # your test set features
test_set_y_orig = np.array(test_dataset["test_set_y"][:]) # your test set labels
classes = np.array(test_dataset["list_classes"][:]) # the list of classes
train_set_y_orig = train_set_y_orig.reshape((1, train_set_y_orig.shape[0]))
test_set_y_orig = test_set_y_orig.reshape((1, test_set_y_orig.shape[0]))
return train_set_x_orig, train_set_y_orig, test_set_x_orig, test_set_y_orig, classes
def random_mini_batches(X, Y, mini_batch_size = 64, seed = 0):
"""
Creates a list of random minibatches from (X, Y)
Arguments:
X -- input data, of shape (input size, number of examples) (m, Hi, Wi, Ci)
Y -- true "label" vector (containing 0 if cat, 1 if non-cat), of shape (1, number of examples) (m, n_y)
mini_batch_size - size of the mini-batches, integer
seed -- this is only for the purpose of grading, so that you're "random minibatches are the same as ours.
Returns:
mini_batches -- list of synchronous (mini_batch_X, mini_batch_Y)
"""
m = X.shape[0] # number of training examples
mini_batches = []
np.random.seed(seed)
# Step 1: Shuffle (X, Y)
permutation = list(np.random.permutation(m))
shuffled_X = X[permutation,:,:,:]
shuffled_Y = Y[permutation,:]
# Step 2: Partition (shuffled_X, shuffled_Y). Minus the end case.
num_complete_minibatches = math.floor(m/mini_batch_size) # number of mini batches of size mini_batch_size in your partitionning
for k in range(0, num_complete_minibatches):
mini_batch_X = shuffled_X[k * mini_batch_size : k * mini_batch_size + mini_batch_size,:,:,:]
mini_batch_Y = shuffled_Y[k * mini_batch_size : k * mini_batch_size + mini_batch_size,:]
mini_batch = (mini_batch_X, mini_batch_Y)
mini_batches.append(mini_batch)
# Handling the end case (last mini-batch < mini_batch_size)
if m % mini_batch_size != 0:
mini_batch_X = shuffled_X[num_complete_minibatches * mini_batch_size : m,:,:,:]
mini_batch_Y = shuffled_Y[num_complete_minibatches * mini_batch_size : m,:]
mini_batch = (mini_batch_X, mini_batch_Y)
mini_batches.append(mini_batch)
return mini_batches
def convert_to_one_hot(Y, C):
Y = np.eye(C)[Y.reshape(-1)].T
return Y
def forward_propagation_for_predict(X, parameters):
"""
Implements the forward propagation for the model: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SOFTMAX
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
b1 = parameters['b1']
W2 = parameters['W2']
b2 = parameters['b2']
W3 = parameters['W3']
b3 = parameters['b3']
# Numpy Equivalents:
Z1 = tf.add(tf.matmul(W1, X), b1) # Z1 = np.dot(W1, X) + b1
A1 = tf.nn.relu(Z1) # A1 = relu(Z1)
Z2 = tf.add(tf.matmul(W2, A1), b2) # Z2 = np.dot(W2, a1) + b2
A2 = tf.nn.relu(Z2) # A2 = relu(Z2)
Z3 = tf.add(tf.matmul(W3, A2), b3) # Z3 = np.dot(W3,Z2) + b3
return Z3
def predict(X, parameters):
W1 = tf.convert_to_tensor(parameters["W1"])
b1 = tf.convert_to_tensor(parameters["b1"])
W2 = tf.convert_to_tensor(parameters["W2"])
b2 = tf.convert_to_tensor(parameters["b2"])
W3 = tf.convert_to_tensor(parameters["W3"])
b3 = tf.convert_to_tensor(parameters["b3"])
params = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2,
"W3": W3,
"b3": b3}
x = tf.placeholder("float", [12288, 1])
z3 = forward_propagation_for_predict(x, params)
p = tf.argmax(z3)
sess = tf.Session()
prediction = sess.run(p, feed_dict = {x: X})
return prediction
# + id="9_d0vVA9UAtc" colab_type="code" colab={}
def identity_block(X, f, filters, stage, block):
"""
Implementation of the identity block as defined in Figure 3
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
Returns:
X -- output of the identity block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value. You'll need this later to add back to the main path.
X_shortcut = X
# First component of main path
X = Conv2D(filters = F1, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path (≈3 lines)
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1,1), padding = 'same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path (≈2 lines)
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
# Final step: Add shortcut value to main path, and pass it through a RELU activation (≈2 lines)
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
# + id="8nJj9oKjUFhB" colab_type="code" colab={}
def convolutional_block(X, f, filters, stage, block, s = 2):
"""
Implementation of the convolutional block as defined in Figure 4
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
stage -- integer, used to name the layers, depending on their position in the network
block -- string/character, used to name the layers, depending on their position in the network
s -- Integer, specifying the stride to be used
Returns:
X -- output of the convolutional block, tensor of shape (n_H, n_W, n_C)
"""
# defining name basis
conv_name_base = 'res' + str(stage) + block + '_branch'
bn_name_base = 'bn' + str(stage) + block + '_branch'
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
##### MAIN PATH #####
# First component of main path
X = Conv2D(F1, (1, 1), strides = (s,s), name = conv_name_base + '2a', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2a')(X)
X = Activation('relu')(X)
# Second component of main path (≈3 lines)
X = Conv2D(filters = F2, kernel_size = (f, f), strides = (1,1), padding = 'same', name = conv_name_base + '2b', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2b')(X)
X = Activation('relu')(X)
# Third component of main path (≈2 lines)
X = Conv2D(filters = F3, kernel_size = (1, 1), strides = (1,1), padding = 'valid', name = conv_name_base + '2c', kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3, name = bn_name_base + '2c')(X)
##### SHORTCUT PATH #### (≈2 lines)
X_shortcut = Conv2D(filters = F3, kernel_size = (1, 1), strides = (s,s), padding = 'valid', name = conv_name_base + '1',
kernel_initializer = glorot_uniform(seed=0))(X_shortcut)
X_shortcut = BatchNormalization(axis = 3, name = bn_name_base + '1')(X_shortcut)
# Final step: Add shortcut value to main path, and pass it through a RELU activation (≈2 lines)
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
# + id="vKBmSsWPUsQh" colab_type="code" colab={}
def ResNet50(input_shape=(64, 64, 3), classes=6):
"""
Implementation of the popular ResNet50 the following architecture:
CONV2D -> BATCHNORM -> RELU -> MAXPOOL -> CONVBLOCK -> IDBLOCK*2 -> CONVBLOCK -> IDBLOCK*3
-> CONVBLOCK -> IDBLOCK*5 -> CONVBLOCK -> IDBLOCK*2 -> AVGPOOL -> TOPLAYER
Arguments:
input_shape -- shape of the images of the dataset
classes -- integer, number of classes
Returns:
model -- a Model() instance in Keras
"""
# Define the input as a tensor with shape input_shape
X_input = Input(input_shape)
# Zero-Padding
#X = ZeroPadding2D((3, 3))(X_input)
# Stage 1
X = Conv2D(64, (1, 1), strides=(2, 2), name='conv1', kernel_initializer=glorot_uniform(seed=0))(X_input)
X = BatchNormalization(axis=3, name='bn_conv1')(X)
X = Activation('relu')(X)
X = MaxPooling2D((3, 3), strides=(2, 2))(X)
# Stage 2
X = convolutional_block(X, f=3, filters=[64, 64, 256], stage=2, block='a', s=1)
X = identity_block(X, 3, [64, 64, 256], stage=2, block='b')
X = identity_block(X, 3, [64, 64, 256], stage=2, block='c')
### START CODE HERE ###
# Stage 3 (≈4 lines)
X = convolutional_block(X, f = 3, filters = [128, 128, 512], stage = 3, block='a', s = 2)
X = identity_block(X, 3, [128, 128, 512], stage=3, block='b')
X = identity_block(X, 3, [128, 128, 512], stage=3, block='c')
X = identity_block(X, 3, [128, 128, 512], stage=3, block='d')
# Stage 4 (≈6 lines)
X = convolutional_block(X, f = 3, filters = [256, 256, 1024], stage = 4, block='a', s = 2)
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='b')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='c')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='d')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='e')
X = identity_block(X, 3, [256, 256, 1024], stage=4, block='f')
# Stage 5 (≈3 lines)
X = convolutional_block(X, f = 3, filters = [512, 512, 2048], stage = 5, block='a', s = 2)
X = identity_block(X, 3, [512, 512, 2048], stage=5, block='b')
X = identity_block(X, 3, [512, 512, 2048], stage=5, block='c')
# AVGPOOL (≈1 line). Use "X = AveragePooling2D(...)(X)"
X = AveragePooling2D((2,2), name="avg_pool")(X)
### END CODE HERE ###
# output layer
X = Flatten()(X)
X = Dense(classes, activation='softmax', name='fc' + str(classes), kernel_initializer = glorot_uniform(seed=0))(X)
# Create model
model = Model(inputs = X_input, outputs = X, name='ResNet50')
return model
# + id="0KoV6eBEUy_-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="a0839d45-0a11-4c3f-9440-63b1c0244f0c" executionInfo={"status": "ok", "timestamp": 1581767103582, "user_tz": -330, "elapsed": 12819, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}}
model = ResNet50(input_shape = (64, 64, 3), classes = 6)
# + id="f0Qio0prU_Ln" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="6fcc292f-1284-468d-c172-a73656e18c46" executionInfo={"status": "ok", "timestamp": 1581767105948, "user_tz": -330, "elapsed": 1062, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}}
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + id="IiIltdRRVCWk" colab_type="code" outputId="a9b7400b-48d5-4c09-83e8-3de3b276891a" executionInfo={"status": "ok", "timestamp": 1581767112371, "user_tz": -330, "elapsed": 3602, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# Normalize image vectors
X_train = X_train_orig/255.
X_test = X_test_orig/255.
# Convert training and test labels to one hot matrices
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
# + id="_68-HGFDcynX" colab_type="code" outputId="725c4c3c-d6b0-4070-9aa2-6d47b2d11330" executionInfo={"status": "ok", "timestamp": 1581767213509, "user_tz": -330, "elapsed": 100747, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.fit(X_train, Y_train, epochs = 25, batch_size = 32)
# + id="qE3Klqrpc0n8" colab_type="code" outputId="61cffeb4-4077-4e86-857c-7fffe969e7f0" executionInfo={"status": "ok", "timestamp": 1581767330038, "user_tz": -330, "elapsed": 3451, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02518986075152188066"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
model.summary()
# + id="O0fk_4stYkGT" colab_type="code" colab={}
Resnet_json = model.to_json()
with open("Resnet.json", "w") as json_file:
json_file.write(Resnet_json)
# + id="UMIpEv5xcVbr" colab_type="code" colab={}
model.save_weights("Resnet.h5")
# + id="1zw_REWFr-hY" colab_type="code" colab={}
| ResNet-50/ResNet_50_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Importando bibliotecas
import pandas as pd
# +
#Importando os dados
# -
data = pd.read_csv('csv/archive/kc_house_data.csv')
data.head(3)
data.shape
# +
#O dataset contém 21.613 casas cadastradas cada uma com 20 atributos e um id único para cada casa
# -
#Nomes de cada atributo
data.columns
#Casa mais cara do cadastro
| modulo1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Kjm9KflC4JPy" outputId="891db49f-6e47-4ce5-c1d8-3443c34485d3"
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print('Your runtime has {:.1f} gigabytes of available RAM\n'.format(ram_gb))
if ram_gb < 20:
print('Not using a high-RAM runtime')
else:
print('You are using a high-RAM runtime!')
# + id="Vn9wuTyn4W1V"
import numpy as np
import pandas as pd
import requests
import json
# + colab={"base_uri": "https://localhost:8080/"} id="jRuEpN9W4qpD" outputId="55b871ca-cc67-479d-d36a-e97643c33c95"
blockNum = 100000
r = requests.get( 'https://blockchain.info/block-height/'+str(blockNum)+'?format-json')
block = r.json()
print (block)
# + colab={"base_uri": "https://localhost:8080/", "height": 394} id="C-IX8BLL6Wmg" outputId="21493720-85fb-4650-cce8-eca7201db8a8"
dataDF = pd.json_normalize(block, record_path= ["blocks"])
txs = pd.json_normalize(dataDF["tx"][0])
txs
#\inputAddresses = pd.json_normalize(txs)
#inputAddresses
# + colab={"base_uri": "https://localhost:8080/", "height": 117} id="m1kI4Rkryijz" outputId="3932d234-f4f6-4799-e749-87dbcca9f4a8"
dataDF
# + colab={"base_uri": "https://localhost:8080/"} id="LdgniWFpsEbk" outputId="c7149c19-723e-41c3-8350-5194c1eb347a"
a= pd.DataFrame(txs,columns=['inputs','out'])
a["inputs"][]
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="Qovy0hHDtjYO" outputId="96b4d174-75d3-4e09-f0f5-b65c9b3511cc"
b = pd.json_normalize(a["out"][1])
b
# + id="SGQi7Xfc-rCw"
def getBlock(blockNum):
r = requests.get( 'https://blockchain.info/block-height/'+str(blockNum)+'?format-json')
block = r.json()
return block
# + id="-_A4l-Ox5Ju0"
def parse(block):
inputAddresses = []
outputAddresses = []
inputValue = []
outputValue = []
timeStamp = []
blockNumber = []
dataDF = pd.json_normalize(block, record_path= ["blocks"])
txs = pd.json_normalize(dataDF["tx"][0])
parsedTxs = pd.DataFrame(txs,columns=['inputs','out'])
#loop through the transactions with inputs and outputs individually
for x in range(1,len(parsedTxs["inputs"])):
Input = pd.json_normalize(parsedTxs["inputs"][x])
Output = pd.json_normalize(parsedTxs["out"][x])
inputAddresses.append(Input["prev_out.addr"])
inputValue.append(Input["prev_out.value"])
outputAddresses.append(Output["addr"])
outputValue.append(Output["value"])
timeStamp.append(dataDF["time"][0])
blockNumber.append(dataDF["height"][0])
data = np.array([inputAddresses,
outputAddresses,
inputValue,
outputValue,
timeStamp,
blockNumber])
#transpose the dataframe, switching the rows and columns
result = data.T
df = pd.DataFrame(result)
df.columns = ["inputAddresses",
"outputAddresses",
"inputValue",
"outputValue",
"timeStamp",
"blockNumber"]
return df
# + colab={"base_uri": "https://localhost:8080/"} id="zWMfwNQMhjLt" outputId="039009ba-c8f7-4d02-efe3-f4dd54e4f05c"
test = getBlock(100000)
df = parse(test)
df.to_csv("/ECON3382/test.csv")
# + id="MhJ1s7s_0_NI"
| ECON3382datacleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/shu65/coffee-tuning/blob/main/coffee_tuning_blog%E7%94%A8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="chLHSr1E6bzv" outputId="04cbd3f0-a429-4711-f42a-bfc73176650b"
# !pip install optuna gspread pandas
# + id="jIqfG-CN7knl"
# Google スプレッドシートの承認
from google.colab import auth
from oauth2client.client import GoogleCredentials
import gspread
auth.authenticate_user()
gc = gspread.authorize(GoogleCredentials.get_application_default())
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="PDd051uL76Iu" outputId="9d020911-a8c2-4e09-97b8-8bdfccf8cd16"
# スプレッドシートからデータを取ってくる
import pandas as pd
ss_name = "shu65コーヒーデータ"
workbook = gc.open(ss_name)
worksheet = workbook.get_worksheet(1)
df = pd.DataFrame(worksheet.get_all_records())
df = df.set_index('淹れた日')
df
# + id="7RHfY57T_Qui"
# 探索空間の定義
import optuna
search_space={
"豆の量(g)": optuna.distributions.IntUniformDistribution(8, 12),
"ミルの時間 (sec)": optuna.distributions.IntUniformDistribution(3, 15),
"トータルの時間 (sec)": optuna.distributions.IntUniformDistribution(180, 300),
"蒸らし時間 (sec)": optuna.distributions.IntUniformDistribution(20, 40),
}
score_column = 'スコア'
# + colab={"base_uri": "https://localhost:8080/"} id="m4UyVOMD7AAn" outputId="b734412f-61ee-4008-aeeb-d180c59e3327"
# 現在までのデータをstudyに登録
import optuna
sampler = optuna.samplers.TPESampler(multivariate=True)
study = optuna.create_study(direction='maximize', sampler=sampler)
for record_i, record in df.iterrows():
print(record.to_dict())
params = {}
for key in search_space.keys():
params[key] = record[key]
trial = optuna.trial.create_trial(
params=params,
distributions=search_space,
value=record[score_column])
study.add_trial(trial)
# + colab={"base_uri": "https://localhost:8080/"} id="_GWb1Pce7CKL" outputId="ea1aeccd-7784-4068-bfce-a9144b515b9b"
# 次のパラメータの出力
trial = study._ask()
new_params = {}
for key, space in search_space.items():
new_params[key] = trial._suggest(key, space)
for key in ["豆の量(g)", "ミルの時間 (sec)", "トータルの時間 (sec)", "蒸らし時間 (sec)"]:
print(key, new_params[key])
| coffee_tuning_blog用.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0";
import ktrain
from ktrain import text as txt
# ## STEP 1: Load and Preprocess the Dataset
#
# A Dutch NER dataset can be downloaded from [here](https://www.clips.uantwerpen.be/conll2002/ner/).
#
# We use the `entities_from_conll2003` function to load and preprocess the data, as the dataset is in a standard **CoNLL** format. (Download the data from the link above to see what the format looks like.)
#
# See the *ktrain* [sequence-tagging tutorial](https://nbviewer.jupyter.org/github/amaiya/ktrain/blob/master/tutorials/tutorial-06-sequence-tagging.ipynb) for more information on how to load data in different ways.
TDATA = 'data/dutch_ner/ned.train'
VDATA = 'data/dutch_ner/ned.testb'
(trn, val, preproc) = txt.entities_from_conll2003(TDATA, val_filepath=VDATA)
# ## STEP 2: Build the Model
#
# Next, we will build a Bidirectional LSTM model that employs the use of transformer embeddings like [BERT word embeddings](https://arxiv.org/abs/1810.04805). By default, the `bilstm-transformer` model will use a pretrained multilingual model (i.e., `bert-base-multilingual-cased`). However, since we are training a Dutch-language model, it is better to select the Dutch pretrained BERT model: `bert-base-dutch-cased`. A full list of available pretrained models is [listed here](https://huggingface.co/transformers/pretrained_models.html). One can also employ the use of [community-uploaded models](https://huggingface.co/models) that focus on specific domains such as the biomedical or scientific domains (e.g, BioBERT, SciBERT). To use SciBERT, for example, set `bert_model` to `allenai/scibert_scivocab_uncased`.
WV_URL='https://dl.fbaipublicfiles.com/fasttext/vectors-crawl/cc.nl.300.vec.gz'
model = txt.sequence_tagger('bilstm-transformer', preproc,
transformer_model='wietsedv/bert-base-dutch-cased', wv_path_or_url=WV_URL)
# In the cell above, notice that we suppied the `wv_path_or_url` argument. This directs *ktrain* to initialized word embeddings with one of the pretrained fasttext (word2vec) word vector sets from [Facebook's fasttext site](https://fasttext.cc/docs/en/crawl-vectors.html). When supplied with a valid URL to a `.vec.gz`, the word vectors will be automatically downloaded, extracted, and loaded in STEP 2 (download location is `<home_directory>/ktrain_data`). To disable pretrained word embeddings, set `wv_path_or_url=None` and randomly initialized word embeddings will be employed. Use of pretrained embeddings will typically boost final accuracy. When used in combination with a model that uses an embedding scheme like BERT (e.g., `bilstm-bert`), the different word embeddings are stacked together using concatenation.
#
# Finally, we will wrap our selected model and datasets in a `Learner` object to facilitate training.
learner = ktrain.get_learner(model, train_data=trn, val_data=val, batch_size=128)
# ## STEP 3: Train the Model
#
# We will train for 5 epochs and decay the learning rate using cosine annealing. This is equivalent to one cycle with a length of 5 epochs. We will save the weights for each epoch in a checkpoint folder. Will train with a learning rate of `0.01`, previously identified using our [learning-rate finder](https://nbviewer.jupyter.org/github/amaiya/ktrain/blob/master/tutorials/tutorial-02-tuning-learning-rates.ipynb).
learner.fit(0.01, 1, cycle_len=5, checkpoint_folder='/tmp/saved_weights')
learner.plot('lr')
# As shown below, our model achieves an F1-Sccore of 83.04 with only a few minutes of training.
learner.validate(class_names=preproc.get_classes())
# ## STEP 4: Make Predictions
predictor = ktrain.get_predictor(learner.model, preproc)
dutch_text = """<NAME> is een Nederlandse politicus die momenteel premier van Nederland is."""
predictor.predict(dutch_text)
predictor.save('/tmp/my_dutch_nermodel')
# The `predictor` can be re-loaded from disk with with `load_predictor`:
predictor = ktrain.load_predictor('/tmp/my_dutch_nermodel')
predictor.predict(dutch_text)
| examples/text/CoNLL2002_Dutch-BiLSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
import re
import pickle
import time
import datetime
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from scipy.sparse import csr_matrix, vstack
# %matplotlib inline
# Custom modules
import const
import func
# -
# ## Load data
print const.TRAIN_FILES
lut = pd.read_csv(const.LOOK_UP_TABLE)
lut.head(3)
cat = func.load_data_file(const.TRAIN_FILES[1])
cat_data = cat['data']['features']
# Load jayjay's features
cat_jay = pd.read_csv('data_jayjay/train.csv')
cat_cols = list(cat_jay.filter(like='CATEGORICAL').columns) + ['L1_L1_Missing value count',
'L3_L3_Missing value count',
'L3_L3_Unique count']
cat_jay = cat_jay[cat_cols]
print cat_jay.shape
cat_jay.head(3)
# ## Reproduce JayJay's features
jay_means = cat_jay.mean()
jay_sums = cat_jay.sum()
print jay_means
# +
def missing_value_count(X):
''' Returns count of missing values per row of sparse matrix X'''
return X.shape[1] - np.diff(X.indptr)
def value_last_element_row(X):
''' Return last value of each row of sparse csr matrix X'''
# Get element where new row starts -1
last = X.indptr[1:] - 1
output = X.data[last]
# Replace row with zero non-zero elements by nan
output[np.diff(X.indptr)==0] = np.nan
return output
def max_element_row(X):
''' Return maximum value of each row of sparse csr matrix X'''
''' nan values are assumed to be encoded as zero'''
output = X.max(1).todense().A1
output[output==0] = np.nan
return output
def alpha_num_max_element_row(X):
''' Return alpha num maximum value of each row of sparse csr matrix X'''
''' nan values are assumed to be encoded as zero'''
''' Lazy, slow implementation, via data/indtptr much faster'''
output= []
for n in range(X.shape[0]):
nz = X[n,:].nonzero()[1]
if nz.shape[0]>0:
data = ['{:d}'.format(int(x)) for x in set(X[n, nz].todense().A1)]
output.append( int(float(max(data))))
else:
#output.append(np.nan)
output.append(0)
return output
def nunique_row(X):
''' Return number of unique per row'''
''' Lazy, slow implementation, via data/indtptr much faster'''
output= []
for n in range(X.shape[0]):
nz = X[n,:].nonzero()[1]
if nz.shape[0]>0:
output.append( len(set(X[n, nz].todense().A1)))
else:
output.append(0)
return output
# +
# 'L1_L1_Missing value count',
col_l1 = [int(i) for i in lut[lut['line']==1].col_cat.values if not np.isnan(i)]
print jay_means['L1_L1_Missing value count']
print pd.Series(missing_value_count(cat_data[:, col_l1])).mean()
# +
# 'L3_L3_Missing value count'
col_l3 = [int(i) for i in lut[lut['line']==3].col_cat.values if not np.isnan(i)]
print jay_means['L3_L3_Missing value count']
print pd.Series(missing_value_count(cat_data[:, col_l3])).mean()
# +
# 'L3_L3_Unique count'
col_l3 = [int(i) for i in lut[lut['line']==3].col_cat.values if not np.isnan(i)]
print jay_means['L3_L3_Unique count']
print pd.Series(nunique_row(cat_data[:, col_l3])).mean()
# -
# CATEGORICAL_Last_____1
n_last = cat_data[n,:].nonzero()[1][-1]
sum([2, 4, 514] == cat_data[n, n_last])
print jay_means['CATEGORICAL_Last_____1']
pd.Series(value_last_element_row(cat_data)).isin([2, 4, 514]).mean()
# CATEGORICAL_Last_____2
print jay_means['CATEGORICAL_Last_____2']
pd.Series(value_last_element_row(cat_data)).isin([16, 48]).mean()
## CATEGORICAL_Missing value count
print jay_means['CATEGORICAL_Missing value count']
pd.Series(cat_data.shape[1] - np.diff(cat_data.indptr)).mean()
# CATEGORICAL_Max______1 (takes a while)
list1 = [2, 8389632, 514]
print jay_means['CATEGORICAL_Max______1']
pd.Series(alpha_num_max_element_row(cat_data)).isin(list1).mean()
# CATEGORICAL_Max______3 (takes a while)
list3 = [3, 145, 4, 143, 8, 512, 6, 32]
print jay_means['CATEGORICAL_Max______3']
pd.Series(alpha_num_max_element_row(cat_data)).isin(list3).mean()
# CATEGORICAL_Unique count
print jay_means['CATEGORICAL_Unique count']
pd.Series(nunique_row(cat_data)).mean()
# CATEGORICAL_out_L3_S32_F3854_class2
# CATEGORICAL_out_out_L3_S32_F3854_class2 0.008123
tmp = np.zeros(d.shape)
tmp[(d==2).values] = 2
tmp[(d==4).values] = 2
tmp.mean()
| feature_set_categorical.ipynb |
# ---
# jupyter:
# accelerator: GPU
# colab:
# collapsed_sections: []
# name: Habitat Interactive Tasks
# provenance: []
# toc_visible: true
# jupytext:
# cell_metadata_filter: -all
# formats: nb_python//py:percent,colabs//ipynb
# notebook_metadata_filter: all
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# language_info:
# codemirror_mode:
# name: ipython
# version: 3
# file_extension: .py
# mimetype: text/x-python
# name: python
# nbconvert_exporter: python
# pygments_lexer: ipython3
# version: 3.7.3
# ---
# %% [markdown]
# # Furniture Rearrangement - How to setup a new interaction task in Habitat-Lab
#
# This tutorial demonstrates how to setup a new task in Habitat that utilizes interaction capabilities in Habitat Simulator.
#
# 
#
# ## Task Definition:
# The working example in this demo will be the task of **Furniture Rearrangement** - The agent will be randomly spawned in an environment in which the furniture are initially displaced from their desired position. The agent is tasked with navigating the environment, picking furniture and putting them in the desired position. To keep the tutorial simple and easy to follow, we will rearrange just a single object.
#
# To setup this task, we will build on top of existing API in Habitat-Simulator and Habitat-Lab. Here is a summary of all the steps involved in setting up this task:
#
# 1. **Setup the Simulator**: Using existing functionalities of the Habitat-Sim, we can add or remove objects from the scene. We will use these methods to spawn the agent and the objects at some pre-defined initial configuration.
# 2. **Create a New Dataset**: We will define a new dataset class to save / load a list of episodes for the agent to train and evaluate on.
# 3. **Grab / Release Action**: We will add the "grab/release" action to the agent's action space to allow the agent to pickup / drop an object under a crosshair.
# 4. **Extend the Simulator Class**: We will extend the Simulator Class to add support for new actions implemented in previous step and add other additional utility functions
# 5. **Create a New Task**: Create a new task definition, implement new *sensors* and *metrics*.
# 6. **Train an RL agent**: We will define rewards for this task and utilize it to train an RL agent using the PPO algorithm.
#
# Let's get started!
# %%
# @title Installation { display-mode: "form" }
# @markdown (double click to show code).
# !curl -L https://raw.githubusercontent.com/facebookresearch/habitat-sim/master/examples/colab_utils/colab_install.sh | NIGHTLY=true bash -s
# %cd /content
# !gdown --id 1Pc-J6pZzXEd8RSeLM94t3iwO8q_RQ853
# !unzip -o /content/coda.zip -d /content/habitat-sim/data/scene_datasets
# reload the cffi version
import sys
if "google.colab" in sys.modules:
import importlib
import cffi
importlib.reload(cffi)
# %%
# @title Path Setup and Imports { display-mode: "form" }
# @markdown (double click to show code).
# %cd /content/habitat-lab
## [setup]
import gzip
import json
import os
import sys
from typing import Any, Dict, List, Optional, Type
import attr
import cv2
import git
import magnum as mn
import numpy as np
# %matplotlib inline
from matplotlib import pyplot as plt
from PIL import Image
import habitat
import habitat_sim
from habitat.config import Config
from habitat.core.registry import registry
from habitat_sim.utils import viz_utils as vut
if "google.colab" in sys.modules:
os.environ["IMAGEIO_FFMPEG_EXE"] = "/usr/bin/ffmpeg"
repo = git.Repo(".", search_parent_directories=True)
dir_path = repo.working_tree_dir
# %cd $dir_path
data_path = os.path.join(dir_path, "data")
output_directory = "data/tutorials/output/" # @param {type:"string"}
output_path = os.path.join(dir_path, output_directory)
if __name__ == "__main__":
import argparse
parser = argparse.ArgumentParser()
parser.add_argument("--no-display", dest="display", action="store_false")
parser.add_argument(
"--no-make-video", dest="make_video", action="store_false"
)
parser.set_defaults(show_video=True, make_video=True)
args, _ = parser.parse_known_args()
show_video = args.display
display = args.display
make_video = args.make_video
else:
show_video = False
make_video = False
display = False
if make_video and not os.path.exists(output_path):
os.makedirs(output_path)
# %%
# @title Util functions to visualize observations
# @markdown - `make_video_cv2`: Renders a video from a list of observations
# @markdown - `simulate`: Runs simulation for a given amount of time at 60Hz
# @markdown - `simulate_and_make_vid` Runs simulation and creates video
def make_video_cv2(
observations, cross_hair=None, prefix="", open_vid=True, fps=60
):
sensor_keys = list(observations[0])
videodims = observations[0][sensor_keys[0]].shape
videodims = (videodims[1], videodims[0]) # flip to w,h order
print(videodims)
video_file = output_path + prefix + ".mp4"
print("Encoding the video: %s " % video_file)
writer = vut.get_fast_video_writer(video_file, fps=fps)
for ob in observations:
# If in RGB/RGBA format, remove the alpha channel
rgb_im_1st_person = cv2.cvtColor(ob["rgb"], cv2.COLOR_RGBA2RGB)
if cross_hair is not None:
rgb_im_1st_person[
cross_hair[0] - 2 : cross_hair[0] + 2,
cross_hair[1] - 2 : cross_hair[1] + 2,
] = [255, 0, 0]
if rgb_im_1st_person.shape[:2] != videodims:
rgb_im_1st_person = cv2.resize(
rgb_im_1st_person, videodims, interpolation=cv2.INTER_AREA
)
# write the 1st person observation to video
writer.append_data(rgb_im_1st_person)
writer.close()
if open_vid:
print("Displaying video")
vut.display_video(video_file)
def simulate(sim, dt=1.0, get_frames=True):
# simulate dt seconds at 60Hz to the nearest fixed timestep
print("Simulating " + str(dt) + " world seconds.")
observations = []
start_time = sim.get_world_time()
while sim.get_world_time() < start_time + dt:
sim.step_physics(1.0 / 60.0)
if get_frames:
observations.append(sim.get_sensor_observations())
return observations
# convenience wrapper for simulate and make_video_cv2
def simulate_and_make_vid(sim, crosshair, prefix, dt=1.0, open_vid=True):
observations = simulate(sim, dt)
make_video_cv2(observations, crosshair, prefix=prefix, open_vid=open_vid)
def display_sample(
rgb_obs,
semantic_obs=np.array([]),
depth_obs=np.array([]),
key_points=None, # noqa: B006
):
from habitat_sim.utils.common import d3_40_colors_rgb
rgb_img = Image.fromarray(rgb_obs, mode="RGB")
arr = [rgb_img]
titles = ["rgb"]
if semantic_obs.size != 0:
semantic_img = Image.new(
"P", (semantic_obs.shape[1], semantic_obs.shape[0])
)
semantic_img.putpalette(d3_40_colors_rgb.flatten())
semantic_img.putdata((semantic_obs.flatten() % 40).astype(np.uint8))
semantic_img = semantic_img.convert("RGBA")
arr.append(semantic_img)
titles.append("semantic")
if depth_obs.size != 0:
depth_img = Image.fromarray(
(depth_obs / 10 * 255).astype(np.uint8), mode="L"
)
arr.append(depth_img)
titles.append("depth")
plt.figure(figsize=(12, 8))
for i, data in enumerate(arr):
ax = plt.subplot(1, 3, i + 1)
ax.axis("off")
ax.set_title(titles[i])
# plot points on images
if key_points is not None:
for point in key_points:
plt.plot(
point[0], point[1], marker="o", markersize=10, alpha=0.8
)
plt.imshow(data)
plt.show(block=False)
# %% [markdown]
# ## 1. Setup the Simulator
#
# ---
#
#
# %%
# @title Setup simulator configuration
# @markdown We'll start with setting up simulator with the following configurations
# @markdown - The simulator will render both RGB, Depth observations of 256x256 resolution.
# @markdown - The actions available will be `move_forward`, `turn_left`, `turn_right`.
def make_cfg(settings):
sim_cfg = habitat_sim.SimulatorConfiguration()
sim_cfg.gpu_device_id = 0
sim_cfg.default_agent_id = settings["default_agent_id"]
sim_cfg.scene_id = settings["scene"]
sim_cfg.enable_physics = settings["enable_physics"]
sim_cfg.physics_config_file = settings["physics_config_file"]
# Note: all sensors must have the same resolution
sensors = {
"rgb": {
"sensor_type": habitat_sim.SensorType.COLOR,
"resolution": [settings["height"], settings["width"]],
"position": [0.0, settings["sensor_height"], 0.0],
},
"depth": {
"sensor_type": habitat_sim.SensorType.DEPTH,
"resolution": [settings["height"], settings["width"]],
"position": [0.0, settings["sensor_height"], 0.0],
},
}
sensor_specs = []
for sensor_uuid, sensor_params in sensors.items():
if settings[sensor_uuid]:
sensor_spec = habitat_sim.SensorSpec()
sensor_spec.uuid = sensor_uuid
sensor_spec.sensor_type = sensor_params["sensor_type"]
sensor_spec.resolution = sensor_params["resolution"]
sensor_spec.position = sensor_params["position"]
sensor_specs.append(sensor_spec)
# Here you can specify the amount of displacement in a forward action and the turn angle
agent_cfg = habitat_sim.agent.AgentConfiguration()
agent_cfg.sensor_specifications = sensor_specs
agent_cfg.action_space = {
"move_forward": habitat_sim.agent.ActionSpec(
"move_forward", habitat_sim.agent.ActuationSpec(amount=0.1)
),
"turn_left": habitat_sim.agent.ActionSpec(
"turn_left", habitat_sim.agent.ActuationSpec(amount=10.0)
),
"turn_right": habitat_sim.agent.ActionSpec(
"turn_right", habitat_sim.agent.ActuationSpec(amount=10.0)
),
}
return habitat_sim.Configuration(sim_cfg, [agent_cfg])
settings = {
"max_frames": 10,
"width": 256,
"height": 256,
"scene": "data/scene_datasets/coda/coda.glb",
"default_agent_id": 0,
"sensor_height": 1.5, # Height of sensors in meters
"rgb": True, # RGB sensor
"depth": True, # Depth sensor
"seed": 1,
"enable_physics": True,
"physics_config_file": "data/default.physics_config.json",
"silent": False,
"compute_shortest_path": False,
"compute_action_shortest_path": False,
"save_png": True,
}
cfg = make_cfg(settings)
# %%
# @title Spawn the agent at a pre-defined location
def init_agent(sim):
agent_pos = np.array([-0.15776923, 0.18244143, 0.2988735])
# Place the agent
sim.agents[0].scene_node.translation = agent_pos
agent_orientation_y = -40
sim.agents[0].scene_node.rotation = mn.Quaternion.rotation(
mn.Deg(agent_orientation_y), mn.Vector3(0, 1.0, 0)
)
cfg.sim_cfg.default_agent_id = 0
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
if make_video:
# Visualize the agent's initial position
simulate_and_make_vid(
sim, None, "sim-init", dt=1.0, open_vid=show_video
)
# %%
# @title Set the object's initial and final position
# @markdown Defines two utility functions:
# @markdown - `remove_all_objects`: This will remove all objects from the scene
# @markdown - `set_object_in_front_of_agent`: This will add an object in the scene in front of the agent at the specified distance.
# @markdown Here we add a chair *3.0m* away from the agent and the task is to place the agent at the desired final position which is *7.0m* in front of the agent.
def remove_all_objects(sim):
for obj_id in sim.get_existing_object_ids():
sim.remove_object(obj_id)
def set_object_in_front_of_agent(sim, obj_id, z_offset=-1.5):
r"""
Adds an object in front of the agent at some distance.
"""
agent_transform = sim.agents[0].scene_node.transformation_matrix()
obj_translation = agent_transform.transform_point(
np.array([0, 0, z_offset])
)
sim.set_translation(obj_translation, obj_id)
obj_node = sim.get_object_scene_node(obj_id)
xform_bb = habitat_sim.geo.get_transformed_bb(
obj_node.cumulative_bb, obj_node.transformation
)
# also account for collision margin of the scene
scene_collision_margin = 0.04
y_translation = mn.Vector3(
0, xform_bb.size_y() / 2.0 + scene_collision_margin, 0
)
sim.set_translation(y_translation + sim.get_translation(obj_id), obj_id)
def init_objects(sim):
# Manager of Object Attributes Templates
obj_attr_mgr = sim.get_object_template_manager()
obj_attr_mgr.load_configs(
str(os.path.join(data_path, "test_assets/objects"))
)
# Add a chair into the scene.
obj_path = "test_assets/objects/chair"
chair_template_id = obj_attr_mgr.load_object_configs(
str(os.path.join(data_path, obj_path))
)[0]
chair_attr = obj_attr_mgr.get_template_by_ID(chair_template_id)
obj_attr_mgr.register_template(chair_attr)
# Object's initial position 3m away from the agent.
object_id = sim.add_object_by_handle(chair_attr.handle)
set_object_in_front_of_agent(sim, object_id, -3.0)
sim.set_object_motion_type(
habitat_sim.physics.MotionType.STATIC, object_id
)
# Object's final position 7m away from the agent
goal_id = sim.add_object_by_handle(chair_attr.handle)
set_object_in_front_of_agent(sim, goal_id, -7.0)
sim.set_object_motion_type(habitat_sim.physics.MotionType.STATIC, goal_id)
return object_id, goal_id
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
init_objects(sim)
# Visualize the scene after the chair is added into the scene.
if make_video:
simulate_and_make_vid(
sim, None, "object-init", dt=1.0, open_vid=show_video
)
# %% [markdown]
# ## Rearrangement Dataset
# 
#
# In the previous section, we created a single episode of the rearrangement task. Let's define a format to store all the necessary information about a single episode. It should store the *scene* the episode belongs to, *initial spawn position and orientation* of the agent, *object type*, object's *initial position and orientation* as well as *final position and orientation*.
#
# The format will be as follows:
# ```
# {
# 'episode_id': 0,
# 'scene_id': 'data/scene_datasets/coda/coda.glb',
# 'goals': {
# 'position': [4.34, 0.67, -5.06],
# 'rotation': [0.0, 0.0, 0.0, 1.0]
# },
# 'objects': {
# 'object_id': 0,
# 'object_template': 'data/test_assets/objects/chair',
# 'position': [1.77, 0.67, -1.99],
# 'rotation': [0.0, 0.0, 0.0, 1.0]
# },
# 'start_position': [-0.15, 0.18, 0.29],
# 'start_rotation': [-0.0, -0.34, -0.0, 0.93]}
# }
# ```
# Once an episode is defined, a dataset will just be a collection of such episodes. For simplicity, in this notebook, the dataset will only contain one episode defined above.
#
# %%
# @title Create a new dataset
# @markdown Utility functions to define and save the dataset for the rearrangement task
def get_rotation(sim, object_id):
quat = sim.get_rotation(object_id)
return np.array(quat.vector).tolist() + [quat.scalar]
def init_episode_dict(episode_id, scene_id, agent_pos, agent_rot):
episode_dict = {
"episode_id": episode_id,
"scene_id": "data/scene_datasets/coda/coda.glb",
"start_position": agent_pos,
"start_rotation": agent_rot,
"info": {},
}
return episode_dict
def add_object_details(sim, episode_dict, obj_id, object_template, object_id):
object_template = {
"object_id": obj_id,
"object_template": object_template,
"position": np.array(sim.get_translation(object_id)).tolist(),
"rotation": get_rotation(sim, object_id),
}
episode_dict["objects"] = object_template
return episode_dict
def add_goal_details(sim, episode_dict, object_id):
goal_template = {
"position": np.array(sim.get_translation(object_id)).tolist(),
"rotation": get_rotation(sim, object_id),
}
episode_dict["goals"] = goal_template
return episode_dict
# set the number of objects to 1 always for now.
def build_episode(sim, episode_num, object_id, goal_id):
episodes = {"episodes": []}
for episode in range(episode_num):
agent_state = sim.get_agent(0).get_state()
agent_pos = np.array(agent_state.position).tolist()
agent_quat = agent_state.rotation
agent_rot = np.array(agent_quat.vec).tolist() + [agent_quat.real]
episode_dict = init_episode_dict(
episode, settings["scene"], agent_pos, agent_rot
)
object_attr = sim.get_object_initialization_template(object_id)
object_path = os.path.relpath(
os.path.splitext(object_attr.render_asset_handle)[0]
)
episode_dict = add_object_details(
sim, episode_dict, 0, object_path, object_id
)
episode_dict = add_goal_details(sim, episode_dict, goal_id)
episodes["episodes"].append(episode_dict)
return episodes
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
object_id, goal_id = init_objects(sim)
episodes = build_episode(sim, 1, object_id, goal_id)
dataset_content_path = "data/datasets/rearrangement/coda/v1/train/"
if not os.path.exists(dataset_content_path):
os.makedirs(dataset_content_path)
with gzip.open(
os.path.join(dataset_content_path, "train.json.gz"), "wt"
) as f:
json.dump(episodes, f)
print(
"Dataset written to {}".format(
os.path.join(dataset_content_path, "train.json.gz")
)
)
# %%
# @title Dataset class to read the saved dataset in Habitat-Lab.
# @markdown To read the saved episodes in Habitat-Lab, we will extend the `Dataset` class and the `Episode` base class. It will help provide all the relevant details about the episode through a consistent API to all downstream tasks.
# @markdown - We will first create a `RearrangementEpisode` by extending the `NavigationEpisode` to include additional information about object's initial configuration and desired final configuration.
# @markdown - We will then define a `RearrangementDatasetV0` class that builds on top of `PointNavDatasetV1` class to read the JSON file stored earlier and initialize a list of `RearrangementEpisode`.
from habitat.core.utils import DatasetFloatJSONEncoder, not_none_validator
from habitat.datasets.pointnav.pointnav_dataset import (
CONTENT_SCENES_PATH_FIELD,
DEFAULT_SCENE_PATH_PREFIX,
PointNavDatasetV1,
)
from habitat.tasks.nav.nav import NavigationEpisode
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementSpec:
r"""Specifications that capture a particular position of final position
or initial position of the object.
"""
position: List[float] = attr.ib(default=None, validator=not_none_validator)
rotation: List[float] = attr.ib(default=None, validator=not_none_validator)
info: Optional[Dict[str, str]] = attr.ib(default=None)
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementObjectSpec(RearrangementSpec):
r"""Object specifications that capture position of each object in the scene,
the associated object template.
"""
object_id: str = attr.ib(default=None, validator=not_none_validator)
object_template: Optional[str] = attr.ib(
default="data/test_assets/objects/chair"
)
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementEpisode(NavigationEpisode):
r"""Specification of episode that includes initial position and rotation
of agent, all goal specifications, all object specifications
Args:
episode_id: id of episode in the dataset
scene_id: id of scene inside the simulator.
start_position: numpy ndarray containing 3 entries for (x, y, z).
start_rotation: numpy ndarray with 4 entries for (x, y, z, w)
elements of unit quaternion (versor) representing agent 3D
orientation.
goal: object's goal position and rotation
object: object's start specification defined with object type,
position, and rotation.
"""
objects: RearrangementObjectSpec = attr.ib(
default=None, validator=not_none_validator
)
goals: RearrangementSpec = attr.ib(
default=None, validator=not_none_validator
)
@registry.register_dataset(name="RearrangementDataset-v0")
class RearrangementDatasetV0(PointNavDatasetV1):
r"""Class inherited from PointNavDataset that loads Rearrangement dataset."""
episodes: List[RearrangementEpisode]
content_scenes_path: str = "{data_path}/content/{scene}.json.gz"
def to_json(self) -> str:
result = DatasetFloatJSONEncoder().encode(self)
return result
def __init__(self, config: Optional[Config] = None) -> None:
super().__init__(config)
def from_json(
self, json_str: str, scenes_dir: Optional[str] = None
) -> None:
deserialized = json.loads(json_str)
if CONTENT_SCENES_PATH_FIELD in deserialized:
self.content_scenes_path = deserialized[CONTENT_SCENES_PATH_FIELD]
for i, episode in enumerate(deserialized["episodes"]):
rearrangement_episode = RearrangementEpisode(**episode)
rearrangement_episode.episode_id = str(i)
if scenes_dir is not None:
if rearrangement_episode.scene_id.startswith(
DEFAULT_SCENE_PATH_PREFIX
):
rearrangement_episode.scene_id = (
rearrangement_episode.scene_id[
len(DEFAULT_SCENE_PATH_PREFIX) :
]
)
rearrangement_episode.scene_id = os.path.join(
scenes_dir, rearrangement_episode.scene_id
)
rearrangement_episode.objects = RearrangementObjectSpec(
**rearrangement_episode.objects
)
rearrangement_episode.goals = RearrangementSpec(
**rearrangement_episode.goals
)
self.episodes.append(rearrangement_episode)
# %%
# @title Load the saved dataset using the Dataset class
config = habitat.get_config("configs/datasets/pointnav/habitat_test.yaml")
config.defrost()
config.DATASET.DATA_PATH = (
"data/datasets/rearrangement/coda/v1/{split}/{split}.json.gz"
)
config.DATASET.TYPE = "RearrangementDataset-v0"
config.freeze()
dataset = RearrangementDatasetV0(config.DATASET)
# check if the dataset got correctly deserialized
assert len(dataset.episodes) == 1
assert dataset.episodes[0].objects.position == [
1.770593523979187,
0.6726829409599304,
-1.9992598295211792,
]
assert dataset.episodes[0].objects.rotation == [0.0, 0.0, 0.0, 1.0]
assert (
dataset.episodes[0].objects.object_template
== "data/test_assets/objects/chair"
)
assert dataset.episodes[0].goals.position == [
4.3417439460754395,
0.6726829409599304,
-5.0634379386901855,
]
assert dataset.episodes[0].goals.rotation == [0.0, 0.0, 0.0, 1.0]
# %% [markdown]
# ## Implement Grab/Release Action
# %%
# @title RayCast utility to implement Grab/Release Under Cross-Hair Action
# @markdown Cast a ray in the direction of crosshair from the camera and check if it collides with another object within a certain distance threshold
def raycast(sim, sensor_name, crosshair_pos=(128, 128), max_distance=2.0):
r"""Cast a ray in the direction of crosshair and check if it collides
with another object within a certain distance threshold
:param sim: Simulator object
:param sensor_name: name of the visual sensor to be used for raycasting
:param crosshair_pos: 2D coordiante in the viewport towards which the
ray will be cast
:param max_distance: distance threshold beyond which objects won't
be considered
"""
render_camera = sim._sensors[sensor_name]._sensor_object.render_camera
center_ray = render_camera.unproject(mn.Vector2i(crosshair_pos))
raycast_results = sim.cast_ray(center_ray, max_distance=max_distance)
closest_object = -1
closest_dist = 1000.0
if raycast_results.has_hits():
for hit in raycast_results.hits:
if hit.ray_distance < closest_dist:
closest_dist = hit.ray_distance
closest_object = hit.object_id
return closest_object
# %%
# Test the raycast utility.
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
obj_attr_mgr = sim.get_object_template_manager()
obj_attr_mgr.load_configs(
str(os.path.join(data_path, "test_assets/objects"))
)
obj_path = "test_assets/objects/chair"
chair_template_id = obj_attr_mgr.load_object_configs(
str(os.path.join(data_path, obj_path))
)[0]
chair_attr = obj_attr_mgr.get_template_by_ID(chair_template_id)
obj_attr_mgr.register_template(chair_attr)
object_id = sim.add_object_by_handle(chair_attr.handle)
print(f"Chair's object id is {object_id}")
set_object_in_front_of_agent(sim, object_id, -1.5)
sim.set_object_motion_type(
habitat_sim.physics.MotionType.STATIC, object_id
)
if make_video:
# Visualize the agent's initial position
simulate_and_make_vid(
sim, [190, 128], "sim-before-grab", dt=1.0, open_vid=show_video
)
# Distance threshold=2 is greater than agent-to-chair distance.
# Should return chair's object id
closest_object = raycast(
sim, "rgb", crosshair_pos=[128, 190], max_distance=2.0
)
print(f"Closest Object ID: {closest_object} using 2.0 threshold")
assert (
closest_object == object_id
), f"Could not pick chair with ID: {object_id}"
# Distance threshold=1 is smaller than agent-to-chair distance .
# Should return -1
closest_object = raycast(
sim, "rgb", crosshair_pos=[128, 190], max_distance=1.0
)
print(f"Closest Object ID: {closest_object} using 1.0 threshold")
assert closest_object == -1, "Agent shoud not be able to pick any object"
# %%
# @title Define a Grab/Release action and create a new action space.
# @markdown Each new action is defined by a `ActionSpec` and an `ActuationSpec`. `ActionSpec` is mapping between the action name and its corresponding `ActuationSpec`. `ActuationSpec` contains all the necessary specifications required to define the action.
from habitat.config.default import _C, CN
from habitat.core.embodied_task import SimulatorTaskAction
from habitat.sims.habitat_simulator.actions import (
HabitatSimActions,
HabitatSimV1ActionSpaceConfiguration,
)
from habitat_sim.agent.controls.controls import ActuationSpec
from habitat_sim.physics import MotionType
# @markdown For instance, `GrabReleaseActuationSpec` contains the following:
# @markdown - `visual_sensor_name` defines which viewport (rgb, depth, etc) to to use to cast the ray.
# @markdown - `crosshair_pos` stores the position in the viewport through which the ray passes. Any object which intersects with this ray can be grabbed by the agent.
# @markdown - `amount` defines a distance threshold. Objects which are farther than the treshold cannot be picked up by the agent.
@attr.s(auto_attribs=True, slots=True)
class GrabReleaseActuationSpec(ActuationSpec):
visual_sensor_name: str = "rgb"
crosshair_pos: List[int] = [128, 128]
amount: float = 2.0
# @markdown Then, we extend the `HabitatSimV1ActionSpaceConfiguration` to add the above action into the agent's action space. `ActionSpaceConfiguration` is a mapping between action name and the corresponding `ActionSpec`
@registry.register_action_space_configuration(name="RearrangementActions-v0")
class RearrangementSimV0ActionSpaceConfiguration(
HabitatSimV1ActionSpaceConfiguration
):
def __init__(self, config):
super().__init__(config)
if not HabitatSimActions.has_action("GRAB_RELEASE"):
HabitatSimActions.extend_action_space("GRAB_RELEASE")
def get(self):
config = super().get()
new_config = {
HabitatSimActions.GRAB_RELEASE: habitat_sim.ActionSpec(
"grab_or_release_object_under_crosshair",
GrabReleaseActuationSpec(
visual_sensor_name=self.config.VISUAL_SENSOR,
crosshair_pos=self.config.CROSSHAIR_POS,
amount=self.config.GRAB_DISTANCE,
),
)
}
config.update(new_config)
return config
# @markdown Finally, we extend `SimualtorTaskAction` which tells the simulator which action to call when a named action ('GRAB_RELEASE' in this case) is predicte by the agent's policy.
@registry.register_task_action
class GrabOrReleaseAction(SimulatorTaskAction):
def step(self, *args: Any, **kwargs: Any):
r"""This method is called from ``Env`` on each ``step``."""
return self._sim.step(HabitatSimActions.GRAB_RELEASE)
_C.TASK.ACTIONS.GRAB_RELEASE = CN()
_C.TASK.ACTIONS.GRAB_RELEASE.TYPE = "GrabOrReleaseAction"
_C.SIMULATOR.CROSSHAIR_POS = [128, 160]
_C.SIMULATOR.GRAB_DISTANCE = 2.0
_C.SIMULATOR.VISUAL_SENSOR = "rgb"
# %% [markdown]
# ##Setup Simulator Class for Rearrangement Task
#
# 
# %%
# @title RearrangementSim Class
# @markdown Here we will extend the `HabitatSim` class for the rearrangement task. We will make the following changes:
# @markdown - define a new `_initialize_objects` function which will load the object in its initial configuration as defined by the episode.
# @markdown - define a `gripped_object_id` property that stores whether the agent is holding any object or not.
# @markdown - modify the `step` function of the simulator to use the `grab/release` action we define earlier.
# @markdown #### Writing the `step` function:
# @markdown Since we added a new action for this task, we have to modify the `step` function to define what happens when `grab/release` action is called. If a simple navigation action (`move_forward`, `turn_left`, `turn_right`) is called, we pass it forward to `act` function of the agent which already defines the behavior of these actions.
# @markdown For the `grab/release` action, if the agent is not already holding an object, we first call the `raycast` function using the values from the `ActuationSpec` to see if any object is grippable. If it returns a valid object id, we put the object in a "invisible" inventory and remove it from the scene.
# @markdown If the agent was already holding an object, `grab/release` action will try release the object at the same relative position as it was grabbed. If the object can be placed without any collision, then the `release` action is successful.
from habitat.sims.habitat_simulator.habitat_simulator import HabitatSim
from habitat_sim.nav import NavMeshSettings
from habitat_sim.utils.common import quat_from_coeffs, quat_to_magnum
@registry.register_simulator(name="RearrangementSim-v0")
class RearrangementSim(HabitatSim):
r"""Simulator wrapper over habitat-sim with
object rearrangement functionalities.
"""
def __init__(self, config: Config) -> None:
self.did_reset = False
super().__init__(config=config)
self.grip_offset = np.eye(4)
agent_id = self.habitat_config.DEFAULT_AGENT_ID
agent_config = self._get_agent_config(agent_id)
self.navmesh_settings = NavMeshSettings()
self.navmesh_settings.set_defaults()
self.navmesh_settings.agent_radius = agent_config.RADIUS
self.navmesh_settings.agent_height = agent_config.HEIGHT
def reconfigure(self, config: Config) -> None:
super().reconfigure(config)
self._initialize_objects()
def reset(self):
sim_obs = super().reset()
if self._update_agents_state():
sim_obs = self.get_sensor_observations()
self._prev_sim_obs = sim_obs
self.did_reset = True
self.grip_offset = np.eye(4)
return self._sensor_suite.get_observations(sim_obs)
def _initialize_objects(self):
objects = self.habitat_config.objects[0]
obj_attr_mgr = self.get_object_template_manager()
obj_attr_mgr.load_configs(
str(os.path.join(data_path, "test_assets/objects"))
)
# first remove all existing objects
existing_object_ids = self.get_existing_object_ids()
if len(existing_object_ids) > 0:
for obj_id in existing_object_ids:
self.remove_object(obj_id)
self.sim_object_to_objid_mapping = {}
self.objid_to_sim_object_mapping = {}
if objects is not None:
object_template = objects["object_template"]
object_pos = objects["position"]
object_rot = objects["rotation"]
object_template_id = obj_attr_mgr.load_object_configs(
object_template
)[0]
object_attr = obj_attr_mgr.get_template_by_ID(object_template_id)
obj_attr_mgr.register_template(object_attr)
object_id = self.add_object_by_handle(object_attr.handle)
self.sim_object_to_objid_mapping[object_id] = objects["object_id"]
self.objid_to_sim_object_mapping[objects["object_id"]] = object_id
self.set_translation(object_pos, object_id)
if isinstance(object_rot, list):
object_rot = quat_from_coeffs(object_rot)
object_rot = quat_to_magnum(object_rot)
self.set_rotation(object_rot, object_id)
self.set_object_motion_type(MotionType.STATIC, object_id)
# Recompute the navmesh after placing all the objects.
self.recompute_navmesh(self.pathfinder, self.navmesh_settings, True)
def _sync_gripped_object(self, gripped_object_id):
r"""
Sync the gripped object with the object associated with the agent.
"""
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
self.set_transformation(
agent_body_transformation, gripped_object_id
)
translation = agent_body_transformation.transform_point(
np.array([0, 2.0, 0])
)
self.set_translation(translation, gripped_object_id)
@property
def gripped_object_id(self):
return self._prev_sim_obs.get("gripped_object_id", -1)
def step(self, action: int):
dt = 1 / 60.0
self._num_total_frames += 1
collided = False
gripped_object_id = self.gripped_object_id
agent_config = self._default_agent.agent_config
action_spec = agent_config.action_space[action]
if action_spec.name == "grab_or_release_object_under_crosshair":
# If already holding an agent
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
T = np.dot(agent_body_transformation, self.grip_offset)
self.set_transformation(T, gripped_object_id)
position = self.get_translation(gripped_object_id)
if self.pathfinder.is_navigable(position):
self.set_object_motion_type(
MotionType.STATIC, gripped_object_id
)
gripped_object_id = -1
self.recompute_navmesh(
self.pathfinder, self.navmesh_settings, True
)
# if not holding an object, then try to grab
else:
gripped_object_id = raycast(
self,
action_spec.actuation.visual_sensor_name,
crosshair_pos=action_spec.actuation.crosshair_pos,
max_distance=action_spec.actuation.amount,
)
# found a grabbable object.
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
self.grip_offset = np.dot(
np.array(agent_body_transformation.inverted()),
np.array(self.get_transformation(gripped_object_id)),
)
self.set_object_motion_type(
MotionType.KINEMATIC, gripped_object_id
)
self.recompute_navmesh(
self.pathfinder, self.navmesh_settings, True
)
else:
collided = self._default_agent.act(action)
self._last_state = self._default_agent.get_state()
# step physics by dt
super().step_world(dt)
# Sync the gripped object after the agent moves.
self._sync_gripped_object(gripped_object_id)
# obtain observations
self._prev_sim_obs = self.get_sensor_observations()
self._prev_sim_obs["collided"] = collided
self._prev_sim_obs["gripped_object_id"] = gripped_object_id
observations = self._sensor_suite.get_observations(self._prev_sim_obs)
return observations
# %% [markdown]
# ## Create the Rearrangement Task
# 
# %%
# @title Implement new sensors and measurements
# @markdown After defining the dataset, action space and simulator functions for the rearrangement task, we are one step closer to training agents to solve this task.
# @markdown Here we define inputs to the policy and other measurements required to design reward functions.
# @markdown **Sensors**: These define various part of the simulator state that's visible to the agent. For simplicity, we'll assume that agent knows the object's current position, object's final goal position relative to the agent's current position.
# @markdown - Object's current position will be made given by the `ObjectPosition` sensor
# @markdown - Object's goal position will be available through the `ObjectGoal` sensor.
# @markdown - Finally, we will also use `GrippedObject` sensor to tell the agent if it's holding any object or not.
# @markdown **Measures**: These define various metrics about the task which can be used to measure task progress and define rewards. Note that measurements are *privileged* information not accessible to the agent as part of the observation space. We will need the following measurements:
# @markdown - `AgentToObjectDistance` which measure the euclidean distance between the agent and the object.
# @markdown - `ObjectToGoalDistance` which measures the euclidean distance between the object and the goal.
from gym import spaces
import habitat_sim
from habitat.config.default import CN, Config
from habitat.core.dataset import Episode
from habitat.core.embodied_task import Measure
from habitat.core.simulator import Observations, Sensor, SensorTypes, Simulator
from habitat.tasks.nav.nav import PointGoalSensor
@registry.register_sensor
class GrippedObjectSensor(Sensor):
cls_uuid = "gripped_object_id"
def __init__(
self, *args: Any, sim: RearrangementSim, config: Config, **kwargs: Any
):
self._sim = sim
super().__init__(config=config)
def _get_uuid(self, *args: Any, **kwargs: Any) -> str:
return self.cls_uuid
def _get_observation_space(self, *args: Any, **kwargs: Any):
return spaces.Discrete(len(self._sim.get_existing_object_ids()))
def _get_sensor_type(self, *args: Any, **kwargs: Any):
return SensorTypes.MEASUREMENT
def get_observation(
self,
observations: Dict[str, Observations],
episode: Episode,
*args: Any,
**kwargs: Any,
):
obj_id = self._sim.sim_object_to_objid_mapping.get(
self._sim.gripped_object_id, -1
)
return obj_id
@registry.register_sensor
class ObjectPosition(PointGoalSensor):
cls_uuid: str = "object_position"
def _get_observation_space(self, *args: Any, **kwargs: Any):
sensor_shape = (self._dimensionality,)
return spaces.Box(
low=np.finfo(np.float32).min,
high=np.finfo(np.float32).max,
shape=sensor_shape,
dtype=np.float32,
)
def get_observation(
self, *args: Any, observations, episode, **kwargs: Any
):
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
rotation_world_agent = agent_state.rotation
object_id = self._sim.get_existing_object_ids()[0]
object_position = self._sim.get_translation(object_id)
pointgoal = self._compute_pointgoal(
agent_position, rotation_world_agent, object_position
)
return pointgoal
@registry.register_sensor
class ObjectGoal(PointGoalSensor):
cls_uuid: str = "object_goal"
def _get_observation_space(self, *args: Any, **kwargs: Any):
sensor_shape = (self._dimensionality,)
return spaces.Box(
low=np.finfo(np.float32).min,
high=np.finfo(np.float32).max,
shape=sensor_shape,
dtype=np.float32,
)
def get_observation(
self, *args: Any, observations, episode, **kwargs: Any
):
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
rotation_world_agent = agent_state.rotation
goal_position = np.array(episode.goals.position, dtype=np.float32)
point_goal = self._compute_pointgoal(
agent_position, rotation_world_agent, goal_position
)
return point_goal
@registry.register_measure
class ObjectToGoalDistance(Measure):
"""The measure calculates distance of object towards the goal."""
cls_uuid: str = "object_to_goal_distance"
def __init__(
self, sim: Simulator, config: Config, *args: Any, **kwargs: Any
):
self._sim = sim
self._config = config
super().__init__(**kwargs)
@staticmethod
def _get_uuid(*args: Any, **kwargs: Any):
return ObjectToGoalDistance.cls_uuid
def reset_metric(self, episode, *args: Any, **kwargs: Any):
self.update_metric(*args, episode=episode, **kwargs)
def _geo_dist(self, src_pos, goal_pos: np.array) -> float:
return self._sim.geodesic_distance(src_pos, [goal_pos])
def _euclidean_distance(self, position_a, position_b):
return np.linalg.norm(
np.array(position_b) - np.array(position_a), ord=2
)
def update_metric(self, episode, *args: Any, **kwargs: Any):
sim_obj_id = self._sim.get_existing_object_ids()[0]
previous_position = np.array(
self._sim.get_translation(sim_obj_id)
).tolist()
goal_position = episode.goals.position
self._metric = self._euclidean_distance(
previous_position, goal_position
)
@registry.register_measure
class AgentToObjectDistance(Measure):
"""The measure calculates the distance of objects from the agent"""
cls_uuid: str = "agent_to_object_distance"
def __init__(
self, sim: Simulator, config: Config, *args: Any, **kwargs: Any
):
self._sim = sim
self._config = config
super().__init__(**kwargs)
@staticmethod
def _get_uuid(*args: Any, **kwargs: Any):
return AgentToObjectDistance.cls_uuid
def reset_metric(self, episode, *args: Any, **kwargs: Any):
self.update_metric(*args, episode=episode, **kwargs)
def _euclidean_distance(self, position_a, position_b):
return np.linalg.norm(
np.array(position_b) - np.array(position_a), ord=2
)
def update_metric(self, episode, *args: Any, **kwargs: Any):
sim_obj_id = self._sim.get_existing_object_ids()[0]
previous_position = np.array(
self._sim.get_translation(sim_obj_id)
).tolist()
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
self._metric = self._euclidean_distance(
previous_position, agent_position
)
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK GRIPPED OBJECT SENSOR
# -----------------------------------------------------------------------------
_C.TASK.GRIPPED_OBJECT_SENSOR = CN()
_C.TASK.GRIPPED_OBJECT_SENSOR.TYPE = "GrippedObjectSensor"
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK ALL OBJECT POSITIONS SENSOR
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_POSITION = CN()
_C.TASK.OBJECT_POSITION.TYPE = "ObjectPosition"
_C.TASK.OBJECT_POSITION.GOAL_FORMAT = "POLAR"
_C.TASK.OBJECT_POSITION.DIMENSIONALITY = 2
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK ALL OBJECT GOALS SENSOR
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_GOAL = CN()
_C.TASK.OBJECT_GOAL.TYPE = "ObjectGoal"
_C.TASK.OBJECT_GOAL.GOAL_FORMAT = "POLAR"
_C.TASK.OBJECT_GOAL.DIMENSIONALITY = 2
# -----------------------------------------------------------------------------
# # OBJECT_DISTANCE_TO_GOAL MEASUREMENT
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_TO_GOAL_DISTANCE = CN()
_C.TASK.OBJECT_TO_GOAL_DISTANCE.TYPE = "ObjectToGoalDistance"
# -----------------------------------------------------------------------------
# # OBJECT_DISTANCE_FROM_AGENT MEASUREMENT
# -----------------------------------------------------------------------------
_C.TASK.AGENT_TO_OBJECT_DISTANCE = CN()
_C.TASK.AGENT_TO_OBJECT_DISTANCE.TYPE = "AgentToObjectDistance"
from habitat.config.default import CN, Config
# %%
# @title Define `RearrangementTask` by extending `NavigationTask`
from habitat.tasks.nav.nav import NavigationTask, merge_sim_episode_config
def merge_sim_episode_with_object_config(
sim_config: Config, episode: Type[Episode]
) -> Any:
sim_config = merge_sim_episode_config(sim_config, episode)
sim_config.defrost()
sim_config.objects = [episode.objects.__dict__]
sim_config.freeze()
return sim_config
@registry.register_task(name="RearrangementTask-v0")
class RearrangementTask(NavigationTask):
r"""Embodied Rearrangement Task
Goal: An agent must place objects at their corresponding goal position.
"""
def __init__(self, **kwargs) -> None:
super().__init__(**kwargs)
def overwrite_sim_config(self, sim_config, episode):
return merge_sim_episode_with_object_config(sim_config, episode)
# %% [markdown]
# ## Implement a hard-coded and an RL agent
#
#
# %%
# @title Load the `RearrangementTask` in Habitat-Lab and run a hard-coded agent
import habitat
config = habitat.get_config("configs/tasks/pointnav.yaml")
config.defrost()
config.ENVIRONMENT.MAX_EPISODE_STEPS = 50
config.SIMULATOR.TYPE = "RearrangementSim-v0"
config.SIMULATOR.ACTION_SPACE_CONFIG = "RearrangementActions-v0"
config.SIMULATOR.GRAB_DISTANCE = 2.0
config.SIMULATOR.HABITAT_SIM_V0.ENABLE_PHYSICS = True
config.TASK.TYPE = "RearrangementTask-v0"
config.TASK.SUCCESS_DISTANCE = 1.0
config.TASK.SENSORS = [
"GRIPPED_OBJECT_SENSOR",
"OBJECT_POSITION",
"OBJECT_GOAL",
]
config.TASK.GOAL_SENSOR_UUID = "object_goal"
config.TASK.MEASUREMENTS = [
"OBJECT_TO_GOAL_DISTANCE",
"AGENT_TO_OBJECT_DISTANCE",
]
config.TASK.POSSIBLE_ACTIONS = ["STOP", "MOVE_FORWARD", "GRAB_RELEASE"]
config.DATASET.TYPE = "RearrangementDataset-v0"
config.DATASET.SPLIT = "train"
config.DATASET.DATA_PATH = (
"data/datasets/rearrangement/coda/v1/{split}/{split}.json.gz"
)
config.freeze()
def print_info(obs, metrics):
print(
"Gripped Object: {}, Distance To Object: {}, Distance To Goal: {}".format(
obs["gripped_object_id"],
metrics["agent_to_object_distance"],
metrics["object_to_goal_distance"],
)
)
try: # Got to make initialization idiot proof
sim.close()
except NameError:
pass
with habitat.Env(config) as env:
obs = env.reset()
obs_list = []
# Get closer to the object
while True:
obs = env.step(1)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
if metrics["agent_to_object_distance"] < 2.0:
break
# Grab the object
obs = env.step(2)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
assert obs["gripped_object_id"] != -1
# Get closer to the goal
while True:
obs = env.step(1)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
if metrics["object_to_goal_distance"] < 2.0:
break
# Release the object
obs = env.step(2)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
assert obs["gripped_object_id"] == -1
if make_video:
make_video_cv2(
obs_list,
[190, 128],
"hard-coded-agent",
fps=5.0,
open_vid=show_video,
)
# %%
# @title Create a task specific RL Environment with a new reward definition.
# @markdown We create a `RearragenmentRLEnv` class and modify the `get_reward()` function.
# @markdown The reward sturcture is as follows:
# @markdown - The agent gets a positive reward if the agent gets closer to the object otherwise a negative reward.
# @markdown - The agent gets a positive reward if it moves the object closer to goal otherwise a negative reward.
# @markdown - The agent gets a positive reward when the agent "picks" up an object for the first time. For all other "grab/release" action, it gets a negative reward.
# @markdown - The agent gets a slack penalty of -0.01 for every action it takes in the environment.
# @markdown - Finally the agent gets a large success reward when the episode is completed successfully.
from typing import Optional, Type
import numpy as np
import habitat
from habitat import Config, Dataset
from habitat_baselines.common.baseline_registry import baseline_registry
from habitat_baselines.common.environments import NavRLEnv
@baseline_registry.register_env(name="RearrangementRLEnv")
class RearrangementRLEnv(NavRLEnv):
def __init__(self, config: Config, dataset: Optional[Dataset] = None):
self._prev_measure = {
"agent_to_object_distance": 0.0,
"object_to_goal_distance": 0.0,
"gripped_object_id": -1,
"gripped_object_count": 0,
}
super().__init__(config, dataset)
self._success_distance = self._core_env_config.TASK.SUCCESS_DISTANCE
def reset(self):
self._previous_action = None
observations = super().reset()
self._prev_measure.update(self.habitat_env.get_metrics())
self._prev_measure["gripped_object_id"] = -1
self._prev_measure["gripped_object_count"] = 0
return observations
def step(self, *args, **kwargs):
self._previous_action = kwargs["action"]
return super().step(*args, **kwargs)
def get_reward_range(self):
return (
self._rl_config.SLACK_REWARD - 1.0,
self._rl_config.SUCCESS_REWARD + 1.0,
)
def get_reward(self, observations):
reward = self._rl_config.SLACK_REWARD
gripped_success_reward = 0.0
episode_success_reward = 0.0
agent_to_object_dist_reward = 0.0
object_to_goal_dist_reward = 0.0
action_name = self._env.task.get_action_name(
self._previous_action["action"]
)
# If object grabbed, add a success reward
# The reward gets awarded only once for an object.
if (
action_name == "GRAB_RELEASE"
and observations["gripped_object_id"] >= 0
):
obj_id = observations["gripped_object_id"]
self._prev_measure["gripped_object_count"] += 1
gripped_success_reward = (
self._rl_config.GRIPPED_SUCCESS_REWARD
if self._prev_measure["gripped_object_count"] == 1
else 0.0
)
# add a penalty everytime grab/action is called and doesn't do anything
elif action_name == "GRAB_RELEASE":
gripped_success_reward += -0.1
self._prev_measure["gripped_object_id"] = observations[
"gripped_object_id"
]
# If the action is not a grab/release action, and the agent
# has not picked up an object, then give reward based on agent to
# object distance.
if (
action_name != "GRAB_RELEASE"
and self._prev_measure["gripped_object_id"] == -1
):
agent_to_object_dist_reward = self.get_agent_to_object_dist_reward(
observations
)
# If the action is not a grab/release action, and the agent
# has picked up an object, then give reward based on object to
# to goal distance.
if (
action_name != "GRAB_RELEASE"
and self._prev_measure["gripped_object_id"] != -1
):
object_to_goal_dist_reward = self.get_object_to_goal_dist_reward()
if (
self._episode_success(observations)
and self._prev_measure["gripped_object_id"] == -1
and action_name == "STOP"
):
episode_success_reward = self._rl_config.SUCCESS_REWARD
reward += (
agent_to_object_dist_reward
+ object_to_goal_dist_reward
+ gripped_success_reward
+ episode_success_reward
)
return reward
def get_agent_to_object_dist_reward(self, observations):
"""
Encourage the agent to move towards the closest object which is not already in place.
"""
curr_metric = self._env.get_metrics()["agent_to_object_distance"]
prev_metric = self._prev_measure["agent_to_object_distance"]
dist_reward = prev_metric - curr_metric
self._prev_measure["agent_to_object_distance"] = curr_metric
return dist_reward
def get_object_to_goal_dist_reward(self):
curr_metric = self._env.get_metrics()["object_to_goal_distance"]
prev_metric = self._prev_measure["object_to_goal_distance"]
dist_reward = prev_metric - curr_metric
self._prev_measure["object_to_goal_distance"] = curr_metric
return dist_reward
def _episode_success(self, observations):
r"""Returns True if object is within distance threshold of the goal."""
dist = self._env.get_metrics()["object_to_goal_distance"]
if (
abs(dist) > self._success_distance
or observations["gripped_object_id"] != -1
):
return False
return True
def _gripped_success(self, observations):
if (
observations["gripped_object_id"] >= 0
and observations["gripped_object_id"]
!= self._prev_measure["gripped_object_id"]
):
return True
return False
def get_done(self, observations):
done = False
action_name = self._env.task.get_action_name(
self._previous_action["action"]
)
if self._env.episode_over or (
self._episode_success(observations)
and self._prev_measure["gripped_object_id"] == -1
and action_name == "STOP"
):
done = True
return done
def get_info(self, observations):
info = self.habitat_env.get_metrics()
info["episode_success"] = self._episode_success(observations)
return info
# %%
import os
import time
from typing import Any, Dict, List, Optional
import numpy as np
from torch.optim.lr_scheduler import LambdaLR
from habitat import Config, logger
from habitat.utils.visualizations.utils import observations_to_image
from habitat_baselines.common.baseline_registry import baseline_registry
from habitat_baselines.common.environments import get_env_class
from habitat_baselines.common.tensorboard_utils import TensorboardWriter
from habitat_baselines.rl.models.rnn_state_encoder import (
build_rnn_state_encoder,
)
from habitat_baselines.rl.ppo import PPO
from habitat_baselines.rl.ppo.policy import Net, Policy
from habitat_baselines.rl.ppo.ppo_trainer import PPOTrainer
from habitat_baselines.utils.common import batch_obs, generate_video
from habitat_baselines.utils.env_utils import make_env_fn
def construct_envs(
config,
env_class,
workers_ignore_signals=False,
):
r"""Create VectorEnv object with specified config and env class type.
To allow better performance, dataset are split into small ones for
each individual env, grouped by scenes.
:param config: configs that contain num_processes as well as information
:param necessary to create individual environments.
:param env_class: class type of the envs to be created.
:param workers_ignore_signals: Passed to :ref:`habitat.VectorEnv`'s constructor
:return: VectorEnv object created according to specification.
"""
num_processes = config.NUM_ENVIRONMENTS
configs = []
env_classes = [env_class for _ in range(num_processes)]
dataset = habitat.datasets.make_dataset(config.TASK_CONFIG.DATASET.TYPE)
scenes = config.TASK_CONFIG.DATASET.CONTENT_SCENES
if "*" in config.TASK_CONFIG.DATASET.CONTENT_SCENES:
scenes = dataset.get_scenes_to_load(config.TASK_CONFIG.DATASET)
if num_processes > 1:
if len(scenes) == 0:
raise RuntimeError(
"No scenes to load, multiple process logic relies on being able to split scenes uniquely between processes"
)
if len(scenes) < num_processes:
scenes = scenes * num_processes
random.shuffle(scenes)
scene_splits = [[] for _ in range(num_processes)]
for idx, scene in enumerate(scenes):
scene_splits[idx % len(scene_splits)].append(scene)
assert sum(map(len, scene_splits)) == len(scenes)
for i in range(num_processes):
proc_config = config.clone()
proc_config.defrost()
task_config = proc_config.TASK_CONFIG
task_config.SEED = task_config.SEED + i
if len(scenes) > 0:
task_config.DATASET.CONTENT_SCENES = scene_splits[i]
task_config.SIMULATOR.HABITAT_SIM_V0.GPU_DEVICE_ID = (
config.SIMULATOR_GPU_ID
)
task_config.SIMULATOR.AGENT_0.SENSORS = config.SENSORS
proc_config.freeze()
configs.append(proc_config)
envs = habitat.ThreadedVectorEnv(
make_env_fn=make_env_fn,
env_fn_args=tuple(zip(configs, env_classes)),
workers_ignore_signals=workers_ignore_signals,
)
return envs
class RearrangementBaselinePolicy(Policy):
def __init__(self, observation_space, action_space, hidden_size=512):
super().__init__(
RearrangementBaselineNet(
observation_space=observation_space, hidden_size=hidden_size
),
action_space.n,
)
def from_config(cls, config, envs):
pass
class RearrangementBaselineNet(Net):
r"""Network which passes the input image through CNN and concatenates
goal vector with CNN's output and passes that through RNN.
"""
def __init__(self, observation_space, hidden_size):
super().__init__()
self._n_input_goal = observation_space.spaces[
ObjectGoal.cls_uuid
].shape[0]
self._hidden_size = hidden_size
self.state_encoder = build_rnn_state_encoder(
2 * self._n_input_goal, self._hidden_size
)
self.train()
@property
def output_size(self):
return self._hidden_size
@property
def is_blind(self):
return False
@property
def num_recurrent_layers(self):
return self.state_encoder.num_recurrent_layers
def forward(self, observations, rnn_hidden_states, prev_actions, masks):
object_goal_encoding = observations[ObjectGoal.cls_uuid]
object_pos_encoding = observations[ObjectPosition.cls_uuid]
x = [object_goal_encoding, object_pos_encoding]
x = torch.cat(x, dim=1)
x, rnn_hidden_states = self.state_encoder(x, rnn_hidden_states, masks)
return x, rnn_hidden_states
@baseline_registry.register_trainer(name="ppo-rearrangement")
class RearrangementTrainer(PPOTrainer):
supported_tasks = ["RearrangementTask-v0"]
def _setup_actor_critic_agent(self, ppo_cfg: Config) -> None:
r"""Sets up actor critic and agent for PPO.
Args:
ppo_cfg: config node with relevant params
Returns:
None
"""
logger.add_filehandler(self.config.LOG_FILE)
self.actor_critic = RearrangementBaselinePolicy(
observation_space=self.envs.observation_spaces[0],
action_space=self.envs.action_spaces[0],
hidden_size=ppo_cfg.hidden_size,
)
self.actor_critic.to(self.device)
self.agent = PPO(
actor_critic=self.actor_critic,
clip_param=ppo_cfg.clip_param,
ppo_epoch=ppo_cfg.ppo_epoch,
num_mini_batch=ppo_cfg.num_mini_batch,
value_loss_coef=ppo_cfg.value_loss_coef,
entropy_coef=ppo_cfg.entropy_coef,
lr=ppo_cfg.lr,
eps=ppo_cfg.eps,
max_grad_norm=ppo_cfg.max_grad_norm,
use_normalized_advantage=ppo_cfg.use_normalized_advantage,
)
def _init_envs(self, config=None):
if config is None:
config = self.config
self.envs = construct_envs(config, get_env_class(config.ENV_NAME))
def train(self) -> None:
r"""Main method for training PPO.
Returns:
None
"""
if self._is_distributed:
raise RuntimeError("This trainer does not support distributed")
self._init_train()
count_checkpoints = 0
lr_scheduler = LambdaLR(
optimizer=self.agent.optimizer,
lr_lambda=lambda _: 1 - self.percent_done(),
)
ppo_cfg = self.config.RL.PPO
with TensorboardWriter(
self.config.TENSORBOARD_DIR, flush_secs=self.flush_secs
) as writer:
while not self.is_done():
if ppo_cfg.use_linear_clip_decay:
self.agent.clip_param = ppo_cfg.clip_param * (
1 - self.percent_done()
)
count_steps_delta = 0
for _step in range(ppo_cfg.num_steps):
count_steps_delta += self._collect_rollout_step()
(
value_loss,
action_loss,
dist_entropy,
) = self._update_agent()
if ppo_cfg.use_linear_lr_decay:
lr_scheduler.step() # type: ignore
losses = self._coalesce_post_step(
dict(value_loss=value_loss, action_loss=action_loss),
count_steps_delta,
)
self.num_updates_done += 1
deltas = {
k: (
(v[-1] - v[0]).sum().item()
if len(v) > 1
else v[0].sum().item()
)
for k, v in self.window_episode_stats.items()
}
deltas["count"] = max(deltas["count"], 1.0)
writer.add_scalar(
"reward",
deltas["reward"] / deltas["count"],
self.num_steps_done,
)
# Check to see if there are any metrics
# that haven't been logged yet
for k, v in deltas.items():
if k not in {"reward", "count"}:
writer.add_scalar(
"metric/" + k,
v / deltas["count"],
self.num_steps_done,
)
losses = [value_loss, action_loss]
for l, k in zip(losses, ["value, policy"]):
writer.add_scalar("losses/" + k, l, self.num_steps_done)
# log stats
if self.num_updates_done % self.config.LOG_INTERVAL == 0:
logger.info(
"update: {}\tfps: {:.3f}\t".format(
self.num_updates_done,
self.num_steps_done / (time.time() - self.t_start),
)
)
logger.info(
"update: {}\tenv-time: {:.3f}s\tpth-time: {:.3f}s\t"
"frames: {}".format(
self.num_updates_done,
self.env_time,
self.pth_time,
self.num_steps_done,
)
)
logger.info(
"Average window size: {} {}".format(
len(self.window_episode_stats["count"]),
" ".join(
"{}: {:.3f}".format(k, v / deltas["count"])
for k, v in deltas.items()
if k != "count"
),
)
)
# checkpoint model
if self.should_checkpoint():
self.save_checkpoint(
f"ckpt.{count_checkpoints}.pth",
dict(step=self.num_steps_done),
)
count_checkpoints += 1
self.envs.close()
def eval(self) -> None:
r"""Evaluates the current model
Returns:
None
"""
config = self.config.clone()
if len(self.config.VIDEO_OPTION) > 0:
config.defrost()
config.NUM_ENVIRONMENTS = 1
config.freeze()
logger.info(f"env config: {config}")
with construct_envs(config, get_env_class(config.ENV_NAME)) as envs:
observations = envs.reset()
batch = batch_obs(observations, device=self.device)
current_episode_reward = torch.zeros(
envs.num_envs, 1, device=self.device
)
ppo_cfg = self.config.RL.PPO
test_recurrent_hidden_states = torch.zeros(
config.NUM_ENVIRONMENTS,
self.actor_critic.net.num_recurrent_layers,
ppo_cfg.hidden_size,
device=self.device,
)
prev_actions = torch.zeros(
config.NUM_ENVIRONMENTS,
1,
device=self.device,
dtype=torch.long,
)
not_done_masks = torch.zeros(
config.NUM_ENVIRONMENTS,
1,
device=self.device,
dtype=torch.bool,
)
rgb_frames = [
[] for _ in range(self.config.NUM_ENVIRONMENTS)
] # type: List[List[np.ndarray]]
if len(config.VIDEO_OPTION) > 0:
os.makedirs(config.VIDEO_DIR, exist_ok=True)
self.actor_critic.eval()
for _i in range(config.TASK_CONFIG.ENVIRONMENT.MAX_EPISODE_STEPS):
current_episodes = envs.current_episodes()
with torch.no_grad():
(
_,
actions,
_,
test_recurrent_hidden_states,
) = self.actor_critic.act(
batch,
test_recurrent_hidden_states,
prev_actions,
not_done_masks,
deterministic=False,
)
prev_actions.copy_(actions)
outputs = envs.step([a[0].item() for a in actions])
observations, rewards, dones, infos = [
list(x) for x in zip(*outputs)
]
batch = batch_obs(observations, device=self.device)
not_done_masks = torch.tensor(
[[not done] for done in dones],
dtype=torch.bool,
device="cpu",
)
rewards = torch.tensor(
rewards, dtype=torch.float, device=self.device
).unsqueeze(1)
current_episode_reward += rewards
# episode ended
if not not_done_masks[0].item():
generate_video(
video_option=self.config.VIDEO_OPTION,
video_dir=self.config.VIDEO_DIR,
images=rgb_frames[0],
episode_id=current_episodes[0].episode_id,
checkpoint_idx=0,
metrics=self._extract_scalars_from_info(infos[0]),
tb_writer=None,
)
print("Evaluation Finished.")
print("Success: {}".format(infos[0]["episode_success"]))
print(
"Reward: {}".format(current_episode_reward[0].item())
)
print(
"Distance To Goal: {}".format(
infos[0]["object_to_goal_distance"]
)
)
return
# episode continues
elif len(self.config.VIDEO_OPTION) > 0:
frame = observations_to_image(observations[0], infos[0])
rgb_frames[0].append(frame)
not_done_masks = not_done_masks.to(device=self.device)
# %%
# %load_ext tensorboard
# %tensorboard --logdir data/tb
# %%
# @title Train an RL agent on a single episode
# !if [ -d "data/tb" ]; then rm -r data/tb; fi
import random
import numpy as np
import torch
import habitat
from habitat import Config
from habitat_baselines.config.default import get_config as get_baseline_config
baseline_config = get_baseline_config(
"habitat_baselines/config/pointnav/ppo_pointnav.yaml"
)
baseline_config.defrost()
baseline_config.TASK_CONFIG = config
baseline_config.TRAINER_NAME = "ddppo"
baseline_config.ENV_NAME = "RearrangementRLEnv"
baseline_config.SIMULATOR_GPU_ID = 0
baseline_config.TORCH_GPU_ID = 0
baseline_config.VIDEO_OPTION = ["disk"]
baseline_config.TENSORBOARD_DIR = "data/tb"
baseline_config.VIDEO_DIR = "data/videos"
baseline_config.NUM_ENVIRONMENTS = 2
baseline_config.SENSORS = ["RGB_SENSOR", "DEPTH_SENSOR"]
baseline_config.CHECKPOINT_FOLDER = "data/checkpoints"
baseline_config.TOTAL_NUM_STEPS = -1.0
if vut.is_notebook():
baseline_config.NUM_UPDATES = 400 # @param {type:"number"}
else:
baseline_config.NUM_UPDATES = 1
baseline_config.LOG_INTERVAL = 10
baseline_config.NUM_CHECKPOINTS = 5
baseline_config.LOG_FILE = "data/checkpoints/train.log"
baseline_config.EVAL.SPLIT = "train"
baseline_config.RL.SUCCESS_REWARD = 2.5 # @param {type:"number"}
baseline_config.RL.SUCCESS_MEASURE = "object_to_goal_distance"
baseline_config.RL.REWARD_MEASURE = "object_to_goal_distance"
baseline_config.RL.GRIPPED_SUCCESS_REWARD = 2.5 # @param {type:"number"}
baseline_config.freeze()
random.seed(baseline_config.TASK_CONFIG.SEED)
np.random.seed(baseline_config.TASK_CONFIG.SEED)
torch.manual_seed(baseline_config.TASK_CONFIG.SEED)
if __name__ == "__main__":
trainer = RearrangementTrainer(baseline_config)
trainer.train()
trainer.eval()
if make_video:
video_file = os.listdir("data/videos")[0]
vut.display_video(os.path.join("data/videos", video_file))
| examples/tutorials/colabs/Habitat_Interactive_Tasks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Lesson 2 Assignment JNB -- Kaggle Galaxy Zoo
#
# 10 May 2017 - <NAME>
#
# Plan:
# 1. Build a Linear Model (no activations) from scratch
# 2. Build a 1-Layer Neural Network using linear model layers + activations
# 3. Build a finetuned DLNN atop VGG16
#
#
# * experiment w/ SGD vs RMSprop
# * experiment w/ sigmoid vs tanh vs ReLU
# * compare scores of ea. model
# * use utils.py & vgg16.py source code + Keras.io documentation for help
#
# Note: I'm pretty sure that by "*from scratch*" what <NAME> means is a fresh linear model atop Vgg16.. Creating a straight Linear Model for image classification... does not sound... very sound..
#
# Whatever, build break learn.
import keras
# Constants
Num_Classes = Num_Classes
batch_size = 4
lr = 0.01
# +
# Helper Functions
# get_batches(..) copied from utils.py
# gen.flow_from_directory() is an iterator that yields batches of images
# from a directory indefinitely.
from keras.preprocessing import image
def get_batches(dirname, gen=image.ImageDataGenerator(), shuffle=True, batche_size=4, class_mode='categorical',
target_size=(224,224)):
return gen.flow_from_directory(dirname, target_size=target_size,
class_mode=class_mode, shuffle=shuffle, batch_size=batch_size)
# fast array saving/loading
import bcolz
def save_array(fname, arr): c=bcolz.carray(arr, rootdir=fname, mode='w'); c.flush()
def load_array(fname): return bcolz.open(fname)[:]
# One-Hot Encoding for Keras
from sklearn.preprocessing import OneHotEncoder
def onehot(x): return np.array(OneHotEncoder().fit_transform(x.reshape(-1, 1))).todense()
# should I use that or from Keras?
# def onehot(x): return keras.utils.np_utils.to_categorical(x)
# from utils.py -- retrieving data saved by bcolz
def get_data(path, target_size=(224,224)):
batches = get_batches(path, shuffle=False, batch_size=1, class_mode=None, target_size=target_size)
return np.concatenate([batches.next() for i in range(batches.nb_sample)])
# -
# ### 1. Basic Linear Model
# +
LM = keras.model.Sequential([Dense(Num_Classes, input_shape=(784,))])
LM.compile(optimizer=SGD(lr=0.01), loss='mse')
# LM.compile(optimizer=RMSprop(lr=0.01), loss='mse')
# -
# ### 2. 1-Layer Neural Network
# ### 3. Finetuned VGG16
import os, sys
sys.path.insert(os.path.join(1, '../utils/'))
import Vgg16
| FAI_old/lesson2/L2HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/medlandolsi/landolsi/blob/main/exe_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="js0-DBzUrQJK" outputId="b6c4a7d9-d618-4110-d851-9cb6976bf5c1"
hello = 'hello world'
print (hello)
# + colab={"base_uri": "https://localhost:8080/"} id="Xgjf6v_Ps1Z4" outputId="5bec50e3-86f3-4cf2-fb00-478a3f08a0fe"
print(hello,'bonjour le monde')
# + colab={"base_uri": "https://localhost:8080/"} id="Zfdm6rQptQfh" outputId="64bad9bd-c2fd-47eb-f9d8-c35039e27e0d"
prix =3000
print(prix)
a=10000; b=20000; c=30000
print(a+b/c)
del prix
print('hnjxdnvjdb')
# + colab={"base_uri": "https://localhost:8080/"} id="eJHx06rnvAxJ" outputId="b7941548-1b3c-403d-df86-7aa1199eabe4"
my_int =2.5
type(my_int)
| exe_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: numSolve_parallel
# language: python
# name: numsolve_parallel
# ---
# ## <NAME>
# ### 16 Jan 2020
# ### Simulate data for training neural network
# ### This uses the "one torque" or the "underactuated" model
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import os
import pandas as pd
import seaborn as sns
from scipy.integrate import odeint
import random
import time
from datetime import datetime
import sys
from multiprocessing import Pool, cpu_count
import simUtils_one_torque # note that this is a custom-written file
import importlib
import functools
import sqlite3
from collections import OrderedDict
print(sys.version)
# +
now = datetime.now()
print("last run on " + str(now))
pythonMadeData = r"D:/Dropbox/AcademiaDropbox/mothMachineLearning_dataAndFigs/PythonGeneratedData_oneTorque"
if not os.path.exists(pythonMadeData):
os.mkdir(pythonMadeData)
# -
np.random.seed(12345)
_ = importlib.reload(simUtils_one_torque)
# +
# save global options
globalDict = OrderedDict({
"bhead": 0.5,
"ahead": 0.9,
"bbutt": 0.75,
"abutt": 1.9,
"rho_head": 0.9,
"rho_butt": 0.4,
"rhoA": 0.00118,
"muA": 0.000186,
"L1": 0.9,
"L2": 1.9,
"L3": 0.75,
"K": 23000,
"c": 14075.8,
"g": 980.0,
"betaR": 0.0,
"nstep": 2, # return start and endpoints
"nrun" : 1000000 # (max) number of trajectories.
})
# Calculated variables
globalDict['m1'] = globalDict['rho_head']*(4/3)*np.pi*(globalDict['bhead']**2)*globalDict['ahead']
globalDict["m2"] = globalDict["rho_butt"]*(4/3)*np.pi*(globalDict["bbutt"]**2)*globalDict["abutt"]
globalDict["echead"] = globalDict["ahead"]/globalDict["bhead"]
globalDict['ecbutt'] = globalDict['abutt']/globalDict['bbutt']
globalDict['I1'] = (1/5)*globalDict['m1']*(globalDict['bhead']**2)*(1 + globalDict['echead']**2)
globalDict['I2'] = (1/5)*globalDict['m2']*(globalDict['bbutt']**2)*(1 + globalDict['ecbutt']**2)
globalDict['S_head'] = np.pi*globalDict['bhead']**2
globalDict['S_butt'] = np.pi*globalDict['bbutt'] **2
t = np.linspace(0, 0.02, num = globalDict["nstep"], endpoint = True)
# convert dict to list, since @jit works better with lists
globalList = [ v for v in globalDict.values() ]
# ranges for control variables
rangeDict = {"Fmin": 0,
"Fmax": 44300,
"alphaMin": 0,
"alphaMax":2*np.pi,
"tau0Min": -100000,
"tau0Max": 100000}
# ranges for controls
ranges = np.array([[rangeDict["Fmin"], rangeDict["Fmax"]],
[rangeDict["alphaMin"], rangeDict["alphaMax"]],
[rangeDict["tau0Min"], rangeDict["tau0Max"] ]])
# ranges for initial conditions
IC_ranges = np.array([[0, 0], #x
[-1500, 1500], #xdot
[0, 0], #y
[-1500, 1500], #ydot
[0, 2*np.pi], #theta
[-25, 25], #theta dot
[0, 2*np.pi], #phi
[-25, 25]]) # phi dot
# +
# generate training data
dataType = "trainingData_"
for ii in np.arange(0,10):
print(ii)
# generate random ICs and controls
# random F, alpha, tau, tau_w
FAlphaTau_list = np.random.uniform(ranges[:, 0], ranges[:, 1],
size=(globalDict["nrun"], ranges.shape[0]))
# random initial conditions for state 0
state0_ICs = np.random.uniform(IC_ranges[:, 0], IC_ranges[:, 1], size=(globalDict["nrun"], IC_ranges.shape[0]))
# run simulations in parallel, "nrun"s at a time
p = Pool(cpu_count() - 2)
stt = time.time()
bb = p.map(functools.partial(simUtils_one_torque.flyBug_listInput_oneTorque, t=t,
state0_ICs = state0_ICs,
FAlphaTau_list= FAlphaTau_list,
globalList = globalList), range(globalDict["nrun"]))
print("time for one run:", time.time() - stt)
p.close()
p.join()
# reshape to put into a pd data frame
bb2 = np.array(bb).reshape(globalDict["nrun"], -1, order = "F")
bb3 = np.hstack([bb2, FAlphaTau_list])
simDF = pd.DataFrame(bb3, columns = ["x_0", "xd_0","y_0","yd_0",
"theta_0","thetad_0","phi_0","phid_0",
"x_f", "xd_f","y_f","yd_f",
"theta_f","thetad_f","phi_f","phid_f",
"F", "alpha", "tau0"])
# write to database,
# makes a new database if it doesn't already exist
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
# get table names from database
try:
cursorObj = con1.cursor()
cursorObj.execute('SELECT name from sqlite_master where type= "table"')
tableNames = cursorObj.fetchall()
cursorObj.close()
except:
print("can't get table names")
# refref: name changed from "trainingData_" to "testingData_" when I generated new data
simDF.to_sql(dataType + str(len(tableNames)).zfill(2), con1, if_exists = "fail", index = False)
# close connection
con1.close()
# -
dataType = "testingData_"
for ii in np.arange(0,5):
print(ii)
# generate random ICs and controls
# random F, alpha, tau, tau_w
FAlphaTau_list = np.random.uniform(ranges[:, 0], ranges[:, 1],
size=(globalDict["nrun"], ranges.shape[0]))
# random initial conditions for state 0
state0_ICs = np.random.uniform(IC_ranges[:, 0], IC_ranges[:, 1], size=(globalDict["nrun"], IC_ranges.shape[0]))
# run simulations in parallel, "nrun"s at a time
p = Pool(cpu_count() - 2)
stt = time.time()
bb = p.map(functools.partial(simUtils_one_torque.flyBug_listInput_oneTorque, t=t,
state0_ICs = state0_ICs,
FAlphaTau_list= FAlphaTau_list,
globalList = globalList), range(globalDict["nrun"]))
print("time for one run:", time.time() - stt)
p.close()
p.join()
# reshape to put into a pd data frame
bb2 = np.array(bb).reshape(globalDict["nrun"], -1, order = "F")
bb3 = np.hstack([bb2, FAlphaTau_list])
simDF = pd.DataFrame(bb3, columns = ["x_0", "xd_0","y_0","yd_0",
"theta_0","thetad_0","phi_0","phid_0",
"x_f", "xd_f","y_f","yd_f",
"theta_f","thetad_f","phi_f","phid_f",
"F", "alpha", "tau0"])
# write to database,
# makes a new database if it doesn't already exist
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
# get table names from database
try:
cursorObj = con1.cursor()
cursorObj.execute('SELECT name from sqlite_master where type= "table"')
tableNames = cursorObj.fetchall()
cursorObj.close()
except:
print("can't get table names")
# refref: name changed from "trainingData_" to "testingData_" when I generated new data
simDF.to_sql(dataType + str(len(tableNames)).zfill(2), con1, if_exists = "fail", index = False)
# close connection
con1.close()
# get table names in database
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
cursorObj = con1.cursor()
res = cursorObj.execute("SELECT name FROM sqlite_master WHERE type='table';")
tableNames = [name[0] for name in res]
con1.close()
print(tableNames)
# Combine testing Data into a single Table
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
con1.execute("DROP TABLE IF EXISTS test")
sqlStatement = "CREATE TABLE test AS " + " UNION ALL ".join(["SELECT * FROM " + tableNames[ii] for ii in range(len(tableNames)) if tableNames[ii].startswith("testingData_")])
print(sqlStatement)
con1.execute(sqlStatement)
con1.close()
# Combine Training Data into a single Table
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
con1.execute("DROP TABLE IF EXISTS train")
sqlStatement = "CREATE TABLE train AS " + " UNION ALL ".join(["SELECT * FROM " + tableNames[ii] for ii in range(len(tableNames)) if tableNames[ii].startswith("trainingData_")])
print(sqlStatement)
con1.execute(sqlStatement)
con1.close()
# +
# print print the max row number
def largestRowNumber(cursor, table_name, print_out=False):
""" Returns the total number of rows in the database """
cursor.execute("SELECT max(rowid) from {}".format(table_name))
n = cursor.fetchone()[0]
if print_out:
print('\nTotal rows: {}'.format(n))
return(n)
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
cursorObj = con1.cursor()
largestRowNumber(cursorObj, "train", print_out=True)
largestRowNumber(cursorObj, "test", print_out=True)
con1.close()
# -
# drop intermediate, smaller training datasets
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
sqlStatement = "".join(["DROP TABLE IF EXISTS " + tableNames[ii] + "; " for ii in range(len(tableNames)) if tableNames[ii].startswith("trainingData_")])
print(sqlStatement)
con1.executescript(sqlStatement)
con1.close()
# drop intermediate, smaller testing datasets
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
sqlStatement = "".join(["DROP TABLE IF EXISTS " + tableNames[ii] + "; " for ii in range(len(tableNames)) if tableNames[ii].startswith("testingData_")])
print(sqlStatement)
con1.executescript(sqlStatement)
con1.close()
# get table names in database
con1 = sqlite3.connect(os.path.join(pythonMadeData, "oneTorqueData.db"))
cursorObj = con1.cursor()
res = cursorObj.execute("SELECT name FROM sqlite_master WHERE type='table';")
tableNames = [name[0] for name in res]
con1.close()
print(tableNames)
| PythonCode/one_torque_model/010_OneTorqueParallelSims.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import collections
# +
Card = collections.namedtuple('Card', ['rank', 'suit'])
class FrenchDeck2(collections.abc.MutableSequence):
ranks = [str(n) for n in range(2, 11)] + list('JQKA')
suits = "spades diamonds clubs hearts".split()
def __init__(self):
self._cards = [Card(rank, suit) for suit in self.suits
for rank in self.ranks]
def __len__(self):
return len(self._cards)
def __getitem__(self, position):
return self._cards[position]
def __setitem__(self, position, value):
self._cards[position] = value
def __delitem__(self, position):
del self._cards[position]
def insert(self, position, value):
self._cards.insert(position, value)
# -
card = FrenchDeck2()
for c in card:
print(c)
Card(rank='2', suit='spades') in card
# ## 猴子补丁
class Monkey:
def __init__(self):
self.a = 5
def func1(self):
print('it is func1')
def func2(self):
print('it is func2')
m = Monkey()
m.func1()
# +
def func3(self):
print(self.a)
print('it is monkey patch')
Monkey.func3 = func3
# -
m.func3()
# **Python会特殊看待看起来像是序列的对象**
# 虽然没有 `__iter__` 方法,但是 Foo 实例是可迭代的对象,因为发现有 `__getitem__` 方法时,Python 会调用它,传入从 0 开始的整数索引, 尝试迭代对象(这是一种后备机制)。尽管没有实现 `__contains__` 方法,但是 Python 足够智能,能迭代 Foo 实例,因此也能使用 in 运算 符:Python 会做全面检查,看看有没有指定的元素。 综上,鉴于序列协议的重要性,如果没有 `__iter__ `和 `__contains__` 方法,Python 会调用 `__getitem__` 方法,设法让迭代和 in 运算符可用。
class FrenchDeck3:
ranks = [str(n) for n in range(2, 11)] + list('JQKA')
suits = "spades diamonds clubs hearts".split()
def __init__(self):
self._cards = [Card(rank, suit) for suit in self.suits
for rank in self.ranks]
def __len__(self):
return len(self._cards)
def __getitem__(self, position):
return self._cards[position]
def __setitem__(self, position, value):
self._cards[position] = value
def __delitem__(self, position):
del self._cards[position]
def insert(self, position, value):
self._cards.insert(position, value)
card2 = FrenchDeck3()
for c in card2:
print(c)
Card(rank='2', suit='spades') in card2
# ## 自定义抽象基类
# + **抽象基类中也可以包含具体方法,抽象基类中的具体方法只能依赖抽象基类中定义的接口(即只能使用抽象基类中的其它具体方法、抽象方法或特性)**
# + **抽象基类中抽象方法也可以有实现代码,但是即便是实现了,子类中也必须要覆盖该方法,子类中可以调用super()方法为它添加功能,而不是从头实现**
# + **抽象方法只存在于抽象基类中**
# + **抽象基类不可以实例化**
# + **继承抽象基类的子类,必须覆盖抽象基类中的所有抽象方法,否则,它也将被当作另一种抽象基类【类似C++中】**
# + **继承抽象基类的子类,可以选择性覆盖抽象基类中的具体方法**
# +
import abc
class Tombola(abc.ABC):
# 定义抽象方法
@abc.abstractmethod
def load(self, iterable):
""""""
# 定义抽象方法
@abc.abstractmethod
def pick(self):
""""""
# 抽象基类中的具体方法
def loaded(self):
return bool(self.inspect())
# 抽象基类中的具体方法
def inspect(self):
items = []
while True:
try:
items.append(self.pick())
except LookupError:
break
self.load(items)
return tuple(sorted(items))
# +
# Fake继承自抽象基类Tombola,但它没有完全覆盖所有抽象方法
class Fake(Tombola):
def pick(self):
return 0
# -
Fake
# +
# Fake被认为是一种抽象基类,因此不能被实例化
f = Fake()
# + **声明抽象基类的最简单方式是继承abc.ABC或其他抽象基类**
# +
import random
# 继承抽象基类Tombola
class BingoCage(Tombola):
def __init__(self, items):
self._randomizer = random.SystemRandom()
self._items = []
self.load(items)
# 实现抽象方法load
def load(self, items):
self._items.extend(items)
self._randomizer.shuffle(self._items)
# 实现抽象方法pick
def pick(self):
try:
return self._items.pop()
except IndexError:
raise LookupError('pick from empty BingoCage')
def _call__(self):
self.pick()
# -
class LotteryBlower(Tombola):
def __init__(self, iterable):
self._balls = list(iterable)
# 实现抽象方法load
def load(self, iterable):
self._balls.extend(iterable)
# 实现抽象方法load
def pick(self):
try:
position = random.randrange(len(self._balls))
except ValueError:
raise LookupError('pick from empty LotteryBlower')
return self._balls.pop(position)
# 重载抽象基类中的具体方法
def loaded(self):
return bool(self._balls)
# 重载抽象基类中的具体方法
def inspect(self):
return tuple(sorted(self._balls))
l = [1,2,3]
s = list(l)
s
# ## 虚拟子类
# 即使不使用继承,也可以把一个类注册为抽象基类的虚拟子类。这样做时,我们必须保证注册的类忠实地实现了抽象基类定义的接口,而Python会选择相信我们,从而不做检查。如果没有实现所有的抽象方法,那么常规的运行时会由于出现异常为而中断程序。
#
# **注册为虚拟子类的类不会从抽象基类中继承任何方法或属性**
@Tombola.register
class TomboList(list):
# 由于TomboList注册为了抽象基类Tombola的虚拟子类,
# 因此,它不会继承Tombola的任何方法和属性,它的__init__方法继承自list类
# 重载Tombola的抽象方法pick()
def pick(self):
if self: # 继承list类的__bool__方法
position = random.randrange(len(self))
return self.pop(position)
else:
raise LookupError('pop from empty TomboList')
# 重载Tombola的抽象方法load()[使用list的extend方法代替]
load = list.extend
# 重载Tombola的具体方法loaded()
def loaded(self):
return bool(self)
# 重载Tombola的具体方法inspect()
def inspect(self):
return tuple(sorted(self))
issubclass(TomboList, Tombola)
t = TomboList(range(100))
isinstance(t, Tombola)
# + **注册之后,可以使用issubclass和isinstance函数判断TomboList是不是Tombola的子类**
#
# + 类的继承关系在一个特殊的类属性中指定【__mro__】(Method Resolution Order for short)
TomboList.__mro__
# **`__subclass__()`返回类的直接子类列表,不含虚拟子类**
for real_class in Tombola.__subclasses__():
print(real_class)
# **虽然现在我们可以把register当做装饰器使用了,但是更常见的做法还是把它当做函数来调用,用于注册在其他地方定义的类。**
#
# 例如,将TomboList注册为Tombola的虚拟子类,通常这样做:
#
# ```python
# Tombola.register(TomboList)
# ```
| Jupyter/11.*.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ml)
# language: python
# name: ml
# ---
# # Test Time Augmentation
# One of the most obvious problems with our model is that it operates on fixed lengths of audio clips while our dataset contains audio clips of various lengths. We would like to improve our model's performance on long clips by re-running it on different portions of the clip and combining the predictions, though it's not obvious how exactly we should combine them.
#
# We're taking inspiration from: https://github.com/fastai/fastai/blob/master/fastai/vision/tta.py#L10-L45
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
import shutil
import pandas as pd
import numpy as np
from sklearn.preprocessing import MultiLabelBinarizer
from iterstrat.ml_stratifiers import MultilabelStratifiedKFold
import PIL
import fastai
from fastai.basic_train import _loss_func2activ
from fastai.vision.data import pil2tensor
from fastai.vision import Path, get_preds, ImageList, Image, imagenet_stats, Learner, cnn_learner, get_transforms, DatasetType, models, load_learner, fbeta
import sklearn.metrics
from functools import partial
import torch
import torch.nn as nn
import torch.nn.functional as F
# -
np.random.seed(42)
torch.manual_seed(0)
# +
NFOLDS = 5
script_name = os.path.basename('01_BasicModel').split('.')[0]
MODEL_NAME = "{0}__folds{1}".format(script_name, NFOLDS)
print("Model: {}".format(MODEL_NAME))
# Make required folders if they're not already present
directories = ['kfolds', 'model_predictions', 'model_source']
for directory in directories:
if not os.path.exists(directory):
os.makedirs(directory)
# +
DATA = Path('data')
WORK = Path('work')
CSV_TRN_MERGED = DATA/'train_merged.csv'
CSV_SUBMISSION = DATA/'sample_submission.csv'
TRN_CURATED = DATA/'train_curated2'
TRN_NOISY = DATA/'train_noisy2'
IMG_TRN_CURATED = WORK/'image/trn_curated2'
IMG_TRN_NOISY = WORK/'image/trn_noisy2'
IMG_TEST = WORK/'image/test'
TEST = DATA/'test'
train = pd.read_csv(DATA/'train_curated.csv')
test = pd.read_csv(DATA/'sample_submission.csv')
train_noisy = pd.read_csv(DATA/'train_noisy.csv')
train_merged = pd.read_csv(DATA/'train_merged.csv')
# +
X = train['fname']
y = train['labels'].apply(lambda f: f.split(','))
y_noisy = train_noisy['labels'].apply(lambda f: f.split(','))
transformed_y = MultiLabelBinarizer().fit_transform(y)
transformed_y_noisy = MultiLabelBinarizer().fit_transform(y_noisy)
filenames = train['fname'].values
filenames = filenames.reshape(-1, 1)
oof_preds = np.zeros((len(train), 80))
test_preds = np.zeros((len(test), 80))
tfms = get_transforms(do_flip=True, max_rotate=0, max_lighting=0.1, max_zoom=0, max_warp=0.)
mskf = MultilabelStratifiedKFold(n_splits=5, random_state=4, shuffle=True)
_, val_index = next(mskf.split(X, transformed_y))
# +
#Our clasifier stuff
src = (ImageList.from_csv(WORK/'image', Path('../../')/DATA/'train_curated.csv', folder='trn_merged', suffix='.jpg')
.split_by_idx(val_index)
#.label_from_df(cols=list(train_merged.columns[1:]))
.label_from_df(label_delim=',')
)
data = (src.transform(tfms, size=128).databunch(bs=64).normalize())
# -
f_score = partial(fbeta, thresh=0.2)
learn = cnn_learner(data, models.xresnet101, pretrained=False, metrics=[f_score]).mixup(stack_y=False)
learn.fit_one_cycle(10, 1e-2)
#Overrides fastai's default 'open_image' method to crop based on our crop counter
def setupNewCrop(counter):
def open_fat2019_image(fn, convert_mode, after_open)->Image:
x = PIL.Image.open(fn).convert('L')
# crop (128x321 for a 5 second long audio clip)
time_dim, base_dim = x.size
#How many crops can we take?
maxCrops = int(np.ceil(time_dim / base_dim))
#What's the furthest point at which we can take a crop without running out of pixels
lastValidCrop = time_dim - base_dim
crop_x = (counter % maxCrops) * base_dim
# We don't want to crop any further than the last 128 pixels
crop_x = min(crop_x, lastValidCrop)
x1 = x.crop([crop_x, 0, crop_x+base_dim, base_dim])
newImage = np.stack([x1,x1,x1], axis=-1)
# standardize
return Image(pil2tensor(newImage, np.float32).div_(255))
fastai.vision.data.open_image = open_fat2019_image
# +
def custom_tta(learn:Learner, ds_type:DatasetType=DatasetType.Valid):
dl = learn.dl(ds_type)
ds = dl.dataset
old_open_image = fastai.vision.data.open_image
try:
maxNumberOfCrops = 25
for i in range(maxNumberOfCrops):
print("starting")
setupNewCrop(i)
yield get_preds(learn.model, dl, activ=_loss_func2activ(learn.loss_func))[0]
finally:
fastai.vision.data.open_image = old_open_image
all_preds = list(custom_tta(learn))
avg_preds = torch.stack(all_preds).mean(0)
# -
avg_preds.shape
# ## Improved TTA
# One problem with the above approach is that we take some number of crops (say 10) of each image and average the results. For smaller images we wrap around to the beginning of the image and begin taking predictions from the start again. This oversampling means that our learner is biased toward sounds that occur that the beginning of the clip.
#
# One approach to fix this might be to only include new predictions in our average so as not to oversample from the start of the image.
# +
def custom_tta(learn:Learner, ds_type:DatasetType=DatasetType.Valid):
dl = learn.dl(ds_type)
ds = dl.dataset
old_open_image = fastai.vision.data.open_image
try:
maxNumberOfCrops = 25
for i in range(maxNumberOfCrops):
print("starting")
setupNewCrop(i)
yield get_preds(learn.model, dl, activ=_loss_func2activ(learn.loss_func))[0]
finally:
fastai.vision.data.open_image = old_open_image
all_preds = list(custom_tta(learn))
# -
stacked = torch.stack(all_preds)
stacked.shape
# +
new_preds = []
for i in range(stacked.shape[1]):
firstPred = stacked[0][i]
for j in range(1, stacked.shape[0]):
currentPred = stacked[j][i]
if torch.all(torch.eq(firstPred, currentPred)):
break
preds = stacked[0:j,i]
avg = preds.mean(0)
new_preds.append(avg)
out = torch.stack(new_preds)
out.shape
| 07_TTA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part3: Create a class with an insert method to insert an int to a list. It should also support calculating the max, min, mean, and mode in O(1).
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Can we assume the inputs are valid?
# * No
# * Is there a range of inputs?
# * 0 <= item <= 100
# * Should mean return a float?
# * Yes
# * Should the other results return an int?
# * Yes
# * If there are multiple modes, what do we return?
# * Any of the modes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * None -> TypeError
# * [] -> ValueError
# * [5, 2, 7, 9, 9, 2, 9, 4, 3, 3, 2]
# * max: 9
# * min: 2
# * mean: 55
# * mode: 9 or 2
# ## Algorithm
#
# We'll init our max and min to None. Alternatively, we can init them to -sys.maxsize and sys.maxsize, respectively.
# - For mean, we'll keep track of the number of items we have inserted so far, as well as the running sum.
# - For mode, we'll keep track of the current mode and an array with the size of the given upper limit
# - Each element in the array will be init to 0
# - Each time we insert, we'll increment the element corresponding to the inserted item's value
# On each insert:
# - Update the max and min
# - Update the mean by calculating running_sum / num_items
# - Update the mode by comparing the mode array's value with the current mode
# ## Code
class Solution(object):
def __init__(self, upper_limit=100):
self.upper_limit = 100
self.max = None
self.min = None
global a
a = []
def insert(self, val):
self.val = val
if val is None:
raise TypeError()
else:
#a = []
a.append(val)
self.max = max(a)
self.min = min(a)
self.mean = sum(a)/len(a)
self.mode = max(set(a), key=a.count)
return self.max, self.min, self.mean, self.mode
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_math_ops.py
from nose.tools import assert_equal, assert_true, assert_raises
class TestMathOps(object):
def test_math_ops(self):
solution = Solution()
assert_raises(TypeError, solution.insert, None)
solution.insert(5)
solution.insert(2)
solution.insert(7)
solution.insert(9)
solution.insert(9)
solution.insert(2)
solution.insert(9)
solution.insert(4)
solution.insert(3)
solution.insert(3)
solution.insert(2)
assert_equal(solution.max, 9)
assert_equal(solution.min, 2)
assert_equal(solution.mean, 5)
assert_true(solution.mode in (2, 9))
print('Success: test_math_ops')
def main():
test = TestMathOps()
test.test_math_ops()
if __name__ == '__main__':
main()
# -
| Exam2Part3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Naive Bayes
# Naive Bayes is one of the most famous classification techniques, one of the most simplest ones, and the easiest to apply.
# ## Evaluation Metrics
# https://developers.google.com/machine-learning/crash-course/classification/precision-and-recall
#
# ### Accuracy
# _Accuracy is one metric for evaluating classification models. Informally, accuracy is the fraction of predictions our model got right._
#
# **A = (TP+TN)/(TP+TN+FP+FN)**
#
#
# ### Recall
# _What proportion of actual positives was identified correctly?_
#
# **R = TP/(TP+FN)**
#
# ### Precision
# _What proportion of positive identifications was actually correct?_
#
# **P = TP/(TP+FP)**
# ## Heart Failure Dataset
# We will be using the Heart Failure Dataset with the out of the box values.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/heart_failure_clinical_records_dataset.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
clf = MultinomialNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'MultinomialNB': MultinomialNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
#
#
#
#
#
#
#
#
#
# ## Heart Failure Dataset (Standardized)
# We will be using the Heart Failure Dataset with the standardized values produced in the Scaling section.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/hf_scaled/HF_standardized.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
# MultinomialNB assumes that features have multinomial distribution which is a generalization of the binomial distribution. Neither binomial nor multinomial distributions can contain negative values.
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
#
#
#
# **Which distribution is more adequate to model our data??**
# - Regardless of the dataset used (with standardization, normalization or without any scaling) we noted that the Gaussian Naive Bayes is the one that obtained the best results (which makes sense since our data follows a Gaussian distribution)
#
# **Is the accuracy achieved good enough??**
# - Most accuracies obtained using the Gaussian Naive Bayes estimator are around 80%, which is not an absolutely amazing score, but not as bad as one would think taking into account how simple this model is
#
# **What is the largest kind of errors??**
# - Most of our problems lied in False Negatives
#
# It should also be noted that the best accuracy obtained was when using the non-scaled dataset and with the Gaussian Naive Bayes estimator.
# <br/>
# <br/>
# <br/>
# <br/>
# <br/>
#
# ## Heart Failure Dataset (Normalized)
# We will be using the Heart Failure Dataset with the normalized values produced in the Scaling section.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/hf_scaled/HF_normalized.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
clf = MultinomialNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'MultinomialNB': MultinomialNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
# ## Summary
# **Which distribution is more adequate to model our data??**
# - Regardless of the dataset used (with standardization, normalization or without any scaling) we noted that the Gaussian Naive Bayes is the one that obtained the best results (which makes sense since our data follows a Gaussian distribution)
#
# **Is the accuracy achieved good enough??**
# - Most accuracies obtained using the Gaussian Naive Bayes estimator are around 80%, which is not an absolutely amazing score, but not as bad as one would think taking into account how simple this model is
#
# **What is the largest kind of errors??**
# - Most of our problems lied in False Negatives
#
# It should also be noted that the best accuracy obtained was when using the non-scaled dataset and with the Gaussian Naive Bayes estimator.
# <br/>
# <br/>
# <br/>
# <br/>
# <br/>
# ## Heart Failure Dataset - Balanced
# We will be using the Balanced Heart Failure Dataset with the out of the box values.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/hf_balanced/HF_balanced.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
clf = MultinomialNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'MultinomialNB': MultinomialNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
#
#
#
#
#
#
#
#
#
# ## Heart Failure Dataset (Standardized) - Balanced
# We will be using the Balanced Heart Failure Dataset with the standardized values produced in the Scaling section.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/HF_balanced_standardized.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
# MultinomialNB assumes that features have multinomial distribution which is a generalization of the binomial distribution. Neither binomial nor multinomial distributions can contain negative values.
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
#
#
#
#
#
#
#
#
# ## Heart Failure Dataset (Normalized) - Balanced
# We will be using the Balanced Heart Failure Dataset with the normalized values produced in the Scaling section.
# ### Imports and data loading
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import sklearn.metrics as metrics
from sklearn.naive_bayes import GaussianNB, MultinomialNB, BernoulliNB
import ds_functions as ds
data: pd.DataFrame = pd.read_csv('../../datasets/hf_scaled/HF_balanced_normalized.csv')
data.head()
# -
# ### Prepare and split data
# +
y: np.ndarray = data.pop('DEATH_EVENT').values # Target Variable
X: np.ndarray = data.values # Values of each feature on each record
labels = pd.unique(y)
if(labels[0] == 1):
temp = labels[0]
labels[0] = labels[1]
labels[1] = temp
train_size = 0.7 # % of records used for train (the remainder will be left for test)
trnX, tstX, trnY, tstY = train_test_split(X, y, train_size=train_size, stratify=y)
# -
# ### Gaussian Naive Bayes Estimator
clf = GaussianNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Multinomial Naive Bayes Estimator
clf = MultinomialNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Bernoulli Naive Bayes Estimator
clf = BernoulliNB()
clf.fit(trnX, trnY)
prd_trn = clf.predict(trnX)
prd_tst = clf.predict(tstX)
ds.plot_evaluation_results(labels, trnY, prd_trn, tstY, prd_tst)
# ### Comparison
# +
estimators = {'GaussianNB': GaussianNB(),
'MultinomialNB': MultinomialNB(),
'BernoulliNB': BernoulliNB()}
# Accuracy
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.accuracy_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Accuracy', percentage=True)
plt.show()
'''
# Precision
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.precision_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Precision', percentage=True)
plt.show()
# Recall
xvalues = []
yvalues = []
for clf in estimators:
xvalues.append(clf)
estimators[clf].fit(trnX, trnY)
prdY = estimators[clf].predict(tstX)
yvalues.append(metrics.recall_score(tstY, prdY))
plt.figure()
ds.bar_chart(xvalues, yvalues, title='Comparison of Naive Bayes Models', ylabel='Recall', percentage=True)
plt.show()
'''
# -
# ## Summary
# **Which distribution is more adequate to model our data??**
# - Regardless of the dataset used (with standardization, normalization or without any scaling) we noted that the Gaussian Naive Bayes is the one that obtained the best results (which makes sense since our data follows a Gaussian distribution)
#
# **Is the accuracy achieved good enough??**
# - Most accuracies obtained using the Gaussian Naive Bayes estimator are around 80%, which is not an absolutely amazing score, but not as bad as one would think taking into account how simple this model is
#
# **What is the largest kind of errors??**
# - Most of our problems lied in False Negatives
| Classification/NaiveBayes/NaiveBayes_HF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering Challenge
#
# Clustering is an *unsupervised* machine learning technique in which you train a model to group similar entities into clusters based on their features.
#
# In this exercise, you must separate a dataset consisting of three numeric features (**A**, **B**, and **C**) into clusters. Run the cell below to load the data.
# +
import pandas as pd
data = pd.read_csv('data/clusters.csv')
data.head()
# -
# Your challenge is to identify the number of discrete clusters present in the data, and create a clustering model that separates the data into that number of clusters. You should also visualize the clusters to evaluate the level of separation achieved by your model.
#
# Add markdown and code cells as required to create your solution.
#
# > **Note**: There is no single "correct" solution. A sample solution is provided in [04 - Clustering Solution.ipynb](04%20-%20Clustering%20Solution.ipynb).
# Your code to create a clustering solution
from sklearn.decomposition import pca
from sklearn.preprocessing import MinMaxScaler
from sklearn.cluster import kmeans, AgglomerativeClustering
| challenges/.ipynb_checkpoints/04 - Clustering Challenge-checkpoint.ipynb |
# %matplotlib inline
from fenics import *
parameters["plotting_backend"] = 'matplotlib'
import pylab
# +
from __future__ import print_function
# Use -02 optimization
parameters["form_compiler"]["cpp_optimize"] = True
# Define mesh and geometry
mesh = Mesh("dolfin-2.xml.gz")
x = SpatialCoordinate(mesh)
n = FacetNormal(mesh)
# Define Taylor--Hood function space W
V = VectorFunctionSpace(mesh, "Lagrange" , 2)
Q = FunctionSpace(mesh, "Lagrange", 1)
P2 = VectorElement("Lagrange", triangle, 2)
P1 = FiniteElement("Lagrange", triangle, 1)
TH = MixedElement([P2, P1])
W = FunctionSpace(mesh, TH)
# Define Function and TestFunction(s)
w = Function(W)
(u, p) = split(w)
(v, q) = TestFunctions(W)
# Define viscosity and bcs
nu = Expression("0.2*(1+pow(x[1],2))", degree=2)
p0 = (1.0 - x[0]) # or Expression("1.0-x[0]", degree=1)
bcs = DirichletBC(W.sub(0), (0.0, 0.0),
"on_boundary && !(near(x[0], 0.0) || near(x[0], 1.0))")
# Define variational form
epsilon = sym(grad(u))
F = (2*nu*inner(epsilon, grad(v)) - div(u)*q - div(v)*p)*dx\
+ p0*dot(v,n)*ds
# Solve problem
solve(F == 0, w, bcs)
# Plot solutions
plot(u, title="Velocity")
pylab.show()
plot(p, title="Pressure")
pylab.show()
interactive()
| notebooks/03_static_nonlinear_pdes/apa/stokes_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Point Data
#
# Here we are going to plot some markers on top of the maps we already made. We will use seismicity information since this is so readily available online (we use GeoJSON files from the [USGS site](http://earthquake.usgs.gov/earthquakes/search/) since these are easy to parse. Typically you have a limit on how many data to grab in each pass so if you want a global dataset you end up with fewer small events or a limited date range. I did this for a few places for you.
#
# +
# %pylab inline
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import matplotlib.pyplot as plt
import gdal
import numpy
# -
base_projection = ccrs.PlateCarree()
global_extent = [-180.0, 180.0, -90.0, 90.0]
globalmag = gdal.Open("Resources/EMAG2_image_V2.tif")
globalmag_img = globalmag.ReadAsArray().transpose(1,2,0)
globalmarble = gdal.Open("Resources/BlueMarbleNG-TB_2004-06-01_rgb_3600x1800.TIFF")
globalmarble_img = globalmarble.ReadAsArray().transpose(1,2,0)
globaletopo = gdal.Open("Resources/color_etopo1_ice_low.tif")
globaletopo_img = globaletopo.ReadAsArray().transpose(1,2,0)
globaletopobw = gdal.Open("Resources/etopo1_grayscale_hillshade.tif")
globaletopobw_img = globaletopobw.ReadAsArray()[::3,::3] / 256.0
# +
# "Features" such as land, ocean, coastlines (50m = the 1:50 million scale)
land = cfeature.NaturalEarthFeature('physical', 'land', '50m',
edgecolor="green",
facecolor="white")
ocean = cfeature.NaturalEarthFeature('physical', 'ocean', '50m',
edgecolor="green",
facecolor="blue")
coastline = cfeature.NaturalEarthFeature('physical', 'coastline', '50m',
edgecolor=(0.0,0.0,0.0),
facecolor="none")
# Add these to the plot object as
# ax.add_feature(coastline, linewidth=4, edgecolor=(1,0,1) zorder=1)
# and so forth. zorder is used to force the layering as required.
# +
# Recent earthquake data (from static downloaded files)
import json
# 1 Global
earthquakes_datafile=open('Resources/Earthquakes-2000-2014-5.5+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes = earthquakes_data["features"]
# Now we have a dictionary of many, many events
eqlon = numpy.empty(len(earthquakes))
eqlat = numpy.empty(len(earthquakes))
eqdep = numpy.empty(len(earthquakes))
eqmag = numpy.empty(len(earthquakes))
for i,eq in enumerate(earthquakes):
eqlon[i], eqlat[i], eqdep[i] = eq["geometry"]["coordinates"]
eqmag[i] = eq["properties"]["mag"]
print "Global depth range: ", eqdep.min()," - ", eqdep.max()
print "Global magnitude range: ", eqmag.min()," - ", eqmag.max()
# 2 Australian
earthquakes_datafile=open('Resources/Earthquakes-AusRegion-2000-2014-4.8-5.5+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes = earthquakes_data["features"]
# Now we have a dictionary of many, many events
ausqlon = numpy.empty(len(earthquakes))
ausqlat = numpy.empty(len(earthquakes))
ausqdep = numpy.empty(len(earthquakes))
ausqmag = numpy.empty(len(earthquakes))
for i, eq in enumerate(earthquakes):
ausqlon[i], ausqlat[i], ausqdep[i] = eq["geometry"]["coordinates"]
ausqmag[i] = eq["properties"]["mag"]
print "Aus Region depth range: ", ausqdep.min()," - ", ausqdep.max()
print "Aus Region magnitude range: ", ausqmag.min()," - ", ausqmag.max()
#3 Japanese - Earthquakes-JapanRegion-2009-2014-4.5+.json
earthquakes_datafile=open('Resources/Earthquakes-JapanRegion-2009-2014-4.5+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes = earthquakes_data["features"]
#3+ South of 30 degrees: Earthquakes-IBMRegion-1990-2014-3+.json
earthquakes_datafile=open('Resources/Earthquakes-IBMRegion-1990-2014-3+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes.extend(earthquakes_data["features"])
jpqlon = numpy.empty(len(earthquakes))
jpqlat = numpy.empty(len(earthquakes))
jpqdep = numpy.empty(len(earthquakes))
jpqmag = numpy.empty(len(earthquakes))
for i, eq in enumerate(earthquakes):
jpqlon[i], jpqlat[i], jpqdep[i] = eq["geometry"]["coordinates"]
jpqmag[i] = eq["properties"]["mag"]
print "Japan Region depth range: ", jpqdep.min()," - ", jpqdep.max()
print "Japan Region magnitude range: ", jpqmag.min()," - ", jpqmag.max()
norm_eqdep = matplotlib.colors.Normalize(vmin = 0.0, vmax = 200, clip = False)
#4 Yakutat EQ
earthquakes_datafile=open('Resources/Earthquakes-YakutatRegion-1990-2014-3+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes = earthquakes_data["features"]
yakqlon = numpy.empty(len(earthquakes))
yakqlat = numpy.empty(len(earthquakes))
yakqdep = numpy.empty(len(earthquakes))
yakqmag = numpy.empty(len(earthquakes))
for i, eq in enumerate(earthquakes):
yakqlon[i], yakqlat[i], yakqdep[i] = eq["geometry"]["coordinates"]
yakqmag[i] = eq["properties"]["mag"]
print "Yakutat Region depth range: ", yakqdep.min()," - ", yakqdep.max()
print "Yakutat Region magnitude range: ", yakqmag.min()," - ", yakqmag.max()
earthquakes_datafile=open('Resources/Earthquakes-MeditRegion-1990-2014-3+.json')
earthquakes_data = json.load(earthquakes_datafile)
earthquakes_datafile.close()
earthquakes = earthquakes_data["features"]
itqlon = numpy.empty(len(earthquakes))
itqlat = numpy.empty(len(earthquakes))
itqdep = numpy.empty(len(earthquakes))
itqmag = numpy.empty(len(earthquakes))
for i, eq in enumerate(earthquakes):
itqlon[i], itqlat[i], itqdep[i] = eq["geometry"]["coordinates"]
itqmag[i] = eq["properties"]["mag"]
print "Vatican Region depth range: ", itqdep.min()," - ", itqdep.max()
print "Vatican Region magnitude range: ", itqmag.min()," - ", itqmag.max()
# -
# ### Plotting points
#
# We add the points to the map using the usual plotting tools from matplotlib plus the transformation argument
# +
projection = ccrs.PlateCarree()
fig = plt.figure(figsize=(12, 12), facecolor="none")
ax = plt.axes(projection=projection)
ax.set_extent([0, 40, 28, 48])
#ax.add_feature(land, edgecolor="black", alpha=0.1, linewidth=2)
ax.add_feature(ocean, alpha=0.1, zorder=1)
ax.imshow(globaletopo_img, origin='upper', transform=base_projection, extent=global_extent)
ax.imshow(globaletopobw_img, origin='upper', cmap=mpl.cm.Greys, transform=base_projection, extent=global_extent, alpha=0.75, zorder=1)
plt.scatter(itqlon, itqlat, c=itqdep, cmap=mpl.cm.jet_r, norm=norm_eqdep, linewidth=0,
s=(itqmag-3.0)*10, transform=ccrs.Geodetic(), alpha=0.333, zorder=2)
plt.show()
# +
# Italy / Mediterranean earthquakes
projection = ccrs.PlateCarree()
fig = plt.figure(figsize=(12, 12), facecolor="none")
ax = plt.axes(projection=projection)
ax.set_extent([0, 40, 28, 48])
ax.add_feature(land, edgecolor="black", alpha=0.1, linewidth=2)
ax.add_feature(ocean, alpha=0.1, zorder=1)
ax.imshow(globaletopo_img, origin='upper', transform=base_projection, extent=global_extent)
ax.imshow(globaletopobw_img, origin='upper', cmap=mpl.cm.Greys, transform=base_projection, extent=global_extent, alpha=0.85, zorder=1)
plt.scatter(itqlon, itqlat, c=itqdep, cmap=mpl.cm.jet_r, norm=norm_eqdep, linewidth=0,
s=(itqmag-3.0)*10, transform=ccrs.Geodetic(), alpha=0.333, zorder=2)
plt.savefig("ItaliaEq.png")
plt.show()
# +
# Seafloor age data and global image - data from Earthbyters
datasize = (1801, 3601, 3)
age_data = np.empty(datasize)
ages = np.load("Resources/global_age_data.3.6.z.npz")["ageData"]
lats = np.linspace(90, -90, datasize[0])
lons = np.linspace(-180.0,180.0, datasize[1])
arrlons,arrlats = np.meshgrid(lons, lats)
age_data[...,0] = arrlons[...]
age_data[...,1] = arrlats[...]
age_data[...,2] = ages[...]
# +
# Global
projection = ccrs.PlateCarree()
bg_projection = ccrs.PlateCarree()
fig = plt.figure(figsize=(10, 10), facecolor="none", edgecolor="none")
ax = plt.axes(projection=projection)
ax.set_extent(global_extent)
ax.add_feature(land, edgecolor="black", alpha=0.2, linewidth=0.25)
# ax.add_feature(ocean, alpha=0.1, zorder=1)
ax.add_feature(coastline, alpha=1.0, linewidth=0.33)
ax.imshow(globaletopobw_img, origin='upper', transform=base_projection, extent=global_extent, zorder=0, cmap="gray")
cf = contourf(age_data[:,:,0], age_data[:,:,1], age_data[:,:,2],
levels = arange(0.5,250,10), vmin=0, vmax=150,
transform=base_projection, cmap="RdYlBu",zorder=2, alpha=0.75)
contour(age_data[:,:,0], age_data[:,:,1], age_data[:,:,2], levels = (0.1,0.5), colors="white", transform=base_projection)
plt.scatter(eqlon, eqlat, c=eqdep, cmap=mpl.cm.jet_r, norm=norm_eqdep, linewidth=0.33,
s=(eqmag-5.5)*10, transform=ccrs.Geodetic(), alpha=0.7, zorder=2)
plt.savefig("GlobalAgeMapEq.png", dpi=600, frameon=False, edgecolor="none", facecolor="none", bbox_inches='tight', pad_inches=0.0)
# +
projection = ccrs.PlateCarree()
fig = plt.figure(figsize=(16, 16), facecolor="none")
ax = plt.axes(projection=projection)
ax.set_extent([90, 180, -50, 5])
# ax.add_feature(ocean, facecolor=(0.4,0.4,0.6), edgecolor="none", linewidth=5, alpha=0.40, zorder=1)
ax.imshow(globaletopobw_img, origin='upper', transform=base_projection, extent=global_extent, zorder=0, cmap="gray")
contourf(age_data[:,:,0], age_data[:,:,1], age_data[:,:,2], levels = arange(0,200,10),
transform=base_projection, cmap="RdYlBu",zorder=2, alpha=0.8)
contour(age_data[:,:,0], age_data[:,:,1], age_data[:,:,2], levels = (0.1,0.5), colors="white", transform=base_projection)
plt.scatter(ausqlon, ausqlat, c=ausqdep, cmap=mpl.cm.jet_r, norm=norm_eqdep, linewidth=0,
s=(ausqmag-4.0)*25, transform=ccrs.Geodetic(), alpha=0.5, zorder=3)
plt.show()
# -
# The plotting of lines is actually a bit more interesting since the tranformation machinery needs to work on all the points between the given end points. We have seen lines and fills in the contouring but it is interesting to see what is actually going on (this is one of the standard examples from cartopy).
# +
ax = plt.axes(projection=ccrs.Robinson())
# ax = plt.axes(projection=ccrs.LambertCylindrical())
# make the map global rather than have it zoom in to
# the extents of any plotted data
ax.set_global()
ax.coastlines()
ax.stock_img()
plt.plot(-0.08, 51.53, 'o', transform=ccrs.PlateCarree())
plt.plot(132 , 43.17, 'o', transform=ccrs.PlateCarree())
plt.plot([-0.08, 132], [51.53, 43.17], transform=ccrs.PlateCarree())
plt.plot([-0.08, 132], [51.53, 43.17], transform=ccrs.Geodetic(), color="Blue", linewidth=3)
# +
# Examples of projections and how to draw/fill a shape
# Inside out / outside in is defined by cw/ccw ordering of points in the filled shape
rotated_pole = ccrs.RotatedPole(pole_latitude=60, pole_longitude=180)
scale = 45
x = [-scale, -scale*1.5, -scale, 0.0, scale, scale*1.5, scale, 0.0, -scale]
y = [-scale, 0.0, scale, scale * 1.5, scale, 0.0, -scale, -scale*1.5, -scale]
xx = x[::-1]
yy = y[::-1]
fig = plt.figure(figsize=(6, 12))
ax = plt.subplot(311, projection=rotated_pole)
ax.stock_img()
ax.coastlines()
ax.plot(x, y, marker='o', transform=ccrs.Geodetic())
ax.fill(x, y, color='coral', transform=ccrs.Geodetic(), alpha=0.4)
ax.gridlines()
ax = plt.subplot(312, projection=ccrs.PlateCarree())
ax.stock_img()
ax.coastlines()
ax.plot(x, y, marker='o', transform=ccrs.Geodetic())
ax.fill(x, y, transform=ccrs.Geodetic(), color='coral', alpha=0.4, closed=True)
ax.gridlines()
ax = plt.subplot(313, projection=rotated_pole)
ax.stock_img()
# ax.coastlines()
ax.plot(x, y, marker='o', transform=ccrs.Geodetic())
ax.fill(xx, yy, color='coral', alpha=0.4, transform=ccrs.Geodetic())
ax.gridlines()
plt.show()
# -
| CourseContent/Notebooks/Mapping/5 - Working with point data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Demonstrate Aggregation of Descriptive Statistics
# Here we create an array of random values and for each row of the array, we create
# a distinct ```pebaystats.dstats``` object to accumulate the descriptive statistics
# for the values in that row.
#
# Once we have the data, we can use the ```numpy``` package to generate the expected
# values for mean and variance of the data for each row. We can also generate the
# expected mean and variance of the total data set.
#
# We then accumulate each column value for each row into its respective ```dstats``` object
# and when we have the data accumulated into these partial results, we can compare with
# the expected row values.
#
# We can then aggregate each of the row values into a final value for mean and variance
# of the entire set of data and compare to the expected value.
# We will need to import the ```numpy``` package as well as the ```dstats``` class from the ```pebaystats``` package. We import the nosetools package to allow direct comparison of expected and generated values.
import numpy as np
import nose.tools as nt
from pebaystats import dstats
# ---
#
# Now we set the parameters, including the random seed for repeatability
# +
np.random.seed(0)
### Random data size
rows = 10
cols = 100
### Each accumulators size
depth = 2
width = 1
# -
# ---
# The test array can now be created and its shape checked.
#
# The individual row statistics and overall mean and variance can be generated as the expected
# values at this time as well.
# +
### Test data -- 10 rows of 100 columns each
test_arr = np.random.random((rows,cols))
print('Test data has shape: %d, %d' % test_arr.shape)
### Expected intermediate output
mid_mean = np.mean(test_arr,axis = 1)
mid_var = np.var(test_arr, axis = 1)
### Expected final output
final_mean = np.mean(test_arr)
final_var = np.var(test_arr)
# -
# ---
# Now we can create a ```dstats``` object for each row and accumulate the row data
# into its respected accumulator. We can print the generated and expected intermediate (row)
# values to check that all is working correctly.
# +
### Create an object for each row and accumulate the data in that row
statsobjects = [ dstats(depth,width) for i in range(0,rows) ]
discard = [ statsobjects[i].add(test_arr[i,j])
for j in range(0,cols)
for i in range(0,rows)]
print('\nIntermediate Results\n')
for i in range(0,rows):
values = statsobjects[i].statistics()
print('Result %d mean: %11g, variance: %11g (M2/N: %11g/%d)' %(i,values[0],values[1],statsobjects[i].moments[1],statsobjects[i].n))
print('Expected mean: %11g, variance: %11g' %(mid_mean[i],mid_var[i]))
nt.assert_almost_equal(values[0], mid_mean[i], places = 14)
nt.assert_almost_equal(values[1], mid_var[i], places = 14)
# -
# ---
# Now we can aggregate each of the intermediate row results into a final mean and
# variance value for the entire data set. And then compare with the ```numpy```
# generated expected values
# +
### Aggregate result into the index 0 accumulator
discard = [ statsobjects[0].aggregate(statsobjects[i]) for i in range(1,rows) ]
values = statsobjects[0].statistics()
print('\nAggregated Results\n')
print('Result mean: %11g, variance: %11g' %(values[0],values[1]))
print('Expected mean: %11g, variance: %11g' %(final_mean,final_var))
nt.assert_almost_equal(values[0], final_mean, places = 14)
nt.assert_almost_equal(values[1], final_var, places = 14)
| examples/aggregation_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from pandas_datareader import data as web
from scipy.stats import norm
import matplotlib.pyplot as plt
# %matplotlib inline
ticker = 'PG'
data = pd.DataFrame()
data[ticker] = web.DataReader(ticker, data_source='iex', start='2015-1-1', end='2017-3-21')['close']
log_returns = np.log(1 + data.pct_change())
# <br /><br />
# $$
# {\LARGE S_t = S_{t-1} \mathbin{\cdot} e^{((r - \frac{1}{2} \cdot stdev^2) \mathbin{\cdot} \delta_t + stdev \mathbin{\cdot} \sqrt{\delta_t} \mathbin{\cdot} Z_t)} }
# $$
# <br /><br />
r = 0.025
stdev = log_returns.std() * 250 ** 0.5
stdev
type(stdev)
stdev = stdev.values
stdev
# +
T = 1.0
t_intervals = 250
delta_t = T / t_intervals
iterations = 10000
# -
Z = np.random.standard_normal((t_intervals + 1, iterations))
S = np.zeros_like(Z)
S0 = data.iloc[-1]
S[0] = S0
# <br /><br />
# $$
# {\LARGE S_t = S_{t-1} \mathbin{\cdot} e^{((r - \frac{1}{2} \cdot stdev^2) \mathbin{\cdot} \delta_t + stdev \mathbin{\cdot} \sqrt{\delta_t} \mathbin{\cdot} Z_t)} }
# $$
# <br /><br />
for t in range(1, t_intervals + 1):
S[t] = S[t-1] * np.exp((r - 0.5 * stdev ** 2) * delta_t + stdev * delta_t ** 0.5 * Z[t])
S
S.shape
plt.figure(figsize=(10, 6))
plt.plot(S[:, :10]);
| Python for Finance - Code Files/109 Monte Carlo - Euler Discretization - Part I/Online Financial Data (APIs)/Python 3 APIs/MC - Euler Discretization - Part I - Lecture_IEX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from google.cloud import bigquery
# %load_ext google.cloud.bigquery
import os
import re
import json
import chardet
import codecs
import time
# -
# # Disambiguation of Attorney Names
# ## 1. Creating a table of raw attorney names
#
# In this step, we first remove any non-alphabetic charactersand then we create a table containing raw attorney names and the processed ones.
# +
client = bigquery.Client()
job_config = bigquery.QueryJobConfig()
job_config.use_query_cache = False
job_config.write_disposition = 'WRITE_TRUNCATE'
# Set Destination
dataset_id = 'data_preparation'
table_id = '5_attorney_raw'
table_ref = client.dataset(dataset_id).table(table_id)
job_config.destination = table_ref
query="""
WITH t1 AS(
SELECT *
FROM(
SELECT UPPER(REGEXP_REPLACE(
REGEXP_REPLACE(
REGEXP_REPLACE(correspondence_name_line_1, r'[^a-zA-Z\s]+', ''),
r'[\s]+', ' '),
r'(^\s+)|(\s+$)', ''
)
) AS lawyer,
correspondence_name_line_1 AS raw_lawyer
FROM `patents-public-data.uspto_oce_pair.correspondence_address`
GROUP BY correspondence_name_line_1
)
GROUP BY raw_lawyer, lawyer
ORDER BY lawyer DESC
)
SELECT *
FROM t1
"""
query_job = client.query(query, location='US', job_config=job_config)
print('Query job has {} started!'.format(query_job.job_id))
query_job.result()
print('Job has finished!')
# -
# ### Extracting the table into a CSV file
# +
# Exctracting table
client = bigquery.Client()
# Set Source table
project_id = 'usptobias'
dataset_id = 'data_preparation'
table_id = '5_attorney_raw'
table_ref = client.dataset(dataset_id, project=project_id).table(table_id)
# Set Destination
dest_bucket = 'uspto-data'
dest_folder = 'data_preparation'
dest_file_name = '5_attorney_raw.csv'
dest_uri = "gs://{0}/{1}/{2}".format(dest_bucket, dest_folder, dest_file_name)
extract_job = client.extract_table(table_ref, dest_uri, location='US')
print('Extract job has {} started!'.format(extract_job.job_id))
extract_job.result()
print('Job has finished and table {} has been exported to {} bucket!'.format(dest_file_name, dest_bucket))
# -
# # 2. Disambiguating Using Standardization Rules
# ***Source***: The standardization rules has been downloaded from the following link:
# https://sites.google.com/site/patentdataproject/Home/posts/namestandardizationroutinesuploaded
#
# We then preprocessed the rules to prepare them for our purpose. The preprocessed rules can be found in the `./stdname_rules/` directory.
# Loading "5_attorney_raw" table in a Pandas dataframe
## You need to first download "5_attorney_raw.csv" into the './data/' folder (located in the current path) ...
## ... from "uspto-data/data_preparation" GCP Bucket
data_folder = './data/'
df_rawlawyer = pd.read_csv(data_folder+'5_attorney_raw.csv', low_memory=False)
print('Number of records: {:,}'.format(df_rawlawyer.shape[0]))
df_rawlawyer.head(2)
# Adding trailing and ending space (for using the rule-based disambiguation)
df_rawlawyer = df_rawlawyer.dropna()
df_rawlawyer.lawyer = df_rawlawyer.lawyer.apply(lambda x: " " + x + " ")
df_rawlawyer.shape
# +
# Extracting the standard rule files
from zipfile import ZipFile
data_folder = './data/'
with ZipFile(data_folder+'stdname_rules.zip', 'r') as file_ref:
file_ref.extractall(data_folder+'stdname_rules/')
files = sorted(os.listdir(data_folder+'stdname_rules/'))
# +
# Loading standard rules into a dictionary
pattern = r'^.*\"(.*?)\".*?\"(.*?)\"'
std_mapper = dict()
decoding = [(2, 1),
(1, 2),
(1, 2),
(2, 1),
(1, 2),
(1, 2)]
for dec, file in zip(decoding, files):
encoding = chardet.detect(open(data_folder+'stdname_rules/'+file, "rb").read())['encoding']
with codecs.open(data_folder+'stdname_rules/'+file, 'r', encoding=encoding) as text_file:
lines = text_file.readlines()
for line in lines:
key = (re.match(pattern, line)[dec[0]]).rstrip()
value = (re.match(pattern, line)[dec[1]]).rstrip()
std_mapper[key] = value
# +
df_mapper = pd.DataFrame(std_mapper, index=['mapped']).T.reset_index(drop=False).rename(columns={'index':'initial'})
df_mapper.mapped = ' '
df_mapper.initial = df_mapper.initial.apply(lambda x: x+' ')
std_mapper = df_mapper.dropna().set_index('initial')['mapped'].to_dict()
df_mapper.head(3)
# -
# Starting standardization
start_t = time.perf_counter()
df_rawlawyer.lawyer = df_rawlawyer.lawyer.replace(std_mapper, regex=True).replace(std_mapper, regex=True)
end_t = time.perf_counter()
diff_t = end_t - start_t
print('Total running time was {:,.0f} hours and {:.0f} minutes!'.format(diff_t//3600, (diff_t%3600)//60))
# +
# Stripping the spaces
df_rawlawyer.lawyer = df_rawlawyer.lawyer.str.strip()
# Getting unique disambiguated lawyers
df_lawyer_id = df_rawlawyer[['lawyer']].drop_duplicates().reset_index(drop=True).copy()
# Adding unique ID to each lawyer
df_lawyer_id = df_lawyer_id.reset_index(drop=False).rename(columns={'index':'lawyer_id'})
df_lawyer_id.lawyer_id = df_lawyer_id.lawyer_id + 100000
print('Number of unique lawyers: {:,}'.format(df_lawyer_id.shape[0]))
df_lawyer_id.head(2)
# -
df_lawyer_merger = pd.merge(df_rawlawyer, df_lawyer_id, on=['lawyer'], how='left')
print('Number of records: {:,}'.format(df_lawyer_merger.shape[0]))
df_lawyer_merger.head(3)
# Saving the resulting dataframes
df_lawyer_id.to_csv('./data/5_attorneyId.csv', encoding='utf-8', index=False)
df_lawyer_merger.to_csv('./data/5_attorney_disambiguated.csv', encoding='utf-8', index=False)
# ## 3. Creating the BigQuery tables using the disambiguated attorney names
# +
# Creating "5_attorneyID" table
# Creating "lawyer_id_fung" table
bq_client = bigquery.Client()
schema = [
bigquery.SchemaField('attorney_id', 'STRING', 'NULLABLE', None, ()),
bigquery.SchemaField('attorney', 'STRING', 'NULLABLE', None, ())
]
dataset_id = 'data_preparation'
dataset_ref = bq_client.dataset(dataset_id)
dest_table_name = '5_attorneyID'
job_config = bigquery.LoadJobConfig()
job_config.schema = schema
job_config.skip_leading_rows = 1
job_config.source_format = bigquery.SourceFormat.CSV
uri = "gs://uspto-data/data_preparation/5_attorneyId.csv"
load_job = bq_client.load_table_from_uri(
uri, dataset_ref.table(dest_table_name), job_config=job_config
)
print("Starting job {}".format(load_job.job_id))
load_job.result()
print('Job has finished!')
# +
# Creating "5_attorney_disambiguated" table
# Creating "lawyer_id_fung" table
bq_client = bigquery.Client()
schema = [
bigquery.SchemaField('attorney', 'STRING', 'NULLABLE', None, ()),
bigquery.SchemaField('raw_attorney', 'STRING', 'NULLABLE', None, ()),
bigquery.SchemaField('attorney_id', 'STRING', 'NULLABLE', None, ())
]
# Setting the destination table path
dataset_id = 'data_preparation'
dataset_ref = bq_client.dataset(dataset_id)
dest_table_name = '5_attorney_disambiguated'
job_config = bigquery.LoadJobConfig()
job_config.schema = schema
job_config.skip_leading_rows = 1
job_config.source_format = bigquery.SourceFormat.CSV
uri = "gs://uspto-data/data_preparation/5_attorney_disambiguated.csv"
load_job = bq_client.load_table_from_uri(
uri, dataset_ref.table(dest_table_name), job_config=job_config
)
print("Starting job {}".format(load_job.job_id))
load_job.result()
print('Job has finished!')
# -
# ## 4. Creating the final table: `5_appln_attorney`
# +
client = bigquery.Client()
job_config = bigquery.QueryJobConfig()
job_config.use_query_cache = False
job_config.write_disposition = 'WRITE_TRUNCATE'
# Set Destination
project_id = 'usptobias'
dataset_id = 'data_preparation'
table_id = '5_appln_attorney'
table_ref = client.dataset(dataset_id).table(table_id)
job_config.destination = table_ref
query="""
WITH rlawyerAppln_table AS(
SELECT
application_number AS appln_nr,
correspondence_name_line_1 AS raw_attorney,
correspondence_region_code AS attorney_region_code,
correspondence_country_code AS attorney_country_code
FROM `patents-public-data.uspto_oce_pair.correspondence_address`
), lawyerMerger_table AS(
SELECT attorney, raw_attorney, attorney_id
FROM `{0}.{1}.5_attorney_disambiguated`
)
SELECT appln_nr, attorney, attorney_id, attorney_region_code, attorney_country_code
FROM rlawyerAppln_table
LEFT JOIN lawyerMerger_table USING(raw_attorney)
WHERE attorney IS NOT NULL
""".format(project_id, dataset_id)
query_job = client.query(query, location='US', job_config=job_config)
print('Query job has {} started!'.format(query_job.job_id))
query_job.result()
print('Job has finished!')
| 03_data_preparation/5_attorney_tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Método CRISP-DM
# - Cross Industry Standard Process - Data Mining
# - Método cíclico para gerenciamento de projeto.
# - 1º Ciclo CRISP completo:
# * 1 versão end-to-end da solução.
# * Velocidade na entrega de valor.
# * Mapeamento de todos os possíveis problemas.
# # Ciclo CRISP-DM
# * Questão de negócio
# - Business model (E-commerce, Software as a Service- SaaS, Free Mobile App, Media Site, User-generated content, Two-sided Marketplaces, consultant company, Startup)
# - Metrics: Custo, receita, Usuários, Conversão, Margem de contribuição, HEART e NPS.
# * Entendimento do negócio
# - Qual a motivação?
# - Qual a causa raíz do problema?
# - Quem é o dono do problema?
# - O formato da solução desejada?
# * Coleta de dados
# - Planilha de dados: CSV
# - Banco de dados: Cloud ou servidor (Snowflake, postgres, Mysql e Oracle)
# - API request: Python (Requests, Urllib e manipulação com o JSON) e Postman (teste de api)
# - Webscrapping: Python (Selenium, BeatifullSoup, Scrappy e Requests)
# * Limpeza dos dados
# - Python: Pandas
# - Spark: Pyspark e Scala
# * Exploração dos dados
# - **Entender de fato os insights que impactam no resultado do negócio**
# - Python: (Pandas, Seaborn, Matplotlib e Plotly)
# - Visualização de dados: Power BI, Tableau e Plotly
# - Spark: Pyspark e ScalaSpark
# - Estatística: Estatística descritiva, Distribuição de probabilidade, Análise bivariada (Correlações: numérica x numérica, numérica x categórica, categórica x categórica, correlação entre time series)
# - Histogramas
# - Hipóteses: Criação e validação de hipóteses
# * Modelagem de dados
# - Rescaling: Standard Scaler, MinMax Scaler, MaxAbs Scaler e Robust Scaler.
# - Encoding: Numérica (Discretização e binarização) e Categórica ( One Hot Enconder, Label encoder, Binary encoder, Hashing encoder, Target encoder, Backward Encoder e polynomial encoder)
# - Seleção de features: Boruta Algorithm, Pearson correlation, Chi-Squared, Lasso Regression Model, Random Forest, Spearman Correlation, ANOVA, Kendall correlation e Mutual Information.
# - Tokenização (NLP)
# * Algoritmos de Machine Learning
# * Avaliação do algoritmo
# - Avaliação de métricas e erros.
# * Modelo em produção
#
| Notebooks/0.0-Introducao.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="06da34af0c189cb0c29beb679dfa202bc76b63df"
# # Within Top 10% with Simple Regression Model.
# + [markdown] _uuid="6f62f70159ddf19687dd1c679055f7cc27630cfb"
# # Step By Step Procedure To Predict House Price
# + [markdown] _uuid="3306a9daf742759fff4e9a6959ab72fec3230e7f"
# # Importing packages
# We have **numpy** and **pandas** to work with numbers and data, and we have **seaborn** and **matplotlib** to visualize data. We would also like to filter out unnecessary warnings. **Scipy** for normalization and skewing of data.
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
#import some necessary librairies
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# %matplotlib inline
import matplotlib.pyplot as plt # Matlab-style plotting
import seaborn as sns
color = sns.color_palette()
sns.set_style('darkgrid')
import warnings
def ignore_warn(*args, **kwargs):
pass
warnings.warn = ignore_warn #ignore annoying warning (from sklearn and seaborn)
from scipy import stats
from scipy.stats import norm, skew #for some statistics
# + [markdown] _uuid="e30ef2b9a5211e7e44031145e7dfbf54a0a429e2"
# # Loading and Inspecting data
# With various Pandas functions, we load our training and test data set as well as inspect it to get an idea of the data we're working with. This is a large dataset we will be working on.
#
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
#Now let's import and put the train and test datasets in pandas dataframe
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
train.describe()
# + _uuid="e79d47658b82d49905c39b7e66e3fcf03a86ced2"
print ("Size of train data : {}" .format(train.shape))
print ("Size of test data : {}" .format(test.shape))
# + [markdown] _uuid="99ad10c6432a461389c5c9b5ffe2f56ceb70d7a9"
# > That is a very large data set! We are going to have to do a lot of work to clean it up
#
# **Drop the Id column because we dont need it currently.**
# + _uuid="1209490182f8356c09e7f43c1834f18e9ec8ba9e"
#Save the 'Id' column
train_ID = train['Id']
test_ID = test['Id']
#Now drop the 'Id' colum since it's unnecessary for the prediction process.
train.drop("Id", axis = 1, inplace = True)
test.drop("Id", axis = 1, inplace = True)
# + _uuid="fca66e4aa038a4310fc6b70122b7e184ee0b765f"
print ("Size of train data after dropping Id: {}" .format(train.shape))
print ("Size of test data after dropping Id: {}" .format(test.shape))
# + [markdown] _uuid="adab25c81ca5515fa0dea82196134266e769e933"
# ## Dealing with outliers
#
# Outlinear in the GrLivArea is recommended by the author of the data to remove it. The author says in documentation “I would recommend removing any houses with more than 4000 square feet from the data set (which eliminates these five unusual observations) before assigning it to students.”
#
# + _uuid="589e1b7432290d42ec3ba5f527000d6de5c7fa90"
fig, ax = plt.subplots()
ax.scatter(x = train['GrLivArea'], y = train['SalePrice'])
plt.ylabel('SalePrice', fontsize=13)
plt.xlabel('GrLivArea', fontsize=13)
plt.show()
# + [markdown] _uuid="190cfe132f4616d905bba693b79b49fde3e89319"
# We can see that there are outlinear with low SalePrice and high GrLivArea. This looks odd.
# We need to remove it.
# + _uuid="e215c8fe40fdcbefa4480a4ac0a8c2adf4f68a5e"
#Deleting outliers
train = train.drop(train[(train['GrLivArea']>4000) & (train['SalePrice']<300000)].index)
# + [markdown] _uuid="502ebe387edeeeb1e4a3f454d5dd004645a07e75"
# ## Correlation Analysis
#
# Let see the most correlated features.
# + _uuid="aa18c6e3a818d58c4a14addedb21888a2ec610cb"
# most correlated features
corrmat = train.corr()
top_corr_features = corrmat.index[abs(corrmat["SalePrice"])>0.5]
plt.figure(figsize=(10,10))
g = sns.heatmap(train[top_corr_features].corr(),annot=True,cmap="RdYlGn")
# + [markdown] _uuid="784b45b84c0d1fe24b7476907a871556f657d1df"
# - From this we can tell which features **(OverallQual, GrLivArea and TotalBsmtSF )** are highly positively correlated with the SalePrice.
# - **GarageCars and GarageArea ** also seems correlated with other, Since the no. of car that will fit into the garage will depend on GarageArea.
# + _uuid="2a7ab1e4534d622b1fd8d1a9c656607c7243b24a"
sns.barplot(train.OverallQual,train.SalePrice)
# + [markdown] _uuid="8c066456b14f788bd5f85f7ffbb271f039df4b18"
# **Scatter plots between 'SalePrice' and correlated variables**
# + _uuid="df041ffe64ef1807796a75237a79fe846d73be82"
sns.set()
cols = ['SalePrice', 'OverallQual', 'GrLivArea', 'GarageCars', 'TotalBsmtSF', 'FullBath', 'YearBuilt']
sns.pairplot(train[cols], size = 2.5)
plt.show();
# + [markdown] _uuid="a3d3e339ff75571f28b9c2787d3c56ab2a9895e9"
# One of the figures we may find interesting is the one between ** 'TotalBsmtSF' and 'GrLiveArea'. **
#
# We can see the dots drawing a linear line, which almost acts like a border. It totally makes sense that the majority of the dots stay below that line. Basement areas can be equal to the above ground living area, but it is not expected a basement area bigger than the above ground living area
# + _uuid="535da1efb310261ff5d59751be4b267ddf1bcd6c"
sns.scatterplot(train.GrLivArea,train.TotalBsmtSF)
# + [markdown] _uuid="7a8333803c5f5efb2fee696d0a2e50429858e575"
# ## Target Variable Transform
# Different features in the data set may have values in different ranges. For example, in this data set, the range of SalePrice feature may lie from thousands to lakhs but the range of values of YearBlt feature will be in thousands. That means a column is more weighted compared to other.
#
# **Lets check the skewness of data**
# 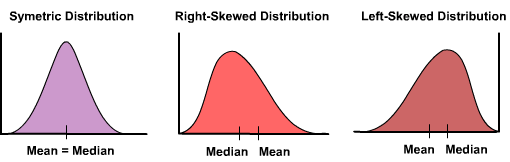
# + _uuid="3e82f95ef5565aadf0ad956f3f70a6fd8a40fe13"
def check_skewness(col):
sns.distplot(train[col] , fit=norm);
fig = plt.figure()
res = stats.probplot(train[col], plot=plt)
# Get the fitted parameters used by the function
(mu, sigma) = norm.fit(train[col])
print( '\n mu = {:.2f} and sigma = {:.2f}\n'.format(mu, sigma))
check_skewness('SalePrice')
# + [markdown] _uuid="c368eceae3e58e9b7ef37bb36737e90a1aa4d87b"
# **This distribution is positively skewed.** Notice that the black curve is more deviated towards the right. If you encounter that your predictive (response) variable is skewed, it is **recommended to fix the skewness** to make good decisions by the model.
#
# ## Okay, So how do I fix the skewness?
# The best way to fix it is to perform a **log transform** of the same data, with the intent to reduce the skewness.
# + _uuid="ad4524d38a8b0c31b3bac90b40daaa558cf7e91c"
#We use the numpy fuction log1p which applies log(1+x) to all elements of the column
train["SalePrice"] = np.log1p(train["SalePrice"])
check_skewness('SalePrice')
# + [markdown] _uuid="fe44621d16a89a2bca2543638690a27e23cee36f"
# After taking logarithm of the same data the curve seems to be normally distributed, although not perfectly normal, this is sufficient to fix the issues from a skewed dataset as we saw before.
#
# **Important : If you log transform the response variable, it is required to also log transform feature variables that are skewed.**
# + [markdown] _uuid="c814c57999220ce366d39db839cb4d4b9374198a"
# # Feature Engineering
# + [markdown] _uuid="c5370e7b469139b439350faed9cf17ef01af1516"
# Here is the [Documentation](http://ww2.amstat.org/publications/jse/v19n3/Decock/DataDocumentation.txt) you can refer , to know more about the dataset.
#
# **Concatenate both train and test values.**
# + _uuid="039f107f3a6b7379ff2627d48ab98c22737825f9"
ntrain = train.shape[0]
ntest = test.shape[0]
y_train = train.SalePrice.values
all_data = pd.concat((train, test)).reset_index(drop=True)
all_data.drop(['SalePrice'], axis=1, inplace=True)
print("all_data size is : {}".format(all_data.shape))
# + [markdown] _uuid="7a6776afb61d2a1637e3003204671d0c0c013ebf"
# # Missing Data
# + _uuid="ad3c34ad8df7a1bbb2b818a1aa0750f91645956f"
all_data_na = (all_data.isnull().sum() / len(all_data)) * 100
all_data_na = all_data_na.drop(all_data_na[all_data_na == 0].index).sort_values(ascending=False)[:30]
missing_data = pd.DataFrame({'Missing Ratio' :all_data_na})
# + _uuid="30af0ffa63d2424affea6028b53c9709e08e0099"
f, ax = plt.subplots(figsize=(15, 12))
plt.xticks(rotation='90')
sns.barplot(x=all_data_na.index, y=all_data_na)
plt.xlabel('Features', fontsize=15)
plt.ylabel('Percent of missing values', fontsize=15)
plt.title('Percent missing data by feature', fontsize=15)
# + _uuid="00773902333384da2515e01f5cd550d5c6a6812f"
all_data.PoolQC.loc[all_data.PoolQC.notnull()]
# + [markdown] _uuid="116000b37a24504895a7b8c7f50bb05831f203c8"
# **GarageType, GarageFinish, GarageQual, GarageCond, GarageYrBlt, GarageArea, GarageCars these all features have same percentage of null values.**
# + [markdown] _uuid="8e2db4ef580f375e4dc7182afd83da4efea5241f"
# # Handle Missing Data
# + [markdown] _uuid="226670f827161cbd4cd5f15c8fae4c83490f851f"
# Since PoolQC has the highest null values according to the data documentation says **null values means 'No Pool.**
# Since majority of houses has no pool.
# So we will replace those null values with 'None'.
# + _uuid="e94513f7cb4dfd3f5e87774b736a39fa1e65a66b"
all_data["PoolQC"] = all_data["PoolQC"].fillna("None")
# + [markdown] _uuid="f0f24b2ecc5283adf1b777b4b6209ecaba69d2f9"
# * **MiscFeature** : Data documentation says NA means "no misc feature"
# + _uuid="f8be0d01feb00595640a0ae90db0d877d361b46a"
all_data["MiscFeature"] = all_data["MiscFeature"].fillna("None")
# + [markdown] _uuid="22dd51000c69deb9b7debe72ec800cbd42d731e4"
# * **Alley** : data description says NA means "no alley access"
#
# + _uuid="09c05ccc29ed49353365d6a0f74d715cb3b4e4da"
all_data["Alley"] = all_data["Alley"].fillna("None")
# + [markdown] _uuid="0b54e59904fc40a3bc1a1a937c890c145cef6f87"
# * **Fence** : data description says NA means "no fence"
#
# + _uuid="c7bfad1d5982ea62ed418b5a556f5da363c8f99d"
all_data["Fence"] = all_data["Fence"].fillna("None")
# + [markdown] _uuid="ebfb8252b25107a6417882fb5244a8e808bdf324"
# * **FireplaceQu** : data description says NA means "no fireplace"
# + _uuid="2eb64243b5e2eb759411e21d0d8e0c80cc6d16f7"
all_data["FireplaceQu"] = all_data["FireplaceQu"].fillna("None")
# + [markdown] _uuid="af166059bccd2bcc56d684f233a7b9a8a9c2e6ea"
# * **LotFrontage** : Since the area of each street connected to the house property most likely have a similar area to other houses in its neighborhood , we can fill in missing values by the median LotFrontage of the neighborhood.
# + _uuid="3ae0bce2d27efc9ed1eb629a73f93c643f309c66" _kg_hide-input=true
# Grouping by Neighborhood and Check the LotFrontage. Most of the grouping has similar areas
grouped_df = all_data.groupby('Neighborhood')['LotFrontage']
for key, item in grouped_df:
print(key,"\n")
print(grouped_df.get_group(key))
break
# + _uuid="1e899c274313d88114288c1e0fa720468da1afee"
#Group by neighborhood and fill in missing value by the median LotFrontage of all the neighborhood
all_data["LotFrontage"] = all_data.groupby("Neighborhood")["LotFrontage"].transform(
lambda x: x.fillna(x.median()))
# + [markdown] _uuid="e1493a84b2e9ebef26d03295511d52830343ace4"
# * **GarageType, GarageFinish, GarageQual and GarageCond** : Replacing missing data with None as per documentation.
# + _kg_hide-output=true _uuid="7a71d08cd1dd160b5184cfd8a681503ec0d92e95"
for col in ['GarageType', 'GarageFinish', 'GarageQual', 'GarageCond']:
all_data[col] = all_data[col].fillna('None')
# + _uuid="0f30a319433e86234355ee6d08ef8e95a23285d3"
abc = ['GarageType', 'GarageFinish', 'GarageQual', 'GarageCond','GarageYrBlt', 'GarageArea', 'GarageCars']
all_data.groupby('GarageType')[abc].count()
# + [markdown] _uuid="197c97edb46edc4ee356d587c66a9a9fca41a2be"
# * **GarageYrBlt, GarageArea and GarageCars** : Replacing missing data with 0 (Since No garage = no cars in such garage.)
# + _uuid="e1c064cc7d91c1f318bbea0be6350a3da14b74cc"
for col in ('GarageYrBlt', 'GarageArea', 'GarageCars'):
all_data[col] = all_data[col].fillna(0)
# + [markdown] _uuid="8cd265faa79ccd0366ec63d691059933153feafc"
# * **BsmtFinSF1, BsmtFinSF2, BsmtUnfSF, TotalBsmtSF, BsmtFullBath and BsmtHalfBath** : missing values are likely zero for having no basement
# + _uuid="ed2d1168ba88ed3f1aed1e0843f9f57e7cd2046d"
for col in ('BsmtFinSF1', 'BsmtFinSF2', 'BsmtUnfSF','TotalBsmtSF', 'BsmtFullBath', 'BsmtHalfBath'):
all_data[col] = all_data[col].fillna(0)
# + [markdown] _uuid="587e73a91191f0ea1cc0fd12b21d8fe45eb403ec"
# * **BsmtQual, BsmtCond, BsmtExposure, BsmtFinType1 and BsmtFinType2** : For all these categorical basement-related features, NaN means that there is no basement.
# + _uuid="6de1156fca72633eb45fbb8c7c926613455ef25f"
for col in ('BsmtQual', 'BsmtCond', 'BsmtExposure', 'BsmtFinType1', 'BsmtFinType2'):
all_data[col] = all_data[col].fillna('None')
# + [markdown] _uuid="737f23b205e2cc84486ddb7a087509303f4b7a5b"
# * **MasVnrArea and MasVnrType** : NA most likely means no masonry veneer for these houses. We can fill 0 for the area and None for the type.
# + _uuid="de69391d40cc5c23765711f4ab248bf61b44bda5"
all_data["MasVnrType"] = all_data["MasVnrType"].fillna("None")
all_data["MasVnrArea"] = all_data["MasVnrArea"].fillna(0)
# + [markdown] _uuid="af7a8d719e79c38f25758488c4f4cb695e3e1976"
# * **MSZoning (The general zoning classification)** : 'RL' is by far the most common value. So we can fill in missing values with 'RL'
# + _uuid="ac0a52cd03ada0f9ab04f219972342753321798e"
all_data['MSZoning'].value_counts()
# + _uuid="7b92b9424d4810edb0d60b03c9316bf8f3a85263"
all_data['MSZoning'] = all_data['MSZoning'].fillna(all_data['MSZoning'].mode()[0])
# + [markdown] _uuid="19602e120fbb71d1fb38290e2330df79b6a053a0"
# * **Utilities** : Since this is a categorical data and most of the data are of same category, Its not gonna effect on model. So we choose to drop it.
# + _uuid="7e44495f79c3deb3757a6f5c23ed597bea0782de"
all_data['Utilities'].value_counts()
# + _uuid="25c2a42c8d23197a1db22b487ef8692da1063e8c"
all_data = all_data.drop(['Utilities'], axis=1)
# + [markdown] _uuid="b82cc3582a64e4d20ead3cd386b2bfb467684353"
# * **Functional** : data description says NA means typical
# + _uuid="af05ef25e7ee2c0e4df51b2df09424112e7fc72f"
all_data["Functional"] = all_data["Functional"].fillna("Typ")
# + [markdown] _uuid="4e9723638293fceec4190dcccc49efd719a28147"
# * **Electrical,KitchenQual, Exterior1st, Exterior2nd, SaleType** : Since this all are categorical values so its better to replace nan values with the most used keyword.
# + _uuid="fd68c6cc53b0f09c99fbaae220455358acba10a5"
mode_col = ['Electrical','KitchenQual', 'Exterior1st', 'Exterior2nd', 'SaleType']
for col in mode_col:
all_data[col] = all_data[col].fillna(all_data[col].mode()[0])
# + [markdown] _uuid="9533bc18b3a5508e78f32471cbacc0d84114a0cc"
# * **MSSubClass** : Na most likely means No building class. We can replace missing values with None
#
# + _uuid="0aa7e338dca4dbb303563fd401bddf69fa517a45"
all_data['MSSubClass'] = all_data['MSSubClass'].fillna("None")
# + [markdown] _uuid="1dece935c4d58ebbbdc20b476ac1ba0c1f49398d"
# ## Lets check for any missing values
# + _uuid="cb249dde6e25900ed562f1106c652c63a2aef72e"
#Check remaining missing values if any
all_data_na = (all_data.isnull().sum() / len(all_data)) * 100
all_data_na = all_data_na.drop(all_data_na[all_data_na == 0].index).sort_values(ascending=False)
missing_data = pd.DataFrame({'Missing Ratio' :all_data_na})
missing_data.head()
# + [markdown] _uuid="fce152364a35ce50b50dc77200f9c25c4709a3c5"
# **Now there any many features that are numerical but categorical.**
# + _uuid="c99d01bd5a5ad8deccf993cc5c571b5f7f11741b"
all_data['OverallCond'].value_counts()
# + [markdown] _uuid="713f061e916a6d7326a86f64657c2a8573cf98b7"
# **Converting some numerical variables that are really categorical type.**
#
# As you can see the category range from 1 to 9 which are numerical (**not ordinal type**). Since its categorical we need to change it to String type.
#
# If we do not convert these to categorical, some model may get affect by this as model will compare the value 1<5<10 . We dont need that to happen with our model.
# + _uuid="62ebb5cfcd932164176a9614bc688a1107799415"
#MSSubClass=The building class
all_data['MSSubClass'] = all_data['MSSubClass'].apply(str)
#Changing OverallCond into a categorical variable
all_data['OverallCond'] = all_data['OverallCond'].astype(str)
#Year and month sold are transformed into categorical features.
all_data['YrSold'] = all_data['YrSold'].astype(str)
all_data['MoSold'] = all_data['MoSold'].astype(str)
# + [markdown] _uuid="4ca8b2ed84fcc54557f4d7b65e4aad367395ab6b"
# ## Label Encoding
# As you might know by now, we can’t have text in our data if we’re going to run any kind of model on it. So before we can run a model, we need to make this data ready for the model.
#
# And to convert this kind of categorical text data into model-understandable numerical data, we use the Label Encoder class.
#
# Suppose, we have a feature State which has 3 category i.e India , France, China . So, Label Encoder will categorize them as 0, 1, 2.
# + _uuid="8308b428bbf30e9e17f17b57a230ed2297081370"
from sklearn.preprocessing import LabelEncoder
cols = ('FireplaceQu', 'BsmtQual', 'BsmtCond', 'GarageQual', 'GarageCond',
'ExterQual', 'ExterCond','HeatingQC', 'PoolQC', 'KitchenQual', 'BsmtFinType1',
'BsmtFinType2', 'Functional', 'Fence', 'BsmtExposure', 'GarageFinish', 'LandSlope',
'LotShape', 'PavedDrive', 'Street', 'Alley', 'CentralAir', 'MSSubClass', 'OverallCond',
'YrSold', 'MoSold')
# process columns, apply LabelEncoder to categorical features
for c in cols:
lbl = LabelEncoder()
lbl.fit(list(all_data[c].values))
all_data[c] = lbl.transform(list(all_data[c].values))
# shape
print('Shape all_data: {}'.format(all_data.shape))
# + [markdown] _uuid="865590a345058e1fd3bb76434441508a0204f6fd"
# Since area related features are very important to determine house prices, we add one more feature which is the total area of basement, first and second floor areas of each house
# + _uuid="b4ddb4163fc92e8a9b2efdb3957965e8d6d65573"
# Adding total sqfootage feature
all_data['TotalSF'] = all_data['TotalBsmtSF'] + all_data['1stFlrSF'] + all_data['2ndFlrSF']
# + [markdown] _uuid="2c4c4d9d412c284b0aa9f34653c193227e36ce07"
# **Lets see the highly skewed features we have**
# + _uuid="0b3f732620e7b42d8c8592c0bef8e9030bcea37c"
numeric_feats = all_data.dtypes[all_data.dtypes != "object"].index
# Check the skew of all numerical features
skewed_feats = all_data[numeric_feats].apply(lambda x: skew(x.dropna())).sort_values(ascending=False)
print("\nSkew in numerical features: \n")
skewness = pd.DataFrame({'Skew' :skewed_feats})
skewness.head(15)
# + [markdown] _uuid="8917aa598b2df98fd173d63bde1f1f941d919d86"
# ## Box Cox Transformation of (highly) skewed features
#
# When you are dealing with real-world data, you are going to deal with features that are heavily skewed. Transformation technique is useful to **stabilize variance**, make the data **more normal distribution-like**, improve the validity of measures of association.
#
# The problem with the Box-Cox Transformation is **estimating lambda**. This value will depend on the existing data, and should be considered when performing cross validation on out of sample datasets.
# + _uuid="04b1a8240b20f470d326c1f480a0061712c30843"
skewness = skewness[abs(skewness) > 0.75]
print("There are {} skewed numerical features to Box Cox transform".format(skewness.shape[0]))
from scipy.special import boxcox1p
skewed_features = skewness.index
lam = 0.15
for feat in skewed_features:
#all_data[feat] += 1
all_data[feat] = boxcox1p(all_data[feat], lam)
# + [markdown] _uuid="aef52487095b221359382ce9135c709541c03958"
# **Getting dummy categorical features**
# + _uuid="d3056bb177dd80797d752d80e0d9d943486e482a"
all_data = pd.get_dummies(all_data)
all_data.shape
# + [markdown] _uuid="142bcae6537641dc5307beb7186a1a9a709fb21a"
# Creating train and test data.
# + _uuid="0c986a9c705e012679c661f03a017399978d6ebd"
train = all_data[:ntrain]
test = all_data[ntrain:]
train.shape
# + [markdown] _uuid="d5c3bdaf7c57955a06d4537e93ad7036af1e54f1"
# ## Lets apply Modelling
#
# 1. Importing Libraries
#
# 2. We will use models
# - Lasso
# - Ridge
# - ElasticNet
# - Gradient Boosting
#
# 3. Find the Cross Validation Score.
# 4. Calculate the mean of all model's prediction.
# 5. Submit the CSV file.
#
# + _uuid="a477c9212c17282e5c8a1767ca6b301e91afcce3"
from sklearn.linear_model import ElasticNet, Lasso, BayesianRidge, LassoLarsIC
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
from sklearn.kernel_ridge import KernelRidge
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.base import BaseEstimator, TransformerMixin, RegressorMixin, clone
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.metrics import mean_squared_error
import xgboost as xgb
import lightgbm as lgb
# + [markdown] _uuid="a907ce9421f8a0b628390c5db13920f9a392d476"
# ## Cross Validation
# It's simple way to calculate error for evaluation.
#
# **KFold( )** splits the train/test data into k consecutive folds, we also have made shuffle attrib to True.
#
# **cross_val_score ( )** evaluate a score by cross-validation.
# + _uuid="d7af5d935df2a27cfc54dc13f746aa64774d010f"
#Validation function
n_folds = 5
def rmsle_cv(model):
kf = KFold(n_folds, shuffle=True, random_state=42).get_n_splits(train.values)
rmse= np.sqrt(-cross_val_score(model, train.values, y_train, scoring="neg_mean_squared_error", cv = kf))
return(rmse)
# + [markdown] _uuid="ef4bf7c58fdfedd15102194023c24b94876f3559"
# # Modelling
# Since in this dataset we have a large set of features. So to make our model avoid Overfitting and noisy we will use Regularization.
# These model have Regularization parameter.
#
# Regularization will reduce the magnitude of the coefficients.
# + [markdown] _uuid="c7cd0953ca1b7021b165aef700d1241732c09d18"
# ## Ridge Regression
# - It shrinks the parameters, therefore it is mostly used to prevent multicollinearity.
# - It reduces the model complexity by coefficient shrinkage.
# - It uses L2 regularization technique.
# + _uuid="b7f81c6d917d9c5325f3d3456bde7adce2899621"
KRR = KernelRidge(alpha=0.6, kernel='polynomial', degree=2, coef0=2.5)
score = rmsle_cv(KRR)
print("Kernel Ridge score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# + [markdown] _uuid="03559fcf57b62b49b0332acd9439274bf8dd9d8a"
# ## Lasso Regression
# LASSO (Least Absolute Shrinkage Selector Operator), is quite similar to ridge.
#
# In case of lasso, even at smaller alpha’s, our coefficients are reducing to absolute zeroes.
# Therefore, lasso selects the only some feature while reduces the coefficients of others to zero. This property is known as feature selection and which is absent in case of ridge.
#
# - Lasso uses L1 regularization technique.
# - Lasso is generally used when we have more number of features, because it automatically does feature selection.
#
# + _uuid="a437402e2a37f26372fc03761fa05c6d7ea4e433"
lasso = make_pipeline(RobustScaler(), Lasso(alpha =0.0005, random_state=1))
score = rmsle_cv(lasso)
print("Lasso score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# + [markdown] _uuid="c854d61f20f4cdab8a37b5037c3908f783b5e644"
# ## Elastic Net Regression
#
# Elastic net is basically a combination of both L1 and L2 regularization. So if you know elastic net, you can implement both Ridge and Lasso by tuning the parameters.
# + _uuid="d06ca9a5f9db49890db7999ffe9db7333f02edc6"
ENet = make_pipeline(RobustScaler(), ElasticNet(alpha=0.0005, l1_ratio=.9, random_state=3))
score = rmsle_cv(ENet)
print("ElasticNet score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# + [markdown] _uuid="120e4a5ed78c963d8278803cc00956781f605691"
# ## Gradient Boosting Regression
# Refer [here](https://medium.com/mlreview/gradient-boosting-from-scratch-1e317ae4587d)
# + _uuid="221e05d63ac4d3f99900b36a9d06e3d8e10f1dc7"
GBoost = GradientBoostingRegressor(n_estimators=3000, learning_rate=0.05,
max_depth=4, max_features='sqrt',
min_samples_leaf=15, min_samples_split=10,
loss='huber', random_state =5)
score = rmsle_cv(GBoost)
print("Gradient Boosting score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# + [markdown] _uuid="7c0a8d859b163d5f10ae59afdd6fd77664bcd907"
# **Fit the training dataset on every model**
# + _uuid="f2a04068b4c93f1a1851708d9f43edcef6990cb8"
LassoMd = lasso.fit(train.values,y_train)
ENetMd = ENet.fit(train.values,y_train)
KRRMd = KRR.fit(train.values,y_train)
GBoostMd = GBoost.fit(train.values,y_train)
# + [markdown] _uuid="b5b2fe94eaa417646e8a91b451db40b400e88bf6"
# ## Mean of all model's prediction.
# np.expm1 ( ) is used to calculate exp(x) - 1 for all elements in the array.
# + _uuid="8b4e59cad1a1499ba00c3206676676658a4b1881"
finalMd = (np.expm1(LassoMd.predict(test.values)) + np.expm1(ENetMd.predict(test.values)) + np.expm1(KRRMd.predict(test.values)) + np.expm1(GBoostMd.predict(test.values)) ) / 4
finalMd
# + [markdown] _uuid="0cd4ca41a0ecde7f82204ac5ed6fe6b502ea4a87"
# ## Submission
# + _uuid="708eb2603a04b74c2c19ed52f5130c5f5704cf0f"
sub = pd.DataFrame()
sub['Id'] = test_ID
sub['SalePrice'] = finalMd
sub.to_csv('submission.csv',index=False)
# + [markdown] _uuid="b6797c8782e8aee1a187da8b2c65b67803853439"
# **If you found this notebook helpful or you just liked it , some upvotes would be very much appreciated.**
#
# **I'll be glad to hear suggestions on improving my models**
| 8 HOUSE PRICES/reach-top-10-with-simple-model-on-housing-prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# +
from dipy.reconst.dti import fractional_anisotropy, color_fa
from argparse import ArgumentParser
from scipy import ndimage
import os
import re
import numpy as np
import nibabel as nb
import sys
import matplotlib
matplotlib.use('Agg') # very important above pyplot import
import matplotlib.pyplot as plt
# -
# cd
# ls
from dipy.reconst.dti import from_lower_triangular
img = nb.load('v100_ch0_tensorfsl_dogsig1_gausig2.3.nii')
data = img.get_data()
output = from_lower_triangular(data)
output_ds = output[4250:4300, 250:300, :, :, :]
# +
print output.shape
print output_ds.shape
# -
FA = fractional_anisotropy(output_ds)
FA = np.clip(FA, 0, 1)
FA[np.isnan(FA)] = 0
# +
print FA.shape
# +
from dipy.reconst.dti import decompose_tensor
# -
evalues, evectors = decompose_tensor(output_ds)
print evectors[..., 0, 0].shape
print evectors.shape[-2:]
print FA[:, :, :, 0].shape
RGB = color_fa(FA[:, :, :, 0], evectors)
nb.save(nb.Nifti1Image(np.array(255 * RGB, 'uint8'), img.get_affine()), 'tensor_rgb_upper.nii.gz')
def plot_rgb(im):
plt.rcParams.update({'axes.labelsize': 'x-large',
'axes.titlesize': 'x-large'})
if im.shape == (182, 218, 182):
x = [78, 90, 100]
y = [82, 107, 142]
z = [88, 103, 107]
else:
shap = im.shape
x = [int(shap[0]*0.35), int(shap[0]*0.51), int(shap[0]*0.65)]
y = [int(shap[1]*0.35), int(shap[1]*0.51), int(shap[1]*0.65)]
z = [int(shap[2]*0.35), int(shap[2]*0.51), int(shap[2]*0.65)]
coords = (x, y, z)
labs = ['Sagittal Slice (YZ fixed)',
'Coronal Slice (XZ fixed)',
'Axial Slice (XY fixed)']
var = ['X', 'Y', 'Z']
idx = 0
for i, coord in enumerate(coords):
for pos in coord:
idx += 1
ax = plt.subplot(3, 3, idx)
ax.set_title(var[i] + " = " + str(pos))
if i == 0:
image = ndimage.rotate(im[pos, :, :], 90)
elif i == 1:
image = ndimage.rotate(im[:, pos, :], 90)
else:
image = im[:, :, pos]
if idx % 3 == 1:
ax.set_ylabel(labs[i])
ax.yaxis.set_ticks([0, image.shape[0]/2, image.shape[0] - 1])
ax.xaxis.set_ticks([0, image.shape[1]/2, image.shape[1] - 1])
plt.imshow(image)
fig = plt.gcf()
fig.set_size_inches(12.5, 10.5, forward=True)
return fig
# +
affine = img.get_affine()
fa = nb.Nifti1Image(np.array(255 * RGB, 'uint8'), affine)
im = fa.get_data()
# -
print np.asarray(fa)
fig = plot_rgb(im)
import os
# cd /root/seelviz/Tony/aut1367/aut1367_raw/v100/ch0
# ls
from PIL import Image
im = plt.imread('RAWoutfileaut1367_3.tiff')
plt.imshow(im)
| Albert_Jupyter/Final+Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Deploy Watson ML (PMML)
# Deploys a PMML model to IBM Watson Machine Learning (WML)
# %pip install ibm-watson-machine-learning
# PLEASE RESTART YOUR KERNAL AFTER THIS LINE HAS BEEN EXECUTED
import wget
from ibm_watson_machine_learning import APIClient
import os
import sys
# +
# IBM Cloud API Key https://cloud.ibm.com/iam/apikeys
api_key = os.environ.get('api_key','<replace with your api key>')
# Machine Learning Model Deployment Space https://dataplatform.cloud.ibm.com/ml-runtime/spaces
space = os.environ.get('space','<replace with your space id>')
# IBM Cloud Region (e.g. us-south)
location = os.environ.get('location','<replace with your location>')
# temporary directory for data
data_dir = os.environ.get('data_dir',
'../../data/')
# +
parameters = list(
map(
lambda s: re.sub('$', '"', s),
map(
lambda s: s.replace('=', '="'),
filter(
lambda s: s.find('=') > -1 and bool(re.match('[A-Za-z0-9_]*=[.\/A-Za-z0-9]*', s)),
sys.argv
)
)
)
)
for parameter in parameters:
logging.warning('Parameter: '+parameter)
exec(parameter)
# -
wml_credentials = {
"apikey": api_key,
"url": 'https://' + location + '.ml.cloud.ibm.com'
}
client = APIClient(wml_credentials)
o = client.software_specifications.get_uid_by_name('spark-mllib_2.4')
software_spec_uid = o
client.set.default_space(space)
# +
model_meta_props = {
client.repository.ModelMetaNames.NAME: 'test_pmml2',
client.repository.ModelMetaNames.TYPE: "pmml_4.2",
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: software_spec_uid
}
published_model = client.repository.store_model(
model=data_dir + 'model.xml',
meta_props=model_meta_props,
)
model_uid = client.repository.get_model_uid(published_model)
| component-library/deploy/deploy_wml_pmml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [webinar]
# language: python
# name: Python [webinar]
# ---
# # Demo 1 - Solving a model file
# ### Step 1: Import functions from the gurobipy module
from gurobipy import *
# ### Step 2: Create model object from model file
model = read("afiro.mps")
# ### Step 3: Solve model to optimality
model.optimize()
# ### Step 4: Display optimal objective value
model.ObjVal
# ### Step 5: Display variable values
model.printAttr('X')
| docs/Gurobi/notebooks/Demo 1 - Solving a model file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.random.seed(42)
from sklearn.datasets import load_iris
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
# Aufgabe 1: Lade das Dataset mit allen Features rein.
dataset = load_iris()
x = dataset.data[:, :]
y = dataset.target
# Aufgabe 2: Wende das k-Means mit einer beliebigen Anzahl an Cluster an.
# Aufgabe 3: Berechne den Score für verschiedene Setups.
| Chapter10_Clustering/KMeansExercise/KMeans_exercise.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// ---
// +
// By default Synapse uses AAD passthrough for authentication
// TokenLibrary is invoked under the hood for obtaining AAD token and using it for
// authenticating against the resource
val df = spark.read.parquet("abfss://..")
// +
// While Gen2 is the default storage for Synapse, AAD passthrough support exists for Gen1 as well. PLEASE NOTE THAT WE DO NOT OFFICIALLY SUPPORT GEN1 IN SYNAPSE AND CUSTOMERS WHO USE IT ARE ON THEIR OWN.
val df = spark.read.parquet("adl://")
// +
// Linked services can be used for storing and retreiving credentials (e.g, account key)
// Example connection string (for storage): "DefaultEndpointsProtocol=https;AccountName=<accountname>;AccountKey=<accountkey>"
val connectionString: String = TokenLibrary.getConnectionString("<linkedServiceName>")
val accountKey: String = TokenLibrary.getConnectionStringAsMap("<linkedServiceName>").get("AccountKey")
// +
// The following feature is in the works. Synapse will have inbuilt integration of linked services with storage Gen2 via TokenLibrary
// For storage Gen2, linkedServiceName can be supplied through config for SAS-key based authentication (in lieu of account-key based authentication)
// Direct invokation of TokenLibrary is not required for obtaining creds and connection info
val sc = spark.sparkContext
sc.hadoopConfiguration.set("spark.storage.synapse.linkedServiceName", "<linkedServiceName>")
sc.hadoopConfiguration.set("fs.azure.account.auth.type", "SAS")
sc.hadoopConfiguration.set("fs.azure.sas.token.provider.type", "com.microsoft.azure.synapse.tokenlibrary.LinkedServiceBasedSASProvider")
val df = spark.read.parquet("abfss://..")
// -
| Notebooks/Scala/Obtain credentials using TokenLibrary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Writing Containers to a tabular format
#
# The `TableWriter`/`TableReader` sub-classes allow you to write a `ctapipe.core.Container` class and its meta-data to an output table. They treat the `Field`s in the `Container` as columns in the output, and automatically generate a schema. Here we will go through an example of writing out data and reading it back with *Pandas*, *PyTables*, and a `ctapipe.io.TableReader`:
#
# In this example, we will use the `HDF5TableWriter`, which writes to HDF5 datasets using *PyTables*. Currently this is the only implemented TableWriter.
# Caveats to think about:
# * vector columns in Containers *can* be written, but some lilbraries like Pandas can not read those (so you must use pytables or astropy to read outputs that have vector columns)
# * units are stored in the table metadata, but some libraries like Pandas ignore them and all other metadata
# ## Create some example Containers
from ctapipe.io import HDF5TableWriter
from ctapipe.core import Container, Field
from astropy import units as u
import numpy as np
class VariousTypesContainer(Container):
a_int = Field(int, 'some int value')
a_float = Field(float, 'some float value with a unit', unit=u.m)
a_bool = Field(bool, 'some bool value')
a_np_int = Field(np.int64, 'a numpy int')
a_np_float = Field(np.float64, 'a numpy float')
a_np_bool = Field(np.bool_, 'np.bool')
# let's also make a dummy stream (generator) that will create a series of these containers
def create_stream(n_event):
data = VariousTypesContainer()
for i in range(n_event):
data.a_int = int(i)
data.a_float = float(i) * u.cm # note unit conversion will happen
data.a_bool = (i % 2) == 0
data.a_np_int = np.int64(i)
data.a_np_float = np.float64(i)
data.a_np_bool = np.bool((i % 2) == 0)
yield data
for data in create_stream(2):
for key, val in data.items():
print('{}: {}, type : {}'.format(key, val, type(val)))
# ## Writing the Data (and good practices)
# ### How not to do it:
# +
h5_table = HDF5TableWriter('container.h5', group_name='data')
for data in create_stream(10):
h5_table.write('table', data)
h5_table.close()
# -
# In that case the file is not garenteed to close properly for instance if one does a mistake in the for loop. Let's just add a stupid mistake and see what happens.
try:
h5_table = HDF5TableWriter('container.h5', group_name='data')
for data in create_stream(10):
h5_table.write('table', data)
0/0 # cause an error
h5_table.close()
except Exception as err:
print("FAILED!", err)
# Now the file did not close properly. So let's try to correct the mistake and execute the code again.
try:
h5_table = HDF5TableWriter('container.h5', group_name='data')
for data in create_stream(10):
h5_table.write('table', data)
0/0 # cause an error
h5_table.close()
except Exception as err:
print("FAILED!", err)
# Ah it seems that the file did not close! Now I am stuck. Maybe I should restart the kernel? ahh no I don't want to loose everything. Can I just close it ?
h5_table.close()
# It worked!
# ### Better to use context management!
try:
with HDF5TableWriter('container.h5', group_name='data') as h5_table:
for data in create_stream(10):
h5_table.write('table', data)
0/0
except Exception as err:
print("FAILED:", err)
print('Done')
# !ls container.h5
# ## Appending new Containers
# To append some new containers we need to set the writing in append mode by using: 'mode=a'. But let's now first look at what happens if we don't.
for i in range(2):
with HDF5TableWriter('container.h5', mode='w', group_name='data_{}'.format(i)) as h5_table:
for data in create_stream(10):
h5_table.write('table', data)
print(h5_table._h5file)
# !rm -f container.h5
# Ok so the writer destroyed the content of the file each time it opens the file. Now let's try to append some data group to it! (using mode='a')
for i in range(2):
with HDF5TableWriter('container.h5', mode='a', group_name='data_{}'.format(i)) as h5_table:
for data in create_stream(10):
h5_table.write('table', data)
print(h5_table._h5file)
# So we can append some data groups. As long as the data group_name does not already exists. Let's try to overwrite the data group : data_1
try:
with HDF5TableWriter('container.h5', mode='a', group_name='data_1') as h5_table:
for data in create_stream(10):
h5_table.write('table', data)
except Exception as err:
print("Failed as expected:", err)
# Good ! I cannot overwrite my data.
print(bool(h5_table._h5file.isopen))
# ## Reading the Data
# ### Reading the whole table at once:
#
# For this, you have several choices. Since we used the HDF5TableWriter in this example, we have at least these options avilable:
#
# * Pandas
# * PyTables
# * Astropy Table
#
# For other TableWriter implementations, others may be possible (depending on format)
#
# #### Reading with Pandas:
#
# Pandas is a convenient way to read the output. **HOWEVER BE WARNED** that so far Pandas does not support reading the table *meta-data* or *units* for colums, so that information is lost!
# +
import pandas as pd
data = pd.read_hdf('container.h5', key='/data_0/table')
data.head()
# -
# #### Reading with PyTables
import tables
h5 = tables.open_file('container.h5')
table = h5.root['data_0']['table']
table
# note that here we can still access the metadata
table.attrs
# ### Reading one-row-at-a-time:
# Rather than using the full-table methods, if you want to read it row-by-row (e.g. to maintain compatibility with an existing event loop), you can use a `TableReader` instance.
#
# The advantage here is that units and other metadata are retained and re-applied
# +
from ctapipe.io import HDF5TableReader
def read(mode):
print('reading mode {}'.format(mode))
with HDF5TableReader('container.h5', mode=mode) as h5_table:
for group_name in ['data_0/', 'data_1/']:
group_name = '/{}table'.format(group_name)
print(group_name)
for data in h5_table.read(group_name, VariousTypesContainer()):
print(data.as_dict())
# -
read('r')
read('r+')
read('a')
read('w')
| docs/examples/table_writer_reader.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dev35]
# language: python
# name: conda-env-dev35-py
# ---
# ## Route between two random nodes
# +
import numpy as np
import osmnx as ox
import networkx as nx
from sklearn.neighbors import KDTree
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# creating a graph by using a point in downtown Omaha
old_market = (41.255676, -95.931338)
G = ox.graph_from_point(old_market, distance=500)
ox.plot_graph(G, fig_height=10, fig_width=10, edge_color='black')
# using NetworkX to calculate the shortest path between two random nodes
route = nx.shortest_path(G, np.random.choice(G.nodes), np.random.choice(G.nodes))
ox.plot_graph_route(G, route, fig_height=10, fig_width=10)
# ## Route between two points
library = ox.geocode('215 S 15th St, Omaha, NE 68102')
museum = ox.geocode('801 S 10th St, Omaha, NE 68108')
# +
fig, ax = ox.plot_graph(G, fig_height=10, fig_width=10,
show=False, close=False,
edge_color='black')
ax.scatter(library[1], library[0], c='red', s=100)
ax.scatter(museum[1], museum[0], c='blue', s=100)
plt.show()
# -
nodes, _ = ox.graph_to_gdfs(G)
nodes.head()
# +
tree = KDTree(nodes[['y', 'x']], metric='euclidean')
lib_idx = tree.query([library], k=1, return_distance=False)[0]
museum_idx = tree.query([museum], k=1, return_distance=False)[0]
closest_node_to_lib = nodes.iloc[lib_idx].index.values[0]
closest_node_to_museum = nodes.iloc[museum_idx].index.values[0]
# +
fig, ax = ox.plot_graph(G, fig_height=10, fig_width=10,
show=False, close=False,
edge_color='black')
ax.scatter(library[1], library[0], c='red', s=100)
ax.scatter(museum[1], museum[0], c='blue', s=100)
ax.scatter(G.node[closest_node_to_lib]['x'], G.node[closest_node_to_lib]['y'], c='green', s=100)
ax.scatter(G.node[closest_node_to_museum]['x'], G.node[closest_node_to_museum]['y'], c='green', s=100)
plt.show()
# +
route = nx.shortest_path(G, closest_node_to_lib, closest_node_to_museum)
ox.plot_graph_route(G, route, fig_height=10, fig_width=10,
show=False, close=False,
edge_color='black')
ax.scatter(library[1], library[0], c='red', s=100)
ax.scatter(museum[1], museum[0], c='blue', s=100)
plt.show()
# -
import folium
m = ox.plot_route_folium(G, route)
folium.Marker(location=library, icon=folium.Icon(color='red')).add_to(m)
folium.Marker(location=museum).add_to(m)
m
| osmnx_routing/OSMnx_routing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''csls'': conda)'
# name: python3
# ---
import numpy as np
import pickle
from itertools import combinations
import matplotlib.pyplot as plt
import sys
sys.path.append("..") # Adds higher directory to python modules path.
from utils.util import *
from dataset import get_loaders
analyze_name = 'analyze_regression'
# +
with open('../../results/%s_results_mlp.P' %(analyze_name), 'rb') as f:
mlp_results = pickle.load(f)
# ----------------------
lesion_p = 0.1
# ----------------------
ctx_order = 'first'
ctx_order_str = 'ctxF'
# ----------------------
with open('../../results/%s_%s_results_rnn_init1.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnn_results_ctxF = pickle.load(f)
with open('../../results/%s_%s_results_rnn_lesionp%s.P' %(analyze_name, ctx_order_str, lesion_p), 'rb') as f:
rnnlesion_results_ctxF = pickle.load(f)
with open('../../results/%s_%s_results_rnncell_init1.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnncell_results_ctxF = pickle.load(f)
with open('../../results/%s_%s_results_rnn_balanced.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnnb_results_ctxF = pickle.load(f)
# ----------------------
ctx_order = 'last'
ctx_order_str = 'ctxL'
# ----------------------
with open('../../results/%s_%s_results_rnn_init1.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnn_results_ctxL = pickle.load(f)
with open('../../results/%s_%s_results_rnn_lesionp%s.P' %(analyze_name, ctx_order_str, lesion_p), 'rb') as f:
rnnlesion_results_ctxL = pickle.load(f)
with open('../../results/%s_%s_results_rnncell_init1.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnncell_results_ctxL = pickle.load(f)
with open('../../results/%s_%s_results_rnn_balanced.P' %(analyze_name, ctx_order_str), 'rb') as f:
rnnb_results_ctxL = pickle.load(f)
# ----------------------
with open('../../results/%s_results_stepwisemlp.P' %(analyze_name), 'rb') as f:
swmlp_results = pickle.load(f)
with open('../../results/%s_results_truncated_stepwisemlp.P' %(analyze_name), 'rb') as f:
swmlp_trunc_results = pickle.load(f)
# ----------------------
with open('../../results/%s_results_mlp_cc.P' %(analyze_name), 'rb') as f:
mlpcc_results = pickle.load(f)
# -
mlp_runs = dict_to_list(mlp_results, analyze_name)
rnn_runs_ctxF = dict_to_list(rnn_results_ctxF, analyze_name)
rnncell_runs_ctxF = dict_to_list(rnncell_results_ctxF, analyze_name)
rnnb_runs_ctxF = dict_to_list(rnnb_results_ctxF, analyze_name)
rnnlesion_runs_ctxF = dict_to_list(rnnlesion_results_ctxF, analyze_name)
rnn_runs_ctxL = dict_to_list(rnn_results_ctxL, analyze_name)
rnncell_runs_ctxL = dict_to_list(rnncell_results_ctxL, analyze_name)
rnnb_runs_ctxL = dict_to_list(rnnb_results_ctxL, analyze_name)
rnnlesion_runs_ctxL = dict_to_list(rnnlesion_results_ctxL, analyze_name)
swmlp_runs = dict_to_list(swmlp_results, analyze_name)
mlpcc_runs = dict_to_list(mlpcc_results, analyze_name)
mlp_runs.keys()
reg_analyze_name = 'cat_reg'
mlp_cat_runs = dict_to_list(mlp_runs, reg_analyze_name)
rnn_cat_runs_ctxF = dict_to_list(rnn_runs_ctxF, reg_analyze_name)
rnncell_cat_runs_ctxF = dict_to_list(rnncell_runs_ctxF, reg_analyze_name)
rnnb_cat_runs_ctxF = dict_to_list(rnnb_runs_ctxF, reg_analyze_name)
rnnlesion_cat_runs_ctxF = dict_to_list(rnnlesion_runs_ctxF, reg_analyze_name)
rnn_cat_runs_ctxL = dict_to_list(rnn_runs_ctxL, reg_analyze_name)
rnncell_cat_runs_ctxL = dict_to_list(rnncell_runs_ctxL, reg_analyze_name)
rnnb_cat_runs_ctxL = dict_to_list(rnnb_runs_ctxL, reg_analyze_name)
rnnlesion_cat_runs_ctxL = dict_to_list(rnnlesion_runs_ctxL, reg_analyze_name)
swmlp_cat_runs = dict_to_list(swmlp_runs, reg_analyze_name)
mlpcc_cat_runs = dict_to_list(mlpcc_runs, reg_analyze_name)
mlp_cat_runs.keys()
data = get_loaders(batch_size=32, meta=False,
use_images=True, image_dir='../images/',
n_episodes=None, N_responses=None, N_contexts=None,
cortical_task='face_task', balanced=False)
train_data, train_loader, test_data, test_loader, analyze_data, analyze_loader = data
y_hat_Es = mlp_cat_runs['y_hat_E'] # [runs, checkpoints, n_combinations]: [20, 21, 120]
ys = mlp_cat_runs['y']
np.asarray(y_hat_Es).shape
def calc_rsa_dis(results, test_data, cp):
y_hat_Es = np.asarray(results['y_hat_E']) # [runs, checkpoints, n_combinations]: [20, 21, 120]
ys = np.asarray(results['y'])
n_states = test_data.n_states
loc2idx = test_data.loc2idx
idxs = [idx for idx in range(n_states)]
locs = [loc for loc, idx in loc2idx.items()]
wE = ys - y_hat_Es
wE = wE.mean(axis=0)[cp]
# print(wE.shape)
rsa_dist = np.zeros(shape=(n_states, n_states))
for i, (idx1, idx2) in enumerate(combinations(idxs, 2)):
rsa_dist[idx1][idx2] = wE[i]
rsa_dist[idx2][idx1] = wE[i]
return rsa_dist
# +
# reg_analyze_name = 'cat_reg'
# mlp_cat_runs = dict_to_list(mlp_runs, reg_analyze_name)
# rnn_cat_runs_ctxF = dict_to_list(rnn_runs_ctxF, reg_analyze_name)
# rnncell_cat_runs_ctxF = dict_to_list(rnncell_runs_ctxF, reg_analyze_name)
# rnnb_cat_runs_ctxF = dict_to_list(rnnb_runs_ctxF, reg_analyze_name)
# rnn_cat_runs_ctxL = dict_to_list(rnn_runs_ctxL, reg_analyze_name)
# rnncell_cat_runs_ctxL = dict_to_list(rnncell_runs_ctxL, reg_analyze_name)
# rnnb_cat_runs_ctxL = dict_to_list(rnnb_runs_ctxL, reg_analyze_name)
# swmlp_cat_runs = dict_to_list(swmlp_runs, reg_analyze_name)
# mlpcc_cat_runs = dict_to_list(mlpcc_runs, reg_analyze_name)
# mlp_cat_runs.keys()
# -
# which checkpoints
cp_mlp = 5
cp_rnn = 20
# +
rsa_dist_mlp = calc_rsa_dis(mlp_cat_runs, test_data, cp=cp_mlp)
rsa_dist_rnn_ctxF = calc_rsa_dis(rnn_cat_runs_ctxF, test_data, cp = cp_rnn)
rsa_dist_rnncell_ctxF = calc_rsa_dis(rnncell_cat_runs_ctxF, test_data, cp = cp_rnn)
rsa_dist_rnnb_ctxF = calc_rsa_dis(rnnb_cat_runs_ctxF, test_data, cp = cp_rnn)
rsa_dist_rnnlesion_ctxF = calc_rsa_dis(rnnlesion_cat_runs_ctxF, test_data, cp = cp_rnn)
rsa_dist_rnn_ctxL = calc_rsa_dis(rnn_cat_runs_ctxL, test_data, cp = cp_rnn)
rsa_dist_rnncell_ctxL = calc_rsa_dis(rnncell_cat_runs_ctxL, test_data, cp = cp_rnn)
rsa_dist_rnnb_ctxL = calc_rsa_dis(rnnb_cat_runs_ctxL, test_data, cp = cp_rnn)
rsa_dist_rnnlesion_ctxL = calc_rsa_dis(rnnlesion_cat_runs_ctxL, test_data, cp = cp_rnn)
# rsa_dist_swmlp_hidd1 = calc_rsa_dis(swmlp_cat_runs[:,0], test_data)
# rsa_dist_swmlp_hidd2 = calc_rsa_dis(swmlp_cat_runs[:,1], test_data)
rsa_dist_mlpcc = calc_rsa_dis(mlpcc_cat_runs, test_data, cp = cp_rnn)
# -
# ### preprating stepwise mlp separately
# +
cp = cp_mlp
y_hat_Es = np.asarray(swmlp_cat_runs['y_hat_E']) # [runs, checkpoints, n_combinations]: [20, 21, 120]
ys = np.asarray(swmlp_cat_runs['y'])
runs, checkpoints, n_combinations, n_hidds = y_hat_Es.shape
ys_swmlp = np.zeros([runs, checkpoints, n_combinations, n_hidds])
yhats_swmlp = np.zeros([runs, checkpoints, n_combinations, n_hidds])
for r in range(runs):
for cp in range(checkpoints):
yhats_swmlp[r,cp,:,:] = y_hat_Es[r,cp]
ys_swmlp[r,cp,:,:] = ys[r,cp]
n_states = test_data.n_states
loc2idx = test_data.loc2idx
idxs = [idx for idx in range(n_states)]
locs = [loc for loc, idx in loc2idx.items()]
wE = ys_swmlp - yhats_swmlp
wE = wE.mean(axis=0)[cp]
rsa_dist_swmlp = np.zeros(shape=(n_states, n_states, n_hidds))
for i, (idx1, idx2) in enumerate(combinations(idxs, 2)):
for h in range(n_hidds):
rsa_dist_swmlp[idx1,idx2,h] = wE[i,h]
rsa_dist_swmlp[idx2,idx1,h] = wE[i,h]
print(rsa_dist_swmlp.shape)
# -
def plot_rsa(ctx_order, ctx_order_str, val_res, model_str, mfig_str, cp):
fig, axs = plt.subplots()
plt.imshow(val_res, vmin=vmin, vmax=vmax)
plt.xticks(idxs, locs, rotation='90')
plt.yticks(idxs, locs)
plt.colorbar()
if ctx_order is not None:
fig.suptitle('RSA Results at Step %s - %s - Ax %s' %(cp, model_str, ctx_order), fontweight='bold', fontsize='18')
else:
fig.suptitle('RSA Results at Step %s - %s' %(cp, model_str), fontweight='bold', fontsize='18')
plt.tight_layout()
fig_str = '%s_rsa_results_%s' %(ctx_order_str, mfig_str)
fig.savefig(('../../figures/' + fig_str + '.pdf'),
bbox_inches = 'tight', pad_inches=0)
fig.savefig(('../../figures/' + fig_str + '.png'),
bbox_inches = 'tight', pad_inches=0)
# # RNN
vmin, vmax = -1.5, 1.5
plot_rsa('first', 'ctxF', rsa_dist_rnn_ctxF, 'RNN', 'rnn', cp=cp_rnn)
plot_rsa('last', 'ctxL', rsa_dist_rnn_ctxL, 'RNN', 'rnn', cp=cp_rnn)
# # Balanced RNN
plot_rsa('first', 'ctxF', rsa_dist_rnnb_ctxF, 'RNN Balanced', 'rnnbalanced', cp=cp_rnn)
plot_rsa('last', 'ctxL', rsa_dist_rnnb_ctxL, 'RNN Balanced', 'rnnbalanced', cp=cp_rnn)
# # RNNCell
plot_rsa('first', 'ctxF', rsa_dist_rnncell_ctxF, 'RNNCell', 'rnncell', cp=cp_rnn)
plot_rsa('last', 'ctxL', rsa_dist_rnncell_ctxL, 'RNNCell', 'rnncell', cp=cp_rnn)
# # MLP
vmin, vmax = -0.5, 0.5
plot_rsa(None, None, rsa_dist_mlp, 'MLP', 'mlp', cp=cp_mlp)
# # Cognitive Controller
plot_rsa(None, None, rsa_dist_mlpcc, 'Cognitive Controller', 'mlpcc', cp=cp_rnn)
# # Stepwise MLP
plot_rsa(None, None, rsa_dist_swmlp[:,:,0], 'Stepwise MLP - Hidden 1', 'rsa_swmlp_hidd1', cp=cp_mlp)
plot_rsa(None, None, rsa_dist_swmlp[:,:,1], 'Stepwise MLP - Hidden 2', 'rsa_swmlp_hidd2', cp=cp_mlp)
| notebooks/results_RSA.ipynb |