code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from jupyterthemes import jtplot
# choose which theme to inherit plotting style from
# onedork | grade3 | oceans16 | chesterish | monokai | solarizedl | solarizedd
jtplot.style(theme='onedork')
jtplot.style(context='talk', fscale=1.4, spines=False, gridlines='--')
jtplot.style(ticks=True, grid=False, figsize=(6, 4.5))
# jtplot.reset()
# +
# Importing the dataset
dataset = pd.read_csv('templates/SVR/Position_Salaries.csv')
dataset.head()
# -
X = dataset.iloc[:, 1:2].values
y = dataset.iloc[:, 2].values
from sklearn.svm import SVR
svr = SVR()
| Tutorials/SVR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading and Processing Stock-Market Time-Series Data
# ### Introduction
#
# Across many problems in finance, one starts with time series. Here, we showcase how to generate pseudo-random time-series, download actual stock-market time series from a number of common providers, and how to compute time-series similarity measures.
# %matplotlib inline
from qiskit_finance import QiskitFinanceError
from qiskit_finance.data_providers import *
import datetime
import matplotlib.pyplot as plt
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
data = RandomDataProvider(tickers=["TICKER1", "TICKER2"],
start = datetime.datetime(2016, 1, 1),
end = datetime.datetime(2016, 1, 30),
seed = 1)
data.run()
# Once the data are loaded, you can run a variety of algorithms on those to aggregate the data. Notably, you can compute the covariance matrix or a variant, which would consider alternative time-series similarity measures based on [dynamic time warping](https://en.wikipedia.org/wiki/Dynamic_time_warping) (DTW). In DTW, changes that vary in speed, e.g., one stock's price following another stock's price with a small delay, can be accommodated.
# +
means = data.get_mean_vector()
print("Means:")
print(means)
rho = data.get_similarity_matrix()
print("A time-series similarity measure:")
print(rho)
plt.imshow(rho)
plt.show()
cov = data.get_covariance_matrix()
print("A covariance matrix:")
print(cov)
plt.imshow(cov)
plt.show()
# -
# If you wish, you can look into the underlying pseudo-random time-series using. Please note that the private class members (starting with underscore) may change in future releases of Qiskit.
# +
print("The underlying evolution of stock prices:")
for (cnt, s) in enumerate(data._tickers):
plt.plot(data._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
for (cnt, s) in enumerate(data._tickers):
print(s)
print(data._data[cnt])
# -
# Clearly, you can adapt the number and names of tickers and the range of dates:
# + tags=["nbsphinx-thumbnail"]
data = RandomDataProvider(tickers=["CompanyA", "CompanyB", "CompanyC"],
start = datetime.datetime(2015, 1, 1),
end = datetime.datetime(2016, 1, 30),
seed = 1)
data.run()
for (cnt, s) in enumerate(data._tickers):
plt.plot(data._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
# -
# ### Access to closing-price time-series
#
# While the access to real-time data usually requires a payment, it is possible
# to access historical (adjusted) closing prices via Wikipedia and Quandl
# free of charge, following registration at:
# https://www.quandl.com/?modal=register
# In the code below, one needs to specify actual tickers of actual NASDAQ
# issues and the access token you obtain from Quandl; by running the code below, you agree to the Quandl terms and
# conditions, including a liability waiver.
# Notice that at least two tickers are required for the computation
# of covariance and time-series matrices, but hundreds of tickers may go
# beyond the fair usage limits of Quandl.
# +
stocks = ["GOOG", "AAPL"]
token = "REPLACE-ME"
if token != "REPLACE-ME":
try:
wiki = WikipediaDataProvider(
token = token,
tickers = stocks,
start = datetime.datetime(2016,1,1),
end = datetime.datetime(2016,1,30))
wiki.run()
except QiskitFinanceError as ex:
print(ex)
print("Error retrieving data.")
# -
# Once the data are loaded, you can again compute the covariance matrix or its DTW variants.
if token != "REPLACE-ME":
if wiki._data:
if wiki._n <= 1:
print("Not enough wiki data to plot covariance or time-series similarity. Please use at least two tickers.")
else:
rho = wiki.get_similarity_matrix()
print("A time-series similarity measure:")
print(rho)
plt.imshow(rho)
plt.show()
cov = wiki.get_covariance_matrix()
print("A covariance matrix:")
print(cov)
plt.imshow(cov)
plt.show()
else:
print('No wiki data loaded.')
# If you wish, you can look into the underlying time-series using:
if token != "REPLACE-ME":
if wiki._data:
print("The underlying evolution of stock prices:")
for (cnt, s) in enumerate(stocks):
plt.plot(wiki._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
for (cnt, s) in enumerate(stocks):
print(s)
print(wiki._data[cnt])
else:
print('No wiki data loaded.')
# ### [Optional] Setup token to access recent, fine-grained time-series
#
# If you would like to download professional data, you will have to set-up a token with one of the major providers. Let us now illustrate the data with NASDAQ Data on Demand, which can supply bid and ask prices in arbitrary resolution, as well as aggregates such as daily adjusted closing prices, for NASDAQ and NYSE issues.
#
# If you don't have NASDAQ Data on Demand license, you can contact NASDAQ (cf. https://business.nasdaq.com/intel/GIS/Nasdaq-Data-on-Demand.html) to obtain a trial or paid license.
#
# If and when you have access to NASDAQ Data on Demand using your own token, you should replace REPLACE-ME below with the token.
# To assure the security of the connection, you should also have your own means of validating NASDAQ's certificates. The DataOnDemandProvider constructor has an optional argument `verify`, which can be `None` or a string or a boolean. If it is `None`, certify certificates will be used (default). If verify is a string, it should be pointing to a certificate for the HTTPS connection to NASDAQ (dataondemand.nasdaq.com), either in the form of a CA_BUNDLE file or a directory wherein to look.
#
token = "REPLACE-ME"
if token != "REPLACE-ME":
try:
nasdaq = DataOnDemandProvider(token = token,
tickers = ["GOOG", "AAPL"],
start = datetime.datetime(2016,1,1),
end = datetime.datetime(2016,1,2))
nasdaq.run()
for (cnt, s) in enumerate(nasdaq._tickers):
plt.plot(nasdaq._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
except QiskitFinanceError as ex:
print(ex)
print("Error retrieving data.")
# Another major vendor of stock market data is Exchange Data International (EDI), whose API can be used to query over 100 emerging and frontier markets that are Africa, Asia, Far East, Latin America and Middle East, as well as the more established ones. See:
# https://www.exchange-data.com/pricing-data/adjusted-prices.php#exchange-coverage
# for an overview of the coverage.
#
# The access again requires a valid access token to replace REPLACE-ME below. The token can be obtained on a trial or paid-for basis at:
# https://www.quandl.com/
token = "REPLACE-ME"
if token != "REPLACE-ME":
try:
lse = ExchangeDataProvider(token = token,
tickers = ["AEO", "ABBY", "ADIG", "ABF",
"AEP", "AAL", "AGK", "AFN", "AAS", "AEFS"],
stockmarket = StockMarket.LONDON,
start=datetime.datetime(2018, 1, 1),
end=datetime.datetime(2018, 12, 31))
lse.run()
for (cnt, s) in enumerate(lse._tickers):
plt.plot(lse._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
except QiskitFinanceError as ex:
print(ex)
print("Error retrieving data.")
# One can also access Yahoo Finance Data, no token needed, from Yahoo! Finance.
try:
data = YahooDataProvider(
tickers = ["AEO", "ABBY", "AEP", "AAL", "AFN"],
start=datetime.datetime(2018, 1, 1),
end=datetime.datetime(2018, 12, 31))
data.run()
for (cnt, s) in enumerate(data._tickers):
plt.plot(data._data[cnt], label=s)
plt.legend()
plt.xticks(rotation=90)
plt.show()
except QiskitFinanceError as ex:
data = None
print(ex)
# For the actual use of the data, please see the <a href="1_portfolio_optimization.ipynb">portfolio_optimization</a> or <a href="2_portfolio_diversification.ipynb">portfolio_diversification</a> notebooks.
import qiskit.tools.jupyter
# %qiskit_version_table
# %qiskit_copyright
| docs/tutorials/11_time_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 4 Class Exercises: Pandas Part 2
# With these class exercises we learn a few new things. When new knowledge is introduced you'll see the icon shown on the right:
# <span style="float:right; margin-left:10px; clear:both;"></span>
# ## Get Started
# Import the Numpy and Pandas packages
import numpy as np
import pandas as pd
# ## Exercise 1: Review of Pandas Part 1
# ### Task 1: Explore the data
# Import the data from the [Lectures in Quantiatives Economics](https://github.com/QuantEcon/lecture-source-py) regarding minimum wages in countries around the world in US Dollars. You can view the data [here](https://github.com/QuantEcon/lecture-source-py/blob/master/source/_static/lecture_specific/pandas_panel/realwage.csv) and you can access the data file here: https://raw.githubusercontent.com/QuantEcon/lecture-source-py/master/source/_static/lecture_specific/pandas_panel/realwage.csv. Then perform the following
#
# Import the data into a variable named `minwages` and print the first 5 lines of data to explore what is there.
minwages = pd.read_csv('https://raw.githubusercontent.com/QuantEcon/lecture-source-py/master/source/_static/lecture_specific/pandas_panel/realwage.csv',)
minwages.head(5)
# Find the shape of the data.
minwages.shape
# List the column names.
minwages.columns
# Identify the data types. Do they match what you would expect?
minwages.dtypes
# Identify columns with missing values.
minwages.isna().sum()
# Identify if there are duplicated entires.
minwages.duplicated().sum()
# How many unique values per row are there. Do these look reasonable for the data type and what you know about what is stored in the column?
minwages.nunique()
# ### Task 2: Explore More
#
# Retrieve descriptive statistics for the data.
minwages.describe()
# Identify all of the countries listed in the data.
minwages['Country'].unique()
# Convert the time column to a datetime object.
minwages['Time'] = pd.to_datetime(minwages['Time'])
minwages.dtypes
# List the time points that were used for data collection. How many years of data collection were there? What time of year were the data collected?
minwages['Time'].unique()
# Because we only have one data point collected per year per country, simplify this by adding a new column with just the year. Print the first 5 rows to confirm the column was added.
minwages['Year'] = minwages['Time'].dt.year
minwages.head()
minwages['Year'].unique()
# There are two pay periods. Retrieve them in a list of just the two strings
minwages['Pay period'].unique()
minwages['Series'].unique()
# ### Task 3: Clean the data
# We have no duplicates in this data so we do not need to consider removing those, but we do have missing values in the `value` column. Lets remove those. Check the dimensions afterwards to make sure they rows with missing values are gone.
minwages.dropna(inplace=True)
minwages.shape
# If your dataframe has an "Unnamed: 0" column remove it, as it's not needed. Note: in the `pd.read_csv()` function you can use the `index_col` argument to set the column in the file that provides the index and that would prevent the "Unnamed: 0" column with this dataset.
# +
#minwages.drop(['Unnamed: 0'], axis=1, inplace=True)
# -
# ### Task 4: Indexing
# Use boolean indexing to retrieve the rows of annual salary in United States
minwages[(minwages['Country'] == 'United States') &
(minwages['Pay period'] == 'Annual')]
# Do we have enough data to calculate descriptive statistics for annual salary in the United States in 2016?
# From here on out, let's only explore the rows that have a "series" value of "In 2015 constant prices at 2015 USD exchange rates"
minwages2 = minwages[minwages['Series'] == 'In 2015 constant prices at 2015 USD exchange rates']
minwages2.shape
# Use `loc` to calculate descriptive statistics for the hourly salary in the United States and then again separately for Ireland. Hint: you will have to set row indexes. Hint: you should reset the index before using `loc`
# Now do the same for Annual salary
# ## Exercise 2: Occurances
# First, reset the indexes back to numeric values. Print the first 10 lines to confirm.
# Get the count of how many rows there are per year?
# ## Exercise 3: Grouping
# ### Task 1: Aggregation
# Calculate the average salary for each country across all years.
groups = minwages2.groupby(['Country', 'Pay period'])
groups.mean().head()
# Calculate the average salary and hourly wage for each country across all years. Save the resulting dataframe containing the means into a new variable named `mwmean`.
# <span style="float:right; margin-left:10px; clear:both;"></span>
#
# Above we saw how to aggregate using built-in functions of the `DataFrameGroupBy` object. For eaxmple we called the `mean` function directly. These handly functions help with writing succint code. However, you can also use the `aggregate` function to do more! You can learn more on the [aggregate description page](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.aggregate.html)
#
# With `aggregate` we can perform operations across rows and columns, and we can perform more than one operation at a time. Explore the online documentation for the function and see how you would calculate the mean, min, and max for each country and pay period type, as well as the total number of records per country and pay period:
#
groups.aggregate(['mean', 'std', 'count'])
# Also you can use the aggregate on a single column of the grouped object. For example:
#
# ```python
# mwgroup = minwages[['Country', 'Pay period', 'value']].groupby(['Country', 'Pay period'])
# mwgroup['value'].aggregate(['mean'])
#
# ```
# Redo the aggregate function in the previous cell but this time apply it to a single column.
# ### Task 2: Slicing/Indexing
# <span style="float:right; margin-left:10px; clear:both;"></span>
#
# In the following code the resulting dataframe should contain only one data column: the mean values. It does, however, have two levels of indexes: Country and Pay period. For example:
#
# ```python
# mwgroup = minwages[['Country', 'Pay period', 'value']].groupby(['Country', 'Pay period'])
# mwmean = mwgroup.mean()
# mwmean
# ```
#
# Try it out:
mwgroup = minwages2[['Country', 'Pay period', 'value']].groupby(['Country', 'Pay period'])
mwmean = mwgroup.mean()
mwmean
# Notice in the output above there are two levels of indexes. This is called MultiIndexing. In reality, there is only one data column and two index levels. So, you can do this:
#
# ```python
# mwmean['value']
# ```
#
# But you can't do this:
#
# ```python
# mwmean['Pay period']
# ```
#
# Why not? Try it:
#
mwmean['value']
mwmean['Pay period']
# The reason we cannot exeucte `mwmean['Pay period']` is because `Pay period` is not a data column. It's an index. Let's learn how to use MultiIndexes to retrieve data. You can learn more about it on the [MultiIndex/advanced indexing page](https://pandas.pydata.org/pandas-docs/stable/user_guide/advanced.html#advanced-indexing-with-hierarchical-index)
#
# First, let's take a look at the indexes using the `index` attribute.
#
# ```python
# mwmean.index
# ```
#
# Try it:
mwmean.index
# Notice that each index is actually a tuple with two levels. The first is the country names and the second is the pay period. Remember, we can use the `loc` function, to slice a dataframe using indexes. We can do so with a MultiIndexed dataframe as well. For example, to extract all elements with they index named 'Australia':
#
# ```python
# mwmean.loc[('Australia')]
# ```
#
# Try it yourself:
# You can specify both indexes to pull out a single row. For example, to find the average hourly salary in Australia:
#
# ```python
# mwmean.loc[('Australia','Hourly')]
# ```
# Try it yourself:
mwmean.loc[('Australia','Hourly')]
# Suppose you wanted to retrieve all of the mean "Hourly" wages. For MultiIndexes, there are multiple ways to slice it, some are not entirely intuitive or flexible enough. Perhaps the easiest is to use the `pd.IndexSlice` object. It allows you to specify an index format that is intuitive to the way you've already learned to slice. For example:
#
# ```python
# idx = pd.IndexSlice
# mwmean.loc[idx[:,'Hourly'],:]
# ```
#
# In the code above the `idx[:, 'Hourly']` portion is used in the "row" indexor position of the `loc` function. It indicates that we want all possible first-level indexes (specified with the `:`) and we want second-level indexes to be restricted to "Hourly".
# Try it out yourself:
idx = pd.IndexSlice
rows = idx[:,'Hourly']
mwmean.loc[rows,:]
# Using what you've learned above about slicing the MultiIndexed dataframe, find out which country has had the highest average annual salary.
# You can move the indexes into the dataframe and reset the index to a traditional single-level numeric index by reseting the indexes:
# ```python
# mwmean.reset_index()
# ```
#
# Try it yourself:
mwmean2 = mwmean.reset_index()
mwmean2[mwmean2['Pay period'] == 'Hourly']
# ### Task 3: Filtering the original data.
# <span style="float:right; margin-left:10px; clear:both;"></span>
#
# Another way we might want to filter is to find records in the dataset that, after grouping meets some criteria. For example, what if we wanted to find the records for all countries with the average annual salary was greater than $35K?
#
# To do this, we can use the `filter` function of the `DataFrameGroupBy` object. The filter function must take a function as an argument (this is new and may seem weird).
#
# ```python
# annualwages = minwages[minwages['Pay period'] == 'Annual']
# annualwages.groupby(['Country']).filter(
# lambda x : x['value'].mean() > 22000
# )
# ```
# Try it:
annualwages = minwages2[minwages2['Pay period'] == 'Annual']
annualwages.groupby(['Country']).filter(
lambda x : x['value'].mean() > 22000
)
# ### Task 4: Reset the index
# If you do not want to use MultiIndexes and you prefer to return any Multiindex dataset back to a traditional 1-level index dataframe you can use the`reset_index` function.
#
# Try it out on the `mwmean` dataframe:
# ## Exercise 4: Task 6d from the practice notebook
# Load the iris dataset.
#
# In the Iris dataset:
# + Create a new column with the label "region" in the iris data frame. This column will indicates geographic regions of the US where measurments were taken. Values should include: 'Southeast', 'Northeast', 'Midwest', 'Southwest', 'Northwest'. Use these randomly.
# + Use `groupby` to get a new data frame of means for each species in each region.
# + Add a `dev_stage` column by randomly selecting from the values "early" and "late".
# + Use `groupby` to get a new data frame of means for each species, in each region and each development stage.
# + Use the `count` function (just like you used the `mean` function) to identify how many rows in the table belong to each combination of species + region + developmental stage.
# ## Exercise 5: Kaggle Titanic Dataset
# A dataset of Titanic passengers and their fates is provided by the online machine learning competition server [Kaggle](https://www.kaggle.com/). See the [Titanic project](https://www.kaggle.com/c/titanic) page for more details.
#
# Let's practice all we have learned thus far to explore and perhaps clean this dataset. You have been provided with the dataset named `Titanic_train.csv`.
#
# ### Task 1: Explore the data
# First import the data and print the first 10 lines.
# Find the shape of the data.
# List the column names.
# Identify the data types. Do they match what you would expect?
# Identify columns with missing values.
# Identify if there are duplicated entires.
# How many unique values per row are there. Do these look reasonable for the data type and what you know about what is stored in the column?
# ### Task 2: Clean the data
# Do missing values need to be removed? If so, remove them.
# Do duplicates need to be removed? If so remove them.
# + active=""
#
# -
# ### Task 3: Find Interesting Facts
# Count the number of passengers that survied and died in each passenger class
# Were men or women more likely to survive?
# What was the average, min and max ticket prices per passenger class?
# Hint: look at the help page for the [agg](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.agg.html) function to help simplify this.
# Give descriptive statistics about the survival age.
| class_exercises/D05-Pandas_Part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.io)
# ## Tipografía.
#
#
# ## Selección del tipo de fuente con *font-family*.
#
# CSS permite seleccionar el tipo de letra que se utilizará en diversos elementos.
#
# La propiedad *font-family*.
#
# Permite elegir la tipografía a partir de las diversas familias de fuentes.
#
# Cabe señalar que es necesario que el navegador tenga instaladas las tipografías a las que se hace referencia.
#
# ### Fuentes genéricas:
#
# Los navegadores pueden ser configurados con fuentes tipográficas genéricas, las cuales son:
#
# * *Serif*.
# * *Sans-serif*.
# * *Monospace*.
#
# ## Importación de una fuente tipográfica externa.
#
#
# ### Importar un documento CSS con *@import*.
#
# La regla *@import* se utiliza para importar otras hojas de estilo.
#
# Para importar una hoja de estilo localizada en un archivo local se usa la siguiente sintaxis:
#
# ```
# @import '(ruta)';
# ```
#
# Para importar una hoja de estilo localizada en una URL se usa la siguiente sintaxis:
#
# ```
# @import url('(URL)');
# ```
#
# Las reglas *@import* siempre dene estar al pricipio del documento CSS.
#
# ### Importar una o varias fuentes tipográficas con *@font-face*.
#
# En caso de utilizar una o varias tipografía que no se encuentre instalada localmente, se utiliza *@font-face*.
#
# Los parámetros que utiliza *@font-face* para descargar e instalar la tipografía que se le indica es:
#
# * *font-family*, indicando el nombre que se le dará a la fuente tipográfica y que se usará como referencia.
# * *src* , incluye las posibles localizaciones de la fuente.
# * Se usa *local* en caso de que la fuente se encuentre localmente.
# * Se usa *url* en caso de que la fuente se encuentre en un sito externo.
# * *format* es el formato de archivo de la fuente tipográfica.
#
# Los formatos de archivo de fuentes tipográficas soportadas son:
# * *eot*
# * *woff*
# * *ttf*
# * *svg*
#
# Cabe hacer notar que algunos estos formatos tienen soporte limitado entre navegadores.
# **Ejemplo:**
# ```
# @font-face {
# font-family: MiPrimeraFuente;
# src: local('Racing Sans One'), local('RacingSansOne-Regular'), url(https://fonts.gstatic.com/s/racingsansone/v5/sykr-yRtm7EvTrXNxkv5jfKKyDCAJnDnapI.woff2) format('woff2');
# }
# ```
# ## Definición del tamaño de una fuente.
#
# Con este atributo es posible definir el tamaño de la tipografía de letra para los elementos seleccionados.
#
# Como ya se mencionó con anterioridad, es posible utilizar varias métricas para definir el tamaño una fuente tipográfica.
#
# * Pixeles (*px*).
# * Porcentajes con respecto al tamaño de la tipografía seleccionada.
# * Unidades *em*, que equivale al ancho de un caracter *m* de la tipografía seleccionada.
# ## Ajustes de texto.
#
# ### "Peso" de los caracteres.
#
# La propiedad *font-weight* define el peso de los caracteres con los siguientes valores:
#
# * *normal*
# * *bold*
#
# ### Estilo de los caracteres.
#
# La propiedad *font-style* define el estilo de los caracteres con los siguientes valores:
#
# * *normal*
# * *italic*
# * *oblique*
#
# ### Transformación de texto.
#
# La propiedad *text-transform* permite realizar algunos cambios a los caracteres de un texto:
#
# * *uppercase* convierte el texto a mayúsculas.
# * *lowecase* convierte el texto a minúsculas.
# * *capitalize* la primera letra de la palabra en mayúsculas.
#
# ### Decoración del texto.
#
# La propiedad *text-decoration* permite definir la decoración(líneas horizontales, colores, parpadeos, etc.) en un texto. Algunos valores de esta propiedad son:
#
# * *none*, el cual impide que existan líneas como las de subrayado.
# * *underline* la cual indica que el texto debe de estar subrayado.
# * *overline* la cual indica que el texto tendrá una raya encima del texto.
# * *line-through* la cual indica que el texto será tachado con una raya cruzando el texto.
# * *blink* la cual indica que el texto se verá parpadeante.
#
# ### Características de párrafo.
#
# #### Tamaños y espacios.
#
# * La propiedad *line-height* define altura de la línea.
# * La propiedad *letter-spacing* define el espaciado entre letras.
# * La propiedad *word-spacing* define el espaciado entre palabras.
#
# #### Alienación del texto.
#
# La propiedad *text-align* define la alineación horizontal del texto con los valores:
#
# * *left*
# * *right*
# * *center*
# * *justify*
#
# La propiedad *vertical-align* define la alineación vertical del texto con los valores:
#
# * *baseline*
# * *sub*
# * *super*
# * *top*
# * *text-top*
# * *middle*
# * *text-botton*
#
# La propiedad *text-indent* permite definir la indentación del párrafo.
# ## Propiedades de las listas.
#
# ### Listas no ordenadas.
#
# Para definir el tipo de "balazo" o marcador se utiliza *list-style-type* con los siguientes valores:
# * *none*
# * *disc*
# * *circle*
# * *square*
#
# ### Listas ordenadas.
#
# Para definir el tipo de numerador también se utiliza *list-style-type* con los siguientes valores:
#
# * *decimal*
# * *decimal-leading-zero*
# * *lower-alpha*
# * *upper-alpha*
# * *lower-roman*
# * *upper-roman*
#
# ### imágenes en los "balazos".
#
# Es posible utilizar imágenes que sustituyan a los caracteres clásicos de los balazos mediante la propiedad *list-style-image*, usánmdola del siguiente modo:
#
# ```
# list-style-image: url("(URL de la imagen)");
# ```
#
# ### Posición de los "balazos" en la línea de texto.
#
# Para definir si hay una indentación del párrafo con respecto al balazo de una lista se uitliza la propiedad *list-style-position* con los siguientes valores:
#
# * *outside* para indicar que el texto posterior a la primera línea del elemento *<li>* no puede ponerse debajo del balazo (se indentan).
# * *inside* para indicar que el texto posterior a la primera línea del elemento *<li>* puede ponerse debajo del balazo (no se indentan).
#
#
# ## Propiedades de las tablas.
#
# Todos los elementos de las tablas siguen el modelo de caja y las propiedades de texto anteriormente discutidas. Sin embargo poseen las siguientes propiedades adicionales:
#
# ### Mostrar bordes en celdas vacías.
#
# Para definir si las celdas vacías presentan bordes se utiliza la propiedad *empty-cells* con los siguientes valores:
#
# * *show* que permite que se vean los bordes.
# * *hide* que esconde los bordes.
# * *inherit* que hereda el comportamiento de la tabla superior si se trata de una tabla dentro de otra.
#
# ### Espaciamiento entre los bordes de una tabla.
#
# Siempre hay un pequeño espacio que separa los bordes de una celda con respecto a otra y para definir el espaciamiento o si debe haberlo se utilizan:
#
# * La propiedad *border-spacing*, la cual define el espaciado de los bordes.
# * La propiedad *border-colapse* la cual indica si existe o no la separación entre los bordes mediante los siguientes valores.
# * *collapse*, indicando que no hay espacios entre bordes.
# * *separate*, indicando que sí hay espacios entre bordes.
#
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2019.</p>
| 02_CSS/05_texto_listas_tablas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import random
import numpy as np
import cupy as cp
from unicode_info.database import generate_supported_consortium_feature_vectors_and_clusters_dict
# +
unicode_codepoint_vectors_dict = pickle.load(open('features_dict_file.pkl', 'rb'))
unicode_index_codepoint_map = list(unicode_codepoint_vectors_dict.keys())
del unicode_codepoint_vectors_dict
unknown_codepoints_simvec_map = pickle.load(open('simmap_pointninethreshold.pkl', 'rb'))
mat = np.stack([vec for vec in unknown_codepoints_simvec_map.values()])
unknown_indices_to_codepoint = [codepoint for codepoint in unknown_codepoints_simvec_map.keys()]
# +
# directly finding homoglyph codepoint sets will blow up memory
def filter_simvec_map_into_row_indices(sim_mat:np.ndarray, threshold:float):
print(sim_mat.shape)
arr = cp.asnumpy(cp.nonzero(cp.sum(cp.array(sim_mat) > threshold, axis=1)>1)[0])
#arr = np.nonzero(np.count_nonzero(sim_mat > threshold, axis=1)>1)[0]
print(arr.shape)
return set(arr)
# these aren't equivalence classes - might be overlapping sets
def randomly_select_indices_and_find_homoglyphs_sets(sim_mat:np.ndarray, simmat_row_indices:set, threshold:float, num_sets:int, full_index_codepoint_map:list):
pairs_for_index = {}
sample = random.sample(simmat_row_indices, k=num_sets)
for row_index in sample:
homoglyph_indices = np.nonzero(sim_mat[row_index] > threshold)[0]
if len(homoglyph_indices) > 1:
pairs_for_index[row_index] = {full_index_codepoint_map[index] for index in homoglyph_indices}
return pairs_for_index
# +
import matplotlib.pyplot as plt
from IPython.display import display
import os
from generate_datasets import try_draw_single_font
min_supported_fonts_dict = pickle.load(open('min_supported_fonts.pkl', 'rb'))
model_info_file = open(os.path.join('model_1', 'model_info.pkl'), 'rb')
model_info_dict = pickle.load(model_info_file)
img_size, font_size = model_info_dict['img_size'], model_info_dict['font_size']
empty_image = np.full((img_size, img_size), 255)
def display_random_findings(threshold, num_set):
str_to_u = lambda s: "U+" + hex(int(s))[2:] + ", " + s
row_indices = filter_simvec_map_into_row_indices(mat, threshold)
pairs_for_index = randomly_select_indices_and_find_homoglyphs_sets(mat, row_indices, threshold, num_set, unicode_index_codepoint_map)
display("Number of unknown homoglyphs: " + str(len(row_indices)))
display("Number of displayed sets: " + str(len(pairs_for_index))+"/" + str(num_set))
i = 1
for index, homoglyphs in pairs_for_index.items():
display("=========================")
display("=========================")
display("Homoglyphs for Codepoint " + str_to_u(unknown_indices_to_codepoint[index]) + ", size: " + str(len(homoglyphs)) + ", #: " + str(i))
i += 1
for homoglyph in homoglyphs:
image = try_draw_single_font(int(homoglyph), min_supported_fonts_dict[homoglyph], empty_image, img_size,
font_size, "./fonts", transform_img=False)
fig, ax = plt.subplots(figsize=(3,2))
ax.imshow(image)
display(fig)
display(str_to_u(homoglyph))
# -
# save findings to file
str_to_u = lambda s: "U+" + hex(int(s))[2:] + ", " + s
row_indices = filter_simvec_map_into_row_indices(mat, 0.93)
row_codepoints = [str_to_u(unknown_indices_to_codepoint[index]) for index in row_indices]
with open('new_predicted_homoglyphs.txt', mode='wt', encoding='utf-8') as myfile:
myfile.write('\n'.join(row_codepoints))
myfile.write('\n')
display_random_findings(0.93, 4)
str_to_u = lambda s: "U+" + hex(int(s))[2:] + ", " + s
vec = unknown_codepoints_simvec_map['66710']
codes = [unicode_index_codepoint_map[index] for index in np.nonzero(vec > 0.93)[0]]
for homoglyph in codes:
image = try_draw_single_font(int(homoglyph), min_supported_fonts_dict[homoglyph], empty_image, img_size,
font_size, "./fonts", transform_img=False)
fig, ax = plt.subplots(figsize=(3,2))
ax.imshow(image)
display(fig)
display(str_to_u(homoglyph))
# +
# debugging purpose
# i = 18816
# print(list(unknown_codepoints_simvec_map.items())[i])
# print([unicode_index_codepoint_map[ind] for ind in np.nonzero(mat[i] > 0.92)[0]])
| viz_unknown_homoglyphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sailon
# language: python
# name: sailon
# ---
# # Notebook Tutorial on an Empty SAIL-ON Detector
#
# In this example, it will show you an example of an empty detector which can be passed into the SAIL-ON-CLIENT protocol via the SAIL-ON Launcher.
# +
# %load_ext autoreload
# %autoreload 2
# Setup Logging to print to the notebook
import logging
import colorlog
handler = colorlog.StreamHandler()
handler.setFormatter(
colorlog.ColoredFormatter(
fmt='[%(cyan)s%(asctime)s%(reset)s][%(blue)s%(name)s%(reset)s]'
'[%(log_color)s%(levelname)s%(reset)s] - %(message)s',
log_colors={
'DEBUG': 'purple',
'INFO': 'green',
'WARNING': 'yellow',
'ERROR': 'red',
'CRITICAL': 'red,bg_white',
},
),
)
log = colorlog.getLogger('notebook')
log.setLevel(logging.DEBUG)
log.addHandler(handler)
# -
# ### Initalize the launcher with hydra to load the config
#
# Note: this config will define the save directory so make sure to edit any configuration values before running the code
# +
from hydra.experimental import initialize, compose
from omegaconf import OmegaConf
import yaml
# Initialize Hydra to get the default config. Overrides of `problem=mock_ond_config` will be selecting
# the 'mock_ond_config' from the folder '<sailon_tinker_launcher_root>/configs/problem/mock_ond_config.yaml'
with initialize(config_path="../configs"):
cfg = compose(config_name="h_config", overrides=['problem=mock_ond_config'])
print(cfg)
# You can also update the parameters here but do it before you convert to a dict (next step).
# Here we update the config with a new novelty class detector name (to match the class in the next cell).
cfg.problem.novelty_detector_class = 'MyDetector'
# Turn the omegaconfig to a dict object and print it nicely
config = OmegaConf.to_container(cfg['problem'])
print(yaml.dump(config, indent=4))
# -
# ### Create Python Class to Run for the experiment.
#
# This is using an empty detector which just says it's name (similar to a pokemon that can't fight). This is actually better since it also said the step it's running.
# +
from sailon_tinker_launcher.deprecated_tinker.basealgorithm import BaseAlgorithm
from evm_based_novelty_detector.condda_12 import condda_without_redlight
from typing import Dict, Any, Tuple
class MyDetector(BaseAlgorithm):
"""Mock Detector for testing image classification protocols."""
def __init__(self, toolset: Dict) -> None:
"""
Detector constructor.
Args:
toolset (dict): Dictionary containing parameters for the constructor
"""
BaseAlgorithm.__init__(self, toolset)
self.step_dict: Dict[str, Callable] = {
"Initialize": self._initialize,
"FeatureExtraction": self._feature_extraction,
"WorldDetection": self._world_detection,
"NoveltyClassification": self._novelty_classification,
"NoveltyAdaption": self._novelty_adaption,
"NoveltyCharacterization": self._novelty_characterization,
}
def execute(self, toolset: Dict, step_descriptor: str) -> Any:
"""
Execute method used by the protocol to run different steps associated with the algorithm.
Args:
toolset (dict): Dictionary containing parameters for different steps
step_descriptor (str): Name of the step
"""
log.info(f"Executing Step: {step_descriptor}")
return self.step_dict[step_descriptor](toolset)
def _initialize(self, toolset: Dict) -> None:
"""
Algorithm Initialization.
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
None
"""
log.info('Initialize')
def _feature_extraction(
self, toolset: Dict
) -> Tuple[Dict[str, Any], Dict[str, Any]]:
"""
Feature extraction step for the algorithm.
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
Tuple of dictionary
"""
self.dataset = toolset["dataset"]
log.info('Feature Extraction')
return {}, {}
def _world_detection(self, toolset: str) -> str:
"""
Detect change in world ( Novelty has been introduced ).
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
path to csv file containing the results for change in world
"""
log.info('World Detection')
return self.dataset
def _novelty_classification(self, toolset: str) -> str:
"""
Classify data provided in known classes and unknown class.
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
path to csv file containing the results for novelty classification step
"""
log.info('Novelty Classification')
return self.dataset
def _novelty_adaption(self, toolset: str) -> None:
"""
Update models based on novelty classification and characterization.
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
None
"""
log.info('Novelty Adaption')
def _novelty_characterization(self, toolset: str) -> str:
"""
Characterize novelty by clustering different novel samples.
Args:
toolset (dict): Dictionary containing parameters for different steps
Return:
path to csv file containing the results for novelty characterization step
"""
log.info('Novelty Characterization')
return self.dataset
# -
# ### launch the detector in the sailon system
#
# The extra plugin parameter takes a dictionary which allows you to add your detector to the list of detectors. The name of the detector in the config needs to match the key in the dictionary. See config above for more details.
# +
from sailon_tinker_launcher.main import LaunchSailonProtocol
launch_protocol = LaunchSailonProtocol()
launch_protocol.run_protocol(config, extra_plugins={'MyDetector': MyDetector})
| scripts/create_empty_sail_on_protocol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# data_prep.py
import pandas as pd
from pathlib import Path
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error, mean_absolute_error
import numpy as np
from joblib import dump
import seaborn as sns
import matplotlib.pyplot as plt
from tensorflow.keras.layers import Dense, LSTM
import tensorflow as tf
# -
tf.__version__
data = Path.cwd().resolve().parents[0] / "data"
df = pd.DataFrame()
FILES = list(data.glob("*.csv"))
print(f'total files found: {len(FILES)}')
for file in FILES:
df_ = pd.read_csv(str(file), parse_dates=["Date"])
df_['Date'] = pd.to_datetime(df_["Date"])
df = df.append(df_)
df.sort_values(by="Date", inplace=True)
df.drop_duplicates(inplace=True)
df.head()
plt.figure(figsize=(15, 8))
sns.scatterplot(x=df.Date, y=df.Close, label="Close", markers="o")
plt.legend()
plt.show()
df.shape
df.sort_values(by="Date")
for lag in range(9):
df["lag"+str(lag+1)] = df["Close"].shift(lag+1)
df.iloc[:, 7:].dropna(inplace=True)
df.dropna(inplace=True)
df.head(10)
df.Date.min(), df.Date.max()
feat = df.iloc[:, 7:].values
mn = MinMaxScaler(feature_range=(0, 1))
features = mn.fit_transform(feat)
target = df.Close.values
features.shape, target.shape
train_len = int(features.shape[0] * 0.95)
train_feat = features[:train_len]
train_target = df.Close.values[:train_len]
val_feat = features[train_len:]
val_target = df.Close.values[train_len:]
train_feat.shape, train_target.shape, val_feat.shape, val_target.shape
train_feat = train_feat.reshape((train_feat.shape[0], 1, train_feat.shape[1]))
train_target = train_target.reshape((-1, 1))
val_feat = val_feat.reshape((val_feat.shape[0], 1, val_feat.shape[1]))
val_target = val_target.reshape((-1, 1))
train_feat.shape, train_target.shape, val_feat.shape, val_target.shape
model = tf.keras.Sequential()
model.add(LSTM(50 , activation="relu", input_shape=(1,train_feat.shape[2]), return_sequences=True))
model.add(LSTM(50, activation="relu", return_sequences=True))
model.add(LSTM(50, activation="relu", return_sequences=True))
model.add(LSTM(50, activation="relu", return_sequences=True))
model.add(LSTM(50, activation="relu"))
model.add(Dense(1))
model.compile(optimizer="adam" , loss="mae", metrics=["mae"])
model.summary()
cb = tf.keras.callbacks.EarlyStopping(
monitor='loss', min_delta=0, patience=5, verbose=0, mode='auto',
baseline=None, restore_best_weights=False
)
history = model.fit(x=train_feat,
y=train_target,
batch_size=32,
validation_data=(val_feat, val_target),
epochs=100,
# callbacks=[cb]
)
plt.figure(figsize=(15,8))
sns.scatterplot(y=history.history["loss"], x=range(len(history.history["loss"])), label="train")
sns.scatterplot(y=history.history["val_loss"], x=range(len(history.history["loss"])), label="val")
plt.legend()
plt.show()
model.save("model.h5")
predictions = model.predict(val_feat)
date = pd.date_range(end="2020-08-31", freq="D", periods=predictions.shape[0])
plt.figure(figsize=(15,8))
sns.scatterplot(y=predictions.reshape((-1)).tolist(), x=date , label="predictions", color="red", markers="o")
sns.scatterplot(y=train_target.reshape((-1)).tolist()[:len(date)], x=date, label="actual", color="blue")
plt.legend()
plt.grid()
plt.show()
mean_squared_error(y_pred=predictions.reshape((-1)).tolist(), y_true=val_target)
mean_absolute_error(y_pred=predictions.reshape((-1)).tolist(), y_true=val_target)
# ## 2nd Try
model = tf.keras.Sequential()
model.add(LSTM(50 , activation="relu", input_shape=(1,train_feat.shape[2])))
model.add(LSTM(50, activation="relu"))
model.add(LSTM(50, activation="relu"))
model.add(LSTM(50, activation="relu"))
model.add(LSTM(50, activation="relu"))
model.add(Dense(1))
model.compile(optimizer="adam" , loss="mae", metrics=["mae"])
model.summary()
| notebook/dev.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from os import path
from enmspring.graphs import GraphAgent
from enmspring import PDB
from enmspring import atom
import MDAnalysis as mda
rootfolder = '/home/yizaochen/codes/dna_rna/fluctmatch_sequence'
enmroot = '/home/yizaochen/codes/dna_rna/enmspring'
# ### Part 1: Initialize
host = 'a_tract_21mer'
g_agent = GraphAgent(host, rootfolder)
# ### Part 2: Show crd in VMD
g_agent.vmd_show_crd()
# ### Part 3: Eigen-decomposition
g_agent.build_node_list_base()
print(f"Thare are {g_agent.n_node} nodes.")
g_agent.build_adjacency_from_df_st()
g_agent.build_degree_from_adjacency()
g_agent.build_laplacian_by_adjacency_degree()
g_agent.eigen_decompose()
# ### Part 4: Select Eigenvector
sele_id = 3
scale_factor = 10.
eigv = g_agent.get_eigenvector_by_id(sele_id)
eigv_scale = scale_factor * eigv
eigv_scale.max()
eigv_scale.min()
# ### Part 5: Convert crd to pdb
u = mda.Universe(g_agent.npt4_crd, g_agent.npt4_crd)
npt4_pdb = path.join(g_agent.input_folder, 'bdna+bdna.nohydrogen.pdb')
with mda.Writer(npt4_pdb, bonds=None, n_atoms=u.atoms.n_atoms) as pdbwriter:
pdbwriter.write(u.atoms)
print(f'vim {npt4_pdb}')
# ### Part 6: Read in to pdbreader
reader = PDB.PDBReader(npt4_pdb, skip_header=9, skip_footer=2, withfragid=True)
# +
#for atg in reader.atomgroup:
# print(atg.tempFactor)
# -
# ### Part 7: Add two dummy atoms to keep color scale in [-1,1]
minimum = eigv_scale.min()
maximum = -eigv_scale.min()
minimum = -eigv_scale.max()
maximum = eigv_scale.max()
# get serial and resid of the last atom
serial = reader.atomgroup[-1].serial + 1
resid = reader.atomgroup[-1].resid + 1
dummy1_data = ['ATOM', serial, 'S1', 'DUM', resid, 0.0, 0.0, 0.0, 0.0, minimum]
dummy2_data = ['ATOM', serial+1, 'S2', 'DUM', resid+1, 0.0, 0.0, 0.0, 0.0, maximum]
reader.atomgroup.append(atom.Atom(dummy1_data, False))
reader.atomgroup.append(atom.Atom(dummy2_data, False))
# ### Part 8: Get nodes idx map to pdb
for cgname, eigv_value in zip(g_agent.node_list, eigv_scale):
atomid = g_agent.atomid_map[cgname]
reader.atomgroup[atomid-1].set_tempFactor(eigv_value)
# ### Part 9: Output PDB for eigenvector
f_out = path.join(g_agent.input_folder, f'eigv_{sele_id}.pdb')
writer = PDB.PDBWriter(f_out, reader.atomgroup)
writer.write_pdb()
print(f'vim {f_out}')
# ### Part 10: Show PDB in vmd
g_agent.vmd_show_crd()
print(f'mol new {f_out} type pdb')
| notebooks/draw_graph_basestack_eigenvector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="mULkvgCWZyyO"
# !rm -r /content/sample_data
# + id="z8sa4Bn9bP46"
# %mkdir an_data
# + id="-yRPmUzkbQ4r" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4beb8f55-8b77-44f4-8e8c-d3b154ecc172"
# %cd /content/an_data
# + id="LYy0E41KbSUr" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 1000} outputId="3542de2e-d139-4995-cf22-0c22949ff1c8"
from google.colab import files #Upload the annotated text files from local drive
files.upload()
# + id="K9Lx4mOFbNOA" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="3102917c-66db-4d9d-ccf6-e2f5416c6b9f"
import nltk, re, string, unicodedata, inflect, glob, os
import re
from nltk.corpus.reader.plaintext import PlaintextCorpusReader
import numpy as np
import pandas as pd #for data manipulation and analysis
from nltk.corpus import stopwords #Stopwords corpus
from nltk.stem import PorterStemmer
from nltk.tokenize import RegexpTokenizer
from string import punctuation
from nltk import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import LancasterStemmer, WordNetLemmatizer
from nltk.tokenize import TreebankWordTokenizer
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('wordnet')
nltk.download('averaged_perceptron_tagger')
# + id="rAg3-xZJbaRG" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="8ff9d0d9-6acd-403f-d9d2-7271f75b4fd8"
for dirname, _, filenames in os.walk('/content/an_data'):
for filename in filenames:
print(os.path.join(dirname, filename))
# + id="Xf8S2tckbapf"
df = pd.concat([pd.read_csv(f, sep = '\t', header = None) for f in glob.glob('*.tsv')], ignore_index = True)#bind the tsv files together in a single file
# + id="1bYdH9rdaC5v"
df.columns = ['sentence','label']
# + id="2kEKloNXbjsB"
with open('manual_sent.txt', 'w') as f:
for text in df['sentence'].tolist():
f.write(text + '\n')
# + id="9xLZZmE4cCiS"
with open("/content/tost.txt", 'rb') as corpus:
text_full = corpus.readlines()
# + id="UqKTY-nkzMKn"
text_full
# + id="LPnwTkbOzMEY"
listA = []
with open('/content/tost.txt') as f:
for line in f:
words = [x.strip(string.punctuation) for x in line.split()]
listA.append(' '.join(w for w in words if w))
# + id="d-Z-46LB2d7-"
filtered_sentence = (" ").join(listA)
# + id="qUpGTheA2qhL"
text_for_spacy = word_tokenize(filtered_sentence)
# + id="TSaBIWuX3iel" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a0749510-22c0-4043-c836-ac30ba3be557"
type(text_for_spacy)
# + id="wMLQsBF62sJt"
for item in text_for_spacy:
item.replace("\\\\x", '')
# + id="jCvOrXK1X-x9"
import pickle
import spacy
# + id="RF-4XWV8X-Ht" colab={"base_uri": "https://localhost:8080/", "height": 683} outputId="fa9211a6-dc14-408e-e5d1-12868a3a03b3"
# !pip install scispacy spacy
# + id="Yr70puhTYAoL" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="7c283a96-ee9f-4764-bbf9-0844e62bb631"
# !pip install https://s3-us-west-2.amazonaws.com/ai2-s2-scispacy/releases/v0.2.3/en_ner_bc5cdr_md-0.2.3.tar.gz
# !pip install https://s3-us-west-2.amazonaws.com/ai2-s2-scispacy/releases/v0.2.3/en_ner_craft_md-0.2.3.tar.gz
# !pip install https://s3-us-west-2.amazonaws.com/ai2-s2-scispacy/releases/v0.2.3/en_ner_jnlpba_md-0.2.3.tar.gz
# !pip install https://s3-us-west-2.amazonaws.com/ai2-s2-scispacy/releases/v0.2.3/en_ner_bionlp13cg_md-0.2.3.tar.gz
# + id="Tx7RS9tAYHCn"
import scispacy
from spacy import displacy
from scispacy.abbreviation import AbbreviationDetector
from scispacy.umls_linking import UmlsEntityLinker
# + id="cBQPzPRaYIIy"
import en_ner_craft_md
import en_ner_jnlpba_md
import en_ner_bc5cdr_md
import en_ner_bionlp13cg_md
# + id="hWSPMlBjYKmR"
nlp = en_ner_craft_md.load()
# + id="mVpYHIBpYLDj"
doc = nlp(str(text_for_spacy))
# + id="4CXkHj5fYMVv" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="42638844-9a30-4566-ff0e-76b9f9814261"
empty_list = []
for entity in doc.ents:
empty_list.append(entity.text)
for entity in doc.ents:
print(entity.label_, ' | ', entity.text)
# + id="9zgo1aS8YNfW"
#len(set(empty_list))
# + id="AjM-h6-53v-7" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9120f253-f0fd-45f3-a91f-c9a851885b74"
empty_list
# + id="KdZJeI51YOiK" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4fb0e39d-c5a6-453a-dfcb-8a7c24bb1f43"
len(empty_list)
| model_sci/SciSpacy_in_Manual_Text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (bl_tf)
# language: python
# name: ml
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
def db(x):
""" Convert linear value to dB value """
return 10*np.log10(x)
import blimpy as bl
import sys
sys.path.insert(0, "/home/bryanb/setigen/")
import setigen as stg
# -
2**25
# +
r'''
Synthesize a Filterbank file.
Motivation: https://github.com/UCBerkeleySETI/turbo_seti/issues/206
'''
import time
from astropy import units as u
import setigen as stg
from blimpy import Waterfall
DIR = '/datax/scratch/bbrzycki/data/'
INFILE = '/datax/scratch/bbrzycki/data/tabby.1.fil'
MAX_DRIFT = 4
MIN_SNR = 25
arg_n_signals = 1
def gen_fil(arg_path, n_signals):
r''' Generate a Filterbank file '''
assert n_signals > 0 and n_signals < 5
# Define parameters.
# fchans = 134217728
fchans = 2**26
#fchans = 1342177
tchans = 5
df = -2.7939677238464355e-06 * u.Hz
dt = 18.253611008 * u.s
fch1 = 19626.464842353016 * u.MHz
noise_std = 0.05 # Gaussian standard deviation
sig_snr_1 = 100.0
sig_width_1 = 1.1 * u.Hz
drate_1 = 1.6 * u.Hz/u.s
f_start_1 = 0
sig_snr_2 = 200.0
sig_width_2 = 1.2 * u.Hz
drate_2 = 1.3 * u.Hz/u.s
f_start_2 = fchans * 0.1
sig_snr_3 = 300.0
sig_width_3 = 1.3 * u.Hz
drate_3 = 2.6 * u.Hz/u.s
f_start_3 = fchans * 0.2
sig_snr_4 = 400.0
sig_width_4 = 1.4 * u.Hz
drate_4 = 3.2 * u.Hz/u.s
f_start_4 = fchans * 0.3
# Generate the frame.
frame = stg.Frame(fchans=fchans,
tchans=tchans,
df=df,
dt=dt,
fch1=fch1)
# Add noise.
frame.add_noise(x_mean=0, x_std=noise_std, noise_type='gaussian')
# Add signal 1.
signal_intensity = frame.get_intensity(snr=sig_snr_1)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_1),
drift_rate=drate_1,
level=signal_intensity,
width=sig_width_1,
f_profile_type='gaussian')
# Add signal 2.
if n_signals > 1:
signal_intensity = frame.get_intensity(snr=sig_snr_2)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_2),
drift_rate=drate_2,
level=signal_intensity,
width=sig_width_2,
f_profile_type='gaussian')
# Add signal 3.
if n_signals > 2:
signal_intensity = frame.get_intensity(snr=sig_snr_3)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_3),
drift_rate=drate_3,
level=signal_intensity,
width=sig_width_3,
f_profile_type='gaussian')
# Add signal 4.
if n_signals > 3:
signal_intensity = frame.get_intensity(snr=sig_snr_4)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_4),
drift_rate=drate_4,
level=signal_intensity,
width=sig_width_4,
f_profile_type='gaussian')
# Save Filterbank file.
frame.save_fil(arg_path)
t1 = time.time()
gen_fil(INFILE, n_signals=1)
t2 = time.time()
print('gen_fil elapsed time = {:.1f}s'.format(t2 - t1))
wf = Waterfall(INFILE)
wf.info()
# -
# !ls /datax/scratch/bbrzycki/data/tabby.1.fil
wf = Waterfall(INFILE)
wf.info()
# +
fchans = 1024
tchans = 5
df = -2.7939677238464355e-06 * u.Hz
dt = 18.253611008 * u.s
fch1 = 19626.464842353016 * u.MHz
noise_std = 0.05 # Gaussian standard deviation
# Generate the frame.
frame = stg.Frame(fchans=fchans,
tchans=tchans,
df=df,
dt=dt,
fch1=fch1)
# -
frame.get_waterfall()
vars(frame.waterfall.container)
vars(frame.waterfall)
# +
wf = frame.get_waterfall()
chunk_dim = wf._get_chunk_dimensions()
blob_dim = wf._get_blob_dimensions(chunk_dim)
n_blobs = wf.container.calc_n_blobs(blob_dim)
chunk_dim, blob_dim, n_blobs
# -
frame.waterfall.container
frame.waterfall.container.calc_n_blobs(blob_dim)
blob_dim[frame.waterfall.container.time_axis]
frame.waterfall.container.selection_shape
updated_blob_dim=(int(frame.waterfall.container.selection_shape[frame.waterfall.container.time_axis] - blob_dim[frame.waterfall.container.time_axis]*1),
1,
int(blob_dim[frame.waterfall.container.freq_axis]))
updated_blob_dim
np.prod(updated_blob_dim)
frame.waterfall.container._d_type
frame.waterfall.container.filename
frame1 = stg.Frame('/home/bryanb/setigen/setigen/assets/sample.fil')
np.prod(frame1.shape)
frame.waterfall.container.MAX_DATA_ARRAY_SIZE
frame.waterfall.container._calc_selection_size()
5*2**26
# +
fchans = 2**26
tchans = 5
df = -2.7939677238464355 * u.Hz
dt = 18.253611008 * u.s
fch1 = 19626.464842353016 * u.MHz
noise_std = 0.05 # Gaussian standard deviation
# Generate the frame.
frame = stg.Frame(fchans=fchans,
tchans=tchans,
df=df,
dt=dt,
fch1=fch1)
frame.get_waterfall()
# -
frame.waterfall.container.MAX_DATA_ARRAY_SIZE, frame.waterfall.container._calc_selection_size()
frame.waterfall.container.selection_shape
5*2**26*32
frame.waterfall.container._n_bytes
frame.waterfall.container.t_stop - frame.waterfall.container.t_start
(frame.waterfall.container.f_stop - frame.waterfall.container.f_start) / abs(frame.waterfall.container.header['foff'])
frame.waterfall.container.header['foff']
frame.waterfall.container.f_stop, frame.waterfall.container.f_start
fchans*5*4
# +
fchans = 2**26
tchans = 5
df = -2.7939677238464355 * u.Hz
dt = 18.253611008 * u.s
fch1 = 19626.464842353016 * u.MHz
noise_std = 0.05 # Gaussian standard deviation
# Generate the frame.
frame = stg.Frame(fchans=fchans,
tchans=tchans,
df=df,
dt=dt,
fch1=fch1)
frame._update_waterfall()
frame.save_fil('example.fil')
# -
INFILE
import os
fn = 'test.txt'
os.path.isabs(fn)
os.path.abspath(fn)
# +
r'''
Synthesize a Filterbank file.
Motivation: https://github.com/UCBerkeleySETI/turbo_seti/issues/206
'''
import time
from astropy import units as u
import setigen as stg
from blimpy import Waterfall
DIR = '/datax/scratch/bbrzycki/data/'
INFILE = '/datax/scratch/bbrzycki/data/tabby.1.fil'
MAX_DRIFT = 4
MIN_SNR = 25
arg_n_signals = 1
def gen_fil(arg_path, n_signals):
r''' Generate a Filterbank file '''
assert n_signals > 0 and n_signals < 5
# Define parameters.
# fchans = 134217728
fchans = 2**27
#fchans = 1342177
tchans = 5
df = -2.7939677238464355e-06 * u.Hz
dt = 18.253611008 * u.s
fch1 = 19626.464842353016 * u.MHz
noise_std = 0.05 # Gaussian standard deviation
sig_snr_1 = 100.0
sig_width_1 = 1.1 * u.Hz
drate_1 = 1.6 * u.Hz/u.s
f_start_1 = 0
sig_snr_2 = 200.0
sig_width_2 = 1.2 * u.Hz
drate_2 = 1.3 * u.Hz/u.s
f_start_2 = fchans * 0.1
sig_snr_3 = 300.0
sig_width_3 = 1.3 * u.Hz
drate_3 = 2.6 * u.Hz/u.s
f_start_3 = fchans * 0.2
sig_snr_4 = 400.0
sig_width_4 = 1.4 * u.Hz
drate_4 = 3.2 * u.Hz/u.s
f_start_4 = fchans * 0.3
# Generate the frame.
frame = stg.Frame(fchans=fchans,
tchans=tchans,
df=df,
dt=dt,
fch1=fch1)
# Add noise.
frame.add_noise(x_mean=0, x_std=noise_std, noise_type='gaussian')
# Add signal 1.
signal_intensity = frame.get_intensity(snr=sig_snr_1)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_1),
drift_rate=drate_1,
level=signal_intensity,
width=sig_width_1,
f_profile_type='gaussian')
# Add signal 2.
if n_signals > 1:
signal_intensity = frame.get_intensity(snr=sig_snr_2)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_2),
drift_rate=drate_2,
level=signal_intensity,
width=sig_width_2,
f_profile_type='gaussian')
# Add signal 3.
if n_signals > 2:
signal_intensity = frame.get_intensity(snr=sig_snr_3)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_3),
drift_rate=drate_3,
level=signal_intensity,
width=sig_width_3,
f_profile_type='gaussian')
# Add signal 4.
if n_signals > 3:
signal_intensity = frame.get_intensity(snr=sig_snr_4)
frame.add_constant_signal(f_start=frame.get_frequency(f_start_4),
drift_rate=drate_4,
level=signal_intensity,
width=sig_width_4,
f_profile_type='gaussian')
# Save Filterbank file.
# frame.get_waterfall()
# frame.waterfall.container.MAX_DATA_ARRAY_SIZE = frame.waterfall.file_size_bytes
frame.save_fil(arg_path, max_load=8)
print(len(sigproc.generate_sigproc_header(frame.waterfall)))
print(frame.waterfall.container.idx_data)
print(frame.waterfall.file_size_bytes)
t1 = time.time()
gen_fil(INFILE, n_signals=1)
t2 = time.time()
print('gen_fil elapsed time = {:.1f}s'.format(t2 - t1))
wf = Waterfall(INFILE)
wf.info()
# -
frame.waterfall.file_size_bytes
2684354560.0/1e9
from blimpy.io import sigproc
sigproc.generate_sigproc_header(wf)
len(sigproc.generate_sigproc_header(wf))
wf.container.idx_data
len(sigproc.generate_sigproc_header(frame.waterfall))
frame.waterfall.container.idx_data
import psutil
stats = psutil.virtual_memory() # returns a named tuple
available = getattr(stats, 'available')
print(available)
available/1e9
| misc/investigate_issue_17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: xpython
# language: python
# name: xpython
# ---
# + [markdown] deletable=false editable=false
# Copyright 2020 <NAME> and made available under [CC BY-SA](https://creativecommons.org/licenses/by-sa/4.0) for text and [Apache-2.0](http://www.apache.org/licenses/LICENSE-2.0) for code.
#
# -
# # Random forests: Problem solving
#
# In this session, we'll use the `boston` dataset, which has been used to examine the relationship between clean air and house prices:
#
#
# | Variable | Type | Description |
# |:----|:-----|:----------|
# |crim | Ratio | per capita crime rate by town |
# |zn | Ratio | proportion of residential land zoned for lots over 25,000 sq. ft. |
# |indus | Ratio | proportion of non-retain business acres per town |
# |chas | Nominal (binary) | Charles River dummy variable (=1 if tract bounds river, =0 otherwise) |
# |nox | Ratio | nitrogen oxides concentration (parts per 10 million) |
# |rm | Ratio | average number of rooms per dwelling |
# |age | Ratio | proportion of owner-occupied units built prior to 1940 |
# |dis | Ratio | weighted mean of distances to fie Boston employment centers |
# |rad | Ordinal | index of accessibility to radial highways |
# |tax | Ratio | full-value proporty tax rate per \$10,000 |
# |ptratio | Ratio | pupil-teacher ratio by town |
# |lstat | Ratio | percent lower status of population (defined as non-high school graduate, manual labor) |
# |medv | Ratio | median value of owner-occupied homes in $1000s |
#
# <div style="text-align:center;font-size: smaller">
# <b>Source:</b> This dataset was taken from the StatLib library which is maintained at Carnegie Mellon University.
# </div>
# <br>
#
# As before, we'll try to predict `medv` using the rest of the variables.
#
# **Because `medv` is a ratio variable, we will do random forest regression trees not decision trees.**
#
# Additionally, we will compare the performance of three models on this problem:
#
# - Regression trees
# - Bagged regression trees
# - Random forest regression trees
# ## Load data
#
# Import `pandas` to load a dataframe.
# Load the dataframe.
# ## Explore data
#
# Some of these steps we've done before with these data, so we'll skip the normal interpretation steps on those parts.
#
# Describe the data.
# Make a correlation heatmap.
#
# First import `plotly.express`.
# Create a correlation matrix.
# And show the correlation heatmap with row/column labels.
# Because these variables are highly correlated (an numeric), a scatterplot matrix would make a lot of sense.
#
# Use `plotly` to make a `scatter_matrix` of the dataframe.
# If you have a hard time reading the labels, you can give it something like `width=1000` and `height=1000` to make it bigger.
# -----------
# **QUESTION:**
#
# Remembering that a perfect correlation is a line, and no correlation is a uniform random scattering of datapoints, what would you say about the pattern of these scatterplots overall?
# Of the scatterplots in the last row (i.e. correlated with `medv`) in particular?
# **ANSWER: (click here to edit)**
#
#
# <hr>
# Ultimately we want to predict median house value (`medv`), so make a histogram of that.
# ------------------
# **QUESTION:**
#
# Do you think we need to transform `medv` to make it more normal? Why or why not?
# **ANSWER: (click here to edit)**
#
#
# <hr>
# ## Prepare train/test sets
#
# If we were just using bagging or random forests, we could use OOB performance instead of splitting the data into training and testing sets.
#
# However, splitting is necessary if we want to compare to regression trees.
#
# Start by dropping the response variable, `medv` to make a new dataframe called `X`.
#
# Save a dataframe with just `medv` in `Y`.
# Import `sklearn.model_selection` to split the data into train/test sets.
# And do the actual split.
# ## Fit models
#
# Fit three models in turn:
#
# - Regression tree
# - Bagged regression tree
# - Random forest regression tree
#
# Import the `sklearn.tree` and `sklearn.ensemble` libraries.
# ### Regression tree
#
# Create the regression tree model.
# Go ahead and create it with a freestyle `random_state=1` so we all get the same results.
# Fit the regression tree model and get predictions.
# ### Bagged regression tree
#
# Next create the bagged regression tree model by using `BaggingRegressor`.
# Just as `BaggingClassifier` uses a decision tree by default, `BaggingRegressor` uses a regression tree by default.
# Use the same parameters as the random forest notebook (e.g. 100 trees, etc).
# Interestingly, for this model, `sklearn` requires us to use `ravel` on `Y` when fitting the model, so import `numpy`.
# Fit the bagged regression tree using `ravel` on `Y` and get predictions.
# ### Random forest regression tree
#
# Next create the random forest regression tree model by using `RandomForestRegressor`, which also uses a regression tree by default.
# Use the same parameters as before.
# Fit the random forest regression tree using `ravel` on `Y` and get predictions.
# ## Evaluate the models
# ### Regression tree
#
# - Get the $r^2$ on the *training* set
#
# - Get the $r^2$ on the *testing* set
# ### Bagged regression tree
#
# - Get the $r^2$ on the *training* set
#
# - Get the $r^2$ on the *testing* set
# ### Random forest regression tree
#
# - Get the $r^2$ on the *training* set
#
# - Get the $r^2$ on the *testing* set
# ------------------
# **QUESTION:**
#
# Compare the three models with respect to their *training data performance*. Which is better?
# Now compare the three models with respect to their *testing data performance*. Which is better?
# What do these differences tell you?
# **ANSWER: (click here to edit)**
#
#
# <hr>
# ## Feature importance
#
# Calculate the feature importance for the three models and plot it as a `plotly` bar chart.
#
# To get the column names for `x=` in the bar chart, you can use `from X get columns` as a shortcut.
# ### Regression tree
# ### Bagged regression tree
#
# For some reason, `sklearn` does not implement `feature_importances_` for `BaggingRegressor`, so use the following freestyle for the `y=` part of your plot:
#
# `np.mean([ tree.feature_importances_ for tree in baggedRegressionTree.estimators_ ], axis=0)`
#
# **You will need to change `baggedRegressionTree` to whatever you called this model above.**
# ### Random forest regression tree
#
# This uses `feature_importances_`, so you can make the plot exactly like you would for the regression tree model.
# -----------
# **QUESTION:**
#
# Look carefully at the three feature importance plots, hovering your mouse over each bar.
# What are the major differences between them?
# **ANSWER: (click here to edit)**
#
#
# <hr>
# **QUESTION:**
#
# What other tool(s) can you think of that we haven't tried that we could use to compare these models?
# **ANSWER: (click here to edit)**
#
#
# <hr>
#
| Random-forests-PS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import nemo
import nemo.collections.nlp as nemo_nlp
from nemo.collections.nlp.data.datasets import BertTextClassificationDataset
from nemo.collections.nlp.nm.data_layers.text_classification_datalayer import BertTextClassificationDataLayer
from nemo.collections.nlp.nm.trainables import SequenceClassifier
from nemo.backends.pytorch.common import CrossEntropyLossNM
from nemo.utils.lr_policies import get_lr_policy
from nemo.collections.nlp.callbacks.text_classification_callback import eval_iter_callback, eval_epochs_done_callback
import os
import json
import math
import numpy as np
import pandas as pd
pd.options.display.max_colwidth = -1
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
# %matplotlib inline
import torch
# -
# This notebook is based on [text_classification_with_bert.py](https://github.com/NVIDIA/NeMo/blob/master/examples/nlp/text_classification/text_classification_with_bert.py) training script, also see Text Classification documentation.
# ## Data Explore
#
# [The SST-2 dataset](https://nlp.stanford.edu/sentiment/index.html) is a standard benchmark for [sentiment classification task](http://nlpprogress.com/english/sentiment_analysis.html). The SST-2 is also a part of the [GLUE Benchmark](https://gluebenchmark.com/tasks). To download the SST-2 dataset as part of GLUE data, you can download [this script](https://gist.github.com/W4ngatang/60c2bdb54d156a41194446737ce03e2e) and then run:
#
# `python download_glue_data.py --data_dir glue_data --tasks SST`
# After running the script, the SST-2 could be found under ``glue_data/SST-2`` folder.
# to get the list of all pretrained BERT Language models run
nemo_nlp.nm.trainables.get_pretrained_lm_models_list()
# +
WORK_DIR = 'output/'
DATA_DIR = 'glue_data/SST-2'
# To use mixed precision, set AMP_OPTIMIZATION_LEVEL to 'O1' or 'O2',
# to train without mixed precision, set it to 'O0'.
AMP_OPTIMIZATION_LEVEL = 'O1'
# select the model name from the list generated in the previous cell
PRETRAINED_MODEL_NAME = 'bert-base-uncased'
MAX_SEQ_LEN = 64 # we will pad with 0's shorter sentences and truncate longer
BATCH_SIZE = 256 # 64 for 'bert-large-uncased'
# -
df = pd.read_csv(DATA_DIR + '/train.tsv', sep='\t')
test_df = pd.read_csv(DATA_DIR + '/test.tsv', sep='\t')
df.head()
test_df.head()
# The dataset comes with a train file (labeled) and a test file (not labeled). We will use part of the train file for model validation
# Split train to train and val and save to disk
np.random.seed(123)
train_mask = np.random.rand((len(df))) < .8
train_df = df[train_mask]
val_df = df[~train_mask]
# In order to take advantage of NeMo's pre-built text classification data layer, the data should be formatted as "sentence\tlabel" (sentence tab label).
# We will add a label column with all 0's (but they will not be used for anything).
test_df['label'] = 0
test_df = test_df[['sentence', 'label']]
test_df.head()
# +
# Save new train, val, and test to disk
SPLIT_DATA_DIR = os.path.join(DATA_DIR, 'split')
os.makedirs(SPLIT_DATA_DIR, exist_ok=True)
train_df.to_csv(os.path.join(SPLIT_DATA_DIR, 'train.tsv'), sep='\t', index=False)
val_df.to_csv(os.path.join(SPLIT_DATA_DIR, 'eval.tsv'), sep='\t', index=False)
test_df.to_csv(os.path.join(SPLIT_DATA_DIR, 'test.tsv'), sep='\t', index=False)
# -
# ## Neural Modules
#
# In NeMo, everything is a Neural Module. Neural modules abstract data and neural network architectures. Where a deep learning framework like PyTorch or Tensorflow is used to combine neural network layers to create a neural network.
# NeMo is used to combine data and neural networks to create AI applications.
# The Neural Module Factory will then manage the neural modules, taking care to flow data through the neural modules, and is also responsible for training (including mixed precision and distributed), logging, and inference.
# instantiate the neural module factory
nf = nemo.core.NeuralModuleFactory(log_dir=WORK_DIR,
create_tb_writer=True,
add_time_to_log_dir=False,
optimization_level=AMP_OPTIMIZATION_LEVEL)
# Pre-trained models will be automatically downloaded and cached.
# Pre-trained BERT
bert = model = nemo_nlp.nm.trainables.get_pretrained_lm_model(pretrained_model_name=PRETRAINED_MODEL_NAME)
tokenizer = nemo.collections.nlp.data.tokenizers.get_tokenizer(tokenizer_name='nemobert', pretrained_model_name=PRETRAINED_MODEL_NAME)
# Note here that the BERT models we are working with are massive. This gives our models a large capacity for learning that is needed to understand the nuance and complexity of natural language.
print(f'{PRETRAINED_MODEL_NAME} has {bert.num_weights} weights')
# Here we define and instantiate the feed forward network that takes as input our BERT embeddings. This network will be used to output the sentence classifications.
# +
# mlp classifier
bert_hidden_size = bert.hidden_size
mlp = SequenceClassifier(hidden_size=bert_hidden_size,
num_classes=2,
num_layers=2,
log_softmax=False,
dropout=0.1)
loss = CrossEntropyLossNM()
# -
# Compared to the BERT model, the MLP is tiny.
print(f'MLP has {mlp.num_weights} weights')
# # Pipelines
#
# Pipelines are used to define how data will flow the different neural networks. In this case, our data will flow through the BERT network and then the MLP network.
#
# We also have different pipelines for training, validation, and inference data.
#
# For training data, we want it to be used for optimization so it must be shuffled and we also need to compute the loss.
#
# For validation data, we won't use it for optimization but we want to know the loss.
#
# And for inference data, we only want the final predictions coming from the model.
# ## Data Layers
# We can gain a lot of efficiency by saving the tokenized data to disk. For future model runs we then don't need to tokenize every time.
# +
USE_CACHE = True
train_data = BertTextClassificationDataLayer(input_file=os.path.join(SPLIT_DATA_DIR, 'train.tsv'),
tokenizer=tokenizer,
max_seq_length=MAX_SEQ_LEN,
shuffle=True,
batch_size=BATCH_SIZE,
use_cache=USE_CACHE)
val_data = BertTextClassificationDataLayer(input_file=os.path.join(SPLIT_DATA_DIR, 'eval.tsv'),
tokenizer=tokenizer,
max_seq_length=MAX_SEQ_LEN,
batch_size=BATCH_SIZE,
use_cache=USE_CACHE)
# -
train_input, train_token_types, train_attn_mask, train_labels = train_data()
val_input, val_token_types, val_attn_mask, val_labels = val_data()
# ## BERT Embeddings
train_embeddings = bert(input_ids=train_input,
token_type_ids=train_token_types,
attention_mask=train_attn_mask)
val_embeddings = bert(input_ids=val_input,
token_type_ids=val_token_types,
attention_mask=val_attn_mask)
# ## Inspect BERT Embeddings
#
# If we want to inspect the data as it flows through our neural factory we can use the .infer method. This method will give us the tensors without performing any optimization.
val_input_tensors = nf.infer(tensors=[val_input])
print(val_input_tensors[0][0][0])
# %%time
val_embeddings_tensors = nf.infer(tensors=[val_embeddings])
# each word is embedded into bert_hidden_size space
# shape: BATCH_SIZE * MAX_SEQ_LEN * bert_hidden_size
val_embeddings_tensors[0][0].shape
print(val_embeddings_tensors[0][0][1][:, 0])
# ## Understanding and Visualizing BERT Embeddings
#
# We are going to look at the BERT embeddings for the words (1-word sentences) in "SPLIT_DATA_DIR/positive_negative.tsv". Since the BERT embeddings are 768 dimensional for BERT base and 1024 dimensional for BERT large, we'll first apply TSNE and reduce the embeddings to two dimensions.
# +
spectrum_words = ['abysmal', 'apalling', 'dreadful', 'awful', 'terrible',
'very bad', 'really bad', 'rubbish', 'unsatisfactory',
'bad', 'poor', 'great', 'really good', 'very good', 'awesome'
'fantastic', 'superb', 'brilliant', 'incredible', 'excellent'
'outstanding', 'perfect']
spectrum_file = os.path.join(SPLIT_DATA_DIR, 'positive_negative.tsv')
with open(spectrum_file, 'w+') as f:
f.write('sentence\tlabel')
for word in spectrum_words:
f.write('\n' + word + '\t0')
# -
spectrum_df = pd.read_csv(spectrum_file, delimiter='\t')
print(spectrum_df.head())
# positive negative spectrum
spectrum_data = BertTextClassificationDataLayer(input_file=spectrum_file,
tokenizer=tokenizer,
max_seq_length=MAX_SEQ_LEN,
batch_size=BATCH_SIZE)
spectrum_input, spectrum_token_types, spectrum_attn_mask, spectrum_labels = spectrum_data()
spectrum_embeddings = bert(input_ids=spectrum_input,
token_type_ids=spectrum_token_types,
attention_mask=spectrum_attn_mask)
spectrum_embeddings_tensors = nf.infer(tensors=[spectrum_embeddings])
spectrum_embeddings_tensors[0][0].shape
plt.figure(figsize=(100,100))
plt.imshow(spectrum_embeddings_tensors[0][0][:,0,:].numpy())
# +
spectrum_activations = spectrum_embeddings_tensors[0][0][:,0,:].numpy()
tsne_spectrum = TSNE(n_components=2, perplexity=10, verbose=1, learning_rate=2,
random_state=123).fit_transform(spectrum_activations)
fig = plt.figure(figsize=(10,10))
plt.plot(tsne_spectrum[0:11, 0], tsne_spectrum[0:11, 1], 'rx')
plt.plot(tsne_spectrum[11:, 0], tsne_spectrum[11:, 1], 'bo')
for (x,y, label) in zip(tsne_spectrum[0:, 0], tsne_spectrum[0:, 1], spectrum_df.sentence.values.tolist() ):
plt.annotate(label, # this is the text
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignment can be left, right or center
# -
# ## Training Pipeline
#
# In order to optimize our network, we need to pass the embeddings through the MLP network and then compute the loss.
# +
train_logits = mlp(hidden_states=train_embeddings)
val_logits = mlp(hidden_states=val_embeddings)
train_loss = loss(logits=train_logits, labels=train_labels)
val_loss = loss(logits=val_logits, labels=val_labels)
# -
# ## Callbacks
#
# Callbacks are used to record and log metrics and save checkpoints for the training and evaluation. We use callbacks to print to screen and also to tensorboard.
#
#
#
# +
NUM_EPOCHS = 1
NUM_GPUS = 1
LEARNING_RATE = 1e-5
OPTIMIZER = 'adam'
train_data_size = len(train_data)
steps_per_epoch = math.ceil(train_data_size / (BATCH_SIZE * NUM_GPUS))
train_callback = nemo.core.SimpleLossLoggerCallback(tensors=[train_loss, train_logits],
print_func=lambda x:nemo.logging.info(f'Train loss: {str(np.round(x[0].item(), 3))}'),
tb_writer=nf.tb_writer,
get_tb_values=lambda x: [["train_loss", x[0]]],
step_freq=steps_per_epoch)
eval_callback = nemo.core.EvaluatorCallback(eval_tensors=[val_logits, val_labels],
user_iter_callback=lambda x, y: eval_iter_callback(x, y, val_data),
user_epochs_done_callback=lambda x:
eval_epochs_done_callback(x, f'{nf.work_dir}/graphs'),
tb_writer=nf.tb_writer,
eval_step=steps_per_epoch)
# Create callback to save checkpoints
ckpt_callback = nemo.core.CheckpointCallback(folder=nf.checkpoint_dir,
epoch_freq=1)
# -
lr_policy_fn = get_lr_policy('WarmupAnnealing',
total_steps=NUM_EPOCHS * steps_per_epoch,
warmup_ratio=0.1)
# %%time
nf.train(tensors_to_optimize=[train_loss],
callbacks=[train_callback, eval_callback, ckpt_callback],
lr_policy=lr_policy_fn,
optimizer=OPTIMIZER,
optimization_params={'num_epochs': NUM_EPOCHS, 'lr': LEARNING_RATE})
# ## Multi-Gpu Training
#
# RESTART KERNEL BEFORE RUNNING THE MULTI-GPU TRAINING
# %%time
num_gpus = 4
# !python -m torch.distributed.launch --nproc_per_node=$NUM_GPUS text_classification_with_bert.py \
# --pretrained_model_name $PRETRAINED_MODEL_NAME \
# --data_dir $SPLIT_DATA_DIR \
# --train_file_prefix 'train' \
# --eval_file_prefix 'eval' \
# --use_cache \
# --batch_size $BATCH_SIZE \
# --max_seq_length 64 \
# --num_gpus $NUM_GPUS \
# --num_epochs $NUM_EPOCHS \
# --amp_opt_level $AMP_OPTIMIZATION_LEVEL \
# --work_dir $WORK_DIR
# ## Inference Pipeline
#
# For inference we instantiate the same neural modules but now we will be using the checkpoints that we just learned.
test_data = BertTextClassificationDataLayer(input_file=os.path.join(SPLIT_DATA_DIR, 'test.tsv'),
tokenizer=tokenizer,
max_seq_length=MAX_SEQ_LEN,
batch_size=BATCH_SIZE)
test_input, test_token_types, test_attn_mask, _ = test_data()
test_embeddings = bert(input_ids=test_input,
token_type_ids=test_token_types,
attention_mask=test_attn_mask)
test_logits = mlp(hidden_states=test_embeddings)
# %%time
test_logits_tensors = nf.infer(tensors=[test_logits])
test_probs = torch.nn.functional.softmax(torch.cat(test_logits_tensors[0])).numpy()[:, 1]
test_df = pd.read_csv(os.path.join(SPLIT_DATA_DIR, 'test.tsv'), sep='\t')
test_df['prob'] = test_probs
inference_file = os.path.join(SPLIT_DATA_DIR, 'test_inference.tsv')
test_df.to_csv(inference_file, sep='\t', index=False)
def sample_classification(data_path):
df = pd.read_csv(data_path, sep='\t')
sample = df.sample()
sentence = sample.sentence.values[0]
prob = sample.prob.values[0]
result = f'{sentence} | {prob}'
return result
num_samples = 10
for _ in range(num_samples):
print(sample_classification(inference_file))
# ## Inference Results:
# the film is just a big , gorgeous , mind-blowing , breath-taking mess . | 0.2738656
#
# a sensual performance from abbass buoys the flimsy story , but her inner journey is largely unexplored and we 're left wondering about this exotic-looking woman whose emotional depths are only hinted at . | 0.48260054
# ## Single sentence classification
def classify_sentence(nf, tokenizer, bert, mlp, sentence):
sentence = sentence.lower()
tmp_file = "/tmp/tmp_sentence.tsv"
with open(tmp_file, 'w+') as tmp_tsv:
header = 'sentence\tlabel\n'
line = sentence + '\t0\n'
tmp_tsv.writelines([header, line])
tmp_data = BertTextClassificationDataLayer(input_file=tmp_file,
tokenizer=tokenizer,
max_seq_length=128,
batch_size=1)
tmp_input, tmp_token_types, tmp_attn_mask, _ = tmp_data()
tmp_embeddings = bert(input_ids=tmp_input,
token_type_ids=tmp_token_types,
attention_mask=tmp_attn_mask)
tmp_logits = mlp(hidden_states=tmp_embeddings)
tmp_logits_tensors = nf.infer(tensors=[tmp_logits, tmp_embeddings])
tmp_probs = torch.nn.functional.softmax(torch.cat(tmp_logits_tensors[0])).numpy()[:, 1]
print(f'{sentence} | {tmp_probs[0]}')
# +
sentences = ['point break is the best movie of all time',
'the movie was a wonderful exercise in understanding the struggles of native americans',
'the performance of diego luna had me excited and annoyed at the same time',
'<NAME>on is the only good thing about this film']
for sentence in sentences:
classify_sentence(nf, tokenizer, bert, mlp, sentence)
# -
# ## Understanding and Visualizing BERT Embeddings
#
# Now that we've fine-tuned our BERT model, let's see if the word embeddings have changed.
# +
spectrum_embeddings = bert(input_ids=spectrum_input,
token_type_ids=spectrum_token_types,
attention_mask=spectrum_attn_mask)
spectrum_embeddings_tensors = nf.infer(tensors=[spectrum_embeddings])
plt.figure(figsize=(100,100))
plt.imshow(spectrum_embeddings_tensors[0][0][:,0,:].numpy())
# +
spectrum_activations = spectrum_embeddings_tensors[0][0][:,0,:].numpy()
tsne_spectrum = TSNE(n_components=2, perplexity=10, verbose=1, learning_rate=2,
random_state=123).fit_transform(spectrum_activations)
fig = plt.figure(figsize=(10,10))
plt.plot(tsne_spectrum[0:11, 0], tsne_spectrum[0:11, 1], 'rx')
plt.plot(tsne_spectrum[11:, 0], tsne_spectrum[11:, 1], 'bo')
for (x,y, label) in zip(tsne_spectrum[0:, 0], tsne_spectrum[0:, 1], spectrum_df.sentence.values.tolist() ):
plt.annotate(label, # this is the text
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignment can be left, right or center
# -
| examples/nlp/text_classification/sentiment_analysis_with_bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# ===================
# Tissot's Indicatrix
# ===================
#
# Creating a flatmap from a folded cortical surface always introduces some
# distortion. This is similar to what happens when a map of the globe is flattened
# into a 2-D map like a Mercator projection. For the cortical surface the amount
# and type of distortion will depend on the curvature of the surface (i.e. whether
# it is on a gyrus or a sulcus) and on the distance to the nearest cut.
#
# In general, we recommend examining data both in flattened and original 3-D space
# using the interactive webGL viewer, but it is also informative to visualize the
# distortion directly.
#
# One method to show distortion is to visualize how geodesic discs, which contain
# all of the points within some geodesic distance of a central point, appear on the
# flattened cortical surface.
#
# This technique is traditionally used to characterize and visualize distortions
# introduced by flattening the globe onto a map:
#
# 
#
#
# +
import cortex
import matplotlib.pyplot as plt
tissot = cortex.db.get_surfinfo("S1", "tissots_indicatrix", radius=10, spacing=30)
tissot.cmap = 'plasma'
cortex.quickshow(tissot, with_labels=False, with_rois=False, with_colorbar=False)
plt.show()
| example-notebooks/surface_analyses/plot_tissots_indicatrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# \title{Continues HDL Sinewave Generator via Chebyshev Polynomial Approximation in Python's myHDL}
# \author{<NAME>}
# \maketitle
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Python-Libraries-Utilized" data-toc-modified-id="Python-Libraries-Utilized-1"><span class="toc-item-num">1 </span>Python Libraries Utilized</a></div><div class="lev1 toc-item"><a href="#Acknowledgments" data-toc-modified-id="Acknowledgments-2"><span class="toc-item-num">2 </span>Acknowledgments</a></div><div class="lev1 toc-item"><a href="#Derivation-of-the-Chebyshev-Polynomial-Approximation-for-Cos" data-toc-modified-id="Derivation-of-the-Chebyshev-Polynomial-Approximation-for-Cos-3"><span class="toc-item-num">3 </span>Derivation of the Chebyshev Polynomial Approximation for Cos</a></div><div class="lev2 toc-item"><a href="#Chebyshev-Polynomials" data-toc-modified-id="Chebyshev-Polynomials-31"><span class="toc-item-num">3.1 </span>Chebyshev Polynomials</a></div><div class="lev3 toc-item"><a href="#Plot-of-Chebyshev-Polynomials" data-toc-modified-id="Plot-of-Chebyshev-Polynomials-311"><span class="toc-item-num">3.1.1 </span>Plot of Chebyshev Polynomials</a></div><div class="lev2 toc-item"><a href="#Chebyshev-Polynomial-cos-Relationship" data-toc-modified-id="Chebyshev-Polynomial-cos-Relationship-32"><span class="toc-item-num">3.2 </span>Chebyshev Polynomial $\cos$ Relationship</a></div><div class="lev2 toc-item"><a href="#Sinusoid-Generator-Algorithm-Development" data-toc-modified-id="Sinusoid-Generator-Algorithm-Development-33"><span class="toc-item-num">3.3 </span>Sinusoid Generator Algorithm Development</a></div><div class="lev3 toc-item"><a href="#Future-additions" data-toc-modified-id="Future-additions-331"><span class="toc-item-num">3.3.1 </span>Future additions</a></div><div class="lev1 toc-item"><a href="#Test-Bench-for-Sinusoid-Generator" data-toc-modified-id="Test-Bench-for-Sinusoid-Generator-4"><span class="toc-item-num">4 </span>Test Bench for Sinusoid Generator</a></div><div class="lev1 toc-item"><a href="#Post-Processing-of-Test-Bench" data-toc-modified-id="Post-Processing-of-Test-Bench-5"><span class="toc-item-num">5 </span>Post Processing of Test Bench</a></div><div class="lev1 toc-item"><a href="#RTL-Synthisis" data-toc-modified-id="RTL-Synthisis-6"><span class="toc-item-num">6 </span>RTL Synthisis</a></div><div class="lev1 toc-item"><a href="#Conclusion" data-toc-modified-id="Conclusion-7"><span class="toc-item-num">7 </span>Conclusion</a></div><div class="lev1 toc-item"><a href="#Refrances" data-toc-modified-id="Refrances-8"><span class="toc-item-num">8 </span>Refrances</a></div>
# -
# # Python Libraries Utilized
# +
import numpy as np
import pandas as pd
from sympy import *
init_printing()
from IPython.display import display, Math, Latex
from myhdl import *
from myhdlpeek import Peeker
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Acknowledgments
# The orgianl Chebyshev Sinwave Genrator written in myHDL was done by ["HARDSOFTLUCID"](https://hardsoftlucid.wordpress.com/various-stuff/myhdl/)
# (myHDL.old version [here](https://github.com/jandecaluwe/site-myhdl-retired/blob/master/_ori/pages/projects/continuous_wave_sinusgenerator.txt))
#
# Author of myHDL [<NAME>](http://www.myhdl.org/users/jandecaluwe.html) and the author of the myHDL Peeker [XESS Corp.](https://github.com/xesscorp/myhdlpeek)
#
#
# # Derivation of the Chebyshev Polynomial Approximation for Cos
# ## Chebyshev Polynomials
# We Start with recalling that the double(n) angle trig identity of $\cos$ for $n=2$ is
# $$\cos(2\theta)= \cos(\theta)^2 -\sin(\theta)^2 = 2\cos(\theta)^2 -1$$
# and for $n=3$ is
# $$\cos(3\theta)= cos(\theta)^3 -3\sin(\theta)^2 \cos(\theta)=4\cos(\theta)^3 -3\cos(\theta)$$
# Now exploiting Chebyshev polynomials that come from the power series solution($y(x)=\sum_{n=0}^{\infty} a_n x^n$) of Chebyshev differential equation:
# $$(1-x^2)y" -xy'+p^2y=0$$
#
# The Power series solution takes on the form of a Recurrence relation for the $a_n$ term in the Power series as
# $$a_{n+2}=\dfrac{(n-p)(n+p)}{(n+1)(n+2)}a_n$$
# for $x\in [-1, 1]$
# that leads to the Chebyshev polynomial defined as
# $$T_0(x)=1$$
# $$T_1(x)=x$$
# $$T_{n+1}(x)=2xT_n(x)-T_{n-1}(x)$$
#
#
#
#
# ### Plot of Chebyshev Polynomials
x=np.linspace(-1.0, 1.0)
fig=plt.figure()
ax=plt.subplot(111)
for i in range(1,8+1):
coeff=[0]*i
coeff[-1]=i
y=np.polynomial.Chebyshev(coeff)(x)
ax.plot(x, y, label=f'$T_{i-1}(x)$')
bbox_to_anchor=ax.get_position()
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
ax.grid()
plt.title(r"Chebyshev Polynomials $T_0(x)-T_1(x), x\in[-1, 1]$" )
None
# ## Chebyshev Polynomial $\cos$ Relationship
#
# If now $T_n(x)=T_n(cos(\theta))=cos(n\theta)$ we have
#
# $$T_0(\cos(0\cdot\theta))=1$$
# $$T_1(\cos(1\cdot\theta))=\cos(\theta)$$
# $$T_{n+1}(\cos(\theta))=2 \cos(\theta)T_n(\cos(\theta))-T_{n-1}(\cos(\theta))$$
# $$\cos((n+1)\theta)=2\cos(\theta)\cos(n\theta)-\cos((n-1)\theta)$$
# solving for $\cos(\theta)$ we get
n, theta=symbols('n, theta')
LHS=cos(theta)
RHS=(cos((n+1)*theta)+cos((n-1)*theta))/(2*cos(n*theta))
Eq(LHS, RHS)
# notice that the RHS can be simplified to
simplify(RHS)
# +
#numericalize symbolic
RHSN=lambdify((n, theta), RHS, dummify=False)
fig=plt.figure()
ax=plt.subplot(111)
thetaN=np.linspace(0, 2*np.pi)
for N in range(1, 8+1):
y=RHSN(N, thetaN)
ax.plot(thetaN, y, label=f'$C_{N-1} aprox$')
ax.plot(thetaN, np.cos(thetaN), label=r'$cos(\theta)$')
ax.grid()
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title(r"Plot of $\cos(\theta), \theta \in[0, 2\pi]$ & $N\in[0, 7]$ CP Approx.")
None
# +
thetaN=np.linspace(0, 2*np.pi)
for N in range(1, 8+1):
y=np.cos(thetaN)-RHSN(N, thetaN)
plt.plot(thetaN, y, label=f'$C_{N-1} error$')
plt.grid()
plt.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title(r"Plot of error of $\cos(\theta), \theta \in[0, 2\pi]$ & $N\in[0, 7]$ CP Aprox")
None
# -
# ## Sinusoid Generator Algorithm Development
# now letting $\cos(\theta)=\cos(2*\pi f_{\cos}/f_{clk})=T_1(x)$ we can translate the recursion relationship for the Chebyshev polynomials into relationships between regestes calls as follows. Let $$T_{n+1}(x)=2xT_n(x)-T_{n-1}(x)$$ become
# $$R_2=K \cdot R_1 -R_0$$
# that is we we replace the $2x$ by a constant factor $K$ and utilize the subscripts as designations of our registers. Furthermore we know that after one call of our recursion relationship $R_2$ ($T_{n+1}(x))$) will now become our old value $R_0$ ($T_{n-1}(x)$) thus we have
# $$R_0=R_2$$
# $$R_2=K \cdot R_1 -R_0$$
#
# further it can be shown that $R_2$ is just the next state value of the $R_1$ so that the above becomes
#
# $$R_0=R_1$$
# $$R_1'=K \cdot R_1 -R_0$$
#
# where $'$ is used to indicate next state
#
# And because the multiplication of two 30 bit numbers will generate a 60 bit number the result needs to be down shifted since the full 30 bits of the register are not being utilized to prevent overflow.
#
# $$R_0=R_1$$
# $$R_1'=((K \cdot R_1)>>(\text{size of}R_1 -1 )) -R_0$$
#
#
#
def SinGenerator(SinFreq_parm, ClkFreq_parm, SinValue_out,
clk, rst, ena):
#contorl byte size and works with -1 to translate cos to sin
INTERNALWIDTH=len(SinValue_out)-2
#qunitited version of the 2x for cos(\theta)
KONSTANT_FACTOR=int(np.cos(2*np.pi * SinFreq_parm /ClkFreq_parm)* 2**(INTERNALWIDTH))
#prep the Needed regesters at sysnthis
Reg_T0=Signal(intbv((2**(INTERNALWIDTH))-1,
min=SinValue_out.min, max=SinValue_out.max))
Reg_T1=Signal(intbv(KONSTANT_FACTOR,
min=SinValue_out.min, max=SinValue_out.max))
#define the polynomal logic
@always(clk.posedge,rst.negedge)
def logicCP():
#clear and prep the regesters
if rst== 0 :
Reg_T0.next=(2**(INTERNALWIDTH))-1
Reg_T1.next=KONSTANT_FACTOR
#run a single recursion iterration of the polynomal
else:
if ena==1:
# recursive Chebyshev formulation for sinus waveform calculation
Reg_T0.next=Reg_T1
#>> shift is a overflow wrapper
Reg_T1.next=((KONSTANT_FACTOR * Reg_T1)>>(INTERNALWIDTH-1)) - Reg_T0
#pole the R1 for the value of the sin function
@always_comb
def comb_logic():
SinValue_out.next=Reg_T1
return instances()
# ### Future additions
# need to add a Amp, Freq, phase modulation to this so as to convert the cos output to any sinusoidal output
# # Test Bench for Sinusoid Generator
# +
SinFreq=0.75e6 # make a 1.45 mhz Sinus
clkFreq=10e6 # 10 mhz
clkPeriod=1.0/clkFreq
OUTPUT_BITWIDTH=30
Peeker.clear()
SinValue_out=Signal(intbv(0, min=-2**OUTPUT_BITWIDTH, max=2**OUTPUT_BITWIDTH))
Peeker(SinValue_out, 'SinVal')
SinValueTracker=[]
clk=Signal(bool(0)); Peeker(clk, 'clk')
ena=Signal(bool(0)); Peeker(ena, 'ena')
rst=Signal(bool(0)); Peeker(rst, 'rst')
DUT=SinGenerator(SinFreq_parm=SinFreq, ClkFreq_parm=clkFreq, SinValue_out=SinValue_out,
clk=clk, rst=rst, ena=ena)
def SinGenerator_TB(TestClkCyc=200):
#clock genrator
@always(delay(int(clkPeriod*0.5*1e9))) ## delay in nano seconds
def clkGen():
clk.next = not clk
# accterla test procdure
@instance
def stimulus():
while 1:
rst.next=0
ena.next=0
#wait one clock cycle
yield clk.posedge
#test reset
rst.next=1
#wait one clock cycle
yield clk.posedge
#run the sin wave genrator
ena.next=1
#run the test for 200 clock cycles
for i in range(TestClkCyc):
#wait for next clock cycle
yield clk.posedge
SinValueTracker.append(int(SinValue_out))
raise StopSimulation
return instances()
# -
# !? Peeker is failing for some reason to capture all these values so having to improvice
N=200
sim = Simulation(DUT, SinGenerator_TB(TestClkCyc=N), *Peeker.instances()).run()
#Peeker.to_wavedrom(start_time=0, stop_time=20, tock=True)
# # Post Processing of Test Bench
# +
SinGenOutDF=pd.DataFrame(columns=['GenValue'], data=SinValueTracker)
SinGenOutDF['Time[s]']=np.arange(0.0,clkPeriod*(len(SinGenOutDF)-0.5),clkPeriod)
SinGenOutDF['GenValueNorm']=SinGenOutDF['GenValue']/SinGenOutDF['GenValue'].max()
SinGenOutDF['f[Hz]']=np.arange(-clkFreq/2.0,clkFreq/2.0,clkFreq/(len(SinValueTracker)))
FFT=np.fft.fftshift(np.fft.fft(SinGenOutDF['GenValueNorm']))
SinGenOutDF['FFTMag']=np.abs(FFT)
SinGenOutDF['FFTPhase']=np.angle(FFT)
SinGenOutDF.head(5)
# -
CosDF=pd.DataFrame(columns=['Time[s]'], data=np.arange(0.0,clkPeriod*(len(SinGenOutDF)-0.5),clkPeriod))
CosDF['Cos']=np.cos(2*np.pi*SinFreq*CosDF['Time[s]'])
CosDF['CosS']=CosDF['Cos']*SinGenOutDF['GenValue'].max()
CosDF['f[Hz]']=np.arange(-clkFreq/2.0,clkFreq/2.0,clkFreq/(len(SinValueTracker)))
FFT=np.fft.fftshift(np.fft.fft(CosDF['Cos']))
CosDF['FFTMag']=np.abs(FFT)
CosDF['FFTPhase']=np.angle(FFT)
CosDF.head(5)
# +
fig, [ax0, ax1]=plt.subplots(nrows=2, ncols=1, sharex=False)
plt.suptitle(f'Plots of Sin Generator output in time for {N} Cycles')
SinGenOutDF.plot(use_index=True ,y='GenValue', ax=ax0)
CosDF.plot(use_index=True, y='CosS', ax=ax0)
ax0.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
ax0.legend(loc='best')
SinGenOutDF.plot(x='Time[s]', y='GenValueNorm', ax=ax1)
CosDF.plot(x='Time[s]', y='CosS', ax=ax1)
ax1.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
ax1.legend(loc='best')
None
# +
fig, [ax0, ax1]=plt.subplots(nrows=2, ncols=1, sharex=True)
plt.suptitle(f'Plots of Sin Generator output in freq for {N} Cycles')
SinGenOutDF.plot(x='f[Hz]' ,y='FFTMag', logy=True, ax=ax0, label='GenFFTMag')
CosDF.plot(x='f[Hz]' ,y='FFTMag', logy=True, ax=ax0, label='SinFFTMag')
ax0.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
ax0.set_ylabel('Amp [dB]')
ax0.legend(loc='best')
SinGenOutDF.plot(x='f[Hz]', y='FFTPhase', ax=ax1, label='GenFFTPhase')
CosDF.plot(x='f[Hz]', y='FFTPhase', ax=ax1, label='CosFFTPhase')
ax1.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
ax1.set_xlabel('f[Hz]'); ax1.set_ylabel('Phase [rad]')
ax1.legend(loc='best')
None
# -
# # RTL Synthisis
# +
SinFreq=0.75e6 # make a 1.45 mhz Sinus
clkFreq=10e6 # 10 mhz
clkPeriod=1.0/clkFreq
OUTPUT_BITWIDTH=30
Peeker.clear()
SinValue_out=Signal(intbv(0, min=-2**OUTPUT_BITWIDTH, max=2**OUTPUT_BITWIDTH))
Peeker(SinValue_out, 'SinVal')
SinValueTracker=[]
clk=Signal(bool(0))
ena=Signal(bool(0))
rst=Signal(bool(0))
toVerilog(SinGenerator, SinFreq, clkFreq, SinValue_out, clk, rst, ena)
toVHDL(SinGenerator, SinFreq, clkFreq, SinValue_out, clk, rst, ena)
None
# -
# Running the lines
# ```
# toVerilog(SinGenerator, SinFreq, clkFreq, SinValue_out, clk, rst, ena)
# toVHDL(SinGenerator, SinFreq, clkFreq, SinValue_out, clk, rst, ena)
# ```
# called **myHDL**'s conversion process that converted the function **SinGenerator(SinFreq_parm, ClkFreq_parm, SinValue_out, clk, rst, ena)** and the signals _SinValue_out, clk, rst, ena_ to be converted and written to _SinGenerator.v_ and _SinGenerator.vhd_ respectively in the folder where this _Jupyter Notebook_ is located.
# +
#helper functions to read in the .v and .vhd generated files into python
def VerilogTextReader(loc, printresult=True):
with open(f'{loc}.v', 'r') as vText:
VerilogText=vText.read()
if printresult:
print(f'***Verilog modual from {loc}.v***\n\n', VerilogText)
return VerilogText
def VHDLTextReader(loc, printresult=True):
with open(f'{loc}.vhd', 'r') as vText:
VerilogText=vText.read()
if printresult:
print(f'***VHDL modual from {loc}.vhd***\n\n', VerilogText)
return VerilogText
# -
_=VerilogTextReader('SinGenerator', True)
_=VHDLTextReader('SinGenerator', True)
# The RTL Schematic from the verilog myHDL synthesis via vavado 2016.1 of the Sine Generator is shown below
# <img style="float: center;" src="SinGenRTL.PNG">
# The RTL Synthesis in **Xilinx**'s _Vivado 2016.1_ shows 65 cells, 34 I/O ports 161 Nets, 2 Register Sets, and 3 RTL Operations (multiply, right shift, subtraction). Where the last two statistics are exactly as predicted from the myHDL (Python) function **SinGenerator**
# # Conclusion
# We can see that by using Python’s myHDL library one can synthesize a working sin generator that can be converted to both Verilog and VHDL making myHDL HDL language agnostic. Furthermore, by conducting the test in python we plot the data and perform the subsequent analysis in the same environment as our HDL function thus allowing for rapid prototyping. And further, with the utilization of the Peeker extension library for myHDL, we can generate a timing diagram to compare from the FPGA synthesis tools to confirm our results. And finally, by utilizing the Jupyter notebook and git, documentation from theoretical development through algorithm design and HDL synthesis can be kept in one easy to read the living digital document that can be shared with ease. Thus banishing obscene separation on code, documentation, and testing that has plagued HDL DSP developers in the past
#
#
# # Refrances
# https://en.wikibooks.org/wiki/Trigonometry/For_Enthusiasts/Chebyshev_Polynomials
# https://www.geophysik.uni-muenchen.de/~igel/Lectures/NMG/05_orthogonalfunctions.pdf
| myHDL_DigitalSignalandSystems/Synthesizers/myHDL_SinewaveGenerator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Spreadsheet
#
# Make a spreadsheet using pinkfish. This is useful for developing trading strategies.
# It can also be used as a tool for buy and sell signals that you then manually execute.
# +
import datetime
import matplotlib.pyplot as plt
import pandas as pd
from talib.abstract import *
import pinkfish as pf
import pinkfish.itable as itable
# Format price data
pd.options.display.float_format = '{:0.2f}'.format
# %matplotlib inline
# -
# Set size of inline plots
'''note: rcParams can't be in same cell as import matplotlib
or %matplotlib inline
%matplotlib notebook: will lead to interactive plots embedded within
the notebook, you can zoom and resize the figure
%matplotlib inline: only draw static images in the notebook
'''
plt.rcParams["figure.figsize"] = (10, 7)
# Some global data
symbol = 'SPY'
start = datetime.datetime(1900, 1, 1)
end = datetime.datetime.now()
# Fetch symbol data from internet; do not use local cache.
ts = pf.fetch_timeseries(symbol, use_cache=False)
ts.tail()
# Select timeseries between start and end. Back adjust prices relative to adj_close for dividends and splits.
ts = pf.select_tradeperiod(ts, start, end, use_adj=False)
ts.head()
# Add technical indicators
# +
# Add 200 day MA.
ts['sma200'] = pf.SMA(ts, timeperiod=200)
# Add ATR.
ts['atr'] = ATR(ts, timeperiod=14)
# Add 5 day high, and 5 day low
ts['high5'] = pd.Series(ts.high).rolling(window=5).max()
ts['low5'] = pd.Series(ts.low).rolling(window=5).min()
# Add RSI, and 2-period cumulative RSI
ts['rsi2'] = RSI(ts, timeperiod=2)
ts['c2rsi2'] = pd.Series(ts.rsi2).rolling(window=2).sum()
# Add midpoint
ts['mp'] = (ts.high + ts.low) / 2
# Add 10 day SMA of midpoint
ts['sma10'] = pd.Series(ts.mp).rolling(window=10).mean()
# Add temporary rolling 10 day Standard Deviation of midpoint
ts['__sd__'] = pd.Series(ts.mp).rolling(window=10).std()
# Add standard deviation envelope or channel around midpoint
ts['upper'] = ts.sma10 + ts['__sd__']*2
ts['lower'] = ts.sma10 - ts['__sd__']*2
# Drop temporary columns.
ts.drop(columns=['__sd__'], inplace=True)
# -
# Finalize timeseries
ts, start = pf.finalize_timeseries(ts, start, dropna=True)
ts.tail()
# Select a smaller time from for use with itable
df = ts['2021-01-01':]
df.head()
# Use itable to format the spreadsheet. New 5 day high has blue highlight; new 5 day low has red highlight.
# +
pt = itable.PrettyTable(
df, tstyle=itable.TableStyle(theme='theme1'), center=True, header_row=True, rpt_header=20)
pt.update_col_header_style(
format_function=lambda x: x.upper(), text_align='right')
pt.update_row_header_style(
format_function=lambda x: pd.to_datetime(str(x)).strftime('%Y/%m/%d'), text_align='right')
for col in range(pt.num_cols):
if pt.df.columns[col] == 'volume':
pt.update_cell_style(cols=[col], format_function=lambda x: format(x, '.0f'), text_align='right')
else:
pt.update_cell_style(cols=[col], format_function=lambda x: format(x, '.2f'), text_align='right')
for row in range(pt.num_rows):
if row == 0:
continue
if (pt.df['high5'][row] == pt.df['high'][row]) and \
(pt.df['high5'][row] > pt.df['high'][row-1]):
col = df.columns.get_loc('high5')
pt.update_cell_style(rows=[row], cols=[col], color='blue')
if (pt.df['low5'][row] == pt.df['low'][row]) and \
(pt.df['low5'][row] < pt.df['low'][row-1]):
col = df.columns.get_loc('low5')
pt.update_cell_style(rows=[row], cols=[col], color='maroon')
# -
pt
| examples/100.spreadsheet/spreadsheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DeepFact Check Project
# Note the dataset is not publicly available.
# +
import pickle
import shutil
from pathlib import Path
import tensorflow as tf
from models.dnn import BasicDNN
from models.recurrent.basic_recurrent import BasicRecurrent
from models.baselines import LogisticRegressionSK, MLP
from models.PositiveLearningElkan.pu_learning import PULogisticRegressionSK
from project_paths import ProjectPaths
from run_files.single_train import single_training
from util.learning_rate_utilities import linear_geometric_curve
from util.tensor_provider import TensorProvider
from util.utilities import ensure_folder
# +
# Initialize tensor-provider (data-source)
the_tensor_provider = TensorProvider(verbose=True)
# Results path
base_path = Path(ProjectPaths.results, "final_model_comparison")
shutil.rmtree(str(base_path), ignore_errors=True)
ensure_folder(base_path)
# Get program IDs
all_program_ids = the_tensor_provider.annotated_program_ids(access_restricted_data=True)
training_programs = the_tensor_provider.accessible_annotated_program_ids
test_programs = set(all_program_ids).difference(set(training_programs))
# Settings
n_runs = 1
# +
# Paths for all models
model_paths = []
# Header
print("-" * 75)
print("-" * 100)
print("\t\t\t\tFINAL MODEL COMPARISON")
print("-" * 100)
print("-" * 75, end="\n\n")
# Runs
for run_nr in range(n_runs):
print("-" * 75)
print("Run {}".format(run_nr))
print("-" * 75, end="\n\n")
####
# List of models
n_batches = 2000
# Models
model_list = [
LogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=True,
use_embedsum=False,
max_iter=300
),
LogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=False,
use_embedsum=True,
max_iter=300
),
LogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=True,
use_embedsum=True,
max_iter=300
),
PULogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=True,
use_embedsum=False
),
PULogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=False,
use_embedsum=True
),
PULogisticRegressionSK(
tensor_provider=the_tensor_provider,
use_bow=True,
use_embedsum=True
),
BasicRecurrent(
tensor_provider=the_tensor_provider,
results_path=base_path,
n_batches=n_batches,
recurrent_units=50,
feedforward_units=[200],
dropouts=[1],
dropout_rate=0.5,
l2_weight_decay=1e-6,
recurrent_neuron_type=tf.nn.rnn_cell.GRUCell,
training_curve_y_limit=1000,
learning_rate_progression=linear_geometric_curve(
n=n_batches,
starting_value=1e-4,
end_value=1e-10,
geometric_component=3. / 4,
geometric_end=5
),
),
BasicRecurrent(
tensor_provider=the_tensor_provider,
results_path=base_path,
n_batches=n_batches,
recurrent_units=400,
feedforward_units=[50],
dropouts=[1],
dropout_rate=0.5,
l2_weight_decay=1e-6,
recurrent_neuron_type=tf.nn.rnn_cell.GRUCell,
training_curve_y_limit=1000,
learning_rate_progression=linear_geometric_curve(
n=n_batches,
starting_value=1e-4,
end_value=1e-10,
geometric_component=3. / 4,
geometric_end=5
),
),
BasicRecurrent(
tensor_provider=the_tensor_provider,
results_path=base_path,
n_batches=n_batches,
recurrent_units=400,
feedforward_units=[200],
dropouts=[1],
dropout_rate=0.5,
l2_weight_decay=1e-6,
recurrent_neuron_type=tf.nn.rnn_cell.GRUCell,
training_curve_y_limit=1000,
learning_rate_progression=linear_geometric_curve(
n=n_batches,
starting_value=1e-4,
end_value=1e-10,
geometric_component=3. / 4,
geometric_end=5
),
),
BasicDNN(
tensor_provider=the_tensor_provider,
units=[150, 50],
n_batches=n_batches,
learning_rate_progression=linear_geometric_curve(
n=n_batches,
starting_value=1e-4,
end_value=1e-10,
geometric_component=3. / 4,
geometric_end=5
),
dropouts=[1, 2],
dropout_rate=0.5,
results_path=base_path
),
]
################
for model_nr, a_model in enumerate(model_list):
print("-" * 40)
print("Model {}: {}\n".format(model_nr, a_model.name))
# Run training on a single model
single_training(
tensor_provider=the_tensor_provider,
model=a_model,
test_split=list(test_programs),
training_split=training_programs,
base_path=base_path,
access_restricted_data=True
)
model_paths.append(a_model.results_path)
print("\nModels created at paths:")
for path in model_paths:
print("\t{}".format(path))
settings = dict(
base_path=base_path,
n_runs=n_runs,
training_programs=training_programs,
test_programs=test_programs,
)
pickle.dump(settings, Path(base_path, "settings.p").open("wb"))
# -
| Notebook-Main-Results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install --upgrade pip
import sys
# !{sys.executable} -m pip install numpy
import sys
# !{sys.executable} -m pip install tensorflow
import sys
# !{sys.executable} -m pip install wheel
import sys
# !{sys.executable} -m pip install joblib
import sys
# !{sys.executable} -m pip install pandas
import sys
# !{sys.executable} -m pip install scipy
import sys
# !{sys.executable} -m pip install scikit-learn
print (sys.executable)
# !pip3 install --upgrade tensorflow-gpu
# !pip3 install "tensorflow-probability>=0.8.0rc0" --pre
# !pip3 install "dm-sonnet>=2.0.0b0" --pre
| Resnet3D/Installation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Solution:
def generate(self, numRows: int) -> List[List[int]]:
result = []
for i in range(numRows):
result.append([])
for j in range(i+1):
if j == 0 or j == i:
result[i].append(1)
else:
result[i].append(result[i-1][j-1]+result[i-1][j])
return result
| easy/pascal's triangle 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prueba para científico de datos
#
# ## Parte 1 - Cargar datos y estadísticas básicas
#
# En la carpeta ```Data``` encontrará un archivo llamado ```diamonds.csv```. Este archivo contiene información de 53940 diamantes. Dentro de la información disponible, está el precio, el color, el peso, etc. Puede consultar las características completas del dataset en [este enlace](https://www.kaggle.com/shivam2503/diamonds).
#
# 1. Cargue el archivo en un dataframe de pandas
# 2. Use los metodos que conozca para describir las propiedades básicas de los datos.
# +
# Respuesta a la parte 1
# -
# ## Parte 2 - Aprendizaje no supervisado
#
# Usted desea encontrar estructura en los datos que le han sido dados.
# 1. A partir del dataframe que cargó en el primer punto, use algún algoritmo de aprendizaje no supervisado para encontrar clusters de diamantes con propiedades similares.
# 2. En una celda de markdown, describa una métrica/método que se pueda utilizar para evaluar la calidad de sus clusters.
# 3. Varie $k$ (la cantidad de clusters) de 1 a 10 y grafique su métrica en función de $k$
# 4. Qué $k$ describe mejor sus datos?
# +
# Respuesta a la parte 2
# -
# ## Parte 3 - Reducción de dimensionalidad y regresión
#
# Usted quiere predecir el precio del diamante a partir de sus características (toda columna en el dataset que no sea el precio). Sin embargo, tiene la intuición que varias columnas son redundantes - es decir - hay columnas que no aportan información nueva.
#
# 1. Realice una reducción de dimensionalidad de los datos para evitar tener información redundante. Procure que en este nuevo espacio se explique por lo menos el 90% de la varianza de los datos.
# 2. En una celda de markdown, describa una métrica que se pueda utilizar para evaluar la calidad de su regresión y su habilidad para explicar los datos.
# 3. Parta los datos en un conjunto de entrenamiento y otro de evaluación.
# 3. Sobre este nuevo espacio, entrene un algoritmo de regresión para predecir el precio de los diamantes sobre el conjunto de entrenamiento. Evalue su algoritmo con su métrica sobre el conjunto de test. ¿Qué tan bien le va a su algoritmo? ¿Lo llevaría a producción? ¿Por qué?
#
# +
# Respuesta a la parte 3
# -
# ## Parte 4 - clasificación
#
# En la carpeta ```Data``` hay un archivo llamado ```emotions.csv``` que contiene informacion sobre las ondas electromagneticas emitidas por los cerebros de 2 pacientes. Hay un total de 2549 columnas con 2132 entradas. Su trabajo es predecir el estado de ánimo de la persona (la columna label): NEUTRAL, POSITIVE o NEGATIVE a partir de las otras columnas. Puede ver una descripción extensa del dataset [aquí](https://www.kaggle.com/birdy654/eeg-brainwave-dataset-feeling-emotions).
#
# Implemente el pipeline que considere necesario para llevar a cabo esta tarea. Es libre de escoger las herramientas y los métodos de clasificación que desee siempre y cuando cumpla lo siguiente:
#
# 1. Implemente por lo menos 2 algoritmos de clasificación.
# 2. Grafique la matriz de confusión y las curvas de precisión y cobertura para cada algoritmo.
#
# Compare los resultados de sus clasificadores.
#
# 3. ¿Cuál algoritmo es mejor?
# 4. ¿Considera que el mejor algoritmo es suficiente para entrar a producción? ¿Por qué? ¿Por qué no?
#
# +
# Respuesta a la parte 4
# -
# ## Parte 5 - Despliegue
#
# Despliegue el mejor clasificador de la etapa anterior en un endpoint. El endpoint debe procesar el objeto JSON del *body* de un POST request. El formato del objeto JSON es el siguiente:
#
# ```
# {"input":[val1,val2,val3, ... ,val2548]}
#
# ```
# El orden de los valores corresponde al orden de las columnas del archivo `emotions.csv`. La lista tiene 2548 valores que corresponden a los 2548 que su clasificador debe tomar como input.
#
# El endpoint debe retornar un json de la siguiente forma si la petición fue exitosa:
#
# ```
# {"output":"clasfOutput"}
# ```
#
# Donde "clasfOutput" corresponde a la predicción del clasificador (NEUTRAL, POSITIVE o NEGATIVE).
# +
# Respuesta a la parte 5 (url del endpoint)
| Prueba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/kiksmeisedwengougent.png" alt="Banner" width="1100"/>
# <div>
# <font color=#690027 markdown="1">
# <h1><NAME></h1>
# </font>
# </div>
# <div class="alert alert-box alert-success">
# In deze notebook wordt een diep neuraal netwerk gebruikt om stomata op een grote afbeelding te herkennen.
# </div>
# <div class="alert alert-block alert-warning">
# Hoe een diep neuraal netwerk werkt en wat de onderdelen zijn kan je terugvinden in de notebook 'Fundamenten van een diep neuraal netwerk voor beeldherkenning'.
# </div>
# Voer onderstaande code-cel uit om van de functies in deze notebook gebruik te kunnen maken.
import imp
with open('.scripts/diep_neuraal_netwerk.py', 'rb') as fp:
diep_neuraal_netwerk = imp.load_module('.scripts', fp, '.scripts/diep_neuraal_netwerk.py', ('.py', 'rb', imp.PY_SOURCE))
# <div>
# <font color=#690027 markdown="1">
# <h2>1. Het referentienetwerk</h2>
# </font>
# </div>
#
# Het netwerk waarvoor we hebben gekozen bestaat uit 3 convolutionele lagen met respectievelijk 32, 64 en 128 filters en heeft 1 feedforward laag met 64 neuronen. Het gebruikt stochastic gradient descent als optimizer met een learning rate van 0,01. Het netwerk gebruikt ook dezelfde regularisatie technieken die beschreven zijn in de notebook 'Overfitting'.
#
# De volgende afbeelding geeft dit netwerk weer zoals de netwerken die we gezien hebben in de notebook 'Fundamenten van een diep neuraal netwerk voor beeldherkenning'.
#
# <img src="images/referentienetwerk.png" width="700"/>
#
# Om dit netwerk in te laden voer je de volgende code-cel uit:
diep_neuraal_netwerk.laad_referentie_model()
# <div>
# <font color=#690027 markdown="1">
# <h2>2. Stomatadetectie op een microfoto van een blad</h2>
# </font>
# </div>
#
# Momenteel kan het getrainde netwerk een classificatie maken tussen 2 klassen ('Stoma' of 'Geen stoma') wanneer het een afbeelding van 120 pixels op 120 pixels krijgt als invoer. We willen echter dat alle stomata in een grote afbeelding (1600 px op 1200 px) gevonden kunnen worden met ons netwerk. Hierbij te verstaan dat de stomata ongeveer 120 op 120 pixels groot zijn op de foto. Om dit te verwezenlijken maken we gebruik van een <b>sliding window</b>. Dit is een vierkantje van 120 px op 120 px dat uit de grote afbeelding wordt geknipt en als invoer van het getrainde netwerk wordt gebruikt. Voor dit vierkantje zal het netwerk dan een voorspelling geven waarna het vierkantje naar rechts opschuift met 10 pixels. Wanneer het vierkantje de volledige breedte van de afbeelding heeft doorlopen zal het opnieuw naar de linkerkant verspringen maar dan 10 pixels lager dan de vorige keer. Dit proces herhaalt zich tot de volledige afbeelding verwerkt werd met telkens sprongen van 10 pixels. Volgende afbeelding geeft een illustratie, rechts van de afbeelding zie je de voorspelling van het netwerk. Een voorspelling van '1' wil zeggen dat het netwerk 100% zeker is dat er een stoma in het vierkantje zit, een voorspelling van '0' wil zeggen dat het netwerk oordeelt dat er zeker geen stoma in het vierkantje zit. Zodra de waarde groter wordt dan 0.5 (de drempelwaarde: zie later) zal het vierkantje geel worden.
#
# <img src="images/slidingwindow.gif" width ="600"/>
#
# Voor elke vierkantje uit de grote afbeelding zal er op deze manier een voorspelling gemaakt worden. Vaak zullen er meerdere vierkantjes rond de effectieve stoma een positieve uitvoer geven ("Stoma") omdat er slechts met 10 pixels wordt opgeschoven en dus niet veel verschil in de vierkantjes zit. Om deze vele positieve voorspellingen samen te voegen, wordt er gebruik gemaakt van <b>clustering</b>. Clustering zal punten die dicht bij elkaar liggen als één cluster beschouwen en het middelpunt van deze clusters is dan het gevonden stoma. Er bestaan verschillende methoden om clustering uit te voeren, wij maken gebruik van <b>mean shift clustering</b>. Bij mean shift clustering zal elk punt iteratief worden verplaatst in de richting waar de punten het dichtst bij elkaar liggen binnen een bepaalde straal (deze straal wordt ook wel de bandwidth genoemd). Uiteindelijk zullen de punten niet (of nauwelijks) meer verplaatsen, de punten die dan op dezelfde plek zijn terechtgekomen, behoren tot dezelfde cluster. Volgende afbeelding geeft een voorbeeld van clustering waarbij de blauwe kruisjes de gevonden punten zijn en de rode kruisjes het middelpunt van elke cluster.
#
# <img src="images/clustering.jpg" width="400"/>
#
# Bij de detectie speelt de <b>drempelwaarde</b> ook een belangrijke rol. Deze drempelwaarde bepaalt voor welke waarden van de uitvoer we de invoer als een stoma beschouwen. Als we bijvoorbeeld 0,5 als drempelwaarde nemen zal alle uitvoer groter dan 0.5 als "Stoma" beschouwd worden en alle uitvoer kleiner dan 0,5 als "Geen stoma".
#
# Een belangrijke afweging die je moet maken bij het kiezen van de drempelwaarde is de verhouding tussen <b>precision</b> en <b>recall</b>.
#
# <ul>
# <li>Precision: het percentage van de gevonden stomata dat ook daadwerkelijk stomata zijn.</li>
# <li>Recall: het percentage van het aantal stomata op een afbeelding dat daadwerkelijk gevonden werd.</li>
# </ul>
#
# Het is duidelijk dat een lage drempelwaarde zal zorgen voor een lage precision (er worden meer punten al stomata beschouwd, dus ook foute punten) maar een hoge recall (er gaan ook meer echte stomata gevonden worden). Andersom geldt dit ook, een hoge drempelwaarde zal zorgen voor een hoge precision maar lage recall.
#
# De volgende code-cel zal voor drie afbeeldingen de stomata detecteren met het referentienetwerk. Je kan de drempelwaarde (thr) aanpassen en het resultaat interpreteren.
diep_neuraal_netwerk.vind_stomata()
# <div>
# <font color=#690027 markdown="1">
# <h2>3. Adversarial learning</h2>
# </font>
# </div>
#
# In het vakgebied van adversarial learning wordt er gezocht naar invoer die een machine kan misleiden en die dus een andere uitvoer oplevert dan de verwachte uitvoer. We kunnen dit zelf toepassen op ons eigen netwerk. Ons netwerk is getraind om stomata te herkennen op een microfoto van een blad. Dus wanneer we een afbeelding invoeren waarop iets volledig anders te zien is, zouden we verwachten dat het netwerk geen enkele stoma vindt. Toch blijkt dit niet het geval, voer maar eens volgende code-cel uit om te zien hoe het netwerk reageert op een afbeelding van een kat.
diep_neuraal_netwerk.misleid_netwerk()
# Je kan zien dat het netwerk niet echt leert wat een stoma is maar gewoon patronen leert herkennen. De ogen van de kat hebben ongeveer dezelfde vorm als een stoma en zullen dus ook als stoma gedetecteerd worden.
#
# Er bestaan tal van voorbeelden van dit soort 'adversarial afbeeldingen'. Eén gebied in adverserial learning gaat op zoek naar afbeeldingen waarmee een (of meerdere) netwerken het moeilijk hebben. Een gekend voorbeeld zijn de afbeeldingen van chihuahua's en muffins waarbij een mens soms al twee keer moet kijken om het onderscheid te maken.
#
# <img src="images/chihuahuamuffin.jpg" width="600"/>
#
# Een andere categorie van adverserial learning zijn de zichtbare manipulaties. In deze categorie is het onderscheid voor de mens duidelijk te zien, maar kan een machine wel misleid worden. Denk hierbij bijvoorbeeld aan een zelfrijdende auto die een mens niet meer herkent omdat de mens bepaalde make-up heeft aangebracht. Onderzoekers zijn er ook in geslaagd om een netwerk voor gezichtsherkenning te misleiden door specifieke patronen op een brilmontuur te plaatsen. Zo konden ze het netwerk laten denken dat het een bepaalde beroemdheid zag ([Accessorize to a Crime: Real and Stealthy Attacks onState-of-the-Art Face Recognition](https://www.cs.cmu.edu/~sbhagava/papers/face-rec-ccs16.pdf)). Volgende afbeelding toont bovenaan de test personen met de speciaal ontworpen bril, onderaan staan de personen die het netwerk denkt te zien.
#
# <img src="images/adversarialglasses.jpg" width="600"/>
#
# Een laatste categorie zijn de onzichtbare manipulaties. Het is mogelijk een afbeelding zo te manipuleren dat het voor de mens dezelfde afbeelding lijkt maar dat een neuraal netwerk een totaal andere conclussie trekt. In een paper ([Explaining and Harnessing Adversarial Examples](https://arxiv.org/abs/1412.6572)) hebben enkele onderzoekers een afbeelding van een panda aangepast door ruiswaarden toe te voegen. Een diep neuraal netwerk dat afbeeldingen classificeert, was voor de aanpassing 57,7% zeker dat er een panda op de afbeelding stond, na de aanpassing oordeelde het netwerk met 99,3% zekerheid dat het om een gibbon ging.
#
# <img src="images/adversarialpanda.jpg" width="600"/>
#
# Deze techniek kan ook misbruikt worden, denk maar aan een zelfrijdende auto die verkeersborden herkent met een diep neuraal netwerk. Iemand met kwade bedoelingen zou een normaal stopbord kunnen vervangen door een aangepast stopbord dat er gelijkaardig uitziet voor de mens maar niet kan gedetecteerd worden door het neurale netwerk van de auto. De auto zou dan alle stopborden negeren en levensgevaarlijk zijn voor de gebruiker.
# <img src="images/cclic.png" alt="Banner" align="left" width="100"/><br><br>
# Notebook KIKS, zie <a href="http://www.aiopschool.be">AI Op School</a>, van <NAME>, <NAME>, T. Neutens & N. Gesquière is in licentie gegeven volgens een <a href="http://creativecommons.org/licenses/by-nc-sa/4.0/">Creative Commons Naamsvermelding-NietCommercieel-GelijkDelen 4.0 Internationaal-licentie</a>.
# <div>
# <h2>Met steun van</h2>
# </div>
# <img src="images/kikssteun.png" alt="Banner" width="1100"/>
| AIOpSchool/KIKS/DeepLearningBasis/0500_StomataDetectie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building The Heisman Dataset
# ***<NAME>***
#
#
# In this notebook, we'll be pulling together the data we need to answer our questions about the Heisman award.
#
# ## Table of Contents
#
# 1. [Data Cleaning and Preparation](#1)
# 1. [Preparing Votes Data](#1a)
# 2. [Setting Up Player and Game APIs](#1b)
# 3. [Pulling Stats for Past Heisman Winners](#1c)
# 2. [Exploratory Data Analysis](#2)
# 3. [Modeling and Machine Learning](#3)
#import libraries
import cfbd
import pandas as pd
import time
# <a id='1'></a><a id='1a'></a>
# ## Preparing Votes Data
#
# I found the results of the past several decades of Heisman voting here: https://www.sports-reference.com/cfb/awards/heisman.html. In order to import it to this notebook, I created a .csv file by hand.
votes_df = pd.read_csv("../data/heisman_votes.csv")
votes_df = votes_df[votes_df["Year"] > 2003]
votes_df
players = votes_df['Player']
players
#clean names
names = []
for player in players:
name = player.split('\\')[0]
if name[-1] == "*": #weird asterisk thing
names.append(name[:-1])
else:
names.append(name)
votes_df["Player"] = names
votes_df["Player"]
# <a id='1b'></a>
# ## Setting Up Player and Game APIs
#
# <a id='1bi'></a>
# **Configuring Writing Functions**
# +
#API Configuration
configuration = cfbd.Configuration()
configuration.api_key['Authorization'] = '<KEY>'
configuration.api_key_prefix['Authorization'] = 'Bearer'
players_api = cfbd.PlayersApi(cfbd.ApiClient(configuration))
games_api = cfbd.GamesApi(cfbd.ApiClient(configuration))
# +
#these functions search the players for a
def pullStatsQB(name, season, team):
qb = {}
qb['NAME'] = name
players = players_api.get_player_season_stats(year=season,team=team)
for player in players: #more accurately, this would be "for stat in stats"
if player.player == name:
if player.category == 'passing':
qb["Pass"+player.stat_type] = player.stat
if player.category == 'rushing':
qb["Rush"+player.stat_type] = player.stat
return qb
def pullStatsRB(name, season, team):
rb = {}
rb['NAME'] = name
players = players_api.get_player_season_stats(year=season,team=team)
for player in players:
if player.player == name:
if player.category == 'rushing':
rb["Rush"+player.stat_type] = player.stat
if player.category == 'receiving':
rb["Rec"+player.stat_type] = player.stat
return rb
def pullStatsWR(name, season, team):
wr = {}
wr['NAME'] = name
players = players_api.get_player_season_stats(year=season,team=team)
for player in players:
if player.player == name:
if player.category == 'rushing':
wr["Rush"+player.stat_type] = player.stat
if player.category == 'receiving':
wr["Rec"+player.stat_type] = player.stat
return wr
# -
#let's make an additional function to add our heisman stats to these dicts
def addVoteStats(d):
d['Year'] = row['Year']
d['School'] = row['School']
d['Class'] = row['Class']
d['1stVotes'] = row['1st']
d['2ndVotes'] = row['2nd']
d['3rdVotes'] = row['3rd']
d['TotalVotes'] = row['Tot']
return d
#let's make one final function that adds the stats of a player's team in
def addTeamStats(d, name, season, team):
records = games_api.get_team_records(year=season, team=team)
try:
d['win_percent'] = records[0].total['wins']/records[0].total['games']
except:
d['win_percent'] = -1
return d
# <a id='1bii'></a>
# **Testing Functions**
kp = pullStatsQB("<NAME>", 2020, "Pittsburgh")
mi = pullStatsRB("<NAME>", 2009, "Alabama")
wr = pullStatsWR("<NAME>", 2015, "Clemson")
kp
mi
wr
# <a id='1biii'></a>
# **Potential Problems**
#
# There were a few cases where the school names from Sports Reference did not align with the CFBD api, in which case I manually changed them (Brigham Young->BYU, Texas Christian->TCU, etc). Additionally, there are a few other cases of key errors I changed manually after exporting the .csv below. Finally, I dropped fumbles as a statistic because of inconsistent access (some players had simple NaN when other sources said they fumbled).
#
# <a id='1c'></a>
# ## Pulling Stats for Past Heisman Winners
# +
qbs = []
rbs = []
wrs = []
total_time = time.time()
for i,row in votes_df.iterrows():
player_time = time.time()
#gather position specific data
if row["Pos"] == "QB":
d = pullStatsQB(row["Player"], row["Year"], row["School"])
d = addVoteStats(d)
d = addTeamStats(d, row["Player"], row["Year"], row["School"])
qbs.append(d)
elif row["Pos"] == "RB":
d = pullStatsRB(row["Player"], row["Year"], row["School"])
d = addVoteStats(d)
d = addTeamStats(d, row["Player"], row["Year"], row["School"])
rbs.append(d)
elif row["Pos"] == "WR":
d = pullStatsWR(row["Player"], row["Year"], row["School"])
d = addVoteStats(d)
d = addTeamStats(d, row["Player"], row["Year"], row["School"])
wrs.append(d)
#print("Player processed in: ", time.time()-player_time)
df_qb = pd.DataFrame(qbs)
df_rb = pd.DataFrame(rbs)
df_wr = pd.DataFrame(wrs)
print("Finished processing in ", (time.time()-total_time)/60, "mins")
# +
#df_qb.to_csv("../data/heisman_QBs_copy.csv")
#df_rb.to_csv("../data/heisman_RBs_copy.csv")
#df_wr.to_csv("../data/heisman_WRs_copy.csv")
# -
df_qb.head(50)
df_rb
df_wr
# <a id='2'></a>
#
# ## Exploratory Data Analysis
#
# As I mentioned before, I did some manual correction of a few players, so we'll reload the data and begin to summarize from there.
# <a id='3'></a>
# ## Modeling and Machine Learning
| notebooks/.ipynb_checkpoints/BuildingHeismanDataset-checkpoint.ipynb |
# ---
# layout: single
# title: "GIS in R: custom legends"
# excerpt: " ."
# authors: ['<NAME>']
# modified: '{:%Y-%m-%d}'.format(datetime.now())
# category: [course-materials]
# class-lesson: ['hw-custom-legend-r']
# permalink: /course-materials/earth-analytics/week-5/r-custom-legend/
# nav-title: 'Create custom map legend'
# module-title: 'Custom plots in R'
# module-description: 'This tutorial covers the basics of creating custom plot legends
# in R'
# module-nav-title: 'Spatial Data: Custom plots in R'
# module-type: 'homework'
# week: 5
# sidebar:
# nav:
# author_profile: false
# comments: true
# order: 1
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
#
# {% include toc title="In This Lesson" icon="file-text" %}
#
#
#
# <div class='notice--success' markdown="1">
#
#
#
# ## <i class="fa fa-graduation-cap" aria-hidden="true"></i> Learning Objectives
#
#
#
# After completing this tutorial, you will be able to:
#
#
#
# * Add a custom legend to a map in `R`.
#
# * Plot a vector dataset by attributes in `R`.
#
#
#
# ## <i class="fa fa-check-square-o fa-2" aria-hidden="true"></i> What you need
#
#
#
# You will need a computer with internet access to complete this lesson and the data for week 5 of the course.
#
#
#
# [<i class="fa fa-download" aria-hidden="true"></i> Download week 5 Data (~500 MB)](https://ndownloader.figshare.com/files/7525363){:data-proofer-ignore='' .btn }
#
#
#
# </div>
#
#
#
# ## Plot Lines by Attribute Value
#
# To plot vector data with the color of each objected determined by it's associated attribute values, the
#
# attribute values must be class = `factor`. A **factor** is similar to a category
#
# - you can group vector objects by a particular category value - for example you
#
# can group all lines of `TYPE=footpath`. However, in `R`, a factor can also have
#
# a determined *order*.
#
#
#
# By default, `R` will import spatial object attributes as `factors`.
#
#
#
# <i class="fa fa-star"></i> **Data Tip:** If our data attribute values are not
#
# read in as factors, we can convert the categorical
#
# attribute values using `as.factor()`.
#
# {: .notice}
#
#
#
# +
```{r echo=FALSE, message=FALSE, warning=FALSE}
# turn off messages everywhere
knitr::opts_chunk$set(message = FALSE, warning = FALSE)
```
# -
#
#
# +
```{r load-libraries, message=FALSE, warning=FALSE}
# load libraries
library(raster)
library(rgdal)
options(stringsAsFactors = FALSE)
```
# -
#
#
# Next, import and explore the data.
#
#
#
# +
```{r convert-to-factor}
# import roads
sjer_roads <- readOGR("data/week5/california/madera-county-roads",
"tl_2013_06039_roads")
# view the original class of the TYPE column
class(sjer_roads$RTTYP)
unique(sjer_roads$RTTYP)
```
# -
#
#
# It looks like we have some missing values in our road types. We want to plot all
#
# road types even those that are NA. Let's change the roads with an `RTTYP` attribute of
#
# NA to "unknown".
#
#
#
# Following, we can convert the road attribute to a factor.
#
#
#
# +
```{r adjust-value-unknown }
# set all NA values to "unknown" so they still plot
sjer_roads$RTTYP[is.na(sjer_roads$RTTYP)] <- "Unknown"
unique(sjer_roads$RTTYP)
# view levels or categories - note that there are no categories yet in our data!
# the attributes are just read as a list of character elements.
levels(sjer_roads$RTTYP)
# Convert the TYPE attribute into a factor
# Only do this IF the data do not import as a factor!
sjer_roads$RTTYP <- as.factor(sjer_roads$RTTYP)
class(sjer_roads$RTTYP)
levels(sjer_roads$RTTYP)
# how many features are in each category or level?
summary(sjer_roads$RTTYP)
```
# -
#
#
# When we use `plot()`, we can specify the colors to use for each attribute using
#
# the `col=` element. To ensure that `R` renders each feature by it's associated
#
# factor / attribute value, we need to create a `vector` or colors - one for each
#
# feature, according to its associated attribute value / `factor` value.
#
#
#
# To create this vector we can use the following syntax:
#
#
#
# `c("colorOne", "colorTwo", "colorThree")[object$factor]`
#
#
#
# Note in the above example we have
#
#
#
# 1. A vector of colors - one for each factor value (unique attribute value)
#
# 2. The attribute itself (`[object$factor]`) of class `factor`.
#
#
#
# Let's give this a try.
#
#
#
#
#
# +
``` {r palette-and-plot, fig.cap="Adjust colors on map by creating a palette."}
# count the number of unique values or levels
length(levels(sjer_roads$RTTYP))
# create a color palette of 4 colors - one for each factor level
roadPalette <- c("blue", "green", "grey", "purple")
roadPalette
# create a vector of colors - one for each feature in our vector object
# according to its attribute value
roadColors <- c("blue", "green", "grey", "purple")[sjer_roads$RTTYP]
head(roadColors)
# plot the lines data, apply a diff color to each factor level)
plot(sjer_roads,
col=roadColors,
lwd=2,
main="Madera County Roads")
```
# -
#
#
# ### Adjust Line Width
#
# We can also adjust the width of our plot lines using `lwd`. We can set all lines
#
# to be thicker or thinner using `lwd=`.
#
#
#
# +
```{r adjust-line-width, fig.cap="map of madera roads"}
# make all lines thicker
plot(sjer_roads,
col=roadColors,
main="Madera County Roads\n All Lines Thickness=6",
lwd=6)
```
# -
#
#
# ### Adjust Line Width by Attribute
#
#
#
# If we want a unique line width for each factor level or attribute category
#
# in our spatial object, we can use the same syntax that we used for colors, above.
#
#
#
# `lwd=c("widthOne", "widthTwo","widthThree")[object$factor]`
#
#
#
# Note that this requires the attribute to be of class `factor`. Let's give it a
#
# try.
#
#
#
# +
```{r line-width-unique, fig.cap="Map with legend that shows unique line widths." }
class(sjer_roads$RTTYP)
levels(sjer_roads$RTTYP)
# create vector of line widths
lineWidths <- (c(1, 2, 3, 4))[sjer_roads$RTTYP]
# adjust line width by level
# in this case, boardwalk (the first level) is the widest.
plot(sjer_roads,
col=roadColors,
main="Madera County Roads \n Line width varies by TYPE Attribute Value",
lwd=lineWidths)
```
# -
#
#
# <div class="notice--warning" markdown="1">
#
#
#
# ## <i class="fa fa-pencil-square-o" aria-hidden="true"></i> Optional challenge: Plot line width by attribute
#
#
#
# We can customize the width of each line, according to specific attribute value,
#
# too. To do this, we create a vector of line width values, and map that vector
#
# to the factor levels - using the same syntax that we used above for colors.
#
# HINT: `lwd=(vector of line width thicknesses)[spatialObject$factorAttribute]`
#
#
#
# Create a plot of roads using the following line thicknesses:
#
#
#
# 1. **unknown** lwd = 3
#
# 2. **M** lwd = 1
#
# 3. **S** lwd = 2
#
# 4. **C** lwd = 1.5
#
#
#
# </div>
#
#
#
# +
```{r roads-map, include=TRUE, results="hide", echo=FALSE, fig.cap="roads map modified"}
# view the factor levels
levels(sjer_roads$RTTYP)
# create vector of line width values
lineWidth <- c(1.5, 1, 2, 3)[sjer_roads$RTTYP]
# view vector
lineWidth
# in this case, boardwalk (the first level) is the widest.
plot(sjer_roads,
col=roadColors,
main="Madera County Roads \n Line width varies by Type Attribute Value",
lwd=lineWidth)
```
# -
#
#
# <i class="fa fa-star"></i> **Data Tip:** Given we have a factor with 4 levels,
#
# we can create an vector of numbers, each of which specifies the thickness of each
#
# feature in our `SpatialLinesDataFrame` by factor level (category): `c(6,4,1,2)[sjer_roads$RTTYP]`
#
# {: .notice}
#
#
#
# ## Add Plot Legend
#
# We can add a legend to our plot too. When we add a legend, we use the following
#
# elements to specify labels and colors:
#
#
#
# * **location**: we can specify an x and Y location of the plot Or generally specify the location e.g. 'bottomright'
#
# keyword. We could also use `top`, `topright`, etc.
#
# * `levels(objectName$attributeName)`: Label the **legend elements** using the
#
# categories of `levels` in an attribute (e.g., levels(sjer_roads$RTTYP) means use
#
# the levels C, S, footpath, etc).
#
# * `fill=`: apply unique **colors** to the boxes in our legend. `palette()` is
#
# the default set of colors that `R` applies to all plots.
#
#
#
# Let's add a legend to our plot.
#
#
#
# +
```{r add-legend-to-plot, fig.cap="SJER roads map with custom legend." }
# add legend to plot
plot(sjer_roads,
col=roadColors,
main="Madera County Roads\n Default Legend")
# we can use the color object that we created above to color the legend objects
roadPalette
# add a legend to our map
legend("bottomright", # location of legend
legend=levels(sjer_roads$RTTYP), # categories or elements to render in
# the legend
fill=roadPalette) # color palette to use to fill objects in legend.
```
# -
#
#
# We can tweak the appearance of our legend too.
#
#
#
# * `bty=n`: turn off the legend BORDER
#
# * `cex`: change the font size
#
#
#
# Let's try it out.
#
#
#
# +
```{r modify-legend-plot, fig.cap="modified custom legend" }
# adjust legend
plot(sjer_roads,
col=roadColors,
main="Madera County Roads \n Modified Legend - smaller font and no border")
# add a legend to our map
legend("bottomright",
legend=levels(sjer_roads$RTTYP),
fill=roadPalette,
bty="n", # turn off the legend border
cex=.8) # decrease the font / legend size
```
# -
#
#
# We can modify the colors used to plot our lines by creating a new color vector,
#
# directly in the plot code too rather than creating a separate object.
#
#
#
# `col=(newColors)[sjer_roads$RTTYP]`
#
#
#
# Let's try it!
#
#
#
# +
```{r plot-different-colors, fig.cap='adjust colors'}
# manually set the colors for the plot!
newColors <- c("springgreen", "blue", "magenta", "orange")
newColors
# plot using new colors
plot(sjer_roads,
col=(newColors)[sjer_roads$RTTYP],
main="Madera County Roads \n Pretty Colors")
# add a legend to our map
legend("bottomright",
levels(sjer_roads$RTTYP),
fill=newColors,
bty="n", cex=.8)
```
# -
#
#
# <i class="fa fa-star"></i> **Data Tip:** You can modify the defaul R color palette
#
# using the palette method. For example `palette(rainbow(6))` or
#
# `palette(terrain.colors(6))`. You can reset the palette colors using
#
# `palette("default")`!
#
# {: .notice}
#
#
#
# ## Plot Lines by Attribute
#
#
#
# Create a plot that emphasizes only roads designated as C or S (County or State).
#
# To emphasize these types of roads, make the lines that are C or S, THICKER than
#
# the other lines.
#
# NOTE: this attribute information is located in the `sjer_roads$RTTYP`
#
# attribute.
#
#
#
# Be sure to add a title and legend to your map! You might consider a color
#
# palette that has all County and State roads displayed in a bright color. All
#
# other lines can be grey.
#
#
#
# +
```{r road-map-2, include=TRUE, fig.cap='emphasize some attributes'}
# view levels
levels(sjer_roads$RTTYP)
# make sure the attribute is of class "factor"
class(sjer_roads$RTTYP)
# convert to factor if necessary
sjer_roads$RTTYP <- as.factor(sjer_roads$RTTYP)
levels(sjer_roads$RTTYP)
# count factor levels
length(levels(sjer_roads$RTTYP))
# set colors so only the allowed roads are magenta
# note there are 3 levels so we need 3 colors
challengeColors <- c("magenta","grey","magenta","grey")
challengeColors
# plot using new colors
plot(sjer_roads,
col=(challengeColors)[sjer_roads$RTTYP],
lwd=c(4,1,1,1)[sjer_roads$RTTYP],
main="SJER Roads")
# add a legend to our map
legend("bottomright",
levels(sjer_roads$RTTYP),
fill=challengeColors,
bty="n", # turn off border
cex=.8) # adjust font size
```
# -
#
#
# Finall, let's adjust the legend. We want the legend SYMBOLS to represent the
#
# actual symbology in the map - which contains lines, not polygons.
#
#
#
# +
```{r final-custom-legend, fig.cap="Custom legend with lines"}
# plot using new colors
plot(sjer_roads,
col=(challengeColors)[sjer_roads$RTTYP],
lwd=c(4,1,2,1)[sjer_roads$RTTYP], # color each line in the map by attribute
main="Madera County Roads\n County and State recognized roads")
# add a legend to our map
legend("bottomright",
levels(sjer_roads$RTTYP),
lty=c(1,1,1,1), # tell are which objects to be drawn as a line in the legend.
lwd=c(4,1,2,1), # set the WIDTH of each legend line
col=challengeColors, # set the color of each legend line
bty="n", # turn off border
cex=.8) # adjust font size
```
# -
#
#
# <!-- C = County
#
# I = Interstate
#
# M = Common Name
#
# O = Other
#
# S = State recognized
#
# U = U.S.-->
#
#
#
# ## Adding point and lines to a legend
#
#
#
# The last step in customizing a legend is adding different types of symbols to
#
# the plot. In the example above, we just added lines. But what if we wanted to add
#
# some POINTS too? We will do that next.
#
#
#
# In the data below, we've create a custom legend where each symbol type and color
#
# is defined using a vector. We have 3 levels: grass, soil and trees. Thus we
#
# need to define 3 symbols and 3 colors for our legend and our plot.
#
#
#
# `pch=c(8,18,8)`
#
#
#
# `plot_colors <- c("chartreuse4", "burlywood4", "darkgreen")`
#
#
#
# +
```{r import-plots, results='hide'}
# import points layer
sjer_plots <- readOGR("data/week5/california/SJER/vector_data",
"SJER_plot_centroids")
```
# -
#
#
# +
```{r legend-points-lines, fig.cap="plot legend with points and lines" }
sjer_plots$plot_type <- as.factor(sjer_plots$plot_type)
levels(sjer_plots$plot_type)
# grass, soil trees
plot_colors <- c("chartreuse4", "burlywood4", "darkgreen")
# plot using new colors
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
pch=8,
main="Madera County Roads\n County and State recognized roads")
# add a legend to our map
legend("bottomright",
legend = levels(sjer_plots$plot_type),
pch=c(8,18,8), # set the WIDTH of each legend line
col=plot_colors, # set the color of each legend line
bty="n", # turn off border
cex=.9) # adjust legend font size
```
# -
#
#
# Next, let's try to plot our roads on top of the plot locations. Then let's create
#
# a custom legend that contains both lines and points. NOTE: in this example i've
#
# fixed the projection for the roads layer and cropped it! You will have to do the same before
#
# this code will work.
#
#
#
# +
```{r reproject-data, echo=FALSE, warning=FALSE, message=FALSE, results='hide', fig.cap="plot legend with points and lines and subheading." }
# load crop extent layer
sjer_aoi <- readOGR("data/week5/california/SJER/vector_data",
"SJER_crop")
# reproject line and point data
sjer_roads_utm <- spTransform(sjer_roads,
crs(sjer_aoi))
sjer_roads_utm <- crop(sjer_roads_utm, sjer_aoi)
```
# -
#
#
# When we create a legend, we will have to add the labels for both the points
#
# layer and the lines layer.
#
#
#
# `c(levels(sjer_plots$plot_type), levels(sjer_roads$RTTYP))`
#
#
#
# +
```{r create-legend-list }
# view all elements in legend
c(levels(sjer_plots$plot_type), levels(sjer_roads$RTTYP))
```
# -
#
#
#
#
# +
```{r custom-legend-points-lines, fig.cap="final plot custom legend."}
# plot using new colors
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
pch=8,
main="Madera County Roads and plot locations")
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend("bottomright",
legend = c(levels(sjer_plots$plot_type), levels(sjer_roads$RTTYP)),
pch=c(8,18,8), # set the WIDTH of each legend line
col=plot_colors, # set the color of each legend line
bty="n", # turn off border
cex=.9) # adjust legend font size
```
# -
#
#
#
#
# Next we have to tell `R`, which symbols are lines and which are point symbols. We
#
# can do that using the lty argument. We have 3 unique point symbols and 4 unique
#
# line symbols. We can include a NA for each element that should not be a line in
#
# the lty argument:
#
#
#
# `lty=c(NA, NA, NA, 1, 1, 1, 1)`
#
#
#
# And we include a `NA` value for each element that should not be a symbol in the
#
# `pch` argument:
#
#
#
# `pch=c(8, 18, 8, NA, NA, NA, NA)``
#
#
#
# +
```{r custom-legend-points-lines-2, fig.cap="Plot with points and lines customized."}
# plot using new colors
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
pch=8,
main="Madera County Roads and plot locations")
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend("bottomright",
legend = c(levels(sjer_plots$plot_type), levels(sjer_roads$RTTYP)),
pch=c(8,18,8, NA, NA, NA, NA), # set the symbol for each point
lty=c(NA,NA, NA, 1, 1, 1, 1),
col=c(plot_colors, challengeColors), # set the color of each legend line
bty="n", # turn off border
cex=.9) # adjust legend font size
```
# -
#
#
#
#
# ## Force the legend to plot next to your plot
#
#
#
# Refining the look of your plots takes a bit of patience in R, but it can be
#
# done! Play with the code below to see if you can make your legend plot NEXT TO
#
# rather than on top of your plot.
#
#
#
# The steps are:
#
#
#
# 1. Place your legend on the OUTSIDE of the plot extent by grabbing the `xmax` and `ymax` values from one of the objects that you are plotting's `extent()`. This allows you to be precise in your legend placement.
#
# 2. Set the `xpd=T` argument in your legend to enforce plotting outside of the plot extent and
#
# 3. OPTIONAL: adjust the plot **PAR**ameters using `par()`. You can set the **mar**gin
#
# of your plot using `mar=`. This provides extra space on the right (if you'd like your legend to plot on the right) for your legend and avoids things being "chopped off". Provide the `mar=` argument in the
#
# format: `c(bottom, left, top, right)`. The code below is telling r to add 7 units
#
# of padding on the RIGHT hand side of our plot. The default units are INCHES.
#
#
#
# **IMPORTANT:** be cautious with margins. Sometimes they can cause problems when you
#
# knit - particularly if they are too large.
#
#
#
# Let's give this a try. First, we grab the northwest corner location x and y. We
#
# will use this to place our legend.
#
#
#
# +
```{r adjust-legend, fig.cap="plot with fixed legend"}
# figure out where the upper RIGHT hand corner of our plot extent is
the_plot_extent <- extent(sjer_aoi)
# grab the upper right hand corner coordinates
furthest_pt_east <- the_plot_extent@xmax
furthest_pt_north <- the_plot_extent@ymax
# view values
furthest_pt_east
furthest_pt_north
# plot using new colors
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
pch=8,
main="Madera County Roads and plot locations")
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend(x=furthest_pt_east, y=furthest_pt_north,
legend = c(levels(sjer_plots$plot_type), levels(sjer_roads$RTTYP)),
pch=c(8, 18, 8, NA, NA, NA, NA), # set the symbol for each point
lty=c(NA, NA, NA, 1, 1, 1, 1) ,
col=c(plot_colors, challengeColors), # set the color of each legend line
bty="n", # turn off border
cex=.9, # adjust legend font size
xpd=T) # force the legend to plot outside of your extent
```
# -
#
#
# +
```{r, echo=FALSE, results='hide', message=FALSE}
dev.off()
```
# -
#
#
# Let's use the margin parameter to clean things up. Also notice i'm using the
#
# AOI extent layer to create a "box" around my plot. Now things are starting to
#
# look much cleaner!
#
#
#
# I've also added some "fake" legend elements to create subheadings like we
#
# might add to a map legend in QGIS or ArcGIS.
#
#
#
# `legend = c("Plots", levels(sjer_plots$plot_type), "Road Types", levels(sjer_roads$RTTYP))`
#
#
#
# +
```{r custom-legend-points-lines-22, fig.cap="final legend with points and lines customized 2ß."}
# adjust margin to make room for the legend
par(mar=c(2, 2, 4, 7))
# plot using new colors
plot(sjer_aoi,
border="grey",
lwd=2,
main="Madera County Roads and plot locations")
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
add=T,
pch=8)
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend(x=(furthest_pt_east+50), y=(furthest_pt_north-15),
legend = c("Plots", levels(sjer_plots$plot_type), "Road Types", levels(sjer_roads$RTTYP)),
pch=c(NA, 8, 18, 8, NA, NA, NA, NA, NA), # set the symbol for each point
lty=c(NA,NA,NA, NA, NA, 1, 1, 1, 1),
col=c(plot_colors, challengeColors), # set the color of each legend line
bty="n", # turn off border
cex=.9, # adjust legend font size
xpd=T)
```
# -
#
#
# +
```{r, echo=FALSE, results='hide'}
dev.off()
```
# -
#
#
# Let's take customization a step further. I can adjust the font styles in the legend
#
# too to make it look **even prettier**. To do this, we use the `text.font` argument.
#
#
#
# The possible values for the `text.font` argument are:
#
#
#
# * 1: normal
#
# * 2: bold
#
# * 3: italic
#
# * 4: bold and italic
#
#
#
# Notice below, i am passing a vector of values, one value to represent each
#
# element in the legend.
#
#
#
# +
```{r custom-legend-points-lines-3, fig.cap="final legend with points and lines customized 2ß."}
# adjust margin to make room for the legend
par(mar=c(2, 2, 4, 7))
# plot using new colors
plot(sjer_aoi,
border="grey",
lwd=2,
main="Madera County Roads and plot locations")
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
add=T,
pch=8)
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend(x=(furthest_pt_east+50), y=(furthest_pt_north-15),
legend = c("Plots", levels(sjer_plots$plot_type), "Road Types", levels(sjer_roads$RTTYP)),
pch=c(NA, 8, 18, 8, NA, NA, NA, NA, NA), # set the symbol for each point
lty=c(NA,NA,NA, NA, NA, 1, 1, 1, 1),
col=c(plot_colors, challengeColors), # set the color of each legend line
bty="n", # turn off border
cex=.9, # adjust legend font size
xpd=T,
text.font =c(2, 1, 1, 1, 2, 1, 1, 1, 1))
```
# -
#
#
# +
```{r, echo=FALSE, results='hide'}
dev.off()
```
# -
#
#
# Now, if you want to move the legend out a bit further, what would you do?
#
#
#
# ## BONUS!
#
#
#
# Any idea how I added a space to the legend below to create "sections"?
#
#
#
# +
```{r custom-legend-points-lines-4, echo=FALSE, fig.cap="final legend with points and lines customized."}
par(mar=c(2,2,4,7))
# plot using new colors
plot(sjer_aoi,
border="grey",
lwd=2,
main="Madera County Roads and plot locations")
plot(sjer_plots,
col=(plot_colors)[sjer_plots$plot_type],
add=T,
pch=8)
# plot using new colors
plot(sjer_roads_utm,
col=(challengeColors)[sjer_plots$plot_type],
pch=8,
add=T)
# add a legend to our map
legend(x=(furthest_pt_east+50), y=(furthest_pt_north-15),
legend = c("Plots",levels(sjer_plots$plot_type), "", "Road Types", levels(sjer_roads$RTTYP)),
pch=c(NA,8,18,8, NA, NA, NA, NA, NA, NA), # set the symbol for each point
lty=c(NA,NA,NA, NA, NA, NA,1, 1, 1, 1),
col=c(plot_colors, challengeColors), # set the color of each legend line
bty="n", # turn off border
cex=.9, # adjust legend font size
xpd=T,
text.font =c(2, 1, 1, 1, 1, 2, 1, 1, 1, 1))
```
# -
#
#
# +
```{r results='hide'}
# important: once you are done, reset par which resets plot margins
# otherwise margins will carry over to your next plot and cause problems!
dev.off()
```
| _posts/course-materials/earth-analytics/python/live/week-5/homework/2016-12-06-plot01-custom-legend-R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID-19 Case Rates for States in Mexico
# [Work in progress]
#
# This notebooks uses data from [COVID-19 Mexico, Gobierno de Mexico](https://coronavirus.gob.mx/datos)
import math
import numpy as np
import pandas as pd
import datetime
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from matplotlib.dates import DateFormatter
from py2neo import Graph
import ipywidgets as widgets
pd.options.display.max_rows = None # display all rows
pd.options.display.max_columns = None # display all columsns
# #### Connect to COVID-19-Net Knowledge Graph
graph = Graph("bolt://172.16.58.3:7687", user="reader", password="<PASSWORD>")
# #### Select Metric to display
metric_widget = widgets.Dropdown(options=('confirmedRate', 'deathRate'), description='Metric')
display(metric_widget)
metric = metric_widget.value
print('Metric:', metric)
# start date for time series
start_date = '2020-04-01'
# #### Get confirmed cases and deaths for all counties in a state
query = """
// get all states (admin1) in Mexico
MATCH (a:Admin1)-[:IN]->(:Country{name: 'Mexico'})
// get COVID-19 cases for all states
MATCH (a)<-[:REPORTED_IN]-(c:Cases{source: 'GOBMX', aggregationLevel: 'Admin1'})
WHERE c.date >= date($start_date)
RETURN a.name AS name, c.date AS date,
c.cases*100000.0/c.population AS confirmedRate,
c.deaths*100000.0/c.population AS deathRate,
c.cases AS cases,
c.deaths AS deaths,
c.population AS population
ORDER BY c.date ASC, a.name
"""
df = graph.run(query, start_date=start_date).to_data_frame()
df.tail(38)
# Reformat data
# convert neo4j date object to datetime
df['date'] = df['date'].astype(str)
df['date'] = pd.to_datetime(df['date'], infer_datetime_format=False)
# pivot table
df_date = df.pivot(index='date', columns='name', values=metric)
df_date.fillna(0, inplace=True)
df_date.head()
ax = df_date.plot(figsize=(16, 8), legend=False, title=f'{metric} for states in Mexico');
ax.set_xlabel('Date');
ax.set_ylabel(f'{metric} per 100,000');
# ### Case rate (per 100,000) by State
# dimensions for subplot layout
cols = 5
rows = math.ceil(df_date.shape[1]/cols)
ax = df_date.plot(subplots=True, layout=(rows,cols), sharey=True, figsize=(16, 2*rows));
| notebooks/obsolete/CovidRatesByStatesMexico.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inexact Move Function
#
# Let's see how we can incorporate **uncertain** motion into our motion update. We include the `sense` function that you've seen, which updates an initial distribution based on whether a robot senses a grid color: red or green.
#
# Next, you're tasked with modifying the `move` function so that it incorporates uncertainty in motion.
#
# <img src='images/uncertain_motion.png' width=50% height=50% />
#
# First let's include our usual resource imports and display function.
# importing resources
import matplotlib.pyplot as plt
import numpy as np
# A helper function for visualizing a distribution.
def display_map(grid, bar_width=1):
if(len(grid) > 0):
x_labels = range(len(grid))
plt.bar(x_labels, height=grid, width=bar_width, color='b')
plt.xlabel('Grid Cell')
plt.ylabel('Probability')
plt.ylim(0, 1) # range of 0-1 for probability values
plt.title('Probability of the robot being at each cell in the grid')
plt.xticks(np.arange(min(x_labels), max(x_labels)+1, 1))
plt.show()
else:
print('Grid is empty')
# You are given the initial variables and the complete `sense` function, below.
# +
# given initial variables
p=[0, 1, 0, 0, 0]
# the color of each grid cell in the 1D world
world=['green', 'red', 'red', 'green', 'green']
# Z, the sensor reading ('red' or 'green')
Z = 'red'
pHit = 0.6
pMiss = 0.2
# You are given the complete sense function
def sense(p, Z):
''' Takes in a current probability distribution, p, and a sensor reading, Z.
Returns a *normalized* distribution after the sensor measurement has been made, q.
This should be accurate whether Z is 'red' or 'green'. '''
q=[]
# loop through all grid cells
for i in range(len(p)):
# check if the sensor reading is equal to the color of the grid cell
# if so, hit = 1
# if not, hit = 0
hit = (Z == world[i])
q.append(p[i] * (hit * pHit + (1-hit) * pMiss))
# sum up all the components
s = sum(q)
# divide all elements of q by the sum to normalize
for i in range(len(p)):
q[i] = q[i] / s
return q
# -
# ### QUIZ: Modify the move function to accommodate the added probabilities of overshooting or undershooting the intended destination.
#
# This function should shift a distribution with the motion, U, with some probability of under/overshooting. For the given, initial `p`, you should see the result for U = 1 and incorporated uncertainties: `[0.0, 0.1, 0.8, 0.1, 0.0]`.
# +
p=[0, 1, 0, 0, 0]
## TODO: Modify the move function to accommodate the added robabilities of overshooting or undershooting
pExact = 0.8
pOvershoot = 0.1
pUndershoot = 0.1
# Complete the move function
def move(p, U):
q=[]
# iterate through all values in p
for i in range(len(p)):
# use the modulo operator to find the new location for a p value
# this finds an index that is shifted by the correct amount
index = (i-U) % len(p)
nextIndex = (index+1) % len(p)
prevIndex = (index-1) % len(p)
s = pExact * p[index]
s = s + pOvershoot * p[nextIndex]
s = s + pUndershoot * p[prevIndex]
# append the correct, modified value of p to q
q.append(s)
return q
## TODO: try this for U = 2 and see the result
p = move(p,1)
print(p)
display_map(p)
| 3-object-tracking-and-localization/activities/2-robot-location/7. Inexact Move Function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Thermal Speed
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from astropy import units as u
from plasmapy.formulary import (
Maxwellian_speed_1D,
Maxwellian_speed_2D,
Maxwellian_speed_3D,
)
from plasmapy.formulary.parameters import thermal_speed
# -
# The thermal_speed function can be used to calculate the thermal velocity for a Maxwellian velocity distribution. There are three common definitions of the thermal velocity, which can be selected using the "method" keyword, which are defined for a 3D velocity distribution as
#
#
# - 'most_probable' <br>
# $v_{th} = \sqrt{\frac{2 k_B T}{m}}$
#
# - 'rms' <br>
# $v_{th} = \sqrt{\frac{3 k_B T}{m}}$
#
# - 'mean_magnitude' <br>
# $v_{th} = \sqrt{\frac{8 k_B T}{m\pi}}$
#
# The differences between these velocities can be seen by plotitng them on a 3D Maxwellian speed distribution
# + nbsphinx-thumbnail={"tooltip": "Thermal Speeds"}
T = 1e5 * u.K
speeds = np.linspace(0, 8e6, num=600) * u.m / u.s
pdf_3D = Maxwellian_speed_3D(speeds, T=T, particle="e-")
fig, ax = plt.subplots(figsize=(4, 3))
v_most_prob = thermal_speed(T=T, particle="e-", method="most_probable", ndim=3)
v_rms = thermal_speed(T=T, particle="e-", method="rms", ndim=3)
v_mean_magnitude = thermal_speed(T=T, particle="e-", method="mean_magnitude", ndim=3)
ax.plot(speeds / v_rms, pdf_3D, color="black", label="Maxwellian")
ax.axvline(x=v_most_prob / v_rms, color="blue", label="Most Probable")
ax.axvline(x=v_rms / v_rms, color="green", label="RMS")
ax.axvline(x=v_mean_magnitude / v_rms, color="red", label="Mean Magnitude")
ax.set_xlim(-0.1, 3)
ax.set_ylim(0, None)
ax.set_title("3D")
ax.set_xlabel("|v|/|v$_{rms}|$")
ax.set_ylabel("f(|v|)")
# -
# Similar speeds are defined for 1D and 2D distributions. The differences between these definitions can be illustrated by plotting them on their respective Maxwellian speed distributions.
# +
pdf_1D = Maxwellian_speed_1D(speeds, T=T, particle="e-")
pdf_2D = Maxwellian_speed_2D(speeds, T=T, particle="e-")
dim = [1, 2, 3]
pdfs = [pdf_1D, pdf_2D, pdf_3D]
plt.tight_layout()
fig, ax = plt.subplots(ncols=3, figsize=(10, 3))
for n, pdf in enumerate(pdfs):
ndim = n + 1
v_most_prob = thermal_speed(T=T, particle="e-", method="most_probable", ndim=ndim)
v_rms = thermal_speed(T=T, particle="e-", method="rms", ndim=ndim)
v_mean_magnitude = thermal_speed(
T=T, particle="e-", method="mean_magnitude", ndim=ndim
)
ax[n].plot(speeds / v_rms, pdf, color="black", label="Maxwellian")
ax[n].axvline(x=v_most_prob / v_rms, color="blue", label="Most Probable")
ax[n].axvline(x=v_rms / v_rms, color="green", label="RMS")
ax[n].axvline(x=v_mean_magnitude / v_rms, color="red", label="Mean Magnitude")
ax[n].set_xlim(-0.1, 3)
ax[n].set_ylim(0, None)
ax[n].set_title("{:d}D".format(ndim))
ax[n].set_xlabel("|v|/|v$_{rms}|$")
ax[n].set_ylabel("f(|v|)")
ax[2].legend(bbox_to_anchor=(1.9, 0.8), loc="upper right")
# -
| docs/notebooks/thermal_speed.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Javascript (Node.js)
// language: javascript
// name: javascript
// ---
// # 16: Notes for Day 8, part 1
//
// ## Creating a simple object
//
// We looked at some Tampa Bay restaurants in Yelp. One of them was Ella’s, which had this information:
//
// * Name: **Ella's**
// * Rating (on a scale of 1 to 5 stars, with half-stars allowed): **⭐️⭐️⭐️⭐️**
// * Cost: **$$**
// * Categories: American (New), Cafes
//
// We used the following object to represent Ella’s in JavaScript:
let restaurant1 = {
name: "Ella's",
rating: 4,
cost: 2,
categories: ["American (New)", "Cafes"]
}
console.log(restaurant1)
// You could visualize the object like this:
//
// 
//
// And think of assigning the object to `restaurant1` like this:
//
// 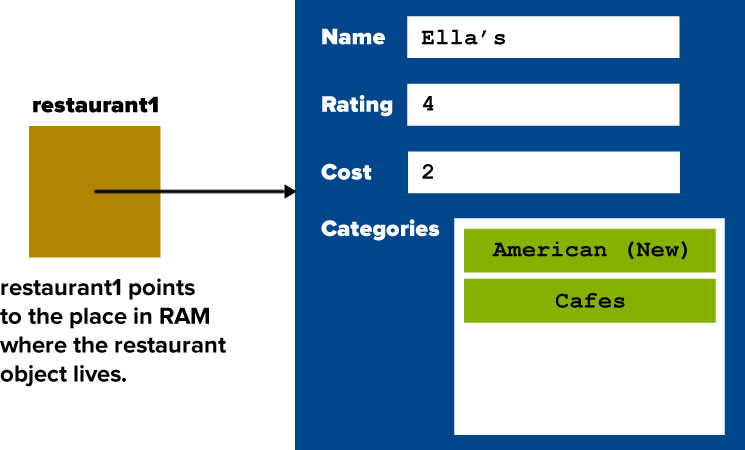
//
//
// ## An object’s properties
//
// As I like to say, objects **know things** and **do things**. Objects can “know” things by storing data inside their own constants and variables. When a constant or variable belongs to an objects, it’s often called a property.
//
// Each property has two parts:
//
// * **key**: This is the name of the property. In the object we just created, the keys are `name`, `rating`, `cost`, and `categories`.
// * **value**: This is what’s contained in a property. The values in the object we just created are listed below for each key:
// * **`name`**: `"Ella's"`
// * **`rating`**: `4`
// * **`cost`**: `2`
// * **`categories`**: `[ "American (New)", "Cafes" ]`
//
//
//
//
// Because `restaurant1` points to the object, we can use `restaurant1` to access the object’s properties.
//
// There are two ways to access an object’s properties. There’s **dot notation**, shown below:
console.log(restaurant1.name)
// The other way is to use **array notation**, shown below. It’s called “array notation” because it’s similar to the way you access elements in an array.
console.log(restaurant1["name"])
// Note that with dot notation, you don’t put the key in quotes. With array notation, you have to put the key in quotes.
//
// Here’s how you change the restaurant’s rating to 3 using dot notation:
restaurant1.rating = 3
console.log(restaurant1.rating)
// And here’s how you change the restaurant’s rating to 2 using array notation:
restaurant1["rating"] = 2
console.log(restaurant1["rating"])
// The restaurant’s `categories` property is an array, which allows it to hold more than one item. This is useful, since restaurants can have more than one category.
//
// Here’s how you access the entire `categories` array:
console.log(restaurant1.categories)
// + active=""
// To access individual elements of the `categories` array, use array indexes:
// -
console.log(restaurant1.categories[0])
// Here’s how you can update the second category:
restaurant1.categories[1] = "Bars"
console.log(restaurant1.categories[1])
// ## More objects
//
// We created a couple more objects:
let rooster = {
name: "<NAME>",
rating: 5,
cost: 3,
categories: ["Fancy", "Precious"]
}
let chicken = {
name: "<NAME> the Coop",
rating: 3,
cost: 1,
categories: ["Chicken"]
}
// We now have three objects, each on representing a restaurant:
//
// 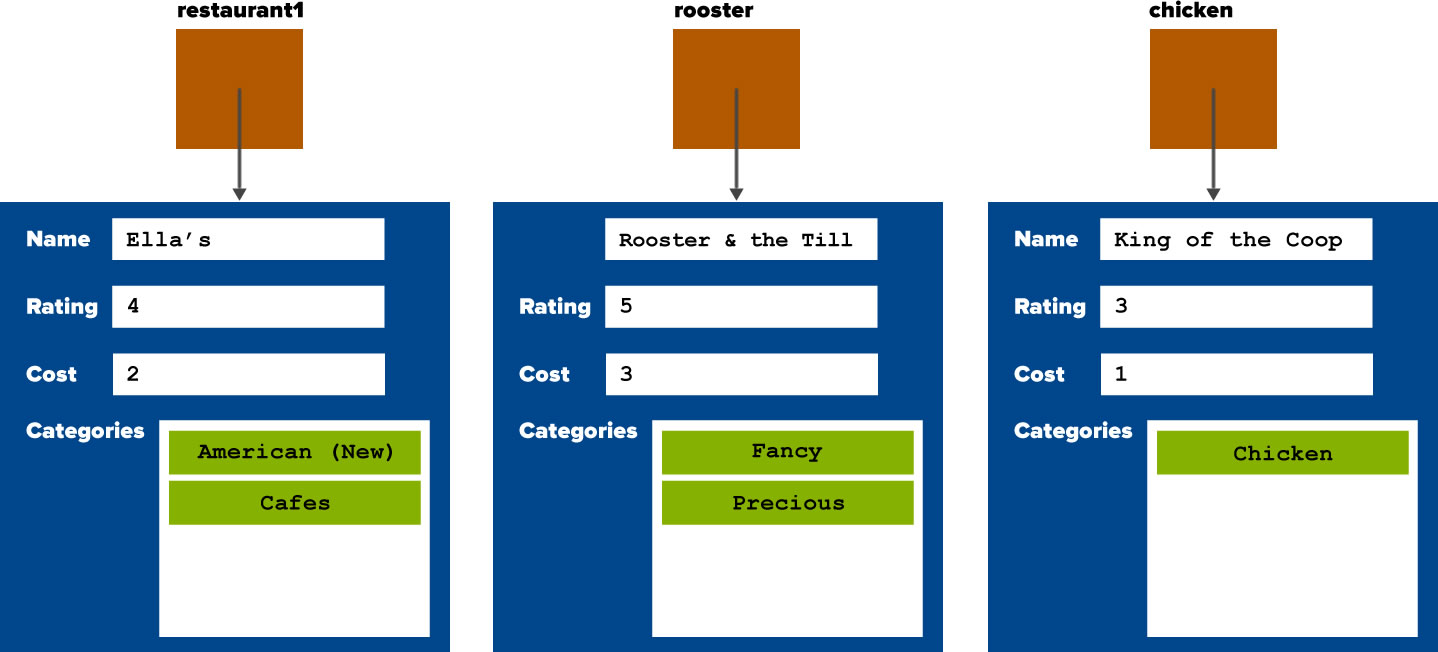
//
//
// ## Putting the objects into an array
//
// A site like Yelp organizes individual restaurants into a catalog of restaurants. We’re going to do the same, and make a mini-Yelp, which means that we’ll also have to organize these invidiual restaurants into a catalog.
//
// Any time you’re working with a collection of objects of the same or similar type, there’s a good chance you’ll need an array. Let’s put our three restaurants into an array named `restaurants`:
let restaurants = [restaurant1, rooster, chicken]
console.log(restaurants)
// Note that the array *doesn’t* contain the literal values `restaurant1`, `rooster`, and `chicken`. That’s because they’re variables that refer to objects. Listing the contents of the array lists those objects.
//
// Here’s a diagram of the objects now:
//
// 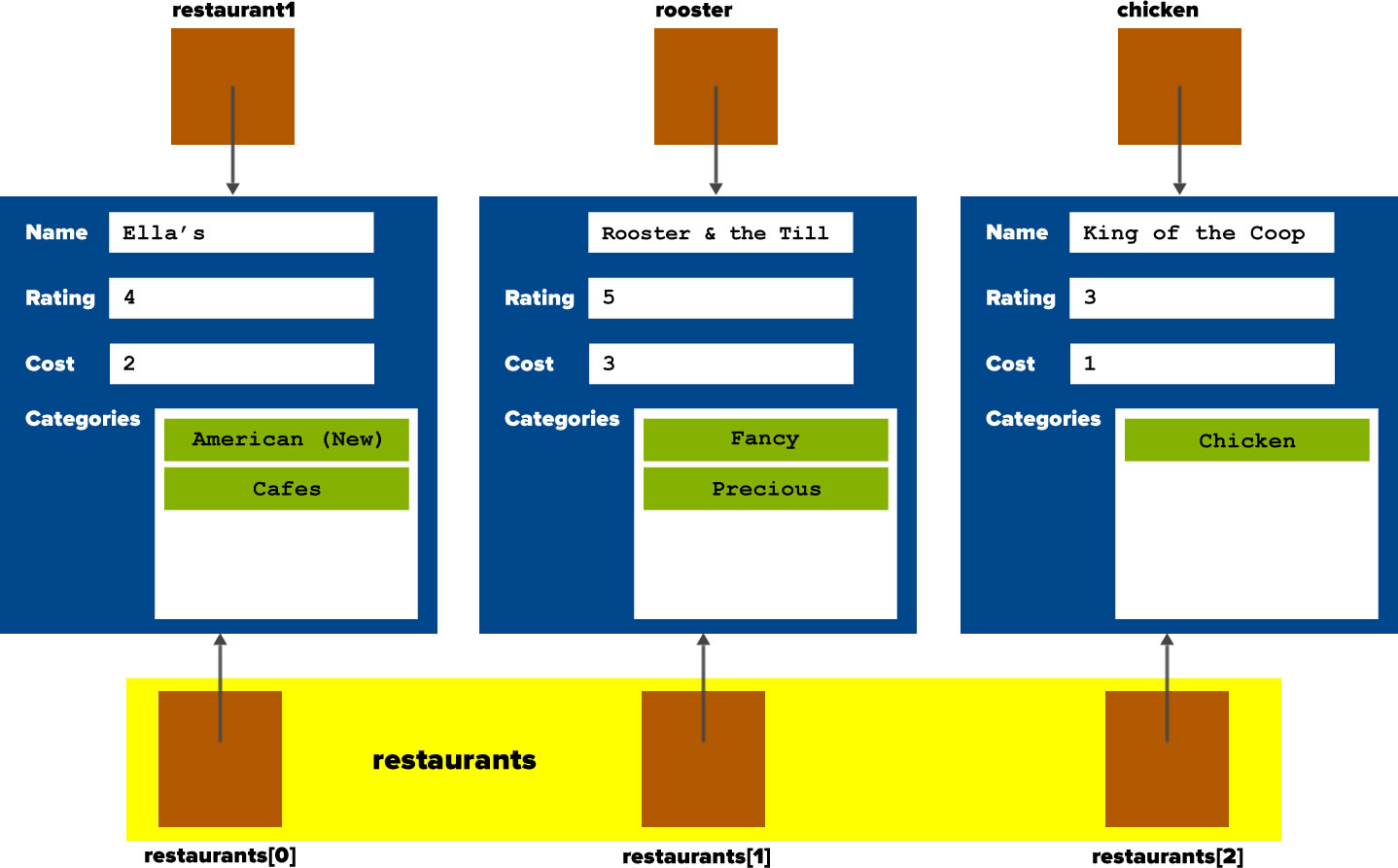
// ### A quick aside
//
// Note that:
//
// * `restaurant1` and `restaurants[0]` both refer to the same object,
// * `rooster` and `restaurants[1]` both refer to the same object, and
// * `chicken` and `restaurants[2]` both refer to the same object.
//
// This will become important later.
//
//
// ### Back to objects and properties
//
// If we wanted to find the name of the first restaurant in `restaurants`, we’d do it like this:
console.log(restaurants[0].name)
// If we wanted to change the rating of the third restaurant to **2**, we’d do it like this:
restaurants[2].rating = 2
console.log(restaurants[2])
// ## Going through the objects in an array, an object at a time
//
// Suppose we wanted to do something with each object in the array, one object at a time. That suggests doing something over and over again, which suggests using a loop.
//
// One way to go through the objects in an array is to set up a `for` loop like this:
for (let index = 0; index < restaurants.length; index++) {
const restaurant = restaurants[index]
console.log(`Here's ${restaurant.name}:`)
console.log(`Its rating is ${restaurant.rating} and its cost is ${restaurant.cost}.`)
}
// There’s another kind of `for` loop that doesn’t require you to set up an index variable — it’s a `for...of` loop. It looks like this:
for (const restaurant of restaurants) {
console.log(`Here's ${restaurant.name}:`)
console.log(`Its rating is ${restaurant.rating} and its cost is ${restaurant.cost}.`)
}
// In this case, the `for...of` loop does the same thing, but with less code and without you having to create an index variable.
// ### Finding a restaurant by name
//
// Suppose we wanted to retrieve a specific restaurant object by name. We could do it by going through the array of restaurants, checking each name for a match.
//
// This sounds like something we’d probably do often, which suggests that we should make it a function. What it does is something repetitive, which suggests using a loop.
//
// We could write this function:
function getRestaurantByName(searchName) {
for (const restaurant of restaurants) {
if (restaurant.name == searchName) {
return restaurant
}
}
return null
}
// In case you were wondering, `null` is a built-in value in JavaScript. It means “no value”, and we’re using it in the `getRestaurantByName()` function to represent the lack of a result. This happens when the we search for a name that doesn’t exist in our list of restaurants.
//
// Let’s put that function to use. Let’s retrieve the restaurant named “King of the Coop”:
console.log(getRestaurantByName("King of the Coop"))
// What happens if we try to get a restaurant by a non-existent name?
console.log(getRestaurantByName("Burger King"))
// ### Making the same change to every restaurant in the list
//
// Suppose we wanted to increase the `cost` property of every restaurant in the list by 1.
//
// This sounds like something that should be generalized into a function that lets you provide a number, and the function then changes the cost of every restaurant in the list by that number. We can even write the function in a way that allows us to either *increase* or *decrease* the cost.
//
// Once again, this is a repetitive process, which suggests that we should use a loop.
//
// Here’s a function that does just that:
function changeCost(changeAmount) {
for (const restaurant of restaurants) {
restaurant.cost += changeAmount
}
}
// Let’s use `changeCost()` to increase the `cost` property of all restaurants by 1:
console.log("Before the increase:")
console.log(restaurants)
changeCost(1)
console.log("After the increase:")
console.log(restaurants)
// We can also use `changeCost()` to descrease the cost, simply by providing it with a negative number:
console.log("Before the decrease:")
console.log(restaurants)
changeCost(-1)
console.log("After the decrease:")
console.log(restaurants)
// ### Selecting only those restaurants that meet some criteria
//
// If you go on Yelp, you can specify “filter criteria” such as “Show me only those restaurants above or below a certain price point” or “Show me only those restaurants with this rating or higher”. We can do that with our array of restaurants.
//
// Let’s try making a function that you give a number, and it returns an array containing only those restaurants with a `cost` of that number or less:
function costAtOrBelow(maximumCost) {
let result = []
for (const restaurant of restaurants) {
if (restaurant.cost <= maximumCost) {
result.push(restaurant)
}
}
return result
}
// Here’s what the function does:
//
// * It accepts a number, `maximumCost`. The function’s job is to return an array that contains only those restaurants whose `cost` property has a value of `maximumCost` or less.
// * It creates an empty array named `result`. This will contain the value that the function will return.
// * It goes through the array of restaurants, one by one, adding any restaurant whose `cost` property has a value of `maximumCost` or less to `result`.
// * At the end of the function, it returns `result`, which should contain only those restaurants `cost` property has a value of `maximumCost` or less. If no such restaurants were found, the function returns an empty array.
//
// Let’s use `costAtOrBelow()` to get an array containing only those restaurants with a `cost` of 2 or less:
console.log(costAtOrBelow(2))
| 16 - Notes for Day 8, part 1.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.4
# language: julia
# name: julia-1.0
# ---
# # Linear Regression
#
# Turing is powerful when applied to complex hierarchical models, but it can also be put to task at common statistical procedures, like [linear regression](https://en.wikipedia.org/wiki/Linear_regression). This tutorial covers how to implement a linear regression model in Turing.
#
# ## Set Up
#
# We begin by importing all the necessary libraries.
# +
# Import Turing and Distributions.
using Turing, Distributions
# Import RDatasets.
using RDatasets
# Import MCMCChains, Plots, and StatPlots for visualizations and diagnostics.
using MCMCChains, Plots, StatsPlots
# Set a seed for reproducibility.
using Random
Random.seed!(0);
# Hide the progress prompt while sampling.
Turing.turnprogress(false);
# -
# We will use the `mtcars` dataset from the [RDatasets](https://github.com/johnmyleswhite/RDatasets.jl) package. `mtcars` contains a variety of statistics on different car models, including their miles per gallon, number of cylinders, and horsepower, among others.
#
# We want to know if we can construct a Bayesian linear regression model to predict the miles per gallon of a car, given the other statistics it has. Lets take a look at the data we have.
# +
# Import the "Default" dataset.
data = RDatasets.dataset("datasets", "mtcars");
# Show the first six rows of the dataset.
first(data, 6)
# -
size(data)
# The next step is to get our data ready for testing. We'll split the `mtcars` dataset into two subsets, one for training our model and one for evaluating our model. Then, we separate the labels we want to learn (`MPG`, in this case) and standardize the datasets by subtracting each column's means and dividing by the standard deviation of that column.
#
# The resulting data is not very familiar looking, but this standardization process helps the sampler converge far easier. We also create a function called `unstandardize`, which returns the standardized values to their original form. We will use this function later on when we make predictions.
# +
# Function to split samples.
function split_data(df, at = 0.70)
r = size(df,1)
index = Int(round(r * at))
train = df[1:index, :]
test = df[(index+1):end, :]
return train, test
end
# A handy helper function to rescale our dataset.
function standardize(x)
return (x .- mean(x, dims=1)) ./ std(x, dims=1), x
end
# Another helper function to unstandardize our datasets.
function unstandardize(x, orig)
return (x .+ mean(orig, dims=1)) .* std(orig, dims=1)
end
# Remove the model column.
select!(data, Not(:Model))
# Standardize our dataset.
(std_data, data_arr) = standardize(Matrix(data))
# Split our dataset 70%/30% into training/test sets.
train, test = split_data(std_data, 0.7)
# Save dataframe versions of our dataset.
train_cut = DataFrame(train, names(data))
test_cut = DataFrame(test, names(data))
# Create our labels. These are the values we are trying to predict.
train_label = train_cut[:, :MPG]
test_label = test_cut[:, :MPG]
# Get the list of columns to keep.
remove_names = filter(x->!in(x, [:MPG, :Model]), names(data))
# Filter the test and train sets.
train = Matrix(train_cut[:,remove_names]);
test = Matrix(test_cut[:,remove_names]);
# -
# ## Model Specification
#
# In a traditional frequentist model using [OLS](https://en.wikipedia.org/wiki/Ordinary_least_squares), our model might look like:
#
# $$
# MPG_i = \alpha + \boldsymbol{\beta}^T\boldsymbol{X_i}
# $$
#
# where $\boldsymbol{\beta}$ is a vector of coefficients and $\boldsymbol{X}$ is a vector of inputs for observation $i$. The Bayesian model we are more concerned with is the following:
#
# $$
# MPG_i \sim \mathcal{N}(\alpha + \boldsymbol{\beta}^T\boldsymbol{X_i}, \sigma^2)
# $$
#
# where $\alpha$ is an intercept term common to all observations, $\boldsymbol{\beta}$ is a coefficient vector, $\boldsymbol{X_i}$ is the observed data for car $i$, and $\sigma^2$ is a common variance term.
#
# For $\sigma^2$, we assign a prior of `TruncatedNormal(0,100,0,Inf)`. This is consistent with [<NAME>'s recommendations](http://www.stat.columbia.edu/~gelman/research/published/taumain.pdf) on noninformative priors for variance. The intercept term ($\alpha$) is assumed to be normally distributed with a mean of zero and a variance of three. This represents our assumptions that miles per gallon can be explained mostly by our assorted variables, but a high variance term indicates our uncertainty about that. Each coefficient is assumed to be normally distributed with a mean of zero and a variance of 10. We do not know that our coefficients are different from zero, and we don't know which ones are likely to be the most important, so the variance term is quite high. The syntax `::Type{T}=Vector{Float64}` allows us to maintain type stability in our model -- for more information, please review the [performance tips](https://turing.ml/dev/docs/using-turing/performancetips#make-your-model-type-stable). Lastly, each observation $y_i$ is distributed according to the calculated `mu` term given by $\alpha + \boldsymbol{\beta}^T\boldsymbol{X_i}$.
# Bayesian linear regression.
@model linear_regression(x, y, n_obs, n_vars, ::Type{T}=Vector{Float64}) where {T} = begin
# Set variance prior.
σ₂ ~ truncated(Normal(0,100), 0, Inf)
# Set intercept prior.
intercept ~ Normal(0, 3)
# Set the priors on our coefficients.
coefficients = T(undef, n_vars)
for i in 1:n_vars
coefficients[i] ~ Normal(0, 10)
end
# Calculate all the mu terms.
mu = intercept .+ x * coefficients
y ~ MvNormal(mu, σ₂)
end;
# With our model specified, we can call the sampler. We will use the No U-Turn Sampler ([NUTS](http://turing.ml/docs/library/#-turingnuts--type)) here.
n_obs, n_vars = size(train)
model = linear_regression(train, train_label, n_obs, n_vars)
chain = sample(model, NUTS(0.65), 3000);
# As a visual check to confirm that our coefficients have converged, we show the densities and trace plots for our parameters using the `plot` functionality.
plot(chain)
# It looks like each of our parameters has converged. We can check our numerical esimates using `describe(chain)`, as below.
describe(chain)
# ## Comparing to OLS
#
# A satisfactory test of our model is to evaluate how well it predicts. Importantly, we want to compare our model to existing tools like OLS. The code below uses the [GLM.jl]() package to generate a traditional OLS multiple regression model on the same data as our probabalistic model.
# +
# Import the GLM package.
using GLM
# Perform multiple regression OLS.
ols = lm(@formula(MPG ~ Cyl + Disp + HP + DRat + WT + QSec + VS + AM + Gear + Carb), train_cut)
# Store our predictions in the original dataframe.
train_cut.OLSPrediction = unstandardize(GLM.predict(ols), data.MPG);
test_cut.OLSPrediction = unstandardize(GLM.predict(ols, test_cut), data.MPG);
# -
# The function below accepts a chain and an input matrix and calculates predictions. We use the mean observation of each parameter in the model starting with sample 200, which is where the warm-up period for the NUTS sampler ended.
# Make a prediction given an input vector.
function prediction(chain, x)
p = get_params(chain[200:end, :, :])
α = mean(p.intercept)
β = collect(mean.(p.coefficients))
return α .+ x * β
end
# When we make predictions, we unstandardize them so they're more understandable. We also add them to the original dataframes so they can be placed in context.
# +
# Calculate the predictions for the training and testing sets.
train_cut.BayesPredictions = unstandardize(prediction(chain, train), data.MPG);
test_cut.BayesPredictions = unstandardize(prediction(chain, test), data.MPG);
# Unstandardize the dependent variable.
train_cut.MPG = unstandardize(train_cut.MPG, data.MPG);
test_cut.MPG = unstandardize(test_cut.MPG, data.MPG);
# Show the first side rows of the modified dataframe.
first(test_cut, 6)
# -
# Now let's evaluate the loss for each method, and each prediction set. We will use sum of squared error function to evaluate loss, given by
#
# $$
# \text{SSE} = \sum{(y_i - \hat{y_i})^2}
# $$
#
# where $y_i$ is the actual value (true MPG) and $\hat{y_i}$ is the predicted value using either OLS or Bayesian linear regression. A lower SSE indicates a closer fit to the data.
# +
bayes_loss1 = sum((train_cut.BayesPredictions - train_cut.MPG).^2)
ols_loss1 = sum((train_cut.OLSPrediction - train_cut.MPG).^2)
bayes_loss2 = sum((test_cut.BayesPredictions - test_cut.MPG).^2)
ols_loss2 = sum((test_cut.OLSPrediction - test_cut.MPG).^2)
println("Training set:
Bayes loss: $bayes_loss1
OLS loss: $ols_loss1
Test set:
Bayes loss: $bayes_loss2
OLS loss: $ols_loss2")
# -
# As we can see above, OLS and our Bayesian model fit our training set about the same. This is to be expected, given that it is our training set. However, the Bayesian linear regression model is less able to predict out of sample -- this is likely due to our selection of priors, and that fact that point estimates were used to forecast instead of the true posteriors.
| 5_LinearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, random, datetime, pickle
from datetime import datetime
import numpy as np
import pandas as pd
import scipy.sparse as sp
import tensorflow as tf
# + code_folding=[]
# reader
class Reader(object):
def read(self, data_path):
handled_path = data_path + 'basic_trainer_saved.pkl'
if os.path.exists(handled_path):
print('load file from local')
(self._entity_num, self._relation_num, self._relation_num_for_eval, self._train_data, self._test_data,
self._valid_data) = pickle.load(open(handled_path, 'rb'))
else:
self.read_data()
self.merge_id()
self.add_reverse()
self.reindex_kb()
self.gen_t_label()
print('start save dfs')
saved = (
self._entity_num, self._relation_num, self._relation_num_for_eval, self._train_data, self._test_data,
self._valid_data)
pickle.dump(saved, open(handled_path, 'wb'))
self.gen_filter_mat()
self._ent_num = self._entity_num
self._rel_num = self._relation_num
self._ent_mapping = pd.DataFrame({'kb_1':{}, 'kb_2':{}})
self._rel_mapping = pd.DataFrame({'kb_1':{}, 'kb_2':{}})
self._ent_testing = pd.DataFrame({'kb_1':{}, 'kb_2':{}})
self._rel_testing = pd.DataFrame({'kb_1':{}, 'kb_2':{}})
self._kb = self._train_data
return
def read_data(self):
pass
def merge_id(self):
self._train_data['h_id'] = self._e_id[self._train_data.h].values
self._train_data['r_id'] = self._r_id[self._train_data.r].values
self._train_data['t_id'] = self._e_id[self._train_data.t].values
self._test_data['h_id'] = self._e_id[self._test_data.h].values
self._test_data['r_id'] = self._r_id[self._test_data.r].values
self._test_data['t_id'] = self._e_id[self._test_data.t].values
self._valid_data['h_id'] = self._e_id[self._valid_data.h].values
self._valid_data['r_id'] = self._r_id[self._valid_data.r].values
self._valid_data['t_id'] = self._e_id[self._valid_data.t].values
def gen_t_label(self):
full = pd.concat([self._train_data, self._test_data, self._valid_data], ignore_index=True)
f_t_labels = full['t_id'].groupby([full['h_id'], full['r_id']]).apply(lambda x: pd.unique(x.values))
f_t_labels.name = 't_label'
self._test_data = self._test_data.join(f_t_labels, on=['h_id', 'r_id'])
self._valid_data = self._valid_data.join(f_t_labels, on=['h_id', 'r_id'])
def add_reverse(self):
def add_reverse_for_data(data):
reversed_data = data.rename(columns={'h_id': 't_id', 't_id': 'h_id'})
reversed_data.r_id += self._relation_num
data = pd.concat(([data, reversed_data]), ignore_index=True)
return data
self._train_data = add_reverse_for_data(self._train_data)
self._test_data = add_reverse_for_data(self._test_data)
self._valid_data = add_reverse_for_data(self._valid_data)
self._relation_num_for_eval = self._relation_num
self._relation_num *= 2
# print (self._relation_num, self._relation_num_for_eval)
def reindex_kb(self):
train_data = self._train_data
test_data = self._test_data
valid_data = self._valid_data
eids = pd.concat([train_data.h_id, train_data.t_id,], ignore_index=True)
tv_eids = np.unique(pd.concat([test_data.h_id, test_data.t_id, valid_data.t_id, valid_data.h_id]))
not_train_eids = tv_eids[~np.in1d(tv_eids, eids)]
rids = pd.concat([train_data.r_id,],ignore_index=True)
def gen_map(eids, rids):
e_num = eids.groupby(eids.values).size().sort_values()[::-1]
not_train = pd.Series(np.zeros_like(not_train_eids), index=not_train_eids)
e_num = pd.concat([e_num, not_train])
r_num = rids.groupby(rids.values).size().sort_values()[::-1]
e_map = pd.Series(range(e_num.shape[0]), index=e_num.index)
r_map = pd.Series(range(r_num.shape[0]), index=r_num.index)
return e_map, r_map
def remap_kb(kb, e_map, r_map):
kb.loc[:, 'h_id'] = e_map.loc[kb.h_id.values].values
kb.loc[:, 'r_id'] = r_map.loc[kb.r_id.values].values
kb.loc[:, 't_id'] = e_map.loc[kb.t_id.values].values
return kb
def remap_id(s, rm):
s = rm.loc[s.values].values
return s
e_map, r_map = gen_map(eids, rids)
self._e_map, self._r_map = e_map, r_map
self._train_data = remap_kb(train_data, e_map, r_map)
self._valid_data = remap_kb(self._valid_data, e_map, r_map)
self._test_data = remap_kb(self._test_data, e_map, r_map)
self._e_id = remap_id(self._e_id, e_map)
self._r_id = remap_id(self._r_id, r_map)
return not_train_eids
def in2d(self, arr1, arr2):
"""Generalisation of numpy.in1d to 2D arrays"""
assert arr1.dtype == arr2.dtype
arr1_view = np.ascontiguousarray(arr1).view(np.dtype((np.void,
arr1.dtype.itemsize * arr1.shape[1])))
arr2_view = np.ascontiguousarray(arr2).view(np.dtype((np.void,
arr2.dtype.itemsize * arr2.shape[1])))
intersected = np.in1d(arr1_view, arr2_view)
return intersected.view(np.bool).reshape(-1)
def gen_filter_mat(self):
def gen_filter_vector(r):
v = np.ones(self._entity_num)
v[r] = -1
return v
print('start gen filter mat')
self._tail_valid_filter_mat = np.stack(self._valid_data.t_label.apply(gen_filter_vector).values)
self._tail_test_filter_mat = np.stack(self._test_data.t_label.apply(gen_filter_vector).values)
def gen_label_mat_for_train(self):
def gen_train_relation_label_vac(r):
c = pd.value_counts(r)
values = 1. * c.values / c.sum()
return np.stack([c.index, values], axis=1)
def gen_train_entity_label_vac(r):
indices = np.stack([r.label_id.values, r.values], axis=1)
values = np.ones_like(r.values, dtype=np.int)
return tf.SparseTensor(indices=indices, values=values, dense_shape=[1, self._entity_num])
tr = self._train_data
print('start gen t_label')
labels = tr['t_id'].groupby([tr['h_id'], tr['r_id']]).size()
labels = pd.Series(range(labels.shape[0]), index=labels.index)
labels.name = 'label_id'
tr = tr.join(labels, on=['h_id', 'r_id'])
self._train_data = tr
sp_tr = tf.SparseTensor(tr[['label_id', 't_id']].values, np.ones([len(tr)], dtype=np.float32), dense_shape=[len(tr), self._entity_num])
self._label_indices, self._label_values = sp_tr.indices[:], sp_tr.values[:]
class FreeBaseReader(Reader):
def read_data(self):
path = self._options.data_path
tr = pd.read_csv(path + 'train.txt', header=None, sep='\t', names=['h', 't', 'r'])
te = pd.read_csv(path + 'test.txt', header=None, sep='\t', names=['h', 't', 'r'])
val = pd.read_csv(path + 'valid.txt', header=None, sep='\t', names=['h', 't', 'r'])
e_id = pd.read_csv(path + 'entity2id.txt', header=None, sep='\t', names=['e', 'eid'])
e_id = pd.Series(e_id.eid.values, index=e_id.e.values)
r_id = pd.read_csv(path + 'relation2id.txt', header=None, sep='\t', names=['r', 'rid'])
r_id = pd.Series(r_id.rid.values, index=r_id.r.values)
self._entity_num = e_id.shape[0]
self._relation_num = r_id.shape[0]
self._train_data = tr
self._test_data = te
self._valid_data = val
self._e_id, self._r_id = e_id, r_id
class WordNetReader(Reader):
def read_data(self):
path = self._options.data_path
tr = pd.read_csv(path+'train.txt', header=None, sep='\t', names=['h', 'r', 't'])
te = pd.read_csv(path + 'test.txt', header=None, sep='\t', names=['h', 'r', 't'])
val = pd.read_csv(path + 'valid.txt', header=None, sep='\t', names=['h', 'r', 't'])
r_list = pd.unique(pd.concat([tr.r, te.r, val.r]))
r_list = pd.Series(r_list, index=np.arange(r_list.shape[0]))
e_list = pd.unique(pd.concat([tr.h, te.h, val.h, tr.t, te.t, val.t, ]))
e_list = pd.Series(e_list, index=np.arange(e_list.shape[0]))
e_id = pd.Series(e_list.index, index=e_list.values)
r_id = pd.Series(r_list.index, index=r_list.values)
self._entity_num = e_id.shape[0]
self._relation_num = r_id.shape[0]
self._train_data = tr
self._test_data = te
self._valid_data = val
self._e_id, self._r_id = e_id, r_id
# + code_folding=[]
# path sampler
class BasicSampler(object):
def sample_paths(self, repeat_times=2):
opts = self._options
kb = self._kb.copy()
kb = kb[['h_id', 'r_id', 't_id']]
# sampling triples with the h_id-(r_id,t_id) form.
rtlist = np.unique(kb[['r_id', 't_id']].values, axis=0)
rtdf = pd.DataFrame(rtlist, columns=['r_id', 't_id'])
rtdf = rtdf.reset_index().rename({'index': 'tail_id'}, axis='columns')
rtkb = kb.merge(
rtdf, left_on=['r_id', 't_id'], right_on=['r_id', 't_id'])
htail = np.unique(rtkb[['h_id', 'tail_id']].values, axis=0)
htailmat = csr_matrix((np.ones(len(htail)), (htail[:, 0], htail[:, 1])),
shape=(model._ent_num, rtlist.shape[0]))
# calulate corss-KG bias at first
em = pd.concat(
[model._ent_mapping.kb_1, model._ent_mapping.kb_2]).values
rtkb['across'] = rtkb.t_id.isin(em)
rtkb.loc[rtkb.across, 'across'] = opts.beta
rtkb.loc[rtkb.across == 0, 'across'] = 1-opts.beta
rtailkb = rtkb[['h_id', 't_id', 'tail_id', 'across']]
def gen_tail_dict(x):
return x.tail_id.values, x.across.values / x.across.sum()
rtailkb = rtailkb.groupby('h_id').apply(gen_tail_dict)
rtailkb = pd.DataFrame({'tails': rtailkb})
# start sampling
hrt = np.repeat(kb.values, repeat_times, axis=0)
# for initial triples
def perform_random(x):
return np.random.choice(x.tails[0], 1, p=x.tails[1].astype(np.float))
# else
def perform_random2(x):
# calculate depth bias
pre_c = htailmat[np.repeat(x.pre, x.tails[0].shape[0]), x.tails[0]]
pre_c[pre_c == 0] = opts.alpha
pre_c[pre_c == 1] = 1-opts.alpha
p = x.tails[1].astype(np.float).reshape(
[-1, ]) * pre_c.A.reshape([-1, ])
p = p / p.sum()
return np.random.choice(x.tails[0], 1, p=p)
rt_x = rtailkb.loc[hrt[:, 2]].apply(perform_random, axis=1)
rt_x = rtlist[np.concatenate(rt_x.values)]
rts = [hrt, rt_x]
c_length = 5
while(c_length < opts.max_length):
curr = rtailkb.loc[rt_x[:, 1]]
curr.loc[:, 'pre'] = hrt[:, 0]
rt_x = curr.apply(perform_random2, axis=1)
rt_x = rtlist[np.concatenate(rt_x.values)]
rts.append(rt_x)
c_length += 2
data = np.concatenate(rts, axis=1)
data = pd.DataFrame(data)
self._train_data = data
data.to_csv('%spaths_%.1f_%.1f' % (opts.data_path, opts.alpha, opts.beta))
# + code_folding=[2, 7, 50, 60, 94, 111]
# model
class RSN4KGC(FreeBaseReader):
def __init__(self, options, session):
self._options = options
self._session = session
def init_variables(self):
options = self._options
hidden_size = options.hidden_size
self._entity_embedding = tf.get_variable(
'entity_embedding',
[self._ent_num, hidden_size],
initializer=tf.contrib.layers.xavier_initializer()
)
self._relation_embedding = tf.get_variable(
'relation_embedding',
[self._rel_num, hidden_size],
initializer=tf.contrib.layers.xavier_initializer()
)
self._rel_w = tf.get_variable(
"relation_softmax_w",
[self._rel_num, hidden_size],
initializer=tf.contrib.layers.xavier_initializer()
)
self._rel_b = tf.get_variable(
"relation_softmax_b",
[self._rel_num],
initializer=tf.constant_initializer(0)
)
self._ent_w = tf.get_variable(
"entity_softmax_w",
[self._ent_num, hidden_size],
initializer=tf.contrib.layers.xavier_initializer()
)
self._ent_b = tf.get_variable(
"entity_softmax_b",
[self._ent_num],
initializer=tf.constant_initializer(0)
)
self._lr = tf.Variable(options.learning_rate, trainable=False)
self._optimizer = tf.train.AdamOptimizer(options.learning_rate)
def bn(self, inputs, is_train=True, reuse=True):
return tf.contrib.layers.batch_norm(inputs,
center=True,
scale=True,
is_training=is_train,
reuse=reuse,
scope='bn',
data_format='NCHW'
)
def lstm_cell(self, drop=True, keep_prob=0.5, num_layers=2, hidden_size=None):
if not hidden_size:
hidden_size = self._options.hidden_size
def basic_lstm_cell():
return tf.contrib.rnn.LSTMCell(
num_units=hidden_size,
initializer=tf.orthogonal_initializer,
forget_bias=1,
reuse=tf.get_variable_scope().reuse,
activation=tf.identity
)
def drop_cell():
return tf.contrib.rnn.DropoutWrapper(
basic_lstm_cell(),
output_keep_prob=keep_prob
)
if drop:
gen_cell = drop_cell
else:
gen_cell = basic_lstm_cell
if num_layers==0:
return gen_cell()
cell = tf.contrib.rnn.MultiRNNCell(
[gen_cell() for _ in range(num_layers)],
state_is_tuple=True,
)
return cell
def sampled_loss(self, inputs, labels, w, b, weight=1, is_entity=False):
num_sampled = min(self._options.num_samples, w.shape[0]//3)
labels = tf.reshape(labels, [-1, 1])
losses = tf.nn.sampled_softmax_loss(
weights=w,
biases=b,
labels=labels,
inputs=tf.reshape(inputs, [-1, int(w.shape[1])]),
num_sampled=num_sampled,
num_classes=w.shape[0],
partition_strategy='div',
)
return losses * weight
def logits(self, input, predict_relation=True):
if not predict_relation:
w = self._ent_w
b = self._ent_b
else:
w = self._rel_w
b = self._rel_b
return tf.nn.bias_add(tf.matmul(input, tf.transpose(w)), b)
def sample(self, data):
choices = np.random.choice(len(data), size=len(data), replace=False)
return data.iloc[choices]
def padding_data(self, data):
padding_num = self._options.batch_size - len(data) % self._options.batch_size
data = np.concatenate([data, np.zeros((padding_num, data.shape[1]), dtype=np.int32)])
return data, padding_num
# + code_folding=[0]
# build tensorflow graph
# build an RSN of length l
def build_sub_graph(self, length=15, reuse=False):
options = self._options
hidden_size = options.hidden_size
batch_size = options.batch_size
seq = tf.placeholder(
tf.int32, [batch_size, length], name='seq'+str(length))
e_em, r_em = self._entity_embedding, self._relation_embedding
# seperately read, and then recover the order
ent = seq[:, :-1:2]
rel = seq[:, 1::2]
ent_em = tf.nn.embedding_lookup(e_em, ent)
rel_em = tf.nn.embedding_lookup(r_em, rel)
em_seq = []
for i in range(length-1):
if i % 2 == 0:
em_seq.append(ent_em[:, i//2])
else:
em_seq.append(rel_em[:, i//2])
# seperately bn
with tf.variable_scope('input_bn'):
if not reuse:
bn_em_seq = [tf.reshape(self.bn(em_seq[i], reuse=(
i is not 0)), [-1, 1, hidden_size]) for i in range(length-1)]
else:
bn_em_seq = [tf.reshape(
self.bn(em_seq[i], reuse=True), [-1, 1, hidden_size]) for i in range(length-1)]
bn_em_seq = tf.concat(bn_em_seq, axis=1)
ent_bn_em = bn_em_seq[:, ::2]
with tf.variable_scope('rnn', reuse=reuse):
cell = self.lstm_cell(True, options.keep_prob, options.num_layers)
outputs, state = tf.nn.dynamic_rnn(cell, bn_em_seq, dtype=tf.float32)
rel_outputs = outputs[:, fd00:c2b6:b24b:be67:2827:688d:e6a1:6a3b, :]
outputs = [outputs[:, i, :] for i in range(length-1)]
ent_outputs = outputs[::2]
# RSN
with tf.variable_scope('resnet', reuse=reuse):
res_rel_outputs = tf.contrib.layers.fully_connected(rel_outputs, hidden_size, biases_initializer=None, activation_fn=None) +\
tf.contrib.layers.fully_connected(
ent_bn_em, hidden_size, biases_initializer=None, activation_fn=None)
# recover the order
res_rel_outputs = [res_rel_outputs[:, i, :] for i in range((length-1)//2)]
outputs = []
for i in range(length-1):
if i % 2 == 0:
outputs.append(ent_outputs[i//2])
else:
outputs.append(res_rel_outputs[i//2])
# output bn
with tf.variable_scope('output_bn'):
if reuse:
bn_outputs = [tf.reshape(
self.bn(outputs[i], reuse=True), [-1, 1, hidden_size]) for i in range(length-1)]
else:
bn_outputs = [tf.reshape(self.bn(outputs[i], reuse=(
i is not 0)), [-1, 1, hidden_size]) for i in range(length-1)]
def cal_loss(bn_outputs, seq):
losses = []
decay = 0.8
for i, output in enumerate(bn_outputs):
if i % 2 == 0:
losses.append(self.sampled_loss(
output, seq[:, i+1], self._rel_w, self._rel_b, weight=decay**(0), is_entity=i))
else:
losses.append(self.sampled_loss(
output, seq[:, i+1], self._ent_w, self._ent_b, weight=decay**(0), is_entity=i))
losses = tf.stack(losses, axis=1)
return losses
seq_loss = cal_loss(bn_outputs, seq)
losses = tf.reduce_sum(seq_loss) / batch_size
return losses, seq
# build the main graph
def build_graph(self):
options = self._options
loss, seq = build_sub_graph(self, length=options.max_length, reuse=False)
tvars = tf.trainable_variables()
grads, _ = tf.clip_by_global_norm(tf.gradients(loss, tvars), 2.0)
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
train_op = self._optimizer.apply_gradients(
zip(grads, tvars),
global_step=tf.train.get_or_create_global_step()
)
self._seq, self._loss, self._train_op = seq, loss, train_op
# + code_folding=[2]
# training procedure
def seq_train(self, data, choices=None, epoch=None):
opts = self._options
# shuffle data
choices = np.random.choice(len(data), size=len(data), replace=True)
batch_size = opts.batch_size
num_batch = len(data) // batch_size
fetches = {
'loss': self._loss,
'train_op': self._train_op
}
losses = 0
for i in range(num_batch):
one_batch_choices = choices[i * batch_size : (i + 1) * batch_size]
one_batch_data = data.iloc[one_batch_choices]
feed_dict = {}
seq = one_batch_data.values[:, :opts.max_length]
feed_dict[self._seq] = seq
vals = self._session.run(fetches, feed_dict)
del one_batch_data
loss = vals['loss']
losses += loss
print('\r%i/%i, batch_loss:%f' % (i, num_batch, loss), end='')
self._last_mean_loss = losses / num_batch
return self._last_mean_loss
# + code_folding=[]
#build the graph for entity prediction (KG completion)
def build_eval_ep(self, length=3, reuse=True):
options = self._options
hidden_size = options.hidden_size
batch_size = options.batch_size
seq = tf.placeholder(tf.int32, [batch_size, length], name='eval_seq')
e_em, r_em = self._entity_embedding, self._relation_embedding
ent = seq[:, :-1:2]
rel = seq[:, 1::2]
ent_em = tf.nn.embedding_lookup(e_em, ent)
rel_em = tf.nn.embedding_lookup(r_em, rel)
em_seq = []
for i in range(length-1):
if i % 2 == 0:
em_seq.append(ent_em[:, i//2])
else:
em_seq.append(rel_em[:, i//2])
with tf.variable_scope('input_bn', reuse=reuse):
if not reuse:
bn_em_seq = [tf.reshape(self.bn(em_seq[i], reuse=(
i is not 0)), [-1, 1, hidden_size]) for i in range(length-1)]
else:
bn_em_seq = [tf.reshape(
self.bn(em_seq[i], reuse=True), [-1, 1, hidden_size]) for i in range(length-1)]
bn_em_seq = tf.concat(bn_em_seq, axis=1)
ent_bn_em = bn_em_seq[:, ::2]
with tf.variable_scope('rnn', reuse=reuse):
cell = self.lstm_cell(True, options.keep_prob, options.num_layers)
outputs, state = tf.nn.dynamic_rnn(cell, bn_em_seq, dtype=tf.float32)
rel_outputs = outputs[:, 1::2, :]
outputs = [outputs[:, i, :] for i in range(length-1)]
ent_outputs = outputs[::2]
with tf.variable_scope('resnet', reuse=reuse):
res_rel_outputs = tf.contrib.layers.fully_connected(rel_outputs, hidden_size, biases_initializer=None, activation_fn=None) +\
tf.contrib.layers.fully_connected(ent_bn_em, hidden_size, biases_initializer=None, activation_fn=None)
res_rel_outputs = [res_rel_outputs[:, i, :] for i in range((length-1)//2)]
outputs=[]
for i in range(length-1):
if i % 2==0:
outputs.append(ent_outputs[i//2])
else:
outputs.append(res_rel_outputs[i//2])
with tf.variable_scope('output_bn', reuse=reuse):
if reuse:
bn_outputs = [tf.reshape(
self.bn(outputs[i], reuse=True), [-1, hidden_size]) for i in range(length-1)]
else:
bn_outputs = [tf.reshape(self.bn(outputs[i], reuse=(
i is not 0)), [-1, hidden_size]) for i in range(length-1)]
logits = self.logits(bn_outputs[1], predict_relation=False)
probs = tf.nn.softmax(logits)
self._eval_seq = seq
self._entity_probs = probs
# + code_folding=[2]
#evaluate the performance on KG completion (entity prediction)
def eval_entity_prediction(model, data, filter_mat, method='min', return_ranks=False, return_probs=False, return_label_probs=False):
options = model._options
batch_size = options.batch_size
label = data[:, 2]
data, padding_num = model.padding_data(data)
num_batch = len(data) // batch_size
eval_seq, fectch_entity_probs = model._eval_seq, model._entity_probs
probs = []
for i in range(num_batch):
feed_dict = {}
feed_dict[eval_seq] = data[i * batch_size:(i + 1) * batch_size]
probs.append(sess.run(fectch_entity_probs, feed_dict))
probs = np.concatenate(probs)[:len(data) - padding_num]
if return_label_probs:
return probs[range(len(label)), label]
if return_probs:
return probs
filter_probs = probs * filter_mat
filter_probs[range(len(label)), label] = probs[range(len(label)), label]
filter_ranks = cal_ranks(filter_probs, method=method, label=label)
if return_ranks:
return filter_ranks
_, f_h_1, _ = cal_performance(filter_ranks, top=1)
f_m_r, f_h_10, f_mrr = cal_performance(filter_ranks)
return (f_h_1, f_h_10, f_mrr, f_m_r)
# + code_folding=[]
# some tool functions
def cal_ranks(probs, method, label):
if method == 'min':
probs = probs - probs[range(len(label)), label].reshape(len(probs), 1)
ranks = (probs > 0).sum(axis=1) + 1
else:
ranks = pd.DataFrame(probs).rank(axis=1, ascending=False, method=method)
ranks = ranks.values[range(len(label)), label]
return ranks
def cal_performance(ranks, top=10):
m_r = sum(ranks) * 1.0 / len(ranks)
h_10 = sum(ranks <= top) * 1.0 / len(ranks)
mrr = (1. / ranks).sum() / len(ranks)
return m_r, h_10, mrr
def padding_data(data, options, batch_size):
padding_num = batch_size - len(data) % batch_size
data = pd.concat([data, pd.DataFrame(np.zeros((padding_num, data.shape[1])), dtype=np.int32, columns=data.columns)],ignore_index=True, axis=0)
return data, padding_num
def in2d(arr1, arr2):
"""Generalisation of numpy.in1d to 2D arrays"""
assert arr1.dtype == arr2.dtype
arr1_view = np.ascontiguousarray(arr1).view(np.dtype((np.void,
arr1.dtype.itemsize * arr1.shape[1])))
arr2_view = np.ascontiguousarray(arr2).view(np.dtype((np.void,
arr2.dtype.itemsize * arr2.shape[1])))
intersected = np.in1d(arr1_view, arr2_view)
return intersected.view(np.bool).reshape(-1)
def write_to_log(path, content):
with open(path, 'a+') as f:
print(content, file=f)
# +
#parameter settings
class Options(object):
pass
opts = Options()
opts.hidden_size = 256
opts.num_layers = 2
opts.batch_size = 2048
opts.learning_rate = 0.0001 # for FB15K-237, the learning rate should decrease to 0.00001
opts.num_samples = 2048*5
opts.keep_prob = 0.5
opts.max_length=7
opts.alpha = 0.7
opts.beta = 0.5
opts.data_path = 'data/FB15k/'
opts.log_file_path = 'logs/%s%dl_%s.log' % (opts.data_path.replace(
'/', '-'), opts.max_length, datetime.now().strftime('%y-%m-%d-%H-%M'))
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
# +
#initial model
sess = tf.InteractiveSession(config=config)
model = RSN4KGC(options=opts, session=sess)
model.read(data_path=model._options.data_path)
model.init_variables()
sequence_datapath = '%spaths_%.1f_%.1f' % (
model._options.data_path, model._options.alpha, model._options.beta)
if not os.path.exists(sequence_datapath):
print('start to sample paths')
model.sample_paths()
train_data = model._train_data
else:
print('load existing training sequences')
train_data = pd.read_csv(sequence_datapath, index_col=0)
# build tensorflow graph and init all tensors
build_graph(model)
build_eval_ep(model)
tf.global_variables_initializer().run()
# +
# initial training settings
write_to_log(opts.log_file_path, opts.__dict__)
epoch = 0
last_mean_loss=0
max_hits1, times, max_times = 0, 0, 3
# we transform the subject entity prediction (?, r, o) to (o, r-, ?) for convenience
test_data = model._test_data[['h_id', 'r_id', 't_id']].values
filter_mat = model._tail_test_filter_mat
valid_data = model._valid_data[['h_id', 'r_id', 't_id']].values
vfilter_mat = model._tail_valid_filter_mat
# + code_folding=[]
r = eval_entity_prediction(model, data=valid_data, filter_mat=vfilter_mat)
msg = 'epoch:%i, Hits@1:%.3f, Hits@10:%.3f, MRR:%.3f, MR:%.3f, mean_loss:%.3f' % (epoch, r[0],r[1],r[2],r[3], last_mean_loss)
print('\n'+msg)
write_to_log(opts.log_file_path, msg)
for i in range(epoch, 200):
last_mean_loss = seq_train(model, train_data)
epoch += 1
# evaluation
if i % 5 ==0:
r = eval_entity_prediction(model, data=valid_data, filter_mat=vfilter_mat)
msg = 'epoch:%i, Hits@1:%.3f, Hits@10:%.3f, MRR:%.3f, MR:%.3f, mean_loss:%.3f' % (i+1, r[0],r[1],r[2],r[3], last_mean_loss)
print('\n'+msg)
write_to_log(opts.log_file_path, msg)
hits1 = r[0]
# early stop
if hits1 > max_hits1:
max_hits1 = hits1
times = 0
else:
times += 1
if times >= max_times:
break
#evaluation on testing data
r = eval_entity_prediction(model, data=test_data, filter_mat=filter_mat, method='average')
msg = 'epoch:%i, Hits@1:%.3f, Hits@10:%.3f, MRR:%.3f, MR:%.3f, mean_loss:%.3f' % (epoch, r[0],r[1],r[2],r[3], last_mean_loss)
print('\n'+msg)
write_to_log(opts.log_file_path, msg)
| RSN4KGC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 한국환경공단 대기오염 API를 이용한 미세먼지 데이터 가져오기
import requests
from bs4 import BeautifulSoup
import pickle
import googlemaps
from gensim.models import Word2Vec
# +
#미세먼지 및 초미세먼지 농도별 등급 판정
def grade(num) :
try :
num = int(num)
if num >=0 and num <= 30 :
result = "좋음"
elif num > 30 and num <= 80 :
result = "보통"
elif num > 80 and num <= 150 :
result = "나쁨"
else :
result = "매우나쁨"
except :
result = "-"
return result
def nano_grade(num) :
try :
num = int(num)
if num >=0 and num <= 15 :
result = "좋음"
elif num > 15 and num <= 35 :
result = "보통"
elif num > 35 and num <= 75 :
result = "나쁨"
else :
result = "매우나쁨"
except :
result = "-"
return result
# -
# ---------
# #### 환경오염공단 API 키 불러오기
with open('secret/get_tm_coor_key.txt', 'rb') as f :
get_tm_coor_key = pickle.load(f)
with open("secret/google_secret_key.txt", "rb") as f :
google_secret_key = pickle.load(f)
# ## 1. 측정소 정보 조회
# - 대기질 측정소 정보를 조회하기 위해 TM 좌표 기반의 가까운 측정소 및 측정소 목록과 측정소의 정보를 조회할 수 있음.
# - 입력값 : 미세먼지 농도를 알고 싶은 위치의 주소명
# - 출력값 : 위치에 대한 TM좌표
# - 환경공단의 API가 인지하지 못하는 지역위치 값 입력시
# > 구글 API로 위치 재조정
model_seoul = Word2Vec.load("model_gwangju")
model_seoul.wv.most_similar("치평동")
def get_tm_coor(loc) :
#가독성을 위한 구글 주소 정보 속 긴 키워드들을 정리
format_add = "formatted_address"
add_compo = "address_components"
ad_level = "administrative_area_level_1" # 광역시, 도
local = "locality" # 시
sub_1 = 'sublocality_level_1' #구
sub_2 = 'sublocality_level_2' #동, 면
korea = {
"서울특별시" : "seoul",
"울산광역시" : "ulsan",
"세종광역시" : "sejong",
"전라남도" : "junnam",
"전라북도" : "junbuk",
"제주특별자치도" : "jeju",
"인천광역시" : "incheon",
"경상남도" : "gyungnam",
"경상북도" : "gyungbuk",
"광주광역시" : "gwangju",
"강원도" : "kangwon",
"대전광역시" : "daejeon",
"충청남도" : "chungnam",
"충청북도" : "chungbuk",
"경기도" : "gyunggi",
"대구광역시" : "daegu",
}
#구글 API를 이용, 사용자가 알고자 하는 지역의 주소 정보 받기
gmaps = googlemaps.Client(key = google_secret_key)
place_info = gmaps.geocode(loc, language = 'ko')
place_info = place_info[0][add_compo]
try :
# 해당 지역의 데이터(json)에서 도(광역시), 시, 동(구) 정리
# 에러 방지, 해당 지역의 데이터를 가져오지 않았는데 선언하는 경우
for idx in range(len(place_info)) :
if ad_level in place_info[idx]['types'] :
do_name = place_info[idx]['long_name']
elif local in place_info[idx]['types'] :
si_name = place_info[idx]['long_name']
elif sub_1 in place_info[idx]['types'] :
gu_name = place_info[idx]['long_name']
elif sub_2 in place_info[idx]['types'] :
dong_name = place_info[idx]['long_name']
#환경공단 API가 동이름을 인식하지 못하는 경우, 주변 동을 대입해 결과물 출력(코사인 유사도 기반)
count = 0
while count != 1 :
model = Word2Vec.load('model_' + korea[do_name])
similar = model.wv.most_similar(dong_name, topn = 10)
for idx in range(len(similar)) :
dong_name = similar[idx][0]
url = "http://openapi.airkorea.or.kr/openapi/services/rest/MsrstnInfoInqireSvc/getTMStdrCrdnt?umdName="+ dong_name +"&pageNo=1&numOfRows=10&ServiceKey=" + get_tm_coor_key
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
count = int(dom.select("totalcount")[0].text)
except Exception as e:
result = "error"
X = dom.select("tmx")[0].text
Y = dom.select("tmy")[0].text
result = X, Y, do_name, dong_name
return result
get_tm_coor("대구 황금동")
dong_name = "황금동"
url = "http://openapi.airkorea.or.kr/openapi/services/rest/MsrstnInfoInqireSvc/getTMStdrCrdnt?umdName="+ dong_name +"&pageNo=1&numOfRows=10&ServiceKey=" + get_tm_coor_key
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
dom
# ----
# ## TM 기준좌표 조회
# - 검색서비스를 사용하여 읍면동 이름을 검색조건으로 기준좌표 (TM좌표)정보를 제공하는 서비스
# - 입력값으로는 TM좌표를 입력
# * TM 좌표는, 위경도와 형태는 비슷하나 내용은 다른 위치 표기법
with open("secret/tm_station_key.txt", "rb") as f :
tm_station_key = pickle.load(f)
def nearest_station(X, Y) :
url = "http://openapi.airkorea.or.kr/openapi/services/rest/MsrstnInfoInqireSvc/getNearbyMsrstnList?tmX=\
" + X + "&tmY=" + Y +"&pageNo=1&numOfRows=10&ServiceKey=" + tm_station_key
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
name = dom.select("stationname")[0].text
distance = dom.select("tm")[0].text
return name, distance + "km"
# --------
# ## 실시간 미세 먼지 조회
# - 위치를 입력받아 실제로 미세먼지를 결과값으로 도출하는 함수
def microdust_1(loc) :
url = "http://openapi.airkorea.or.kr/openapi/services/rest/ArpltnInforInqireSvc/getMsrstnAcctoRltmMesureDnsty?stationName="+loc+"&dataTerm=month&pageNo=1&numOfRows=10&ServiceKey="+ tm_station_key + "&ver=1.3"
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
date = dom.select("datatime")[0].text
pm10 = dom.select("pm10value")[0].text
pm25 = dom.select("pm25value")[0].text
result = '''
{}에 관측했을 때,
미세먼지 농도 : {}({}),
초 미세먼지 농도 : {}({})였습니다!
선택하신 지역의 인접지역 미세먼지는 아래 버튼을 클릭하세요.
'''.format(date, pm10, grade(pm10), pm25, nano_grade(pm25))
return result
# ## 인접 지역 추천
# - 주소를 벡터화, 코사인 유사도를 이용한 지형적 근접 지역 추천
# ## 최종 함수
# - 알고 싶은 위치에서 가장 가까운 관측소에서 측정한 미세먼지 및 초미세먼지 농도를 출력
def get_microdust(loc) :
try :
X, Y = get_tm_coor(loc)
name, distance = nearest_station(X, Y)
output_ls = []
result = microdust_1(name)
except Exception as e :
result = "오타이거나 혹은 개발중인 기능을 요구하셨습니다."
return result
get_microdust("수원 영통동")
# #### 클로즈 베타테스트 후 추가한 기능
# - 동이름을 말했는데 국내에 중복되는 지명이 있는 경우가 있음
# (흑석동 입력 시 -> 대전광역시 흑석동, 서울특별시 흑석동, 광주 광역시 흑석동 등)
# - 위와 같은 경우, 리스트로 사용자에게 어느 지역인지 입력하게 함
def check_detail(loc) :
url = "http://openapi.airkorea.or.kr/openapi/services/rest/MsrstnInfoInqireSvc/getTMStdrCrdnt?umdName="+ loc +"&pageNo=1&numOfRows=10&ServiceKey=" + get_tm_coor_key
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
name_ls = [name.text + " " + loc for name in dom.select("sidoname")]
return name_ls
| chatbot/microdust/Total_Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
]activate .
using Semagrams, Catlab.Present, Catlab.Theories, Catlab.CSetDataStructures
# +
@present TheoryPetri(FreeSchema) begin
(S,I,T,O)::Ob
is::Hom(I,S)
it::Hom(I,T)
os::Hom(O,S)
ot::Hom(O,T)
end
@semagramschema PetriSema(TheoryPetri) begin
@box S Circle
@box T Square
@wire I(is,it)
@wire O(ot,os)
end
Petri = CSetType(TheoryPetri);
# -
petri = Semagram{Petri}(PetriSema)
# +
@present TheoryCircuit(FreeSchema) begin
V::Ob
S::Ob
R::Ob
I::Ob
ssrc::Hom(S,V)
stgt::Hom(S,V)
rsrc::Hom(R,V)
rtgt::Hom(R,V)
isrc::Hom(I,V)
itgt::Hom(I,V)
end
@semagramschema CircuitSema(TheoryCircuit) begin
@box V Boxes.SmallCircle
@wire S(ssrc,stgt)
@wire R(rsrc,rtgt)
@wire I(isrc,itgt)
end
Circuit = CSetType(TheoryCircuit);
# -
c = Semagrams.ControlSemagram{Circuit}(CircuitSema)
c.scope
# +
@present TheoryDirectedPortGraph(FreeSchema) begin
Box::Ob
IPort::Ob
OPort::Ob
Wire::Ob
ibox::Hom(IPort,Box)
obox::Hom(OPort,Box)
src::Hom(Wire,OPort)
tgt::Hom(Wire,IPort)
end
@semagramschema DirectedPortGraphSema(TheoryDirectedPortGraph) begin
@box Box Square
@port IPort(ibox) "Input"
@port OPort(obox) "Output"
@wire Wire(src,tgt)
end
DirectedPortGraph = CSetType(TheoryDirectedPortGraph);
# -
# + jupyter={"outputs_hidden": true}
dpg = Semagrams.ControlSemagram{DirectedPortGraph}(DirectedPortGraphSema)
dpg.scope
# +
@present TheoryCircularPortGraph(FreeSchema) begin
Box::Ob
Port::Ob
Wire::Ob
box::Hom(Port,Box)
src::Hom(Wire,Port)
tgt::Hom(Wire,Port)
end
@semagramschema CPGSema(TheoryCircularPortGraph) begin
@box Box Circle
@port Port(box) "Circular"
@wire Wire(src,tgt)
end
CPG = CSetType(TheoryCircularPortGraph);
# -
cpg = Semagrams.ControlSemagram{CPG}(CPGSema)
cpg.scope
enzyme_acset = Semagrams.get_acset(enzyme)
u0 = [100.,0.,10.]
p = [0.01]
prob = ODEProblem(vectorfield(enzyme_acset),u0,(0.,100.),p)
sol = solve(prob,Tsit5(),abstol=1e-8);
plot(sol)
| notebooks/SemagramsTalkExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: brandon_env
# language: python
# name: brandon_env
# ---
# +
import requests as r
import json
import base64
from dotenv import load_dotenv
import os
import urllib.request
import itertools
import pandas as pd
from bs4 import BeautifulSoup
import string
load_dotenv()
oauth_token = os.getenv('GITHUB_TOKEN')
#Get all repositories from Github
base_url = "https://api.github.com/user/repos"
response = r.get(base_url, headers={'Authorization': 'token {}'.format(oauth_token)})
for x in response.json():
print(x["name"])
# -
# +
import itertools
logins = ["brandonleekramer", "uva-bi-sdad", "facebook"]
repos = [ "diversity", "oss-2020", "react"]
for login, repo in zip(logins, repos):
url = f'https://github.com/{login}/{repo}/'
urllib.request.urlretrieve(url, f'readme_{login}_{repo}.txt')
print(f'Finished scraping: {login}/{repo}')
# -
html_text = open("readme_facebook_react.txt", "r")
soup = BeautifulSoup(html_text, 'html.parser')
clean_html = ''.join(soup.article.findAll(text=True))
clean_html
# +
import os
for filename in os.listdir('/sfs/qumulo/qhome/kb7hp/git/oss-2020/src/09_repository-scraping/'):
if filename.endswith('.txt'):
with open(os.path.join('/sfs/qumulo/qhome/kb7hp/git/oss-2020/src/09_repository-scraping', filename)) as f:
content = f.read()
content
# -
html_text = open("readme_brandonleekramer_diversity.txt", "r")
soup = BeautifulSoup(html_text, 'html.parser')
clean_html = ''.join(soup.article.findAll(text=True))
clean_html
# +
html_text = open("readme_facebook_react.txt", "r")
soup = BeautifulSoup(html_text)
clean_html = ''.join(soup.findAll(text=True))
lines = clean_html.split("\n")
non_empty_lines = [line for line in lines if line.strip() != ""]
string_without_empty_lines = ""
for line in non_empty_lines:
string_without_empty_lines += line + "\n"
print(string_without_empty_lines)
type(string_without_empty_lines)
# -
html_text = open("readme_facebook_react.txt", "r")
soup = BeautifulSoup(html_text, 'html.parser')
clean_html = ''.join(soup.article.findAll(text=True))
clean_html
string_without_empty_lines.split('README.md')[3]
# +
login_list = ["brandonleekramer", "bayoan", "gkorkmaz"]
repo_list = [ "diversity", "oss", "another_thing"]
a = []
b = []
for login, repo in zip(login_list, repo_list):
a.append(login)
b.append(repo)
df = pd.DataFrame({'login': a, 'repo': b}, columns=["login", "repo"])
df
# -
# # use this later after we clean the html from the README files
#
# https://www.postgresqltutorial.com/postgresql-python/create-tables/
| src/10_scrape-repos/readme_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import Counter
import matplotlib.pyplot as plt
from typing import List
import sys
# +
from scratch.linear_algebra import sum_of_squares
# +
num_friends = [100.0,49,41,40,25,21,21,19,19,18,18,16,15,15,15,15,14,14,13,13,13,13,12,12,11,10,10,10,10,10,10,10,10,10,10,10,10,10,10,10,9,9,9,9,9,9,9,9,9,9,9,9,9,9,9,9,9,9,8,8,8,8,8,8,8,8,8,8,8,8,8,7,7,7,7,7,7,7,7,7,7,7,7,7,7,7,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,6,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,5,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,2,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1]
# +
friend_counts = Counter(num_friends)
xs = range(101) # largest value is 100
ys = [friend_counts[x] for x in xs] # height is just # of friends
plt.bar(xs, ys)
plt.axis([0, 101, 0, 25])
plt.title("Histogram of Friend Counts")
plt.xlabel("# of friends")
plt.ylabel("# of people")
plt.show()
# +
def mean(xs: List[float])-> float:
return sum(xs)/len(xs)
mean(num_friends)
# -
def _median_odd(xs: List[float]) -> float:
'''
Si el largo de xs es imapr la media es elemento medio
'''
return sorted(xs)[len(xs)//2]
# +
def _median_even(xs: List[float])-> float:
'''
Si el largo de xs es par , ela media es el promedio e los dos elementos medios
'''
sorted_xs = sorted(xs)
upper_midpoint= len(xs)//2
return (sorted_xs[upper_midpoint-1]+sorted_xs[upper_midpoint])/2
# -
def median(v: List[float])->float:
'''
Encuentra el vaor más medio para v
'''
return _median_even(v) if len(v)%2 ==0 else _median_odd(v)
assert median([1,10,2,9,5]) == 5
median_friends = median(num_friends)
median_friends
# Una generalización de la mediana es el cuantil, que representa el valor
# bajo el cual se encuentra un cierto percentil de los datos (la mediana representa el
# valor por debajo del cual se encuentra el 50% de los datos)
def quantile(xs: List[float], p: float) -> float:
'''
Returns the percentil p en el valor de x
'''
p_index = int(p*len(xs))
return sorted(xs)[p_index]
assert quantile(num_friends, 0.10) == 1
assert quantile(num_friends, 0.25) == 3
assert quantile(num_friends, 0.75) == 9
assert quantile(num_friends, 0.90) == 13
quantile(num_friends,0.10)
# +
#La moda. El valor más frecuente.
def mode(x:List[float])->List[float]:
'''
Retorna una lita de los más frecuentes.
'''
count = Counter(x)
max_count = max(count.values())
return [x_i for x_i, count in count.items() if count == max_count]
# -
mode(num_friends)
assert set(mode(num_friends)) == {1,6}
# # Dispersión
#
# Es la medida de la como está distribuido la data, valores cecaos a 0 significan que no están dispersos, y valores grandes significa que están muy dispersos. Una medida simple es el rango. **Rango** es la diferencia entre el más largo y el más corto de lso elementos:
def data_range(xs: List[float]) -> float:
return max(xs) -min(xs)
'''
El rango es muy influenciado por la diferencia d elos valores que poseen, el caso donde el rango es 0 es cuando %%latex
los datos son exactamente los mismos.
El rango, como la media, no depende realmente de todo el dataset.
'''
data_range(num_friends)
# # Varianza
#
#
| NoteBooks/Curso de Ciencia de datos/Statistics/DSFS - ESTADÍSTICA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# based on: https://github.com/essentialsofparallelcomputing/Chapter3/blob/master/JupyterNotebook/HardwarePlatformCharaterization.ipynb
# ## Operator Benchmark on Gauss3
# ### Theoretical Parameters
Sockets=2
ProcessorFrequency=2.35
ProcessorCores=64
Hyperthreads=2
VectorWidth=256
WordSizeBits=64
FMA=2
DataTransferRate=3200
MemoryChannels=8
BytesTransferredPerAccess=8
TheoreticalMaximumFlops=Sockets*ProcessorCores*Hyperthreads*ProcessorFrequency*VectorWidth/WordSizeBits*FMA
TheoreticalMemoryBandwidth=Sockets*DataTransferRate*MemoryChannels*BytesTransferredPerAccess/1000
# ### Empirically Determined Parameters
#
# based on `src_master_thesis/node_characterization/likwid-bench_gauss3.out`:
emp_flops_max=2459.32383
emp_mem_band=233.172
# ### Benchmark data
# #### Previous Versions
# +
FLOPS_apply = [
90.8279029,
205.09239259999998,
208.7462992,
875.2892422,
742.2407463,
725.7962939,
733.7519474
]
AI_apply = [
0.43017927686062557,
0.9082995369414192,
0.929154764181568,
4.924579091929559,
10.63773242520501,
14.481824627676085,
22.428474680608744
]
labels_apply = [
"11_several_coeff_precalc",
"nonlinear_mul2",
"nonlinear_pow2",
"nonlinear_exp",
"nonlinear_pow4_3",
"nonlinear_costly_0",
"nonlinear_costly_1"
]
# -
# #### Current Versions
# +
import csv
def read_Results(p_path):
results = []
with open(p_path, 'r') as f:
reader = csv.reader(f)
#
# NOTE skip the header
#
next(reader, None)
for row in reader:
#
# NOTE select row
#
if row[4] == 'apply':
performance = float(row[10])
memory_bandwidth = float(row[11])
result = {
#
# NOTE MFLOP/s => GFLOP/s
#
'flops': performance/1000,
'ai': performance/memory_bandwidth,
'label': f'{row[2]}_{row[3]}'
}
results.append(result)
return results
# +
results = read_Results('./e_roofline.csv')
print(results)
for result in results:
FLOPS_apply.append(result['flops'])
AI_apply.append(result['ai'])
labels_apply.append(result['label'])
# -
# ### Roofline Model
# +
# roofline_flops=TheoreticalMaximumFlops
# roofline_mem_band=TheoreticalMemoryBandwidth
roofline_flops=emp_flops_max
roofline_mem_band=emp_mem_band
roofline_ma_bal=roofline_flops/roofline_mem_band
# +
# # Install a pip package in the current Jupyter kernel
# import sys
# # !{sys.executable} -m pip install matplotlib
# # !{sys.executable} -m pip install numpy
# # %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
font = { 'size' : 20}
plt.rc('font', **font)
markersize = 16
#
# NOTE: color scheme 'Tableau 20'
# source: https://help.tableau.com/current/pro/desktop/de-de/formatting_create_custom_colors.htm
#
colors = (
'#17becf',
'#dbdb8d',
'#bcbd22',
'#c7c7c7',
'#7f7f7f',
'#f7b6d2',
'#e377c2',
'#c49c94',
'#8c564b',
'#c5b0d5',
'#9467bd',
'#ff9896',
'#d62728',
'#98df8a',
'#2ca02c',
'#ffbb78',
'#ff7f0e',
'#aec7e8',
'#1f77b4'
)
styles = ['o','s','v','^','D',">","<","*","h","H","+","1","2","3","4","8","p","d","|","_",".",","]
roofline_color = 'r'
fig = plt.figure(1,figsize=(15,10))
plt.clf()
ax = fig.gca()
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel('Arithmetic Intensity [FLOPS/Byte]')
ax.set_ylabel('Performance [GFLOP/sec]')
ax.grid()
ax.grid(which='minor', linestyle=':', linewidth=0.5, color='black')
nx = 10000
xmin = -3
xmax = 2
ymin = 0.1
ymax = 10000
ax.set_xlim(10**xmin, 10**xmax)
ax.set_ylim(ymin, ymax)
ixx = int(nx*0.02)
xlim = ax.get_xlim()
ylim = ax.get_ylim()
scomp_x_elbow = []
scomp_ix_elbow = []
smem_x_elbow = []
smem_ix_elbow = []
x = np.logspace(xmin,xmax,nx)
#
# rooflines
#
for ix in range(1,nx):
if roofline_mem_band * x[ix] >= roofline_flops and roofline_mem_band * x[ix-1] < roofline_flops:
theoMem_ix_elbow = ix-1
break
for ix in range(1,nx):
if (roofline_flops <= roofline_mem_band * x[ix] and roofline_flops > roofline_mem_band * x[ix-1]):
theoFlops_ix_elbow = ix-1
break
y = np.ones(len(x)) * roofline_flops
ax.plot(x[theoFlops_ix_elbow:],y[theoFlops_ix_elbow:],c=roofline_color,ls='--',lw='2')
ax.text(x[-ixx],roofline_flops*0.95,
'node1: P_T',
horizontalalignment='right',
verticalalignment='top',
c=roofline_color)
y = x * roofline_mem_band
ax.plot(x[:theoMem_ix_elbow+1],y[:theoMem_ix_elbow+1],c=roofline_color,ls='--',lw='2')
ang = np.arctan(np.log10(xlim[1]/xlim[0]) / np.log10(ylim[1]/ylim[0])
* fig.get_size_inches()[1]/fig.get_size_inches()[0] )
ax.text(x[ixx],x[ixx]*roofline_mem_band*(1+0.25*np.sin(ang)**2),
'node1: B_T',
horizontalalignment='left',
verticalalignment='bottom',
rotation=180/np.pi*ang,
c=roofline_color)
plt.vlines(roofline_ma_bal, 0, roofline_flops, colors=roofline_color, linestyles='dashed', linewidth=2)
ax.text(roofline_ma_bal,2*ymin,
'node1: MB_T',
horizontalalignment='right',
verticalalignment='bottom',
rotation=90,
c=roofline_color)
marker_handles = list()
for i in range(0,len(AI_apply)):
ax.plot(float(AI_apply[i]),float(FLOPS_apply[i]),c=colors[i],marker=styles[0],linestyle='None',ms=markersize,label=labels_apply[i])
marker_handles.append(ax.plot([],[],c=colors[i],marker=styles[0],linestyle='None',ms=markersize,label=labels_apply[i])[0])
#
# NOTE: put legend out of the plot: https://stackoverflow.com/a/43439132
#
leg1 = plt.legend(handles = marker_handles,
ncol=2,
bbox_to_anchor=(0.5,-0.25),
bbox_transform=fig.transFigure,
loc="lower center")
ax.add_artist(leg1)
# plt.savefig('roofline.png')
# plt.savefig('roofline.eps')
# plt.savefig('roofline.pdf')
# plt.savefig('roofline.svg')
plt.show()
| src_optimization/40_nonlinear_cg/e_roofline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MyPython
# language: python
# name: mypython
# ---
//%log:0
//%onlyruncmd
//%command:node --version
//%log:0
//%onlyruncmd
//%npmcmd: --version
# +
//%log:0
///%overwritefile
///%file:src/mlinetest1.js
var x = "我的\
公司市值\
2000亿美元";
console.log(x);
# -
//%log:0
console.log("Hello World");
console.log( __filename );
console.log( __dirname );
function printHello1(){
console.log( "1.Hello, World!");
}
// 两秒后执行以上函数
setTimeout(printHello1, 2000);
function printHello2(){
console.log( "2.Hello, World!");
}
// 两秒后执行以上函数
var t = setTimeout(printHello2, 2000);
// 清除定时器
clearTimeout(t);
function printHello3(){
console.log( "3.Hello, World!");
}
// 两秒后执行以上函数
setInterval(printHello3, 2000);
//hello.js
//%log:1
//%noruncode
//%overwritefile
//%file:src/hello.js
function Hello() {
var name;
this.setName = function(thyName) {
name = thyName;
};
this.sayHello = function() {
console.log('Hello ' + name);
};
};
module.exports = Hello;
//main.js
//%log:0
//%overwritefile
//%file:src/main.js
var Hello = require('./hello');
hello = new Hello();
hello.setName('BYVoid');
hello.sayHello();
# +
//main.js
//%log:0
//%overwritefile
//%file:src/test1.js
// 引入 events 模块
var events = require('events');
// 创建 eventEmitter 对象
var eventEmitter = new events.EventEmitter();
// 创建事件处理程序
var connectHandler = function connected() {
console.log('连接成功。');
// 触发 data_received 事件
eventEmitter.emit('data_received');
}
//---------------------------------
// 绑定 connection 事件处理程序
eventEmitter.on('connection', connectHandler);
// 使用匿名函数绑定 data_received 事件
eventEmitter.on('data_received', function(){
console.log('数据接收成功。');
});
//---------------------------------
// 触发 connection 事件
eventEmitter.emit('connection');
console.log("程序执行完毕。");
# +
// ES6出来可以通过Promise来进行异步处理
// 方式1
var p = new Promise(function (resolve, reject) {
// resolve 表示执行成功
// reject 表示执行失败
setTimeout(() => {
var name = "小明";
resolve(name)
})
})
p.then( (data) => {
console.log(data);// 小明
})
# +
// ES6出来可以通过Promise来进行异步处理
// 方式2 ,封装getData函数
function getData(resolve, reject) {
setTimeout(function (){
var name = "Tom"
resolve(name);
}, 5000)
}
var p = new Promise(getData)
p.then((data)=> {
console.log(data); //Tom
})
# -
//%log:0
//%overwritefile
//%file:src/test4.js
async function test () {
return "hello world!"
}
console.log(test())
// Promise { 'hello world!' }
async function main () {
var data = await test();
console.log(data);// hello world!
}
main()
# +
async function test () {
return new Promise((resolve, reject) => {
setTimeout(function () {
var name = "Lucy";
resolve(name);
}, 1000)
})
}
async function main () {
var data = await test();
console.log(data);//Lucy
}
main()
# +
var fs = require("fs")
async function isDir(path) {
return new Promise((resolve, reject)=> {
fs.stat(path, (err,stats)=> {
if (err) {
console.log(err);
reject(err);
return
}
if (stats.isDirectory()) {
resolve(true);
}else {
resolve(false);
}
})
})
}
function main() {
var path = "../MyNodejs"
var dirArr = []
fs.readdir(path, async (err, data)=> {
if (err) {
console.log(err);
return
}
for (var i=0;i<data.length;i++){
if (await isDir(path + '/' + data[i])) {
dirArr.push(data[i])
}
}
console.log(dirArr)
})
console.log('Gameover!')
}
main()
# -
print("MyPython")
| example/Nodejs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Outlier
# An outlier is an observation that lies outside the overall pattern of a distribution (Moore and McCabe 1999). It can be found by residual plots and scatter plot of x, y points.
# +
#This outlier analysis is adapted from our work Outlier_analysis.py, with change of order of some code lines
from regression import GenerateData, UniformX #our work
from Outlier import Outlier, Outlier_position #our work
from simulation import print_coef, simulation, change_factor #our work
import numpy as np
from numpy.random import default_rng
import matplotlib.pyplot as plt
from mpl_toolkits import mplot3d
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from yellowbrick.regressor import ResidualsPlot #need prior installation
rng = np.random.default_rng()
#initialize for object of GenerateData sub class Outlier
set1=UniformX(N=1000)
set1.generate_dataset()
X=set1.X
y=set1.y
#setting number and magnitude of outliers
outlier_number = 10
outlier_magnitude = 200
#residual plot - library reference: https://www.scikit-yb.org/en/latest/api/regressor/residuals.html
#initialize for object of GenerateData sub class Outlier
test4=Outlier(N=1000)
test4.generate_dataset(magnitude=outlier_magnitude, original_X=X, original_y=y, original_beta=set1.beta, positions=positions)
X_train, X_test, y_train, y_test = train_test_split(test4.X, test4.y, test_size=0.2, random_state=42) # split the train and test data
model = LinearRegression() # Instantiate the linear model and visualizer
visualizer = ResidualsPlot(model)
visualizer.fit(X_train, y_train) # Fit the training data to the visualizer
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.show() # Finalize and render the figure
print("Fig. 1 Residual Plot")
#how to detect outliers
#initialize for object of GenerateData sub class Outlier
test3=Outlier(N=1000)
positions=Outlier_position([-10,-10], N=outlier_number)
test3.generate_dataset(magnitude=outlier_magnitude, original_X=X, original_y=y, original_beta=set1.beta, positions=positions)
test3.fit()
test3.plot2D()
plt.show()
print("Fig. 2 Scatter plot of x, y points")
# -
# # Detection of outliers
# In Fig. 1 outliers are found in the far negative area of the two residual plots. In the histogram of residual plot below, most residuals are around zero in a shape similar to normal distribution, except those in far negative area, hence those are outliers. If standardized, residuals beyond +/- 3 may be regarded as outliers. Residual scatter plot against predicted values of y can also display obvious outliers e.g. bottom left of Fig. 1.
# In Fig. 2, scatter plot of x and y also visualizes outliers in the top left. However, note that outlier in scatter plot of two independent variables x1 and x2 (i.e. not showing y) may not affect the model if the predicted value y is not too far from actual value.
# # Impact of outliers
# An outlier can affect the linear regression model greatly as the difference is squared for minimization. We will study how outliers affect the intercept, coefficients, residuals and R2 score of the model by holding the same underlying true linear model y = 1 + 2$x_{1}$ + 3$x_{2}$ + e (where e is random noise variable ~N(0,1)), but with varying magnitude, number and position of outliers. Simulations are done 1000 times for calculating the variance and mean.
#
# +
#testing impact
#test1 - test impact of Num of Outliers, holding magnitude constant at 500
test1=Outlier(N=1000)
outlier_magnitude = 500
#assuming i)evenly spread, ii)high end value of input X, iii)low end, iv)centre value of input X
#initialize 4 different scenarios of outlier positions, and set varying number of outliers for each scenario
Number_of_outliers = np.arange(0,11)
p = ['even', 'high','low','centre']
pos = {k:None for k in p}
num = {k:None for k in p}
pos['even']=[Outlier_position([10,10],[-10,-10],N=i) for i in Number_of_outliers]
pos['high']=[Outlier_position([10,10],N=i) for i in Number_of_outliers]
pos['low']=[Outlier_position([-10,-10],N=i) for i in Number_of_outliers]
pos['centre']=[Outlier_position([0,0],N=i) for i in Number_of_outliers]
#create dictionary of dictionary storing results based on varying number of outliers for 4 position scenarios
for position in p:
num[position] = change_factor(test1, 1000,factor={"positions":pos[position]}, magnitude=outlier_magnitude, original_X=X, original_y=y, original_beta=set1.beta,)
#create chart to show impact on coefficients against number of outliers
print('testing impact of number of outliers')
for estimate_key in ["b0_mean", "b1_mean", "b2_mean", "b0_variance", "b1_variance", "b2_variance","score_mean", "score_variance", "ssr_mean", "ssr_variance"]:
for position in p:
plt.plot(Number_of_outliers, num[position][estimate_key], label = position)
plt.title(estimate_key)
plt.xlabel('Number of Outliers')
plt.legend()
plt.show()
#test2 - test impact of magnitude of Outliers, holding number of outliers constant at 10
#assuming i)evenly spread, ii)high end value of input X, iii)low end, iv)centre value of input X
#initialize for object of GenerateData sub class Outlier
test2=Outlier(N=1000)
outlier_number = 10
#initialize 4 different scenarios of outlier positions, and set varying number of outliers for each scenario
Magnitude_of_outliers = np.arange(-1000,1000, step=100)
p = ['even', 'high','low','centre']
pos = {k:None for k in p}
mag = {k:None for k in p}
pos['even']=Outlier_position([10,10],[-10,-10], N=outlier_number)
pos['high']=Outlier_position([10,10], N=outlier_number)
pos['low']=Outlier_position([-10,-10], N=outlier_number)
pos['centre']=Outlier_position([0,0], N=outlier_number)
#create dictionary of dictionary storing results based on varying magnitude of outliers for 4 position scenarios
for position in p:
mag[position] = change_factor(test2, 1000,factor={"magnitude":Magnitude_of_outliers},positions=pos[position], original_X=X, original_y=y, original_beta=set1.beta,)
#create chart to show impact on coefficients against magnitude of outliers
print('testing impact of magnitude of outliers')
for estimate_key in ["b0_mean", "b1_mean", "b2_mean", "b0_variance", "b1_variance", "b2_variance","score_mean", "score_variance", "ssr_mean", "ssr_variance"]:
for position in p:
plt.plot(Magnitude_of_outliers, mag[position][estimate_key], label = position)
plt.title(estimate_key)
plt.xlabel('Magnitude of Outliers')
plt.legend()
plt.show()
# -
# # Summary of findings from 20 charts above
# The fitted intercept increases linearly with magnitude and number of outliers, regardless of the positions of outliers. Interestingly, while the variance of the intercept increases with the number of outliers, it only fluctuates with the magnitude of the outliers.
#
# Position matters a lot - outliers at centre of feature space do not affect b1 at all! Evenly distributed outliers across the feature space may slightly change b1, but the mean of b1 remain the same as the true model and the variance is smaller than if outliers are concentrated all at the high end (or all at the low end) of the feature space. The mean of b1 is in linear relationship with magnitude and number of outliers that are not evenly distributed. The variance of b1 increases with number of outliers but only fluctuates with the magnitude.
#
# We measure residuals by sum of squares of residuals (ssr). The mean of ssr increases linearly with number of outliers and quadratically with the magnitude, regardless of position of outliers. The ssr variance has the same shape as the mean of ssr, though less smoothly.
#
# R2 score = 1-ssr/(Var(Y)) , interpreted as proportion of explained variance, is commonly used as one indicator of fitness of the model, though it may need adjustment when compared with models of different number of parameters. Its mean decreases with absolute magnitude and number of outliers, which implies that ssr increases more than the total variance. Its variance also have same shape except at the point of zero magnitude or zero number of outliers.
| task2/src/.ipynb_checkpoints/Linear Regression Model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Who voted what? - A dive into the Colombia's peace plebiscite outcome
#
# **_by <NAME>_**
#
# Last October 2th was a milestone in the Colombian history. On that Sunday, Colombians casted votes approving or rejecting the peace agreement signed between the official government and FARC guerrilla. The outcome came as a setback to pollsters’ predictions as the negative vote prevailed by a razor-thin margin of 50.21% to 49.78%, leaving the country immersed in a massive sea of uncertainty. This plebiscite was important since the peace accord represents an opportunity to halt an armed conflict prolonged for over 50 years, which so far has left the an estimated of 218,000 people killed, 27,000 kidnappings, 5.7 million forced displaces and many more human rights violations and war crimes (see plot below).
#
# In this tutorial, the results from the plebiscite are collected, processed and analyzed. Theis dataset is crossed with adjacent country statistics (e.g. poverty rate and conflict impact region-wise), to answer insightful questions and draw conclusions using data analytics techniques.
#
# [<img src="1402confl.png" width="600">](https://www.wola.org/files/images/1402confl.png)
#
# The tutorial is organized as follows:
# - [Background](#Background)
# - [Setting up the data](#Setting-up-the-data)
# - [How did "YES/"NO" votes distribute?](#How-did-"YES/"NO"-votes-distribute?)
# - [What did the most vulnerable regions vote?](#What-did-the-most-vulnerable-regions-vote?)
# - [Did people actually vote?](#Did-people-actually-vote?)
# - [Main takeaways](#Main-takeaways)
# - [References](#References)
#
# ## Background
#
# FARC is the oldest and largest a guerrilla in Latin America. This leftist group is known for employing a variety of military tactics including terrorism, children recruitment and sexual assault, among other unconventional methods. Colombian governments have combatted this group along history in order to protect the population and civil infrastructure. Not surprisingly, the poorest regions of the country have suffered the most the fallouts from this conflict.
#
# Attempts to negotiate with FARC have been undertaken in the past with unsuccessful results. On August 2012, the government of President <NAME> embarked in a new negotiation with FARC seeking to terminate this conflict once and for all. After four years of negations both parties crafted an agreement, the first ever reached, which renders a true possibility to end this half-century-long warfare.
#
# To boost public support and preserve legitimacy, President Santos promised to give people the opportunity to back or reject the accord through this plebiscite. Voters were asked a single question in the ballot:
#
# _**“Do you support the final accord to end the conflict and build a stable and lasting peace?”**_
#
# The possible answers were: **“YES”** or **“NO”**. _Any other answers would make the vote invalid_. For further information about the conflict and peace agreement in Colombia, please refer to the links in [References](#References).
#
# ## Setting up the data
#
# A web scraper was programmed to extract the data since results from the plebiscite are available in the form of infrographics at the [Colombia's National Registry website](http://plebiscito.registraduria.gov.co/99PL/DPLZZZZZZZZZZZZZZZZZ_L1.htm). The country is divided in 32 regions plus the Capital District. Here, the consulates are treated as a region, then totaling 34 regions. Furthermore, regions are divided into cities and municipalities - for the sake of simplicity, _municipality_ will refer to both categories. There are 1,098 municipalities in the database.
#
# The following python libraries are used in this tutorial:
import numpy as np, math, requests, re, pandas as pd, json, geopandas as gpd
import matplotlib, shapely, rtree, matplotlib.pyplot as plt, seaborn as sns
from bs4 import BeautifulSoup
from scipy.stats import norm
from geopy.geocoders import GoogleV3
# %matplotlib inline
matplotlib.rc("figure", figsize=(8,6));matplotlib.rc("axes", labelsize=16, titlesize=16)
matplotlib.rc("xtick", labelsize=14);matplotlib.rc("ytick", labelsize=14)
matplotlib.rc("legend", fontsize=14);matplotlib.rc("font", size=14);sns.set()
# Regional and municipal results are in different `url` addresses with an standardized HTML format. The `read_data_page` function parses the HTML content to extract the desired information using `BeautifulSoup and regular expressions`. This function returns a dictionary:
# ```python
# {
# 'total_voters': (int) Total eligible voters
# 'voters': (int) Total actual votes
# 'yes_votes': (int) Number of YES votes
# 'yes_votes_p': (float) Proportion of YES votes
# 'no_votes': (int) Number of NO votes
# 'no_votes_p': (float) Proportion of NO votes
# 'valid_votes': (int) Total valid votes
# 'unmarked_votes': (int) Votes not marked
# 'null_votes': (int) Null votes
# }
# ```
def read_data_page(url):
# This function reads the content of number of votes, type of votes, number of voters, etc...
output = {} # Dictionary containing the retrieved data
raw = requests.get(url)
pinput = BeautifulSoup(raw.content, "html.parser")
# List of municipalities as xml tags
try:
muni_list = pinput.find('select', id = 'combo3').find_all('option')
except AttributeError:
muni_list = []
# Number of voters vs. number of people allowed to vote
total_voters = pinput.find('div', class_ = 'cajaSupSegundaContainer').find('span', class_ = 'descripcionCaja').get_text()
total_voters = total_voters.replace('.','')
nums = re.compile(r"\d+").findall(total_voters)
output['voters'] = int(nums[0])
output['total_voters'] = int(nums[1])
#Positive and negative votes
votes = pinput.find_all('div', class_ = 'skill-bar-percent')
temp = votes[0].get_text().replace('%','').replace(',','.')
output['yes_votes_p'] = float(temp)/100
temp = votes[1].get_text().replace('.','')
output['yes_votes'] = int(re.compile(r"\d+").findall(temp)[0])
temp = votes[2].get_text().replace('%','').replace(',','.')
output['no_votes_p'] = float(temp)/100
temp = votes[3].get_text().replace('.','')
output['no_votes'] = int(re.compile(r"\d+").findall(temp)[0])
#Valid and invalid votes
temp = pinput.find('div', class_ = 'cajaInfPrimera').find('div', class_ = 'contenido').get_text().replace('.','')
output['valid_votes'] = int(re.compile(r"\d+").findall(temp)[0])
temp = pinput.find('div', class_ = 'cajaInfSegunda').find('div', class_ = 'contenido').get_text().replace('.','')
output['unmarked_votes'] = int(re.compile(r"\d+").findall(temp)[0])
temp = pinput.find('div', class_ = 'cajaInfTercera').find('div', class_ = 'contenido').get_text().replace('.','')
output['null_votes'] = int(re.compile(r"\d+").findall(temp)[0])
return output, muni_list
# Two dictionaries of dictionaries are created: `regions` and `munis`, representing regions and municipalities respectively. The following script fills up the information in both dictionaries by iterating over the websites and calling the `read_data_page` function.
# Creating dictionies for regions and municipalities with name, url votes statistics for each one
# This script takes approximately 4.5 minutes
root_url = 'http://plebiscito.registraduria.gov.co'
url = root_url + "/99PL/DPLZZZZZZZZZZZZZZZZZ_L1.htm"
rurl = requests.get(url)
pinput = BeautifulSoup(rurl.content, "html.parser")
reg_list = pinput.find('select', id = 'combo2').find_all('option') # List of regions as xml tags
regions = {}; munis = {}
for dpt in reg_list:
reg_name = dpt.get_text().replace('.','').replace(',','')
if reg_name == 'Todos':
reg_name = 'COLOMBIA'
reg_url = root_url + dpt['value'][2:]
regions[reg_name] = {}
regions[reg_name]['url'] = reg_url
rdata = read_data_page(reg_url) # Extarcting data for the specific region
regions[reg_name].update(rdata[0])
if reg_name == 'COLOMBIA':
continue
# Creating dictionary for municipalities
for muni in rdata[1]:
muni_name = muni.get_text().replace('.','').replace(',','')
if muni_name == 'Todos':
continue
munis[muni_name] = {}
muni_url = root_url + muni['value'][2:]
munis[muni_name]['region'] = reg_name
munis[muni_name]['url'] = muni_url
rdata2 = read_data_page(muni_url) # Extarcting data for the specific municipality
munis[muni_name].update(rdata2[0])
pass
# An example of the dictionary structure for the municipality of SOACHA:
# ```python
# region: 'CUNDINAMARCA'
# url: 'http://plebiscito.registraduria.gov.co/99PL/DPL15247ZZZZZZZZZZZZ_L1.htm'
# total_voters : 201745
# voters : 90969
# yes_votes : 42449
# yes_votes_p : 0.4758
# no_votes : 46767
# no_votes_p: 0.5241
# valid_votes: 89216
# unmarked_votes: 289
# null_votes: 1464```
# The two dictionaries are then transformed into dataframes with rows listing the different regions/municipalities and columns correspond to the keys.
df_regions = pd.DataFrame.from_dict(regions, orient='index'); df_munis = pd.DataFrame.from_dict(munis, orient='index')
df_regions.drop('url', axis=1, inplace=True); df_regions.drop('COLOMBIA', inplace=True);
df_munis.drop(df_munis[df_munis.no_votes == 0].index, axis=0, inplace=True)
# Finally, additional data was incorporated to enhance the analysis. While a comprehensive assessment should consider a swath of socio-economic factors, given the scope of this tutorial and the availability of data, the present analysis only includes three additional factors:
# - _Poverty:_ The latest estimations of economic poverty rate per region were downloaded from the National Statistics Department ([DANE](https://www.dane.gov.co/index.php/estadisticas-por-tema/pobreza-y-condiciones-de-vida/pobreza-y-desigualdad)) statistics database.
# - _Violence intensity index:_ This [map](http://www.kienyke.com/politica/las-zonas-de-mas-violencia-en-colombia) prepared by the Humanitarian Attention Unit (OCHA, acronym in Spanish), resumes the violent events occurred during 2012-2013 related to the armed conflict by region. A _violence index_ was computed by dividing the number of violent events into the total voters per region. These values were further normalized by setting the region with the highest score as “1” and then scaling the others accordingly.
# - _Vulnerability index:_ The [Foundation of Peace and Reconciliation]( http://www.pares.com.co/) has identified the municipalities most vulnerable to relapse into violence in a post-peace-agreement scenario given the historical strong presence of armed illegal groups. This [map]( http://www.las2orillas.co/wp-content/uploads/2015/02/mapa-.png) classifies the municipalities in four categories of vulnerability: _Low, Medium, High and Extreme_.
# +
# Load and incorporate information in terms of poverty rate, income per capita and number of violent events per region
v_events = pd.read_csv(r'regional_data.csv',encoding='utf8'); v_events.set_index('Reg', inplace = 'True')
df_regions = df_regions.join(v_events)
# Load and incorporate information in terms conflict vulnerability index per municipality/city
vulner = pd.read_csv(r'vulnerability_index.csv',encoding='utf8'); vulner.set_index('muni', inplace = 'True');
df_munis = df_munis.join(vulner)
# Useful calculations
df_regions['yes_no_ratio'] = df_regions["yes_votes_p"]/df_regions["no_votes_p"]
df_munis['yes_no_ratio'] = df_munis["yes_votes_p"]/df_munis["no_votes_p"]
df_regions['elec_part'] = df_regions["voters"]/df_regions["total_voters"]
df_munis['elec_part'] = df_munis["voters"]/df_munis["total_voters"]
temp = (df_regions["v_events"]/df_regions["total_voters"]); temp = temp/max(temp);df_regions['conflict'] = temp;df_regions.tail(3)
# -
# ## How did "YES/"NO" votes distribute?
# To get a sense upon how Colombians voted overall, Figures 1 and 2 show the histograms of "YES" and "NO" votes participation across municipalities, respectively. It is worth to highlight that the **"NO" votes won by a very tight margin of 24,213 votes (less than 0.2% of total votes)**. The histograms approximate to a triangular distribution in both categories, with mode around 55% for "NO" and 45% for "YES", although the average share of "YES" votes was higher. Interestingly, the standard deviation was the same in both cases, however, the coefficient of variation was higher for the "NO" meaning the it slightly variated more across the territory.
# Generations histograms and descriptive statistics of "Yes" and "No" votes
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(20,10))
axes[0].hist(df_munis['yes_votes_p'],color='g',bins=50,alpha=0.7)
axes[0].text(0.7,56,r'Total votes: {:,}'.format(np.sum(df_munis['yes_votes'])),fontsize=14)
axes[0].text(0.7,54,r'Average: {0:.3g}%'.format(np.mean(df_munis['yes_votes_p'])*100),fontsize=14)
axes[0].text(0.7,52,r'Std: {0:.3g}'.format(np.std(df_munis['yes_votes_p'])),fontsize=14)
axes[0].text(0.7,50,r'Var. coefficient: {0:.3g}'.format(np.std(df_munis['yes_votes_p'])/np.mean(df_munis['yes_votes_p'])),fontsize=14)
axes[0].text(0.7,48,r'Max: {0:.3g}%'.format(np.max(df_munis['yes_votes_p'])*100),fontsize=14)
axes[0].text(0.7,46,r'Min: {0:.3g}%'.format(np.min(df_munis['yes_votes_p'])*100),fontsize=14)
axes[0].set_title('Fig. 1: Histogram and stats of "YES" votes participation in municipalities/cities',fontsize=15)
axes[1].hist(df_munis['no_votes_p'],color='r',bins=50,alpha=0.7);
axes[1].text(0.1,56,r'Total votes: {:,}'.format(np.sum(df_munis['no_votes'])),fontsize=14)
axes[1].text(0.1,54,r'Average: {0:.3g}%'.format(np.mean(df_munis['no_votes_p'])*100),fontsize=14)
axes[1].text(0.1,52,r'Std: {0:.3g}'.format(np.std(df_munis['no_votes_p'])),fontsize=14)
axes[1].text(0.1,50,r'Var. coefficient: {0:.3g}'.format(np.std(df_munis['no_votes_p'])/np.mean(df_munis['no_votes_p'])),fontsize=14)
axes[1].text(0.1,48,r'Max: {0:.3g}%'.format(np.max(df_munis['no_votes_p'])*100),fontsize=14)
axes[1].text(0.1,46,r'Min: {0:.3g}%'.format(np.min(df_munis['no_votes_p'])*100),fontsize=14)
axes[1].set_title('Fig. 2: Histogram and stats of "NO" votes participation in municipalities/cities',fontsize=15);
# Data from Figures 1 and 2 show that even though on average Colombians voted favorably at the municipal level, the amount of "YES" votes gathered in pro-peace-agreement regions was not enough to outweigh the negative votes in the regions of opposition overall.
#
# ## What did the most vulnerable regions vote?
# Considering that the armed conflict with FARC has spread geographically in an asymmetric fashion, there are zones largely more affected than others. Therefore, it is worthwhile to find out what is the stand of people directly impacted by this conflict with regards the peace agreement. A geospatial visualization comes handy in order to analyze variables that are geographically related. Using [`Geopandas`](http://geopandas.org/index.html) package and the [`geojson` map of Colombia](https://bl.ocks.org/john-guerra/43c7656821069d00dcbc), Figures 3 and 4 are generated to illustrate the ratio of YES/NO votes as well as the violence intensity index region-wise, respectively.
# +
# Importing geojson file containing Colombian coordinates and merging data from df_regions
geo_regions = gpd.read_file('Colombia.geo.json.txt')
geo_regions.set_index('NOMBRE_DPT', inplace = True)
geo_regions = geo_regions.join(df_regions.loc[:,['yes_no_ratio','conflict']],how='inner')
# Map plot of YES/NO ratios and violence index across Colombia
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(20,10))
ax1 = geo_regions.plot(column='yes_no_ratio', scheme='quantiles',figsize = (10,10),cmap='Greens',ax=axes[0]);
ax1.tick_params(axis='x', colors='white');ax1.tick_params(axis='y', colors='white')
ax1.set_title('Fig. 3: Ratio of YES vs. NO votes across regions',fontsize=14)
fig1 = ax1.get_figure();cax = fig1.add_axes([0.126, 0.868, 0.35, 0.03])
sm = plt.cm.ScalarMappable(cmap='Greens', norm=plt.Normalize(vmin=4, vmax=0)); sm._A = []
fig1.colorbar(sm, cax=cax, orientation='horizontal')
ax2 = geo_regions.plot(column='conflict', scheme='quantiles',figsize = (10,10),cmap='Reds',ax=axes[1]);
ax2.tick_params(axis='x', colors='white');ax2.tick_params(axis='y', colors='white')
ax2.set_title('Fig. 4: Scale of violence intensity during 2012-2013 across regions',fontsize=14)
fig2 = ax2.get_figure();cax = fig2.add_axes([0.548, 0.868, 0.35, 0.03])
sm2 = plt.cm.ScalarMappable(cmap='Reds', norm=plt.Normalize(vmin=1, vmax=0)); sm2._A = []
cb2 = fig2.colorbar(sm2, cax=cax, orientation='horizontal')
# -
# At the regional level, a modest positive correlation between the two variables plotted could be grasped by visually contrasting Figures 3 and 4. In regions like the West-coast and the South-west, where the intensity of violent events is high, "YES" votes outweighed "NO" votes by a factor of 2x to 4x. However, there also some regions highly affected by violence as well, like the Mid-south and Mid-north, where the "NO" votes prevailed. To dig deeper into this issue, the data could be also correlated with the vulnerability index at the municipal level. The following script generates Figure 5 which presents the average and standard deviation of “YES” and “NO” voting participation in municipalities classified by vulnerability category.
# Erorbar plot for YES/NO votes participation in munipalities grouped by vulnerability index
plt.figure(figsize=(10,5)); df_munis_gv = df_munis.groupby(['vulner_idx']);leng = np.arange(len(df_munis_gv)); wd = 0.3
yes_means = df_munis_gv['yes_votes_p'].mean()*100; yes_stds = df_munis_gv['yes_votes_p'].std()*100
no_means = df_munis_gv['no_votes_p'].mean()*100; no_stds = df_munis_gv['no_votes_p'].std()*100
plt.bar(leng,yes_means.values,wd,color='g',yerr=yes_stds.values,error_kw=dict(ecolor='black',capthick=2,label=''),alpha=0.7)
plt.bar(leng+wd,no_means.values,wd,color='r',yerr=no_stds.values, error_kw=dict(ecolor='black',capthick=2),alpha=0.7)
plt.ylabel('Voting share (%)');plt.xlabel('Vulnerability Index (Percentage of total voters)',fontsize = 13);
plt.legend(['Standard deviation','Avg. Yes votes','Avg. NO votes'],fontsize = 12,loc=2)
plt.title('Fig. 5: Comparison of YES/NO votes participation in munipalities grouped by vulnerability index',fontsize = 14)
vot = df_munis_gv['voters'].sum()/np.sum(df_munis['voters'])*100
plt.xticks(leng+wd); plt.xticks(leng+wd,('Low ({0:.2g}%)'.format(vot[0]), 'Medium ({0:.1g}%)'.format(vot[1]),'High ({0:.1g}%)'.format(vot[2]+1), 'Extreme ({0:.1g}%)'.format(vot[3])),fontsize = 12);
mm = df_munis_gv['yes_no_ratio'].mean();
plt.text(0.15,7,r'Ratio: {:,.2g}'.format(mm[0]));plt.text(1.15,7,r'Ratio: {:,.2g}'.format(mm[1]))
plt.text(2.15,7,r'Ratio: {:,.2g}'.format(mm[2]));plt.text(3.10,7,r'Ratio: {:,.3g}'.format(mm[3]));
# Figure 5 conveys a conclusive message: **Colombians living in _High_ and _Extreme_ vulnerable municipalities across the country remarkably supported the peace agreement in the ballot**. This means that victims who unfortunately have suffered the most the stark consequences of this war (e.g. massacres, land displacement, child recruitment), are willing to back the government efforts to terminate the long-lasting armed conflict. However, the population in these zones only represents 6% of voters, therefore, people living in less vulnerable regions are the ones who actually made the final decision. While the average YES/NO ratio was also greater than one in Low and Medium vulnerable municipalities, negative votes casted primarily in the country’s center (i.e. urban areas) slightly surpassed the positive scenario portrayed specially in vulnerable zones (i.e. rural areas).
#
# Furthermore, the YES/NO ratio is also correlated with data of poverty and electoral participation (i.e. voters/total_voters). Figure 6 and 7 illustrate this relation at the regional level for both variables respectively. **There is a positive correlation between the ratio of YES/NO votes and the poverty rate per region**. This result is coherent with the former claims, as people in least favorable regions from a socio-economic standpoint (e.g. rural areas or low income cities), were keener towards the agreement than people living in areas whit lower poverty rates. On the other hand, the YES/NO ratio also renders a negative correlation with the electoral participation, meaning that as votes grew in magnitude the negative share strengthened.
rho = df_regions.corr(method='pearson');fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(15,5))
ax1=sns.regplot(x='poverty',y='yes_no_ratio',data=df_regions,ci=False,scatter_kws={"s": 50, "alpha": 1,'color':'grey'},ax=axes[0])
ax1.text(0.1,3,r'$\rho = {0:.3g}$'.format(rho['yes_no_ratio'][10]),fontsize=20);
ax1.set_title('Fig. 6: Relation of YES/No ratio and poverty rate',fontsize=14)
ax2=sns.regplot(x='elec_part',y='yes_no_ratio',data=df_regions,ci=False,scatter_kws={"s": 50, "alpha": 1,'color':'grey'},ax=axes[1])
ax2.text(0.12,3,r'$\rho = {0:.4g}$'.format(rho['yes_no_ratio'][14]),fontsize=20);
ax2.set_title('Fig. 7: Relation of YES/No ratio and electoral participation',fontsize=14);
# ## Did people actually vote?
# Short answer: Not really!. Figure 8 shows the histogram of electoral participation at the municipal/city level. **Approximately 21 million voters didn’t show up to the ballot, representing 75% of the electoral population**. The maximum participation rate was 62% and the minimum 3%. To provide a reference, in the 2014 presidential elections the participation rate was about 40%. Several political analyst have associated this lackluster electoral participation with the low levels of popularity held by the current government. Another likely reason is the unfortunate effect of <NAME> in its journey near the North-cost during that weekend.
#
# To assess the likelihood of this meteorological event affecting the electoral participation rate, a dummy variable named `h_affect` is created to identify the regions and municipalities harmed by the pass of Hurricane Matthew.
# +
# Identifying regions affected by hurricane Mathews
dpts = ['ATLANTICO', '<NAME>','<NAME>','BOLIVAR', 'MAGDALENA']
df_regions['h_affect'] = [1 if i in dpts else 0 for i in df_regions.index]
df_munis['h_affect'] = [1 if i in dpts else 0 for i in df_munis['region']]
# Distribution of electoral participation
plt.figure(figsize=(10,5))
sns.distplot(df_munis["elec_part"],fit=norm, kde=False)
plt.title('Fig. 8: Distribution and stats of electoral participation', fontsize = 14)
plt.text(0.0001,5,r'Total elegible voters: {:,}'.format(np.sum(df_munis['total_voters'])))
plt.text(0.0001,4.7,r'Total missing votes: {:,}'.format(np.sum(df_munis['total_voters']-df_munis['voters'])))
plt.text(0.0001,4.4,r'Avg. participation: {0:.3g}%'.format(np.mean(df_munis['elec_part'])*100))
plt.text(0.0001,4.1,r'Std: {0:.3g}'.format(np.std(df_munis['elec_part'])))
plt.text(0.0001,3.8,r'Max: {0:.3g}%'.format(np.max(df_munis['elec_part'])*100))
plt.text(0.0001,3.5,r'Min: {0:.3g}%'.format(np.min(df_munis['elec_part'])*100))
# Plot electoral participation across regions
part = df_regions["elec_part"]; part = part.sort_values(ascending = False)
plt.figure(figsize=(20,5))
bar_plot1 = plt.bar(range(len(part)),part.values, 0.6); plt.xticks(np.arange(len(part))+0.3, part.index, rotation = 90);
[bar_plot1[i].set_color('orange') if v in dpts else '' for i,v in enumerate(part.index)];
plt.legend(['Not Affected by <NAME>', "Affected by <NAME>"], fontsize = 14);
plt.title('Fig. 9: Electoral participation across regions', fontsize = 14)
# Bottom 10% of municipalities/cities in terms of electoral participation
part = df_munis["elec_part"]; part = part.sort_values(ascending = False)[-100:]
plt.figure(figsize=(20,5))
bar_plot1 = plt.bar(range(len(part)),part.values, 0.6); plt.xticks(np.arange(len(part))+0.3, part.index, rotation = 90);
[bar_plot1[i].set_color('orange') if df_munis.loc[v,'h_affect'] == 1 else '' for i,v in enumerate(part.index)];
plt.title('Fig. 10: Bottom 10% of municipalities/cities in terms of electoral participation', fontsize = 14)
plt.legend(["Affected by <NAME>"], fontsize = 14);
# -
# Figures 9 shows the electoral participation rate at the regional level in a descending order. Orange colored bars correspond to the areas affected by the hurricane. Noticeably, four out of the eight regions with the lowest participation were coastal regions likely blocked-off because of the hurricane. At the municipal level, Figure 10 presents the bottom 10% of municipalities in terms of participation, and the impacted zones account for a big chunk as well. The average participation in this zones was _25%_ which contrats with the _35%_ national average. Moreover, the YES/NO ratio in those zones was _2.7x_ compared to _1.8x_ in the rest. **Therefore, given the slim difference between “YES” and “NO” votes, it could be argued that the Hurricane Mathew did have an effect in the electoral participation and such effect might in turn signified a tipping point in the final outcome of the plebiscite.**
#
# Whatever the underlying reasons were for this drab participation rate, the fact is that it is rather disappointing and discouraging given the importance of what was at stake.
#
# ## Main takeaways
# While a deeper study is required to further break-down these results, some conclusions can be drawn from the above analysis:
# - Colombians rejected the peace agreement by a margin less than 0.2% of total votes. Even though on average the ratio of YES/NO votes was greater than one municipally-wise, positive votes weren't enough overall to outweigh negative votes.
# - People living in _High_ and _Extreme_ vulnerable municipalities largely supported the peace agreement. Likewise, data shows a positive correlation between the YES/NO ratio and poverty rate region-wise.
# - Electoral participation was very low with 75% of absence.
# - Data suggest that Hurricane Mathew did have a negative effect in this participation rate, and it might entailed a turning point in the plebiscite’s outcome given thin margin of difference.
#
# As you read this tutorial, the Colombian government and FARC are sifting through the proposals submitted by opposition parties, seeking to incorporate them in a re-negotiated agreement. Colombians are eagerly awaiting for this process as a country loudly yelling that this conflict cannot be bore any longer.
#
# ## References
# - (In spanish) http://lasillavacia.com/silla-blanca
# - https://internationaljusticeathunter.wordpress.com/2012/03/18/brief-summary-of-armed-conflict-in-colombia-3/
# - https://www.washingtonpost.com/news/monkey-cage/wp/2016/10/01/sunday-colombians-vote-on-the-historic-peace-accord-with-the-farc-heres-what-you-need-to-know/
# - https://www.wola.org/analysis/ending-50-years-of-conflict-in-colombia-a-new-report-from-wola/
| 2016/tutorial_final/59/2016_Colombian_plebiscite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit
# language: python
# name: python37664bit52c43a8b57e04c11b7ae243a4050fcee
# ---
# +
import time
import numpy as np
x1 = [9, 2, 5, 0, 0, 7, 5, 0, 0, 0, 9, 2, 5, 0, 0]
x2 = [9, 2, 2, 9, 0, 9, 2, 5, 0, 0, 9, 2, 5, 0, 0]
### CLASSIC DOT PRODUCT OF VECTORS IMPLEMENTATION ###
tic = time.process_time()
dot = 0
for i in range(len(x1)):
dot += x1[i] * x2[i]
toc = time.process_time()
print ("dot = " + str(dot) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
### CLASSIC OUTER PRODUCT IMPLEMENTATION ###
tic = time.process_time()
outer = np.zeros((len(x1), len(x2))) # we create a len(x1)*len(x2) matrix with only zeros
for i in range(len(x1)):
for j in range(len(x2)):
outer[i,j] = x1[i] * x2[j]
toc = time.process_time()
print ("outer = " + str(outer) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
### CLASSIC ELEMENTWISE IMPLEMENTATION ###
tic = time.process_time()
mul = np.zeros(len(x1))
for i in range(len(x1)):
mul[i] = x1[i] * x2[i]
toc = time.process_time()
print ("elementwise multiplication = " + str(mul) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
### CLASSIC GENERAL DOT PRODUCT IMPLEMENTATION ###
W = np.random.rand(3,len(x1)) # Random 3*len(x1) numpy array
tic = time.process_time()
gdot = np.zeros(W.shape[0])
for i in range(W.shape[0]):
for j in range(len(x1)):
gdot[i] += W[i,j] * x1[j]
toc = time.process_time()
print ("gdot = " + str(gdot) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
# +
### VECTORIZED DOT PRODUCT OF VECTORS ###
tic = time.process_time()
dot = np.dot(x1,x2)
toc = time.process_time()
print ("dot = " + str(dot) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
### VECTORIZED OUTER PRODUCT ###
tic = time.process_time()
outer = np.outer(x1,x2)
toc = time.process_time()
print ("outer = " + str(outer) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
### VECTORIZED ELEMENTWISE MULTIPLICATION ###
tic = time.process_time()
mul = np.multiply(x1,x2)
toc = time.process_time()
print ("elementwise multiplication = " + str(mul) + "\n ----- Computation time = " + str(1000*(toc - tic)) + "ms")
### VECTORIZED GENERAL DOT PRODUCT ###
tic = time.process_time()
dot = np.dot(W,x1)
toc = time.process_time()
print ("gdot = " + str(dot) + "\n ----- Computation time = " + str(1000 * (toc - tic)) + "ms")
| NN_DL(coursera)/Week2/vectorization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''upp'': conda)'
# metadata:
# interpreter:
# hash: 4c6c1704d8a03e4775c7322f86036425ff380d055473196d495e2bd41cd3b29a
# name: python3
# ---
# Percentiles
#
# In a data set, a given percentile is the point at which that percent of the data is less than the point we are.
#
# Example : income distribution
#
# We got some income data of the country and we sorted the data by income. The 99th percentile will be the income amount that which 99% of the rest was making less than that, at the same time that means 1 percenter makes more than that.
# Percentiles in a normal distribution
#
# Q1, Q3 (quartiles) are the points that contain together 50% of the data, like 25% each side of the median.
#
# IQR (Inter quartile range) is the erea of the distribution that contains 50% of the data
#
# For the practice, we will generate some random normal distributed data points, and then compute the Percentiles.
# +
import numpy as np
import matplotlib.pyplot as plt
vals = np.random.normal(0, 0.5, 10000)
# we generate 10000 data points centered around 0 (mu= 0) and with a std = 0.5
plt.hist(vals, 50)
plt.show()
# -
np.percentile(vals, 50) # this is the 50th percentile value (median percentile). this represent the point where 50% of the data is less than that value above
np.percentile(vals, 90) # 90% of the data is less than 0.627
np.percentile(vals, 20) # 20% of the data is less than -0.4 and 80% is greater
# If we want to figure out where breaking data are in the data set, the percentile is the measure to use
# Moments
#
# Basically, we use moments to measure the shape of a distribution (here we talk about pdf)
#
# * The first moment of the data set is the mean of the data we're looking at.
# * The second moment of the data set is the variance
# * The third moment is "skew" of how "lopsided" is the distribution.
#
# - A distribution with a longer tail on the left will be skewed left and have a negative skew
# - A distribution with a longer tail on the right will be skewed right and have a positive skew
#
# * The fourth moment is "kurtosis" or how thick is the tail, and how sharp is the peak, compared to a normal distribution.
#
# Higher peaks have higher kurtosis
# The first moment is the mean
#
# This data should average out to about 0:
np.mean(vals)
# The second moment is the variance
np.var(vals)
# The third moment is the skew
#
# Since our data is nicely centered around 0, it should be almost 0:
import scipy.stats as sp
sp.skew(vals)
# The fourth momentg is "kurtosis"
#
# It discribes the shape of the tail. For a normal distribution, this is 0:
sp.kurtosis(vals)
| notebooks/statistics/percenctal_moments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hungrycarpet/Quakedet/blob/main/speech_recognition_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="gAb7do6uQAAT"
import speech_recognition as sr
import pyaudio
r=sr.Recognizer()
mic=sr.Microphone()
mic=sr.Microphone(device_index=3)
with mic as source:
r.adjust_for_ambient_noise(source)
audio=r.listen(source)
string=r.recognize_google(audio)
| speech_recognition_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.formula.api as smf
import warnings
warnings.filterwarnings('ignore')
sns.set_theme()
plt.rcParams['figure.figsize'] = [8,8]
africa = pd.read_csv("../datasets/africa.csv")
n_countries = len( africa )
n_countries
africa
sns.scatterplot(data=africa, x="Literacy", y="IMR")
plt.title("Infant mortality rates (IMR) versus Literacy for young African women")
# plt.savefig("literacy_imr.png")
x_bar = africa.Literacy.mean()
x_bar
y_bar = africa.IMR.mean()
y_bar
sns.scatterplot(data=africa, x="Literacy", y="IMR")
plt.axvline( x_bar, linestyle='--')
plt.axhline( y_bar, linestyle='--')
plt.title("IMR vs Literacy: quadrants determined by averages")
# plt.savefig("quadrants.png")
africa_corr = africa.Literacy.corr( africa.IMR )
africa_corr
sns.scatterplot(data=africa, x="Literacy", y="IMR")
# null model:
plt.axhline( y_bar, color="black")
plt.title("IMR vs Literacy: Null model")
# plt.savefig("africa_null.png")
africa_tss = ((africa.IMR - y_bar)**2).sum()
africa_tss
africa.IMR.var()
(n_countries - 1)*africa.IMR.var()
s_x = africa.Literacy.std()
s_x
s_y = africa.IMR.std()
s_y
m_sd = -s_y / s_x
m_sd
b_sd = y_bar - m_sd*x_bar
b_sd
sns.scatterplot(data=africa, x="Literacy", y="IMR")
# SD line
plt.axline( (x_bar, y_bar), slope = m_sd, linestyle='--', color="black")
plt.title("IMR vs Literacy: SD line")
# plt.savefig("literacy_imr_sdline.png")
# +
# africa = africa.assign( null_resid = africa.IMR - y_bar, sd_pred = b_sd + m_sd*africa.Literacy, sd_resid = africa.IMR - b_sd - m_sd*africa.Literacy)
# -
m = africa_corr*(s_y/s_x)
m
b = y_bar - m*x_bar
b
sns.scatterplot(data=africa, x="Literacy", y="IMR")
# line of best fit
plt.axline( (x_bar,y_bar), slope = m, color="black")
plt.title("IMR vs Literacy: Line of best fit")
# plt.savefig("literacy_imr_regression_line.png")
sns.scatterplot(data=africa, x="Literacy", y="IMR")
plt.axline( (x_bar,y_bar), slope = m, color="black")
plt.axline( (x_bar, y_bar), slope = m_sd, linestyle='--', color="black")
plt.title("IMR vs Literacy: Regression line (solid) and SD line (dashed)")
plt.plot(x_bar,y_bar,'ok')
# plt.savefig("literacy_imr_both_lines.png")
africa_model = smf.ols('IMR ~ Literacy', data = africa)
africa_fit = africa_model.fit()
africa_fit.params
africa_null = smf.ols('IMR ~ 1', data = africa)
africa_null = africa_null.fit()
africa_null.params
print( africa_fit.summary() )
africa_ssr = (africa_fit.resid**2).sum()
africa_ssr
africa_fit.ssr
africa_ssr / (n_countries - 2)
np.sqrt( africa_ssr / (n_countries - 2) )
africa_fit.scale
africa_rse = np.sqrt( africa_fit.scale )
africa_rse
africa_fit.rsquared
1 - (africa_ssr / africa_tss)
sns.residplot(data=africa, x="Literacy", y="IMR", line_kws={"color":"black"})
plt.axhline( 2*africa_rse, linestyle=":", color="black")
plt.axhline( -2*africa_rse, linestyle=":", color="black")
plt.title("IMR versus Literacy residuals")
# plt.savefig("literacy_imr_residuals.png")
import scipy.stats as st
st.norm.ppf(.975 )
st.t.ppf(.975, df=45)
st.t.ppf(.995, df=45)
2*st.t.cdf(-5.26,df=45)
africa_fit.conf_int()
new = pd.DataFrame(data = {"Literacy":[50,80]})
africa_fit.predict( new )
africa_fit.get_prediction( new ).summary_frame()
sns.regplot(data=africa, x="Literacy", y="IMR", line_kws={"color":"black"})
plt.title("IMR vs Literacy with confidence bands")
# plt.savefig("literacy_imr_with_ci.png")
x = africa.Literacy.sort_values(ignore_index=True)
endpts = africa_fit.get_prediction(x).summary_frame()
sns.regplot(data=africa, x="Literacy", y="IMR", line_kws={"color":"black"})
plt.plot(x, endpts.obs_ci_upper, ':', color="black")
plt.plot(x, endpts.obs_ci_lower, ':', color="black")
plt.title("IMR vs Literacy with confidence and prediction bands")
# plt.savefig("literacy_imr_ci_preds.png")
| jupyter_notebooks/.ipynb_checkpoints/africa-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Groupby
#
# **The groupby method allows you to group rows of data together and call aggregate functions.**
# ## What is Groupby ?
#
# > In case you don't have SQL experience, we'll explain what groupby does.
# Simply put, Groupby **allows you to group together rows based off of a column and perform an aggregate function on them.**
#
# # **Let's consider the image below as an example :**
# 
# * In the image above we have 3 Partitions of ID's 1, 2 and 3.
# * We can groupby the ID column and aggregate them using some sort of aggregate function.
# * Aggregate function is sort of a fancy term for any function that takes in many values and splits out a single value. Other examples include :
# 1. Taking the sum of input values and outputting the result.
# 2. Average
# 3. Standard Deviation etc.
# * **Choose a column to groupby >> Gather all those rows together >> Perform some sort of agg. function on it.**
#
# # Groupby with Pandas
import numpy as np
import pandas as pd
# Create dataframe
data = {'Company':['GOOG','GOOG','MSFT','MSFT','FB','FB'],
'Person':['Sam','Charlie','Amy','Vanessa','Carl','Sarah'],
'Sales':[200,120,340,124,243,350]}
# +
# For having gridlines
# -
# %%HTML
<style type="text/css">
table.dataframe td, table.dataframe th {
border: 1px black solid !important;
color: black !important;
}
df = pd.DataFrame(data) # Metadata with company column with 3 company codes, Person col, Sales column
df
# We can now use groupby method to group rows together based off of a column name.
df.groupby('Company') # Points to where this is stored in memory.
# DataFrame_name.groupby('col_name')
byComp = df.groupby('Company') # Store the memory location in a variable.
# Calling some sort of aggregate function on the variable holding the memory location.
# To get mean by company
byComp.mean() # Looks at column and gives us average sales by the company.
# It can't perform mean on column Person as it is of string type, and mean wouldn't make sense.
byComp.sum()
byComp.std()
byComp.sum().loc['FB'] # As we get a DataFrame in return, so calling FB sum of sales.
# Calling in one line
df.groupby('Company').sum().loc['GOOG']
# **Other useful aggregate functions :**
# * count : Counts the number of instances per column.
# * max : Returns the maximum number from number column, and alphabetically max value from string type column.
# * min : Returns the minimum number from number column, and alphabetically min value from string type column.
df.groupby('Company').count()
df.groupby('Company').max()
df.groupby('Company').min()
# **A lot of times we can use groupby with the describe method and it gives us useful info in one go.**
df.groupby('Company').describe()
df.groupby('Company').describe().transpose()
#Call Single Companies using following
# df.groupby('Company').describe().transpose()['FB']
# **You just call the DataFrame ==> Specify .groupby ==> ('Column_you_want_to_group_by').aggregate_method() making the entire syntax for using Groupby = DataFrame.groupby('Column_you_want_to_group_by').aggregate_method()**
| 06. Python for Data Analysis - Pandas/6.5 groupby.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import the necessary packages
from tensorflow.keras.models import load_model
from skimage import transform
from skimage import exposure
from skimage import io
from imutils import paths
import numpy as np
import argparse
import imutils
import random
import cv2
import os
from tkinter import *
from tkinter import ttk
print("[INFO] loading model...")
model = load_model("output/trafficsignnet.model")
labelNames = open("signnames.csv").read().strip().split("\n")[1:]
labelNames = [l.split(",")[1] for l in labelNames]
print("[INFO] predicting...")
imagePaths = list(paths.list_images("gtsrb-german-traffic-sign/Test"))
random.shuffle(imagePaths)
imagePaths = imagePaths[:25]
imagePaths = ['gtsrb-german-traffic-sign/Test\\00067.png']
label = "null"
# loop over the image paths
for (i, imagePath) in enumerate(imagePaths):
# load the image, resize it to 32x32 pixels, and then apply
# Contrast Limited Adaptive Histogram Equalization (CLAHE),
# just like we did during training
image = io.imread(imagePath)
image = transform.resize(image, (32, 32))
image = exposure.equalize_adapthist(image, clip_limit=0.1)
# preprocess the image by scaling it to the range [0, 1]
image = image.astype("float32") / 255.0
image = np.expand_dims(image, axis=0)
# make predictions using the traffic sign recognizer CNN
preds = model.predict(image)
j = preds.argmax(axis=1)[0]
label = labelNames[j]
print(label)
# load the image using OpenCV, resize it, and draw the label
# on it
image = cv2.imread(imagePath)
image = imutils.resize(image, width=128)
while(True):
cv2.putText(image, label, (5, 15), cv2.FONT_HERSHEY_SIMPLEX,
0.45, (0, 255, 0), 2)
cv2.imshow("image", image)
key = cv2.waitKey(1)
if key & 0xFF == ord('q'):
break
p = os.path.sep.join(["examples", "{}.png".format(i)])
cv2.imwrite(p, image)
cv2.destroyAllWindows()
#variable that stores current speed value
speed = 0
#variable that stores speed limit value
speed_lim = int(label[13:15])
print(speed_lim)
speed_json = {"speed_limit" : speed_lim}
with open('person.txt', 'w') as json_file:
json.dump(person_dict, json_file)
#creating the main window
root = Tk()
#changing background color to grey
root.configure(background='grey')
speedLimLabel = Label(root, text = "Current Speed limit = " + str(speed_lim) + " Km/hr", fg = "white", bg = "grey", padx = 20, pady = 40)
speedLimLabel.config(font=("Courier", 44))
speedLimLabel.grid(row = 0, sticky = 'w')
ttk.Separator(root,orient=HORIZONTAL).grid(row=1, columnspan=5, sticky = 'ew')
currentSpeedLabel = Label(root, text = str(speed) + " Km/hr", fg = "white", bg = "grey", padx = 20, pady = 40)
currentSpeedLabel.config(font=("Courier", 44))
currentSpeedLabel.grid(row = 2, sticky = 'w')
#increaments speed by 5 on left click and checks if speed is breached
def leftClick(event):
global speed
speed = speed + 5
currentSpeedLabel['text'] = str(speed) + " Km/hr"
if speed > speed_lim:
warningLabel['bg'] = "red"
warningLabel['text'] = "Speed Limit breached"
else:
warningLabel['bg'] = "white"
warningLabel['text'] = "Under Speed Limit"
#decreaments speed by 5 on left click and checks if speed is under normal
def rightClick(event):
global speed
if speed > 0:
speed = speed - 5
else:
speed = speed
currentSpeedLabel['text'] = str(speed) + " Km/hr"
if speed > speed_lim:
warningLabel['bg'] = "red"
warningLabel['text'] = "Speed Limit breached"
else:
warningLabel['bg'] = "white"
warningLabel['text'] = "Under Speed Limit"
ttk.Separator(root,orient=HORIZONTAL).grid(row=3, columnspan=5, sticky = 'ew')
warningLabel = Label(root, text = "Under Speed Limit", fg = "black", bg = "white", padx = 20, pady = 40)
warningLabel.config(font=("Courier", 44))
warningLabel.grid(row = 4)
#calls left click function when anywhere left clicked on the window
root.bind('<Button-1>', leftClick)
#calls right click function when anywhere left clicked on the window
root.bind('<Button-3>', rightClick)
#runs the window until closed
root.mainloop()
# -
| src/road sign with python gui.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # FABRIC API Example: Create a Simple Slice
#
#
# ## Configure the Environment
import os
from fabrictestbed.slice_manager import SliceManager, Status, SliceState
import json
# +
ssh_key_file_priv=os.environ['HOME']+"/.ssh/id_rsa"
ssh_key_file_pub=os.environ['HOME']+"/.ssh/id_rsa.pub"
ssh_key_pub = None
with open (ssh_key_file_pub, "r") as myfile:
ssh_key_pub=myfile.read()
ssh_key_pub=ssh_key_pub.strip()
# +
credmgr_host = os.environ['FABRIC_CREDMGR_HOST']
print(f"FABRIC Credential Manager : {credmgr_host}")
orchestrator_host = os.environ['FABRIC_ORCHESTRATOR_HOST']
print(f"FABRIC Orchestrator : {orchestrator_host}")
# -
# ## Create Slice Manager Object
# Users can request tokens with different Project and Scopes by altering project_name and scope parameters in the refresh call below.
#
# +
slice_manager = SliceManager(oc_host=orchestrator_host, cm_host=credmgr_host, project_name='all', scope='all')
# Initialize the slice manager
slice_manager.initialize()
# + [markdown] tags=[]
# ## Configure Slice Parameters
#
#
# -
slice_name='MySlice'
node_name='node1'
site='RENC'
image_name='default_centos_8'
image_type='qcow2'
# ## Build the Slice
# +
from fabrictestbed.slice_editor import ExperimentTopology, Capacities, ComponentType, ComponentModelType, ServiceType
# Create topology
experiment = ExperimentTopology()
# Add node
node = experiment.add_node(name=node_name, site=site)
# Set capacities
cap = Capacities()
cap.set_fields(core=2, ram=16, disk=100)
# Set Properties
node.set_properties(capacities=cap, image_type=image_type, image_ref=image_name)
# -
# ## Submit the Slice
# +
# Generate Slice Graph
slice_graph = experiment.serialize()
# Request slice from Orchestrator
return_status, slice_reservations = slice_manager.create(slice_name=slice_name,
slice_graph=slice_graph,
ssh_key=ssh_key_file_pub)
if return_status == Status.OK:
slice_id = slice_reservations[0].get_slice_id()
print("Submitted slice creation request. Slice ID: {}".format(slice_id))
else:
print(f"Failure: {slice_reservations}")
# -
# ## Query Slices
#
# You can get a list of all your slices from the slice manager. If this is your first slice, it should return only one slice.
#
# Note that the status returned by the call to slices indicates the success or failure of the call to the FABRIC control framework. The status is not the status of the slices. The status of each slice is included in the list of slices.
# +
return_status, slices = slice_manager.slices(excludes=[SliceState.Dead])
if return_status == Status.OK:
for slice in slices:
print("{}:".format(slice.slice_name))
print(" ID : {}".format(slice.slice_id))
print(" State : {}".format(slice.slice_state))
print(" Lease End : {}".format(slice.lease_end))
print()
else:
print(f"Failure: {slices}")
# -
# ## Get the New Slice
#
# You new slice is in the list of all your slices. You can loop through the list of slices to get the new slice. Python has a standard tool to filter lists. Try using a lambda function to filter out your slice using its name.
# +
slice = list(filter(lambda x: x.slice_name == slice_name, slices))[0]
print("Slice Name : {}".format(slice.slice_name))
print("ID : {}".format(slice.slice_id))
print("State : {}".format(slice.slice_state))
print("Lease End : {}".format(slice.lease_end))
# -
# ## Delete Slice
# +
return_status, result = slice_manager.delete(slice_object=slice)
print("Response Status {}".format(return_status))
print("Response received {}".format(result))
# -
| fabric_examples/basic_examples/create_slice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: U4-S2-NNF (Python3)
# language: python
# name: u4-s2-nnf
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Jaydenzk/DS-ML-Build-Week/blob/master/NN_Model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab_type="code" id="pcxEynIzRAOm" colab={}
import numpy as np
import pandas as pd
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import GridSearchCV
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from sklearn.metrics import mean_absolute_error
# + colab_type="code" id="0bSoxhNx7ZFr" outputId="4385cdff-88fc-4c0c-9a9a-dcb5c9979cab" colab={"base_uri": "https://localhost:8080/", "height": 222}
train = pd.read_csv("/content/test_prop_2017.csv")
test = pd.read_csv("/content/train_prop_2017.csv")
print(train.shape)
#df = df.drop(columns=['Unnamed: 0'])
train.head()
# + colab_type="code" id="ovdZs4VN_vmg" outputId="89938f1c-900b-4879-9933-4134f3a8e23b" colab={"base_uri": "https://localhost:8080/", "height": 222}
print(test.shape)
test.head()
# + colab_type="code" id="lh6C3Hx97ZIr" colab={}
X_train = train[['bathroomcnt', 'bedroomcnt', 'calculatedfinishedsquarefeet',
'yearbuilt']].values
y_train = train['taxvaluedollarcnt'].values
X_test = test[['bathroomcnt', 'bedroomcnt', 'calculatedfinishedsquarefeet',
'yearbuilt']].values
y_test = test['taxvaluedollarcnt'].values
# + colab_type="code" id="d5XOOV0g7pG_" colab={}
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
#X = scaler.fit_transform(X)
# + colab_type="code" id="5EQFLwHO7pLh" outputId="8c9230bd-ecf0-4f42-f7d1-ea20e6ad5635" colab={"base_uri": "https://localhost:8080/", "height": 35}
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# + colab_type="code" id="gNbou5w5RLxM" outputId="8eb1c6b0-8d68-4550-b8f3-67c0d9379488" colab={"base_uri": "https://localhost:8080/", "height": 269}
nnmodel = Sequential()
# Input => Hidden
nnmodel.add(Dense(32, input_dim=4, activation='relu'))
# Hidden
nnmodel.add(Dense(32, activation='relu'))
#nnmodel.add(Dense(32, activation='relu'))
#nnmodel.add(Dense(32, activation='relu'))
# Output Layer
nnmodel.add(Dense(1, activation='linear'))
# Compile
nnmodel.compile(loss='mean_absolute_error',
optimizer='adam',
metrics=['mean_absolute_error'])
nnmodel.summary()
# + colab_type="code" id="T0NZCyb6RP6F" outputId="c5cae5a6-4493-44b2-f8d4-835daf5258e3" colab={"base_uri": "https://localhost:8080/", "height": 431}
nnmodel.fit(X_train, y_train, epochs=10, batch_size=32, validation_split=.3, verbose=1)
scores = nnmodel.evaluate(X_test, y_test)
print(f'{nnmodel.metrics_names[1]}: {scores[1]*100}')
# + colab_type="code" id="iFqjmheFRP8W" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="ddead809-6f69-490f-8adc-99d2f40697cc"
# Function to create model
def create_model():
# create model
model = Sequential()
# Input => Hidden
model.add(Dense(32, input_dim=4, activation='relu'))
# Hidden
model.add(Dense(32, activation='relu'))
model.add(Dense(32, activation='relu'))
model.add(Dense(32, activation='relu'))
# Output Layer
model.add(Dense(1, activation='linear'))
# Compile
model.compile(loss='mean_absolute_error',
optimizer='adam',
metrics=['mean_absolute_error'])
return model
model = KerasRegressor(build_fn=create_model, verbose=1)
# define the grid search parameters
param_grid = {'batch_size': [10, 20],
'epochs': [10]}
# Create Grid Search
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=5)
grid_result = grid.fit(X_train, y_train)
# + id="JlLmh1nbgD8_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="d430a54f-cea0-484f-85cc-45a62900e817"
# Report Results
print(f"Best: {grid_result.best_score_} using {grid_result.best_params_}")
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print(f"Means: {mean}, Stdev: {stdev} with: {param}")
# + colab_type="code" id="w8WinkXiRP-d" colab={}
def baseline_model(optimizer='adam'):
# create model
model = Sequential()
model.add(Dense(32, activation='relu',
kernel_regularizer = 'l2',
kernel_initializer = 'normal',
input_shape=(4,)))
# model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(32, activation='relu',
kernel_regularizer = 'l2',
kernel_initializer = 'normal'))
#model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(1, activation='linear',
kernel_regularizer = 'l2',
kernel_initializer='normal'))
model.compile(loss='mean_absolute_error', optimizer=optimizer, metrics=['mean_absolute_error'])
return model
def gridSearch_neural_network(X_train, y_train):
print("Train Data:", X_train.shape)
print("Train label:", y_train.shape)
# evaluate model with standardized dataset
estimator = KerasRegressor(build_fn=baseline_model, nb_epoch=10, batch_size=5, verbose=1)
# grid search epochs, batch size and optimizer
optimizers = ['rmsprop', 'adam']
#dropout_rate = [0.0, 0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]
init = ['glorot_uniform', 'normal', 'uniform']
epochs = [50]
batches = [5, 10, 20]
weight_constraint = [1, 2, 3, 4, 5]
param_grid = dict(optimizer=optimizers,
#dropout_rate=dropout_rate,
epochs=epochs,
batch_size=batches,
#weight_constraint=weight_constraint,
#init=init
)
grid = GridSearchCV(estimator=estimator, param_grid=param_grid)
grid_result = grid.fit(X_train, y_train)
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + id="RLjSS5LPLKtE" colab_type="code" outputId="50ea92cb-13a0-4c72-c481-809a7a1c17f3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
gridSearch_neural_network(X_train, y_train)
# + colab_type="code" id="zjw9V6UARQCI" colab={}
import pickle
# + colab_type="code" id="gPW59MTv73yv" colab={}
pickle.dump(model, open('zillow_nn_model.pkl','wb'))
| NN_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
from Bio.Blast import NCBIWWW
fasta_string = 'MPEVTGCAATVSGRVWDSLHLLVDAGVDLQLRTTLWRDSVISQHLPELQHLVSEQGFDLVIQQARAADGSPFQLV'
result_handle = NCBIWWW.qblast("blastp", "nr", fasta_string)
from Bio.Blast import NCBIXML
blast_record = NCBIXML.read(result_handle)
E_VALUE_THRESH = 0.04
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
if hsp.expect < E_VALUE_THRESH:
print('****Alignment****')
print('sequence:', alignment.title)
print('length:', alignment.length)
print('e value:', hsp.expect)
print(hsp.query[0:75] + '...')
print(hsp.match[0:75] + '...')
print(hsp.sbjct[0:75] + '...')
# +
## now you should run the smith-waterman alignment on the top scoring blast hits (to locally align your protein of interest with the blastp hits)
# -
import swalign
match = 2
mismatch = -1
scoring = swalign.NucleotideScoringMatrix(match, mismatch)
sw = swalign.LocalAlignment(scoring)
alignment = sw.align(fasta_string,hsp.match)
alignment.dump()
# +
read_to_struct = []
E_VALUE_THRESH = 0.04
for alignment in blast_record.alignments:
for hsp in alignment.hsps:
if hsp.expect < E_VALUE_THRESH:
sw_tmp = sw.align(fasta_string,hsp.match)
score = sw_tmp.score
desc = alignment.title.split('|')[4]
ref = alignment.title.split('|')[3]
id_type = alignment.title.split('|')[2]
org = alignment.title.split('[')[1].split(']')[0]
matches = sw_tmp.matches
read_to_struct.append({'desc':desc, 'id_type':id_type ,'id':ref, 'organism':org, 'sw-score':score,'matches':matches, 'e_val':hsp.expect, 'alignment_length':alignment.length})
DF_alignment = pd.DataFrame(read_to_struct)
DF_alignment.head(2)
# +
# you might need to do a sensitivity check here- to see if the top scoring (sw-score) also has the lowest e val.
# +
#Choose the top score as the gene id to go on to the next one...
# -
| Biopython_blast_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Wrapper for CMR
# `A python library to interface with CMR - Collection Search Demo`
#
# This demo will show how to preform a **collection** search against CMR while inside a notebook.
# ## Loading the library
# From the command line, make sure you call `runme.sh -p -i` to both backage and install the library through pip3.n
# ## Load modules
import cmr.search.collection as coll
# ## Get Online Help
#
# At least some understanding of the CMR API will be needed from time to time, to assist with that the following call can be used to open a browser window to the API. For the fun of it, you can pass in an HTML anchor tag on the page and jump directly there.
coll.open_api()
# ## Searching
#
# ### Perform A Basic Searches
# Search for all records that contain the word 'salt'.
results = coll.search({'keyword':'salt'})
print("Found {} records.".format(len(results)))
for i in results:
print (i)
# ### A Search with a columns filtered from result
# Reduce the result columns by only showing the collection curration fields and drop the entry title.
#
# This search also searches UAT
# +
params = {}
#params['provider'] = 'SEDAC' # 276 records
#params['keyword'] = 'fish food' # 131 records
params['keyword'] = 'salt' # 290 records
config={'env':'uat'} # 290 in prod, 49 in UAT as of 2020-12-01
results = coll.search(params,
filters=[coll.collection_core_fields,
coll.drop_fields('EntryTitle')],
limit=1000,
config=config)
print("Found {} records.".format(len(results)))
for i in results:
print (i)
# -
# ### Find a lot of collection records
# This should find just over a full page (2000) of results.
params = {}
results = coll.search(params,
filters=[coll.collection_core_fields, coll.drop_fields('EntryTitle')],
limit=2048,
config={'env':'uat'})
print("Found {} records.".format(len(results)))
for i in results:
print (i)
# ## Applying Filters after a search
# Internally the code calls apply_filters() but it can be called manually as show below. One reason to do this is to download the data once and then apply filters as needed.
params = {}
raw_results = coll.search(params, limit=2, config={'env':'uat'})
# +
clean_results = coll.apply_filters([coll.collection_core_fields,coll.drop_fields('EntryTitle')], raw_results)
print("Found {} records.".format(len(clean_results)))
for i in clean_results:
print (i)
# -
# ## Sorting
# +
def sorted_search(params):
results = coll.search(params, filters=[coll.collection_core_fields], limit=11)
print("Found {} records.".format(len(results)))
for i in results:
print (i)
#params = {'keyword':'modis', 'sort_key': 'instrument'}
sorted_search({'keyword':'modis', 'sort_key': 'instrument'})
print('\nvs\n')
sorted_search({'keyword':'modis', 'sort_key': '-instrument'})
# -
# ### Help with Sort Keys
# Can not remember the sort keys, look them up
coll.open_api("#sorting-collection-results")
# ## Getting Help
# print out all the docstrings, you can filter by a prefix if you want
print(coll.print_help())
# ----
# EOF
| CMR/python/demos/collections.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Do Package Manager stuff
# > Use the right tool for the job: there are some things that aren't worth re-engineering in Python. Here are some tools in python and javascript that will help your project be more maintainable
# !cd .. && pip install -r requirements-dev.txt
# !cd .. && npm install
# !cd .. && bower install
# # Check Style
# !flake8 ../ipythond3sankey/*.py ../*.py
# !cd .. && npm run jshint
# # Transpile JS/CSS
# !cd .. && npm run less
# ## Run Tests
# !cd .. && PYTHONWARNINGS=ignore python setup.py test
# # Generate READMEs
# !ipython nbconvert README.ipynb --to=rst --stdout > ../README.rst
# # PyPi: release
# !rm ../dist/ipython-d3-sankey-*.tar.gz
# !cd .. && python setup.py sdist
# !cd .. && twine upload dist/ipython-d3-sankey-*.tar.gz
| notebooks/Automation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (CS189)
# language: python
# name: myenv
# ---
# + deletable=true editable=true
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + deletable=true editable=true
save_path = '/Users/dianchen/state.npy'
data = np.load(save_path)
n, d = data.shape
# + deletable=true editable=true
for i in range(d):
plt.figure()
plt.hist(data[:, i])
plt.title('Dim %d' % i)
# + deletable=true editable=true
| notebooks/plot_state_dist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
suppressWarnings(suppressPackageStartupMessages(library(ggplot2)))
suppressWarnings(suppressPackageStartupMessages(library(RColorBrewer)))
suppressWarnings(suppressPackageStartupMessages(library(scran)))
suppressWarnings(suppressPackageStartupMessages(library(dplyr)))
suppressWarnings(suppressPackageStartupMessages(library(ggbeeswarm)))
# +
# load dataset
# -
sce = readRDS("/hps/nobackup/hipsci/scratch/singlecell_endodiff/data_processed/merged/20180618/sce_merged_afterqc_filt_allexpts.rds")
sce = normalize(sce)
design = model.matrix(~ experiment, data = colData(sce))
alt.fit = trendVar(sce, design = design, use.spikes = FALSE)
alt.decomp = decomposeVar(sce, alt.fit)
# get top 500
top.500.hvgs <- rownames(alt.decomp[order(alt.decomp$bio, decreasing = TRUE),][1:500,])
# get PC1 from expression of the top 500 hvgs
sce_500hvg = sce[rownames(sce) %in% top.500.hvgs,]
p1 = as.data.frame(prcomp(t(logcounts(sce_500hvg)))$x[,1])
colnames(p1) = "PC1_top500hvgs"
head(p1)
sce$pc1top500hvgs = p1$PC1_top500hvgs
# +
# define pseudotime as PC1 from top 500 HVGs, scaled between 0 and 1
# -
range01 <- function(x){(x-min(x))/(max(x)-min(x))}
sce$pseudotime <- range01(sce$pc1top500hvgs)
# +
# make time point (day of collection) specific scesets
# -
sce_0 = sce[,sce$day == "day0"]
sce_1 = sce[,sce$day == "day1"]
sce_2 = sce[,sce$day == "day2"]
sce_3 = sce[,sce$day == "day3"]
# +
#######################################
######### Figure 3A, left #############
#######################################
# +
# to quantify average pseudotime per donot, we first make data frames per day
# -
df_pseudo0 = data.frame(pseudo = sce_0$pseudotime, donor = sce_0$donor_short_id, batch = sce_0$experiment)
df_pseudo1 = data.frame(pseudo = sce_1$pseudotime, donor = sce_1$donor_short_id, batch = sce_1$experiment)
df_pseudo2 = data.frame(pseudo = sce_2$pseudotime, donor = sce_2$donor_short_id, batch = sce_2$experiment)
df_pseudo3 = data.frame(pseudo = sce_3$pseudotime, donor = sce_3$donor_short_id, batch = sce_3$experiment)
# count the number of cells and get average pseudotime value per (donor, day) group
pseudo0 <- df_pseudo0 %>% group_by(donor) %>% summarize(ncells = n(), avg_pseudo = mean(as.numeric(pseudo)))
pseudo1 <- df_pseudo1 %>% group_by(donor) %>% summarize(ncells = n(), avg_pseudo = mean(as.numeric(pseudo)))
pseudo2 <- df_pseudo2 %>% group_by(donor) %>% summarize(ncells = n(), avg_pseudo = mean(as.numeric(pseudo)))
pseudo3 <- df_pseudo3 %>% group_by(donor) %>% summarize(ncells = n(), avg_pseudo = mean(as.numeric(pseudo)))
# only trust average if done with at least 10 cells
N = 10
pseudo0 = pseudo0[pseudo0$ncells > N,]
pseudo1 = pseudo1[pseudo1$ncells > N,]
pseudo2 = pseudo2[pseudo2$ncells > N,]
pseudo3 = pseudo3[pseudo3$ncells > N,]
# combine
pseudo0_bis = inner_join(pseudo0, pseudo3, by = "donor", suffix = c(".day0",".day3"))
colnames(pseudo0_bis)[c(3,5)] <- c("avg_pseudo","diff_capacity")
pseudo1_bis = inner_join(pseudo1, pseudo3, by = "donor", suffix = c(".day1",".day3"))
colnames(pseudo1_bis)[c(3,5)] <- c("avg_pseudo","diff_capacity")
pseudo2_bis = inner_join(pseudo2, pseudo3, by = "donor", suffix = c(".day2",".day3"))
colnames(pseudo2_bis)[c(3,5)] <- c("avg_pseudo","diff_capacity")
pseudo3$diff_capacity = pseudo3$avg_pseudo
head(pseudo3)
pseudo0_bis$day = "day0"
pseudo1_bis$day = "day1"
pseudo2_bis$day = "day2"
pseudo3$day = "day3"
cols = c("donor","day","avg_pseudo","diff_capacity")
head(pseudo0_bis)
# make dataframe for plotting
df = rbind(pseudo0_bis[,cols], pseudo1_bis[,cols], pseudo2_bis[,cols], pseudo3[,cols])
nrow(df)
head(df[order(df$diff_capacity),])
length(unique(df$donor)) # donors with data (at least 10 cells) across all time points
options(repr.plot.width = 9, repr.plot.height = 6)
mid <- min(df$diff_capacity) + 0.45*(max(df$diff_capacity)-min(df$diff_capacity))
ggplot(df, aes(x = day, y = avg_pseudo, col = as.numeric(diff_capacity), group = donor)) +
# scale_colour_gradientn(colors = rev(brewer.pal(9,"Spectral"))) +
# scale_colour_gradientn(colors = brewer.pal("Spectral", n = 9)) +
scale_colour_gradient2(midpoint = mid, low = "firebrick", mid = "goldenrod1", high = "dodgerblue", space = "Lab") +
# scale_colour_gradient(low = "red", high = "blue") +
geom_point(alpha = 0.8) +
geom_line(alpha = 0.5) +
theme_minimal() #+
# theme(panel.background = element_rect(fill = 'grey86'),
# panel.grid.major = element_line(size = 0.5, linetype = 'solid', colour = "gray90"),
# panel.grid.minor = element_line(size = 0.5, linetype = 'solid', colour = "gray90"),
# panel.border = element_rect(colour = "gray88", fill = NA, size = 0.75))
# +
#######################################
######### Figure 3A, right ############
#######################################
# -
pseudo3_batch <- df_pseudo3 %>% group_by(donor,batch) %>% summarize(ncells = n(), avg_pseudo = mean(as.numeric(pseudo)))
options(repr.plot.width = 7, repr.plot.height = 6)
df3 = pseudo3_batch
df3$diff_capacity = df3$avg_pseudo
ggplot(df3, aes(x = reorder(donor, -diff_capacity), y = diff_capacity, colour = diff_capacity)) +
# scale_colour_gradientn(colors = brewer.pal("Spectral", n = 9)) +
scale_colour_gradient2(midpoint = mid, low = "firebrick", mid = "goldenrod1", high = "dodgerblue", space = "Lab") +
geom_point(alpha = 0.8) + geom_line(alpha = 0.7) +
xlab("donors ordered by differentiation efficiency") + ylab("differentiation efficiency") +
theme_bw() + theme(axis.text.x = element_blank(), axis.ticks.x = element_blank(),
# axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
panel.border = element_rect(colour = "gray88", fill = NA, size = 0.75),
panel.grid.major = element_line(size = 0.5, linetype = 'solid', colour = "gray95"),
panel.grid.minor = element_line(size = 0.25, linetype = 'solid',colour = "gray88"))
# +
########################################
######### Figure 3B, Manhattan #########
########################################
# -
minigwas_filename = "/nfs/leia/research/stegle/dseaton/hipsci/singlecell_endodiff/data/supplementary_tables/SuppTable_S3_differentiation_association_tests_genetic_markers.tsv"
minigwas = read.csv(minigwas_filename, sep = "\t")
head(minigwas)
df = minigwas
feature_highlight = "DPH3"
# +
# Prepare the dataset
don <- df %>%
# Compute chromosome size
group_by(snp_chromosome) %>%
summarise(chr_len = max(snp_position)) %>%
# Calculate cumulative position of each chromosome
dplyr::mutate(tot = cumsum(chr_len) - chr_len) %>%
select(-chr_len) %>%
# Add this info to the initial dataset
left_join(df, ., by = c("snp_chromosome" = "snp_chromosome")) %>%
# Add a cumulative position of each SNP
dplyr::arrange(snp_chromosome, snp_position) %>%
dplyr::mutate( BPcum = snp_position + tot) %>%
# Add highlight and annotation information
# dplyr::mutate( is_highlight1 = ifelse(hgnc_gene_symbol %in% feature_highlight, "yes", "no")) %>%
# Filter SNP to make the plot lighter
dplyr::filter(-log10(pval) > 0)
# Prepare X axis
axisdf <- don %>% group_by(snp_chromosome) %>% summarize(center =( max(BPcum) + min(BPcum) ) / 2 )
# Make the plot
p <- ggplot(don, aes(x = BPcum, y = -log10(pval), text = "")) +
# Show all points
geom_point( aes(color = as.factor(snp_chromosome)), alpha = 0.6, size = 2) +
scale_color_manual(values = rep(c("grey75", "dimgrey"), 22 )) +
# custom X axis:
scale_x_continuous( label = axisdf$snp_chromosome, breaks = axisdf$center ) +
scale_y_continuous(expand = c(0, 0) ) + # remove space between plot area and x axis
ylim(0,-log10(min(don$pval))+0.5)+
xlab("SNP location") +
ylab("-log10(P)") +
# Add highlighted points
# geom_point(data = subset(don, is_highlight1 == "yes"), col = "dodgerblue", size = 1.2, alpha = 0.3) +
# Add significance threshold
geom_hline(yintercept = -log10(2.3e-04), alpha = 0.5, col = "firebrick") +
# Custom the theme:
theme_bw() +
theme(
legend.position="none",
panel.border = element_blank(),
panel.grid.major.x = element_blank(),
panel.grid.minor.x = element_blank(),
panel.grid.major.y = element_blank(),
text = element_text(size = 20)
)
options(repr.plot.width = 10, repr.plot.height = 4)
p
# +
######################################
######### Figure 3B, Boxplot #########
######################################
# -
genos = read.table("/hps/nobackup/hipsci/scratch/genotypes/imputed/2017-03-27/selectionAnna_July3rd.dosages.txt", header = T, row.names = 1)
leads.mesendo = read.csv("/nfs/leia/research/stegle/acuomo/mean/mesendo_est_June20/leads.csv", row.names = 1)
res = leads.mesendo
head(res)
i = grep("DPH3",res$feature)
qtl = res[i,]
qtl
gene = qtl$feature
snp = qtl$snp_id
snp_name = "rs73138519"
ref = unlist(strsplit(snp,"_"))[3]
alt = unlist(strsplit(snp,"_"))[4]
sce = sce_3
df0 = data.frame(donor = sce$donor_short_id, expt = sce$experiment, day = sce$day, pseudotime = sce$pseudotime)
# sce = sce_mesendo
# df0 = data.frame(donor = sce$donor_short_id, expt = sce$experiment, day = sce$day, exprs = logcounts(sce)[gene,])
df0 <- dplyr::mutate(df0, donor.expt.day = paste0(donor, "-", expt, "-", day))
df1 <- df0 %>% group_by(donor.expt.day) %>% summarize(mean.pseudo = mean(as.numeric(pseudotime)))
# df1 <- df0 %>% group_by(donor.expt.day) %>% summarize(mean.exprs.sc = mean(as.numeric(exprs)))
df1$donor <- gsub("*-.*","",df1$donor.expt.day)
df1$expt <- gsub("*-.*","",gsub(".*-e","e",df1$donor.expt.day))
df1$day <- gsub(".*-","",df1$donor.expt.day)
# select snp
geno = genos[rownames(genos) == snp,]
df2 = as.data.frame(t(rbind(geno,colnames(geno))))
df2[,2] = gsub("\\.","-",df2[,2])
df2[,2] = gsub(".*-","",df2[,2])
colnames(df2) = c("dosages","donor")
df2 <- dplyr::mutate(df2, genotypes = round(as.numeric(dosages)))
head(df2)
# make dataframe
df = as.data.frame(left_join(df1, df2, by = "donor"))
head(df)
col1 = "grey45"
col2 = "grey45"
df$alleles = c()
df$alleles[df$genotypes == 0] <- paste0(ref,ref)
df$alleles[df$genotypes == 1] <- paste0(ref,alt)
df$alleles[df$genotypes == 2] <- paste0(alt,alt)
options(repr.plot.width = 3, repr.plot.height = 4)
ggplot(df[df$genotypes %in% c(0,1,2),], aes(x = as.factor(alleles), y = as.numeric(mean.pseudo))) +
geom_violin(col = col1)+
geom_quasirandom(aes(x = as.factor(alleles), y = as.numeric(mean.pseudo)), col = col2, alpha = 0.5) +
geom_boxplot(width = 0.1, col = col2) + xlab(snp_name) +
labs(y = "differentiation efficiency") + theme_classic()
# +
########################################
######### Figure 3C, Manhattan #########
########################################
# -
x_chrom_predict_filename = "/nfs/leia/research/stegle/dseaton/hipsci/singlecell_endodiff/data/differentiation_prediction/differentiation_marker_LMM_w_donor_results.transcriptome_wide_scan_pseudotime.tsv"
x_chrom_predict = read.csv(x_chrom_predict_filename, sep = "\t")
head(x_chrom_predict)
x_chrom_predict_day0 = x_chrom_predict[x_chrom_predict$timepoint1 == "day0",]
x_chrom_predict_day0$hgnc_symbol = x_chrom_predict_day0$gene1
head(x_chrom_predict_day0)
## add gene info
gene_info_filename = "/nfs/leia/research/stegle/dseaton/genomes/hg19/annotation/Homo_sapiens.GRCh37.75.genes.bed"
gene_info = read.table(gene_info_filename)
colnames(gene_info) = c("chromosome_name","start_position","end_position","ensembl_gene_id","gene_biotype","strand")
head(gene_info)
gene_annos_filename = "/hps/nobackup/hipsci/scratch/singlecell_endodiff/data_processed/scQTLs/annos/combined_feature_id_annos.tsv"
gene_annos = read.csv(gene_annos_filename, sep = "\t")
head(gene_annos)
df.gene = inner_join(gene_info, gene_annos, by = "ensembl_gene_id")
head(df.gene)
nrow(x_chrom_predict_day0)
df = left_join(x_chrom_predict_day0, df.gene, by = "hgnc_symbol")
nrow(df)
df = df[!(is.na(df$start_position)),]
nrow(df)
add_TSS <- function(res){
res[res$strand == "+","TSS"] = res[res$strand == "+","start_position"]
res[res$strand == "-","TSS"] = res[res$strand == "-","end_position"]
res
}
df = add_TSS(df)
df$corr_pval = p.adjust(df$pval, method = "BH")
## features to highlight in the Manhattan plot
# significant, NOT on X chromosome
featuresOfInterestNotX = df[df$corr_pval < 0.1 & df$chromosome_name != "X", "feature_id"]
# significant, ON X chromosome
featuresOfInterestX = df[df$corr_pval < 0.1 & df$chromosome_name == "X", "feature_id"]
example_X = rownames(sce)[grep("ZDHHC9",rownames(sce))]
example_X
## P value corresponding to largest significant corrected P
df_sign = df[df$corr_pval < 0.1,]
mythresh = round(-log10(tail(df_sign[order(df_sign$corr_pval),],1)[,"pval"]),digits = 2)
mythresh
df = df[df$chromosome_name %in% c(1:22,"X"),]
# re-order chromosome names
df$CHR = as.factor(df$chromosome)
levels(df$CHR)
df$CHR =factor(df$CHR, levels(df$CHR)[c(1,12,16:22,2:11,13:15,23)])
levels(df$CHR)
# +
# Prepare the dataset
don <- df %>%
# Compute chromosome size
group_by(CHR) %>%
summarise(chr_len = max(TSS)) %>%
# Calculate cumulative position of each chromosome
dplyr::mutate(tot = cumsum(chr_len) - chr_len) %>%
select(-chr_len) %>%
# Add this info to the initial dataset
left_join(df, ., by = c("CHR" = "CHR")) %>%
# Add a cumulative position of each SNP
dplyr::arrange(CHR, TSS) %>%
dplyr::mutate( BPcum = TSS + tot) %>%
# Add highlight and annotation information
dplyr::mutate( is_highlight1 = ifelse(feature_id %in% featuresOfInterestNotX, "yes", "no")) %>%
dplyr::mutate( is_highlight2 = ifelse(feature_id %in% featuresOfInterestX, "yes", "no")) %>%
# specific examples
dplyr::mutate( is_highlight3 = ifelse(feature_id == example_X, "yes", "no")) %>%
# Filter SNP to make the plot lighter
dplyr::filter(-log10(pval) > 0)
# Prepare X axis
axisdf <- don %>% group_by(CHR) %>% summarize(center =( max(BPcum) + min(BPcum) ) / 2 )
# Make the plot
p <- ggplot(don, aes(x = BPcum, y = -log10(pval)*sign(coefficient), text = "")) +
# Show all points
geom_point( aes(color = as.factor(CHR)), alpha = 0.3, size = 0.8) +
scale_color_manual(values = rep(c("grey75", "dimgrey"), 22 )) +
# custom X axis:
scale_x_continuous( label = axisdf$CHR, breaks = axisdf$center ) +
scale_y_continuous(expand = c(0, 0) ) + # remove space between plot area and x axis
ylim(-9,9)+
xlab("gene location (TSS)") +
ylab("-log10(P) X \neffect direction") +
# Add highlighted points
geom_point(data = subset(don, is_highlight1 == "yes"), col = "dodgerblue", size = 1.2, alpha = 0.3) +
geom_point(data = subset(don, is_highlight2 == "yes"), col = "firebrick", size = 1.2, alpha = 0.3) +
geom_point(data = subset(don, is_highlight3 == "yes"), col = "darkorange3", size = 1.9) +
# geom_point(data = subset(don, is_highlight4 == "yes"), col = "#344d90", size = 1.8) +
# geom_point(data = subset(don, is_highlight3 == "yes"), col = "green", size = 3) +
# Add significance threshold
geom_hline(yintercept = mythresh, alpha = 0.3, col = "dodgerblue") +
geom_hline(yintercept = -mythresh, alpha = 0.3, col = "dodgerblue") +
# Custom the theme:
theme_bw() +
theme(
legend.position="none",
panel.border = element_blank(),
panel.grid.major.x = element_blank(),
panel.grid.minor.x = element_blank(),
panel.grid.major.y = element_blank(),
text = element_text(size = 16)
)
options(repr.plot.width = 7, repr.plot.height = 5)
p
# +
##########################################
######### Figure 3C, Scatterplot #########
##########################################
# -
ZDHHC9_filename = "/nfs/leia/research/stegle/dseaton/hipsci/singlecell_endodiff/data/differentiation_prediction/differentiation_marker_LMM_w_donor_results_subset_of_day0_markers_data/day0_ZDHHC9_day3_pseudotime.tsv"
ZDHHC9 = read.csv(ZDHHC9_filename, sep = "\t")
head(ZDHHC9)
options(repr.plot.width = 4, repr.plot.height = 4)
col = "gray27"
p = ggplot(ZDHHC9, aes(x = phenotype, y = fixed_effect)) + geom_point(col = col)
p = p + stat_smooth(colour = col, linetype = 2, method = "lm", alpha = 0.05)
p = p + theme_minimal()
p = p + xlab(paste0("\niPSC expression \nof ZDHHC9")) + ylab("differentiation efficacy\n")
p + theme(plot.title = element_text(hjust = 0.5, colour = col),
text = element_text(size = 16))
| plotting_notebooks/differentiation_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kfp
# language: python
# name: kfp
# ---
# # Part 1 - Experimentation
#
#
# +
import os
import numpy as np
import pandas as pd
import uuid
import time
import tempfile
from googleapiclient import discovery
from googleapiclient import errors
from google.cloud import bigquery
from jinja2 import Template
from kfp.components import func_to_container_op
from typing import NamedTuple
from sklearn.metrics import accuracy_score
from sklearn.linear_model import SGDClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.compose import ColumnTransformer
# -
# ## Configure environment settings
# Make sure to update the constants to reflect your environment settings.
PROJECT_ID = 'mlops-workshop'
DATASET_LOCATION = 'US'
CLUSTER_NAME = 'mlops-workshop-cluster'
CLUSTER_ZONE = 'us-central1-a'
REGION = 'us-central1'
DATASET_ID = 'lab_11'
SOURCE_TABLE_ID = 'covertype'
TRAINING_TABLE_ID = 'training_split'
VALIDATION_TABLE_ID = 'validation_split'
TESTING_TABLE_ID = 'testing_split'
LAB_GCS_BUCKET='gs://mlops-workshop-lab-11'
TRAINING_FILE_PATH = LAB_GCS_BUCKET + '/datasets/training/data.csv'
VALIDATION_FILE_PATH = LAB_GCS_BUCKET + '/datasets/validation/data.csv'
TESTING_FILE_PATH = LAB_GCS_BUCKET + '/datasets/testing/data.csv'
# ## Explore the source dataset
# Bring a few rows from the source dataset.
# +
client = bigquery.Client(project=PROJECT_ID, location=DATASET_LOCATION)
query_template = """
SELECT *
FROM `{{ source_table }}`
LIMIT 10
"""
query = Template(query_template).render(
source_table='{}.{}.{}'.format(PROJECT_ID, DATASET_ID, SOURCE_TABLE_ID))
df = client.query(query).to_dataframe()
num_of_columns = len(df.columns)
df
# -
# Count the number of rows and columns in the source.
# +
query_template = """
SELECT count(*)
FROM `{{ source_table }}`
"""
query = Template(query_template).render(
source_table='{}.{}.{}'.format(PROJECT_ID, DATASET_ID, SOURCE_TABLE_ID))
df = client.query(query).to_dataframe()
number_of_rows_in_full_dataset = df.iloc[0,0]
print('{} x {}'.format(number_of_rows_in_full_dataset, num_of_columns))
# -
# ## Create the training, validation and testing splits
# Define the sampling query template.
sampling_query_template = """
SELECT *
FROM
`{{ source_table }}` AS cover
WHERE
MOD(ABS(FARM_FINGERPRINT(TO_JSON_STRING(cover))), {{ num_lots }}) in {{ lots_to_select }}
"""
# Configure the sampling query job settings.
job_config = bigquery.QueryJobConfig()
job_config.create_disposition = bigquery.job.CreateDisposition.CREATE_IF_NEEDED
job_config.write_disposition = bigquery.job.WriteDisposition.WRITE_TRUNCATE
dataset_ref = client.dataset(DATASET_ID)
# Create the training split table.
# +
query = Template(sampling_query_template).render(
source_table='{}.{}.{}'.format(PROJECT_ID, DATASET_ID, SOURCE_TABLE_ID),
num_lots=10,
lots_to_select='(1, 2, 3)')
job_config.destination = dataset_ref.table(TRAINING_TABLE_ID)
client.query(query, job_config).result()
# -
# Extract the training split table to GCS.
client.extract_table(dataset_ref.table(TRAINING_TABLE_ID), TRAINING_FILE_PATH).result()
# Inspect the extracted file.
# !gsutil cat -r 0-500 {TRAINING_FILE_PATH}
# Create the validation split table.
# +
query = Template(sampling_query_template).render(
source_table='{}.{}.{}'.format(PROJECT_ID, DATASET_ID, SOURCE_TABLE_ID),
num_lots=10,
lots_to_select='(8)')
job_config.destination = dataset_ref.table(VALIDATION_TABLE_ID)
client.query(query, job_config).result()
# -
# Extract the validation split table to GCS.
client.extract_table(dataset_ref.table(VALIDATION_TABLE_ID), VALIDATION_FILE_PATH).result()
# Create the testing split table.
# +
query = Template(sampling_query_template).render(
source_table='{}.{}.{}'.format(PROJECT_ID, DATASET_ID, SOURCE_TABLE_ID),
num_lots=10,
lots_to_select='(9)')
job_config.destination = dataset_ref.table(TESTING_TABLE_ID)
client.query(query, job_config).result()
# -
# Extract the testing split table to GCS.
client.extract_table(dataset_ref.table(TESTING_TABLE_ID), TESTING_FILE_PATH).result()
# ## Develop the training script
# Configure the `sklearn` training pipeline.
# +
numeric_features = ['Elevation', 'Aspect', 'Slope', 'Horizontal_Distance_To_Hydrology',
'Vertical_Distance_To_Hydrology', 'Horizontal_Distance_To_Roadways',
'Hillshade_9am', 'Hillshade_Noon', 'Hillshade_3pm',
'Horizontal_Distance_To_Fire_Points']
categorical_features = ['Wilderness_Area', 'Soil_Type']
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_features),
('cat', OneHotEncoder(), categorical_features)
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', SGDClassifier(loss='log'))
])
# -
# Run the pipeline locally.
# +
df_train = pd.read_csv(TRAINING_FILE_PATH)
df_validation = pd.read_csv(VALIDATION_FILE_PATH)
X_train = df_train.drop('Cover_Type', axis=1)
y_train = df_train['Cover_Type']
X_validation = df_validation.drop('Cover_Type', axis=1)
y_validation = df_validation['Cover_Type']
pipeline.set_params(classifier__alpha=0.001, classifier__max_iter=200)
pipeline.fit(X_train, y_train)
pipeline.score(X_validation, y_validation)
# -
# #### Prepare the hyperparameter tuning application.
# Since the training run on this dataset is computationally expensive you can benefit from running a distributed hyperparameter tuning job on AI Platform Training.
TRAINING_APP_FOLDER = 'training_app'
os.makedirs(TRAINING_APP_FOLDER, exist_ok=True)
# Write the tuning script.
# +
# %%writefile {TRAINING_APP_FOLDER}/train.py
import os
import subprocess
import sys
import fire
import pickle
import numpy as np
import pandas as pd
import hypertune
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import SGDClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
def train_evaluate(job_dir, training_dataset_path, validation_dataset_path, alpha, max_iter, hptune):
df_train = pd.read_csv(training_dataset_path)
df_validation = pd.read_csv(validation_dataset_path)
if not hptune:
df_train = pd.concat([df_train, df_validation])
numeric_features = ['Elevation', 'Aspect', 'Slope', 'Horizontal_Distance_To_Hydrology',
'Vertical_Distance_To_Hydrology', 'Horizontal_Distance_To_Roadways',
'Hillshade_9am', 'Hillshade_Noon', 'Hillshade_3pm',
'Horizontal_Distance_To_Fire_Points']
categorical_features = ['Wilderness_Area', 'Soil_Type']
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_features),
('cat', OneHotEncoder(), categorical_features)
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', SGDClassifier(loss='log'))
])
print('Starting training: alpha={}, max_iter={}'.format(alpha, max_iter))
X_train = df_train.drop('Cover_Type', axis=1)
y_train = df_train['Cover_Type']
pipeline.set_params(classifier__alpha=alpha, classifier__max_iter=max_iter)
pipeline.fit(X_train, y_train)
if hptune:
X_validation = df_validation.drop('Cover_Type', axis=1)
y_validation = df_validation['Cover_Type']
accuracy = pipeline.score(X_validation, y_validation)
print('Model accuracy: {}'.format(accuracy))
# Log it with hypertune
hpt = hypertune.HyperTune()
hpt.report_hyperparameter_tuning_metric(
hyperparameter_metric_tag='accuracy',
metric_value=accuracy
)
# Save the model
if not hptune:
model_filename = 'model.pkl'
with open(model_filename, 'wb') as model_file:
pickle.dump(pipeline, model_file)
gcs_model_path = "{}/{}".format(job_dir, model_filename)
subprocess.check_call(['gsutil', 'cp', model_filename, gcs_model_path], stderr=sys.stdout)
print("Saved model in: {}".format(gcs_model_path))
if __name__ == "__main__":
fire.Fire(train_evaluate)
# -
# Package the script into a docker image.
# +
# %%writefile {TRAINING_APP_FOLDER}/Dockerfile
FROM gcr.io/deeplearning-platform-release/base-cpu
RUN pip install -U fire cloudml-hypertune
WORKDIR /app
COPY train.py .
ENTRYPOINT ["python", "train.py"]
# -
# Build the docker image.
# +
IMAGE_NAME='trainer_image'
IMAGE_TAG='latest'
IMAGE_URI='gcr.io/{}/{}:{}'.format(PROJECT_ID, IMAGE_NAME, IMAGE_TAG)
# !gcloud builds submit --tag $IMAGE_URI $TRAINING_APP_FOLDER
# -
# Create the hyperparameter configuration file.
# +
# %%writefile {TRAINING_APP_FOLDER}/hptuning_config.yaml
trainingInput:
hyperparameters:
goal: MAXIMIZE
maxTrials: 6
maxParallelTrials: 3
hyperparameterMetricTag: accuracy
enableTrialEarlyStopping: TRUE
params:
- parameterName: max_iter
type: DISCRETE
discreteValues: [
200,
500
]
- parameterName: alpha
type: DOUBLE
minValue: 0.00001
maxValue: 0.001
scaleType: UNIT_LINEAR_SCALE
# -
# #### Submit the hyperparameter tuning job.
# +
JOB_NAME = "JOB_{}".format(time.strftime("%Y%m%d_%H%M%S"))
JOB_DIR = "{}/{}".format(LAB_GCS_BUCKET, JOB_NAME)
SCALE_TIER = "BASIC"
# !gcloud ai-platform jobs submit training $JOB_NAME \
# --region=$REGION \
# --job-dir=$LAB_GCS_BUCKET/$JOB_NAME \
# --master-image-uri=$IMAGE_URI \
# --scale-tier=$SCALE_TIER \
# --config $TRAINING_APP_FOLDER/hptuning_config.yaml \
# -- \
# --training_dataset_path=$TRAINING_FILE_PATH \
# --validation_dataset_path=$VALIDATION_FILE_PATH \
# --hptune
# -
# #### Monitor the job.
# !gcloud ai-platform jobs describe $JOB_NAME
# !gcloud ai-platform jobs stream-logs $JOB_NAME
# ### Retrieve HP-tuning results.
# Call AI Platform Training end-point.
# +
ml = discovery.build('ml', 'v1')
job_id = 'projects/{}/jobs/{}'.format(PROJECT_ID, JOB_NAME)
request = ml.projects().jobs().get(name=job_id)
try:
response = request.execute()
except errors.HttpError as err:
print(err)
except:
print("Unexpected error")
response
# -
# Retrieve the best run.
response['trainingOutput']['trials'][0]
| Lab-11-KFP-CAIP/notebooks/covertype_experimentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cseveriano/evolving_clustering/blob/master/notebooks/Evolving_Clustering_Static_DS3_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="lKynvns2FzaT" colab_type="code" outputId="7a8779ef-5bdb-4cef-d5b9-7bc2c04c986d" colab={"base_uri": "https://localhost:8080/", "height": 122}
path = "/content/gdrive/My Drive/Evolving_Results/"
from google.colab import drive
drive.mount("/content/gdrive")
# + colab_type="code" cellView="both" id="4fB7iiO4qvqY" outputId="d1041b97-79f4-45b7-a1ca-d81017cd33b8" colab={"base_uri": "https://localhost:8080/", "height": 18805}
#@title
# !apt-get update
# !apt-get install r-base
# !pip install rpy2
# !apt-get install libmagick++-dev
# #!apt-get install r-cran-rjava
import os #importing os to set environment variable
def install_java():
# !apt-get install -y openjdk-8-jdk-headless -qq > /dev/null #install openjdk
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64" #set environment variable
os.environ["LD_LIBRARY_PATH"] = "/usr/lib/jvm/java-8-openjdk-amd64/jre/lib/amd64:/usr/lib/jvm/java-8-openjdk-amd64/jre/lib/amd64/server"
# !java -version #check java version
install_java()
# !R CMD javareconf
# #!apt-get install r-cran-rjava
# #!apt-get install libgdal-dev libproj-dev
# !R -e 'install.packages(c("magick", "animation", "stream", "rJava", "streamMOA"))'
# + [markdown] colab_type="text" id="USb2KES7qvqf"
# ##Install R Packages
# + colab_type="code" id="I0ekyA9bqvqh" colab={}
# enables the %%R magic, not necessary if you've already done this
# %load_ext rpy2.ipython
# + colab_type="code" id="0OEaC7mNqvqk" outputId="bec8cd0c-1240-4f32-ad3a-178d18388aa3" colab={"base_uri": "https://localhost:8080/", "height": 479} language="R"
# dyn.load("/usr/lib/jvm/java-8-openjdk-amd64/jre/lib/amd64/server/libjvm.so")
# library("stream")
# library("streamMOA")
# + [markdown] colab_type="text" id="9IcAjfliqvqo"
# ##Generate Concept Drift Data Stream
#
#
# * Choose concept drift benchmark (read Bolanos paper)
# * send outputs:
# * stream_df
# * prequential window size
#
#
#
#
# + colab_type="code" id="x-8ehC-vqvqp" colab={} language="R"
# experiment <- function(){
# df <- read.csv("https://query.data.world/s/faarcpz24ukr2g53druxljp4nghhzn", header=TRUE);
# nsamples <- nrow(df)
# df <- df[sample(nsamples),]
# stream <- DSD_Memory(df[,c("x", "y")], class=df[,"class"], k=max(df[,"class"]))
# return (get_points(stream, n = nsamples, class = TRUE))
# }
# + [markdown] colab_type="text" id="t1nqh3j-qvqx"
# ##Run Benchmark Models
#
# ### Benchmark methods:
# * DBStream
# * DenStream
# * ClusStream
# * Stream KM++
#
# ### Benchmark metrics:
# * Precision
#
#
# + colab_type="code" id="KhJqTP_Sqvqy" colab={}
# Experiment parameters
nclusters = 9
nsamples = 3000
train_size = 1800
window_size = 100
metric = "cRand"
trials = 30
# + colab_type="code" id="YFXp5kaZqvq8" outputId="d791d666-341a-4677-c0ed-3b4f2f6dee8b" colab={"base_uri": "https://localhost:8080/", "height": 527} magic_args="-i metric -i window_size -i trials -i path -i nclusters" language="R"
#
# alg_names <- c("DenStream", "Clustream", "StreamKM")
# trials_df <- data.frame(matrix(ncol = length(alg_names), nrow = 0))
# colnames(trials_df) <- alg_names
#
# for (i in 1:(trials)){
# algorithms <- list("DenStream" = DSC_DenStream(epsilon=0.01, mu=4, beta=0.4),
# "Clustream" = DSC_CluStream(m = 10, horizon = 10, t = 1, k=NULL),
# "StreamKM" = DSC_StreamKM(sizeCoreset = 100, numClusters = nclusters)
# )
# writeLines(sprintf("Trial: %d", i))
#
# evaluation <- sapply(algorithms, FUN = function(alg) {
#
# df <- read.csv("https://query.data.world/s/faarcpz24ukr2g53druxljp4nghhzn", header=TRUE);
# nsamples <- nrow(df)
# df <- df[sample(nsamples),]
# stream <- DSD_Memory(df[,c("x", "y")], class=df[,"class"], k=max(df[,"class"]))
# update(alg, stream, n=nsamples)
# reset_stream(stream)
# evaluate(alg, stream, measure = metric, n = nsamples, type = "macro", assign = "macro")
# })
#
# trials_df[nrow(trials_df) + 1,] = as.data.frame(evaluation)[,'evaluation']
# }
#
# write.csv(trials_df, paste(path,"results_DS3_benchmark.csv"))
# + [markdown] colab_type="text" id="IrW6hxL2qvrG"
# ##Run Evolving Clustering
#
# * Convert to X,y format
# * run prequential routine
# * plot results
# + colab_type="code" id="GQN8GerDqvrG" outputId="8b6411db-cb37-4de8-d1be-75330d742b71" colab={"base_uri": "https://localhost:8080/", "height": 170}
# !pip install -U git+https://github.com/cseveriano/evolving_clustering
# + colab_type="code" id="zkPU7anLqvrJ" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from evolving import EvolvingClustering, load_dataset, Metrics, Benchmarks, util
from sklearn.metrics import adjusted_rand_score
import time
import rpy2.robjects as robjects
from rpy2.robjects import pandas2ri
pandas2ri.activate()
r = robjects.r
# + colab_type="code" id="Z3SHLq_FqvrV" outputId="9bde4658-d1bb-4b52-8050-69c2566ab438" colab={"base_uri": "https://localhost:8080/", "height": 615}
evol_trials_df = pd.DataFrame(columns=["microTEDAclus"])
for i in np.arange(trials):
named_tuple = time.localtime() # get struct_time
time_string = time.strftime("%m/%d/%Y, %H:%M:%S", named_tuple)
print("Trial: ",i ," at ",time_string)
stream_df = pandas2ri.ri2py_dataframe(r.experiment())
X = stream_df[['x', 'y']].values
y = stream_df['class'].values
evol_model = EvolvingClustering.EvolvingClustering(variance_limit=0.0006, debug=False)
evol_model.fit(X)
y_hat = evol_model.predict(X)
error = adjusted_rand_score(y, y_hat)
evol_trials_df = evol_trials_df.append({'microTEDAclus': error}, ignore_index=True)
evol_trials_df.to_csv(path+'results_DS3_evolving.csv', index=False)
| notebooks/Evolving_Clustering_Static_DS3_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
import seaborn as sns
import string
import math
# +
datapath = "/Users/aavattikutis/Documents/epidemicmodel/cccruns/fits/fit2/tables/"
models = ["fulllinearmodel_fit_table.csv","reducedlinearmodelNegBinom_fit_table.csv",
"reducedlinearmodelq0_fit_table.csv","reducedlinearmodelq0ctime_fit_table.csv",
"nonlinearmodelq0ctime_fit_table.csv"]
# model_hash = {}
# k = -1
# for model in models:
# k += 1
# model_hash[model] = string.ascii_uppercase[k]
# df = pd.DataFrame.from_dict(model_hash, orient='index')
# df.to_csv('../postmodel_derivatives/model_hash.csv', header=False)
# +
rois = []
for model in models:
df = pd.read_csv(datapath + model) #get rois in all tables (some may have failed)
rois += list(df.roi.unique())
rois = list(set(rois))
#get inferred
theta = df.columns[2:]
ntheta = len(theta)
# +
#get rois
roi_us = np.sort([i for i in rois if i[:2]=='US'])[::-1]
roi_other = np.sort([i for i in rois if i[:2]!='US'])[::-1]
rois = list(roi_us) + list(roi_other)
# +
font = {'family' : 'sans-serif',
'weight' : 'normal',
'size' : 22}
matplotlib.rc('font', **font)
def simpleaxis(ax):
ax.spines['top'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
return
fs=24
theta_ = ["R0","car (week 0)","ifr (week 0)"]
label_ = [r'R$_{0}$','CAR (week=0)','IFR (week=0)']
xticks_ = [[0,5,10],[0,0.1,0.2],[0,0.005,0.01]]
# xlim_ = [[0,5,10],[0,1,2],[0,0.5,1],[0,2,5]]
def histrois(ax,theta,label,histcolor,xticks):
x = []
dfbest = pd.read_csv("../postfit_derivatives/"+theta+"_summary.csv")
for roi in rois:
model = dfbest.loc[(dfbest.Region==roi), "Model"].values[0] + "_fit_table.csv"
df = pd.read_csv(datapath + model)
try:
x2 = df.loc[(df.roi==roi)&(df['quantile']==0.5), theta].values[0]
if np.isfinite(x2):
if theta in ["car (week 0)","ifr (week 0)"]:
x.append(x2)
else:
x.append(x2)
except:
print()
mu = np.median(x)
print(mu)
f = sns.distplot(x,hist=True,kde=False,ax=ax,color=histcolor)
simpleaxis(ax)
ax.set_title(label,fontsize=fs)
ax.axvline(mu)
ax.text(mu,20,str(np.round(mu,3)))
ax.get_yaxis().set_visible(False)
ax.set_xticks(xticks)
ax.set_xlim((min(xticks),max(xticks)))
return
f,ax = plt.subplots(1,3,figsize=(20,5))
c_ = sns.color_palette("cubehelix")
for i in range(len(theta_)):
histrois(ax.flatten()[i],theta_[i],label_[i],c_[i],xticks_[i])
plt.subplots_adjust(wspace=0.5)
plt.savefig('../postfit_derivatives/fig_regionaverages.png')
# -
| notebooks/model_reports/zeroplot.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 3.0.2
# language: ruby
# name: ruby
# ---
require 'bio'
require 'rest-client' # this is how you access the Web
def fetch(url, headers = {accept: "*/*"}, user = "", pass="")
response = RestClient::Request.execute({
method: :get,
url: url.to_s,
user: user,
password: <PASSWORD>,
headers: headers})
return response
rescue RestClient::ExceptionWithResponse => e
$stderr.puts e.inspect
response = false
return response # now we are returning 'False', and we will check that with an \"if\" statement in our main code
rescue RestClient::Exception => e
$stderr.puts e.inspect
response = false
return response # now we are returning 'False', and we will check that with an \"if\" statement in our main code
rescue Exception => e
$stderr.puts e.inspect
response = false
return response # now we are returning 'False', and we will check that with an \"if\" statement in our main code
end
address = 'http://www.ebi.ac.uk/Tools/dbfetch/dbfetch?db=ensemblgenomesgene&format=embl&id=At3g54340'
response = fetch(address)
record = response.body
# create a local file with this data
File.open('./embl_records/At3g54340.embl', 'w') do |myfile| # w makes it writable
myfile.puts record
end
end
# +
datafile2 = Bio::FlatFile.auto('./embl_records/At3g54340.embl')
puts datafile2.class
datafile2.each_entry do |entry| # the FILE is not the same as the RECORD - multiple records can exist in a file
# shows accession and organism
next unless entry.accession # Specific DB ID.
puts entry.class # Bio::EMBL
puts "# #{entry.accession} - #{entry.species}"
# iterates over each element in 'features' # features method finds all features
entry.features.each do |feature|
next unless feature.feature == "exon"
position = feature.position # Look at Bio::Feature object
puts "\n\n\n\nPOSITION = #{position}"
puts position.class
puts "\n\n\n\nFeature = #{feature.feature}"
qual = feature.assoc # feature.assoc gives you a hash of Bio::Feature::Qualifier objects
# i.e. qualifier['key'] = value for example qualifier['gene'] = "CYP450")
puts "Associations = #{qual}"
# skips the entry if "/translation=" is not found
# next unless qual['translation'] # this is an indication that the feature is a transcript
# collects gene name and so on and joins it into a string
gene_info = [
qual['gene'], qual['product'], qual['note'], qual['function']
].compact.join(', ')
puts "TRANSCRIPT FOUND!\nGene Info: #{gene_info}"
# shows nucleic acid sequence
puts "\n\n>Exon('#{position}') : #{gene_info}" # Transcripts are the only things that have translation.
# Each transcript position tells the splice structure relative to entire seq
puts entry.naseq.class # this is a Bio::Sequence::NA Look at the documentation to understand the .splicing() method
puts entry.naseq.splice(position) # http://bioruby.org/rdoc/Bio/Sequence/Common.html#method-i-splice
end
puts "\n\nNumber of features #{entry.features.length}" # How many features are there
end
# -
require './Gene.rb'
gene = Gene.new(id: "AT2G46340")
puts
gene.embl.accession
acc_regex = Regexp.new(/chromosome:TAIR10:(\d):(\d*):(\d*):1/)
match = acc_regex.match(gene.embl.accession)
puts match[1]
puts 19022154 + 1990 - 1
puts 19022154 + 1995 - 1
from, to = gene.features['exons']['AT2G46340.1.exon1'].locations.span
puts from
puts to
seq = gene.sequence.splicing(gene.features['exons']['AT2G46340.1.exon1'].position)
gene.sequence.splicing("#{101 + 5009 - 1}..#{106 + 5009 - 1}")
gene.sequence.splicing("complement(F10M23_4:122538..122685)")
seq.splicing("5121 5126")
gene.features['exons'].each do |id, value|
to, from = value.locations.span
puts to
end
puts
repeats_array = Array.new
gene.sequence.features.each do |feature|
next unless feature.feature == 'repeat'
puts feature.assoc['note'].class
repeats << feature.assoc['note']
# next if gene.features['exons'].key?(idx)
# puts gene.features['exons'][idx]
end
puts
puts repeats.uniq.join(' ')
puts
puts gene.features["exons"].keys.join(' ')
gene.features["exons"].keys - repeats.uniq
gene.features['exons']["AT2G46340.2.exon7"]
gene.features["exons"].keys
# Search exons for CTTCTT
repeat = "CTTCTT"
repeat_regex = Regexp.new(/cttctt/i)
gene.features["exons"].each do |exon_id, exon|
# sequence = gene.embl.naseq.splicing(exon.position)
# For testing
sequence = "atgcttcttaaacttctt"
matches = repeat_regex.match(sequence)
if matches
puts matches.to_a
abort
matches.each do |match|
puts match.offset[1]
puts sequence[matches.offset[0]...matches.offset[1]]
end
end
end
puts
repeat = Bio::Sequence::NA.new("CTTCTT")
puts repeat.complement
# Search exons for CTTCTT
repeat = Bio::Sequence::NA.new("CTTCTT")
reverse_comp = "aagaag"
repeat_regex = Regexp.new(/(?=#{repeat})/i)
reverse_repeat_regex = Regexp.new(/(?=#{repeat.complement})/i)
gene.features["exons"].each do |exon_id, exon|
#sequence = gene.sequence.splice(exon.position)
# For testing
sequence = "cttaacttcttctt"
# sequence = "aagaagaagtttaag"
match_datas = sequence.to_enum(:scan, repeat_regex).map {Regexp.last_match}
if match_datas
match_datas.each do |match|
# puts match.offset(0)[0]
# puts match.offset(0)[1]
position = "#{match.offset(0)[0]+1}..#{match.offset(0)[0]+1 + 5}"
puts position
# puts sequence.splicing(position)
# repeat_ft = Bio::Feature::new('repeat', position)
# repeat_ft.append(Bio::Feature::Qualifier.new('note', exon_id))
# gene.sequence.features << repeat_ft
# puts exon.position
# puts match.offset(0)
# puts sequence[match.offset(0)[0]...match.offset(0)[1]]
# puts sequence[match.offset(0)[0]...match.offset(0)[1]]
# puts
end
end
rev_match_datas = sequence.to_enum(:scan, reverse_repeat_regex).map {Regexp.last_match}
if rev_match_datas
rev_match_datas.each do |match|
position = "complement(#{match.offset(0)[0]+1}..#{match.offset(0)[0]+1 + 5})"
puts position
# # puts sequence.splicing(position)
# rev_repeat_ft = Bio::Feature::new('repeat', position)
# rev_repeat_ft.append(Bio::Feature::Qualifier.new('note', exon_id))
# gene.sequence.features << rev_repeat_ft
# puts exon.position
# puts match.offset(0)
# puts sequence[match.offset(0)[0]...match.offset(0)[1]]
# puts sequence[match.offset(0)[0]...match.offset(0)[1]]
# puts
end
end
end
puts
def matching_substrings(string, regex)
string.size.times.each_with_object([]) do |start_index, maching_substrings|
start_index.upto(string.size.pred) do |end_index|
substring = string[start_index..end_index]
maching_substrings.push(substring) if substring =~ /^#{regex}$/
end
end
end
tests = 'cttcttcttctt'
repeat_regex = Regexp.new(/ctt/i)
match_datas = tests.to_enum(:scan, repeat_regex).map {Regexp.last_match}
match_datas.each_cons(2).to_a
gene.sequence.features.each do |feature|
puts feature
end
puts
puts sequence.length
puts sequence.splicing('complement(258..263)').complement
repeat = "CTTCTT"
sequence = "atgcttcttaaacttctt"
repeat_regex = Regexp.new(/#{repeat]/i)
s = Bio::Sequence::NA.new("atgcttcttaaacttctt")
puts s
puts s.splicing('4..9')
puts s.splicing('complement(4..9)')
# +
# iterates over each element in 'features' # features method finds all features
features_hash = Hash.new
entry.features.each do |feature|
# FILTER EXONS
next unless feature.feature == "exon"
puts feature.class
puts "FEATURE = #{feature.feature}"
position = feature.position # Look at Bio::Feature object
puts "POSITION = #{position}"
qual = feature.assoc #
puts "Associations = #{qual}"
puts "Associations note = #{qual["note"]}"
exon_id_regex = Regexp.new(/exon_id=(.*)/)
if exon_id_regex.match(qual["note"])
features_hash[$1] = feature
end
puts
end
puts 'end'
# -
puts features_hash.keys
features_hash["AT2G46340.2.exon8"].class
require './Gene.rb'
gene = Gene.new(id: "AT5G15850")
puts
puts gene.write_report
| Assignment3/assignment3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
import csv
import json
import matplotlib.pyplot as plt
# +
columns = ['test_accuracy_mean', 'test_precision_mean', 'test_recall_mean', 'test_f1-score_mean']
#columns = ['test_f1-score_mean']
metric_count = len(columns)
data = {}
category = 'Reinforcement Learning'
#category = 'Audio'
with open('../../results/final_results/final_results_main_majority_correct.csv') as f:
reader = csv.DictReader(f, delimiter=';')
for row in reader:
if row['PipelineID'] not in data:
data[row['PipelineID']] = {}
data[row['PipelineID']][row['Pipeline']] = [row[key] for key in columns]
#print(json.dumps(data, indent=4))
data1 = data[category]
data = {}
with open('../../results/final_results/final_results_main_negative_samples_majority_correct.csv') as f:
reader = csv.DictReader(f, delimiter=';')
for row in reader:
if row['PipelineID'] not in data:
data[row['PipelineID']] = {}
data[row['PipelineID']][row['Pipeline']] = [row[key] for key in columns]
data2 = data[category]
data = {}
with open('../../results/final_results/final_results_main_structured.csv') as f:
reader = csv.DictReader(f, delimiter=';')
for row in reader:
if row['PipelineID'] not in data:
data[row['PipelineID']] = {}
data[row['PipelineID']][row['Pipeline']] = [row[key] for key in columns]
data3 = data[category]
# +
labels = ['LR_CV_RU', 'LR_TFIDF_RU', 'SVC_CV_RU', 'SVC_TFIDF_RU', 'KNN_CV_RU', 'KNN_TFIDF_RU', 'RFC_CV_RU', 'RFC_TFIDF_RU', 'LSVC_CV_RU', 'LSVC_TFIDF_RU']
xticknum = len(labels)
y1 = []
x1 = []
i = 0
y2 = []
x2 = []
y3 = []
x3 = []
plt.figure(figsize=(20,8))
count = 0
for key in data1.keys():
for ind in range(len(data1[key])):
y1.append(float(data1[key][ind]))
y2.append(float(data2[key][ind]))
y3.append(float(data3[key][ind]))
x1.append(i)
x2.append(i)
x3.append(i)
if count % 4 == 0:
plt.bar(i, float(data1[key][ind]), 3, alpha=0.7, color='red')
plt.bar(i+3, float(data2[key][ind]), 3, alpha=0.7, color='blue')
plt.bar(i+6, float(data3[key][ind]), 3, alpha=0.7, color='green')
elif count % 4 == 1:
plt.plot(i, float(data1[key][ind]), 'p', alpha=0.7, c='red')
plt.plot(i, float(data2[key][ind]), 'p', alpha=0.7, c='blue')
plt.plot(i, float(data3[key][ind]), 'p', alpha=0.7, c='green')
elif count % 4 == 2:
plt.plot(i, float(data1[key][ind]), '*', alpha=0.7, c='red')
plt.plot(i, float(data2[key][ind]), '*', alpha=0.7, c='blue')
plt.plot(i, float(data3[key][ind]), '*', alpha=0.7, c='green')
elif count % 4 == 3:
plt.plot(i, float(data1[key][ind]), 'h', alpha=0.7, c='red')
plt.plot(i, float(data2[key][ind]), 'h', alpha=0.7, c='blue')
plt.plot(i, float(data3[key][ind]), 'h', alpha=0.7, c='green')
count += 1
i += metric_count
i += metric_count*metric_count
#plt.show()
j = 0
for u in range(len(data1.keys())-1):
j += metric_count*metric_count*2
plt.axvline(x = j-metric_count*2+1, ymin = 0, ymax=1, c='lightgray')
plt.xticks(range(0, i, int(i/xticknum)), labels, rotation=60)
plt.title(category +' experiments')
plt.xlabel('Pipeline')
plt.ylabel('Score')
#plt.plot(x1, y1, 'x', label='Baseline', alpha=0.7, c='red')
#plt.plot(x2, y2, 'o', label='Negative samples', alpha=0.7, c='blue')
#plt.plot(x3, y3, 'og', label='Base structured')
#plt.tight_layout()
plt.ylim(0.0,1.0)
plt.subplots_adjust(bottom=0.2)
legend1 = plt.legend(['Baseline'], bbox_to_anchor=(0,0,1,1.2))
plt.legend(['Accuracy', 'Precision'], bbox_to_anchor=(0,0,0,1.2))
plt.gca().add_artist(legend1)
#plt.savefig("../../results/pics/"+"_".join(category.lower().split(' '))+"_negative_samples.svg")
#plt.legend(loc='upper left')
#plt.tight_layout()
plt.show()
plt.clf()
| src/util/graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ASpBp5GzOYGT" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras import datasets, layers, models
import matplotlib.pyplot as plt
# + id="SeUhQXZCOgiy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="afaf05b2-1661-4d14-b74e-d1b3be043570"
(train_images, train_labels), (test_images, test_labels) = datasets.cifar10.load_data()
train_images, test_images = train_images / 255.0, test_images / 255.0
# + id="rr5xkcsGOr4F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="ec064b75-572a-4ea1-e728-9f0d02c6f293"
class_names = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i][0]])
plt.show()
# + id="vDG2_ji7Pb5r" colab_type="code" colab={}
model = models.Sequential()
model.add(layers.Conv2D(32, (3,3), activation="relu", input_shape=(32,32,3)))
model.add(layers.MaxPooling2D((2,2)))
model.add(layers.Conv2D(64, (3,3), activation="relu"))
model.add(layers.MaxPooling2D((2,2)))
model.add(layers.Conv2D(64, (3,3), activation="relu"))
# + id="4I32xdpGQBl8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="94dcee8e-0e0f-45b2-de7f-812d999aacee"
model.summary()
# + id="-rl8wYWOQG34" colab_type="code" colab={}
model.add(layers.Flatten())
model.add(layers.Dense(64, activation="relu"))
model.add(layers.Dense(10))
# + id="6hhze8tnQZcY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 433} outputId="fbbffc86-8f41-4891-c41f-80719f0fed57"
model.summary()
# + id="hu7eDUqYQayf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 363} outputId="09d0525f-3313-4fc9-be25-e450665fee43"
model.compile(optimizer="adam",
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=["accuracy"])
history = model.fit(train_images, train_labels, epochs=10, validation_data=(test_images, test_labels))
# + id="pqNAHMHwQr_x" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="1c8b41a5-1151-4335-c222-4056707617eb"
plt.plot(history.history["accuracy"], label="accuracy")
plt.plot(history.history["val_accuracy"], label="val_accuracy")
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
plt.ylim([0.5, 1])
plt.legend(loc="lower right")
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
# + id="YJOoPZvsRku-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0165aedf-211f-4a8a-e29d-7f2f83ac80e1"
print(test_acc)
| tensorflow/images/CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_19cls48"
# # Graph Breadth First Search
# In this exercise, you'll see how to do a breadth first search on a graph. To start, let's create a graph class in Python.
# + graffitiCellId="id_pmkelaq"
class GraphNode(object):
def __init__(self, val):
self.value = val
self.children = []
def add_child(self,new_node):
self.children.append(new_node)
def remove_child(self,del_node):
if del_node in self.children:
self.children.remove(del_node)
class Graph(object):
def __init__(self,node_list):
self.nodes = node_list
def add_edge(self,node1,node2):
if(node1 in self.nodes and node2 in self.nodes):
node1.add_child(node2)
node2.add_child(node1)
def remove_edge(self,node1,node2):
if(node1 in self.nodes and node2 in self.nodes):
node1.remove_child(node2)
node2.remove_child(node1)
# + [markdown] graffitiCellId="id_dk66y5p"
# Now let's create the graph.
# + graffitiCellId="id_4twme6x"
nodeG = GraphNode('G')
nodeR = GraphNode('R')
nodeA = GraphNode('A')
nodeP = GraphNode('P')
nodeH = GraphNode('H')
nodeS = GraphNode('S')
graph1 = Graph([nodeS,nodeH,nodeG,nodeP,nodeR,nodeA] )
graph1.add_edge(nodeG,nodeR)
graph1.add_edge(nodeA,nodeR)
graph1.add_edge(nodeA,nodeG)
graph1.add_edge(nodeR,nodeP)
graph1.add_edge(nodeH,nodeG)
graph1.add_edge(nodeH,nodeP)
graph1.add_edge(nodeS,nodeR)
# + [markdown] graffitiCellId="id_iplss81"
# ## Implement BFS
# Using what you know about BFS for trees and DFS for graphs, let's do BFS for graphs. Implement the `bfs_search` to return the `GraphNode` with the value `search_value` starting at the `root_node`.
# + graffitiCellId="id_1vk7aeh"
import copy
def bfs_search(root_node, search_value):
goal = False
frontier_nodes = root_node.children
#print("frontier_nodes.value")
#print(frontier_nodes[0].value)
#print(len(frontier_nodes))
history_nodes = []
history_nodes.append(root_node)
while goal == False:
print("-----------")
for node in frontier_nodes:
print("node.value")
print(node.value)
if node not in history_nodes:
if(node.value == search_value):
goal = True
print("Achieved goal!")
return node
else:
history_nodes.append(node)
next_nodes = []
for node in frontier_nodes:
for child_node in node.children:
if child_node not in history_nodes:
print("child_node.value")
print(child_node.value)
next_nodes.append(child_node)
frontier_nodes = copy.deepcopy(next_nodes)
print("len(frontier_nodes)")
print(len(frontier_nodes))
# + [markdown] graffitiCellId="id_fg1wpq1"
# <span class="graffiti-highlight graffiti-id_fg1wpq1-id_g7fi7m5"><i></i><button>Hide Solution</button></span>
# + graffitiCellId="id_g7fi7m5"
# Solution
def bfs_search(root_node, search_value):
visited = set() # Sets are faster while lookup. Lists are faster to iterate.
queue = [root_node]
while len(queue) > 0:
current_node = queue.pop(0)
visited.add(current_node)
if current_node.value == search_value:
return current_node
for child in current_node.children:
if child not in visited: # Lookup
queue.append(child)
# + [markdown] graffitiCellId="id_ej37296"
# ### Tests
# + graffitiCellId="id_ajsx9hw"
assert nodeA == bfs_search(nodeS, 'A')
assert nodeS == bfs_search(nodeP, 'S')
assert nodeR == bfs_search(nodeH, 'R')
# + graffitiCellId="id_8z41elb"
| Exercises/2_Search/1_graph_bfs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numerical and logical functions for working with iterators
#
# These functions are always available. You don't need to import them.
#
# ## `any()`: checks if at least one element evaluates to `True`
#
# Without `any()`:
# +
none_true = [0, 0, 0]
some_true = [0, 1, 0]
all_true = [1, 1, 1]
def check_any(i):
for e in i:
if e:
return True
return False
check_any(none_true)
# -
# With `any()`:
any(none_true)
# An equivalent implementation using a generator expression:
True in (bool(e) for e in none_true)
# ## `all(): checks if all elements evaluates to `True`
#
# Without `all()`:
# +
def check_all(i):
for e in i:
if not e:
return False
return True
check_all(none_true)
# -
# With `all()`:
all(none_true)
# An equivalent implementation using a generator expression:
False not in (bool(e) for e in none_true)
# ## sorted(), min(), max(), and sum()
# `sorted()` takes an Iterator with numeric elements, sorts it, and returns a `list`:
numbers = [2, -1, 2, 4]
sorted(numbers)
# Without `min()` and `max()`:
sorted(numbers)[-1]
# With `min()` and `max()`:
max(numbers)
# Without `sum()`:
# +
def get_sum(i):
total = 0
for e in i:
total += e
return total
get_sum(numbers)
# -
# With `sum()`:
sum(numbers)
| Functional_Thinking/Lab/30B-numerical-and-logical-functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sonify
import numpy as np
#reading the data
water_data = pd.read_csv('adaSensorData.csv')
water = pd.DataFrame(water_data)
#playing with pH data
ph = water['pH']
ph = ph[:10]
index = [0,1,2,3,4,5,6,7,8,9]
normalized_index = sonify.scale_list_to_range(index, new_min=0, new_max=9)
normalized_pH = sonify.scale_list_to_range(ph, new_min=0, new_max=10)
normed_ph_data = list(zip(index, normalized_pH))
sonify.play_midi_from_data(normed_ph_data)
| examples/Sonification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("../")
import numpy as np
from quantum_compiler.ShapeModule import setFunc
from quantum_compiler.WaveModule import Wave, Waveform,QubitChannel
from instruments.drivers.Tektronix.Tektronix_AWG import AWG5208
# +
# connect to AWG
awg = AWG5208(
inst_name='AWG5208',
inst_address='TCPIP0::192.168.10.42::inst0::INSTR')
# help(awg)
# set sampling rate and reference clock
awg.set_sample_rate(sample_rate=1.0E9)
awg.set_extref_source(ref_freq=10E6)
awg.clr_wfm()
awg.clr_seq()
# +
exp_peak = 5e-6
tau = 280e-9
flat = 900e-9
num_cases = 11
amp_array = np.linspace(0,1,num_cases)
marker = Wave(setFunc('square', {'start': 10e-9,'flat':10e-9}, 10e-6))
for i in amp_array:
a = ~(i*Wave(setFunc('exp_rising', {'peak_x': exp_peak,'tau':tau}, 10e-6)))
a.plot()
b = ~Wave(setFunc('square', {'start': exp_peak,'flat':flat}, 10e-6))
# a.plot()
marker.plot()
b.plot()
# +
#make exp rising pulse (change amp)
for i in amp_array:
awg.set_wfm(wfm_name=f'rising_amp_{i}',
wfm = (~(i*Wave(setFunc('exp_rising', {'peak_x': exp_peak,'tau':tau}, 10e-6)))).y,
mkr1= (~Wave(setFunc('square', {'start': 10e-9,'flat':10e-9}, 10e-6))).y
)
#make flux tunning pulse
awg.set_wfm(wfm_name= 'square_test',
wfm = (~Wave(setFunc('square', {'start': exp_peak,'flat':flat}, 10e-6))).y,
)
awg.upload_wfm()
# +
seq_name='exp_rising'
trackList = ["exp_rising_amp"]
# clear, set, assign and upload
awg.clr_seq()
awg.set_seq(seq_name, num_track=len(trackList), num_step=num_cases)
for track_idx, track in enumerate(trackList):
for step_idx, amp in enumerate(amp_array):
awg.assign_seq(f'rising_amp_{amp}', seq_name, track_idx+1, step_idx+1)
awg.upload_seq(seq_name)
# -
awg.assign_ch(1, seq_name, track_index=1)
awg.assign_ch(2,'square_test')
| measure_scripts/exp_rising_change_amp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# +
import glob
import os
dirs = glob.glob(os.path.join('./BlobStorage/test_data/*/*jpg'))
#print(dirs)
# +
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
for file in dirs:
try:
img = mpimg.imread(file)
#print(img.shape)
except:
os.remove(file)
# -
| azure_code/cleanFiles.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Geomorphometry II: Spatial and Temporal Terrain Analysis
#
# Resources: [
# GRASS GIS overview and manual](https://grass.osgeo.org/grass76/manuals/index.html),
# [GRASSbook](http://www.grassbook.org/).
#
#
# Download Mapset and color tables:
#
# * Download [
# NagsHead time series](http://fatra.cnr.ncsu.edu/geospatial-modeling-course/data/NagsHead_series.zip) and copy it into your nc_spm_08_grass7 Location
# (it should be the same level directory as PERMANENT).
# Do not let your unzipping program create additional level directory with the same name!
# If you are not sure about GRASS GIS Database structure read about it in
# [the manual](https://grass.osgeo.org/grass76/manuals/grass_database.html).
# * Custom color table for time series standard deviations map [stddev_color.txt](data/stddev_color.txt)
#
#
#
# ### Start GRASS GIS
# Start GRASS - click on GRASS icon or type
# using Python to initialize GRASS GIS
# This is a quick introduction into Jupyter Notebook.
# Python code can be executed like this:
a = 6
b = 7
c = a * b
print("Answer is", c)
# Python code can be mixed with command line code (Bash).
# It is enough just to prefix the command line with an exclamation mark:
# !echo "Answer is $c"
# Use Shift+Enter to execute this cell. The result is below.
# +
# using Python to initialize GRASS GIS
import os
import sys
import subprocess
from IPython.display import Image
# create GRASS GIS runtime environment
gisbase = subprocess.check_output(["grass", "--config", "path"], text=True).strip()
os.environ['GISBASE'] = gisbase
sys.path.append(os.path.join(gisbase, "etc", "python"))
# do GRASS GIS imports
import grass.script as gs
import grass.script.setup as gsetup
# set GRASS GIS session data
rcfile = gsetup.init(gisbase, "/home/jovyan/grassdata", "nc_spm_08_grass7", "user1")
# -
# using Python to initialize GRASS GIS
# default font displays
os.environ['GRASS_FONT'] = 'sans'
# overwrite existing maps
os.environ['GRASS_OVERWRITE'] = '1'
gs.set_raise_on_error(True)
gs.set_capture_stderr(True)
# using Python to initialize GRASS GIS
# set display modules to render into a file (named map.png by default)
os.environ['GRASS_RENDER_IMMEDIATE'] = 'cairo'
os.environ['GRASS_RENDER_FILE_READ'] = 'TRUE'
os.environ['GRASS_LEGEND_FILE'] = 'legend.txt'
# In startup pannel set GRASS GIS Database Directory to path to datasets,
# for example on MS Windows, `C:\Users\myname\grassdata`.
# For GRASS Location select nc_spm_08_grass7 (North Carolina, State Plane, meters) and
# for GRASS Mapset create a new mapset (called e.g. HW_terrain_analysis).
# Click Start GRASS session.
#
#
# Change working directory:
# _Settings_ > _GRASS working environment_ > _Change working directory_ > select/create any directory
# or type `cd` (stands for change directory) into the GUI
# _Console_ and hit Enter:
# a proper directory is already set, download files
import urllib.request
urllib.request.urlretrieve("http://ncsu-geoforall-lab.github.io/geospatial-modeling-course/grass/data/stddev_color.txt", "stddev_color.txt")
# Download all text files (see above)
# to the selected directory. Now you can use the commands from the assignment requiring the text file
# without the need to specify the full path to the file.
#
#
# ### Compute basic topographic parameters: slope and aspect
# !g.region raster=elevation -p
# !r.slope.aspect elevation=elevation slope=myslope aspect=myaspect
# Display resulting maps with legend using GUI.
# !d.rast myslope
# !d.legend myslope at=2,40,2,6
Image(filename="map.png")
# !d.rast myaspect
# !d.legend myaspect at=2,40,2,6
Image(filename="map.png")
# Show impact of integer values in meters on slope and aspect pattern.
# Compute integer DEM and derive its slope and aspect.
# Use GUI to display the histogram: in _Map Display_ > _Analyze_ > _Create histogram_:
# !r.mapcalc "elev_int = int(elevation)"
# !r.slope.aspect elevation=elev_int aspect=aspect_int_10m slope=slope_int_10m
# !d.erase
# !d.histogram slope_int_10m
# !d.histogram myslope
# !d.histogram aspect_int_10m
# !d.histogram myaspect
# !d.erase
# !d.rast myslope
Image(filename="map.png")
# Zoom into NW area of the current region (relatively flat area near large interchange).
# Can you explain the difference in slope maps derived from integer (m vertical resolution)
# and floating point (mm vertical resolution) DEMs?
# !d.rast slope_int_10m
Image(filename="map.png")
# ### Compute slope along road
# First set the region to the extent of the bus route #11 and to 10m resolution.
# Then convert the vector line of the route to raster using the direction of the route.
# !g.region vect=busroute11 align=elevation res=10 -p
# !v.to.rast input=busroute11 type=line output=busroute11_dir use=dir
# !r.colors map=busroute11_dir color=aspect
# !d.rast busroute11_dir
# !d.legend busroute11_dir
Image(filename="map.png")
# Compute the steepest slope of the topography along the route by assigning
# the values of slope derived from a DEM in the first part of this assignment
# to the grid cells along the route.
# !r.mapcalc "route_slope = if(busroute11_dir, myslope)"
# Then compute the slope in the direction of the route using the difference between aspect of the topography
# and the route direction angles.
# !r.mapcalc "route_slope_dir = abs(atan(tan(myslope) * cos(myaspect - busroute11_dir)))"
# !r.colors map=route_slope,route_slope_dir color=slope
# Display the results along with the elevation contours and compute univariate statistics.
# Comment on the difference of the two results.
# !r.contour input=elevation output=contours step=2
# !d.vect contours
# !d.rast route_slope
# !d.legend route_slope
Image(filename="map.png")
# !d.rast route_slope_dir
Image(filename="map.png")
# !r.univar route_slope
# !r.univar route_slope_dir
Image(filename="map.png")
# ### Curvatures
#
#
#
# Compute slope, aspect and curvatures simultaneously with interpolation.
# You can do the examples below for the bare earth data only (first example),
# multiple return example is optional (if you are curious how it differs from BE).
# !g.region rural_1m -p
# !v.surf.rst input=elev_lid792_bepts elevation=elev_lid_1m aspect=asp_lid_1m pcurvature=pc_lid_1m tcurvature=tc_lid_1m npmin=120 segmax=25
# !v.surf.rst input=elev_lidrural_mrpts elevation=elev_lidmr_1m aspect=asp_lidmr_1m pcurvature=pc_lidmr_1m tcurvature=tc_lidmr_1m npmin=120 segmax=25 tension=300 smooth=1.
# Display the results as 2D images or in 3D view.
# For 3D view, switch off everything except for elevation surface that you want to view.
# Drape topographic parameters raster maps over DEMs as color.
# !d.erase
# !d.rast elev_lid_1m
# !d.rast pc_lid_1m
Image(filename="map.png")
# !d.rast elev_lidmr_1m
# !d.rast pc_lidmr_1m
Image(filename="map.png")
# The curvature maps reflect survey pattern rather than topographic features.
# So we lower the tension and increase the smoothing.
# You can use multiple displays to compare the results side-by-side.
# !g.region rural_1m -p
# !v.surf.rst input=elev_lid792_bepts elevation=elev_lidt15_1m aspect=asp_lidt15_1m pcurvature=pc_lidt15_1m tcurvature=tc_lidt15_1m npmin=120 segmax=25 tension=15 smooth=1.
# !v.surf.rst input=elev_lidrural_mrpts elevation=elev_lidmrt15_1m aspect=asp_lidmrt15_1m pcurvature=pc_lidmrt15_1m tcurvature=tc_lidmrt15_1m npmin=120 segmax=25 tension=15 smooth=1.
# !d.erase
# !d.rast elev_lidt15_1m
# !d.rast pc_lidt15_1m
Image(filename="map.png")
# !d.rast tc_lidt15_1m
# !d.rast elev_lidmrt15_1m
# !d.rast pc_lidmrt15_1m
Image(filename="map.png")
# ### Landforms
#
#
# Extract landforms at different levels of detail by adjusting the size of moving window.
# Set rural subregion at 1m resolution,
# compute landforms using 9m and 45m neighborhood: read the manual to learn more.
# Explain types of landforms and the role of the neighborhood size.
# !g.region rural_1m -p
# !r.param.scale elev_lid792_1m output=feature9c_1m size=9 method=feature
# !r.param.scale elev_lid792_1m output=feature45c_1m size=45 method=feature
# Display with legend, save images for the report.
# Optionally display the feature maps draped over elev_lid792_1m as color.
# !d.rast feature9c_1m
# !d.legend feature9c_1m at=2,20,2,6
# !d.rast feature45c_1m
# !d.legend feature45c_1m at=2,20,2,6
# !d.vect elev_lid792_cont1m color=brown
Image(filename="map.png")
# ### Raster time series analysis
#
# For this exercise we will use NagsHead_series Mapset you downloaded.
# You have to first make the mapset accessible.
# In GUI: menu _ Settings_ -> _GRASS working environment_ -> _Mapset access_
# or by using a command:
# !g.mapsets operation=add mapset=NagsHead_series
# If you don't see the mapset, make sure you downloaded it and unzipped it correctly.
#
#
# Run the series analysis and explain the results:
# Which maps are core and envelope?
# Which landforms have high standard deviation and what does it mean?
# !g.region raster=NH_2008_1m -p
# !d.erase
# !d.rast NH_2008_1m
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_min method=minimum
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_max method=maximum
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_mintime method=min_raster
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_maxtime method=max_raster
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_range method=range
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_avg method=average
# !r.series NH_1999_1m,NH_2001_1m,NH_2004_1m,NH_2005_1m,NH_2007_1m,NH_2008_1m out=NH_9908_stddev method=stddev
# !r.colors NH_9908_stddev rules=stddev_color.txt
# !d.rast NH_9908_stddev
# !d.rast NH_9908_range
Image(filename="map.png")
# Use cutting plane in 3D view to show the core and envelope.
# Add constant elevation plane at -1m for reference,
# set zexag somewhere 3-5 (the default is too high).
# Assign surfaces constant color, use top or bottom surface for crossection color.
# When using top for color, lower the light source to make
# top surface dark and highlight the crossection.
#
#
#
#
# ### Optional: Cut and fill and volume
#
#
#
# Compute cut and fill for 4m deep excavation to build a facility.
# First, set the region and display facility on top of orthophoto.
# !g.region rural_1m -p
# !d.erase
# !d.rast ortho_2001_t792_1m
# !d.rast facility
Image(filename="map.png")
# Then set (raster) mask to the facility map and find minimum elevation
# within the facility:
# !r.mask raster=facility
# !r.univar elevation
# Minimum which you obtain should be 123.521m.
# Bottom of 4m excavation will be then
#
# ```
# 123.52 - 4 = 119.52
# ```
#
# Use raster algebra to create the excavation:
# !r.mapcalc "excavation = elevation - 119.52"
# !r.univar excavation
# !d.rast excavation
Image(filename="map.png")
# Minimum you get should be 4.00057 and maximum 9.50554.
# Note that the excavation is limited by the mask we set earlier,
# so we can now do global operation to compute the volume
# which applies just the the facility.
# !r.volume excavation
# Now remove mask. This is important so that
# your future work is not affected.
# !r.mask -r
# end the GRASS session
os.remove(rcfile)
| notebooks/terrain_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
from IPython.core.display import HTML
css = open('style-table.css').read() + open('style-notebook.css').read()
HTML('<style>{}</style>'.format(css))
titles = pd.read_csv('data/titles.csv', index_col=None)
titles.head()
cast = pd.read_csv('data/cast.csv', index_col=None)
cast.head()
# ### How many movies are listed in the titles dataframe?
# ### What are the earliest two films listed in the titles dataframe?
# ### How many movies have the title "Hamlet"?
# ### How many movies are titled "North by Northwest"?
# ### When was the first movie titled "Hamlet" made?
# ### List all of the "Treasure Island" movies from earliest to most recent.
# ### How many movies were made in the year 1950?
# ### How many movies were made in the year 1960?
# ### How many movies were made from 1950 through 1959?
# ### In what years has a movie titled "Batman" been released?
# ### How many roles were there in the movie "Inception"?
# ### How many roles in the movie "Inception" are NOT ranked by an "n" value?
# ### But how many roles in the movie "Inception" did receive an "n" value?
# ### Display the cast of "North by Northwest" in their correct "n"-value order, ignoring roles that did not earn a numeric "n" value.
# ### Display the entire cast, in "n"-order, of the 1972 film "Sleuth".
# ### Now display the entire cast, in "n"-order, of the 2007 version of "Sleuth".
# ### How many roles were credited in the silent 1921 version of Hamlet?
# ### How many roles were credited in Branagh’s 1996 Hamlet?
# ### How many "Hamlet" roles have been listed in all film credits through history?
# ### How many people have played an "Ophelia"?
# ### How many people have played a role called "The Dude"?
# ### How many people have played a role called "The Stranger"?
# ### How many roles has <NAME> played throughout his career?
# ### How many roles has <NAME> played?
# ### List the supporting roles (having n=2) played by <NAME> in the 1940s, in order by year.
# ### List the leading roles that <NAME> played in the 1940s in order by year.
# ### How many roles were available for actors in the 1950s?
# ### How many roles were avilable for actresses in the 1950s?
# ### How many leading roles (n=1) were available from the beginning of film history through 1980?
# ### How many non-leading roles were available through from the beginning of film history through 1980?
# ### How many roles through 1980 were minor enough that they did not warrant a numeric "n" rank?
t = titles.head()
t.sort_values('year')
| Exercises-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iBiha3UzKmVX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0e6b4f9f-4e75-4e67-ee94-f5951f6ea954" executionInfo={"status": "ok", "timestamp": 1545002621511, "user_tz": 180, "elapsed": 2092, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}}
import numpy as np
import matplotlib.pyplot as plt
import mnist
# %matplotlib inline
# + id="IAha9QNTQomd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="5f57f398-5e99-446a-e2ce-767382b860e1" executionInfo={"status": "ok", "timestamp": 1545002625123, "user_tz": 180, "elapsed": 5665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}}
from keras import datasets
(X_train, y_train), (X_test, y_test) = datasets.mnist.load_data()
# + id="z3UcNr8VQqJQ" colab_type="code" colab={}
from keras.utils import to_categorical
X_train = np.expand_dims(X_train, axis=3)
X_test = np.expand_dims(X_test, axis=3)
X_train = X_train / 255
X_test = X_test / 255
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
# + id="MbxXPjqDQaCl" colab_type="code" colab={}
model = mnist.build_model()
# + id="T2zC8UxyRsOe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 578} outputId="37e4e9ea-1c45-4c04-f5bd-c8ecb1592cf9" executionInfo={"status": "ok", "timestamp": 1545002627491, "user_tz": 180, "elapsed": 7968, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}}
model.summary()
# + id="K4OZR6l3pu05" colab_type="code" colab={}
from keras.callbacks import ModelCheckpoint
checkpointer = ModelCheckpoint('weights.hdf5', save_best_only=True, verbose=1)
# + id="jIMLw0WBRcow" colab_type="code" outputId="1f6181c7-5198-459d-d49b-ef6be5c9e5c6" executionInfo={"status": "ok", "timestamp": 1545002898396, "user_tz": 180, "elapsed": 278822, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}} colab={"base_uri": "https://localhost:8080/", "height": 734}
training = model.fit(X_train, y_train, batch_size=32, epochs=10,
validation_data=(X_test, y_test), callbacks=[checkpointer])
# + id="9TFZmD6nT-kl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 403} outputId="735e12ee-27dc-43bf-a149-492c261503a1" executionInfo={"status": "ok", "timestamp": 1545002898400, "user_tz": 180, "elapsed": 278797, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}}
plt.figure(figsize=(9,6))
plt.plot(training.history['loss'], label='Train')
plt.plot(training.history['val_loss'], label='Validation')
plt.title('Model Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# + id="4kNnhhYvUWTc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 403} outputId="236ce2e6-a7b3-4238-c24b-bb14f3b7d084" executionInfo={"status": "ok", "timestamp": 1545002899567, "user_tz": 180, "elapsed": 279928, "user": {"displayName": "<NAME>", "photoUrl": "https://lh6.googleusercontent.com/-rzXqGNbVTGA/AAAAAAAAAAI/AAAAAAAAAbw/sFCkhgEPong/s64/photo.jpg", "userId": "04022106560278464488"}}
plt.figure(figsize=(9,6))
plt.plot(training.history['acc'], label='Train')
plt.plot(training.history['val_acc'], label='Validation')
plt.title('Model Accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
| exemples/mnist/model_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Scikit-Learn PCA and Logistic Regression Pipeline
# ### Using BREASTCANCER_VIEW from DWC. This view has 569 records
# ## Install fedml aws library
pip install fedml-aws --force-reinstall
# ## Import Libraries
from fedml_aws import DwcSagemaker
from fedml_aws import DbConnection
import numpy as np
import pandas as pd
import json
# ## Create DwcSagemaker instance to access libraries functions
dwcs = DwcSagemaker(prefix='<prefix>', bucket_name='<bucket_name>')
# ## Create DbConnection instance to get data from DWC
# Before running the following cell, you should have a config.json file in the same directory as this notebook with the specified values to allow you to access to DWC.
#
# You should also have the follow view `BREASTCANCER_VIEW` created in your DWC. To gather this data, please refer to https://www.kaggle.com/uciml/breast-cancer-wisconsin-data
# %%time
db = DbConnection()
res, column_headers = db.get_data_with_headers(table_name="BREASTCANCER_VIEW", size=1)
data = pd.DataFrame(res, columns=column_headers)
data
data.columns
# ## Train SciKit Model¶
# `train_data` is the data you want to train your model with.
#
# In order to deploy a model to AWS using the Scikit-learn Sagemaker SDK, you must have a script that tells Sagemaker how to train and deploy the model. The path to the script is passed to the `train_sklearn_model` function in the `train_script` parameter.
#
# `instance_type` specifies how much computing power we want AWS to allocate for our services.
clf = dwcs.train_sklearn_model(data,
train_script='pca_pipeline_script.py',
instance_type='ml.c4.xlarge',
wait=True,
download_output=False,
hyperparameters={'n_components':3})
# ## Using the fedml_aws deploy to kyma function
# !aws configure set aws_access_key_id '<aws_access_key_id>' --profile 'sample-pr'
# !aws configure set aws_secret_access_key '<aws_secret_access_key>' --profile 'sample-pr'
# !aws configure set region '<region>' --profile 'sample-pr'
dwcs.deploy_to_kyma(clf, initial_instance_count=1, profile_name='sample-pr')
# ## Using the fedml_aws invoke kyma endpoint function
org_data = data.sample(frac=1).reset_index(drop=True)
org_data = org_data[500:]
org_data.fillna(0, inplace=True)
y = org_data['diagnosis']
X = org_data.drop(['diagnosis'], axis=1)
result = dwcs.invoke_kyma_endpoint(api='<endpoint>',
payload=X.to_json(),
content_type='application/json')
result = result.content.decode()
result
# ## Write back to DWC
X.columns
X.dtypes
# +
# ['ID', 'Units_Sold', 'Unit_Price', 'Unit_Cost', 'Total_Revenue','Total_Cost', 'totalprofit']
db.create_table("CREATE TABLE PCA_Pipeline_Table (ID INTEGER PRIMARY KEY, radius_mean FLOAT(2), texture_mean FLOAT(2), perimeter_mean FLOAT(2), area_mean FLOAT(2), smoothness_mean FLOAT(2), compactness_mean FLOAT(2), concavity_mean FLOAT(2), concave_points_mean FLOAT(2), symmetry_mean FLOAT(2), fractal_dimension_mean FLOAT(2), radius_se FLOAT(2), texture_se FLOAT(2), perimeter_se FLOAT(2), area_se FLOAT(2), smoothness_se FLOAT(2), compactness_se FLOAT(2), concavity_se FLOAT(2), concave_points_se FLOAT(2), symmetry_se FLOAT(2), fractal_dimension_se FLOAT(2), radius_worst FLOAT(2), texture_worst FLOAT(2), perimeter_worst FLOAT(2), area_worst FLOAT(2), smoothness_worst FLOAT(2), compactness_worst FLOAT(2), concavity_worst FLOAT(2), concave_points_worst FLOAT(2), symmetry_worst FLOAT(2), fractal_dimension_worst FLOAT(2), column32 INTEGER, diagnosis_predict VARCHAR(100))")
# -
res = result.strip('][').split(', ')
res
dwc_data = X
dwc_data = dwc_data.assign(diagnosis_predict = res)
dwc_data.columns = ['id', 'radius_mean', 'texture_mean', 'perimeter_mean', 'area_mean',
'smoothness_mean', 'compactness_mean', 'concavity_mean',
'concave_points_mean', 'symmetry_mean', 'fractal_dimension_mean',
'radius_se', 'texture_se', 'perimeter_se', 'area_se', 'smoothness_se',
'compactness_se', 'concavity_se', 'concave_points_se', 'symmetry_se',
'fractal_dimension_se', 'radius_worst', 'texture_worst',
'perimeter_worst', 'area_worst', 'smoothness_worst',
'compactness_worst', 'concavity_worst', 'concave_points_worst',
'symmetry_worst', 'fractal_dimension_worst', 'column32', 'diagnosis_predict']
for i in dwc_data.columns[1:-1]:
dwc_data[i] = dwc_data[i].astype('float64')
dwc_data
dwc_data.dtypes
db.insert_into_table('PCA_Pipeline_Table', dwc_data)
| AWS/sample-notebooks/PCAPipeline/PCAPipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 3 Conditional Execution
# ### 3.1 Boolean expressions
# True and false are values of data type bool.
type(True)
type(False)
5==5
5==7
# Statements using comparison operators result in boolean values: true or false.
x = 5
y = 7
x != y
x > y
x < y
x >= y
x <= y
x is y
x is not y
x = 5
y = 5
x is y
x is not y
# ### 3.2 Logical operators
# There are 3 logical operators: and, or, not
x = 5
x > 0 and x < 10
n = 8
n%2==0 and n%3==0
n%2==0 or n%3==0
# ### 3.3 Conditional execution
x = 9
if x > 0:
print('x is positive')
if x < 0:
pass
# ### 3.4 Alternative execution
x = 13
if x%2 == 0:
print('x is even')
else:
print('x is odd')
# ### 3.5 Chained conditionals
x = 9
y = 20
if x < y:
print('x is less than y')
elif x > y:
print('x is greater than y')
else:
print('x and y are equal')
choice = int(input('Guess a number between 1 and 10'))
correct = 6
if choice < correct - 1:
print('Too low')
elif choice == correct - 1 or choice == correct + 1:
print('Close, but not quite')
elif choice > correct + 1:
print('Too high')
elif choice == correct:
print('Correct!')
# ### 3.6 Nested conditionals
x = 9
if x%2 == 1:
print('x is odd')
if x%3 == 0:
print('x is divisible by 3 but x is not 6')
if 0 < x:
if x < 10:
print('x is a positive single-digit number')
if 0 < x and x < 10:
print('x is a positive single-digit number')
# ### 3.6 Catching exceptions using try and except
inp = input('Enter Fahrenheit temperature:')
fahr = float(inp)
cel = (fahr - 32.0) * 5.0/9.0
print(cel)
inp = input('Enter Fahrenheit temperature:')
fahr = float(inp)
cel = (fahr - 32.0) * 5.0/9.0
print(cel)
inp = input('Enter Fahrenheit temperature:')
try:
fahr = float(inp)
cel = (fahr - 32.0) * 5.0/9.0
print(cel)
except:
print('Please enter a number.')
| 03-conditional-execution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Quandl: US GDP, Implicit Price Deflator
#
# In this notebook, we'll take a look at data set , available on [Quantopian](https://www.quantopian.com/data). This dataset spans from 1961 through the current day. It contains the value for the United States' inflation rate, using the price deflator. We access this data via the API provided by [Quandl](https://www.quandl.com). [More details](https://www.quandl.com/data/UGID/INFL_USA-Inflation-GDP-deflator-United-States-of-America) on this dataset can be found on Quandl's website.
#
# ### Blaze
# Before we dig into the data, we want to tell you about how you generally access Quantopian partner data sets. These datasets are available using the [Blaze](http://blaze.pydata.org) library. Blaze provides the Quantopian user with a convenient interface to access very large datasets.
#
# Some of these sets (though not this one) are many millions of records. Bringing that data directly into Quantopian Research directly just is not viable. So Blaze allows us to provide a simple querying interface and shift the burden over to the server side.
#
# To learn more about using Blaze and generally accessing Quantopian partner data, clone [this tutorial notebook](https://www.quantopian.com/clone_notebook?id=561827d21777f45c97000054).
#
# With preamble in place, let's get started:
# +
# import the dataset
from quantopian.interactive.data.quandl import ugid_infl_usa
# Since this data is public domain and provided by Quandl for free, there is no _free version of this
# data set, as found in the premium sets. This import gets you the entirety of this data set.
# import data operations
from odo import odo
# import other libraries we will use
import pandas as pd
import matplotlib.pyplot as plt
# -
ugid_infl_usa.sort('asof_date')
# The data goes all the way back to 1961 and is updated annually.
#
# Blaze provides us with the first 10 rows of the data for display. Just to confirm, let's just count the number of rows in the Blaze expression:
ugid_infl_usa.count()
# Let's go plot it for fun. This data set is definitely small enough to just put right into a Pandas DataFrame
# +
inf_df = odo(ugid_infl_usa, pd.DataFrame)
inf_df.plot(x='asof_date', y='value')
plt.xlabel("As Of Date (asof_date)")
plt.ylabel("Inflation Rate")
plt.title("US Inflation, Implicit Price Deflator")
plt.legend().set_visible(False)
# -
| docs/memo/notebooks/data/quandl.ugid_infl_usa/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
from numpy.fft import fft, ifft, fftfreq
from scipy import signal
from astropy.stats import LombScargle
from nfft import ndft, nfft, ndft_adjoint, nfft_adjoint
from gatspy.periodic import LombScargleFast
import time
import pdb
plt.style.use('seaborn')
# +
def signal_no_equip(N, fixed=True):
# 3 parts separated in time, one with slight irregularities in time sampling
# another with change of spacing and the last one with big outlier in spacing
T = np.zeros(N)
dt_implicit = 1 / N
t0 = np.linspace(0, 2*int(N/6)-1, 2*int(N/6))
if fixed:
np.random.seed(1)
e = np.random.normal(0, dt_implicit * 0.5, 2*int(N/6))
T[0:2*int(N/6)] = t0 * dt_implicit + e
shift = 30 * dt_implicit
if fixed:
np.random.seed(2)
t0 = np.linspace(2*int(N/6), 3*int(N/6)-1, int(N/6))
e = np.random.normal(0, dt_implicit * 0.5, int(N/6))
T[2*int(N/6):3*int(N/6)] = shift + t0 * dt_implicit / 2 + e
if fixed:
np.random.seed(3)
t0 = np.linspace(3*int(N/6), 4*int(N/6)-1, int(N/6))
e = np.random.normal(0, dt_implicit * 0.5, int(N/6))
T[3*int(N/6):4*int(N/6)] = t0 * 2 * dt_implicit + e
if fixed:
np.random.seed(4)
t0 = np.linspace(4*int(N/6), N-1, N - 4*int(N/6))
e = np.random.normal(0, dt_implicit * 0.5, N - 4*int(N/6))
T[4*int(N/6):N] = 2 * shift + t0 * dt_implicit / 2 + e
T.sort()
# signal is sinusoidal again with same frequency
temp = np.zeros(N)
segment_duration = int(N/3)
init = int(N/10)
times_segment = T[init: init + segment_duration]
times_segment = times_segment - min(times_segment)
freq_sin = 2 / (max(times_segment) - min(times_segment))
# print("choosen freq is: ", freq_sin)
temp[init: init + segment_duration] = np.sin(freq_sin * 2 * np.pi * times_segment)
return temp, freq_sin, T
def get_nfft(Nf, data, temp, t):
dwindow = signal.tukey(len(temp), alpha=1./8)
nfft_d = nfft_adjoint(t, dwindow * data, Nf)
nfft_t = nfft_adjoint(t, dwindow * temp, Nf)
k = -(Nf // 2) + np.arange(Nf)
return nfft_d, nfft_t, k
# -
N = 1200
temp2, freq_sin, t2 = signal_no_equip(N, fixed=False)
print(freq_sin)
temp = temp2[:800]
t = t2[:800]
data = np.random.normal(0, 0.3, 800) + temp
N = 800
plt.plot(t, temp, '.')
plt.plot(t, data, alpha=0.5)
# +
# get the nfft and normalize to unitary
nfft_d, nfft_t, k = get_nfft(N, data, temp, t)
## plot not normalized:
plt.figure()
plt.title("nfff of data/template not normalized")
plt.plot(k, np.abs(nfft_d), 'r', label='data')
plt.plot(k, np.abs(nfft_t), 'b', label='temp')
plt.legend()
## normalize for L2
norm = np.sqrt((np.abs(nfft_t)**2).sum())
nfft_d /= norm
nfft_t /= norm
plt.figure()
plt.title("nfft of data/template normalized by L2")
plt.plot(k, np.abs(nfft_d), 'r', label='data')
plt.plot(k, np.abs(nfft_t), 'b', label='temp')
plt.legend()
# +
## normalize data to unity by L2
norm_data = np.sqrt((np.abs(temp)**2).sum())
data /= norm_data
temp /= norm_data
plt.figure()
plt.title("normalized data/template by L2")
plt.plot(t, data, 'r', label='data')
plt.plot(t, temp, 'b', label='temp')
plt.legend()
# -
# check the parseval theorem for the inverse fourier transform
print(np.sum(np.abs(data)**2), np.sum(np.abs(nfft_d)**2))
# +
## try doing the SNR with this normalization
def get_psd(k, t, data, min_freq=None, data_per_peak=1):
df = 1 / ((max(t) - min(t)) * data_per_peak)
if min_freq is None:
min_freq = 0.5 * df
NK = len(k)
if NK % 2 == 0: # par
N = int(NK / 2)
else:
N = int((NK-1) / 2)
max_freq = (N - 1) * df + min_freq
frequency, power = LombScargle(t, data).autopower(maximum_frequency=max_freq, minimum_frequency=min_freq,
samples_per_peak=data_per_peak)
if len(frequency) != N:
raise ValueError("algo malo")
return frequency, power, df
def snr_no_equip(N, only_noise=False, fixed=False):
temp, freq_sin, t = signal_no_equip(N, fixed=fixed)
if only_noise:
if fixed:
np.random.seed(12312)
data = np.random.normal(0, 0.3, N)
else:
data = np.random.normal(0, 0.3, N) + temp
## get the first 800 data generated
temp = temp[:800]
data = data[:800]
t = t[:800]
N = 800
## normalize the data/temp by L2
norm_data = np.sqrt((np.abs(temp)**2).sum())
temp /= norm_data
data /= norm_data
# calcula la psd
Nf = 4 * N
k = -(Nf // 2) + np.arange(Nf)
freqs, pw, df = get_psd(k, t, data)
# repite la psd para obtener los datos con frequencias negativas, si Nf es par entonces el ultimo no se repite
pw = np.append(pw, pw)
if Nf % 2 == 0:
pw = np.delete(pw, len(pw) - 1)
nfft_d, nfft_t, k = get_nfft(Nf, data, temp, t)
nfft_d = np.delete(nfft_d, 0) # remving the value corresponding to 0 frequency
nfft_t = np.delete(nfft_t, 0)
## to get this as even remove another freq, for this time it will be the last one
last_one = len(pw)-1
nfft_d = np.delete(nfft_d, last_one)
nfft_t = np.delete(nfft_t, last_one)
pw = np.delete(pw, last_one)
## normaliza las nfft por L2
norm_nfft = np.sqrt((np.abs(nfft_t)**2).sum())
nfft_d /= norm_nfft
nfft_t /= norm_nfft
# check the parseval
print("parseval theorem: ", np.sum(np.abs(data)**2), np.sum(np.abs(nfft_d)**2))
norm_sigma = 4 * df
h_norm = (nfft_t * nfft_t.conjugate() / pw).sum()
norm_corr = 4 * df / np.sqrt(h_norm.real * norm_sigma)
corr = nfft_d * nfft_t.conjugate() / pw / (2*Nf)
inv_nfft = nfft(t, corr)
# check parseval again
print("parseval: ", np.sum(np.abs(data)**2), np.sum(np.abs(inv_nfft)**2))
snr = inv_nfft * norm_corr * (max(t) - min(t)) * (len(nfft_d) - 1) / N
snr = np.roll(snr, len(snr) // 2)
return t, np.abs(snr), data, temp
# -
N = 1200
t, snr, data, temp = snr_no_equip(N, only_noise=False, fixed=False)
plt.figure()
plt.title("data")
plt.plot(t, data, 'r', alpha=0.5, label='data')
plt.plot(t, temp, 'b.', label='temp')
plt.legend()
plt.figure()
plt.title("SNR")
plt.plot(t-t[len(t)//2], snr)
| developing/notebooks/ParsevalTheorem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><i>This notebook was put together by [<NAME>](http://www.vanderplas.com). Source and license info is on [GitHub](https://github.com/jakevdp/sklearn_tutorial/).</i></small>
# # An Introduction to scikit-learn: Machine Learning in Python
# ## Goals of this Tutorial
# - **Introduce the basics of Machine Learning**, and some skills useful in practice.
# - **Introduce the syntax of scikit-learn**, so that you can make use of the rich toolset available.
# ## Schedule:
# **Preliminaries: Setup & introduction** (15 min)
# * Making sure your computer is set-up
#
# **Basic Principles of Machine Learning and the Scikit-learn Interface** (45 min)
# * What is Machine Learning?
# * Machine learning data layout
# * Supervised Learning
# - Classification
# - Regression
# - Measuring performance
# * Unsupervised Learning
# - Clustering
# - Dimensionality Reduction
# - Density Estimation
# * Evaluation of Learning Models
# * Choosing the right algorithm for your dataset
#
# **Supervised learning in-depth** (1 hr)
# * Support Vector Machines
# * Decision Trees and Random Forests
#
# **Unsupervised learning in-depth** (1 hr)
# * Principal Component Analysis
# * K-means Clustering
# * Gaussian Mixture Models
#
# **Model Validation** (1 hr)
# * Validation and Cross-validation
# ## Preliminaries
# This tutorial requires the following packages:
#
# - Python version 2.7 or 3.4+
# - `numpy` version 1.8 or later: http://www.numpy.org/
# - `scipy` version 0.15 or later: http://www.scipy.org/
# - `matplotlib` version 1.3 or later: http://matplotlib.org/
# - `scikit-learn` version 0.15 or later: http://scikit-learn.org
# - `ipython`/`jupyter` version 3.0 or later, with notebook support: http://ipython.org
# - `seaborn`: version 0.5 or later, used mainly for plot styling
#
# The easiest way to get these is to use the [conda](http://store.continuum.io/) environment manager.
# I suggest downloading and installing [miniconda](http://conda.pydata.org/miniconda.html).
#
# The following command will install all required packages:
# ```
# $ conda install numpy scipy matplotlib scikit-learn ipython-notebook
# ```
#
# Alternatively, you can download and install the (very large) Anaconda software distribution, found at https://store.continuum.io/.
# ### Checking your installation
#
# You can run the following code to check the versions of the packages on your system:
#
# (in IPython notebook, press `shift` and `return` together to execute the contents of a cell)
# +
from __future__ import print_function
import IPython
print('IPython:', IPython.__version__)
import numpy
print('numpy:', numpy.__version__)
import scipy
print('scipy:', scipy.__version__)
import matplotlib
print('matplotlib:', matplotlib.__version__)
import sklearn
print('scikit-learn:', sklearn.__version__)
import seaborn
print('seaborn', seaborn.__version__)
# -
# ## Useful Resources
# - **scikit-learn:** http://scikit-learn.org (see especially the narrative documentation)
# - **matplotlib:** http://matplotlib.org (see especially the gallery section)
# - **IPython:** http://ipython.org (also check out http://nbviewer.ipython.org)
| Scikit-learn/sklearn_tutorial_notebooks/01-Preliminaries 24-10-2017.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="XO0nr0mHm3y8" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + id="XBh8lo6vnFkJ" colab_type="code" colab={}
x=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_train.csv')
y=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_labels_train.csv')
x_test=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/dengue_features_test.csv')
# + id="o9tXwvsKoZZ0" colab_type="code" outputId="07e32651-a808-4dfd-f298-8177c00ef4be" colab={"base_uri": "https://localhost:8080/", "height": 1181}
x.info()
y.info()
x_test.info()
# + id="Vxu-4W5OoeJy" colab_type="code" outputId="e29780c7-11ca-4f1d-811c-f1c18ab275b5" colab={"base_uri": "https://localhost:8080/", "height": 69}
x['city'].value_counts()
# + id="a4lrj0FBqqQy" colab_type="code" outputId="0ba4bc2c-67df-48d9-8e59-68c2c75174ce" colab={"base_uri": "https://localhost:8080/", "height": 69}
x_test['city'].value_counts()
# + id="3CWrDd8wrSTh" colab_type="code" outputId="f01c211b-b447-4c3a-ba44-22cfe124f76d" colab={"base_uri": "https://localhost:8080/", "height": 1024}
x.fillna(method='ffill', inplace=True)
x_test.fillna(method='ffill', inplace=True)
x.info()
x_test.info()
# + id="6KE15nsIsb_m" colab_type="code" colab={}
x1=x
# + id="Al9kadyUs2lI" colab_type="code" outputId="a5a6c15d-8938-4dc3-f75c-472a1cef4867" colab={"base_uri": "https://localhost:8080/", "height": 521}
x1.info()
# + id="wL59Y4qys4eH" colab_type="code" colab={}
x1.drop(columns='city',inplace=True)
x1.drop(columns='year',inplace=True)
x1.drop(columns='weekofyear',inplace=True)
# + id="SmxOL2AbGldX" colab_type="code" outputId="3ec379aa-6413-496f-f83e-b859168b9a4b" colab={"base_uri": "https://localhost:8080/", "height": 173}
y.head()
y.info()
# + id="Cogm_rMgtDM0" colab_type="code" outputId="8f71843b-29e1-4ee7-ec99-3e5980f39e81" colab={"base_uri": "https://localhost:8080/", "height": 538}
data=pd.concat([x1,y],axis=1)
data.head()
data.info()
# + id="P--ObURGyNv6" colab_type="code" colab={}
from sklearn.utils import shuffle
data = shuffle(data)
# + id="OZpUngIIt4qx" colab_type="code" outputId="e3017d2e-13cb-44c7-a368-ab0e7840d5f1" colab={"base_uri": "https://localhost:8080/", "height": 1070}
corr=data.corr()
plt.figure(figsize=(20,15))
ax=sns.heatmap(corr,vmin=-1,vmax=1,annot=True)
# + id="UmXVS_qHuOV8" colab_type="code" outputId="4c17a638-0a5c-45bf-d5ca-007e788abce3" colab={"base_uri": "https://localhost:8080/", "height": 538}
data.info()
# + id="nsFiLml-wSy7" colab_type="code" outputId="f07821e6-0048-440b-f4c6-00092d176751" colab={"base_uri": "https://localhost:8080/", "height": 226}
data.head()
# + id="wPeHxgCb0Ezv" colab_type="code" colab={}
data['city1']=np.where(data['city']=='sj',1,-1)
data.drop(columns='city',inplace=True)
# + id="egtitL_ly95S" colab_type="code" outputId="be3054fc-2821-48ad-b66c-d9f7e6593bcb" colab={"base_uri": "https://localhost:8080/", "height": 538}
data.info()
# + id="SNhF_n2Zy_5H" colab_type="code" colab={}
data.drop(columns='week_start_date',inplace=True)
# + id="ArH3_9jBzJur" colab_type="code" outputId="b3304395-1bf1-45cf-8d82-87cec5408455" colab={"base_uri": "https://localhost:8080/", "height": 521}
data.info()
# + id="Hbf302ZzzMFS" colab_type="code" colab={}
x1=data
y1=data[['total_cases']]
x1.drop(columns='total_cases',inplace=True)
# + id="cbeDy4FGzxkA" colab_type="code" outputId="31526dff-eaa3-40e7-cd58-f63f280191d5" colab={"base_uri": "https://localhost:8080/", "height": 817}
y1.info()
x1.info()
x1.tail()
# + id="_tdca0qfz1Y6" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
seed = 0
test_size = 0.3
X_train, X_test, y_train, y_test = train_test_split(x1, y1, test_size=test_size, random_state=seed)
# + id="nMtrh3JYs3d1" colab_type="code" colab={}
# + id="FIv1N13w6EPK" colab_type="code" outputId="beca053f-638a-43ac-cc50-cb4c1ce88a6e" colab={"base_uri": "https://localhost:8080/", "height": 156}
from xgboost import XGBRegressor
my_model = XGBRegressor(learning_rate=0.1,subsample=0.75,max_depth=6)
my_model.fit(X_train, y_train)
# + id="6tmtUyZA7-2l" colab_type="code" outputId="56be698c-5a98-4142-d0e7-734ae2a71fe7" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import mean_absolute_error
predictions = my_model.predict(X_test)
print("Mean Absolute Error: " + str(mean_absolute_error(predictions, y_test)))
# + id="GQoSGYSvDquD" colab_type="code" outputId="3e7505e0-282b-44bf-962a-a65512d95db1" colab={"base_uri": "https://localhost:8080/", "height": 156}
model_full=XGBRegressor()
model_full.fit(x1,y1)
# + id="zCqqNsTVDrpM" colab_type="code" colab={}
x_test.drop(columns='week_start_date',inplace=True)
z=x_test[['year','weekofyear']]
x_test.drop(columns='year',inplace=True)
x_test.drop(columns='weekofyear',inplace=True)
x_test=pd.concat([x_test,z],axis=1)
x_test['city1']=np.where(x_test['city']=='sj',1,-1)
x_test.drop(columns='city',inplace=True)
# + id="xTYp2G7CDsB5" colab_type="code" outputId="c3404e60-5da3-41ed-b046-4aa317b8a93a" colab={"base_uri": "https://localhost:8080/", "height": 226}
x_test.head()
# + id="_ge4dhPgDsuu" colab_type="code" outputId="2584f073-effa-49de-d572-a2a59f19f9f7" colab={"base_uri": "https://localhost:8080/", "height": 503}
x_test.info()
# + id="TOXZuVmjDtFi" colab_type="code" colab={}
test_pred=model_full.predict(x_test).astype(int)
# + id="AuLDuws5LXhP" colab_type="code" colab={}
submission=pd.read_csv('https://s3.amazonaws.com/drivendata/data/44/public/submission_format.csv',index_col=[0,1,2])
submission.total_cases=test_pred
submission.to_csv('values.csv')
# + id="lywxJqJxs5zP" colab_type="code" colab={}
features=['ndvi_ne','year','weekofyear','ndvi_se','ndvi_nw','reanalysis_air_temp_k','precipitation_amt_mm','ndvi_sw','reanalysis_dew_point_temp_k','reanalysis_precip_amt_kg_per_m2','station_precip_mm','station_avg_temp_c','station_diur_temp_rng_c']
# + id="BphbHDKa8EXq" colab_type="code" outputId="51df47f6-0954-4386-8e13-79312fcec995" colab={"base_uri": "https://localhost:8080/", "height": 295}
from xgboost import plot_importance
plot_importance(my_model)
plt.show()
# + id="B0aNWZDK-Ofp" colab_type="code" colab={}
features=['year','weekofyear','ndvi_se','station_avg_temp_c','ndvi_sw','reanalysis_dew_point_temp_k','reanalysis_air_temp_k','ndvi_ne','reanalysis_max_air_temp_k','reanalysis_min_air_temp_k','ndvi_nw','reanalysis_tdtr_k','precipitation_amt_mm','reanalysis_precip_amt_kg_per_m2','station_precip_mm']
# + id="SgXqDAviuMC1" colab_type="code" outputId="57656a07-6745-40ac-d5aa-0061362bd21e" colab={"base_uri": "https://localhost:8080/", "height": 330}
X=x[features]
X.info()
# + id="lyEs--Guvqs1" colab_type="code" colab={}
features=['city','year','weekofyear','ndvi_se','station_avg_temp_c','ndvi_sw','reanalysis_dew_point_temp_k','reanalysis_air_temp_k','ndvi_ne','reanalysis_max_air_temp_k','reanalysis_min_air_temp_k','ndvi_nw','reanalysis_tdtr_k','precipitation_amt_mm','reanalysis_precip_amt_kg_per_m2','station_precip_mm']
| my_ml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EJERCICIOS
# ## CAPITULO 1
#
# El capitulo 1 del Manual está diseñado para dar una leve introducción a python. Es posible que al comienzo no sea tan intuitivo seguir la lógica de programar, pero lo importante es aprender y mejorar. En esta carpeta hay un script de prueba en python llamado **prueba_python.py** con un modelo muy básico de como deberia estar estructurado y funcionar un script.
#
# Recuerde que para *correr* un script de python en **un computador con Python instalado** basta con ejecutarlo desde la consola como:
# ```
# python prueba_python.py
# ```
# o si se quiere correr con la ultima version de python exclusivamente :
# ```
# python3 prueba_python.py
# ```
# **RECUERDE** que esto solo es valido si la consola está abierta en la misma carpeta que el script.
#
# En realidad, basta con usar el **path absoluto** del script para correrlo desde cualquier ubicación en consola:
#
# ```
# python C:\user\santaclaus\Documents\python\prueba_python.py
# ```
# #### *EJERCICIO 0*
#
# Ejecutar el script **prueba_python.py** en su computador. Una vez hecho esto, ya puede seguir con los demás ejercicios.
#
#
#
# **DISCLAIMER:** En varios ejercicios se solicita *"Escribir un script"*. NO tiene que escribir un script para cada numeral de cada clase. Con un solo script por clase no se acaba el mundo.
# ### CLASE 1 - VARIABLES
#
#
# * **A)** Escriba un script que imprima en consola los numeros del 1 al 10.
# * **B)** Escriba un script que imprima en consola un diccionario con los siguientes precios:
#
# | Televisor | Precio USD |
# |------------|------------|
# | Kalley | 524 |
# | Samsung | 760 |
# | LG | 650 |
# | Panasonic | 600 |
# | Challenger | 489 |
# | Sony | 836 |
# | Vaio | 744 |
# | Daewoo | 546 |
#
# * **C)** Escriba un script que imprima en consola el valor de la función $\sin (4x) \cos(x) + \left( \frac{x}{5} \right) ^2 +5 $ para **cien puntos** en el intervalo [ -4$\pi$ , 4$\pi$ ]. También debe imprimir por aparte solo los valores que sean inferiores a 7, imprimiendo antes *"Estos valores son menores a 7"*.
#
# * **D)** Escriba un script que imprima en consola una [matriz mágica](https://es.wikipedia.org/wiki/Cuadrado_m%C3%A1gico) de 4x4 . Cabe resaltar que no debe hacer ningùn cálculo, solo escribirla. También debe imprimir en consola la matriz transpuesta.
# * **E)** En la clase se vio cómo crear un numero complejo en python. Escriba una matriz de 3x3 con valores complejos y reales, tanto decimales como enteros. Imprima dicha matriz en consola , al igual que dicha matriz multiplicada por otro numero complejo de su elección también. ¿Obtiene lo esperado? Recuerde que la forma general de los numeros complejos es $a + ib$ donde **a** y __b__ son numeros reales. i es la unidad compleja tal que $i^2 = -1$. Así:
#
# $(a+ib) \cdot (c+id) = (ac-bd)+i\cdot(ad+bc)$
# ### CLASE 2 - LOOPS
#
#
# * **A)** Escriba un script que imprima en consola los numeros del 1 al 10 usando loops.
# * **B)** Escriba un script que imprima para cada numero entre 0 y 150 si se trata de un numero para o impar.
| Chapter_1/Ejercicios/.ipynb_checkpoints/Ejercicios_C1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.1 64-bit
# metadata:
# interpreter:
# hash: 63fd5069d213b44bf678585dea6b12cceca9941eaf7f819626cde1f2670de90d
# name: python3
# ---
# # Python dictionary
# - A dictionary consists of a collection of key:value pairs.
#
# - Each key:value pair maps the key to its associated value.
#
# - A dictionary is a collection which is ordered, changeable and does not allow duplicates.
# empty dictionary
d = {}
type(d)
d2 = {'name':'John', 'last_name':'Doe', 'age':30}
d2
# accesing items
student_name = d2['name']
student_name
# Other way: using get
student_name = d2.get('name')
student_name
# Change value
d2['age'] = 33
d2
# check if key exists
'name' in d2
'middle_name' in d2
# Remember: dictionaries are accessed by key, not by the position of the items.
#
# It doesn’t make sense to slice a dictionary.
d2['name':'last_name'] # This will raise an error
# ## Python methods for working with dictionaries
# len: lenght of a dictionary
len(d2)
# Adding items. It is done by using a new key and assigning a value to it.
d2['weight'] = 65
d2
# update(): Updates the dictionary with the specified key-value pairs
d2.update({'height':5.8})
d2
# items(): Returns a list containing a tuple for each key value pair
d2.items()
# keys(): Returns a list containing the dictionary's keys
d2.keys()
# values(): Returns a list of all the values in the dictionary
d2.values()
# pop(): removes the item with specified key name
d2.pop('weight')
d2
# popitem(): Removes the last inserted key-value pair
d2.popitem()
# You cannot copy a dictionary simply by typing dict2 = dict1, because: dict2 will only be a reference to dict1, and changes made in dict1 will automatically also be made in dict2.
#
# If you want to copy the dict (which is rare), you have to do so explicitly with one of these two options:
d3 = dict(d2)
d3
# copy: make a copy of a dictionary
d3 = d2.copy()
d3
# clear: empties the dictionary
d3.clear()
d3
d2
# del: removes the item with the specified key name
del d2['name']
d2
# del can also delete the dictionary completely
del d2
d2 # This will raise an error
# Nested Dictionaries
child1 = {
'name':'Hazel',
'year': 2001,
'gender':'F'
}
child2 = {
'name':'Helen',
'year': 2003,
'gender':'F'
}
child3 = {
'name':'Abel',
'year': 2006,
'gender':'M'
}
child4 = {
'name':'Diana',
'year': 2012,
'gender':'F'
}
child1
family = {
'child1':child1,
'child2':child2,
'child3':child3,
'child4':child4
}
family
| 01-Basic Python/05-Dictionaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/abhishek-parashar/Reddit-flair-detection/blob/master/scripts/model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="exhielZubZ9v" colab_type="text"
# ## Importing the Required libraries
# + id="gXBPJXyVPTSn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="edd60ba2-41d3-4648-a418-af522928c77c"
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.externals import joblib
from xgboost import XGBClassifier
import pickle
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="40GM5UYLcZ3B" colab_type="text"
# ##loading the data
# + id="efumd_cycPBc" colab_type="code" colab={}
data=pd.read_csv('datafinal.csv')
# + id="M4zaAejIkttw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 602} outputId="ffcabbe5-3e47-4ac0-cb78-e875e660614b"
data.head()
# + id="oz1F_coOcV0X" colab_type="code" colab={}
data.fillna("",inplace = True)
# + id="zpEaY8P6xCjH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="87239db9-38a9-40b2-cb2d-f8243519725f"
data.shape
# + id="3kHtrl0-lHaX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="09fccfcc-95fa-47d8-afff-9af65fb8cbce"
data['flair'].unique()
# + id="Gub4fbYHk_ME" colab_type="code" colab={}
flair = ["AskIndia", "Non-Political", "[R]eddiquette",
"Scheduled", "Photography", "Science/Technology",
"Politics", "Business/Finance", "Policy/Economy",
"Sports", "Food", "AMA","Coronavirus"]
# + [markdown] id="AhlqsfLHd3cJ" colab_type="text"
# ## Trying different models
# ## though there are different models but cince this is a cassification task so I am using some of the models which work with classification and thn comparing their results
# + [markdown] id="RHMJkbNeedSw" colab_type="text"
# svm
# + id="6_NTnvMEcjZM" colab_type="code" colab={}
def linear_svm(X_train, X_test, y_train, y_test):
sgd = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', SGDClassifier(loss='hinge',alpha=1e-4, random_state=16, max_iter=5, tol=None)),
])
sgd.fit(X_train, y_train)
y_pred = sgd.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=flair))
# + [markdown] id="a996ktiree0f" colab_type="text"
# random forest
# + id="8b_L6wrJeYrM" colab_type="code" colab={}
def randomforest(X_train, X_test, y_train, y_test):
ranfor = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', RandomForestClassifier(n_estimators = 1000, random_state = 42)),
])
ranfor.fit(X_train, y_train)
y_pred = ranfor.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=flair))
# + [markdown] id="AguxwAACetky" colab_type="text"
# mlp classifier
# + id="PkKkxbTWelo3" colab_type="code" colab={}
def mlpclassifier(X_train, X_test, y_train, y_test):
mlp = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MLPClassifier(hidden_layer_sizes=(30,30,30))),
])
mlp.fit(X_train, y_train)
y_pred = mlp.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=flair))
# + [markdown] id="aij7yIkCexgg" colab_type="text"
# xgboost
# + id="mI_lTZhvewue" colab_type="code" colab={}
def xgbclassifier(X_train, X_test, y_train, y_test):
xgb_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', XGBClassifier(random_state=42, seed=2,n_estimators=1000,verbosity=1,objective='multi:softmax')),
])
xgb_clf.fit(X_train, y_train)
y_pred = xgb_clf.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=flair))
# + [markdown] id="6muBjUqskyuK" colab_type="text"
# logistic regression
# + id="nKnFdvSGtas2" colab_type="code" colab={}
def logisticreg(X_train, X_test, y_train, y_test):
from sklearn.linear_model import LogisticRegression
logreg = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', LogisticRegression(n_jobs=1, C=1e5)),
])
logreg.fit(X_train, y_train)
y_pred = logreg.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred, y_test))
print(classification_report(y_test, y_pred,target_names=flair))
# + [markdown] id="J2lPaWWOjue1" colab_type="text"
# ## Let's Evaluate the model
# + id="8AQjx5Bxi53r" colab_type="code" colab={}
def train_test(X,y):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state = 42)
print("Results of Linear Support Vector Machine")
linear_svm(X_train, X_test, y_train, y_test)
print("Results of Logistic Regression")
logisticreg(X_train, X_test, y_train, y_test)
print("Results of Random Forest")
randomforest(X_train, X_test, y_train, y_test)
print("Results of MLP Classifier")
mlpclassifier(X_train, X_test, y_train, y_test)
print("Results of XGB Classifier")
xgbclassifier(X_train, X_test, y_train, y_test)
# + [markdown] id="_JXGVd7cj7ez" colab_type="text"
# for evaluation comments, title, url, body and combined features are used.
# + id="vbsHzK0gjp1P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9725bbc0-4203-4da0-db5c-fd9f19f075a5"
cat = data['flair']
V = data['combined_features']
W = data['comments']
X = data['title']
Y = data['body']
Z = data['url']
print("Flair Detection using Title as Feature")
train_test(X,cat)
print("Flair Detection using Body as Feature")
train_test(Y,cat)
print("Flair Detection using URL as Feature")
train_test(Z,cat)
print("Flair Detection using Comments as Feature")
train_test(W,cat)
print("Flair Detection using Combined Features")
train_test(V,cat)
# + [markdown] id="CNSHWwap33J6" colab_type="text"
# ## saving the model
# + [markdown] id="mrPxeYx636dH" colab_type="text"
# I have used job lib to save the model earlier I used pickle but it gave errors
# + id="Uzyhwbgbkg_T" colab_type="code" colab={}
import xgboost as xgb
X_train, X_test, y_train, y_test = train_test_split(V, cat, test_size=0.2, random_state = 42)
model = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', xgb.sklearn.XGBClassifier(random_state=42,n_estimators=1000,verbosity=1, seed=2, colsample_bytree=0.6, subsample=0.7,objective='multi:softmax')),
])
XGB = model.fit(X_train, y_train)
# + id="5-JNmzR24dss" colab_type="code" colab={}
joblib.dump(XGB, open('xgb.bin', 'wb'))
# + [markdown] id="GrDLeKvG6jcd" colab_type="text"
# ## Before deploying the model to webiste first let's check the prediction
# + [markdown] id="hvq9b-LZ6v8z" colab_type="text"
# loading the model and all the files
# + id="w3JmVLjA44Mg" colab_type="code" colab={}
model_final=joblib.load(open('xgb.bin', 'rb'))
# + id="bLEdnRnj7OmU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 339} outputId="54228b84-36d5-4834-97f2-d46d1d03b59e"
# !pip install praw
# + id="CK5blW3C7V0t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3c50a435-1bc4-4101-9bff-76b3cc700225"
import praw
import numpy as np
import pandas as pd
import re
import nltk
from nltk.corpus import stopwords
import datetime as dt
nltk.download('all')
from bs4 import BeautifulSoup
# + id="3FXrSqqaQkME" colab_type="code" colab={}
model = model_final
# + id="26BlzPmd7aRE" colab_type="code" colab={}
reddit = praw.Reddit(client_id='QPdCUgBcp4WinA', client_secret='<KEY>', user_agent='reddit-flair', username='reddit-flair', password='<PASSWORD>')
# + id="7OWVGnBO7l0U" colab_type="code" colab={}
REPLACE_BY_SPACE_RE = re.compile('[/(){}\[\]\|@,;]')
BAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')
STOPWORDS = set(stopwords.words('english'))
def text_cleaning(text):
text = BeautifulSoup(text, "lxml").text
text = text.lower()
text = REPLACE_BY_SPACE_RE.sub(' ', text)
text = BAD_SYMBOLS_RE.sub('', text)
text = ' '.join(word for word in text.split() if word not in STOPWORDS)
return text
def string(value):
return str(value)
# + id="6bbmZdDrHrRP" colab_type="code" colab={}
def prediction(url):
submission = reddit.submission(url = url)
data = {}
data["title"] =str(submission.title)
data["url"] = str(submission.url)
data["body"] = str(submission.selftext)
submission.comments.replace_more(limit=None)
comment = ''
count = 0
for top_level_comment in submission.comments:
comment = comment + ' ' + top_level_comment.body
count+=1
if(count > 10):
break
data["comment"] = str(comment)
data['title'] = text_cleaning(str(data['title']))
data['body'] = text_cleaning(str(data['body']))
data['comment'] = text_cleaning(str(data['comment']))
combined_features = data["title"] + data["comment"] + data["body"] + data["url"]
return str(model.predict([combined_features]))[2:-2]
# + id="DFUIwtCgOEKa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dc0ae2b2-49a2-4501-902a-8508d091d80e"
prediction("https://www.reddit.com/r/india/comments/d1m9ld/iran_removes_antiindia_banners_from_pak_consulate/")
# + id="Rh5R5M6iQHsQ" colab_type="code" colab={}
| scripts/model.ipynb |
#!/usr/bin/env python
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfrl-cookbook
# language: python
# name: tfrl-cookbook
# ---
# Proximal Policy Optimizatin (PPO) agent training script
# Chapter 3, TensorFlow 2 Reinforcement Learning Cookbook | <NAME>
import argparse
import os
from datetime import datetime
import gym
import numpy as np
import tensorflow as tf
from tensorflow.keras.layers import Dense, Input, Lambda
tf.keras.backend.set_floatx("float64")
parser = argparse.ArgumentParser(prog="TFRL-Cookbook-Ch3-PPO")
parser.add_argument("--env", default="Pendulum-v0")
parser.add_argument("--update-freq", type=int, default=5)
parser.add_argument("--epochs", type=int, default=3)
parser.add_argument("--actor-lr", type=float, default=0.0005)
parser.add_argument("--critic-lr", type=float, default=0.001)
parser.add_argument("--clip-ratio", type=float, default=0.1)
parser.add_argument("--gae-lambda", type=float, default=0.95)
parser.add_argument("--gamma", type=float, default=0.99)
parser.add_argument("--logdir", default="logs")
args = parser.parse_args([])
logdir = os.path.join(
args.logdir, parser.prog, args.env, datetime.now().strftime("%Y%m%d-%H%M%S")
)
print(f"Saving training logs to:{logdir}")
writer = tf.summary.create_file_writer(logdir)
class Actor:
def __init__(self, state_dim, action_dim, action_bound, std_bound):
self.state_dim = state_dim
self.action_dim = action_dim
self.action_bound = action_bound
self.std_bound = std_bound
self.model = self.nn_model()
self.opt = tf.keras.optimizers.Adam(args.actor_lr)
def nn_model(self):
state_input = Input((self.state_dim,))
dense_1 = Dense(32, activation="relu")(state_input)
dense_2 = Dense(32, activation="relu")(dense_1)
out_mu = Dense(self.action_dim, activation="tanh")(dense_2)
mu_output = Lambda(lambda x: x * self.action_bound)(out_mu)
std_output = Dense(self.action_dim, activation="softplus")(dense_2)
return tf.keras.models.Model(state_input, [mu_output, std_output])
def get_action(self, state):
state = np.reshape(state, [1, self.state_dim])
mu, std = self.model.predict(state)
action = np.random.normal(mu[0], std[0], size=self.action_dim)
action = np.clip(action, -self.action_bound, self.action_bound)
log_policy = self.log_pdf(mu, std, action)
return log_policy, action
def log_pdf(self, mu, std, action):
std = tf.clip_by_value(std, self.std_bound[0], self.std_bound[1])
var = std ** 2
log_policy_pdf = -0.5 * (action - mu) ** 2 / var - 0.5 * tf.math.log(
var * 2 * np.pi
)
return tf.reduce_sum(log_policy_pdf, 1, keepdims=True)
def compute_loss(self, log_old_policy, log_new_policy, actions, gaes):
ratio = tf.exp(log_new_policy - tf.stop_gradient(log_old_policy))
gaes = tf.stop_gradient(gaes)
clipped_ratio = tf.clip_by_value(
ratio, 1.0 - args.clip_ratio, 1.0 + args.clip_ratio
)
surrogate = -tf.minimum(ratio * gaes, clipped_ratio * gaes)
return tf.reduce_mean(surrogate)
def train(self, log_old_policy, states, actions, gaes):
with tf.GradientTape() as tape:
mu, std = self.model(states, training=True)
log_new_policy = self.log_pdf(mu, std, actions)
loss = self.compute_loss(log_old_policy, log_new_policy, actions, gaes)
grads = tape.gradient(loss, self.model.trainable_variables)
self.opt.apply_gradients(zip(grads, self.model.trainable_variables))
return loss
class Critic:
def __init__(self, state_dim):
self.state_dim = state_dim
self.model = self.nn_model()
self.opt = tf.keras.optimizers.Adam(args.critic_lr)
def nn_model(self):
return tf.keras.Sequential(
[
Input((self.state_dim,)),
Dense(32, activation="relu"),
Dense(32, activation="relu"),
Dense(16, activation="relu"),
Dense(1, activation="linear"),
]
)
def compute_loss(self, v_pred, td_targets):
mse = tf.keras.losses.MeanSquaredError()
return mse(td_targets, v_pred)
def train(self, states, td_targets):
with tf.GradientTape() as tape:
v_pred = self.model(states, training=True)
# assert v_pred.shape == td_targets.shape
loss = self.compute_loss(v_pred, tf.stop_gradient(td_targets))
grads = tape.gradient(loss, self.model.trainable_variables)
self.opt.apply_gradients(zip(grads, self.model.trainable_variables))
return loss
class Agent:
def __init__(self, env):
self.env = env
self.state_dim = self.env.observation_space.shape[0]
self.action_dim = self.env.action_space.shape[0]
self.action_bound = self.env.action_space.high[0]
self.std_bound = [1e-2, 1.0]
self.actor_opt = tf.keras.optimizers.Adam(args.actor_lr)
self.critic_opt = tf.keras.optimizers.Adam(args.critic_lr)
self.actor = Actor(
self.state_dim, self.action_dim, self.action_bound, self.std_bound
)
self.critic = Critic(self.state_dim)
def gae_target(self, rewards, v_values, next_v_value, done):
n_step_targets = np.zeros_like(rewards)
gae = np.zeros_like(rewards)
gae_cumulative = 0
forward_val = 0
if not done:
forward_val = next_v_value
for k in reversed(range(0, len(rewards))):
delta = rewards[k] + args.gamma * forward_val - v_values[k]
gae_cumulative = args.gamma * args.gae_lambda * gae_cumulative + delta
gae[k] = gae_cumulative
forward_val = v_values[k]
n_step_targets[k] = gae[k] + v_values[k]
return gae, n_step_targets
def train(self, max_episodes=1000):
with writer.as_default():
for ep in range(max_episodes):
state_batch = []
action_batch = []
reward_batch = []
old_policy_batch = []
episode_reward, done = 0, False
state = self.env.reset()
while not done:
# self.env.render()
log_old_policy, action = self.actor.get_action(state)
next_state, reward, done, _ = self.env.step(action)
state = np.reshape(state, [1, self.state_dim])
action = np.reshape(action, [1, 1])
next_state = np.reshape(next_state, [1, self.state_dim])
reward = np.reshape(reward, [1, 1])
log_old_policy = np.reshape(log_old_policy, [1, 1])
state_batch.append(state)
action_batch.append(action)
reward_batch.append((reward + 8) / 8)
old_policy_batch.append(log_old_policy)
if len(state_batch) >= args.update_freq or done:
states = np.array([state.squeeze() for state in state_batch])
actions = np.array(
[action.squeeze() for action in action_batch]
)
rewards = np.array(
[reward.squeeze() for reward in reward_batch]
)
old_policies = np.array(
[old_pi.squeeze() for old_pi in old_policy_batch]
)
v_values = self.critic.model.predict(states)
next_v_value = self.critic.model.predict(next_state)
gaes, td_targets = self.gae_target(
rewards, v_values, next_v_value, done
)
actor_losses, critic_losses = [], []
for epoch in range(args.epochs):
actor_loss = self.actor.train(
old_policies, states, actions, gaes
)
actor_losses.append(actor_loss)
critic_loss = self.critic.train(states, td_targets)
critic_losses.append(critic_loss)
# Plot mean actor & critic losses on every update
tf.summary.scalar("actor_loss", np.mean(actor_losses), step=ep)
tf.summary.scalar(
"critic_loss", np.mean(critic_losses), step=ep
)
state_batch = []
action_batch = []
reward_batch = []
old_policy_batch = []
episode_reward += reward[0][0]
state = next_state[0]
print(f"Episode#{ep} Reward:{episode_reward}")
tf.summary.scalar("episode_reward", episode_reward, step=ep)
if __name__ == "__main__":
env_name = "Pendulum-v0"
env = gym.make(env_name)
agent = Agent(env)
agent.train(max_episodes=2) # Increase max_episodes value
| Chapter03/6_ppo_continuous.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/python4phys1cs/physics-problems/blob/main/calculating-bandgap/calculating-bandgap.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bSjrD_Qhn7GU"
# # Calculating the bangap of material from transmittance data
# + [markdown] id="uuNTc5qUn2Dr"
# Importing necessary libraries and modules
# + id="O9HRq-T3-_Lc"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.signal import savgol_filter
from sklearn.metrics import mean_squared_error
# + [markdown] id="bWVKQN2vF3tF"
# Reading Transmittance data to a data frame using read_csv module
# + id="Awq-39NLnvO1"
data = "https://raw.githubusercontent.com/python4phys1cs/physics-problems/main/calculating-bandgap/data/znse-data.csv"
df = pd.read_csv(data, sep=",")
# + [markdown] id="lsPAl1C3F_VV"
# Converting the Transmittance values from percentage to absolute values
# $$ T= \frac {T\%}{100}$$
# + id="L_hQD1RpFvfS"
#T = T%/100
T = df["Transmittance"]/100
# + [markdown] id="X5NIogqSUQ7z"
# Converting Wavelength to meters\
# **Conversion factor:** $ 1\ m = 10^{-9}\ nm $
# + id="JG-um34CUUu5"
wavelength = df['Wavelength']*1e-9
# + [markdown] id="NPXCIYxyGQXr"
# Calculating the energy associated with wavelength of light
# + id="dontTqybFzE1"
h = 6.626e-34 #planck's constant
c = 3e8 #velocity of light
eV = 1.602e-19 # 1 electron-volt
Eg = h*c/(wavelength*eV)
# + [markdown] id="b76-h-7fHBF5"
# Calculating molar absorption coefficient $ \alpha $
#
# $$ \alpha = - \frac{ln(T)}{t} $$
# where $ t $ is the thickness of sample
# + id="siLwazbiHX6k"
t = 2e-4 #thickness of sample in meter
alpha = - np.log(T)/t
# + [markdown] id="uDAeOaUxJtb7"
# Calculating $ (\alpha h \nu ) ^ {1/2} $
#
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="Ud3rPchmKQss" outputId="0b21ba48-847f-45e2-efbf-3efcf1b2f05a"
#setting power for direct or indirect semiconductor
n=2
#evaluating the values for Tauc Plot
TP = (alpha*Eg)**n
plt.plot(Eg,TP)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 275} id="CnGQkWF4WWkJ" outputId="9df9f10c-fb73-4bff-f82e-2a83577f2d9d"
#smoothening the data using Savitzky-Golay Filter
sg = savgol_filter(TP, 9, 4)
plt.plot(Eg, sg)
plt.show()
# + id="XpUcLFLjkUu8"
#calculating the maximum value of Tauc plot for rescaling
sgmax = max(sg)
#rescaling the Tauc plot
sgre = sg/sgmax
# + colab={"base_uri": "https://localhost:8080/"} id="HVXP0cOwWxKb" outputId="d7658658-a98a-4e33-f817-073a0bfcd18c"
#initiating arrays to store values
sgpred = []
slope = []
intercept = []
for i in range(len(Eg)):
#calculating slope and intercept of line for every 10 points
m, b = np.polyfit(Eg[i:i+10],sgre[i:i+10],1)
slope.append(m)
intercept.append(b)
#calculating the predicted value from the line parameters
sgpred.append(m*Eg[i]+b)
# + id="b7WJ1LPOcUkP"
rmse = []
#calculating the root mean square error between predicted and actual values
for i in range(len(sg)):
mse = mean_squared_error(sgre, sgpred)
rmse.append(mse**0.5)
# + id="M0S4SdaBdGjF"
selseg = []
#selecting only those segments for which rmse<0.75
for i in range(len(slope)):
if(rmse[i]<0.75):
selseg.append(slope[i])
else:
selseg.append(0)
# + id="SIE4S8lteoDj"
#finding the maximum slope within the selected segments
max_slope = max(selseg)
#find the index for which slope is maximum
max_slope_index = selseg.index(max_slope)
# + colab={"base_uri": "https://localhost:8080/"} id="VDz_hnAegGwL" outputId="0cb1c4df-cf32-4cdb-e532-ef5f13d95208"
#calculating the bandgap of material
bg = (max_slope*Eg[max_slope_index]-sgre[max_slope_index])/max_slope
print("The band gap of material is:", bg)
| calculating-bandgap/calculating-bandgap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="ocn-alaCjZBc" colab_type="text"
# ## Part I: Implement and train on Cifar10 dataset a simple baseline for image classification using standard 2D convolutions given the structure below
# - conv output channels 64, 64, 128, 128, 128, 256, 256, 256, 512, 512, 512
# - kernel shape (3,3)
# - strides: 1, 1, 2, 1, 1, 2, 1, 1, 2, 1, 1
# - padding: SAME (snt.SAME)
# - num_output_classes = 10
# - After each conv layer, add BatchNorm and ReLU.
# - Use `tf.reduce_mean` to pool spatially the activations at the end -- this way the network can run on inputs of any size.
# - Project final activations into label space using `snt.Linear`.
#
#
#
# #### Exercises:
#
# 1. Fill in the code for the Sonnet module which defines the network, the predictions ops and the loss function ops
#
# 2. Train the network and see the loss going down. Pay attention to the data augmentation and learning schedule.
#
# 3. Understand how BatchNorm works:
#
#
# > * remove the update operations and see if the model performs well;
# > * keep the update ops, but use `test_local_stats=True`. This will work, but the accuracy will be lower than when using the (training) moving averages.
#
#
#
#
# + [markdown] id="FhWI4Pix5GJw" colab_type="text"
# ### Imports
# + id="na0VvPXmYKp1" colab_type="code" colab={}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import math
import time
import tensorflow as tf
# Don't forget to select GPU runtime environment in Runtime -> Change runtime type
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# we will use Sonnet on top of TF
# !pip install -q dm-sonnet
import sonnet as snt
import numpy as np
# Plotting library.
from matplotlib import pyplot as plt
import pylab as pl
from IPython import display
# + id="1xlKHOLbhvY7" colab_type="code" colab={}
# Reset graph
tf.reset_default_graph()
# + [markdown] id="8g16XweXs2Uq" colab_type="text"
# ### Download dataset to be used for training and testing
# - Cifar-10 equivalent of MNIST for natural RGB images
# - 60000 32x32 colour images in 10 classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck
# - train: 50000; test: 10000
# + id="1g_EOx07s1XZ" colab_type="code" colab={}
cifar10 = tf.keras.datasets.cifar10
# (down)load dataset
(train_images, train_labels), (test_images, test_labels) = cifar10.load_data()
# + id="rt6hU4aQtwpZ" colab_type="code" colab={}
# Check sizes of tensors
print ('Size of training images')
print (train_images.shape)
print ('Size of training labels')
print (train_labels.shape)
print ('Size of test images')
print (test_images.shape)
print ('Size of test labels')
print (test_labels.shape)
assert train_images.shape[0] == train_labels.shape[0]
# + [markdown] id="ME1oz3sJkAKX" colab_type="text"
# ### Display the images
#
# The gallery function below shows sample images from the data, together with their labels.
# + id="eO1xIgRtjvXU" colab_type="code" colab={}
MAX_IMAGES = 10
def gallery(images, label, title='Input images'):
class_dict = [u'airplane', u'automobile', u'bird', u'cat', u'deer', u'dog', u'frog', u'horse', u'ship', u'truck']
num_frames, h, w, num_channels = images.shape
num_frames = min(num_frames, MAX_IMAGES)
ff, axes = plt.subplots(1, num_frames,
figsize=(num_frames, 1),
subplot_kw={'xticks': [], 'yticks': []})
for i in range(0, num_frames):
if num_channels == 3:
axes[i].imshow(np.squeeze(images[i]))
else:
axes[i].imshow(np.squeeze(images[i]), cmap='gray')
axes[i].set_title(class_dict[label[i][0]])
plt.setp(axes[i].get_xticklabels(), visible=False)
plt.setp(axes[i].get_yticklabels(), visible=False)
ff.subplots_adjust(wspace=0.1)
plt.show()
# + id="dN5Kq_xweBhe" colab_type="code" colab={}
gallery(train_images, train_labels)
# + [markdown] id="JHAggitWu94_" colab_type="text"
# ### Prepare the data for training and testing
# - for training, we use stochastic optimizers (e.g. SGD, Adam), so we need to sample at random mini-batches from the training dataset
# - for testing, we iterate sequentially through the test set
# + id="iZofMjOuUEOF" colab_type="code" colab={}
# define dimension of the batches to sample from the datasets
BATCH_SIZE_TRAIN = 32 #@param
BATCH_SIZE_TEST = 100 #@param
# create Dataset objects using the data previously downloaded
dataset_train = tf.data.Dataset.from_tensor_slices((train_images, train_labels))
# we shuffle the data and sample repeatedly batches for training
batched_dataset_train = dataset_train.shuffle(100000).repeat().batch(BATCH_SIZE_TRAIN)
# create iterator to retrieve batches
iterator_train = batched_dataset_train.make_one_shot_iterator()
# get a training batch of images and labels
(batch_train_images, batch_train_labels) = iterator_train.get_next()
# check that the shape of the training batches is the expected one
print ('Shape of training images')
print (batch_train_images)
print ('Shape of training labels')
print (batch_train_labels)
# + id="yWtdQ0ESxkBQ" colab_type="code" colab={}
# we do the same for test dataset
dataset_test = tf.data.Dataset.from_tensor_slices((test_images, test_labels))
batched_dataset_test = dataset_test.repeat().batch(BATCH_SIZE_TEST)
iterator_test = batched_dataset_test.make_one_shot_iterator()
(batch_test_images, batch_test_labels) = iterator_test.get_next()
print ('Shape of test images')
print (batch_test_images)
print ('Shape of test labels')
print (batch_test_labels)
# + id="P4iKpKNB-c74" colab_type="code" colab={}
# Squeeze labels and convert from uint8 to int32 - required below by the loss op
batch_test_labels = tf.cast(tf.squeeze(batch_test_labels), tf.int32)
batch_train_labels = tf.cast(tf.squeeze(batch_train_labels), tf.int32)
# + [markdown] id="i5rWCfPp-50y" colab_type="text"
# ### # Preprocess input for training and testing
# + id="_PS2GjTxRZx9" colab_type="code" colab={}
# Data augmentation used for train preprocessing
# - scale image to [-1 , 1]
# - get a random crop
# - apply horizontal flip randomly
def train_image_preprocess(h, w, random_flip=True):
"""Image processing required for training the model."""
def random_flip_left_right(image, flip_index, seed=None):
shape = image.get_shape()
if shape.ndims == 3 or shape.ndims is None:
uniform_random = tf.random_uniform([], 0, 1.0, seed=seed)
mirror_cond = tf.less(uniform_random, .5)
result = tf.cond(
mirror_cond,
lambda: tf.reverse(image, [flip_index]),
lambda: image
)
return result
elif shape.ndims == 4:
uniform_random = tf.random_uniform(
[tf.shape(image)[0]], 0, 1.0, seed=seed
)
mirror_cond = tf.less(uniform_random, .5)
return tf.where(
mirror_cond,
image,
tf.map_fn(lambda x: tf.reverse(x, [flip_index]), image, dtype=image.dtype)
)
else:
raise ValueError("\'image\' must have either 3 or 4 dimensions.")
def fn(image):
# Ensure the data is in range [-1, 1].
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
image = image * 2.0 - 1.0
# Randomly choose a (h, w, 3) patch.
image = tf.random_crop(image, size=(BATCH_SIZE_TRAIN, h, w, 3))
# Randomly flip the image.
image = random_flip_left_right(image, 2)
return image
return fn
# Test preprocessing: only scale to [-1,1].
def test_image_preprocess():
def fn(image):
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
image = image * 2.0 - 1.0
return image
return fn
# + [markdown] id="vTsLFmVCF585" colab_type="text"
# ### Define the network
#
#
#
# + id="2scBoc09ZsO4" colab_type="code" colab={}
class Baseline(snt.AbstractModule):
def __init__(self, num_classes, name="baseline"):
super(Baseline, self).__init__(name=name)
self._num_classes = num_classes
self._output_channels = [
64, 64, 128, 128, 128, 256, 256, 256, 512, 512, 512
]
#############
# #
# YOUR CODE #
# #
#############
def _build(self, inputs, is_training=None, test_local_stats=False):
#############
# #
# YOUR CODE #
# #
#############
return logits
# + id="OD8IR90m_0-r" colab_type="code" colab={}
# Get number of parameters in a scope by iterating through the trainable variables
def get_num_params(scope):
total_parameters = 0
for variable in tf.trainable_variables(scope):
# shape is an array of tf.Dimension
shape = variable.get_shape()
variable_parameters = 1
for dim in shape:
variable_parameters *= dim.value
total_parameters += variable_parameters
return total_parameters
# + [markdown] id="inX9OlHW5xWR" colab_type="text"
# ### Instantiate the model and connect to data
#
# + id="TZzlpO0oJFZy" colab_type="code" colab={}
# First define the preprocessing ops for the train/test data
crop_height = 24 #@param
cropt_width = 24 #@param
preprocess_fn_train = train_image_preprocess(crop_height, cropt_width)
preprocess_fn_test = test_image_preprocess()
num_classes = 10 #@param
# + id="WZlB8Ao5CZJn" colab_type="code" colab={}
# Instantiate the model
with tf.variable_scope("baseline"):
baseline_model = Baseline(num_classes)
# + id="rt87AMfB6VKt" colab_type="code" colab={}
# Get predictions from the model; use the corresponding preprocess ops and is_training flag
train_predictions = ############## YOUR CODE ##############
print (train_predictions)
test_predictions = ############## YOUR CODE ##############
print (test_predictions)
# + id="_JUYuIR3Bv8P" colab_type="code" colab={}
# Get number of parameters in the model. Can you obtain this number by hand?
print ("Total number of parameters of baseline model")
print (get_num_params("baseline"))
# + [markdown] id="MpxLmb3sJa0o" colab_type="text"
# ### Define the loss to be minimized during training
# + id="9juCwywKqe5X" colab_type="code" colab={}
def get_loss(logits=None, labels=None):
# We reduce over batch dimension, to ensure the loss is a scalar.
return tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
labels=labels, logits=logits))
# + id="LQBnrZPb7XcI" colab_type="code" colab={}
# Define train and test loss ops
train_loss = ############## YOUR CODE ##############
test_loss = ############## YOUR CODE ##############
# + id="5K4VMXej8Fem" colab_type="code" colab={}
# for evaluation, we look at top_k_accuracy since it's easier to interpret; normally k=1 or k=5
def top_k_accuracy(k, labels, logits):
in_top_k = tf.nn.in_top_k(predictions=tf.squeeze(logits), targets=labels, k=k)
return tf.reduce_mean(tf.cast(in_top_k, tf.float32))
# + [markdown] id="lGyLJwJ408ZZ" colab_type="text"
# ### Create the optimizer
#
# We will use the Momentum optimizer, but other optimizers such as Adam or AdaGrad can be used.
# + id="f8V7fy_U2yY2" colab_type="code" colab={}
def get_optimizer(step):
"""Get the optimizer used for training."""
lr_init = 0.1 # initial value for the learning rate
lr_schedule = (40e3, 60e3, 80e3) # after how many iterations to reduce the learning rate
lr_schedule = tf.to_int64(lr_schedule)
lr_factor = 0.1 # reduce learning rate by this factor
num_epochs = tf.reduce_sum(tf.to_float(step >= lr_schedule))
lr = lr_init * lr_factor**num_epochs
return tf.train.MomentumOptimizer(learning_rate=lr, momentum=0.9)
# + [markdown] id="PTFLYiWv8Z_n" colab_type="text"
# ### Set up the training
# + id="i5tjjaENpgmj" colab_type="code" colab={}
# Create a global step that is incremented during training; useful for e.g. learning rate annealing
global_step = tf.train.get_or_create_global_step()
# instantiate the optimizer
optimizer = get_optimizer(global_step)
# + [markdown] id="Mvw-WbFDeTl0" colab_type="text"
# ### BatchNorm ops
#
# Batch normalization requires updating the moving averages during training, so they can be used during testing instead of the statistics of the test batch. However, there is no direct dependency between the moving averages and the train ops. Hence running only the train ops will not update the moving averages.
#
# The Sonnet BatchNorm module ensures that the moving average updates are added to the global UPDATE_OPS collections. So all we need to do is to group the train ops with the update ops.
#
# To find out more about collections: https://www.tensorflow.org/api_guides/python/framework#Graph_collections
#
# + id="nTAm5Nfsc2Ug" colab_type="code" colab={}
# Get training ops
training_baseline_op = optimizer.minimize(train_loss, global_step)
# Retrieve the update ops, which contain the moving average ops
update_ops = tf.group(*tf.get_collection(tf.GraphKeys.UPDATE_OPS))
# Manually add the update ops to the dependency path executed at each training iteration
training_baseline_op = tf.group(training_baseline_op, update_ops)
# For exercise 3, comment the line above and see if the model performs well.
# + id="7aGft4Zhppdz" colab_type="code" colab={}
# Get test ops
test_acc_baseline_op = top_k_accuracy(1, batch_test_labels, test_predictions)
# + id="pF0oEXrHFB7W" colab_type="code" colab={}
# Function that takes a list of losses and plots them.
def plot_losses(loss_list, steps):
display.clear_output(wait=True)
display.display(pl.gcf())
pl.plot(steps, loss_list, c='b')
time.sleep(1.0)
# + [markdown] id="x2unavvlpBFn" colab_type="text"
# ### Training params
#
#
# + id="s-m8-e5vpUIQ" colab_type="code" colab={}
# Define number of training iterations and reporting intervals
TRAIN_ITERS = 90e3 #@param
REPORT_TRAIN_EVERY = 10 #@param
PLOT_EVERY = 500 #@param
REPORT_TEST_EVERY = 1000 #@param
TEST_ITERS = 10 #@param
# + [markdown] id="Os8cSIn7xZiJ" colab_type="text"
# ### Train the model (you can stop the training once you observe the loss going down and the test accuracy going up). Running the full training gives around 94% accuracy on the test set.
# + id="elUiAs1S1gkv" colab_type="code" colab={}
# Create the session and initialize variables
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# Question: What is the accuracy of the model at iteration 0, i.e. before training starts?
train_iter = 0
losses = []
steps = []
for train_iter in range(int(TRAIN_ITERS)):
_, train_loss_np = sess.run([training_baseline_op, train_loss])
if (train_iter % REPORT_TRAIN_EVERY) == 0:
losses.append(train_loss_np)
steps.append(train_iter)
if (train_iter % PLOT_EVERY) == 0:
plot_losses(losses, steps)
if (train_iter % REPORT_TEST_EVERY) == 0:
avg_acc = 0.0
for test_iter in range(TEST_ITERS):
acc = sess.run(test_acc_baseline_op)
avg_acc += acc
avg_acc /= (TEST_ITERS)
print ('Test acc at iter {0:5d} out of {1:5d} is {2:.2f}%'.format(int(train_iter), int(TRAIN_ITERS), avg_acc*100.0))
| vision/baseline_start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Свёрточные нейросети и POS-теггинг
#
# POS-теггинг - определение частей речи (снятие частеречной неоднозначности)
# +
# Если Вы запускаете ноутбук на colab,
# выполните следующие строчки, чтобы подгрузить библиотеку dlnlputils:
# # !git clone https://github.com/Samsung-IT-Academy/stepik-dl-nlp.git
# import sys; sys.path.append('/content/stepik-dl-nlp')
# +
# # !pip install pyconll
# +
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
from sklearn.metrics import classification_report
import numpy as np
import pyconll
import torch
from torch import nn
from torch.nn import functional as F
from torch.utils.data import TensorDataset
import dlnlputils
from dlnlputils.data import tokenize_corpus, build_vocabulary, \
character_tokenize, pos_corpus_to_tensor, POSTagger
from dlnlputils.pipeline import train_eval_loop, predict_with_model, init_random_seed
init_random_seed()
# -
# ## Загрузка текстов и разбиение на обучающую и тестовую подвыборки
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
# !wget -O ./datasets/ru_syntagrus-ud-train.conllu https://raw.githubusercontent.com/UniversalDependencies/UD_Russian-SynTagRus/master/ru_syntagrus-ud-train.conllu
# !wget -O ./datasets/ru_syntagrus-ud-dev.conllu https://raw.githubusercontent.com/UniversalDependencies/UD_Russian-SynTagRus/master/ru_syntagrus-ud-dev.conllu
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
full_train = pyconll.load_from_file('./datasets/ru_syntagrus-ud-train.conllu')
full_test = pyconll.load_from_file('./datasets/ru_syntagrus-ud-dev.conllu')
for sent in full_train[:2]:
for token in sent:
print(token.form, token.upos)
print()
MAX_SENT_LEN = max(len(sent) for sent in full_train)
MAX_ORIG_TOKEN_LEN = max(len(token.form) for sent in full_train for token in sent)
print('Наибольшая длина предложения', MAX_SENT_LEN)
print('Наибольшая длина токена', MAX_ORIG_TOKEN_LEN)
all_train_texts = [' '.join(token.form for token in sent) for sent in full_train]
print('\n'.join(all_train_texts[:10]))
train_char_tokenized = tokenize_corpus(all_train_texts, tokenizer=character_tokenize)
char_vocab, word_doc_freq = build_vocabulary(train_char_tokenized, max_doc_freq=1.0, min_count=5, pad_word='<PAD>')
print("Количество уникальных символов", len(char_vocab))
print(list(char_vocab.items())[:10])
UNIQUE_TAGS = ['<NOTAG>'] + sorted({token.upos for sent in full_train for token in sent if token.upos})
label2id = {label: i for i, label in enumerate(UNIQUE_TAGS)}
label2id
# +
train_inputs, train_labels = pos_corpus_to_tensor(full_train, char_vocab, label2id, MAX_SENT_LEN, MAX_ORIG_TOKEN_LEN)
train_dataset = TensorDataset(train_inputs, train_labels)
test_inputs, test_labels = pos_corpus_to_tensor(full_test, char_vocab, label2id, MAX_SENT_LEN, MAX_ORIG_TOKEN_LEN)
test_dataset = TensorDataset(test_inputs, test_labels)
# -
train_inputs[1][:5]
train_labels[1]
# ## Вспомогательная свёрточная архитектура
class StackedConv1d(nn.Module):
def __init__(self, features_num, layers_n=1, kernel_size=3, conv_layer=nn.Conv1d, dropout=0.0):
super().__init__()
layers = []
for _ in range(layers_n):
layers.append(nn.Sequential(
conv_layer(features_num, features_num, kernel_size, padding=kernel_size//2),
nn.Dropout(dropout),
nn.LeakyReLU()))
self.layers = nn.ModuleList(layers)
def forward(self, x):
"""x - BatchSize x FeaturesNum x SequenceLen"""
for layer in self.layers:
x = x + layer(x)
return x
# ## Предсказание частей речи на уровне отдельных токенов
class SingleTokenPOSTagger(nn.Module):
def __init__(self, vocab_size, labels_num, embedding_size=32, **kwargs):
super().__init__()
self.char_embeddings = nn.Embedding(vocab_size, embedding_size, padding_idx=0)
self.backbone = StackedConv1d(embedding_size, **kwargs)
self.global_pooling = nn.AdaptiveMaxPool1d(1)
self.out = nn.Linear(embedding_size, labels_num)
self.labels_num = labels_num
def forward(self, tokens):
"""tokens - BatchSize x MaxSentenceLen x MaxTokenLen"""
batch_size, max_sent_len, max_token_len = tokens.shape
tokens_flat = tokens.view(batch_size * max_sent_len, max_token_len)
char_embeddings = self.char_embeddings(tokens_flat) # BatchSize*MaxSentenceLen x MaxTokenLen x EmbSize
char_embeddings = char_embeddings.permute(0, 2, 1) # BatchSize*MaxSentenceLen x EmbSize x MaxTokenLen
features = self.backbone(char_embeddings)
global_features = self.global_pooling(features).squeeze(-1) # BatchSize*MaxSentenceLen x EmbSize
logits_flat = self.out(global_features) # BatchSize*MaxSentenceLen x LabelsNum
logits = logits_flat.view(batch_size, max_sent_len, self.labels_num) # BatchSize x MaxSentenceLen x LabelsNum
logits = logits.permute(0, 2, 1) # BatchSize x LabelsNum x MaxSentenceLen
return logits
single_token_model = SingleTokenPOSTagger(len(char_vocab), len(label2id), embedding_size=64, layers_n=3, kernel_size=3, dropout=0.3)
print('Количество параметров', sum(np.product(t.shape) for t in single_token_model.parameters()))
(best_val_loss,
best_single_token_model) = train_eval_loop(single_token_model,
train_dataset,
test_dataset,
F.cross_entropy,
lr=5e-3,
epoch_n=10,
batch_size=64,
device='cuda',
early_stopping_patience=5,
max_batches_per_epoch_train=500,
max_batches_per_epoch_val=100,
lr_scheduler_ctor=lambda optim: torch.optim.lr_scheduler.ReduceLROnPlateau(optim, patience=2,
factor=0.5,
verbose=True))
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
torch.save(best_single_token_model.state_dict(), './models/single_token_pos.pth')
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
single_token_model.load_state_dict(torch.load('./models/single_token_pos.pth'))
# +
train_pred = predict_with_model(single_token_model, train_dataset)
train_loss = F.cross_entropy(torch.tensor(train_pred),
torch.tensor(train_labels))
print('Среднее значение функции потерь на обучении', float(train_loss))
print(classification_report(train_labels.view(-1), train_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
print()
test_pred = predict_with_model(single_token_model, test_dataset)
test_loss = F.cross_entropy(torch.tensor(test_pred),
torch.tensor(test_labels))
print('Среднее значение функции потерь на валидации', float(test_loss))
print(classification_report(test_labels.view(-1), test_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
# -
# ## Предсказание частей речи на уровне предложений (с учётом контекста)
class SentenceLevelPOSTagger(nn.Module):
def __init__(self, vocab_size, labels_num, embedding_size=32, single_backbone_kwargs={}, context_backbone_kwargs={}):
super().__init__()
self.embedding_size = embedding_size
self.char_embeddings = nn.Embedding(vocab_size, embedding_size, padding_idx=0)
self.single_token_backbone = StackedConv1d(embedding_size, **single_backbone_kwargs)
self.context_backbone = StackedConv1d(embedding_size, **context_backbone_kwargs)
self.global_pooling = nn.AdaptiveMaxPool1d(1)
self.out = nn.Conv1d(embedding_size, labels_num, 1)
self.labels_num = labels_num
def forward(self, tokens):
"""tokens - BatchSize x MaxSentenceLen x MaxTokenLen"""
batch_size, max_sent_len, max_token_len = tokens.shape
tokens_flat = tokens.view(batch_size * max_sent_len, max_token_len)
char_embeddings = self.char_embeddings(tokens_flat) # BatchSize*MaxSentenceLen x MaxTokenLen x EmbSize
char_embeddings = char_embeddings.permute(0, 2, 1) # BatchSize*MaxSentenceLen x EmbSize x MaxTokenLen
char_features = self.single_token_backbone(char_embeddings)
token_features_flat = self.global_pooling(char_features).squeeze(-1) # BatchSize*MaxSentenceLen x EmbSize
token_features = token_features_flat.view(batch_size, max_sent_len, self.embedding_size) # BatchSize x MaxSentenceLen x EmbSize
token_features = token_features.permute(0, 2, 1) # BatchSize x EmbSize x MaxSentenceLen
context_features = self.context_backbone(token_features) # BatchSize x EmbSize x MaxSentenceLen
logits = self.out(context_features) # BatchSize x LabelsNum x MaxSentenceLen
return logits
sentence_level_model = SentenceLevelPOSTagger(len(char_vocab), len(label2id), embedding_size=64,
single_backbone_kwargs=dict(layers_n=3, kernel_size=3, dropout=0.3),
context_backbone_kwargs=dict(layers_n=3, kernel_size=3, dropout=0.3))
print('Количество параметров', sum(np.product(t.shape) for t in sentence_level_model.parameters()))
(best_val_loss,
best_sentence_level_model) = train_eval_loop(sentence_level_model,
train_dataset,
test_dataset,
F.cross_entropy,
lr=5e-3,
epoch_n=10,
batch_size=64,
device='cuda',
early_stopping_patience=5,
max_batches_per_epoch_train=500,
max_batches_per_epoch_val=100,
lr_scheduler_ctor=lambda optim: torch.optim.lr_scheduler.ReduceLROnPlateau(optim, patience=2,
factor=0.5,
verbose=True))
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
torch.save(best_sentence_level_model.state_dict(), './models/sentence_level_pos.pth')
# Если Вы запускаете ноутбук на colab, добавьте в начало пути /content/stepik-dl-nlp
sentence_level_model.load_state_dict(torch.load('./models/sentence_level_pos.pth'))
# +
train_pred = predict_with_model(sentence_level_model, train_dataset)
train_loss = F.cross_entropy(torch.tensor(train_pred),
torch.tensor(train_labels))
print('Среднее значение функции потерь на обучении', float(train_loss))
print(classification_report(train_labels.view(-1), train_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
print()
test_pred = predict_with_model(sentence_level_model, test_dataset)
test_loss = F.cross_entropy(torch.tensor(test_pred),
torch.tensor(test_labels))
print('Среднее значение функции потерь на валидации', float(test_loss))
print(classification_report(test_labels.view(-1), test_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
# -
# ## Применение полученных теггеров и сравнение
single_token_pos_tagger = POSTagger(single_token_model, char_vocab, UNIQUE_TAGS, MAX_SENT_LEN, MAX_ORIG_TOKEN_LEN)
sentence_level_pos_tagger = POSTagger(sentence_level_model, char_vocab, UNIQUE_TAGS, MAX_SENT_LEN, MAX_ORIG_TOKEN_LEN)
test_sentences = [
'Мама мыла раму.',
'Косил косой косой косой.',
'Глокая куздра штеко будланула бокра и куздрячит бокрёнка.',
'Сяпала Калуша с Калушатами по напушке.',
'Пирожки поставлены в печь, мама любит печь.',
'Ведро дало течь, вода стала течь.',
'Три да три, будет дырка.',
'Три да три, будет шесть.',
'Сорок сорок'
]
test_sentences_tokenized = tokenize_corpus(test_sentences, min_token_size=1)
for sent_tokens, sent_tags in zip(test_sentences_tokenized, single_token_pos_tagger(test_sentences)):
print(' '.join('{}-{}'.format(tok, tag) for tok, tag in zip(sent_tokens, sent_tags)))
print()
for sent_tokens, sent_tags in zip(test_sentences_tokenized, sentence_level_pos_tagger(test_sentences)):
print(' '.join('{}-{}'.format(tok, tag) for tok, tag in zip(sent_tokens, sent_tags)))
print()
# ## Свёрточный модуль своими руками
class MyConv1d(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size, padding=0):
super().__init__()
self.in_channels = in_channels
self.out_channels = out_channels
self.kernel_size = kernel_size
self.padding = padding
self.weight = nn.Parameter(torch.randn(in_channels * kernel_size, out_channels) / (in_channels * kernel_size),
requires_grad=True)
self.bias = nn.Parameter(torch.zeros(out_channels), requires_grad=True)
def forward(self, x):
"""x - BatchSize x InChannels x SequenceLen"""
batch_size, src_channels, sequence_len = x.shape
if self.padding > 0:
pad = x.new_zeros(batch_size, src_channels, self.padding)
x = torch.cat((pad, x, pad), dim=-1)
sequence_len = x.shape[-1]
chunks = []
chunk_size = sequence_len - self.kernel_size + 1
for offset in range(self.kernel_size):
chunks.append(x[:, :, offset:offset + chunk_size])
in_features = torch.cat(chunks, dim=1) # BatchSize x InChannels * KernelSize x ChunkSize
in_features = in_features.permute(0, 2, 1) # BatchSize x ChunkSize x InChannels * KernelSize
out_features = torch.bmm(in_features, self.weight.unsqueeze(0).expand(batch_size, -1, -1)) + self.bias.unsqueeze(0).unsqueeze(0)
out_features = out_features.permute(0, 2, 1) # BatchSize x OutChannels x ChunkSize
return out_features
sentence_level_model_my_conv = SentenceLevelPOSTagger(len(char_vocab), len(label2id), embedding_size=64,
single_backbone_kwargs=dict(layers_n=3, kernel_size=3, dropout=0.3, conv_layer=MyConv1d),
context_backbone_kwargs=dict(layers_n=3, kernel_size=3, dropout=0.3, conv_layer=MyConv1d))
print('Количество параметров', sum(np.product(t.shape) for t in sentence_level_model_my_conv.parameters()))
(best_val_loss,
best_sentence_level_model_my_conv) = train_eval_loop(sentence_level_model_my_conv,
train_dataset,
test_dataset,
F.cross_entropy,
lr=5e-3,
epoch_n=10,
batch_size=64,
device='cuda',
early_stopping_patience=5,
max_batches_per_epoch_train=500,
max_batches_per_epoch_val=100,
lr_scheduler_ctor=lambda optim: torch.optim.lr_scheduler.ReduceLROnPlateau(optim, patience=2,
factor=0.5,
verbose=True))
# +
train_pred = predict_with_model(best_sentence_level_model_my_conv, train_dataset)
train_loss = F.cross_entropy(torch.tensor(train_pred),
torch.tensor(train_labels))
print('Среднее значение функции потерь на обучении', float(train_loss))
print(classification_report(train_labels.view(-1), train_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
print()
test_pred = predict_with_model(best_sentence_level_model_my_conv, test_dataset)
test_loss = F.cross_entropy(torch.tensor(test_pred),
torch.tensor(test_labels))
print('Среднее значение функции потерь на валидации', float(test_loss))
print(classification_report(test_labels.view(-1), test_pred.argmax(1).reshape(-1), target_names=UNIQUE_TAGS))
| task3_cnn_postag.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # cuDF Cheat Sheets sample code
#
# (c) 2020 NVIDIA, Blazing SQL
#
# Distributed under Apache License 2.0
# ### Imports
import cudf
import numpy as np
# ### Sample DataFrame
df = cudf.DataFrame(
[
(39, 6.88, np.datetime64('2020-10-08T12:12:01'), np.timedelta64(14378,'s'), 'C', 'D', 'data'
, 'RAPIDS.ai is a suite of open-source libraries that allow you to run your end to end data science and analytics pipelines on GPUs.')
, (11, 4.21, None, None , 'A', 'D', 'cuDF'
, 'cuDF is a Python GPU DataFrame (built on the Apache Arrow columnar memory format)')
, (31, 4.71, np.datetime64('2020-10-10T09:26:43'), np.timedelta64(12909,'s'), 'U', 'D', 'memory'
, 'cuDF allows for loading, joining, aggregating, filtering, and otherwise manipulating tabular data using a DataFrame style API.')
, (40, 0.93, np.datetime64('2020-10-11T17:10:00'), np.timedelta64(10466,'s'), 'P', 'B', 'tabular'
, '''If your workflow is fast enough on a single GPU or your data comfortably fits in memory on
a single GPU, you would want to use cuDF.''')
, (33, 9.26, np.datetime64('2020-10-15T10:58:02'), np.timedelta64(35558,'s'), 'O', 'D', 'parallel'
, '''If you want to distribute your workflow across multiple GPUs or have more data than you can fit
in memory on a single GPU you would want to use Dask-cuDF''')
, (42, 4.21, np.datetime64('2020-10-01T10:02:23'), np.timedelta64(20480,'s'), 'U', 'C', 'GPUs'
, 'BlazingSQL provides a high-performance distributed SQL engine in Python')
, (36, 3.01, np.datetime64('2020-09-30T14:36:26'), np.timedelta64(24409,'s'), 'T', 'D', None
, 'BlazingSQL is built on the RAPIDS GPU data science ecosystem')
, (38, 6.44, np.datetime64('2020-10-10T08:34:36'), np.timedelta64(90171,'s'), 'X', 'B', 'csv'
, 'BlazingSQL lets you ETL raw data directly into GPU memory as a GPU DataFrame (GDF)')
, (17, 5.28, np.datetime64('2020-10-09T08:34:40'), np.timedelta64(30532,'s'), 'P', 'D', 'dataframes'
, 'Dask is a flexible library for parallel computing in Python')
, (10, 8.28, np.datetime64('2020-10-03T03:31:21'), np.timedelta64(23552,'s'), 'W', 'B', 'python'
, None)
]
, columns = ['num', 'float', 'datetime', 'timedelta', 'char', 'category', 'word', 'string']
)
df['category'] = df['category'].astype('category')
# ---
#
# # Querying
#
# ---
# #### cudf.core.dataframe.DataFrame.head()
df.head()
df.head(2)
# #### cudf.core.dataframe.DataFrame.info()
df.info()
# #### cudf.core.dataframe.DataFrame.memory_usage()
df.memory_usage()
f'Total memory usage of df: {sum(df.memory_usage().to_pandas())/1024}kB'
# #### cudf.core.dataframe.DataFrame.nlargest()
df.nlargest(3, ['num'])
# #### cudf.core.dataframe.DataFrame.nsmallest()
df.nsmallest(3, ['num'])
# #### cudf.core.dataframe.DataFrame.query()
df.query('num == 10')
df.query('num >= 40')
# #### cudf.core.dataframe.DataFrame.sample()
df.sample()
df.sample(3)
| cheatsheets/cuDF/cuDF_Query.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Starter Notebook
#
# The goal of this notebook is to provide a brief introduction to the challenge and the data associated with it. It won't go into detail on a solution approach since this is a challenge which may be approached in a number of ways.
import pandas as pd
import geopandas as gpd # For loading the map of road segments
# # The Challenge
#
# The main dataset for this challenge (in Train.csv) is a record of crashes within Nairobi over 2018-19. Each crash has an associated datetime and location. Let's load the data and take a look.
# Load the data
df = pd.read_csv('Data/Train.csv', parse_dates=['datetime'])
print(df.shape)
df.head()
# Plot the crashes
df.plot(x='longitude', y='latitude', kind='scatter', figsize=(12, 12), alpha=0.3, title='Crash Locations')
# Rather than trying to predict accident locations or the number of accidents in a given timeframe, we are instead asked to come up with an ambulance deployment schedule that will minimise distance travelled.
#
# This is of course a simplification of the problem, but by solving this you will hopefully develop strategies that can be used going forward.
# View the submission format
ss = pd.read_csv('Data/SampleSubmission.csv', parse_dates=['date'])
ss.head()
# # Scoring
#
# You can re-create the scoring process by creating your own test set and using the following function:
def score(sub, ref):
total_distance = 0
for date, c_lat, c_lon in ref[['datetime', 'latitude', 'longitude']].values:
row = sub.loc[sub.date < date].tail(1) # Prior to Oct 2 this was incorrectly .head(1)
dists = []
for a in range(6):
dist = ((c_lat - row[f'A{a}_Latitude'].values[0])**2+(c_lon - row[f'A{a}_Longitude'].values[0])**2)**0.5
dists.append(dist)
total_distance += min(dists)
return total_distance
# +
# Example:
# Test set
reference = df.loc[df.datetime > '2019-01-01'] # Using 2019 as our test set
# Date rane covering test set
dates = pd.date_range('2019-01-01', '2020-01-01', freq='3h')
# Create submission dataframe
sub = pd.DataFrame({
'date':dates
})
for ambulance in range(6):
sub['A'+str(ambulance)+'_Latitude'] = 0
sub['A'+str(ambulance)+'_Longitude'] = 0
# Place an ambulance in the center of the city:
sub['A'+str(ambulance)+'_Latitude'] = 36.82
sub['A'+str(ambulance)+'_Longitude'] = -1.3
score(sub, reference)
# -
# This scoring function can be used for local testing without needing to submit on Zindi. Lower is better. Experiment with specifying set locations for all 6 ambulances, and see how low you can get your score.
# # Weather Data
#
# Some weatehr variables form the GFS dataset are provided, covering the greater Nairobi area. Let's take a quick look at the dataset before moving on:
weather = pd.read_csv('Data/Weather_Nairobi_Daily_GFS.csv', parse_dates=['Date'])
weather.head()
# Interestingly, Kenya has two rainy seasons per year!
weather.set_index('Date')['precipitable_water_entire_atmosphere'].rolling(10).mean().plot()
# # Road Survey Data
#
# The road survey data contains many different measurements for each surveyed road segment. Unfortunately we can't access the raw data, and must work 'in the dark' in terms of what the column headings mean. However, the naming convention does at least show which columns may be related.
#
# To locate the road segments geographically, a geojson file is provided. Here is an example of merging the survey data with the segments and visualizing the result.
# Load the survey data
road_surveys = pd.read_csv('Data/Segment_info.csv')
road_surveys.head()
# Load the map
road_segment_locs = gpd.read_file('Data/segments_geometry.geojson')
road_segment_locs.head()
# Because this is a geodataframe, we can plot it
road_segment_locs.plot()
# Let's merge the two and color by one of the survey columns
segments_merged = pd.merge(road_segment_locs, road_surveys, on='segment_id', how='left')
segments_merged.plot(column='55_90', figsize=(20, 8))
# This data will come in useful if you're trying to predict the number of crashes along a particular road, or within a specified region.
# # Adding Uber Data
#
# You can access speed data from https://movement.uber.com/explore/nairobi/speeds and clicking 'Download Data'. Here we'll load in the speeds from January 2019 and explore the data briefly.
speeds = pd.read_csv('../uber_data/movement-speeds-hourly-nairobi-2019-1.csv', parse_dates=['utc_timestamp'])
speeds.head()
# Plot speeds over time for a specific OSM way
way = speeds.loc[speeds.osm_way_id == 133901081]
way = way.set_index('utc_timestamp')
way.resample('1h').mean().plot(y='speed_kph_mean')
way.resample('1D').mean().plot(y='speed_kph_mean')
# You can use any and all data available through Uber Movement, including linking the speeds data to OSM segments, downloading quarterly statistics, using the Movement Data Toolkit etc.
# You can find much more info about working with the speeds data here: https://medium.com/uber-movement/working-with-uber-movement-speeds-data-cc01d35937b3 - including how to link the segment_id with OSM data
# # What Next?
#
# There are many ways you could approach this challenge. For example, you could:
# - Create a model to predict the liklihood of an accident given a location, the features of the nearby road segments, the weather, the traffic speed and the time of day
# - Use this model to predict the probability of crashes for different times+locations over the test period, and then sample from this probability distribution to generate plausible crash locations.
# - Use an optimization technique to minimise the travel distance for the ambulances to your fake crash points.
#
# This might seem too complicated - perhaps simply picking 6 fixed locations based on the existing data will be enough? Or maybe it's worth setting separate locations during busy traffic times?
#
# Perhaps a Reinforcement Learning aproach will do well?
#
# Or maybe you can see a totally different way to solve this!
#
# Share your ideas on the discussion boards, and let's see how innovative we can be :)
| Ambualnce-Perabulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dataset - Movielens data
# https://medium.com/@iliazaitsev/how-to-implement-a-recommendation-system-with-deep-learning-and-pytorch-2d40476590f9
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
import io
import os
import math
import copy
import pickle
import zipfile
from textwrap import wrap
from pathlib import Path
from itertools import zip_longest
from collections import defaultdict
from urllib.error import URLError
from urllib.request import urlopen
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import torch
from torch import nn
from torch import optim
from torch.nn import functional as F
from torch.optim.lr_scheduler import _LRScheduler
# -
plt.style.use('ggplot')
def set_random_seed(state=1):
gens = (np.random.seed, torch.manual_seed, torch.cuda.manual_seed)
for set_state in gens:
set_state(state)
RANDOM_STATE = 1
set_random_seed(RANDOM_STATE)
def try_download(url, download_path):
archive_name = url.split('/')[-1]
folder_name, _ = os.path.splitext(archive_name)
try:
r = urlopen(url)
except URLError as e:
print('Cannot download the data. Error: %s' % s)
return
assert r.status == 200
data = r.read()
with zipfile.ZipFile(io.BytesIO(data)) as arch:
arch.extractall(download_path)
print('The archive is extracted into folder: %s' % download_path)
def read_data(path):
files = {}
for filename in path.glob('*'):
if filename.suffix == '.csv':
files[filename.stem] = pd.read_csv(filename)
elif filename.suffix == '.dat':
if filename.stem == 'ratings':
columns = ['userId', 'movieId', 'rating', 'timestamp']
else:
columns = ['movieId', 'title', 'genres']
data = pd.read_csv(filename, sep='::', names=columns, engine='python')
files[filename.stem] = data
return files['ratings'], files['movies']
# pick any other dataset instead
archive_url = f'http://files.grouplens.org/datasets/movielens/ml-1m.zip'
download_path = '/Users/varunn/Documents/NLP-data'
try_download(archive_url, download_path)
# !ls -l $download_path
# pick one of the available folders
ratings, movies = read_data(Path(os.path.join(download_path, 'ml-1m')))
print(ratings.shape)
ratings.head()
print(movies.shape)
movies.head()
def tabular_preview(ratings, n=15):
"""Creates a cross-tabular view of users vs movies."""
user_groups = ratings.groupby('userId')['rating'].count()
top_users = user_groups.sort_values(ascending=False)[:15]
movie_groups = ratings.groupby('movieId')['rating'].count()
top_movies = movie_groups.sort_values(ascending=False)[:15]
top = (
ratings.
join(top_users, rsuffix='_r', how='inner', on='userId').
join(top_movies, rsuffix='_r', how='inner', on='movieId'))
return pd.crosstab(top.userId, top.movieId, top.rating,
aggfunc=np.sum)
tabular_preview(ratings, movies)
def create_dataset(ratings, top=None):
if top is not None:
ratings.groupby('userId')['rating'].count()
unique_users = ratings.userId.unique()
user_to_index = {old: new for new, old in enumerate(unique_users)}
new_users = ratings.userId.map(user_to_index)
unique_movies = ratings.movieId.unique()
movie_to_index = {old: new for new, old in enumerate(unique_movies)}
new_movies = ratings.movieId.map(movie_to_index)
n_users = unique_users.shape[0]
n_movies = unique_movies.shape[0]
X = pd.DataFrame({'user_id': new_users, 'movie_id': new_movies})
y = ratings['rating'].astype(np.float32)
return (n_users, n_movies), (X, y), (user_to_index, movie_to_index)
(n, m), (X, y), _ = create_dataset(ratings)
print(f'Embeddings: {n} users, {m} movies')
print(f'Dataset shape: {X.shape}')
print(f'Target shape: {y.shape}')
X.head()
class ReviewsIterator:
def __init__(self, X, y, batch_size=32, shuffle=True):
X, y = np.asarray(X), np.asarray(y)
if shuffle:
index = np.random.permutation(X.shape[0])
X, y = X[index], y[index]
self.X = X
self.y = y
self.batch_size = batch_size
self.shuffle = shuffle
self.n_batches = int(math.ceil(X.shape[0] // batch_size))
self._current = 0
def __iter__(self):
return self
def __next__(self):
return self.next()
def next(self):
if self._current >= self.n_batches:
raise StopIteration()
k = self._current
self._current += 1
bs = self.batch_size
return self.X[k*bs:(k + 1)*bs], self.y[k*bs:(k + 1)*bs]
def batches(X, y, bs=32, shuffle=True):
for xb, yb in ReviewsIterator(X, y, bs, shuffle):
xb = torch.LongTensor(xb)
yb = torch.FloatTensor(yb)
yield xb, yb.view(-1, 1)
for x_batch, y_batch in batches(X, y, bs=4):
print(x_batch)
print(y_batch)
break
# +
class EmbeddingNet(nn.Module):
"""
Creates a dense network with embedding layers.
Args:
n_users:
Number of unique users in the dataset.
n_movies:
Number of unique movies in the dataset.
n_factors:
Number of columns in the embeddings matrix.
embedding_dropout:
Dropout rate to apply right after embeddings layer.
hidden:
A single integer or a list of integers defining the number of
units in hidden layer(s).
dropouts:
A single integer or a list of integers defining the dropout
layers rates applyied right after each of hidden layers.
"""
def __init__(self, n_users, n_movies,
n_factors=50, embedding_dropout=0.02,
hidden=10, dropouts=0.2):
super().__init__()
hidden = get_list(hidden)
dropouts = get_list(dropouts)
n_last = hidden[-1]
def gen_layers(n_in):
"""
A generator that yields a sequence of hidden layers and
their activations/dropouts.
Note that the function captures `hidden` and `dropouts`
values from the outer scope.
"""
nonlocal hidden, dropouts
assert len(dropouts) <= len(hidden)
for n_out, rate in zip_longest(hidden, dropouts):
yield nn.Linear(n_in, n_out)
yield nn.ReLU()
if rate is not None and rate > 0.:
yield nn.Dropout(rate)
n_in = n_out
self.u = nn.Embedding(n_users, n_factors)
self.m = nn.Embedding(n_movies, n_factors)
self.drop = nn.Dropout(embedding_dropout)
self.hidden = nn.Sequential(*list(gen_layers(n_factors * 2)))
self.fc = nn.Linear(n_last, 1)
self._init()
def forward(self, users, movies, minmax=None):
features = torch.cat([self.u(users), self.m(movies)], dim=1)
x = self.drop(features)
x = self.hidden(x)
out = torch.sigmoid(self.fc(x))
if minmax is not None:
min_rating, max_rating = minmax
out = out*(max_rating - min_rating + 1) + min_rating - 0.5
return out
def _init(self):
"""
Setup embeddings and hidden layers with reasonable initial values.
"""
def init(m):
if type(m) == nn.Linear:
torch.nn.init.xavier_uniform_(m.weight)
m.bias.data.fill_(0.01)
self.u.weight.data.uniform_(-0.05, 0.05)
self.m.weight.data.uniform_(-0.05, 0.05)
self.hidden.apply(init)
init(self.fc)
def get_list(n):
if isinstance(n, (int, float)):
return [n]
elif hasattr(n, '__iter__'):
return list(n)
raise TypeError('layers configuraiton should be a single number or a list of numbers')
# -
EmbeddingNet(n, m, n_factors=150, hidden=100, dropouts=0.5)
EmbeddingNet(n, m, n_factors=150, hidden=[100, 200, 300], dropouts=[0.25, 0.5])
class CyclicLR(_LRScheduler):
def __init__(self, optimizer, schedule, last_epoch=-1):
assert callable(schedule)
self.schedule = schedule
super().__init__(optimizer, last_epoch)
def get_lr(self):
return [self.schedule(self.last_epoch, lr) for lr in self.base_lrs]
def triangular(step_size, max_lr, method='triangular', gamma=0.99):
def scheduler(epoch, base_lr):
period = 2 * step_size
cycle = math.floor(1 + epoch/period)
x = abs(epoch/step_size - 2*cycle + 1)
delta = (max_lr - base_lr)*max(0, (1 - x))
if method == 'triangular':
pass # we've already done
elif method == 'triangular2':
delta /= float(2 ** (cycle - 1))
elif method == 'exp_range':
delta *= (gamma**epoch)
else:
raise ValueError('unexpected method: %s' % method)
return base_lr + delta
return scheduler
def cosine(t_max, eta_min=0):
def scheduler(epoch, base_lr):
t = epoch % t_max
return eta_min + (base_lr - eta_min)*(1 + math.cos(math.pi*t/t_max))/2
return scheduler
def plot_lr(schedule):
ts = list(range(1000))
y = [schedule(t, 0.001) for t in ts]
plt.plot(ts, y)
plot_lr(triangular(250, 0.005))
plot_lr(triangular(250, 0.005, 'triangular2'))
plot_lr(triangular(250, 0.005, 'exp_range', gamma=0.999))
plot_lr(cosine(t_max=500, eta_min=0.0005))
# ### Training Loop
X_train, X_valid, y_train, y_valid = train_test_split(
X, y, test_size=0.2, random_state=RANDOM_STATE)
datasets = {'train': (X_train, y_train), 'val': (X_valid, y_valid)}
dataset_sizes = {'train': len(X_train), 'val': len(X_valid)}
dataset_sizes
minmax = (float(ratings.rating.min()),
float(ratings.rating.max()))
minmax
net = EmbeddingNet(
n_users=n, n_movies=m,
n_factors=150, hidden=[500, 500, 500],
embedding_dropout=0.05, dropouts=[0.5, 0.5, 0.25])
net
# +
lr = 1e-3
wd = 1e-5
bs = 2000
n_epochs = 100
patience = 10
no_improvements = 0
best_loss = np.inf
best_weights = None
history = []
lr_history = []
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
net.to(device)
criterion = nn.MSELoss(reduction='sum')
optimizer = optim.Adam(net.parameters(), lr=lr, weight_decay=wd)
iterations_per_epoch = int(math.ceil(dataset_sizes['train'] // bs))
scheduler = CyclicLR(optimizer, cosine(t_max=iterations_per_epoch * 2,
eta_min=lr/10))
for epoch in range(n_epochs):
stats = {'epoch': epoch + 1, 'total': n_epochs}
for phase in ('train', 'val'):
training = phase == 'train'
running_loss = 0.0
n_batches = 0
for batch in batches(*datasets[phase], shuffle=training, bs=bs):
x_batch, y_batch = [b.to(device) for b in batch]
optimizer.zero_grad()
# compute gradients only during 'train' phase
with torch.set_grad_enabled(training):
outputs = net(x_batch[:, 0], x_batch[:, 1], minmax)
loss = criterion(outputs, y_batch)
print(loss)
# don't update weights and rates when in 'val' phase
if training:
scheduler.step()
loss.backward()
optimizer.step()
lr_history.extend(scheduler.get_lr())
running_loss += loss.item()
epoch_loss = running_loss / dataset_sizes[phase]
stats[phase] = epoch_loss
# early stopping: save weights of the best model so far
if phase == 'val':
if epoch_loss < best_loss:
print('loss improvement on epoch: %d' % (epoch + 1))
best_loss = epoch_loss
best_weights = copy.deepcopy(net.state_dict())
no_improvements = 0
else:
no_improvements += 1
history.append(stats)
print('[{epoch:03d}/{total:03d}] train: {train:.4f} - val: {val:.4f}'.format(**stats))
if no_improvements >= patience:
print('early stopping after epoch {epoch:03d}'.format(**stats))
break
# -
ax = pd.DataFrame(history).drop(columns='total').plot(x='epoch')
_ = plt.plot(lr_history[:2*iterations_per_epoch])
net.load_state_dict(best_weights)
# +
groud_truth, predictions = [], []
with torch.no_grad():
for batch in batches(*datasets['val'], shuffle=False, bs=bs):
x_batch, y_batch = [b.to(device) for b in batch]
outputs = net(x_batch[:, 0], x_batch[:, 1], minmax)
groud_truth.extend(y_batch.tolist())
predictions.extend(outputs.tolist())
groud_truth = np.asarray(groud_truth).ravel()
predictions = np.asarray(predictions).ravel()
# -
final_loss = np.sqrt(np.mean((predictions - groud_truth)**2))
print(f'Final RMSE: {final_loss:.4f}')
with open(os.path.join(download_path, 'ml-1m/pytorch_model_best.weights'),
'wb') as file:
pickle.dump(best_weights, file)
# ### Embeddings Visualization
from sklearn.decomposition import PCA
with open(os.path.join(download_path, 'ml-1m/pytorch_model_best.weights'),
'rb') as file:
best_weights = pickle.load(file)
net.load_state_dict(best_weights)
def to_numpy(tensor):
return tensor.cpu().numpy()
_, _, (user_id_map, movie_id_map) = create_dataset(ratings)
embed_to_original = {v: k for k, v in movie_id_map.items()}
popular_movies = ratings.groupby('movieId').movieId.count().sort_values(
ascending=False).values[:1000]
embed = to_numpy(net.m.weight.data)
pca = PCA(n_components=5)
components = pca.fit(embed[popular_movies].T).components_
print(components.shape)
components
components_df = pd.DataFrame(components.T, columns=[f'fc{i}' for i in range(pca.n_components_)])
movie_ids = [embed_to_original[idx] for idx in components_df.index]
meta = movies.set_index('movieId')
components_df['movie_id'] = movie_ids
components_df['title'] = meta.loc[movie_ids].title.values
components_df['genres'] = meta.loc[movie_ids].genres.values
components_df.sample(4)
def plot_components(components, component, ascending=False):
fig, ax = plt.subplots(figsize=(18, 12))
subset = components.sort_values(by=component, ascending=ascending).iloc[:12]
columns = components_df.columns
features = columns[columns.str.startswith('fc')].tolist()
fc = subset[features]
titles = ['\n'.join(wrap(t, width=10)) for t in subset.title]
genres = subset.genres.str.replace('|', '\n')
labels = [f'{t}\n\n{g}' for t, g in zip(titles, genres)]
fc.plot(ax=ax, kind='bar')
y_ticks = [f'{t:2.2f}' for t in ax.get_yticks()]
ax.set_xticklabels(labels, rotation=0, fontsize=14)
ax.set_yticklabels(y_ticks, fontsize=14)
ax.legend(loc='best', fontsize=14)
plot_title = f"Movies with {['highest', 'lowest'][ascending]} '{component}' component values"
ax.set_title(plot_title, fontsize=20)
plot_components(components_df, 'fc0', ascending=False)
plot_components(components_df, 'fc0', ascending=True)
# ## Matrix Factorization in pytorch
import torch
from torch.autograd import Variable
from torch import nn
from torch.nn import functional as F
class MatrixFactorization(nn.Module):
def __init__(self, n_users, n_items, n_factors=20):
super().__init__()
self.user_factors = nn.Embedding(n_users, n_factors)
self.item_factors = nn.Embedding(n_items, n_factors)
def forward(self, user, item):
return (self.user_factors(user)*self.item_factors(item)).sum(1)
def predict(self, user, item):
return self.forward(user, item)
print('num users: ', n)
print('num movies: ', m)
model = MatrixFactorization(n_users=n, n_items=m, n_factors=150)
model
# +
x_train, y_train = datasets['train']
print(x_train.shape)
print(y_train.shape)
print(type(x_train))
print(x_train.head())
x_val, y_val = datasets['val']
print(x_val.shape)
print(y_val.shape)
print(type(x_val))
print(x_val.head())
# -
count = 0
for row in X.itertuples():
print(row, '\t', row[1], '\t', row[2])
if count > 2:
break
count += 1
# +
import os
import requests
import zipfile
import numpy as np
import pandas as pd
import scipy.sparse as sp
"""
Shamelessly stolen from
https://github.com/maciejkula/triplet_recommendations_keras
"""
def train_test_split(interactions, n=10):
"""
Split an interactions matrix into training and test sets.
Parameters
----------
interactions : np.ndarray
n : int (default=10)
Number of items to select / row to place into test.
Returns
-------
train : np.ndarray
test : np.ndarray
"""
test = np.zeros(interactions.shape)
train = interactions.copy()
for user in range(interactions.shape[0]):
if interactions[user, :].nonzero()[0].shape[0] > n:
test_interactions = np.random.choice(interactions[user, :].nonzero()[0],
size=n,
replace=False)
train[user, test_interactions] = 0.
test[user, test_interactions] = interactions[user, test_interactions]
# Test and training are truly disjoint
assert(np.all((train * test) == 0))
return train, test
def _get_data_path():
"""
Get path to the movielens dataset file.
"""
data_path = os.path.join(os.path.dirname(os.path.abspath(__file__)),
'data')
if not os.path.exists(data_path):
print('Making data path')
os.mkdir(data_path)
return data_path
def _download_movielens(dest_path):
"""
Download the dataset.
"""
url = 'http://files.grouplens.org/datasets/movielens/ml-100k.zip'
req = requests.get(url, stream=True)
print('Downloading MovieLens data')
with open(os.path.join(dest_path, 'ml-100k.zip'), 'wb') as fd:
for chunk in req.iter_content(chunk_size=None):
fd.write(chunk)
with zipfile.ZipFile(os.path.join(dest_path, 'ml-100k.zip'), 'r') as z:
z.extractall(dest_path)
def get_movielens_interactions(ratings):
user_ids = ratings.userId.unique()
item_ids = ratings.movieId.unique()
n_users, n_items = user_ids.shape[0], item_ids.shape[0]
user2index = {value: i for i, value in enumerate(user_ids)}
item2index = {value: i for i, value in enumerate(item_ids)}
interactions = np.zeros((n_users, n_items))
for row in ratings.itertuples():
interactions[user2index[row[1]], item2index[row[2]]] = row[3]
return interactions, user2index, item2index
def get_movielens_train_test_split(ratings, implicit=False):
interactions, user2index, item2index = get_movielens_interactions(
ratings)
if implicit:
interactions = (interactions >= 4).astype(np.float32)
train, test = train_test_split(interactions)
train = sp.coo_matrix(train)
test = sp.coo_matrix(test)
return train, test, user2index, item2index
# -
train, test, user2index, item2index = get_movielens_train_test_split(
ratings)
print(train.shape)
print(test.shape)
train
test
test.astype(np.float32).tocoo().nnz
test.nnz
test.row[0], test.col[0], test.data[0]
| src/recommendation_system_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Frozen Lake Introduction
# - [Fronze Lake](https://gym.openai.com/envs/FrozenLake-v0/) is a simple game where you are on a frozen lake and you need to retrieve an item on the frozen lake where some parts are frozen and some parts are holes (if you walk into them you die)
# - Actions: $\mathcal{A} = \{0, 1, 2, 3\}$
# 1. LEFT: 0
# 2. DOWN = 1
# 3. RIGHT = 2
# 4. UP = 3
# - Whole lake is a 4 x 4 grid world, $\mathcal{S} = \{0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15\}$
# - 
# - On each grid, there are 4 possibilities
# - S: starting point, safe (code = 'SFFF')
# - F: frozen surface, safe (code = 'FHFH')
# - H: hole, fall to your doom (code = 'FFFH')
# - G: goal, where the frisbee is located ('HFFG')
# - 
#
# ### Goal of Frozen Lake
# The key here is we want to get to **G** without falling into the hole **H** in the shortest amount of time
#
# ### Why Dynamic Programming?
# In this game, we know our transition probability function and reward function, essentially the whole environment, allowing us to turn this game into a simple planning problem via dynamic programming through 4 simple functions: (1) policy evaluation (2) policy improvement (3) policy iteration or (4) value iteration
#
# Before we explore how to solve this game, let's first understand how the game works in detail.
# ### Deterministic Policy Environment
# Import gym, installable via `pip install gym`
import gym
# +
# Environment environment Slippery (stochastic policy, move left probability = 1/3) comes by default!
# If we want deterministic policy, we need to create new environment
# Make environment No Slippery (deterministic policy, move left = left)
gym.envs.register(
id='FrozenLakeNotSlippery-v0',
entry_point='gym.envs.toy_text:FrozenLakeEnv',
kwargs={'map_name' : '4x4', 'is_slippery': False},
max_episode_steps=100,
reward_threshold=0.78, # optimum = .8196
)
# You can only register once
# To delete any new environment
# del gym.envs.registry.env_specs['FrozenLakeNotSlippery-v0']
# -
# Make the environment based on deterministic policy
env = gym.make('FrozenLakeNotSlippery-v0')
# State space
print(env.observation_space)
S_n = env.observation_space.n
print(S_n)
# We should expect to see 15 possible grids from 0 to 15 when
# we uniformly randomly sample from our observation space
for i in range(10):
print(env.observation_space.sample())
# Action space
print(env.action_space)
A_n = env.action_space.n
print(A_n)
# We should expect to see 4 actions when
# we uniformly randomly sample:
# 1. LEFT: 0
# 2. DOWN = 1
# 3. RIGHT = 2
# 4. UP = 3
for i in range(10):
print(env.action_space.sample())
# #### Making Steps
# This sets the initial state at S, our starting point
# We can render the environment to see where we are on the 4x4 frozenlake gridworld
env.reset()
env.render()
# +
# Go left (action=0), nothing should happen, and we should stay at the starting point, because there's no grid on the left
env.reset()
action = 0
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 1: move to grid number 1 (unchanged)
# Prob = 1: deterministic policy, if we choose to go left, we'll go left
print(observation, reward, done, prob)
# +
# Go down (action = 1), we should be safe as we step on frozen grid
env.reset()
action = 1
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 4: move to grid number 4
# Prob = 1: deterministic policy, if we choose to go down we'll go down
print(observation, reward, done, prob)
# +
# Go right (action = 2), we should be safe as we step on frozen grid
env.reset()
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 1: move to grid number 1
# Prob = 1: deterministic policy, if we choose to go right we'll go right
print(observation, reward, done, prob)
# +
# Go right twice (action = 2), we should be safe as we step on 2 frozen grids
env.reset()
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 2: move to the right twice from grid 0 to grid 2
# Prob = 1: deterministic policy, if we choose to go right twice we'll go right twice
print(observation, reward, done, prob)
# -
# #### Dying: drop in hole grid 12, H
# +
# Go down thrice (action = 1), we will die as we step onto the grid with a hole
env.reset()
action = 1
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 2: move to the right twice from grid 0 to grid 2
# Prob = 1: deterministic policy, if we choose to go right twice we'll go right twice
# Done = True because the game ends when we die (go onto hole grid (H) or finish the game (G))
print(observation, reward, done, prob)
# -
# #### Winning: get to grid 15, G
# +
# Go right twice (action = 2), go down thrice (action = 1), go right once (action = 2)
env.reset()
# Right Twice
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
# Down Thrice
action = 1
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
(observation, reward, done, prob) = env.step(action)
env.render()
# Right Once
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 2: move to the right twice from grid 0 to grid 2
# Prob = 1: deterministic policy, if we choose to go right twice we'll go right twice
# Done = True because the game ends when we die (go onto hole grid (H) or finish the game (G))
print(observation, reward, done, prob)
# -
# ### Non-deterministic Policy Environment
# Make the environment based on deterministic policy
env = gym.make('FrozenLake-v0')
# +
# Go right once (action = 2), we should go to the right but we did not!
env.seed(8)
env.reset()
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
# Observation = 0: move to the right once from grid 0 to grid 1
# Prob = 1/3: non-deterministic policy, if we choose to go right, there's only a 1/3 probability we would go to the right and with this environment seed we did not
print(observation, reward, done, prob)
# -
# Try to go to the right 10 times, let's see how many times it goes to the right, by right we won't die because we would end up at the extreme right of grid 3
# See how it can go down/left/up/nothing instead of just right?
# Intuitively when we are moving on a frozen lake, some times when we want to walk one direction we may end up in another direction as it's slippery
# Setting seed here of the environment so you can reproduce my results, otherwise stochastic policy will yield different results for each run
env.seed(8)
env.reset()
for i in range(10):
action = 2
(observation, reward, done, prob) = env.step(action)
env.render()
# ### Custom Frozen Lake Non-deterministic Policy Environment
# - Because original code from [OpenAI](https://github.com/openai/gym/blob/master/gym/envs/toy_text/frozen_lake.py) only allows us to run `env.step(action)`, this is challenging if we want to do some visualization of our state-value and action-value (q-value) functions for learning
# - Hence, we'll be copying the whole code from [OpenAI Frozen Lake implementation](https://github.com/openai/gym/blob/master/gym/envs/toy_text/frozen_lake.py) and adding just one line to make sure we can get P via `self.P = P`
# - This code is not important, you can just copy it
# +
import sys
from contextlib import closing
import numpy as np
from six import StringIO, b
from gym import utils
from gym.envs.toy_text import discrete
LEFT = 0
DOWN = 1
RIGHT = 2
UP = 3
MAPS = {
"4x4": [
"SFFF",
"FHFH",
"FFFH",
"HFFG"
],
"8x8": [
"SFFFFFFF",
"FFFFFFFF",
"FFFHFFFF",
"FFFFFHFF",
"FFFHFFFF",
"FHHFFFHF",
"FHFFHFHF",
"FFFHFFFG"
],
}
# Generates a random valid map (one that has a path from start to goal)
# @params size, size of each side of the grid
# @prams p, probability that a tile is frozen
def generate_random_map(size=8, p=0.8):
valid = False
#BFS to check that it's a valid path
def is_valid(arr, r=0, c=0):
if arr[r][c] == 'G':
return True
tmp = arr[r][c]
arr[r][c] = "#"
if r+1 < size and arr[r+1][c] not in '#H':
if is_valid(arr, r+1, c) == True:
arr[r][c] = tmp
return True
if c+1 < size and arr[r][c+1] not in '#H':
if is_valid(arr, r, c+1) == True:
arr[r][c] = tmp
return True
if r-1 >= 0 and arr[r-1][c] not in '#H':
if is_valid(arr, r-1, c) == True:
arr[r][c] = tmp
return True
if c-1 >= 0 and arr[r][c-1] not in '#H':
if is_valid(arr,r, c-1) == True:
arr[r][c] = tmp
return True
arr[r][c] = tmp
return False
while not valid:
p = min(1, p)
res = np.random.choice(['F','H'], (size, size), p=[p, 1-p])
res[0][0] = 'S'
res[-1][-1] = 'G'
valid = is_valid(res)
return ["".join(x) for x in res]
class FrozenLakeEnv(discrete.DiscreteEnv):
"""
Winter is here. You and your friends were tossing around a frisbee at the park
when you made a wild throw that left the frisbee out in the middle of the lake.
The water is mostly frozen, but there are a few holes where the ice has melted.
If you step into one of those holes, you'll fall into the freezing water.
At this time, there's an international frisbee shortage, so it's absolutely imperative that
you navigate across the lake and retrieve the disc.
However, the ice is slippery, so you won't always move in the direction you intend.
The surface is described using a grid like the following
SFFF
FHFH
FFFH
HFFG
S : starting point, safe
F : frozen surface, safe
H : hole, fall to your doom
G : goal, where the frisbee is located
The episode ends when you reach the goal or fall in a hole.
You receive a reward of 1 if you reach the goal, and zero otherwise.
"""
metadata = {'render.modes': ['human', 'ansi']}
def __init__(self, desc=None, map_name="4x4",is_slippery=True):
if desc is None and map_name is None:
desc = generate_random_map()
elif desc is None:
desc = MAPS[map_name]
self.desc = desc = np.asarray(desc,dtype='c')
self.nrow, self.ncol = nrow, ncol = desc.shape
self.reward_range = (0, 1)
nA = 4
nS = nrow * ncol
isd = np.array(desc == b'S').astype('float64').ravel()
isd /= isd.sum()
P = {s : {a : [] for a in range(nA)} for s in range(nS)}
def to_s(row, col):
return row*ncol + col
def inc(row, col, a):
if a==0: # left
col = max(col-1,0)
elif a==1: # down
row = min(row+1,nrow-1)
elif a==2: # right
col = min(col+1,ncol-1)
elif a==3: # up
row = max(row-1,0)
return (row, col)
for row in range(nrow):
for col in range(ncol):
s = to_s(row, col)
for a in range(4):
li = P[s][a]
letter = desc[row, col]
if letter in b'GH':
li.append((1.0, s, 0, True))
else:
if is_slippery:
for b in [(a-1)%4, a, (a+1)%4]:
newrow, newcol = inc(row, col, b)
newstate = to_s(newrow, newcol)
newletter = desc[newrow, newcol]
done = bytes(newletter) in b'GH'
rew = float(newletter == b'G')
li.append((1.0/3.0, newstate, rew, done))
else:
newrow, newcol = inc(row, col, a)
newstate = to_s(newrow, newcol)
newletter = desc[newrow, newcol]
done = bytes(newletter) in b'GH'
rew = float(newletter == b'G')
li.append((1.0, newstate, rew, done))
# New change because environment only allows step without
# specific state for learning environment!
self.P = P
super(FrozenLakeEnv, self).__init__(nS, nA, P, isd)
def render(self, mode='human'):
outfile = StringIO() if mode == 'ansi' else sys.stdout
row, col = self.s // self.ncol, self.s % self.ncol
desc = self.desc.tolist()
desc = [[c.decode('utf-8') for c in line] for line in desc]
desc[row][col] = utils.colorize(desc[row][col], "red", highlight=True)
if self.lastaction is not None:
outfile.write(" ({})\n".format(["Left","Down","Right","Up"][self.lastaction]))
else:
outfile.write("\n")
outfile.write("\n".join(''.join(line) for line in desc)+"\n")
if mode != 'human':
with closing(outfile):
return outfile.getvalue()
# -
# ## Policy Evaluation
# ### Transition Probability Function
# - $\mathcal{P}_{ss'}^a = \mathcal{P}(s' \vert s, a) = \mathbb{P} [S_{t+1} = s' \vert S_t = s, A_t = a]$
# #### Deterministic Environment
# - There's no probability distribution, if you decide to go left you'll go left
# - Hence in this example, given `current_state = 8` and `action = 0` which is left, we will end up with `probability = 1` in `new_state = 9`
# +
# Deterministic
env = FrozenLakeEnv(is_slippery=False)
current_state = 10 # State from S_n=16 State space
action = 0 # Left action from A_n=4 Action space
[(probability, new_state, reward, done)] = env.P[current_state][action]
print('Probability {}, New State {}'.format(probability, new_state))
# -
# #### Stochastic Environment
# - Given $S_t = 10, A_t = 0$ in a stochastic environment, the transition probability functions indicate you can end up in grid 6, 9, 14 each with 1/3 probability:
# - $\mathbb{P} [S_{t+1} = 6 \vert S_t = 10, A_t = 0] = \frac{1}{3}$
# - $\mathbb{P} [S_{t+1} = 9 \vert S_t = 10, A_t = 0] = \frac{1}{3}$
# - $\mathbb{P} [S_{t+1} = 14 \vert S_t = 10, A_t = 0] = \frac{1}{3}$
# +
# Stochastic
env = FrozenLakeEnv(is_slippery=True)
current_state = 10 # State from S_n=16 State space
action = 0 # Left action from A_n=4 Action space
env.P[current_state][action]
# -
# ### Random Policy Function
# +
# Random policy generation
def generate_random_policy(S_n, A_n):
# return np.random.randint(A_n, size=(S_n, A_n))
return np.ones([S_n, A_n]) / A_n
# Given the total number of states S_n = 16
# For each state out of 16 states, we can take 4 actions
# Since this is a stochastic environment, we'll initialize a policy to have equal probabilities 0.25 of doing each action each state
policy = generate_random_policy(S_n, A_n)
print(policy.shape)
# -
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(5, 16))
sns.heatmap(policy, cmap="YlGnBu", annot=True, cbar=False);
# ### Policy Evaluation Function comprising State-value Function
# - How: $\mathcal{V}_{\pi}(s) = \sum_{a \in \mathcal{A}} \pi(a | s) \sum_{s' \in \mathcal{S}} \mathcal{P}_{ss'}^a \big[\mathcal{R}_s^a + \gamma {V}_{\pi}(s')\big]$
# - Simple code equation:
# - Values of state given policy = sum ( action probability \* transition probability \* [reward + discount \* value of new state] )
# - **Aim: getting state-values**
# +
import numpy as np
def policy_evaluation(env, policy, gamma=1., theta=1e-8):
r"""Policy evaluation function. Loop until state values stable, delta < theta.
Returns V comprising values of states under given policy.
Args:
env (gym.env): OpenAI environment class instantiated and assigned to an object.
policy (np.array): policy array to evaluate
gamma (float): discount rate for rewards
theta (float): tiny positive number, anything below it indicates value function convergence
"""
# 1. Create state-value array (16,)
V = np.zeros(S_n)
while True:
delta = 0
# 2. Loop through states
for s in range(S_n):
Vs = 0
# 2.1 Loop through actions for the unique state
# Given each state, we've 4 actions associated with different probabilities
# 0.25 x 4 in this case, so we'll be looping 4 times (4 action probabilities) at each state
for a, action_prob in enumerate(policy[s]):
# 2.1.1 Loop through to get transition probabilities, next state, rewards and whether the game ended
for prob, next_state, reward, done in env.P[s][a]:
# State-value function to get our values of states given policy
Vs += action_prob * prob * (reward + gamma * V[next_state])
# This simple equation allows us to stop this loop when we've converged
# How do we know? The new value of the state is smaller than a tiny positive value we set
# State value change is tiny compared to what we have so we just stop!
delta = max(delta, np.abs(V[s]-Vs))
# 2.2 Update our state value for that state
V[s] = Vs
# 3. Stop policy evaluation if our state values changes are smaller than our tiny positive number
if delta < theta:
break
return V
# +
# Generate random policy with equal probabilities of each action given any state
rand_policy = generate_random_policy(S_n, A_n)
# Evaluate the policy to get state values
V = policy_evaluation(env, rand_policy)
# -
# Plot heatmap
plt.figure(figsize=(8, 8))
sns.heatmap(V.reshape(4, 4), cmap="YlGnBu", annot=True, cbar=False);
# This is our environment
# Notice how the state values near the goal have higher values?
# Those with "H" = hole, where you die if you step, have 0 values indicating those are bad areas to be in
env.render()
# ## Policy Improvement
# ### Action-value (Q-value) function from State-value function
# - How: $\mathcal{Q}_{\pi}(s, a) = \sum_{s' \in \mathcal{S}} \mathcal{P}_{ss'}^a \big[ \mathcal{R}_s^a + \gamma \mathcal{V}_{\pi}(s') \big]$
# - Code equation
# - Values of action = sum ( transition probability \* [reward + discount \* value of next state] )
# - **Aim: getting q-values (action-values)**
def q_value(env, V, s, gamma=1):
r"""Q-value (action-value) function from state-value function
Returns Q values, values of actions.
Args:
env (gym.env): OpenAI environment class instantiated and assigned to an object.
V (np.array): array of state-values obtained from policy evaluation function.
s (integer): integer representing current state in the gridworld
gamma (float): discount rate for rewards.
"""
# 1. Create q-value array for one state
# We have 4 actions, so let's create an array with the size of 4
q = np.zeros(A_n)
# 2. Loop through each action
for a in range(A_n):
# 2.1 For each action, we've our transition probabilities, next state, rewards and whether the game ended
for prob, next_state, reward, done in env.P[s][a]:
# 2.1.1 Get our action-values from state-values
q[a] += prob * (reward + gamma * V[next_state])
# Return action values
return q
# +
# For every state, we've 4 actions, hence we've 16 x 4 q values
Q = np.zeros([S_n, A_n])
# Loop through each state out of 16
# For each state, we will get the 4 q-values associated with the 4 actions
for s in range(env.nS):
Q[s] = q_value(env, V, s)
# -
plt.figure(figsize=(5, 16))
sns.heatmap(Q, cmap="YlGnBu", annot=True, cbar=False);
# Notice how 13/14, those in the last row of the gridworld just before reaching the goal of finishing the game, their action values are large?
env.render()
# ### Policy Improvement Function
# - How: maximizing q-values per state by choosing actions with highest q-values
# - Aim: get improved policy
def policy_improvement(env, V, gamma=1.):
r"""Function to improve the policy by utilizing state values and action (q) values.
Args:
env (gym.env): OpenAI environment class instantiated and assigned to an objects
V (np.array): array of state-values obtained from policy evaluation function
gamma (float): discount of rewards
"""
# 1. Blank policy
policy = np.zeros([env.nS, env.nA]) / env.nA
# 2. For each state in 16 states
for s in range(env.nS):
# 2.1 Get q values: q.shape returns (4,)
q = q_value(env, V, s, gamma)
# 2.2 Find best action based on max q-value
# np.argwhere(q==np.max(q)) gives the position of largest q value
# given array([0.00852356, 0.01163091, 0.0108613 , 0.01550788]), this would return array([[3]]) of shape (1, 1)
# .flatten() reduces the shape to (1,) where we've array([3])
best_a = np.argwhere(q==np.max(q)).flatten()
# 2.3 One-hot encode best action and store into policy array's row for that state
# In our case where the best action is array([3]), this would return
# array([0., 0., 0., 1.]) where position 3 is the best action
# Now we can store the best action into our policy
policy[s] = np.sum([np.eye(env.nA)[i] for i in best_a], axis=0)/len(best_a)
return policy
new_policy = policy_improvement(env, V)
plt.figure(figsize=(5, 16))
sns.heatmap(new_policy, cmap="YlGnBu", annot=True, cbar=False);
# Compared to this equiprobable policy, the one above is making some improvements by maximizing q-values per state
plt.figure(figsize=(5, 16))
sns.heatmap(rand_policy, cmap="YlGnBu", annot=True, cbar=False);
# ### Policy Iteration Function
# - How: loop through policy evaluation (get state-values) and policy improvement functions (use state-values to calculate q-values to improve policy) until optimal policy obtained
# - Aim: improve policy until convergence
# - Convergence: difference of state values between old and new policies is very small (less than theta, a very small positive number)
import copy
def policy_iteration(env, gamma=1, theta=1e-8):
# 1. Create equiprobable policy where every state has 4 actions with equal probabilities as a starting policy
policy = np.ones([env.nS, env.nA]) / env.nA
# 2. Loop through policy_evaluation and policy_improvement functions
while True:
# 2.1 Get state-values
V = policy_evaluation(env, policy, gamma, theta)
# 2.2 Get new policy by getting q-values and maximizing q-values per state to get best action per state
new_policy = policy_improvement(env, V)
# 2.3 Stop if the value function estimates for successive policies has converged
if np.max(abs(policy_evaluation(env, policy) - policy_evaluation(env, new_policy))) < theta * 1e2:
break;
# 2.4 Replace policy with new policy
policy = copy.copy(new_policy)
return policy, V
# obtain the optimal policy and optimal state-value function
policy_pi, V_pi = policy_iteration(env)
# Optimal policy (pi)
# LEFT = 0, DOWN = 1, RIGHT = 2, UP = 3
plt.figure(figsize=(5, 16))
sns.heatmap(policy_pi, cmap="YlGnBu", annot=True, cbar=False, square=True);
# State values
plt.figure(figsize=(8, 8))
sns.heatmap(V_pi.reshape(4, 4), cmap="YlGnBu", annot=True, cbar=False, square=True);
# State values without policy improvement, just evaluation
plt.figure(figsize=(8, 8))
sns.heatmap(V.reshape(4, 4), cmap="YlGnBu", annot=True, cbar=False);
# ### Value iteration
# - Alternative to policy iteration
# - How: loop through to find optimal value function then get one-off policy
# - Aim: improve value function until convergence
# - Convergence: until difference in new and old state values are small (smaller than theta, small positive number)
def value_iteration(env, gamma=1, theta=1e-8):
# 1. Create state values of shape (16,)
V = np.zeros(env.nS)
# 2. Loop through q-value function until convergence
while True:
delta = 0
# 2.1 Loop through each state
for s in range(env.nS):
# 2.2 Archive old state value
v = V[s]
# 2.3 New state value = max of q-value
V[s] = max(q_value(env, V, s, gamma))
delta = max(delta, abs(V[s] - v))
# 2.2 If state value changes small, converged
if delta < theta:
break
# 3. Extract one-off policy with optimal state values
policy = policy_improvement(env, V, gamma)
return policy, V
policy_vi, V_vi = value_iteration(env)
# Optimal policy
# LEFT = 0, DOWN = 1, RIGHT = 2, UP = 3
plt.figure(figsize=(5, 16))
sns.heatmap(policy_vi, cmap="YlGnBu", annot=True, cbar=False, square=True);
# State values
plt.figure(figsize=(8, 8))
sns.heatmap(V_vi.reshape(4, 4), cmap="YlGnBu", annot=True, cbar=False, square=True);
| docs/deep_learning/deep_reinforcement_learning_pytorch/dynamic_programming_frozenlake.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pathlib import Path
import os
import numpy as np
# + pycharm={"name": "#%%\n"}
experiments_dataframe_path = Path(os.path.abspath("trained_clf_exploration.ipynb")).parent.parent / 'clf_experiments_dataframe_bartholin.csv'
# + pycharm={"name": "#%%\n"}
df = pd.read_csv(experiments_dataframe_path)
print(df.columns.tolist())
# + pycharm={"name": "#%%\n"}
eval_columns = ['dice', 'mean_AP_total', 'precision', 'specificity', 'accuracy']
# + [markdown] pycharm={"name": "#%% md\n"}
# # Best methods for bin labels
# + pycharm={"name": "#%%\n"}
show_cols = ['pred_count_Finding', 'gt_count_Finding', 'img_clf_type', 'mean_AP_total', 'clf_loss','initial_learning_rate', 'weighted_sampler','binary_labels', 'modality', 'text_encoding', 'total_epochs',*eval_columns ]
temp_exp_df = df.dropna(subset= [*eval_columns])
sub_df = temp_exp_df.loc[(temp_exp_df['binary_labels'] == 1)& (temp_exp_df['total_epochs'] > 4), show_cols]
df_stats = sub_df[[ *eval_columns]].apply(pd.DataFrame.describe, axis=1)
sub_dfdf = sub_df.merge(df_stats, left_index=True, right_index = True)
sub_dfdf[['mean',*show_cols]].sort_values(by=['mean'], ascending=False)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Best methods for 3 labels
# + pycharm={"name": "#%%\n"}
show_cols = ['clf_loss','initial_learning_rate', 'weighted_sampler','binary_labels', 'modality', 'total_epochs',*eval_columns ]
temp_exp_df = df.dropna(subset= [*eval_columns])
sub_df = temp_exp_df.loc[(temp_exp_df['binary_labels'] != 1), show_cols]
df_stats = sub_df[[ *eval_columns]].apply(pd.DataFrame.describe, axis=1)
sub_dfdf = sub_df.merge(df_stats, left_index=True, right_index = True)
sub_dfdf[['mean',*show_cols]].sort_values(by=['mean'], ascending=False)
# + [markdown] pycharm={"name": "#%% md\n"}
# # Which method has the highest average precision?
# ### modality text with word encoding
# + pycharm={"name": "#%%\n"}
print(np.max(df['dice']))
df.loc[df['dice'] == np.max(df['dice']), ['weighted_sampler','binary_labels', 'modality', 'text_encoding', 'total_epochs', 'dice']]
# + [markdown] pycharm={"name": "#%% md\n"}
# # Overall the densenet performs better than the resnet
# ### If the fixed_extractor is NaN of False, the whole densenet was trained (fine_tuning). If true, the extractor part was frozen (transfer-learning).
# + pycharm={"name": "#%%\n"}
temp = df.loc[(df['modality'] == 'PA') & (df['img_clf_type'] == 'densenet'), ['modality', 'mean_AP', 'total_epochs', 'fixed_extractor']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
temp = df.loc[(df['modality'] == 'PA') & (df['img_clf_type'] == 'resnet'), ['modality', 'mean_AP', 'total_epochs']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
# + pycharm={"name": "#%%\n"}
temp = df.loc[(df['modality'] == 'text') & (df['img_clf_type'] == 'densenet'), ['modality', 'mean_AP', 'total_epochs']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
temp = df.loc[(df['modality'] == 'text') & (df['img_clf_type'] == 'resnet'), ['modality', 'mean_AP', 'total_epochs']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
# + pycharm={"name": "#%%\n"}
temp = df.loc[(df['modality'] == 'Lateral') & (df['img_clf_type'] == 'densenet'), ['modality', 'mean_AP', 'total_epochs']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
temp = df.loc[(df['modality'] == 'Lateral') & (df['img_clf_type'] == 'resnet'), ['modality', 'mean_AP', 'total_epochs']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
# + [markdown] pycharm={"name": "#%% md\n"}
# # What works best for the densenet, transfer learning or fine-tuning?
# ### Fine tuning works better
# + pycharm={"name": "#%%\n"}
temp =df.loc[((df['img_clf_type'] == 'densenet') & (df['modality'] != 'text')&(df['fixed_extractor'] == 1.0)), ['modality', 'mean_AP', 'total_epochs', 'fixed_extractor']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
temp =df.loc[((df['img_clf_type'] == 'densenet') & (df['modality'] != 'text')&(df['fixed_extractor'] != 1.0)), ['modality', 'mean_AP', 'total_epochs', 'fixed_extractor']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
# + [markdown] pycharm={"name": "#%% md\n", "is_executing": true}
# # Does a bigger image size improve the score by a lot?
# + pycharm={"name": "#%%\n"}
temp = df.loc[((df['img_size'] == 128) & (df['modality'] != 'text')), ['modality', 'mean_AP', 'total_epochs', 'fixed_extractor']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
temp = df.loc[((df['img_size'] == 256) & (df['modality'] != 'text')), ['modality', 'mean_AP', 'total_epochs', 'fixed_extractor']]
display(temp.loc[temp['mean_AP'] == temp['mean_AP'].max()])
# + pycharm={"name": "#%%\n"}
# Does a larger
| mimic/notebooks/trained_clf_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <div align="center">Classification of DT Signals</div>
# ---------------------------------------------------------------------
#
# you can Find me on Github:
# > ###### [ GitHub](https://github.com/lev1khachatryan)
#
# Just like Continuous time signals, Discrete time signals can be classified according to the conditions or operations on the signals.
#
# # <div align="center">Even and Odd Signals</div>
# ---------------------------------------------------------------------
#
# ## Even Signal
# A signal is said to be even or symmetric if it satisfies the following condition:
#
# $x(-t) = x(t)$
#
# <img src='asset/5/1.png'>
#
# Here, we can see that x(-1) = x(1), x(-2) = x(2) and x(-n) = x(n). Thus, it is an even signal.
#
#
#
# ## Odd Signal
# A signal is said to be odd if it satisfies the following condition:
#
# $x(-t) = -x(t)$
#
# <img src='asset/5/2.png'>
#
# From the figure, we can see that x(1) = -x(-1), x(2) = -x(2) and x(n) = -x(-n). Hence, it is an odd as well as anti-symmetric signal.
#
#
#
# # <div align="center">Periodic and Non-Periodic Signals</div>
# ---------------------------------------------------------------------
#
# <img src='asset/5/3.png'>
#
#
#
# # <div align="center">Energy and Power Signals</div>
# ---------------------------------------------------------------------
#
# <img src='asset/5/4.png'>
#
#
#
#
#
| Lectures/Basics/5. Classification of DT Signals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from matplotlib import pyplot as plt
data = pd.read_csv('countries.csv')
data.head()
data_1997 = data[data.year == 1997]
europe_1997 = data_1997[data_1997.continent == 'Europe']
americas_1997 = data_1997[data_1997.continent == 'Americas']
# +
print("Europe's life Expectancy mean in 1997")
print(europe_1997.lifeExpectancy.mean())
print("Americas's life Expectancy mean in 1997")
print(americas_1997.lifeExpectancy.mean())
print("Europe's life Expectancy median in 1997")
print(europe_1997.lifeExpectancy.median())
print("Americas's life Expectancy median in 1997")
print(americas_1997.lifeExpectancy.median())
# +
minrange = min(min(americas_1997.lifeExpectancy), min(europe_1997.lifeExpectancy))
maxrange = max(max(americas_1997.lifeExpectancy), max(europe_1997.lifeExpectancy))
plt.subplot(2, 1, 1)
plt.title('Distribution of Life Expectancy')
plt.hist(europe_1997.lifeExpectancy, range=(minrange, maxrange), edgecolor='black')
plt.ylabel('Europe')
plt.subplot(2, 1, 2)
plt.hist(americas_1997.lifeExpectancy, range=(minrange, maxrange), edgecolor='black')
plt.ylabel('Americas')
plt.show()
# -
americas_1997[americas_1997.lifeExpectancy < 65]
| pluralsight/data_visualization_YK_Sugi/Module 4 - Histograms/Practice Problem 1 - Compare Europe and Americas' Life Expectancy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# # Statistical inference
#
#
# Here we will briefly cover multiple concepts of inferential statistics in an
# introductory manner, and demonstrate how to use some MNE statistical functions.
# :depth: 3
#
# +
# Authors: <NAME> <<EMAIL>>
# License: BSD (3-clause)
from functools import partial
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # noqa, analysis:ignore
import mne
from mne.stats import (ttest_1samp_no_p, bonferroni_correction, fdr_correction,
permutation_t_test, permutation_cluster_1samp_test)
print(__doc__)
# -
# Hypothesis testing
# ------------------
# Null hypothesis
# ^^^^^^^^^^^^^^^
# From `Wikipedia <https://en.wikipedia.org/wiki/Null_hypothesis>`__:
#
# In inferential statistics, a general statement or default position that
# there is no relationship between two measured phenomena, or no
# association among groups.
#
# We typically want to reject a **null hypothesis** with
# some probability (e.g., p < 0.05). This probability is also called the
# significance level $\alpha$.
# To think about what this means, let's follow the illustrative example from
# [1]_ and construct a toy dataset consisting of a 40 x 40 square with a
# "signal" present in the center with white noise added and a Gaussian
# smoothing kernel applied.
#
#
# +
width = 40
n_subjects = 10
signal_mean = 100
signal_sd = 100
noise_sd = 0.01
gaussian_sd = 5
sigma = 1e-3 # sigma for the "hat" method
n_permutations = 'all' # run an exact test
n_src = width * width
# For each "subject", make a smoothed noisy signal with a centered peak
rng = np.random.RandomState(2)
X = noise_sd * rng.randn(n_subjects, width, width)
# Add a signal at the center
X[:, width // 2, width // 2] = signal_mean + rng.randn(n_subjects) * signal_sd
# Spatially smooth with a 2D Gaussian kernel
size = width // 2 - 1
gaussian = np.exp(-(np.arange(-size, size + 1) ** 2 / float(gaussian_sd ** 2)))
for si in range(X.shape[0]):
for ri in range(X.shape[1]):
X[si, ri, :] = np.convolve(X[si, ri, :], gaussian, 'same')
for ci in range(X.shape[2]):
X[si, :, ci] = np.convolve(X[si, :, ci], gaussian, 'same')
# -
# The data averaged over all subjects looks like this:
#
#
fig, ax = plt.subplots()
ax.imshow(X.mean(0), cmap='inferno')
ax.set(xticks=[], yticks=[], title="Data averaged over subjects")
# In this case, a null hypothesis we could test for each voxel is:
#
# There is no difference between the mean value and zero
# ($H_0 \colon \mu = 0$).
#
# The alternative hypothesis, then, is that the voxel has a non-zero mean
# ($H_1 \colon \mu \neq 0$).
# This is a *two-tailed* test because the mean could be less than
# or greater than zero, whereas a *one-tailed* test would test only one of
# these possibilities, i.e. $H_1 \colon \mu \geq 0$ or
# $H_1 \colon \mu \leq 0$.
#
# <div class="alert alert-info"><h4>Note</h4><p>Here we will refer to each spatial location as a "voxel".
# In general, though, it could be any sort of data value,
# including cortical vertex at a specific time, pixel in a
# time-frequency decomposition, etc.</p></div>
#
# Parametric tests
# ^^^^^^^^^^^^^^^^
# Let's start with a **paired t-test**, which is a standard test
# for differences in paired samples. Mathematically, it is equivalent
# to a 1-sample t-test on the difference between the samples in each condition.
# The paired t-test is **parametric**
# because it assumes that the underlying sample distribution is Gaussian, and
# is only valid in this case. This happens to be satisfied by our toy dataset,
# but is not always satisfied for neuroimaging data.
#
# In the context of our toy dataset, which has many voxels
# ($40 \cdot 40 = 1600$), applying the paired t-test is called a
# *mass-univariate* approach as it treats each voxel independently.
#
#
# +
titles = ['t']
out = stats.ttest_1samp(X, 0, axis=0)
ts = [out[0]]
ps = [out[1]]
mccs = [False] # these are not multiple-comparisons corrected
def plot_t_p(t, p, title, mcc, axes=None):
if axes is None:
fig = plt.figure(figsize=(6, 3))
axes = [fig.add_subplot(121, projection='3d'), fig.add_subplot(122)]
show = True
else:
show = False
p_lims = [0.1, 0.001]
t_lims = -stats.distributions.t.ppf(p_lims, n_subjects - 1)
p_lims = [-np.log10(p) for p in p_lims]
# t plot
x, y = np.mgrid[0:width, 0:width]
surf = axes[0].plot_surface(x, y, np.reshape(t, (width, width)),
rstride=1, cstride=1, linewidth=0,
vmin=t_lims[0], vmax=t_lims[1], cmap='viridis')
axes[0].set(xticks=[], yticks=[], zticks=[],
xlim=[0, width - 1], ylim=[0, width - 1])
axes[0].view_init(30, 15)
cbar = plt.colorbar(ax=axes[0], shrink=0.75, orientation='horizontal',
fraction=0.1, pad=0.025, mappable=surf)
cbar.set_ticks(t_lims)
cbar.set_ticklabels(['%0.1f' % t_lim for t_lim in t_lims])
cbar.set_label('t-value')
cbar.ax.get_xaxis().set_label_coords(0.5, -0.3)
if not show:
axes[0].set(title=title)
if mcc:
axes[0].title.set_weight('bold')
# p plot
use_p = -np.log10(np.reshape(np.maximum(p, 1e-5), (width, width)))
img = axes[1].imshow(use_p, cmap='inferno', vmin=p_lims[0], vmax=p_lims[1],
interpolation='nearest')
axes[1].set(xticks=[], yticks=[])
cbar = plt.colorbar(ax=axes[1], shrink=0.75, orientation='horizontal',
fraction=0.1, pad=0.025, mappable=img)
cbar.set_ticks(p_lims)
cbar.set_ticklabels(['%0.1f' % p_lim for p_lim in p_lims])
cbar.set_label(r'$-\log_{10}(p)$')
cbar.ax.get_xaxis().set_label_coords(0.5, -0.3)
if show:
text = fig.suptitle(title)
if mcc:
text.set_weight('bold')
plt.subplots_adjust(0, 0.05, 1, 0.9, wspace=0, hspace=0)
mne.viz.utils.plt_show()
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# -
# "Hat" variance adjustment
# ~~~~~~~~~~~~~~~~~~~~~~~~~
# The "hat" technique regularizes the variance values used in the t-test
# calculation [1]_ to compensate for implausibly small variances.
#
#
ts.append(ttest_1samp_no_p(X, sigma=sigma))
ps.append(stats.distributions.t.sf(np.abs(ts[-1]), len(X) - 1) * 2)
titles.append(r'$\mathrm{t_{hat}}$')
mccs.append(False)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# Non-parametric tests
# ^^^^^^^^^^^^^^^^^^^^
# Instead of assuming an underlying Gaussian distribution, we could instead
# use a **non-parametric resampling** method. In the case of a paired t-test
# between two conditions A and B, which is mathematically equivalent to a
# one-sample t-test between the difference in the conditions A-B, under the
# null hypothesis we have the principle of **exchangeability**. This means
# that, if the null is true, we can exchange conditions and not change
# the distribution of the test statistic.
#
# When using a paired t-test, exchangeability thus means that we can flip the
# signs of the difference between A and B. Therefore, we can construct the
# **null distribution** values for each voxel by taking random subsets of
# samples (subjects), flipping the sign of their difference, and recording the
# absolute value of the resulting statistic (we record the absolute value
# because we conduct a two-tailed test). The absolute value of the statistic
# evaluated on the veridical data can then be compared to this distribution,
# and the p-value is simply the proportion of null distribution values that
# are smaller.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>In the case of a true one-sample t-test, i.e. analyzing a single
# condition rather than the difference between two conditions,
# it is not clear where/how exchangeability applies; see
# `this FieldTrip discussion <ft_exch_>`_.</p></div>
#
# In the case where ``n_permutations`` is large enough (or "all") so
# that the complete set of unique resampling exchanges can be done
# (which is $2^{N_{samp}}-1$ for a one-tailed and
# $2^{N_{samp}-1}-1$ for a two-tailed test, not counting the
# veridical distribution), instead of randomly exchanging conditions
# the null is formed from using all possible exchanges. This is known
# as a permutation test (or exact test).
#
#
# +
# Here we have to do a bit of gymnastics to get our function to do
# a permutation test without correcting for multiple comparisons:
X.shape = (n_subjects, n_src) # flatten the array for simplicity
titles.append('Permutation')
ts.append(np.zeros(width * width))
ps.append(np.zeros(width * width))
mccs.append(False)
for ii in range(n_src):
ts[-1][ii], ps[-1][ii] = permutation_t_test(X[:, [ii]], verbose=False)[:2]
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# -
# Multiple comparisons
# --------------------
# So far, we have done no correction for multiple comparisons. This is
# potentially problematic for these data because there are
# $40 \cdot 40 = 1600$ tests being performed. If we use a threshold
# p < 0.05 for each individual test, we would expect many voxels to be declared
# significant even if there were no true effect. In other words, we would make
# many **type I errors** (adapted from `here <errors_>`_):
#
# .. rst-class:: skinnytable
#
# +----------+--------+------------------+------------------+
# | | Null hypothesis |
# | +------------------+------------------+
# | | True | False |
# +==========+========+==================+==================+
# | | | Type I error | Correct |
# | | Yes | False positive | True positive |
# + Reject +--------+------------------+------------------+
# | | | Correct | Type II error |
# | | No | True Negative | False negative |
# +----------+--------+------------------+------------------+
#
# To see why, consider a standard $\alpha = 0.05$.
# For a single test, our probability of making a type I error is 0.05.
# The probability of making at least one type I error in
# $N_{\mathrm{test}}$ independent tests is then given by
# $1 - (1 - \alpha)^{N_{\mathrm{test}}}$:
#
#
N = np.arange(1, 80)
alpha = 0.05
p_type_I = 1 - (1 - alpha) ** N
fig, ax = plt.subplots(figsize=(4, 3))
ax.scatter(N, p_type_I, 3)
ax.set(xlim=N[[0, -1]], ylim=[0, 1], xlabel=r'$N_{\mathrm{test}}$',
ylabel=u'Probability of at least\none type I error')
ax.grid(True)
fig.tight_layout()
fig.show()
# To combat this problem, several methods exist. Typically these
# provide control over either one of the following two measures:
#
# 1. `Familywise error rate (FWER) <fwer_>`_
# The probability of making one or more type I errors:
#
# .. math::
# \mathrm{P}(N_{\mathrm{type\ I}} >= 1 \mid H_0)
#
# 2. `False discovery rate (FDR) <fdr_>`_
# The expected proportion of rejected null hypotheses that are
# actually true:
#
# .. math::
# \mathrm{E}(\frac{N_{\mathrm{type\ I}}}{N_{\mathrm{reject}}}
# \mid N_{\mathrm{reject}} > 0) \cdot
# \mathrm{P}(N_{\mathrm{reject}} > 0 \mid H_0)
#
# We cover some techniques that control FWER and FDR below.
#
# Bonferroni correction
# ^^^^^^^^^^^^^^^^^^^^^
# Perhaps the simplest way to deal with multiple comparisons, `Bonferroni
# correction <https://en.wikipedia.org/wiki/Bonferroni_correction>`__
# conservatively multiplies the p-values by the number of comparisons to
# control the FWER.
#
#
titles.append('Bonferroni')
ts.append(ts[-1])
ps.append(bonferroni_correction(ps[0])[1])
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# False discovery rate (FDR) correction
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
# Typically FDR is performed with the Benjamini-Hochberg procedure, which
# is less restrictive than Bonferroni correction for large numbers of
# comparisons (fewer type II errors), but provides less strict control of type
# I errors.
#
#
titles.append('FDR')
ts.append(ts[-1])
ps.append(fdr_correction(ps[0])[1])
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# Non-parametric resampling test with a maximum statistic
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
# **Non-parametric resampling tests** can also be used to correct for multiple
# comparisons. In its simplest form, we again do permutations using
# exchangeability under the null hypothesis, but this time we take the
# *maximum statistic across all voxels* in each permutation to form the
# null distribution. The p-value for each voxel from the veridical data
# is then given by the proportion of null distribution values
# that were smaller.
#
# This method has two important features:
#
# 1. It controls FWER.
# 2. It is non-parametric. Even though our initial test statistic
# (here a 1-sample t-test) is parametric, the null
# distribution for the null hypothesis rejection (the mean value across
# subjects is indistinguishable from zero) is obtained by permutations.
# This means that it makes no assumptions of Gaussianity
# (which do hold for this example, but do not in general for some types
# of processed neuroimaging data).
#
#
titles.append(r'$\mathbf{Perm_{max}}$')
out = permutation_t_test(X, verbose=False)[:2]
ts.append(out[0])
ps.append(out[1])
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# Clustering
# ^^^^^^^^^^
# Each of the aforementioned multiple comparisons corrections have the
# disadvantage of not fully incorporating the correlation structure of the
# data, namely that points close to one another (e.g., in space or time) tend
# to be correlated. However, by defining the connectivity/adjacency/neighbor
# structure in our data, we can use **clustering** to compensate.
#
# To use this, we need to rethink our null hypothesis. Instead
# of thinking about a null hypothesis about means per voxel (with one
# independent test per voxel), we consider a null hypothesis about sizes
# of clusters in our data, which could be stated like:
#
# The distribution of spatial cluster sizes observed in two experimental
# conditions are drawn from the same probability distribution.
#
# Here we only have a single condition and we contrast to zero, which can
# be thought of as:
#
# The distribution of spatial cluster sizes is independent of the sign
# of the data.
#
# In this case, we again do permutations with a maximum statistic, but, under
# each permutation, we:
#
# 1. Compute the test statistic for each voxel individually.
# 2. Threshold the test statistic values.
# 3. Cluster voxels that exceed this threshold (with the same sign) based on
# adjacency.
# 4. Retain the size of the largest cluster (measured, e.g., by a simple voxel
# count, or by the sum of voxel t-values within the cluster) to build the
# null distribution.
#
# After doing these permutations, the cluster sizes in our veridical data
# are compared to this null distribution. The p-value associated with each
# cluster is again given by the proportion of smaller null distribution
# values. This can then be subjected to a standard p-value threshold
# (e.g., p < 0.05) to reject the null hypothesis (i.e., find an effect of
# interest).
#
# This reframing to consider *cluster sizes* rather than *individual means*
# maintains the advantages of the standard non-parametric permutation
# test -- namely controlling FWER and making no assumptions of parametric
# data distribution.
# Critically, though, it also accounts for the correlation structure in the
# data -- which in this toy case is spatial but in general can be
# multidimensional (e.g., spatio-temporal) -- because the null distribution
# will be derived from data in a way that preserves these correlations.
#
# However, there is a drawback. If a cluster significantly deviates from
# the null, no further inference on the cluster (e.g., peak location) can be
# made, as the entire cluster as a whole is used to reject the null.
# Moreover, because the test statistic concerns the full data, the null
# hypothesis (and our rejection of it) refers to the structure of the full
# data. For more information, see also the comprehensive
# `FieldTrip tutorial <ft_cluster_>`_.
#
# Defining the connectivity/neighbor/adjacency matrix
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# First we need to define our connectivity/neighbor/adjacency matrix.
# This is a square array (or sparse matrix) of shape ``(n_src, n_src)`` that
# contains zeros and ones to define which spatial points are connected, i.e.,
# which voxels are adjacent to each other. In our case this
# is quite simple, as our data are aligned on a rectangular grid.
#
# Let's pretend that our data were smaller -- a 3 x 3 grid. Thinking about
# each voxel as being connected to the other voxels it touches, we would
# need a 9 x 9 connectivity matrix. The first row of this matrix contains the
# voxels in the flattened data that the first voxel touches. Since it touches
# the second element in the first row and the first element in the second row
# (and is also a neighbor to itself), this would be::
#
# [1, 1, 0, 1, 0, 0, 0, 0, 0]
#
# :mod:`sklearn.feature_extraction` provides a convenient function for this:
#
#
from sklearn.feature_extraction.image import grid_to_graph # noqa: E402
mini_connectivity = grid_to_graph(3, 3).toarray()
assert mini_connectivity.shape == (9, 9)
print(mini_connectivity[0])
# In general the connectivity between voxels can be more complex, such as
# those between sensors in 3D space, or time-varying activation at brain
# vertices on a cortical surface. MNE provides several convenience functions
# for computing connectivity/neighbor/adjacency matrices (see the
# `Statistics API <api_reference_statistics>`).
#
# Standard clustering
# ~~~~~~~~~~~~~~~~~~~
# Here, since our data are on a grid, we can use ``connectivity=None`` to
# trigger optimized grid-based code, and run the clustering algorithm.
#
#
titles.append('Clustering')
# Reshape data to what is equivalent to (n_samples, n_space, n_time)
X.shape = (n_subjects, width, width)
# Compute threshold from t distribution (this is also the default)
threshold = stats.distributions.t.ppf(1 - alpha, n_subjects - 1)
t_clust, clusters, p_values, H0 = permutation_cluster_1samp_test(
X, n_jobs=1, threshold=threshold, connectivity=None,
n_permutations=n_permutations)
# Put the cluster data in a viewable format
p_clust = np.ones((width, width))
for cl, p in zip(clusters, p_values):
p_clust[cl] = p
ts.append(t_clust)
ps.append(p_clust)
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# "Hat" variance adjustment
# ~~~~~~~~~~~~~~~~~~~~~~~~~
# This method can also be used in this context to correct for small
# variances [1]_:
#
#
titles.append(r'$\mathbf{C_{hat}}$')
stat_fun_hat = partial(ttest_1samp_no_p, sigma=sigma)
t_hat, clusters, p_values, H0 = permutation_cluster_1samp_test(
X, n_jobs=1, threshold=threshold, connectivity=None,
n_permutations=n_permutations, stat_fun=stat_fun_hat, buffer_size=None)
p_hat = np.ones((width, width))
for cl, p in zip(clusters, p_values):
p_hat[cl] = p
ts.append(t_hat)
ps.append(p_hat)
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
#
# Threshold-free cluster enhancement (TFCE)
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
# TFCE eliminates the free parameter initial ``threshold`` value that
# determines which points are included in clustering by approximating
# a continuous integration across possible threshold values with a standard
# `Riemann sum <https://en.wikipedia.org/wiki/Riemann_sum>`__ [2]_.
# This requires giving a starting threshold ``start`` and a step
# size ``step``, which in MNE is supplied as a dict.
# The smaller the ``step`` and closer to 0 the ``start`` value,
# the better the approximation, but the longer it takes.
#
# A significant advantage of TFCE is that, rather than modifying the
# statistical null hypothesis under test (from one about individual voxels
# to one about the distribution of clusters in the data), it modifies the *data
# under test* while still controlling for multiple comparisons.
# The statistical test is then done at the level of individual voxels rather
# than clusters. This allows for evaluation of each point
# independently for significance rather than only as cluster groups.
#
#
titles.append(r'$\mathbf{C_{TFCE}}$')
threshold_tfce = dict(start=0, step=0.2)
t_tfce, _, p_tfce, H0 = permutation_cluster_1samp_test(
X, n_jobs=1, threshold=threshold_tfce, connectivity=None,
n_permutations=n_permutations)
ts.append(t_tfce)
ps.append(p_tfce)
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# We can also combine TFCE and the "hat" correction:
#
#
titles.append(r'$\mathbf{C_{hat,TFCE}}$')
t_tfce_hat, _, p_tfce_hat, H0 = permutation_cluster_1samp_test(
X, n_jobs=1, threshold=threshold_tfce, connectivity=None,
n_permutations=n_permutations, stat_fun=stat_fun_hat, buffer_size=None)
ts.append(t_tfce_hat)
ps.append(p_tfce_hat)
mccs.append(True)
plot_t_p(ts[-1], ps[-1], titles[-1], mccs[-1])
# Visualize and compare methods
# -----------------------------
# Let's take a look at these statistics. The top row shows each test statistic,
# and the bottom shows p-values for various statistical tests, with the ones
# with proper control over FWER or FDR with bold titles.
#
#
fig = plt.figure(facecolor='w', figsize=(14, 3))
assert len(ts) == len(titles) == len(ps)
for ii in range(len(ts)):
ax = [fig.add_subplot(2, 10, ii + 1, projection='3d'),
fig.add_subplot(2, 10, 11 + ii)]
plot_t_p(ts[ii], ps[ii], titles[ii], mccs[ii], ax)
fig.tight_layout(pad=0, w_pad=0.05, h_pad=0.1)
plt.show()
# The first three columns show the parametric and non-parametric statistics
# that are not corrected for multiple comparisons:
#
# - Mass univariate **t-tests** result in jagged edges.
# - **"Hat" variance correction** of the t-tests produces less peaky edges,
# correcting for sharpness in the statistic driven by low-variance voxels.
# - **Non-parametric resampling tests** are very similar to t-tests. This is to
# be expected: the data are drawn from a Gaussian distribution, and thus
# satisfy parametric assumptions.
#
# The next three columns show multiple comparison corrections of the
# mass univariate tests (parametric and non-parametric). These
# too conservatively correct for multiple comparisons because neighboring
# voxels in our data are correlated:
#
# - **Bonferroni correction** eliminates any significant activity.
# - **FDR correction** is less conservative than Bonferroni.
# - A **permutation test with a maximum statistic** also eliminates any
# significant activity.
#
# The final four columns show the non-parametric cluster-based permutation
# tests with a maximum statistic:
#
# - **Standard clustering** identifies the correct region. However, the whole
# area must be declared significant, so no peak analysis can be done.
# Also, the peak is broad.
# - **Clustering with "hat" variance adjustment** tightens the estimate of
# significant activity.
# - **Clustering with TFCE** allows analyzing each significant point
# independently, but still has a broadened estimate.
# - **Clustering with TFCE and "hat" variance adjustment** tightens the area
# declared significant (again FWER corrected).
#
# Statistical functions in MNE
# ----------------------------
# The complete listing of statistical functions provided by MNE are in
# the `Statistics API list <api_reference_statistics>`, but we will give
# a brief overview here.
#
# MNE provides several convenience parametric testing functions that can be
# used in conjunction with the non-parametric clustering methods. However,
# the set of functions we provide is not meant to be exhaustive.
#
# If the univariate statistical contrast of interest is not listed here
# (e.g., interaction term in an unbalanced ANOVA), consider checking out the
# :mod:`statsmodels` package. It offers many functions for computing
# statistical contrasts, e.g., :func:`statsmodels.stats.anova.anova_lm`.
# To use these functions in clustering:
#
# 1. Determine which test statistic (e.g., t-value, F-value) you would use
# in a univariate context to compute your contrast of interest. In other
# words, if there were only a single output such as reaction times, what
# test statistic might you compute on the data?
# 2. Wrap the call to that function within a function that takes an input of
# the same shape that is expected by your clustering function,
# and returns an array of the same shape without the "samples" dimension
# (e.g., :func:`mne.stats.permutation_cluster_1samp_test` takes an array
# of shape ``(n_samples, p, q)`` and returns an array of shape ``(p, q)``).
# 3. Pass this wrapped function to the ``stat_fun`` argument to the clustering
# function.
# 4. Set an appropriate ``threshold`` value (float or dict) based on the
# values your statistical contrast function returns.
#
# Parametric methods provided by MNE
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# - :func:`mne.stats.ttest_1samp_no_p`
# Paired t-test, optionally with hat adjustment.
# This is used by default for contrast enhancement in paired cluster tests.
#
# - :func:`mne.stats.f_oneway`
# One-way ANOVA for independent samples.
# This can be used to compute various F-contrasts. It is used by default
# for contrast enhancement in non-paired cluster tests.
#
# - :func:`mne.stats.f_mway_rm`
# M-way ANOVA for repeated measures and balanced designs.
# This returns F-statistics and p-values. The associated helper function
# :func:`mne.stats.f_threshold_mway_rm` can be used to determine the
# F-threshold at a given significance level.
#
# - :func:`mne.stats.linear_regression`
# Compute ordinary least square regressions on multiple targets, e.g.,
# sensors, time points across trials (samples).
# For each regressor it returns the beta value, t-statistic, and
# uncorrected p-value. While it can be used as a test, it is
# particularly useful to compute weighted averages or deal with
# continuous predictors.
#
# Non-parametric methods
# ^^^^^^^^^^^^^^^^^^^^^^
#
# - :func:`mne.stats.permutation_cluster_test`
# Unpaired contrasts with connectivity.
#
# - :func:`mne.stats.spatio_temporal_cluster_test`
# Unpaired contrasts with spatio-temporal connectivity.
#
# - :func:`mne.stats.permutation_t_test`
# Paired contrast with no connectivity.
#
# - :func:`mne.stats.permutation_cluster_1samp_test`
# Paired contrasts with connectivity.
#
# - :func:`mne.stats.spatio_temporal_cluster_1samp_test`
# Paired contrasts with spatio-temporal connectivity.
#
# <div class="alert alert-danger"><h4>Warning</h4><p>In most MNE functions, data has shape
# ``(..., n_space, n_time)``, where the spatial dimension can
# be e.g. sensors or source vertices. But for our spatio-temporal
# clustering functions, the spatial dimensions need to be **last**
# for computational efficiency reasons. For example, for
# :func:`mne.stats.spatio_temporal_cluster_1samp_test`, ``X``
# needs to be of shape ``(n_samples, n_time, n_space)``. You can
# use :func:`numpy.transpose` to transpose axes if necessary.</p></div>
#
# References
# ----------
# .. [1] Ridgway et al. 2012, "The problem of low variance voxels in
# statistical parametric mapping; a new hat avoids a 'haircut'",
# NeuroImage. 2012 Feb 1;59(3):2131-41.
#
# .. [2] <NAME> 2009, "Threshold-free cluster enhancement:
# addressing problems of smoothing, threshold dependence, and
# localisation in cluster inference", NeuroImage 44 (2009) 83-98.
#
# .. include:: ../../tutorial_links.inc
#
#
| stable/_downloads/846c9103e2de535e71d8b76a75654d36/plot_background_statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from rsna_retro.imports import *
from rsna_retro.metadata import *
from rsna_retro.preprocess import *
from rsna_retro.train import *
from rsna_retro.train3d import *
from rsna_retro.trainfull3d_labels import *
torch.cuda.set_device(3)
dls = get_3d_dls_aug(Meta.df_comb, sz=128, bs=32, grps=Meta.grps_stg1)
# ## Model
m = xres3d().cuda()
learn = get_learner(dls, m, get_loss())
learn.add_cb(RowLoss())
# +
# learn.load(f'runs/baseline_stg1_xresnet18-3', strict=False)
# -
name = 'trainfull3d_labels_full3d'
# ## Training
learn.lr_find()
do_fit(learn, 12, 1e-2)
learn.save(f'runs/{name}-1')
learn.load(f'runs/{name}-1')
learn.dls = get_3d_dls_aug(Meta.df_comb, sz=256, bs=12, grps=Meta.grps_stg1)
do_fit(learn, 12, 1e-3)
learn.save(f'runs/{name}-2')
learn.load(f'runs/{name}-2')
learn.dls = get_3d_dls_aug(Meta.df_comb, sz=384, bs=4, path=path_jpg, grps=Meta.grps_stg1)
do_fit(learn, 4, 1e-4)
learn.save(f'runs/{name}-3')
| 04_trainfull3d/04_trainfull3d_labels_02_full3d_old.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## CIFAR 10
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# +
import argparse
import os
import shutil
import time
from fastai.transforms import *
from fastai.dataset import *
from fastai.fp16 import *
from fastai.conv_learner import *
from pathlib import *
from fastai import io
import tarfile
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.distributed as dist
import torch.optim
import torch.utils.data
import torch.utils.data.distributed
import torchvision.transforms as transforms
import torchvision.datasets as datasets
import models
import models.cifar10 as cifar10models
from distributed import DistributedDataParallel as DDP
from datetime import datetime
# print(models.cifar10.__dict__)
model_names = sorted(name for name in models.__dict__
if name.islower() and not name.startswith("__")
and callable(models.__dict__[name]))
cifar10_names = sorted(name for name in cifar10models.__dict__
if name.islower() and not name.startswith("__")
and callable(cifar10models.__dict__[name]))
model_names = cifar10_names + model_names
# +
#print(models.cifar10.__dict__)
#print(model_names)
# +
# Example usage: python run_fastai.py /home/paperspace/ILSVRC/Data/CLS-LOC/ -a resnext_50_32x4d --epochs 1 -j 4 -b 64 --fp16
parser = argparse.ArgumentParser(description='PyTorch Cifar10 Training')
parser.add_argument('data', metavar='DIR',
help='path to dataset')
parser.add_argument('--save-dir', type=str, default=Path.home()/'imagenet_training',
help='Directory to save logs and models.')
parser.add_argument('--arch', '-a', metavar='ARCH', default='resnet56',
choices=model_names,
help='model architecture: ' +
' | '.join(model_names) +
' (default: resnet56)')
parser.add_argument('-dp', '--data-parallel', action='store_true', help='Use DataParallel')
parser.add_argument('-j', '--workers', default=7, type=int, metavar='N',
help='number of data loading workers (default: 4)')
parser.add_argument('--epochs', default=1, type=int, metavar='N',
help='number of total epochs to run')
parser.add_argument('--cycle-len', default=95, type=float, metavar='N',
help='Length of cycle to run')
# parser.add_argument('--start-epoch', default=0, type=int, metavar='N',
# help='manual epoch number (useful on restarts)')
parser.add_argument('-b', '--batch-size', default=512, type=int,
metavar='N', help='mini-batch size (default: 512)')
parser.add_argument('--lr', '--learning-rate', default=0.8, type=float,
metavar='LR', help='initial learning rate')
parser.add_argument('--momentum', default=0.9, type=float, metavar='M', help='momentum')
parser.add_argument('--weight-decay', '--wd', default=1e-4, type=float,
metavar='W', help='weight decay (default: 1e-4)')
# parser.add_argument('--print-freq', '-p', default=10, type=int,
# metavar='N', help='print frequency (default: 10)')
# parser.add_argument('--resume', default='', type=str, metavar='PATH',
# help='path to latest checkpoint (default: none)')
# parser.add_argument('-e', '--evaluate', dest='evaluate', action='store_true',
# help='evaluate model on validation set')
parser.add_argument('--pretrained', dest='pretrained', action='store_true', help='use pre-trained model')
parser.add_argument('--fp16', action='store_true', help='Run model fp16 mode.')
parser.add_argument('--use-tta', default=False, type=bool, help='Validate model with TTA at the end of traiing.')
parser.add_argument('--sz', default=32, type=int, help='Size of transformed image.')
# parser.add_argument('--decay-int', default=30, type=int, help='Decay LR by 10 every decay-int epochs')
parser.add_argument('--use-clr', default='10,13.68,0.95,0.85', type=str,
help='div,pct,max_mom,min_mom. Pass in a string delimited by commas. Ex: "20,2,0.95,0.85"')
parser.add_argument('--loss-scale', type=float, default=128,
help='Loss scaling, positive power of 2 values can improve fp16 convergence.')
parser.add_argument('--warmup', action='store_true', help='Do a warm-up epoch first')
parser.add_argument('--prof', dest='prof', action='store_true', help='Only run a few iters for profiling.')
parser.add_argument('--dist-url', default='file://sync.file', type=str,
help='url used to set up distributed training')
parser.add_argument('--dist-backend', default='nccl', type=str, help='distributed backend')
parser.add_argument('--world-size', default=1, type=int,
help='Number of GPUs to use. Can either be manually set ' +
'or automatically set by using \'python -m multiproc\'.')
parser.add_argument('--rank', default=0, type=int,
help='Used for multi-process training. Can either be manually set ' +
'or automatically set by using \'python -m multiproc\'.')
# +
def pad(img, p=4, padding_mode='reflect'):
return Image.fromarray(np.pad(np.asarray(img), ((p, p), (p, p), (0, 0)), padding_mode))
class TorchModelData(ModelData):
def __init__(self, path, sz, trn_dl, val_dl, aug_dl=None):
super().__init__(path, trn_dl, val_dl)
self.aug_dl = aug_dl
self.sz = sz
def download_cifar10(data_path):
# (AS) TODO: put this into the fastai library
def untar_file(file_path, save_path):
if file_path.endswith('.tar.gz') or file_path.endswith('.tgz'):
obj = tarfile.open(file_path)
obj.extractall(save_path)
obj.close()
os.remove(file_path)
cifar_url = 'http://files.fast.ai/data/cifar10.tgz' # faster download
# cifar_url = 'http://pjreddie.com/media/files/cifar.tgz'
io.get_data(cifar_url, args.data+'/cifar10.tgz')
untar_file(data_path+'/cifar10.tgz', data_path)
# Loader expects train and test folders to be outside of cifar10 folder
shutil.move(data_path+'/cifar10/train', data_path)
shutil.move(data_path+'/cifar10/test', data_path)
def torch_loader(data_path, size):
if not os.path.exists(data_path+'/train'): download_cifar10(data_path)
# Data loading code
traindir = os.path.join(data_path, 'train')
valdir = os.path.join(data_path, 'test')
normalize = transforms.Normalize(mean=[0.4914 , 0.48216, 0.44653], std=[0.24703, 0.24349, 0.26159])
tfms = [transforms.ToTensor(), normalize]
scale_size = 40
padding = int((scale_size - size) / 2)
train_tfms = transforms.Compose([
pad, # TODO: use `padding` rather than assuming 4
transforms.RandomCrop(size),
transforms.ColorJitter(.25,.25,.25),
transforms.RandomRotation(2),
transforms.RandomHorizontalFlip(),
] + tfms)
val_tfms = transforms.Compose(tfms)
train_dataset = datasets.ImageFolder(traindir, train_tfms)
val_dataset = datasets.ImageFolder(valdir, val_tfms)
train_sampler = (torch.utils.data.distributed.DistributedSampler(train_dataset) if args.distributed else None)
val_sampler = (torch.utils.data.distributed.DistributedSampler(val_dataset) if args.distributed else None)
train_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=args.batch_size, shuffle=(train_sampler is None),
num_workers=args.workers, pin_memory=True, sampler=train_sampler)
val_loader = torch.utils.data.DataLoader(
val_dataset, batch_size=args.batch_size*2, shuffle=False,
num_workers=args.workers, pin_memory=True, sampler=val_sampler)
aug_loader = torch.utils.data.DataLoader(
datasets.ImageFolder(valdir, train_tfms),
batch_size=args.batch_size, shuffle=False,
num_workers=args.workers, pin_memory=True)
train_loader = DataPrefetcher(train_loader)
val_loader = DataPrefetcher(val_loader)
aug_loader = DataPrefetcher(aug_loader)
if args.prof:
train_loader.stop_after = 200
val_loader.stop_after = 0
data = TorchModelData(data_path, args.sz, train_loader, val_loader, aug_loader)
if train_sampler:
data.trn_sampler = train_sampler
data.val_sampler = val_sampler
return data
# Seems to speed up training by ~2%
class DataPrefetcher():
def __init__(self, loader, stop_after=None):
self.loader = loader
self.dataset = loader.dataset
self.stream = torch.cuda.Stream()
self.stop_after = stop_after
self.next_input = None
self.next_target = None
def __len__(self):
return len(self.loader)
def preload(self):
try:
self.next_input, self.next_target = next(self.loaditer)
except StopIteration:
self.next_input = None
self.next_target = None
return
with torch.cuda.stream(self.stream):
self.next_input = self.next_input.cuda(async=True)
self.next_target = self.next_target.cuda(async=True)
def __iter__(self):
count = 0
self.loaditer = iter(self.loader)
self.preload()
while self.next_input is not None:
torch.cuda.current_stream().wait_stream(self.stream)
input = self.next_input
target = self.next_target
self.preload()
count += 1
yield input, target
if type(self.stop_after) is int and (count > self.stop_after):
break
# +
def top5(output, target):
"""Computes the precision@k for the specified values of k"""
top5 = 5
batch_size = target.size(0)
_, pred = output.topk(top5, 1, True, True)
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
correct_k = correct[:top5].view(-1).float().sum(0, keepdim=True)
return correct_k.mul_(1.0 / batch_size)
class ImagenetLoggingCallback(Callback):
start_time = 0
def __init__(self, save_path, print_every=50):
super().__init__()
self.save_path=save_path
self.print_every=print_every
self.start_time = datetime.now()
def on_train_begin(self):
self.batch = 0
self.epoch = 0
self.f = open(self.save_path, "a", 1)
self.log("epoch\thours\ttop1Accuracy")
def on_epoch_end(self, metrics):
current_time = datetime.now()
time_diff = current_time - self.start_time
log_str = f'{self.epoch}\t{float(time_diff.total_seconds() / 3600.0)}\t{100 * metrics[1]}'
#for (k,v) in zip(['val_loss', 'acc'], metrics): if k=='acc': log_str += f'\t{k}:{v}'
self.log(log_str)
self.epoch += 1
def on_batch_end(self, metrics):
self.last_loss = metrics
self.batch += 1
#if self.batch % self.print_every == 0:
#self.log(f'Epoch: {self.epoch} Batch: {self.batch} Metrics: {metrics}')
def on_train_end(self): self.f.close()
def log(self, string): self.f.write(string+"\n")
# Logging + saving models
def save_args(name, save_dir):
if (args.rank != 0) or not args.save_dir: return {}
log_dir = f'{save_dir}/training_logs'
os.makedirs(log_dir, exist_ok=True)
return {
'best_save_name': f'{name}_best_model',
'cycle_save_name': f'{name}',
'callbacks': [
ImagenetLoggingCallback(f'{log_dir}/{name}_log.txt')
]
}
def save_sched(sched, save_dir):
if (args.rank != 0) or not args.save_dir: return {}
log_dir = f'{save_dir}/training_logs'
sched.save_path = log_dir
sched.plot_loss()
sched.plot_lr()
def update_model_dir(learner, base_dir):
learner.tmp_path = f'{base_dir}/tmp'
os.makedirs(learner.tmp_path, exist_ok=True)
learner.models_path = f'{base_dir}/models'
os.makedirs(learner.models_path, exist_ok=True)
# -
# ### Configuration
args_input = [
'/home/paperspace/data/cifar10',
'--save-dir', '/home/paperspace/data/cifar_training/model_merge',
'-a', 'wrn_22',
'-b', '512',
'--loss-scale', '512',
'--fp16',
'--cycle-len', '30',
'--use-clr', '20,20,0.95,0.85',
'--wd', '1e-4',
'--lr', '1.5',
# '--train-half' # With fp16, iterations are so fast this doesn't matter
]
# This is important for speed
cudnn.benchmark = True
global arg
args = parser.parse_args(args_input); args
if args.cycle_len > 1: args.cycle_len = int(args.cycle_len)
# +
args.distributed = args.world_size > 1
args.gpu = 0
if args.distributed: args.gpu = args.rank % torch.cuda.device_count()
if args.distributed:
torch.cuda.set_device(args.gpu)
dist.init_process_group(backend=args.dist_backend, init_method=args.dist_url,
world_size=args.world_size)
if args.fp16: assert torch.backends.cudnn.enabled, "missing cudnn"
# -
model = cifar10models.__dict__[args.arch] if args.arch in cifar10_names else models.__dict__[args.arch]
if args.pretrained: model = model(pretrained=True)
else: model = model()
model = model.cuda()
if args.distributed: model = DDP(model)
if args.data_parallel:
n_dev = 4
model = nn.DataParallel(model, range(n_dev))
args.batch_size *= n_dev
# +
data = torch_loader(args.data, args.sz)
learner = Learner.from_model_data(model, data)
learner.crit = F.cross_entropy
learner.metrics = [accuracy]
if args.fp16: learner.half()
if args.prof: args.epochs,args.cycle_len = 1,0.01
if args.use_clr: args.use_clr = tuple(map(float, args.use_clr.split(',')))
# +
# Full size
update_model_dir(learner, args.save_dir)
sargs = save_args('first_run', args.save_dir)
if args.warmup:
learner.fit(args.lr/10, 1, cycle_len=1, wds=args.weight_decay,
use_clr_beta=(100,1,0.9,0.8), loss_scale=args.loss_scale, **sargs)
learner.fit(args.lr,args.epochs, cycle_len=args.cycle_len,
wds=args.weight_decay,
use_clr_beta=args.use_clr, loss_scale=args.loss_scale,
**sargs)
save_sched(learner.sched, args.save_dir)
# -
learner.save('wrn_22_dawn')
# +
# learner.get_layer_groups()
# -
m1 = learner.model
arch2 = 'wrn_28'
m2 = cifar10models.__dict__[arch2] if arch2 in cifar10_names else models.__dict__[arch2]
m2 = m2()
sd1 = m1.state_dict()
sd2 = m2.state_dict()
sd2.update(sd1)
m2.load_state_dict(sd2)
m1
m2
new_params = set(sd2) - set(sd1); new_params
for k,v in m2.named_parameters():
if k in sd1:
v.required_grad=False
if k in new_params:
if k.
m2 = m2.cuda()
learner2 = Learner.from_model_data(m2, data)
learner2.crit = F.cross_entropy
learner2.metrics = [accuracy]
if args.fp16: learner2.half()
# +
# Full size
update_model_dir(learner2, args.save_dir)
sargs = save_args('depth28', args.save_dir)
learner2.fit(args.lr,args.epochs, cycle_len=args.cycle_len,
wds=args.weight_decay,
use_clr_beta=args.use_clr, loss_scale=args.loss_scale,
**sargs)
save_sched(learner2.sched, args.save_dir)
# -
| cifar10/cifar10-double-resnet-size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/evansekeful/syncasanacanvas/blob/main/AsanaCanvasSync.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5QrQXz_RSKRD"
# ## Setup Tasks
# + id="-i3MGTPJDTkg"
# install required packages
# %pip install asana
# %pip install icalendar
# + id="AxmCCPZMEoEJ"
# import libraries
import requests
import json
import pandas as pd
import asana
from icalendar import Calendar, Event
from datetime import datetime
from datetime import timezone
from pytz import all_timezones
import pytz
# + id="klUVmLpyZQOZ"
# TODO prompt users to upload config.json
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="bBsMiCU2HX9E"
# ## Attention!
#
#
# ---
#
#
# It is important that you map the location of the config.json below and that both packages above have been installed correctly before moving to the next step.
# + id="8hRvp_hAbhnN"
# read config variables into environment
config_path = "/content/drive/MyDrive/Colab Notebooks/config.json"
with open(config_path) as json_file:
config = json.load(json_file)
# TODO validate config.json schema
canvas = config["canvas"]["url"]
token = config["asana"]["token"]
project = config["asana"]["project_gid"]
assignee = config["asana"]["assignee"]
workspace = config["asana"]["workspace"]
# + id="6t3_G5Dr6nrJ"
# set up Asana client
client = asana.Client.access_token(token)
# + [markdown] id="BVbRiyyQ6H4H"
# ## Run Extract + Transform
# + [markdown] id="8iiiWBh_SRwF"
# Read Canvas Calendar
# + id="c_c7NK75xyBI"
# set up events dictionary
events = {k:[] for k in ["uid","name","start","end"]}
# + id="1Eg6_QsdErio"
# import Canvas ics
cal = Calendar.from_ical(requests.get(canvas).text)
# set timezone to standardize timeawareness
utc = pytz.utc
# read ics into dictionary of events
for comp in cal.walk():
if comp.get("UID") == None: continue # skip blank items
if comp.get("SUMMARY") == None: continue # skip blank items
if comp.get("UID").startswith("event-assignment"):
events["uid"].append(str(comp.get("UID")))
else: continue # skip events that are not assignments
events["name"].append(str(comp.get("SUMMARY")))
if hasattr(comp.get("dtstart"), "dt"):
if comp.get("dtstart").dt.tzname() == None:
events["start"].append(utc.localize(comp.get("dtstart").dt))
else:
events["start"].append(comp.get("dtstart").dt)
if hasattr(comp.get("dtend"), "dt"):
if comp.get("dtend").dt.tzname() == None:
events["end"].append(utc.localize(comp.get("dtend").dt))
else:
events["end"].append(comp.get("dtend").dt)
# + id="55tu4cp1jq4B"
# create extract dataframe
extract = pd.DataFrame(events)
# + [markdown] id="xHCZIbbsSufC"
# Read Project + Clean Canvas Data
# + id="DOXQPgrbS9As"
# set up task list
tasks = []
# set up Asana client
homework = {"project": project}
options = "notes"
result = client.tasks.get_tasks(homework,opt_fields=options)
# read homework project
for task in result:
tasks.append(task["notes"])
# + id="zAU28tZm383i"
# delete duplicates from extract table
duplicates = tuple(tasks)
extract = extract[~extract["uid"].isin(duplicates)]
# + id="jKj-HTo2DK2k"
#TODO create condition to update due dates
# + id="ctGkVUJFVCUw"
# delete past due entries
extract = extract[extract["end"] > datetime.now(tz=timezone.utc)]
# + id="deVPuwTkgGeL"
# delete unused columns
extract = extract[["name","uid","start","end"]]
# + id="bnFx0ql2pxFl"
# set timezone to localize due dates TODO: add to config file
extract["start"] = extract["start"].dt.tz_convert('US/Pacific')
extract["end"] = extract["end"].dt.tz_convert('US/Pacific')
# + [markdown] id="7rT2j9n_Sar6"
# Format Calendar into JSON
# + id="44cnfhB_P_o0"
# iterate over rows to create list of dictionaries
load = []
for index, row in extract.iterrows():
temp = {}
temp["assignee"] = assignee
temp["due_on"] = row["end"].strftime("%Y-%m-%d")
temp["name"] = row["name"]
temp["notes"] = row["uid"]
temp["projects"] = [project]
temp["resource_subtype"] = "default_task"
#temp["start_on"] = row["start"].strftime("%Y-%m-%d") # paid feature
temp["workspace"] = workspace
load.append(temp)
# + id="z1lyPbbCYPfe"
# set up validation list
validate = []
for task in load:
validate.append(task["notes"])
# + [markdown] id="EU7_4tmSWI6n"
# ## Check Data
#
# ---
#
# For running as an automation, be sure to wrap the load tasks with the following check.
# + id="XGelTLKRW14I"
len(load) > 0
# + [markdown] id="Ork28sbCXH-G"
# ## Run Load
# + id="K5rb96osDZUc"
#TODO split new vs update
# + [markdown] id="PKiVdi1HTIW7"
# Post New Assignments to Asana
# + id="I6zP936FE5YR"
# post new assignments to Asana
for data in load:
result = client.tasks.create_task(data)
# + [markdown] id="E3c7J9nMTUMO"
# Validate New Assignments Posted
# + id="PQRPaUKCFF0R"
# set up task list
tasks = []
# read homework project
result = client.tasks.get_tasks(homework,opt_fields=options)
for task in result:
tasks.append(task["notes"])
# compare list with validation
test = set(tasks).intersection(validate)
len(validate) == len(test)
# + id="IvBWGR_6dSn1"
# TODO print missing assignment ids to file
| AsanaCanvasSync.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.121193, "end_time": "2021-03-25T14:49:37.871225", "exception": false, "start_time": "2021-03-25T14:49:37.750032", "status": "completed"} tags=[]
# ## Summary => <br>
# This notebook includes the following topics. <br><br>
#
# The notebook will be constructed in two stages. <br>
# * 1st Stage -> Complete python implementations along with brief descriptions. (Est. Date of Completion - 28-03-2021)
# * 2nd Stage -> Solving questions on these topics using python. (Est. Date of Completion - 10-03-2021)
# + [markdown] papermill={"duration": 0.12668, "end_time": "2021-03-25T14:49:38.115755", "exception": false, "start_time": "2021-03-25T14:49:37.989075", "status": "completed"} tags=[]
# ## Table of Contents
#
# * Understanding Data types
# * Interval Scale
# * Binary
# * Categorical
# * Ordinal
# * Ratio Scaled
# * Mixed Type
# * Different types of distances
# * Simmilarity and Dissimilarity Matrix
# * Handling Missing data values
# * Central Tendency & Dispersion
# * Descriptive Statistics
# * [Sample](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#16.-Sample-Statistics) vs [Population statistics](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#17.-Population-Statistics)
# * Random Variables
# * Probability Distribution Function
# * [Measuring p-value](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#13.-Calculating-p-Value)
# * [Measuring Correlation](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#14.-Measuring-Correlation)
# * [Measuring Variance](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#15.-Measuring-Variance)
# * Expected Value
# * [Binomial Distribution](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#8.-Binomial-Distribution)
# * [Normal Distributions](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#9.-Normal-Distribution)
# * [Poisson Distributions](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#10.-Poisson-Distribution)
# * [Bernoulli Distribution](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#11.-Bernoulli-Distribution)
# * [z-score](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#5.-Z-Test)
# * Hypothesis Testing
# * Null & Alternate Hypothesis
# * Type 1 Error; Type 2 Error
# * Various Approaches
# * p-value
# * critical value
# * confidence interval value
# * z-stats vs t-stats
#
# * Two Sample Tests
# * Confidence Interval
# * Similarity & Dissimilarity Matrices
# * [Central Limit Theorem](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#12.-Central-Limit-Theorem)
# * [Chi Square Test](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#3.-Chi-Square-Test)
# * [T Test](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#4.-T-Test)
# * [ANOVA Test](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#6.-ANOVA-Test)
# * [One Way Anova Test](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#6.1-One-Way-ANOVA-Test)
# * F Test (LSD Test)
# * Tukey Kramer Test
# * [Two Way Anova Test](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#6.2-Two-Way-ANOVA-Test)
# * Interaction Effects
# * [F Stats](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#7.-F-Stats-Test)
# * [Regressions (Linear, Multiple) + ROC](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#2.-Regressions)
# * Logistic Regression
# * Python Implementation
# * Calculating G Statistics
# * Residual Analysis
# * Maximum Likelihood Estimation
# * Cluster Analysis
# * Partitioning Cluster Methods
# * K-Means
# * K Mediods
# * Hierarchial Cluster Methods
# * Agglomerative
# * [CART Algorithms](https://www.kaggle.com/antoreepjana/statistics-for-ml-data-analysis/#1.-CART-Algorithms)
# * Python Implementation
# * various Calculations involved
# * Information Gain
# * Gain Ratio
# * Gini Index
# * Confusion Metrics, ROC & Regression Analysis
# + papermill={"duration": 0.114088, "end_time": "2021-03-25T14:49:38.346151", "exception": false, "start_time": "2021-03-25T14:49:38.232063", "status": "completed"} tags=[]
# + papermill={"duration": 0.116777, "end_time": "2021-03-25T14:49:38.577091", "exception": false, "start_time": "2021-03-25T14:49:38.460314", "status": "completed"} tags=[]
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.133579, "end_time": "2021-03-25T14:49:38.836472", "exception": false, "start_time": "2021-03-25T14:49:38.702893", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import os
import random as rnd
# + papermill={"duration": 1.314954, "end_time": "2021-03-25T14:49:40.267307", "exception": false, "start_time": "2021-03-25T14:49:38.952353", "status": "completed"} tags=[]
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# + [markdown] papermill={"duration": 0.111519, "end_time": "2021-03-25T14:49:40.494896", "exception": false, "start_time": "2021-03-25T14:49:40.383377", "status": "completed"} tags=[]
# ### 1. CART Algorithms
# + [markdown] papermill={"duration": 0.113988, "end_time": "2021-03-25T14:49:40.720460", "exception": false, "start_time": "2021-03-25T14:49:40.606472", "status": "completed"} tags=[]
# Brief Description ->
# + [markdown] papermill={"duration": 0.112846, "end_time": "2021-03-25T14:49:40.952249", "exception": false, "start_time": "2021-03-25T14:49:40.839403", "status": "completed"} tags=[]
# ##### Tools Used
#
# Dataset Used -> Boston Dataset (UCI Machine Learning Repository)
# + papermill={"duration": 0.629537, "end_time": "2021-03-25T14:49:41.698647", "exception": false, "start_time": "2021-03-25T14:49:41.069110", "status": "completed"} tags=[]
from sklearn.datasets import load_boston
boston_dataset = load_boston()
# + papermill={"duration": 0.129119, "end_time": "2021-03-25T14:49:41.944306", "exception": false, "start_time": "2021-03-25T14:49:41.815187", "status": "completed"} tags=[]
boston = pd.DataFrame(boston_dataset.data, columns = boston_dataset.feature_names)
# + papermill={"duration": 0.180957, "end_time": "2021-03-25T14:49:42.247142", "exception": false, "start_time": "2021-03-25T14:49:42.066185", "status": "completed"} tags=[]
boston.head()
# + papermill={"duration": 0.125974, "end_time": "2021-03-25T14:49:42.488222", "exception": false, "start_time": "2021-03-25T14:49:42.362248", "status": "completed"} tags=[]
boston['MEDV'] = boston_dataset.target
# + papermill={"duration": 0.122964, "end_time": "2021-03-25T14:49:42.724202", "exception": false, "start_time": "2021-03-25T14:49:42.601238", "status": "completed"} tags=[]
names = boston_dataset.feature_names
# + papermill={"duration": 0.477656, "end_time": "2021-03-25T14:49:43.318777", "exception": false, "start_time": "2021-03-25T14:49:42.841121", "status": "completed"} tags=[]
from sklearn.tree import DecisionTreeRegressor
# + papermill={"duration": 0.12613, "end_time": "2021-03-25T14:49:43.562653", "exception": false, "start_time": "2021-03-25T14:49:43.436523", "status": "completed"} tags=[]
array = boston.values
X = array[:, 0:13]
Y = array[:, 13]
# + papermill={"duration": 0.139088, "end_time": "2021-03-25T14:49:43.816118", "exception": false, "start_time": "2021-03-25T14:49:43.677030", "status": "completed"} tags=[]
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X,Y, test_size = 0.3, random_state = 1234)
# + papermill={"duration": 0.135304, "end_time": "2021-03-25T14:49:44.070288", "exception": false, "start_time": "2021-03-25T14:49:43.934984", "status": "completed"} tags=[]
model = DecisionTreeRegressor(max_leaf_nodes = 20)
# + papermill={"duration": 0.136381, "end_time": "2021-03-25T14:49:44.326557", "exception": false, "start_time": "2021-03-25T14:49:44.190176", "status": "completed"} tags=[]
model.fit(X_train, Y_train)
# + papermill={"duration": 0.132755, "end_time": "2021-03-25T14:49:44.575393", "exception": false, "start_time": "2021-03-25T14:49:44.442638", "status": "completed"} tags=[]
from sklearn.metrics import r2_score
# + papermill={"duration": 0.131001, "end_time": "2021-03-25T14:49:44.826852", "exception": false, "start_time": "2021-03-25T14:49:44.695851", "status": "completed"} tags=[]
YHat = model.predict(X_test)
# + papermill={"duration": 0.130999, "end_time": "2021-03-25T14:49:45.079444", "exception": false, "start_time": "2021-03-25T14:49:44.948445", "status": "completed"} tags=[]
r2 = r2_score(Y_test, YHat)
print("R2 Score -> ", r2)
# + [markdown] papermill={"duration": 0.11655, "end_time": "2021-03-25T14:49:45.316234", "exception": false, "start_time": "2021-03-25T14:49:45.199684", "status": "completed"} tags=[]
# ### plot the decision tree as a graph
# + papermill={"duration": 0.150426, "end_time": "2021-03-25T14:49:45.585320", "exception": false, "start_time": "2021-03-25T14:49:45.434894", "status": "completed"} tags=[]
import graphviz
from sklearn import tree
# + [markdown] papermill={"duration": 0.12147, "end_time": "2021-03-25T14:49:45.828883", "exception": false, "start_time": "2021-03-25T14:49:45.707413", "status": "completed"} tags=[]
# method 1
# + papermill={"duration": 2.045958, "end_time": "2021-03-25T14:49:47.992370", "exception": false, "start_time": "2021-03-25T14:49:45.946412", "status": "completed"} tags=[]
fig = plt.figure(figsize=(25,20))
_ = tree.plot_tree(model,
feature_names=names,
class_names=boston_dataset.target,
filled=True)
# + [markdown] papermill={"duration": 0.12984, "end_time": "2021-03-25T14:49:48.247511", "exception": false, "start_time": "2021-03-25T14:49:48.117671", "status": "completed"} tags=[]
# method 2
# + papermill={"duration": 0.784986, "end_time": "2021-03-25T14:49:49.155313", "exception": false, "start_time": "2021-03-25T14:49:48.370327", "status": "completed"} tags=[]
plt.figure(figsize = (20,20))
dot_data = tree.export_graphviz(model, out_file=None,
feature_names=names,
class_names=boston_dataset.target,
filled=True, rounded= True)
# Draw graph
graph = graphviz.Source(dot_data, format="png")
graph
# + [markdown] papermill={"duration": 0.123843, "end_time": "2021-03-25T14:49:49.476917", "exception": false, "start_time": "2021-03-25T14:49:49.353074", "status": "completed"} tags=[]
# We'll learn how to custom paint your graph from the default settings (coming soon)
# + papermill={"duration": 0.126376, "end_time": "2021-03-25T14:49:49.728507", "exception": false, "start_time": "2021-03-25T14:49:49.602131", "status": "completed"} tags=[]
# + papermill={"duration": 0.141625, "end_time": "2021-03-25T14:49:49.999951", "exception": false, "start_time": "2021-03-25T14:49:49.858326", "status": "completed"} tags=[]
"""import pydotplus
graph = pydotplus.graph_from_dot_data(dot_data)
nodes = graph.get_node_list()
for node in nodes:
if node.get_label():
print(node.get_label())
node.set_fillcolor('yellow')
graph.write_png('colored_tree.png')
"""
# + papermill={"duration": 0.153491, "end_time": "2021-03-25T14:49:50.282208", "exception": false, "start_time": "2021-03-25T14:49:50.128717", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.127446, "end_time": "2021-03-25T14:49:50.547926", "exception": false, "start_time": "2021-03-25T14:49:50.420480", "status": "completed"} tags=[]
# ### 2. Regressions
# + [markdown] papermill={"duration": 0.128826, "end_time": "2021-03-25T14:49:50.801274", "exception": false, "start_time": "2021-03-25T14:49:50.672448", "status": "completed"} tags=[]
# Useful Resources -> <br>
#
# * https://www.maths.usyd.edu.au/u/UG/SM/STAT3022/r/current/Lecture/lecture03_2020JC.html#1
# * https://towardsdatascience.com/maximum-likelihood-estimation-explained-normal-distribution-6207b322e47f#:~:text=%E2%80%9CA%20method%20of%20estimating%20the,observed%20data%20is%20most%20probable.%E2%80%9D&text=By%20assuming%20normality%2C%20we%20simply,the%20popular%20Gaussian%20bell%20curve.
# * https://online.stat.psu.edu/stat462/node/207/
# * https://psychscenehub.com/psychpedia/odds-ratio-2/
# * http://statkat.com/stat-tests/logistic-regression.php#:~:text=Logistic%20regression%20analysis%20tests%20the,%3D%CE%B2K%3D0
# + [markdown] papermill={"duration": 0.129085, "end_time": "2021-03-25T14:49:51.059776", "exception": false, "start_time": "2021-03-25T14:49:50.930691", "status": "completed"} tags=[]
# 1. Linear Regression Analysis
# + papermill={"duration": 0.124892, "end_time": "2021-03-25T14:49:51.310078", "exception": false, "start_time": "2021-03-25T14:49:51.185186", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.130425, "end_time": "2021-03-25T14:49:51.565923", "exception": false, "start_time": "2021-03-25T14:49:51.435498", "status": "completed"} tags=[]
# 2. Multiple Regression Analysis
# + papermill={"duration": 0.126574, "end_time": "2021-03-25T14:49:51.821750", "exception": false, "start_time": "2021-03-25T14:49:51.695176", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.13649, "end_time": "2021-03-25T14:49:52.088680", "exception": false, "start_time": "2021-03-25T14:49:51.952190", "status": "completed"} tags=[]
# ### 3. Chi Square Test
# + [markdown] papermill={"duration": 0.126273, "end_time": "2021-03-25T14:49:52.346061", "exception": false, "start_time": "2021-03-25T14:49:52.219788", "status": "completed"} tags=[]
# background ->
# + [markdown] papermill={"duration": 0.124694, "end_time": "2021-03-25T14:49:52.595349", "exception": false, "start_time": "2021-03-25T14:49:52.470655", "status": "completed"} tags=[]
# degrees of freedom for the chi-squared distribution -> (rows -1) * (cols -1)
# + [markdown] papermill={"duration": 0.132542, "end_time": "2021-03-25T14:49:52.855836", "exception": false, "start_time": "2021-03-25T14:49:52.723294", "status": "completed"} tags=[]
# a. Understanding Contigency Tables (also known as crosstab)
# + [markdown] papermill={"duration": 0.128638, "end_time": "2021-03-25T14:49:53.113257", "exception": false, "start_time": "2021-03-25T14:49:52.984619", "status": "completed"} tags=[]
# Contigency tables are the pivot tables obtained by utilizing the categorical variable. The contigency here is whether a variable affects the values of the caegorical variable. <br>
#
# + papermill={"duration": 0.130674, "end_time": "2021-03-25T14:49:53.380491", "exception": false, "start_time": "2021-03-25T14:49:53.249817", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.125892, "end_time": "2021-03-25T14:49:53.633761", "exception": false, "start_time": "2021-03-25T14:49:53.507869", "status": "completed"} tags=[]
# b. Performing Chi-Square Tests
# + papermill={"duration": 0.127877, "end_time": "2021-03-25T14:49:53.889054", "exception": false, "start_time": "2021-03-25T14:49:53.761177", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.126769, "end_time": "2021-03-25T14:49:54.144709", "exception": false, "start_time": "2021-03-25T14:49:54.017940", "status": "completed"} tags=[]
# c. Chi-Square Tests for Feature Selection
# + [markdown] papermill={"duration": 0.125722, "end_time": "2021-03-25T14:49:54.397106", "exception": false, "start_time": "2021-03-25T14:49:54.271384", "status": "completed"} tags=[]
# 
# + [markdown] papermill={"duration": 0.12594, "end_time": "2021-03-25T14:49:54.648355", "exception": false, "start_time": "2021-03-25T14:49:54.522415", "status": "completed"} tags=[]
# #### Note:- Used only for Categorical Features.
# + [markdown] papermill={"duration": 0.127215, "end_time": "2021-03-25T14:49:54.901403", "exception": false, "start_time": "2021-03-25T14:49:54.774188", "status": "completed"} tags=[]
# Dataset used -> https://www.kaggle.com/c/cat-in-the-dat
# + papermill={"duration": 2.515862, "end_time": "2021-03-25T14:49:57.543995", "exception": false, "start_time": "2021-03-25T14:49:55.028133", "status": "completed"} tags=[]
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
data = pd.read_csv('../input/cat-in-the-dat/train.csv')
# + papermill={"duration": 0.16604, "end_time": "2021-03-25T14:49:57.835517", "exception": false, "start_time": "2021-03-25T14:49:57.669477", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.201549, "end_time": "2021-03-25T14:49:58.164705", "exception": false, "start_time": "2021-03-25T14:49:57.963156", "status": "completed"} tags=[]
data.drop(['id'], axis = 1, inplace = True)
# + papermill={"duration": 0.144321, "end_time": "2021-03-25T14:49:58.435441", "exception": false, "start_time": "2021-03-25T14:49:58.291120", "status": "completed"} tags=[]
data.dtypes
# + papermill={"duration": 1.210199, "end_time": "2021-03-25T14:49:59.772193", "exception": false, "start_time": "2021-03-25T14:49:58.561994", "status": "completed"} tags=[]
for col in data.columns:
print(col, data[col].nunique())
# + papermill={"duration": 1.527259, "end_time": "2021-03-25T14:50:01.430075", "exception": false, "start_time": "2021-03-25T14:49:59.902816", "status": "completed"} tags=[]
for col in data.columns:
print(col, '\n\n',data[col].value_counts())
print('-'*10)
# + [markdown] papermill={"duration": 0.131437, "end_time": "2021-03-25T14:50:01.694149", "exception": false, "start_time": "2021-03-25T14:50:01.562712", "status": "completed"} tags=[]
# bin_3, bin_4 has T/F values. <br>
# nom_0, nom_1, nom_2, nom_3, nom_4 have 3-6 unique values. <br>
# nom_5, nom_6, nom_7, nom_8, nom_9 have many unique values <br>
# THen comes the ordinal variables
# + papermill={"duration": 0.130864, "end_time": "2021-03-25T14:50:01.957124", "exception": false, "start_time": "2021-03-25T14:50:01.826260", "status": "completed"} tags=[]
# + papermill={"duration": 0.364058, "end_time": "2021-03-25T14:50:02.463715", "exception": false, "start_time": "2021-03-25T14:50:02.099657", "status": "completed"} tags=[]
data['bin_3'] = data['bin_3'].map({"T" : 1, "F" : 0})
data['bin_4'] = data['bin_4'].map({"Y" : 1, "N" : 0})
# + papermill={"duration": 0.17292, "end_time": "2021-03-25T14:50:02.775870", "exception": false, "start_time": "2021-03-25T14:50:02.602950", "status": "completed"} tags=[]
data.head()
# + [markdown] papermill={"duration": 0.135858, "end_time": "2021-03-25T14:50:03.046201", "exception": false, "start_time": "2021-03-25T14:50:02.910343", "status": "completed"} tags=[]
# We're done with dealing of binary variables. <br>
# Now we're left to deal with the nominals & ordinals.
# + papermill={"duration": 0.133141, "end_time": "2021-03-25T14:50:03.313175", "exception": false, "start_time": "2021-03-25T14:50:03.180034", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.132527, "end_time": "2021-03-25T14:50:03.581581", "exception": false, "start_time": "2021-03-25T14:50:03.449054", "status": "completed"} tags=[]
# We have 5 ordinal variables of which 4 have few unique values and can be dealt in a similar manner. <br>
# ord_5 has multiple unique values and needs to be handled separately.
# + papermill={"duration": 1.794114, "end_time": "2021-03-25T14:50:05.508159", "exception": false, "start_time": "2021-03-25T14:50:03.714045", "status": "completed"} tags=[]
for col in ['ord_1', 'ord_2', 'ord_3', 'ord_4']:
print(col, list(np.unique(data[col])))
# + papermill={"duration": 0.254726, "end_time": "2021-03-25T14:50:05.900783", "exception": false, "start_time": "2021-03-25T14:50:05.646057", "status": "completed"} tags=[]
m1_ord1 = {'Novice' : 0, 'Contributor' : 1, 'Expert' : 2, 'Master' : 3, 'Grandmaster' : 4}
data['ord_1'] = data['ord_1'].map(m1_ord1)
# + papermill={"duration": 0.176128, "end_time": "2021-03-25T14:50:06.219320", "exception": false, "start_time": "2021-03-25T14:50:06.043192", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.241618, "end_time": "2021-03-25T14:50:06.597422", "exception": false, "start_time": "2021-03-25T14:50:06.355804", "status": "completed"} tags=[]
m2_ord2 = {'Boiling Hot' : 0, 'Cold' : 1, 'Freezing' : 2, 'Hot' : 3, 'Lava Hot' : 4, 'Warm' : 5}
data['ord_2'] = data['ord_2'].map(m2_ord2)
# + papermill={"duration": 0.172314, "end_time": "2021-03-25T14:50:06.917167", "exception": false, "start_time": "2021-03-25T14:50:06.744853", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.143811, "end_time": "2021-03-25T14:50:07.218397", "exception": false, "start_time": "2021-03-25T14:50:07.074586", "status": "completed"} tags=[]
# + papermill={"duration": 0.729978, "end_time": "2021-03-25T14:50:08.082811", "exception": false, "start_time": "2021-03-25T14:50:07.352833", "status": "completed"} tags=[]
data['ord_3'] = data['ord_3'].apply(lambda x : ord(x) - ord('a'))
data['ord_4'] = data['ord_4'].apply(lambda x : ord(x) - ord('A'))
# + papermill={"duration": 0.170352, "end_time": "2021-03-25T14:50:08.392283", "exception": false, "start_time": "2021-03-25T14:50:08.221931", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 1.447318, "end_time": "2021-03-25T14:50:09.977622", "exception": false, "start_time": "2021-03-25T14:50:08.530304", "status": "completed"} tags=[]
data['ord_5a'] = data['ord_5'].str[0]
data['ord_5b'] = data['ord_5'].str[1]
data['ord_5a'] = data['ord_5a'].map({val : idx for idx, val in enumerate(np.unique(data['ord_5a']))})
data['ord_5b'] = data['ord_5b'].map({val : idx for idx, val in enumerate(np.unique(data['ord_5b']))})
# + papermill={"duration": 0.169551, "end_time": "2021-03-25T14:50:10.285989", "exception": false, "start_time": "2021-03-25T14:50:10.116438", "status": "completed"} tags=[]
data.head()
# + [markdown] papermill={"duration": 0.138753, "end_time": "2021-03-25T14:50:10.561704", "exception": false, "start_time": "2021-03-25T14:50:10.422951", "status": "completed"} tags=[]
# Let's deal the nominal variables.
# + papermill={"duration": 0.252937, "end_time": "2021-03-25T14:50:10.956788", "exception": false, "start_time": "2021-03-25T14:50:10.703851", "status": "completed"} tags=[]
data[['nom_0', 'nom_2', 'nom_3', 'nom_4', 'nom_5', 'nom_6', 'nom_7', 'nom_8', 'nom_9']]
# + papermill={"duration": 0.236142, "end_time": "2021-03-25T14:50:11.344476", "exception": false, "start_time": "2021-03-25T14:50:11.108334", "status": "completed"} tags=[]
data['nom_1'].value_counts()
# + papermill={"duration": 0.233586, "end_time": "2021-03-25T14:50:11.782330", "exception": false, "start_time": "2021-03-25T14:50:11.548744", "status": "completed"} tags=[]
data['nom_2'].value_counts()
# + papermill={"duration": 0.262018, "end_time": "2021-03-25T14:50:12.187724", "exception": false, "start_time": "2021-03-25T14:50:11.925706", "status": "completed"} tags=[]
data['nom_3'].value_counts()
# + papermill={"duration": 0.234912, "end_time": "2021-03-25T14:50:12.582148", "exception": false, "start_time": "2021-03-25T14:50:12.347236", "status": "completed"} tags=[]
data['nom_4'].value_counts()
# + papermill={"duration": 0.239117, "end_time": "2021-03-25T14:50:12.961710", "exception": false, "start_time": "2021-03-25T14:50:12.722593", "status": "completed"} tags=[]
data['nom_5'].value_counts()
# + papermill={"duration": 0.233273, "end_time": "2021-03-25T14:50:13.348316", "exception": false, "start_time": "2021-03-25T14:50:13.115043", "status": "completed"} tags=[]
data['nom_6'].value_counts()
# + papermill={"duration": 0.237634, "end_time": "2021-03-25T14:50:13.728652", "exception": false, "start_time": "2021-03-25T14:50:13.491018", "status": "completed"} tags=[]
data['nom_7'].value_counts()
# + papermill={"duration": 0.197324, "end_time": "2021-03-25T14:50:14.067806", "exception": false, "start_time": "2021-03-25T14:50:13.870482", "status": "completed"} tags=[]
data.drop(['ord_5', 'nom_5', 'nom_6', 'nom_7', 'nom_8', 'nom_9'], axis = 1, inplace = True)
# + papermill={"duration": 0.141161, "end_time": "2021-03-25T14:50:14.353315", "exception": false, "start_time": "2021-03-25T14:50:14.212154", "status": "completed"} tags=[]
# + papermill={"duration": 0.155063, "end_time": "2021-03-25T14:50:14.654106", "exception": false, "start_time": "2021-03-25T14:50:14.499043", "status": "completed"} tags=[]
"""data['day'] = data['day'] / 7.0
data['month'] = data['month'] / 12.0"""
# + papermill={"duration": 0.172005, "end_time": "2021-03-25T14:50:14.974223", "exception": false, "start_time": "2021-03-25T14:50:14.802218", "status": "completed"} tags=[]
data.head()
# + [markdown] papermill={"duration": 0.144937, "end_time": "2021-03-25T14:50:15.264359", "exception": false, "start_time": "2021-03-25T14:50:15.119422", "status": "completed"} tags=[]
# Let's encode the remaining of the nominal values
# + papermill={"duration": 0.233609, "end_time": "2021-03-25T14:50:15.642800", "exception": false, "start_time": "2021-03-25T14:50:15.409191", "status": "completed"} tags=[]
data['nom_1'].value_counts()
# + papermill={"duration": 0.224219, "end_time": "2021-03-25T14:50:16.016223", "exception": false, "start_time": "2021-03-25T14:50:15.792004", "status": "completed"} tags=[]
m1_nom1 = {'Trapezoid' : 0, 'Square' : 1, 'Star' : 2, 'Circle' : 3, 'Polygon' : 4, 'Triangle' : 5}
data['nom_1'] = data['nom_1'].map(m1_nom1)
# + papermill={"duration": 0.234149, "end_time": "2021-03-25T14:50:16.392891", "exception": false, "start_time": "2021-03-25T14:50:16.158742", "status": "completed"} tags=[]
data['nom_2'].value_counts()
# + papermill={"duration": 0.217767, "end_time": "2021-03-25T14:50:16.759865", "exception": false, "start_time": "2021-03-25T14:50:16.542098", "status": "completed"} tags=[]
m2_nom2 = {'Lion' : 0, 'Cat' : 1, 'Snake' : 2, 'Dog' : 3, 'Axolotl' : 4, 'Hamster' : 5}
data['nom_2'] = data['nom_2'].map(m2_nom2)
# + papermill={"duration": 0.249077, "end_time": "2021-03-25T14:50:17.159053", "exception": false, "start_time": "2021-03-25T14:50:16.909976", "status": "completed"} tags=[]
data['nom_3'].value_counts()
# + papermill={"duration": 0.205612, "end_time": "2021-03-25T14:50:17.512765", "exception": false, "start_time": "2021-03-25T14:50:17.307153", "status": "completed"} tags=[]
m3_nom3 = {'Russia' : 0, 'Canada' : 1, 'China' : 2, 'Finland' : 3, 'Costa Rica' : 4, 'India' : 5}
data['nom_3'] = data['nom_3'].map(m3_nom3)
# + papermill={"duration": 0.23588, "end_time": "2021-03-25T14:50:17.897379", "exception": false, "start_time": "2021-03-25T14:50:17.661499", "status": "completed"} tags=[]
data['nom_4'].value_counts()
# + papermill={"duration": 0.205583, "end_time": "2021-03-25T14:50:18.251334", "exception": false, "start_time": "2021-03-25T14:50:18.045751", "status": "completed"} tags=[]
m4_nom4 = {'Oboe' : 0, 'Piano' : 1, 'Bassoon' : 2, 'Theremin' : 3}
data['nom_4'] = data['nom_4'].map(m4_nom4)
# + papermill={"duration": 0.170497, "end_time": "2021-03-25T14:50:18.566460", "exception": false, "start_time": "2021-03-25T14:50:18.395963", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.232772, "end_time": "2021-03-25T14:50:18.945230", "exception": false, "start_time": "2021-03-25T14:50:18.712458", "status": "completed"} tags=[]
data['nom_0'].value_counts()
# + papermill={"duration": 0.200713, "end_time": "2021-03-25T14:50:19.293854", "exception": false, "start_time": "2021-03-25T14:50:19.093141", "status": "completed"} tags=[]
m0_nom0 = {'Green' : 0, 'Blue' : 1, 'Red' : 2}
data['nom_0'] = data['nom_0'].map(m0_nom0)
# + papermill={"duration": 0.144929, "end_time": "2021-03-25T14:50:19.587002", "exception": false, "start_time": "2021-03-25T14:50:19.442073", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.144947, "end_time": "2021-03-25T14:50:19.876804", "exception": false, "start_time": "2021-03-25T14:50:19.731857", "status": "completed"} tags=[]
# Perform One Hot Encoding of the ordinal features
# + [markdown] papermill={"duration": 0.148646, "end_time": "2021-03-25T14:50:20.176474", "exception": false, "start_time": "2021-03-25T14:50:20.027828", "status": "completed"} tags=[]
# Label Encoding multiple columns
# + papermill={"duration": 0.278193, "end_time": "2021-03-25T14:50:20.601963", "exception": false, "start_time": "2021-03-25T14:50:20.323770", "status": "completed"} tags=[]
df_copy = data.copy()
df_copy.drop(['target'], axis = 1, inplace = True)
# + papermill={"duration": 0.63611, "end_time": "2021-03-25T14:50:21.384479", "exception": false, "start_time": "2021-03-25T14:50:20.748369", "status": "completed"} tags=[]
df_copy = pd.get_dummies(df_copy, columns = df_copy.columns)
df_copy
# + papermill={"duration": 0.146805, "end_time": "2021-03-25T14:50:21.679980", "exception": false, "start_time": "2021-03-25T14:50:21.533175", "status": "completed"} tags=[]
# + papermill={"duration": 0.177942, "end_time": "2021-03-25T14:50:22.005434", "exception": false, "start_time": "2021-03-25T14:50:21.827492", "status": "completed"} tags=[]
data.head()
# + papermill={"duration": 0.15653, "end_time": "2021-03-25T14:50:22.331824", "exception": false, "start_time": "2021-03-25T14:50:22.175294", "status": "completed"} tags=[]
# + papermill={"duration": 0.157883, "end_time": "2021-03-25T14:50:22.638294", "exception": false, "start_time": "2021-03-25T14:50:22.480411", "status": "completed"} tags=[]
#X = data.drop(['target'], axis = 1)
X = df_copy
y = data.target
# + papermill={"duration": 0.161834, "end_time": "2021-03-25T14:50:22.952777", "exception": false, "start_time": "2021-03-25T14:50:22.790943", "status": "completed"} tags=[]
# perform feature engineering to encode categorical variables so as to be processed by chi2_feature transform
# + papermill={"duration": 0.188539, "end_time": "2021-03-25T14:50:23.302576", "exception": false, "start_time": "2021-03-25T14:50:23.114037", "status": "completed"} tags=[]
# + papermill={"duration": 3.072693, "end_time": "2021-03-25T14:50:26.524336", "exception": false, "start_time": "2021-03-25T14:50:23.451643", "status": "completed"} tags=[]
chi2_features = SelectKBest(chi2, k = 10)
X_kbest_features = chi2_features.fit_transform(X,y)
print("Original Number of Features -> (shape)", X.shape[1])
print("K Best Features (shape)-> ",X_kbest_features.shape[1])
# + papermill={"duration": 0.160522, "end_time": "2021-03-25T14:50:26.834138", "exception": false, "start_time": "2021-03-25T14:50:26.673616", "status": "completed"} tags=[]
X_kbest_features
# + papermill={"duration": 0.149123, "end_time": "2021-03-25T14:50:27.137254", "exception": false, "start_time": "2021-03-25T14:50:26.988131", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.149136, "end_time": "2021-03-25T14:50:27.447176", "exception": false, "start_time": "2021-03-25T14:50:27.298040", "status": "completed"} tags=[]
# ### 4. T-Test
# + [markdown] papermill={"duration": 0.154854, "end_time": "2021-03-25T14:50:27.755019", "exception": false, "start_time": "2021-03-25T14:50:27.600165", "status": "completed"} tags=[]
# t-test also known as Student's t-test compares the two averages (means) and tells you if they are different from each other. <br>
# Can also tell you how significant the differences are.
# + [markdown] papermill={"duration": 0.156144, "end_time": "2021-03-25T14:50:28.064055", "exception": false, "start_time": "2021-03-25T14:50:27.907911", "status": "completed"} tags=[]
# **t-score**
# + [markdown] papermill={"duration": 0.152328, "end_time": "2021-03-25T14:50:28.368404", "exception": false, "start_time": "2021-03-25T14:50:28.216076", "status": "completed"} tags=[]
# **T-Values vs P-Values**
# + papermill={"duration": 0.153208, "end_time": "2021-03-25T14:50:28.677659", "exception": false, "start_time": "2021-03-25T14:50:28.524451", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.157276, "end_time": "2021-03-25T14:50:28.995642", "exception": false, "start_time": "2021-03-25T14:50:28.838366", "status": "completed"} tags=[]
# Types of T-Test <br>
# * Independent Samples t-test
# * Paired Sample t-test
# * One Sample t-test
# + [markdown] papermill={"duration": 0.158045, "end_time": "2021-03-25T14:50:29.311645", "exception": false, "start_time": "2021-03-25T14:50:29.153600", "status": "completed"} tags=[]
# ### 5. Z-Test
# + papermill={"duration": 0.151967, "end_time": "2021-03-25T14:50:29.617771", "exception": false, "start_time": "2021-03-25T14:50:29.465804", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.154502, "end_time": "2021-03-25T14:50:29.927210", "exception": false, "start_time": "2021-03-25T14:50:29.772708", "status": "completed"} tags=[]
# ### 6. ANOVA Test
# + [markdown] papermill={"duration": 0.151305, "end_time": "2021-03-25T14:50:30.236328", "exception": false, "start_time": "2021-03-25T14:50:30.085023", "status": "completed"} tags=[]
# ANOVA -> Analysis of Variance. <br>
# Helps to compare the means of more than 2 groups. <br>
# ANOVA F Test is also called omnibus test. <br><br><br>
#
# Main types of ANOVA Test ->
# * One-way or One-factor
# * Two-way or Two-factor
# + [markdown] papermill={"duration": 0.15106, "end_time": "2021-03-25T14:50:30.541101", "exception": false, "start_time": "2021-03-25T14:50:30.390041", "status": "completed"} tags=[]
# ANOVA Hypotheses -> <br>
# * Null Hypotheses = Group means are equal. No variation in the groups.
# * Alternative Hypothesis = At least, one group is different from other groups.
# + [markdown] papermill={"duration": 0.15616, "end_time": "2021-03-25T14:50:30.850115", "exception": false, "start_time": "2021-03-25T14:50:30.693955", "status": "completed"} tags=[]
# ANOVA Assumptions -> <br><br>
# * Residuals(experimental error) are normally distributed.(Shapiro-Wilks Test)
# * Homogenity of variances (variances are equal between treatment groups) (Levene's or Bartlett's Test)
# * Observations are sampled independently from each other.
# + [markdown] papermill={"duration": 0.15567, "end_time": "2021-03-25T14:50:31.163235", "exception": false, "start_time": "2021-03-25T14:50:31.007565", "status": "completed"} tags=[]
# ANOVA Working -> <br><br>
# * Check sample sizes, i.e., Equal number of observations in each group.
# * Calculate Mean Square for each group (MS) (SS of group/degrees of freedom-1)
# * Calc Mean Sq. Error (SS Error / df of residuals)
# * Calc F value (MS of group / MSE)
# + [markdown] papermill={"duration": 0.151686, "end_time": "2021-03-25T14:50:31.467320", "exception": false, "start_time": "2021-03-25T14:50:31.315634", "status": "completed"} tags=[]
# #### 6.1 One-Way ANOVA Test
# + papermill={"duration": 0.165767, "end_time": "2021-03-25T14:50:31.788107", "exception": false, "start_time": "2021-03-25T14:50:31.622340", "status": "completed"} tags=[]
import random
# + papermill={"duration": 0.178544, "end_time": "2021-03-25T14:50:32.123306", "exception": false, "start_time": "2021-03-25T14:50:31.944762", "status": "completed"} tags=[]
random.seed(2021)
# + papermill={"duration": 0.164328, "end_time": "2021-03-25T14:50:32.444942", "exception": false, "start_time": "2021-03-25T14:50:32.280614", "status": "completed"} tags=[]
df = pd.DataFrame([random.sample(range(1, 1000), 4) , random.sample(range(1, 1000), 4), random.sample(range(1, 1000), 4), random.sample(range(1, 1000), 4)], columns = ['A', 'B', 'C', "D"])
# + papermill={"duration": 0.166058, "end_time": "2021-03-25T14:50:32.761874", "exception": false, "start_time": "2021-03-25T14:50:32.595816", "status": "completed"} tags=[]
df
# + papermill={"duration": 0.171775, "end_time": "2021-03-25T14:50:33.084484", "exception": false, "start_time": "2021-03-25T14:50:32.912709", "status": "completed"} tags=[]
df_melt = pd.melt(df.reset_index(), id_vars = ['index'], value_vars = ['A','B','C','D'])
df_melt.columns = ['index', 'treatments', 'value']
# + papermill={"duration": 0.166957, "end_time": "2021-03-25T14:50:33.405018", "exception": false, "start_time": "2021-03-25T14:50:33.238061", "status": "completed"} tags=[]
df_melt
# + papermill={"duration": 0.39042, "end_time": "2021-03-25T14:50:33.947303", "exception": false, "start_time": "2021-03-25T14:50:33.556883", "status": "completed"} tags=[]
sns.boxplot(x='treatments', y='value', data=df_melt, color='#99c2a2')
sns.swarmplot(x="treatments", y="value", data=df_melt, color='#7d0013')
plt.show()
# + papermill={"duration": 0.163952, "end_time": "2021-03-25T14:50:34.270516", "exception": false, "start_time": "2021-03-25T14:50:34.106564", "status": "completed"} tags=[]
from scipy import stats
# + papermill={"duration": 0.170511, "end_time": "2021-03-25T14:50:34.611442", "exception": false, "start_time": "2021-03-25T14:50:34.440931", "status": "completed"} tags=[]
fvalue, pvalue = stats.f_oneway(df['A'], df['B'], df['C'], df['D'])
print("f Value -> ", fvalue)
print("p value -> ", pvalue)
# + papermill={"duration": 1.137213, "end_time": "2021-03-25T14:50:35.921965", "exception": false, "start_time": "2021-03-25T14:50:34.784752", "status": "completed"} tags=[]
import statsmodels.api as sm
from statsmodels.formula.api import ols
model = ols('value ~ C(treatments)', data = df_melt).fit()
anova_table = sm.stats.anova_lm(model, typ = 2)
anova_table
# + papermill={"duration": 0.154402, "end_time": "2021-03-25T14:50:36.233447", "exception": false, "start_time": "2021-03-25T14:50:36.079045", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.153479, "end_time": "2021-03-25T14:50:36.539622", "exception": false, "start_time": "2021-03-25T14:50:36.386143", "status": "completed"} tags=[]
# ##### Interpretation
# + [markdown] papermill={"duration": 0.155465, "end_time": "2021-03-25T14:50:36.847263", "exception": false, "start_time": "2021-03-25T14:50:36.691798", "status": "completed"} tags=[]
# p-value obtained from ANOVA Analysis is not significant (p > 0.05), and therefore, we conclude that there are no significant differences amongst the groups.
# + [markdown] papermill={"duration": 0.159849, "end_time": "2021-03-25T14:50:37.163641", "exception": false, "start_time": "2021-03-25T14:50:37.003792", "status": "completed"} tags=[]
# #### 6.2 Two-Way ANOVA Test
# + [markdown] papermill={"duration": 0.16378, "end_time": "2021-03-25T14:50:37.489646", "exception": false, "start_time": "2021-03-25T14:50:37.325866", "status": "completed"} tags=[]
# In Two-Way ANOVA Test, we have 2 independent variables and their different levels
# + papermill={"duration": 0.171259, "end_time": "2021-03-25T14:50:37.827839", "exception": false, "start_time": "2021-03-25T14:50:37.656580", "status": "completed"} tags=[]
data = pd.DataFrame(list(zip(['A','A','A','B','B','B', 'C', 'C', 'C', 'D', 'D', 'D'], [np.random.ranf() for _ in range(12)], [np.random.ranf() for _ in range(12)], [np.random.ranf() for _ in range(12)])), columns = ['Genotype', '1_year', '2_year', '3_year'])
# + papermill={"duration": 0.191894, "end_time": "2021-03-25T14:50:38.191143", "exception": false, "start_time": "2021-03-25T14:50:37.999249", "status": "completed"} tags=[]
data
# + papermill={"duration": 0.17465, "end_time": "2021-03-25T14:50:38.523757", "exception": false, "start_time": "2021-03-25T14:50:38.349107", "status": "completed"} tags=[]
data_melt = pd.melt(data, id_vars = ['Genotype'], value_vars = ['1_year', '2_year', '3_year'])
# + papermill={"duration": 0.181457, "end_time": "2021-03-25T14:50:38.860671", "exception": false, "start_time": "2021-03-25T14:50:38.679214", "status": "completed"} tags=[]
data_melt.head()
# + papermill={"duration": 0.17217, "end_time": "2021-03-25T14:50:39.202050", "exception": false, "start_time": "2021-03-25T14:50:39.029880", "status": "completed"} tags=[]
data_melt.columns = ['Genotype', 'years', 'value']
# + papermill={"duration": 0.73694, "end_time": "2021-03-25T14:50:40.096016", "exception": false, "start_time": "2021-03-25T14:50:39.359076", "status": "completed"} tags=[]
sns.boxplot(x = 'Genotype', y = 'value', hue = 'years', data = data_melt, palette = ['r', 'k', 'w'])
# + papermill={"duration": 0.176754, "end_time": "2021-03-25T14:50:40.431992", "exception": false, "start_time": "2021-03-25T14:50:40.255238", "status": "completed"} tags=[]
model = ols('value ~ C(Genotype) + C(years) + C(Genotype) : C(years)', data = data_melt).fit()
# + papermill={"duration": 0.213422, "end_time": "2021-03-25T14:50:40.805096", "exception": false, "start_time": "2021-03-25T14:50:40.591674", "status": "completed"} tags=[]
anova_table = sm.stats.anova_lm(model, typ = 2)
anova_table
# + papermill={"duration": 0.176307, "end_time": "2021-03-25T14:50:41.142282", "exception": false, "start_time": "2021-03-25T14:50:40.965975", "status": "completed"} tags=[]
# + papermill={"duration": 0.160091, "end_time": "2021-03-25T14:50:41.474000", "exception": false, "start_time": "2021-03-25T14:50:41.313909", "status": "completed"} tags=[]
# + papermill={"duration": 0.157853, "end_time": "2021-03-25T14:50:41.790489", "exception": false, "start_time": "2021-03-25T14:50:41.632636", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.17378, "end_time": "2021-03-25T14:50:42.128287", "exception": false, "start_time": "2021-03-25T14:50:41.954507", "status": "completed"} tags=[]
# ##### Post-Hoc Analysis (Tukey's Test)
# + papermill={"duration": 14.380589, "end_time": "2021-03-25T14:50:56.670437", "exception": false, "start_time": "2021-03-25T14:50:42.289848", "status": "completed"} tags=[]
# !pip install bioinfokit
from bioinfokit.analys import stat
# + papermill={"duration": 0.338274, "end_time": "2021-03-25T14:50:57.177898", "exception": false, "start_time": "2021-03-25T14:50:56.839624", "status": "completed"} tags=[]
res = stat()
res.tukey_hsd(df = df_melt, res_var = 'value', xfac_var = 'treatments', anova_model = 'value ~ C(treatments)')
output = res.tukey_summary
# + papermill={"duration": 0.182483, "end_time": "2021-03-25T14:50:57.526676", "exception": false, "start_time": "2021-03-25T14:50:57.344193", "status": "completed"} tags=[]
output
# + [markdown] papermill={"duration": 0.164781, "end_time": "2021-03-25T14:50:57.857293", "exception": false, "start_time": "2021-03-25T14:50:57.692512", "status": "completed"} tags=[]
# All the values are in accordance to the condition p > 0.05 <br>
# Hence, aren't statistically significant.
# + papermill={"duration": 0.170801, "end_time": "2021-03-25T14:50:58.193305", "exception": false, "start_time": "2021-03-25T14:50:58.022504", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.163681, "end_time": "2021-03-25T14:50:58.522490", "exception": false, "start_time": "2021-03-25T14:50:58.358809", "status": "completed"} tags=[]
# ### 7. F Stats Test
# + papermill={"duration": 0.16538, "end_time": "2021-03-25T14:50:58.853227", "exception": false, "start_time": "2021-03-25T14:50:58.687847", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.166432, "end_time": "2021-03-25T14:50:59.184039", "exception": false, "start_time": "2021-03-25T14:50:59.017607", "status": "completed"} tags=[]
# ### 8. Binomial Distribution
# + papermill={"duration": 0.188458, "end_time": "2021-03-25T14:50:59.537263", "exception": false, "start_time": "2021-03-25T14:50:59.348805", "status": "completed"} tags=[]
from scipy.stats import binom
n = 6
p = 0.6
r_values = list(range(n + 1))
mean, var = binom.stats(n, p)
dist = [binom.pmf(r, n, p) for r in r_values]
df = pd.DataFrame(list(zip(r_values, dist)), columns = ['r', 'p(r)'], index = None)
df
# + papermill={"duration": 0.361882, "end_time": "2021-03-25T14:51:00.066197", "exception": false, "start_time": "2021-03-25T14:50:59.704315", "status": "completed"} tags=[]
df['p(r)'].plot.bar()
# + [markdown] papermill={"duration": 0.166276, "end_time": "2021-03-25T14:51:00.401316", "exception": false, "start_time": "2021-03-25T14:51:00.235040", "status": "completed"} tags=[]
# ### 9. Normal Distribution
# + [markdown] papermill={"duration": 0.166821, "end_time": "2021-03-25T14:51:00.735234", "exception": false, "start_time": "2021-03-25T14:51:00.568413", "status": "completed"} tags=[]
# also known as
# * Gaussian Distribution
# * Bell Curve
#
#
# <br><br> Below is the probability distribution function (pdf) for Normal Distribution ->
# + [markdown] papermill={"duration": 0.170293, "end_time": "2021-03-25T14:51:01.074260", "exception": false, "start_time": "2021-03-25T14:51:00.903967", "status": "completed"} tags=[]
# 
# + [markdown] papermill={"duration": 0.232286, "end_time": "2021-03-25T14:51:01.472268", "exception": false, "start_time": "2021-03-25T14:51:01.239982", "status": "completed"} tags=[]
# * x -> input value
# * mu -> mean
# * sigma -> std deviation
# + [markdown] papermill={"duration": 0.167195, "end_time": "2021-03-25T14:51:01.805753", "exception": false, "start_time": "2021-03-25T14:51:01.638558", "status": "completed"} tags=[]
# 
# + papermill={"duration": 0.191789, "end_time": "2021-03-25T14:51:02.164296", "exception": false, "start_time": "2021-03-25T14:51:01.972507", "status": "completed"} tags=[]
mu, sigma = 0.5, 1
# + papermill={"duration": 0.177506, "end_time": "2021-03-25T14:51:02.509312", "exception": false, "start_time": "2021-03-25T14:51:02.331806", "status": "completed"} tags=[]
data = np.random.normal(mu, sigma, 10000)
# + papermill={"duration": 0.376779, "end_time": "2021-03-25T14:51:03.053364", "exception": false, "start_time": "2021-03-25T14:51:02.676585", "status": "completed"} tags=[]
count, bins, ignored = plt.hist(data, 20)
# + [markdown] papermill={"duration": 0.16682, "end_time": "2021-03-25T14:51:03.393027", "exception": false, "start_time": "2021-03-25T14:51:03.226207", "status": "completed"} tags=[]
#
# + papermill={"duration": 0.166641, "end_time": "2021-03-25T14:51:03.726004", "exception": false, "start_time": "2021-03-25T14:51:03.559363", "status": "completed"} tags=[]
# + papermill={"duration": 0.170747, "end_time": "2021-03-25T14:51:04.063293", "exception": false, "start_time": "2021-03-25T14:51:03.892546", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.165925, "end_time": "2021-03-25T14:51:04.398763", "exception": false, "start_time": "2021-03-25T14:51:04.232838", "status": "completed"} tags=[]
# ### 10. Poisson Distribution
# + papermill={"duration": 0.168575, "end_time": "2021-03-25T14:51:04.735669", "exception": false, "start_time": "2021-03-25T14:51:04.567094", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.170305, "end_time": "2021-03-25T14:51:05.077673", "exception": false, "start_time": "2021-03-25T14:51:04.907368", "status": "completed"} tags=[]
# ### 11. Bernoulli Distribution
# + papermill={"duration": 0.166282, "end_time": "2021-03-25T14:51:05.413098", "exception": false, "start_time": "2021-03-25T14:51:05.246816", "status": "completed"} tags=[]
# + papermill={"duration": 0.165876, "end_time": "2021-03-25T14:51:05.746814", "exception": false, "start_time": "2021-03-25T14:51:05.580938", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.175089, "end_time": "2021-03-25T14:51:06.087910", "exception": false, "start_time": "2021-03-25T14:51:05.912821", "status": "completed"} tags=[]
# ### 12. Central Limit Theorem
# + [markdown] papermill={"duration": 0.165772, "end_time": "2021-03-25T14:51:06.425001", "exception": false, "start_time": "2021-03-25T14:51:06.259229", "status": "completed"} tags=[]
# **What it states?** <br><br>
# Even when a sample is not normally distributed, if you draw multiple samples and take each of their averages, the averages will represent a normal distribution.<br><br>
# Which means repeated sampling from a not normally distributed sample and taking the means of those repeated samples will end up being a normally distributed sample. <br><br>
#
# 100 samples in total which are not normally distributed. Take random 10 samples say 50 times and take the mean of these samples. It will come out to be a normally distributed sample.
# + papermill={"duration": 0.166602, "end_time": "2021-03-25T14:51:06.758161", "exception": false, "start_time": "2021-03-25T14:51:06.591559", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.172863, "end_time": "2021-03-25T14:51:07.099002", "exception": false, "start_time": "2021-03-25T14:51:06.926139", "status": "completed"} tags=[]
# The following is an experiment of dice roll for 1000 times. <br>
# for 1000 times, we make samples of samples size 100 where possible outcomes are 1,2,3,4,5,6 <br><br>
# By plotting the histogram of the sample means, we obtain a normally distributed plot. <br>
# This is Central Limit Theorem
# + papermill={"duration": 0.394474, "end_time": "2021-03-25T14:51:07.681227", "exception": false, "start_time": "2021-03-25T14:51:07.286753", "status": "completed"} tags=[]
means = [np.mean(np.random.randint(1, 7, 100)) for _ in range(1000)]
plt.hist(means)
plt.show()
# + [markdown] papermill={"duration": 0.174503, "end_time": "2021-03-25T14:51:08.026007", "exception": false, "start_time": "2021-03-25T14:51:07.851504", "status": "completed"} tags=[]
# ##### Key Takeaways :- <br><br>
#
# 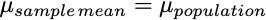
#
# 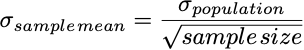
# + [markdown] papermill={"duration": 0.17407, "end_time": "2021-03-25T14:51:08.381106", "exception": false, "start_time": "2021-03-25T14:51:08.207036", "status": "completed"} tags=[]
# You can never experiment with all your customers (population). However, to draw a conclusion for an experiment which is a good representaion of your customers, you need to perform repeated experiments on different set of customers (different samples of the not normally distributed population/sample as per the context) and confirm your hypotheses.
# + papermill={"duration": 0.170499, "end_time": "2021-03-25T14:51:08.720380", "exception": false, "start_time": "2021-03-25T14:51:08.549881", "status": "completed"} tags=[]
# + papermill={"duration": 0.181764, "end_time": "2021-03-25T14:51:09.076684", "exception": false, "start_time": "2021-03-25T14:51:08.894920", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.168611, "end_time": "2021-03-25T14:51:09.421553", "exception": false, "start_time": "2021-03-25T14:51:09.252942", "status": "completed"} tags=[]
# ### 13. Calculating p-Value
# + papermill={"duration": 0.181859, "end_time": "2021-03-25T14:51:09.774003", "exception": false, "start_time": "2021-03-25T14:51:09.592144", "status": "completed"} tags=[]
# + papermill={"duration": 0.193465, "end_time": "2021-03-25T14:51:10.164609", "exception": false, "start_time": "2021-03-25T14:51:09.971144", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.169116, "end_time": "2021-03-25T14:51:10.506380", "exception": false, "start_time": "2021-03-25T14:51:10.337264", "status": "completed"} tags=[]
# ### 14. Measuring Correlation
# + papermill={"duration": 0.180977, "end_time": "2021-03-25T14:51:10.861997", "exception": false, "start_time": "2021-03-25T14:51:10.681020", "status": "completed"} tags=[]
# + papermill={"duration": 0.196327, "end_time": "2021-03-25T14:51:11.242797", "exception": false, "start_time": "2021-03-25T14:51:11.046470", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.180991, "end_time": "2021-03-25T14:51:11.603871", "exception": false, "start_time": "2021-03-25T14:51:11.422880", "status": "completed"} tags=[]
# ### 15. Measuring Variance
# + papermill={"duration": 0.182534, "end_time": "2021-03-25T14:51:11.961612", "exception": false, "start_time": "2021-03-25T14:51:11.779078", "status": "completed"} tags=[]
# + papermill={"duration": 0.187434, "end_time": "2021-03-25T14:51:12.352582", "exception": false, "start_time": "2021-03-25T14:51:12.165148", "status": "completed"} tags=[]
# + papermill={"duration": 0.175788, "end_time": "2021-03-25T14:51:12.709285", "exception": false, "start_time": "2021-03-25T14:51:12.533497", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.17964, "end_time": "2021-03-25T14:51:13.059023", "exception": false, "start_time": "2021-03-25T14:51:12.879383", "status": "completed"} tags=[]
# ### 16. Sample Statistics
# + papermill={"duration": 0.178305, "end_time": "2021-03-25T14:51:13.437853", "exception": false, "start_time": "2021-03-25T14:51:13.259548", "status": "completed"} tags=[]
# + papermill={"duration": 0.169735, "end_time": "2021-03-25T14:51:13.783099", "exception": false, "start_time": "2021-03-25T14:51:13.613364", "status": "completed"} tags=[]
# + papermill={"duration": 0.182085, "end_time": "2021-03-25T14:51:14.137815", "exception": false, "start_time": "2021-03-25T14:51:13.955730", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.16952, "end_time": "2021-03-25T14:51:14.496194", "exception": false, "start_time": "2021-03-25T14:51:14.326674", "status": "completed"} tags=[]
# ### 17. Population Statistics
# + papermill={"duration": 0.171278, "end_time": "2021-03-25T14:51:14.836753", "exception": false, "start_time": "2021-03-25T14:51:14.665475", "status": "completed"} tags=[]
# + papermill={"duration": 0.173728, "end_time": "2021-03-25T14:51:15.185828", "exception": false, "start_time": "2021-03-25T14:51:15.012100", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.169682, "end_time": "2021-03-25T14:51:15.595066", "exception": false, "start_time": "2021-03-25T14:51:15.425384", "status": "completed"} tags=[]
# ### 18. Maximum Likehood Estimation
# + papermill={"duration": 0.169775, "end_time": "2021-03-25T14:51:15.941859", "exception": false, "start_time": "2021-03-25T14:51:15.772084", "status": "completed"} tags=[]
# + papermill={"duration": 0.171196, "end_time": "2021-03-25T14:51:16.283745", "exception": false, "start_time": "2021-03-25T14:51:16.112549", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.185889, "end_time": "2021-03-25T14:51:16.640814", "exception": false, "start_time": "2021-03-25T14:51:16.454925", "status": "completed"} tags=[]
# ### 19. Cluster Analysis
# + papermill={"duration": 0.170575, "end_time": "2021-03-25T14:51:16.994860", "exception": false, "start_time": "2021-03-25T14:51:16.824285", "status": "completed"} tags=[]
# + papermill={"duration": 0.179835, "end_time": "2021-03-25T14:51:17.368185", "exception": false, "start_time": "2021-03-25T14:51:17.188350", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.17362, "end_time": "2021-03-25T14:51:17.713576", "exception": false, "start_time": "2021-03-25T14:51:17.539956", "status": "completed"} tags=[]
# ### 20. Hypothesis Testing
# + papermill={"duration": 0.178033, "end_time": "2021-03-25T14:51:18.064628", "exception": false, "start_time": "2021-03-25T14:51:17.886595", "status": "completed"} tags=[]
# + papermill={"duration": 0.196189, "end_time": "2021-03-25T14:51:18.461416", "exception": false, "start_time": "2021-03-25T14:51:18.265227", "status": "completed"} tags=[]
# + papermill={"duration": 0.173982, "end_time": "2021-03-25T14:51:18.809146", "exception": false, "start_time": "2021-03-25T14:51:18.635164", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.169035, "end_time": "2021-03-25T14:51:19.149724", "exception": false, "start_time": "2021-03-25T14:51:18.980689", "status": "completed"} tags=[]
# ### 21. Type-I Error & Type-II Error
# + papermill={"duration": 0.167427, "end_time": "2021-03-25T14:51:19.488645", "exception": false, "start_time": "2021-03-25T14:51:19.321218", "status": "completed"} tags=[]
# + papermill={"duration": 0.172267, "end_time": "2021-03-25T14:51:19.828102", "exception": false, "start_time": "2021-03-25T14:51:19.655835", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.173227, "end_time": "2021-03-25T14:51:20.170883", "exception": false, "start_time": "2021-03-25T14:51:19.997656", "status": "completed"} tags=[]
# ### 22. Z-Stats & T-Stats
# + papermill={"duration": 0.169026, "end_time": "2021-03-25T14:51:20.512895", "exception": false, "start_time": "2021-03-25T14:51:20.343869", "status": "completed"} tags=[]
# + papermill={"duration": 0.174153, "end_time": "2021-03-25T14:51:20.857235", "exception": false, "start_time": "2021-03-25T14:51:20.683082", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.173901, "end_time": "2021-03-25T14:51:21.204498", "exception": false, "start_time": "2021-03-25T14:51:21.030597", "status": "completed"} tags=[]
# ### 23. Confidence Interval
# + papermill={"duration": 0.177078, "end_time": "2021-03-25T14:51:21.569933", "exception": false, "start_time": "2021-03-25T14:51:21.392855", "status": "completed"} tags=[]
# + papermill={"duration": 0.172883, "end_time": "2021-03-25T14:51:21.914847", "exception": false, "start_time": "2021-03-25T14:51:21.741964", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.196939, "end_time": "2021-03-25T14:51:22.288580", "exception": false, "start_time": "2021-03-25T14:51:22.091641", "status": "completed"} tags=[]
# ### 24. Confusion Metrics, ROC & Regression Analysis
# + papermill={"duration": 0.167494, "end_time": "2021-03-25T14:51:22.632679", "exception": false, "start_time": "2021-03-25T14:51:22.465185", "status": "completed"} tags=[]
# + papermill={"duration": 0.169913, "end_time": "2021-03-25T14:51:22.980037", "exception": false, "start_time": "2021-03-25T14:51:22.810124", "status": "completed"} tags=[]
# + papermill={"duration": 0.175715, "end_time": "2021-03-25T14:51:23.342408", "exception": false, "start_time": "2021-03-25T14:51:23.166693", "status": "completed"} tags=[]
# + [markdown] papermill={"duration": 0.169867, "end_time": "2021-03-25T14:51:23.684331", "exception": false, "start_time": "2021-03-25T14:51:23.514464", "status": "completed"} tags=[]
# ### Notebook in Making. <br>
# Est. Date of Completion - 28-03-2021
| statistics-for-ml-data-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda3]
# language: python
# name: conda-env-anaconda3-py
# ---
# +
import pandas as pd
import os
from pysam import VariantFile
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from functools import reduce
plt.style.use('aa_paper')
# %matplotlib inline
# -
# # Reformat VCFs in parallel
#
# Using a script called `get_gens_df.py` in `AlleleAnalyzer/generate_gens_dfs/get_gens_df.py`, we reformat the 1000 Genomes VCFs in order to more easily annotate variants for whether they are near or in PAM sites. This is necessary because in ordinary VCF files, variants can have multiple alleles listed on one line, and these need to be split up for annotation based on each individual allele.
#
# For the 1000 Genomes analysis, we parallelized this process by splitting the genome into 10kb windows. (Will this make too many files? Maybe 500kb would be more feasible, then redo any that don't work, similar to ExAc approach. This approach is used because 1000 Genomes data is whole-genome rather than exome, like ExAc.
#
# Make 10 kb windows of the genome using `bedtools makewindows`.
#
# ## hg38
#
# `bedtools makewindows -g hg38.sizes -w 10000 > hg38.10kbwindows.bed`
#
# 321,184 regions for hg38.
#
# ### Add unique regions IDs
# +
hg38_regions = pd.read_csv('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/dat/hg38.10kbwindows.bed',
sep='\t', header=None, names=['chrom','start','stop'])
hg38_regions['region_id'] = 'region_' + hg38_regions.index.astype(str)
# # hg38_regions.to_csv('dat/1kgp_hg38_regions.bed', sep='\t', index=False, header=False)
# -
hg38_regions = pd.read_csv('dat/1kgp_hg38_regions.bed', sep='\t', header=None,
names=['chrom','start','stop','region_id'])
# ## Check whether all regions were appropriately reformatted
# +
hg38_regions['gens_fname'] = '/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_formatted_variants/' + hg38_regions['region_id'] + '.h5'
# hg38_regions['gens_complete'] = hg38_regions['gens_fname'].map(os.path.isfile)
# -
hg38_regions.query('~gens_complete')
len(hg38_regions.query('~gens_complete'))
# ## Check whether annotations were completed for appropriate regions
# +
hg38_regions['region_id'] = hg38_regions['region_id'].str.replace('_','')
hg38_regions['annots_fname'] = '/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_annotated_variants/' + \
hg38_regions['region_id'] + '.h5'
# -
hg38_regions['annots_file_exists'] = hg38_regions['annots_fname'].map(os.path.isfile)
hg38_regions.to_csv('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/region_annots.tsv',
sep='\t', index=False)
len(hg19_regions.query('~annots_file_exists'))
# ## hg38
#
# `bedtools makewindows -g hg38.sizes -w 10000 > hg38.10kbwindows.bed`
#
# 321,184 regions for hg38.
#
# ### add unique region IDs
hg38_regions = pd.read_csv('dat/1kgp_hg38_regions.bed', sep='\t', header=None,
names=['chrom','start','stop','region_id'])
# +
# check that gens file completed
hg38_regions['gens_fname'] = '/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_formatted_variants/' + hg38_regions['region_id'] + '.h5'
# hg38_regions['gens_complete'] = hg38_regions['gens_fname'].map(os.path.isfile)
# -
len(hg38_regions.query('gens_complete'))
hg38_regions.to_csv('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/region_annots.tsv',
sep='\t', index=False)
# # ExcisionFinder results
#
#
#
# `python gene_targ_variation.py /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/get_gene_list/gene_list_hg19.tsv /pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg19_analysis/1kgp_excisionfinder_results/results_by_chrom/ /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/src/hg19_analysis/plotting/`
#
# Targetable genes per person, just change the dir where h5 files are pulled to do 5kb window analysis.
#
# `python targ_genes_per_person.py /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/get_gene_list/gene_list_hg38.tsv /pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom/ targ_genes_per_person`
#
# with 5kb flanking
#
# `python targ_genes_per_person.py /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/get_gene_list/gene_list_hg38.tsv /pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom_5kb_window targ_genes_per_person_5kb`
#
# # Determine mean # putatively targetable autosomal genes per person in 1000 Genomes Cohort
def translate_gene_name(gene_name):
"""
HDF5 throws all sort of errors when you have weird punctuation in the gene name, so
this translates it to a less offensive form.
"""
repls = ("-", "dash"), (".", "period")
trans_gene_name = reduce(lambda a, kv: a.replace(*kv), repls, str(gene_name))
return trans_gene_name
genes = pd.read_csv('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/get_gene_list/gene_list_hg38.tsv',
sep='\t')
autosomal_genes = genes.query('(chrom != "chrX") and (chrom != "chrY")')
protein_coding_autosomal_genes = set(genes[genes['name'].str.startswith('NM')]['official_gene_symbol'].tolist())
genes.query('official_gene_symbol == "BEST1"')
targ_genes_per_person = np.load('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom/targ_genes_per_persongenes_per_person.npy').item()
targ_genes_per_person_5kb = np.load('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom_5kb_window/targ_genes_per_person_5kbgenes_per_person.npy').item()
# +
gene_dict = {}
genes_eval = 0
# for c_dir in os.listdir('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom'):
for chrom in list(range(1,23)):
c_dir = os.path.join('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom',f'chr{chrom}_ef_results/')
genes_in_dir = os.listdir(os.path.join('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom', c_dir))
for gene in genes_in_dir:
if gene.endswith('.h5'):
genes_eval += 1
gene_dict[gene[:-3]] = translate_gene_name(gene[:-3])
# print(genes_in_dir[:5])
# +
gene_dict = {}
genes_eval = 0
# for c_dir in os.listdir('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom'):
for chrom in list(range(1,23)):
c_dir = os.path.join('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom_5kb_window/',f'chr{chrom}_ef_results/')
genes_in_dir = os.listdir(os.path.join('/pollard/data/projects/AlleleAnalyzer_data/1kgp_data/hg38_analysis/1kgp_excisionfinder_results/results_by_chrom', c_dir))
for gene in genes_in_dir:
if gene.endswith('.h5'):
genes_eval += 1
gene_dict[gene[:-3]] = translate_gene_name(gene[:-3])
# print(genes_in_dir[:5])
# +
gene_dict_df = pd.DataFrame.from_dict(gene_dict, orient='index')
gene_dict_df['gene'] = gene_dict_df.index
gene_dict_df.columns = ['translated_gene','gene']
gene_dict_df.head()
gene_dict_df.to_csv('/pollard/data/projects/AlleleAnalyzer_data/AlleleAnalyzer_supporting_data/1000_genomes_analysis/hg38_analysis/excisionFinder_results/5kb_window/gene_dict.tsv',
sep='\t', index=False)
# -
print(genes_eval)
len(genes)
# ## Gene only (autosomal)
# +
ppl = []
num_targ_genes = []
cas = []
for key in targ_genes_per_person:
ppl.append(key)
num_targ_genes.append(len(protein_coding_autosomal_genes.intersection(set(targ_genes_per_person[key]))))
targ_genes_per_person_df = pd.DataFrame({'ppl':ppl, 'num_targ_genes':num_targ_genes})
targ_genes_per_person_df['perc_targ_genes'] = targ_genes_per_person_df['num_targ_genes'].divide(len(protein_coding_autosomal_genes)) * 100.0
targ_genes_per_person_df['perc_targ_genes'].mean()
# -
# ## Gene + 5 kb
# +
ppl = []
num_targ_genes = []
cas = []
for key in targ_genes_per_person_5kb:
ppl.append(key)
num_targ_genes.append(len(protein_coding_autosomal_genes.intersection(set(targ_genes_per_person_5kb[key]))))
targ_genes_per_person_df = pd.DataFrame({'ppl':ppl, 'num_targ_genes':num_targ_genes})
targ_genes_per_person_df['perc_targ_genes'] = targ_genes_per_person_df['num_targ_genes'].divide(len(protein_coding_autosomal_genes)) * 100.0
targ_genes_per_person_df['perc_targ_genes'].mean()
# -
# # people targetable
#
# In this faceted density plot, height of the colored portion indicates the proportion of genes where the specified percentage of the 1000 genomes cohort is putatively targetable.
# +
# inspired and helped by this page: https://seaborn.pydata.org/examples/kde_joyplot.html
plot_df = pd.read_csv('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/1000genomes_analysis/src/hg19_analysis/plotting/targ_per_gene_and_cas.tsv',
sep='\t')
cas_dict = np.load('/pollard/data/projects/AlleleAnalyzer_data/cas_abbrev_dict.npy').item()
cas_dict['StCas9'] = 'StCas9'
cas_dict['all'] = 'all'
plot_df['% people targetable per gene'] = plot_df['% people targetable']*100.0
sns.set(style="white", rc={"axes.facecolor": (0, 0, 0, 0)}, font_scale=1.2)
cas_list = plot_df.Cas.drop_duplicates().tolist()
pal = sns.cubehelix_palette(len(cas_list), rot=-.25, light=.7)
g = sns.FacetGrid(plot_df, row='Cas', hue='Cas', aspect=10, size=.5, palette=pal)
g.map(sns.kdeplot, '% people targetable per gene', shade=True, alpha=1, lw=1.5, bw=.2)
g.map(sns.kdeplot, '% people targetable per gene', color="w", lw=2, bw=.2)
g.map(plt.axhline, y=0, lw=2)
# Define and use a simple function to label the plot in axes coordinates
def label(x, color, label):
ax = plt.gca()
ax.text(0, .2, cas_dict[label], fontweight="bold",
ha="left", va="center", fontsize=11,transform=ax.transAxes)
g.map(label, '% people targetable per gene')
# Set the subplots to overlap
g.fig.subplots_adjust(hspace=-.25)
# Remove axes details that don't play well with overlap
g.set_titles("")
g.set(yticks=[])
g.despine(bottom=True, left=True)
# g.savefig('people_targetable_per_gene_per_cas.pdf', dpi=300, bbox_inches='tight')
# -
| manuscript_analyses/1000genomes_analysis/1000 Genomes Analysis hg38.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # %matplotlib inline
from matplotlib.gridspec import GridSpec
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
mpl.style.use('seaborn-white')
import multiprocess as mp
import numpy as np
import pandas as pd
import bioframe
import cooltools
import cooler
#import bbi
from cooltools import snipping
import sys
# +
dot_file_FA ='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-ESC4DN-FA-DpnII-R1-R2_hg38/combineddots/cloops_U54-ESC4DN-FA-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
dot_file_DSG='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-ESC4DN-DSG-DpnII-R1-R2_hg38/combineddots/cloops_U54-ESC4DN-DSG-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
dot_file_MNase='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-H1ESC4DN-FA-DSG-MNase-R1-R2_hg38/combineddots/cloops_U54-H1ESC4DN-FA-DSG-MNase-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_FA ='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-FA-DpnII-R1-R2_hg38/combineddots/cloops_U54-HFFc6-FA-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_DSG='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-DSG-DpnII-R1-R2_hg38/combineddots/cloops_U54-HFFc6-DSG-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_MNase='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-FA-DSG-MNase-R1-R3.hg38/combineddots/cloops_U54-HFFc6-FA-DSG-MNase-R1-R3.hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
ddd = [dot_file_FA,dot_file_DSG,dot_file_MNase]
lll = ['Hi-C-FA-DpnII','Hi-C-DSG-DpnII','Micro-C-DSG-MNase']
col=['r','b','g']
def dot_size_hist(dot_files,labels):
resolution=10000
plt.figure(figsize=(3.5 , 3.5))
w=7
s=(2*w-2)*resolution
bins=np.geomspace(s,20000000,num=50)
hist_keys = {"histtype":'step',
"cumulative":True,
"density":True,
"linewidth":2,}
ax=plt.subplot(1,1,1)
i=0
for dot_file,label in zip(dot_files,labels):
sites = pd.read_table(dot_file)
# print(sites.head(6))
dists=(sites['start2']-sites['start1'])
#ax.set_xlim(0, 3000000)
ax.hist(dists,bins=bins,color=col[i],label=label,**hist_keys)
print(sites.shape)
i=i+1
ax.set_xscale('log')
ax.set_ylabel('Frequency')
ax.set_xlabel('Dot size')
#plt.savefig('U54-HFF-DSG_dotsize.png')
ax.set_xlim(0, 3000000)
#plt.savefig('U54-HFF-DSG_dotsize.png')
# plt.legend(loc="best")
plt.legend(loc="lower right")
dot_size_hist(ddd,lll)
plt.savefig("Dotsize_ESC.pdf")
# +
#dot_file_FA = "/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/comp_dots/U54_ESC4DN_FA_DpnII_R1_R2_hg38_uniq_comp_to_U54_ESC4DN_DSG_DpnII_R1_R2_hg38.txt"
#dot_file_DSG = "/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/comp_dots/U54_ESC4DN_DSG_DpnII_R1_R2_hg38_uniq_comp_to_U54_ESC4DN_FA_DpnII_R1_R2_hg38.txt"
#dot_file_FA ='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-ESC4DN-FA-DpnII-R1-R2_hg38/combineddots/cloops_U54-ESC4DN-FA-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_DSG='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-ESC4DN-DSG-DpnII-R1-R2_hg38/combineddots/cloops_U54-ESC4DN-DSG-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_MNase='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-H1ESC4DN-FA-DSG-MNase-R1-R2_hg38/combineddots/cloops_U54-H1ESC4DN-FA-DSG-MNase-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
dot_file_FA ='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-FA-DpnII-R1-R2_hg38/combineddots/cloops_U54-HFFc6-FA-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
dot_file_DSG='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-DSG-DpnII-R1-R2_hg38/combineddots/cloops_U54-HFFc6-DSG-DpnII-R1-R2_hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
#dot_file_MNase='/nl/umw_job_dekker/users/ba69w/HiC_Analysis/U54_deep/snakedots/U54-HFFc6-FA-DSG-MNase-R1-R3.hg38/combineddots/cloops_U54-HFFc6-FA-DSG-MNase-R1-R3.hg38.mapq_30.1000.mcool.combined.bedpe.postproc'
ddd = [dot_file_DSG,dot_file_MNase]
lll = ['Hi-C-FA-DpnII','Hi-C-DSG-DpnII']
col=['r','b']
def dot_size_hist(dot_files,labels):
resolution=10000
plt.figure(figsize=(3.5 , 3.5))
w=7
s=(2*w-2)*resolution
bins=np.geomspace(s,20000000,num=50)
hist_keys = {"histtype":'step',
"cumulative":True,
"density":True,
"linewidth":2,}
ax=plt.subplot(1,1,1)
i=0
for dot_file,label in zip(dot_files,labels):
sites = pd.read_table(dot_file)
# print(sites.head(6))
dists=(sites['start2']-sites['start1'])
#ax.set_xlim(0, 3000000)
ax.hist(dists,bins=bins,color=col[i],label=label,**hist_keys)
print(sites.shape)
i=i+1
ax.set_xscale('log')
ax.set_ylabel('Frequency')
ax.set_xlabel('Dot size')
#plt.savefig('U54-HFF-DSG_dotsize.png')
ax.set_xlim(0, 3000000)
#plt.savefig('U54-HFF-DSG_dotsize.png')
# plt.legend(loc="best")
plt.legend(loc="lower right")
dot_size_hist(ddd,lll)
# -
| notebooks/plot_dot_size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml-project
# language: python
# name: ml-project
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="EXmhjBIll5g2" executionInfo={"status": "ok", "timestamp": 1610895624263, "user_tz": -60, "elapsed": 3141, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="ef659e09-7b11-4d4d-c866-e88353faff7a"
# !pip install ipython-autotime
# # !pip install -U scikit-learn
# %load_ext autotime
# + colab={"base_uri": "https://localhost:8080/"} id="bBKWty5P66sP" executionInfo={"status": "ok", "timestamp": 1610895624264, "user_tz": -60, "elapsed": 3124, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="b5955ab4-52da-405c-e8ca-e47ae504d528"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="7hWZupWyjh6i" executionInfo={"status": "ok", "timestamp": 1610895624518, "user_tz": -60, "elapsed": 2279, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="d598a2cf-2240-4afd-9663-41348833bd33"
# !pwd
import os
os.chdir('/content/drive/My Drive/ML/ml-project-master/src/')
# !pwd
# + colab={"base_uri": "https://localhost:8080/"} id="fnQOdVCePeHx" executionInfo={"status": "ok", "timestamp": 1610895627091, "user_tz": -60, "elapsed": 4073, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="991502d2-eb15-4e0e-d784-a38ff2d5c953"
from sklearn.datasets import fetch_openml
from sklearn.preprocessing import OneHotEncoder
import matplotlib.pyplot as plt
import tensorflow as tf
from utility import *
# + [markdown] id="ynMmZQI-PeID"
# # Import Dataset
# + id="mL96Zw5qPeIE" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1610895627092, "user_tz": -60, "elapsed": 2210, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="d75b6969-d834-4ddb-887b-03e60120522f"
Xtrain, Xtest, ytrain, ytest = load_monk("3")
print(Xtrain.shape)
print(Xtest.shape)
print(ytrain.shape)
print(ytest.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="hLG-zsJC2SHs" executionInfo={"status": "ok", "timestamp": 1610895627094, "user_tz": -60, "elapsed": 1075, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="a1b34327-2104-42e5-bff0-e54afa4c5823"
# X, y = fetch_openml('monks-problems-3', return_X_y=True)
# y = y.astype(np.float32)
# X = OneHotEncoder(sparse=False).fit_transform(X)
# y = y.reshape(y.shape[0], 1)
# + [markdown] id="RBI2gd0E2g1f"
#
# + [markdown] id="O3OOFhaO2ncY"
# # Grid Search - Gradient Descent
# + colab={"base_uri": "https://localhost:8080/"} id="06rvDXOo2nca" executionInfo={"status": "ok", "timestamp": 1610895630112, "user_tz": -60, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="5919003b-7296-4965-a7dc-737010f6d4df"
best_results_filename = "best_results_monk3.txt"
single_grid_results_filename = 'grid_results_monk3.1.csv'
# + id="0vDdoiUG2ncd"
from sklearn.model_selection import GridSearchCV
from tensorflow.keras.models import Sequential
from tensorflow.keras import layers
from tensorflow.keras.layers import Dense
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from tensorflow.keras.optimizers import SGD
# L2 = tf.keras.regularizers.L2(l2=0.02)
# Function to create model, required for KerasClassifier
def create_model(learn_rate=0.01, momentum=0, neurons=1, init_mode='uniform', activation='relu'):
# create model
model = Sequential()
model.add(layers.InputLayer(input_shape=(17)))
model.add(Dense(neurons, activation=activation, kernel_initializer=init_mode))
model.add(Dense(1, activation='sigmoid', kernel_initializer=init_mode))
optimizer = SGD(
learning_rate=learn_rate, momentum=momentum, nesterov=False)
# Compile model
model.compile(optimizer=optimizer,
loss=tf.keras.losses.MSE,
metrics='accuracy')
return model
BATCH_SIZE = len(Xtrain)
# create model
model = KerasClassifier(build_fn=create_model, verbose=0)
# define the grid search parameters
batch_size = [32, 64, BATCH_SIZE]
epochs = [1000]
learn_rate = [0.001, 0.01, 0.1]
momentum = [0.0, 0.2, 0.6, 0.8, 0.9]
neurons = [4, 5]
init_mode = ['glorot_uniform']
activation = ['relu', 'tanh', 'sigmoid']
# momentum = [0.0, 0.2, 0.4, 0.6, 0.8, 0.9]
# neurons = [2, 3, 4, 5]
# init_mode = ['uniform', 'normal', 'zero', 'glorot_uniform']
# optimizer = ['SGD', 'RMSprop', 'Adagrad', 'Adadelta', 'Adam', 'Adamax', 'Nadam']
param_grid = dict(batch_size=batch_size, epochs=epochs, learn_rate=learn_rate, momentum=momentum, neurons=neurons, init_mode=init_mode, activation=activation)#, optimizer=optimizer)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1, cv=3)
grid_result = grid.fit(Xtrain, ytrain)
# + id="NLg-Ybd82ncf"
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
grid_values_string = f'Grid on: batch_size: {batch_size}, epochs: {epochs}, learn_rate: {learn_rate}, momentum: {momentum}, init_mode = {init_mode} \n'
grid_results_string = f'Best: {grid_result.best_score_} using {grid_result.best_params_} \n'
grid_correspondant_parameter_name = f'Results in: {single_grid_results_filename} \n'
# Open a file with access mode 'a'
with open(best_results_filename, "a") as file_object:
# Append 'hello' at the end of file
file_object.write(grid_correspondant_parameter_name)
file_object.write(grid_values_string)
file_object.write(grid_results_string)
# + id="yoGMbYYX2ncg"
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + id="SQzBbsYE2nch"
import csv
with open(single_grid_results_filename, mode='w') as grid:
grid = csv.writer(grid, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
for mean, stdev, param in zip(means, stds, params):
grid.writerow([mean, stdev, param])
# + id="9t0UZcUP2nci"
# from sklearn.externals import joblib
# # #save your model or results
# # joblib.dump(grid_result, 'gs_object_monk3.pkl')
# # #load your model for further usage
# # boh = joblib.load("gs_object_monk3.pkl")
# # joblib.dump(grid.best_estimator_, 'gs_best_estimator_monk3.pkl', compress = 1)
# # filename = 'finalized_model.sav'
# # pickle.dump(model, open(filename, 'wb'))
# + [markdown] id="mboN1SB4xod2"
# # Grid Search - Gradient Descent with regularization
# + colab={"base_uri": "https://localhost:8080/"} id="HZ10SJ5ybRml" executionInfo={"status": "ok", "timestamp": 1610892633434, "user_tz": -60, "elapsed": 5527, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="bad477ae-8d17-4013-9d71-766e6382f8f8"
best_results_filename = "best_results_monk3.txt"
single_grid_results_filename = 'grid_results_monk3.2.csv'
# + colab={"base_uri": "https://localhost:8080/", "height": 370} id="wc_dfjUHxCHS" executionInfo={"status": "error", "timestamp": 1610895102831, "user_tz": -60, "elapsed": 133439, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiItemmzluuiJcy95qoZ5g1w4YMcvBQA0c2FQxbpg=s64", "userId": "11115762631469754742"}} outputId="1447da97-8b46-41cb-dae5-ac188920c812"
from sklearn.model_selection import GridSearchCV
from tensorflow.keras.models import Sequential
from tensorflow.keras import layers
from tensorflow.keras.layers import Dense
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.regularizers import L2
# L2 = tf.keras.regularizers.L2(l2=0.02)
# Function to create model, required for KerasClassifier
def create_model(learn_rate=0.01, momentum=0, neurons=1, init_mode='uniform', activation='relu', reg_L2 = 0.0):
# create model
model = Sequential()
model.add(layers.InputLayer(input_shape=(17)))
model.add(Dense(neurons, activation=activation, kernel_initializer= init_mode, kernel_regularizer= L2(reg_L2) ))
model.add(Dense(1, activation='sigmoid', kernel_initializer=init_mode, kernel_regularizer= L2(reg_L2)))
optimizer = SGD(
learning_rate=learn_rate, momentum=momentum, nesterov=False)
# Compile model
model.compile(optimizer=optimizer,
loss=tf.keras.losses.MSE,
metrics='accuracy')
return model
BATCH_SIZE = len(Xtrain)
# create model
model = KerasClassifier(build_fn=create_model, verbose=0)
# define the grid search parameters
batch_size = [32, 64, BATCH_SIZE]
epochs = [1000]
learn_rate = [0.1, 0.001, 0.01, 0.1]
momentum = [0.0, 0.2, 0.6, 0.8, 0.9]
neurons = [4, 5]
init_mode = ['glorot_uniform']
activation = ['tanh'] #['relu', 'tanh', 'sigmoid']
#L2_list = [0.0, 0.1, 0.2, 0.01]
# init_mode = ['uniform', 'normal', 'zero', 'glorot_uniform']
# optimizer = ['SGD', 'RMSprop', 'Adagrad', 'Adadelta', 'Adam', 'Adamax', 'Nadam']
param_grid = dict(batch_size=batch_size, epochs=epochs, learn_rate=learn_rate, momentum=momentum, neurons=neurons, init_mode=init_mode, activation=activation, reg_L2 = L2_list )#, optimizer=optimizer)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1, cv=3)
grid_result = grid.fit(Xtrain, ytrain)
# + id="Lo7rzmewiUaG"
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
grid_values_string = f'Grid on: batch_size: {batch_size}, epochs: {epochs}, learn_rate: {learn_rate}, momentum: {momentum}, init_mode = {init_mode} \n'
grid_results_string = f'Best: {grid_result.best_score_} using {grid_result.best_params_} \n'
grid_correspondant_parameter_name = f'Results in: {single_grid_results_filename} \n'
# Open a file with access mode 'a'
with open(best_results_filename, "a") as file_object:
# Append 'hello' at the end of file
file_object.write(grid_correspondant_parameter_name)
file_object.write(grid_values_string)
file_object.write(grid_results_string)
# + id="SA9qd_IA524a"
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
params = grid_result.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# + id="dUtpM0u02yll"
import csv
with open(single_grid_results_filename, mode='w') as grid:
grid = csv.writer(grid, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
for mean, stdev, param in zip(means, stds, params):
grid.writerow([mean, stdev, param])
# + id="ONVYVykW6s3V"
# from sklearn.externals import joblib
# # #save your model or results
# # joblib.dump(grid_result, 'gs_object_monk3.pkl')
# # #load your model for further usage
# # boh = joblib.load("gs_object_monk3.pkl")
# # joblib.dump(grid.best_estimator_, 'gs_best_estimator_monk3.pkl', compress = 1)
# # filename = 'finalized_model.sav'
# # pickle.dump(model, open(filename, 'wb'))
# + id="kNuP8YzW4A7e"
| src/keras/Monk3_GS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# <a href="https://colab.research.google.com/github/institutohumai/cursos-python/blob/master/Introduccion/4_Intro_Poo/intro-poo-solucion.ipynb"> <img src='https://colab.research.google.com/assets/colab-badge.svg' /> </a>
# <div align="center"> Recordá abrir en una nueva pestaña </div>
# <h1 id="tocheading">Programación orientada a objetos</h1>
#
# I. Terminologia de clases y objetos
# II. Creando la primera clase
# I. Herencia
# II. Protección de acceso
# III. Métodos Especiales
# IV. Métodos Estáticos
# V. Duck typing y monkey patching
# I. Duck typing
# II. Monkey Patching
# + [markdown] colab_type="text" id="JWZi0Jz7myZr"
# En el paradigma de programación orientada a objetos los programas se estructuran organizando el código en entidades llamadas objetos. Estos nos permiten encapsular data, funciones y variables dentro de una misma clase. Veamos de qué se trata.
# + [markdown] colab_type="text" id="8facntg8myZs"
# ## Terminologia de clases y objetos
# + [markdown] colab_type="text" id="f4-ocEflmyZs"
# 1. Una **clase** es un prototipo de objeto, que engloba atributos que poseen todos los objetos de esa clase. Los atributos pueden ser datos como variables de clase y de instancia, y métodos (funciones). Se acceden con un punto.
#
# 2. Una **instancia** es un objeto en particular que pertenece a una clase.
#
# 3. Una variable de clase o **class variable** es un atributo compartido por todas las instancias de la clase. Se definen dentro de una clase pero fuera de un método.
#
# 4. La **herencia** es la transferencia de atributos de una clase a otra clase
#
# 5. Un **método** es una función contenida dentro de un objeto.
#
# 6. Un **objeto** es una instancia única de una estructura definida por su clase. Posee de atributos variables de clase, de instancia y métodos.
#
#
# + [markdown] colab_type="text" id="WL8GajTfmyZt"
# ## Creando la primera clase
# -
import math
# + slideshow={"slide_type": "slide"}
#La sintáxis es:
class Ejemplo:
pass
# Instancio la clase
x = Ejemplo()
print(type(x))
# -
# Por convención, las clases se nombran empleando "upper camel case". Es decir, con mayúscula para cada término que sea parte del nombre.
# + [markdown] slideshow={"slide_type": "subslide"}
# Una librería famosa en Python por sus clases es "requests". Esta ĺibrería se usa para acceder a información web por HTTP. Algunas de sus clases son:
#
# - Session
# - Request
# - ConnectionError
# - ConnectTimeout
#
# Las últimas dos clases son para especificar errores, noten que se repiten las mayúsculas.
# -
# Podemos pensar a una clase como un molde, el cual usamos para generar objetos o instancias que tienen ciertos atributos o métodos (funciones) que deseamos mantener.
#
# Aquellos atributos y métodos que queremos que los objetos conserven son definidos como parte del constructor. El constructor en Python es el método reservado **\_\_init\_\_()**. Este método se llama cuando se instancia la clase y en ese momento se inicializan los atributos de la clase, para lo cual podemos pasar parámetros.
#
# Además, vamos a emplear el término reservado **self** para indicar aquellos atributos y métodos que van a ser propios de los objetos. Veámoslo con un ejemplo.
# + colab={} colab_type="code" id="ru6DucmamyZ7"
class Persona():
def __init__(self, nombre, apellido, edad, contacto):
# Este método puede tomar parámetros que asignamos a los atributos, que luego podemos acceder
self.edad = edad # este es un atributo
self.contacto = contacto # este es otro atributo
self.nombre = nombre
self.apellido = apellido
def nombre_completo(self):
# este método toma el nombre completo y lo separa en nombre y apellido
nombre_completo = ', '.join([self.apellido,self.nombre])
return nombre_completo
def saludar(self):
print(f'Hola mi nombre es {self.nombre_completo()}',
f'y te dejo mi mail por si necesitás algo: {self.contacto}')
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="-3AWihITmyZ-" outputId="cbe535a4-e3bc-499f-a956-027f7affc2ac"
instancia_ejemplo = Persona('<NAME>','Ripley', 24, '<EMAIL>')
instancia_ejemplo.saludar()
# -
# Ahora veamos una clase menú que administra los platos y los precios
class Menu():
def __init__(self, items):
self.items = items
def precio(self, lista_items):
precio = 0
for nombre_item in lista_items:
precio = precio + self.items[nombre_item]
return precio
def tamaño(self):
return len(self.items)
mi_menu = Menu({'latte':25, 'medialuna':15})
# ¿Cuánto salen un latte y dos madialunas? ¿Cuántos ítems tenemos?
mi_menu.precio(['latte','medialuna','medialuna'])
mi_menu.tamaño()
# Ejercicio: Vamos a mejorar la clase anterior... En lugar de que el método precio reciba una lista de strings, hagamos que reciba una lista de diccionarios cada uno con dos claves nombre y cantidad que querríamos ordenar ¿Cuántos cuestan 10 lattes y 30 medialunas?
class Menu():
def __init__(self, items):
self.items = items
def precio(self, lista_items):
precio = 0
for pedido in lista_items:
# Leemos el diccionario
nombre = pedido['nombre']
cantidad = pedido['cantidad']
# Buscamos el precio en los ítems del menú
precio_unidad = self.items[nombre]
# Agregamos el total de cada ítem
precio = precio + precio_unidad * cantidad
return precio
def tamaño(self):
return len(self.items)
mi_menu = Menu({'latte':25, 'medialuna':15})
mi_menu.precio([{'nombre':'latte','cantidad':10},
{'nombre':'medialuna','cantidad':30}])
# Los atributos también son conocidos como variables de instancia, en contraposición a las variables de clase. Las variables de instancia toman un valor específico a una instancia en particular (por eso se emplea el término **self**), por su parte, las variables de clase tienen un valor común para todas las instancias de una clase. Por convención las variables de clase se definen antes del constructor y no llevan **self** en su definición pero sí cuando se la quiere llamar.
# + colab={} colab_type="code" id="ffB3W0b9pz3g"
class Curso:
max_alumnos = 35 # definimos variable de clase
def __init__(self, nombre, duracion, alumnos = None, costo=10):
self.nombre = nombre
self.duracion = duracion
if alumnos is None:
self.alumnos = []
else:
self.alumnos = alumnos
self.costo = costo # costo tiene un valor por default
"""¿Por qué ese if? Las variables por default sólo se evalúan a la hora de ejecutar la sentencia def.
En nuestro caso necesitamos que self.alumnos sea una lista y las listas son objetos mutables.
Esto quiere decir que podemos modificarla sin volver a asignarla. Si en vez de 'alumnos = None' usáramos
alumnos = [], entonces con cada nueva instancia del objeto estaríamos compartiendo los alumnos.
Para evitar eso, en general la forma pythónica de hacerlo es usando None por default y asignando el valor
deseado dentro de la función y no en el 'def' """
def inscribir_alumno(self, nombre):
self.alumnos.append(nombre) # para poder llamar a alumnos tengo que usar self.
print(f'Se agregó al alumno/a {nombre}')
def tomar_lista(self):
for a in self.alumnos:
print(f'Alumno: {a}')
def resumen(self):
print(f'Curso {self.nombre}, {self.duracion} clases pensadas para {len(self.alumnos)} alumnos\n'
f'Por el muy módico precio de {self.costo} rupias.',
# llamo variable de clase:
f'La ocupación actual es del {round(len(self.alumnos)/self.max_alumnos,2)*100}%')
# + colab={} colab_type="code" id="XmX4l4IF_eTI"
curso_python = Curso('Python', 6)
# -
curso_python.alumnos
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="MnRKu_2M_sxP" outputId="380c19ec-65b3-4840-bf2e-d0711fbde5e6"
# Llamamos metodos de la instancia
curso_python.inscribir_alumno('Diotimia')
curso_python.inscribir_alumno('Aritófanes')
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="gFvzeuDn_yNf" outputId="2c62cced-2a98-41e6-a2c0-7a372038c02c"
curso_python.tomar_lista()
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="AqkOrzAavtQf" outputId="3a20c0cd-a11d-49b2-b1e7-05c241aae760"
curso_python.resumen()
# -
curso_ml = Curso('Machine Learning', 8)
curso_ml.alumnos # vean que el curso está vacío!
curso_ml = Curso('Machine Learning', 8)
curso_ml.inscribir_alumno('Agatón')
curso_ml.inscribir_alumno('Erixímaco')
curso_ml.inscribir_alumno('Sócrates')
curso_ml.resumen()
curso_ml.alumnos
# Ejercicios:
#
# 1- Defina una clase Punto que tome como parámetros x e y (las coordenadas) y constante que se puede instanciar correctamente.
#
# 2- En Python existen los llamados métodos mágicos (magic methods) o dunder (Double Underscores). Estos métodos se caracterizan, justamente, por comenzar y terminar con "\_\_". Uno de los más comunes es el que permite darle estilo a la función **print**. Para que nuestro objeto entonces tenga un lindo print tenemos que definir una función "\_\_str\_\_" que sólo toma "self" como parámetro y que torne un string. Eso que retorna es el string que queremos que muestra cuando hagamos "print" del objeto. Dicho ésto, te invitamos a que lo intentes de la siguiente manera:
#
# a. Definí una función "\_\_str\_\_" que sólo toma self como parámetro.
#
# b. La función debe retornar el string que querés mostrar, recordá que podés usar los valores de "x" y de "y"
class Punto:
def __init__(self, x, y):
self.x = x
self.y = y
def __str__(self):
return f"({self.x}, {self.y})"
punto = Punto(1.0, 2.0)
print(punto)
# + [markdown] colab_type="text" id="tZhkGmH4zp4t"
# ### Herencia
# La herencia se emplea cuando queremos que una clase tome los atributos y características de otra clase.
# En este caso, la clase derivada (Alumno) **hereda** atributos y métodos de la clase base (Persona).
# Para acceder a los métodos de la clase previa vamos a emplear el método reservado **super()**. Con este método podemos invocar el constructor y así acceder a los atributos de esa clase.
# + colab={} colab_type="code" id="20SpMkEzDDNB"
# Clase derivada
class Alumno(Persona):
def __init__(self, curso: Curso, *args):
"""
Alumno pertence a un Curso (una instancia de la clase Curso) y, además, tiene otros atributos que pasaremos
a la clase previa
"""
self.curso = curso
super().__init__(*args) # inicializamos la clase 'madre'. La llamamos usando super() y ejecutamos el constructor
# Nótese también que desempacamos args
def saludar(self): # Sobrecarga de métodos, ver abajo
super().saludar() # ejecutamos el método de Persona .saludar() y agregamos más cosas a este método
print('Estoy cursando:')
self.curso.resumen()
def estudiar(self, dato): # También podemos definir nuevos métodos
self.conocimiento = dato
# -
# La clase Persona cuenta con un método saludar() y para Alumno también definimos un método saludar(). Cuando instanciemos un Alumno y ejecutemos el método saludar() lo que va a ejecutarse es el método saludar() de Alumno, no de Persona. Esto no quita que el método saludar() de Alumno llame al de Persona. Además, vale la pena mencionar que los dos tienen los mismos parámetros (ninguno en este caso). Este patrón de diseño es lo que se llama sobrecarga de métodos o overriding.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="eWN6A1VebZDF" outputId="b0eac13b-0865-4958-c64a-9de6d8e94956"
scott = Alumno(curso_python, 'Scott', 'Henderson', 49, '<EMAIL>')
scott.saludar()
# + colab={} colab_type="code" id="yaActu69GauJ"
scott.estudiar('Se puede heredar de otra clase y extender sus métodos')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="TxnvrBvsHGzW" outputId="572c6a52-53af-444d-80a0-de6228446c5d"
scott.conocimiento
# -
# Ejercicio:
#
# 1- Listar cuáles son los atributos y los métodos de scott y especificar cuáles provienen de Persona y cuáles están definidos por ser Alumno.
# + [markdown] colab_type="text" id="6BBCTXGskrBp"
# ### Protección de acceso
#
# Podemos cambiar el acceso (publico, no publico, protejido) de los métodos y variables.
# + [markdown] colab_type="text" id="xY3x3-5Co8-F"
# Dos formas distintas de encapsulamiento:
#
# - `_nopublico`
# - `__protegido`
#
# Los atributos o método no públicos pueden ser accedidos desde el objeto y llevan el prefijo "\_". La utilidad de este es indicarle al usuario que es una variable o método privado, de uso interno en el código de la clase y que no está pensando que sea usado desde afuera, por el usuario.
#
# Por otra parte, en el caso de usar como prefijo "\_\_" (doble "\_") directamente vamos a ocultar la variable o método de la lista de sugerencias para el usuario y tampoco va a poder invocarlo desde el objeto. Por este motivo, decimos que el atributo o método está protegido.
# + colab={} colab_type="code" id="-MxLTGBfkqTN"
class Auto():
def __init__(self, color, marca, velocidad_maxima):
self.color = color
self.marca = marca
self.__velocidad_maxima = 200
self.velocidad = 0
self.__contador = 0 # kilometros recorridos
def avanzar(self, horas=1, velocidad=10):
if self._chequear_velocidad(velocidad):
self.velocidad = velocidad
print(f'avanzando durante {horas} horas')
self.__contador += horas*self.velocidad
else:
print(f"Tu auto no puede llegar a tanta velocidad, el máximo es {self.__velocidad_maxima}")
def _chequear_velocidad(self, velocidad):
es_valida = False
if velocidad < self.__velocidad_maxima:
es_valida = True
if self.velocidad < velocidad:
print("Vas a acelerar!")
else:
print("Vas a desaceler!")
else:
print("Tu motor no permite ir tan rápido")
es_valida = False
return es_valida
def status(self):
print(f"Vas a una velocidad de {self.velocidad} y llevás {self.__contador} km. recorridos")
# + colab={} colab_type="code" id="pHGKm6iondA5"
superauto = Auto('rojo','Ferraudi', 200)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="2VWZRkQ6pfZz" outputId="c70d1485-807d-4f9e-dcf2-331706a789e5"
# Atributo no publ
superauto.avanzar(10)
# -
superauto.status()
# + colab={"base_uri": "https://localhost:8080/", "height": 164} colab_type="code" id="WSafkUzHnp8y" outputId="10939014-e8b5-4444-f0c1-ed514e0344c2"
# No se puede acceder a un atributo protegido
superauto.__contador
# -
# Pero sí se puede acceder a un método no público:
superauto._chequear_velocidad(10)
# Ejercicio:
#
# A continuación se define una clase Linea. Esta clase toma como parámetros dos objetos Punto() (instancias de la clase que definieron antes).
#
# 1- Agregar un método 'largo' que permita calcular el largo de la línea. Para ello vale la pena recordar que ésta se puede calcular como la hipotenusa del triángulo rectángulo que se forma con los dos puntos.
# \\[ a = \sqrt{b^2 + c^2} \\]
# <img src="https://static1.abc.es/media/ciencia/2019/10/31/TeoremadePitagorasABC-kW8F-U3032581527206JG-620x450@abc.jpg" width=250/>
# 2- Agregar un método 'pendiente' que permita calcular la pendiente de la línea. Recordar que ésta se puede calcular como el cociente entre las diferencias de 'y' y de 'x'.
#
# La fórmula es :
# \\[ m = (y_2 - y_1)/(x_2 - x_1) \\]
class Linea(object):
def __init__(self, p1: Punto, p2: Punto):
self.p1 = Punto(x0,y0)
self.p2 = Punto(x1,y1)
def __str__(self):
x1, y1 = self.p1.x, self.p1.y
x2, y2 = self.p2.x, self.p2.y
linea = "((%f,%f),(%f,%f))" % (x0, y0, x1, y1)
return linea
def largo(self):
dist_x = self.p2.x - self.p1.x
dist_y = self.p2.y - self.p1.y
dist_x_squared = dist_x ** 2
dist_y_squared = dist_y ** 2
largo = (dist_x_squared + dist_y_squared) ** 0.5
return largo
def pendiente(self):
dist_y = self.p2.y - self.p1.y
dist_x = self.p2.x - self.p1.x
pendiente = dist_y/dist_x
return pendiente
# + slideshow={"slide_type": "subslide"}
x0,y0 = 7,5
x1,y1 = 4,1
# -
p1 = Punto(x0,y0)
p2 = Punto(x1,y1)
linea = Linea(p1,p2)
linea.pendiente()
linea.largo()
# ## Métodos Especiales
#
# Las clases en Python cuentan con múltiples métodos especiales, los cuales se encuentran entre dobles guiones bajos __<metodo>__().
#
# Los métodos especiales más utilizados son <strong> \_\_init\_\_(), \_\_str\_\_() y \_\_del\_\_() </strong>.
#
# \_\_init\_\_ sirve para inicializar la clase y \_\_del\_\_ sirve para eliminar completamente el objeto del compilador.
#
# Veamos un ejemplo de uso de \_\_str\_\_. Una vez que definimos este método, responde a la sintaxis reservada de Python str().
# Clase derivada
class Alumno(Persona):
def __init__(self, curso: Curso, *args):
"""
Alumno pertence a un Curso (una instancia de la clase Curso) y, además, tiene otros atributos que pasaremos
a la clase previa
"""
self.curso = curso
super().__init__(*args) # inicializamos la clase 'madre'. La llamamos usando super() y ejecutamos el constructor
# Nótese también que desempacamos args
def saludar(self): # Sobrecarga de métodos, ver abajo
super().saludar() # ejecutamos el método de Persona .saludar() y agregamos más cosas a este método
print('Estoy cursando:')
self.curso.resumen()
def estudiar(self, dato): # También podemos definir nuevos métodos
self.conocimiento = dato
def __str__(self):
#Devuelve un string representativo del alumno
return f'Alumno {self.nombre_completo()}'
un_alumno = Alumno(curso_python, 'Scott', 'Henderson', 49, '<EMAIL>')
str(un_alumno)
# ## Métodos Estáticos
#
# ¿Qué pasa si no queremos instanciar los objetos a la hora de usarlos? En algunos diseños, tiene sentido utilizar las clases como simples repositorios de métodos. Por ejemplo, si necesito resolver varias operaciones geométricas puedo crear una clase Geometria que contenga todos los métodos necesarios.
#
# Para crear este tipo de métodos en una clase utilizamos el decorador @staticmethod.
#
#
class Geomtria():
"""Resuelve operaciones geométricas"""
@staticmethod
def pendiente(x1,y1,x2,y2):
return ((y2 - y1)/(x2-x1))
@staticmethod
def area_circulo(radio):
return math.pi * (radio**2)
Geomtria.area_circulo(3)
# ## Duck typing y monkey patching
#
# Dos características de la programación orientada a objetos con Python son el duck tiping y el monkey patching. Este tipo de flexibilidad es el que le permitió a Python crecer tanto en su adopción porque reducen la cantidad de palabras que es necesario escribir para desarrollar código, lo cual ahorra tiempo y también disminuyen la complejidad.
#
# ### Duck typing
#
#
# +
class ElHobbit:
def __init__(self,nombre):
self.nombre = nombre
def __len__(self):
return 95022
def saludar(self):
return f'Hola soy {self.nombre}'
el_hobbit = ElHobbit('Frodo')
# -
len(el_hobbit)
mi_str = "Hello World"
mi_list = [34, 54, 65, 78]
mi_dict = {"a": 123, "b": 456, "c": 789}
# <i> “If it walks like a duck, and it quacks like a duck, then it must be a duck.”</i>
#
# Duck typing significa que a diferencia de otros lenguajes, las funciones especiales no están definidas para una lista específica de clases y tipos, si no que se pueden usar para cualquier objeto que las implemente. Esto no es así para la mayoría de los lenguajes.
len(mi_str)
len(mi_list)
len(mi_dict)
len(el_hobbit)
mi_int = 7
mi_float = 42.3
len(mi_int)
len(mi_float)
# ### Monkey Patching
#
# Guerrilla, gorilla, ¿monkey?... Este término viene de uno anterior, "guerrilla patching", que hace referencia a emparchar el código rápido y cuando es necesario.
#
# Se refiere a la posibilidad en Python de sobreescribir clases después de haberlas instanciado y por qué no también la funcionalidad de los módulos.
el_hobbit.saludar()
def saludo_largo(self):
return f'Hola mi nombre es {self.nombre}'
ElHobbit.saludar = saludo_largo
el_hobbit.saludar()
# Esto es especialmente útil cuando queremos sobre-escribir ligeramente módulos hechos por terceros (¡o por nosotros mismos en otro momento!)
import math
math.pi
math.pi = 2
math.pi
# Los cambios se sobre-escriben si REINICIAMOS EL KERNEL y volvemos a importar el módulo
import math
math.pi
| Introduccion/4_Intro_Poo/intro-poo-solucion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7AdJo7mYXw68"
# # Setup and Imports
# We will download the necessary libraries and embedding models
# + id="KXxJYKamcB3-"
# !pip install ratelimit
# !pip install flair
# !pip install transformers
# !pip install rgf_python
# !pip install textstat
# !pip install pytorch-tabnet
# !pip install shap
# !python -m spacy download en_core_web_lg
# !wget https://conceptnet.s3.amazonaws.com/downloads/2019/numberbatch/numberbatch-en-19.08.txt.gz
# !gzip -d ./numberbatch-en-19.08.txt.gz
# !wget https://www.dropbox.com/s/1m0ofr06zy0nnbm/gpt2glecmed.zip?dl=0
# !unzip gpt2glecmed.zip\?dl\=0
# + id="voJ3xqU9XX-3" colab={"base_uri": "https://localhost:8080/"} outputId="e27457d7-7b1a-4874-dd0e-487a1b3e1f58"
import pandas as pd
import numpy as np
import regex as re
import requests
from nltk.corpus import wordnet as wn
import nltk
from nltk.stem import SnowballStemmer
from tqdm.notebook import tqdm
import spacy
import en_core_web_lg
import gensim.downloader
from flair.embeddings import TransformerDocumentEmbeddings
from flair.data import Sentence
from scipy import spatial
from json import JSONDecodeError
from sklearn.metrics.pairwise import cosine_similarity
from sklearn.decomposition import PCA
from ratelimit import limits, sleep_and_retry
from flair.embeddings import TransformerWordEmbeddings,TransformerDocumentEmbeddings,WordEmbeddings,FlairEmbeddings
from flair.data import Sentence
nlp =en_core_web_lg.load()
stemmer = nltk.stem.SnowballStemmer('english')
nltk.download('wordnet')
# + [markdown] id="ivCDo35h26rh"
# ## Read and Preprocess the corpus
# + id="nV9g-ZxN9lzm"
from preprocess import read_text,preprocess_column
df=read_text('./464katz_mets_nader.txt')
df.label=df.label.astype(int)
df['processed_metaphor']=preprocess_column(df.M.values)
# + [markdown] id="o88frEg7OulK"
# # Document based features
# + [markdown] id="pZmC3u2xjp7L"
# In this section, we're going to use an ensemble of transformer based document embedding to embed the metaphors, then use the concatenated vector to predict the binary target (literary/non-literary)
# The ensemble uses transformers fine tuned on different applications, this includes: part-of-speech tagging, language modeling and sentiment classifiaction. The intuition behind this is that finetuned transformers on different task would encompass different semantic features of the metaphors.
# + id="TzmA0GiN07Kl"
from flair.embeddings import TransformerDocumentEmbeddings
from flair.data import Sentence
# compute the embeddings for each model and stack them together
docs_e=[]
models=['vblagoje/bert-english-uncased-finetuned-pos','gpt2-xl','facebook/bart-large-cnn','bhadresh-savani/distilbert-base-uncased-emotion']
lengths=[]
for model in models:
doc_embedding = TransformerDocumentEmbeddings(model)
#store the embedding length for later usage
lengths.append(doc_embedding.embedding_length)
doc_e=[]
for d in tqdm(df.M):
sent=Sentence(d)
doc_embedding.embed(sent)
doc_e.append(sent.embedding.detach().cpu().numpy())
docs_e.append(doc_e)
stacked_embeddings=docs_e[0]
for d in docs_e[1:]:
stacked_embeddings=np.concatenate((stacked_embeddings,d),axis=1)
# + id="FDgXSl1j6pDp"
X=np.array(stacked_embeddings)
y=df.label.values
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="qxU9UM76Dci0" outputId="e3646167-3178-45c7-aebe-461fdf2e3b53"
from utils import plot_auc_cv
plot_auc_cv(LogisticRegressionCV(max_iter=2000),X,y)
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["b44f7f57f85646c1a0b819b2fda63598", "2a7c591146f54fdaafeaebcc553be0fe", "1fa30cd8cbeb48e48f9b7362a8c80ff2", "62f7bf0bcd2c4c7ca80993d86881a5f9", "d702044b49354f43b782de5f32b72826", "33ad2a4bad3a4110854d478db82c1a2d", "eb7b6769afe44542ad84aa596d755462", "84966c09019c43b78ef009810eda738a", "c4f6ec09eca141708a2d8ed30c9ee08d", "e2c3c0ffad1c4bb6ae02c174caf3a56c", "9e899ad616604372b3b7d34a9e88aaa4"]} id="JOoVV_9yoEji" outputId="803e9653-495b-4681-a8b2-c70a716fa9c5"
from classifiers import classifiers_auc
classifiers_auc(X,y,2)
# + [markdown] id="pK8FmB1_3Du3"
# On 2 cross validation iterations only eg. only half of the data is used for training, the model can reach 0.8445 AUC.
# + [markdown] id="wwlv9cDXJCpQ"
# ### visualizing embedding model importance
# this section fits an ERT to the ensembled Transformer embeddings, and then aggregates the feature importance for each transformer model.
# We note that gpt2-xl contains the most information about literariness of metaphors.
# + colab={"base_uri": "https://localhost:8080/", "height": 485} id="E6Qz7zhXI-kF" outputId="6313928b-3189-4dbc-da6f-772526d0065a"
from utils import plot_hbar
clf=ExtraTreesClassifier(n_estimators=1000,max_depth=15)
clf.fit(X,y)
print("ERT Fitness: ",clf.score(X,y))
feature_importances=clf.feature_importances_
init=0
model_names=['bert-english-uncased-finetuned-pos','gpt2-xl','bart-large-cnn','distilbert-base-uncased-emotion']
#aggregate feature importance for each transformer
model_importances=[]
for i in np.cumsum(lengths):
model_imp=np.sum(feature_importances[init:i])
model_importances.append(model_imp)
init=i
plot_hbar(data)
# + [markdown] id="PND9Q1ZUTePx"
# # Flair Classifier
# In this section we train a flair classifier with the gpt2-medium finetuned document embedding, we report 5 cross validation results on the dataset, and notice the surprising good performance (excellent auc .90-1) of this classifier.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["9692036adc964ef89d87329e2bcab2a0"]} id="50Azr_xXBWml" outputId="fdd8734f-f60a-47ba-fbf9-89db1b71504e"
from flair.data_fetcher import NLPTaskDataFetcher
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import accuracy_score,auc,roc_auc_score
from flair.trainers import ModelTrainer
from flair.models import TextClassifier
from flair.datasets import CSVClassificationCorpus
from pathlib import Path
import sklearn.metrics as metrics
import os
import matplotlib.pyplot as plt
tprs = []
aucs = []
mean_fpr = np.linspace(0, 1, 100)
fig, ax = plt.subplots(figsize=(8,8))
column_name_map = {0: "text", 1: "label"}
label_map={'0':'nonliterary','1':'literary'}
skf = StratifiedKFold(n_splits=5)
document_embedding=TransformerDocumentEmbeddings('./gpt2_medium_glec/')
model_results=[]
X=df.M.values
y=df.label.values
for train_index, test_index in tqdm(skf.split(X, y)):
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
tdf=pd.DataFrame({0:X_train,1:y_train.astype(str)})
tdf[1]=tdf[1].map(label_map)
tdf.to_csv('./train.csv',index=False)
tdf2=pd.DataFrame({0:X_test,1:y_test.astype(str)})
tdf2[1]=tdf2[1].map(label_map)
tdf2.to_csv('./test.csv',index=False)
# corpus = NLPTaskDataFetcher.load_classification_corpus(Path('./'), test_file='test.csv', train_file='train.csv')
corpus=CSVClassificationCorpus(Path('./'),column_name_map,label_type='class',skip_header=True)
classifier = TextClassifier(document_embedding,label_type='class', label_dictionary=corpus.make_label_dictionary('class'), multi_label=False)
trainer = ModelTrainer(classifier, corpus)
trainer.train('./', max_epochs=10,mini_batch_size=4,learning_rate=.01,anneal_against_dev_loss=True)
model_score=[]
for s in X_test:
s=Sentence(s)
classifier.predict(s)
score=(s.labels[0].score if s.labels[0].value =='literary' else 1-s.labels[0].score)
model_score.append(score)
fpr, tpr, thresholds = metrics.roc_curve(y_test, model_score)
interp_tpr = np.interp(mean_fpr, fpr, tpr)
interp_tpr[0] = 0.0
tprs.append(interp_tpr)
roc_auc=roc_auc_score(y_test,model_score)
model_results.append(roc_auc)
aucs.append(roc_auc)
os.remove("./dev.tsv")
os.remove("./test.tsv")
os.remove("./loss.tsv")
os.remove("training.log")
os.remove("final-model.pt")
os.remove("./best-model.pt")
os.remove("./weights.txt")
ax.plot([0, 1], [0, 1], linestyle="--", lw=2, color="r", label="Chance", alpha=0.8)
mean_tpr = np.mean(tprs, axis=0)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
std_auc = np.std(aucs)
ax.plot(
mean_fpr,
mean_tpr,
color="b",
label=r"Mean ROC (AUC = %0.2f $\pm$ %0.2f)" % (np.mean(aucs), np.std(aucs)),
lw=2,
alpha=0.8,
)
std_tpr = np.std(tprs, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
ax.fill_between(
mean_fpr,
tprs_lower,
tprs_upper,
color="grey",
alpha=0.2,
label=r"$\pm$ 1 std. dev.",
)
ax.set(
xlim=[-0.05, 1.05],
ylim=[-0.05, 1.05],
title="Receiver operating characteristic example",
)
ax.legend(loc="lower right")
plt.show()
# + [markdown] id="FTo0ok69OomP"
# # Word based embeddings
# + [markdown] id="wXOq6S2glNNy"
# This section investigates word level features, the features include the word tenor vehicle relation with different embedding models, the cosine similarity between the metaphor words from different embedding models, sentiment and confidence score, ease of readness, and the gltr word statistics.
# There are 14 features in total which makes this approach explainable.
# + [markdown] id="srZWFlnqCm3a"
# ### Featurize the dataset
# + id="7Dbx5iBeX9td"
from featurize import featurize
labels=[
'gltr_mean','gltr_std','gltr_max','gltr_cond_p','flair_mean','flair_std','flair_explaind_sum','conceptnet_t/v_sim','sentiment','sentiment_confidence','flescher_reading_index','ARI'
]
X=featurize(df)
X1=np.array(X)
X2=df[df.columns[4:]].values
y=df.label.values
# + id="q3sntQAdVeaZ"
df_met = pd.DataFrame(data=X1, index=np.arange(X1.shape[0]), columns=labels)
df_met['label']=y
df_met.to_csv("metaphors_data.csv")
# + [markdown] id="TWgvGgBUCpZq"
# ### Tabnet Benchmark using the generated features
# + id="nI6jzP7NUPjE" colab={"base_uri": "https://localhost:8080/", "height": 236, "referenced_widgets": ["b4bc35725eb940c4a76177545dcbcaec", "fa7824ee3ab0405c8e17929d37b0a16b", "858e5fae0af44302b17dd4eb60ea5be0", "2c92f8cc70a047aca44f09b3e27a55ef", "f15edaccea78423eac1ede72306abca0", "8bf96630a084483d911824b527cb9f68", "ccf3085df0f84d438b1cc1a5ee0b0417", "c35a4c41d82f4d70a8ee7ee83cfc608e", "adc5a5abdf0c44a0ad4106a5e97d1262", "627f033bb55541938ef98ead724b8996", "80d0c5749c3e41299b4ad18f1d74fd72"]} outputId="cdd3539f-b7c2-430c-b069-9287d7ae58e2"
from classifiers import tabnet_auc
results,feature_importances=tabnet_auc(X1,y,10)
# + colab={"base_uri": "https://localhost:8080/", "height": 468} id="XNInQ3UT_K9d" outputId="0f2973e1-f7e7-4f6a-80a3-0558fc3fcf67"
data = {k:v for k,v in zip(labels,feature_importances)}
plot_hbar(data)
# + [markdown] id="51vP7TZsCtk0"
# ### Benchmarking different machine learning models
# + colab={"base_uri": "https://localhost:8080/", "height": 202, "referenced_widgets": ["4d3bf43208fd43e68011449a354747c9", "cdaeec99b8f248bb9a182d406b6a2535", "a4b0a81e7bfa48659289932fddb883fd", "af3d455709cf4a8a9aa33f19d3858126", "26242e1bb10c46b0911c93c4d5bec937", "8e4e96cb7e984cf2be21b4e9bae0ac77", "43a10aef334f48a08c7005a379345b80", "<KEY>", "9e9d08cf47d348708b1e7d7ddec6962f", "245b192ad00a4eff9298510383a8310d", "ef2a00e5f13f4f048f236aa571dc1f73"]} id="-xkNuFn-vkQU" outputId="5d9a92c8-2e45-40a3-f7c7-ed1969d5a9ea"
from classifiers import classifiers_auc
classifiers_auc(X1,y)
# + [markdown] id="trew64KLkLh8"
# # Visualizing Feature importances and effects for ERT and LogisticRegression classifiers
# + [markdown] id="ReUk1nmhCzOB"
# Since ERT and Logisitic regression are the best performers , we will analyze their prediction and the feature impact on the models predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 468} id="hxu2ZH--rTzo" outputId="48c9167a-5d6b-443f-817e-9dd54c0e043d"
scl=StandardScaler()
X=scl.fit_transform(X1)
clf=ExtraTreesClassifier(n_estimators=1000,max_depth=6).fit(X,y)
data = {k:v for k,v in zip(df_met.columns[:-1],clf.feature_importances_)}
plot_hbar(data)
# + [markdown] id="NzNiqX83xVAO"
# #### LogisticRegression Feature Impact
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="Sau2maV3-90K" outputId="a0c4e99e-68d6-4d41-c3ed-cfb730c32dd5"
scl=StandardScaler()
X=scl.fit_transform(X1)
clf=LogisticRegressionCV().fit(X,y)
explainer = shap.LinearExplainer(clf, X)
shap_values = explainer.shap_values(X)
shap.summary_plot(shap_values, X, feature_names=df_met.columns[:-1])
# + [markdown] id="yotZrBEwyqCL"
# ### Insights:
# 1- Flescher reading index is informative for literary metaphors detection, literary metaphors tend to be less confusing.
# 2- Literary metaphors tend to have greater mean embedding distance, which means they spann wider concepts.
# 3- Literary metaphors tend to be more surprising for a transformer model
# + [markdown] id="ineUlCKYMJ-k"
# # Regression Tasks
# The 14 features seem to only somewhat work for SRL, MGD, ALT and MET ratings only.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["1f0368c2cd8c44ff9d9bddc7570aadd4", "d5b21102b48f42dd8553d9697d7e2deb", "96c53f8df56f42ad936f4a6ae5c68466", "61d593fe63104552a296e9932bac5cc5", "c4eee7b6ad334924b56fdef65693fe84", "64b6880e3ed840f0ad6b37e3451549ea", "e1a03fda9c00498f89bda6a52a0f3947", "<KEY>", "0f3e964939de40e78b3cda675ce247af", "<KEY>", "<KEY>"]} id="WVK2MByRMI8P" outputId="6e7dfeec-57da-42d8-9985-0bdafb71d075"
columns=["CMP","ESI","MET","MGD","SRL","MIM","IMS","IMP","FAM","ALT"]
from sklearn.model_selection import StratifiedKFold
from sklearn.ensemble import GradientBoostingRegressor,RandomForestRegressor
from sklearn.naive_bayes import GaussianNB
from sklearn.neighbors import KNeighborsRegressor
from sklearn.linear_model import LogisticRegressionCV
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import StratifiedKFold,KFold
from sklearn.metrics import accuracy_score,auc,roc_auc_score,mean_absolute_error
from sklearn.svm import SVR
from xgboost import XGBRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.neural_network import MLPRegressor
from sklearn.linear_model import RidgeCV,LinearRegression
for col in columns:
y=df[col].values
skf = KFold(n_splits=10)
models=[
("SVM",SVR(kernel='rbf')),
("LinearRegression",LinearRegression()),
("Ridge",RidgeCV()),
('Knn',KNeighborsRegressor(n_neighbors=3)),
("ERT",ExtraTreesRegressor(n_estimators=1000,max_depth=9)),
]
model_results={k[0]:[] for k in models}
for train_index, test_index in tqdm(skf.split(X1, y)):
X_train, X_test = X1[train_index], X1[test_index]
y_train, y_test = y[train_index], y[test_index]
scl=StandardScaler()
X_train=scl.fit_transform(X_train)
X_test=scl.transform(X_test)
for m in models:
m[1].fit(X_train,y_train)
model_results[m[0]].append(mean_absolute_error(y_test,m[1].predict(X_test)))
print("Predicting: ",col)
for label,res in model_results.items():
print(label,' mean absolute error: ',np.mean(res))
| src/Literary_Metaphor_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Recommendation to clients
# Based on the analysis below, the cryptocurrencies can be clustered. I recommend using three to five clusters.
# Import dependencies
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
# Read in raw data
raw_crypto_data_df = pd.read_csv("crypto_data.csv")
raw_crypto_data_df
# Filter by IsTrading == True
traded_crypto_df = raw_crypto_data_df.loc[raw_crypto_data_df.IsTrading == True, :]
traded_crypto_df.head()
# Remove IsTrading column
traded_crypto_df = traded_crypto_df.drop(columns="IsTrading")
traded_crypto_df.head()
# Remove all rows with at least one null value
traded_crypto_not_null_df = traded_crypto_df.dropna()
traded_crypto_not_null_df.head()
# Filter for only mined currencies
mined_crypto_df = traded_crypto_not_null_df.loc[traded_crypto_not_null_df.TotalCoinsMined > 0, :]
mined_crypto_df.head()
# Remove columns that do not contribute to data analysis
mined_crypto_clean_df = mined_crypto_df.drop(columns = ["Unnamed: 0", "CoinName"])
mined_crypto_clean_df.head()
mined_crypto_clean_df.shape
# Transforme categorical data to numeric
mined_crypto_dummies_df = pd.get_dummies(data=mined_crypto_clean_df, columns=["Algorithm", "ProofType"])
mined_crypto_dummies_df.head()
mined_crypto_dummies_df.shape
# ### Observation
# After transforming the categorical data to numeric, the dataframe changed for 532 rows and four columns to the same number of rows but 98 columns.
# Scaling the data to prevent any one feature from having undue weight
scaler = StandardScaler()
scaled_data = scaler.fit_transform(mined_crypto_dummies_df)
scaled_data_df = pd.DataFrame(scaled_data, columns=mined_crypto_dummies_df.columns)
scaled_data_df
# Applying PCA to reduce the feature dimentionality
pca = PCA(n_components=0.9)
crypto_pca = pca.fit_transform(mined_crypto_dummies_df)
crypto_pca.shape
# ### Observation
# The number of features decreased from 98 to two.
# Apply t-SNE to further reduce the dimentionality of the dataset
tsne = TSNE(learning_rate=35)
tsne_features = tsne.fit_transform(crypto_pca)
tsne_features.shape
# ### Observation
# The number of features remained the same at two.
# +
# The first column of transformed features
x = tsne_features[:,0]
# The second column of transformed features
y = tsne_features[:,1]
# Visualize the clusters
plt.scatter(x, y)
plt.show()
# -
# ### Observation
# Upon visitual inspection, there seems to be five clusters.
# +
# Finding the best k
inertia = []
k = [1,2,3,4,5,6,7,8,9,10]
for i in k:
km = KMeans(n_clusters=i, random_state=0)
km.fit(tsne_features)
inertia.append(km.inertia_)
# +
# Create a dataframe and plot the elbow curve
elbow_data = {"k": k, "inertia": inertia}
elbow_df = pd.DataFrame(elbow_data)
plt.plot(df_elbow["k"], elbow_df["inertia"])
plt.xticks(range(1,11))
plt.xlabel("Number of clusters")
plt.ylabel("Inertia")
plt.show()
# -
# ### Observation
# The "elbow" of the plot seems to be at k=3; however, given the steepness of the line at k=3, I would be inclined to propose that the best k is 5.
| crypto_clusters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example 1: Setting Up Your Archive
#
# This first example runs through the basics of getting started with `MetaViz`. This includes a run-down on the organization of the code, an introduction to the `config.py` file, and instantiating an `Archive` object that you can begin playing with.
#
# More complex interactions with your `Archive` will be covered in Example 2.
# ## Setting up the `config` file
#
# Before we can begin using the tools and routines available in `MetaViz`, we need to tell it a few details about our collection. The easiest way to do this is by updating the `config.py` file inside the `MetaViz` folder prior to running `setup.py`, which will be read whenever the package is imported.
#
# Inside the `config` file, you will find the following variables which should be updated to reflect your file collection:
#
# - **`CollectionPath`**: Absolute path to the folder containing your media collection
#
# - **`ExcludeFolders`**: Optional list of folders *inside* `CollectionPath` which are to be excluded from `MetaViz` processing. Every folder named one of the terms in this list will be ignored.
#
# - **`csvPath`**: Absolute path to the folder in which to save intermediate `.csv` files containing the metadata from your collection. This is done for both speed and convenience, as will be explained later.
#
# - **`fields`**: List of XMP/EXIF metadata fields which are of interest in your collection. Usually, files will contain many more metadata fields than are of any real interest most of the time, and this list allows you to refine the metadata you save to the fields you care about. *(Note: If you ever plan to use `MetaViz` to update the metadata in your files, you should be specific when specifying these fields, e.g. writing "XMP-dc:Title" instead of just "Title")*
#
# - **`BackupPath`**: (Optional) Absolute path to the folder in which to save compressed backup files of your media collection. Not required for using the package, but I **strongly recommend** you make sure to have backups of all your files! I've never had an issue with these functions causing trouble, but that doesn't mean it couldn't happen.
#
# Lastly, the `config` file will also loop through `CollectionPath` and store the names of all the subfolders in your collection.
#
# Whenever the `config` file has been updated, you'll want to run (or re-run) `python setup.py install` to make sure details have been updated in the Python directory.
# ## Import `MetaViz`
#
# Now that everything has been setup, let's import the `MetaViz` package under the name `mv`. Let's also instantiate an `Archive` object, which will make use of the info stored in the newly-updated `config` file. You can think of this object a bit like a photo album, so we'll give it the name `album`.
import MetaViz as mv
album = mv.Archive()
# Most tools and plotting routines are accessible directly under `mv`, which we will return to later. However, for now, let's focus on getting everything setup for those functions, which we will do via the `album` object.
#
# First, let's check and make sure we've properly read in the details from the updated `config` file. If the `CollectionPath` differs from what was entered earlier, check and make sure things were updated properly.
print(album.CollectionPath)
# ## Initializing with ExifTool
#
# *A bit of background*: In order to access the metadata inside our media files, `MetaViz` relies on an installation of the perl package [`exiftool`](https://exiftool.org/), which we interface with via the command line behind the scenes. The way we do this is not particularly sophisticated, but it gets the job done. Because calling `exiftool` to access file metadata is computationally expensive, we only do this once ahead of time, after which we save all of the metadata fields of interest (specified in `config.fields`) inside a folder of `.csv` files (at the location specified in `config.csvPath`). Then, to make use of this metadata in all our functions, we simply use `Pandas` to quickly read all that information into a DataFrame.
#
# Therefore, in order to access the metadata in your files, we first need to utilize `exiftool` and extract a bunch of metadata from your files. We do this by calling `UpdateCSV()`:
album.UpdateCSV()
# Behind the scenes, what we're doing here is calling exiftool repeatedly with the following bash command:
# ```
# exiftool -csv foldername > csvname
# ```
# which is run for every subfolder inside `CollectionPath`. We break this up by subfolder (rather than doing this recursively with `-r`) because it offers greater flexibility in how we read and update metadata later.
#
# If your collection is unchanging through time, running this function once will be enough. If your collection gets updated through time, this function needs to be re-run to reflect that, at least for the subfolders which are changing. To update only specific subfolders, specify them as a list of strings, i.e. `UpdateCSV(['path/to/subfolder1', 'path/to/subfolder2'])`
#
# ## Creating a backup
#
# Before getting carried away with anything else in this package, now seems like another good time to recommend **backing up your files**. For simplicity, we've included a function for doing just that inside the `album` object.
#
# Here, we will zip up each folder one level below `CollectionPath` and store those backups inside the folder specified in `config.BackupPath`.
#
# *Note: This built-in function may have difficulty compressing files with timestamps older than 1980, which is the beginning of the zip epoch*
album.CreateBackup()
# ## Updating metadata
#
# Should the time come that you wish to update the metadata inside your actual media files, we offer a function similar to `UpdateCSV` in reverse: given some modified `.csv` files, the function `UpdateMetadata()` can update the metadata inside your raw files. This again makes use of `exiftool` behind the scenes, using the following bash command:
#
# ```
# exiftool -csv=csvname foldername -overwrite_original_in_place -P -F
# ```
#
# For details on these default flags, please see the [`exiftool` FAQ](https://exiftool.org/faq.html). The gist is that, rather than copying your files, we are updating them in place with their existing metadata, and fixing any minor errors should they arise.
#
# Note that this function accepts a list of specific subfolders to update, which can be much faster (and safer!) than the default of updating every file.
album.UpdateMetadata()
# ## Looking forward: Structure of the code
#
# Now that we've finished setting things up and are ready to begin searching and plotting, I think it's a good idea to give a brief overview of the structure of the code. Scripts fall roughly into the following heirarchy, which will show up in later examples:
#
# 0. `config` and `tools`: At the base of the package are the config file and several scripts of all-purpose tools, used throughout the rest of the package.
#
# 1. `Archive` class: One step above is these base-level scripts is the `Archive` class we've been using to interface with our collection metadata. Interacting with exiftool and searching through metadata are all achieved by methods inside the `Archive` class.
#
# 2. `plot_` scripts: Plotting routines built on top of the previous two levels. These make use of outputs from the previous two levels, but generally don't interface with metadata directly, unless done through an `Archive` object. These are broken into the category of information they display, such as `_timeseries` or `_statistics`
#
# Lastly, we've seperately made a `scripts` folder, which includes some useful non-Python bash scripts for the exiftool commands used behind the scenes here, as well as a few others I've found useful.
| examples/1_ArchiveSetup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # ML job market segmentation in India
# In this project we will be using a kaggle dataset to perform following tasks:-
#
# 1.To enable the company to recognize best fit candidate for ml engineer/data analyst with respect to his /her skills.
#
# 2.To analyse the ml job market in India and outline the most optimal segment to apply for ml jobs.
# # **Import dependencies**
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# # Import dataset
df = pd.read_csv('jobs1.csv')
# # Exploratory data analysis
df.head()
# checking for null values
df.isnull().sum()
# checking columns of our dataset
df.columns
df.shape
df.info()
# # Feature Engineering
# Dropping irrelevant columns
df.drop(['Job URL','Salary','Posted','Company URL'],axis = 1,inplace = True)
df.head()
# +
df.Location.apply(pd.Series).stack().str.strip().value_counts()[:10].plot.pie(figsize=(12,10),startangle=50,autopct='%1.1f%%',fontsize=15)
plt.title("Location Wise Machine Learning Jobs",fontsize=30)
circle = plt.Circle((0,0),0.72,color='gray', fc='white',linewidth=1.25)
fig = plt.gcf()
fig.gca().add_artist(circle)
plt.axis('equal')
plt.show()
# -
# Company wise distribution of jobs
df["Company Name"].value_counts()[:10].plot.pie(figsize=(12,10),explode=[0.03,0.04,0.05,0.05,0.05,0.05,0.05,0.05,0.05,0.05],shadow=True, startangle=40,autopct='%1.1f%%',fontsize=20)
plt.title("Companies wise job postings",fontsize=35)
plt.show()
#Experience wise distribution of jobs
df["Exp"].value_counts()[:10].plot.barh(figsize=(8,5),fontsize=13,color="b")
plt.xlabel("No.of Vacancies",fontsize=18)
plt.ylabel("Experience",fontsize=18)
plt.show()
# Job wise distribution
df["Job Title"].value_counts()[:10].plot.bar(figsize=(8,5),fontsize=10,color="y")#(figsize=(10,10),explode=[0.05,0.04,0.05,0,0,0,0,0,0,0],shadow=True, startangle=50,autopct='%1.1f%%')
plt.xticks(rotation=45,ha='right')
plt.title("Machine Learning Roles",fontsize=20)
plt.ylabel("No of Vacancies",fontsize=15,rotation=90)
plt.xlabel("Roles",fontsize=15)
plt.show()
# Skills wise distribution
df.Skills.apply(pd.Series).stack().value_counts()[:32].plot(kind="bar",figsize=(18,6),fontsize=15,color="r")
plt.xticks(rotation=50,ha='right')
plt.ylabel("No.of Vacancies",fontsize=20)
plt.xlabel("Top Skills for Machine Learning",fontsize=25)
plt.show()
# Feature scaling
from sklearn.preprocessing import LabelEncoder
le=LabelEncoder()
df['Job Title']=le.fit_transform(df['Job Title'])
df['Company Name']=le.fit_transform(df['Company Name'])
df['Exp']=le.fit_transform(df['Exp'])
df['Location']=le.fit_transform(df['Location'])
df['Skills']=le.fit_transform(df['Skills'])
df.head()
# # Finding Number of Cluster
# Using elbow method
X=df.values
from sklearn.cluster import KMeans
wcss=[]
for i in range(1,11):
kmeans=KMeans(n_clusters=i,init='k-means++',random_state=42)
kmeans.fit(X)
wcss.append(kmeans.inertia_)
plt.plot(range(1,11),wcss)
plt.title('Elbow method')
plt.xlabel('no of clusters')
plt.ylabel('wcss')
plt.show()
# # Fitting K-means algorithm
# +
# It is seen that 4 is ideal number of clusters for our dataset
kmeans=KMeans(n_clusters=4,init='k-means++',random_state=42)
y_kmeans=kmeans.fit_predict(X)
# +
# Visualizing clusters
plt.scatter(X[y_kmeans==0,0],X[y_kmeans==0,1],s=20,c='yellow',label='cluster1')
plt.scatter(X[y_kmeans==1,0],X[y_kmeans==1,1],s=20,c='blue',label='cluster1')
plt.scatter(X[y_kmeans==2,0],X[y_kmeans==2,1],s=20,c='green',label='cluster1')
plt.scatter(X[y_kmeans==3,0],X[y_kmeans==3,1],s=20,c='cyan',label='cluster1')
# -
| Machine Learning job market segmentation in India( Feynn Labs final project).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import the Libraries
from keras.models import Sequential
from keras.layers import Conv2D
from keras.layers import MaxPool2D
from keras.layers import Flatten
from keras.layers import Dense
# Initialising the CNN
classifier = Sequential()
# Convolution
classifier.add(Conv2D(32, (3, 3), input_shape = (64, 64, 3), activation = 'relu'))
classifier.add(Conv2D(32, (3, 3), input_shape = (64, 64, 3), activation = 'relu'))
classifier.add(Conv2D(32, (3, 3), input_shape = (64, 64, 3), activation = 'relu'))
# Pooling
classifier.add(MaxPool2D(pool_size = (2, 2)))
classifier.add(Conv2D(32, (3, 3), activation = 'relu'))
classifier.add(MaxPool2D(pool_size = (2, 2)))
# Flattening
classifier.add(Flatten())
# +
# Full ANN connection
classifier.add(Dense(units = 128, activation = 'relu'))
classifier.add(Dense(units = 128, activation = 'relu'))
classifier.add(Dense(units = 128, activation = 'relu'))
classifier.add(Dense(units = 1, activation = 'sigmoid'))
# Compiling the CNN
classifier.compile(optimizer = 'SGD', loss = 'binary_crossentropy', metrics = ['accuracy'])
# +
# This block is part of Activity 3
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
test_datagen = ImageDataGenerator(rescale = 1./255)
training_set = train_datagen.flow_from_directory('dataset/training_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
test_set = test_datagen.flow_from_directory('dataset/test_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
classifier.fit_generator(training_set,
steps_per_epoch = 10000,
epochs = 2,
validation_data = test_set,
validation_steps = 2500)
# +
import numpy as np
from keras.preprocessing import image
new_image = image.load_img('D:/test_image_1.jpg', target_size = (64, 64))
new_image = image.img_to_array(new_image)
new_image = np.expand_dims(new_image, axis = 0)
result = classifier.predict(new_image)
training_set.class_indices
if result[0][0] == 1:
prediction = 'It is a Dog'
else:
prediction = 'It is a Cat'
print(prediction)
| Lesson07/Exercise21.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="bOChJSNXtC9g" colab_type="text"
# # Data and Models
# + [markdown] id="OLIxEDq6VhvZ" colab_type="text"
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/logo.png" width=150>
#
# So far we've seen a varity of different models on different datasets for different tasks (regression/classification) and we're going to learn about even more algorithsm in subsequent lessons. But we've ignored a fundamental concept about data and modeling: quality and quantity. In a nutshe, a machine learning model consumes input data and produces predictions. The quality of the predictions directly corresponds to the quality and quantity of data you train the model with; garbage in, garbage out.
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/nutshell.png" width=500>
#
#
#
#
# + [markdown] id="FLH7kzZl8wnf" colab_type="text"
# # Set Up
# + [markdown] id="qAE9BjMH8x4q" colab_type="text"
# We're going to go through all the concepts with concrete code examples. We'll first synthesize some data to train our models on. The task is to determine wether a tumor will be benign (harmless) or malignant (harmful) based on leukocyte (white blood cells) count and blood pressure.
# + id="m0nzLDcVXJTx" colab_type="code" colab={}
# Load PyTorch library
# !pip3 install torch torchvision
# + id="N9uu2nngKDrW" colab_type="code" colab={}
from argparse import Namespace
import collections
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import random
import torch
# + id="gPHmsndLdUOH" colab_type="code" colab={}
# Set Numpy and PyTorch seeds
def set_seeds(seed, cuda):
np.random.seed(seed)
torch.manual_seed(seed)
if cuda:
torch.cuda.manual_seed_all(seed)
# + id="0-dXQiLlTIgz" colab_type="code" outputId="3f81662d-d58e-4667-e0bf-cedd5747b5f6" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Arguments
args = Namespace(
seed=1234,
cuda=False,
shuffle=True,
data_file="tumors.csv",
reduced_data_file="tumors_reduced.csv",
train_size=0.75,
test_size=0.25,
num_hidden_units=100,
learning_rate=1e-3,
num_epochs=100,
)
# Set seeds
set_seeds(seed=args.seed, cuda=args.cuda)
# Check CUDA
if not torch.cuda.is_available():
args.cuda = False
args.device = torch.device("cuda" if args.cuda else "cpu")
print("Using CUDA: {}".format(args.cuda))
# + [markdown] id="RV2IddoZde-r" colab_type="text"
# # Data
# + id="5wDazzQdaoy2" colab_type="code" colab={}
import re
import urllib
# + id="GbsXoFVgdh6K" colab_type="code" colab={}
# Upload data from GitHub to notebook's local drive
url = "https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/data/tumors.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(args.data_file, 'wb') as fp:
fp.write(html)
# + id="y6LNWmoidh8q" colab_type="code" outputId="dd5f8f63-2d55-4d56-a126-1fe1a33d7b42" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Raw data
df = pd.read_csv(args.data_file, header=0)
df.head()
# + id="YVo6CuZLC3h2" colab_type="code" colab={}
def plot_tumors(df):
i = 0; colors=['r', 'b']
for name, group in df.groupby("tumor"):
plt.scatter(group.leukocyte_count, group.blood_pressure, edgecolors='k',
color=colors[i]); i += 1
plt.xlabel('leukocyte count')
plt.ylabel('blood pressure')
plt.legend(['0 - benign', '1 - malignant'], loc="upper right")
plt.show()
# + id="nXFUmnfte6z6" colab_type="code" outputId="c65d8763-affb-446a-c0fe-e4144138a891" colab={"base_uri": "https://localhost:8080/", "height": 361}
# Plot data
plot_tumors(df)
# + id="237OzHqlNQ-D" colab_type="code" colab={}
# Convert to PyTorch tensors
X = df.as_matrix(columns=['leukocyte_count', 'blood_pressure'])
y = df.as_matrix(columns=['tumor'])
X = torch.from_numpy(X).float()
y = torch.from_numpy(y.ravel()).long()
# + id="0pahDv9WLD2S" colab_type="code" outputId="5f560a0a-b815-4815-958d-d09c56985312" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Shuffle data
shuffle_indicies = torch.LongTensor(random.sample(range(0, len(X)), len(X)))
X = X[shuffle_indicies]
y = y[shuffle_indicies]
# Split datasets
test_start_idx = int(len(X) * args.train_size)
X_train = X[:test_start_idx]
y_train = y[:test_start_idx]
X_test = X[test_start_idx:]
y_test = y[test_start_idx:]
print("We have %i train samples and %i test samples." % (len(X_train), len(X_test)))
# + [markdown] id="owLnzReJJdpj" colab_type="text"
# # Model
# + [markdown] id="zlPe1lXEJfcA" colab_type="text"
# Let's fit a model on this synthetic data.
# + id="0WhYfDOjJdIV" colab_type="code" colab={}
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
# + id="nTtsFHZY_G45" colab_type="code" colab={}
# Multilayer Perceptron
class MLP(nn.Module):
def __init__(self, input_dim, hidden_dim, output_dim):
super(MLP, self).__init__()
self.fc1 = nn.Linear(input_dim, hidden_dim)
self.fc2 = nn.Linear(hidden_dim, output_dim)
def forward(self, x_in, apply_softmax=False):
a_1 = F.relu(self.fc1(x_in)) # activaton function added!
y_pred = self.fc2(a_1)
if apply_softmax:
y_pred = F.softmax(y_pred, dim=1)
return y_pred
# + id="1kXlfHpPJ5Vq" colab_type="code" colab={}
# Initialize model
model = MLP(input_dim=len(df.columns)-1,
hidden_dim=args.num_hidden_units,
output_dim=len(set(df.tumor)))
# + id="Ncxbef0yJ6pD" colab_type="code" colab={}
# Optimization
loss_fn = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=args.learning_rate)
# + id="srlaBr8oiftE" colab_type="code" colab={}
# Accuracy
def get_accuracy(y_pred, y_target):
n_correct = torch.eq(y_pred, y_target).sum().item()
accuracy = n_correct / len(y_pred) * 100
return accuracy
# + id="Mjg4u-zCK90q" colab_type="code" outputId="68cd90f4-5349-4041-95b3-aa31ebaa8987" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Training
for t in range(args.num_epochs):
# Forward pass
y_pred = model(X_train)
# Accuracy
_, predictions = y_pred.max(dim=1)
accuracy = get_accuracy(y_pred=predictions.long(), y_target=y_train)
# Loss
loss = loss_fn(y_pred, y_train)
# Verbose
if t%20==0:
print ("epoch: {0} | loss: {1:.4f} | accuracy: {2:.1f}%".format(t, loss, accuracy))
# Zero all gradients
optimizer.zero_grad()
# Backward pass
loss.backward()
# Update weights
optimizer.step()
# + id="wHCvuSEaK-2x" colab_type="code" colab={}
# Predictions
_, pred_train = model(X_train, apply_softmax=True).max(dim=1)
_, pred_test = model(X_test, apply_softmax=True).max(dim=1)
# + id="5whE6K0rOmGN" colab_type="code" outputId="2b29834b-0a4e-4514-9c2a-e74b20f40776" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Train and test accuracies
train_acc = get_accuracy(y_pred=pred_train, y_target=y_train)
test_acc = get_accuracy(y_pred=pred_test, y_target=y_test)
print ("train acc: {0:.1f}%, test acc: {1:.1f}%".format(train_acc, test_acc))
# + id="bzFb90SJOmI2" colab_type="code" colab={}
# Visualization
def plot_multiclass_decision_boundary(model, X, y):
x_min, x_max = X[:, 0].min() - 0.1, X[:, 0].max() + 0.1
y_min, y_max = X[:, 1].min() - 0.1, X[:, 1].max() + 0.1
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 101), np.linspace(y_min, y_max, 101))
cmap = plt.cm.Spectral
X_test = torch.from_numpy(np.c_[xx.ravel(), yy.ravel()]).float()
y_pred = model(X_test, apply_softmax=True)
_, y_pred = y_pred.max(dim=1)
y_pred = y_pred.reshape(xx.shape)
plt.contourf(xx, yy, y_pred, cmap=plt.cm.Spectral, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, s=40, cmap=plt.cm.RdYlBu)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
# + [markdown] id="ViwfNFOYRDkm" colab_type="text"
# We're going to plot a white point, which we know belongs to the malignant tumor class. Our well trained model here would accurate predict that it is indeed a malignant tumor!
# + id="_oEf6XRmOsJE" colab_type="code" outputId="aae143af-8d74-4053-ee8e-c9315c18e490" colab={"base_uri": "https://localhost:8080/", "height": 335}
# Visualize the decision boundary
plt.figure(figsize=(12,5))
plt.subplot(1, 2, 1)
plt.title("Train")
plot_multiclass_decision_boundary(model=model, X=X_train, y=y_train)
plt.scatter(np.mean(df.leukocyte_count), np.mean(df.blood_pressure), s=200,
c='b', edgecolor='w', linewidth=2)
plt.subplot(1, 2, 2)
plt.title("Test")
plot_multiclass_decision_boundary(model=model, X=X_test, y=y_test)
plt.scatter(np.mean(df.leukocyte_count), np.mean(df.blood_pressure), s=200,
c='b', edgecolor='w', linewidth=2)
plt.show()
# + [markdown] id="o231eJaQPi5E" colab_type="text"
# Great! We received great performances on both our train and test data splits. We're going to use this dataset to show the important of data quality and quantity.
# + [markdown] id="pZ3rnGH8PtBu" colab_type="text"
# # Data Quality and Quantity
# + [markdown] id="ONRP3WQgR3zc" colab_type="text"
# Let's remove some training data near the decision boundary and see how robust the model is now.
# + id="Sxd2S63EYtxt" colab_type="code" colab={}
# Upload data from GitHub to notebook's local drive
url = "https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/data/tumors_reduced.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(args.reduced_data_file, 'wb') as fp:
fp.write(html)
# + id="sU69PjH3Z4bm" colab_type="code" outputId="6b938dfe-43b7-478d-b0f6-0b37904b8dcd" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Raw reduced data
df_reduced = pd.read_csv(args.reduced_data_file, header=0)
df_reduced.head()
# + id="1OwgEJSsZ4g5" colab_type="code" outputId="61175ac9-5c21-4b96-d116-c5c46b3c241b" colab={"base_uri": "https://localhost:8080/", "height": 361}
# Plot data
plot_tumors(df_reduced)
# + id="r9xlQme0beTY" colab_type="code" colab={}
# Convert to PyTorch tensors
X = df_reduced.as_matrix(columns=['leukocyte_count', 'blood_pressure'])
y = df_reduced.as_matrix(columns=['tumor'])
X = torch.from_numpy(X).float()
y = torch.from_numpy(y.ravel()).long()
# + id="RerzDWJQbeVz" colab_type="code" outputId="9e1c42ac-6e1d-42b4-ae3c-019d8f75d48c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Shuffle data
shuffle_indicies = torch.LongTensor(random.sample(range(0, len(X)), len(X)))
X = X[shuffle_indicies]
y = y[shuffle_indicies]
# Split datasets
test_start_idx = int(len(X) * args.train_size)
X_train = X[:test_start_idx]
y_train = y[:test_start_idx]
X_test = X[test_start_idx:]
y_test = y[test_start_idx:]
print("We have %i train samples and %i test samples." % (len(X_train), len(X_test)))
# + id="JCZ7yDl1OsdU" colab_type="code" colab={}
# Initialize model
model = MLP(input_dim=len(df_reduced.columns)-1,
hidden_dim=args.num_hidden_units,
output_dim=len(set(df_reduced.tumor)))
# + id="-IZ4YOKtSCRk" colab_type="code" colab={}
# Optimization
loss_fn = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=args.learning_rate)
# + id="7NBWLKDISDj8" colab_type="code" outputId="23799fc4-5ccd-416f-fac9-3e6f692b4197" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Training
for t in range(args.num_epochs):
# Forward pass
y_pred = model(X_train)
# Accuracy
_, predictions = y_pred.max(dim=1)
accuracy = get_accuracy(y_pred=predictions.long(), y_target=y_train)
# Loss
loss = loss_fn(y_pred, y_train)
# Verbose
if t%20==0:
print ("epoch: {0} | loss: {1:.4f} | accuracy: {2:.1f}%".format(t, loss, accuracy))
# Zero all gradients
optimizer.zero_grad()
# Backward pass
loss.backward()
# Update weights
optimizer.step()
# + id="uGWbZlhUSFOz" colab_type="code" colab={}
# Predictions
_, pred_train = model(X_train, apply_softmax=True).max(dim=1)
_, pred_test = model(X_test, apply_softmax=True).max(dim=1)
# + id="Gz2Sh4JpSFT9" colab_type="code" outputId="b1653546-a06d-4732-f406-327f4d06b3c8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Train and test accuracies
train_acc = get_accuracy(y_pred=pred_train, y_target=y_train)
test_acc = get_accuracy(y_pred=pred_test, y_target=y_test)
print ("train acc: {0:.1f}%, test acc: {1:.1f}%".format(train_acc, test_acc))
# + id="DmTCz8OnSFRn" colab_type="code" outputId="ea6a32ff-03c9-457f-a1e6-7b11aafba4ff" colab={"base_uri": "https://localhost:8080/", "height": 335}
# Visualize the decision boundary
plt.figure(figsize=(12,5))
plt.subplot(1, 2, 1)
plt.title("Train")
plot_multiclass_decision_boundary(model=model, X=X_train, y=y_train)
plt.scatter(np.mean(df.leukocyte_count), np.mean(df.blood_pressure), s=200,
c='b', edgecolor='w', linewidth=2)
plt.subplot(1, 2, 2)
plt.title("Test")
plot_multiclass_decision_boundary(model=model, X=X_test, y=y_test)
plt.scatter(np.mean(df.leukocyte_count), np.mean(df.blood_pressure), s=200,
c='b', edgecolor='w', linewidth=2)
plt.show()
# + [markdown] id="kdP98xnlbvVn" colab_type="text"
# This is a very scary but highly realistic scenario. Based on our reduced synthetic dataset, we have achieved a model that generalized really well on the test data. But when we ask for the prediction for the same white point earlier (which we known is a tumor), the prediction is now a benign tumor. We would have completely missed the tumor.
#
# **MODELS ARE NOT CRYSTAL BALLS**
# It's so important that before any machine learning, we really look at our data and ask ourselves if it is truly representative for the task we want solve. The model itself may fit really well and generalize well on your data but if the data is of poor quality to begin with, the model cannot be trusted.
# + [markdown] id="yWzAC39adTwk" colab_type="text"
# # Models
# + [markdown] id="cR45QpjQdY6N" colab_type="text"
# Once you are confident that you data is of good quality and quantity, you can findally start thinking about modeling. The type of model you choose depends on many factors, including the task, type of data, complexity required, etc.
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/practicalAI/master/images/models1.png" width=550>
#
# So once you figure out what type of model your task needs, start with simple models and then slowly add complexity. You don’t want to start with neural networks right away because that may not be right model for your data and task. Striking this balance in model complexity is one of the key tasks of your data scientists. **simple models → complex models**
| notebooks/09_Data_and_Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from PyEMD import EMD, Visualisation
import scipy
import math
import scipy.io
import scipy.linalg
import sklearn.metrics
import sklearn.neighbors
from sklearn import metrics
from sklearn import svm
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch.utils.data import DataLoader, Dataset, TensorDataset
import ipdb
# -
# # Utils
# +
def normalize(V):
return ( V - min(V.flatten()) ) / ( max(V.flatten()) - min(V.flatten()) )
def sliding_window(T, T_org, seq_len, label_seq_len):
# seq_len is equal to window_size
# T (np.array) has dim: population, seq_len (window length)
TT = T.reshape(-1, 1)
K = TT.shape[0] - seq_len - label_seq_len + 1 # Li, et al., 2021, TRJ part C, pp. 8
TT_org = T_org.reshape(-1, 1)
# TT has dim: n, 1
# assemble the data into 2D
x_set = np.vstack(TT[i : K+i, 0] for i in range(seq_len)).T
y_set = np.vstack(TT_org[i+seq_len : K+seq_len+i, 0] for i in range(label_seq_len)).T
assert x_set.shape[0] == y_set.shape[0]
# return size: n_samp, seq_len
return x_set, y_set
def var_name(var, all_var=locals()):
# get the name of the variable
return [var_name for var_name in all_var if all_var[var_name] is var][0]
def np2csv(A):
# store numpy to local csv file
if type(A) == torch.Tensor:
np.savetxt('./outputs/BDA/'+var_name(A)+'.csv', A.detach().numpy(), delimiter=',')
elif type(A) == np.ndarray:
np.savetxt('./outputs/BDA/'+var_name(A)+'.csv', A, delimiter=',')
# -
# # 1. BDA Part
# ## 1.a. Define BDA methodology
# +
def kernel(ker, X1, X2, gamma):
K = None
if not ker or ker == 'primal':
K = X1
elif ker == 'linear':
if X2 is not None:
K = sklearn.metrics.pairwise.linear_kernel(
np.asarray(X1).T, np.asarray(X2).T)
else:
K = sklearn.metrics.pairwise.linear_kernel(np.asarray(X1).T)
elif ker == 'rbf':
if X2 is not None:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, np.asarray(X2).T, gamma)
else:
K = sklearn.metrics.pairwise.rbf_kernel(
np.asarray(X1).T, None, gamma)
return K
def proxy_a_distance(source_X, target_X):
"""
Compute the Proxy-A-Distance of a source/target representation
"""
nb_source = np.shape(source_X)[0]
nb_target = np.shape(target_X)[0]
train_X = np.vstack((source_X, target_X))
train_Y = np.hstack((np.zeros(nb_source, dtype=int),
np.ones(nb_target, dtype=int)))
clf = svm.LinearSVC(random_state=0)
clf.fit(train_X, train_Y)
y_pred = clf.predict(train_X)
error = metrics.mean_absolute_error(train_Y, y_pred)
dist = 2 * (1 - 2 * error)
return dist
def estimate_mu(_X1, _Y1, _X2, _Y2):
adist_m = proxy_a_distance(_X1, _X2)
C = len(np.unique(_Y1))
epsilon = 1e-3
list_adist_c = []
for i in range(1, C + 1):
ind_i, ind_j = np.where(_Y1 == i), np.where(_Y2 == i)
Xsi = _X1[ind_i[0], :]
Xtj = _X2[ind_j[0], :]
adist_i = proxy_a_distance(Xsi, Xtj)
list_adist_c.append(adist_i)
adist_c = sum(list_adist_c) / C
mu = adist_c / (adist_c + adist_m)
if mu > 1:
mu = 1
if mu < epsilon:
mu = 0
return mu
# -
class BDA:
def __init__(self, kernel_type='primal', dim=30, lamb=1, mu=0.5, gamma=1, T=10, mode='BDA', estimate_mu=False):
'''
Init func
:param kernel_type: kernel, values: 'primal' | 'linear' | 'rbf'
:param dim: dimension after transfer
:param lamb: lambda value in equation
:param mu: mu. Default is -1, if not specificied, it calculates using A-distance
:param gamma: kernel bandwidth for rbf kernel
:param T: iteration number
:param mode: 'BDA' | 'WBDA'
:param estimate_mu: True | False, if you want to automatically estimate mu instead of manally set it
'''
self.kernel_type = kernel_type
self.dim = dim
self.lamb = lamb
self.mu = mu
self.gamma = gamma
self.T = T
self.mode = mode
self.estimate_mu = estimate_mu
def fit(self, Xs, Ys, Xt, Yt):
'''
Transform and Predict using 1NN as JDA paper did
:param Xs: ns * n_feature, source feature
:param Ys: ns * 1, source label
:param Xt: nt * n_feature, target feature
:param Yt: nt * 1, target label
:return: acc, y_pred, list_acc
'''
#ipdb.set_trace()
list_acc = []
X = np.hstack((Xs.T, Xt.T)) # X.shape: [n_feature, ns+nt]
X /= np.linalg.norm(X, axis=0) # why it's axis=0?
m, n = X.shape
ns, nt = len(Xs), len(Xt)
e = np.vstack((1 / ns * np.ones((ns, 1)), -1 / nt * np.ones((nt, 1))))
C = len(np.unique(Ys))
H = np.eye(n) - 1 / n * np.ones((n, n))
mu = self.mu
M = 0
Y_tar_pseudo = None
Xs_new = None
for t in range(self.T):
N = 0
M0 = e * e.T * C
if Y_tar_pseudo is not None and len(Y_tar_pseudo) == nt:
for c in range(1, C + 1):
e = np.zeros((n, 1))
Ns = len(Ys[np.where(Ys == c)])
Nt = len(Y_tar_pseudo[np.where(Y_tar_pseudo == c)])
if self.mode == 'WBDA':
Ps = Ns / len(Ys)
Pt = Nt / len(Y_tar_pseudo)
alpha = Pt / Ps
mu = 1
else:
alpha = 1
tt = Ys == c
e[np.where(tt == True)] = 1 / Ns
yy = Y_tar_pseudo == c
ind = np.where(yy == True)
inds = [item + ns for item in ind]
e[tuple(inds)] = -alpha / Nt
e[np.isinf(e)] = 0 # ?
N = N + np.dot(e, e.T)
# In BDA, mu can be set or automatically estimated using A-distance
# In WBDA, we find that setting mu=1 is enough
if self.estimate_mu and self.mode == 'BDA':
if Xs_new is not None:
mu = estimate_mu(Xs_new, Ys, Xt_new, Y_tar_pseudo)
else:
mu = 0
M = (1 - mu) * M0 + mu * N
M /= np.linalg.norm(M, 'fro')
K = kernel(self.kernel_type, X, None, gamma=self.gamma)
n_eye = m if self.kernel_type == 'primal' else n
a, b = np.linalg.multi_dot(
[K, M, K.T]) + self.lamb * np.eye(n_eye), np.linalg.multi_dot([K, H, K.T])
w, V = scipy.linalg.eig(a, b)
ind = np.argsort(w)
A = V[:, ind[:self.dim]]
Z = np.dot(A.T, K)
Z /= np.linalg.norm(Z, axis=0) # why it's axis=0?
Xs_new, Xt_new = Z[:, :ns].T, Z[:, ns:].T
'''
clf = sklearn.neighbors.KNeighborsClassifier(n_neighbors=1)
clf.fit(Xs_new, Ys.ravel())
Y_tar_pseudo = clf.predict(Xt_new)
acc = sklearn.metrics.accuracy_score(Yt, Y_tar_pseudo)
list_acc.append(acc)
print('{} iteration [{}/{}]: Acc: {:.4f}'.format(self.mode, t + 1, self.T, acc))
'''
return Xs_new, Xt_new, A #, acc, Y_tar_pseudo, list_acc
# ## 1.b. Load Data
# +
weekdays = np.array([np.arange(2+7*i,7+7*i,1) for i in range(4)]).flatten()
weekends = np.array([np.arange(7+7*i,9+7*i,1) for i in range(3)]).flatten()[:-1]
src_domain = np.array(pd.read_csv('../TCA_traffic/data/siteM4_2168B_20210101_20210131.csv'))[np.array([5,6,7,8]), :]
data_target = np.array(pd.read_csv('../TCA_traffic/data/siteM4_2188B_20210101_20210131.csv'))[20:25, :]
date_choosen = 10
num_test_day = 4
#tar_domain = data_target[weekdays[date_choosen:date_choosen+1 + num_test_day], :].reshape(-1, 96)
tar_domain = data_target.copy()
tgt_validation = tar_domain[1:num_test_day+1, :]
Xs = normalize(src_domain.flatten())
Xt = normalize(tar_domain.flatten())
# -
# ## 1.d. Hyperparameters
# +
label_seq_len = 1
# batch_size = full batch
seq_len = 10
reduced_dim = 1
inp_dim = seq_len
label_dim = seq_len
hid_dim = 64
layers = 3
lamb = 3
hyper = {
'inp_dim':inp_dim,
'label_dim':label_dim,
'label_seq_len':label_seq_len,
'seq_len':seq_len,
'reduced_dim':reduced_dim,
'hid_dim':hid_dim,
'layers':layers,
'lamb':lamb}
hyper = pd.DataFrame(hyper, index=['Values'])
# -
hyper
# ## 1.e. Apply BDA and get $Xs_{new}$, $Xt_{new}$
# +
Xs, Ys = sliding_window(Xs, Xs, seq_len, label_seq_len)
Xt, Yt = sliding_window(Xt, Xt, seq_len, label_seq_len)
inp_dim -= reduced_dim
label_dim -= reduced_dim
# -
Xs.shape
# +
bda = BDA(kernel_type='linear', dim=inp_dim, lamb=lamb, mu=0.6, gamma=1)
Xs_new, Xt_new, A = bda.fit(Xs, Ys, Xt, Yt) # input shape: ns, n_feature | ns, 1
Xt_new_valid = Xt_new.copy()[int(96):, :]
Xt_new = Xt_new.copy()[:int(96), :]
Yt_valid = Yt.copy()[int(96):, :]
Yt = Yt.copy()[:int(96), :]
print(Xs_new.shape)
print(Xt_new.shape)
print(Xt_new_valid.shape)
np2csv(Xs_new)
np2csv(Xt_new)
# -
# # 2. Learning Part
# ## 2.a. Build network
class LSTM(nn.Module):
def __init__(self, inp_dim, out_dim, hid_dim, layers):
super(LSTM, self).__init__()
self.out_dim = out_dim
self.lstm = nn.LSTM(inp_dim, hid_dim, layers, dropout=0.3, batch_first=True)
self.fc = nn.Sequential(
nn.ReLU(),
nn.Linear(hid_dim, hid_dim*2),
nn.ReLU(),
nn.Linear(hid_dim*2, out_dim)
) # regression
def forward(self, x):
# input: (batchsize, seq_len, input_dim)
# output: (batchsize, seq_len, hid_dim)
#ipdb.set_trace()
y = self.lstm(x)[0] # y, (h, c) = self.rnn(x)
y = self.fc(y[:, :, :]) # fully connected layer
return y[:, -1, :]
# ## 2.b. Assemble Dataloader
Xt_new.shape
# +
batch_size = 960
train_x = np.vstack([Xs_new, Xt_new])[:, :, np.newaxis]
train_y = np.vstack([Ys, Yt])
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
train_x = torch.tensor(train_x, dtype=torch.float32).to(device)
train_y = torch.tensor(train_y, dtype=torch.float32).to(device)
Xt_new_valid = torch.tensor(Xt_new_valid[:, :, np.newaxis], dtype=torch.float32).to(device)
Yt_valid = torch.tensor(Yt_valid, dtype=torch.float32).to(device)
train_dataset = TensorDataset(train_x, train_y)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size, shuffle=True)
train_iter = iter(train_loader)
# -
# ## 2.c. Learn
# build model
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
net = LSTM(1, 1, hid_dim, layers).to(device)
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(net.parameters())
#scheduler = torch.optim.lr_scheduler.StepLR(optimizer, 0.7)
# +
# train
net.train()
epoches = 1000
train_loss_set = []
val_loss_set = []
for e in range(epoches):
for i in range(len(train_loader)):
try:
data, label = train_iter.next()
except:
train_iter = iter(train_loader)
data, label = train_iter.next()
out = net(data)
loss = criterion(out, label)
optimizer.zero_grad()
loss.backward()
optimizer.step()
val_out = net(Xt_new_valid)
val_loss = criterion(val_out, Yt_valid)
val_loss_set.append(val_loss.cpu().detach().numpy())
train_loss_set.append(loss.cpu().detach().numpy())
if e%20==0:
print('Epoch No. %i success, loss: %.5f, val loss: %.5f'%(e, loss.cpu().detach().numpy(), val_loss.cpu().detach().numpy()))
# -
fig = plt.figure(figsize = [16, 4])
ax1 = fig.add_subplot(121)
ax1.plot(train_loss_set)
ax2 = fig.add_subplot(122)
ax2.plot(val_loss_set)
# # 3. Evaluation
# +
def mape_loss_func(preds, labels):
try:
if preds.device.type == 'cuda':
preds = preds.cpu().detach().numpy()
if labels.device.type == 'cuda':
labels = labels.cpu().detach().numpy()
except:
None
mask = labels > .05
return np.mean(np.fabs(labels[mask]-preds[mask])/labels[mask])
def smape_loss_func(preds, labels):
try:
if preds.device.type == 'cuda':
preds = preds.cpu().detach().numpy()
if labels.device.type == 'cuda':
labels = labels.cpu().detach().numpy()
except:
None
mask= labels > .05
return np.mean(2*np.fabs(labels[mask]-preds[mask])/(np.fabs(labels[mask])+np.fabs(preds[mask])))
def mae_loss_func(preds, labels):
try:
if preds.device.type == 'cuda':
preds = preds.cpu().detach().numpy()
if labels.device.type == 'cuda':
labels = labels.cpu().detach().numpy()
except:
None
mask= labels > .05
return np.fabs((labels[mask]-preds[mask])).mean()
def eliminate_nan(b):
a = np.array(b)
c = a[~np.isnan(a)]
return c
# +
net.eval()
print('MAPE: %.5f'%mape_loss_func(val_out, Yt_valid))
print('SMAPE: %.5f'%smape_loss_func(val_out, Yt_valid))
print('MAE: %.5f'%mae_loss_func(val_out, Yt_valid))
# -
| BDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
from datetime import datetime
import numpy as np
def numpysum(n):
a = np.arange(n) ** 2
b = np.arange(n) ** 3
c = a + b
return c
def pythonsum(n):
a = [i for i in range(n)]
b = [i for i in range(n)]
c = []
for i in range(len(a)):
a[i] = i ** 2
b[i] = i ** 3
c.append(a[i] + b[i])
return c
size = int(sys.argv[1])
start = datetime.now()
c = pythonsum(size)
delta = datetime.now() - start
print("The last 2 elements of the sum", c[-2:])
print("PythonSum elapsed time in microseconds", delta.microseconds)
start - datetime.now()
c = numpysum(size)
delta = datetime.now() - start
print("The last 2 elements of the sum", c[-2:])
print("NumpySum elapsed time in microseconds", delta.microseconds)
# -
| pda/pda_ch01.ipynb |